File size: 3,407 Bytes
7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 b6d7045 7fe8119 | 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 | ---
license: bsd-3-clause
library_name: braindecode
pipeline_tag: feature-extraction
tags:
- eeg
- biosignal
- pytorch
- neuroscience
- braindecode
- convolutional
- transformer
---
# EEGConformer
EEG Conformer from Song et al (2022) [song2022].
> **Architecture-only repository.** Documents the
> `braindecode.models.EEGConformer` class. **No pretrained weights are
> distributed here.** Instantiate the model and train it on your own
> data.
## Quick start
```bash
pip install braindecode
```
```python
from braindecode.models import EEGConformer
model = EEGConformer(
n_chans=22,
sfreq=250,
input_window_seconds=4.0,
n_outputs=4,
)
```
The signal-shape arguments above are illustrative defaults — adjust to
match your recording.
## Documentation
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.EEGConformer.html>
- Interactive browser (live instantiation, parameter counts):
<https://huggingface.co/spaces/braindecode/model-explorer>
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/eegconformer.py#L14>
## Architecture
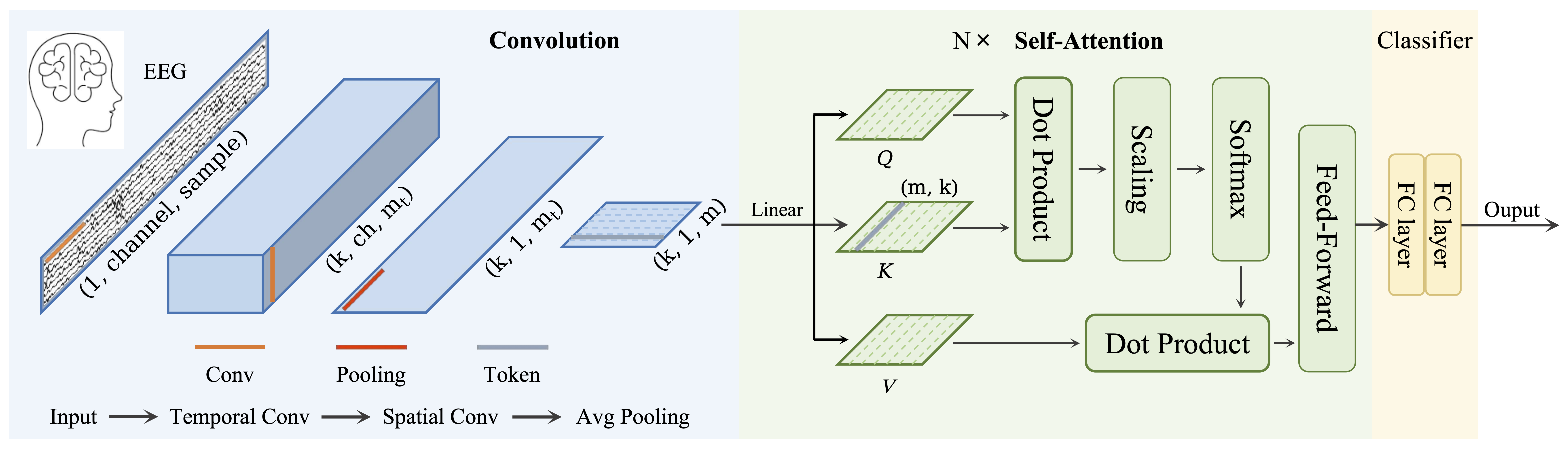
## Parameters
| Parameter | Type | Description |
|---|---|---|
| `n_filters_time: int` | — | Number of temporal filters, defines also embedding size. |
| `filter_time_length: int` | — | Length of the temporal filter. |
| `pool_time_length: int` | — | Length of temporal pooling filter. |
| `pool_time_stride: int` | — | Length of stride between temporal pooling filters. |
| `drop_prob: float` | — | Dropout rate of the convolutional layer. |
| `num_layers: int` | — | Number of self-attention layers. |
| `num_heads: int` | — | Number of attention heads. |
| `att_drop_prob: float` | — | Dropout rate of the self-attention layer. |
| `final_fc_length: int | str` | — | The dimension of the fully connected layer. |
| `return_features: bool` | — | If True, the forward method returns the features before the last classification layer. Defaults to False. |
| `activation: nn.Module` | — | Activation function as parameter. Default is nn.ELU |
| `activation_transfor: nn.Module` | — | Activation function as parameter, applied at the FeedForwardBlock module inside the transformer. Default is nn.GeLU |
## References
1. Song, Y., Zheng, Q., Liu, B. and Gao, X., 2022. EEG conformer: Convolutional transformer for EEG decoding and visualization. IEEE Transactions on Neural Systems and Rehabilitation Engineering, 31, pp.710-719. https://ieeexplore.ieee.org/document/9991178
2. Song, Y., Zheng, Q., Liu, B. and Gao, X., 2022. EEG conformer: Convolutional transformer for EEG decoding and visualization. https://github.com/eeyhsong/EEG-Conformer.
## Citation
Cite the original architecture paper (see *References* above) and braindecode:
```bibtex
@article{aristimunha2025braindecode,
title = {Braindecode: a deep learning library for raw electrophysiological data},
author = {Aristimunha, Bruno and others},
journal = {Zenodo},
year = {2025},
doi = {10.5281/zenodo.17699192},
}
```
## License
BSD-3-Clause for the model code (matching braindecode).
Pretraining-derived weights, if you fine-tune from a checkpoint,
inherit the licence of that checkpoint and its training corpus.
|