Update README.md
Browse files
README.md
CHANGED
|
@@ -18,13 +18,7 @@ Custom fine-tuned version of NVIDIA's segformer model for colony slides in micro
|
|
| 18 |
Xie, E., Wang, W., Yu, Z., Anandkumar, A., Alvarez, J. M., & Luo, P. (2021). SegFormer: Simple and efficient design for semantic segmentation with transformers. arXiv preprint arXiv:2105.15203. https://arxiv.org/abs/2105.15203
|
| 19 |
|
| 20 |
The Segformer-Cityscapes model was changed to a ternary classifier and fine-tuned on custom training data, where colonies and necrosis were made as separate masks and then merged with different grayscale values.
|
| 21 |
-

|
| 22 |
-

|
| 23 |
-

|
| 24 |
|
| 25 |
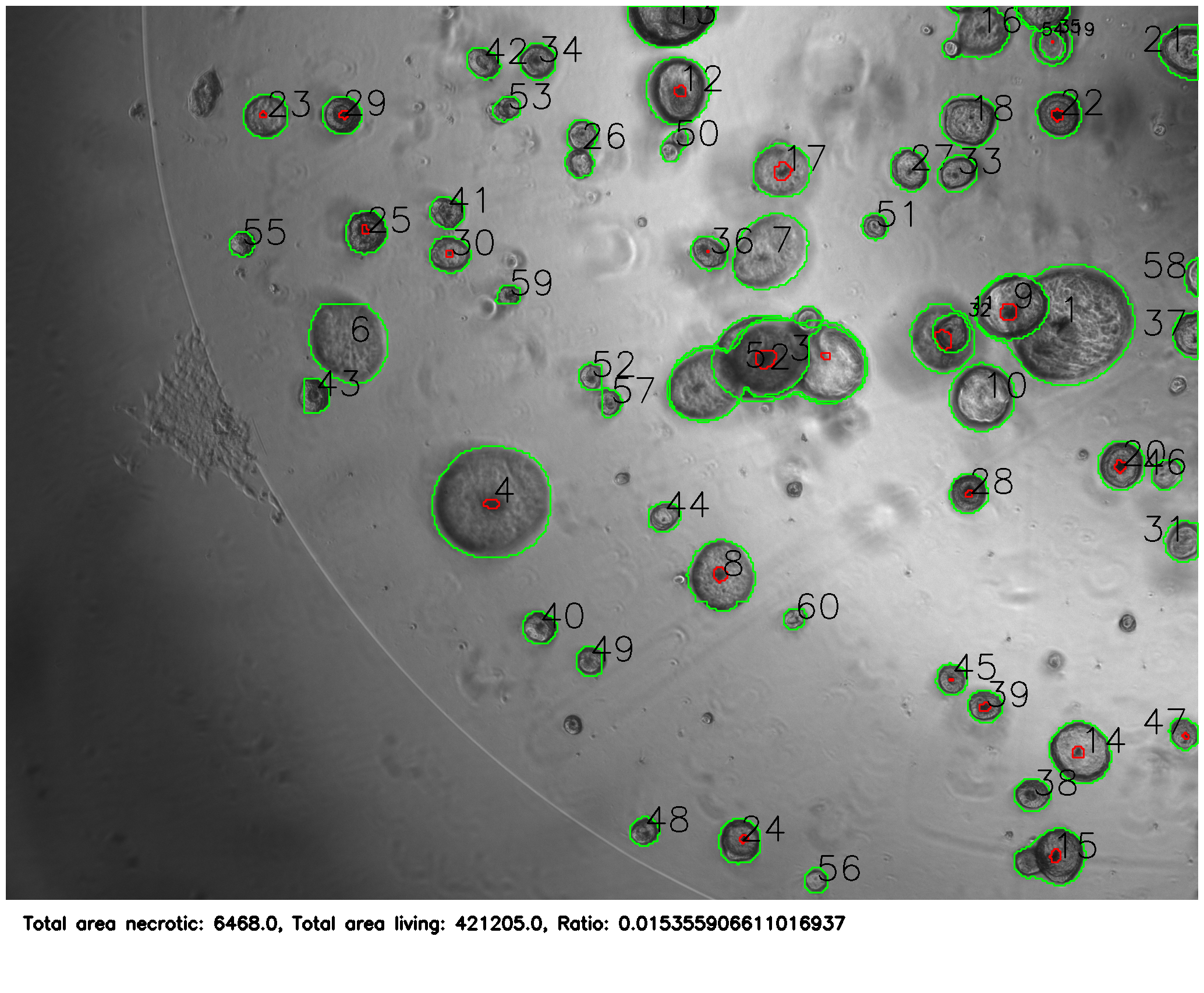
After training, it was able to correctly identify organoids and necrosis.
|
| 26 |
-

|
| 27 |
-

|
| 28 |
|
| 29 |
-
The python program (see linked GitHub) then uses the masks to annotate the images and provide statistics about the colonies.
|
| 30 |
-
|
|
|
|
| 18 |
Xie, E., Wang, W., Yu, Z., Anandkumar, A., Alvarez, J. M., & Luo, P. (2021). SegFormer: Simple and efficient design for semantic segmentation with transformers. arXiv preprint arXiv:2105.15203. https://arxiv.org/abs/2105.15203
|
| 19 |
|
| 20 |
The Segformer-Cityscapes model was changed to a ternary classifier and fine-tuned on custom training data, where colonies and necrosis were made as separate masks and then merged with different grayscale values.
|
|
|
|
|
|
|
|
|
|
| 21 |
|
| 22 |
After training, it was able to correctly identify organoids and necrosis.
|
|
|
|
|
|
|
| 23 |
|
| 24 |
+
The python program (see linked GitHub) then uses the masks to annotate the images and provide statistics about the colonies.
|
|
|