Upload MONAI segmentation models
Browse files- README.md +37 -0
- wholeBody_ct_segmentation/.gitattributes +35 -0
- wholeBody_ct_segmentation/LICENSE +201 -0
- wholeBody_ct_segmentation/configs/evaluate.json +80 -0
- wholeBody_ct_segmentation/configs/inference.json +171 -0
- wholeBody_ct_segmentation/configs/inference_trt.json +12 -0
- wholeBody_ct_segmentation/configs/logging.conf +21 -0
- wholeBody_ct_segmentation/configs/metadata.json +203 -0
- wholeBody_ct_segmentation/configs/multi_gpu_evaluate.json +32 -0
- wholeBody_ct_segmentation/configs/multi_gpu_train.json +43 -0
- wholeBody_ct_segmentation/configs/train.json +424 -0
- wholeBody_ct_segmentation/docs/README.md +264 -0
- wholeBody_ct_segmentation/docs/data_license.txt +6 -0
- wholeBody_ct_segmentation/models/model.pt +3 -0
- wholeBody_ct_segmentation/models/model_lowres.pt +3 -0
README.md
ADDED
|
@@ -0,0 +1,37 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
---
|
| 2 |
+
license: apache-2.0
|
| 3 |
+
tags:
|
| 4 |
+
- medical-imaging
|
| 5 |
+
- segmentation
|
| 6 |
+
- monai
|
| 7 |
+
- ct-scan
|
| 8 |
+
---
|
| 9 |
+
|
| 10 |
+
# NeuroScan AI - Medical Imaging Models
|
| 11 |
+
|
| 12 |
+
This repository contains pretrained models for NeuroScan AI medical imaging analysis platform.
|
| 13 |
+
|
| 14 |
+
## Models
|
| 15 |
+
|
| 16 |
+
### wholeBody_ct_segmentation
|
| 17 |
+
- **Description**: Whole body CT segmentation model
|
| 18 |
+
- **Framework**: MONAI
|
| 19 |
+
- **Organs**: 104 anatomical structures
|
| 20 |
+
- **Input**: CT scan (NIfTI format)
|
| 21 |
+
|
| 22 |
+
## Usage
|
| 23 |
+
|
| 24 |
+
```python
|
| 25 |
+
from monai.bundle import download
|
| 26 |
+
|
| 27 |
+
# Download the model
|
| 28 |
+
download(name="wholeBody_ct_segmentation", bundle_dir="./models")
|
| 29 |
+
```
|
| 30 |
+
|
| 31 |
+
## License
|
| 32 |
+
|
| 33 |
+
Apache 2.0
|
| 34 |
+
|
| 35 |
+
## Citation
|
| 36 |
+
|
| 37 |
+
If you use these models, please cite NeuroScan AI project.
|
wholeBody_ct_segmentation/.gitattributes
ADDED
|
@@ -0,0 +1,35 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
*.7z filter=lfs diff=lfs merge=lfs -text
|
| 2 |
+
*.arrow filter=lfs diff=lfs merge=lfs -text
|
| 3 |
+
*.bin filter=lfs diff=lfs merge=lfs -text
|
| 4 |
+
*.bz2 filter=lfs diff=lfs merge=lfs -text
|
| 5 |
+
*.ckpt filter=lfs diff=lfs merge=lfs -text
|
| 6 |
+
*.ftz filter=lfs diff=lfs merge=lfs -text
|
| 7 |
+
*.gz filter=lfs diff=lfs merge=lfs -text
|
| 8 |
+
*.h5 filter=lfs diff=lfs merge=lfs -text
|
| 9 |
+
*.joblib filter=lfs diff=lfs merge=lfs -text
|
| 10 |
+
*.lfs.* filter=lfs diff=lfs merge=lfs -text
|
| 11 |
+
*.mlmodel filter=lfs diff=lfs merge=lfs -text
|
| 12 |
+
*.model filter=lfs diff=lfs merge=lfs -text
|
| 13 |
+
*.msgpack filter=lfs diff=lfs merge=lfs -text
|
| 14 |
+
*.npy filter=lfs diff=lfs merge=lfs -text
|
| 15 |
+
*.npz filter=lfs diff=lfs merge=lfs -text
|
| 16 |
+
*.onnx filter=lfs diff=lfs merge=lfs -text
|
| 17 |
+
*.ot filter=lfs diff=lfs merge=lfs -text
|
| 18 |
+
*.parquet filter=lfs diff=lfs merge=lfs -text
|
| 19 |
+
*.pb filter=lfs diff=lfs merge=lfs -text
|
| 20 |
+
*.pickle filter=lfs diff=lfs merge=lfs -text
|
| 21 |
+
*.pkl filter=lfs diff=lfs merge=lfs -text
|
| 22 |
+
*.pt filter=lfs diff=lfs merge=lfs -text
|
| 23 |
+
*.pth filter=lfs diff=lfs merge=lfs -text
|
| 24 |
+
*.rar filter=lfs diff=lfs merge=lfs -text
|
| 25 |
+
*.safetensors filter=lfs diff=lfs merge=lfs -text
|
| 26 |
+
saved_model/**/* filter=lfs diff=lfs merge=lfs -text
|
| 27 |
+
*.tar.* filter=lfs diff=lfs merge=lfs -text
|
| 28 |
+
*.tar filter=lfs diff=lfs merge=lfs -text
|
| 29 |
+
*.tflite filter=lfs diff=lfs merge=lfs -text
|
| 30 |
+
*.tgz filter=lfs diff=lfs merge=lfs -text
|
| 31 |
+
*.wasm filter=lfs diff=lfs merge=lfs -text
|
| 32 |
+
*.xz filter=lfs diff=lfs merge=lfs -text
|
| 33 |
+
*.zip filter=lfs diff=lfs merge=lfs -text
|
| 34 |
+
*.zst filter=lfs diff=lfs merge=lfs -text
|
| 35 |
+
*tfevents* filter=lfs diff=lfs merge=lfs -text
|
wholeBody_ct_segmentation/LICENSE
ADDED
|
@@ -0,0 +1,201 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
Apache License
|
| 2 |
+
Version 2.0, January 2004
|
| 3 |
+
http://www.apache.org/licenses/
|
| 4 |
+
|
| 5 |
+
TERMS AND CONDITIONS FOR USE, REPRODUCTION, AND DISTRIBUTION
|
| 6 |
+
|
| 7 |
+
1. Definitions.
|
| 8 |
+
|
| 9 |
+
"License" shall mean the terms and conditions for use, reproduction,
|
| 10 |
+
and distribution as defined by Sections 1 through 9 of this document.
|
| 11 |
+
|
| 12 |
+
"Licensor" shall mean the copyright owner or entity authorized by
|
| 13 |
+
the copyright owner that is granting the License.
|
| 14 |
+
|
| 15 |
+
"Legal Entity" shall mean the union of the acting entity and all
|
| 16 |
+
other entities that control, are controlled by, or are under common
|
| 17 |
+
control with that entity. For the purposes of this definition,
|
| 18 |
+
"control" means (i) the power, direct or indirect, to cause the
|
| 19 |
+
direction or management of such entity, whether by contract or
|
| 20 |
+
otherwise, or (ii) ownership of fifty percent (50%) or more of the
|
| 21 |
+
outstanding shares, or (iii) beneficial ownership of such entity.
|
| 22 |
+
|
| 23 |
+
"You" (or "Your") shall mean an individual or Legal Entity
|
| 24 |
+
exercising permissions granted by this License.
|
| 25 |
+
|
| 26 |
+
"Source" form shall mean the preferred form for making modifications,
|
| 27 |
+
including but not limited to software source code, documentation
|
| 28 |
+
source, and configuration files.
|
| 29 |
+
|
| 30 |
+
"Object" form shall mean any form resulting from mechanical
|
| 31 |
+
transformation or translation of a Source form, including but
|
| 32 |
+
not limited to compiled object code, generated documentation,
|
| 33 |
+
and conversions to other media types.
|
| 34 |
+
|
| 35 |
+
"Work" shall mean the work of authorship, whether in Source or
|
| 36 |
+
Object form, made available under the License, as indicated by a
|
| 37 |
+
copyright notice that is included in or attached to the work
|
| 38 |
+
(an example is provided in the Appendix below).
|
| 39 |
+
|
| 40 |
+
"Derivative Works" shall mean any work, whether in Source or Object
|
| 41 |
+
form, that is based on (or derived from) the Work and for which the
|
| 42 |
+
editorial revisions, annotations, elaborations, or other modifications
|
| 43 |
+
represent, as a whole, an original work of authorship. For the purposes
|
| 44 |
+
of this License, Derivative Works shall not include works that remain
|
| 45 |
+
separable from, or merely link (or bind by name) to the interfaces of,
|
| 46 |
+
the Work and Derivative Works thereof.
|
| 47 |
+
|
| 48 |
+
"Contribution" shall mean any work of authorship, including
|
| 49 |
+
the original version of the Work and any modifications or additions
|
| 50 |
+
to that Work or Derivative Works thereof, that is intentionally
|
| 51 |
+
submitted to Licensor for inclusion in the Work by the copyright owner
|
| 52 |
+
or by an individual or Legal Entity authorized to submit on behalf of
|
| 53 |
+
the copyright owner. For the purposes of this definition, "submitted"
|
| 54 |
+
means any form of electronic, verbal, or written communication sent
|
| 55 |
+
to the Licensor or its representatives, including but not limited to
|
| 56 |
+
communication on electronic mailing lists, source code control systems,
|
| 57 |
+
and issue tracking systems that are managed by, or on behalf of, the
|
| 58 |
+
Licensor for the purpose of discussing and improving the Work, but
|
| 59 |
+
excluding communication that is conspicuously marked or otherwise
|
| 60 |
+
designated in writing by the copyright owner as "Not a Contribution."
|
| 61 |
+
|
| 62 |
+
"Contributor" shall mean Licensor and any individual or Legal Entity
|
| 63 |
+
on behalf of whom a Contribution has been received by Licensor and
|
| 64 |
+
subsequently incorporated within the Work.
|
| 65 |
+
|
| 66 |
+
2. Grant of Copyright License. Subject to the terms and conditions of
|
| 67 |
+
this License, each Contributor hereby grants to You a perpetual,
|
| 68 |
+
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
|
| 69 |
+
copyright license to reproduce, prepare Derivative Works of,
|
| 70 |
+
publicly display, publicly perform, sublicense, and distribute the
|
| 71 |
+
Work and such Derivative Works in Source or Object form.
|
| 72 |
+
|
| 73 |
+
3. Grant of Patent License. Subject to the terms and conditions of
|
| 74 |
+
this License, each Contributor hereby grants to You a perpetual,
|
| 75 |
+
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
|
| 76 |
+
(except as stated in this section) patent license to make, have made,
|
| 77 |
+
use, offer to sell, sell, import, and otherwise transfer the Work,
|
| 78 |
+
where such license applies only to those patent claims licensable
|
| 79 |
+
by such Contributor that are necessarily infringed by their
|
| 80 |
+
Contribution(s) alone or by combination of their Contribution(s)
|
| 81 |
+
with the Work to which such Contribution(s) was submitted. If You
|
| 82 |
+
institute patent litigation against any entity (including a
|
| 83 |
+
cross-claim or counterclaim in a lawsuit) alleging that the Work
|
| 84 |
+
or a Contribution incorporated within the Work constitutes direct
|
| 85 |
+
or contributory patent infringement, then any patent licenses
|
| 86 |
+
granted to You under this License for that Work shall terminate
|
| 87 |
+
as of the date such litigation is filed.
|
| 88 |
+
|
| 89 |
+
4. Redistribution. You may reproduce and distribute copies of the
|
| 90 |
+
Work or Derivative Works thereof in any medium, with or without
|
| 91 |
+
modifications, and in Source or Object form, provided that You
|
| 92 |
+
meet the following conditions:
|
| 93 |
+
|
| 94 |
+
(a) You must give any other recipients of the Work or
|
| 95 |
+
Derivative Works a copy of this License; and
|
| 96 |
+
|
| 97 |
+
(b) You must cause any modified files to carry prominent notices
|
| 98 |
+
stating that You changed the files; and
|
| 99 |
+
|
| 100 |
+
(c) You must retain, in the Source form of any Derivative Works
|
| 101 |
+
that You distribute, all copyright, patent, trademark, and
|
| 102 |
+
attribution notices from the Source form of the Work,
|
| 103 |
+
excluding those notices that do not pertain to any part of
|
| 104 |
+
the Derivative Works; and
|
| 105 |
+
|
| 106 |
+
(d) If the Work includes a "NOTICE" text file as part of its
|
| 107 |
+
distribution, then any Derivative Works that You distribute must
|
| 108 |
+
include a readable copy of the attribution notices contained
|
| 109 |
+
within such NOTICE file, excluding those notices that do not
|
| 110 |
+
pertain to any part of the Derivative Works, in at least one
|
| 111 |
+
of the following places: within a NOTICE text file distributed
|
| 112 |
+
as part of the Derivative Works; within the Source form or
|
| 113 |
+
documentation, if provided along with the Derivative Works; or,
|
| 114 |
+
within a display generated by the Derivative Works, if and
|
| 115 |
+
wherever such third-party notices normally appear. The contents
|
| 116 |
+
of the NOTICE file are for informational purposes only and
|
| 117 |
+
do not modify the License. You may add Your own attribution
|
| 118 |
+
notices within Derivative Works that You distribute, alongside
|
| 119 |
+
or as an addendum to the NOTICE text from the Work, provided
|
| 120 |
+
that such additional attribution notices cannot be construed
|
| 121 |
+
as modifying the License.
|
| 122 |
+
|
| 123 |
+
You may add Your own copyright statement to Your modifications and
|
| 124 |
+
may provide additional or different license terms and conditions
|
| 125 |
+
for use, reproduction, or distribution of Your modifications, or
|
| 126 |
+
for any such Derivative Works as a whole, provided Your use,
|
| 127 |
+
reproduction, and distribution of the Work otherwise complies with
|
| 128 |
+
the conditions stated in this License.
|
| 129 |
+
|
| 130 |
+
5. Submission of Contributions. Unless You explicitly state otherwise,
|
| 131 |
+
any Contribution intentionally submitted for inclusion in the Work
|
| 132 |
+
by You to the Licensor shall be under the terms and conditions of
|
| 133 |
+
this License, without any additional terms or conditions.
|
| 134 |
+
Notwithstanding the above, nothing herein shall supersede or modify
|
| 135 |
+
the terms of any separate license agreement you may have executed
|
| 136 |
+
with Licensor regarding such Contributions.
|
| 137 |
+
|
| 138 |
+
6. Trademarks. This License does not grant permission to use the trade
|
| 139 |
+
names, trademarks, service marks, or product names of the Licensor,
|
| 140 |
+
except as required for reasonable and customary use in describing the
|
| 141 |
+
origin of the Work and reproducing the content of the NOTICE file.
|
| 142 |
+
|
| 143 |
+
7. Disclaimer of Warranty. Unless required by applicable law or
|
| 144 |
+
agreed to in writing, Licensor provides the Work (and each
|
| 145 |
+
Contributor provides its Contributions) on an "AS IS" BASIS,
|
| 146 |
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or
|
| 147 |
+
implied, including, without limitation, any warranties or conditions
|
| 148 |
+
of TITLE, NON-INFRINGEMENT, MERCHANTABILITY, or FITNESS FOR A
|
| 149 |
+
PARTICULAR PURPOSE. You are solely responsible for determining the
|
| 150 |
+
appropriateness of using or redistributing the Work and assume any
|
| 151 |
+
risks associated with Your exercise of permissions under this License.
|
| 152 |
+
|
| 153 |
+
8. Limitation of Liability. In no event and under no legal theory,
|
| 154 |
+
whether in tort (including negligence), contract, or otherwise,
|
| 155 |
+
unless required by applicable law (such as deliberate and grossly
|
| 156 |
+
negligent acts) or agreed to in writing, shall any Contributor be
|
| 157 |
+
liable to You for damages, including any direct, indirect, special,
|
| 158 |
+
incidental, or consequential damages of any character arising as a
|
| 159 |
+
result of this License or out of the use or inability to use the
|
| 160 |
+
Work (including but not limited to damages for loss of goodwill,
|
| 161 |
+
work stoppage, computer failure or malfunction, or any and all
|
| 162 |
+
other commercial damages or losses), even if such Contributor
|
| 163 |
+
has been advised of the possibility of such damages.
|
| 164 |
+
|
| 165 |
+
9. Accepting Warranty or Additional Liability. While redistributing
|
| 166 |
+
the Work or Derivative Works thereof, You may choose to offer,
|
| 167 |
+
and charge a fee for, acceptance of support, warranty, indemnity,
|
| 168 |
+
or other liability obligations and/or rights consistent with this
|
| 169 |
+
License. However, in accepting such obligations, You may act only
|
| 170 |
+
on Your own behalf and on Your sole responsibility, not on behalf
|
| 171 |
+
of any other Contributor, and only if You agree to indemnify,
|
| 172 |
+
defend, and hold each Contributor harmless for any liability
|
| 173 |
+
incurred by, or claims asserted against, such Contributor by reason
|
| 174 |
+
of your accepting any such warranty or additional liability.
|
| 175 |
+
|
| 176 |
+
END OF TERMS AND CONDITIONS
|
| 177 |
+
|
| 178 |
+
APPENDIX: How to apply the Apache License to your work.
|
| 179 |
+
|
| 180 |
+
To apply the Apache License to your work, attach the following
|
| 181 |
+
boilerplate notice, with the fields enclosed by brackets "[]"
|
| 182 |
+
replaced with your own identifying information. (Don't include
|
| 183 |
+
the brackets!) The text should be enclosed in the appropriate
|
| 184 |
+
comment syntax for the file format. We also recommend that a
|
| 185 |
+
file or class name and description of purpose be included on the
|
| 186 |
+
same "printed page" as the copyright notice for easier
|
| 187 |
+
identification within third-party archives.
|
| 188 |
+
|
| 189 |
+
Copyright [yyyy] [name of copyright owner]
|
| 190 |
+
|
| 191 |
+
Licensed under the Apache License, Version 2.0 (the "License");
|
| 192 |
+
you may not use this file except in compliance with the License.
|
| 193 |
+
You may obtain a copy of the License at
|
| 194 |
+
|
| 195 |
+
http://www.apache.org/licenses/LICENSE-2.0
|
| 196 |
+
|
| 197 |
+
Unless required by applicable law or agreed to in writing, software
|
| 198 |
+
distributed under the License is distributed on an "AS IS" BASIS,
|
| 199 |
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
| 200 |
+
See the License for the specific language governing permissions and
|
| 201 |
+
limitations under the License.
|
wholeBody_ct_segmentation/configs/evaluate.json
ADDED
|
@@ -0,0 +1,80 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"validate#postprocessing": {
|
| 3 |
+
"_target_": "Compose",
|
| 4 |
+
"transforms": [
|
| 5 |
+
{
|
| 6 |
+
"_target_": "Activationsd",
|
| 7 |
+
"keys": "pred",
|
| 8 |
+
"softmax": true
|
| 9 |
+
},
|
| 10 |
+
{
|
| 11 |
+
"_target_": "Invertd",
|
| 12 |
+
"keys": [
|
| 13 |
+
"pred",
|
| 14 |
+
"label"
|
| 15 |
+
],
|
| 16 |
+
"transform": "@validate#preprocessing",
|
| 17 |
+
"orig_keys": "image",
|
| 18 |
+
"meta_key_postfix": "meta_dict",
|
| 19 |
+
"nearest_interp": [
|
| 20 |
+
true,
|
| 21 |
+
true
|
| 22 |
+
],
|
| 23 |
+
"to_tensor": true
|
| 24 |
+
},
|
| 25 |
+
{
|
| 26 |
+
"_target_": "AsDiscreted",
|
| 27 |
+
"keys": [
|
| 28 |
+
"pred",
|
| 29 |
+
"label"
|
| 30 |
+
],
|
| 31 |
+
"argmax": [
|
| 32 |
+
true,
|
| 33 |
+
false
|
| 34 |
+
],
|
| 35 |
+
"to_onehot": 105
|
| 36 |
+
},
|
| 37 |
+
{
|
| 38 |
+
"_target_": "SaveImaged",
|
| 39 |
+
"_disabled_": true,
|
| 40 |
+
"keys": "pred",
|
| 41 |
+
"meta_keys": "pred_meta_dict",
|
| 42 |
+
"output_dir": "@output_dir",
|
| 43 |
+
"resample": false,
|
| 44 |
+
"squeeze_end_dims": true
|
| 45 |
+
}
|
| 46 |
+
]
|
| 47 |
+
},
|
| 48 |
+
"validate#handlers": [
|
| 49 |
+
{
|
| 50 |
+
"_target_": "CheckpointLoader",
|
| 51 |
+
"load_path": "$@ckpt_dir + '/model.pt'",
|
| 52 |
+
"load_dict": {
|
| 53 |
+
"model": "@network"
|
| 54 |
+
}
|
| 55 |
+
},
|
| 56 |
+
{
|
| 57 |
+
"_target_": "StatsHandler",
|
| 58 |
+
"iteration_log": false
|
| 59 |
+
},
|
| 60 |
+
{
|
| 61 |
+
"_target_": "MetricsSaver",
|
| 62 |
+
"save_dir": "@output_dir",
|
| 63 |
+
"metrics": [
|
| 64 |
+
"val_mean_dice",
|
| 65 |
+
"val_acc"
|
| 66 |
+
],
|
| 67 |
+
"metric_details": [
|
| 68 |
+
"val_mean_dice"
|
| 69 |
+
],
|
| 70 |
+
"batch_transform": "$lambda x: [xx['image'].meta for xx in x]",
|
| 71 |
+
"summary_ops": "*"
|
| 72 |
+
}
|
| 73 |
+
],
|
| 74 |
+
"initialize": [
|
| 75 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)"
|
| 76 |
+
],
|
| 77 |
+
"run": [
|
| 78 |
+
"$@validate#evaluator.run()"
|
| 79 |
+
]
|
| 80 |
+
}
|
wholeBody_ct_segmentation/configs/inference.json
ADDED
|
@@ -0,0 +1,171 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"displayable_configs": {
|
| 3 |
+
"highres": true,
|
| 4 |
+
"sw_overlap": 0.25,
|
| 5 |
+
"sw_batch_size": 1
|
| 6 |
+
},
|
| 7 |
+
"imports": [
|
| 8 |
+
"$import glob",
|
| 9 |
+
"$import numpy",
|
| 10 |
+
"$import os"
|
| 11 |
+
],
|
| 12 |
+
"bundle_root": ".",
|
| 13 |
+
"image_key": "image",
|
| 14 |
+
"output_dir": "$@bundle_root + '/eval'",
|
| 15 |
+
"output_ext": ".nii.gz",
|
| 16 |
+
"output_dtype": "$numpy.float32",
|
| 17 |
+
"output_postfix": "trans",
|
| 18 |
+
"separate_folder": true,
|
| 19 |
+
"load_pretrain": true,
|
| 20 |
+
"dataset_dir": "sampledata",
|
| 21 |
+
"datalist": "$list(sorted(glob.glob(@dataset_dir + '/imagesTs/*.nii.gz')))",
|
| 22 |
+
"device": "$torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')",
|
| 23 |
+
"pixdim": "$[1.5, 1.5, 1.5] if @displayable_configs#highres else [3.0, 3.0, 3.0]",
|
| 24 |
+
"modelname": "$'model.pt' if @displayable_configs#highres else 'model_lowres.pt'",
|
| 25 |
+
"network_def": {
|
| 26 |
+
"_target_": "SegResNet",
|
| 27 |
+
"spatial_dims": 3,
|
| 28 |
+
"in_channels": 1,
|
| 29 |
+
"out_channels": 105,
|
| 30 |
+
"init_filters": 32,
|
| 31 |
+
"blocks_down": [
|
| 32 |
+
1,
|
| 33 |
+
2,
|
| 34 |
+
2,
|
| 35 |
+
4
|
| 36 |
+
],
|
| 37 |
+
"blocks_up": [
|
| 38 |
+
1,
|
| 39 |
+
1,
|
| 40 |
+
1
|
| 41 |
+
],
|
| 42 |
+
"dropout_prob": 0.2
|
| 43 |
+
},
|
| 44 |
+
"network": "$@network_def.to(@device)",
|
| 45 |
+
"preprocessing": {
|
| 46 |
+
"_target_": "Compose",
|
| 47 |
+
"transforms": [
|
| 48 |
+
{
|
| 49 |
+
"_target_": "LoadImaged",
|
| 50 |
+
"keys": "@image_key"
|
| 51 |
+
},
|
| 52 |
+
{
|
| 53 |
+
"_target_": "EnsureTyped",
|
| 54 |
+
"keys": "@image_key"
|
| 55 |
+
},
|
| 56 |
+
{
|
| 57 |
+
"_target_": "EnsureChannelFirstd",
|
| 58 |
+
"keys": "@image_key"
|
| 59 |
+
},
|
| 60 |
+
{
|
| 61 |
+
"_target_": "Orientationd",
|
| 62 |
+
"keys": "@image_key",
|
| 63 |
+
"axcodes": "RAS"
|
| 64 |
+
},
|
| 65 |
+
{
|
| 66 |
+
"_target_": "Spacingd",
|
| 67 |
+
"keys": "@image_key",
|
| 68 |
+
"pixdim": "@pixdim",
|
| 69 |
+
"mode": "bilinear"
|
| 70 |
+
},
|
| 71 |
+
{
|
| 72 |
+
"_target_": "NormalizeIntensityd",
|
| 73 |
+
"keys": "@image_key",
|
| 74 |
+
"nonzero": true
|
| 75 |
+
},
|
| 76 |
+
{
|
| 77 |
+
"_target_": "ScaleIntensityd",
|
| 78 |
+
"keys": "@image_key",
|
| 79 |
+
"minv": -1.0,
|
| 80 |
+
"maxv": 1.0
|
| 81 |
+
}
|
| 82 |
+
]
|
| 83 |
+
},
|
| 84 |
+
"dataset": {
|
| 85 |
+
"_target_": "Dataset",
|
| 86 |
+
"data": "$[{'image': i} for i in @datalist]",
|
| 87 |
+
"transform": "@preprocessing"
|
| 88 |
+
},
|
| 89 |
+
"dataloader": {
|
| 90 |
+
"_target_": "DataLoader",
|
| 91 |
+
"dataset": "@dataset",
|
| 92 |
+
"batch_size": 1,
|
| 93 |
+
"shuffle": false,
|
| 94 |
+
"num_workers": 1
|
| 95 |
+
},
|
| 96 |
+
"inferer": {
|
| 97 |
+
"_target_": "SlidingWindowInferer",
|
| 98 |
+
"roi_size": [
|
| 99 |
+
96,
|
| 100 |
+
96,
|
| 101 |
+
96
|
| 102 |
+
],
|
| 103 |
+
"sw_batch_size": "@displayable_configs#sw_batch_size",
|
| 104 |
+
"overlap": "@displayable_configs#sw_overlap",
|
| 105 |
+
"padding_mode": "replicate",
|
| 106 |
+
"mode": "gaussian",
|
| 107 |
+
"device": "$torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')"
|
| 108 |
+
},
|
| 109 |
+
"postprocessing": {
|
| 110 |
+
"_target_": "Compose",
|
| 111 |
+
"transforms": [
|
| 112 |
+
{
|
| 113 |
+
"_target_": "Activationsd",
|
| 114 |
+
"keys": "pred",
|
| 115 |
+
"softmax": true
|
| 116 |
+
},
|
| 117 |
+
{
|
| 118 |
+
"_target_": "AsDiscreted",
|
| 119 |
+
"keys": "pred",
|
| 120 |
+
"argmax": true
|
| 121 |
+
},
|
| 122 |
+
{
|
| 123 |
+
"_target_": "Invertd",
|
| 124 |
+
"keys": "pred",
|
| 125 |
+
"transform": "@preprocessing",
|
| 126 |
+
"orig_keys": "@image_key",
|
| 127 |
+
"nearest_interp": true,
|
| 128 |
+
"to_tensor": true
|
| 129 |
+
},
|
| 130 |
+
{
|
| 131 |
+
"_target_": "SaveImaged",
|
| 132 |
+
"keys": "pred",
|
| 133 |
+
"output_dir": "@output_dir",
|
| 134 |
+
"output_ext": "@output_ext",
|
| 135 |
+
"output_dtype": "@output_dtype",
|
| 136 |
+
"output_postfix": "@output_postfix",
|
| 137 |
+
"separate_folder": "@separate_folder"
|
| 138 |
+
}
|
| 139 |
+
]
|
| 140 |
+
},
|
| 141 |
+
"handlers": [
|
| 142 |
+
{
|
| 143 |
+
"_target_": "StatsHandler",
|
| 144 |
+
"iteration_log": false
|
| 145 |
+
}
|
| 146 |
+
],
|
| 147 |
+
"evaluator": {
|
| 148 |
+
"_target_": "SupervisedEvaluator",
|
| 149 |
+
"device": "@device",
|
| 150 |
+
"val_data_loader": "@dataloader",
|
| 151 |
+
"network": "@network",
|
| 152 |
+
"inferer": "@inferer",
|
| 153 |
+
"postprocessing": "@postprocessing",
|
| 154 |
+
"val_handlers": "@handlers",
|
| 155 |
+
"amp": true
|
| 156 |
+
},
|
| 157 |
+
"checkpointloader": {
|
| 158 |
+
"_target_": "CheckpointLoader",
|
| 159 |
+
"load_path": "$@bundle_root + '/models/' + @modelname",
|
| 160 |
+
"load_dict": {
|
| 161 |
+
"model": "@network"
|
| 162 |
+
}
|
| 163 |
+
},
|
| 164 |
+
"initialize": [
|
| 165 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)",
|
| 166 |
+
"$@checkpointloader(@evaluator) if @load_pretrain else None"
|
| 167 |
+
],
|
| 168 |
+
"run": [
|
| 169 |
+
"$@evaluator.run()"
|
| 170 |
+
]
|
| 171 |
+
}
|
wholeBody_ct_segmentation/configs/inference_trt.json
ADDED
|
@@ -0,0 +1,12 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"imports": [
|
| 3 |
+
"$import glob",
|
| 4 |
+
"$import os",
|
| 5 |
+
"$import torch_tensorrt"
|
| 6 |
+
],
|
| 7 |
+
"network_def": "$torch.jit.load(@bundle_root + '/models/model_trt.ts')",
|
| 8 |
+
"evaluator#amp": false,
|
| 9 |
+
"initialize": [
|
| 10 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)"
|
| 11 |
+
]
|
| 12 |
+
}
|
wholeBody_ct_segmentation/configs/logging.conf
ADDED
|
@@ -0,0 +1,21 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
[loggers]
|
| 2 |
+
keys=root
|
| 3 |
+
|
| 4 |
+
[handlers]
|
| 5 |
+
keys=consoleHandler
|
| 6 |
+
|
| 7 |
+
[formatters]
|
| 8 |
+
keys=fullFormatter
|
| 9 |
+
|
| 10 |
+
[logger_root]
|
| 11 |
+
level=INFO
|
| 12 |
+
handlers=consoleHandler
|
| 13 |
+
|
| 14 |
+
[handler_consoleHandler]
|
| 15 |
+
class=StreamHandler
|
| 16 |
+
level=INFO
|
| 17 |
+
formatter=fullFormatter
|
| 18 |
+
args=(sys.stdout,)
|
| 19 |
+
|
| 20 |
+
[formatter_fullFormatter]
|
| 21 |
+
format=%(asctime)s - %(name)s - %(levelname)s - %(message)s
|
wholeBody_ct_segmentation/configs/metadata.json
ADDED
|
@@ -0,0 +1,203 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"schema": "https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/meta_schema_20240725.json",
|
| 3 |
+
"version": "0.2.7",
|
| 4 |
+
"changelog": {
|
| 5 |
+
"0.2.7": "enhance metadata with improved descriptions",
|
| 6 |
+
"0.2.6": "update to huggingface hosting",
|
| 7 |
+
"0.2.5": "use monai 1.4 and update large files",
|
| 8 |
+
"0.2.4": "update to use monai 1.3.1",
|
| 9 |
+
"0.2.3": "add load_pretrain flag for infer",
|
| 10 |
+
"0.2.2": "add checkpoint loader for infer",
|
| 11 |
+
"0.2.1": "remove meta_dict usage",
|
| 12 |
+
"0.2.0": "add support for TensorRT conversion and inference",
|
| 13 |
+
"0.1.9": "fix the wrong GPU index issue of multi-node",
|
| 14 |
+
"0.1.8": "Update evalaute doc, GPU usage details, and dataset preparation instructions",
|
| 15 |
+
"0.1.7": "remove error dollar symbol in readme",
|
| 16 |
+
"0.1.6": "add RAM usage with CacheDataset and GPU consumtion warning",
|
| 17 |
+
"0.1.5": "fix mgpu finalize issue",
|
| 18 |
+
"0.1.4": "Update README Formatting",
|
| 19 |
+
"0.1.3": "add non-deterministic note",
|
| 20 |
+
"0.1.2": "Update figure with links",
|
| 21 |
+
"0.1.1": "adapt to BundleWorkflow interface and val metric",
|
| 22 |
+
"0.1.0": "complete the model package",
|
| 23 |
+
"0.0.1": "initialize the model package structure"
|
| 24 |
+
},
|
| 25 |
+
"monai_version": "1.4.0",
|
| 26 |
+
"pytorch_version": "2.4.0",
|
| 27 |
+
"numpy_version": "1.24.4",
|
| 28 |
+
"required_packages_version": {
|
| 29 |
+
"itk": "5.4.0",
|
| 30 |
+
"nibabel": "5.2.1",
|
| 31 |
+
"pytorch-ignite": "0.4.11",
|
| 32 |
+
"tensorboard": "2.17.0"
|
| 33 |
+
},
|
| 34 |
+
"supported_apps": {},
|
| 35 |
+
"name": "Whole Body CT Segmentation",
|
| 36 |
+
"task": "Multi-organ Segmentation in Whole Body CT Scans",
|
| 37 |
+
"description": "A SegResNet-based volumetric segmentation model that segments 104 distinct anatomical structures from CT scans. The model processes 96x96x96 pixel patches and provides segmentation masks for major organs, bones, muscles, and vascular structures throughout the body, trained on TotalSegmentator data.",
|
| 38 |
+
"authors": "MONAI team",
|
| 39 |
+
"copyright": "Copyright (c) MONAI Consortium",
|
| 40 |
+
"data_source": "TotalSegmentator",
|
| 41 |
+
"data_type": "nibabel",
|
| 42 |
+
"image_classes": "104 foreground channels, 0 channel for the background, intensity scaled to [0, 1]",
|
| 43 |
+
"label_classes": "0 is the background, others are whole body segments",
|
| 44 |
+
"pred_classes": "0 is the background, 104 other chanels are whole body segments",
|
| 45 |
+
"eval_metrics": {
|
| 46 |
+
"mean_dice": 0.8
|
| 47 |
+
},
|
| 48 |
+
"intended_use": "This is an example, not to be used for diagnostic purposes",
|
| 49 |
+
"references": [
|
| 50 |
+
"Wasserthal, J., Meyer, M., Breit, H.C., Cyriac, J., Yang, S. and Segeroth, M., 2022. TotalSegmentator: robust segmentation of 104 anatomical structures in CT images. arXiv preprint arXiv:2208.05868.",
|
| 51 |
+
"Myronenko, A., Siddiquee, M.M.R., Yang, D., He, Y. and Xu, D., 2022. Automated head and neck tumor segmentation from 3D PET/CT. arXiv preprint arXiv:2209.10809.",
|
| 52 |
+
"Tang, Y., Gao, R., Lee, H.H., Han, S., Chen, Y., Gao, D., Nath, V., Bermudez, C., Savona, M.R., Abramson, R.G. and Bao, S., 2021. High-resolution 3D abdominal segmentation with random patch network fusion. Medical image analysis, 69, p.101894."
|
| 53 |
+
],
|
| 54 |
+
"network_data_format": {
|
| 55 |
+
"inputs": {
|
| 56 |
+
"image": {
|
| 57 |
+
"type": "image",
|
| 58 |
+
"format": "hounsfield",
|
| 59 |
+
"modality": "CT",
|
| 60 |
+
"num_channels": 1,
|
| 61 |
+
"spatial_shape": [
|
| 62 |
+
96,
|
| 63 |
+
96,
|
| 64 |
+
96
|
| 65 |
+
],
|
| 66 |
+
"dtype": "float32",
|
| 67 |
+
"value_range": [
|
| 68 |
+
0,
|
| 69 |
+
1
|
| 70 |
+
],
|
| 71 |
+
"is_patch_data": true,
|
| 72 |
+
"channel_def": {
|
| 73 |
+
"0": "image"
|
| 74 |
+
}
|
| 75 |
+
}
|
| 76 |
+
},
|
| 77 |
+
"outputs": {
|
| 78 |
+
"pred": {
|
| 79 |
+
"type": "image",
|
| 80 |
+
"format": "segmentation",
|
| 81 |
+
"num_channels": 105,
|
| 82 |
+
"spatial_shape": [
|
| 83 |
+
96,
|
| 84 |
+
96,
|
| 85 |
+
96
|
| 86 |
+
],
|
| 87 |
+
"dtype": "float32",
|
| 88 |
+
"value_range": [
|
| 89 |
+
0,
|
| 90 |
+
104
|
| 91 |
+
],
|
| 92 |
+
"is_patch_data": true,
|
| 93 |
+
"channel_def": {
|
| 94 |
+
"0": "background",
|
| 95 |
+
"1": "spleen",
|
| 96 |
+
"2": "kidney_right",
|
| 97 |
+
"3": "kidney_left",
|
| 98 |
+
"4": "gallbladder",
|
| 99 |
+
"5": "liver",
|
| 100 |
+
"6": "stomach",
|
| 101 |
+
"7": "aorta",
|
| 102 |
+
"8": "inferior_vena_cava",
|
| 103 |
+
"9": "portal_vein_and_splenic_vein",
|
| 104 |
+
"10": "pancreas",
|
| 105 |
+
"11": "adrenal_gland_right",
|
| 106 |
+
"12": "adrenal_gland_left",
|
| 107 |
+
"13": "lung_upper_lobe_left",
|
| 108 |
+
"14": "lung_lower_lobe_left",
|
| 109 |
+
"15": "lung_upper_lobe_right",
|
| 110 |
+
"16": "lung_middle_lobe_right",
|
| 111 |
+
"17": "lung_lower_lobe_right",
|
| 112 |
+
"18": "vertebrae_L5",
|
| 113 |
+
"19": "vertebrae_L4",
|
| 114 |
+
"20": "vertebrae_L3",
|
| 115 |
+
"21": "vertebrae_L2",
|
| 116 |
+
"22": "vertebrae_L1",
|
| 117 |
+
"23": "vertebrae_T12",
|
| 118 |
+
"24": "vertebrae_T11",
|
| 119 |
+
"25": "vertebrae_T10",
|
| 120 |
+
"26": "vertebrae_T9",
|
| 121 |
+
"27": "vertebrae_T8",
|
| 122 |
+
"28": "vertebrae_T7",
|
| 123 |
+
"29": "vertebrae_T6",
|
| 124 |
+
"30": "vertebrae_T5",
|
| 125 |
+
"31": "vertebrae_T4",
|
| 126 |
+
"32": "vertebrae_T3",
|
| 127 |
+
"33": "vertebrae_T2",
|
| 128 |
+
"34": "vertebrae_T1",
|
| 129 |
+
"35": "vertebrae_C7",
|
| 130 |
+
"36": "vertebrae_C6",
|
| 131 |
+
"37": "vertebrae_C5",
|
| 132 |
+
"38": "vertebrae_C4",
|
| 133 |
+
"39": "vertebrae_C3",
|
| 134 |
+
"40": "vertebrae_C2",
|
| 135 |
+
"41": "vertebrae_C1",
|
| 136 |
+
"42": "esophagus",
|
| 137 |
+
"43": "trachea",
|
| 138 |
+
"44": "heart_myocardium",
|
| 139 |
+
"45": "heart_atrium_left",
|
| 140 |
+
"46": "heart_ventricle_left",
|
| 141 |
+
"47": "heart_atrium_right",
|
| 142 |
+
"48": "heart_ventricle_right",
|
| 143 |
+
"49": "pulmonary_artery",
|
| 144 |
+
"50": "brain",
|
| 145 |
+
"51": "iliac_artery_left",
|
| 146 |
+
"52": "iliac_artery_right",
|
| 147 |
+
"53": "iliac_vena_left",
|
| 148 |
+
"54": "iliac_vena_right",
|
| 149 |
+
"55": "small_bowel",
|
| 150 |
+
"56": "duodenum",
|
| 151 |
+
"57": "colon",
|
| 152 |
+
"58": "rib_left_1",
|
| 153 |
+
"59": "rib_left_2",
|
| 154 |
+
"60": "rib_left_3",
|
| 155 |
+
"61": "rib_left_4",
|
| 156 |
+
"62": "rib_left_5",
|
| 157 |
+
"63": "rib_left_6",
|
| 158 |
+
"64": "rib_left_7",
|
| 159 |
+
"65": "rib_left_8",
|
| 160 |
+
"66": "rib_left_9",
|
| 161 |
+
"67": "rib_left_10",
|
| 162 |
+
"68": "rib_left_11",
|
| 163 |
+
"69": "rib_left_12",
|
| 164 |
+
"70": "rib_right_1",
|
| 165 |
+
"71": "rib_right_2",
|
| 166 |
+
"72": "rib_right_3",
|
| 167 |
+
"73": "rib_right_4",
|
| 168 |
+
"74": "rib_right_5",
|
| 169 |
+
"75": "rib_right_6",
|
| 170 |
+
"76": "rib_right_7",
|
| 171 |
+
"77": "rib_right_8",
|
| 172 |
+
"78": "rib_right_9",
|
| 173 |
+
"79": "rib_right_10",
|
| 174 |
+
"80": "rib_right_11",
|
| 175 |
+
"81": "rib_right_12",
|
| 176 |
+
"82": "humerus_left",
|
| 177 |
+
"83": "humerus_right",
|
| 178 |
+
"84": "scapula_left",
|
| 179 |
+
"85": "scapula_right",
|
| 180 |
+
"86": "clavicula_left",
|
| 181 |
+
"87": "clavicula_right",
|
| 182 |
+
"88": "femur_left",
|
| 183 |
+
"89": "femur_right",
|
| 184 |
+
"90": "hip_left",
|
| 185 |
+
"91": "hip_right",
|
| 186 |
+
"92": "sacrum",
|
| 187 |
+
"93": "face",
|
| 188 |
+
"94": "gluteus_maximus_left",
|
| 189 |
+
"95": "gluteus_maximus_right",
|
| 190 |
+
"96": "gluteus_medius_left",
|
| 191 |
+
"97": "gluteus_medius_right",
|
| 192 |
+
"98": "gluteus_minimus_left",
|
| 193 |
+
"99": "gluteus_minimus_right",
|
| 194 |
+
"100": "autochthon_left",
|
| 195 |
+
"101": "autochthon_right",
|
| 196 |
+
"102": "iliopsoas_left",
|
| 197 |
+
"103": "iliopsoas_right",
|
| 198 |
+
"104": "urinary_bladder"
|
| 199 |
+
}
|
| 200 |
+
}
|
| 201 |
+
}
|
| 202 |
+
}
|
| 203 |
+
}
|
wholeBody_ct_segmentation/configs/multi_gpu_evaluate.json
ADDED
|
@@ -0,0 +1,32 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"device": "$torch.device('cuda:' + os.environ['LOCAL_RANK'])",
|
| 3 |
+
"network": {
|
| 4 |
+
"_target_": "torch.nn.parallel.DistributedDataParallel",
|
| 5 |
+
"module": "$@network_def.to(@device)",
|
| 6 |
+
"device_ids": [
|
| 7 |
+
"@device"
|
| 8 |
+
]
|
| 9 |
+
},
|
| 10 |
+
"validate#sampler": {
|
| 11 |
+
"_target_": "DistributedSampler",
|
| 12 |
+
"dataset": "@validate#dataset",
|
| 13 |
+
"even_divisible": false,
|
| 14 |
+
"shuffle": false
|
| 15 |
+
},
|
| 16 |
+
"validate#dataloader#sampler": "@validate#sampler",
|
| 17 |
+
"validate#handlers#1#_disabled_": "$dist.get_rank() > 0",
|
| 18 |
+
"initialize": [
|
| 19 |
+
"$import torch.distributed as dist",
|
| 20 |
+
"$dist.is_initialized() or dist.init_process_group(backend='nccl')",
|
| 21 |
+
"$torch.cuda.set_device(@device)",
|
| 22 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)",
|
| 23 |
+
"$import logging",
|
| 24 |
+
"$@validate#evaluator.logger.setLevel(logging.WARNING if dist.get_rank() > 0 else logging.INFO)"
|
| 25 |
+
],
|
| 26 |
+
"run": [
|
| 27 |
+
"$@validate#evaluator.run()"
|
| 28 |
+
],
|
| 29 |
+
"finalize": [
|
| 30 |
+
"$dist.is_initialized() and dist.destroy_process_group()"
|
| 31 |
+
]
|
| 32 |
+
}
|
wholeBody_ct_segmentation/configs/multi_gpu_train.json
ADDED
|
@@ -0,0 +1,43 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"device": "$torch.device('cuda:' + os.environ['LOCAL_RANK'])",
|
| 3 |
+
"network": {
|
| 4 |
+
"_target_": "torch.nn.parallel.DistributedDataParallel",
|
| 5 |
+
"module": "$@network_def.to(@device)",
|
| 6 |
+
"device_ids": [
|
| 7 |
+
"@device"
|
| 8 |
+
]
|
| 9 |
+
},
|
| 10 |
+
"train#sampler": {
|
| 11 |
+
"_target_": "DistributedSampler",
|
| 12 |
+
"dataset": "@train#dataset",
|
| 13 |
+
"even_divisible": true,
|
| 14 |
+
"shuffle": true
|
| 15 |
+
},
|
| 16 |
+
"train#dataloader#sampler": "@train#sampler",
|
| 17 |
+
"train#dataloader#shuffle": false,
|
| 18 |
+
"train#trainer#train_handlers": "$@train#handlers[: -2 if dist.get_rank() > 0 else None]",
|
| 19 |
+
"validate#sampler": {
|
| 20 |
+
"_target_": "DistributedSampler",
|
| 21 |
+
"dataset": "@validate#dataset",
|
| 22 |
+
"even_divisible": false,
|
| 23 |
+
"shuffle": false
|
| 24 |
+
},
|
| 25 |
+
"validate#dataloader#sampler": "@validate#sampler",
|
| 26 |
+
"validate#evaluator#val_handlers": "$None if dist.get_rank() > 0 else @validate#handlers",
|
| 27 |
+
"initialize": [
|
| 28 |
+
"$import torch.distributed as dist",
|
| 29 |
+
"$dist.is_initialized() or dist.init_process_group(backend='nccl')",
|
| 30 |
+
"$torch.cuda.set_device(@device)",
|
| 31 |
+
"$monai.utils.set_determinism(seed=123)",
|
| 32 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)",
|
| 33 |
+
"$import logging",
|
| 34 |
+
"$@train#trainer.logger.setLevel(logging.WARNING if dist.get_rank() > 0 else logging.INFO)",
|
| 35 |
+
"$@validate#evaluator.logger.setLevel(logging.WARNING if dist.get_rank() > 0 else logging.INFO)"
|
| 36 |
+
],
|
| 37 |
+
"run": [
|
| 38 |
+
"$@train#trainer.run()"
|
| 39 |
+
],
|
| 40 |
+
"finalize": [
|
| 41 |
+
"$dist.is_initialized() and dist.destroy_process_group()"
|
| 42 |
+
]
|
| 43 |
+
}
|
wholeBody_ct_segmentation/configs/train.json
ADDED
|
@@ -0,0 +1,424 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"displayable_configs": {
|
| 3 |
+
"highres": true,
|
| 4 |
+
"init_LR": 0.0001
|
| 5 |
+
},
|
| 6 |
+
"imports": [
|
| 7 |
+
"$import glob",
|
| 8 |
+
"$import os",
|
| 9 |
+
"$import ignite"
|
| 10 |
+
],
|
| 11 |
+
"bundle_root": ".",
|
| 12 |
+
"ckpt_dir": "$@bundle_root + '/models'",
|
| 13 |
+
"output_dir": "$@bundle_root + '/eval'",
|
| 14 |
+
"dataset_dir": "sampledata",
|
| 15 |
+
"images": "$list(sorted(glob.glob(@dataset_dir + '/imagesTr/*.nii.gz')))",
|
| 16 |
+
"labels": "$list(sorted(glob.glob(@dataset_dir + '/labelsTr/*.nii.gz')))",
|
| 17 |
+
"highres": true,
|
| 18 |
+
"val_interval": 20,
|
| 19 |
+
"init_LR": 0.0001,
|
| 20 |
+
"batch_size": 4,
|
| 21 |
+
"pixdim": "$[1.5, 1.5, 1.5] if @displayable_configs#highres else [3.0, 3.0, 3.0]",
|
| 22 |
+
"modelname": "$'model.pt' if @displayable_configs#highres else 'model_lowres.pt'",
|
| 23 |
+
"device": "$torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')",
|
| 24 |
+
"network_def": {
|
| 25 |
+
"_target_": "SegResNet",
|
| 26 |
+
"spatial_dims": 3,
|
| 27 |
+
"in_channels": 1,
|
| 28 |
+
"out_channels": 105,
|
| 29 |
+
"init_filters": 32,
|
| 30 |
+
"blocks_down": [
|
| 31 |
+
1,
|
| 32 |
+
2,
|
| 33 |
+
2,
|
| 34 |
+
4
|
| 35 |
+
],
|
| 36 |
+
"blocks_up": [
|
| 37 |
+
1,
|
| 38 |
+
1,
|
| 39 |
+
1
|
| 40 |
+
],
|
| 41 |
+
"dropout_prob": 0.2
|
| 42 |
+
},
|
| 43 |
+
"network": "$@network_def.to(@device)",
|
| 44 |
+
"loss": {
|
| 45 |
+
"_target_": "DiceCELoss",
|
| 46 |
+
"to_onehot_y": true,
|
| 47 |
+
"softmax": true
|
| 48 |
+
},
|
| 49 |
+
"optimizer": {
|
| 50 |
+
"_target_": "torch.optim.AdamW",
|
| 51 |
+
"params": "$@network.parameters()",
|
| 52 |
+
"lr": "@displayable_configs#init_LR",
|
| 53 |
+
"weight_decay": 1e-05
|
| 54 |
+
},
|
| 55 |
+
"train": {
|
| 56 |
+
"deterministic_transforms": [
|
| 57 |
+
{
|
| 58 |
+
"_target_": "LoadImaged",
|
| 59 |
+
"keys": [
|
| 60 |
+
"image",
|
| 61 |
+
"label"
|
| 62 |
+
]
|
| 63 |
+
},
|
| 64 |
+
{
|
| 65 |
+
"_target_": "EnsureChannelFirstd",
|
| 66 |
+
"keys": [
|
| 67 |
+
"image",
|
| 68 |
+
"label"
|
| 69 |
+
]
|
| 70 |
+
},
|
| 71 |
+
{
|
| 72 |
+
"_target_": "EnsureTyped",
|
| 73 |
+
"keys": [
|
| 74 |
+
"image",
|
| 75 |
+
"label"
|
| 76 |
+
]
|
| 77 |
+
},
|
| 78 |
+
{
|
| 79 |
+
"_target_": "Orientationd",
|
| 80 |
+
"keys": [
|
| 81 |
+
"image",
|
| 82 |
+
"label"
|
| 83 |
+
],
|
| 84 |
+
"axcodes": "RAS"
|
| 85 |
+
},
|
| 86 |
+
{
|
| 87 |
+
"_target_": "Spacingd",
|
| 88 |
+
"keys": [
|
| 89 |
+
"image",
|
| 90 |
+
"label"
|
| 91 |
+
],
|
| 92 |
+
"pixdim": "@pixdim",
|
| 93 |
+
"mode": [
|
| 94 |
+
"bilinear",
|
| 95 |
+
"nearest"
|
| 96 |
+
]
|
| 97 |
+
},
|
| 98 |
+
{
|
| 99 |
+
"_target_": "NormalizeIntensityd",
|
| 100 |
+
"keys": "image",
|
| 101 |
+
"nonzero": true
|
| 102 |
+
},
|
| 103 |
+
{
|
| 104 |
+
"_target_": "CropForegroundd",
|
| 105 |
+
"keys": [
|
| 106 |
+
"image",
|
| 107 |
+
"label"
|
| 108 |
+
],
|
| 109 |
+
"source_key": "image",
|
| 110 |
+
"margin": 10,
|
| 111 |
+
"k_divisible": [
|
| 112 |
+
96,
|
| 113 |
+
96,
|
| 114 |
+
96
|
| 115 |
+
]
|
| 116 |
+
},
|
| 117 |
+
{
|
| 118 |
+
"_target_": "GaussianSmoothd",
|
| 119 |
+
"keys": [
|
| 120 |
+
"image"
|
| 121 |
+
],
|
| 122 |
+
"sigma": 0.4
|
| 123 |
+
},
|
| 124 |
+
{
|
| 125 |
+
"_target_": "ScaleIntensityd",
|
| 126 |
+
"keys": "image",
|
| 127 |
+
"minv": -1.0,
|
| 128 |
+
"maxv": 1.0
|
| 129 |
+
},
|
| 130 |
+
{
|
| 131 |
+
"_target_": "EnsureTyped",
|
| 132 |
+
"keys": [
|
| 133 |
+
"image",
|
| 134 |
+
"label"
|
| 135 |
+
]
|
| 136 |
+
}
|
| 137 |
+
],
|
| 138 |
+
"random_transforms": [
|
| 139 |
+
{
|
| 140 |
+
"_target_": "RandSpatialCropd",
|
| 141 |
+
"keys": [
|
| 142 |
+
"image",
|
| 143 |
+
"label"
|
| 144 |
+
],
|
| 145 |
+
"roi_size": [
|
| 146 |
+
96,
|
| 147 |
+
96,
|
| 148 |
+
96
|
| 149 |
+
],
|
| 150 |
+
"random_size": false
|
| 151 |
+
}
|
| 152 |
+
],
|
| 153 |
+
"preprocessing": {
|
| 154 |
+
"_target_": "Compose",
|
| 155 |
+
"transforms": "$@train#deterministic_transforms + @train#random_transforms"
|
| 156 |
+
},
|
| 157 |
+
"dataset": {
|
| 158 |
+
"_target_": "CacheDataset",
|
| 159 |
+
"data": "$[{'image': i, 'label': l} for i, l in zip(@images[:-10], @labels[:-10])]",
|
| 160 |
+
"transform": "@train#preprocessing",
|
| 161 |
+
"cache_rate": 0.4,
|
| 162 |
+
"num_workers": 4
|
| 163 |
+
},
|
| 164 |
+
"dataloader": {
|
| 165 |
+
"_target_": "DataLoader",
|
| 166 |
+
"dataset": "@train#dataset",
|
| 167 |
+
"batch_size": "@batch_size",
|
| 168 |
+
"shuffle": true,
|
| 169 |
+
"num_workers": 4
|
| 170 |
+
},
|
| 171 |
+
"inferer": {
|
| 172 |
+
"_target_": "SimpleInferer"
|
| 173 |
+
},
|
| 174 |
+
"postprocessing": {
|
| 175 |
+
"_target_": "Compose",
|
| 176 |
+
"transforms": [
|
| 177 |
+
{
|
| 178 |
+
"_target_": "Activationsd",
|
| 179 |
+
"keys": "pred",
|
| 180 |
+
"softmax": true
|
| 181 |
+
},
|
| 182 |
+
{
|
| 183 |
+
"_target_": "AsDiscreted",
|
| 184 |
+
"keys": [
|
| 185 |
+
"pred",
|
| 186 |
+
"label"
|
| 187 |
+
],
|
| 188 |
+
"argmax": [
|
| 189 |
+
true,
|
| 190 |
+
false
|
| 191 |
+
],
|
| 192 |
+
"to_onehot": 105
|
| 193 |
+
}
|
| 194 |
+
]
|
| 195 |
+
},
|
| 196 |
+
"handlers": [
|
| 197 |
+
{
|
| 198 |
+
"_target_": "ValidationHandler",
|
| 199 |
+
"validator": "@validate#evaluator",
|
| 200 |
+
"epoch_level": true,
|
| 201 |
+
"interval": "@val_interval"
|
| 202 |
+
},
|
| 203 |
+
{
|
| 204 |
+
"_target_": "StatsHandler",
|
| 205 |
+
"tag_name": "train_loss",
|
| 206 |
+
"output_transform": "$monai.handlers.from_engine(['loss'], first=True)"
|
| 207 |
+
},
|
| 208 |
+
{
|
| 209 |
+
"_target_": "TensorBoardStatsHandler",
|
| 210 |
+
"log_dir": "@output_dir",
|
| 211 |
+
"tag_name": "train_loss",
|
| 212 |
+
"output_transform": "$monai.handlers.from_engine(['loss'], first=True)"
|
| 213 |
+
}
|
| 214 |
+
],
|
| 215 |
+
"key_metric": {
|
| 216 |
+
"train_accuracy": {
|
| 217 |
+
"_target_": "ignite.metrics.Accuracy",
|
| 218 |
+
"output_transform": "$monai.handlers.from_engine(['pred', 'label'])"
|
| 219 |
+
}
|
| 220 |
+
},
|
| 221 |
+
"trainer": {
|
| 222 |
+
"_target_": "SupervisedTrainer",
|
| 223 |
+
"max_epochs": 4000,
|
| 224 |
+
"device": "@device",
|
| 225 |
+
"train_data_loader": "@train#dataloader",
|
| 226 |
+
"network": "@network",
|
| 227 |
+
"loss_function": "@loss",
|
| 228 |
+
"optimizer": "@optimizer",
|
| 229 |
+
"inferer": "@train#inferer",
|
| 230 |
+
"postprocessing": "@train#postprocessing",
|
| 231 |
+
"key_train_metric": "@train#key_metric",
|
| 232 |
+
"train_handlers": "@train#handlers",
|
| 233 |
+
"amp": true
|
| 234 |
+
}
|
| 235 |
+
},
|
| 236 |
+
"validate": {
|
| 237 |
+
"preprocessing": {
|
| 238 |
+
"_target_": "Compose",
|
| 239 |
+
"transforms": [
|
| 240 |
+
{
|
| 241 |
+
"_target_": "LoadImaged",
|
| 242 |
+
"keys": [
|
| 243 |
+
"image",
|
| 244 |
+
"label"
|
| 245 |
+
]
|
| 246 |
+
},
|
| 247 |
+
{
|
| 248 |
+
"_target_": "EnsureChannelFirstd",
|
| 249 |
+
"keys": [
|
| 250 |
+
"image",
|
| 251 |
+
"label"
|
| 252 |
+
]
|
| 253 |
+
},
|
| 254 |
+
{
|
| 255 |
+
"_target_": "EnsureTyped",
|
| 256 |
+
"keys": [
|
| 257 |
+
"image",
|
| 258 |
+
"label"
|
| 259 |
+
]
|
| 260 |
+
},
|
| 261 |
+
{
|
| 262 |
+
"_target_": "Orientationd",
|
| 263 |
+
"keys": [
|
| 264 |
+
"image",
|
| 265 |
+
"label"
|
| 266 |
+
],
|
| 267 |
+
"axcodes": "RAS"
|
| 268 |
+
},
|
| 269 |
+
{
|
| 270 |
+
"_target_": "Spacingd",
|
| 271 |
+
"keys": [
|
| 272 |
+
"image",
|
| 273 |
+
"label"
|
| 274 |
+
],
|
| 275 |
+
"pixdim": "@pixdim",
|
| 276 |
+
"mode": [
|
| 277 |
+
"bilinear",
|
| 278 |
+
"nearest"
|
| 279 |
+
]
|
| 280 |
+
},
|
| 281 |
+
{
|
| 282 |
+
"_target_": "NormalizeIntensityd",
|
| 283 |
+
"keys": "image",
|
| 284 |
+
"nonzero": true
|
| 285 |
+
},
|
| 286 |
+
{
|
| 287 |
+
"_target_": "CropForegroundd",
|
| 288 |
+
"keys": [
|
| 289 |
+
"image",
|
| 290 |
+
"label"
|
| 291 |
+
],
|
| 292 |
+
"source_key": "image",
|
| 293 |
+
"margin": 10,
|
| 294 |
+
"k_divisible": [
|
| 295 |
+
96,
|
| 296 |
+
96,
|
| 297 |
+
96
|
| 298 |
+
]
|
| 299 |
+
},
|
| 300 |
+
{
|
| 301 |
+
"_target_": "GaussianSmoothd",
|
| 302 |
+
"keys": [
|
| 303 |
+
"image"
|
| 304 |
+
],
|
| 305 |
+
"sigma": 0.4
|
| 306 |
+
},
|
| 307 |
+
{
|
| 308 |
+
"_target_": "ScaleIntensityd",
|
| 309 |
+
"keys": "image",
|
| 310 |
+
"minv": -1.0,
|
| 311 |
+
"maxv": 1.0
|
| 312 |
+
},
|
| 313 |
+
{
|
| 314 |
+
"_target_": "CenterSpatialCropd",
|
| 315 |
+
"keys": [
|
| 316 |
+
"image",
|
| 317 |
+
"label"
|
| 318 |
+
],
|
| 319 |
+
"roi_size": [
|
| 320 |
+
160,
|
| 321 |
+
160,
|
| 322 |
+
160
|
| 323 |
+
]
|
| 324 |
+
}
|
| 325 |
+
]
|
| 326 |
+
},
|
| 327 |
+
"postprocessing": {
|
| 328 |
+
"_target_": "Compose",
|
| 329 |
+
"transforms": [
|
| 330 |
+
{
|
| 331 |
+
"_target_": "Activationsd",
|
| 332 |
+
"keys": "pred",
|
| 333 |
+
"softmax": true
|
| 334 |
+
},
|
| 335 |
+
{
|
| 336 |
+
"_target_": "AsDiscreted",
|
| 337 |
+
"keys": [
|
| 338 |
+
"pred",
|
| 339 |
+
"label"
|
| 340 |
+
],
|
| 341 |
+
"argmax": [
|
| 342 |
+
true,
|
| 343 |
+
false
|
| 344 |
+
],
|
| 345 |
+
"to_onehot": 105
|
| 346 |
+
}
|
| 347 |
+
]
|
| 348 |
+
},
|
| 349 |
+
"dataset": {
|
| 350 |
+
"_target_": "Dataset",
|
| 351 |
+
"data": "$[{'image': i, 'label': l} for i, l in zip(@images[-10:], @labels[-10:])]",
|
| 352 |
+
"transform": "@validate#preprocessing"
|
| 353 |
+
},
|
| 354 |
+
"dataloader": {
|
| 355 |
+
"_target_": "DataLoader",
|
| 356 |
+
"dataset": "@validate#dataset",
|
| 357 |
+
"batch_size": 1,
|
| 358 |
+
"shuffle": false,
|
| 359 |
+
"num_workers": 4
|
| 360 |
+
},
|
| 361 |
+
"inferer": {
|
| 362 |
+
"_target_": "SlidingWindowInferer",
|
| 363 |
+
"roi_size": [
|
| 364 |
+
96,
|
| 365 |
+
96,
|
| 366 |
+
96
|
| 367 |
+
],
|
| 368 |
+
"sw_batch_size": 1,
|
| 369 |
+
"overlap": 0.25
|
| 370 |
+
},
|
| 371 |
+
"handlers": [
|
| 372 |
+
{
|
| 373 |
+
"_target_": "StatsHandler",
|
| 374 |
+
"iteration_log": false
|
| 375 |
+
},
|
| 376 |
+
{
|
| 377 |
+
"_target_": "TensorBoardStatsHandler",
|
| 378 |
+
"log_dir": "@output_dir",
|
| 379 |
+
"iteration_log": false
|
| 380 |
+
},
|
| 381 |
+
{
|
| 382 |
+
"_target_": "CheckpointSaver",
|
| 383 |
+
"save_dir": "@ckpt_dir",
|
| 384 |
+
"save_dict": {
|
| 385 |
+
"model": "@network"
|
| 386 |
+
},
|
| 387 |
+
"save_key_metric": true,
|
| 388 |
+
"key_metric_filename": "@modelname"
|
| 389 |
+
}
|
| 390 |
+
],
|
| 391 |
+
"key_metric": {
|
| 392 |
+
"val_mean_dice": {
|
| 393 |
+
"_target_": "MeanDice",
|
| 394 |
+
"include_background": false,
|
| 395 |
+
"output_transform": "$monai.handlers.from_engine(['pred', 'label'])"
|
| 396 |
+
}
|
| 397 |
+
},
|
| 398 |
+
"additional_metrics": {
|
| 399 |
+
"val_accuracy": {
|
| 400 |
+
"_target_": "ignite.metrics.Accuracy",
|
| 401 |
+
"output_transform": "$monai.handlers.from_engine(['pred', 'label'])"
|
| 402 |
+
}
|
| 403 |
+
},
|
| 404 |
+
"evaluator": {
|
| 405 |
+
"_target_": "SupervisedEvaluator",
|
| 406 |
+
"device": "@device",
|
| 407 |
+
"val_data_loader": "@validate#dataloader",
|
| 408 |
+
"network": "@network",
|
| 409 |
+
"inferer": "@validate#inferer",
|
| 410 |
+
"postprocessing": "@validate#postprocessing",
|
| 411 |
+
"key_val_metric": "@validate#key_metric",
|
| 412 |
+
"additional_metrics": "@validate#additional_metrics",
|
| 413 |
+
"val_handlers": "@validate#handlers",
|
| 414 |
+
"amp": true
|
| 415 |
+
}
|
| 416 |
+
},
|
| 417 |
+
"initialize": [
|
| 418 |
+
"$monai.utils.set_determinism(seed=123)",
|
| 419 |
+
"$setattr(torch.backends.cudnn, 'benchmark', True)"
|
| 420 |
+
],
|
| 421 |
+
"run": [
|
| 422 |
+
"$@train#trainer.run()"
|
| 423 |
+
]
|
| 424 |
+
}
|
wholeBody_ct_segmentation/docs/README.md
ADDED
|
@@ -0,0 +1,264 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Model Overview
|
| 2 |
+
Body CT segmentation models are evolving. Starting from abdominal multi-organ segmentation model [1]. Now the community is developing hundreds of target anatomies. In this bundle, we provide re-trained models for (3D) segmentation of 104 whole-body segments.
|
| 3 |
+
|
| 4 |
+
This model is trained using the SegResNet [3] network. The model is trained using TotalSegmentator datasets [2].
|
| 5 |
+
|
| 6 |
+

|
| 7 |
+
|
| 8 |
+
Figure source from the TotalSegmentator [2].
|
| 9 |
+
|
| 10 |
+
### MONAI Label Showcase
|
| 11 |
+
|
| 12 |
+
- We highlight the use of this bundle to use and visualize in MONAI Label + 3D Slicer integration.
|
| 13 |
+
|
| 14 |
+
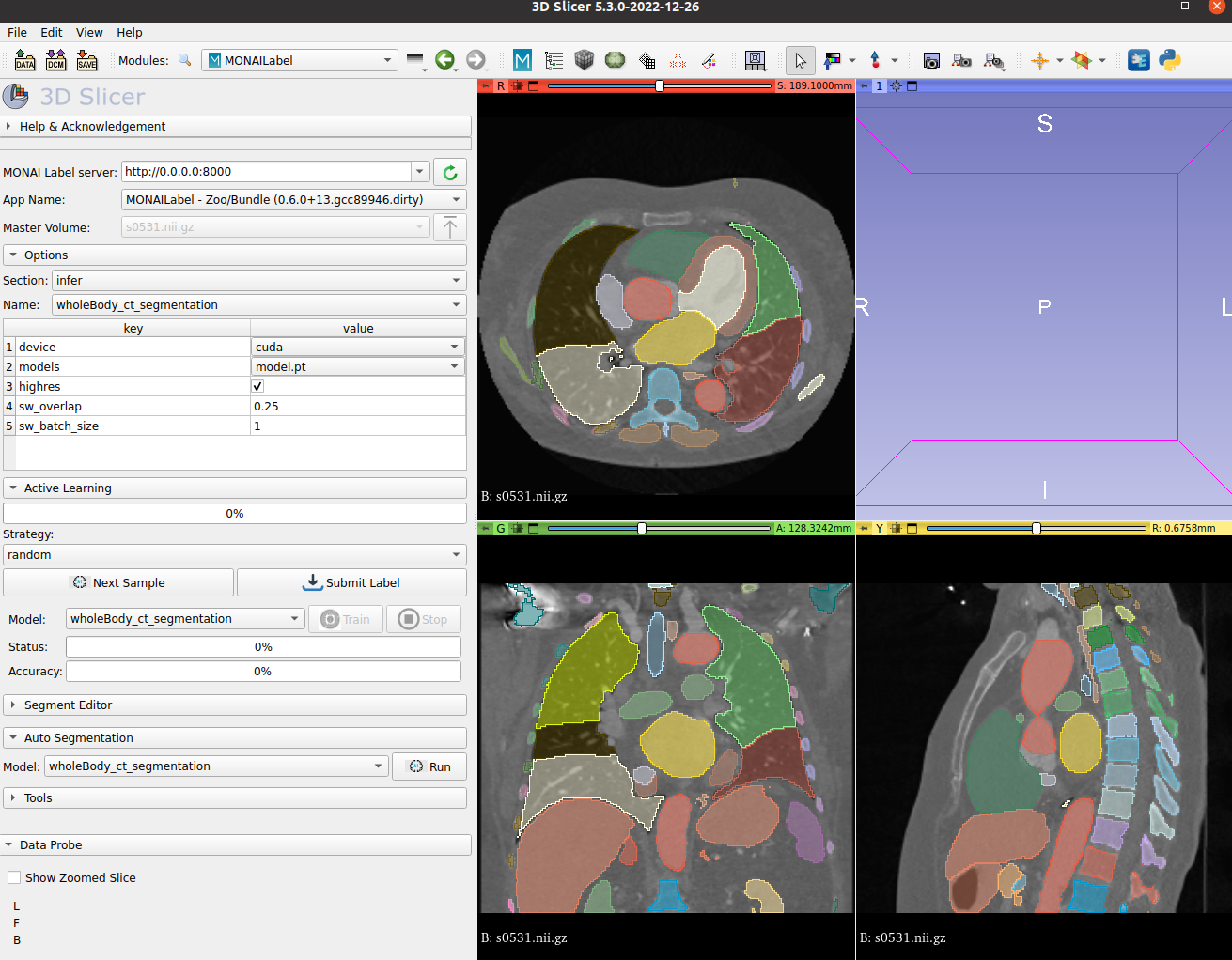 <br>
|
| 15 |
+
|
| 16 |
+
## Data
|
| 17 |
+
|
| 18 |
+
The training set is the 104 whole-body structures from the TotalSegmentator released datasets. Users can find more details on the datasets at https://github.com/wasserth/TotalSegmentator. All rights and licenses are reserved to the original authors.
|
| 19 |
+
|
| 20 |
+
- Target: 104 structures
|
| 21 |
+
- Modality: CT
|
| 22 |
+
- Source: TotalSegmentator
|
| 23 |
+
- Challenge: Large volumes of structures in CT images
|
| 24 |
+
|
| 25 |
+
### Preprocessing
|
| 26 |
+
|
| 27 |
+
To use the bundle, users need to download the data and merge all annotated labels into one NIFTI file. Each file contains 0-104 values, each value represents one anatomy class. We provide sample datasets and step-by-step instructions on how to get prepared:
|
| 28 |
+
|
| 29 |
+
Instruction on how to start with the prepared sample dataset:
|
| 30 |
+
|
| 31 |
+
1. Download the sample set with this [link](https://drive.google.com/file/d/1DtDmERVMjks1HooUhggOKAuDm0YIEunG/view?usp=share_link).
|
| 32 |
+
2. Unzip the dataset into a workspace folder.
|
| 33 |
+
3. There will be three sub-folders, each with several preprocessed CT volumes:
|
| 34 |
+
- imagesTr: 20 samples of training scans and validation scans.
|
| 35 |
+
- labelsTr: 20 samples of pre-processed label files.
|
| 36 |
+
- imagesTs: 5 samples of sample testing scans.
|
| 37 |
+
4. Usage: users can add `--dataset_dir <totalSegmentator_mergedLabel_samples>` to the bundle run command to specify the data path.
|
| 38 |
+
|
| 39 |
+
Instruction on how to merge labels with the raw dataset:
|
| 40 |
+
|
| 41 |
+
- There are 104 binary masks associated with each CT scan, each mask corresponds to anatomy. These pixel-level labels are class-exclusive, users can assign each anatomy a class number then merge to a single NIFTI file as the ground truth label file. The order of anatomies can be found [here](https://github.com/Project-MONAI/model-zoo/blob/dev/models/wholeBody_ct_segmentation/configs/metadata.json).
|
| 42 |
+
|
| 43 |
+
## Training Configuration
|
| 44 |
+
|
| 45 |
+
The segmentation of 104 tissues is formulated as voxel-wise multi-label segmentation. The model is optimized with the gradient descent method minimizing Dice + cross-entropy loss between the predicted mask and ground truth segmentation.
|
| 46 |
+
|
| 47 |
+
The training was performed with the following:
|
| 48 |
+
|
| 49 |
+
- GPU: 48 GB of GPU memory
|
| 50 |
+
- Actual Model Input: 96 x 96 x 96
|
| 51 |
+
- AMP: True
|
| 52 |
+
- Optimizer: AdamW
|
| 53 |
+
- Learning Rate: 1e-4
|
| 54 |
+
- Loss: DiceCELoss
|
| 55 |
+
|
| 56 |
+
## Evaluation Configuration
|
| 57 |
+
|
| 58 |
+
The model predicts 105 channels output at the same time using softmax and argmax. It requires higher GPU memory when calculating
|
| 59 |
+
metrics between predicted masked and ground truth. The consumption of hardware requirements, such as GPU memory is dependent on the input CT volume size.
|
| 60 |
+
|
| 61 |
+
The recommended evaluation configuration and the metrics were acquired with the following hardware:
|
| 62 |
+
|
| 63 |
+
- GPU: equal to or larger than 48 GB of GPU memory
|
| 64 |
+
- Model: high resolution model pre-trained at a slice thickness of 1.5 mm.
|
| 65 |
+
|
| 66 |
+
Note: there are two pre-trained models provided. The default is the high resolution model, evaluation pipeline at slice thickness of **1.5mm**,
|
| 67 |
+
users can use the lower resolution model if out of memory (OOM) occurs, which the model is pre-trained with CT scans at a slice thickness of **3.0mm**.
|
| 68 |
+
|
| 69 |
+
Users can also use the inference pipeline for predicted masks, we provide detailed GPU memory consumption in the following sections.
|
| 70 |
+
|
| 71 |
+
### Memory Consumption
|
| 72 |
+
|
| 73 |
+
- Dataset Manager: CacheDataset
|
| 74 |
+
- Data Size: 1000 3D Volumes
|
| 75 |
+
- Cache Rate: 0.4
|
| 76 |
+
- Single GPU - System RAM Usage: 83G
|
| 77 |
+
- Multi GPU (8 GPUs) - System RAM Usage: 666G
|
| 78 |
+
|
| 79 |
+
### Memory Consumption Warning
|
| 80 |
+
|
| 81 |
+
If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate `cache_rate` in the configurations within range [0, 1] to minimize the System RAM requirements.
|
| 82 |
+
|
| 83 |
+
### Input
|
| 84 |
+
|
| 85 |
+
One channel
|
| 86 |
+
- CT image
|
| 87 |
+
|
| 88 |
+
### Output
|
| 89 |
+
|
| 90 |
+
105 channels
|
| 91 |
+
- Label 0: Background (everything else)
|
| 92 |
+
- label 1-105: Foreground classes (104)
|
| 93 |
+
|
| 94 |
+
## Resource Requirements and Latency Benchmarks
|
| 95 |
+
|
| 96 |
+
### GPU Consumption Warning
|
| 97 |
+
|
| 98 |
+
The model is trained with 104 classes in single instance, for predicting 104 structures, the GPU consumption can be large.
|
| 99 |
+
|
| 100 |
+
For inference pipeline, please refer to the following section for benchmarking results. Normally, a CT scans with 300 slices will take about 27G memory, if your CT is larger, please prepare larger GPU memory or use CPU for inference.
|
| 101 |
+
|
| 102 |
+
### High-Resolution and Low-Resolution Models
|
| 103 |
+
|
| 104 |
+
We retrained two versions of the totalSegmentator models, following the original paper and implementation.
|
| 105 |
+
To meet multiple demands according to computation resources and performance, we provide a 1.5 mm model and a 3.0 mm model, both models are trained with 104 foreground output channels.
|
| 106 |
+
|
| 107 |
+
In this bundle, we configured a parameter called `highres`, users can set it to `true` when using 1.5 mm model, and set it to `false` to use the 3.0 mm model. The high-resolution model is named `model.pt` by default, the low-resolution model is named `model_lowres.pt`.
|
| 108 |
+
|
| 109 |
+
In MONAI Label use case, users can set the parameter in 3D Slicer plugin to control which model to infer and train.
|
| 110 |
+
|
| 111 |
+
- Pretrained Checkpoints
|
| 112 |
+
- 1.5 mm model: [Download link](https://drive.google.com/file/d/1PHpFWboimEXmMSe2vBra6T8SaCMC2SHT/view?usp=share_link)
|
| 113 |
+
- 3.0 mm model: [Download link](https://drive.google.com/file/d/1c3osYscnr6710ObqZZS8GkZJQlWlc7rt/view?usp=share_link)
|
| 114 |
+
|
| 115 |
+
Latencies and memory performance of using the bundle with MONAI Label:
|
| 116 |
+
|
| 117 |
+
Tested Image Dimension: **(512, 512, 397)**, the slice thickness is **1.5mm** in this case. After resample to **1.5** isotropic resolution, the dimension is **(287, 287, 397)**
|
| 118 |
+
|
| 119 |
+
### 1.5 mm (highres) model (Single Model with 104 foreground classes)
|
| 120 |
+
|
| 121 |
+
Benchmarking on GPU: Memory: **28.73G**
|
| 122 |
+
|
| 123 |
+
- `++ Latencies => Total: 6.0277; Pre: 1.6228; Inferer: 4.1153; Invert: 0.0000; Post: 0.0897; Write: 0.1995`
|
| 124 |
+
|
| 125 |
+
Benchmarking on CPU: Memory: **26G**
|
| 126 |
+
|
| 127 |
+
- `++ Latencies => Total: 38.3108; Pre: 1.6643; Inferer: 30.3018; Invert: 0.0000; Post: 6.1656; Write: 0.1786`
|
| 128 |
+
|
| 129 |
+
### 3.0 mm (lowres) model (single model with 104 foreground classes)
|
| 130 |
+
|
| 131 |
+
GPU: Memory: **5.89G**
|
| 132 |
+
|
| 133 |
+
- `++ Latencies => Total: 1.9993; Pre: 1.2363; Inferer: 0.5207; Invert: 0.0000; Post: 0.0358; Write: 0.2060`
|
| 134 |
+
|
| 135 |
+
CPU: Memory: **2.3G**
|
| 136 |
+
|
| 137 |
+
- `++ Latencies => Total: 6.6138; Pre: 1.3192; Inferer: 3.6746; Invert: 0.0000; Post: 1.4431; Write: 0.1760`
|
| 138 |
+
|
| 139 |
+
## Performance
|
| 140 |
+
|
| 141 |
+
### 1.5 mm Model Training
|
| 142 |
+
|
| 143 |
+
#### Training Accuracy
|
| 144 |
+
|
| 145 |
+
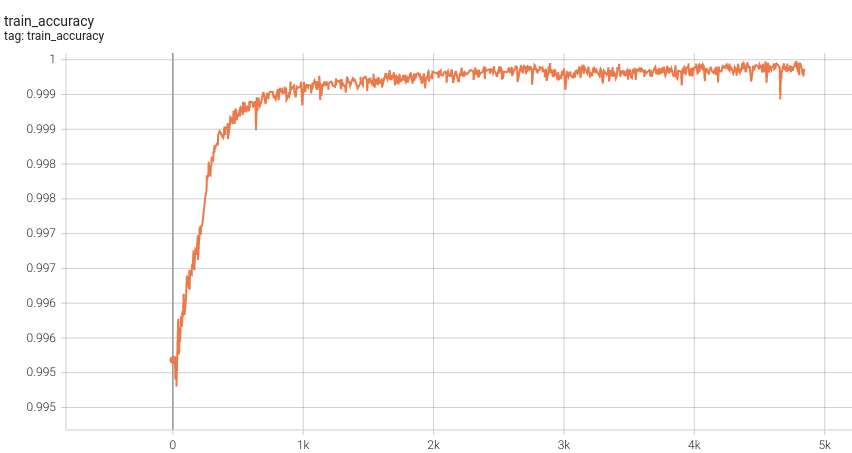 <br>
|
| 146 |
+
|
| 147 |
+
#### Validation Dice
|
| 148 |
+
|
| 149 |
+
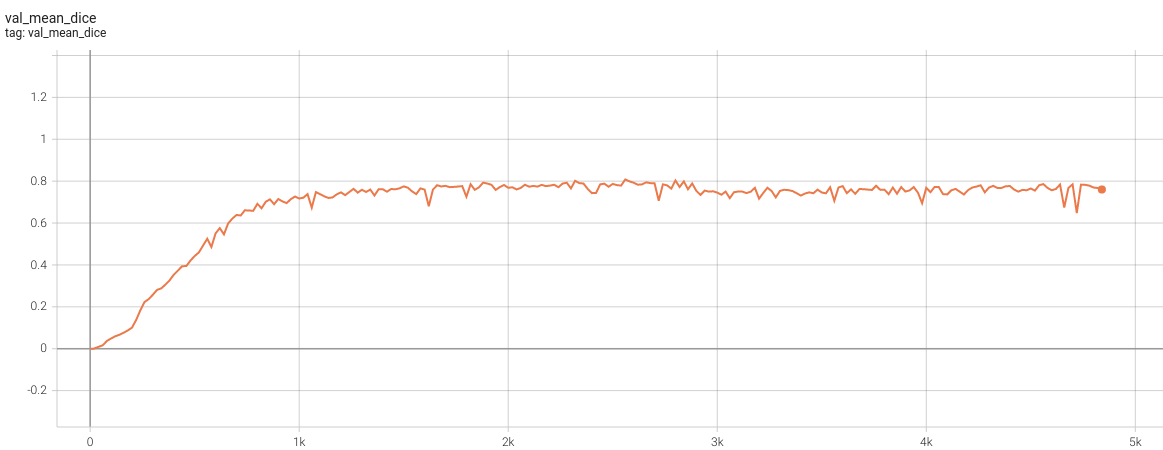 <br>
|
| 150 |
+
|
| 151 |
+
Please note that this bundle is non-deterministic because of the trilinear interpolation used in the network. Therefore, reproducing the training process may not get exactly the same performance.
|
| 152 |
+
Please refer to https://pytorch.org/docs/stable/notes/randomness.html#reproducibility for more details about reproducibility.
|
| 153 |
+
|
| 154 |
+
#### TensorRT speedup
|
| 155 |
+
This bundle supports acceleration with TensorRT. The table below displays the speedup ratios observed on an A100 80G GPU.
|
| 156 |
+
|
| 157 |
+
| method | torch_fp32(ms) | torch_amp(ms) | trt_fp32(ms) | trt_fp16(ms) | speedup amp | speedup fp32 | speedup fp16 | amp vs fp16|
|
| 158 |
+
| :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: |
|
| 159 |
+
| model computation | 88.20 | 37.1 | 39.2 | 36.9 | 2.38 | 2.25 | 2.39 | 1.01 |
|
| 160 |
+
| end2end | 3717.14 | 2596.77 | 2517.29 | 2501.37 | 1.43 | 1.48 | 1.49 | 1.04 |
|
| 161 |
+
|
| 162 |
+
Where:
|
| 163 |
+
- `model computation` means the speedup ratio of model's inference with a random input without preprocessing and postprocessing
|
| 164 |
+
- `end2end` means run the bundle end-to-end with the TensorRT based model.
|
| 165 |
+
- `torch_fp32` and `torch_amp` are for the PyTorch models with or without `amp` mode.
|
| 166 |
+
- `trt_fp32` and `trt_fp16` are for the TensorRT based models converted in corresponding precision.
|
| 167 |
+
- `speedup amp`, `speedup fp32` and `speedup fp16` are the speedup ratios of corresponding models versus the PyTorch float32 model
|
| 168 |
+
- `amp vs fp16` is the speedup ratio between the PyTorch amp model and the TensorRT float16 based model.
|
| 169 |
+
|
| 170 |
+
This result is benchmarked under:
|
| 171 |
+
- TensorRT: 8.6.1+cuda12.0
|
| 172 |
+
- Torch-TensorRT Version: 1.4.0
|
| 173 |
+
- CPU Architecture: x86-64
|
| 174 |
+
- OS: ubuntu 20.04
|
| 175 |
+
- Python version:3.8.10
|
| 176 |
+
- CUDA version: 12.1
|
| 177 |
+
- GPU models and configuration: A100 80G
|
| 178 |
+
|
| 179 |
+
## MONAI Bundle Commands
|
| 180 |
+
In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file.
|
| 181 |
+
|
| 182 |
+
For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html).
|
| 183 |
+
|
| 184 |
+
#### Execute training:
|
| 185 |
+
|
| 186 |
+
```
|
| 187 |
+
python -m monai.bundle run --config_file configs/train.json
|
| 188 |
+
```
|
| 189 |
+
|
| 190 |
+
Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`:
|
| 191 |
+
|
| 192 |
+
```
|
| 193 |
+
python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path>
|
| 194 |
+
```
|
| 195 |
+
|
| 196 |
+
#### Override the `train` config to execute multi-GPU training:
|
| 197 |
+
|
| 198 |
+
```
|
| 199 |
+
torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']"
|
| 200 |
+
```
|
| 201 |
+
|
| 202 |
+
Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html).
|
| 203 |
+
|
| 204 |
+
#### Override the `train` config to execute evaluation with the trained model:
|
| 205 |
+
|
| 206 |
+
```
|
| 207 |
+
python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']"
|
| 208 |
+
```
|
| 209 |
+
|
| 210 |
+
#### Override the `train` config and `evaluate` config to execute multi-GPU evaluation:
|
| 211 |
+
|
| 212 |
+
```
|
| 213 |
+
torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json','configs/multi_gpu_evaluate.json']"
|
| 214 |
+
```
|
| 215 |
+
|
| 216 |
+
#### Execute inference:
|
| 217 |
+
|
| 218 |
+
```
|
| 219 |
+
python -m monai.bundle run --config_file configs/inference.json
|
| 220 |
+
```
|
| 221 |
+
#### Execute inference with Data Samples:
|
| 222 |
+
|
| 223 |
+
```
|
| 224 |
+
python -m monai.bundle run --config_file configs/inference.json --datalist "['sampledata/imagesTr/s0037.nii.gz','sampledata/imagesTr/s0038.nii.gz']"
|
| 225 |
+
```
|
| 226 |
+
|
| 227 |
+
#### Export checkpoint to TensorRT based models with fp32 or fp16 precision:
|
| 228 |
+
|
| 229 |
+
```
|
| 230 |
+
python -m monai.bundle trt_export --net_id network_def --filepath models/model_trt.ts --ckpt_file models/model.pt --meta_file configs/metadata.json --config_file configs/inference.json --precision <fp32/fp16> --use_trace "True"
|
| 231 |
+
```
|
| 232 |
+
|
| 233 |
+
#### Execute inference with the TensorRT model:
|
| 234 |
+
|
| 235 |
+
```
|
| 236 |
+
python -m monai.bundle run --config_file "['configs/inference.json', 'configs/inference_trt.json']"
|
| 237 |
+
```
|
| 238 |
+
|
| 239 |
+
|
| 240 |
+
# References
|
| 241 |
+
|
| 242 |
+
[1] Tang, Y., Gao, R., Lee, H.H., Han, S., Chen, Y., Gao, D., Nath, V., Bermudez, C., Savona, M.R., Abramson, R.G. and Bao, S., 2021. High-resolution 3D abdominal segmentation with random patch network fusion. Medical image analysis, 69, p.101894.
|
| 243 |
+
|
| 244 |
+
[2] Wasserthal, J., Meyer, M., Breit, H.C., Cyriac, J., Yang, S. and Segeroth, M., 2022. TotalSegmentator: robust segmentation of 104 anatomical structures in CT images. arXiv preprint arXiv:2208.05868.
|
| 245 |
+
|
| 246 |
+
[3] Myronenko, A., Siddiquee, M.M.R., Yang, D., He, Y. and Xu, D., 2022. Automated head and neck tumor segmentation from 3D PET/CT. arXiv preprint arXiv:2209.10809.
|
| 247 |
+
|
| 248 |
+
|
| 249 |
+
|
| 250 |
+
# License
|
| 251 |
+
|
| 252 |
+
Copyright (c) MONAI Consortium
|
| 253 |
+
|
| 254 |
+
Licensed under the Apache License, Version 2.0 (the "License");
|
| 255 |
+
you may not use this file except in compliance with the License.
|
| 256 |
+
You may obtain a copy of the License at
|
| 257 |
+
|
| 258 |
+
http://www.apache.org/licenses/LICENSE-2.0
|
| 259 |
+
|
| 260 |
+
Unless required by applicable law or agreed to in writing, software
|
| 261 |
+
distributed under the License is distributed on an "AS IS" BASIS,
|
| 262 |
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
| 263 |
+
See the License for the specific language governing permissions and
|
| 264 |
+
limitations under the License.
|
wholeBody_ct_segmentation/docs/data_license.txt
ADDED
|
@@ -0,0 +1,6 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
Third Party Licenses
|
| 2 |
+
-----------------------------------------------------------------------
|
| 3 |
+
|
| 4 |
+
/*********************************************************************/
|
| 5 |
+
i. TotalSegmentator
|
| 6 |
+
https://zenodo.org/record/6802614#.Y9iTydLMJ6I
|
wholeBody_ct_segmentation/models/model.pt
ADDED
|
@@ -0,0 +1,3 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
version https://git-lfs.github.com/spec/v1
|
| 2 |
+
oid sha256:80b429fb4b080df11c9ed0b0bdaa8a615ff083921bb213a512cf285afbc4e3fe
|
| 3 |
+
size 75225922
|
wholeBody_ct_segmentation/models/model_lowres.pt
ADDED
|
@@ -0,0 +1,3 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
version https://git-lfs.github.com/spec/v1
|
| 2 |
+
oid sha256:c3ab55eb979785fdcb30690872c210bbeee73d79a170c32fdaa1eca117779f90
|
| 3 |
+
size 75225922
|