Upload brats_mri_segmentation version 0.5.3
Browse files- .gitattributes +1 -0
- LICENSE +201 -0
- configs/evaluate.json +94 -0
- configs/inference.json +149 -0
- configs/inference_trt.json +12 -0
- configs/logging.conf +21 -0
- configs/metadata.json +111 -0
- configs/multi_gpu_train.json +40 -0
- configs/train.json +329 -0
- docs/README.md +163 -0
- docs/data_license.txt +49 -0
- models/model.pt +3 -0
- models/model.ts +3 -0
- scripts/prepare_datalist.py +72 -0
.gitattributes
CHANGED
|
@@ -33,3 +33,4 @@ saved_model/**/* filter=lfs diff=lfs merge=lfs -text
|
|
| 33 |
*.zip filter=lfs diff=lfs merge=lfs -text
|
| 34 |
*.zst filter=lfs diff=lfs merge=lfs -text
|
| 35 |
*tfevents* filter=lfs diff=lfs merge=lfs -text
|
|
|
|
|
|
| 33 |
*.zip filter=lfs diff=lfs merge=lfs -text
|
| 34 |
*.zst filter=lfs diff=lfs merge=lfs -text
|
| 35 |
*tfevents* filter=lfs diff=lfs merge=lfs -text
|
| 36 |
+
models/model.ts filter=lfs diff=lfs merge=lfs -text
|
LICENSE
ADDED
|
@@ -0,0 +1,201 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
Apache License
|
| 2 |
+
Version 2.0, January 2004
|
| 3 |
+
http://www.apache.org/licenses/
|
| 4 |
+
|
| 5 |
+
TERMS AND CONDITIONS FOR USE, REPRODUCTION, AND DISTRIBUTION
|
| 6 |
+
|
| 7 |
+
1. Definitions.
|
| 8 |
+
|
| 9 |
+
"License" shall mean the terms and conditions for use, reproduction,
|
| 10 |
+
and distribution as defined by Sections 1 through 9 of this document.
|
| 11 |
+
|
| 12 |
+
"Licensor" shall mean the copyright owner or entity authorized by
|
| 13 |
+
the copyright owner that is granting the License.
|
| 14 |
+
|
| 15 |
+
"Legal Entity" shall mean the union of the acting entity and all
|
| 16 |
+
other entities that control, are controlled by, or are under common
|
| 17 |
+
control with that entity. For the purposes of this definition,
|
| 18 |
+
"control" means (i) the power, direct or indirect, to cause the
|
| 19 |
+
direction or management of such entity, whether by contract or
|
| 20 |
+
otherwise, or (ii) ownership of fifty percent (50%) or more of the
|
| 21 |
+
outstanding shares, or (iii) beneficial ownership of such entity.
|
| 22 |
+
|
| 23 |
+
"You" (or "Your") shall mean an individual or Legal Entity
|
| 24 |
+
exercising permissions granted by this License.
|
| 25 |
+
|
| 26 |
+
"Source" form shall mean the preferred form for making modifications,
|
| 27 |
+
including but not limited to software source code, documentation
|
| 28 |
+
source, and configuration files.
|
| 29 |
+
|
| 30 |
+
"Object" form shall mean any form resulting from mechanical
|
| 31 |
+
transformation or translation of a Source form, including but
|
| 32 |
+
not limited to compiled object code, generated documentation,
|
| 33 |
+
and conversions to other media types.
|
| 34 |
+
|
| 35 |
+
"Work" shall mean the work of authorship, whether in Source or
|
| 36 |
+
Object form, made available under the License, as indicated by a
|
| 37 |
+
copyright notice that is included in or attached to the work
|
| 38 |
+
(an example is provided in the Appendix below).
|
| 39 |
+
|
| 40 |
+
"Derivative Works" shall mean any work, whether in Source or Object
|
| 41 |
+
form, that is based on (or derived from) the Work and for which the
|
| 42 |
+
editorial revisions, annotations, elaborations, or other modifications
|
| 43 |
+
represent, as a whole, an original work of authorship. For the purposes
|
| 44 |
+
of this License, Derivative Works shall not include works that remain
|
| 45 |
+
separable from, or merely link (or bind by name) to the interfaces of,
|
| 46 |
+
the Work and Derivative Works thereof.
|
| 47 |
+
|
| 48 |
+
"Contribution" shall mean any work of authorship, including
|
| 49 |
+
the original version of the Work and any modifications or additions
|
| 50 |
+
to that Work or Derivative Works thereof, that is intentionally
|
| 51 |
+
submitted to Licensor for inclusion in the Work by the copyright owner
|
| 52 |
+
or by an individual or Legal Entity authorized to submit on behalf of
|
| 53 |
+
the copyright owner. For the purposes of this definition, "submitted"
|
| 54 |
+
means any form of electronic, verbal, or written communication sent
|
| 55 |
+
to the Licensor or its representatives, including but not limited to
|
| 56 |
+
communication on electronic mailing lists, source code control systems,
|
| 57 |
+
and issue tracking systems that are managed by, or on behalf of, the
|
| 58 |
+
Licensor for the purpose of discussing and improving the Work, but
|
| 59 |
+
excluding communication that is conspicuously marked or otherwise
|
| 60 |
+
designated in writing by the copyright owner as "Not a Contribution."
|
| 61 |
+
|
| 62 |
+
"Contributor" shall mean Licensor and any individual or Legal Entity
|
| 63 |
+
on behalf of whom a Contribution has been received by Licensor and
|
| 64 |
+
subsequently incorporated within the Work.
|
| 65 |
+
|
| 66 |
+
2. Grant of Copyright License. Subject to the terms and conditions of
|
| 67 |
+
this License, each Contributor hereby grants to You a perpetual,
|
| 68 |
+
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
|
| 69 |
+
copyright license to reproduce, prepare Derivative Works of,
|
| 70 |
+
publicly display, publicly perform, sublicense, and distribute the
|
| 71 |
+
Work and such Derivative Works in Source or Object form.
|
| 72 |
+
|
| 73 |
+
3. Grant of Patent License. Subject to the terms and conditions of
|
| 74 |
+
this License, each Contributor hereby grants to You a perpetual,
|
| 75 |
+
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
|
| 76 |
+
(except as stated in this section) patent license to make, have made,
|
| 77 |
+
use, offer to sell, sell, import, and otherwise transfer the Work,
|
| 78 |
+
where such license applies only to those patent claims licensable
|
| 79 |
+
by such Contributor that are necessarily infringed by their
|
| 80 |
+
Contribution(s) alone or by combination of their Contribution(s)
|
| 81 |
+
with the Work to which such Contribution(s) was submitted. If You
|
| 82 |
+
institute patent litigation against any entity (including a
|
| 83 |
+
cross-claim or counterclaim in a lawsuit) alleging that the Work
|
| 84 |
+
or a Contribution incorporated within the Work constitutes direct
|
| 85 |
+
or contributory patent infringement, then any patent licenses
|
| 86 |
+
granted to You under this License for that Work shall terminate
|
| 87 |
+
as of the date such litigation is filed.
|
| 88 |
+
|
| 89 |
+
4. Redistribution. You may reproduce and distribute copies of the
|
| 90 |
+
Work or Derivative Works thereof in any medium, with or without
|
| 91 |
+
modifications, and in Source or Object form, provided that You
|
| 92 |
+
meet the following conditions:
|
| 93 |
+
|
| 94 |
+
(a) You must give any other recipients of the Work or
|
| 95 |
+
Derivative Works a copy of this License; and
|
| 96 |
+
|
| 97 |
+
(b) You must cause any modified files to carry prominent notices
|
| 98 |
+
stating that You changed the files; and
|
| 99 |
+
|
| 100 |
+
(c) You must retain, in the Source form of any Derivative Works
|
| 101 |
+
that You distribute, all copyright, patent, trademark, and
|
| 102 |
+
attribution notices from the Source form of the Work,
|
| 103 |
+
excluding those notices that do not pertain to any part of
|
| 104 |
+
the Derivative Works; and
|
| 105 |
+
|
| 106 |
+
(d) If the Work includes a "NOTICE" text file as part of its
|
| 107 |
+
distribution, then any Derivative Works that You distribute must
|
| 108 |
+
include a readable copy of the attribution notices contained
|
| 109 |
+
within such NOTICE file, excluding those notices that do not
|
| 110 |
+
pertain to any part of the Derivative Works, in at least one
|
| 111 |
+
of the following places: within a NOTICE text file distributed
|
| 112 |
+
as part of the Derivative Works; within the Source form or
|
| 113 |
+
documentation, if provided along with the Derivative Works; or,
|
| 114 |
+
within a display generated by the Derivative Works, if and
|
| 115 |
+
wherever such third-party notices normally appear. The contents
|
| 116 |
+
of the NOTICE file are for informational purposes only and
|
| 117 |
+
do not modify the License. You may add Your own attribution
|
| 118 |
+
notices within Derivative Works that You distribute, alongside
|
| 119 |
+
or as an addendum to the NOTICE text from the Work, provided
|
| 120 |
+
that such additional attribution notices cannot be construed
|
| 121 |
+
as modifying the License.
|
| 122 |
+
|
| 123 |
+
You may add Your own copyright statement to Your modifications and
|
| 124 |
+
may provide additional or different license terms and conditions
|
| 125 |
+
for use, reproduction, or distribution of Your modifications, or
|
| 126 |
+
for any such Derivative Works as a whole, provided Your use,
|
| 127 |
+
reproduction, and distribution of the Work otherwise complies with
|
| 128 |
+
the conditions stated in this License.
|
| 129 |
+
|
| 130 |
+
5. Submission of Contributions. Unless You explicitly state otherwise,
|
| 131 |
+
any Contribution intentionally submitted for inclusion in the Work
|
| 132 |
+
by You to the Licensor shall be under the terms and conditions of
|
| 133 |
+
this License, without any additional terms or conditions.
|
| 134 |
+
Notwithstanding the above, nothing herein shall supersede or modify
|
| 135 |
+
the terms of any separate license agreement you may have executed
|
| 136 |
+
with Licensor regarding such Contributions.
|
| 137 |
+
|
| 138 |
+
6. Trademarks. This License does not grant permission to use the trade
|
| 139 |
+
names, trademarks, service marks, or product names of the Licensor,
|
| 140 |
+
except as required for reasonable and customary use in describing the
|
| 141 |
+
origin of the Work and reproducing the content of the NOTICE file.
|
| 142 |
+
|
| 143 |
+
7. Disclaimer of Warranty. Unless required by applicable law or
|
| 144 |
+
agreed to in writing, Licensor provides the Work (and each
|
| 145 |
+
Contributor provides its Contributions) on an "AS IS" BASIS,
|
| 146 |
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or
|
| 147 |
+
implied, including, without limitation, any warranties or conditions
|
| 148 |
+
of TITLE, NON-INFRINGEMENT, MERCHANTABILITY, or FITNESS FOR A
|
| 149 |
+
PARTICULAR PURPOSE. You are solely responsible for determining the
|
| 150 |
+
appropriateness of using or redistributing the Work and assume any
|
| 151 |
+
risks associated with Your exercise of permissions under this License.
|
| 152 |
+
|
| 153 |
+
8. Limitation of Liability. In no event and under no legal theory,
|
| 154 |
+
whether in tort (including negligence), contract, or otherwise,
|
| 155 |
+
unless required by applicable law (such as deliberate and grossly
|
| 156 |
+
negligent acts) or agreed to in writing, shall any Contributor be
|
| 157 |
+
liable to You for damages, including any direct, indirect, special,
|
| 158 |
+
incidental, or consequential damages of any character arising as a
|
| 159 |
+
result of this License or out of the use or inability to use the
|
| 160 |
+
Work (including but not limited to damages for loss of goodwill,
|
| 161 |
+
work stoppage, computer failure or malfunction, or any and all
|
| 162 |
+
other commercial damages or losses), even if such Contributor
|
| 163 |
+
has been advised of the possibility of such damages.
|
| 164 |
+
|
| 165 |
+
9. Accepting Warranty or Additional Liability. While redistributing
|
| 166 |
+
the Work or Derivative Works thereof, You may choose to offer,
|
| 167 |
+
and charge a fee for, acceptance of support, warranty, indemnity,
|
| 168 |
+
or other liability obligations and/or rights consistent with this
|
| 169 |
+
License. However, in accepting such obligations, You may act only
|
| 170 |
+
on Your own behalf and on Your sole responsibility, not on behalf
|
| 171 |
+
of any other Contributor, and only if You agree to indemnify,
|
| 172 |
+
defend, and hold each Contributor harmless for any liability
|
| 173 |
+
incurred by, or claims asserted against, such Contributor by reason
|
| 174 |
+
of your accepting any such warranty or additional liability.
|
| 175 |
+
|
| 176 |
+
END OF TERMS AND CONDITIONS
|
| 177 |
+
|
| 178 |
+
APPENDIX: How to apply the Apache License to your work.
|
| 179 |
+
|
| 180 |
+
To apply the Apache License to your work, attach the following
|
| 181 |
+
boilerplate notice, with the fields enclosed by brackets "[]"
|
| 182 |
+
replaced with your own identifying information. (Don't include
|
| 183 |
+
the brackets!) The text should be enclosed in the appropriate
|
| 184 |
+
comment syntax for the file format. We also recommend that a
|
| 185 |
+
file or class name and description of purpose be included on the
|
| 186 |
+
same "printed page" as the copyright notice for easier
|
| 187 |
+
identification within third-party archives.
|
| 188 |
+
|
| 189 |
+
Copyright [yyyy] [name of copyright owner]
|
| 190 |
+
|
| 191 |
+
Licensed under the Apache License, Version 2.0 (the "License");
|
| 192 |
+
you may not use this file except in compliance with the License.
|
| 193 |
+
You may obtain a copy of the License at
|
| 194 |
+
|
| 195 |
+
http://www.apache.org/licenses/LICENSE-2.0
|
| 196 |
+
|
| 197 |
+
Unless required by applicable law or agreed to in writing, software
|
| 198 |
+
distributed under the License is distributed on an "AS IS" BASIS,
|
| 199 |
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
| 200 |
+
See the License for the specific language governing permissions and
|
| 201 |
+
limitations under the License.
|
configs/evaluate.json
ADDED
|
@@ -0,0 +1,94 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"validate#postprocessing": {
|
| 3 |
+
"_target_": "Compose",
|
| 4 |
+
"transforms": [
|
| 5 |
+
{
|
| 6 |
+
"_target_": "Activationsd",
|
| 7 |
+
"keys": "pred",
|
| 8 |
+
"sigmoid": true
|
| 9 |
+
},
|
| 10 |
+
{
|
| 11 |
+
"_target_": "Invertd",
|
| 12 |
+
"keys": "pred",
|
| 13 |
+
"transform": "@validate#preprocessing",
|
| 14 |
+
"orig_keys": "image",
|
| 15 |
+
"meta_keys": "pred_meta_dict",
|
| 16 |
+
"nearest_interp": false,
|
| 17 |
+
"to_tensor": true,
|
| 18 |
+
"device": "@validate#evaluator#device"
|
| 19 |
+
},
|
| 20 |
+
{
|
| 21 |
+
"_target_": "AsDiscreted",
|
| 22 |
+
"keys": "pred",
|
| 23 |
+
"threshold": 0.5
|
| 24 |
+
},
|
| 25 |
+
{
|
| 26 |
+
"_target_": "SplitDimd",
|
| 27 |
+
"keys": [
|
| 28 |
+
"pred",
|
| 29 |
+
"label"
|
| 30 |
+
],
|
| 31 |
+
"output_postfixes": [
|
| 32 |
+
"tc",
|
| 33 |
+
"wt",
|
| 34 |
+
"et"
|
| 35 |
+
]
|
| 36 |
+
},
|
| 37 |
+
{
|
| 38 |
+
"_target_": "CopyItemsd",
|
| 39 |
+
"keys": "pred",
|
| 40 |
+
"names": "pred_combined",
|
| 41 |
+
"times": 1
|
| 42 |
+
},
|
| 43 |
+
{
|
| 44 |
+
"_target_": "Lambdad",
|
| 45 |
+
"keys": "pred_combined",
|
| 46 |
+
"func": "$lambda x: torch.where(x[[2]] > 0, 4, torch.where(x[[0]] > 0, 1, torch.where(x[[1]] > 0, 2, 0)))"
|
| 47 |
+
},
|
| 48 |
+
{
|
| 49 |
+
"_target_": "SaveImaged",
|
| 50 |
+
"keys": "pred_combined",
|
| 51 |
+
"meta_keys": "pred_meta_dict",
|
| 52 |
+
"output_dir": "@output_dir",
|
| 53 |
+
"output_postfix": "seg",
|
| 54 |
+
"output_dtype": "uint8",
|
| 55 |
+
"resample": false,
|
| 56 |
+
"squeeze_end_dims": true
|
| 57 |
+
}
|
| 58 |
+
]
|
| 59 |
+
},
|
| 60 |
+
"validate#handlers": [
|
| 61 |
+
{
|
| 62 |
+
"_target_": "CheckpointLoader",
|
| 63 |
+
"load_path": "$@ckpt_dir + '/model.pt'",
|
| 64 |
+
"load_dict": {
|
| 65 |
+
"model": "@network"
|
| 66 |
+
}
|
| 67 |
+
},
|
| 68 |
+
{
|
| 69 |
+
"_target_": "StatsHandler",
|
| 70 |
+
"iteration_log": false
|
| 71 |
+
},
|
| 72 |
+
{
|
| 73 |
+
"_target_": "MetricsSaver",
|
| 74 |
+
"save_dir": "@output_dir",
|
| 75 |
+
"metrics": [
|
| 76 |
+
"val_mean_dice",
|
| 77 |
+
"val_mean_dice_tc",
|
| 78 |
+
"val_mean_dice_wt",
|
| 79 |
+
"val_mean_dice_et"
|
| 80 |
+
],
|
| 81 |
+
"metric_details": [
|
| 82 |
+
"val_mean_dice"
|
| 83 |
+
],
|
| 84 |
+
"batch_transform": "$monai.handlers.from_engine(['image_meta_dict'])",
|
| 85 |
+
"summary_ops": "*"
|
| 86 |
+
}
|
| 87 |
+
],
|
| 88 |
+
"initialize": [
|
| 89 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)"
|
| 90 |
+
],
|
| 91 |
+
"run": [
|
| 92 |
+
"$@validate#evaluator.run()"
|
| 93 |
+
]
|
| 94 |
+
}
|
configs/inference.json
ADDED
|
@@ -0,0 +1,149 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"imports": [
|
| 3 |
+
"$import glob",
|
| 4 |
+
"$import numpy",
|
| 5 |
+
"$import os"
|
| 6 |
+
],
|
| 7 |
+
"bundle_root": ".",
|
| 8 |
+
"image_key": "image",
|
| 9 |
+
"ckpt_dir": "$@bundle_root + '/models'",
|
| 10 |
+
"output_dir": "$@bundle_root + '/eval'",
|
| 11 |
+
"output_ext": ".nii.gz",
|
| 12 |
+
"output_dtype": "uint8",
|
| 13 |
+
"output_postfix": "seg",
|
| 14 |
+
"separate_folder": true,
|
| 15 |
+
"load_pretrain": true,
|
| 16 |
+
"data_list_file_path": "$@bundle_root + '/configs/datalist.json'",
|
| 17 |
+
"dataset_dir": "/workspace/data/medical/brats2018challenge",
|
| 18 |
+
"test_datalist": "$monai.data.load_decathlon_datalist(@data_list_file_path, data_list_key='testing', base_dir=@dataset_dir)",
|
| 19 |
+
"device": "$torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')",
|
| 20 |
+
"amp": true,
|
| 21 |
+
"network_def": {
|
| 22 |
+
"_target_": "SegResNet",
|
| 23 |
+
"blocks_down": [
|
| 24 |
+
1,
|
| 25 |
+
2,
|
| 26 |
+
2,
|
| 27 |
+
4
|
| 28 |
+
],
|
| 29 |
+
"blocks_up": [
|
| 30 |
+
1,
|
| 31 |
+
1,
|
| 32 |
+
1
|
| 33 |
+
],
|
| 34 |
+
"init_filters": 16,
|
| 35 |
+
"in_channels": 4,
|
| 36 |
+
"out_channels": 3,
|
| 37 |
+
"dropout_prob": 0.2
|
| 38 |
+
},
|
| 39 |
+
"network": "$@network_def.to(@device)",
|
| 40 |
+
"preprocessing": {
|
| 41 |
+
"_target_": "Compose",
|
| 42 |
+
"transforms": [
|
| 43 |
+
{
|
| 44 |
+
"_target_": "LoadImaged",
|
| 45 |
+
"keys": "@image_key",
|
| 46 |
+
"image_only": false
|
| 47 |
+
},
|
| 48 |
+
{
|
| 49 |
+
"_target_": "NormalizeIntensityd",
|
| 50 |
+
"keys": "@image_key",
|
| 51 |
+
"nonzero": true,
|
| 52 |
+
"channel_wise": true
|
| 53 |
+
}
|
| 54 |
+
]
|
| 55 |
+
},
|
| 56 |
+
"dataset": {
|
| 57 |
+
"_target_": "Dataset",
|
| 58 |
+
"data": "@test_datalist",
|
| 59 |
+
"transform": "@preprocessing"
|
| 60 |
+
},
|
| 61 |
+
"dataloader": {
|
| 62 |
+
"_target_": "DataLoader",
|
| 63 |
+
"dataset": "@dataset",
|
| 64 |
+
"batch_size": 1,
|
| 65 |
+
"shuffle": true,
|
| 66 |
+
"num_workers": 4
|
| 67 |
+
},
|
| 68 |
+
"inferer": {
|
| 69 |
+
"_target_": "SlidingWindowInferer",
|
| 70 |
+
"roi_size": [
|
| 71 |
+
240,
|
| 72 |
+
240,
|
| 73 |
+
160
|
| 74 |
+
],
|
| 75 |
+
"sw_batch_size": 1,
|
| 76 |
+
"overlap": 0.5
|
| 77 |
+
},
|
| 78 |
+
"postprocessing": {
|
| 79 |
+
"_target_": "Compose",
|
| 80 |
+
"transforms": [
|
| 81 |
+
{
|
| 82 |
+
"_target_": "Activationsd",
|
| 83 |
+
"keys": "pred",
|
| 84 |
+
"sigmoid": true
|
| 85 |
+
},
|
| 86 |
+
{
|
| 87 |
+
"_target_": "Invertd",
|
| 88 |
+
"keys": "pred",
|
| 89 |
+
"transform": "@preprocessing",
|
| 90 |
+
"orig_keys": "@image_key",
|
| 91 |
+
"meta_keys": "pred_meta_dict",
|
| 92 |
+
"nearest_interp": false,
|
| 93 |
+
"to_tensor": true
|
| 94 |
+
},
|
| 95 |
+
{
|
| 96 |
+
"_target_": "AsDiscreted",
|
| 97 |
+
"keys": "pred",
|
| 98 |
+
"threshold": 0.5
|
| 99 |
+
},
|
| 100 |
+
{
|
| 101 |
+
"_target_": "Lambdad",
|
| 102 |
+
"keys": "pred",
|
| 103 |
+
"func": "$lambda x: torch.where(x[[2]] > 0, 4, torch.where(x[[0]] > 0, 1, torch.where(x[[1]] > 0, 2, 0)))"
|
| 104 |
+
},
|
| 105 |
+
{
|
| 106 |
+
"_target_": "SaveImaged",
|
| 107 |
+
"keys": "pred",
|
| 108 |
+
"meta_keys": "pred_meta_dict",
|
| 109 |
+
"output_dir": "@output_dir",
|
| 110 |
+
"output_ext": "@output_ext",
|
| 111 |
+
"output_dtype": "@output_dtype",
|
| 112 |
+
"output_postfix": "@output_postfix",
|
| 113 |
+
"separate_folder": "@separate_folder",
|
| 114 |
+
"resample": false,
|
| 115 |
+
"squeeze_end_dims": true
|
| 116 |
+
}
|
| 117 |
+
]
|
| 118 |
+
},
|
| 119 |
+
"handlers": [
|
| 120 |
+
{
|
| 121 |
+
"_target_": "StatsHandler",
|
| 122 |
+
"iteration_log": false
|
| 123 |
+
}
|
| 124 |
+
],
|
| 125 |
+
"evaluator": {
|
| 126 |
+
"_target_": "SupervisedEvaluator",
|
| 127 |
+
"device": "@device",
|
| 128 |
+
"val_data_loader": "@dataloader",
|
| 129 |
+
"network": "@network",
|
| 130 |
+
"inferer": "@inferer",
|
| 131 |
+
"postprocessing": "@postprocessing",
|
| 132 |
+
"val_handlers": "@handlers",
|
| 133 |
+
"amp": true
|
| 134 |
+
},
|
| 135 |
+
"checkpointloader": {
|
| 136 |
+
"_target_": "CheckpointLoader",
|
| 137 |
+
"load_path": "$@bundle_root + '/models/model.pt'",
|
| 138 |
+
"load_dict": {
|
| 139 |
+
"model": "@network"
|
| 140 |
+
}
|
| 141 |
+
},
|
| 142 |
+
"initialize": [
|
| 143 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)",
|
| 144 |
+
"$@checkpointloader(@evaluator) if @load_pretrain else None"
|
| 145 |
+
],
|
| 146 |
+
"run": [
|
| 147 |
+
"$@evaluator.run()"
|
| 148 |
+
]
|
| 149 |
+
}
|
configs/inference_trt.json
ADDED
|
@@ -0,0 +1,12 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"imports": [
|
| 3 |
+
"$import glob",
|
| 4 |
+
"$import os",
|
| 5 |
+
"$import torch_tensorrt"
|
| 6 |
+
],
|
| 7 |
+
"network_def": "$torch.jit.load(@bundle_root + '/models/model_trt.ts')",
|
| 8 |
+
"evaluator#amp": false,
|
| 9 |
+
"initialize": [
|
| 10 |
+
"$monai.utils.set_determinism(seed=123)"
|
| 11 |
+
]
|
| 12 |
+
}
|
configs/logging.conf
ADDED
|
@@ -0,0 +1,21 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
[loggers]
|
| 2 |
+
keys=root
|
| 3 |
+
|
| 4 |
+
[handlers]
|
| 5 |
+
keys=consoleHandler
|
| 6 |
+
|
| 7 |
+
[formatters]
|
| 8 |
+
keys=fullFormatter
|
| 9 |
+
|
| 10 |
+
[logger_root]
|
| 11 |
+
level=INFO
|
| 12 |
+
handlers=consoleHandler
|
| 13 |
+
|
| 14 |
+
[handler_consoleHandler]
|
| 15 |
+
class=StreamHandler
|
| 16 |
+
level=INFO
|
| 17 |
+
formatter=fullFormatter
|
| 18 |
+
args=(sys.stdout,)
|
| 19 |
+
|
| 20 |
+
[formatter_fullFormatter]
|
| 21 |
+
format=%(asctime)s - %(name)s - %(levelname)s - %(message)s
|
configs/metadata.json
ADDED
|
@@ -0,0 +1,111 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"schema": "https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/meta_schema_20240725.json",
|
| 3 |
+
"version": "0.5.3",
|
| 4 |
+
"changelog": {
|
| 5 |
+
"0.5.3": "update to huggingface hosting",
|
| 6 |
+
"0.5.2": "use monai 1.4 and update large files",
|
| 7 |
+
"0.5.1": "update to use monai 1.3.1",
|
| 8 |
+
"0.5.0": "add load_pretrain flag for infer",
|
| 9 |
+
"0.4.9": "add checkpoint loader for infer",
|
| 10 |
+
"0.4.8": "fix the wrong GPU index issue of multi-node",
|
| 11 |
+
"0.4.7": "enhance prepare datalist file",
|
| 12 |
+
"0.4.6": "add dataset dir example",
|
| 13 |
+
"0.4.5": "update ONNX-TensorRT descriptions",
|
| 14 |
+
"0.4.4": "update error links",
|
| 15 |
+
"0.4.3": "add the ONNX-TensorRT way of model conversion",
|
| 16 |
+
"0.4.2": "fix mgpu finalize issue",
|
| 17 |
+
"0.4.1": "add non-deterministic note",
|
| 18 |
+
"0.4.0": "adapt to BundleWorkflow interface",
|
| 19 |
+
"0.3.9": "black autofix format and add name tag",
|
| 20 |
+
"0.3.8": "modify dataset key name",
|
| 21 |
+
"0.3.7": "restructure readme to match updated template",
|
| 22 |
+
"0.3.6": "added train/val graphs",
|
| 23 |
+
"0.3.5": "update prepare datalist function",
|
| 24 |
+
"0.3.4": "update output format of inference",
|
| 25 |
+
"0.3.3": "update to use monai 1.0.1",
|
| 26 |
+
"0.3.2": "enhance readme on commands example",
|
| 27 |
+
"0.3.1": "fix license Copyright error",
|
| 28 |
+
"0.3.0": "update license files",
|
| 29 |
+
"0.2.1": "fix network_data_format error",
|
| 30 |
+
"0.2.0": "unify naming",
|
| 31 |
+
"0.1.1": "update for MetaTensor",
|
| 32 |
+
"0.1.0": "complete the model package"
|
| 33 |
+
},
|
| 34 |
+
"monai_version": "1.4.0",
|
| 35 |
+
"pytorch_version": "2.4.0",
|
| 36 |
+
"numpy_version": "1.24.4",
|
| 37 |
+
"required_packages_version": {
|
| 38 |
+
"nibabel": "5.2.1",
|
| 39 |
+
"pytorch-ignite": "0.4.11",
|
| 40 |
+
"scikit-learn": "1.5.1",
|
| 41 |
+
"tensorboard": "2.17.0"
|
| 42 |
+
},
|
| 43 |
+
"supported_apps": {},
|
| 44 |
+
"name": "BraTS MRI segmentation",
|
| 45 |
+
"task": "Multimodal Brain Tumor segmentation",
|
| 46 |
+
"description": "A pre-trained model for volumetric (3D) segmentation of brain tumor subregions from multimodal MRIs based on BraTS 2018 data",
|
| 47 |
+
"authors": "MONAI team",
|
| 48 |
+
"copyright": "Copyright (c) MONAI Consortium",
|
| 49 |
+
"data_source": "https://www.med.upenn.edu/sbia/brats2018/data.html",
|
| 50 |
+
"data_type": "nibabel",
|
| 51 |
+
"image_classes": "4 channel data, T1c, T1, T2, FLAIR at 1x1x1 mm",
|
| 52 |
+
"label_classes": "3 channel data, channel 0 for Tumor core, channel 1 for Whole tumor, channel 2 for Enhancing tumor",
|
| 53 |
+
"pred_classes": "3 channels data, same as label_classes",
|
| 54 |
+
"eval_metrics": {
|
| 55 |
+
"val_mean_dice": 0.8518,
|
| 56 |
+
"val_mean_dice_tc": 0.8559,
|
| 57 |
+
"val_mean_dice_wt": 0.9026,
|
| 58 |
+
"val_mean_dice_et": 0.7905
|
| 59 |
+
},
|
| 60 |
+
"intended_use": "This is an example, not to be used for diagnostic purposes",
|
| 61 |
+
"references": [
|
| 62 |
+
"Myronenko, Andriy. '3D MRI brain tumor segmentation using autoencoder regularization.' International MICCAI Brainlesion Workshop. Springer, Cham, 2018. https://arxiv.org/abs/1810.11654"
|
| 63 |
+
],
|
| 64 |
+
"network_data_format": {
|
| 65 |
+
"inputs": {
|
| 66 |
+
"image": {
|
| 67 |
+
"type": "image",
|
| 68 |
+
"format": "magnitude",
|
| 69 |
+
"modality": "MR",
|
| 70 |
+
"num_channels": 4,
|
| 71 |
+
"spatial_shape": [
|
| 72 |
+
"8*n",
|
| 73 |
+
"8*n",
|
| 74 |
+
"8*n"
|
| 75 |
+
],
|
| 76 |
+
"dtype": "float32",
|
| 77 |
+
"value_range": [],
|
| 78 |
+
"is_patch_data": true,
|
| 79 |
+
"channel_def": {
|
| 80 |
+
"0": "T1c",
|
| 81 |
+
"1": "T1",
|
| 82 |
+
"2": "T2",
|
| 83 |
+
"3": "FLAIR"
|
| 84 |
+
}
|
| 85 |
+
}
|
| 86 |
+
},
|
| 87 |
+
"outputs": {
|
| 88 |
+
"pred": {
|
| 89 |
+
"type": "image",
|
| 90 |
+
"format": "segmentation",
|
| 91 |
+
"num_channels": 3,
|
| 92 |
+
"spatial_shape": [
|
| 93 |
+
"8*n",
|
| 94 |
+
"8*n",
|
| 95 |
+
"8*n"
|
| 96 |
+
],
|
| 97 |
+
"dtype": "float32",
|
| 98 |
+
"value_range": [
|
| 99 |
+
0,
|
| 100 |
+
1
|
| 101 |
+
],
|
| 102 |
+
"is_patch_data": true,
|
| 103 |
+
"channel_def": {
|
| 104 |
+
"0": "Tumor core",
|
| 105 |
+
"1": "Whole tumor",
|
| 106 |
+
"2": "Enhancing tumor"
|
| 107 |
+
}
|
| 108 |
+
}
|
| 109 |
+
}
|
| 110 |
+
}
|
| 111 |
+
}
|
configs/multi_gpu_train.json
ADDED
|
@@ -0,0 +1,40 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"device": "$torch.device('cuda:' + os.environ['LOCAL_RANK'])",
|
| 3 |
+
"network": {
|
| 4 |
+
"_target_": "torch.nn.parallel.DistributedDataParallel",
|
| 5 |
+
"module": "$@network_def.to(@device)",
|
| 6 |
+
"device_ids": [
|
| 7 |
+
"@device"
|
| 8 |
+
]
|
| 9 |
+
},
|
| 10 |
+
"train#sampler": {
|
| 11 |
+
"_target_": "DistributedSampler",
|
| 12 |
+
"dataset": "@train#dataset",
|
| 13 |
+
"even_divisible": true,
|
| 14 |
+
"shuffle": true
|
| 15 |
+
},
|
| 16 |
+
"train#dataloader#sampler": "@train#sampler",
|
| 17 |
+
"train#dataloader#shuffle": false,
|
| 18 |
+
"train#trainer#train_handlers": "$@train#handlers[: -2 if dist.get_rank() > 0 else None]",
|
| 19 |
+
"validate#sampler": {
|
| 20 |
+
"_target_": "DistributedSampler",
|
| 21 |
+
"dataset": "@validate#dataset",
|
| 22 |
+
"even_divisible": false,
|
| 23 |
+
"shuffle": false
|
| 24 |
+
},
|
| 25 |
+
"validate#dataloader#sampler": "@validate#sampler",
|
| 26 |
+
"validate#evaluator#val_handlers": "$None if dist.get_rank() > 0 else @validate#handlers",
|
| 27 |
+
"initialize": [
|
| 28 |
+
"$import torch.distributed as dist",
|
| 29 |
+
"$dist.is_initialized() or dist.init_process_group(backend='nccl')",
|
| 30 |
+
"$torch.cuda.set_device(@device)",
|
| 31 |
+
"$monai.utils.set_determinism(seed=123)",
|
| 32 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)"
|
| 33 |
+
],
|
| 34 |
+
"run": [
|
| 35 |
+
"$@train#trainer.run()"
|
| 36 |
+
],
|
| 37 |
+
"finalize": [
|
| 38 |
+
"$dist.is_initialized() and dist.destroy_process_group()"
|
| 39 |
+
]
|
| 40 |
+
}
|
configs/train.json
ADDED
|
@@ -0,0 +1,329 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"imports": [
|
| 3 |
+
"$import glob",
|
| 4 |
+
"$import os"
|
| 5 |
+
],
|
| 6 |
+
"bundle_root": ".",
|
| 7 |
+
"ckpt_dir": "$@bundle_root + '/models'",
|
| 8 |
+
"output_dir": "$@bundle_root + '/eval'",
|
| 9 |
+
"data_list_file_path": "$@bundle_root + '/configs/datalist.json'",
|
| 10 |
+
"dataset_dir": "/workspace/data/medical/brats2018challenge",
|
| 11 |
+
"train_datalist": "$monai.data.load_decathlon_datalist(@data_list_file_path, data_list_key='training', base_dir=@dataset_dir)",
|
| 12 |
+
"val_datalist": "$monai.data.load_decathlon_datalist(@data_list_file_path, data_list_key='validation', base_dir=@dataset_dir)",
|
| 13 |
+
"device": "$torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')",
|
| 14 |
+
"epochs": 300,
|
| 15 |
+
"val_interval": 1,
|
| 16 |
+
"learning_rate": 0.0001,
|
| 17 |
+
"amp": true,
|
| 18 |
+
"network_def": {
|
| 19 |
+
"_target_": "SegResNet",
|
| 20 |
+
"blocks_down": [
|
| 21 |
+
1,
|
| 22 |
+
2,
|
| 23 |
+
2,
|
| 24 |
+
4
|
| 25 |
+
],
|
| 26 |
+
"blocks_up": [
|
| 27 |
+
1,
|
| 28 |
+
1,
|
| 29 |
+
1
|
| 30 |
+
],
|
| 31 |
+
"init_filters": 16,
|
| 32 |
+
"in_channels": 4,
|
| 33 |
+
"out_channels": 3,
|
| 34 |
+
"dropout_prob": 0.2
|
| 35 |
+
},
|
| 36 |
+
"network": "$@network_def.to(@device)",
|
| 37 |
+
"loss": {
|
| 38 |
+
"_target_": "DiceLoss",
|
| 39 |
+
"smooth_nr": 0,
|
| 40 |
+
"smooth_dr": 1e-05,
|
| 41 |
+
"squared_pred": true,
|
| 42 |
+
"to_onehot_y": false,
|
| 43 |
+
"sigmoid": true
|
| 44 |
+
},
|
| 45 |
+
"optimizer": {
|
| 46 |
+
"_target_": "torch.optim.Adam",
|
| 47 |
+
"params": "$@network.parameters()",
|
| 48 |
+
"lr": "@learning_rate",
|
| 49 |
+
"weight_decay": 1e-05
|
| 50 |
+
},
|
| 51 |
+
"lr_scheduler": {
|
| 52 |
+
"_target_": "torch.optim.lr_scheduler.CosineAnnealingLR",
|
| 53 |
+
"optimizer": "@optimizer",
|
| 54 |
+
"T_max": "@epochs"
|
| 55 |
+
},
|
| 56 |
+
"train": {
|
| 57 |
+
"preprocessing_transforms": [
|
| 58 |
+
{
|
| 59 |
+
"_target_": "LoadImaged",
|
| 60 |
+
"keys": [
|
| 61 |
+
"image",
|
| 62 |
+
"label"
|
| 63 |
+
],
|
| 64 |
+
"image_only": false
|
| 65 |
+
},
|
| 66 |
+
{
|
| 67 |
+
"_target_": "ConvertToMultiChannelBasedOnBratsClassesd",
|
| 68 |
+
"keys": "label"
|
| 69 |
+
},
|
| 70 |
+
{
|
| 71 |
+
"_target_": "NormalizeIntensityd",
|
| 72 |
+
"keys": "image",
|
| 73 |
+
"nonzero": true,
|
| 74 |
+
"channel_wise": true
|
| 75 |
+
}
|
| 76 |
+
],
|
| 77 |
+
"random_transforms": [
|
| 78 |
+
{
|
| 79 |
+
"_target_": "RandSpatialCropd",
|
| 80 |
+
"keys": [
|
| 81 |
+
"image",
|
| 82 |
+
"label"
|
| 83 |
+
],
|
| 84 |
+
"roi_size": [
|
| 85 |
+
224,
|
| 86 |
+
224,
|
| 87 |
+
144
|
| 88 |
+
],
|
| 89 |
+
"random_size": false
|
| 90 |
+
},
|
| 91 |
+
{
|
| 92 |
+
"_target_": "RandFlipd",
|
| 93 |
+
"keys": [
|
| 94 |
+
"image",
|
| 95 |
+
"label"
|
| 96 |
+
],
|
| 97 |
+
"prob": 0.5,
|
| 98 |
+
"spatial_axis": 0
|
| 99 |
+
},
|
| 100 |
+
{
|
| 101 |
+
"_target_": "RandFlipd",
|
| 102 |
+
"keys": [
|
| 103 |
+
"image",
|
| 104 |
+
"label"
|
| 105 |
+
],
|
| 106 |
+
"prob": 0.5,
|
| 107 |
+
"spatial_axis": 1
|
| 108 |
+
},
|
| 109 |
+
{
|
| 110 |
+
"_target_": "RandFlipd",
|
| 111 |
+
"keys": [
|
| 112 |
+
"image",
|
| 113 |
+
"label"
|
| 114 |
+
],
|
| 115 |
+
"prob": 0.5,
|
| 116 |
+
"spatial_axis": 2
|
| 117 |
+
},
|
| 118 |
+
{
|
| 119 |
+
"_target_": "RandScaleIntensityd",
|
| 120 |
+
"keys": "image",
|
| 121 |
+
"factors": 0.1,
|
| 122 |
+
"prob": 1.0
|
| 123 |
+
},
|
| 124 |
+
{
|
| 125 |
+
"_target_": "RandShiftIntensityd",
|
| 126 |
+
"keys": "image",
|
| 127 |
+
"offsets": 0.1,
|
| 128 |
+
"prob": 1.0
|
| 129 |
+
}
|
| 130 |
+
],
|
| 131 |
+
"preprocessing": {
|
| 132 |
+
"_target_": "Compose",
|
| 133 |
+
"transforms": "$@train#preprocessing_transforms + @train#random_transforms"
|
| 134 |
+
},
|
| 135 |
+
"dataset": {
|
| 136 |
+
"_target_": "Dataset",
|
| 137 |
+
"data": "@train_datalist",
|
| 138 |
+
"transform": "@train#preprocessing"
|
| 139 |
+
},
|
| 140 |
+
"dataloader": {
|
| 141 |
+
"_target_": "DataLoader",
|
| 142 |
+
"dataset": "@train#dataset",
|
| 143 |
+
"batch_size": 1,
|
| 144 |
+
"shuffle": true,
|
| 145 |
+
"num_workers": 4
|
| 146 |
+
},
|
| 147 |
+
"inferer": {
|
| 148 |
+
"_target_": "SimpleInferer"
|
| 149 |
+
},
|
| 150 |
+
"postprocessing": {
|
| 151 |
+
"_target_": "Compose",
|
| 152 |
+
"transforms": [
|
| 153 |
+
{
|
| 154 |
+
"_target_": "Activationsd",
|
| 155 |
+
"keys": "pred",
|
| 156 |
+
"sigmoid": true
|
| 157 |
+
},
|
| 158 |
+
{
|
| 159 |
+
"_target_": "AsDiscreted",
|
| 160 |
+
"keys": "pred",
|
| 161 |
+
"threshold": 0.5
|
| 162 |
+
}
|
| 163 |
+
]
|
| 164 |
+
},
|
| 165 |
+
"handlers": [
|
| 166 |
+
{
|
| 167 |
+
"_target_": "LrScheduleHandler",
|
| 168 |
+
"lr_scheduler": "@lr_scheduler",
|
| 169 |
+
"print_lr": true
|
| 170 |
+
},
|
| 171 |
+
{
|
| 172 |
+
"_target_": "ValidationHandler",
|
| 173 |
+
"validator": "@validate#evaluator",
|
| 174 |
+
"epoch_level": true,
|
| 175 |
+
"interval": "@val_interval"
|
| 176 |
+
},
|
| 177 |
+
{
|
| 178 |
+
"_target_": "StatsHandler",
|
| 179 |
+
"tag_name": "train_loss",
|
| 180 |
+
"output_transform": "$monai.handlers.from_engine(['loss'], first=True)"
|
| 181 |
+
},
|
| 182 |
+
{
|
| 183 |
+
"_target_": "TensorBoardStatsHandler",
|
| 184 |
+
"log_dir": "@output_dir",
|
| 185 |
+
"tag_name": "train_loss",
|
| 186 |
+
"output_transform": "$monai.handlers.from_engine(['loss'], first=True)"
|
| 187 |
+
}
|
| 188 |
+
],
|
| 189 |
+
"key_metric": {
|
| 190 |
+
"train_mean_dice": {
|
| 191 |
+
"_target_": "MeanDice",
|
| 192 |
+
"include_background": true,
|
| 193 |
+
"output_transform": "$monai.handlers.from_engine(['pred', 'label'])"
|
| 194 |
+
}
|
| 195 |
+
},
|
| 196 |
+
"trainer": {
|
| 197 |
+
"_target_": "SupervisedTrainer",
|
| 198 |
+
"max_epochs": "@epochs",
|
| 199 |
+
"device": "@device",
|
| 200 |
+
"train_data_loader": "@train#dataloader",
|
| 201 |
+
"network": "@network",
|
| 202 |
+
"loss_function": "@loss",
|
| 203 |
+
"optimizer": "@optimizer",
|
| 204 |
+
"inferer": "@train#inferer",
|
| 205 |
+
"postprocessing": "@train#postprocessing",
|
| 206 |
+
"key_train_metric": "@train#key_metric",
|
| 207 |
+
"train_handlers": "@train#handlers",
|
| 208 |
+
"amp": "@amp"
|
| 209 |
+
}
|
| 210 |
+
},
|
| 211 |
+
"validate": {
|
| 212 |
+
"preprocessing": {
|
| 213 |
+
"_target_": "Compose",
|
| 214 |
+
"transforms": "$@train#preprocessing_transforms"
|
| 215 |
+
},
|
| 216 |
+
"dataset": {
|
| 217 |
+
"_target_": "Dataset",
|
| 218 |
+
"data": "@val_datalist",
|
| 219 |
+
"transform": "@validate#preprocessing"
|
| 220 |
+
},
|
| 221 |
+
"dataloader": {
|
| 222 |
+
"_target_": "DataLoader",
|
| 223 |
+
"dataset": "@validate#dataset",
|
| 224 |
+
"batch_size": 1,
|
| 225 |
+
"shuffle": false,
|
| 226 |
+
"num_workers": 4
|
| 227 |
+
},
|
| 228 |
+
"inferer": {
|
| 229 |
+
"_target_": "SlidingWindowInferer",
|
| 230 |
+
"roi_size": [
|
| 231 |
+
240,
|
| 232 |
+
240,
|
| 233 |
+
160
|
| 234 |
+
],
|
| 235 |
+
"sw_batch_size": 1,
|
| 236 |
+
"overlap": 0.5
|
| 237 |
+
},
|
| 238 |
+
"postprocessing": {
|
| 239 |
+
"_target_": "Compose",
|
| 240 |
+
"transforms": [
|
| 241 |
+
{
|
| 242 |
+
"_target_": "Activationsd",
|
| 243 |
+
"keys": "pred",
|
| 244 |
+
"sigmoid": true
|
| 245 |
+
},
|
| 246 |
+
{
|
| 247 |
+
"_target_": "AsDiscreted",
|
| 248 |
+
"keys": "pred",
|
| 249 |
+
"threshold": 0.5
|
| 250 |
+
},
|
| 251 |
+
{
|
| 252 |
+
"_target_": "SplitDimd",
|
| 253 |
+
"keys": [
|
| 254 |
+
"pred",
|
| 255 |
+
"label"
|
| 256 |
+
],
|
| 257 |
+
"output_postfixes": [
|
| 258 |
+
"tc",
|
| 259 |
+
"wt",
|
| 260 |
+
"et"
|
| 261 |
+
]
|
| 262 |
+
}
|
| 263 |
+
]
|
| 264 |
+
},
|
| 265 |
+
"handlers": [
|
| 266 |
+
{
|
| 267 |
+
"_target_": "StatsHandler",
|
| 268 |
+
"iteration_log": false
|
| 269 |
+
},
|
| 270 |
+
{
|
| 271 |
+
"_target_": "TensorBoardStatsHandler",
|
| 272 |
+
"log_dir": "@output_dir",
|
| 273 |
+
"iteration_log": false
|
| 274 |
+
},
|
| 275 |
+
{
|
| 276 |
+
"_target_": "CheckpointSaver",
|
| 277 |
+
"save_dir": "@ckpt_dir",
|
| 278 |
+
"save_dict": {
|
| 279 |
+
"model": "@network"
|
| 280 |
+
},
|
| 281 |
+
"save_key_metric": true,
|
| 282 |
+
"key_metric_filename": "model.pt"
|
| 283 |
+
}
|
| 284 |
+
],
|
| 285 |
+
"key_metric": {
|
| 286 |
+
"val_mean_dice": {
|
| 287 |
+
"_target_": "MeanDice",
|
| 288 |
+
"include_background": true,
|
| 289 |
+
"output_transform": "$monai.handlers.from_engine(['pred', 'label'])"
|
| 290 |
+
}
|
| 291 |
+
},
|
| 292 |
+
"additional_metrics": {
|
| 293 |
+
"val_mean_dice_tc": {
|
| 294 |
+
"_target_": "MeanDice",
|
| 295 |
+
"include_background": true,
|
| 296 |
+
"output_transform": "$monai.handlers.from_engine(['pred_tc', 'label_tc'])"
|
| 297 |
+
},
|
| 298 |
+
"val_mean_dice_wt": {
|
| 299 |
+
"_target_": "MeanDice",
|
| 300 |
+
"include_background": true,
|
| 301 |
+
"output_transform": "$monai.handlers.from_engine(['pred_wt', 'label_wt'])"
|
| 302 |
+
},
|
| 303 |
+
"val_mean_dice_et": {
|
| 304 |
+
"_target_": "MeanDice",
|
| 305 |
+
"include_background": true,
|
| 306 |
+
"output_transform": "$monai.handlers.from_engine(['pred_et', 'label_et'])"
|
| 307 |
+
}
|
| 308 |
+
},
|
| 309 |
+
"evaluator": {
|
| 310 |
+
"_target_": "SupervisedEvaluator",
|
| 311 |
+
"device": "@device",
|
| 312 |
+
"val_data_loader": "@validate#dataloader",
|
| 313 |
+
"network": "@network",
|
| 314 |
+
"inferer": "@validate#inferer",
|
| 315 |
+
"postprocessing": "@validate#postprocessing",
|
| 316 |
+
"key_val_metric": "@validate#key_metric",
|
| 317 |
+
"additional_metrics": "@validate#additional_metrics",
|
| 318 |
+
"val_handlers": "@validate#handlers",
|
| 319 |
+
"amp": "@amp"
|
| 320 |
+
}
|
| 321 |
+
},
|
| 322 |
+
"initialize": [
|
| 323 |
+
"$monai.utils.set_determinism(seed=123)",
|
| 324 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)"
|
| 325 |
+
],
|
| 326 |
+
"run": [
|
| 327 |
+
"$@train#trainer.run()"
|
| 328 |
+
]
|
| 329 |
+
}
|
docs/README.md
ADDED
|
@@ -0,0 +1,163 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Model Overview
|
| 2 |
+
A pre-trained model for volumetric (3D) segmentation of brain tumor subregions from multimodal MRIs based on BraTS 2018 data.
|
| 3 |
+
|
| 4 |
+
The model is trained to segment 3 nested subregions of primary brain tumors (gliomas): the "enhancing tumor" (ET), the "tumor core" (TC), the "whole tumor" (WT) based on 4 aligned input MRI scans (T1c, T1, T2, FLAIR).
|
| 5 |
+
- The ET is described by areas that show hyper intensity in T1c when compared to T1, but also when compared to "healthy" white matter in T1c.
|
| 6 |
+
- The TC describes the bulk of the tumor, which is what is typically resected. The TC entails the ET, as well as the necrotic (fluid-filled) and the non-enhancing (solid) parts of the tumor.
|
| 7 |
+
- The WT describes the complete extent of the disease, as it entails the TC and the peritumoral edema (ED), which is typically depicted by hyper-intense signal in FLAIR.
|
| 8 |
+
|
| 9 |
+

|
| 10 |
+
|
| 11 |
+
## Data
|
| 12 |
+
The training data is from the [Multimodal Brain Tumor Segmentation Challenge (BraTS) 2018](https://www.med.upenn.edu/sbia/brats2018.html).
|
| 13 |
+
|
| 14 |
+
- Target: 3 tumor subregions
|
| 15 |
+
- Task: Segmentation
|
| 16 |
+
- Modality: MRI
|
| 17 |
+
- Size: 285 3D volumes (4 channels each)
|
| 18 |
+
|
| 19 |
+
The provided labelled data was partitioned, based on our own split, into training (200 studies), validation (42 studies) and testing (43 studies) datasets.
|
| 20 |
+
|
| 21 |
+
### Preprocessing
|
| 22 |
+
The data list/split can be created with the script `scripts/prepare_datalist.py`.
|
| 23 |
+
|
| 24 |
+
```
|
| 25 |
+
python scripts/prepare_datalist.py --path your-brats18-dataset-path
|
| 26 |
+
```
|
| 27 |
+
|
| 28 |
+
## Training configuration
|
| 29 |
+
This model utilized a similar approach described in 3D MRI brain tumor segmentation using autoencoder regularization, which was a winning method in BraTS2018 [1]. The training was performed with the following:
|
| 30 |
+
|
| 31 |
+
- GPU: At least 16GB of GPU memory.
|
| 32 |
+
- Actual Model Input: 224 x 224 x 144
|
| 33 |
+
- AMP: True
|
| 34 |
+
- Optimizer: Adam
|
| 35 |
+
- Learning Rate: 1e-4
|
| 36 |
+
- Loss: DiceLoss
|
| 37 |
+
|
| 38 |
+
## Input
|
| 39 |
+
4 channel aligned MRIs at 1 x 1 x 1 mm
|
| 40 |
+
- T1c
|
| 41 |
+
- T1
|
| 42 |
+
- T2
|
| 43 |
+
- FLAIR
|
| 44 |
+
|
| 45 |
+
## Output
|
| 46 |
+
3 channels
|
| 47 |
+
- Label 0: TC tumor subregion
|
| 48 |
+
- Label 1: WT tumor subregion
|
| 49 |
+
- Label 2: ET tumor subregion
|
| 50 |
+
|
| 51 |
+
## Performance
|
| 52 |
+
Dice score was used for evaluating the performance of the model. This model achieved Dice scores on the validation data of:
|
| 53 |
+
- Tumor core (TC): 0.8559
|
| 54 |
+
- Whole tumor (WT): 0.9026
|
| 55 |
+
- Enhancing tumor (ET): 0.7905
|
| 56 |
+
- Average: 0.8518
|
| 57 |
+
|
| 58 |
+
Please note that this bundle is non-deterministic because of the trilinear interpolation used in the network. Therefore, reproducing the training process may not get exactly the same performance.
|
| 59 |
+
Please refer to https://pytorch.org/docs/stable/notes/randomness.html#reproducibility for more details about reproducibility.
|
| 60 |
+
|
| 61 |
+
#### Training Loss and Dice
|
| 62 |
+
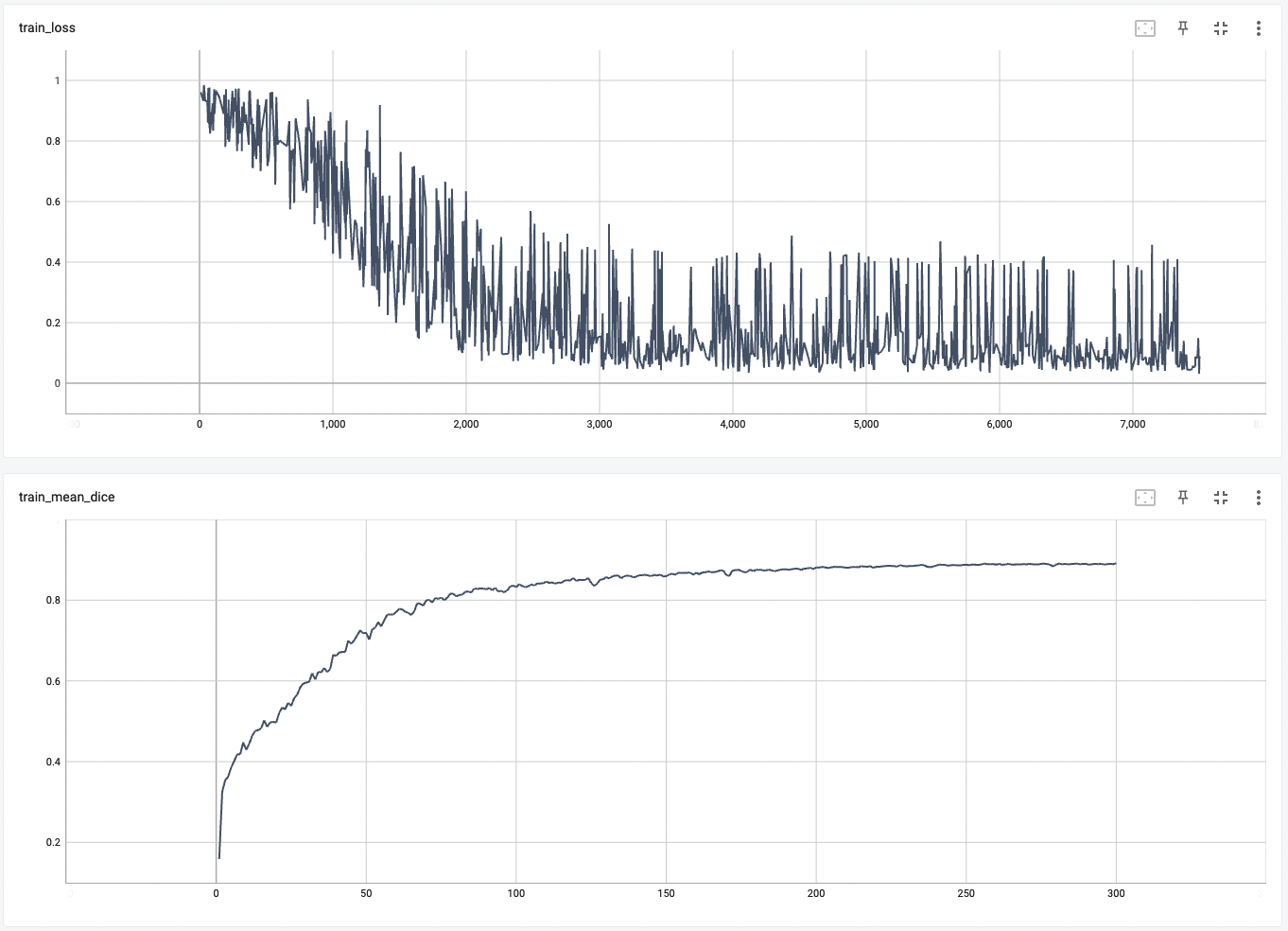
|
| 63 |
+
|
| 64 |
+
#### Validation Dice
|
| 65 |
+
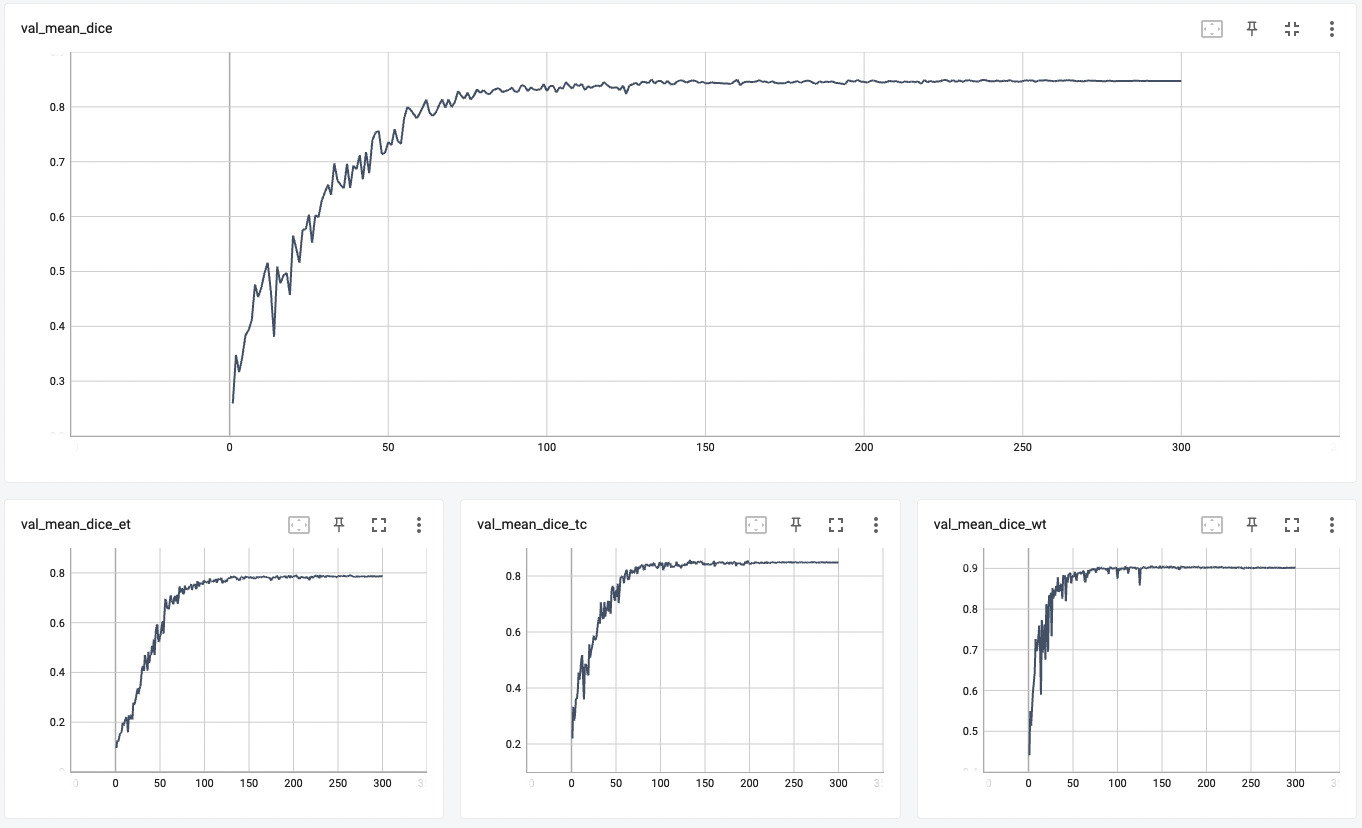
|
| 66 |
+
|
| 67 |
+
#### TensorRT speedup
|
| 68 |
+
The `brats_mri_segmentation` bundle supports acceleration with TensorRT through the ONNX-TensorRT method. The table below displays the speedup ratios observed on an A100 80G GPU.
|
| 69 |
+
|
| 70 |
+
| method | torch_fp32(ms) | torch_amp(ms) | trt_fp32(ms) | trt_fp16(ms) | speedup amp | speedup fp32 | speedup fp16 | amp vs fp16|
|
| 71 |
+
| :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: |
|
| 72 |
+
| model computation | 5.49 | 4.36 | 2.35 | 2.09 | 1.26 | 2.34 | 2.63 | 2.09 |
|
| 73 |
+
| end2end | 592.01 | 434.59 | 395.73 | 394.93 | 1.36 | 1.50 | 1.50 | 1.10 |
|
| 74 |
+
|
| 75 |
+
Where:
|
| 76 |
+
- `model computation` means the speedup ratio of model's inference with a random input without preprocessing and postprocessing
|
| 77 |
+
- `end2end` means run the bundle end-to-end with the TensorRT based model.
|
| 78 |
+
- `torch_fp32` and `torch_amp` are for the PyTorch models with or without `amp` mode.
|
| 79 |
+
- `trt_fp32` and `trt_fp16` are for the TensorRT based models converted in corresponding precision.
|
| 80 |
+
- `speedup amp`, `speedup fp32` and `speedup fp16` are the speedup ratios of corresponding models versus the PyTorch float32 model
|
| 81 |
+
- `amp vs fp16` is the speedup ratio between the PyTorch amp model and the TensorRT float16 based model.
|
| 82 |
+
|
| 83 |
+
Currently, the only available method to accelerate this model is through ONNX-TensorRT. However, the Torch-TensorRT method is under development and will be available in the near future.
|
| 84 |
+
|
| 85 |
+
This result is benchmarked under:
|
| 86 |
+
- TensorRT: 8.5.3+cuda11.8
|
| 87 |
+
- Torch-TensorRT Version: 1.4.0
|
| 88 |
+
- CPU Architecture: x86-64
|
| 89 |
+
- OS: ubuntu 20.04
|
| 90 |
+
- Python version:3.8.10
|
| 91 |
+
- CUDA version: 12.0
|
| 92 |
+
- GPU models and configuration: A100 80G
|
| 93 |
+
|
| 94 |
+
## MONAI Bundle Commands
|
| 95 |
+
In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file.
|
| 96 |
+
|
| 97 |
+
For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html).
|
| 98 |
+
|
| 99 |
+
#### Execute training:
|
| 100 |
+
|
| 101 |
+
```
|
| 102 |
+
python -m monai.bundle run --config_file configs/train.json
|
| 103 |
+
```
|
| 104 |
+
|
| 105 |
+
Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`:
|
| 106 |
+
|
| 107 |
+
```
|
| 108 |
+
python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path>
|
| 109 |
+
```
|
| 110 |
+
|
| 111 |
+
#### Override the `train` config to execute multi-GPU training:
|
| 112 |
+
|
| 113 |
+
```
|
| 114 |
+
torchrun --standalone --nnodes=1 --nproc_per_node=8 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']"
|
| 115 |
+
```
|
| 116 |
+
|
| 117 |
+
Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html).
|
| 118 |
+
|
| 119 |
+
#### Override the `train` config to execute evaluation with the trained model:
|
| 120 |
+
|
| 121 |
+
```
|
| 122 |
+
python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']"
|
| 123 |
+
```
|
| 124 |
+
|
| 125 |
+
#### Execute inference:
|
| 126 |
+
|
| 127 |
+
```
|
| 128 |
+
python -m monai.bundle run --config_file configs/inference.json
|
| 129 |
+
```
|
| 130 |
+
|
| 131 |
+
#### Export checkpoint to TensorRT based models with fp32 or fp16 precision:
|
| 132 |
+
|
| 133 |
+
```bash
|
| 134 |
+
python -m monai.bundle trt_export --net_id network_def \
|
| 135 |
+
--filepath models/model_trt.ts --ckpt_file models/model.pt \
|
| 136 |
+
--meta_file configs/metadata.json --config_file configs/inference.json \
|
| 137 |
+
--precision <fp32/fp16> --input_shape "[1, 4, 240, 240, 160]" --use_onnx "True" \
|
| 138 |
+
--use_trace "True"
|
| 139 |
+
```
|
| 140 |
+
|
| 141 |
+
#### Execute inference with the TensorRT model:
|
| 142 |
+
|
| 143 |
+
```
|
| 144 |
+
python -m monai.bundle run --config_file "['configs/inference.json', 'configs/inference_trt.json']"
|
| 145 |
+
```
|
| 146 |
+
|
| 147 |
+
# References
|
| 148 |
+
[1] Myronenko, Andriy. "3D MRI brain tumor segmentation using autoencoder regularization." International MICCAI Brainlesion Workshop. Springer, Cham, 2018. https://arxiv.org/abs/1810.11654.
|
| 149 |
+
|
| 150 |
+
# License
|
| 151 |
+
Copyright (c) MONAI Consortium
|
| 152 |
+
|
| 153 |
+
Licensed under the Apache License, Version 2.0 (the "License");
|
| 154 |
+
you may not use this file except in compliance with the License.
|
| 155 |
+
You may obtain a copy of the License at
|
| 156 |
+
|
| 157 |
+
http://www.apache.org/licenses/LICENSE-2.0
|
| 158 |
+
|
| 159 |
+
Unless required by applicable law or agreed to in writing, software
|
| 160 |
+
distributed under the License is distributed on an "AS IS" BASIS,
|
| 161 |
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
| 162 |
+
See the License for the specific language governing permissions and
|
| 163 |
+
limitations under the License.
|
docs/data_license.txt
ADDED
|
@@ -0,0 +1,49 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
Third Party Licenses
|
| 2 |
+
-----------------------------------------------------------------------
|
| 3 |
+
|
| 4 |
+
/*********************************************************************/
|
| 5 |
+
i. Multimodal Brain Tumor Segmentation Challenge 2018
|
| 6 |
+
https://www.med.upenn.edu/sbia/brats2018/data.html
|
| 7 |
+
/*********************************************************************/
|
| 8 |
+
|
| 9 |
+
Data Usage Agreement / Citations
|
| 10 |
+
|
| 11 |
+
You are free to use and/or refer to the BraTS datasets in your own
|
| 12 |
+
research, provided that you always cite the following two manuscripts:
|
| 13 |
+
|
| 14 |
+
[1] Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby
|
| 15 |
+
[J, Burren Y, Porz N, Slotboom J, Wiest R, Lanczi L, Gerstner E, Weber
|
| 16 |
+
[MA, Arbel T, Avants BB, Ayache N, Buendia P, Collins DL, Cordier N,
|
| 17 |
+
[Corso JJ, Criminisi A, Das T, Delingette H, Demiralp Γ, Durst CR,
|
| 18 |
+
[Dojat M, Doyle S, Festa J, Forbes F, Geremia E, Glocker B, Golland P,
|
| 19 |
+
[Guo X, Hamamci A, Iftekharuddin KM, Jena R, John NM, Konukoglu E,
|
| 20 |
+
[Lashkari D, Mariz JA, Meier R, Pereira S, Precup D, Price SJ, Raviv
|
| 21 |
+
[TR, Reza SM, Ryan M, Sarikaya D, Schwartz L, Shin HC, Shotton J,
|
| 22 |
+
[Silva CA, Sousa N, Subbanna NK, Szekely G, Taylor TJ, Thomas OM,
|
| 23 |
+
[Tustison NJ, Unal G, Vasseur F, Wintermark M, Ye DH, Zhao L, Zhao B,
|
| 24 |
+
[Zikic D, Prastawa M, Reyes M, Van Leemput K. "The Multimodal Brain
|
| 25 |
+
[Tumor Image Segmentation Benchmark (BRATS)", IEEE Transactions on
|
| 26 |
+
[Medical Imaging 34(10), 1993-2024 (2015) DOI:
|
| 27 |
+
[10.1109/TMI.2014.2377694
|
| 28 |
+
|
| 29 |
+
[2] Bakas S, Akbari H, Sotiras A, Bilello M, Rozycki M, Kirby JS,
|
| 30 |
+
[Freymann JB, Farahani K, Davatzikos C. "Advancing The Cancer Genome
|
| 31 |
+
[Atlas glioma MRI collections with expert segmentation labels and
|
| 32 |
+
[radiomic features", Nature Scientific Data, 4:170117 (2017) DOI:
|
| 33 |
+
[10.1038/sdata.2017.117
|
| 34 |
+
|
| 35 |
+
In addition, if there are no restrictions imposed from the
|
| 36 |
+
journal/conference you submit your paper about citing "Data
|
| 37 |
+
Citations", please be specific and also cite the following:
|
| 38 |
+
|
| 39 |
+
[3] Bakas S, Akbari H, Sotiras A, Bilello M, Rozycki M, Kirby J,
|
| 40 |
+
[Freymann J, Farahani K, Davatzikos C. "Segmentation Labels and
|
| 41 |
+
[Radiomic Features for the Pre-operative Scans of the TCGA-GBM
|
| 42 |
+
[collection", The Cancer Imaging Archive, 2017. DOI:
|
| 43 |
+
[10.7937/K9/TCIA.2017.KLXWJJ1Q
|
| 44 |
+
|
| 45 |
+
[4] Bakas S, Akbari H, Sotiras A, Bilello M, Rozycki M, Kirby J,
|
| 46 |
+
[Freymann J, Farahani K, Davatzikos C. "Segmentation Labels and
|
| 47 |
+
[Radiomic Features for the Pre-operative Scans of the TCGA-LGG
|
| 48 |
+
[collection", The Cancer Imaging Archive, 2017. DOI:
|
| 49 |
+
[10.7937/K9/TCIA.2017.GJQ7R0EF
|
models/model.pt
ADDED
|
@@ -0,0 +1,3 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
version https://git-lfs.github.com/spec/v1
|
| 2 |
+
oid sha256:860ccb3f1c21c99d0410ad8a1ac4ef6b8fab60cec0a503b0ba42675741a750ae
|
| 3 |
+
size 18840620
|
models/model.ts
ADDED
|
@@ -0,0 +1,3 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
version https://git-lfs.github.com/spec/v1
|
| 2 |
+
oid sha256:729980a0bd9347bf2397701eb329e12517918dc282a2d09c40458e95b24ceed9
|
| 3 |
+
size 18911784
|
scripts/prepare_datalist.py
ADDED
|
@@ -0,0 +1,72 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
import argparse
|
| 2 |
+
import glob
|
| 3 |
+
import json
|
| 4 |
+
import os
|
| 5 |
+
|
| 6 |
+
import monai
|
| 7 |
+
from sklearn.model_selection import train_test_split
|
| 8 |
+
|
| 9 |
+
|
| 10 |
+
def produce_sample_dict(line: str):
|
| 11 |
+
names = os.listdir(line)
|
| 12 |
+
seg, t1ce, t1, t2, flair = [], [], [], [], []
|
| 13 |
+
for name in names:
|
| 14 |
+
name = os.path.join(line, name)
|
| 15 |
+
if "_seg.nii" in name:
|
| 16 |
+
seg.append(name)
|
| 17 |
+
elif "_t1ce.nii" in name:
|
| 18 |
+
t1ce.append(name)
|
| 19 |
+
elif "_t1.nii" in name:
|
| 20 |
+
t1.append(name)
|
| 21 |
+
elif "_t2.nii" in name:
|
| 22 |
+
t2.append(name)
|
| 23 |
+
elif "_flair.nii" in name:
|
| 24 |
+
flair.append(name)
|
| 25 |
+
|
| 26 |
+
return {"label": seg[0], "image": t1ce + t1 + t2 + flair}
|
| 27 |
+
|
| 28 |
+
|
| 29 |
+
def produce_datalist(dataset_dir: str, train_size: int = 200):
|
| 30 |
+
"""
|
| 31 |
+
This function is used to split the dataset.
|
| 32 |
+
It will produce "train_size" number of samples for training, and the other samples
|
| 33 |
+
are divided equally into val and test sets.
|
| 34 |
+
"""
|
| 35 |
+
|
| 36 |
+
samples = sorted(glob.glob(os.path.join(dataset_dir, "*", "*"), recursive=True))
|
| 37 |
+
datalist = []
|
| 38 |
+
for line in samples:
|
| 39 |
+
datalist.append(produce_sample_dict(line))
|
| 40 |
+
train_list, other_list = train_test_split(datalist, train_size=train_size)
|
| 41 |
+
val_list, test_list = train_test_split(other_list, train_size=0.5)
|
| 42 |
+
|
| 43 |
+
return {"training": train_list, "validation": val_list, "testing": test_list}
|
| 44 |
+
|
| 45 |
+
|
| 46 |
+
def main(args):
|
| 47 |
+
"""
|
| 48 |
+
split the dataset and output the data list into a json file.
|
| 49 |
+
"""
|
| 50 |
+
data_file_base_dir = os.path.join(os.path.abspath(args.path), "training")
|
| 51 |
+
# produce deterministic data splits
|
| 52 |
+
monai.utils.set_determinism(seed=123)
|
| 53 |
+
datalist = produce_datalist(dataset_dir=data_file_base_dir, train_size=args.train_size)
|
| 54 |
+
with open(args.output, "w") as f:
|
| 55 |
+
json.dump(datalist, f)
|
| 56 |
+
|
| 57 |
+
|
| 58 |
+
if __name__ == "__main__":
|
| 59 |
+
parser = argparse.ArgumentParser(description="")
|
| 60 |
+
parser.add_argument(
|
| 61 |
+
"--path",
|
| 62 |
+
type=str,
|
| 63 |
+
default="/workspace/data/medical/brats2018challenge",
|
| 64 |
+
help="root path of brats 2018 dataset.",
|
| 65 |
+
)
|
| 66 |
+
parser.add_argument(
|
| 67 |
+
"--output", type=str, default="configs/datalist.json", help="relative path of output datalist json file."
|
| 68 |
+
)
|
| 69 |
+
parser.add_argument("--train_size", type=int, default=200, help="number of training samples.")
|
| 70 |
+
args = parser.parse_args()
|
| 71 |
+
|
| 72 |
+
main(args)
|