diff --git "a/3554.jsonl" "b/3554.jsonl"
new file mode 100644--- /dev/null
+++ "b/3554.jsonl"
@@ -0,0 +1,110 @@
+{"seq_id":"42938276185","text":"# ### Importing libraries\n#\n\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n# %matplotlib inline\n\ndf=pd.read_csv(r'C:\\Users\\Admin\\Downloads\\Advertising(1).csv')\n\n# ### Dataframe check for count,sum, null values\n\ndf.head()\n\ndf.describe()\n\ndf.info()\n\ndf.isnull().sum()\n\ndf.head()\n\ndf. rename(columns = {'Unnamed: 0':'Index'},inplace = True)\n\ndf.head()\n\ndf=df.set_index('Index')\n\ndf.head()\n\n# ### Visualisation of data\n\nTV=df['TV']\nradio=df['radio']\nNP=df['newspaper']\nsales=df['sales']\n\nsns.distplot(sales)\n\n# ###### Distribution plot shows sales is following normal distribution\n\nsns.distplot(TV)\n\nsns.distplot(radio)\n\n# ###### Distribution for both TV and Radio are following uniform distribution.\n\nsns.distplot(NP)\n\n# ###### Newspaper distribution is right-skewed data\n\n# +\nsns.pairplot(df,x_vars=['TV','radio','newspaper'],y_vars=['sales'],height = 5, aspect=1.0,kind = 'reg')\n\n#reg : to perform linear regression on the graph\n# -\n\n# The TV and sales show a very strong relationship through this linear regression model. whereas, somewhat, weaker relationship is assumed between radio and sales and newspaper and sales have the weakest relationship\n\n#MAX VALUE OF CORRELATION IS BETWEEN TV AND SALES\ndf.TV.corr(df.sales)\n\n# #### Plotting correlation matrix and heat map to validate the results\n\n# +\nprint(df.corr())\n \n# plotting correlation heatmap\ndataplot = sns.heatmap(df.corr(), cmap=\"YlGnBu\", annot=True)\n \n# displaying heatmap\nplt.rcParams['figure.figsize']=(4,4)\nplt.show()\n\n# -\n\n# #### The results proved above seems similar to the correlation matrix with strongest relation between TV and SALES.\n\n# ### BUILDING LINEAR REGRESSION MODEL\n\nX=df[['TV']]\nprint(X.shape)\nprint(type(X))\n\nprint(X.head())\n\nY=df['sales']\n\nprint(type(Y))\nprint(Y.shape)\n\n\nY.head()\n\nfrom sklearn.model_selection import train_test_split\n\nX_train,X_test,Y_train,Y_test = train_test_split(X,Y,test_size=0.3,random_state=1)\n# using 30 percent data for test\n\n# ##### Printing X_train,Y_train and X_test and Y_test\n\nprint(X_train.shape)\n\nprint(Y_train.shape)\n\nprint(X_test.shape)\n\nprint(Y_test.shape)\n\nfrom sklearn.linear_model import LinearRegression\nmodel=LinearRegression()\nmodel.fit(X_train,Y_train)\n\nmodel.get_params()\n\n# ##### Interpret model\n\n# +\nprint(model.intercept_)\nprint(model.coef_)\n\n#Model intercept tells us the constant term in linear regression equation\n#Model coefficient tells us the Coefficient of x term in linear regression equation\n# -\n\n# #### Predictions with the Model\n\ny_pred = model.predict(X_test)\n\n#To check first few values of pred\nprint(y_pred[:5])\n\n# ##### FINAL PART OF MODEL\n# ### MODEL EVALUATION METRICS\n\n# ##### 1. Mean square Error\n# ##### 2. Mean Absolute Error\n# ##### 3. Root Mean Squared Error\n\n# #### MEAN ABSOLUTE ERROR:\n\nk=abs(Y_test-y_pred)\n\nsumma=0\nfor i in k:\n summa=summa+i\nprint(summa)\n\nn=60\nMAE=(1/n)*summa\n\nprint(f\"MEAN ABSOLUTE ERROR IS :{MAE}\")\n\n# #### MEAN SQUARE ERROR :\n#\n\nSD=(y_pred-Y_test)**2 # squared difference\nsu=0\nfor j in SD:\n su=su+j\nprint(su)\n\nn=60\nMSE=(1/n)*su\n\nprint(f\"MEAN SQUARED ERROR IS :{MSE}\")\n\n# #### ROOT MEAN SQUARED ERROR:\n#\n\nRMSE=(MSE)**0.5\nprint(f\"ROOT MEAN SQUARED ERROR IS :{RMSE}\")\n\nfrom sklearn import metrics\nprint(metrics)\n\n# #### Check answers with help of Metrics\n\nprint(metrics.mean_absolute_error(Y_test,y_pred))\n\nprint(metrics.mean_squared_error(Y_test,y_pred))\n\nprint(np.sqrt(metrics.mean_squared_error(Y_test,y_pred)))\n\n# ##### ALL VALUES ARE VERIFIED\n\n\n","repo_name":"Himanshu2102/Machine-Learning-Projects","sub_path":"Predicting sales and Revenue.ipynb","file_name":"Predicting sales and Revenue.ipynb","file_ext":"py","file_size_in_byte":3480,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"41885297054","text":"# +\nimport scapy\nfrom scapy.all import *\n\nimport pandas as pd\n\npackets=rdpcap('../data/web.pcap')\n# -\n\nimport numpy as np\n\nprint(len(packets))\n\n# ### Features to extract\n#\n# 1. FW Packet length max\n# flow duration\n# flow packets/s\n# fwd IAt total\n# flow IAT std\n# Fwd packet length std\n# fwd packet length mean\n# total length of backward packets\n# flow bytes/s\n# total length of fwd packets\n# flow iat mean\n# Bot(Bwd Packet Length Mean)\n# DDos(Bwd Packet Length Std)\n# DDos golden eye (Flow IAT Max)\n# Total Fwd Packets\n# Total Backward Packets'\n\n# +\nlist_of_all_dictionaries=[]\n\nfor i in range(len(packets)):\n\n dictionary_of_data={}\n pkt=packets[i]\n\n # Features from IP\n try :\n dictionary_of_data['sender_ip']=pkt['IP'].src\n dictionary_of_data['destination_ip']=pkt['IP'].dst\n dictionary_of_data['protocol_ip']=pkt['IP'].proto\n dictionary_of_data['length']=pkt['IP'].len\n except:\n continue# Features from TCP\n\n try:\n dictionary_of_data['source_port']=pkt['TCP'].sport\n dictionary_of_data['destination_port']=pkt['TCP'].dport\n dictionary_of_data['data_of_s']=pkt['TCP'].dataofs\n dictionary_of_data['seq']=pkt['TCP'].seq\n dictionary_of_data['ack']=pkt['TCP'].ack\n dictionary_of_data['flags']=pkt['TCP'].flags\n dictionary_of_data['window']=pkt['TCP'].window\n dictionary_of_data['chksum']=pkt['TCP'].chksum\n dictionary_of_data['urgptr']=pkt['TCP'].urgptr\n dictionary_of_data['start_time']=pkt['TCP'].options[2][1][1]\n dictionary_of_data['end_time']=pkt['TCP'].options[2][1][0]\n except:\n continue\n# elif pkt['IP'].proto==17:\n# try:\n# dictionary_of_data['source_port']=pkt['UDP'].sport\n# dictionary_of_data['destination_port']=pkt['UDP'].dport\n# dictionary_of_data['chksum']=pkt['UDP'].chksum\n# except:\n# continue\n\n list_of_all_dictionaries.append(dictionary_of_data)# Only store data if TCP data is available\n\n\n \n\n\n\n\nprint(len(list_of_all_dictionaries))\n\n# -\n\npackets[30]\n\ndf=pd.DataFrame(list_of_all_dictionaries)\n\ndf.to_csv('../data/web.csv')\n\ndf.head()\n\n# #### Fwd Packet Length Max , Flow Duration\n\n# +\nsessions = []\ndata = {}\ntime_data = {}\ndata_size = {}\nfor row in df.iterrows():\n \n address = row[1]['sender_ip'] + row[1]['destination_ip']\n address2 = row[1]['destination_ip'] + row[1]['sender_ip']\n \n if row[1]['flags'] == 'S': # SYN frst step in conection setup\n sessions.append(address)\n data[address] = []\n time_data[address] = []\n data_size[address] = 0\n \n if row[1]['flags'] == 'A' or row[1]['flags'] == 'PA':\n \n \n if address in sessions: # forward\n \n if len(data[address])<1: # start time\n time_data[address].append(row[1]['start_time'])\n\n value = row[1]['length']\n data[address].append(value)\n \n data_size[address] = data_size[address] + row[1]['data_of_s']\n \n if address2 in sessions: # Backward\n data_size[address2] = data_size[address2] + row[1]['data_of_s']\n\n \n if row[1]['flags'] == 'F' or row[1]['flags'] == 'FA':\n \n if address in sessions:\n time_data[address].append(row[1]['end_time'])\n data_size[address] = data_size[address] + row[1]['data_of_s']\n\n elif address2 in sessions:\n time_data[address2].append(row[1]['end_time'])\n data_size[address2] = data_size[address2] + row[1]['data_of_s']\n\n \n\n \n \n \n\n# -\n\ndata\n\n# +\n# Forward Packet length max\n\nfor key, values in data.items():\n if len(values)>0:\n print(np.max(values))\n# -\n\ntime_data\n\n# Duration\nflow_duration = {}\nfor key, values in time_data.items():\n if len(values)>1:\n flow_duration[key] = values[-1]-values[0]\n \n\n# Flow bytes per second\nflow_bytes_per_second = {}\nfor key, values in data_size.items():\n if key in flow_duration:\n flow_bytes_per_second[key] = (values/flow_duration[key]) * 8\n\nflow_bytes_per_second\n\n\n","repo_name":"SaqlainHussainShah/real-time-anomaly-detection","sub_path":"extract features.ipynb","file_name":"extract features.ipynb","file_ext":"py","file_size_in_byte":4206,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"7285500385","text":"# **2.HAFTA ÖDEV KONUSU : Bu hafta öğrendiğimiz Partial Dependence’ı ve Extrapolation Sorununu giderme konularını kendi seçtiğiniz bir veriye uygulamak.**\n\n# Geçen hafta üzerinde çalıştığım bike sharing demand veri seti üzerinde partial dependence ve extrapolasyon konularını inceliyor olacağım\n\n# +\n# Gerekli kütüphanelerin import edilmesi\n\nfrom sklearn.ensemble import RandomForestRegressor, RandomForestClassifier\nfrom IPython.display import display\nimport scipy\nfrom pdpbox import pdp\nfrom plotnine import *\nfrom matplotlib import pyplot as plt\nfrom sklearn import metrics\nimport pandas as pd\nimport numpy as np\nimport math\nimport re\nfrom pandas.api.types import is_string_dtype\nfrom pandas.api.types import is_numeric_dtype\nfrom sklearn.ensemble import forest\nimport warnings\nwarnings.filterwarnings('ignore')\n# -\n\nPATH = \"data/bike_sharing_demand/\" # dosya yolunu path ile tutuyorum\n\ndf_main= pd.read_csv(f'{PATH}train.csv', low_memory=False, \n parse_dates=[\"count\"]) # dosyayı csv formatında okuyup üzerinde çalışacağımız dataframe haline getiriyorum.\n\ndf_main.head(10) # ilk 10 satırı gözlemleyelim\n\ndf_main.datetime = pd.to_datetime(df_main.datetime)\n\ndf_main.drop(['casual','registered'],axis=1,inplace=True) # count feature'ı hedef değişken olduğu için bu 2 değişkenin toplamı doğrudan countu vereceği için silebiliriz.\n\n\n# +\n# tüm kolonları görebilmek için display_all_dataframe fonksiyonunu oluşturalım\n\ndef display_all_dataframe(df):\n with pd.option_context(\"display.max_rows\", 1000, \"display.max_columns\", 1000): \n display(df)\n\n\n# -\n\ndf_main[\"count\"] = df_main[\"count\"].astype(\"float\") # hedef değişkenin veri tipini objectten floata dönüştürelim\n\ndf_main[\"count\"] = np.log1p(df_main[\"count\"]) # hedef değişkenin logaritmasını alalım (log1p olduğu için her değere 1 ekleyip log değeri hesaplanır)\n\nweather=pd.get_dummies(df_main['weather'],prefix='weather')\ndf_=pd.concat([df_main,weather],axis=1)\n#df_main.drop(['wheather','season'],axis=1,inplace=True)\n# her bir sezon değeri için ayrı bir feature oluşturalım. Bunun nedeni özellikleri geliştirecek olmasıdır.\nseason=pd.get_dummies(df_main['season'],prefix='season')\ndf_main=pd.concat([df_main,season],axis=1)\ndf_main.head()\n\n\n# +\n# bu fonksiyon sayesinde modele tarih ile ilgili yeni sütunlar eklenir ve model için anlamlı fraturelar üzerinde öalışılmaya yardımcı olunur\n\ndef add_info_datepart(df, dt_name, drop=True, time=True):\n \"Creates new columns from our datetime column\"\n \n \n dt_column = df[dt_name]\n column_dtype = dt_column.dtype\n \n\n targ_name = re.sub('[Dd]ate$', '', dt_name)\n \n \n attr = ['Year', 'Month', 'Week', 'Day', 'Dayofweek', 'Dayofyear',\n 'Is_month_end', 'Is_month_start', 'Is_quarter_end', 'Is_quarter_start', 'Is_year_end', 'Is_year_start']\n \n if time: \n attr = attr + ['Hour'] # verilen datada sadece saat dataları var olduğundan dakika ve saniye bilgilerini bilmiyoruz\n \n for a in attr: \n df[targ_name + a] = getattr(dt_column.dt, a.lower())\n \n # ne kadar zaman geçtiğini hesaplama için 1 sütun daha oluşturulalım\n df[targ_name + 'Elapsed'] = dt_column.astype(np.int64) // 10 ** 9\n \n if drop: \n df.drop(dt_name, axis=1, inplace=True)\n# -\n\nadd_info_datepart(df_main, \"datetime\") # fonksiyona gidildikten sonra datetine sütunu silinir bu sütından kazanılan bilgiler yeni kolonlara aktarılır.\n\n\n# +\n# veri setimizin ilk n satırı train diğer satırlar validasyon datasına aktarılır\n\ndef split_train_val(df,n): \n \n return df[:n].copy(), df[n:].copy()\n\n\n# +\n# kullanacağımız metrik formülü\n\ndef rmse(x,y): \n return math.sqrt(((x-y)**2).mean())\n\n\n# -\n\n# skor hesabı verilen veri seti için rmsle olduğundan log alınan verilere rmse uygulanır.\ndef print_score(m):\n \n print(f\"RMSE of train set {rmse(m.predict(x), y)}\")\n print(f\"RMSE of validation set {rmse(m.predict(x_valid), y_valid)}\")\n print(f\"R^2 of train set {m.score(x, y)}\")\n print(f\"R^2 of validation set {m.score(x_valid, y_valid)}\")\n\n\n# +\n#Scikit'in rastgele ormanlarını öğrenip her ağaca rastgele bir örnek verecek şekilde değiştirir.\n\ndef set_rf_samples(n):\n\n forest._generate_sample_indices = (lambda rs, n_samples:\n forest.check_random_state(rs).randint(0, n_samples, n))\n\n# +\n# Set_rf_samples tarafından üretilen değişiklikleri geri alır\n\n\ndef reset_rf_samples():\n\n forest._generate_sample_indices = (lambda rs, n_samples:\n forest.check_random_state(rs).randint(0, n_samples, n_samples))\n\n\n# -\n\ndef numericalize(df, col, name, max_cat):\n if not is_numeric_dtype(col) and (max_cat is None or len(col.cat.categories)>max_cat):\n df[name] = pd.Categorical(col).codes+1\n\n\n# +\n\ndef fix_missing(df, col, name, nan_dict, is_train):\n \n if is_train:\n if is_numeric_dtype(col):\n if pd.isnull(col).sum():\n df[name+\"_na\"] = pd.isnull(col)\n nan_dict[name] = col.median()\n df[name] = col.fillna(nan_dict[name])\n \n \n else:\n if is_numeric_dtype(col):\n if name in nan_dict:\n df[name+\"_na\"] = pd.isnull(col)\n df[name] = col.fillna(nan_dict[name])\n \n \n else:\n df[name] = col.fillna(col.median())\n \n# -\n\ndef apply_proc_df(df, y_fld,nan_dict=None,is_train=True, max_cat=None):\n \n df = df.copy()\n y = df[y_fld].values\n \n df.drop(y_fld, axis = 1, inplace = True)\n \n if nan_dict is None:\n nan_dict = {}\n \n for n, c in df.items():\n fix_missing(df, c, n,nan_dict, is_train)\n numericalize(df, c, n, max_cat)\n \n df = pd.get_dummies(df, dummy_na=True)\n \n \n return df, y, nan_dict\n\n\n# +\n# tüm verileri ele alarak model oluşturalım\nn_valid = int(len(df_main)/10) \nn_trn = len(df_main)-n_valid\n\ndf_train, y_train, nan_dict = apply_proc_df(df_main, 'count', max_cat=7)\n\nx, x_valid = split_train_val(df_train, n_trn)\ny, y_valid = split_train_val(y_train, n_trn)\n\nm = RandomForestRegressor(n_estimators=40, min_samples_leaf=3, max_features=0.6, n_jobs=-1, oob_score=True)\nm.fit(x, y)\nprint_score(m)\n# -\n\nx.shape,y.shape,x_valid.shape,y_valid.shape # tüm veri setini kullanarak 1/10'u val kalanını eğitim datası olarak ayırıp shape ile veri sayısı kontrolü yapıldı.\n\n\n# +\n#Subsampling yaparak model oluşturalım\n\n# veri setimizden büyüklüğünü kendimiz bir örneklem kümesi alalım:\ndef get_sample(df,n):\n\n idxs = np.random.permutation(len(df))[:n]\n return idxs, df.iloc[idxs].copy()\n\n\n# -\n\nidxs, xs = get_sample(x, 8000) # Veri setinden 800 tane örneklem alalım\nys = y[idxs]\n\nxs.shape, ys.shape, x_valid.shape, y_valid.shape # xs:x_sample and ys:y_sample\n\n# +\n# Oluşturduğumuz sample ile model kuruyoruz\n\nm = RandomForestRegressor(n_estimators=40, min_samples_leaf=3, max_features=0.6, n_jobs=-1, oob_score=True)\nm.fit(xs, ys)\nprint_score(m)\n\n\n# +\n\ndef randomforest_feat_importance(m, df): # veri setindeki her kolonu önem derecesine göre sıralar\n return pd.DataFrame({'columns':df.columns, 'importance':m.feature_importances_}\n ).sort_values('importance', ascending=False)\n\n\ndef plotting_importance(fi): # belirlenen önemli featureları görüntüler\n return fi.plot('columns', 'importance', 'barh', figsize=(12,7), legend=False)\n\n\ndef get_samples(df,n):\n \n idxs = sorted(np.random.permutation(len(df))[:n])\n return df.iloc[idxs].copy()\n\n\n# +\n# En önemli featureları 1.ödevdeki feature importance ile belirledik\n\nfeature_importance = randomforest_feat_importance(m, df_train) #tüm kolonların importancelerını döndürür \n\nkeep_columns = feature_importance[feature_importance[\"importance\"]>0.005][\"columns\"]\n\ndf_keep = df_train[keep_columns].copy()\n\nprint(\"önemli görülenkolon sayısı\\n\",len(keep_columns))\nprint(keep_columns)\n\n# +\n# Önemlilik derecelerine göre featureları çizdirelim.\n# -\n\nplotting_importance(randomforest_feat_importance(m, df_train)[:15]) \n\n# +\n# mainplot yardımıyla datetimeHour değişkeninin hedef değişken ile ilgili etkisi belli bir aralıkta gösterilebilir.\n# Bu örneğimde 0-24 saatleri arasında ortalama yapılan bisikleeet satışları gösterilmektedir(log olarak)\n\ndf_main.plot('datetimeHour', 'count', 'scatter', alpha=1, figsize=(10,8));\n# -\n\nx_all = get_samples(df_main[df_main.datetimeHour>0], df_main.shape[0])\nx_all.shape\n\ndisplay_all_dataframe(x_all.head())\n\n# +\n#ggplot yardımıyla saat bazında count değişimini göstrelim.\n\nggplot(x_all, aes('datetimeHour', 'count'))+stat_smooth(se=True, method='lowess' )\n# -\n\n# # KISMİ BAĞIMLILIKLARIN İNCELENMESİ\n\n#\n# Rastgele orman ve güçlendirme gibi kara kutu makine öğrenimi algoritmalarını kullanırken, tahmin ediciler ve model sonucu arasındaki ilişkileri anlamak zordur. Örneğin, rastgele orman açısından, aldığımız tek şey özellik önemidir. Önem hesaplamasına dayanarak hangi özelliğin sonucu önemli ölçüde etkilediğini bilsek de, hangi yönü etkilediğini bilmiyor olmamız gerçekten kötü bir durumdur. Ve gerçek vakaların çoğunda, etki monoton değildir. Tahmin ediciler ve model tahmini arasındaki karmaşık ilişkileri anlamaya yardımcı olacak bazı güçlü araçlara ihtiyacımız var.\n\n# +\nfrom pdpbox import pdp\nfrom plotnine import *\n\n# # !pip install PDPbox==0.2.0 versiyonu kullanılmıştır. 0.1.0'da bazı hatalar ile karşılaşılmıştır.\n# -\n\nx_sample = get_samples(x[x[\"datetimeHour\"]>0], 1000)\n\nx.columns.shape,x_sample.columns.shape,\n\n\ndef plot_pdp(feat, clusters=None, feat_name=None):\n feat_name = feat_name or feat\n p = pdp.pdp_isolate(m, x_sample, x_sample.columns, feat)\n return pdp.pdp_plot(p, feat_name, plot_lines=True,\n cluster=clusters is not None,\n n_cluster_centers=clusters)\n\n\nfeat = 'datetimeHour'\n\nplot_pdp('datetimeHour') # Modelde en önemli feature olan datetimeHour'un tüm gözlem değerleri kullanılarak tahminleri ve ortalamayı çizdirelim.\n\nplot_pdp('datetimeHour', clusters = 10) # 10 gözlem değeri ve bunun ortalamasını çizdirelim\n\nplot_pdp('atemp') # Modelimizde diğer bir önemli feature olan atemp için tahminleri görselleştirelim.\n\n# # Çoklu Değişkenler Bağımlı Değişkeni Nasıl Etkiler?\n\n# +\n# # ?pdp_interact_plot\n#pdp_interact kullanılarak belirlenen featurelar için etkileşim görselleştirilebilir.\n\n# +\nfrom pdpbox.pdp import pdp_interact, pdp_interact_plot\n\nfeatures = ['datetimeHour', 'atemp'] # Modeli en çok etkileyen featureların birbiri ile ilişkisini aynı grafikte gösterelim\n\ninteraction = pdp_interact(\n model=m,\n dataset=x_sample,\n model_features=x_sample.columns,\n features=features \n)\n\npdp_interact_plot(interaction, plot_type='grid', feature_names=features); \n# -\n\n\n\n# # Ekstrapolasyon Sorunu İle Başa Çıkma\n\n#\n# Validation kümesinden gelip gelmediğini anlamak için is_valid olarak yeni bir sütun ekliyoruz\n\ndf_ext = df_keep.copy()\ndf_ext['is_valid'] = 1\ndf_ext.is_valid[:n_trn] = 0\nx, y, nas = apply_proc_df(df_ext, 'is_valid')\n\nm = RandomForestClassifier(n_estimators=40, min_samples_leaf=3, max_features=0.5, n_jobs=-1, oob_score=True)\nm.fit(x, y);\nm.oob_score_\n\n# Bu modelin feature önemini bulursak, bu bize doğrulamadaki verileri eğitimden ayıran önemli parametreleri söyleyecektir.\n\nfi = randomforest_feat_importance(m, x); fi[:10]\n\nfeats=['datetimeElapsed', 'datetimeDayofyear', 'datetimeWeek']\n\n(x[feats]/1000).describe()\n\n(x_valid[feats]/1000).describe()\n\nx.drop(feats, axis=1, inplace=True)\n\nm = RandomForestClassifier(n_estimators=40, min_samples_leaf=3, max_features=0.5, n_jobs=-1, oob_score=True)\nm.fit(x, y);\nm.oob_score_\n\nfi = randomforest_feat_importance(m, x); fi[:10]\n\n#\n# Bike sharing demand yarışmanın test verilerini eğitim setinize koyup ve \"is_test\" adlı bir sütun oluşturalım, bu sütunu tahmin eden bir model oluşturalım. Modelimiz makul bir aralıkta tahmin yapabiliyorsa, bu, test setinin rastgele olmadığı anlamına gelir! Dolayısıyla, çapraz doğrulama ve yerleşik bölme yöntemleri test setini temsil etmeyecektir, bu nedenle doğrulama setinin nasıl oluşturulacağını bulmamız gerekir \n\ncols_feats=['humidity', 'temp', 'datetimeHour', 'windspeed', 'datetimeDayofweek', 'atemp']\n\n#\n# cols_featsde belirtilen kolonları almak zararlı değilse, bu daha iyidir çünkü bu şekilde zaman içinde genelleyen daha iyi bir modele sahip olacağız.\n\ny, y_valid = split_train_val(y_train, n_trn)\n\nx, x_valid = split_train_val(df_keep, n_trn)\nm = RandomForestRegressor(n_estimators=40, min_samples_leaf=3, max_features=0.5, n_jobs=-1, oob_score=True)\nm.fit(x, y)\nprint_score(m)\n\nfor f in cols_feats:\n df_subs = df_keep.drop(f, axis=1)\n x, x_valid = split_train_val(df_subs, n_trn)\n m = RandomForestRegressor(n_estimators=40, min_samples_leaf=3, max_features=0.5, n_jobs=-1, oob_score=True)\n m.fit(x, y)\n print(f)\n print_score(m)\n\nreset_rf_samples()\n\n# +\n#final modelimizin sonuçları :\n\ndf_subs = df_keep.drop(['temp', 'windspeed', 'atemp'], axis=1)\nx, x_valid = split_train_val(df_subs, n_trn)\nm = RandomForestRegressor(n_estimators=40, min_samples_leaf=3, max_features=0.5, n_jobs=-1, oob_score=True)\nm.fit(x, y)\nprint_score(m)\n","repo_name":"AYSE-DUMAN/Kodluyoruz-Uygulamal-Veri-Bilimi-102-Bootcamp-odev-ve-Uygulamalar","sub_path":"Ayşe-Duman-homework2/2.hafta-Ayşe-Duman.ipynb","file_name":"2.hafta-Ayşe-Duman.ipynb","file_ext":"py","file_size_in_byte":13448,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"74798799178","text":"# +\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport os\nimport glob\nimport pandas as pd\nimport seaborn as sns\nimport SimpleITK as sitk\nimport matplotlib.backends.backend_pdf\nimport pandas as pd\n\nplt.rcParams['figure.figsize'] = [18, 15]\n# -\n\nroot_path = \"/Users/amithkamath/repo/deep-planner\"\nresults_path = os.path.join(root_path, \"results\")\n\n# +\nperturbation_data = os.path.join(results_path, \"perturbation_data_new.csv\")\n\nresults = pd.read_csv(perturbation_data)\nresults = results.sort_values(by=['Structure'])\n# -\n\nresults.loc[results['Structure'] == \"BrainStem\"]\n\n# +\n#ax = sns.boxplot(data=results, x=\"Structure\", y=\"AvgDiff\")\n#plt.grid() #just add this\n#sns.move_legend(ax, \"upper left\", bbox_to_anchor=(1, 1))\n# -\n\nax = sns.boxplot(data=results, x=\"Structure\", y=\"XCorrDist\")\nplt.grid() #just add this\n#sns.move_legend(ax, \"upper left\", bbox_to_anchor=(1, 1))\n\nresults.groupby('Structure').mean()\n\nax = sns.boxplot(data=results, x=\"Structure\", y=\"XCorrGrad\")\nplt.grid() #just add this\n#sns.move_legend(ax, \"upper left\", bbox_to_anchor=(1, 1))\n\n# +\ngeometry_data = os.path.join(results_path, \"target_geometry.csv\")\n\ntarget_geometry = pd.read_csv(geometry_data)\n# -\n\ntarget_geometry.columns\n\n# +\nax = sns.scatterplot(data=target_geometry, x=\"Volume\", y=\"Dose\", s=1000)\nplt.grid()\n\nfor i in range(target_geometry.shape[0]):\n plt.text(x=target_geometry.Volume[i],y=target_geometry.Dose[i],s=target_geometry.Case[i], \n fontdict=dict(color='red',size=10),\n bbox=dict(facecolor='yellow',alpha=0.5))\n\nplt.xlabel('Volume fraction of Target (to Brain)', fontsize=18)\nplt.ylabel('Dose Score', fontsize=18)\n\nprops = dict(boxstyle='round', facecolor='wheat', alpha=0.5)\nax.text(0.5, 0.1, \"Correlation Coefficient: 0.6622\", transform=ax.transAxes, fontsize=28,\n verticalalignment='top', bbox=props)\n\n# +\nax = sns.scatterplot(data=target_geometry, x=\"Volume\", y=\"DVH\", s=1000)\nplt.grid()\nfor i in range(target_geometry.shape[0]):\n plt.text(x=target_geometry.Volume[i],y=target_geometry.DVH[i],s=target_geometry.Case[i], \n fontdict=dict(color='red',size=10),\n bbox=dict(facecolor='yellow',alpha=0.5))\n\nplt.xlabel('Volume fraction of Target (to Brain)', fontsize=18)\nplt.ylabel('DVH Score', fontsize=18)\n\nprops = dict(boxstyle='round', facecolor='wheat', alpha=0.5)\nax.text(0.5, 0.1, \"Correlation Coefficient: 0.8061\", transform=ax.transAxes, fontsize=28,\n verticalalignment='top', bbox=props)\n","repo_name":"amithjkamath/astra","sub_path":"notebooks/3.0-ajk-transformation-graphs.ipynb","file_name":"3.0-ajk-transformation-graphs.ipynb","file_ext":"py","file_size_in_byte":2447,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"42184944234","text":"# + hideCode=true hideOutput=true hidePrompt=true jupyter={\"source_hidden\": true} slideshow={\"slide_type\": \"skip\"} tags=[\"remove-cell\"]\n# %config InlineBackend.figure_formats = ['svg']\nimport os\n\nSTATIC_WEB_PAGE = {\"EXECUTE_NB\", \"READTHEDOCS\"}.intersection(os.environ)\n# -\n\n# ```{autolink-concat}\n# ```\n\n# + [markdown] tags=[\"physics\", \"sympy\"]\n# ::::{margin}\n# :::{card} Symbolic phase space boundary for a three-body decay\n# TR-017\n# ^^^\n# This reports shows how define the physical phase space region on a Dalitz plot using a Kibble function.\n# # +++\n# ✅ [compwa-org#139](https://github.com/ComPWA/compwa-org/issues/139)\n# :::\n# ::::\n# -\n\n# # Phase space for a three-body decay\n\n# + jupyter={\"source_hidden\": true} tags=[\"remove-cell\"]\n# %pip install -q ampform==0.14.0 sympy==1.10.1 tensorwaves==0.4.5\n\n# + jupyter={\"source_hidden\": true} tags=[\"hide-cell\"]\nfrom __future__ import annotations\n\nimport warnings\nfrom typing import TYPE_CHECKING\n\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport sympy as sp\nfrom ampform.sympy import (\n UnevaluatedExpression,\n create_expression,\n implement_doit_method,\n make_commutative,\n)\nfrom IPython.display import Math\nfrom ipywidgets import FloatSlider, VBox, interactive_output\nfrom matplotlib.patches import Patch\nfrom tensorwaves.function.sympy import create_parametrized_function\n\nif TYPE_CHECKING:\n from matplotlib.axis import Axis\n from matplotlib.contour import QuadContourSet\n from matplotlib.lines import Line2D\n\nwarnings.filterwarnings(\"ignore\")\n\n\n# -\n\n# Kinematics for a three-body decay $0 \\to 123$ can be fully described by two **Mandelstam variables** $\\sigma_1, \\sigma_2$, because the third variable $\\sigma_3$ can be expressed in terms $\\sigma_1, \\sigma_2$, the mass $m_0$ of the initial state, and the masses $m_1, m_2, m_3$ of the final state. As can be seen, the roles of $\\sigma_1, \\sigma_2, \\sigma_3$ are interchangeable.\n#\n# ```{margin}\n# See Eq. (1.2) in {cite}`bycklingParticleKinematics1973`\n# ```\n\n# + jupyter={\"source_hidden\": true} tags=[\"hide-input\", \"keep_output\"]\ndef compute_third_mandelstam(sigma1, sigma2, m0, m1, m2, m3) -> sp.Expr:\n return m0**2 + m1**2 + m2**2 + m3**2 - sigma1 - sigma2\n\n\nm0, m1, m2, m3 = sp.symbols(\"m:4\")\ns1, s2, s3 = sp.symbols(\"sigma1:4\")\ncomputed_s3 = compute_third_mandelstam(s1, s2, m0, m1, m2, m3)\nMath(Rf\"{sp.latex(s3)} = {sp.latex(computed_s3)}\")\n\n\n# -\n\n# The phase space is defined by the closed area that satisfies the condition $\\phi(\\sigma_1,\\sigma_2) \\leq 0$, where $\\phi$ is a **Kibble function**:\n#\n#\n# ```{margin}\n# See §V.2 in {cite}`bycklingParticleKinematics1973`\n# ```\n\n# + jupyter={\"source_hidden\": true} tags=[\"scroll-input\", \"hide-input\", \"keep_output\"]\n@make_commutative\n@implement_doit_method\nclass Kibble(UnevaluatedExpression):\n def __new__(cls, sigma1, sigma2, sigma3, m0, m1, m2, m3, **hints) -> Kibble:\n return create_expression(\n cls, sigma1, sigma2, sigma3, m0, m1, m2, m3, **hints\n )\n\n def evaluate(self) -> sp.Expr:\n sigma1, sigma2, sigma3, m0, m1, m2, m3 = self.args\n return Källén(\n Källén(sigma2, m2**2, m0**2),\n Källén(sigma3, m3**2, m0**2),\n Källén(sigma1, m1**2, m0**2),\n )\n\n def _latex(self, printer, *args):\n sigma1, sigma2, *_ = map(printer._print, self.args)\n return Rf\"\\phi\\left({sigma1}, {sigma2}\\right)\"\n\n\n@make_commutative\n@implement_doit_method\nclass Källén(UnevaluatedExpression):\n def __new__(cls, x, y, z, **hints) -> Källén:\n return create_expression(cls, x, y, z, **hints)\n\n def evaluate(self) -> sp.Expr:\n x, y, z = self.args\n return x**2 + y**2 + z**2 - 2 * x * y - 2 * y * z - 2 * z * x\n\n def _latex(self, printer, *args):\n x, y, z = map(printer._print, self.args)\n return Rf\"\\lambda\\left({x}, {y}, {z}\\right)\"\n\n\nkibble = Kibble(s1, s2, s3, m0, m1, m2, m3)\nMath(Rf\"{sp.latex(kibble)} = {sp.latex(kibble.doit(deep=False))}\")\n# -\n\n# and $\\lambda$ is the **Källén function**:\n\n# + jupyter={\"source_hidden\": true} tags=[\"hide-input\", \"keep_output\"]\nx, y, z = sp.symbols(\"x:z\")\nexpr = Källén(x, y, z)\nMath(f\"{sp.latex(expr)} = {sp.latex(expr.doit())}\")\n\n\n# -\n\n# Any distribution over the phase space can now be defined using a two-dimensional grid over a Mandelstam pair $\\sigma_1,\\sigma_2$ of choice, with the condition $\\phi(\\sigma_1,\\sigma_2)<0$ selecting the values that are physically allowed.\n\n# + jupyter={\"source_hidden\": true} tags=[\"hide-input\", \"keep_output\"]\ndef is_within_phasespace(\n sigma1, sigma2, m0, m1, m2, m3, outside_value=sp.nan\n) -> sp.Piecewise:\n sigma3 = compute_third_mandelstam(sigma1, sigma2, m0, m1, m2, m3)\n kibble = Kibble(sigma1, sigma2, sigma3, m0, m1, m2, m3)\n return sp.Piecewise(\n (1, sp.LessThan(kibble, 0)),\n (outside_value, True),\n )\n\n\nis_within_phasespace(s1, s2, m0, m1, m2, m3)\n# -\n\nphsp_expr = is_within_phasespace(s1, s2, m0, m1, m2, m3, outside_value=0)\nphsp_func = create_parametrized_function(\n phsp_expr.doit(),\n parameters={m0: 2.2, m1: 0.2, m2: 0.4, m3: 0.4},\n backend=\"numpy\",\n)\n\n# + tags=[\"remove-input\"]\n# %matplotlib widget\n\n# + jupyter={\"source_hidden\": true} tags=[\"scroll-input\", \"hide-input\", \"remove-output\"]\nsliders = {\n \"m0\": FloatSlider(description=\"m0\", max=3, value=2.1, step=0.01),\n \"m1\": FloatSlider(description=\"m1\", max=2, value=0.2, step=0.01),\n \"m2\": FloatSlider(description=\"m2\", max=2, value=0.4, step=0.01),\n \"m3\": FloatSlider(description=\"m3\", max=2, value=0.4, step=0.01),\n}\n\nresolution = 300\nX, Y = np.meshgrid(\n np.linspace(0, 4, num=resolution),\n np.linspace(0, 4, num=resolution),\n)\ndata = {\"sigma1\": X, \"sigma2\": Y}\n\nsidebar_ratio = 0.15\nfig, ((ax1, _), (ax, ax2)) = plt.subplots(\n figsize=(7, 7),\n ncols=2,\n nrows=2,\n gridspec_kw={\n \"height_ratios\": [sidebar_ratio, 1],\n \"width_ratios\": [1, sidebar_ratio],\n },\n)\n_.remove()\nax.set_xlim(0, 4)\nax.set_ylim(0, 4)\nax.set_xlabel(R\"$\\sigma_1$\")\nax.set_ylabel(R\"$\\sigma_2$\")\nax.set_xticks(range(5))\nax.set_yticks(range(5))\nax1.set_xlim(0, 4)\nax2.set_ylim(0, 4)\nfor a in [ax1, ax2]:\n a.set_xticks([])\n a.set_yticks([])\n a.axis(\"off\")\nfig.tight_layout()\nfig.subplots_adjust(wspace=0, hspace=0)\n\nfig.canvas.toolbar_visible = False\nfig.canvas.header_visible = False\nfig.canvas.footer_visible = False\n\nMESH: QuadContourSet | None = None\nPROJECTIONS: tuple[Line2D, Line2D] = None\nBOUNDARIES: list[Line2D] | None = None\n\n\ndef plot(**parameters):\n draw_boundaries(\n parameters[\"m0\"],\n parameters[\"m1\"],\n parameters[\"m2\"],\n parameters[\"m3\"],\n )\n global MESH, PROJECTIONS\n if MESH is not None:\n for coll in MESH.collections:\n ax.collections.remove(coll)\n phsp_func.update_parameters(parameters)\n Z = phsp_func(data)\n MESH = ax.contour(X, Y, Z, colors=\"black\")\n contour = MESH.collections[0]\n contour.set_facecolor(\"lightgray\")\n x = X[0]\n y = Y[:, 0]\n Zx = np.nansum(Z, axis=0)\n Zy = np.nansum(Z, axis=1)\n if PROJECTIONS is None:\n PROJECTIONS = (\n ax1.plot(x, Zx, c=\"black\", lw=2)[0],\n ax2.plot(Zy, y, c=\"black\", lw=2)[0],\n )\n else:\n PROJECTIONS[0].set_data(x, Zx)\n PROJECTIONS[1].set_data(Zy, y)\n ax1.relim()\n ax2.relim()\n ax1.autoscale_view(scalex=False)\n ax2.autoscale_view(scaley=False)\n create_legend(ax)\n fig.canvas.draw()\n\n\ndef draw_boundaries(m0, m1, m2, m3) -> None:\n global BOUNDARIES\n s1_min = (m2 + m3) ** 2\n s1_max = (m0 - m1) ** 2\n s2_min = (m1 + m3) ** 2\n s2_max = (m0 - m2) ** 2\n if BOUNDARIES is None:\n BOUNDARIES = [\n ax.axvline(s1_min, c=\"red\", ls=\"dotted\", label=\"$(m_2+m_3)^2$\"),\n ax.axhline(s2_min, c=\"blue\", ls=\"dotted\", label=\"$(m_1+m_3)^2$\"),\n ax.axvline(s1_max, c=\"red\", ls=\"dashed\", label=\"$(m_0-m_1)^2$\"),\n ax.axhline(s2_max, c=\"blue\", ls=\"dashed\", label=\"$(m_0-m_2)^2$\"),\n ]\n else:\n BOUNDARIES[0].set_data(get_line_data(s1_min))\n BOUNDARIES[1].set_data(get_line_data(s2_min, horizontal=True))\n BOUNDARIES[2].set_data(get_line_data(s1_max))\n BOUNDARIES[3].set_data(get_line_data(s2_max, horizontal=True))\n\n\ndef create_legend(ax: Axis):\n if ax.get_legend() is not None:\n return\n label = Rf\"${sp.latex(kibble)}\\leq0$\"\n ax.legend(\n handles=[\n Patch(label=label, ec=\"black\", fc=\"lightgray\"),\n *BOUNDARIES,\n ],\n loc=\"upper right\",\n facecolor=\"white\",\n framealpha=1,\n )\n\n\ndef get_line_data(value, horizontal: bool = False) -> np.ndarray:\n pair = (value, value)\n if horizontal:\n return np.array([(0, 1), pair])\n return np.array([pair, (0, 1)])\n\n\noutput = interactive_output(plot, controls=sliders)\nVBox([output, *sliders.values()])\n# -\n\n# {{ run_interactive }}\n#\n# ```{image} https://user-images.githubusercontent.com/29308176/165257660-af66c9c1-2da9-475f-b4c4-4a52950acd00.gif\n# :alt: Cell output - interactive Dalitz plot\n# :align: center\n# ```\n#
\n\n# The phase space boundary can be described analytically in terms of $\\sigma_1$ or $\\sigma_2$, in which case there are two solutions:\n\nsol1, sol2 = sp.solve(kibble.doit().subs(s3, computed_s3), s2)\n\n# + jupyter={\"source_hidden\": true} tags=[\"remove-input\", \"keep_output\", \"full-width\"]\nlatex = R\"\\begin{array}{c}\" + \"\\n\"\nfor sol in [sol1, sol2]:\n latex += Rf\" {sp.latex(sol)} \\\\\" + \"\\n\"\nlatex += R\"\\end{array}\"\nMath(latex)\n# -\n\n# The boundary cannot be parametrized analytically in polar coordinates, but there is a numeric solution. The idea is to solve the condition $\\phi(\\sigma_1,\\sigma_2)=0$ after the following substitutions:\n#\n# ```{margin}\n# See {cite}`bycklingParticleKinematics1973`, pp. 109–112\n# ```\n\n# + tags=[\"hide-input\"]\nT0, T1, T2, T3 = sp.symbols(\"T0:4\")\nr, theta = sp.symbols(\"r theta\", nonnegative=True)\nsubstitutions = {\n s1: (m2 + m3) ** 2 + T1,\n s2: (m1 + m3) ** 2 + T2,\n s3: (m1 + m2) ** 2 + T3,\n T1: T0 / 3 - r * sp.cos(theta),\n T2: T0 / 3 - r * sp.cos(theta + 2 * sp.pi / 3),\n T3: T0 / 3 - r * sp.cos(theta + 4 * sp.pi / 3),\n T0: (\n m0**2\n + m1**2\n + m2**2\n + m3**2\n - (m2 + m3) ** 2\n - (m1 + m3) ** 2\n - (m1 + m2) ** 2\n ),\n}\n\n# + jupyter={\"source_hidden\": true} tags=[\"remove-input\", \"keep_output\"]\nlatex = R\"\\begin{array}{rcl}\" + \"\\n\"\nfor item in substitutions.items():\n lhs, rhs = map(sp.latex, item)\n latex += Rf\" {lhs} &=& {rhs} \\\\\" + \"\\n\"\nlatex += R\"\\end{array}\"\nMath(latex)\n# -\n\n# For every value of $\\theta \\in [0, 2\\pi)$, the value of $r$ can now be found by solving the condition $\\phi(r, \\theta)=0$. Note that $\\phi(r, \\theta)$ is a **cubic polynomial of $r$**. For instance, if we take $m_0=5, m_1=2, m_{2,3}=1$:\n\n# + tags=[\"hide-input\", \"remove-output\"]\nphi_r = (\n kibble.doit()\n .subs(substitutions) # substitute sigmas\n .subs(substitutions) # substitute T123\n .subs(substitutions) # substitute T0\n .subs({m0: 5, m1: 2, m2: 1, m3: 1})\n .simplify()\n .collect(r)\n)\n\n# + jupyter={\"source_hidden\": true} tags=[\"remove-input\", \"keep_output\"]\nlhs = sp.Symbol(R\"\\phi(r, \\theta)\")\nlatex = sp.multiline_latex(lhs, phi_r, environment=\"eqnarray\")\nMath(latex)\n# -\n\n# The lowest value of $r$ that satisfies $\\phi(r,\\theta)=0$ defines the phase space boundary.\n","repo_name":"ComPWA/compwa-org","sub_path":"docs/report/017.ipynb","file_name":"017.ipynb","file_ext":"py","file_size_in_byte":11395,"program_lang":"python","lang":"en","doc_type":"code","stars":5,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"20744334212","text":"# + [markdown] id=\"1wEUPnr2aHgk\"\n# Question 2\n#\n# • Create an empty dictionary called dog\n#\n# • Add name, color, breed, legs, age to the dog dictionary\n#\n# • Create a student dictionary and add first_name, last_name, gender, age, marital status,skills, country, city and address as keys for the dictionary\n#\n# • Get the length of the student dictionary\n#\n# • Get the value of skills and check the data type, it should be a list\n#\n# • Modify the skills values by adding one or two skills\n#\n# • Get the dictionary keys as a list\n#\n# • Get the dictionary values as a list\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"BRuZOcA1aTV2\" outputId=\"8c908abf-6a7e-488d-e500-d10940f7c703\"\ndog={}\ndog['name']=\"Jacky\"\ndog[\"color\"]=\"Black & Brown\"\ndog[\"breed\"]=\"German sheperd\"\ndog[\"legs\"]=\"long\"\ndog[\"age\"]=3\nprint('updated Dictionary \"dog\":',dog)\nstudent={\"Name\": \"Vinod Yadav\",\"Last_name\":\"Kannaboina\",\"Gender\":\"Male\",\"Marital status\": \"Unmarried\",\"Skills\":[\"python,SQL\"],\"Country\":\"India\",\"City\":\"Karimnagar\",\"Address\":\"HNO:12-137/1,Huzurabad\"}\nprint('\\n',student)\nprint('Length of dictionary:',len(student))\nx=student[\"Skills\"]\nprint(x)\nprint('data type of skils:',type(x))\nstudent['Skills']=[\"Machine Learning, Deep Learning\"]\nprint('Student Dictionary :',student)\ny=list(student.keys()) #returns keys of student dict as list\nz=list(student.values()) #returns values as list\nprint('Keys:',y,'\\n','Values:',z)\n","repo_name":"vinodyadav2413/Machine_Learning_Assignment_1","sub_path":"Question_2.ipynb","file_name":"Question_2.ipynb","file_ext":"py","file_size_in_byte":1425,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43129621195","text":"import numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nimport re, string, math\n\ndf = pd.read_csv('stock_market.csv')\n\ndf.head()\n\ndf.shape\n\n# The dataframe has 12,420 records and 4 columns\n\n# +\n# Dropping id column as it is same as index\n\ndf.drop('id', axis=1, inplace=True)\n# -\n\ndf.info()\n\n# Attribute Description:- \n#\n# \n# - created_at: The date on which the review was given
\n# - full_text: Complete review
\n# - score: The score given to review
\n#
\n\n# +\n# Changing datatype of created_at to datetime\n\ndf['created_at'] = pd.to_datetime(df['created_at'])\n\n# +\n# Checking for null values\n\ndf.isnull().sum()\n# -\n\n# The dataset doesn't have any null values\n\ndf.dtypes\n\n# +\n# Expanding contractions\n\n# Dictionary of English Contractions\ncontractions_dict = { \"ain't\": \"are not\",\"'s\":\" is\",\"aren't\": \"are not\",\n \"can't\": \"cannot\",\"can't've\": \"cannot have\",\n \"'cause\": \"because\",\"could've\": \"could have\",\"couldn't\": \"could not\",\n \"couldn't've\": \"could not have\", \"didn't\": \"did not\",\"doesn't\": \"does not\",\n \"don't\": \"do not\",\"hadn't\": \"had not\",\"hadn't've\": \"had not have\",\n \"hasn't\": \"has not\",\"haven't\": \"have not\",\"he'd\": \"he would\",\n \"he'd've\": \"he would have\",\"he'll\": \"he will\", \"he'll've\": \"he will have\",\n \"how'd\": \"how did\",\"how'd'y\": \"how do you\",\"how'll\": \"how will\",\n \"I'd\": \"I would\", \"I'd've\": \"I would have\",\"I'll\": \"I will\",\n \"I'll've\": \"I will have\",\"I'm\": \"I am\",\"I've\": \"I have\", \"isn't\": \"is not\",\n \"it'd\": \"it would\",\"it'd've\": \"it would have\",\"it'll\": \"it will\",\n \"it'll've\": \"it will have\", \"let's\": \"let us\",\"ma'am\": \"madam\",\n \"mayn't\": \"may not\",\"might've\": \"might have\",\"mightn't\": \"might not\", \n \"mightn't've\": \"might not have\",\"must've\": \"must have\",\"mustn't\": \"must not\",\n \"mustn't've\": \"must not have\", \"needn't\": \"need not\",\n \"needn't've\": \"need not have\",\"o'clock\": \"of the clock\",\"oughtn't\": \"ought not\",\n \"oughtn't've\": \"ought not have\",\"shan't\": \"shall not\",\"sha'n't\": \"shall not\",\n \"shan't've\": \"shall not have\",\"she'd\": \"she would\",\"she'd've\": \"she would have\",\n \"she'll\": \"she will\", \"she'll've\": \"she will have\",\"should've\": \"should have\",\n \"shouldn't\": \"should not\", \"shouldn't've\": \"should not have\",\"so've\": \"so have\",\n \"that'd\": \"that would\",\"that'd've\": \"that would have\", \"there'd\": \"there would\",\n \"there'd've\": \"there would have\", \"they'd\": \"they would\",\n \"they'd've\": \"they would have\",\"they'll\": \"they will\",\n \"they'll've\": \"they will have\", \"they're\": \"they are\",\"they've\": \"they have\",\n \"to've\": \"to have\",\"wasn't\": \"was not\",\"we'd\": \"we would\",\n \"we'd've\": \"we would have\",\"we'll\": \"we will\",\"we'll've\": \"we will have\",\n \"we're\": \"we are\",\"we've\": \"we have\", \"weren't\": \"were not\",\"what'll\": \"what will\",\n \"what'll've\": \"what will have\",\"what're\": \"what are\", \"what've\": \"what have\",\n \"when've\": \"when have\",\"where'd\": \"where did\", \"where've\": \"where have\",\n \"who'll\": \"who will\",\"who'll've\": \"who will have\",\"who've\": \"who have\",\n \"why've\": \"why have\",\"will've\": \"will have\",\"won't\": \"will not\",\n \"won't've\": \"will not have\", \"would've\": \"would have\",\"wouldn't\": \"would not\",\n \"wouldn't've\": \"would not have\",\"y'all\": \"you all\", \"y'all'd\": \"you all would\",\n \"y'all'd've\": \"you all would have\",\"y'all're\": \"you all are\",\n \"y'all've\": \"you all have\", \"you'd\": \"you would\",\"you'd've\": \"you would have\",\n \"you'll\": \"you will\",\"you'll've\": \"you will have\", \"you're\": \"you are\",\n \"you've\": \"you have\"}\n\n# Regular expression for finding contractions\ncontractions_re=re.compile('(%s)' % '|'.join(contractions_dict.keys()))\n\n# Function for expanding contractions\ndef expand_contractions(text,contractions_dict=contractions_dict):\n def replace(match):\n return contractions_dict[match.group(0)]\n return contractions_re.sub(replace, text)\n\n# Expanding Contractions in the reviews\ndf['full_text'] = df['full_text'].apply(lambda x:expand_contractions(x))\n\n# +\n# Converting text to lowercase\n\ndf['full_text'] = df['full_text'].apply(lambda x:x.lower())\n\n# +\n# Removing digits and words containing digits\n\ndf['full_text'] = df['full_text'].apply(lambda x: re.sub('\\w*\\d\\w*','', x))\n\n# +\n# Removing punctuations\n\ndf['full_text'] = df['full_text'].apply(lambda x: re.sub(r'[^\\w\\s]', '', x))\n\n# +\n# Removing extra spaces\n\ndf['full_text']=df['full_text'].apply(lambda x: re.sub(' +',' ',x))\n\n# +\n# Displaying the text after cleaning\n\nfor index,text in enumerate(df['full_text'][35:40]):\n print('Review %d:\\n'%(index+1), text)\n\n# +\n# Applying stopwords and lemmatization\n\nimport nltk\nfrom nltk.corpus import stopwords\n\nw_tokenizer = nltk.tokenize.WhitespaceTokenizer()\nlemmatizer = nltk.stem.WordNetLemmatizer()\n\ndef lemmatize_text(text):\n rev = [lemmatizer.lemmatize(w) for w in w_tokenizer.tokenize(text) if w not in stopwords.words('english')]\n rev = ' '.join(rev)\n return rev\n\ndf['text_lemmatized'] = df.full_text.apply(lemmatize_text)\n# -\n\n# Getting hour from created_at\ndf['hour'] = df['created_at'].dt.hour\n\n# +\n# Grouping according to hour of day\n\ndf_grouped=df[['hour','text_lemmatized']].groupby(by='hour').agg(lambda x:' '.join(x))\ndf_grouped.head()\n\n# +\n# Creating Document Term Matrix\n\nfrom sklearn.feature_extraction.text import CountVectorizer\n\ncv = CountVectorizer(analyzer='word')\n\ndata = cv.fit_transform(df_grouped['text_lemmatized'])\n\ndf_dtm = pd.DataFrame(data.toarray(), columns=cv.get_feature_names())\ndf_dtm.index = df_grouped.index\n\ndf_dtm.head(3)\n\n# +\ndf_dtm = df_dtm.astype(str)\n\n# Transposing document term matrix\ndf_dtm = df_dtm.transpose()\n\ndf_dtm\n\n# +\n# # Importing wordcloud for plotting word clouds and textwrap for wrapping longer text\n\n# from wordcloud import WordCloud\n# from textwrap import wrap\n\n# # Function for generating word clouds\n# def generate_wordcloud(data,title):\n# wc = WordCloud(width=400, height=330, max_words=150, colormap=\"Dark2\").generate_from_frequencies(data)\n# plt.figure(figsize=(10,8))\n# plt.imshow(wc, interpolation='bilinear')\n# plt.axis(\"off\")\n# plt.title('\\n'.join(wrap(title,60)),fontsize=13)\n# plt.show()\n\n# # Plotting word cloud for each hour\n# for index,hour in enumerate(df_dtm.columns):\n# generate_wordcloud(df_dtm[hour], index)\n\n# +\nfrom textblob import TextBlob\n\ndf['polarity']=df['text_lemmatized'].apply(lambda x:TextBlob(x).sentiment.polarity)\n# -\n\nprint(\"3 Random Reviews with Highest Polarity:\")\nfor index,review in enumerate(df.iloc[df['polarity'].sort_values(ascending=False)[:3].index]['full_text']):\n print('Review {}:\\n'.format(index+1),review)\n\nprint(\"3 Random Reviews with Lowest Polarity:\")\nfor index,review in enumerate(df.iloc[df['polarity'].sort_values(ascending=True)[:3].index]['full_text']):\n print('Review {}:\\n'.format(index+1),review)\n\n# +\n# Polarity by each hour\n\na = df.groupby('hour').agg({'polarity':'mean'})\na.plot(kind='bar', color='green')\nplt.legend()\n# -\n\n\n","repo_name":"devanshiipatel/21_Days_Devanshi_Patel","sub_path":"Day 3_Amazon EDA.ipynb","file_name":"Day 3_Amazon EDA.ipynb","file_ext":"py","file_size_in_byte":7560,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"25657205017","text":"# + [markdown] id=\"l_k4ldTl6xI6\"\n# # Overfitting(과적합)\n#\n# 신경망이 훈련 데이터에만 지나치게 적응되어 그 외의 데이터에는 제대로 대응하지 못하는 상태\n\n# + [markdown] id=\"F-JzSIeO6366\"\n# # 데이터 준비\n\n# + id=\"4-BR6aOX6XSY\"\nfrom sklearn.datasets import make_moons\nfrom tensorflow.keras.layers import Dense\nfrom tensorflow.keras.models import Sequential\nfrom matplotlib import pyplot\nfrom pandas import DataFrame\n\n\n# + id=\"yCz9rq_Q66Ak\"\ndef create_data_with_total_size(total_size=100, train_ratio=0.8):\n x, y = make_moons(n_samples=total_size, noise=0.2, random_state=1)\n\n df = DataFrame(dict(x=x[:,0], y=x[:,1], label=y))\n colors = {0:'red', 1:'blue'}\n fig, ax = pyplot.subplots()\n grouped = df.groupby('label')\n for key, group in grouped:\n group.plot(ax=ax, kind='scatter', x='x', y='y', label=key, color=colors[key])\n pyplot.show()\n\n split_index = int(x.shape[0]*train_ratio)\n train_x, test_x = x[:split_index], x[split_index:]\n train_y, test_y = y[:split_index], y[split_index:]\n\n return (train_x, train_y), (test_x, test_y)\n\n\n# + [markdown] id=\"0lCTH8uo9WUL\"\n# # 데이터 개수와 오버피팅\n\n# + id=\"07z5viJd66Cp\"\ndef show_overfitting(total_data_count, train_ratio):\n\n (train_x, train_y), (test_x, test_y) = create_data_with_total_size(total_data_count, train_ratio)\n\n model = Sequential()\n model.add(Dense(500, input_shape=(2,), activation='relu'))\n model.add(Dense(1, activation='sigmoid'))\n model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])\n\n history = model.fit(train_x, train_y, validation_data=(test_x, test_y), epochs=2000, verbose=0, batch_size=64)\n\n pyplot.plot(history.history['loss'], label='train_loss')\n pyplot.plot(history.history['val_loss'], label='test_loss')\n pyplot.legend()\n pyplot.show()\n\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"t0IERG7566Ef\" outputId=\"c30fe3d7-c2a9-4da5-f512-d0a8e0eb7f05\"\n# data 100개\nshow_overfitting(total_data_count=100, train_ratio=0.3)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 530} id=\"KL8rUEas66IN\" outputId=\"a1f8caa7-75c1-4ee9-9fc7-9ff56f743558\"\n# data 200개\nshow_overfitting(total_data_count=200, train_ratio=0.3)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 528} id=\"cD4sUfBH66Kj\" outputId=\"695c42ba-144f-4fee-b8c7-e4bbf13c2a45\"\n# data 500개\nshow_overfitting(total_data_count=500, train_ratio=0.3)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"q0yRp3pT66MZ\" outputId=\"0275bd5d-242f-4207-8543-968543ae2914\"\n# data 1000개\nshow_overfitting(total_data_count=1000, train_ratio=0.3)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"GrF6zCkk66OC\" outputId=\"0653a31e-b068-493e-faea-7f99ed23a6d6\"\n# data 2000개\nshow_overfitting(total_data_count=2000, train_ratio=0.3)\n\n# + [markdown] id=\"07H_Yi7iXGJ1\"\n# 전체적으로 보았을 때 데이터 개수를 늘릴수록 오버피팅이 줄어드는 것을 확인할 수 있음\n\n# + [markdown] id=\"xRg83v3uXCqV\"\n# # 데이터 비율과 오버피팅\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"GPRv0BZsYII4\" outputId=\"7328797b-ab15-406d-c5ee-cdf00e2c7f89\"\n# 비율 0.3\nshow_overfitting(total_data_count=1000, train_ratio=0.3)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"pXmws5HQYIGa\" outputId=\"35d1cc34-7caa-4813-da2d-6d4ab8c9da6e\"\n# 비율 0.4\nshow_overfitting(total_data_count=1000, train_ratio=0.4)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"DhwnYDEAYIDu\" outputId=\"354c5b14-ee31-4df3-a54e-682bf7bf00cc\"\n# 비율 0.5\nshow_overfitting(total_data_count=1000, train_ratio=0.5)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"WtXOH55ZYIA1\" outputId=\"bd506a24-4505-46f8-f8d1-5b551f9cbc3c\"\n# 비율 0.6\nshow_overfitting(total_data_count=1000, train_ratio=0.6)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"DtO9Q3Q4YH9P\" outputId=\"e66cd144-1f10-4b86-8ddc-43944639a672\"\n# 비율 0.7\nshow_overfitting(total_data_count=1000, train_ratio=0.7)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 527} id=\"SMEQb8qrYH6K\" outputId=\"54b0b979-ce3e-461b-fd13-ff1c67a91c35\"\n# 비율 0.8\nshow_overfitting(total_data_count=1000, train_ratio=0.8)\n\n# + [markdown] id=\"V-tVWHpWYcln\"\n# 학습 데이터 비율을 높일수록 오버피팅이 줄어드는 것을 확인할 수 있음\n","repo_name":"yunsongyi/Deep-Learning","sub_path":"10_Overfitting.ipynb","file_name":"10_Overfitting.ipynb","file_ext":"py","file_size_in_byte":4410,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"1875025300","text":"# ### 주사위 게임\n# ### 1에서부터 6까지의 눈을 가진 3개의 주사위를 던져서 다음과 같은 규칙에 따라 상금을 받는 게\n# ### 임이 있다.\n# ### 규칙(1) 같은 눈이 3개가 나오면 10,000원+(같은 눈)*1,000원의 상금을 받게 된다.\n# ### 규칙(2) 같은 눈이 2개만 나오는 경우에는 1,000원+(같은 눈)*100원의 상금을 받게 된다.\n# ### 규칙(3) 모두 다른 눈이 나오는 경우에는 (그 중 가장 큰 눈)*100원의 상금을 받게 된다.\n# ### 예를 들어, 3개의 눈 3, 3, 6이 주어지면 상금은 1,000+3*100으로 계산되어 1,300원을 받게 된\n# ### 다. 또 3개의 눈이 2, 2, 2로 주어지면 10,000+2*1,000 으로 계산되어 12,000원을 받게 된다.\n# ### 3개의 눈이 6, 2, 5로 주어지면 그 중 가장 큰 값이 6이므로 6*100으로 계산되어 600원을 상금\n# ### 으로 받게 된다.\n# ### N 명이 주사위 게임에 참여하였을 때, 가장 많은 상금을 받은 사람의 상금을 출력하는 프로그램\n# ### 을 작성하시오\n# ### ▣ 입력설명\n# ### 첫째 줄에는 참여하는 사람 수 N(2<=N<=1,000)이 주어지고 그 다음 줄부터 N개의 줄에 사람\n# ### 들이 주사위를 던진 3개의 눈이 빈칸을 사이에 두고 각각 주어진다.\n# ### ▣ 출력설명\n# ### 첫째 줄에 가장 많은 상금을 받은 사람의 상금을 출력한다.\n# ### ▣ 입력예제 1\n# ### 3\n# ### 3 3 6\n# ### 2 2 2\n# ### 6 2 5\n# ### ▣ 출력예제 1\n# ### 12000\n#\n\nN = int(input())\n\n\ndef how_much(dice):\n temp = set(dice)\n a,b,c = dice\n if len(temp) ==1:\n return 10000 + 1000*a\n elif a==b and a !=c:\n return 1000 + 100*a\n elif a==c and a !=b:\n return 1000 + 100*b\n elif b==c and b !=a:\n return 1000 + 100*b\n else:\n return max(dice)*100\n\n\nanswer = -1\nfor _ in range(N):\n dice = [int(x) for x in input().split(' ')]\n temp = how_much(dice)\n if answer < temp:\n answer = temp\n\nprint(answer)\n","repo_name":"TrendSeminar/DailyStudy","sub_path":"coding_test/코드 구현 능력 기르기/주사위 게임.ipynb","file_name":"주사위 게임.ipynb","file_ext":"py","file_size_in_byte":2020,"program_lang":"python","lang":"ko","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"10195883946","text":"# +\nimport matplotlib.pyplot as plt\nimport numpy as np\n\nimport matplotlib.image as mpimg\n\n\n# +\nm = [[0,0,0.5,0,5],\n [0,0,-1,0.7,5],\n [0,-2.8,0,0,5.5]\n ]\n\nplt.imshow(m)\n# -\n\n\n\n\n\nm.describe()\n\nnp.sin(np.array((0., 30., 45., 60., 90.)) * np.pi / 180. )\n\nx = np.sin(np.array((0., 30., 45., 60., 90.)) * np.pi / 180. ) + np.sin(np.array((0., 30., 45., 60., 90.)) * np.pi / 90. )\nx\n\nplt.imshow(x)\n\n# +\n\nimg=mpimg.imread('img2.png')\nimg\n# -\n\n\n\nplt.imshow(img)\n\n\n\nlum_img = img[:,:,0]\nplt.imshow(lum_img)\n\n\n\nplt.imshow(lum_img, cmap=\"hot\")\n\n\n\n\nimgplot = plt.imshow(lum_img)\nplt.colorbar()\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n","repo_name":"NitinSharma2017/BasicDS","sub_path":"ImgShow Example.ipynb","file_name":"ImgShow Example.ipynb","file_ext":"py","file_size_in_byte":620,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"9396760345","text":"# # Curiefense performance report\n# ## Set measurement folder\n\n# +\nimport os\nimport json\nimport glob\nimport statistics\n\nimport pandas as pd\nfrom collections import defaultdict\nimport os\nimport matplotlib.pyplot as plt\n\nDEFAULT_RESULTS_DIR = \"./results\"\nRESULTS_DIR = os.environ.get(\"RESULTS_DIR\", DEFAULT_RESULTS_DIR).rstrip(\"/\")\nprint(f\"Performance measurements will be read from {RESULTS_DIR}\")\n# -\n\n# ## Sample data\n# ### Sample fortio measurement\n\nj = json.load(open(os.path.join(RESULTS_DIR, \"with_cf/fortio-30-50-250.json\")))\nj\n\n# ### Sample jaeger measurement\n# Two *spans* are present for each query: one for service `istio-ingressgateway`, one for service `ratings.bookinfo`. Related traces have the same `traceID` attribute. The span for `ratings.bookinfo` has a non-empty `references` attribute, and is smaller than the span for `istio-ingressgateway`.\n\nj = json.load(open(os.path.join(RESULTS_DIR, \"with_cf/jaeger-30-50-250.json\")))\nj\n\n\n# ## Create a pandas dataframe from jaeger & fortio outputs\n\ndef folder2pd(folder):\n res = defaultdict(dict)\n for f in glob.glob(os.path.join(RESULTS_DIR,folder,\"fortio-*.json\")):\n tname = \"-\".join(f.split(\"-\")[-3:])\n d = res[tname]\n j = json.load(open(f))\n d[\"Connections\"] = j[\"NumThreads\"]\n d[\"RequestedQPS\"] = int(j[\"RequestedQPS\"])\n dh = j[\"DurationHistogram\"]\n d[\"SizeMin\"] = j[\"Sizes\"][\"Min\"]\n d[\"SizeMax\"] = j[\"Sizes\"][\"Max\"]\n d[\"ActualQPS\"] = j[\"ActualQPS\"]\n pct = dh[\"Percentiles\"]\n d[\"P50\"] = [x[\"Value\"] for x in pct if x[\"Percentile\"] == 50][0]\n d[\"P75\"] = [x[\"Value\"] for x in pct if x[\"Percentile\"] == 75][0]\n d[\"P90\"] = [x[\"Value\"] for x in pct if x[\"Percentile\"] == 90][0]\n d[\"P99\"] = [x[\"Value\"] for x in pct if x[\"Percentile\"] == 99][0]\n d[\"P99.9\"] = [x[\"Value\"] for x in pct if x[\"Percentile\"] == 99.9][0]\n d[\"TimeMin\"] = dh[\"Min\"]\n d[\"TimeMax\"] = dh[\"Max\"]\n d[\"TimeAvg\"] = dh[\"Avg\"]\n d[\"TimeStdDev\"] = dh[\"StdDev\"]\n for f in glob.glob(os.path.join(RESULTS_DIR,folder,\"jaeger-*.json\")):\n tname = \"-\".join(f.split(\"-\")[-3:])\n d = res[tname]\n j = json.load(open(f))\n trace_data = defaultdict(lambda: [None, None])\n for span in j[\"data\"]:\n for s in span[\"spans\"]:\n trace_id = s[\"traceID\"]\n duration = s[\"duration\"]\n if len(s[\"references\"]) == 0:\n # istio trace\n trace_data[trace_id][0] = float(duration)/1e6\n else:\n # app trace\n trace_data[trace_id][1] = float(duration)/1e6\n trace_difference = []\n for (i, a) in trace_data.values():\n if i is None or a is None:\n continue\n trace_difference.append(i-a)\n d[\"JIstioTimeAvg\"] = statistics.fmean(trace_difference)\n d[\"JIstioTimeP50\"] = statistics.quantiles(trace_difference, n=2)[-1]\n d[\"JIstioTimeP75\"] = statistics.quantiles(trace_difference, n=4)[-1]\n d[\"JIstioTimeP90\"] = statistics.quantiles(trace_difference, n=10)[-1]\n d[\"JIstioTimeP99\"] = statistics.quantiles(trace_difference, n=100)[-1]\n d[\"JIstioTimeMin\"] = min(trace_difference)\n d[\"JIstioTimeMax\"] = max(trace_difference)\n return pd.DataFrame(res.values())\n\n\nfolders = [\"with_cf\", \"without_cf\"]\ndata = {f: folder2pd(f) for f in folders}\ndata[\"without_cf\"]\n\n\n# ## Draw QPS vs time percentile\n# As measured from fortio\n\n# +\ndef qps_vs(cnx, col, args, ylim=None):\n lbls, vals = list(zip(*args.items()))\n for m in vals:\n t = (m[(m[\"Connections\"]==cnx)]).sort_values(by=[\"ActualQPS\"]) \n p = plt.plot(t[\"ActualQPS\"],t[col])\n plt.legend(lbls)\n plt.xlabel(\"Actual QPS\")\n plt.ylabel(f\"{col} (s)\")\n if ylim:\n plt.ylim(ylim)\n plt.title(f\"connections {cnx} -- ActualQPS vs {col}\")\n\ndef big_plot(*args, **kargs):\n fig = plt.figure(figsize=(20,20), dpi=72)\n qps_vs(*args, **kargs)\n\n\n# -\n\nqps_vs(250, \"P90\", data, ylim=(0,1))\n\nbig_plot(500, \"P90\", data)\n\n\ndef plot_all(args, all_cols=[\"P50\",\"P90\",\"P99\"], ylim=None):\n fig = plt.figure(figsize=(16,8), dpi= 100)\n plt.subplots_adjust(top=2)\n all_cnx = [10,70,125,250]\n \n i = 0\n for cnx in all_cnx:\n for col in all_cols:\n i += 1\n fig.add_subplot(len(all_cnx),len(all_cols),i)\n qps_vs(cnx,col,args, ylim=ylim)\n\n\nplot_all(data,ylim=(0,8))\n\n# ## Draw QPS vs latency introduced by istio\n# As measured from jaeger\n\nplot_all(data, all_cols=[\"JIstioTimeP50\", \"JIstioTimeP90\", \"JIstioTimeP99\"])\n\n\n","repo_name":"magnologan/curiefense","sub_path":"e2e/latency/Curiefense performance report.ipynb","file_name":"Curiefense performance report.ipynb","file_ext":"py","file_size_in_byte":4631,"program_lang":"python","lang":"en","doc_type":"code","dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"15981334273","text":"# # Equations of Motion for a Dubins Vehicle\n\nfrom sympy import symbols, cos, sin\nfrom sympy.physics.mechanics import dynamicsymbols, ReferenceFrame, Point, Particle, KanesMethod\n\n# +\nx, y, th, ux, uy, uth = dynamicsymbols('x y theta ux uy u_theta') # generalized coordinates and velocities\nxdot, ydot, thdot, uxdot, uydot, uthdot = dynamicsymbols('x y theta ux uy u_theta', 1) # derivatives of gen. coor. and vel.\nm = symbols('m') # mass of the particle\n\n# position, velocity, and acceleration commands\nth_com = dynamicsymbols('pc')\nthdot_com = dynamicsymbols('pc', 1)\nthddot_com = dynamicsymbols('pc', 2)\n\nbf_speed = dynamicsymbols('sp')\n# f = dynamicsymbols('f')\n\nN = ReferenceFrame('N')\nTF = N.orientnew('TF', 'Axis', [th, N.z]) # frame for direction of travel\nTF.set_ang_vel(N, uth * N.z) # cable 1 rotates at ang vel. of u2 about N.z\nP = Point('P') # point defining the particle\n\n# P.set_vel(N,ux*N.x)\nP.set_vel(N, (ux**2+uy**2)**(1/2)*TF.x)\n\nkd = [xdot - ux, ydot - uy, thdot - uth] # kinematic differential equation\n\n# Define constraints\nconf_const = [th - th_com] # configuration constraints\nvel_const = [bf_speed*cos(th)-ux, bf_speed*sin(th)-uy, uth - thdot_com] # velocity constraints\nacc_const = [uxdot - thddot_com] # acceleration constraints\n\npa = Particle('pa',P,m)\n# FL = [(P,f*N.x)] # force list\nBL = [pa] # body list\n\n# -\n\n# KM = KanesMethod(N, q_ind=[x], u_ind=[ux], kd_eqs=kd)\n# KM = KanesMethod(N, q_ind=[x, y], u_ind=[ux, uy], kd_eqs=kd, q_dependent=[th], configuration_constraints=conf_const, u_dependent=[uth], velocity_constraints=vel_const, acceleration_constraints=acc_const)\nKM = KanesMethod(N, q_ind=[x, y], kd_eqs=kd, q_dependent=[th], configuration_constraints=conf_const, u_dependent=[ux, uy, uth], velocity_constraints=vel_const)\n(fr, frstar) = KM.kanes_equations(BL)\n\nKM.rhs()\n\n\n","repo_name":"gge0866/Gerald-E-Personal","sub_path":"EOMs/.ipynb_checkpoints/DubinsVehicle_EOM-checkpoint.ipynb","file_name":"DubinsVehicle_EOM-checkpoint.ipynb","file_ext":"py","file_size_in_byte":1813,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"31113681793","text":"# %%time\nimport cv2\nimport os\npath = './daum_img/' # 다음 썸네일\nfile_list = os.listdir(path)\nfile_list.remove('.DS_Store')\nfile_list.sort()\nlen(file_list)\nerror = []\ncontour_list_daum = []\nfor file in file_list:\n# print(file)\n try:\n img = cv2.imread(f'{path}/{file}')\n imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)\n contour = cv2.Canny(img, 50, 100) # 100, 200은 변경 가능한 인자 값\n contour_list_daum.append(contour)\n# plt.imshow(contour, cmap='gray') # binary\n# plt.xticks([]); plt.yticks([])\n# plt.savefig(f'./contour_naver/{file}', dpi=200)\n except:\n continue\n\ncontour_daum = np.array(contour_list_daum)\n\nnp.save('./contour_vector/contour_daum', contour_daum)\n\nloaded_contour_d = np.load('./contour_vector/contour_daum.npy', allow_pickle=True)\nloaded_contour_d.shape\n\nfile[:5]\n\n\n\n# %%time\nimport cv2\nimport os\npath = './naver_img/' # 네이버 썸네일\nfile_list = os.listdir(path)\nfile_list.remove('.DS_Store')\nfile_list.sort()\nlen(file_list)\nerror = []\ncontour_list_naver = [] # 네이버 이미지 벡터\nfor file in file_list:\n# print(file)\n try:\n img = cv2.imread(f'{path}/{file}')\n imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)\n contour = cv2.Canny(img, 50, 100) # 100, 200은 변경 가능한 인자 값\n contour_list_naver.append(contour)\n# print(contour)\n# plt.imshow(contour, cmap='gray') # binary\n# plt.xticks([]); plt.yticks([])\n# plt.savefig(f'./contour_naver/{file}', dpi=200)\n except:\n continue\n\ncontour_naver = np.array(contour_list_naver)\n\nnp.save('./contour_vector/contour_naver', contour_naver)\n\nloaded_contour_n = np.load('./contour_vector/contour_naver.npy', allow_pickle=True)\nloaded_contour_n.shape\n\nplt.imshow(loaded_contour_n[26062])\n\nwebtoon = pd.read_csv('./total_img_info.csv')\nwebtoon\n\nwebtoon.describe()\n\nlen(webtoon['genre'].unique())\n\npd.read_csv('./webtoon_info.csv')\n\n\n","repo_name":"yeejun007/machine_learning_project","sub_path":"test/naver & daum_img_Pictorial.ipynb","file_name":"naver & daum_img_Pictorial.ipynb","file_ext":"py","file_size_in_byte":1969,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"35438744494","text":"# %matplotlib inline\nfrom matplotlib import style\nstyle.use('fivethirtyeight')\nimport matplotlib.pyplot as plt\n\nimport numpy as np\nimport pandas as pd\n\nimport datetime as dt\n\n# # Reflect Tables into SQLAlchemy ORM\n\n# Python SQL toolkit and Object Relational Mapper\nimport sqlalchemy\nfrom sqlalchemy.ext.automap import automap_base\nfrom sqlalchemy.orm import Session\nfrom sqlalchemy import create_engine, func\n\nengine = create_engine(\"sqlite:///Resources/hawaii.sqlite\")\n\n# reflect an existing database into a new model\nBase = automap_base()\n# reflect the tables\nBase.prepare(engine, reflect=True)\n\n\n# We can view all of the classes that automap found\nBase.classes.keys()\n\n# Save references to each table\nMeasurement = Base.classes.measurement\nStation = Base.classes.station\n\n# Create our session (link) from Python to the DB\nsession = Session(engine)\n\n# # Exploratory Climate Analysis\n\n# Design a query to retrieve the last 12 months of precipitation data and plot the results\nlast_date = session.query(Measurement.date).order_by(Measurement.date.desc()).first()\nprint(last_date)\n\n\n# Calculate the date 1 year ago from the last data point in the database\nlast_year = dt.date(2017, 8, 23) - dt.timedelta(days=366)\nprint(last_year)\n\nrain = session.query(Measurement.date, Measurement.prcp).\\\n filter(Measurement.date > last_year).\\\n order_by(Measurement.date).all()\n\n# Perform a query to retrieve the data and precipitation scores\nrain_df = pd.DataFrame(rain)\nrain_df.head()\n\n# Sort the dataframe by date\nrain_df.set_index('date').head()\n\n# +\n# Use Pandas Plotting with Matplotlib to plot the data\n\nrain_df.plot('date', 'prcp')\nplt.xlabel(\"Date\")\nplt.ylabel(\"Rain in Inches\")\nplt.title(\"Precipitation Analysis (8/23/16 to 8/22/17)\")\nplt.legend([\"Precipitation\"])\nplt.xticks(rotation=90)\nplt.savefig('Images/Precipitation_Analysis.png')\nplt.show()\n# -\n\n# Use Pandas to calcualte the summary statistics for the precipitation data\nrain_df.describe()\n\n# Design a query to show how many stations are available in this dataset?\nlocations = session.query(Measurement).group_by(Measurement.station).count()\nprint(\"There are {} stations.\".format(locations))\n\n# +\n# What are the most active stations? (i.e. what stations have the most rows)?\n# List the stations and the counts in descending order.\nactive_station = session.query(Measurement.station, func.count(Measurement.tobs)).group_by(Measurement.station).\\\n order_by(func.count(Measurement.tobs).desc()).all()\n\nbusiest = active_station[0][0] \nprint(\"The busiest Station was\",busiest,\"with\",active_station[0][1],\"weather observations.\")\nprint()\nprint(\"Here are all of the Stations (in descending order) with their number of observations:\")\nfor station, count in active_station:\n print(\"Station\",station,\"had\",count, \"weather observations.\")\n# -\n\n# Using the station id from the previous query, calculate the lowest temperature recorded, \n# highest temperature recorded, and average temperature of the most active station?\nmost_active_station = 'USC00519281'\nsummary_temps = session.query(func.min(Measurement.tobs), func.max(Measurement.tobs), func.avg(Measurement.tobs)).\\\n filter(Measurement.station == most_active_station).all()\nprint(f\"Lowest Temperature: {summary_temps[0][0]} Fahrenheit\")\nprint(f\"Highest Temperature: {summary_temps[0][1]} Fahrenheit\")\nprint(f\"Average Temperature: {round(summary_temps[0][2], 2)} Fahrenheit\")\n\n# Choose the station with the highest number of temperature observations.\n# Query the last 12 months of temperature observation data for this station and plot the results as a histogram\ntemp_obs = session.query(Measurement.tobs).\\\n filter(Measurement.station==most_active_station).\\\n filter(Measurement.date >= last_year).\\\n order_by(Measurement.date.desc()).all()\ntemp_obs_df = pd.DataFrame(temp_obs)\ntemp_obs_df.head()\n\nplt.hist(temp_obs_df['tobs'], bins=12)\nplt.xlabel(\"Temperature\")\nplt.ylabel(\"Frequency\")\nplt.title(\"Station Analysis (8/23/16 to 8/22/17) for Station \" + most_active_station)\nplt.legend([\"tobs\"])\nplt.savefig('Images/Station_Analysis.png')\nplt.show()\n\n\n# ## Bonus Challenge Assignment\n\n# +\n# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d' \n# and return the minimum, average, and maximum temperatures for that range of dates\ndef calc_temps(start_date, end_date):\n \"\"\"TMIN, TAVG, and TMAX for a list of dates.\n \n Args:\n start_date (string): A date string in the format %Y-%m-%d\n end_date (string): A date string in the format %Y-%m-%d\n \n Returns:\n TMIN, TAVE, and TMAX\n \"\"\"\n \n return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\\\n filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()\n\n# function usage example\nprint(calc_temps('2012-02-28', '2012-03-05'))\n# -\n\n# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax \n# for your trip using the previous year's data for those same dates.\ntemps = calc_temps('2011-02-28', '2011-03-05')\ntemps\n\n# +\n# Plot the results from your previous query as a bar chart. \n# Use \"Trip Avg Temp\" as your Title\n# Use the average temperature for the y value\n# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)\nlower = temps[0][1] - temps[0][0]\nupper = temps[0][2] - temps[0][1]\n\nplt.figure(figsize=(3,6))\nplt.bar(0, temps[0][1], yerr=[upper-lower], color='salmon', alpha=0.5)\nplt.title('Trip Avg Temp')\nplt.xticks([])\nplt.ylabel('Temp (F)')\nplt.ylim(60)\nplt.savefig('Images/Trip_Avg_Temp.png')\nplt.show()\n\n# -\n\n# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.\n# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation\nsel=[Station.station,Station.name,Station.latitude,Station.longitude,Station.elevation,func.sum(Measurement.prcp)]\nresults=session.query(*sel).filter(Measurement.station==Station.station).filter(Measurement.date>='2011-02-28').filter(Measurement.date<='2011-03-05')\\\n.group_by(Station.name).order_by(func.sum(Measurement.prcp).desc()).all()\nprint(results)\n\n\n# +\n# Create a query that will calculate the daily normals \n# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)\n\ndef daily_normals(date):\n \"\"\"Daily Normals.\n \n Args:\n date (str): A date string in the format '%m-%d'\n \n Returns:\n A list of tuples containing the daily normals, tmin, tavg, and tmax\n \n \"\"\"\n \n sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]\n return session.query(*sel).filter(func.strftime(\"%m-%d\", Measurement.date) == date).all()\n \ndaily_normals(\"01-01\")\n\n# +\n# calculate the daily normals for your trip\n# push each tuple of calculations into a list called `normals`\n\n# Set the start and end date of the trip\n\n# Use the start and end date to create a range of dates\n\n# Stip off the year and save a list of %m-%d strings\n\n# Loop through the list of %m-%d strings and calculate the normals for each date\n\n# -\n\ntrip_dates = [\"01-01\", \"01-02\", \"01-03\", \"01-04\", \"01-05\", \"01-06\", \"01-07\"]\n\n# +\nnormals = []\n\nfor trip_date in trip_dates:\n day = {}\n day['Date'] = f\"2018-{trip_date}\"\n daily_norm = daily_normals(trip_date)\n day['Min'] = daily_norm[0][0]\n day['Avg'] = daily_norm[0][1]\n day['Max'] = daily_norm[0][2]\n normals.append(day)\n \nnormals\n\n# +\n#load into a data frame\ndaily_norms_df = pd.DataFrame(normals)\n\n#set index\ndaily_norms_df.set_index('Date', inplace = True)\n\n#display\ndaily_norms_df\n\n# +\n# Plot the daily normals as an area plot with `stacked=False`\nfig, ax = plt.subplots(figsize = (13, 10))\ndaily_norms_df.plot.area(ax = ax, stacked = False)\n\n#set labels\nax.set_xlabel('Date')\nax.set_ylabel('Temperature (Fahrenheit)')\nax.set_title('Daily Normal Temperatures for Honolulu, HI')\n\n#set ticks\ndates = [f\"2018-{trip_date}\" for trip_date in trip_dates]\nax.set_xticks(np.arange(len(dates)))\nax.set_xticklabels(dates, rotation = 45)\n\n#save figure\nplt.savefig('Images/dailynormarea.png')\n\n#show\nplt.grid()\nplt.tight_layout()\nplt.show()\n# -\n\n\n","repo_name":"Monalipatel/sqlalchemy-challenge","sub_path":"climate_starter.ipynb","file_name":"climate_starter.ipynb","file_ext":"py","file_size_in_byte":8316,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"16063971075","text":"# + colab={\"base_uri\": \"https://localhost:8080/\"} executionInfo={\"elapsed\": 34381, \"status\": \"ok\", \"timestamp\": 1630746637606, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"WgTDpAoJlqd6\" outputId=\"2d38c485-0e19-4480-decd-a58d517f21e8\"\nfrom google.colab import drive\ndrive.mount('/content/drive')\n\n# + id=\"qMoDPcDhycHJ\"\nimport pandas as pd\nimport numpy as np \nimport gc\n\n\n# + id=\"FMQDH4a9uyHd\"\ndef summary(df, pred=None):\n obs = df.shape[0]\n Types = df.dtypes\n Counts = df.apply(lambda x: x.count())\n Min = df.min()\n Max = df.max()\n Uniques = df.apply(lambda x: x.unique().shape[0])\n Nulls = df.apply(lambda x: x.isnull().sum())\n print('Data shape:', df.shape)\n\n if pred is None:\n cols = ['Types', 'Counts', 'Uniques', 'Nulls', 'Min', 'Max']\n str = pd.concat([Types, Counts, Uniques, Nulls, Min, Max], axis = 1, sort=True)\n\n str.columns = cols\n print('___________________________\\nData Types:')\n print(str.Types.value_counts())\n print('___________________________')\n display(str.sort_values(by='Uniques', ascending=False))\n\n\n\n# + id=\"HRdI6HlHjT8o\"\n#Function used for calculate aggregation statistics (mean, max, min, count) for numeraical variables\n\ndef agg_numeric2(df, group_var, df_name):\n \"\"\"Aggregates the numeric values in a dataframe. This can\n be used to create features for each instance of the grouping variable.\n \n Parameters\n --------\n df (dataframe): \n the dataframe to calculate the statistics on\n group_var (string): \n the variable by which to group df\n df_name (string): \n the variable used to rename the columns\n \n Return\n --------\n agg (dataframe): \n a dataframe with the statistics aggregated for \n all numeric columns. Each instance of the grouping variable will have \n the statistics (mean, min, max, sum; currently supported) calculated. \n The columns are also renamed to keep track of features created.\n \n \"\"\"\n # Remove id variables other than grouping variable\n for col in df:\n if col != group_var and 'SK_ID' in col:\n df = df.drop(columns = col)\n \n group_ids = df[group_var]\n numeric_df = df.select_dtypes('number')\n numeric_df[group_var] = group_ids\n\n # Group by the specified variable and calculate the statistics\n agg = numeric_df.groupby(group_var).agg(['count', 'mean', 'max', 'min', 'sum']).reset_index()\n\n # Need to create new column names\n columns = [group_var]\n\n # Iterate through the variables names\n for var in agg.columns.levels[0]:\n # Skip the grouping variable\n if var != group_var:\n # Iterate through the stat names\n for stat in agg.columns.levels[1][:-1]:\n # Make a new column name for the variable and stat\n columns.append('%s_%s_%s' % (df_name, var, stat))\n\n agg.columns = columns\n return agg\n\n\n\n# + id=\"Z44G-VtFbsL6\"\ndef aggregate_client(df, group_vars, df_names):\n \"\"\"Aggregate a dataframe with data at the loan level \n at the client level\n \n Args:\n df (dataframe): data at the loan level\n group_vars (list of two strings): grouping variables for the loan \n and then the client (example ['SK_ID_PREV', 'SK_ID_CURR'])\n names (list of two strings): names to call the resulting columns\n (example ['cash', 'client'])\n \n Returns:\n df_client (dataframe): aggregated numeric stats at the client level. \n Each client will have a single row with all the numeric data aggregated\n \"\"\"\n \n # Aggregate the numeric columns\n df_agg = agg_numeric(df, parent_var = group_vars[0], df_name = df_names[0])\n \n # If there are categorical variables\n if any(df.dtypes == 'category'):\n \n # Count the categorical columns\n df_counts = agg_categorical(df, parent_var = group_vars[0], df_name = df_names[0])\n\n # Merge the numeric and categorical\n df_by_loan = df_counts.merge(df_agg, on = group_vars[0], how = 'outer')\n\n gc.enable()\n del df_agg, df_counts\n gc.collect()\n\n # Merge to get the client id in dataframe\n df_by_loan = df_by_loan.merge(df[[group_vars[0], group_vars[1]]], on = group_vars[0], how = 'left')\n\n # Remove the loan id\n df_by_loan = df_by_loan.drop(columns = [group_vars[0]])\n\n # Aggregate numeric stats by column\n df_by_client = agg_numeric(df_by_loan, parent_var = group_vars[1], df_name = df_names[1])\n\n \n # No categorical variables\n else:\n # Merge to get the client id in dataframe\n df_by_loan = df_agg.merge(df[[group_vars[0], group_vars[1]]], on = group_vars[0], how = 'left')\n \n gc.enable()\n del df_agg\n gc.collect()\n \n # Remove the loan id\n df_by_loan = df_by_loan.drop(columns = [group_vars[0]])\n \n # Aggregate numeric stats by column\n df_by_client = agg_numeric(df_by_loan, parent_var = group_vars[1], df_name = df_names[1])\n \n # Memory management\n gc.enable()\n del df, df_by_loan\n gc.collect()\n\n return df_by_client\n\n\n# + id=\"9sIi9Ktr-Zc-\"\ndef missing_values_table(df):\n mis_val = df.isnull().sum()\n mis_val_percent = 100 * df.isnull().sum() / len(df)\n mis_val_table = pd.concat([mis_val, mis_val_percent], axis=1)\n mis_val_table_ren_columns = mis_val_table.rename(\n columns = {0 : 'Missing Values', 1 : '% of Total Values'})\n mis_val_table_ren_columns = mis_val_table_ren_columns[\n mis_val_table_ren_columns.iloc[:,1] != 0].sort_values(\n '% of Total Values', ascending=False).round(1)\n \n print (\"Your selected dataframe has \" + str(df.shape[1]) + \" columns.\\n\" \n \"There are \" + str(mis_val_table_ren_columns.shape[0]) +\n \" columns that have missing values.\")\n \n return mis_val_table_ren_columns\n\n\n# + id=\"NbF-as04-c6I\"\n# function for diminishing memory used. (optional) \nimport sys\n\ndef convert_types(df, print_info = False):\n \n original_memory = df.memory_usage().sum()\n \n # Iterate through each column\n for c in df:\n \n # Convert ids and booleans to integers\n if ('SK_ID' in c):\n df[c] = df[c].fillna(0).astype(np.int32)\n \n # Convert objects to category\n elif (df[c].dtype == 'object') and (df[c].nunique() < df.shape[0]):\n df[c] = df[c].astype('category')\n \n # Booleans mapped to integers\n elif list(df[c].unique()) == [1, 0]:\n df[c] = df[c].astype(bool)\n \n # Float64 to float32\n elif df[c].dtype == float:\n df[c] = df[c].astype(np.float32)\n \n # Int64 to int32\n elif df[c].dtype == int:\n df[c] = df[c].astype(np.int32)\n \n new_memory = df.memory_usage().sum()\n \n if print_info:\n print(f'Original Memory Usage: {round(original_memory / 1e9, 2)} gb.')\n print(f'New Memory Usage: {round(new_memory / 1e9, 2)} gb.')\n \n return df\n\n\n# + id=\"nOfj3a37cn7T\"\ndef agg_numeric(df, parent_var, df_name):\n \"\"\"\n Groups and aggregates the numeric values in a child dataframe\n by the parent variable.\n \n Parameters\n --------\n df (dataframe): \n the child dataframe to calculate the statistics on\n parent_var (string): \n the parent variable used for grouping and aggregating\n df_name (string): \n the variable used to rename the columns\n \n Return\n --------\n agg (dataframe): \n a dataframe with the statistics aggregated by the `parent_var` for \n all numeric columns. Each observation of the parent variable will have \n one row in the dataframe with the parent variable as the index. \n The columns are also renamed using the `df_name`. Columns with all duplicate\n values are removed. \n \n \"\"\"\n \n # Remove id variables other than grouping variable\n for col in df:\n if col != parent_var and 'SK_ID' in col:\n df = df.drop(columns = col)\n \n # Only want the numeric variables\n parent_ids = df[parent_var].copy()\n numeric_df = df.select_dtypes('number').copy()\n numeric_df[parent_var] = parent_ids\n\n # Group by the specified variable and calculate the statistics\n agg = numeric_df.groupby(parent_var).agg(['count', 'mean', 'max', 'min', 'sum'])\n\n # Need to create new column names\n columns = []\n\n # Iterate through the variables names\n for var in agg.columns.levels[0]:\n if var != parent_var:\n # Iterate through the stat names\n for stat in agg.columns.levels[1]:\n # Make a new column name for the variable and stat\n columns.append('%s_%s_%s' % (df_name, var, stat))\n \n agg.columns = columns\n \n # Remove the columns with all redundant values\n _, idx = np.unique(agg, axis = 1, return_index=True)\n agg = agg.iloc[:, idx]\n \n return agg\n\n\n# + [markdown] id=\"efovAgw1pm0T\"\n# # Installments Payments\n\n# + [markdown] id=\"KXOLhnAUj78S\"\n# Installments Payments data include payment history for previous loans at Home Credit. \n# There is one row for every made payment and one row for every missed payment.\n#\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} executionInfo={\"elapsed\": 15833, \"status\": \"ok\", \"timestamp\": 1630746719006, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"3LwlY7EGmD0F\" outputId=\"1f44f628-7be1-4e75-ff3f-e53963992271\"\ndfinst = pd.read_csv('C:\\\\Users\\\\kbc\\\\Downloads\\\\home-credit-default-risk\\\\installments_payments.csv')\nprint(dfinst.shape)\n\n# + [markdown] id=\"Eemnrv1EkLYL\"\n# ## Exploration of Dataframe\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 421} executionInfo={\"elapsed\": 2809, \"status\": \"ok\", \"timestamp\": 1630747333424, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"KrdByrFlazZ7\" outputId=\"693fbfb1-6cd5-4aa6-8fb4-b56ad69b2304\"\nsummary(dfinst)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 300} executionInfo={\"elapsed\": 4445, \"status\": \"ok\", \"timestamp\": 1630747339211, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"-Ka06xgkazqM\" outputId=\"0dafafb2-f312-4ad2-88b4-7ee2e3739642\"\ndfinst.describe().T\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} executionInfo={\"elapsed\": 396, \"status\": \"ok\", \"timestamp\": 1630747339586, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"oReOgeNqzDbC\" outputId=\"472e2973-c8a5-415f-950a-53e29c0564b7\"\ndfinst.columns\n\n# + [markdown] id=\"YOEglfDKkbwa\"\n# ## Missing Values\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 146} executionInfo={\"elapsed\": 799, \"status\": \"ok\", \"timestamp\": 1630747343276, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"dXEGsMhyahyS\" outputId=\"a0ef4304-bb5f-4202-a365-a340c6d3bd80\"\n# No need to drop any data or column as the missing values percentages are very small.\nmissing_values_table(dfinst)\n# -\n\n#We can see amount of missing values in bar grafic\nimport missingno as msno\nmsno.bar(dfinst);\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 538} executionInfo={\"elapsed\": 2030, \"status\": \"ok\", \"timestamp\": 1630094260171, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"iC-cCCmjxBTO\" outputId=\"67dc932a-bab3-429f-da12-1428fb446f60\"\nsummary(installments_by_client)\n\n# + [markdown] id=\"gHRXqtb5ki09\"\n# ## Feature Generation\n# -\n\n# We used a function from Will Koehrsen`s code to aggregate the variables.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 444} executionInfo={\"elapsed\": 7347, \"status\": \"ok\", \"timestamp\": 1630747401506, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"FZVeelgU-XjN\" outputId=\"f9240b6d-5afb-4a56-d67f-23b575b725f6\"\ndfinstagg = agg_numeric2(dfinst, group_var = 'SK_ID_CURR', df_name = 'instpay')\n\ndfinstagg\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 286} executionInfo={\"elapsed\": 73161, \"status\": \"ok\", \"timestamp\": 1630747499315, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"aFEKCSkvbA0O\" outputId=\"7fd860f2-20d9-455a-92e0-3a0e2972cd91\"\ninstallments_by_client = aggregate_client(dfinst, group_vars = ['SK_ID_PREV', 'SK_ID_CURR'], df_names = ['installments', 'client'])\ninstallments_by_client.head()\n# -\n\n# According to the missing value results, there is nothing to do about missing results in installments paymenst data.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 641} executionInfo={\"elapsed\": 679, \"status\": \"ok\", \"timestamp\": 1630094294861, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"Nrg94oAi1Tak\" outputId=\"0cb5bc11-3610-4691-b7ed-2aa262b49475\"\nmissing_values_table(installments_by_client)\n# -\n\n# Ensure there is no object variable.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} executionInfo={\"elapsed\": 227, \"status\": \"ok\", \"timestamp\": 1630747741529, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"v4k7Mo6Q1Klg\" outputId=\"f6515474-d871-4da9-cfe2-ef7a16c80cfe\"\ninstallments_by_client.select_dtypes('object').apply(pd.Series.nunique, axis = 0)\n\n# + [markdown] id=\"WIF41cwolCFW\"\n# ## Converting data types \n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} executionInfo={\"elapsed\": 8135, \"status\": \"ok\", \"timestamp\": 1630747757519, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"4YL3XAwtqw3Y\" outputId=\"3e3e8a14-36a8-4dae-aa74-f833a82f718b\"\ndfinstagg = convert_types(installments_by_client, print_info=True)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} executionInfo={\"elapsed\": 235, \"status\": \"ok\", \"timestamp\": 1630249926374, \"user\": {\"displayName\": \"abdullah\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14GiHOvvcSx7L5oa6zZYPPlUZmJtOU1F4pPzqlNmALvY=s64\", \"userId\": \"04818964644513229474\"}, \"user_tz\": -120} id=\"r15_wLxjP25e\" outputId=\"3ddf640a-c3c9-443d-be17-36118825b4c7\"\ndfinstagg.dtypes.value_counts()\n\n# + [markdown] id=\"MlH2reUulMYJ\"\n# ## Saving new file\n\n# + id=\"oPMzZ7B6y6-X\"\ndfinstagg.to_csv('/content/drive/MyDrive/home-credit-default-risk/instpay_ready.csv')\n","repo_name":"abdullahkbc/Home-Credit-Default-Risk","sub_path":"installment_payments.ipynb","file_name":"installment_payments.ipynb","file_ext":"py","file_size_in_byte":15813,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"10385435015","text":"import torchvision\nfrom fastai import *\nfrom fastai.vision import *\nfrom skimage.io import imread, imshow\n\n\n# +\nclass Colors(Dataset):\n def __init__(self, seed, size):\n red=np.stack([np.ones((224,224)), np.zeros((224,224)), np.zeros((224,224))], axis=0).astype(np.float32)\n blue=np.stack([np.zeros((224,224)), np.zeros((224,224)), np.ones((224,224))], axis=0).astype(np.float32)\n self.cards = [red, blue]\n np.random.seed(seed)\n self.size = size\n self.list = np.random.random(size)\n\n def __getitem__(self, index):\n cat = int(self.list[index]>.5)\n return self.cards[cat], cat\n \n def __len__(self):\n return self.size\n\ndltrain = DataLoader(Colors(0, 500), batch_size=8)\ndlvalid = DataLoader(Colors(1, 50), batch_size=8)\ndb = ImageDataBunch(dltrain, dlvalid)\ndb.c = 2\n# -\n\nimshow(db.train_ds[0][0].transpose(1,2,0))\n\nlearn = create_cnn(db, arch=torchvision.models.vgg16_bn, metrics=accuracy, loss_func=F.cross_entropy)\n\nlearn.fit_one_cycle(1, .01)\n","repo_name":"simonm3/published","sub_path":"colortest.ipynb","file_name":"colortest.ipynb","file_ext":"py","file_size_in_byte":1018,"program_lang":"python","lang":"en","doc_type":"code","stars":4,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"4184438863","text":"import pandas as pd\nimport numpy as np\n\ndf = pd.read_json('../data/raw/april_movie_scores_v2.json')\n\ndf.head()\n\n# View how many rows are missing critic scores\ndf[df.Critic_score == \"\"].count()\n\n# View how many rows are missing audience scores\ndf[df.Audience_score == \"\"].count()\n\n# Determine count of missing values across dataframe\ndf.replace(\"\", np.NaN, inplace=True)\ndf.isnull().sum()\n\n# Replace missing Critic_score, Audience_score, and Rating values with NR\ndf.Critic_score.replace(np.NaN, \"NR\", inplace=True)\ndf.Audience_score.replace(np.NaN, \"NR\", inplace=True)\ndf.Rating.replace(np.NaN, \"NR\", inplace=True)\n\ndf.isnull().sum()\n\n# View first value of Details column\ndf.Details[0]\n\n# +\n# Extract values from Details column to create new columns\ndf['Year'] = df.Details.str.split(\",\").str[0].str.strip()\ndf['Title_category'] = df.Details.str.split(\",\").str[1]\ndf['Title_category'] = df.Title_category.str.strip()\ndf['Runtime'] = df.Details.str.split(\",\").str[2].str.strip()\n\n# Drop Details from df\ndf.drop('Details', axis=1, inplace=True)\ndf.head(2)\n# -\n\n# Reorder dataframe to match SQL table\ncolumns = ['Title', 'Year', 'Critic_score', 'Audience_score', 'Title_category', 'Rating', 'Runtime', 'URL', 'Poster_URL']\ndf = df[columns]\ndf.head(2)\n\n# Check count of duplicates based on Title, Year, Critic_score, Audience_score\ndf.duplicated(subset=['Title', 'Year', 'Critic_score', 'Audience_score']).value_counts()\n\n# View duplicated values based on Title, Year, Critic_score, Audience_score\ndf[df.duplicated(subset=['Title', 'Year', 'Critic_score', 'Audience_score'], keep=False)]\n\n# Drop those duplicated values, keeping the last one\ndf = df.drop_duplicates(subset=['Title', 'Year', 'Critic_score', 'Audience_score'], keep='last')\n\n# Omit Poster_URL values with .gif as these are bad URLs\ndf.loc[df.Poster_URL.str.contains('.gif'), 'Poster_URL'] = \"\"\n","repo_name":"charlesdungy/new-to-streaming-scraper","sub_path":"notebooks/rotten_tomatoes_data_clean_test_v2.ipynb","file_name":"rotten_tomatoes_data_clean_test_v2.ipynb","file_ext":"py","file_size_in_byte":1855,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"28233353678","text":"# +\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\npd.set_option('display.max_columns', None)\npd.set_option('display.max_rows', None)\n# -\n\ndf = pd.read_csv('test.csv')\nprint(df.shape)\ndf.head()\n\nsns.heatmap(df.isnull(), yticklabels = False, cbar = False, cmap= 'viridis')\n\n\n# +\ncols, per = [],[]\nfor col in df.columns:\n \n if(df[col].isnull().sum() > 0):\n print(col, 100*(df[col].isnull().sum()/len(df)).round(2),\"%\")\n cols.append(col)\n per.append(100*(df[col].isnull().sum()/len(df)).round(2))\n \nplt.figure(figsize =(8,8))\nplt.barh(range(len(cols)),per, tick_label = cols)\nplt.show()\n# -\n\ndf.drop(['Alley', 'MiscFeature','PoolQC','Fence'],axis = 1, inplace = True)\n\n# +\nnum_cols = []\nobj_cols = []\n\nfor col in df.select_dtypes(include = 'object').columns:\n if df[col].isnull().sum() > 0:\n obj_cols.append(col)\n \nfor col in df.select_dtypes(include = ['int64','float64']).columns:\n if df[col].isnull().sum() > 0:\n num_cols.append(col)\n \n\nfor col in num_cols:\n df[col].fillna(df[col].mean(),inplace = True)\n \nfor col in obj_cols:\n df[col].fillna(df[col].mode()[0], inplace = True)\n \n# -\n\ndf.shape\n\n# +\ncols, per = [],[]\nfor col in df.columns:\n \n if(df[col].isnull().sum() > 0):\n print(col, 100*(df[col].isnull().sum()/len(df)).round(2),\"%\")\n cols.append(col)\n per.append(100*(df[col].isnull().sum()/len(df)).round(2))\n \nplt.figure(figsize =(8,8))\nplt.barh(range(len(cols)),per, tick_label = cols)\nplt.show()\n# -\n\nsns.heatmap(df.isnull(),yticklabels=False,cbar=False,cmap='coolwarm')\n\n\ndf.drop('Id', axis = 1, inplace = True)\n\ndf.shape\n\ndf.to_csv('formulated_test.csv')\n\n\n\nlen(df)\n\n\n\n\n\n\n","repo_name":"aleemsiddiqui0299/Advanced-Regression","sub_path":"handling_test_data.ipynb","file_name":"handling_test_data.ipynb","file_ext":"py","file_size_in_byte":1741,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"19141766825","text":"# +\n# %load_ext autoreload\n# %autoreload 2\n# %matplotlib inline\n# %config InlineBackend.figure_format='retina'\n\nfrom FRLM import network\n\nfrom matplotlib import pyplot as plt\nimport matplotlib as mpl\nfrom network2tikz import plot\nimport pandas as pd\nimport numpy as np\nimport json\nimport compress_json\n\nnetwork_name = '25NODE'\nsolution_fn = 'results/test/25NODE_10_15_25.json.gz'\n\n# +\njson_data = compress_json.load(solution_fn)\n\nparams = json_data['params']\nsolution = json_data['solution']\n\nnet = network.Net(dataDir=f\"instance/{params['network_name']}\", dist_multiplier=params['dist_multiplier'])\n\nnode_weight_l = [[i,net.G.nodes[i]['weight']] for i in net.G.nodes if net.G.nodes[i]['weight'] > 0]\nweighted_nodes = [i for i,w in sorted(node_weight_l,key=lambda l:l[1], reverse=True)][:params['num_node']]\n\narcs = {}\nfor k,(p,l,t,f) in json_data['covered_full_paths_k'].items():\n prev_i = p[0]\n for i in p[1:]:\n arc = tuple(sorted((prev_i, i)))\n if arc not in arcs:\n arcs[arc] = f\n else:\n arcs[arc] += f\n prev_i = i\n\nlayout = {i:net.G.nodes[i]['pos'] for i in net.G.nodes}\n\nvisual_style = {}\n\nvisual_style['node_label_off'] = False\nvisual_style['node_opacity'] = [0.5 for i in net.G.nodes]\nvisual_style['node_label'] = [net.G.nodes[i]['weight'] if i in weighted_nodes else '' for i in net.G.nodes]\n# visual_style['node_label_color'] = ['white' for i in net.G.nodes]\nvisual_style['node_size'] = [0.4 if i in weighted_nodes else 0.2 for i in net.G.nodes]\nvisual_style['node_color'] = []\nvisual_style['node_shape'] = []\nfor i in net.G.nodes:\n if i in solution[0]:\n if solution[1][solution[0].index(i)] == 1:\n shape = 'diamond'\n visual_style['node_size'][i] = 0.5\n else:\n shape = 'circle'\n color = 'red'\n else:\n color = 'white!50!gray'\n shape = 'circle'\n visual_style['node_color'].append(color)\n visual_style['node_shape'].append(shape)\n\nvisual_style['edge_width'] = []\nvisual_style['edge_color'] = []\nfor (i1,i2) in net.G.edges:\n if i1 > i2:\n continue\n if (i1,i2) in arcs:\n visual_style['edge_width'].append(max(arcs[(i1,i2)]/250, 1))\n visual_style['edge_color'].append('red')\n else:\n visual_style['edge_width'].append(1)\n visual_style['edge_color'].append('black')\n\nvisual_style['edge_opacity'] = 0.8\nvisual_style['edge_label'] = [net.G.edges[i]['weight'] for i in net.G.edges]\nvisual_style['edge_label_color'] = ['black' for i in net.G.edges]\nvisual_style['edge_label_position'] = ['below' for i in net.G.edges]\n\nvisual_style['canvas'] = (18,18)\nvisual_style['margin'] = 1\n\nvisual_style['layout'] = layout\n\nplot(net.G, 'solution.pdf', **visual_style)\n# plot(net.G, 'figure/TEXAS_solution.csv', **visual_style)\n","repo_name":"optboy/FRLMwCT","sub_path":"read_result.ipynb","file_name":"read_result.ipynb","file_ext":"py","file_size_in_byte":2792,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"11407263711","text":"# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"4SQPiTRsrppA\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674110851066, \"user_tz\": -540, \"elapsed\": 2641, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"de02872b-3821-48ab-a1d8-92e92d02db6b\"\nfrom google.colab import drive\ndrive.mount('/content/drive')\n\n# + id=\"i97gsg6urwy3\"\n# 필요한 모듈 import\n\nimport pandas as pd\nimport numpy as np\nfrom numpy import dot\nfrom numpy.linalg import norm\nimport collections, operator\n\n# + id=\"DpxraS_1vrz9\"\n# 모든 경우의 수에 대한 원 핫 인코딩 DataFrame 만들기\n'''\n이 DataFrame으로 User가 웹에서 만들어낼 수 있는 모든 경우의 수에 ���한 결과를 저장해놓고,\n바로바로 사용자가 추천 결과를 확인할 수 있도록 하였음.\n\n이 Dataframe의 각 line마다 인플루언서 100명과의 코사인 유사도를 계산한 후,\n코사인 유사도가 가장 높은 상위 3명을 옆 column으로 추가할 것임.\n'''\n\ndef user_encoding(style_num, tone_num, season_num):\n '''\n input 값\n - style_num: 스타일 카테고리 개수\n - tone_num: 톤 카테고리 개수\n - season_num: 계절 카테고리 개수\n '''\n\n # 각 카테고리의 개수만큼의 길이를 가진 dummy list를 생성\n style_list = [0] * style_num\n tone_list = [0] * tone_num\n season_list = [0] * season_num\n\n # 각 카테고리의 개수만큼의 길이를 가진 column 이름을 만든 후, list에 저장\n style_column = [('style_' + str(i)) for i in range(len(style_list))]\n tone_column = [('tone_' + str(i)) for i in range(len(tone_list))]\n season_column = [('season_' + str(i)) for i in range(len(season_list))]\n\n # 위에서 만든 column들을 가지는 빈 Dataframe을 생성\n df = pd.DataFrame(columns = (style_column + tone_column + season_column))\n\n # 모든 경우의 수를 가지는 원 핫 인코딩 Dataframe을 생성\n # 각 카테고리마다 하나의 태그는 1을, 나머지는 0을 가지게 됨. \n num = 0\n for i in range(len(style_list)):\n style_list = [0] * style_num\n style_list[i] = 1\n for j in range(len(tone_list)):\n tone_list = [0] * tone_num\n tone_list[j] = 1\n for k in range(len(season_list)):\n season_list[k] = 1\n df.loc[num] = (style_list + tone_list + season_list)\n num += 1\n season_list = [0] * season_num\n\n # 최종적으로 만들어진 Dataframe 반환\n return df\n\n\n# + [markdown] id=\"rdINqaGrV_IN\"\n# # 남자 인플루언서 추천 알고리즘\n\n# + id=\"kGk9FLyj9FYV\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674110966386, \"user_tz\": -540, \"elapsed\": 369, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 424} outputId=\"ecaf3ff0-71fc-4b4b-aa2c-bfb0f455774c\"\n# 남자 인플루언서 Database 불러오기\nman_data = pd.read_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/final_man_influ.csv', index_col = 0)\nman_data\n\n# + id=\"U_J5alfYB_kI\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674110971806, \"user_tz\": -540, \"elapsed\": 278, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} colab={\"base_uri\": \"https://localhost:8080/\"} outputId=\"a86a5024-6e19-42e2-c506-d21d33e52ee3\"\n# 남자 인플루언서들의 인스타그램 아이디를 추출\nman_ids = list(set(list(man_data['influ_id'])))\nman_ids\n\n# + [markdown] id=\"8X0XCB-qBstp\"\n# ## Man / Style\n\n# + id=\"5F6SQsPC_i1z\"\n# 남자의 스타일에 해당하는 스타일 태그를 list type으로 저장\nstyle_tag_list = ['americancasual','casual','dandy','formal','loose','street']\n\n# 위의 list를 column으로 가지는 빈 Dataframe 생성\ndf_man_style = pd.DataFrame(columns = style_tag_list)\n\n# Classifier를 통해 각 인플루언서의 피드 이미지마다의 스타일을 추출 -> 각 스타일의 빈도 수를 전체 빈도 수로 나누어 Dataframe에 추가\nfor i in range(len(man_ids)):\n a = man_data[man_data['influ_id'] == man_ids[i]]\n result = dict(a['style'].value_counts())\n df_man_style.loc[i] = result\n df_man_style.loc[i] /= len(a)\n# NaN은 인플루언서가 해당 스타일을 가진 피드 이미지가 없다는 것\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} id=\"UZa9CLY0A4VL\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674110998049, \"user_tz\": -540, \"elapsed\": 266, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"d812818c-9ab2-4ae8-e938-7714c8e59274\"\ndf_man_style\n\n# + [markdown] id=\"39aNKrvyBx6C\"\n# ## Man / Tone\n\n# + id=\"m6P0DhQg02r-\"\n# 남자의 톤에 해당하는 톤 태그를 list type으로 저장\ntone_tag_list = ['mono','neutral','pastel','toneon']\n\n# 위의 list를 column으로 가지는 빈 Dataframe 생성\ndf_man_tone = pd.DataFrame(columns = tone_tag_list)\n\n# Classifier를 통해 각 인플루언서의 피드 이미지마다의 톤을 추출 -> 각 톤의 빈도 수를 전체 빈도 수로 나누어 Dataframe에 추가\nfor i in range(len(man_ids)):\n a = man_data[man_data['influ_id'] == man_ids[i]]\n result = dict(a['tone'].value_counts())\n df_man_tone.loc[i] = result\n df_man_tone.loc[i] /= len(a)\n# NaN은 인플루언서가 해당 톤을 가진 피드 이미지가 없다는 것\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} id=\"5e-n7jwXCF86\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674111034845, \"user_tz\": -540, \"elapsed\": 8, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"1f62f516-df76-4400-e00d-2314f0a634c2\"\ndf_man_tone\n\n# + [markdown] id=\"oUrOTfLzCIM-\"\n# ## Man / Season\n\n# + id=\"QGwbVZlXCPmr\"\n# 계절감에 해당하는 계절 태그를 list type으로 저장 ('autumn'은 봄과 가을 모두에 해당하는 태그)\nseason_tag_list = ['summer','autumn','winter']\n\n# 위의 list를 column으로 가지는 빈 Dataframe 생성\ndf_man_season = pd.DataFrame(columns = season_tag_list)\n\n# Classifier를 통해 각 인플루언서의 피드 이미지마다의 계절감을 추출 -> 각 계절감의 빈도 수를 전체 빈도 수로 나누어 Dataframe에 추가\nfor i in range(len(man_ids)):\n a = man_data[man_data['influ_id'] == man_ids[i]]\n result = dict(a['season'].value_counts())\n df_man_season.loc[i] = result\n df_man_season.loc[i] /= len(a)\n# NaN은 인플루언서가 해당 스타일을 가진 계절감 이미지가 없다는 것\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} id=\"r3IPm3HECP0m\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674111158145, \"user_tz\": -540, \"elapsed\": 299, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"ef43e46f-e0d2-4d0c-efda-ee0b9bfde782\"\ndf_man_season\n\n# + [markdown] id=\"-Zqd1SagCfOV\"\n# ## Man DataFrame Concat\n\n# + id=\"XJvaVGqDChZ2\"\n# 3개의 카테고리에 해당하는 Dataframe을 병합\ndf_man_total = pd.concat([df_man_style, df_man_tone, df_man_season], axis = 1)\n\n# NaN 값을 0으로 치환\ndf_man_total = df_man_total.fillna(0)\n\n# 각 row에 인스타그램 id 매칭\ndf_man_total['ids'] = man_ids\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} id=\"223eGfUyCrsh\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674111172696, \"user_tz\": -540, \"elapsed\": 296, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"5e1471d4-ca52-48e7-fa54-c1caa9891c71\"\ndf_man_total\n\n\n# + [markdown] id=\"igZVbJtYFfJx\"\n# ## Man 정답표 구성하기\n\n# + id=\"VNch71gY5Pig\"\n# 코사인 유사도를 계산하는 함수\ndef cos_sim(a, b):\n return dot(a, b)/(norm(a)*norm(b))\n\n\n# + id=\"kQMQgH8jGFFo\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674111217060, \"user_tz\": -540, \"elapsed\": 365, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 424} outputId=\"1cb55ee6-0d72-4df1-8111-358c2147e1a6\"\n# user(사용자로부터 입력받는 input 데이터) -> 더미변수를 이용하여 모든 경우의 수를 데이터프레임으로 생성\nusers = user_encoding(6, 4, 3)\n'''\n남성의 경우,\n\nstyle: 6개\ntone: 4개\nseason: 3개\n'''\nusers\n\n# + id=\"SEzqFyz8Fu9T\"\n# 상위 3명의 인플루언서 정보를 저장할 빈 Dataframe 생성\n# 인플루언서 인스타그램 id와 코사인 유사도 계산 값을 저장\ni_result_df = pd.DataFrame(columns = ['first_influ','first_score','second_influ','second_score','third_influ','third_score'])\n\nfor i in range(len(users)):\n i_result_dict = {}\n i_line = []\n user = users.iloc[i]\n for j in range(len(df_man_total)):\n target = df_man_total.iloc[j].to_list()[:-1]\n result_cos = cos_sim(user, target)\n i_result_dict[j] = result_cos\n i_result_dict = sorted(i_result_dict.items(), key = operator.itemgetter(1), reverse = True)\n for rank in range(3):\n i_line.append(df_man_total['ids'].iloc[i_result_dict[rank][0]])\n i_line.append(round(i_result_dict[rank][1], 2))\n i_result_df.loc[i] = i_line\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 424} id=\"sQ0lcGOiMIi2\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674111288066, \"user_tz\": -540, \"elapsed\": 6, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"0dae8724-12a5-45ed-acc1-d2062e2e4f74\"\ni_result_df\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 488} id=\"JcSTX2qrdyh7\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674112833720, \"user_tz\": -540, \"elapsed\": 265, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"0a6a1774-6e3c-42af-b85d-d52d42451e48\"\n# 모든 경우의 수를 담은 Dataframe과 결과 Dataframe을 매칭하여 하나의 Dataframe으로 저장 후\n# csv 파일로 내보내서 최종적인 Database 구축\nman_answer = pd.concat([users, i_result_df], axis = 1)\nman_answer.to_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/final_man_dataset.csv')\nman_answer\n\n# + [markdown] id=\"lKiagEQXfJ-v\"\n# # 여자 인플루언서 추천 알고리즘\n#\n# 위의 남자 인플루언서 추천 알고리즘과 동일한 방식과 코드를 사용하므로 추가적으로 자세한 주석을 생략함.\n\n# + executionInfo={\"status\": \"ok\", \"timestamp\": 1674111315374, \"user_tz\": -540, \"elapsed\": 378, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 424} outputId=\"642b7a00-d051-4db4-f3d7-a63b407f08e1\" id=\"MYD-DmpgfJ-v\"\nwoman_data = pd.read_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/final_woman_influ.csv', index_col = 0)\nwoman_data\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"tjszxgFSfJ-v\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674111317017, \"user_tz\": -540, \"elapsed\": 295, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"dba5599d-c49c-4b9f-f16f-daba9de0650c\"\n# 여자 인플루언서들의 인스타그램 아이디를 추출\nwoman_ids = list(set(list(woman_data['influ_id'])))\nwoman_ids\n\n# + [markdown] id=\"FVmteo2QfJ-w\"\n# ## Woman / Style\n\n# + id=\"e6TIkTMOfJ-w\"\n# 여자의 스타일에 해당하는 스타일 태그를 list type으로 저장\nstyle_tag_list = ['casual','romantic','girlish','chic','street','formal','loose']\n\n# 위의 list를 column으로 가지는 빈 Dataframe 생성\ndf_woman_style = pd.DataFrame(columns = style_tag_list)\n\n# Classifier를 통해 각 인플루언서의 피드 이미지마다의 스타일을 추출 -> 각 스타일의 빈도 수를 전체 빈도 수로 나누어 Dataframe에 추가\nfor i in range(len(woman_ids)):\n a = woman_data[woman_data['influ_id'] == woman_ids[i]]\n result = dict(a['style'].value_counts())\n df_woman_style.loc[i] = result\n df_woman_style.loc[i] /= len(a)\n# NaN은 인플루언서가 해당 스타일을 가진 피드 이미지가 없다는 것\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} executionInfo={\"status\": \"ok\", \"timestamp\": 1674111596822, \"user_tz\": -540, \"elapsed\": 10, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"336ff104-4ec9-48f7-9233-1210707aeed0\" id=\"9RrlkmptfJ-w\"\ndf_woman_style\n\n# + [markdown] id=\"oMSGfPSAfJ-w\"\n# ## Woman / Tone\n\n# + id=\"qaAnoAhXfJ-w\"\n# 여자의 톤에 해당하는 톤 태그를 list type으로 저장\ntone_tag_list = ['mono','neutral','pastel','toneon']\n\n# 위의 list를 column으로 가지는 빈 Dataframe 생성\ndf_woman_tone = pd.DataFrame(columns = tone_tag_list)\n\n# Classifier를 통해 각 인플루언서의 피드 이미지마다의 톤을 추출 -> 각 톤의 빈도 수를 전체 빈도 수로 나누어 Dataframe에 추가\nfor i in range(len(woman_ids)):\n a = woman_data[woman_data['influ_id'] == woman_ids[i]]\n result = dict(a['tone'].value_counts())\n df_woman_tone.loc[i] = result\n df_woman_tone.loc[i] /= len(a)\n# NaN은 인플루언서가 해당 톤을 가진 피드 이미지가 없다는 것\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} executionInfo={\"status\": \"ok\", \"timestamp\": 1674111643049, \"user_tz\": -540, \"elapsed\": 7, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"8ac10194-7fa4-457b-80df-fb5e3e99ab44\" id=\"fRWmOVAefJ-w\"\ndf_woman_tone\n\n# + [markdown] id=\"SGJy2yV6fJ-w\"\n# ## Woman / Season\n\n# + id=\"lFEtfut0fJ-w\"\n# 계절감에 해당하는 계절 태그를 list type으로 저장 ('autumn'은 봄과 가을 모두에 해당하는 태그)\nseason_tag_list = ['summer','autumn','winter']\n\n# 위의 list를 column으로 가지는 빈 Dataframe 생성\ndf_woman_season = pd.DataFrame(columns = season_tag_list)\n\n# Classifier를 통해 각 인플루언서의 피드 이미지마다의 계절감을 추출 -> 각 계절감의 빈도 수를 전체 빈도 수로 나누어 Dataframe에 추가\nfor i in range(len(woman_ids)):\n a = woman_data[woman_data['influ_id'] == woman_ids[i]]\n result = dict(a['season'].value_counts())\n df_woman_season.loc[i] = result\n df_woman_season.loc[i] /= len(a)\n# NaN은 인플루언서가 해당 스타일을 가진 계절감 이미지가 없다는 것\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} executionInfo={\"status\": \"ok\", \"timestamp\": 1674111647424, \"user_tz\": -540, \"elapsed\": 8, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"39399977-227a-4a24-f15a-648220ec9fc0\" id=\"VJY03G8vfJ-x\"\ndf_woman_season\n\n# + [markdown] id=\"8AbPFrpBfJ-x\"\n# ## Woman DataFrame Concat\n\n# + id=\"2Eo3VFb_fJ-x\"\n# 3개의 카테고리에 해당하는 Dataframe을 병합\ndf_woman_total = pd.concat([df_woman_style, df_woman_tone, df_woman_season], axis = 1)\n\n# NaN 값을 0으로 치환\ndf_woman_total = df_woman_total.fillna(0)\n\n# 각 row에 인스타그램 id 매칭\ndf_woman_total['ids'] = woman_ids\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000} executionInfo={\"status\": \"ok\", \"timestamp\": 1674111660140, \"user_tz\": -540, \"elapsed\": 528, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"9cf19a6e-194a-46c4-a704-8cdcc2736fbb\" id=\"JTDXBreHfJ-x\"\ndf_woman_total\n\n\n# + [markdown] id=\"mv8zo7cyfJ-x\"\n# ## Woman 정답표 구성하기\n\n# + id=\"wubSgO3PfJ-x\"\n# 코사인 유사도를 계산하는 함수\ndef cos_sim(a, b):\n return dot(a, b)/(norm(a)*norm(b))\n\n\n# + executionInfo={\"status\": \"ok\", \"timestamp\": 1674111730414, \"user_tz\": -540, \"elapsed\": 260, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 424} outputId=\"18fe4bf4-09b4-4bfa-9513-769654473fb8\" id=\"-Cxt9wfefJ-x\"\n# user(사용자로부터 입력받는 input 데이터) -> 더미변수를 이용하여 모든 경우의 수를 데이터프레임으로 생성\nusers_woman = user_encoding(7, 4, 3)\n'''\n여자의 경우,\n\nstyle: 7개\ntone: 4개\nseason: 3개\n'''\nusers_woman\n\n# + id=\"455HMrCTfJ-x\"\ni_result_df_w = pd.DataFrame(columns = ['first_influ','first_score','second_influ','second_score','third_influ','third_score'])\n\nfor i in range(len(users_woman)):\n i_result_dict = {}\n i_line = []\n user_w = users_woman.iloc[i]\n for j in range(len(df_woman_total)):\n target = df_woman_total.iloc[j].to_list()[:-1]\n result_cos = cos_sim(user_w, target)\n i_result_dict[j] = result_cos\n i_result_dict = sorted(i_result_dict.items(), key = operator.itemgetter(1), reverse = True)\n for rank in range(3):\n i_line.append(df_woman_total['ids'].iloc[i_result_dict[rank][0]])\n i_line.append(round(i_result_dict[rank][1], 2))\n i_result_df_w.loc[i] = i_line\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 424} executionInfo={\"status\": \"ok\", \"timestamp\": 1674111787973, \"user_tz\": -540, \"elapsed\": 278, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"be8e2ba8-7bb5-4b90-866f-0b91c4d825b4\" id=\"Q5vzlVQCfJ-x\"\ni_result_df_w\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 488} executionInfo={\"status\": \"ok\", \"timestamp\": 1674112811492, \"user_tz\": -540, \"elapsed\": 296, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"df75a2e9-1d49-4b7d-af13-34b391f33e73\" id=\"686tYIdMfJ-y\"\nwoman_answer = pd.concat([users_woman, i_result_df_w], axis = 1)\nwoman_answer.to_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/final_woman_dataset.csv')\nwoman_answer\n\n# + [markdown] id=\"k4EwmqsvjWz4\"\n# # 사용자 input 읽어서 결과 출력\n#\n# 아래의 코드는 일부 수정 후, 프론트엔드 코드와 연결할 수 있도록 'code.py'로 내보낸 파일이 존재함.\n#\n# 따라서 프론트엔드 코드 폴더와 함께 있는 code.py와 일부 코드 구성이 다를 수 있음.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 81} id=\"5fcW2Qq6jZan\" executionInfo={\"status\": \"ok\", \"timestamp\": 1674113054711, \"user_tz\": -540, \"elapsed\": 870, \"user\": {\"displayName\": \"\\uc815\\uc9c0\\ud6c8\", \"userId\": \"16483350189460332753\"}} outputId=\"2ddf2b07-8c84-49cc-fe3a-cbf20b8a072f\"\n# 웹을 통해서 사용자가 선택한 input_data.csv 파일 불러오기\ninput_data = pd.read_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/input_data.csv')\ninput_data\n\n\n# + id=\"Oe3PoJx1kTBB\"\n# 사용자의 input data를 받아서 결과를 출력하는 함수 정의\ndef result_print(input_data):\n\n input_data = input_data.loc[0].to_dict()\n\n # 남자, 여자 Database 불러오기\n man_answer = pd.read_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/final_man_dataset.csv')\n woman_answer = pd.read_csv('/content/drive/MyDrive/2023 KUIAI 해커톤/final_woman_dataset.csv')\n\n # 웹에서 전달받은 input과 기존에 만들어진 Database 간의 매칭을 시키기 위한 Dictionary 선언\n man_style_match = {\n '아메리칸 캐주얼':'style_0',\n '캐주얼':'style_1',\n '댄디':'style_2',\n '포멀':'style_3',\n '루즈핏':'style_4',\n '스트릿':'style_5'\n }\n\n woman_style_match = {\n '캐주얼':'style_0',\n '로맨틱':'style_1',\n '걸리시':'style_2',\n '시크':'style_3',\n '스트릿':'style_4',\n '포멀':'style_5',\n '루즈핏':'style_6'\n }\n\n tone_match = {\n '모노톤':'tone_0',\n '뉴트럴톤':'tone_1',\n '파스텔톤':'tone_2',\n '톤온톤':'tone_3'\n }\n\n season_match = {\n '여름':'season_0',\n '봄/가을':'season_1',\n '겨울':'season_2'\n }\n\n # input_data csv 파일의 'Gender' 정보를 이용하여 남자, 여자를 나누어 코드가 진행되도록 구현함\n # 남자 function과 여자 function의 경우 구성이 거의 동일함\n if input_data['Gender'] == '남자':\n result_func_man(input_data, man_answer, man_style_match, tone_match, season_match)\n else: # 여자의 경우\n result_func_woman(input_data, woman_answer, woman_style_match, tone_match, season_match)\n\n\n# 남자\ndef result_func_man(input_data, man_answer, man_style_match, tone_match, season_match):\n '''\n - input_data: 사용자가 입력한 input_data\n - man_answer: 남자 인플루언서 Database\n - man_style_match, tone_match, season_match: Database 매칭을 위한 Dictionary\n '''\n\n # 사용자가 입력한 'style', 'tone', 'season' 값을 각각의 변수에 저장\n user_style = input_data['Style']\n user_tone = input_data['Tone']\n user_season = input_data['Season']\n\n # 사용자가 입력한 태그의 조건과 일치하는 정보를 Database에서 가져오기\n # 사용자가 입력한 태그의 조건을 'condition' 변수에 저장 -> 입력한 조건을 1로 찾음\n condition = (man_answer[man_style_match[user_style]] == 1) & (man_answer[tone_match[user_tone]] == 1) & (man_answer[season_match[user_season]] == 1)\n result = man_answer[condition]\n columns = ['first_influ','second_influ','third_influ']\n\n send = result[columns].iloc[0].to_list()\n\n # 최종 상위 3명의 인플루언서 id를 .txt 파일로 구성하여 내보내기\n f = open(\"/content/drive/MyDrive/2023 KUIAI 해커톤/recommendation_result.txt\", 'w')\n for i in range(len(send)):\n data = str(i+1) + \"번째 맞춤 인플루언서: \" + send[i] + '\\n'\n f.write(data)\n f.close()\n\n\n# 여자\ndef result_func_woman(input_data, woman_answer, woman_style_match, tone_match, season_match):\n '''\n - input_data: 사용자가 입력한 input_data\n - woman_answer: 여자 인플루언서 Database\n - woman_style_match, tone_match, season_match: Database 매칭을 위한 Dictionary\n '''\n user_style = input_data['Style']\n user_tone = input_data['Tone']\n user_season = input_data['Season']\n\n condition = (woman_answer[woman_style_match[user_style]] == 1) & (woman_answer[tone_match[user_tone]] == 1) & (woman_answer[season_match[user_season]] == 1)\n result = woman_answer[condition]\n columns = ['first_influ','second_influ','third_influ']\n\n send = result[columns].iloc[0].to_list()\n\n f = open(\"/content/drive/MyDrive/2023 KUIAI 해커톤/recommendation_result.txt\", 'w')\n for i in range(len(send)):\n data = str(i+1) + \"번째 맞춤 인플루언서: \" + send[i] + '\\n'\n f.write(data)\n f.close()\n\n\n# + id=\"A87vqECB5TJ-\"\n# 최종적으로 실행해야하는 코드 -> 이 코드 하나를 실행시키면 자동적으로 3명의 인플루언서 정보가 담긴 텍스트 파일을 저장함\nresult_print(input_data)\n","repo_name":"starnam11/2023_influ_recommendation_hackerton","sub_path":"I_추천 알고리즘.ipynb","file_name":"I_추천 알고리즘.ipynb","file_ext":"py","file_size_in_byte":22325,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"9532247631","text":"import cv2\nimport matplotlib.pyplot as plt\nimport os\n\npath='Fnt'\nfolders=os.listdir(path)\nfor i in range(0, len(folders)):\n loopPath=path+'/'+folders[i]\n images=os.listdir(loopPath)\n for j in range(0, len(images)):\n imgPath=loopPath+'/'+images[j]\n im=cv2.imread(imgPath)\n im=cv2.resize(im,(28,28))\n im=cv2.bitwise_not(im)\n cv2.imwrite(imgPath,im)\nprint('done')\n\npath='data/train'\nfolders=os.listdir(path)\nfor i in range(0, len(folders)):\n loopPath=path+'/'+folders[i]\n images=os.listdir(loopPath)\n for j in range(0, len(images)):\n imgPath=loopPath+'/'+images[j]\n destPath=loopPath+'/'+folders[i]+'_'+str(j)+'.png'\n os.rename(imgPath,destPath)\nprint('done')\n\nimport keras\nfrom keras import backend as k\nfrom keras.models import Sequential\nfrom keras.layers import Activation\nfrom keras.layers.core import Dense, Flatten, Dropout\nfrom keras.optimizers import Adam\nfrom keras.preprocessing.image import ImageDataGenerator\nfrom keras.layers.normalization import BatchNormalization\nfrom keras.layers.convolutional import *\n\ntrainPath='data/train'\nvalidationPath='data/val'\nclassList=os.listdir(trainPath)\n\nprint(classList)\n\ntrainBatches=ImageDataGenerator().flow_from_directory(trainPath, target_size=(28,28), batch_size=1, class_mode='categorical')\nvalidationBatches=ImageDataGenerator().flow_from_directory(validationPath, target_size=(28,28), batch_size=1, class_mode='categorical')\n\nprint(validationBatches.class_indices)\n\nmodel=Sequential([\n Conv2D(32,(24,24), input_shape=(28,28,3),activation='relu', padding='same'),\n MaxPooling2D(pool_size=(2,2)),\n Dropout(0.5),\n Flatten(),\n Dense(128, activation='relu'),\n Dense(36,activation='softmax'),\n])\n\nmodel.compile(Adam(lr=0.0001), loss='categorical_crossentropy', metrics=['accuracy'])\n\nhistory=model.fit_generator(trainBatches, steps_per_epoch=4945,validation_data=validationBatches, validation_steps=216,epochs=20)\n\nmodel.save('myanpr.h5')\nprint('SAVED')\n\nprint(history.history.keys())\n\n# +\nplt.plot(history.history['accuracy'])\nplt.plot(history.history['val_accuracy'])\nplt.title('Model Accuracy')\nplt.ylabel('Accuracy')\nplt.xlabel('Epoch')\nplt.legend(['train','test'], loc='upper left')\nplt.show()\n\nplt.plot(history.history['loss'])\nplt.plot(history.history['val_loss'])\nplt.title('Model Loss')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['train','test'], loc='upper left')\nplt.show()\n# -\n\nfrom keras.models import load_model\npMq=load_model('Rupammyanpr.h5')\nprint(pM.summary())\n\nimport pandas as pd\nimport numpy as np\n\n\ndef extractPlate(img):\n copyImg=img.copy()\n casVar=cv2.CascadeClassifier('indian_license_plate.xml')\n plateCoordinates=casVar.detectMultiScale(copyImg, scaleFactor=1.3, minNeighbors=7)\n print(plateCoordinates)\n for (x,y,w,h) in plateCoordinates:\n plate=copyImg[y:y+h, x:x+w, :]\n return plate\n\n\nimg=cv2.imread('img1.png')\nplt.imshow(img)\nout=extractPlate(img)\nplt.imshow(out)\n\ncv2.imwrite('outputPlate.jpg',out)\n\n\nins=cv2.imread('outputPlate.jpg')\nins=cv2.resize(ins,(333,75))\nins=cv2.cvtColor(ins,cv2.COLOR_BGR2GRAY)\ndim,img_binary=cv2.threshold(ins,200,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU) \nins=cv2.erode(img_binary,(3,3))\nins=cv2.dilate(ins,(3,3))\nplt.imshow(ins)\ncv2.imwrite('dialated.jpg',ins)\n\nplateWidth=ins.shape[0]\nplateHeight=ins.shape[1]\nprint(plateHeight, plateWidth)\n\ndimentions=plateWidth/6, plateWidth/2, plateHeight/10, 2*plateHeight/3\nprint(dimentions)\n\ncntrs,dim=cv2.findContours(ins.copy(), cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)\ncntrs=sorted(cntrs, key=cv2.contourArea, reverse=True)[:15]\ncoImg=ins.copy()\ncv2.drawContours(coImg,cntrs, -1, (233,255,255),3)\n\n\nxcx=[]\ntc=[]\nfor cntr in cntrs:\n ix, iy, iw, ih= cv2.boundingRect(cntr)\n if iw > dimentions[0] and iw < dimentions[1] and ih > dimentions[2] and ih < dimentions[3]:\n xcx.append(ix)\n chcop=np.zeros((44,24))\n char= ins[iy:iy+ih, ix: ix+iw]\n char=cv2.resize(char,(20,40))\n char=cv2.subtract(255, char)\n chcop[2:42, 2:22]=char\n tc.append(chcop)\nindices= sorted(range(len(xcx)), key=lambda k: xcx[k])\nimgResCopy=[]\nfor idx in indices:\n imgResCopy.append(tc[idx])\ntc=np.array(imgResCopy)\nfor i in range(0, len(tc)):\n name='Segment_'+str(i)+'.png'\n cv2.imwrite(name,tc[i])\n\nch='0123456789ABCDEFGHIJKLMNOPQRSTUVWXYZ'\nplt.figure(figsize=(10,6))\nfor i in range(0,len(tc)):\n path='Prediction'\n images=os.listdir(path)\n imgPath=path+'/'+images[i]\n print(imgPath)\n pre=cv2.imread(imgPath)\n pre=cv2.resize(pre, (28,28))\n pre1=pre.reshape(1,28,28,3)\n Y=pMq.predict_classes(pre1)[0]\n print(ch[Y])\n plt.subplot(3,4,i+1)\n plt.imshow(pre)\n plt.title(ch[Y])\n plt.axis('off')\n\n\n","repo_name":"rupamgangulydev/Automatic_Number_Plate_Recognition_PYTHON","sub_path":"Code.ipynb","file_name":"Code.ipynb","file_ext":"py","file_size_in_byte":4737,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"6584709804","text":"# +\nfrom sklearn.datasets import load_iris\ndata = load_iris()\nprint(dir(data)) # 查看data所具有的属性或方法\nprint(data.DESCR) # 查看数据集的简介\n\nimport pandas as pd\n#直接读到pandas的数据框中\ntest = pd.DataFrame(data=data.data, columns=data.feature_names)\n\n# -\n\ntest.head(5)\n\n\n# ### 1. 导入数据集\n#\n# 【知识点:输入和输出】\n#\n# 如何导入存在数字和文本的数据集?\n\ndef change_dtype(x):\n return float(x) \n\n\n# +\nimport numpy as np\n\n\n# 导入数字和文本数据集\ndatapath = './iris.data'\ndata = np.genfromtxt(datapath,delimiter=\",\",dtype='object',names=True,converters={0:change_dtype,1:change_dtype,2:change_dtype,3:change_dtype})\n\ndata['sepallength']\n# -\n\ndata['species']==b'Iris-setosa'\n\n# ### 2. 求出鸢尾属植物萼片长度的平均值、中位数和标准差(第1列,sepallength)\n#\n# 【知识点:统计相关】\n#\n# 如何计算numpy数组的均值,中位数,标准差?\n\n# +\n# 均值\nprint(\"the mean of sepallength:\",np.mean(data['sepallength']),)\n\n#中位数\nprint(\"the median of sepallength:\",np.median(data['sepallength']),)\n\n#标准差\nprint(\"the std of sepallength:\",np.std(data['sepallength']),)\n# -\n\n# ### 3. 创建一种标准化形式的鸢尾属植物萼片长度,其值正好介于0和1之间,这样最小值为0,最大值为1(第1列,sepallength)。\n#\n# 【知识点:统计相关】\n#\n# 如何标准化数组?\n\n# +\n# 最大最小值初始化\ns_max,s_min = np.amax(data['sepallength']), np.amin(data['sepallength'])\n\nnew_sepallength = (data['sepallength'] - s_min)/(s_max - s_min)\n\n\ndata['sepallength'] = new_sepallength\n# -\n\n# ### 4. 找到鸢尾属植物萼片长度的第5和第95百分位数(第1列,sepallength)。\n#\n# 【知识点:统计相关】\n#\n# 如何找到numpy数组的百分位数?\n\n# +\n# 5,95的分位数\n\nprint(\"the 5% and 95% pre of sepallength: \", np.percentile(data['sepallength'],[5,95]))\n# -\n\n# ### 5. 把iris_data数据集中的20个随机位置修改为np.nan值。\n#\n# 【知识点:随机抽样】\n#\n# 如何在数组中的随机位置修改值?\n\n# +\n# 随机位置修改\n\ndata[20]\n# -\n\ndata[20]=np.nan\ndata[20]\n\n# ### 6. 在iris_data的sepallength中查找缺失值的个数和位置(第1列)。\n#\n# 【知识点:逻辑函数、搜索】\n#\n# 如何���numpy数组中找到缺失值的位置?\n\n\n\n# ### 7. 筛选具有 sepallength(第1列)< 5.0 并且 petallength(第3列)> 1.5 的 iris_data行。\n#\n# 【知识点:搜索】\n#\n# 如何根据两个或多个条件筛选numpy数组?\n\ndata[(data['sepallength']<5.0 ) & (data['petallength'] >1.5)]\n\n# ### 8. 选择没有任何 nan 值的 iris_data行。\n#\n# 【知识点:逻辑函数、搜索】\n#\n# 如何从numpy数组中删除包含缺失值的行?\n\n\n\n# ### 9. 计算 iris_data 中sepalLength(第1列)和petalLength(第3列)之间的相关系数。\n#\n# 【知识点:统计相关】\n#\n# 如何计算numpy数组两列之间的相关系数?\n\n# +\nnp.corrcoef(data['sepallength'].astype(float),data['petallength'].astype(float))\n\n# 如果没有astype(float) 会出现 np.corrcoef 'float' object has no attribute 'shape' 的问题\n# -\n\n# ### 10. 找出iris_data是否有任何缺失值。\n# 【知识点:逻辑函数】\n#\n# 如何查找给定数组是否具有空值?\n\n# +\nx = data['sepallength'].astype(float)\n\nnp.isnan(x)\n# -\n\n# ### 11. 在numpy数组中将所有出现的nan替换为0。\n#\n# 【知识点:逻辑函数】\n#\n# 如何在numpy数组中用0替换所有缺失值?\n\n# +\n## 方法一,使用逻辑函数 nan_to_num\nnp.nan_to_num(data)\n\n\n# -\n\n# ### 12. 找出鸢尾属植物物种中的唯一值和唯一值出现的数量。\n#\n# 【知识点:数组操作】\n#\n# 如何在numpy数组中查找唯一值的计数?\n\n# +\ns_counts = {}\nfor s in data['species']:\n s_counts[s]=s_counts.get(s,0)+1\n\ns_counts\n# -\n\n# ### 13. 将 iris_data 的花瓣长度(第3列)以形成分类变量的形式显示。定义:Less than 3 --> ‘small’;3-5 --> ‘medium’;’>=5 --> ‘large’。\n#\n# 【知识点:统计相关】\n#\n# 如何将数字转换为分类(文本)数组?\n\n# +\n\n\n# 排序\npetallength = np.digitize(data['petallength'].astype('float'), [0,3,5,10], right=False)\n\nlabel_map = {1: 'small', 2: 'medium', 3: 'large', 4: np.nan}\n\npetallength_map = [label_map[x] for x in petallength]\n\npetallength_map \n# -\n\n# ### 14. 在 iris_data 中创建一个新列,其中 volume 是 (pi x petallength x sepallength ^ 2)/ 3。\n#\n# 【知识点:数组操作】\n#\n# 如何从numpy数组的现有列创建新列?\n\ndata['petallength']\n\n# +\n# 计算新的一列的数值\n\npetallength = data['petallength'].astype('float')\nsepallength = data['sepallength'].astype('float')\n\nnew_volume = (np.pi * petallength * (sepallength **2))/3\n\nnew_volume = new_volume.astype('object')\n# -\n\n# \nnew_data = np.column_stack((data,new_volume.T))\n\nnew_data[:4,1]\n\n# ### 15. 随机抽鸢尾属植物的种类,使得Iris-setosa的数量是Iris-versicolor和Iris-virginica数量的两倍。\n#\n# 【知识点:随机抽样】\n#\n# 如何在numpy中进行概率抽样?\n\n\n\n# ### 16. 根据 sepallength 列对数据集进行排序。\n#\n# 【知识点:排序】\n#\n# 如何按列对2D数组进行排序?\n\nsort_data = sorted(data,key = lambda x:x[0])\n\n# ### 17. 在鸢尾属植物数据集中找到最常见的花瓣长度值(第3列)。\n#\n# 【知识点:数组操作】\n#\n# 如何在numpy数组中找出出现次数最多的值?\n\npetallength = list( data['petallength'])\n\n# +\npetallength_count = {}\n\nfor p in petallength :\n petallength_count[p] = petallength_count.get(p,0)+1\n \npetallength_count = sorted(petallength_count.items(),key = lambda x:-x[1])\npetallength_count[0][0]\n\n# +\n\nvals, counts = np.unique(data['petallength'], return_counts=True)\nprint(vals,counts)\n\nnp.argmax()\n# -\n\n# ### 18. 在鸢尾花数据集的 petalwidth(第4列)中查找第一次出现的值大于1.0的位置。\n#\n# 【知识点:搜索】\n#\n# 如何找到第一次出现大于给定值的位置?\n\n# +\n\nnp.argwhere(data['petalwidth'].astype(float) > 1.0)[0]\n\n# -\n\n\n","repo_name":"StevenZhou5/CuteCodeV2","sub_path":"Numpy/task10_大作业/大作业.ipynb","file_name":"大作业.ipynb","file_ext":"py","file_size_in_byte":6108,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"26239385799","text":"import numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.linear_model import LinearRegression\nfrom sklearn import metrics\n\ndf = pd.read_csv('insurance.csv')\ndf.head()\n\ndf.shape\n\n\ndf.info()\n\ndf.isnull().sum()\n\ndf.describe()\n\n# +\n# to plot a distribution of each, density of maximin age densiy.\n# -\n\n#to make distrubtion of age (density) \nsns.set()\nplt.figure(figsize = (6,6))\nsns. distplot(df['age'])\nplt.title(\"Age Distribution\")\nplt.show()\n\nplt.figure(figsize=(6,6))\nsns.countplot(x = 'sex',data = df)\nplt.title('Sex Distribution')\nplt.show()\n\ndf['sex'].value_counts()\n\nsns.distplot(df['bmi'])\nplt.title('BMI')\nplt.show()\n\ndf['region'].value_counts()\n\ndf.replace({'sex': {'male':0 , 'female':1}},inplace = True)\n\ndf.replace({'smoker':{'yes':0, 'no':1}},inplace =True)\n\ndf.replace({'region':{'southeast':0, 'southwest':1, 'northwest' :2, 'northeast' :3 }} ,inplace = True)\n\n# slpitting indepedent and dependent features\nx = df.drop(columns = \"charges\", axis = 1)\ny = df['charges']\n\nx\n\ny\n\nx_train,x_test,y_train,y_test = train_test_split(x,y,test_size = 0.2)\n\nx_train.shape\n\nx_test.shape\n\nreg = LinearRegression()\n\nreg.fit(x_train,y_train)\n\ntraining_data_prediction = reg.predict(x_train)\n\nr2_train = metrics.r2_score(y_train, training_data_prediction)\n\nr2_train\n\ntest_data_prediction = reg.predict(x_test)\nmetrics.r2_score(y_test, test_data_prediction)\n\nsample_input_data = (30,1,22.7,0,1,0)\n\ninput_data_as_numpy = np.asarray(sample_input_data)\n\ninput_data_as_numpy\n\ninput_data_reshaped = input_data_as_numpy.reshape(1,-1)\n\ninput_data_reshaped\n\nprediction = reg.predict(input_data_reshaped)\n\nprint(\"the insurance cost is\", prediction)\n\n\n","repo_name":"Danish159/Data-Science-and-Machine-Learning-","sub_path":"Medical Insurance Cost Prediction.ipynb","file_name":"Medical Insurance Cost Prediction.ipynb","file_ext":"py","file_size_in_byte":1741,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"33080693240","text":"# # Propulsion Module Examples\n# Solving the course examples of AC701.\n\n# +\nimport model_reference as mf\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport warnings\n\nplt.style.use('seaborn')\nwarnings.filterwarnings('ignore')\n# -\n\n# ## Example 2.1: TurboJet\n\n# ### Exercise Information\n#\n# Calculate the specific thrust and specific consumption (TSFC) for a turbojet engine operating at Mach 0.85 and at 12,200 m (Pa = 18.75kPa and Ta = 216.7 K). Use the data below for efficiencies and specific heat ratios.\n#\n# | Component | Adiabatic Efficiency | Specific Heat Ratio |\n# |------------|----------------------|---------------------|\n# | diffuser | 0,97 | 1,40 |\n# | compressor | 0,85 | 1,37 |\n# | combustion | 1,00 | 1,35 |\n# | turbine | 0.90 | 1,33 |\n# | nozzle | 0,98 | 1,36 |\n#\n# Operating Parameters:\n#\n# | Pressure Ration in Compressor | 30 |\n# |------------------------------------------|-------------------|\n# | Outlet temperature in combustion chamber | 1600 K |\n# | Lower Heating Value of the Fuel | 45000 kJ/kg |\n# | Mean R | 288,3 m^2/(s^2.K) |\n# | specific heat in the combustion chamber | 1,11 kJ/(kg.K) |\n#\n#\n\n# ### Exercise Computation\n\n# +\n\nvalues_ex2_1 = {\n 'ta':216.7,\n 'pa':18750,\n 'n_d':0.97,\n 'gamma_d':1.4,\n 'n_c':0.85,\n 'gamma_c':1.37,\n 'n_b':1,\n 'gamma_b':1.35,\n 'n_t':0.9,\n 'gamma_t':1.33,\n 'n_n':0.98,\n 'gamma_n':1.36,\n 'prc':30,\n 'pc_fuel':45000,\n 'cp_fuel':1.110,\n 'r':288.3,\n 't04':1600,\n 'mach':0.85\n }\n\nturbojet = mf.TurboJet(values_ex2_1)\ntargets = ['t02','p02','t03', 'p03', 'f', 't05', 'p05', 'u_s', 'TSFC', 'u_i', 'specific_thrust']\nresult_2_1 = (\n turbojet.sumarise().\n append(turbojet.sumarise_results()).\n loc[targets]\n)\nclean_names = ['Temperatura na Saída Difusor', 'Pressão na Saída Difusor', \n'Temperatura na Saída Compressor', 'Pressão na Saída Compressor', \n'Razão combustível/ar', 'Temperatura na Saída Turbina', \n'Pressão na Saída Turbina', 'Velocidade de Saída do ar', \n'Consumo específico', 'Velocidade de voo', 'Empuxo específico']\nresult_2_1.rename(index = dict(zip(targets, clean_names)))\n# -\n\n# ## Example 2.2: TurboFan\n\n# ### Exercise Information\n#\n# Calculate the specific thrust and specific consumption (TSFC) for a turbofan engine operating at Mach 0.85 and at 12,200 m (Pa = 18.75kPa and Ta = 216.7 K). Use the data below for efficiencies and specific heat ratios.\n#\n# | Component | Adiabatic Efficiency | Specific Heat Ratio |\n# |--------------------|----------------------|---------------------|\n# | intake | 0,97 | 1,40 |\n# | fan | 0,85 | 1,40 |\n# | compressor | 0,85 | 1,37 |\n# | combustion | 1,00 | 1,35 |\n# | compressor turbine | 0.90 | 1,33 |\n# | compressor fan | 0.90 | 1,33 |\n# | Hot air nozzle | 0,98 | 1,36 |\n# | Fan nozzle | 0,98 | 1,40 |\n#\n# Operating Parameters:\n#\n# | Outlet temperature in combustion chamber | 1600 K |\n# |------------------------------------------|-------------------|\n# | Pressure Ration in Compressor | 20 |\n# | Pressure Ration in Fan | 1.5 |\n# | Air passsage ratio | 5.0 |\n# | Lower Heating Value of the Fuel | 45000 kJ/kg |\n# | Mean R | 288,3 m^2/(s^2.K) |\n# | specific heat in the combustion chamber | 1,11 kJ/(kg.K) |\n#\n#\n\n# ### Exercise Computation\n\n# +\nvalues_ex2_2 = {\n 'ta':216.7,\n 'pa':18.750,\n 'n_d':0.97,\n 'n_c':0.85,\n 'n_f':0.85,\n 'n_b':1,\n 'n_t':0.9,\n 'n_tf':0.9,\n 'n_n':0.98,\n 'n_nf':0.98,\n 'gamma_d':1.4,\n 'gamma_f':1.4,\n 'gamma_c':1.37,\n 'gamma_b':1.35,\n 'gamma_t':1.33,\n 'gamma_tf':1.33,\n 'gamma_n':1.36,\n 'gamma_nf':1.4,\n 'prc':20,\n 'prf':1.5,\n 'bypass_ratio':5,\n 'pc_fuel':45000,\n 'cp_fuel':1.110,\n 'r':288.3,\n 't04':1600,\n 'mach':0.85\n }\n\nturbofan = mf.TurboFan(values_ex2_2)\ntarget = ['t02','p02','t08','p08','t03', 'p03', 'f','tet','pet', 't05', 'p05', 'u_s', 'u_sf','u_i', 'specific_thrust', 'TSFC']\nresult_2_2 = (\n turbofan.sumarise().\n append(turbofan.sumarise_results()).\n loc[target]\n)\nresult_2_2.columns = ['turbofan']\nresult_2_2['turbojet'] = result_2_1\nresult_2_2\n# -\n\n# ## Example 2.3: TurboFan - Baseline Project Computation\n\n# ### Exercise Information\n#\n# Operating Engine Parameters:\n#\n# | Baseline Operating Condition | |\n# |------------------------------------------|-------------------|\n# | Mass Flux | 756 kg/s |\n# | Outlet temperature in combustion chamber | 1550 K |\n# | Pressure Ration in Compressor | 17.2 |\n# | Pressure Ration in Booster | 1.00 |\n# | Pressure Ration in Fan | 1.69 |\n# | Air bypass ratio | 4.69 |\n# | Pressure loss in combustion chamber | 5 % |\n#\n# Operating Condition:\n#\n# | Flight Operation Condition | |\n# |------------------------------------------|-------------------|\n# | Flight Mach | 0 |\n# | Atmospheric Pressure | 101.63 kPa |\n# | Atmospheric Temperature | 290 K |\n# | % of maximum compressor rotation | 1.00 |\n#\n#\n# Efficiency and specific heat ratio data:\n#\n# | Component | Adiabatic Efficiency | Specific Heat Ratio |\n# |--------------------|----------------------|---------------------|\n# | intake | 0,97 | 1,40 |\n# | fan | 0,93 | 1,40 |\n# | compressor | 0,90 | 1,37 |\n# | combustion | 0,9995 | 1,35 |\n# | compressor turbine | 0.95 | 1,33 |\n# | compressor fan | 0.932 | 1,33 |\n# | Hot air nozzle | 0,98 | 1,36 |\n# | Fan nozzle | 0,98 | 1,40 |\n#\n# Thermodynamic Data:\n#\n# | Thermodynamic Data | |\n# |------------------------------------------|-------------------|\n# | Lower Heating Value of the Fuel | 45000 kJ/kg |\n# | Mean R | 288,3 m^2/(s^2.K) |\n# | specific heat in the combustion chamber | 1,11 kJ/(kg.K) |\n#\n#\n\n# ### Exercise Computation\n\n# +\nvalues_ex2_3 = {\n 'ta':290,\n 'pa':101.630,\n 'n_d':0.97,\n 'n_f':0.93,\n 'n_c':0.90,\n 'n_b':0.9995,\n 'n_t':0.95,\n 'n_tf':0.932,\n 'n_n':0.98,\n 'n_nf':0.98,\n 'gamma_d':1.4,\n 'gamma_f':1.4,\n 'gamma_c':1.37,\n 'gamma_b':1.35,\n 'gamma_t':1.33,\n 'gamma_tf':1.33,\n 'gamma_n':1.36,\n 'gamma_nf':1.4,\n 'prc':17.2,\n 'prf':1.69,\n 'bypass_ratio':4.749,\n 'pc_fuel':45000,\n 'cp_fuel':1.110,\n 'r':288.3,\n 't04':1550,\n 'mach':0,\n 'mass_flow':756,\n 'pressure_loss':0.05,\n }\n\nturbofan23 = mf.TurboFan(values_ex2_3)\n\nsummarise_target = ['n1', 'thrust_total', 'TSFC', 'prf', 'prc', 't04','hot_mass_flow','bypass_ratio']\n\n\nresults = (\n turbofan23.sumarise().\n append(turbofan23.sumarise_results()).\n loc[summarise_target]\n)\n\nfor i in np.arange(0.9,0.4,-0.1):\n turbofan23.set_n2(i)\n results[i] = (\n turbofan23.sumarise().\n append(turbofan23.sumarise_results()).\n loc[summarise_target]\n )\n\nresults.loc['total_fuel_comsuntion'] = results.loc['TSFC'] * results.loc['thrust_total']\nresults.loc['pr_global'] = results.loc['prc'] * results.loc['prf']\n\nordered_target = ['n1', 'thrust_total', 'TSFC', 'prf', 'prc', 'pr_global', 't04','hot_mass_flow','total_fuel_comsuntion','bypass_ratio']\nclean_names = ['Rotação do Fan N1', 'Empuxo Total', 'TSFC','Prf', 'Prc', 'Taxa de comp. Global', 'Temp. Saída do Combustor','Vazão mássica de ar quente', 'Consumo', 'Razão de Passagem']\nresults = results.loc[ordered_target]\n\n# -\n\nresults.rename(index=dict(zip(ordered_target, clean_names)))\n\ny_labels = ['Rotação N1 [%P.P]', 'Empuxo [kN]', 'TSFC [kg/(kN.s)]', '$P_{rf}$', '$P_{rc}$', '$P_r$ Global', 'T04 [K]', '$\\dot{m}_{a_H}$ [kg/s]', 'Consumo [kg/s]','B']\ntitles = ['N1 x N2', 'Empuxo x N2', 'Consumo Específico x N2', '$P_{rf}$ x N1', '$P_{rc}$ x N2','$P_r$ Global x N2', 'T04 x N2', '$\\dot{m}_{a_H}$ x N2', 'Consumo de Combustível x N2','Razão de Passagem x N1']\nyticks = [np.arange(0,1.3,0.2), np.arange(0,301,50),np.arange(0,0.017,0.002),np.arange(0,1.9,0.2),np.arange(0,21,2),np.arange(0,36,5),np.arange(0,1801,200),np.arange(0,161,20),np.arange(0,3.1,0.5),np.arange(2,9,1)]\n\nfontsize=14\ncols = 2\nrows=5\nfig, ax = plt.subplots(nrows=rows, ncols=cols, figsize=(16,20))\nfig.subplots_adjust(hspace=.5)\naux=0\nfor i in range(0,rows):\n for j in range(0,cols):\n plt.sca(ax[i,j])\n if results.index[aux] == 'prf':\n plt.plot(results.loc['n1'], results.iloc[aux], marker='o')\n plt.axis([0.2,1.01,0,max(results.iloc[aux])*1.2])\n plt.xlabel('Rotação N1', fontsize=fontsize)\n\n elif results.index[aux] == 'bypass_ratio':\n plt.plot(results.loc['n1'], results.iloc[aux], marker='o')\n plt.axis([0.2,1.01,2,max(results.iloc[aux])*1.2])\n plt.xlabel('Rotação N1', fontsize=fontsize)\n else:\n plt.plot(results.iloc[aux], marker='o')\n plt.axis([0.4,1.01,0,max(results.iloc[aux])*1.2])\n plt.xlabel('Rotação N2', fontsize=fontsize)\n plt.ylabel(y_labels[aux], fontsize=fontsize)\n plt.title(titles[aux], fontsize=fontsize)\n plt.yticks(yticks[aux])\n aux += 1\n\n\n# ## Example 7.1: TurboProp - Calibrating Model\n\n# ### Exercise Information\n#\n# Taking the data for the PW127 engine at sea level and stationary: free turbine shaft power 2457 kW, compressor air flow 8.49 kg/s, compressor pressure ratio 15.77 and combustion chamber outlet temperature 1600K. Calibrate the engine in the performance simulation to obtain shaft power close to the reported by the manufacturer.\n#\n# Operating Engine Parameters:\n#\n# | Baseline Operating Condition | |\n# |------------------------------------------|-------------------|\n# | Mass Flux | 8.49 kg/s |\n# | Outlet temperature in combustion chamber | 1600 K |\n# | Pressure Ration in Compressor | 15.77 |\n#\n#\n# Operating Condition:\n#\n# | Flight Operation Condition | |\n# |------------------------------------------|-------------------|\n# | Flight Mach | 0 |\n# | Atmospheric Pressure | 101.30 kPa |\n# | Atmospheric Temperature | 288.15 K |\n# | % of maximum compressor rotation | 1.00 |\n#\n#\n# Efficiency and specific heat ratio data:\n#\n# | Component | Adiabatic Efficiency | Specific Heat Ratio |\n# |--------------------|----------------------|---------------------|\n# | intake | 0,85 | 1,40 |\n# | compressor | 0,75 | 1,37 |\n# | combustion | 1,00 | 1,35 |\n# | compressor turbine | 0,90 | 1,33 |\n# | free turbine | 0,90 | 1,33 |\n# | hot air nozzle | 0,98 | 1,36 |\n#\n# Thermodynamic Data:\n#\n# | Thermodynamic Data | |\n# |------------------------------------------|-------------------|\n# | Lower Heating Value of the Fuel | 45000 kJ/kg |\n# | Mean R | 288,3 m^2/(s^2.K) |\n# | specific heat in the combustion chamber | 1,11 kJ/(kg.K) |\n#\n#\n#\n#\n#\n#\n#\n\n# ### Part 1: Exercise computation\n# Calibrating the model to the manufacturer report with pressure ratio in the turbine equal to 2.87 and % of maximum rotation equals to 1.00.\n\n# +\nimport model_reference as mf\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport pandas as pd\nimport warnings\n# warnings.filterwarnings('ignore')\n\nvalues = {\n 'mass_flow': 8.49,\n 't04': 1600,\n 'prc': 15.77,\n 'u_in': 0,\n 'ta': 288.15,\n 'pa': 101.30,\n 'n_d': 0.85,\n 'gamma_d': 1.4,\n 'n_c': 0.75,\n 'gamma_c': 1.37,\n 'n_b': 1,\n 'gamma_b': 1.35,\n 'n_t': 0.9,\n 'gamma_t': 1.33,\n 'n_tl': 0.9,\n 'gamma_tl': 1.33,\n 'n_n': 0.98,\n 'gamma_n': 1.36,\n 'pc_fuel': 45000,\n 'cp_fuel': 1.110,\n 'cp_tls': 1.160,\n 'cp_tl': 1.160,\n 'r': 288.3,\n 'pr_tl': 2.87,\n 'gearbox_power_ratio': 0.98,\n 'propeller_efficiency': 0.85,\n }\n\n\ntarget = [\n 'specific_power_turbine', \n 'turbine_power', \n 'gearbox_power', \n 'specific_thrust', \n 'BSFC', \n 'EBSFC', \n 'TSFC', \n 'thrust_hot_air', \n 'thrust_propeller', \n 'fuel_consumption', \n 'thrust_total',\n 'mass_flow',\n 'aircraft_speed'\n ]\n\nlist_of_turbine_pressure_ratio = np.linspace(1.8, 3.4, 10)\nturboprop = mf.TurboProp(values)\nperformance_results = turboprop.sumarise_results()\nperformance_results = performance_results.drop(columns=[1])\n\nfor turbine_pressure_ratio in list_of_turbine_pressure_ratio:\n values[\"pr_tl\"] = turbine_pressure_ratio\n turboprop = mf.TurboProp(values)\n performance_result = turboprop.sumarise_results()\n performance_results[turbine_pressure_ratio] = performance_result\n\n\nfig1 = plt.figure()\nfig1.suptitle(\"Turbine Power\")\naxis1 = fig1.add_subplot(111)\naxis1 = performance_results.loc['turbine_power'].plot()\naxis1.set_ylabel(\"Turbine Power [kW] \")\naxis1.set_xlabel(\"Turbine Expansion Rate [ ]\")\naxis1.axhline(2457, linestyle='--', color='gray', label='given turbine power')\naxis1.axvline(2.87, linestyle='--', color='gray')\naxis1.legend()\n\nperformance_results\n# -\n\n# ### Part 1 Results:\n# We have varied the turbine expansion rate to find the turbine power that corresponds to the manufcaturer given turbine power.\n#\n# From the results above, we find the following parameters to calibrate the model:\n#\n# | Engine Parameters | |\n# |------------------------------------------|-------------------|\n# | Pressure Ratio in the Free Turbine | 2.87 |\n# | % of maximum rotation in the compressor | 1.00 |\n\n# ### Part 2: Comparing to Manufacturer Report\n#\n# The thermodynamic power of the turbine is superior to the limit power of the gearbox 2050 kW.\n# Thus, the solution is to reduce the compressor rotation.\n\n# +\nimport model_reference as mf\nimport numpy as np\nimport pandas as pd\nimport warnings\n# warnings.filterwarnings('ignore')\n\nvalues = {\n 'mass_flow': 8.49,\n 't04': 1600,\n 'prc': 15.77,\n 'u_in': 0,\n 'ta': 288.15,\n 'pa': 101.30,\n 'n_d': 0.85,\n 'gamma_d': 1.4,\n 'n_c': 0.75,\n 'gamma_c': 1.37,\n 'n_b': 1,\n 'gamma_b': 1.35,\n 'n_t': 0.9,\n 'gamma_t': 1.33,\n 'n_tl': 0.9,\n 'gamma_tl': 1.33,\n 'n_n': 0.98,\n 'gamma_n': 1.36,\n 'pc_fuel': 45000,\n 'cp_fuel': 1.110,\n 'cp_tl': 1.160,\n 'cp_tls': 1.160,\n 'r': 288.3,\n 'pr_tl': 2.87,\n 'gearbox_power_ratio': 0.98,\n 'propeller_efficiency': 0.85,\n }\n\n\nlist_of_n2 = np.linspace(0.9, 1.0, 21)\nturboprop = mf.TurboProp(values)\nperformance_results = turboprop.sumarise_results()\nperformance_results = performance_results.drop(columns=[1])\n\nfor n2 in list_of_n2:\n turboprop.set_n2(n2)\n performance_result = turboprop.sumarise_results()\n performance_results[n2] = performance_result\n\n\nfig1 = plt.figure()\nfig1.suptitle(\"Turbine Power\")\naxis1 = fig1.add_subplot(111)\naxis1 = performance_results.loc['turbine_power'].plot()\naxis1.set_ylabel(\"Turbine Power [kW] \")\naxis1.set_xlabel(\"Rotation (N2 / N2 @ P.P) [ ]\")\naxis1.axhline(2050, linestyle='--', color='gray', label='Gearbox Limit Power')\naxis1.axvline(0.941, linestyle='--', color='gray')\naxis1.legend()\n\nperformance_results\n\n# -\n\n# Engine Parameters varied to achieve the manufacture perfomance:\n#\n# | Engine Parameters | |\n# |------------------------------------------|-------------------|\n# | Pressure Ratio in the Free Turbine | 2.87 |\n# | % of maximum rotation in the compressor | 0.941 |\n\n# ## Example 7.2: TurboProp - Flight Condition Simulation\n\n# ### Exercise Information\n#\n# Now simulate the engine operating at 7,000 m and Mach 0.45, keeping other input parameters constant compared to Example 8.1. As operating strategy consider N1 = 100% and N2 = 85%.\n#\n# Operating Engine Parameters:\n#\n# | Baseline Operating Condition | |\n# |------------------------------------------|-------------------|\n# | Mass Flux | 8.49 kg/s |\n# | Outlet temperature in combustion chamber | 1600 K |\n# | Pressure Ration in Compressor | 15.77 |\n#\n#\n# Operating Condition:\n#\n# | Flight Operation Condition | |\n# |------------------------------------------|-------------------|\n# | Flight Mach | 0.45 |\n# | Atmospheric Pressure | 41.00 kPa |\n# | Atmospheric Temperature | 246.55 K |\n# | % of maximum compressor rotation | 0.85 |\n#\n#\n# Efficiency and specific heat ratio data:\n#\n# | Component | Adiabatic Efficiency | Specific Heat Ratio |\n# |--------------------|----------------------|---------------------|\n# | intake | 0,85 | 1,40 |\n# | compressor | 0,75 | 1,37 |\n# | combustion | 1,00 | 1,35 |\n# | compressor turbine | 0,90 | 1,33 |\n# | free turbine | 0,90 | 1,33 |\n# | hot air nozzle | 0,98 | 1,36 |\n#\n# Thermodynamic Data:\n#\n# | Thermodynamic Data | |\n# |------------------------------------------|-------------------|\n# | Lower Heating Value of the Fuel | 45000 kJ/kg |\n# | Mean R | 288,3 m^2/(s^2.K) |\n# | specific heat in the combustion chamber | 1,11 kJ/(kg.K) |\n#\n#\n#\n#\n#\n#\n#\n\n# ### Exercise computation\n# Calibrating the model to the manufacturer report with pressure ratio in the turbine equal to 2.87 and % of maximum rotation equals to 1.00.\n\n# +\nimport model_reference as mf\nimport numpy as np\nimport pandas as pd\nimport warnings\n# warnings.filterwarnings('ignore')\n\nvalues = {\n 'mass_flow': 8.49,\n 't04': 1600,\n 'prc': 15.77,\n 'mach': 0.45,\n 'ta': 246.55,\n 'pa': 41,\n 'n_d': 0.85,\n 'gamma_d': 1.4,\n 'n_c': 0.75,\n 'gamma_c': 1.37,\n 'n_b': 1,\n 'gamma_b': 1.35,\n 'n_t': 0.9,\n 'gamma_t': 1.33,\n 'n_tl': 0.9,\n 'gamma_tl': 1.33,\n 'n_n': 0.98,\n 'gamma_n': 1.36,\n 'pc_fuel': 45000,\n 'cp_fuel': 1.110,\n 'cp_tl': 1.160,\n 'cp_tls': 1.160,\n 'r': 288.3,\n 'pr_tl': 2.87,\n 'gearbox_power_ratio': 0.98,\n 'propeller_efficiency': 0.85,\n }\n\nturboprop = mf.TurboProp(values)\nturboprop.set_n2(0.85)\nperformance_results1 = turboprop.sumarise_results()\nthermodynamic_results1 = turboprop.sumarise()\n\nperformance_results1\n# -\n\n# ## Example 7.3: TurboFan - Comparing to TurboProp 7.2\n\n# ### Exercise Information\n#\n# Simulate a turbofan engine to fulfill the same mission\n# of the turbofan engine in example 8.2, i.e. generate thrust of 4.26 kN at 7,000 m and flight Mach 0.45. To make the simulations we took as reference data from the PW545B engine, with a total pressure ratio of 12.5 and a passage ratio of 4.2 in the take-off condition at sea level. Some assumptions were made for the simulation as will be discussed in the following.\n#\n# Operating Engine Parameters:\n#\n# | Baseline Operating Condition | |\n# |------------------------------------------|-------------------|\n# | Outlet temperature in combustion chamber | 1550 K |\n# | Pressure Ration in Compressor | 8.33 |\n# | Pressure Ration in Booster | 1.00 |\n# | Pressure Ration in Fan | 1.50 |\n# | Air bypass ratio | 4.2 |\n# | Pressure loss in combustion chamber | 0 % |\n#\n# Operating Condition:\n#\n# | Flight Operation Condition | |\n# |------------------------------------------|-------------------|\n# | Flight Mach | 0.45 |\n# | Atmospheric Pressure | 41.00 kPa |\n# | Atmospheric Temperature | 246.55 K |\n# | % of maximum compressor rotation | 0.85 |\n#\n#\n# Efficiency and specific heat ratio data:\n#\n# | Component | Adiabatic Efficiency | Specific Heat Ratio |\n# |--------------------|----------------------|---------------------|\n# | intake | 0,97 | 1,40 |\n# | fan | 0,90 | 1,40 |\n# | compressor | 0,75 | 1,37 |\n# | combustion | 0,9995 | 1,35 |\n# | compressor turbine | 0.90 | 1,33 |\n# | compressor fan | 0.90 | 1,33 |\n# | Hot air nozzle | 0,98 | 1,36 |\n# | Fan nozzle | 0,98 | 1,40 |\n#\n# Thermodynamic Data:\n#\n# | Thermodynamic Data | |\n# |------------------------------------------|-------------------|\n# | Lower Heating Value of the Fuel | 45000 kJ/kg |\n# | Mean R | 288,3 m^2/(s^2.K) |\n# | specific heat in the combustion chamber | 1,11 kJ/(kg.K) |\n#\n# Engine Parameters varied to achieve the TurboProp example 7.2 perfomance:\n#\n# | Engine Parameters | |\n# |------------------------------------------|-------------------|\n# | Mass Flux | 88.42 kg/s |\n#\n#\n\n# ### Exercise Computation\n\n# +\nimport model_reference as mf\nimport numpy as np\nimport pandas as pd\nimport warnings\n# warnings.filterwarnings('ignore')\n\nvalues = {\n 'pa': 41,\n 'ta': 246.55,\n 'n_d': 0.97,\n 'n_f': 0.90,\n 'n_c': 0.75,\n 'n_b': 0.9995,\n 'n_t': 0.9,\n 'n_tf': 0.9,\n 'n_n': 0.98,\n 'n_nf': 0.98,\n 'gamma_d': 1.4,\n 'gamma_f': 1.4,\n 'gamma_c': 1.37,\n 'gamma_b': 1.35,\n 'gamma_t': 1.33,\n 'gamma_tf': 1.33,\n 'gamma_n': 1.36,\n 'gamma_nf': 1.40,\n 'prc': 8.33,\n 'prf': 1.5,\n 'bypass_ratio': 4.2,\n 'pc_fuel': 45000,\n 'cp_fuel': 1.110,\n 'r': 288.3,\n 't04': 1550,\n 'mach': 0.45,\n 'mass_flow': 88.42,\n }\n\nturboprop = mf.TurboFan(values)\nturboprop.set_n2(0.85)\nperformance_results1 = turboprop.sumarise_results()\nthermodynamic_results1 = turboprop.sumarise()\n\nperformance_results1\n","repo_name":"jonasdegrave/gas-turbine","sub_path":"notebook_course_examples.ipynb","file_name":"notebook_course_examples.ipynb","file_ext":"py","file_size_in_byte":23824,"program_lang":"python","lang":"en","doc_type":"code","stars":3,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"22381912256","text":"# +\n# tutorial https://blog.socialcops.com/engineering/machine-learning-python/\n\nimport matplotlib.pyplot as plt\n# %matplotlib inline\nimport random\nimport numpy as np\nimport pandas as pd\nfrom sklearn import datasets, svm, cross_validation, tree, preprocessing, metrics\nimport sklearn.ensemble as ske\n\n#can't use tensorflow as described in the last part of the tutorial,\n#because skflow package has been removed; try \"learn\":\n\nimport tensorflow as tf\n#from tensorflow.contrib import skflow\nimport tensorflow.contrib.learn.python.learn as learn\n# -\n\n#read the spreadsheet file into a Pandas dataframe\ntitanic_df = pd.read_excel('titanic3.xls', 'titanic3', index_col=None, na_values=['NA'])\n\n# .head() shows first 5 rows\ntitanic_df.head()\n\n#show chance of survival for a passenger (it's 38%)\ntitanic_df['survived'].mean()\n\n#group passengers by class and show average survival changes for each class\ntitanic_df.groupby('pclass').mean()\n\n#group by a list of two parameters and assign it to \"class_sex_grouping\"\nclass_sex_grouping = titanic_df.groupby(['pclass','sex']).mean()\nclass_sex_grouping\n\nclass_sex_grouping['survived'].plot.bar()\n\ngroup_by_age = pd.cut(titanic_df[\"age\"], np.arange(0, 90, 10))\nage_grouping = titanic_df.groupby([group_by_age, 'pclass']).mean()\nage_grouping['survived'].plot.bar()\n\n# +\n# preparing data for machine learning\n\n# first: see which columns have missing values\ntitanic_df.count()\n# -\n\n# drop 3 columns\ntitanic_df = titanic_df.drop(['body','cabin','boat'], axis=1)\n\ntitanic_df[\"home.dest\"] = titanic_df[\"home.dest\"].fillna(\"NA\")\n\n# drop lines without age\ntitanic_df = titanic_df.dropna()\n\ntitanic_df.count()\n\n\n# function: format remaining data in a way that the machine learning algorithms will accept\n# i.e. male and female become 0 and 1\n# drop name, ticket and home.dest because we can't break them down to 0 and 1\ndef preprocess_titanic_df(df):\n processed_df = df.copy()\n le = preprocessing.LabelEncoder()\n processed_df.sex = le.fit_transform(processed_df.sex)\n processed_df.embarked = le.fit_transform(processed_df.embarked)\n processed_df = processed_df.drop(['name','ticket','home.dest'],axis=1)\n return processed_df\n\n\n# use the function defined above\nprocessed_df = preprocess_titanic_df(titanic_df)\n\n# separate data set into 2 arrays: X with everything but survived, Y only with survived\nX = processed_df.drop(['survived'], axis=1).values\ny = processed_df['survived'].values\n\n# divide X and Y in a training set (80%) and in a test group (20%)\nX_train, X_test, y_train, y_test = cross_validation.train_test_split(X,y,test_size=0.2)\n\n# +\n# make an untrained decision tree classifier named clf_dt,\n# that examines one variable after the other\n\n# our tree splits max. 10 times beginning with the most important\nclf_dt = tree.DecisionTreeClassifier(max_depth=10)\n\n# +\n# create the tree with our trainig set\nclf_dt.fit (X_train, y_train)\n\n# test the tree and see, how accurate it is\nclf_dt.score (X_test, y_test)\n\n# +\n# Problem: depending on the choosen text data the results vary\n# Solution: a shuffle validator that generates 20 unique permutations of the 20/80 split\n\nshuffle_validator = cross_validation.ShuffleSplit(len(X), n_iter=20, test_size=0.2, random_state=0)\ndef test_classifier(clf):\n scores = cross_validation.cross_val_score(clf, X, y, cv=shuffle_validator)\n print(\"Accuracy: %0.4f (+/- %0.2f)\" % (scores.mean(), scores.std()))\n\n\n# -\n\ntest_classifier(clf_dt)\n\n# +\n# next try: the “Random Forest” classification algorithm\n# generates several decision trees and returns the most returnd prediction\n\nclf_rf = ske.RandomForestClassifier(n_estimators=50)\ntest_classifier(clf_rf)\n# -\n\nclf_gb = ske.GradientBoostingClassifier(n_estimators=50)\ntest_classifier(clf_gb)\n\neclf = ske.VotingClassifier([('dt', clf_dt), ('rf', clf_rf), ('gb', clf_gb)])\ntest_classifier(eclf)\n\n# +\n# Deep Neural Network (DNN)\n\n# DNNs don't just pass data between nodes, but between layers of nodes\n# are able to find patterns in unstructured data\n# TensorFlow = open source libary for programing neural networks\n# Problem: skflow isn't working anymore \n# -\n\ntf_clf_dnn = skflow.TensorFlowDNNClassifier(hidden_units=[20, 40, 20], n_classes=2, batch_size=256, steps=1000, learning_rate=0.05)\ntf_clf_dnn.fit(X_train, y_train)\ntf_clf_dnn.score(X_test, y_test)\n\n\ndef custom_model(X, y):\n layers = skflow.ops.dnn(X, [20, 40, 20], tf.tanh)\n return skflow.models.logistic_regression(layers, y)\n\n\ntf_clf_c = skflow.TensorFlowEstimator(model_fn=custom_model, n_classes=2, batch_size=256, steps=1000, learning_rate=0.05)\ntf_clf_c.fit(X_train, y_train)\nmetrics.accuracy_score(y_test, tf_clf_c.predict(X_test))\n\n\n","repo_name":"id-gue/ml-titanic","sub_path":"ml-titanic-jupyter.ipynb","file_name":"ml-titanic-jupyter.ipynb","file_ext":"py","file_size_in_byte":4648,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"19127821951","text":"# # Loading Source Data from Google Sheets\n# The review data supporting this project was collected by me as part of a Google Sheets spreadsheet. This notebook shows how to properly connect to that sheet to download the data into a local CSV.\n\n# Importing required Python libraries\nimport pandas as pd\n\n# Defining the ID of the Google Sheet with the movie ratings\nsheet_id = '1-8tdDUtm0iBrCdCRAsYCw2KOimecrHcmsnL-aqG-l0E'\n\n\n# Creating a small function to load the data sheet by ID and sheet name\ndef load_google_sheet(sheet_id, sheet_name):\n url = f'https://docs.google.com/spreadsheets/d/{sheet_id}/gviz/tq?tqx=out:csv&sheet={sheet_name}'\n df = pd.read_csv(url)\n return df\n\n\n# Loading all the sheets and joining them together\ndf_main = load_google_sheet(sheet_id, 'main')\ndf_patreon = load_google_sheet(sheet_id, 'patreon')\ndf_mnight = load_google_sheet(sheet_id, 'movie_night')\ndf = pd.concat([df_main, df_patreon, df_mnight], axis = 0)\n\n# Viewing the first few rows of the combined DataFrame\ndf.head()\n\n# Keeping only the movies\ndf_movies = df[df['Category'] == 'Movie']\ndf_movies.head()\n\n# Removing the columns we do not need for the model\ndf_movies.drop(columns = ['Category', 'Episode Number', 'Notes'], inplace = True)\ndf_movies.head()\n\n# Creating a dictionary to rename the columns\nnew_col_names = {'Name': 'movie_name',\n 'Rating': 'biehn_scale_rating',\n 'Flickable': 'biehn_yes_or_no'}\n\n# Applying the renaming to the DataFrame\ndf_movies.rename(columns = new_col_names, inplace = True)\n\n# Viewing the first few rows of the final DataFrame\ndf_movies.head()\n\n# Saving the dataset to an external directory\ndf_movies.to_csv('../data/raw/caelan_reviews.csv', index = False)\n\n\n","repo_name":"dkhundley/movie-ratings-model","sub_path":"notebooks/load-from-google-sheets.ipynb","file_name":"load-from-google-sheets.ipynb","file_ext":"py","file_size_in_byte":1720,"program_lang":"python","lang":"en","doc_type":"code","stars":4,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"9057008111","text":"import matplotlib.pyplot as plt\nfrom sklearn.preprocessing import normalize\nimport scipy.io.wavfile\n\nrate, data = scipy.io.wavfile.read('voy.wav')\n\ndata = normalize(data)\ndatal = [s[0] for s in data]\ndatar = [s[1] for s in data]\n\nrate\n\nprint(\"{} samps\".format(len(data)))\nprint(\"{:.2f} samps per s\".format(len(data)/(15*60+45)))\n\n0.017*rate\n\nimg = []\nwidth = 734.4\noffset = int((31.3363)*rate) + int(0*width)\nfor i in range(512): #512\n offset2 = int(i*width)\n img.append(datal[offset + offset2 : offset + offset2 + int(width)])\nplt.figure(dpi=200)\nplt.imshow(img, cmap=plt.get_cmap(\"Greys\"))\nplt.show()\n\nplt.figure(dpi=200)\nplt.imshow(img, cmap=plt.get_cmap(\"Greys\"))\nplt.show()\n\n\n\n60*16*rate/55/512\n","repo_name":"pracowniaPK/VoyagerDecoder","sub_path":"voyager.ipynb","file_name":"voyager.ipynb","file_ext":"py","file_size_in_byte":706,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"70218668637","text":"# # Data Consistency Checks\n#\n# ## - Mixed-type Data\n# ## - Missing Values\n# ## - Duplicates\n\n# import libraries\nimport pandas as pd\nimport numpy as np\nimport os\n\n# +\n# import data: products.csv from Original Data\npath = r'/Users/azadehrezaei/Desktop/Career Foundry/Achievement 4/08-2021 Instacart Basket Analysis'\ndf_prods = pd.read_csv(os.path.join(path, '02 Data', 'Original Data', 'products.csv'), index_col = False)\n\n#import data: orders_wrangled.csv from Prepared data\ndf_ords = pd.read_csv(os.path.join(path, '02 Data', 'Prepared Data', 'orders_wrangled.csv'), index_col = False)\n# -\n\n# Create a dataframe\ndf_test = pd.DataFrame()\n\n# Create a mixed type column\ndf_test['mix'] = ['a', 'b', 1, True]\n\ndf_test.head()\n\n# Check for mixed data types\nfor col in df_test.columns.tolist():\n weird = (df_test[[col]].applymap(type) != df_test[[col]].iloc[0].apply(type)).any(axis = 1)\n if len (df_test[weird]) > 0:\n print (col)\n\n# Change data type to string \ndf_test['mix'] = df_test['mix'].astype('str')\n\n# # Finding Missing Values:\n\n# Fidning the columns that contain missing values, and how many values they're missing:\ndf_prods.isnull().sum()\n\n# Creating a subset for the dataframe, containing the missing records:\ndf_nan = df_prods[df_prods['product_name'].isnull() == True]\n\ndf_nan\n\ndf_prods.shape\n\ndf_prods_clean = df_prods[df_prods['product_name'].isnull() == False]\n\ndf_prods_clean.shape\n\n# Finding duplicates and assigning them to a new subset\ndf_dups = df_prods_clean[df_prods_clean.duplicated()]\n\ndf_dups\n\n# Dropping duplicates \ndf_prods_clean_no_dups = df_prods_clean.drop_duplicates()\n\ndf_prods_clean_no_dups.shape\n\n# # Exercise 4.5:\n\n# +\n# Question 1: All consistency checks are done in the above section. \n# -\n\n# Question 2: \ndf_prods.describe()\n\n# ##I don't think anything looks off in this dataframe in the first 3 columns based on the results of df.describe(), the only think I'd probably change is the data type of product_id, aisle_id and department_id columns since these columns are simply identifiers and although the values are integers they don't need to be treated as integers and there's no value in performing descriptive analysis on them. I would change these datatypes to string instead. \n#\n# The price column however shows inconsistency. The maximum price value is recorded as 9999 although the mean value is still pretty low. This means that this value might be an outlier.\n\n# Question 3: Checking for mixed-type data in df_ords\ndf_ords.describe()\n\n# Question 3: Checking for mixed type data in df_ords:\nfor col in df_ords.columns.tolist():\n weird = (df_ords[[col]].applymap(type) != df_ords[[col]].iloc[0].apply(type)).any(axis = 1)\n if len(df_ords[weird]) > 0:\n print(col + ' has mixed data type')\n else:\n print(col + ' has one data type!')\n \n\n# df_ords columns all have uniform data types. \n\n# Question 5: Checking for missing values in df_ords: \ndf_ords.isnull().sum()\n\n# The column \"days_since_prior_order\" has 206,209 rows of missing values. \n# This number can be the first time users placed their order which means since they are the very first order, this row doesn't get any value!\n#\n\ndf_ords.shape\n\n# To check this hypothesis we can compare the number of missing values in this column with the number of order_id's that are marked as 1:\n\ndf_ords_firstorders = df_ords[df_ords['order_number'] == 1]\n\ndf_ords_firstorders.shape\n\n# The number of missing values in the \"days_since_prior_order\" column and the number of first orders are both 206,209. So this shows that the empty/missing values in \"days_since_prior_order\" column are simply the users' first orders. \n# There are a total of 3,421,083 rows of data and the missing values makes about 6% of the total rows of data. Since we are trying to use this analysis to investigate customer purchasing behavior, I am not sure how the rows with the missing value for the \"days_since_prior_order\" can add value to our analysis and I don't think removing these rows would damage the data; So I decided to removed these rows:\n#\n#\n# *** Correction: I undid the removal of the null values after the feedback from pedro:\"These are all valid data points so removing them would be a mistake. Here I would leave it as null or use a number like -1 to indicate the first order and so the data is not mixed.\"\n\ndf_ords_firstorders_replaced = df_ords['days_since_prior_order'].fillna(value = -1, inplace = True)\n\n# I noticed here that instead of having null values I have string NaN values that don't show up as null here:\ndf_ords.isnull().sum()\n\n# so to fix the NaN values, I replaced them with -1 using the .replace() function.\ndf_ords['days_since_prior_order'].replace({'Nan': -1})\n\ndf_ords\n\n# to check my work I counted the values in this column and I have the same number of -1s as I had nulls and NaNs! \ndf_ords['days_since_prior_order'].value_counts()\n\n# CORRECTION: Leaving this here but I am not going to remove the null values in the clean dataframe. Replaced them with -1\n# Removing the rows with missing values:\ndf_ords_clean = df_ords[df_ords['days_since_prior_order'].isnull() == False]\n\ndf_ords_clean.shape\n\n# Question 8: Address the duplicates\ndf_ords_dups = df_ords_clean[df_ords_clean.duplicated()]\n\ndf_ords_dups.shape\n\n# There are no full duplicates found in this set, which is great. There might be rows that share the same order_id and user_id or even the department_id and this is normal since it can simply show that a customer purchase multiple items from the same department in the same order.\n\ndf_ords.head()\n\ndf_ords.columns\n\n# +\n# df_ords = df_ords.drop(columns = ['Unnamed: 0'])\n# -\n\ndf_ords.head()\n\n# +\n# Question 9: Export the final cleaned datafarmes: \n# CORRECTION: I am replacing the df_ords_clean with \n\ndf_prods_clean_no_dups.to_csv(os.path.join(path, '02 Data', 'Prepared Data', 'prods_clean.csv'))\n\ndf_ords.to_csv(os.path.join(path, '02 Data', 'Prepared Data', 'orders_clean.csv'))\n# -\n\ndf_ords\n\ndf_test = pd.read_csv(os.path.join(path, '02 Data', 'Prepared Data', 'orders_clean.csv'), index_col = False)\n\ndf_test.head()\n\n\n","repo_name":"Azadeh-Rezaei/Python-InstacartGroceryBasketAnalysis","sub_path":"03 Scripts/Exercise 4.5.ipynb","file_name":"Exercise 4.5.ipynb","file_ext":"py","file_size_in_byte":6062,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"50"}
+{"seq_id":"16380105203","text":"# # Markov Decision Process\n\nimport numpy as np\n\n\nclass GameShow:\n def __init__(self):\n # Define the states of the game\n self.states = ['Q1', 'Q2', 'Q3', 'Q4', 'end', 'lost']\n \n # Define the available actions and their corresponding values\n self.actions = {'play': 1, 'quit': 0}\n \n # Define the rewards associated with each state\n self.rewards = {'Q1': 100, 'Q2': 1000, 'Q3': 10000, 'Q4': 50000, 'end': 0, 'lost': 0}\n \n # Define the probabilities of success for transitioning to the next state\n self.probs = {'Q1': 0.9, 'Q2': 0.75, 'Q3': 0.5, 'Q4': 0.1}\n \n # Initialize the current state and current return variables\n self.currentState = 'Q1'\n self.currentReturn = 0\n \n # Initialize the random number generator\n self.rng = np.random.default_rng()\n \n def resetGame(self):\n # Reset the game to the initial state\n self.currentState = 'Q1'\n self.currentReturn = 0\n \n def play(self, action):\n if self.currentState == 'end':\n print('You cannot play anymore.')\n elif action == 0:\n # If action is 'quit', transition to the end state\n self.currentState = 'end'\n elif action == 1:\n if self.rng.random() < self.probs[self.currentState]:\n # If action is 'play' and success probability is met, transition to the next state\n newIndex = self.states.index(self.currentState) + 1\n self.currentState = self.states[newIndex]\n else:\n # If action is 'play' and success probability is not met, transition to the lost state\n self.currentState = 'lost'\n self.currentReturn = 0\n \n # Update the current return with the reward associated with the current state\n self.currentReturn += self.rewards[self.currentState]\n\n\n\n# Create a GameShow instance\nmyShow = GameShow()\n\n# +\n# Print the initial state and current return of the game\nprint(myShow.currentState, myShow.currentReturn)\n\n# Play the game for 10 iterations\nfor _ in range(10):\n # Take action 'play'\n myShow.play(action=1)\n \n # Print the current state and current return of the game\n print(myShow.currentState, myShow.currentReturn)\n \n # Check if the game has reached an end state or lost state\n if myShow.currentState in ['end', 'lost']:\n # Reset the game to its initial state\n myShow.resetGame()\n \n # Print the current state and current return after resetting the game\n print(myShow.currentState, myShow.currentReturn)\n\n\n# -\n\nclass Agent:\n def __init__(self, task, initPredMat=True):\n self.task = task\n self.rng = np.random.default_rng()\n \n if initPredMat:\n self.predMat = np.full(5, 0.5) # Initialize the prediction matrix with default values\n else:\n self.predMat = initPredMat # Use the provided prediction matrix\n \n def playGame(self, T, out=True):\n t = 0\n if out:\n print(self.task.currentState, self.task.currentReturn) # Print the initial state and current return of the game\n \n while t < T:\n state = self.task.currentState # Get the current state of the game\n stateId = self.task.states.index(self.task.currentState) # Get the index of the current state\n \n # Generate a random number and compare it with the corresponding prediction probability\n cond = self.rng.random() < self.predMat[stateId]\n \n action = 1 if cond else 0 # Choose the action based on the condition\n self.task.play(action) # Play the game by taking the chosen action\n \n if out:\n print(self.task.currentState, self.task.currentReturn) # Print the current state and current return of the game\n \n if self.task.currentState in ['end', 'lost']:\n self.task.resetGame() # Reset the game if it has reached an end or lost state\n \n t += 1 # Increment the time step\n\n\n# Create an agent\nplayer = Agent(myShow)\n\nmyShow.resetGame() # reset the game\nplayer.playGame(100) # play the game for 100 questions\n\n\n# ## Q-values\n\n# We define a new mehtod of the agent to estimate value of a given state and a new method to compute the value of a state\n\nclass Agent:\n def __init__(self, task, initPredMat=True):\n self.task = task\n self.rng = np.random.default_rng()\n \n if initPredMat:\n self.actionPredMat = np.full(5, 0.5) # Initialize the prediction matrix with default values\n self.statePredMat = np.full((2,5), 0.5) \n else:\n self.predMat = initPredMat # Use the provided prediction matrix\n \n def playGame(self, T, out=True):\n t = 0\n if out:\n print(self.task.currentState, self.task.currentReturn) # Print the initial state and current return of the game\n \n while t < T:\n state = self.task.currentState # Get the current state of the game\n stateId = self.task.states.index(self.task.currentState) # Get the index of the current state\n \n # Generate a random number and compare it with the corresponding prediction probability\n cond = self.rng.random() < self.predMat[stateId]\n \n action = 1 if cond else 0 # Choose the action based on the condition\n self.task.play(action) # Play the game by taking the chosen action\n \n if out:\n print(self.task.currentState, self.task.currentReturn) # Print the current state and current return of the game\n \n if self.task.currentState in ['end', 'lost']:\n self.task.resetGame() # Reset the game if it has reached an end or lost state\n \n t += 1 # Increment the time step\n \n def computeStateValue(self):\n pass\n\n\n(player.task.currentState, player.task.currentReturn)\n\n# +\nindex = player.task.states.index(player.task.currentState)\n\nplayer.predMat[index] \n# -\n\nnp.full((2,5), 0.5)\n\n\n\n\n\n\n\n\n\n\n\n\n\n# # Reinforcement Learning\n#\n# - you are a robot in a world controlled by an MDP\n# - you are not given the rewards (R) or transition probabilities (T)\n# - Method 1: Certainty-Equivalent Learning\n# - learn R and T from observation\n# - possible, but takes many repititions\n# - Method 2: Value sampling\n# - Compute expected return at each time step\n# - similarly long\n# - Method 3: Stochastic Approximation Theory - Temporal Difference Learning (TDL)\n# - $newEstimate = (1-a)OldEstimate + a*newSample$\n# - Method 4: Q-learning\n# - TDL applied to q-values\n# - \n\n# - at the end, we make the decision through a soft-max, i.e. we select action $i$ with probability $\\frac{\\exp(Q(s_t, i))}{\\sum_{j\\neq i}\\exp(Q(s_t, j)}$ .\n\n\n","repo_name":"Mod4NeuCog/lif-neuron-JakobGruen","sub_path":"Ashwin/.ipynb_checkpoints/mdp_game-checkpoint.ipynb","file_name":"mdp_game-checkpoint.ipynb","file_ext":"py","file_size_in_byte":7062,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"50"}
+{"seq_id":"15748654717","text":"# %pip install tensorflow\n\nimport tensorflow as tf\n\n\n# +\nfrom tensorflow.keras import datasets\n\n# 케라스 데이터셋을 다운받아 변수에 각각 넣어준다.\n(train_images, train_labels), (test_images,\n test_labels) = datasets.cifar10.load_data()\n# -\n\nfrom tensorflow.keras.applications.resnet50 import ResNet50\nbase_model = ResNet50(include_top=True, input_shape = (224, 224 ,3), weights = 'imagenet')\n\nbase_model.summary()\n\n# +\nbase_model.compile(\n optimizer='adam', loss='sparse_categorical_crossentropy', metrics='accuracy')\n\nfor i in range(5):\n inputs = test_images[i] \n inputs = tf.expand_dims(inputs, axis=0)\n x = tf.keras.layers.experimental.preprocessing.Resizing(\n 224, 224)(inputs) \n \n x = tf.keras.applications.resnet50.preprocess_input(x)\n base_model.evaluate(x, test_labels[i])\n\n\n# +\nimport matplotlib.pyplot as plt\nfrom tensorflow.keras.applications.resnet50 import decode_predictions\n\nfor i in range(5):\n\n inputs = test_images[i]\n inputs = tf.expand_dims(inputs, axis=0)\n x = tf.keras.layers.experimental.preprocessing.Resizing(224, 224)(inputs)\n x = tf.keras.applications.resnet50.preprocess_input(x)\n\n preds = base_model.predict(x) \n\n fig = plt.figure(figsize=(3, 3))\n plt.title(decode_predictions(preds, top=1)[0][0][1]) \n plt.imshow(test_images[i]) \n plt.show()\n\n","repo_name":"jjack-lee/cbnu","sub_path":"projects/TensorFlow_resnet50_base/resnet50_base.ipynb","file_name":"resnet50_base.ipynb","file_ext":"py","file_size_in_byte":1367,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"50"}
+{"seq_id":"30008360859","text":"# # Исследование надёжности заёмщиков\n#\n# Заказчик — кредитный отдел банка. Нужно разобраться, влияет ли семейное положение и количество детей клиента на факт погашения кредита в срок. Входные данные от банка — статистика о платёжеспособности клиентов.\n#\n# Результаты исследования будут учтены при построении модели **кредитного скоринга** — специальной системы, которая оценивает способность потенциального заёмщика вернуть кредит банку.\n\n# **Описание данных:**\n#\n# - *children* — количество детей в семье\n# - *days_employed* — общий трудовой стаж в днях\n# - *dob_years* — возраст клиента в годах\n# - *education* — уровень образования клиента\n# - *education_id* — идентификатор уровня образования\n# - *family_status* — семейное положение\n# - *family_status_id* — идентификатор семейного положения\n# - *gender* — пол клиента\n# - *income_type* — тип занятости\n# - *debt* — имел ли задолженность по возврату кредитов\n# - *total_income* — ежемесячный доход\n# - *purpose* — цель получения кредита\n\n# ### Шаг 1. Файл с данными. \n\nimport pandas as pd\nfrom pymystem3 import Mystem\n\n# +\nm = Mystem()\n\ndata = pd.read_csv('/datasets/data.csv')\ndata.info()\n# -\n\ndata\n\n# ### Вывод\n\n# Данные содержат ошибки и пропущенные значения:\n# * столбы days_employed и total_income содержат пропуски;\n# * также дробное представление данных в этих столбцах излишне, стаж удобнее считать в целых днях, а доход без копеек;\n# * в столбце days_employed присутствуют аномально большие значения. Сомневаюсь, что типусы могут иметь стаж более 900 лет, они бы уже за 300 лет выстроли свою экономику так, чтобы им больше не пришлось работать;\n# * также в этом столбце содержится большое количество отрицательных значений;\n# * в education и family_status следует все значения привести к нижнему регистру.\n\n# ### Шаг 2. Предобработка данных\n\n# ### Обработка пропусков\n\ndata[data['days_employed'].isna()] #посмотрим строки, содержащие NaN значения\n\n# Гипотеза о том, что данные пропущены у пользователей, которые никогда не работали не подходит, так как значения в столбце income_type свидетельствуют об обратном. Скорее всего при заполнении таблицы часть данных была потеряна. \n# Заполним пропуски средними значениями или медианой. \n# Для начала избавимся от минусов и аномально больших значений.\n\ndata[data['days_employed'] > 0]\n\ndata[data['days_employed'] > 0].describe()\n\n# Все положительные значения в *days_employed* являются аномальными.\n# Также есть клиенты с возрастом 0 лет, таких уберем из данных.\n\ndata[data['income_type'] == 'пенсионер'] #хм. пенсионеры какие-то подозрительные, посмотрим на них поближе\n\nprint(data[data['income_type'] == 'пенсионер']['income_type'].count()) # количество пенсионеров\ndata[(data['income_type'] == 'пенсионер') & (data['days_employed'] > 0)]['income_type'].count() # количество пенсионеров с аномально большими значниями стажа\n\n# Похоже все строки с \"пенсионерами\" заполняются некорректно. Причем удаление строк с аномально большими значениями стажа приведет к удалению большей части пенсионеров, тогда заменим их средними значениями. \n# Но для начала избавимся от отрицательных значений.\n\ndata.loc[data['days_employed'] > 0, 'days_employed'] = 0\n\ndata = data[data['dob_years'] > 0]\n\ndata.loc[:,['days_employed']] = data['days_employed'] * (-1)\ndata\n\ndays_employed_avg = data['days_employed'].mean()\ndays_employed_avg\n\n# +\nimport warnings\n\nwarnings.filterwarnings('ignore')\n# -\n\ndata['days_employed'] = data.loc[:,['days_employed']].fillna(days_employed_avg) # заменим отсутствующие значения на среднии\ndata['days_employed'] = data.loc[:,['days_employed']].replace(0, days_employed_avg) # заменим аномальные значения на среднии\ndata.head(10)\n\n# аналогично заменим пропуски в столбце total_income, использую медиану, т.к. высокие зарплаты могут сместить среднее вверх\ntotal_income_median = data['total_income'].median()\ndata['total_income'] = data['total_income'].fillna(total_income_median)\n\ndata.describe()\n\n# ### Вывод\n\n# Пропуски устранены. Выявлено, что столбец days_employed для пенсионеров заполняется некоррекно.\n\n# ### Замена типа данных\n\n# Заменим тип данных столбцов days_employed и total_income на целочисленный\ndata[['days_employed', 'total_income']] = data[['days_employed', 'total_income']].astype('int')\ndata.head(10)\n\n# ### Вывод\n\n# Значения в days_employed и total_income теперь имеют удобный вид\n\n# ### Обработка дубликатов\n\ndata['education'] = data['education'].str.lower() # значения столбца education к нижнему регистру\ndata['family_status'] = data['family_status'].str.lower() # значения столбца family_status к нижнему регистру\n\n# Так как мы не можем отобрать уникальных пользователей, будем искать дубли по полной строке\ndata.duplicated().sum()\n\n# удалим дубли\ndata = data.drop_duplicates().reset_index(drop=True)\n\n# ### Вывод\n\n# Дубли обнаружены в незначительном количестве и удалены.\n\n# ### Лемматизация\n\ndata['purpose'].unique() # уникальные цели получения кредита\n\n\n# функция лемматизирует значение столбца purpose и проверяя по ключевому слову выдает значение нового столбца purpose_category\ndef purpose_lem(purpose):\n lemmas = m.lemmatize(purpose)\n if 'недвижимость' in lemmas or 'жилье' in lemmas:\n return 'недвижимость'\n if 'автомобиль' in lemmas:\n return 'автомобиль'\n if 'образование' in lemmas:\n return 'образование'\n if 'свадьба' in lemmas:\n return 'свадьба'\n else:\n return 'другое'\n \n\n\ndata['purpose_category'] = data['purpose'].apply(purpose_lem)\ndata.head(10)\n\ndata['purpose_category'].unique()\n\n# ### Вывод\n\n# Цели получения кредита поделены на 4 категории.\n\n# ### Категоризация данных\n\n# Выделим два словаря: по семейному положению и по уровню образования.\n\nfamily_status = data[['family_status_id', 'family_status']]\neducation_level = data[['education_id', 'education']]\n\n\n# разделим доход на 4 категории: до 60к от 60к до 100к, от 100к до 200к и более 200к\ndef profit_people(total_income):\n if total_income < 60000:\n return 'до 60к'\n elif 60000 <= total_income < 100000:\n return '60-100'\n elif total_income >= 200000:\n return 'более 200к'\n else:\n return 'от 100 до 200'\n \n\n\ndata['profit_cat'] = data['total_income'].apply(profit_people)\n\n# ### Вывод\n\n# Данные категоризированы на два словаря.\n\n# ### Шаг 3. Исследование\n\n# - Проверим, есть ли зависимость между наличием детей и возвратом кредита в срок\n\n# +\n# в children значения -1 и 20 - скорее всего опечатки, заменим их на 1 и 2 соответственно\ndata['children'] = data['children'].replace(-1, 1)\ndata['children'] = data['children'].replace(20, 2)\n\ndata_group_children = data.groupby('children').agg({'debt': ['count', 'sum']}).reset_index() # групировка по children с подсчетом суммы невозвратов\ndata_group_children['ratio'] = data_group_children['debt']['sum'] / data_group_children['debt']['count'] # ratio - индекс невозврата кредита в срок\ndata_group_children\n# -\n\nchildren_pivot = data_group_children.pivot_table(index='children', values='ratio', aggfunc='sum')\nchildren_pivot\n\n# ### Вывод\n\n# Индекс невозврата для клиентов у которых есть дети при любом количестве детей, за исключением 5 детей (слишком малочисленная группа для исследования) больше чем у клиентов без детей.\n\n# - Проверим, есть ли зависимость между семейным положением и возвратом кредита в срок\n\ndata_group_family_status = data.groupby('family_status').agg({'debt': ['count', 'sum']}).reset_index() # групировка по family_status с подсчетом суммы невозвратов\ndata_group_family_status['ratio'] = data_group_family_status['debt']['sum'] / data_group_family_status['debt']['count']\ndata_group_family_status\n\n# ### Вывод\n\n# Наибольший результат показывают люди, живущие в гражданском браке и неженатые/незамужнии.\n\n# - Есть ли зависимость между уровнем дохода и возвратом кредита в срок?\n\ndata_group_profit = data.groupby('profit_cat').agg({'debt': ['count', 'sum']}).reset_index() # групировка по profit_cat с подсчетом суммы невозвратов\ndata_group_profit['ratio'] = data_group_profit['debt']['sum'] / data_group_profit['debt']['count']\ndata_group_profit\n\n# ### Вывод\n\n# Не у одной категории доходов нет невозвратов более 9%. Наибольшую вероятность невозврата имеют люди, получающие от 100к до 200к. Причины неизвестны.\n\n# - Как разные цели кредита влияют на его возврат в срок?\n\ndata_group_purpose = data.groupby('purpose_category').agg({'debt': ['count', 'sum']}).reset_index() # групировка по data_group_purpose с подсчетом суммы невозвратов\ndata_group_purpose['ratio'] = data_group_purpose['debt']['sum'] / data_group_purpose['debt']['count']\ndata_group_purpose\n\n# ### Вывод\n\n# Примерно одинаковые наибольшие индексы невозврата имеют цели для приобретения автомобиля и получение образования. Меньший результат покаывают \"недвижимость\" и \"свадьба\", скорее всего люди относятся к покупке недвижимости более серьезно, чем к покупке автомобиля, а кридит за свадьбу могут оплатить подарками гостей.\n\n# ### Шаг 4. Общий вывод\n\n# Индекс невозврата, показывающий для данной категории процент людей не вернувших кредит в срок, принимает наибольшие значения приближенные к 9 %. \n# Люди имеющие детей чаще не возвращают кредит в срок, чем бездетные. \n# Однако живущие гражданским браком или не состоявшие в браке также имеют высокий индекс невозврата (9,7 % для последних). \n# Исследование зависимости между уровнем дохода и возвратом кредита показало, что чаще склонны просрочить кредит люди, получающие ежемесячный доход от 100 до 200 тысяч.\n# А исследование целей получения кредита, показало, что стоит уделить внимание людям имеющих цели, связанные с автомобилем или образованием.\n\n\n","repo_name":"NColemann/yandex-praktikum-projects","sub_path":"1.Исследование_надёжности_заёмщиков/1.Исследование_надёжности_заёмщиков.ipynb","file_name":"1.Исследование_надёжности_заёмщиков.ipynb","file_ext":"py","file_size_in_byte":14120,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"50"}
+{"seq_id":"12692462325","text":"# + id=\"wdmzgoLq9sk7\"\nimport pandas as pd\nimport numpy as np\n\n# + id=\"gNN9P2vQzA4H\"\nmarkup = pd.read_csv('markup.csv', sep = ';')\nvalid = pd.read_csv('valid.csv', sep = ';')\n\n# + id=\"uA4WVQ8DooYl\"\nmarkup = pd.read_csv('markup.csv', sep = ';')\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 142} id=\"oyMMPg8rVDfR\" outputId=\"c562f212-7a68-4ae2-87ef-7748d04946ee\"\nvalid\n\n# + [markdown] id=\"717O9FaOmIjo\"\n# # Делаем обязательную часть из условия задачи:\n# 1. переводим markup_dt в datetime\n# 2. проставляем порядковый номер ans_set для ответа размечтика\n\n# + id=\"BvdZQ780VLB3\"\n# присваиваем ans_set\n\nmarkup['markup_dt'] = pd.to_datetime(markup['markup_dt']) # трансформируем в нужный тип данных\nmarkup.sort_values(by=['text_id', 'markup_dt'], ascending=[True, True], \n inplace=True) # сортируем по каждому тексту ответы разметчиков по времени\nmarkup['ans_set'] = markup.groupby(['text_id'])['markup_dt'].rank() # присваиваем ans_set от 1 до 3 для каждого текста\nmarkup['ans_set'] = markup['ans_set'].astype(int)\n\n# + [markdown] id=\"I3-T9P6JmVC7\"\n# # Решаем задачу №1 - создать витрину правильных ответов по каждому тексту, который пойдет на разметку\n\n# + id=\"MhX-NjEQc0fy\"\n# создаем датасет, куда будем складывать правильные ответы\n\nmarkup_answers = pd.DataFrame()\ntexts = list(set(markup['text_id'].values))\nmarkup_answers['text_id'] = texts\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 173} id=\"aCw-B5oGbmZu\" outputId=\"6e6f7b12-4898-415e-a1c0-ab5c0b9dcc7c\"\nmarkup_answers\n\n# + id=\"Pk7YriSMdIRU\"\nmarkup_answers.sort_values(by='text_id', inplace=True) # сортируем в нужном порядке текста\n\n# + id=\"4NoD9LlPY_8q\"\nmarkup_answers['subject1'] = markup.groupby('text_id')['subject1'].agg(\n lambda x: x.value_counts().index[0] \n if x.value_counts().values.max() >= 2 else np.nan\n).values # порядок соответсвует ответам, потому что изначальный датафрейм markup мы отсортировали по текстам\n\n# + id=\"hAkoxKgPdzEt\"\nmarkup_answers['subject2'] = markup.groupby('text_id')['subject2'].agg(\n lambda x: x.value_counts().index[0] \n if x.value_counts().values.max() >= 2 else np.nan\n).values\n\n# + id=\"vWrrS4gMd0kh\"\nmarkup_answers['subject3'] = markup.groupby('text_id')['subject3'].agg(\n lambda x: x.value_counts().index[0] \n if x.value_counts().values.max() >= 2 else np.nan\n).values\n\n# + id=\"vO1a6FTBd2T-\"\nmarkup_answers['subject4'] = markup.groupby('text_id')['subject4'].agg(\n lambda x: x.value_counts().index[0] \n if x.value_counts().values.max() >= 2 else np.nan\n).values\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 173} id=\"-l_P4eOJg7yE\" outputId=\"b78f2e29-505a-4775-96e3-62ac5798f490\"\nmarkup_answers # получен ли консенсус по текстам и subject, если есть консенсус по всем subject, то не будет nan в строке по тексту\n\n# + id=\"hmCXOOvVh4TO\"\nconsensus = markup_answers.dropna()\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 80} id=\"D60G20mliZUR\" outputId=\"cc4bee90-eacb-4e5e-ba58-e98a2d615acb\"\nconsensus # те тексты по которым был консенсус по всем subject\n\n# + id=\"cp-ESVLkhN1b\"\nconsensus_valid = pd.concat([valid, consensus])\n\n# + id=\"dRR8Rr0BiWM1\"\nconsensus_valid.reset_index(drop=True, inplace=True)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 173} id=\"oKSinjr3irJL\" outputId=\"d8bef4d0-a5fc-4007-96cc-f16d48d2d7e3\"\nconsensus_valid # датафрейм который состоит из valid и тех строк где получен консенсус\n\n# + id=\"sNt4pFdLiuqL\"\nfinal = markup.merge(consensus_valid, how='left', on='text_id', suffixes=['_mark', '_valid']) \n\n# + id=\"TcYJeb01kjjS\"\n# проверка на то указал ли валидатор несколько значений или нет: если несколько взять меньший\nfinal['ans_set_valid'] = np.where(\n len(final['ans_set_valid']) > 1, # если больше 1 значения\n final['ans_set_valid'].astype(str).apply(lambda x: x.split(';')[0]), # вытащим первый номер\n final['ans_set_valid'] # оставим также\n)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 645} id=\"_ZjUBBRWknz-\" outputId=\"bbd7508b-368c-4061-e09d-52a730bbd9bf\"\nfinal\n\n# + id=\"xqKP-cCEkD9l\"\nfinal['ans_set_mark'] = final['ans_set_mark'].astype(float)\nfinal['ans_set_valid'] = final['ans_set_valid'].astype(float)\n\n# + id=\"rP6S8x1tjGi3\"\n# получаем таким образом итоговые значение по каждому тексту, некоторые будут с nan но их удалим позже\nfinal['subject1_final'] = np.where(\n final['ans_set_valid'] == final['ans_set_mark'], \n final['subject1_mark'],\n final['subject1_valid']\n)\nfinal['subject2_final'] = np.where(\n final['ans_set_valid'] == final['ans_set_mark'], \n final['subject2_mark'],\n final['subject2_valid']\n)\nfinal['subject3_final'] = np.where(\n final['ans_set_valid'] == final['ans_set_mark'], \n final['subject3_mark'],\n final['subject3_valid']\n)\nfinal['subject4_final'] = np.where(\n final['ans_set_valid'] == final['ans_set_mark'], \n final['subject4_mark'],\n final['subject4_valid']\n)\n\n# + id=\"wdqELPIijaTv\"\n# вот тут очищаем данные\nresult_answers = final.dropna(\n subset=['text_id', 'subject1_final', 'subject2_final', 'subject3_final', 'subject4_final']\n ).drop_duplicates(subset=['text_id']\n )[['text_id', 'subject1_final', 'subject2_final', 'subject3_final', 'subject4_final']]\n\n# + [markdown] id=\"Z70eedh5nK51\"\n# ## Финальная витрина правильных ответов\n\n# + id=\"TPPzYMS0nPvA\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 173} outputId=\"78aba180-efdc-4a2e-9908-20aa7a88d276\"\nresult_answers\n\n# + [markdown] id=\"QbnEdJoknQyF\"\n# # Решаем задачу №2 - считаем среднюю точность разметчиков\n\n# + id=\"v1lyme3XjFkb\"\nquality = markup.merge(result_answers, how='inner', on='text_id') # соединиям данные и маркап, чтобы потом посчитать качество разметчиков\n\n# + id=\"BFjutYE4mQDD\"\n# ставим один если совпал ответ маркапера и итогового значения после валидации и 0 если не совпали\nquality['subject1_right'] = np.where(quality['subject1'] == quality['subject1_final'], 1, 0)\nquality['subject2_right'] = np.where(quality['subject2'] == quality['subject2_final'], 1, 0)\nquality['subject3_right'] = np.where(quality['subject3'] == quality['subject3_final'], 1, 0)\nquality['subject4_right'] = np.where(quality['subject4'] == quality['subject4_final'], 1, 0)\n\n# + id=\"BwR3rbknnlfk\"\n# делим на 4 чтобы получить процент правильных ответов\nquality['score'] = (\n (quality['subject1_right'] + quality['subject2_right'] + quality['subject3_right'] + quality['subject4_right']) / 4\n)\n\n# + id=\"_NGyGlfSmfJo\"\nresult = quality.merge(valid[['text_id', 'ans_set']], \n how='left', \n on='text_id', \n suffixes=['_mark', '_valid'])\n\n# + id=\"FdJygJy1rw2g\"\nresult['ans_set_mark'] = result['ans_set_mark'].astype(str)\n\n# + id=\"tEtmvgZ1r0o8\"\nresult['ans_set_valid'].fillna('0', inplace=True)\n\n# + id=\"5fEm4sBTso35\"\n# проверяем выбирал ли валидатор значения валидатор как правильные у более чем 1 разметчика, если выбрал, то ставим 1 в графу эффективность\nresult['score'] = np.where(\n result.apply(lambda x: x.ans_set_mark in x.ans_set_valid, axis=1), # если ответ разметчика валидатор посчитал верным, то ставим 1\n 1, result['score']\n)\n\n# + id=\"0LErwtXVrwBk\"\nmarkupers_df = result.groupby('markuper')['score'].mean().reset_index() # итоговые значения эффективности разметк для markuper\n\n# + [markdown] id=\"z2t1t90HnqWg\"\n# ## Финальный ответ по задаче №2\n\n# + id=\"1yFSxtDIns9Q\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 142} outputId=\"c0c46e72-f16f-420d-fd4c-fcef8743c03c\"\nmarkupers_df\n\n# + id=\"aWMbmLBws2M4\"\nmarkupers_df.to_csv('markupers_quality.csv', index=False)\n","repo_name":"rgimanov/test_tasks","sub_path":"СБЕР/test2/final_result_test_2.ipynb","file_name":"final_result_test_2.ipynb","file_ext":"py","file_size_in_byte":8900,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"50"}
+{"seq_id":"7116646245","text":"# ## Vector Plots in Physics\n#\n# Seems like this should be pretty straight forward but who knows.....\n\n# ### Dependencies\n#\n# The usual...\n\nimport numpy as np\nimport matplotlib.pyplot as plt\n\n\n# ### Example\n#\n# Once you know what the call for a quiver plot expects it seems like it should be pretty direct....\n# [quiver documentation](https://matplotlib.org/api/_as_gen/matplotlib.axes.Axes.quiver.html)\n#\n# ```ax.quiver(0,0,1,1, units = 'xy', scale = 1)``` \n# In the previous expression the first pair of arguments is the origin of the vector. The next two are the x and y components repectively. Then there are the usual \\**kwargs options for color and units etc. \n#\n# ```ax.quiver((0,1), (0,0), (1,0), (1,2.5), units = 'xy', scale = 1)``` \n# In this expression the same system applies but two vectors are being plotted. The first argument is the x value of the origin for the first vector and the x value of the origin for the second vector. The arguments to the quiver call have the same meaning but they can be tuples that quiver interprets sequentially.\n\nfig, ax = plt.subplots()\n#fig = plt.figure()\n#ax = fig.add_subplot()\n#ax.quiver(0,0,2.4,-3.1,\n# color = \"green\", \n# units = 'xy', \n# scale = 1, \n# width = .1)\nax.quiver((0,1), (0,0), (1,0), (1,2.3), color = (\"red\",\"green\"),units = 'xy', scale = 1)\n#ax.quiver((0,0), (-2,-3), units = 'xy', scale = 1)\nplt.axis('equal')\nplt.xticks(range(-5,6))\nplt.yticks(range(-5,6))\nplt.grid()\nplt.show()\n\n# ### Components of a vector\n#\n# Let's see if this works...\n\n# +\n# vector\nvectorcomp = [2.4, 3.3]\nvectorlocate = [0.,0.]\n\n# color choices and font size\ncolorcomp = \"red\"\ncolorresult = \"green\"\nplt.rcParams.update({'font.size': 16})\n\n# plot figure\nfig1, ax1 = plt.subplots()\nax1.quiver((vectorlocate[0],vectorcomp[0],vectorlocate[0]),\n (vectorlocate[1],vectorlocate[1],vectorlocate[1]),\n (vectorcomp[0], 0, vectorcomp[0]),\n (0.,vectorcomp[1],vectorcomp[1]),\n color = (colorcomp,colorcomp,colorresult), \n units = 'xy', \n scale = 1, \n width = .1)\n\n# locate labels\noffset = .5 # this will need to scale depending on problem\nxcomp_xlabel = (vectorcomp[0] - vectorlocate[0])/2\nxcomp_ylabel = vectorlocate[0]- offset\nycomp_xlabel = vectorcomp[0] + offset\nycomp_ylabel = (vectorcomp[1] - vectorlocate[1])/2\nresult_xlabel = xcomp_xlabel - offset\nresult_ylabel = ycomp_ylabel + offset\n\nplt.text(xcomp_xlabel,xcomp_ylabel, \"Ax\", color = colorcomp)\nplt.text(ycomp_xlabel, ycomp_ylabel, \"Ay\", color = colorcomp)\nplt.text(result_xlabel,result_ylabel, \"A\", color = colorresult)\n\nplt.axis('equal')\nplt.xlim(-3,6)\nplt.ylim(-3,6)\n#plt.xticks(range(-5,6))\n#plt.yticks(range(-5,6))\nplt.grid()\nplt.show()\n# -\n\n\n\n\n\n\n","repo_name":"smithrockmaker/JupyterLabReference","sub_path":"VectorQuiver.ipynb","file_name":"VectorQuiver.ipynb","file_ext":"py","file_size_in_byte":2746,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"33351261367","text":"# +\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nsns.set(style=\"whitegrid\")\n\n#Define Dataset specific missing value\nmissing_values = [\"NaN\", \"#NAME?\"]\n\n#Read data with missing value handling\ndf = pd.read_csv(\"googleplaystore.csv\", na_values = missing_values)\n# -\n\n#Get data set info\nprint(df.head(3))\nprint()\nprint(df.info())\n\n# +\n# Rating\ndf[\"Rating\"] = pd.to_numeric(df[\"Rating\"])\ndf.Rating.describe().round(2)\n\nfigure = plt.figure(figsize=(9,6))\naxes = figure.add_subplot(1,1,1)\naxes.hist(df.Rating,color='Steelblue')\naxes.set_title(\"App Rating Distribution\")\naxes.set_xlabel(\"Rating\")\naxes.set_ylabel(\"Counts\")\naxes.yaxis.grid( b=True, which=\"major\")\naxes.xaxis.grid(False)\n\nplt.show()\nplt.close()\n\n\n# +\n# Category\ndf.Category.describe()\npd.DataFrame(df.Category.value_counts( normalize=True).sort_index())\n\ncounts = df.Category.value_counts().sort_index()\nx = range( len( counts))\nwidth = 1/1.5\nfigure = plt.figure(figsize=(12,12))\n\naxes = figure.add_subplot(1, 1,1)\naxes.barh(x, counts, width, color=\"steelblue\", align=\"center\")\naxes.set_yticks(range( len( counts)))\naxes.set_yticklabels(counts.axes[0])\naxes.set_title( \"Category Counts\")\naxes.set_xlabel( \"Counts\")\naxes.yaxis.grid(False)\n\n\n# +\n# Reviews\n\nfigure = plt.figure(figsize=(10, 6))\naxes = figure.add_subplot(1, 1, 1)\naxes.boxplot(df.Reviews, patch_artist=True, zorder=1, vert=False)\naxes.set_xlabel(\"Reviews\")\naxes.set_title(\"Boxplot - Reviews\")\nplt.show()\nplt.close()\n\n\n# +\n# Rating By Category\n\ngrouped = df.groupby( \"Category\")\ngrouped_y = grouped[\"Rating\"].describe().round(2)\nprint(grouped_y)\n\n","repo_name":"gwang39/Social-Media-Analytics","sub_path":"A7- EDA/.ipynb_checkpoints/A7gwang-checkpoint.ipynb","file_name":"A7gwang-checkpoint.ipynb","file_ext":"py","file_size_in_byte":1595,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"20007747706","text":"# # Interactive Accumulator Models with `Evidently`\n#\n# This notebook contains interactive visualisations for a number of evidence models\n# created using the [evidently](https://github.com/EoinTravers/Evidently) package for python.\n#\n# If you're running this notebook on [mybinder.org](https://mybinder.org/),\n# and find things to be a little slow,\n# you can run the code on your own machine instead by downloading the \n# [GitHub repository](https://github.com/EoinTravers/Evidently).\n#\n# If you're unfamiliar with [Jupyter notebooks](https://jupyter.org/),\n# you can:\n# - Run each code cell by clicking on it to select it,\n# then clicking the \n# [ Run]\n# button in the toolbar above.\n# - Run everything by clicking [Cell] → [Run All]\n#\n# Either way, some of the cells below will generate interactive plots with sliders attached.\n# Go ahead and play with the sliders, and see what happens!\n\n# +\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport matplotlib as mpl\nimport evidently\n\nfrom ipywidgets import interact, FloatSlider\ndef fs(v, low, high, step, desc=''):\n return FloatSlider(value=v, min=low, max=high, step=step, description=desc, continuous_update=False)\n\n\n# -\n\n# # Simple Wald Model\n#\n\nprint(evidently.Wald.__doc__) # Print the documentation for this model\n\n\n# +\ndef wald_simulation_plot(t0=1., v=.5, z=0, a=1., c=1.):\n model = evidently.Wald(pars=[t0, v, z, a, c], max_time=5)\n evidently.viz.visualise_model(model, model_type='wald', measure='raw')\n title = 'Start time = %.1f, Drift = %.1f, Start = %.1f, Threshold = %.1f' % (t0, v, z, a)\n plt.suptitle(title, y=1.01)\n plt.show()\n\ninteract(wald_simulation_plot,\n t0 = fs(1., 0, 5., .1, 'Start time'),\n v = fs(.5, 0, 2., .1, 'Drift'),\n z = fs(0, 0, .5, .1, 'Starting point'),\n a = fs(.5, 0., 2., .1, 'Threshold'),\n c = fs(.1, 0., 2., .1, 'Noise SD'));\n# -\n\n# # Simple Drift Diffusion Model\n#\n\nprint(evidently.Diffusion.__doc__)\n\n\n# +\ndef ddm_simulation_plot(t0=1., v=.5, z=0., a=.5, c=.1):\n model = evidently.Diffusion(pars=[t0, v, z, a, c])\n evidently.viz.visualise_model(model)\n title = 't0 = %.1f, Drift = %.1f, Bias = %.1f, Threshold = %.1f' % (t0, v, z, a)\n plt.suptitle(title, y=1.01)\n plt.show()\n\ninteract(ddm_simulation_plot,\n t0 = fs(1., 0, 2., .1, 't0'),\n v = fs(.5, 0, 2., .1, 'Drift'),\n z = fs(-.5, -1., 1., .1, 'Bias'),\n a = fs(.5, 0., 2., .1, 'Threshold'),\n c = fs(.2, 0., 1., .1, 'Noise SD'));\n# -\n\n# # Hierarchical Drift Diffusion Model\n\nprint(evidently.HDiffusion.__doc__)\n\n\n# +\ndef hddm_simulation_plot(m_t0=1., m_v=.5, m_z=0., m_a=.5,\n s_t0=.1, s_v=.1, s_z=.1, s_a=.1, c=.1):\n model = evidently.HDiffusion(pars=[m_t0, m_v, m_z, m_a, s_t0, s_v, s_z, s_a, c])\n evidently.viz.visualise_model(model, measure='means', threshold=m_a)\n title = 't0 = %.1f (%.1f), Drift = %.1f (%.1f), Bias = %.1f (%.1f), Threshold = %.1f (%.1f), Noise = %.1f' % (\n m_t0, s_t0, m_v, s_v, m_z, s_z, m_a, s_a, c)\n plt.suptitle(title, y=1.05)\n plt.show()\n\ninteract(hddm_simulation_plot,\n m_t0 = fs(1., 0., 1., .1, 'T0'),\n m_v = fs(.5, -1., 1., .1, 'Drift'),\n m_z = fs(-.5, -1., 1., .1, 'Bias'),\n m_a = fs(1., 0., 2., .1, 'Threshold'),\n s_t0 = fs(.1, 0., 1., .1, 'SD T0'),\n s_v = fs(.5, 0., 1., .1, 'SD Drift'),\n s_z = fs(.1, 0., 1., .1, 'SD Bias'),\n s_a = fs(.1, 0., 1., .1, 'SD Threshold'),\n c = fs(.1, 0., 1., .1, 'Noise'));\n# -\n\n# # Race\n\nprint(evidently.Race.__doc__)\n\n\n# +\ndef race_simulation_plot(t1, v1, z1, c1, t2, v2, z2, c2, a):\n model = evidently.Race(pars=[t1, v1, z1, c1, t2, v2, z2, c2, a])\n evidently.viz.visualise_model(model, model_type='race', measure='means')\n title = 'Option #1: t0 = %.1f, Drift = %.1f, Bias = %.1f, Noise = %.1f' % (t1, v1, z1, c1)\n title += '\\nOption #2: t0 = %.1f, Drift = %.1f, Bias = %.1f, Noise = %.1f' % (t2, v2, z2, c2)\n title += '\\nThreshold: %.1f' % a \n plt.suptitle(title, y=1.2)\n plt.show()\n\ninteract(race_simulation_plot,\n t1 = fs(1., 0, 2., .1, '#1: t0'),\n v1 = fs(.5, 0, 2., .1, '#1: Drift'),\n z1 = fs(0., 0, 1., .1, '#1: Bias'),\n c1 = fs(.1, 0., 1., .1, '#1: Noise SD'),\n t2 = fs(2., 0, 2., .1, '#2: t0'),\n v2 = fs(.75, 0, 2., .1, '#2: Drift'),\n z2 = fs(.2, 0., 1., .1, '#2: Bias'),\n c2 = fs(.2, 0., 1., .1, '#2: Noise SD'),\n a = fs(.5, 0., 2., .1, 'Threshold'));\n# -\n\n# ## Schurger Model\n\nprint(evidently.models.Schurger.__doc__)\n\n\n# +\n\ndef schurger_simulation_plot(t0=0., v=.5, k=.6, z=0., a=1., c=1.):\n model = evidently.models.Schurger(pars=[t0, v, k, z, a, c], max_time=20.)\n evidently.viz.visualise_schurger(model, rp_duration=3)\n title = 'Start time = %.1f, Input = %.1f, Decay = %.1f, Start = %.1f, Threshold = %.1f, Noise = %.1f' % (\n t0, v, k, z, a, c)\n plt.suptitle(title, y=1.05)\n plt.show()\n\ninteract(schurger_simulation_plot,\n t0 = fs(0., 0, 5., .1, 'Start time'),\n v = fs(.5, 0, 2., .1, 'Drift'),\n k = fs(.6, 0, 2., .1, 'Decay'),\n z = fs(0, 0, 1., .1, 'Starting point'),\n a = fs(1., 0., 2., .1, 'Threshold'),\n c = fs(.1, 0., 1., .1, 'Noise SD'));\n","repo_name":"EoinTravers/Evidently","sub_path":"dashboards/Interactive Models.ipynb","file_name":"Interactive Models.ipynb","file_ext":"py","file_size_in_byte":5377,"program_lang":"python","lang":"en","doc_type":"code","stars":22,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"13244205381","text":"# +\nimport pandas as pd\n\nfrom sklearn.metrics import precision_score\nfrom sklearn.metrics import accuracy_score\nfrom sklearn.metrics import recall_score\nfrom sklearn.metrics import classification_report\n\nimport env\nimport acquire\nimport prepare\n\nfrom sklearn.model_selection import train_test_split\n\nfrom sklearn.tree import DecisionTreeClassifier\nfrom sklearn.tree import export_graphviz\nfrom sklearn.metrics import confusion_matrix\nfrom sklearn.ensemble import RandomForestClassifier\nfrom sklearn.linear_model import LogisticRegression\n\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport graphviz\nfrom graphviz import Graph\n# -\n\ntitanic = acquire.get_titanic_data()\ntitanic_df = titanic.copy()\ntitanic_df.head()\n\n\ndef prep_titanic(df):\n '''\n take in titanc dataframe, remove all rows where age or embarked is null, \n get dummy variables for sex and embark_town, \n and drop sex, deck, passenger_id, class, and embark_town. \n '''\n\n df = df[(df.age.notna()) & (df.embarked.notna())]\n df = df.drop(columns=['deck', 'passenger_id', 'class'])\n\n dummy_df = pd.get_dummies(df[['sex', 'embark_town']], prefix=['sex', 'embark'])\n\n df = pd.concat([df, dummy_df.drop(columns=['sex_male'])], axis=1)\n\n df = df.drop(columns=['sex', 'embark_town', 'embarked']) \n\n df = df.rename(columns={\"sex_female\": \"is_female\"})\n\n return df\n\n\ntitanic_df = prep_titanic(titanic_df)\ntitanic_df.head()\n\n\ndef train_validate_test_split(df, target):\n '''\n This function takes in a dataframe, the name of the target variable\n (for stratification purposes)\n and splits the data into train, validate and test. \n Test is 20% of the original dataset, validate is .25*.90= 22.5% of the \n original dataset, and train is .70*.80= 56% of the original dataset. \n The function returns, in this order, train, validate and test dataframes. \n '''\n train_validate, test = train_test_split(df, test_size=0.2, \n stratify=df[target])\n \n \n train, validate = train_test_split(train_validate, test_size=0.3, \n stratify=train_validate[target])\n return train, validate, test\n\ntrain, validate, test = train_validate_test_split(titanic_df, target = 'survived')\n\ntrain.shape, validate.shape, test.shape\n\ntrain.info()\n\n# +\ntrain['baseline1'] = 0\n\nvalidate['baseline1'] = 0\n\ntest['baseline1'] = 0\n# -\n\ntrain.survived.value_counts()\n\n\n\n# +\nbaseline_accuracy = (train.survived == train.baseline1).mean()\nprint(f'baseline accuracy: {baseline_accuracy:.2%}')\n\nbaseline_accuracy = (validate.survived == validate.baseline1).mean()\nprint(f'baseline accuracy: {baseline_accuracy:.2%}')\n\nbaseline_accuracy = (test.survived == test.baseline1).mean()\nprint(f'baseline accuracy: {baseline_accuracy:.2%}')\n# -\n\n\n\n# +\nX_train = train.drop(columns=['survived', 'baseline1'])\ny_train = train.survived\n\nX_validate = validate.drop(columns=['survived', 'baseline1'])\ny_validate = validate.survived\n\nX_test = test.drop(columns=['survived', 'baseline1'])\ny_test = test.survived\n# -\n\nclf = DecisionTreeClassifier(max_depth=3)\n\nclf2 = DecisionTreeClassifier(max_depth=8)\n\nclf = clf.fit(X_train, y_train)\n\nclf2 = clf2.fit(X_train, y_train)\n\n# +\ndot_data = export_graphviz(clf, feature_names= X_train.columns, rounded=True, filled=True, out_file=None)\ngraph = graphviz.Source(dot_data) \n\ngraph.render('titanic_decision_tree', view=True)\n\n# +\ndot_data = export_graphviz(clf2, feature_names= X_train.columns, rounded=True, filled=True, out_file=None)\ngraph = graphviz.Source(dot_data) \n\ngraph.render('titanic_decision_tree', view=True)\n# -\n\ny_pred2 = clf2.predict(X_train)\ny_pred2[0:4]\n\ny_pred = clf.predict(X_train)\ny_pred[0:4]\n\ny_pred_proba = clf.predict_proba(X_train)\ny_pred_proba[0:4]\n\ny_pred_proba2 = clf2.predict_proba(X_train)\ny_pred_proba2[0:4]\n\nprint('Accuracy of Decision Tree classifier on training set: {:.2f}'\n .format(clf.score(X_train, y_train)))\n\nprint('Accuracy of Decision Tree classifier on training set: {:.2f}'\n .format(clf2.score(X_train, y_train)))\n\n# +\n# confusion matrix\n\nconfusion_matrix(y_train, y_pred)\n\n# +\nprint(\"First Descision Tree Model\")\nprint(\"-------------------------------\")\n#TP/TP+FN\nprint(\"True Positive Rate: \", round((193/ (193+21)), 2))\n\n#FP / FP+TN\nprint(\"False Positive Rate: \", round((44 / (44 + 140)), 2))\n\n#TN / TN+FP\nprint(\"True Negative Rate: \", round((140 / (140+44)), 2))\n\n#FN / FN + TP\nprint(\"False Negative Rate: \", round((21 / (21+193)), 2))\n\n# +\n# confusion matrix\n\nconfusion_matrix(y_train, y_pred2)\n\n# +\nprint(\"Second Descision Tree Model\")\nprint(\"-------------------------------\")\n#TP/TP+FN\nprint(\"True Positive Rate: \", round((236/ (236+25)), 2))\n\n#FP / FP+TN\nprint(\"False Positive Rate: \", round((1 / (1 + 136)), 2))\n\n#TN / TN+FP\nprint(\"True Negative Rate: \", round((136/ (136+1)), 2))\n\n#FN / FN + TP\nprint(\"False Negative Rate: \", round((25/ (25+236)), 2))\n# -\n\ny_train.value_counts()\n\nlabels = sorted(y_train.unique())\nprint('Actual on the left, predicted on the top')\npd.DataFrame(confusion_matrix(y_train, y_pred), index=labels, columns=labels)\n\nlabels = sorted(y_train.unique())\nprint('Actual on the left, predicted on the top')\npd.DataFrame(confusion_matrix(y_train, y_pred2), index=labels, columns=labels)\n\nprint(classification_report(y_train, y_pred))\n\nprint(classification_report(y_train, y_pred2))\n\n# # Evaluate Validate\n\nprint('Accuracy of Decision Tree classifier on validate set: {:.2f}'\n .format(clf.score(X_validate, y_validate)))\n\nprint('Accuracy of Decision Tree classifier on validate set: {:.2f}'\n .format(clf2.score(X_validate, y_validate)))\n\n# +\n# And since accuracy isn't everything\n\n# Produce y_predictions that come from the X_validate\ny_pred = clf.predict(X_validate)\n\n# Compare actual y values (from validate) to predicted y_values from the model run on X_validate\nprint(classification_report(y_validate, y_pred))\n\n\n# +\n# And since accuracy isn't everything\n\n# Produce y_predictions that come from the X_validate\ny_pred2 = clf2.predict(X_validate)\n\n# Compare actual y values (from validate) to predicted y_values from the model run on X_validate\nprint(classification_report(y_validate, y_pred2))\n\n# -\n\n# # Exercises: random forest\n# Continue working in your model file with titanic data to do the following:\n#\n# Fit the Random Forest classifier to your training sample and transform (i.e. make predictions on the training sample) setting the random_state accordingly and setting min_samples_leaf = 1 and max_depth = 10.\n#\n# Evaluate your results using the model score, confusion matrix, and classification report.\n#\n# Print and clearly label the following: Accuracy, true positive rate, false positive rate, true negative rate, false negative rate, precision, recall, f1-score, and support.\n#\n# Run through steps increasing your min_samples_leaf and decreasing your max_depth.\n#\n# What are the differences in the evaluation metrics? Which performs better on your in-sample data? Why?\n#\n# After making a few models, which one has the best performance (or closest metrics) on both train and validate?\n\nclf3 = RandomForestClassifier(max_depth=10, min_samples_leaf=1)\n\nclf3 = clf3.fit(X_train, y_train)\n\nprint(clf3.feature_importances_)\n\ny_pred = clf3.predict(X_train)\ny_pred\n\ny_pred_proba = clf3.predict_proba(X_train)\ny_pred_proba\n\nprint('Accuracy of random forest classifier on training set: {:.2f}'\n .format(clf3.score(X_train, y_train)))\n\nprint(confusion_matrix(y_train, y_pred))\n\n# +\nTP = 236\nFP = 1\nFN = 10\nTN = 151\n\nprint(\"Random Forest Model\")\nprint(\"-------------------------------\")\n#TP/TP+FN\nprint(\"True Positive Rate: \", round((TP/ (TP+FN)), 2))\n\n#FP / FP+TN\nprint(\"False Positive Rate: \", round((FP / (FP + TN)), 2))\n\n#TN / TN+FP\nprint(\"True Negative Rate: \", round((TN/ (TN+FP)), 2))\n\n#FN / FN + TP\nprint(\"False Negative Rate: \", round((FN/ (FN+TP)), 2))\n# -\n\nprint(classification_report(y_train, y_pred))\n\n# ### Chaning max_depth and min_samples_leaf parameters\n\n# +\nclf4 = RandomForestClassifier(max_depth=5, min_samples_leaf=3)\nclf4 = clf4.fit(X_train, y_train)\n\ny_pred2 = clf4.predict(X_train)\n#y_pred\ny_pred_proba2 = clf4.predict_proba(X_train)\n#y_pred_proba2\n\nprint('Accuracy of random forest classifier on training set: {:.2f}'\n .format(clf4.score(X_train, y_train)))\n\nprint(\"---------------------------------------------------------\")\nprint(\"---------------------------------------------------------\\n\")\nprint(confusion_matrix(y_train, y_pred2))\n\nprint(\"---------------------------------------------------------\")\nprint(\"---------------------------------------------------------\\n\")\nprint(classification_report(y_train, y_pred2))\n\n# +\nTP = 225\nFP = 12\nFN = 37\nTN = 124\n\nprint(\"Random Forest Model\")\nprint(\"-------------------------------\")\n#TP/TP+FN\nprint(\"True Positive Rate: \", round((TP/ (TP+FN)), 2))\n\n#FP / FP+TN\nprint(\"False Positive Rate: \", round((FP / (FP + TN)), 2))\n\n#TN / TN+FP\nprint(\"True Negative Rate: \", round((TN/ (TN+FP)), 2))\n\n#FN / FN + TP\nprint(\"False Negative Rate: \", round((FN/ (FN+TP)), 2))\n\n# +\n\n\nclf4 = RandomForestClassifier(max_depth=5, min_samples_leaf=3)\nclf4 = clf4.fit(X_train, y_train)\n\ny_pred2 = clf4.predict(X_train)\n#y_pred\ny_pred_proba2 = clf4.predict_proba(X_train)\n#y_pred_proba2\n\nprint('Accuracy of random forest classifier on training set: {:.2f}'\n .format(clf4.score(X_train, y_train)))\n\nprint(\"---------------------------------------------------------\")\nprint(\"---------------------------------------------------------\\n\")\nprint(confusion_matrix(y_train, y_pred2))\n\nprint(\"---------------------------------------------------------\")\nprint(\"---------------------------------------------------------\\n\")\nprint(classification_report(y_train, y_pred2))\n# -\n\nfor i in range(10, 0, -1):\n print(i)\n\nfor i in range(3,10):\n for n in range(7, 1, -1):\n clf4 = RandomForestClassifier(max_depth=n, min_samples_leaf=i)\n clf4 = clf4.fit(X_train, y_train)\n\n y_pred2 = clf4.predict(X_train)\n #y_pred\n y_pred_proba2 = clf4.predict_proba(X_train)\n #y_pred_proba2\n\n print('Accuracy of random forest classifier on training set: {:.2f}'\n .format(clf4.score(X_train, y_train)))\n\n print(\"---------------------------------------------------------\")\n print(\"---------------------------------------------------------\\n\")\n print(confusion_matrix(y_train, y_pred2))\n\n print(\"---------------------------------------------------------\")\n print(\"---------------------------------------------------------\\n\")\n print(classification_report(y_train, y_pred2))\n \n\npd.set_option('display.max_rows', 10)\n\n# +\n# Let's continue getting loopy, so we can compare in-sample to out-of-sample\nmetrics = []\nnumber_depths = 20\n\nfor i in range(1,number_depths):\n \n \n depth = number_depths - i\n \n clf4 = RandomForestClassifier(max_depth = depth, min_samples_leaf = i)\n clf4 = clf4.fit(X_train, y_train)\n\n y_pred2 = clf4.predict(X_train)\n #y_pred\n y_pred_proba2 = clf4.predict_proba(X_train)\n #y_pred_proba2\n\n in_sample_accuracy = clf4.score(X_train, y_train)\n \n out_of_sample_accuracy = clf4.score(X_validate, y_validate)\n\n output = {\n \"max_depth\": depth,\n \"min_sample_leaf\": i,\n \"train_accuracy\": in_sample_accuracy,\n \"validate_accuracy\": out_of_sample_accuracy\n }\n \n metrics.append(output)\n \n \ndf = pd.DataFrame(metrics)\ndf[\"difference\"] = df.train_accuracy - df.validate_accuracy\n#df.sort_values(by = ['max_depth', 'min_sample_leaf'], ascending = [False, True])\ndf\n# -\n\ndf.sort_values(by = ['max_depth', 'min_sample_leaf'], ascending = [False, True])\n\ndf1 = df.sort_values(by = ['max_depth', 'min_sample_leaf'], ascending = [True, False])\n\ndf1.head(10)\n\ndf1.set_index('max_depth')[['train_accuracy', 'validate_accuracy', 'difference']].plot()\n\ndf[df.difference <= 0.05].sort_values(by=['validate_accuracy', 'difference'], ascending = [False,True])\n\n# # KNN Exercises\n\n# 1. Fit a K-Nearest Neighbors classifier to your training sample and transform (i.e. make predictions on the training sample)\n#\n# 2. Evaluate your results using the model score, confusion matrix, and classification report.\n#\n# 3. Print and clearly label the following: Accuracy, true positive rate, false positive rate, true negative rate, false negative rate, precision, recall, f1-score, and support.\n#\n# 4. Run through steps 2-4 setting k to 10\n#\n# 5. Run through setps 2-4 setting k to 20\n#\n# 6. What are the differences in the evaluation metrics? Which performs better on your in-sample data? Why?\n#\n# 7. Which model performs best on our out-of-sample data from validate?\n\nfrom sklearn.neighbors import KNeighborsClassifier\n\ntitanic = acquire.get_titanic_data()\ntitanic_df = titanic.copy()\ntitanic_df.head()\n\n\ndef prep_titanic(df):\n '''\n take in titanc dataframe, remove all rows where age or embarked is null, \n get dummy variables for sex and embark_town, \n and drop sex, deck, passenger_id, class, and embark_town. \n '''\n\n df = df[(df.age.notna()) & (df.embarked.notna())]\n df = df.drop(columns=['deck', 'passenger_id', 'class'])\n\n dummy_df = pd.get_dummies(df[['sex', 'embark_town']], prefix=['sex', 'embark'])\n\n df = pd.concat([df, dummy_df.drop(columns=['sex_male'])], axis=1)\n\n df = df.drop(columns=['sex', 'embark_town', 'embarked']) \n\n df = df.rename(columns={\"sex_female\": \"is_female\"})\n\n return df\n\n\ntitanic_df = prep_titanic(titanic_df)\n\n\ndef train_validate_test_split(df, target):\n '''\n This function takes in a dataframe, the name of the target variable\n (for stratification purposes)\n and splits the data into train, validate and test. \n Test is 20% of the original dataset, validate is .25*.90= 22.5% of the \n original dataset, and train is .70*.80= 56% of the original dataset. \n The function returns, in this order, train, validate and test dataframes. \n '''\n train_validate, test = train_test_split(df, test_size=0.2, \n stratify=df[target])\n \n \n train, validate = train_test_split(train_validate, test_size=0.3, \n stratify=train_validate[target])\n return train, validate, test\n\ntrain, validate, test = train_validate_test_split(titanic_df, target = 'survived')\n\ntrain.shape\n\n# +\ntrain['baseline1'] = 0\n\nvalidate['baseline1'] = 0\n\ntest['baseline1'] = 0\n\n# +\nX_train = train.drop(columns=['survived', 'baseline1'])\ny_train = train.survived\n\nX_validate = validate.drop(columns=['survived', 'baseline1'])\ny_validate = validate.survived\n\nX_test = test.drop(columns=['survived', 'baseline1'])\ny_test = test.survived\n# -\n\nknn = KNeighborsClassifier()\n\nknn.fit(X_train, y_train)\n\ny_pred = knn.predict(X_train)\n\n# +\naccuracy = knn.score(X_train, y_train)\n\nprint(\"This is the accuracy: \", round(accuracy, 3))\n# -\n\nprint(classification_report(y_train, y_pred))\n\nprint(confusion_matrix(y_train, y_pred))\n\nprint(knn.score(X_train, y_train))\n\n# +\nTP= 189\nTN = 113\nFP = 48\nFN = 48\n\nALL = TP + TN + FP + FN\n\n\n\nprint(\"KNN Model\")\nprint(\"-------------------------------\")\n#TP/TP+FN\nprint(\"True Positive Rate: \", round((TP/ (TP+FN)), 2))\n\n#FP / FP+TN\nprint(\"False Positive Rate: \", round((FP / (FP + TN)), 2))\n\n#TN / TN+FP\nprint(\"True Negative Rate: \", round((TN/ (TN+FP)), 2))\n\n#FN / FN + TP\nprint(\"False Negative Rate: \", round((FN/ (FN+TP)), 2))\n\naccuracy = (TP + TN)/ALL\nprecision = TP/(TP+FP) \nrecall = TP/(TP+FN)\nprint(\"-------------------------------\")\nprint(\"-------------------------------\")\n\nprint(\"Accuracy : \", round((accuracy), 2))\nprint(\"Precision : \", round((precision), 2))\nprint(\"Recall : \", round((recall), 2))\n\nf1_score = 2*(precision*recall)/(precision+recall) \nsupport_pos = TP + FN \nsupport_neg = FP + TN \nprint(\"-------------------------------\")\nprint(\"-------------------------------\")\n\nprint(\"F1_Score : \", round((f1_score), 2))\nprint(\"Support Positive : \", support_pos)\nprint(\"Support Negative : \", support_neg)\n\n# +\nknn2 = KNeighborsClassifier(10)\nknn2.fit(X_train, y_train)\ny_pred = knn2.predict(X_train)\n\nprint(classification_report(y_train, y_pred))\nprint(\"------------------------------------\")\nprint(knn2.score(X_train, y_train))\n\n\n# +\nknn3 = KNeighborsClassifier(20)\nknn3.fit(X_train, y_train)\ny_pred = knn3.predict(X_train)\n\n\nprint(classification_report(y_train, y_pred))\nprint(\"------------------------------------\")\nprint(knn3.score(X_train, y_train))\n\n# +\nmetrics = []\n\n# loop through different values of k\nfor k in range(1, 21):\n \n # define the thing\n knn = KNeighborsClassifier(n_neighbors=k)\n \n # fit the thing (remmeber only fit on training data)\n knn.fit(X_train, y_train)\n \n # use the thing (calculate accuracy)\n train_accuracy = knn.score(X_train, y_train)\n validate_accuracy = knn.score(X_validate, y_validate)\n \n output = {\n \"k\": k,\n \"train_accuracy\": train_accuracy,\n \"validate_accuracy\": validate_accuracy\n }\n \n metrics.append(output)\n\n# make a dataframe\nresults = pd.DataFrame(metrics)\n\n# plot the data\nresults.set_index('k').plot(figsize = (16,9))\nplt.ylabel('Accuracy')\nplt.xticks(np.arange(0,21,1))\nplt.grid()\n# -\n\n# # Regression Exercises: \n\ntitanic_df = acquire.get_titanic_data()\ntitanic_df.head()\n\n# +\n# Handle missing ages\navg_age = titanic_df.age.mean()\ntitanic_df.age = titanic_df.age.fillna(avg_age)\n\n# Encode the gender column\ntitanic_df[\"is_female\"] = (titanic_df.sex == \"female\").astype('int')\n\n# Encode the embarked_town\n# Embark_Town values are Southampton, Cherbourg, and Queenstown\ndummy_df = pd.get_dummies(titanic_df[['embark_town']], dummy_na=False, drop_first=True)\ntitanic_df = pd.concat([titanic_df, dummy_df], axis=1)\n\n# Drop unnecessary columns\ntitanic_df = titanic_df.drop(columns=[\"passenger_id\", \"deck\", \"class\", \"embarked\", \"sex\", \"embark_town\"])\n\ntitanic_df.head(3)\n# -\n\ntrain, validate, test = prepare.split_data(titanic_df)\n\ntrain.head()\n\n# +\nX_train = train.drop(columns=[\"survived\"])\ny_train = train.survived\n\nX_validate = validate.drop(columns=[\"survived\"])\ny_validate = validate.survived\n\nX_test = test.drop(columns=[\"survived\"])\ny_test = test.survived\n# -\n\nbaseline_accuracy = (train.survived == 0).mean()\n\n# +\nlogit = LogisticRegression()\n\nfeatures = [\"age\", \"pclass\", \"fare\"]\n\nlogit.fit(X_train[features], y_train)\n\ny_pred = logit.predict(X_train[features])\n\nprint(\"Baseline is\", round(baseline_accuracy, 2))\nprint(\"Logistic Regression using age, pclass, and fare features\")\nprint(\"Accuracy of Logistic Regression classifier on training set: \", \n round(logit.score(X_train[features], y_train), 2))\n\n# +\nlogit1 = LogisticRegression()\n\nfeatures = [\"age\", \"pclass\", \"fare\", \"is_female\"]\n\nlogit1.fit(X_train[features], y_train)\n\ny_pred = logit1.predict(X_train[features])\n\nprint(\"Logistic Regression using age, pclass, fare, and gender features\")\nprint(\"Accuracy of Logistic Regression classifier on training set: \", \n round(logit1.score(X_train[features], y_train), 2))\n\n# +\nlogit2 = LogisticRegression()\n\nlogit2.fit(X_train, y_train)\n\ny_pred = logit2.predict(X_train)\n\nprint(\"Model trained on all features\")\nprint(\"Accuracy of Logistic Regression classifier on training set: \",\n (logit2.score(X_train, y_train)))\n\n# +\nfeatures = [\"age\", \"pclass\", \"fare\"]\n\ny_pred = logit.predict(X_validate[features])\n\nprint(classification_report(y_validate, y_pred))\n\n# +\nfeatures = [\"age\", \"pclass\", \"fare\", \"is_female\"]\n\ny_pred = logit1.predict(X_validate[features])\n\nprint(classification_report(y_validate, y_pred))\n\n# +\ny_pred = logit2.predict(X_validate)\n\nprint(classification_report(y_validate, y_pred))\n\n# +\n# Logit 1 regression has the best results\n","repo_name":"PGuajardo/classification-exercises","sub_path":"model.ipynb","file_name":"model.ipynb","file_ext":"py","file_size_in_byte":19666,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"12639221775","text":"import pandas as pd\nimport geopandas as gpd\nfrom shapely.geometry import Point\nfrom geopandas.tools import geocode\nimport os\n\n# +\n#IMPORTANDO CSV COM OS DADOS DOS PROJETOS LANÇADOS DE 2021\n# encoding='utf-8'\n# encoding='latin1'\n# encoding='utf8'\n# encoding='iso-8859-1'\n\narquivo = 'projetos-1.csv'\nrelacaoProjetos = pd.read_csv(arquivo, encoding='utf8', delimiter=';')\nrelacaoProjetos.columns\n# -\n\n#Simplifica a tabela com o esencial (Numero de identificação do projeto, Endereço(Logradouro, numero e cidade) e o Estado)\nprojetos = relacaoProjetos[['Nº PROJ','ENDEREÇO', 'ESTADO']]\nprojetos.head(5)\n\n# +\n#Adiciona a geolocalização (latitude longitude) em formato de ponto a patir do endereço fornecido\n\ngeo = geocode(projetos['ENDEREÇO'], provider='Nominatim', user_agent='myGeocoder', timeout=1)\n# -\n\n#Junta a tabela geolocalizada a tabela com a relação de projetos e outras informações\njuntar = geo.join(relacaoProjetos)\njuntar.head\ntype(juntar)\n\n#Retira dessa tabela a coluna \"Address\" \njuntar.drop('address', axis=1, inplace=True)\n\n\nprojetosGeoreferenciados = juntar\n\n#Pega o nome do arquivo usado para procurar o endereço\nnomeArquivo = os.path.splitext(arquivo)\nnomeArquivo = nomeArquivo[0]\n\n\n# +\n#Salva o arquivo em formato GeoJSON com o nome do arquivo 'base' (arquivo de entrada + Json)\n\nsaida = nomeArquivo + '.json'\nprojetosGeoreferenciados.to_file(saida, driver='GeoJSON')\n","repo_name":"wesleyclzns/endereco-o-para-lat-long","sub_path":"latitude_longitude.ipynb","file_name":"latitude_longitude.ipynb","file_ext":"py","file_size_in_byte":1399,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"72320393099","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#  \n\n# + [markdown] id=\"CBE1c8fzOncZ\"\n# #Semantic Similarity\n#\n# Implementation of a semantic similarity score between texts using the [Universal Sentence Encoder](https://github.com/tensorflow/hub/blob/master/examples/colab/semantic_similarity_with_tf_hub_universal_encoder.ipynb). The Universal Sentence Encoder is employed to produce an embedding of the texts to be compared, and the similarity is measured by the inner product between the embeddings.\n#\n#\n\n# + [markdown] id=\"sR0MU_EQPxob\"\n# # Python class to compute semantic similarity score\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} cellView=\"form\" id=\"y9L_Eu7QlIMz\" outputId=\"98bea815-e351-4922-e118-be8dee03a9b7\"\n#@title double-click to see the code\n\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns\nimport tensorflow_hub as hub\n\nfrom absl import logging\nfrom dataclasses import dataclass\n\nsns.set()\n\nlogging.set_verbosity(logging.ERROR)\n\n@dataclass\nclass TextSim:\n ''' Class to compute semantic similarity among two list of strings'''\n module_url = \"https://tfhub.dev/google/universal-sentence-encoder/4\"\n model = hub.load(module_url)\n\n @staticmethod\n def clean(input):\n '''Convert inputs to list of string elements'''\n input = input if isinstance(input, list) else [input]\n return [str(element) for element in input] \n\n def embed(self, input):\n '''Convert text to embedded representation'''\n return self.model(input) \n\n def plot_all_results(self, inputA, inputB, corr):\n for i, itemA in enumerate(inputA):\n data = pd.DataFrame({'inputB': inputB, 'corr': corr[i]})\n data = data.sort_values(by='corr').reset_index(drop=True)\n self.plot_one_result(itemA, data)\n \n @staticmethod\n def plot_one_result(itemA, data):\n fig, g = plt.subplots(figsize = (8, 1*data.shape[0]))\n g = sns.barplot(x='corr', y='inputB', data=data, palette=\"Blues\")\n g.set(xlim=(0, 1))\n g.set_xlabel(None)\n g.set_ylabel(None)\n g.set_title(f\"Similarity to '{itemA}'\")\n plt.show()\n\n def compare(self, inputA, inputB, display_score=True, return_scores=False):\n '''Compute pearson correlation among two list of inputs'''\n inputA, inputB = self.clean(inputA), self.clean(inputB)\n embedA, embedB = self.embed(inputA), self.embed(inputB)\n corr = np.inner(embedA, embedB)\n\n if display_score:\n self.plot_all_results(inputA, inputB, corr)\n \n if return_scores:\n return corr\n\nts = TextSim()\n\nprint(f'Model loaded: {ts.module_url}')\n\n# + [markdown] id=\"rYQBMhWuQGzk\"\n# # Semantic Similarity Between Words\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 229} id=\"9BiulCsGx1I2\" outputId=\"bdded523-ef88-4fad-8e35-e4ba0e3e83e3\"\ncompare_from = ['animal']\ncompare_to = ['cat', 'table', 'apple']\nts.compare(compare_from, compare_to)\n\n# + [markdown] id=\"jtmW5Omjvlq1\"\n# # Semantic Similarity Between Sentences\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 393} id=\"q96oo1t0JdiJ\" outputId=\"1db64ea8-3c72-4552-f73d-96437ae3eb26\"\ncompare_from = ['The animal walks in the zoo']\ncompare_to = ['The animal walks in the zoo', \n 'The cat walks in the zoo', \n 'The cat strolls in the zoo', \n 'My cat strolls in the zoo', \n 'My cat strolls around the zoo', \n 'My cat strolls around the park']\n \nts.compare(compare_from, compare_to)\n\n# + [markdown] id=\"he5tomszvql5\"\n# # Semantic Similarity Between Questions\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 338} id=\"89Q68cUpmBWJ\" outputId=\"0b4dbd4b-ab2b-4857-9450-15670ab487a3\"\ncompare_from = ['Which brush head fits my Philips Sonicare Toothbrush?']\ncompare_to = ['How to pick a head for my toothbrush?',\n 'toothbrush I dont know what replacement I need',\n 'I dont know what replacement I need',\n 'What head should I buy for my toothbrush',\n 'My headphones are not charging']\nts.compare(compare_from, compare_to)\n","repo_name":"ossskkar/NLP","sub_path":"Semantic_Similarity_with_Universal_Sentence_Encoder.ipynb","file_name":"Semantic_Similarity_with_Universal_Sentence_Encoder.ipynb","file_ext":"py","file_size_in_byte":4378,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"38569353035","text":"# ```\n# This notebook contains summary statistics for the drugs.com depression drug reviews dataset. \n# In particular, it contains the following:\n#\n# Summary stats:\n# - Total number of reviews\n# - Total number of reviews per rating\n# - Total number of reviews per duration\n# - Total number of unique drugs\n# - Average length of comments\n# - Average length of comments per rating\n# - Average rating\n# - Average number of upvotes\n# - Average number of upvotes per rating\n# - Highest length comment\n# - Highest upvote count review\n#\n# Subpopulations to look at:\n# - Reviews with comments longer than 300 characters (arbitrary number)\n# - Reviews with more than 15 upvotes (arbitrary number)\n# \n# Relationships of interest\n# - Length of review vs number of upvotes\n# - Length of review vs rating\n# - Upvotes vs time a review has been up (current date minus review date)\n# (so we can see upvotes per review \"duration\", essentially)\n# ```\n\nimport pandas as pd\nimport numpy as np\nfrom sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer\n# %matplotlib inline\n\ndf = pd.read_csv('./data/ddc_depression_drug_reviews.csv')\ndf = df[pd.notnull(df['genRating'])] #bye bye non-rated reviews!\ndf['comment length'] = df['comment'].apply(lambda x: len(x)) #let's look at this, too\n\n# Summary statistics for the whole dataset\ndf.describe()\n\n# The longest comment\ndf[df['comment length'] == 3000].values[0][0]\n\n# The shortest comment\ndf[df['comment length'] == 6].values[0][0]\n\n# The most upvoted comment\ndf[df['upVotes'] == 1291].values[0][0]\n\n# Not very correlated...\ndf['upVotes'].corr(df['comment length'])\n\ndf.plot.scatter(x='comment length', y='upVotes')\n\n# These aren't particularly correlated, either.\ndf['genRating'].corr(df['comment length'])\n\ncomments = df['comment'].tolist()\nratings = df['genRating'].tolist()\n\n\n# +\ndef get_wordcounts(text, vectorizer=None):\n \"\"\"\n Takes a string or list of strings and returns term frequencies for each word in\n a vectorizer's vocabulary. If no vectorizer is passed, the vocabulary will be generated\n from the input text's unigrams and bigrams.\n \"\"\"\n if vectorizer is None:\n vectorizer = CountVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\n vectorizer.fit(text)\n term_counts = zip(vectorizer.get_feature_names(),\n np.asarray(vectorizer.transform(text).sum(axis=0)).ravel())\n return sorted(term_counts, key=lambda x: x[1], reverse=True)\n\ndef get_tfidf_weights(text, vectorizer=None):\n if vectorizer is None:\n vectorizer = TfidfVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\n vectorizer.fit(text)\n tfidf_weights = zip(vectorizer.get_feature_names(),\n np.asarray(vectorizer.transform(text).sum(axis=0)).ravel())\n return sorted(tfidf_weights, key=lambda x: x[1], reverse=True)\n\n\n# +\ncount_vec = CountVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\ncount_vec.fit(comments)\n\ntfidf_vec = TfidfVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\ntfidf_vec.fit(comments)\n\ntest_counts = get_wordcounts(comments, count_vec)[:10]\ntest_tfidf = get_tfidf_weights(comments, tfidf_vec)[:10]\n# -\n\nfor x in range(1,11):\n df_slice = df[df['genRating'] == x]\n print('Summary stats for reviews with a rating of {} \\n'.format(x))\n print(df_slice.describe())\n print()\n coms = df_slice['comment'].tolist()\n top_words_tfidf = get_tfidf_weights(coms, tfidf_vec)[:10]\n print('Top words (by tf-idf) for reviews with a rating of {} \\n'.format(x))\n print(str(top_words_tfidf))\n print()\n top_words = get_wordcounts(coms, count_vec)[:10]\n print('Top words (by raw count) for reviews with a rating of {} \\n'.format(x))\n print(str(top_words))\n print()\n print('********************************************************************')\n print('********************************************************************')\n print()\n","repo_name":"ebertucc/snooprx","sub_path":"Depression-Drugs-Data-Exploration.ipynb","file_name":"Depression-Drugs-Data-Exploration.ipynb","file_ext":"py","file_size_in_byte":4056,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"39934883115","text":"import sys\nsys.path.append('/Library/Frameworks/Python.framework/Versions/3.9/lib/python3.9/site-packages')\n\nfrom os import listdir\nimport numpy as np\nimport pandas as pd\nfrom AudioFeatureExtractor import AudioFeatureExtractor\n\ngenres = ['Blues', 'Classical', 'Country', 'Disco', 'Hiphop', 'Jazz', 'Metal', 'Pop', 'Rock', 'Reggae']\ncsv_name = 'features_gtzan.csv'\ngtzan_dir = '/Users/griffinbholt/Downloads/GTZAN_split_data/'\n\n# Create the AudioFeatureExtractor Object\nafe = AudioFeatureExtractor()\n\n# Get the column names for the data\nfeature_list = afe.generate_feature_list()\n\nall_feature_data = []\n\n# Extract features from each audio file in the Blues folder\n# Done: i = 0, 1, 2, 3, 4, 5, 6, 7, 8, 9\nfor i in range(10):\n folder_path = gtzan_dir + genres[i].lower()\n audio_files = listdir(folder_path)\n\n feature_data_for_this_genre = []\n\n for audio_file in audio_files:\n if '.wav' in audio_file:\n audio_data = afe.extract((folder_path + '/' + audio_file), genres[i])\n feature_data_for_this_genre.append(audio_data)\n\n all_feature_data.append(np.array(feature_data_for_this_genre))\n\nall_feature_data[9].shape\n# Blues, Classical, Country, Disco, Hiphop, Jazz, Metal, Pop, Rock, Reggae\n\nall_feature_data_stacked = np.vstack(all_feature_data)\n\nall_feature_df = pd.DataFrame(data=all_feature_data_stacked, columns=feature_list)\n\n# +\nnan_columns = []\n\nfor column in all_feature_df.columns.values:\n unique = all_feature_df[column].unique()\n if 'nan' in unique:\n nan_columns.append(column)\n# -\n\nfor column in nan_columns:\n all_feature_df[column] = all_feature_df[column].replace('nan', 0)\n\nnan_columns\n\nfor column in all_feature_df.columns.values[:-1]:\n all_feature_df[column] = pd.to_numeric(all_feature_df[column])\n\n## Save the data to a CSV\nall_feature_df.to_csv('/Users/griffinbholt/SchoolProjects/MusicGenreIdentification/gtzan_features.csv', sep=',', index=False)\n\n\n","repo_name":"griffinbholt/CS472MusicGenreIdentification","sub_path":"Code/GTZAN Feature Extraction.ipynb","file_name":"GTZAN Feature Extraction.ipynb","file_ext":"py","file_size_in_byte":1929,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"12940227247","text":"# Let's now examine the performance of integration methods for the following stiff, non-linear ODE system, **Van der Pol's oscillator**. This file solves This file solves the following ordinary differential equation (ODE) system:\n# \n# \\begin{eqnarray*}\n# \t\\frac{\\displaystyle {dy_1(t)} }{\\displaystyle {dt} } &=& y_2(t)\\\\\n# \t\\frac{\\displaystyle {dy_2(t)} }{\\displaystyle {dt} } &=& \\lambda\\left[1 - y^2_1(t)\\right]y_2(t) - y_1(t)\n# \\end{eqnarray*}\n#\n# where $\\lambda \\ge 0$ and $[y_1(0), y_2(0)]^T = [2, 1]^T$.\n\n# Import\n\n# +\nimport sys\nsys.path.append('..//source')\n\nimport matplotlib.pylab as plt\nimport numpy as np\nimport numpy.linalg as LA\n\nimport scipy, time \nfrom scipy.interpolate import CubicSpline\n\nimport analysis, integrator, params, plotter\n\nimport points\n# -\n\n# %matplotlib notebook\n\n# Functions\n\n# +\n# the stiffness parameter\nK_stiff = 20\n# https://www.cs.ox.ac.uk/files/1175/lecture06-handout.pdf\n\n# backward euler, implicit tolerance. Also sused in SDC\nBE_TOL = 1e-12\nSDC_TOL = 1e-18 #1e-12\nN_ITER_MAX_NEWTON=50\n\ndef f_eval(t, y):\n \n \"\"\"\n sovling\n dy1/dt = y2\n dy2/dt = k * (1 - y1^2)y2 - y1\n \n :param float t: the time node\n :param numpy.ndarray: y the approximate solution length( m) at time node t\n \"\"\"\n\n F = np.zeros( y.shape )\n m = len(y)\n \n # get the component-wise interaction\n F[0] = y[1]\n F[1] = K_stiff * (1 - y[0]**2) * y[1] - y[0]\n \n return F\n\ndef get_corrections_norm(D):\n \n \"\"\"\n :param numpy.ndarray D: dimensions (the number of iterations, the number of nodes, the dimension of the problem)\n \"\"\"\n \n y = np.vstack( [ np.linalg.norm(x, axis=0) for x in D] )\n \n return y\n\ndef get_stuff(p, dt):\n \n if p.t[-1] != 1:\n t = dt * np.hstack( [p.t, 1])\n else:\n t = dt * p.t\n \n S, S_p = dt * p.S, dt * p.S_p\n \n spectral_radius = p.spectral_radius\n node_type = p.node_type\n \n return t, S, S_p, spectral_radius, node_type\n\n\n# -\n\n# Parameters\n\n# +\n#\n# set up integration parameters\n#\n\n# the parameters of the simulation\nn_nodes = 10\n\n# the time step\ndt = .25\n\n# the dimension of the problem\nm = 2\n\n# the initial solution\ny0 = np.array( (2, 1) )\n\n# create the parameter object\np_lobatto = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_LOBATTO)\n\n# create paramter for radau\n#p_radau = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_RADAU)\n\n# create paramter for radau 2a\n#p_radau_2a = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_RADAU_2A)\n\n# legendre nodes\n#p_legendre = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_LEGENDRE)\n\n\n# -\n\n# ---\n# Run one step\n#\n# ---\n\n# Backward Euler\n\n# +\nnodes, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\ny_be = integrator.backward_euler(f_eval, nodes, y0, S_p, be_tol=BE_TOL, do_print=True)\n\nt = nodes\n#LA.norm( y_be - exact(t), ord=np.inf)\n# -\n\n# SDC\n\n# +\nt, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\n\ny_be = integrator.backward_euler(f_eval, t, y0, S_p, be_tol=BE_TOL)\n\ny_sdc, Y_sdc, D_sdc, is_converged_sdc = integrator.sdc(f_eval, t, y0, y_be, S, S_p, be_tol=BE_TOL, sdc_tol=SDC_TOL, do_print=True)\nprint('SDC converged: %s ' % is_converged_sdc)\n\n# the norm of the corrections\nd_norm_sdc, log_d_norm_sdc, d_norm_rel_sdc, log_d_norm_rel_sdc = analysis.analyze_corrections(D_sdc, Y_sdc)\n# -\n\n# JFNK\n\n# +\nt, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\n\ny_be = integrator.backward_euler(f_eval, t, y0, S_p, be_tol=BE_TOL)\n\ny_jfnk, Y_jfnk, D_jfnk, is_converged_jfnk, is_stiff,ratios = integrator.jfnk(f_eval, t, y0, y_be, S, S_p, spectral_radius, be_tol=BE_TOL, \\\n sdc_tol=SDC_TOL, do_print=True)\nprint('JFNK converged: %s ' % is_converged_jfnk)\n\n# the norm of the corrections\nd_norm_jfnk, log_d_norm_jfnk, d_norm_rel_jfnk, log_d_norm_rel_jfnk = analysis.analyze_corrections(D_jfnk, Y_jfnk)\n# -\n\n# Spectral Solution\n\n# +\nt, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\n\ny_be = integrator.backward_euler(f_eval, t, y0, S_p, be_tol=BE_TOL)\n\ny_spect = integrator.run_spectral(f_eval, t, y0, S, tol=1e-10, y_approx=y_be, verbose=True)\n# -\n\n# Plot the corrections\n\n# +\nplt.figure()\nplt.title('Magnitude of corrections ')\n\nplt.plot( range(log_d_norm_sdc.shape[0]), log_d_norm_sdc.max(axis=1), '-o', label='sdc')\nplt.plot( range(log_d_norm_jfnk.shape[0]), log_d_norm_jfnk.max(axis=1), '-o', label='jfnk')\n\nplt.xlabel('SDC iteration')\nplt.ylabel('log10 (error)')\nplt.legend(loc='best')\nplt.show()\n# -\n\n# ---\n# Run Multiple Steps\n#\n# ---\n\n# +\nt_final =1\ntol = 1e-5\ndt_init = 4e-1\n\nprint(t_final, dt_init)\n# -\n\n# **Adaptive** JFNK\n\nt_adapt, y_adapt, Y_adapt, D_adapt, h_adapt = integrator.jfnk_adaptive(f_eval, t_init=0, t_final=t_final, dt_init=dt_init, p=p_lobatto, y0=y0, \\\n tol=tol, do_print=True)\n\n# the time steps\nh_adapt\n\n# **Uniform** JFNK\n\nt_uni, y_uni, Y_uni, D_uni = integrator.jfnk_uniform(f_eval, t_init=0, t_final=t_final, n_steps=12, p=p_lobatto, y0=y0, do_print=True)\n\n# **Reference** solution\n\n# +\n# the reference solution\nn_steps_ref = 100\np_ref = params.Params(n_nodes=10, m=m, node_type=points.GAUSS_LOBATTO)\n\nt_ref, y_ref, Y_ref, D_ref = integrator.jfnk_uniform(f_eval, t_init=0, t_final=t_final, n_steps=n_steps_ref, p=p_ref, y0=y0, do_print=True)\n\n# spline\ny_spline = CubicSpline(t_ref, y_ref)\n\n# errors\nf = lambda t, y: ( LA.norm( y_spline(t) - y, ord=np.inf), integrator.relative_norm(y_spline(t) - y, y_spline(t)) )\n# -\n\n# Comparing the absolute and relative erros of the **uniform** solution to the reference solution\n\nf(t_uni, y_uni)\n\n# Comparing the absolute and relative errors of the **adaptive** solution to the reference solution\n\nf(t_adapt, y_adapt), tol\n\n# ---\n# Plotting\n#\n# ---\n\n# Plot the JFNK corrections for each time step\n\n# +\nplotter.plot_correction_time_steps(D_uni, Y_uni)\n\nplt.show()\n# -\n\n# Plot norm of corrections\n\n# +\ny_spect_norm = LA.norm(y_spect, axis=0)\n\nlabels=['y1', 'y2']\n\nplotter.corrections(d_norm_sdc, d_norm_jfnk, y_spect_norm, do_rerr=True, labels=labels, do_legend=True)\nplt.show()\n# -\n\n# ---\n# Errors\n#\n# ---\n\n#\n# error\n#\naerr_be, rerr_be = analysis.error_analysis(y_be, y_spect)\naerr_sdc, rerr_sdc = analysis.error_analysis(y_sdc, y_spect)\naerr_jfnk, rerr_jfnk = analysis.error_analysis(y_jfnk, y_spect)\n\n\n","repo_name":"namdi/JFNK-integrator","sub_path":"examples/van_der_pol.ipynb","file_name":"van_der_pol.ipynb","file_ext":"py","file_size_in_byte":6416,"program_lang":"python","lang":"en","doc_type":"code","stars":5,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"11900144276","text":"# ## Please first install these libraries\n# pip install pythreejs pyntcloud pandas numpy\n\nfrom pyntcloud import PyntCloud\nimport pandas as pd\nimport os\nimport numpy as np\nimport PIL.Image as Image\n# %matplotlib inline\n\ndef load_velo_scan(velo_filename):\n scan = np.fromfile(velo_filename, dtype=np.float32)\n scan = scan.reshape((-1, 4))\n return scan\n\n\n#point cloud 형식으로 만들기 위해서 rgb 데이터 입히기\ndef paint_points(points, color = [0,0,100]):\n # color = [r, g, b]\n color = np.array([color])\n new_pts = np.zeros([points.shape[0],6])\n new_pts[:,:3] = points\n new_pts[:, 3:] = new_pts[:, 3:] + color\n return new_pts\n\n\npath = '/home/joonsu0109/Joon/Image_to_Points/depth_code.bin'\n\npoints = load_velo_scan(path)[:,:3]\n\npoints.shape\n\npd_points = pd.DataFrame(paint_points(points, img_edit), columns=['x','y','z','red','green','blue'])\n\ncloud = PyntCloud(pd_points)\ncloud.plot(initial_point_size=0.00001)\n\ncloud\n\ncloud.to_file(\"depth_color.ply\")\n\n# +\nimport open3d as o3d\n\npcd = o3d.io.read_point_cloud(\"/home/joonsu0109/Joon/Image_to_Point/depth_color.ply\")\no3d.io.write_point_cloud(\"depth_color.pcd\", pcd)\n# -\n\n# ---\n\nimport cv2\nimg = cv2.imread('/home/joonsu0109/Joon/Image_to_Point/set/000002.png')\n\n\n","repo_name":"joonsu0109gh/Image_to_Points","sub_path":"Image_to_Points/bin_to_pcd.ipynb","file_name":"bin_to_pcd.ipynb","file_ext":"py","file_size_in_byte":1250,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"5966523361","text":"# +\nimport os\nimport numpy as np\nimport decoy_mnist\nfrom matplotlib import pyplot as plt\n\nfrom keras.layers import Input, Conv2D, MaxPool2D, Dense, Flatten, UpSampling2D, Multiply, Reshape\nfrom keras.models import Model, load_model\nfrom keras.utils import to_categorical\nfrom keras.optimizers import SGD\nfrom keras.regularizers import l1, l2\nfrom keras.losses import kld\nfrom keras.datasets import mnist\nfrom keras import backend as K\n# -\n\n# ### Load decoy MNIST\n\nXr, x_train, y_train, E, Xtr, x_test, y_test, Et = decoy_mnist.generate_dataset(cachefile='data/decoy-mnist.npz')\nx_train = x_train / 255 / 5 + 4/5\nx_test = x_test / 255 / 5 + 4/5\nx_train = x_train.reshape((60000, 28, 28, 1))\nx_test = x_test.reshape((10000, 28, 28, 1))\ny_train = to_categorical(y_train, 10)\ny_test = to_categorical(y_test, 10)\n\n\n# ### Load normal MNIST\n\n# +\ndef load_mnist():\n print('Loading MNIST')\n (x_train, y_train), (x_test, y_test) = mnist.load_data()\n x_train = np.expand_dims(x_train, -1) / 255 / 5 + 4 / 5\n x_test = np.expand_dims(x_test, -1) / 255 / 5 + 4 / 5\n y_train = to_categorical(y_train, 10)\n y_test = to_categorical(y_test, 10)\n return (x_train, y_train), (x_test, y_test)\n\n\n(x_train_g, y_train_g), (x_test_g, y_test_g) = load_mnist()\n\n\n# -\n\n# ## Model looking at corner splotches\n\ndef ConvNet(train):\n if train or not os.path.isfile('classifier_d.hdf5'):\n print('Classifier model is being trained')\n inputs = Input([28, 28, 1])\n x = Conv2D(32, (3, 3), activation='relu', name='conv1')(inputs)\n x = Conv2D(64, (3, 3), activation='relu', name='conv2')(x)\n x = MaxPool2D((2, 2), name='mp1')(x)\n x = Conv2D(64, (3, 3), activation='relu', name='conv3')(x)\n x = Conv2D(128, (3, 3), activation='relu', name='conv4')(x)\n x = Flatten(name='flatten')(x)\n x = Dense(10, activation='softmax', name='dense1')(x)\n \n model = Model(inputs, x)\n model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])\n model.fit(x_train, y_train, batch_size=1000, epochs=15, validation_split=1/6)\n model.save('classifier_d.hdf5')\n else:\n print('Classifier model loaded')\n model = load_model('classifier_d.hdf5')\n return model\n\n\nconvnet1 = ConvNet(train=True)\nconvnet1.trainable = False\nprint(convnet1.evaluate(x_test, y_test))\nprint(convnet1.evaluate(x_test_g, y_test_g))\n\n# We see that model is robust and performs well on both decoy MNIST and regular MNIST test sets.\n\n# ### Explaining robust model\n\nmodel = convnet1\n\n\n# +\ndef normalized_kld(p, q):\n return 1 - K.exp(-kld(p, q))\n\ndef ExplanationModel(reg, train):\n if train or not os.path.isfile('exp_model_d.hdf5'):\n inputs = Input(shape=x_train.shape[1:])\n\n x = Conv2D(32, (3, 3), activation='relu', padding='same')(inputs)\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n x = MaxPool2D((2, 2))(x)\n\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n x = Conv2D(128, (3, 3), activation='relu', padding='same')(x)\n x = MaxPool2D((2, 2))(x)\n\n x = UpSampling2D((2, 2))(x)\n x = Conv2D(128, (3, 3), activation='relu', padding='same')(x)\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n\n x = UpSampling2D((2, 2))(x)\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)\n\n x = Conv2D(1, (1, 1), activation='sigmoid', padding='same', name='attention', activity_regularizer=l2(reg))(x)\n\n x = Multiply()([x, inputs])\n x = model(x)\n\n exp_model = Model(inputs, x)\n\n y_train = model.predict(x_train)\n exp_model.compile(optimizer='adam', loss=normalized_kld)\n exp_model.summary()\n for l in exp_model.layers[-1].layers:\n l.trainable = False\n exp_model.fit(x_train, y_train, epochs=5)\n exp_model.save('exp_model_d.hdf5')\n else:\n exp_model = load_model('exp_model_d.hdf5', compile=False)\n exp_model.compile(optimizer='sgd', loss=normalized_kld)\n return exp_model\n\n\n# -\n\nexp_model = ExplanationModel(reg=0.0001, train=True)\n\n# +\nexp_model = load_model('exp_model_d.hdf5', compile=False)\nsaliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\nsaliency_values = saliency([x_test])[0]\nprediction = model.predict(x_test, batch_size=1000)\ny = model.predict(x_test)\n\nfor i in range(5):\n print('Processing image #{}'.format(i+1))\n exp_model = load_model('exp_model_d.hdf5', compile=False)\n opt = SGD(lr=0.01)\n exp_model.compile(optimizer=opt, loss=normalized_kld)\n saliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n\n img = x_test[i, :, :, 0]\n\n plt.subplot(3, 3, 2)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n orig_sal = saliency_values[i].reshape((28, 28))\n sal_interp = (orig_sal - orig_sal.min()) / (orig_sal.max() - orig_sal.min())\n plt.imshow(sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(3, 3, 1)\n plt.axis('off')\n plt.title('Original saliency')\n plt.imshow(orig_sal, cmap='jet', vmin=0, vmax=1)\n\n for l in exp_model.layers[-1].layers:\n l.trainable = False\n hist = exp_model.fit(x_test[i:i + 1], y[i:i + 1], epochs=100, verbose=False)\n saliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n final_sal = saliency([x_test[i:i + 1]])[0].reshape((28, 28))\n final_sal_interp = (final_sal - final_sal.min()) / (final_sal.max() - final_sal.min())\n\n plt.subplot(2, 2, 1)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n plt.imshow(final_sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(2, 2, 2)\n plt.axis('off')\n plt.title('Final saliency')\n plt.imshow(final_sal, cmap='jet', vmin=0, vmax=1)\n\n\n plt.subplot(2, 2, 3)\n plt.title('Fine-tuning loss')\n plt.plot(hist.history['loss'])\n\n plt.subplot(2, 2, 4)\n plt.title('Saliency histogram')\n plt.hist(orig_sal.flatten())\n\n plt.show()\n\n\n# -\n\n# Saliency maps show that model is not paying attention to corner splotches.\n\n# ## Model looking at corner splotches\n\ndef ConvNet2(train):\n if train or not os.path.isfile('classifier_d2.hdf5'):\n print('Classifier model is being trained')\n inputs = Input([28, 28, 1])\n x = Flatten(name='flatten')(inputs)\n x = Dense(1024, activation='relu', name='dense1')(x)\n x = Dense(10, activation='softmax', name='dense2')(x)\n \n model = Model(inputs, x)\n model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])\n model.fit(x_train, y_train, batch_size=100, epochs=30, validation_split=1/6)\n model.save('classifier_d2.hdf5')\n else:\n print('Classifier model loaded')\n model = load_model('classifier_d2.hdf5')\n return model\n\n\nconvnet2 = ConvNet2(train=True)\nconvnet2.trainable = False\nprint(convnet2.evaluate(x_test, y_test))\nprint(convnet2.evaluate(x_test_g, y_test_g))\n\n# Model performs well on training decoy data but fails badly on test decoy data and even worse on normal test data.\n\n# ### Explaining bad model\n\nmodel = convnet2\n\nexp_model = ExplanationModel(reg=0.00001, train=True)\n\n# +\nexp_model = load_model('exp_model_d.hdf5', compile=False)\nsaliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n\nsaliency_values = saliency([x_test])[0]\nprediction = model.predict(x_test, batch_size=1000)\ny = model.predict(x_test)\n\nfor i in range(15):\n print('Processing image #{}'.format(i+1))\n exp_model = load_model('exp_model_d.hdf5', compile=False)\n opt = SGD(lr=0.01)\n exp_model.compile(optimizer=opt, loss=normalized_kld)\n\n img = x_test[i, :, :, 0]\n\n plt.subplot(2, 2, 2)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n orig_sal = saliency_values[i].reshape((28, 28))\n sal_interp = (orig_sal - orig_sal.min()) / (orig_sal.max() - orig_sal.min())\n plt.imshow(sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(2, 2, 1)\n plt.axis('off')\n plt.title('Original saliency')\n plt.imshow(orig_sal, cmap='jet', vmin=0, vmax=1)\n\n hist = exp_model.fit(x_test[i:i + 1], y[i:i + 1], epochs=100, verbose=False)\n saliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n final_sal = saliency([x_test[i:i + 1]])[0].reshape((28, 28))\n final_sal_interp = (final_sal - final_sal.min()) / (final_sal.max() - final_sal.min())\n\n plt.subplot(2, 2, 4)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n plt.imshow(final_sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(2, 2, 3)\n plt.axis('off')\n plt.title('Final saliency')\n plt.imshow(final_sal, cmap='jet', vmin=0, vmax=1)\n plt.show()\n \n plt.subplot(1, 2, 1)\n plt.title('Fine-tuning loss')\n plt.plot(hist.history['loss'])\n\n plt.subplot(1, 2, 2)\n plt.title('Saliency histogram')\n plt.hist(orig_sal.flatten())\n\n plt.show()\n# -\n\n# Explanations show that this bad model is indeed looking at corner splotches.\n","repo_name":"eclique/learning-saliency","sub_path":"mnist/Decoy MNIST.ipynb","file_name":"Decoy MNIST.ipynb","file_ext":"py","file_size_in_byte":9261,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"27823260185","text":"# + deletable=true editable=true\nimport cv2\nimport tensorflow\nimport os\nimport matplotlib.pyplot as plt\nos.environ[\"CUDA_DEVICE_ORDER\"]=\"PCI_BUS_ID\"\nos.environ[\"CUDA_VISIBLE_DEVICES\"]=\"4\"\n\n# + deletable=true editable=true\n# the data path\ndata_path ='../../data/MPIIGaze/Data/Original/'\nimg_list = []\ndef Read_img(data_path):\n for data in os.listdir(data_path):\n path = os.path.join(data_path, data)\n or_img = cv2.imread(path)\n if or_img != None:\n# plt.imshow(or_img[180:540, 320:960]) # 显示图片\n# plt.axis('off') # 不显示坐标轴\n# plt.show()\n \n# img_get = or_img.getdata()\n# print(img_get)\n or_img.tolist()\n# print(or_img)\n img_list.append(or_img)\n if os.path.isdir(path):\n Read_img(path)\n \n\n\n# + deletable=true editable=true\nRead_img(data_path)\n\n# + deletable=true editable=true\n# img1 = cv2.imread(filenames,cv2.IMREAD_GRAYSCALE) #读取图片,第二个参数表示以灰度图像读入\n# if img1 is None: #判断读入的img1是否为空,为空就继续下一轮循环\n# continue\n# res1= cv2.resize(img1,(28,28)) #对图片进行缩放,第一个参数是读入的图片,第二个是制定的缩放大小\n# res1_1 = res1.reshape(1,784)/255 #将表示图片的二维矩阵转换成一维\n# res1_1_1 = res1_1.tolist() #将numpy.narray类型的矩阵转换成list\n# train_set_x.append(res1_1_1)\n\n# + deletable=true editable=true\n\n","repo_name":"qbqandyxq/My_gaze","sub_path":"code/Read_img.ipynb","file_name":"Read_img.ipynb","file_ext":"py","file_size_in_byte":1576,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"37780962312","text":"# # Lab | Matplotlib & Seaborn\n#\n# #### Import all the necessary libraries here:\n\n# Libraries\nimport numpy as np\n\n# ## Challenge 1\n#\n# #### The data we will use in this challenge is:\n\nx = np.arange(0,100)\ny = x*2\nz = x**2\n\n# #### Plot (x, y) and (x, z).\n# There are 2 ways of doing this. Do in both ways.\n#\n# **Hint**: Check out the nrows, ncols and index arguments of subplots. \n#\n# Also, play around with the linewidth and style. Use the ones you're most happy with.\n\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\n# +\n# Method 1\n# 1 Figure with 2 subplots, together \nfig = plt.figure()\n\n# (x, y) on the first plot\nplt.plot(x, y, color='orange', label='(x, y)')\n\n# (x, z) on the second plot\nplt.plot(x, z, color='blue', label='(x, z)')\n\nplt.legend()\n\nplt.show()\n# -\n\n# #### Use plt.subplots(nrows=1, ncols=2) to create the plot.\n\n# +\n# Method 2\n# Figure with 1 row and 2 columns of subplots\nfig, ax = plt.subplots(nrows=1, ncols=2)\n\n# (x, y) on the first subplot\nax[0].plot(x, y, color='orange', label='(x, y)')\nax[0].legend()\n\n# (x, z) on the second subplot\nax[1].plot(x, z, color='blue', label='(x, z)')\nax[1].legend()\n\n# Show the figure\nplt.show()\n# -\n\n# #### Use your previous code but now, resize your plot.\n# **Hint**: Add the figsize argument in plt.subplots().\n#\n# If you want, try to add a title to the plot or even axes labels. You can also play with the fontweight and fontsize of the titles and labels. \n\n# +\n# your code here\n# Resizing the plot and doing it all\nfig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))\n\n# (x, y) on the first subplot\nax[0].plot(x, y, color='orange', label='(x, y)')\nax[0].legend()\nax[0].set_title(\"(x,y)\", fontweight='bold', fontsize=16)\nax[0].set_xlabel(\"X\", fontsize=10)\nax[0].set_ylabel(\"Y\", fontsize=10)\n\n# (x, z) on the second subplot\nax[1].plot(x, z, color='blue', label='(x, z)')\nax[1].legend()\nax[1].set_title(\"(x,z)\", fontweight='bold', fontsize=16)\nax[1].set_xlabel(\"X\", fontsize=10)\nax[1].set_ylabel(\"Z\", fontsize=10)\n\n# Show the figure\nplt.show()\n# -\n\n# #### Plot both $y=x^2$ and $y=e^x$ in the same plot using normal and logarithmic scale.\n# **Hint**: Use `set_xscale` and `set_yscale`.\n\n# +\n# your code here\n# -\n\n# #### As a bonus challenge, try to add a legend to the plot.\n\n# +\n# your code here\nx = np.arange(0,100)\ny = x**2\nz = np.exp(x)\n\nfig, axs = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))\n\naxs[0].plot(x, y, label='y = x^2')\naxs[0].plot(x, z, label='z = e^x')\naxs[0].set_title('Normal Scale')\naxs[0].legend()\n\naxs[1].plot(x, y, label='y = x^2')\naxs[1].plot(x, z, label='z = e^x')\naxs[1].set_title('Logarithmic Scale')\naxs[1].set_xscale('log')\naxs[1].set_yscale('log')\naxs[1].legend()\n\nplt.show()\n# -\n\n# ## Challenge 2\n# #### Import the `Fitbit2` dataset and store it in a variable called `fitbit`. You can find the dataset in Ironhack's database:\n# * db: `fitbit`\n# * table: `fitbit2`\n\n# +\n# your code here\nimport pandas as pd\n\n# Read the CSV file\nfitbit = pd.read_csv('Fitbit2.csv')\nfitbit\n# -\n\nfitbit.columns\n\n# #### From the Fitbit data, we want to visually understand:\n#\n# How the average number of steps change by month. Use the appropriate visualization to show the median steps by month. Is Fitbitter more active on weekend or workdays? All plots must be in the same jupyter notebook cell.\n#\n# **Hints**:\n#\n# * Use Months_encoded and Week_or Weekend columns.\n# * Use matplolib.pyplot object oriented API.\n# * Set your size figure to 12,4\n# * Explore plt.sca\n# * Explore plt.xticks\n# * Save your figures in a folder called `figures` in your repo. \n\n# +\n# your code here\n# MEDIAN STEPS BY MONTH_ENCODED\nfig, axs = plt.subplots(1, 2, figsize=(12, 4))\nmonthly_steps = fitbit.groupby(['Months_encoded'])['Steps'].median()\nmonthly_steps.plot(kind='bar', ax=axs[0], color = 'grey')\naxs[0].set_title('Median Steps by Month')\naxs[0].set_xlabel('Month')\naxs[0].set_ylabel('Steps')\n\n# MEDIAN STEPS BY WORK_OR_WEEKEND\nweekly_steps = fitbit['Steps'].groupby(fitbit['Work_or_Weekend']).median()\nplt.barh(weekly_steps.index, weekly_steps.values, height= 0.9, color= 'black', align='center', )\nplt.yticks([0, 1], ['Weekend', 'Work'])\nplt.show()\n\n# Save the figure\nplt.show()\n# -\n\n# #### Write a loop to plot 3 scatter plots of the following features:\n#\n# * Minutes Lightly Active vs Steps \n# * Minutes Very Active vs Steps \n# * Minutes Sedentary vs Steps \n\n# +\n# your code here\nfig, axs = plt.subplots(1, 3, figsize=(12, 4))\n\nfeatures = ['Minutes Lightly Active', 'Minutes Very Active', 'Minutes Sedentary']\n\nfor i, feature in enumerate(features):\n axs[i].scatter(fitbit[feature], fitbit['Steps'], color = 'purple')\n axs[i].set_xlabel(feature)\n axs[i].set_ylabel('Steps')\nplt.show()\n# -\n\n# ## Challenge 3\n#\n# #### Import the `titanic` dataset and store it in a variable called `titanic`. You can find the dataset in Ironhack's database:\n# * db: `titanic`\n# * table: `titanic`\n\n# your code here\n# Read the CSV file\ntitanic = pd.read_csv('titanic.csv')\ntitanic\n\n# #### Explore the titanic dataset using Pandas dtypes.\n\n# your code here\ntitanic.dtypes\n\n# #### What are your numerical variables? What are your categorical variables?\n# **Hint**: Use Pandas select_dtypes.\n\n# NUMERICAL VARIABLES\n# your code here\ntitanic.select_dtypes(include='number')\n\n# CATEGORICAL VARIABLES\n# your code here\ntitanic.select_dtypes(include='object')\n\n# #### Set the plot style to classic and the figure size to (12,6).\n# **Hint**: To set the style you can use matplotlib or seaborn functions. Do some research on the matter.\n\n# your code here\nplt.style.use('classic')\nsns.set_style(\"darkgrid\")\nplt.rcParams['figure.figsize'] = (12, 6)\n\n# #### Use the right visulalization to show the distribution of column `Age`.\n\n# +\n# your code here\ntitanic['Age'].hist(bins=50, color = 'green')\n\nplt.title(\"Distribution\")\nplt.xlabel(\"Age\")\nplt.ylabel(\"Count\")\n\nplt.show()\n# -\n\n# #### Use subplots and plot the distribution of the `Age` with bins equal to 10, 20 and 50.\n\n# +\n# your code here\nplt.subplot(1,3,1)\ntitanic['Age'].hist(bins = 10, color = \"orange\")\nplt.subplot(1,3,2)\ntitanic['Age'].hist(bins=20, color = \"green\")\nplt.subplot(1,3,3)\ntitanic['Age'].hist(bins=30, color = \"red\")\n\nplt.show()\n# -\n\n# #### How does the bin size affect your plot?\n\n\"\"\"\nyour comments here \nAs the bin size increases, each bin will represent a larger range of values. This makes the plot more difficult to read.\n\"\"\"\n\n# #### Use seaborn to show the distribution of column `Age`.\n\n# your code here\nsns.distplot(titanic['Age'])\n\n# #### Use the right plot to visualize column `Gender`. There are 2 ways of doing it. Do it both ways.\n# **Hint**: Use matplotlib and seaborn.\n\n# +\n# Method 1 - matplotlib\n# your code here\ngender_counts = titanic['Gender'].value_counts()\n\nplt.bar(gender_counts.index, gender_counts.values, color = \"grey\")\nplt.xlabel('Gender')\nplt.ylabel('Occurrences')\nplt.show()\n\n# +\n# Method 2 - seaborn\n# your code here\n\nsns.countplot(x='Gender', data=titanic, color = \"grey\")\nplt.xlabel('Gender')\nplt.ylabel('Number of occurrences')\nplt.show()\n# -\n\n# #### Use the right plot to visualize the column `Pclass`.\n\n# +\n# your code here\npclass_counts = titanic['Pclass'].value_counts()\n\n# Create a bar plot\nplt.bar(pclass_counts.index, pclass_counts.values, color = \"grey\")\nplt.xlabel('Passenger Class')\nplt.ylabel('Count')\nplt.title('Distribution of Passenger Classes')\nplt.show()\n# -\n\n# #### We would like to have in one single plot the summary statistics of the feature `Age`. What kind of plot would you use? Plot it. \n\n# your code here\nsns.boxplot(x=titanic['Age'], color = \"orange\")\n\n# +\nQ1 = titanic[\"Age\"].quantile(0.25)\nQ2 = titanic[\"Age\"].quantile(0.50)\nQ3 = titanic[\"Age\"].quantile(0.75)\n\nprint(Q1)\nprint(Q2) # median\nprint(Q3)\n# -\n\n\"\"\"\nyour comments here \nThis plot shows the median, quartiles, and outliers of the data and can provide a quick visual representation of \nthe distribution of the data.\n\n\"\"\"\n\n# #### What does the last plot tell you about the feature `Age`?\n\n\"\"\"\nyour comments here\n\n25% of the values in the Age feature are less than or equal to 22.0, 50% of the values are less than or equal to 28.5, \nand 75% of the values are less than or equal to 37.0.\n\n\"\"\"\n\n# #### Now in addition to the summary statistics, we want to have in the same plot the distribution of `Age`. What kind of plot would you use? Plot it. \n\n# your code here\nplt.figure(figsize=(10,5))\nsns.violinplot(data=titanic, x='Age', color = \"purple\")\nplt.show()\n\n# #### What additional information does the last plot provide about feature `Age`?\n\n\"\"\"\nyour comments here\nIt shows the distribution of the data through the width of the violin and the summary statistics - Minimum and maximum at both \nends; median, and quartiles through the white dot in the center and the black bar inside the violin).\n\"\"\"\n\n# #### We suspect that there is a linear relationship between `Fare` and `Age`. Use the right plot to show the relationship between these 2 features. There are 2 ways, please do it both ways.\n# **Hint**: Use matplotlib and seaborn.\n\n# Method 1 - matplotlib\n# your code here\nplt.scatter(titanic['Age'], titanic['Fare'], s = 100)\nplt.xlabel('Age')\nplt.ylabel('Fare')\nplt.show()\n\n# Method 2 - seaborn\n# your code here\nsns.scatterplot(x='Age', y='Fare', data=titanic, s = 100)\n\n# #### Plot the correlation matrix using seaborn.\n\n# your code here\ncorr = titanic.corr()\nsns.heatmap(corr, annot=True, cmap='Reds')\n\n# #### What are the most correlated features?\n\n\"\"\"\nyour comments here\n- Fare & Survived\n- Parch & SibSp\n\"\"\"\n\n# #### Use the most appropriate plot to display the summary statistics of `Age` depending on `Pclass`.\n\n# your code here\nsns.boxplot(x='Pclass', y='Age', data=titanic)\nplt.title('Summary Statistics of Age by Passenger Class')\nplt.xlabel('Passenger Class')\nplt.ylabel('Age')\nplt.show()\n\n# #### Use seaborn to plot the distribution of `Age` based on the `Gender`.\n# **Hint**: Use Facetgrid.\n\n# your code here\ngender = sns.FacetGrid(titanic, col=\"Gender\")\ngender.map(sns.histplot, \"Age\", kde=False)\ngender.add_legend()\nplt.xlabel('Age')\nplt.ylabel('Frequency')\nplt.show()\n","repo_name":"sfmar15/lab-matplotlib-seaborn","sub_path":"your-code/main.ipynb","file_name":"main.ipynb","file_ext":"py","file_size_in_byte":10034,"program_lang":"python","lang":"en","doc_type":"code","dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"23779972301","text":"# #### Imports\n\n# +\nimport os\nimport csv\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport tensorflow as tf\nimport seaborn as sns\nfrom tensorflow.keras import datasets, layers, models\n\nfrom sklearn.model_selection import train_test_split, KFold\nfrom sklearn.metrics import confusion_matrix, roc_curve, auc, classification_report, precision_recall_curve\nfrom sklearn.utils import shuffle\nfrom PIL import Image\n\n\n# -\n\n# ## Methods for Experiments\n\ndef get_distr(targets, labels):\n vals = {}\n for label in labels.keys():\n vals[label] = 0\n \n for target in targets:\n vals[target] += 1\n \n for label in labels.keys():\n print(str(labels[label]) + \": \" + str(vals[label]))\n \n fig = plt.figure()\n ax = fig.add_axes([0,0,1,1])\n langs = labels.values()\n students = vals.values()\n ax.bar(langs,students)\n plt.show()\n\n\ndef plot_ROC(y_test, y_pred, title):\n fpr, tpr, thresholds = roc_curve(y_test, y_pred)\n auc_val = auc(fpr, tpr)\n plt.plot(fpr,tpr) \n plt.plot([0, 1], [0, 1], linestyle='dashed')\n plt.axis([-0.05, 1.05, -0.05, 1.05])\n plt.xlabel('False Positive Rate') \n plt.ylabel('True Positive Rate') \n title += \"\\nAUC: \" + str(round(auc_val, 4))\n plt.title(title)\n plt.show()\n\n\ndef plot_cm(y_test, y_pred, title):\n cm = confusion_matrix(y_test, y_pred)\n cm_df = pd.DataFrame(cm)\n cm_df = (cm_df.T/np.sum(cm_df, axis=1)).T\n sns.heatmap(cm_df, annot=True, cmap=plt.cm.Blues, fmt=\".2%\")\n plt.imshow(cm)\n plt.xlabel(\"Predicted Label\")\n plt.ylabel(\"True Label\")\n plt.title(title)\n\n\ndef plot_pr(y_test, y_pred, title):\n new_pred = []\n for val in y_pred:\n if val >= 0.5:\n new_pred.append(val)\n else:\n new_pred.append(0)\n prec, recall, _ = precision_recall_curve(y_test, new_pred)\n plt.plot(recall, prec)\n plt.ylim([0, 1])\n plt.xlabel(\"Recall\")\n plt.ylabel(\"Precision\")\n plt.title(title + \" Precision vs Recall\")\n\n\ndef convert_pred(y_pred, threshold):\n new_pred = np.empty((y_pred.size), dtype='int32')\n for i in range(y_pred.size):\n if y_pred[i] >= threshold:\n new_pred[i] = 1\n else:\n new_pred[i] = 0\n \n return new_pred\n\n\n# # Part 0: Load in csv data and format it to images and target data\n#\n# Read in csv of metadata for the images and select target data. For our target data we are selecting COVID/Non-COVID, Survival, Intubation, and Admission to the ICU - resulting in 28 classes where class 0 = Non-COVID and class > 0 is COVID and some unique combination of the classes above. For the initial training and testing of this data, any class > 0 we set to 1 to train a binary classifier for COVID/Non-COVID\n\n# +\ndata = []\nwith open(\"metadata.csv\") as file:\n for row in csv.reader(file):\n data.append(row)\n\ndata = data[1:]\n\nd = {'filename': [], 'target': []}\n\nfor image in data:\n # skip volume data\n if image[22] == \"volumes\":\n continue\n \n # ignore CT scan data\n if image[19] == \"CT\":\n continue\n \n diagnosis = image[4].split(\"/\")\n diagnosis = diagnosis[len(diagnosis) - 1].lower()\n # set target based on meta data\n if diagnosis != \"covid-19\":\n if diagnosis == \"todo\":\n # if not classified yet target = -1\n # d['filename'].append(image[23])\n # d['target'].append(-1)\n # ignore unknown data\n continue\n else:\n # if not COVID-19 target = 0\n d['filename'].append(image[23])\n d['target'].append(0)\n else:\n # if it is COVID-19\n if image[6] == \"Y\":\n # if they survived\n if image[7] == \"Y\":\n # if they were intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(1)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(2)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(3)\n elif image[7] == \"N\":\n # if they were not intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(4)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(5)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(6)\n else:\n # if they were intubated is unknown\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(7)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(8)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(9)\n elif image[6] == \"N\":\n # if they did not survive\n if image[7] == \"Y\":\n # if they were intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(10)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(11)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(12)\n elif image[7] == \"N\":\n # if they were not intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(13)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(14)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(15)\n else:\n # if they were intubated is unknown\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(16)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(17)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(18)\n else:\n # if their survival is unknown\n if image[7] == \"Y\":\n # if they were intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(19)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(20)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(21)\n elif image[7] == \"N\":\n # if they were not intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(22)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(23)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(23)\n else:\n # if they were intubated is unknown\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(24)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(25)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(26)\n\n \n# add normal chest xrays to data\ndir_path = os.path.abspath(os.getcwd())\ndir_path = dir_path+\"/chest_xray/test/Normal\"\nlist = os.listdir(dir_path)\nfor pic in list:\n d['filename'].append(pic)\n d['target'].append(0)\ntotal = [0] * 28\n\nfor target in d['target']:\n if target == -1:\n total[27] += 1\n else:\n total[target] += 1\n\n# set up binary classification (covid vs non-covid)\n# all targets > 0 are covid\nfor i in range(len(d['target'])):\n if d['target'][i] > 0:\n d['target'][i] = 1\n\nprint(total)\n#print(d)\n\nfile.close()\n\n# -\n\nlabels = {0: \"Non COVID\", 1: \"COVID\"}\nget_distr(d['target'], labels)\n\nprint(len(d['target']))\nprint(d['target'].count(1))\nprint(d['target'].count(0))\n\n# # Part 1: Preprocessing for images\n# Load in images for the corresponding target data with PIL. Reformat all images to the same dimensions and convert them to a numpy array for the models to use (where each is a (# pixels, # pixels) array).\n\n# +\n# Load in image data from filenames with PIL\nimg_dim = 128\nimages = np.empty((len(d['target']), img_dim, img_dim, 3))\n\nfor i in range(len(d['filename'])):\n image = Image.open(\"model_images/\" + d['filename'][i])\n \n # resize image shape\n image = image.resize((img_dim, img_dim))\n \n # convert to grayscale\n image = image.convert('RGB')\n \n images[i] = np.array(image)\n\n# normalize values\nimages = images / 255.0\n# -\n\ncallbacks = [\n tf.keras.callbacks.EarlyStopping(\n monitor=\"val_loss\",\n min_delta=0.01,\n patience=2,\n verbose=1\n )\n] \n\n# # Part 2: Deep Learning Classifier with Keras\n\n# Split the data into training and testing with sklearn\n\n# split data\nx_train, x_test, y_train, y_test = train_test_split(images, d['target'], test_size=0.2, random_state=0)\ny_train = np.asarray(y_train)\ny_test = np.asarray(y_test)\n\n# Build and compile the deep learning model\n\n# +\n# build model\ndl_model = tf.keras.Sequential([\n tf.keras.layers.Flatten(input_shape=(img_dim,img_dim,2)),\n tf.keras.layers.Dense(512, activation='relu'),\n tf.keras.layers.Dense(1, activation='sigmoid')\n])\n\n# compile model\ndl_model.compile(optimizer='adam',\n loss=tf.keras.losses.BinaryCrossentropy(),\n metrics=['accuracy'])\n# -\n\ndl_model.summary()\n\n# Train the model\n\nhistory = dl_model.fit(x_train, y_train, epochs=10,\n validation_data=(x_test, y_test),\n callbacks=callbacks,\n batch_size=16)\n\n# Evaluate the model with testing data\n\n# +\nplt.plot(history.history['accuracy'], label='accuracy')\nplt.plot(history.history['val_accuracy'], label='val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title('Deep Learning Classifier')\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\ntest_loss, test_acc = dl_model.evaluate(x_test, y_test, verbose=2)\nprint(\"Test Accuracy:\", round(test_acc, 4))\n# -\n\ndly_pred = dl_model.predict(x_test)\n\ndly_pred_norm = convert_pred(dly_pred, 0.5)\n\nprint(classification_report(y_test, dly_pred_norm))\n\nplot_cm(y_test, dly_pred_norm, \"Not COVID(0)/COVID(1) - MLP\")\n\nplot_ROC(y_test, dly_pred, \"Not COVID/COVID - MLP\")\n\n# # Part 3: Classifier with a Convolution Neural Network\n\n# Build and compile the model\n\n# +\n# build model for convolution network\ncnn_model = models.Sequential()\ncnn_model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(img_dim, img_dim, 2), padding='same'))\ncnn_model.add(layers.MaxPooling2D((2, 2)))\ncnn_model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))\ncnn_model.add(layers.MaxPooling2D((2, 2)))\ncnn_model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))\n\n# add more layers\ncnn_model.add(layers.Flatten())\ncnn_model.add(layers.Dense(64, activation='relu'))\ncnn_model.add(layers.Dense(1, activation='sigmoid'))\n\n# compile \ncnn_model.compile(optimizer='adam',\n loss=tf.keras.losses.BinaryCrossentropy(),\n metrics=['accuracy'])\n# -\n\ncnn_model.summary()\n\n# Train the model\n\nhistory = cnn_model.fit(x_train, y_train, epochs=10, \n validation_data=(x_test, y_test),\n callbacks=callbacks,\n batch_size=16)\n\n# Evaluate the model with testing data and plot results\n\n# +\nplt.plot(history.history['accuracy'], label='accuracy')\nplt.plot(history.history['val_accuracy'], label = 'val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title('CNN Classifier')\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\ntest_loss, test_acc = cnn_model.evaluate(x_test, y_test, verbose=2)\nprint(\"Test Accuracy:\", round(test_acc, 4))\n# -\n\ncnny_pred = cnn_model.predict(x_test)\n\ncnny_pred_norm = convert_pred(cnny_pred, 0.5)\n\nprint(classification_report(y_test, cnny_pred_norm))\n\nplot_cm(y_test, cnny_pred_norm, \"Not COVID(0)/COVID(1) - CNN\")\n\nplot_ROC(y_test, cnny_pred, \"Not COVID/COVID - CNN\")\n\n# Compare models\nfpr_dl, tpr_dl, thresholds = roc_curve(y_test, dly_pred)\nfpr_cnn, tpr_cnn, thresholds = roc_curve(y_test, cnny_pred)\nauc_valdl = auc(fpr_dl, tpr_dl)\nauc_valcnn = auc(fpr_cnn, tpr_cnn)\nplt.plot(fpr_dl,tpr_dl) \nplt.plot(fpr_cnn, tpr_cnn)\nplt.plot([0, 1], [0, 1], linestyle='dashed')\nplt.axis([-0.05, 1.05, -0.05, 1.05])\nplt.xlabel('False Positive Rate') \nplt.ylabel('True Positive Rate')\nplt.title(\"ROC Curve\\nMLP AUC: \" + str(round(auc_valdl, 4)) + \"\\nCNN AUC: \" + str(round(auc_valcnn, 4)))\nplt.legend([\"MLP\", \"CNN\"])\nplt.show()\n\n# # Part 4: Optimization\n# Based on the results of the above models, we will now attempt to optimize our algorithms. We will be using cross validation to find a find the best split between our training/testing data.\n\n# +\n# split data into 10 folds\nfolds = 10\nkfold = KFold(n_splits=folds)\n\n# shuffle initial data\nrimages, rtargets = shuffle(images, d['target'], random_state=0)\n# -\n\n# Repeat Deep Learning and CNN models above with KFold technique and compare results\n\n# ### Deep Learning - Cross Validation\n\n# +\nresults = []\ntest_accuracies = []\n\nfor train, test in kfold.split(rimages):\n x_train, x_test = rimages[train], rimages[test]\n y_train, y_test = np.array(rtargets)[train], np.array(rtargets)[test]\n \n # build model\n dl_model = tf.keras.Sequential([\n tf.keras.layers.Flatten(input_shape=(img_dim,img_dim,2)),\n tf.keras.layers.Dense(512, activation='relu'),\n tf.keras.layers.Dense(2)\n ])\n\n # compile model\n dl_model.compile(optimizer='adam',\n loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),\n metrics=['accuracy'])\n \n # train model\n history = dl_model.fit(x_train, y_train, epochs=10,\n validation_data=(x_test, y_test))\n \n test_loss, test_acc = model.evaluate(x_test, y_test, verbose=2)\n \n results.append(history)\n test_accuracies.append(test_acc)\n# -\n\n# Get best model and plot results\n\n# choose model with highest test accuracy\nbest_model = results[0]\nbest_acc = test_accuracies[0]\nfor i in range(len(results) - 1):\n if test_accuracies[i + 1] > test_accuracies[i]:\n best_model = results[i+1]\n\n# +\nplt.plot(best_model.history['accuracy'], label='accuracy')\nplt.plot(best_model.history['val_accuracy'], label = 'val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title('Deep Learning Classifier - CV')\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\nprint(\"Test Accuracy:\", round(best_acc, 4))\n# -\n\n# ### CNN - Cross Validation\n\n# +\nresults = []\ntest_accuracies = []\n\nfor train, test in kfold.split(rimages):\n x_train, x_test = rimages[train], rimages[test]\n y_train, y_test = np.array(rtargets)[train], np.array(rtargets)[test]\n \n # build model for convolusion network\n cnn_model = models.Sequential()\n cnn_model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(img_dim, img_dim, 2)))\n cnn_model.add(layers.MaxPooling2D((2, 2)))\n cnn_model.add(layers.Conv2D(64, (3, 3), activation='relu'))\n cnn_model.add(layers.MaxPooling2D((2, 2)))\n cnn_model.add(layers.Conv2D(64, (3, 3), activation='relu'))\n\n # add more layers\n cnn_model.add(layers.Flatten())\n cnn_model.add(layers.Dense(64, activation='relu'))\n cnn_model.add(layers.Dense(2))\n\n # compile \n cnn_model.compile(optimizer='adam',\n loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),\n metrics=['accuracy'])\n \n history = cnn_model.fit(x_train, y_train, epochs=10,\n validation_data=(x_test, y_test))\n \n results.append(history)\n \n \n test_accuracies.append(test_acc)\n# -\n\n# Get best model and plot results\n\n# choose model with highest test accuracy\nbest_model = results[0]\nbest_acc = test_accuracies[0]\nfor i in range(len(results) - 1):\n if test_accuracies[i + 1] > test_accuracies[i]:\n best_model = results[i+1]\n\n# +\nplt.plot(best_model.history['accuracy'], label='accuracy')\nplt.plot(best_model.history['val_accuracy'], label = 'val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title(\"CNN Classifier - CV\")\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\nprint(\"Test Accuracy:\", round(best_acc, 4))\n# -\n\n\n","repo_name":"gdosulli/CS254_COVID","sub_path":"GOBT_CS254_InitialModels.ipynb","file_name":"GOBT_CS254_InitialModels.ipynb","file_ext":"py","file_size_in_byte":18545,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"30443140186","text":"# # Importing libraries and Exploring the Dataset\n\nimport numpy as np \nimport pandas as pd \nfrom matplotlib import pyplot as plt\nimport seaborn as sns\nimport scipy.stats as stats\nsns.set() \n\n# Read the Dataset\ninsurance_df = pd.read_csv(\"insurance-data.csv\")\n\n# +\n#Display the first five dataset\n\ninsurance_df.head()\n\n# +\n#Info about the data shape of data, type of individual columns\n\ninsurance_df.info()\n# -\n\n# # Checking missing values and Outliers\n\n# +\n# Check for the null values \n\ninsurance_df.isna().apply(pd.value_counts)\n# -\n\n# Checking for the outliers\nfig, ax = plt.subplots(3, 1, figsize=(10,7))\nax[0].set_title(\"BMI\")\nax[1].set_title(\"AGE\")\nax[2].set_title(\"CHARGES\")\nsns.boxplot(data = insurance_df.bmi, orient=\"h\", ax=ax[0], color = \"b\")\nsns.boxplot(data = insurance_df.age, orient=\"h\", ax=ax[1], color = \"r\")\nsns.boxplot(data = insurance_df.charges, orient=\"h\", ax=ax[2], color = \"g\")\nplt.tight_layout()\n\n# +\n# a brief summary for the dataset\n\ninsurance_df.describe().T\n# -\n\n# # Creating visual methods to analyze the data.\n\n# +\n# Plots to see the distribution of the continuous features individually\n\nplt.figure(figsize=(20,15))\nplt.subplot(3,3,1)\nplt.hist(insurance_df.bmi, color=\"lightblue\", edgecolor=\"black\", alpha=0.7)\nplt.xlabel(\"bmi\")\n\nplt.subplot(3,3,2)\nplt.hist(insurance_df.age, color='red', edgecolor = 'black', alpha = 0.7)\nplt.xlabel('age')\n\nplt.subplot(3,3,3)\nplt.hist(insurance_df.charges, color='lightgreen', edgecolor = 'black', alpha = 0.7)\nplt.xlabel('charges')\n\nplt.show()\n# -\n\n# Fatih hocamızın gönderdiği kod:\nfig, ax = plt.subplots(1, 3, figsize=(15,4))\nsns.distplot(insurance_df.bmi, ax=ax[0], color = \"b\", bins=10, kde=False)\nsns.distplot(insurance_df.age, ax=ax[1], color = \"r\", bins=10, kde=False)\nsns.distplot(insurance_df.charges, ax=ax[2], color = \"g\", bins=10, kde=False)\n\n# +\nplt.figure(figsize=(17,20))\nx = insurance_df.smoker.value_counts().index\ny = [insurance_df['smoker'].value_counts()[i] for i in x]\nplt.subplot(4,2,1)\nplt.bar(x,y, align='center',color = 'lightblue',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('Is Smoker?')\nplt.ylabel('Count ')\nplt.title('Smoker distribution')\n\nx1 = insurance_df.sex.value_counts().index #Values for x-axis\ny1 = [insurance_df['sex'].value_counts()[j] for j in x1] # Count of each class on y-axis\n\nplt.subplot(4,2,2)\nplt.bar(x1,y1, align='center',color = 'red',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('Gender')\nplt.ylabel('Count')\nplt.title('Gender distribution')\n\nx2 = insurance_df.region.value_counts().index #Values for x-axis\ny2 = [insurance_df['region'].value_counts()[k] for k in x2] # Count of each class on y-axis\n\nplt.subplot(4,2,3)\nplt.bar(x2,y2, align='center',color = 'green',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('Region')\nplt.ylabel('Count ')\nplt.title(\"Regions' distribution\")\n\nx3 = insurance_df.children.value_counts().index #Values for x-axis\ny3 = [insurance_df['children'].value_counts()[l] for l in x3] # Count of each class on y-axis\n\nplt.subplot(4,2,4)\nplt.bar(x3,y3, align='center',color = 'purple',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('No. of children')\nplt.ylabel('Count ')\nplt.title(\"Children distribution\")\nplt.tight_layout()\nplt.show()\n# -\n\nsns.countplot(insurance_df.region)\n\nsns.pairplot(insurance_df)\n\n# # Analyzing trends, patterns, and relationships in the Data.\n\n# Do charges of people who smoke differ significantly from the people who don't?\n\n\nsns.heatmap(insurance_df.corr(), cmap = 'coolwarm', annot = True)\n\n# Scatter plot to look for visual evidence of dependency between attributes smoker and charges accross different ages\nplt.figure(figsize=(12,7))\nsns.scatterplot(insurance_df.age, insurance_df.charges, hue=insurance_df.smoker, palette=[\"red\", \"green\"], alpha=0.7)\nplt.show()\n\nplt.figure(figsize=(12,7))\nsns.scatterplot(insurance_df.age, insurance_df.charges, hue=insurance_df.sex, palette=[\"red\", \"blue\"], alpha=0.7)\nplt.show()\n\ninsurance_df.sex.value_counts()\n\n# +\n# T-test to check dependency of smoking on charges\nH0=\"Charges of smoker and non-smoker are the same\"\nHa=\"Charges of smoker and non-smoker are not the same\"\n\na=np.array(insurance_df[insurance_df.smoker==\"yes\"].charges)\nb=np.array(insurance_df[insurance_df.smoker==\"no\"].charges)\n\nt, p_value=stats.ttest_ind(a,b, axis=0)\n\nif p_value < 0.05: # Setting our significance level at 5%\n print(f'{Ha} as the p_value ({p_value}) < 0.05')\nelse:\n print(f'{Ho} as the p_value ({p_value}) > 0.05')\n\n\n# +\n#Does bmi of males differ significantly from that of females?\nH0=\"gender has no effect on bmi\"\nHa=\"gender has effect on bmi\"\n\na=np.array(insurance_df[insurance_df.sex==\"male\"].bmi)\nb=np.array(insurance_df[insurance_df.sex==\"female\"].bmi)\n\nt, p_value=stats.ttest_ind(a,b, axis=0)\n\nif p_value < 0.05: # Setting our significance level at 5%\n print(f'{Ha} as the p_value ({p_value}) < 0.05')\nelse:\n print(f'{H0} as the p_value ({p_value}) > 0.05')\n# -\n\n\n\n\n# Hasan hocamızın gönrderdiği kod:\nttest2=stats.ttest_ind(insurance_df.bmi[insurance_df[\"sex\"]==\"male\"],insurance_df.bmi[insurance_df[\"sex\"]==\"female\"])\nif ttest2[1]<0.05:\n print(f\"Gender has effect on bmi as the p_value {ttest2[1]} < 0.05\")\nelse:\n print(f\"Gender has no effect on bmi as the p_value {round(ttest2[1],2)} > 0.05\")\n\n\n","repo_name":"hersany/DataScience","sub_path":"EDA/EDA of Insurance Company's Dataset.ipynb","file_name":"EDA of Insurance Company's Dataset.ipynb","file_ext":"py","file_size_in_byte":5312,"program_lang":"python","lang":"en","doc_type":"code","stars":6,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"12715736122","text":"# * Matplotlib \n# Matplotlib是一个最常用的2D绘图包,可以完成绝大部分的绘图任务,当然也可以完成部分简单的3D绘图任务,效果并不如其他的库那么理想\n\n# 获取数据\n# %matplotlib inline\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\nplt.plot(x, c, x, s)\n\n# +\nplt.figure(figsize=(8, 6), dpi=80) # (8, 6) 英寸,每英寸2,54cm, 分辨率是80\nplt.subplot(1, 1, 1) # 传建一个子图 1 * 1,占据第1个网格\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\n# color 设置颜色,设置线宽,设置线的样式,alpha还可以设置我们的透明度\nplt.plot(x, c, color='blue', linewidth=1.0, linestyle='-')\nplt.plot(x, s, color='green', linewidth=1.0, linestyle='-')\n# 设置x轴的范围\nplt.xlim(-4.0, 4.0)\n# 设置轴的说明\nplt.xlabel('这是x')\n# 设置坐标轴的刻度,支持使用Latex标注\nplt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi],[r'$-\\pi$', r'$-\\pi/2$', r'$0$', r'$+\\pi/2$', r'$+\\pi$'])\nplt.ylim(-1.0, 1.0)\nplt.ylabel('这是y')\nplt.yticks(np.linspace(-1, 1, 5, endpoint=True))\n\nax = plt.gca()\n\n# 为了可以让我们的刻度显示出来不被线条遮盖\nfor label in ax.get_xticklabels() + ax.get_yticklabels():\n # 将刻度的字体大小调整\n label.set_fontsize(16)\n # 字体的矩形显示框\n label.set_bbox(dict(facecolor='blue', edgecolor='None', alpha=0.65))\n\n\nplt.show()\n\n# plt.savefig('filename.png', dpi=80) # 保存分辨率是80的图图片到对应的路径上\n# -\n\n# spines的移动\nax = plt.gca() # 获取当前的坐标轴\n# 单个运行下面的语句,将右边的边界置空\nax.spines['right'].set_color('none') # right, top, left, bottom\nax.spines['top'].set_color('none')\nax.xaxis.set_ticks_position('bottom') # 将xaxis, x坐标轴设定成底边的bottom spines\nax.yaxis.set_ticks_position('left')\nax.spines['bottom'].set_position(('data', 0.5)) # 将位置设定在y轴的0.5上\nax.spines['left'].set_position(('data', 0.5)) # 将位置设定在x轴的0.5刻度上\nplt.show()\n\n# 添加标签\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\nplt.plot(x, c, color='blue', linewidth=2.5, linestyle='-', label='cosline') # label属性制定之后的legend的内容\nplt.plot(x, s, color='red', linewidth=2.5, linestyle='-', label='sine')\nplt.legend(loc='upper left') # 将lengend图标显示在左上角,不加的话,默认将其添加在最合适的位置上\n\n# +\n# 注释 annotate\nax = plt.gca() # 获取当前的坐标轴\n# 单个运行下面的语句,将右边的边界置空\nax.spines['right'].set_color('none') # right, top, left, bottom\nax.spines['top'].set_color('none')\nax.xaxis.set_ticks_position('bottom') # 将xaxis, x坐标轴设定成底边的bottom spines\nax.yaxis.set_ticks_position('left')\nax.spines['bottom'].set_position(('data', 0)) # 将位置设定在y轴的0.5上\nax.spines['left'].set_position(('data', 0)) # 将位置设定在x轴的0.5刻度上\n\n# 为了可以让我们的刻度显示出来不被线条遮盖\nfor label in ax.get_xticklabels() + ax.get_yticklabels():\n # 将刻度的字体大小调整\n label.set_fontsize(16)\n # 字体的矩形显示框\n label.set_bbox(dict(facecolor='white', edgecolor='None', alpha=0.65))\n\n# 注释的代码\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\nplt.plot(x, c, color='blue', linewidth=10, linestyle='-', label='cosline', alpha=0.5) # label属性制定之后的legend的内容\nplt.plot(x, s, color='red', linewidth=2.5, linestyle='-', label='sine')\nplt.legend(loc='upper left') # 将lengend图标显示在左上角,不加的话,默认将其添加在最合适的位置上\n\nt = 2 * np.pi / 3\nplt.plot([t, t], [0, np.cos(t)], color='blue', linewidth=2.5, linestyle='--') # 显示直线(t,0)->(t,cos(t))\nplt.scatter([t, ], [np.cos(t), ], 50, color='blue') # 将(t,np.cos(t))处打上点,显示\n\n# 第一个参数是注释的文本Latex嵌入,注释的目标点是(t, cos(t)), 注释的文本坐标是(-90, -100), 距离使用offset point相对与终点的\n# arrowprops箭头的属性 , arrowstyle是箭头的格式,还有连接的方式\nplt.annotate(r'$cos(\\frac{2\\pi}{3})=-\\frac{1}{2}$', \\\n xy=(t, np.cos(t)), xycoords='data', xytext=(-90, -100), \\\n textcoords='offset points', fontsize=16, arrowprops=dict(arrowstyle='->', connectionstyle='arc3,rad=.2'))\n\nt = 2 * np.pi / 3\nplt.plot([t, t], [0, np.sin(t)], color='blue', linewidth=2.5, linestyle='--') # 显示直线(t,0)->(t,cos(t))\nplt.scatter([t, ], [np.sin(t), ], 50, color='blue') # 将(t,np.cos(t))处打上点,显示\n\nplt.annotate(r'$sin(\\frac{2\\pi}{3})=-\\frac{\\sqrt{3}}{2}$', \\\n xy=(t, np.sin(t)), xycoords='data', xytext=(-90, -100), \\\n textcoords='offset points', fontsize=16, arrowprops=dict(arrowstyle='->', connectionstyle='arc3,rad=.2'))\n# -\n\n# * 数字,子图,轴的技巧使用\n# * figure : 是一个窗口\n# * subplots : 是一个窗口的子图,默认使用figure创建的是subplots(111)\n\n# %matplotlib inline\n# num是图的编号,不指定默认自增,figsize是图片的边框大小,facecolor是前景色,edgecolor是后景色,frameon是是否绘制图框\nplt.figure(num=1, figsize=(80, 80), dpi=100, facecolor='green', edgecolor='blue', frameon=True)\nplt.show()\nplt.close()\n\n# 子图命令\n# 使用子图可以在常规网格中排雷图,并按照指示制定行数和列数以及绘制图\nplt.subplot(2, 1, 1)\nplt.subplot(2, 1, 2)\n\n# Axes 很类似子图但是语序将图放置在图的任何位置\nplt.axes([0, 0, 0.5, 0.5]) # x, y, xs, ys (x, y)坐标,xs, ys是长宽\nplt.axes([0.1, 0.1, 0.5, 0.5])\nplt.axes([0.2, 0.2, 0.5, 0.5])\n\n# +\n# 颜色填充\nn = 256\nX = np.linspace(-np.pi, np.pi, n, endpoint=True)\nY= np.sin(2 * X)\n\nplt.axes([0.025, 0.025, 0.95, 0.95])\n\nplt.plot(X, Y + 1, color='blue', alpha=1.00)\nplt.fill_between(X, 1, Y + 1, color='blue', alpha=0.25) # 1 - Y + 1之间使用蓝色填充\n\nplt.plot(X, Y - 1, color='blue', alpha=1.00)\nplt.fill_between(X, -1, Y - 1, (Y - 1) > -1, color='blue', alpha=0.25) # where数组选择性填充\nplt.fill_between(X, -1, Y - 1, (Y - 1) < -1, color='red', alpha=0.25)\n\n# +\n# 散点图\nn = 1024\nX = np.random.normal(0, 1, n) # 高斯分布,均值,标准差,规模\nY = np.random.normal(0, 1, n)\nT = np.arctan2(Y, X)\n\nplt.axes([0.25, 2.5, 0.95, 0.95])\nplt.scatter(X, Y, s=75, c=T, alpha=0.5) # size大小,c颜色,alpha是透明度\n\n# +\n# 条形图\nn = 12\nx = np.arange(n)\ny1 = (1 - x / float(n)) * np.random.uniform(0.5, 1.0, n) # 均匀分布中随机采样,[low, high),size是规模\ny2 = (1 - x / float(n)) * np.random.uniform(0.5, 1.0, n)\n\nplt.bar(x, +y1, facecolor='#9999ff', edgecolor='white') # 条形图\nplt.bar(x, -y2, facecolor='#ff9999', edgecolor='white')\n\nfor x1, y in zip(x, y1):\n plt.text(x1, y + 0.05, '%.2f' % y) # x,y是文本注释的坐标,第三个参数显示的内容,alpha是透明度,family设置字体\nfor x1, y in zip(x, y2):\n plt.text(x1 + 0.4, - y - 0.05, '%.2f' % y, ha='center', va='top')\n\n\n# +\n# 等高线\n# np.meshgrid函数用来用两个一维数组(x, y)生成两个二维网格(X, Y)\n# 其中X是x的简单复制,Y是y的简单复制,两个矩阵的规模都是n * m(x - m, y - n)\ndef f(x, y):\n return (1 - x / 2 + x ** 5 + y ** 3) * np.exp(- x ** 2 - y ** 2)\n\nn = 256\nx = np.linspace(-3, 3, n)\ny = np.linspace(-3, 3, n)\nX, Y = np.meshgrid(x, y)\nZ = f(X, Y)\n\nplt.axes([0.025, 0.025, 0.95, 0.95])\nplt.imshow(Z, interpolation='nearest', cmap='bone', origin='lower')\nplt.colorbar(shrink=0.92)\n\n# +\n# 饼图\nn = 20\nZ = np.ones(n)\nplt.axes([0.025, 0.025, 0.95, 0.95])\n\nplt.pie(Z, explode=Z*0.05, colors=['%f' % (i / float(n)) for i in range(n)])\nplt.axis('equal') # 刻度对齐,保证是标准圆形\nplt.xticks()\nplt.yticks()\n\nplt.show()\n","repo_name":"gmftbyGMFTBY/Study","sub_path":"Python/AllForPython/Python-Scipy/.ipynb_checkpoints/Matplotlib-checkpoint.ipynb","file_name":"Matplotlib-checkpoint.ipynb","file_ext":"py","file_size_in_byte":7933,"program_lang":"python","lang":"en","doc_type":"code","stars":10,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"35935036017","text":"# # Author: Yoonhyuck WOO / JBNU_Industrial Information system Engineering\n# # Date; 3. 7. 2022 - 3. 17. 2022\n# # Title: Korean_NER\n# # Professor: Seung-Hoon Na\n\nimport os\nimport json\nimport numpy as np\nfrom functools import partial\nimport torch\nimport torch.nn as nn\nfrom torch.utils.data import DataLoader\nfrom torch.optim import AdamW\nfrom transformers import BertTokenizer, BertModel, FlaubertModel\n# from transformers import BertModel\nfrom sklearn.metrics import classification_report\n\nfrom DataPreprocessing import Data_NER, ner_collate_fn\nfrom modeling_ner import Bert_NER\n\nfrom ner_collate_fn import collate_fn_custom\n\nPRETAINED_MODEL_NAME = 'bert-base-multilingual-cased'\ntokenizer = BertTokenizer.from_pretrained(PRETAINED_MODEL_NAME)\nbert = BertModel.from_pretrained(PRETAINED_MODEL_NAME)\n# flaubert = FlaubertModel.from_pretrained(language_model_dir, output_loading_info=True)\n\n# # BertTokenizer\n# - return tensors = \"pt\" : Finally, you want the tokenizer to return the actual tensors that are fed to the model.\n# \n# # BertModel\n# - Attention_mask : 1 where you care and 0 where you don't care.\n# - Input_ids : the IDs of the sentence morpheme.\n# - Token_type_ids : for the question problem, but it's enough to set it to zero now.\n#\n# reference: https://huggingface.co/docs/transformers/model_doc/bert\n#\n# # Example\n\n# + active=\"\"\n# tokenizer = BertTokenizer.from_pretrained(\"bert-base-uncased\")\n# model = BertModel.from_pretrained(\"bert-base-uncased\")\n#\n# inputs = tokenizer(\"Hello, my dog is cute\", return_tensors=\"pt\")\n# outputs = model(**inputs)\n#\n# last_hidden_states = outputs.last_hidden_state\n#\n# print(inputs)\n#\n# => {'input_ids': tensor([[ 101, 7592, 1010, 2026, 3899, 2003, 10140, 102]]), 'token_type_ids': tensor([[0, 0, 0, 0, 0, 0, 0, 0]]), 'attention_mask': tensor([[1, 1, 1, 1, 1, 1, 1, 1]])}\n# -\n\n# # Loading data\n\nPATH_dir = 'C:\\\\Users\\\\LG\\\\Desktop\\\\github\\\\JBNU-2022-SPRING\\\\English world class tagging & Korean_Named Entity Recognition\\\\Ko_En_NER_POStag_data\\Ko_NER_POS'\nPATH_ko_train = os.path.join(PATH_dir, 'prepro_train.json')\nPATH_ko_test = os.path.join(PATH_dir, 'prepro_test.json')\nPATH_ko_dev = os.path.join(PATH_dir, 'prepro_dev.json')\ntotal_tag = os.path.join(PATH_dir, 'total_tag.json')\n\nwith open(PATH_ko_train) as f: \n dataset_train = json.load(f)\nwith open(PATH_ko_test) as f: \n dataset_test = json.load(f)\nwith open(PATH_ko_dev) as f: \n dataset_dev = json.load(f)\nwith open(total_tag, 'r') as f:\n tag_converter = json.load(f)\n\nprint('train', len(dataset_train))\nprint('test', len(dataset_test))\nprint('dev', len(dataset_dev))\n# tag_converter.id_to_tag\n\n# # collate_fn\n# - Tie it in batches.\n# - If the data set is variable length, it cannot be tied right away and causes an error, so you have to make a 'collate_fn' and hand it over.\n# - partial_collate_fn: when collating, with padding\n# \n# \n# - reference: https://hulk89.github.io/pytorch/2019/09/30/pytorch_dataset/\n\n# # collate_fn_custom\n\n# + jupyter={\"source_hidden\": true}\ndef collate_fn_custom(data):\n \n input_sent = [sample[0] for sample in data]\n labels = [sample[2] for sample in data]\n \n padded_inputs = torch.nn.utils.rnn.pad_sequence(input_sent, batch_first = True)\n \n return {'input': padded_inputs.contiguous(),\n 'label': torch.stack(labels).contiguous()}\n\n\n# -\n\n# # 0309\n\nfrom ner_collate_fn import make_same_len\n\ntag_to_id ={'PAD': 0, 'B-<휠': 1, 'B-OG': 2, 'I-조선': 3, 'I-PS': 4, 'B-LC': 5, 'B-1': 6, 'I-<휠': 7, 'I-LC': 8, 'B-PS': 9, 'I-TI': 10, \n 'B-목소': 11, 'O': 12, 'I-목소': 13, 'I-1': 14, 'I-': 15, 'B-조선': 16, 'I-OG': 17, 'B-': 18, 'I-DT': 19, 'B-TI': 20, 'B-DT': 21}\n\n\n# + active=\"\"\n# convert_to = []\n# def make_same_len(batch):\n# each_len_list = [len(sample) for sample in batch]\n# max_len = max(each_len_list)\n# \n# padded_batch = []\n# pad_id = 0\n# special_token = 'PAD'\n#\n# for sample in batch:\n# padded_batch.append(['CLS'] + sample+ ['PAD'] * (max_len - len(sample)) + ['SEP'])\n# \n# return padded_batch\n#\n# -\n\ndef tag_to_id2(batch):\n total = []\n convert_to = []\n for sample in batch:\n for j in sample:\n j = tag_to_id.get(j)\n convert_to.append(j)\n total.append(convert_to)\n convert_to = []\n \n return total\n\n\n\n# + active=\"\"\n# def make_same_len(batch):\n# each_len_list = [len(sample) for sample in batch]\n# max_len = max(each_len_list)\n# total = []\n# convert_to = []\n# print(max_len)\n# padded_batch = []\n# pad_id = 0\n# special_token = 'PAD'\n#\n# for sample in batch:\n# for j in sample:\n# j = tag_to_id.get(j)\n# convert_to.append(j)\n# total.append(convert_to)\n# convert_to = []\n# sample = total\n# print(len(total))\n# padded_batch.append(['PAD'] + sample + ['PAD'] * (max_len - 2) + ['PAD'])\n# total = []\n# return padded_batch\n#\n# -\n\na = [['PAD'],['B-OG','I-조선'],['I-PS','PAD','I-OG','O']]\ntag_to_id2(a)\n\nprint(make_same_len(a))\ntag_to_id2(make_same_len(a))\n\n\n\nconvert_to = []\ndef tag_to_id(data):\n for j in data:\n j = tag_to_id.get(j)\n convert_to.append(j)\n convert_to = torch.tensor(convert_to)\n \n \n return convert_to\n\n\ndef make_same_len(batch):\n each_len_list = [len(sample) for sample in batch]\n max_len = max(each_len_list)\n \n padded_batch = []\n pad_id = 0\n special_token = 'PAD'\n\n for sample in batch:\n padded_batch.append(['PAD'] + sample + ['PAD'] * (max_len - len(sample)) + ['PAD'])\n \n return padded_batch\n\n\n\nb = [[1],[2,2],[3,3,3],[4,4,4,4],[5,5,5,5,5],[6,6,6,6,6,6]]\nmake_same_len(b)\n\n# +\nconvert_to = []\ndef make_same_len(batch):\n each_len_list = [len(sample) for sample in batch]\n max_len = max(each_len_list)\n \n padded_batch = []\n pad_id = 0\n special_token = 'PAD'\n\n for sample in batch:\n padded_batch.append(['CLS'] + sample+ ['PAD'] * (max_len - len(sample)) + ['SEP'])\n \n for tag in padded_batch:\n print(tag)\n for j in tag:\n j = tag_to_id.get(j)\n convert_to.append(j)\n \n padded_batch = convert_to\n# print('before',padded_batch)\n# for tag in padded_batch:\n# print(tag)\n# print('='*90)\n# for j in tag:\n# # print(j)\n# j = tag_to_id.get(j)\n# convert_to.append(j)\n \n# padded_batch = convert_to\n# # conver_to = []\n# print('aftre')\n return padded_batch\n\n\n\n# -\n\na = [['PAD'],['B-OG','I-조선'],[2,2,2]]\nmake_same_len(a)\n\n\ndef ner_fn_custom(tokenizer, data):\n \n input_sent = [sample[0] for sample in data]\n labels = [sample[2] for sample in data]\n\n batch_inputs = tokenizer(input_sent, padding = True, return_tensors = \"pt\")\n batch_labels = make_same_len(labels)\n batch_labels = tag_to_id2(batch_labels)\n batch_labels = torch.tensor(batch_labels)\n \n return batch_inputs, batch_labels\n# return {'input': padded_inputs.contiguous(),\n# 'label': torch.stack(labels).contiguous()}\n\n\ndef ner_fn_custom(tokenizer, data):\n \n input_sent = [sample[0] for sample in data]\n labels = [sample[2] for sample in data]\n\n batch_inputs = tokenizer(input_sent, padding = True, return_tensors = \"pt\")\n batch_labels = make_same_len(labels)\n \n \n return batch_inputs, batch_labels\n# return {'input': padded_inputs.contiguous(),\n# 'label': torch.stack(labels).contiguous()}\n\nbatch_size = 10\n\n# +\nbatch_size = 1\n# partial (func, /, *args, **keywords) positional argument: args, keyword argument: keywords\n\n# partial_collate_fn = partial(collate_fn_custom, tokenizer, tag_converter)\n# partial_collate_fn = collate_fn_custom\n\npartial_collate_fn = partial(ner_fn_custom, tokenizer)\n\n'''\npartial_collate_fn = partial(collate_fn_custom, tokenizer, tag_converter)\n'''\n# ner_collate_fn: padding & making batch?\n'''\ndef ner_collate_fn(tokenizer, tag_converter):\n \n'''\n\n# +\ndataloader_train = DataLoader(\n dataset_train,\n batch_size = batch_size,\n shuffle = True,\n collate_fn = partial_collate_fn)\n\ndataloader_test = DataLoader(\n dataset_test,\n batch_size = batch_size,\n shuffle = False,\n collate_fn = partial_collate_fn)\n\ndataloader_dev = DataLoader(\n dataset_dev,\n batch_size = batch_size,\n shuffle = False,\n collate_fn = partial_collate_fn)\n\n\n# -\n\n# # BERT_NER\n# - Dropout: method to solve overfitting, one of the problems in deep learning learning. In summary, how to prevent some units of the hidden layer from being overfitted by inoperative
\n\n# + [markdown] id=\"CBE1c8fzOncZ\"\n# #Semantic Similarity\n#\n# Implementation of a semantic similarity score between texts using the [Universal Sentence Encoder](https://github.com/tensorflow/hub/blob/master/examples/colab/semantic_similarity_with_tf_hub_universal_encoder.ipynb). The Universal Sentence Encoder is employed to produce an embedding of the texts to be compared, and the similarity is measured by the inner product between the embeddings.\n#\n#\n\n# + [markdown] id=\"sR0MU_EQPxob\"\n# # Python class to compute semantic similarity score\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} cellView=\"form\" id=\"y9L_Eu7QlIMz\" outputId=\"98bea815-e351-4922-e118-be8dee03a9b7\"\n#@title double-click to see the code\n\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns\nimport tensorflow_hub as hub\n\nfrom absl import logging\nfrom dataclasses import dataclass\n\nsns.set()\n\nlogging.set_verbosity(logging.ERROR)\n\n@dataclass\nclass TextSim:\n ''' Class to compute semantic similarity among two list of strings'''\n module_url = \"https://tfhub.dev/google/universal-sentence-encoder/4\"\n model = hub.load(module_url)\n\n @staticmethod\n def clean(input):\n '''Convert inputs to list of string elements'''\n input = input if isinstance(input, list) else [input]\n return [str(element) for element in input] \n\n def embed(self, input):\n '''Convert text to embedded representation'''\n return self.model(input) \n\n def plot_all_results(self, inputA, inputB, corr):\n for i, itemA in enumerate(inputA):\n data = pd.DataFrame({'inputB': inputB, 'corr': corr[i]})\n data = data.sort_values(by='corr').reset_index(drop=True)\n self.plot_one_result(itemA, data)\n \n @staticmethod\n def plot_one_result(itemA, data):\n fig, g = plt.subplots(figsize = (8, 1*data.shape[0]))\n g = sns.barplot(x='corr', y='inputB', data=data, palette=\"Blues\")\n g.set(xlim=(0, 1))\n g.set_xlabel(None)\n g.set_ylabel(None)\n g.set_title(f\"Similarity to '{itemA}'\")\n plt.show()\n\n def compare(self, inputA, inputB, display_score=True, return_scores=False):\n '''Compute pearson correlation among two list of inputs'''\n inputA, inputB = self.clean(inputA), self.clean(inputB)\n embedA, embedB = self.embed(inputA), self.embed(inputB)\n corr = np.inner(embedA, embedB)\n\n if display_score:\n self.plot_all_results(inputA, inputB, corr)\n \n if return_scores:\n return corr\n\nts = TextSim()\n\nprint(f'Model loaded: {ts.module_url}')\n\n# + [markdown] id=\"rYQBMhWuQGzk\"\n# # Semantic Similarity Between Words\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 229} id=\"9BiulCsGx1I2\" outputId=\"bdded523-ef88-4fad-8e35-e4ba0e3e83e3\"\ncompare_from = ['animal']\ncompare_to = ['cat', 'table', 'apple']\nts.compare(compare_from, compare_to)\n\n# + [markdown] id=\"jtmW5Omjvlq1\"\n# # Semantic Similarity Between Sentences\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 393} id=\"q96oo1t0JdiJ\" outputId=\"1db64ea8-3c72-4552-f73d-96437ae3eb26\"\ncompare_from = ['The animal walks in the zoo']\ncompare_to = ['The animal walks in the zoo', \n 'The cat walks in the zoo', \n 'The cat strolls in the zoo', \n 'My cat strolls in the zoo', \n 'My cat strolls around the zoo', \n 'My cat strolls around the park']\n \nts.compare(compare_from, compare_to)\n\n# + [markdown] id=\"he5tomszvql5\"\n# # Semantic Similarity Between Questions\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 338} id=\"89Q68cUpmBWJ\" outputId=\"0b4dbd4b-ab2b-4857-9450-15670ab487a3\"\ncompare_from = ['Which brush head fits my Philips Sonicare Toothbrush?']\ncompare_to = ['How to pick a head for my toothbrush?',\n 'toothbrush I dont know what replacement I need',\n 'I dont know what replacement I need',\n 'What head should I buy for my toothbrush',\n 'My headphones are not charging']\nts.compare(compare_from, compare_to)\n","repo_name":"ossskkar/NLP","sub_path":"Semantic_Similarity_with_Universal_Sentence_Encoder.ipynb","file_name":"Semantic_Similarity_with_Universal_Sentence_Encoder.ipynb","file_ext":"py","file_size_in_byte":4378,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"38569353035","text":"# ```\n# This notebook contains summary statistics for the drugs.com depression drug reviews dataset. \n# In particular, it contains the following:\n#\n# Summary stats:\n# - Total number of reviews\n# - Total number of reviews per rating\n# - Total number of reviews per duration\n# - Total number of unique drugs\n# - Average length of comments\n# - Average length of comments per rating\n# - Average rating\n# - Average number of upvotes\n# - Average number of upvotes per rating\n# - Highest length comment\n# - Highest upvote count review\n#\n# Subpopulations to look at:\n# - Reviews with comments longer than 300 characters (arbitrary number)\n# - Reviews with more than 15 upvotes (arbitrary number)\n# \n# Relationships of interest\n# - Length of review vs number of upvotes\n# - Length of review vs rating\n# - Upvotes vs time a review has been up (current date minus review date)\n# (so we can see upvotes per review \"duration\", essentially)\n# ```\n\nimport pandas as pd\nimport numpy as np\nfrom sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer\n# %matplotlib inline\n\ndf = pd.read_csv('./data/ddc_depression_drug_reviews.csv')\ndf = df[pd.notnull(df['genRating'])] #bye bye non-rated reviews!\ndf['comment length'] = df['comment'].apply(lambda x: len(x)) #let's look at this, too\n\n# Summary statistics for the whole dataset\ndf.describe()\n\n# The longest comment\ndf[df['comment length'] == 3000].values[0][0]\n\n# The shortest comment\ndf[df['comment length'] == 6].values[0][0]\n\n# The most upvoted comment\ndf[df['upVotes'] == 1291].values[0][0]\n\n# Not very correlated...\ndf['upVotes'].corr(df['comment length'])\n\ndf.plot.scatter(x='comment length', y='upVotes')\n\n# These aren't particularly correlated, either.\ndf['genRating'].corr(df['comment length'])\n\ncomments = df['comment'].tolist()\nratings = df['genRating'].tolist()\n\n\n# +\ndef get_wordcounts(text, vectorizer=None):\n \"\"\"\n Takes a string or list of strings and returns term frequencies for each word in\n a vectorizer's vocabulary. If no vectorizer is passed, the vocabulary will be generated\n from the input text's unigrams and bigrams.\n \"\"\"\n if vectorizer is None:\n vectorizer = CountVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\n vectorizer.fit(text)\n term_counts = zip(vectorizer.get_feature_names(),\n np.asarray(vectorizer.transform(text).sum(axis=0)).ravel())\n return sorted(term_counts, key=lambda x: x[1], reverse=True)\n\ndef get_tfidf_weights(text, vectorizer=None):\n if vectorizer is None:\n vectorizer = TfidfVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\n vectorizer.fit(text)\n tfidf_weights = zip(vectorizer.get_feature_names(),\n np.asarray(vectorizer.transform(text).sum(axis=0)).ravel())\n return sorted(tfidf_weights, key=lambda x: x[1], reverse=True)\n\n\n# +\ncount_vec = CountVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\ncount_vec.fit(comments)\n\ntfidf_vec = TfidfVectorizer(stop_words='english', min_df = 2, lowercase=True, ngram_range=(1,2))\ntfidf_vec.fit(comments)\n\ntest_counts = get_wordcounts(comments, count_vec)[:10]\ntest_tfidf = get_tfidf_weights(comments, tfidf_vec)[:10]\n# -\n\nfor x in range(1,11):\n df_slice = df[df['genRating'] == x]\n print('Summary stats for reviews with a rating of {} \\n'.format(x))\n print(df_slice.describe())\n print()\n coms = df_slice['comment'].tolist()\n top_words_tfidf = get_tfidf_weights(coms, tfidf_vec)[:10]\n print('Top words (by tf-idf) for reviews with a rating of {} \\n'.format(x))\n print(str(top_words_tfidf))\n print()\n top_words = get_wordcounts(coms, count_vec)[:10]\n print('Top words (by raw count) for reviews with a rating of {} \\n'.format(x))\n print(str(top_words))\n print()\n print('********************************************************************')\n print('********************************************************************')\n print()\n","repo_name":"ebertucc/snooprx","sub_path":"Depression-Drugs-Data-Exploration.ipynb","file_name":"Depression-Drugs-Data-Exploration.ipynb","file_ext":"py","file_size_in_byte":4056,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"39934883115","text":"import sys\nsys.path.append('/Library/Frameworks/Python.framework/Versions/3.9/lib/python3.9/site-packages')\n\nfrom os import listdir\nimport numpy as np\nimport pandas as pd\nfrom AudioFeatureExtractor import AudioFeatureExtractor\n\ngenres = ['Blues', 'Classical', 'Country', 'Disco', 'Hiphop', 'Jazz', 'Metal', 'Pop', 'Rock', 'Reggae']\ncsv_name = 'features_gtzan.csv'\ngtzan_dir = '/Users/griffinbholt/Downloads/GTZAN_split_data/'\n\n# Create the AudioFeatureExtractor Object\nafe = AudioFeatureExtractor()\n\n# Get the column names for the data\nfeature_list = afe.generate_feature_list()\n\nall_feature_data = []\n\n# Extract features from each audio file in the Blues folder\n# Done: i = 0, 1, 2, 3, 4, 5, 6, 7, 8, 9\nfor i in range(10):\n folder_path = gtzan_dir + genres[i].lower()\n audio_files = listdir(folder_path)\n\n feature_data_for_this_genre = []\n\n for audio_file in audio_files:\n if '.wav' in audio_file:\n audio_data = afe.extract((folder_path + '/' + audio_file), genres[i])\n feature_data_for_this_genre.append(audio_data)\n\n all_feature_data.append(np.array(feature_data_for_this_genre))\n\nall_feature_data[9].shape\n# Blues, Classical, Country, Disco, Hiphop, Jazz, Metal, Pop, Rock, Reggae\n\nall_feature_data_stacked = np.vstack(all_feature_data)\n\nall_feature_df = pd.DataFrame(data=all_feature_data_stacked, columns=feature_list)\n\n# +\nnan_columns = []\n\nfor column in all_feature_df.columns.values:\n unique = all_feature_df[column].unique()\n if 'nan' in unique:\n nan_columns.append(column)\n# -\n\nfor column in nan_columns:\n all_feature_df[column] = all_feature_df[column].replace('nan', 0)\n\nnan_columns\n\nfor column in all_feature_df.columns.values[:-1]:\n all_feature_df[column] = pd.to_numeric(all_feature_df[column])\n\n## Save the data to a CSV\nall_feature_df.to_csv('/Users/griffinbholt/SchoolProjects/MusicGenreIdentification/gtzan_features.csv', sep=',', index=False)\n\n\n","repo_name":"griffinbholt/CS472MusicGenreIdentification","sub_path":"Code/GTZAN Feature Extraction.ipynb","file_name":"GTZAN Feature Extraction.ipynb","file_ext":"py","file_size_in_byte":1929,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"12940227247","text":"# Let's now examine the performance of integration methods for the following stiff, non-linear ODE system, **Van der Pol's oscillator**. This file solves This file solves the following ordinary differential equation (ODE) system:\n# \n# \\begin{eqnarray*}\n# \t\\frac{\\displaystyle {dy_1(t)} }{\\displaystyle {dt} } &=& y_2(t)\\\\\n# \t\\frac{\\displaystyle {dy_2(t)} }{\\displaystyle {dt} } &=& \\lambda\\left[1 - y^2_1(t)\\right]y_2(t) - y_1(t)\n# \\end{eqnarray*}\n#\n# where $\\lambda \\ge 0$ and $[y_1(0), y_2(0)]^T = [2, 1]^T$.\n\n# Import\n\n# +\nimport sys\nsys.path.append('..//source')\n\nimport matplotlib.pylab as plt\nimport numpy as np\nimport numpy.linalg as LA\n\nimport scipy, time \nfrom scipy.interpolate import CubicSpline\n\nimport analysis, integrator, params, plotter\n\nimport points\n# -\n\n# %matplotlib notebook\n\n# Functions\n\n# +\n# the stiffness parameter\nK_stiff = 20\n# https://www.cs.ox.ac.uk/files/1175/lecture06-handout.pdf\n\n# backward euler, implicit tolerance. Also sused in SDC\nBE_TOL = 1e-12\nSDC_TOL = 1e-18 #1e-12\nN_ITER_MAX_NEWTON=50\n\ndef f_eval(t, y):\n \n \"\"\"\n sovling\n dy1/dt = y2\n dy2/dt = k * (1 - y1^2)y2 - y1\n \n :param float t: the time node\n :param numpy.ndarray: y the approximate solution length( m) at time node t\n \"\"\"\n\n F = np.zeros( y.shape )\n m = len(y)\n \n # get the component-wise interaction\n F[0] = y[1]\n F[1] = K_stiff * (1 - y[0]**2) * y[1] - y[0]\n \n return F\n\ndef get_corrections_norm(D):\n \n \"\"\"\n :param numpy.ndarray D: dimensions (the number of iterations, the number of nodes, the dimension of the problem)\n \"\"\"\n \n y = np.vstack( [ np.linalg.norm(x, axis=0) for x in D] )\n \n return y\n\ndef get_stuff(p, dt):\n \n if p.t[-1] != 1:\n t = dt * np.hstack( [p.t, 1])\n else:\n t = dt * p.t\n \n S, S_p = dt * p.S, dt * p.S_p\n \n spectral_radius = p.spectral_radius\n node_type = p.node_type\n \n return t, S, S_p, spectral_radius, node_type\n\n\n# -\n\n# Parameters\n\n# +\n#\n# set up integration parameters\n#\n\n# the parameters of the simulation\nn_nodes = 10\n\n# the time step\ndt = .25\n\n# the dimension of the problem\nm = 2\n\n# the initial solution\ny0 = np.array( (2, 1) )\n\n# create the parameter object\np_lobatto = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_LOBATTO)\n\n# create paramter for radau\n#p_radau = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_RADAU)\n\n# create paramter for radau 2a\n#p_radau_2a = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_RADAU_2A)\n\n# legendre nodes\n#p_legendre = params.Params(n_nodes=n_nodes, m=m, node_type=points.GAUSS_LEGENDRE)\n\n\n# -\n\n# ---\n# Run one step\n#\n# ---\n\n# Backward Euler\n\n# +\nnodes, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\ny_be = integrator.backward_euler(f_eval, nodes, y0, S_p, be_tol=BE_TOL, do_print=True)\n\nt = nodes\n#LA.norm( y_be - exact(t), ord=np.inf)\n# -\n\n# SDC\n\n# +\nt, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\n\ny_be = integrator.backward_euler(f_eval, t, y0, S_p, be_tol=BE_TOL)\n\ny_sdc, Y_sdc, D_sdc, is_converged_sdc = integrator.sdc(f_eval, t, y0, y_be, S, S_p, be_tol=BE_TOL, sdc_tol=SDC_TOL, do_print=True)\nprint('SDC converged: %s ' % is_converged_sdc)\n\n# the norm of the corrections\nd_norm_sdc, log_d_norm_sdc, d_norm_rel_sdc, log_d_norm_rel_sdc = analysis.analyze_corrections(D_sdc, Y_sdc)\n# -\n\n# JFNK\n\n# +\nt, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\n\ny_be = integrator.backward_euler(f_eval, t, y0, S_p, be_tol=BE_TOL)\n\ny_jfnk, Y_jfnk, D_jfnk, is_converged_jfnk, is_stiff,ratios = integrator.jfnk(f_eval, t, y0, y_be, S, S_p, spectral_radius, be_tol=BE_TOL, \\\n sdc_tol=SDC_TOL, do_print=True)\nprint('JFNK converged: %s ' % is_converged_jfnk)\n\n# the norm of the corrections\nd_norm_jfnk, log_d_norm_jfnk, d_norm_rel_jfnk, log_d_norm_rel_jfnk = analysis.analyze_corrections(D_jfnk, Y_jfnk)\n# -\n\n# Spectral Solution\n\n# +\nt, S, S_p, spectral_radius, node_type = get_stuff(p_lobatto, dt)\n\ny_be = integrator.backward_euler(f_eval, t, y0, S_p, be_tol=BE_TOL)\n\ny_spect = integrator.run_spectral(f_eval, t, y0, S, tol=1e-10, y_approx=y_be, verbose=True)\n# -\n\n# Plot the corrections\n\n# +\nplt.figure()\nplt.title('Magnitude of corrections ')\n\nplt.plot( range(log_d_norm_sdc.shape[0]), log_d_norm_sdc.max(axis=1), '-o', label='sdc')\nplt.plot( range(log_d_norm_jfnk.shape[0]), log_d_norm_jfnk.max(axis=1), '-o', label='jfnk')\n\nplt.xlabel('SDC iteration')\nplt.ylabel('log10 (error)')\nplt.legend(loc='best')\nplt.show()\n# -\n\n# ---\n# Run Multiple Steps\n#\n# ---\n\n# +\nt_final =1\ntol = 1e-5\ndt_init = 4e-1\n\nprint(t_final, dt_init)\n# -\n\n# **Adaptive** JFNK\n\nt_adapt, y_adapt, Y_adapt, D_adapt, h_adapt = integrator.jfnk_adaptive(f_eval, t_init=0, t_final=t_final, dt_init=dt_init, p=p_lobatto, y0=y0, \\\n tol=tol, do_print=True)\n\n# the time steps\nh_adapt\n\n# **Uniform** JFNK\n\nt_uni, y_uni, Y_uni, D_uni = integrator.jfnk_uniform(f_eval, t_init=0, t_final=t_final, n_steps=12, p=p_lobatto, y0=y0, do_print=True)\n\n# **Reference** solution\n\n# +\n# the reference solution\nn_steps_ref = 100\np_ref = params.Params(n_nodes=10, m=m, node_type=points.GAUSS_LOBATTO)\n\nt_ref, y_ref, Y_ref, D_ref = integrator.jfnk_uniform(f_eval, t_init=0, t_final=t_final, n_steps=n_steps_ref, p=p_ref, y0=y0, do_print=True)\n\n# spline\ny_spline = CubicSpline(t_ref, y_ref)\n\n# errors\nf = lambda t, y: ( LA.norm( y_spline(t) - y, ord=np.inf), integrator.relative_norm(y_spline(t) - y, y_spline(t)) )\n# -\n\n# Comparing the absolute and relative erros of the **uniform** solution to the reference solution\n\nf(t_uni, y_uni)\n\n# Comparing the absolute and relative errors of the **adaptive** solution to the reference solution\n\nf(t_adapt, y_adapt), tol\n\n# ---\n# Plotting\n#\n# ---\n\n# Plot the JFNK corrections for each time step\n\n# +\nplotter.plot_correction_time_steps(D_uni, Y_uni)\n\nplt.show()\n# -\n\n# Plot norm of corrections\n\n# +\ny_spect_norm = LA.norm(y_spect, axis=0)\n\nlabels=['y1', 'y2']\n\nplotter.corrections(d_norm_sdc, d_norm_jfnk, y_spect_norm, do_rerr=True, labels=labels, do_legend=True)\nplt.show()\n# -\n\n# ---\n# Errors\n#\n# ---\n\n#\n# error\n#\naerr_be, rerr_be = analysis.error_analysis(y_be, y_spect)\naerr_sdc, rerr_sdc = analysis.error_analysis(y_sdc, y_spect)\naerr_jfnk, rerr_jfnk = analysis.error_analysis(y_jfnk, y_spect)\n\n\n","repo_name":"namdi/JFNK-integrator","sub_path":"examples/van_der_pol.ipynb","file_name":"van_der_pol.ipynb","file_ext":"py","file_size_in_byte":6416,"program_lang":"python","lang":"en","doc_type":"code","stars":5,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"11900144276","text":"# ## Please first install these libraries\n# pip install pythreejs pyntcloud pandas numpy\n\nfrom pyntcloud import PyntCloud\nimport pandas as pd\nimport os\nimport numpy as np\nimport PIL.Image as Image\n# %matplotlib inline\n\ndef load_velo_scan(velo_filename):\n scan = np.fromfile(velo_filename, dtype=np.float32)\n scan = scan.reshape((-1, 4))\n return scan\n\n\n#point cloud 형식으로 만들기 위해서 rgb 데이터 입히기\ndef paint_points(points, color = [0,0,100]):\n # color = [r, g, b]\n color = np.array([color])\n new_pts = np.zeros([points.shape[0],6])\n new_pts[:,:3] = points\n new_pts[:, 3:] = new_pts[:, 3:] + color\n return new_pts\n\n\npath = '/home/joonsu0109/Joon/Image_to_Points/depth_code.bin'\n\npoints = load_velo_scan(path)[:,:3]\n\npoints.shape\n\npd_points = pd.DataFrame(paint_points(points, img_edit), columns=['x','y','z','red','green','blue'])\n\ncloud = PyntCloud(pd_points)\ncloud.plot(initial_point_size=0.00001)\n\ncloud\n\ncloud.to_file(\"depth_color.ply\")\n\n# +\nimport open3d as o3d\n\npcd = o3d.io.read_point_cloud(\"/home/joonsu0109/Joon/Image_to_Point/depth_color.ply\")\no3d.io.write_point_cloud(\"depth_color.pcd\", pcd)\n# -\n\n# ---\n\nimport cv2\nimg = cv2.imread('/home/joonsu0109/Joon/Image_to_Point/set/000002.png')\n\n\n","repo_name":"joonsu0109gh/Image_to_Points","sub_path":"Image_to_Points/bin_to_pcd.ipynb","file_name":"bin_to_pcd.ipynb","file_ext":"py","file_size_in_byte":1250,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"5966523361","text":"# +\nimport os\nimport numpy as np\nimport decoy_mnist\nfrom matplotlib import pyplot as plt\n\nfrom keras.layers import Input, Conv2D, MaxPool2D, Dense, Flatten, UpSampling2D, Multiply, Reshape\nfrom keras.models import Model, load_model\nfrom keras.utils import to_categorical\nfrom keras.optimizers import SGD\nfrom keras.regularizers import l1, l2\nfrom keras.losses import kld\nfrom keras.datasets import mnist\nfrom keras import backend as K\n# -\n\n# ### Load decoy MNIST\n\nXr, x_train, y_train, E, Xtr, x_test, y_test, Et = decoy_mnist.generate_dataset(cachefile='data/decoy-mnist.npz')\nx_train = x_train / 255 / 5 + 4/5\nx_test = x_test / 255 / 5 + 4/5\nx_train = x_train.reshape((60000, 28, 28, 1))\nx_test = x_test.reshape((10000, 28, 28, 1))\ny_train = to_categorical(y_train, 10)\ny_test = to_categorical(y_test, 10)\n\n\n# ### Load normal MNIST\n\n# +\ndef load_mnist():\n print('Loading MNIST')\n (x_train, y_train), (x_test, y_test) = mnist.load_data()\n x_train = np.expand_dims(x_train, -1) / 255 / 5 + 4 / 5\n x_test = np.expand_dims(x_test, -1) / 255 / 5 + 4 / 5\n y_train = to_categorical(y_train, 10)\n y_test = to_categorical(y_test, 10)\n return (x_train, y_train), (x_test, y_test)\n\n\n(x_train_g, y_train_g), (x_test_g, y_test_g) = load_mnist()\n\n\n# -\n\n# ## Model looking at corner splotches\n\ndef ConvNet(train):\n if train or not os.path.isfile('classifier_d.hdf5'):\n print('Classifier model is being trained')\n inputs = Input([28, 28, 1])\n x = Conv2D(32, (3, 3), activation='relu', name='conv1')(inputs)\n x = Conv2D(64, (3, 3), activation='relu', name='conv2')(x)\n x = MaxPool2D((2, 2), name='mp1')(x)\n x = Conv2D(64, (3, 3), activation='relu', name='conv3')(x)\n x = Conv2D(128, (3, 3), activation='relu', name='conv4')(x)\n x = Flatten(name='flatten')(x)\n x = Dense(10, activation='softmax', name='dense1')(x)\n \n model = Model(inputs, x)\n model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])\n model.fit(x_train, y_train, batch_size=1000, epochs=15, validation_split=1/6)\n model.save('classifier_d.hdf5')\n else:\n print('Classifier model loaded')\n model = load_model('classifier_d.hdf5')\n return model\n\n\nconvnet1 = ConvNet(train=True)\nconvnet1.trainable = False\nprint(convnet1.evaluate(x_test, y_test))\nprint(convnet1.evaluate(x_test_g, y_test_g))\n\n# We see that model is robust and performs well on both decoy MNIST and regular MNIST test sets.\n\n# ### Explaining robust model\n\nmodel = convnet1\n\n\n# +\ndef normalized_kld(p, q):\n return 1 - K.exp(-kld(p, q))\n\ndef ExplanationModel(reg, train):\n if train or not os.path.isfile('exp_model_d.hdf5'):\n inputs = Input(shape=x_train.shape[1:])\n\n x = Conv2D(32, (3, 3), activation='relu', padding='same')(inputs)\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n x = MaxPool2D((2, 2))(x)\n\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n x = Conv2D(128, (3, 3), activation='relu', padding='same')(x)\n x = MaxPool2D((2, 2))(x)\n\n x = UpSampling2D((2, 2))(x)\n x = Conv2D(128, (3, 3), activation='relu', padding='same')(x)\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n\n x = UpSampling2D((2, 2))(x)\n x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)\n x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)\n\n x = Conv2D(1, (1, 1), activation='sigmoid', padding='same', name='attention', activity_regularizer=l2(reg))(x)\n\n x = Multiply()([x, inputs])\n x = model(x)\n\n exp_model = Model(inputs, x)\n\n y_train = model.predict(x_train)\n exp_model.compile(optimizer='adam', loss=normalized_kld)\n exp_model.summary()\n for l in exp_model.layers[-1].layers:\n l.trainable = False\n exp_model.fit(x_train, y_train, epochs=5)\n exp_model.save('exp_model_d.hdf5')\n else:\n exp_model = load_model('exp_model_d.hdf5', compile=False)\n exp_model.compile(optimizer='sgd', loss=normalized_kld)\n return exp_model\n\n\n# -\n\nexp_model = ExplanationModel(reg=0.0001, train=True)\n\n# +\nexp_model = load_model('exp_model_d.hdf5', compile=False)\nsaliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\nsaliency_values = saliency([x_test])[0]\nprediction = model.predict(x_test, batch_size=1000)\ny = model.predict(x_test)\n\nfor i in range(5):\n print('Processing image #{}'.format(i+1))\n exp_model = load_model('exp_model_d.hdf5', compile=False)\n opt = SGD(lr=0.01)\n exp_model.compile(optimizer=opt, loss=normalized_kld)\n saliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n\n img = x_test[i, :, :, 0]\n\n plt.subplot(3, 3, 2)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n orig_sal = saliency_values[i].reshape((28, 28))\n sal_interp = (orig_sal - orig_sal.min()) / (orig_sal.max() - orig_sal.min())\n plt.imshow(sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(3, 3, 1)\n plt.axis('off')\n plt.title('Original saliency')\n plt.imshow(orig_sal, cmap='jet', vmin=0, vmax=1)\n\n for l in exp_model.layers[-1].layers:\n l.trainable = False\n hist = exp_model.fit(x_test[i:i + 1], y[i:i + 1], epochs=100, verbose=False)\n saliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n final_sal = saliency([x_test[i:i + 1]])[0].reshape((28, 28))\n final_sal_interp = (final_sal - final_sal.min()) / (final_sal.max() - final_sal.min())\n\n plt.subplot(2, 2, 1)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n plt.imshow(final_sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(2, 2, 2)\n plt.axis('off')\n plt.title('Final saliency')\n plt.imshow(final_sal, cmap='jet', vmin=0, vmax=1)\n\n\n plt.subplot(2, 2, 3)\n plt.title('Fine-tuning loss')\n plt.plot(hist.history['loss'])\n\n plt.subplot(2, 2, 4)\n plt.title('Saliency histogram')\n plt.hist(orig_sal.flatten())\n\n plt.show()\n\n\n# -\n\n# Saliency maps show that model is not paying attention to corner splotches.\n\n# ## Model looking at corner splotches\n\ndef ConvNet2(train):\n if train or not os.path.isfile('classifier_d2.hdf5'):\n print('Classifier model is being trained')\n inputs = Input([28, 28, 1])\n x = Flatten(name='flatten')(inputs)\n x = Dense(1024, activation='relu', name='dense1')(x)\n x = Dense(10, activation='softmax', name='dense2')(x)\n \n model = Model(inputs, x)\n model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])\n model.fit(x_train, y_train, batch_size=100, epochs=30, validation_split=1/6)\n model.save('classifier_d2.hdf5')\n else:\n print('Classifier model loaded')\n model = load_model('classifier_d2.hdf5')\n return model\n\n\nconvnet2 = ConvNet2(train=True)\nconvnet2.trainable = False\nprint(convnet2.evaluate(x_test, y_test))\nprint(convnet2.evaluate(x_test_g, y_test_g))\n\n# Model performs well on training decoy data but fails badly on test decoy data and even worse on normal test data.\n\n# ### Explaining bad model\n\nmodel = convnet2\n\nexp_model = ExplanationModel(reg=0.00001, train=True)\n\n# +\nexp_model = load_model('exp_model_d.hdf5', compile=False)\nsaliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n\nsaliency_values = saliency([x_test])[0]\nprediction = model.predict(x_test, batch_size=1000)\ny = model.predict(x_test)\n\nfor i in range(15):\n print('Processing image #{}'.format(i+1))\n exp_model = load_model('exp_model_d.hdf5', compile=False)\n opt = SGD(lr=0.01)\n exp_model.compile(optimizer=opt, loss=normalized_kld)\n\n img = x_test[i, :, :, 0]\n\n plt.subplot(2, 2, 2)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n orig_sal = saliency_values[i].reshape((28, 28))\n sal_interp = (orig_sal - orig_sal.min()) / (orig_sal.max() - orig_sal.min())\n plt.imshow(sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(2, 2, 1)\n plt.axis('off')\n plt.title('Original saliency')\n plt.imshow(orig_sal, cmap='jet', vmin=0, vmax=1)\n\n hist = exp_model.fit(x_test[i:i + 1], y[i:i + 1], epochs=100, verbose=False)\n saliency = K.function(inputs=exp_model.inputs, outputs=[exp_model.get_layer('attention').output])\n final_sal = saliency([x_test[i:i + 1]])[0].reshape((28, 28))\n final_sal_interp = (final_sal - final_sal.min()) / (final_sal.max() - final_sal.min())\n\n plt.subplot(2, 2, 4)\n plt.axis('off')\n plt.title('Interpolated')\n plt.imshow(img, cmap='gray')\n plt.imshow(final_sal_interp, alpha=0.5, cmap='jet')\n\n plt.subplot(2, 2, 3)\n plt.axis('off')\n plt.title('Final saliency')\n plt.imshow(final_sal, cmap='jet', vmin=0, vmax=1)\n plt.show()\n \n plt.subplot(1, 2, 1)\n plt.title('Fine-tuning loss')\n plt.plot(hist.history['loss'])\n\n plt.subplot(1, 2, 2)\n plt.title('Saliency histogram')\n plt.hist(orig_sal.flatten())\n\n plt.show()\n# -\n\n# Explanations show that this bad model is indeed looking at corner splotches.\n","repo_name":"eclique/learning-saliency","sub_path":"mnist/Decoy MNIST.ipynb","file_name":"Decoy MNIST.ipynb","file_ext":"py","file_size_in_byte":9261,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"27823260185","text":"# + deletable=true editable=true\nimport cv2\nimport tensorflow\nimport os\nimport matplotlib.pyplot as plt\nos.environ[\"CUDA_DEVICE_ORDER\"]=\"PCI_BUS_ID\"\nos.environ[\"CUDA_VISIBLE_DEVICES\"]=\"4\"\n\n# + deletable=true editable=true\n# the data path\ndata_path ='../../data/MPIIGaze/Data/Original/'\nimg_list = []\ndef Read_img(data_path):\n for data in os.listdir(data_path):\n path = os.path.join(data_path, data)\n or_img = cv2.imread(path)\n if or_img != None:\n# plt.imshow(or_img[180:540, 320:960]) # 显示图片\n# plt.axis('off') # 不显示坐标轴\n# plt.show()\n \n# img_get = or_img.getdata()\n# print(img_get)\n or_img.tolist()\n# print(or_img)\n img_list.append(or_img)\n if os.path.isdir(path):\n Read_img(path)\n \n\n\n# + deletable=true editable=true\nRead_img(data_path)\n\n# + deletable=true editable=true\n# img1 = cv2.imread(filenames,cv2.IMREAD_GRAYSCALE) #读取图片,第二个参数表示以灰度图像读入\n# if img1 is None: #判断读入的img1是否为空,为空就继续下一轮循环\n# continue\n# res1= cv2.resize(img1,(28,28)) #对图片进行缩放,第一个参数是读入的图片,第二个是制定的缩放大小\n# res1_1 = res1.reshape(1,784)/255 #将表示图片的二维矩阵转换成一维\n# res1_1_1 = res1_1.tolist() #将numpy.narray类型的矩阵转换成list\n# train_set_x.append(res1_1_1)\n\n# + deletable=true editable=true\n\n","repo_name":"qbqandyxq/My_gaze","sub_path":"code/Read_img.ipynb","file_name":"Read_img.ipynb","file_ext":"py","file_size_in_byte":1576,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"37780962312","text":"# # Lab | Matplotlib & Seaborn\n#\n# #### Import all the necessary libraries here:\n\n# Libraries\nimport numpy as np\n\n# ## Challenge 1\n#\n# #### The data we will use in this challenge is:\n\nx = np.arange(0,100)\ny = x*2\nz = x**2\n\n# #### Plot (x, y) and (x, z).\n# There are 2 ways of doing this. Do in both ways.\n#\n# **Hint**: Check out the nrows, ncols and index arguments of subplots. \n#\n# Also, play around with the linewidth and style. Use the ones you're most happy with.\n\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\n# +\n# Method 1\n# 1 Figure with 2 subplots, together \nfig = plt.figure()\n\n# (x, y) on the first plot\nplt.plot(x, y, color='orange', label='(x, y)')\n\n# (x, z) on the second plot\nplt.plot(x, z, color='blue', label='(x, z)')\n\nplt.legend()\n\nplt.show()\n# -\n\n# #### Use plt.subplots(nrows=1, ncols=2) to create the plot.\n\n# +\n# Method 2\n# Figure with 1 row and 2 columns of subplots\nfig, ax = plt.subplots(nrows=1, ncols=2)\n\n# (x, y) on the first subplot\nax[0].plot(x, y, color='orange', label='(x, y)')\nax[0].legend()\n\n# (x, z) on the second subplot\nax[1].plot(x, z, color='blue', label='(x, z)')\nax[1].legend()\n\n# Show the figure\nplt.show()\n# -\n\n# #### Use your previous code but now, resize your plot.\n# **Hint**: Add the figsize argument in plt.subplots().\n#\n# If you want, try to add a title to the plot or even axes labels. You can also play with the fontweight and fontsize of the titles and labels. \n\n# +\n# your code here\n# Resizing the plot and doing it all\nfig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))\n\n# (x, y) on the first subplot\nax[0].plot(x, y, color='orange', label='(x, y)')\nax[0].legend()\nax[0].set_title(\"(x,y)\", fontweight='bold', fontsize=16)\nax[0].set_xlabel(\"X\", fontsize=10)\nax[0].set_ylabel(\"Y\", fontsize=10)\n\n# (x, z) on the second subplot\nax[1].plot(x, z, color='blue', label='(x, z)')\nax[1].legend()\nax[1].set_title(\"(x,z)\", fontweight='bold', fontsize=16)\nax[1].set_xlabel(\"X\", fontsize=10)\nax[1].set_ylabel(\"Z\", fontsize=10)\n\n# Show the figure\nplt.show()\n# -\n\n# #### Plot both $y=x^2$ and $y=e^x$ in the same plot using normal and logarithmic scale.\n# **Hint**: Use `set_xscale` and `set_yscale`.\n\n# +\n# your code here\n# -\n\n# #### As a bonus challenge, try to add a legend to the plot.\n\n# +\n# your code here\nx = np.arange(0,100)\ny = x**2\nz = np.exp(x)\n\nfig, axs = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))\n\naxs[0].plot(x, y, label='y = x^2')\naxs[0].plot(x, z, label='z = e^x')\naxs[0].set_title('Normal Scale')\naxs[0].legend()\n\naxs[1].plot(x, y, label='y = x^2')\naxs[1].plot(x, z, label='z = e^x')\naxs[1].set_title('Logarithmic Scale')\naxs[1].set_xscale('log')\naxs[1].set_yscale('log')\naxs[1].legend()\n\nplt.show()\n# -\n\n# ## Challenge 2\n# #### Import the `Fitbit2` dataset and store it in a variable called `fitbit`. You can find the dataset in Ironhack's database:\n# * db: `fitbit`\n# * table: `fitbit2`\n\n# +\n# your code here\nimport pandas as pd\n\n# Read the CSV file\nfitbit = pd.read_csv('Fitbit2.csv')\nfitbit\n# -\n\nfitbit.columns\n\n# #### From the Fitbit data, we want to visually understand:\n#\n# How the average number of steps change by month. Use the appropriate visualization to show the median steps by month. Is Fitbitter more active on weekend or workdays? All plots must be in the same jupyter notebook cell.\n#\n# **Hints**:\n#\n# * Use Months_encoded and Week_or Weekend columns.\n# * Use matplolib.pyplot object oriented API.\n# * Set your size figure to 12,4\n# * Explore plt.sca\n# * Explore plt.xticks\n# * Save your figures in a folder called `figures` in your repo. \n\n# +\n# your code here\n# MEDIAN STEPS BY MONTH_ENCODED\nfig, axs = plt.subplots(1, 2, figsize=(12, 4))\nmonthly_steps = fitbit.groupby(['Months_encoded'])['Steps'].median()\nmonthly_steps.plot(kind='bar', ax=axs[0], color = 'grey')\naxs[0].set_title('Median Steps by Month')\naxs[0].set_xlabel('Month')\naxs[0].set_ylabel('Steps')\n\n# MEDIAN STEPS BY WORK_OR_WEEKEND\nweekly_steps = fitbit['Steps'].groupby(fitbit['Work_or_Weekend']).median()\nplt.barh(weekly_steps.index, weekly_steps.values, height= 0.9, color= 'black', align='center', )\nplt.yticks([0, 1], ['Weekend', 'Work'])\nplt.show()\n\n# Save the figure\nplt.show()\n# -\n\n# #### Write a loop to plot 3 scatter plots of the following features:\n#\n# * Minutes Lightly Active vs Steps \n# * Minutes Very Active vs Steps \n# * Minutes Sedentary vs Steps \n\n# +\n# your code here\nfig, axs = plt.subplots(1, 3, figsize=(12, 4))\n\nfeatures = ['Minutes Lightly Active', 'Minutes Very Active', 'Minutes Sedentary']\n\nfor i, feature in enumerate(features):\n axs[i].scatter(fitbit[feature], fitbit['Steps'], color = 'purple')\n axs[i].set_xlabel(feature)\n axs[i].set_ylabel('Steps')\nplt.show()\n# -\n\n# ## Challenge 3\n#\n# #### Import the `titanic` dataset and store it in a variable called `titanic`. You can find the dataset in Ironhack's database:\n# * db: `titanic`\n# * table: `titanic`\n\n# your code here\n# Read the CSV file\ntitanic = pd.read_csv('titanic.csv')\ntitanic\n\n# #### Explore the titanic dataset using Pandas dtypes.\n\n# your code here\ntitanic.dtypes\n\n# #### What are your numerical variables? What are your categorical variables?\n# **Hint**: Use Pandas select_dtypes.\n\n# NUMERICAL VARIABLES\n# your code here\ntitanic.select_dtypes(include='number')\n\n# CATEGORICAL VARIABLES\n# your code here\ntitanic.select_dtypes(include='object')\n\n# #### Set the plot style to classic and the figure size to (12,6).\n# **Hint**: To set the style you can use matplotlib or seaborn functions. Do some research on the matter.\n\n# your code here\nplt.style.use('classic')\nsns.set_style(\"darkgrid\")\nplt.rcParams['figure.figsize'] = (12, 6)\n\n# #### Use the right visulalization to show the distribution of column `Age`.\n\n# +\n# your code here\ntitanic['Age'].hist(bins=50, color = 'green')\n\nplt.title(\"Distribution\")\nplt.xlabel(\"Age\")\nplt.ylabel(\"Count\")\n\nplt.show()\n# -\n\n# #### Use subplots and plot the distribution of the `Age` with bins equal to 10, 20 and 50.\n\n# +\n# your code here\nplt.subplot(1,3,1)\ntitanic['Age'].hist(bins = 10, color = \"orange\")\nplt.subplot(1,3,2)\ntitanic['Age'].hist(bins=20, color = \"green\")\nplt.subplot(1,3,3)\ntitanic['Age'].hist(bins=30, color = \"red\")\n\nplt.show()\n# -\n\n# #### How does the bin size affect your plot?\n\n\"\"\"\nyour comments here \nAs the bin size increases, each bin will represent a larger range of values. This makes the plot more difficult to read.\n\"\"\"\n\n# #### Use seaborn to show the distribution of column `Age`.\n\n# your code here\nsns.distplot(titanic['Age'])\n\n# #### Use the right plot to visualize column `Gender`. There are 2 ways of doing it. Do it both ways.\n# **Hint**: Use matplotlib and seaborn.\n\n# +\n# Method 1 - matplotlib\n# your code here\ngender_counts = titanic['Gender'].value_counts()\n\nplt.bar(gender_counts.index, gender_counts.values, color = \"grey\")\nplt.xlabel('Gender')\nplt.ylabel('Occurrences')\nplt.show()\n\n# +\n# Method 2 - seaborn\n# your code here\n\nsns.countplot(x='Gender', data=titanic, color = \"grey\")\nplt.xlabel('Gender')\nplt.ylabel('Number of occurrences')\nplt.show()\n# -\n\n# #### Use the right plot to visualize the column `Pclass`.\n\n# +\n# your code here\npclass_counts = titanic['Pclass'].value_counts()\n\n# Create a bar plot\nplt.bar(pclass_counts.index, pclass_counts.values, color = \"grey\")\nplt.xlabel('Passenger Class')\nplt.ylabel('Count')\nplt.title('Distribution of Passenger Classes')\nplt.show()\n# -\n\n# #### We would like to have in one single plot the summary statistics of the feature `Age`. What kind of plot would you use? Plot it. \n\n# your code here\nsns.boxplot(x=titanic['Age'], color = \"orange\")\n\n# +\nQ1 = titanic[\"Age\"].quantile(0.25)\nQ2 = titanic[\"Age\"].quantile(0.50)\nQ3 = titanic[\"Age\"].quantile(0.75)\n\nprint(Q1)\nprint(Q2) # median\nprint(Q3)\n# -\n\n\"\"\"\nyour comments here \nThis plot shows the median, quartiles, and outliers of the data and can provide a quick visual representation of \nthe distribution of the data.\n\n\"\"\"\n\n# #### What does the last plot tell you about the feature `Age`?\n\n\"\"\"\nyour comments here\n\n25% of the values in the Age feature are less than or equal to 22.0, 50% of the values are less than or equal to 28.5, \nand 75% of the values are less than or equal to 37.0.\n\n\"\"\"\n\n# #### Now in addition to the summary statistics, we want to have in the same plot the distribution of `Age`. What kind of plot would you use? Plot it. \n\n# your code here\nplt.figure(figsize=(10,5))\nsns.violinplot(data=titanic, x='Age', color = \"purple\")\nplt.show()\n\n# #### What additional information does the last plot provide about feature `Age`?\n\n\"\"\"\nyour comments here\nIt shows the distribution of the data through the width of the violin and the summary statistics - Minimum and maximum at both \nends; median, and quartiles through the white dot in the center and the black bar inside the violin).\n\"\"\"\n\n# #### We suspect that there is a linear relationship between `Fare` and `Age`. Use the right plot to show the relationship between these 2 features. There are 2 ways, please do it both ways.\n# **Hint**: Use matplotlib and seaborn.\n\n# Method 1 - matplotlib\n# your code here\nplt.scatter(titanic['Age'], titanic['Fare'], s = 100)\nplt.xlabel('Age')\nplt.ylabel('Fare')\nplt.show()\n\n# Method 2 - seaborn\n# your code here\nsns.scatterplot(x='Age', y='Fare', data=titanic, s = 100)\n\n# #### Plot the correlation matrix using seaborn.\n\n# your code here\ncorr = titanic.corr()\nsns.heatmap(corr, annot=True, cmap='Reds')\n\n# #### What are the most correlated features?\n\n\"\"\"\nyour comments here\n- Fare & Survived\n- Parch & SibSp\n\"\"\"\n\n# #### Use the most appropriate plot to display the summary statistics of `Age` depending on `Pclass`.\n\n# your code here\nsns.boxplot(x='Pclass', y='Age', data=titanic)\nplt.title('Summary Statistics of Age by Passenger Class')\nplt.xlabel('Passenger Class')\nplt.ylabel('Age')\nplt.show()\n\n# #### Use seaborn to plot the distribution of `Age` based on the `Gender`.\n# **Hint**: Use Facetgrid.\n\n# your code here\ngender = sns.FacetGrid(titanic, col=\"Gender\")\ngender.map(sns.histplot, \"Age\", kde=False)\ngender.add_legend()\nplt.xlabel('Age')\nplt.ylabel('Frequency')\nplt.show()\n","repo_name":"sfmar15/lab-matplotlib-seaborn","sub_path":"your-code/main.ipynb","file_name":"main.ipynb","file_ext":"py","file_size_in_byte":10034,"program_lang":"python","lang":"en","doc_type":"code","dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"23779972301","text":"# #### Imports\n\n# +\nimport os\nimport csv\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport tensorflow as tf\nimport seaborn as sns\nfrom tensorflow.keras import datasets, layers, models\n\nfrom sklearn.model_selection import train_test_split, KFold\nfrom sklearn.metrics import confusion_matrix, roc_curve, auc, classification_report, precision_recall_curve\nfrom sklearn.utils import shuffle\nfrom PIL import Image\n\n\n# -\n\n# ## Methods for Experiments\n\ndef get_distr(targets, labels):\n vals = {}\n for label in labels.keys():\n vals[label] = 0\n \n for target in targets:\n vals[target] += 1\n \n for label in labels.keys():\n print(str(labels[label]) + \": \" + str(vals[label]))\n \n fig = plt.figure()\n ax = fig.add_axes([0,0,1,1])\n langs = labels.values()\n students = vals.values()\n ax.bar(langs,students)\n plt.show()\n\n\ndef plot_ROC(y_test, y_pred, title):\n fpr, tpr, thresholds = roc_curve(y_test, y_pred)\n auc_val = auc(fpr, tpr)\n plt.plot(fpr,tpr) \n plt.plot([0, 1], [0, 1], linestyle='dashed')\n plt.axis([-0.05, 1.05, -0.05, 1.05])\n plt.xlabel('False Positive Rate') \n plt.ylabel('True Positive Rate') \n title += \"\\nAUC: \" + str(round(auc_val, 4))\n plt.title(title)\n plt.show()\n\n\ndef plot_cm(y_test, y_pred, title):\n cm = confusion_matrix(y_test, y_pred)\n cm_df = pd.DataFrame(cm)\n cm_df = (cm_df.T/np.sum(cm_df, axis=1)).T\n sns.heatmap(cm_df, annot=True, cmap=plt.cm.Blues, fmt=\".2%\")\n plt.imshow(cm)\n plt.xlabel(\"Predicted Label\")\n plt.ylabel(\"True Label\")\n plt.title(title)\n\n\ndef plot_pr(y_test, y_pred, title):\n new_pred = []\n for val in y_pred:\n if val >= 0.5:\n new_pred.append(val)\n else:\n new_pred.append(0)\n prec, recall, _ = precision_recall_curve(y_test, new_pred)\n plt.plot(recall, prec)\n plt.ylim([0, 1])\n plt.xlabel(\"Recall\")\n plt.ylabel(\"Precision\")\n plt.title(title + \" Precision vs Recall\")\n\n\ndef convert_pred(y_pred, threshold):\n new_pred = np.empty((y_pred.size), dtype='int32')\n for i in range(y_pred.size):\n if y_pred[i] >= threshold:\n new_pred[i] = 1\n else:\n new_pred[i] = 0\n \n return new_pred\n\n\n# # Part 0: Load in csv data and format it to images and target data\n#\n# Read in csv of metadata for the images and select target data. For our target data we are selecting COVID/Non-COVID, Survival, Intubation, and Admission to the ICU - resulting in 28 classes where class 0 = Non-COVID and class > 0 is COVID and some unique combination of the classes above. For the initial training and testing of this data, any class > 0 we set to 1 to train a binary classifier for COVID/Non-COVID\n\n# +\ndata = []\nwith open(\"metadata.csv\") as file:\n for row in csv.reader(file):\n data.append(row)\n\ndata = data[1:]\n\nd = {'filename': [], 'target': []}\n\nfor image in data:\n # skip volume data\n if image[22] == \"volumes\":\n continue\n \n # ignore CT scan data\n if image[19] == \"CT\":\n continue\n \n diagnosis = image[4].split(\"/\")\n diagnosis = diagnosis[len(diagnosis) - 1].lower()\n # set target based on meta data\n if diagnosis != \"covid-19\":\n if diagnosis == \"todo\":\n # if not classified yet target = -1\n # d['filename'].append(image[23])\n # d['target'].append(-1)\n # ignore unknown data\n continue\n else:\n # if not COVID-19 target = 0\n d['filename'].append(image[23])\n d['target'].append(0)\n else:\n # if it is COVID-19\n if image[6] == \"Y\":\n # if they survived\n if image[7] == \"Y\":\n # if they were intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(1)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(2)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(3)\n elif image[7] == \"N\":\n # if they were not intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(4)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(5)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(6)\n else:\n # if they were intubated is unknown\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(7)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(8)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(9)\n elif image[6] == \"N\":\n # if they did not survive\n if image[7] == \"Y\":\n # if they were intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(10)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(11)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(12)\n elif image[7] == \"N\":\n # if they were not intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(13)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(14)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(15)\n else:\n # if they were intubated is unknown\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(16)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(17)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(18)\n else:\n # if their survival is unknown\n if image[7] == \"Y\":\n # if they were intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(19)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(20)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(21)\n elif image[7] == \"N\":\n # if they were not intubated\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(22)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(23)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(23)\n else:\n # if they were intubated is unknown\n if image[8] == \"Y\":\n # if they were put in the icu\n d['filename'].append(image[23])\n d['target'].append(24)\n elif image[8] == \"N\":\n # if they were not put in the icu\n d['filename'].append(image[23])\n d['target'].append(25)\n else:\n # if they were put in the icu is unknown\n d['filename'].append(image[23])\n d['target'].append(26)\n\n \n# add normal chest xrays to data\ndir_path = os.path.abspath(os.getcwd())\ndir_path = dir_path+\"/chest_xray/test/Normal\"\nlist = os.listdir(dir_path)\nfor pic in list:\n d['filename'].append(pic)\n d['target'].append(0)\ntotal = [0] * 28\n\nfor target in d['target']:\n if target == -1:\n total[27] += 1\n else:\n total[target] += 1\n\n# set up binary classification (covid vs non-covid)\n# all targets > 0 are covid\nfor i in range(len(d['target'])):\n if d['target'][i] > 0:\n d['target'][i] = 1\n\nprint(total)\n#print(d)\n\nfile.close()\n\n# -\n\nlabels = {0: \"Non COVID\", 1: \"COVID\"}\nget_distr(d['target'], labels)\n\nprint(len(d['target']))\nprint(d['target'].count(1))\nprint(d['target'].count(0))\n\n# # Part 1: Preprocessing for images\n# Load in images for the corresponding target data with PIL. Reformat all images to the same dimensions and convert them to a numpy array for the models to use (where each is a (# pixels, # pixels) array).\n\n# +\n# Load in image data from filenames with PIL\nimg_dim = 128\nimages = np.empty((len(d['target']), img_dim, img_dim, 3))\n\nfor i in range(len(d['filename'])):\n image = Image.open(\"model_images/\" + d['filename'][i])\n \n # resize image shape\n image = image.resize((img_dim, img_dim))\n \n # convert to grayscale\n image = image.convert('RGB')\n \n images[i] = np.array(image)\n\n# normalize values\nimages = images / 255.0\n# -\n\ncallbacks = [\n tf.keras.callbacks.EarlyStopping(\n monitor=\"val_loss\",\n min_delta=0.01,\n patience=2,\n verbose=1\n )\n] \n\n# # Part 2: Deep Learning Classifier with Keras\n\n# Split the data into training and testing with sklearn\n\n# split data\nx_train, x_test, y_train, y_test = train_test_split(images, d['target'], test_size=0.2, random_state=0)\ny_train = np.asarray(y_train)\ny_test = np.asarray(y_test)\n\n# Build and compile the deep learning model\n\n# +\n# build model\ndl_model = tf.keras.Sequential([\n tf.keras.layers.Flatten(input_shape=(img_dim,img_dim,2)),\n tf.keras.layers.Dense(512, activation='relu'),\n tf.keras.layers.Dense(1, activation='sigmoid')\n])\n\n# compile model\ndl_model.compile(optimizer='adam',\n loss=tf.keras.losses.BinaryCrossentropy(),\n metrics=['accuracy'])\n# -\n\ndl_model.summary()\n\n# Train the model\n\nhistory = dl_model.fit(x_train, y_train, epochs=10,\n validation_data=(x_test, y_test),\n callbacks=callbacks,\n batch_size=16)\n\n# Evaluate the model with testing data\n\n# +\nplt.plot(history.history['accuracy'], label='accuracy')\nplt.plot(history.history['val_accuracy'], label='val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title('Deep Learning Classifier')\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\ntest_loss, test_acc = dl_model.evaluate(x_test, y_test, verbose=2)\nprint(\"Test Accuracy:\", round(test_acc, 4))\n# -\n\ndly_pred = dl_model.predict(x_test)\n\ndly_pred_norm = convert_pred(dly_pred, 0.5)\n\nprint(classification_report(y_test, dly_pred_norm))\n\nplot_cm(y_test, dly_pred_norm, \"Not COVID(0)/COVID(1) - MLP\")\n\nplot_ROC(y_test, dly_pred, \"Not COVID/COVID - MLP\")\n\n# # Part 3: Classifier with a Convolution Neural Network\n\n# Build and compile the model\n\n# +\n# build model for convolution network\ncnn_model = models.Sequential()\ncnn_model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(img_dim, img_dim, 2), padding='same'))\ncnn_model.add(layers.MaxPooling2D((2, 2)))\ncnn_model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))\ncnn_model.add(layers.MaxPooling2D((2, 2)))\ncnn_model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))\n\n# add more layers\ncnn_model.add(layers.Flatten())\ncnn_model.add(layers.Dense(64, activation='relu'))\ncnn_model.add(layers.Dense(1, activation='sigmoid'))\n\n# compile \ncnn_model.compile(optimizer='adam',\n loss=tf.keras.losses.BinaryCrossentropy(),\n metrics=['accuracy'])\n# -\n\ncnn_model.summary()\n\n# Train the model\n\nhistory = cnn_model.fit(x_train, y_train, epochs=10, \n validation_data=(x_test, y_test),\n callbacks=callbacks,\n batch_size=16)\n\n# Evaluate the model with testing data and plot results\n\n# +\nplt.plot(history.history['accuracy'], label='accuracy')\nplt.plot(history.history['val_accuracy'], label = 'val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title('CNN Classifier')\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\ntest_loss, test_acc = cnn_model.evaluate(x_test, y_test, verbose=2)\nprint(\"Test Accuracy:\", round(test_acc, 4))\n# -\n\ncnny_pred = cnn_model.predict(x_test)\n\ncnny_pred_norm = convert_pred(cnny_pred, 0.5)\n\nprint(classification_report(y_test, cnny_pred_norm))\n\nplot_cm(y_test, cnny_pred_norm, \"Not COVID(0)/COVID(1) - CNN\")\n\nplot_ROC(y_test, cnny_pred, \"Not COVID/COVID - CNN\")\n\n# Compare models\nfpr_dl, tpr_dl, thresholds = roc_curve(y_test, dly_pred)\nfpr_cnn, tpr_cnn, thresholds = roc_curve(y_test, cnny_pred)\nauc_valdl = auc(fpr_dl, tpr_dl)\nauc_valcnn = auc(fpr_cnn, tpr_cnn)\nplt.plot(fpr_dl,tpr_dl) \nplt.plot(fpr_cnn, tpr_cnn)\nplt.plot([0, 1], [0, 1], linestyle='dashed')\nplt.axis([-0.05, 1.05, -0.05, 1.05])\nplt.xlabel('False Positive Rate') \nplt.ylabel('True Positive Rate')\nplt.title(\"ROC Curve\\nMLP AUC: \" + str(round(auc_valdl, 4)) + \"\\nCNN AUC: \" + str(round(auc_valcnn, 4)))\nplt.legend([\"MLP\", \"CNN\"])\nplt.show()\n\n# # Part 4: Optimization\n# Based on the results of the above models, we will now attempt to optimize our algorithms. We will be using cross validation to find a find the best split between our training/testing data.\n\n# +\n# split data into 10 folds\nfolds = 10\nkfold = KFold(n_splits=folds)\n\n# shuffle initial data\nrimages, rtargets = shuffle(images, d['target'], random_state=0)\n# -\n\n# Repeat Deep Learning and CNN models above with KFold technique and compare results\n\n# ### Deep Learning - Cross Validation\n\n# +\nresults = []\ntest_accuracies = []\n\nfor train, test in kfold.split(rimages):\n x_train, x_test = rimages[train], rimages[test]\n y_train, y_test = np.array(rtargets)[train], np.array(rtargets)[test]\n \n # build model\n dl_model = tf.keras.Sequential([\n tf.keras.layers.Flatten(input_shape=(img_dim,img_dim,2)),\n tf.keras.layers.Dense(512, activation='relu'),\n tf.keras.layers.Dense(2)\n ])\n\n # compile model\n dl_model.compile(optimizer='adam',\n loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),\n metrics=['accuracy'])\n \n # train model\n history = dl_model.fit(x_train, y_train, epochs=10,\n validation_data=(x_test, y_test))\n \n test_loss, test_acc = model.evaluate(x_test, y_test, verbose=2)\n \n results.append(history)\n test_accuracies.append(test_acc)\n# -\n\n# Get best model and plot results\n\n# choose model with highest test accuracy\nbest_model = results[0]\nbest_acc = test_accuracies[0]\nfor i in range(len(results) - 1):\n if test_accuracies[i + 1] > test_accuracies[i]:\n best_model = results[i+1]\n\n# +\nplt.plot(best_model.history['accuracy'], label='accuracy')\nplt.plot(best_model.history['val_accuracy'], label = 'val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title('Deep Learning Classifier - CV')\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\nprint(\"Test Accuracy:\", round(best_acc, 4))\n# -\n\n# ### CNN - Cross Validation\n\n# +\nresults = []\ntest_accuracies = []\n\nfor train, test in kfold.split(rimages):\n x_train, x_test = rimages[train], rimages[test]\n y_train, y_test = np.array(rtargets)[train], np.array(rtargets)[test]\n \n # build model for convolusion network\n cnn_model = models.Sequential()\n cnn_model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(img_dim, img_dim, 2)))\n cnn_model.add(layers.MaxPooling2D((2, 2)))\n cnn_model.add(layers.Conv2D(64, (3, 3), activation='relu'))\n cnn_model.add(layers.MaxPooling2D((2, 2)))\n cnn_model.add(layers.Conv2D(64, (3, 3), activation='relu'))\n\n # add more layers\n cnn_model.add(layers.Flatten())\n cnn_model.add(layers.Dense(64, activation='relu'))\n cnn_model.add(layers.Dense(2))\n\n # compile \n cnn_model.compile(optimizer='adam',\n loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),\n metrics=['accuracy'])\n \n history = cnn_model.fit(x_train, y_train, epochs=10,\n validation_data=(x_test, y_test))\n \n results.append(history)\n \n \n test_accuracies.append(test_acc)\n# -\n\n# Get best model and plot results\n\n# choose model with highest test accuracy\nbest_model = results[0]\nbest_acc = test_accuracies[0]\nfor i in range(len(results) - 1):\n if test_accuracies[i + 1] > test_accuracies[i]:\n best_model = results[i+1]\n\n# +\nplt.plot(best_model.history['accuracy'], label='accuracy')\nplt.plot(best_model.history['val_accuracy'], label = 'val_accuracy')\nplt.xlabel('Epoch')\nplt.ylabel('Accuracy')\nplt.title(\"CNN Classifier - CV\")\nplt.ylim([0.5, 1])\nplt.legend(loc='lower right')\n\nprint(\"Test Accuracy:\", round(best_acc, 4))\n# -\n\n\n","repo_name":"gdosulli/CS254_COVID","sub_path":"GOBT_CS254_InitialModels.ipynb","file_name":"GOBT_CS254_InitialModels.ipynb","file_ext":"py","file_size_in_byte":18545,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"30443140186","text":"# # Importing libraries and Exploring the Dataset\n\nimport numpy as np \nimport pandas as pd \nfrom matplotlib import pyplot as plt\nimport seaborn as sns\nimport scipy.stats as stats\nsns.set() \n\n# Read the Dataset\ninsurance_df = pd.read_csv(\"insurance-data.csv\")\n\n# +\n#Display the first five dataset\n\ninsurance_df.head()\n\n# +\n#Info about the data shape of data, type of individual columns\n\ninsurance_df.info()\n# -\n\n# # Checking missing values and Outliers\n\n# +\n# Check for the null values \n\ninsurance_df.isna().apply(pd.value_counts)\n# -\n\n# Checking for the outliers\nfig, ax = plt.subplots(3, 1, figsize=(10,7))\nax[0].set_title(\"BMI\")\nax[1].set_title(\"AGE\")\nax[2].set_title(\"CHARGES\")\nsns.boxplot(data = insurance_df.bmi, orient=\"h\", ax=ax[0], color = \"b\")\nsns.boxplot(data = insurance_df.age, orient=\"h\", ax=ax[1], color = \"r\")\nsns.boxplot(data = insurance_df.charges, orient=\"h\", ax=ax[2], color = \"g\")\nplt.tight_layout()\n\n# +\n# a brief summary for the dataset\n\ninsurance_df.describe().T\n# -\n\n# # Creating visual methods to analyze the data.\n\n# +\n# Plots to see the distribution of the continuous features individually\n\nplt.figure(figsize=(20,15))\nplt.subplot(3,3,1)\nplt.hist(insurance_df.bmi, color=\"lightblue\", edgecolor=\"black\", alpha=0.7)\nplt.xlabel(\"bmi\")\n\nplt.subplot(3,3,2)\nplt.hist(insurance_df.age, color='red', edgecolor = 'black', alpha = 0.7)\nplt.xlabel('age')\n\nplt.subplot(3,3,3)\nplt.hist(insurance_df.charges, color='lightgreen', edgecolor = 'black', alpha = 0.7)\nplt.xlabel('charges')\n\nplt.show()\n# -\n\n# Fatih hocamızın gönderdiği kod:\nfig, ax = plt.subplots(1, 3, figsize=(15,4))\nsns.distplot(insurance_df.bmi, ax=ax[0], color = \"b\", bins=10, kde=False)\nsns.distplot(insurance_df.age, ax=ax[1], color = \"r\", bins=10, kde=False)\nsns.distplot(insurance_df.charges, ax=ax[2], color = \"g\", bins=10, kde=False)\n\n# +\nplt.figure(figsize=(17,20))\nx = insurance_df.smoker.value_counts().index\ny = [insurance_df['smoker'].value_counts()[i] for i in x]\nplt.subplot(4,2,1)\nplt.bar(x,y, align='center',color = 'lightblue',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('Is Smoker?')\nplt.ylabel('Count ')\nplt.title('Smoker distribution')\n\nx1 = insurance_df.sex.value_counts().index #Values for x-axis\ny1 = [insurance_df['sex'].value_counts()[j] for j in x1] # Count of each class on y-axis\n\nplt.subplot(4,2,2)\nplt.bar(x1,y1, align='center',color = 'red',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('Gender')\nplt.ylabel('Count')\nplt.title('Gender distribution')\n\nx2 = insurance_df.region.value_counts().index #Values for x-axis\ny2 = [insurance_df['region'].value_counts()[k] for k in x2] # Count of each class on y-axis\n\nplt.subplot(4,2,3)\nplt.bar(x2,y2, align='center',color = 'green',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('Region')\nplt.ylabel('Count ')\nplt.title(\"Regions' distribution\")\n\nx3 = insurance_df.children.value_counts().index #Values for x-axis\ny3 = [insurance_df['children'].value_counts()[l] for l in x3] # Count of each class on y-axis\n\nplt.subplot(4,2,4)\nplt.bar(x3,y3, align='center',color = 'purple',edgecolor = 'black',alpha = 0.7) #plot a bar chart\nplt.xlabel('No. of children')\nplt.ylabel('Count ')\nplt.title(\"Children distribution\")\nplt.tight_layout()\nplt.show()\n# -\n\nsns.countplot(insurance_df.region)\n\nsns.pairplot(insurance_df)\n\n# # Analyzing trends, patterns, and relationships in the Data.\n\n# Do charges of people who smoke differ significantly from the people who don't?\n\n\nsns.heatmap(insurance_df.corr(), cmap = 'coolwarm', annot = True)\n\n# Scatter plot to look for visual evidence of dependency between attributes smoker and charges accross different ages\nplt.figure(figsize=(12,7))\nsns.scatterplot(insurance_df.age, insurance_df.charges, hue=insurance_df.smoker, palette=[\"red\", \"green\"], alpha=0.7)\nplt.show()\n\nplt.figure(figsize=(12,7))\nsns.scatterplot(insurance_df.age, insurance_df.charges, hue=insurance_df.sex, palette=[\"red\", \"blue\"], alpha=0.7)\nplt.show()\n\ninsurance_df.sex.value_counts()\n\n# +\n# T-test to check dependency of smoking on charges\nH0=\"Charges of smoker and non-smoker are the same\"\nHa=\"Charges of smoker and non-smoker are not the same\"\n\na=np.array(insurance_df[insurance_df.smoker==\"yes\"].charges)\nb=np.array(insurance_df[insurance_df.smoker==\"no\"].charges)\n\nt, p_value=stats.ttest_ind(a,b, axis=0)\n\nif p_value < 0.05: # Setting our significance level at 5%\n print(f'{Ha} as the p_value ({p_value}) < 0.05')\nelse:\n print(f'{Ho} as the p_value ({p_value}) > 0.05')\n\n\n# +\n#Does bmi of males differ significantly from that of females?\nH0=\"gender has no effect on bmi\"\nHa=\"gender has effect on bmi\"\n\na=np.array(insurance_df[insurance_df.sex==\"male\"].bmi)\nb=np.array(insurance_df[insurance_df.sex==\"female\"].bmi)\n\nt, p_value=stats.ttest_ind(a,b, axis=0)\n\nif p_value < 0.05: # Setting our significance level at 5%\n print(f'{Ha} as the p_value ({p_value}) < 0.05')\nelse:\n print(f'{H0} as the p_value ({p_value}) > 0.05')\n# -\n\n\n\n\n# Hasan hocamızın gönrderdiği kod:\nttest2=stats.ttest_ind(insurance_df.bmi[insurance_df[\"sex\"]==\"male\"],insurance_df.bmi[insurance_df[\"sex\"]==\"female\"])\nif ttest2[1]<0.05:\n print(f\"Gender has effect on bmi as the p_value {ttest2[1]} < 0.05\")\nelse:\n print(f\"Gender has no effect on bmi as the p_value {round(ttest2[1],2)} > 0.05\")\n\n\n","repo_name":"hersany/DataScience","sub_path":"EDA/EDA of Insurance Company's Dataset.ipynb","file_name":"EDA of Insurance Company's Dataset.ipynb","file_ext":"py","file_size_in_byte":5312,"program_lang":"python","lang":"en","doc_type":"code","stars":6,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"12715736122","text":"# * Matplotlib \n# Matplotlib是一个最常用的2D绘图包,可以完成绝大部分的绘图任务,当然也可以完成部分简单的3D绘图任务,效果并不如其他的库那么理想\n\n# 获取数据\n# %matplotlib inline\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\nplt.plot(x, c, x, s)\n\n# +\nplt.figure(figsize=(8, 6), dpi=80) # (8, 6) 英寸,每英寸2,54cm, 分辨率是80\nplt.subplot(1, 1, 1) # 传建一个子图 1 * 1,占据第1个网格\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\n# color 设置颜色,设置线宽,设置线的样式,alpha还可以设置我们的透明度\nplt.plot(x, c, color='blue', linewidth=1.0, linestyle='-')\nplt.plot(x, s, color='green', linewidth=1.0, linestyle='-')\n# 设置x轴的范围\nplt.xlim(-4.0, 4.0)\n# 设置轴的说明\nplt.xlabel('这是x')\n# 设置坐标轴的刻度,支持使用Latex标注\nplt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi],[r'$-\\pi$', r'$-\\pi/2$', r'$0$', r'$+\\pi/2$', r'$+\\pi$'])\nplt.ylim(-1.0, 1.0)\nplt.ylabel('这是y')\nplt.yticks(np.linspace(-1, 1, 5, endpoint=True))\n\nax = plt.gca()\n\n# 为了可以让我们的刻度显示出来不被线条遮盖\nfor label in ax.get_xticklabels() + ax.get_yticklabels():\n # 将刻度的字体大小调整\n label.set_fontsize(16)\n # 字体的矩形显示框\n label.set_bbox(dict(facecolor='blue', edgecolor='None', alpha=0.65))\n\n\nplt.show()\n\n# plt.savefig('filename.png', dpi=80) # 保存分辨率是80的图图片到对应的路径上\n# -\n\n# spines的移动\nax = plt.gca() # 获取当前的坐标轴\n# 单个运行下面的语句,将右边的边界置空\nax.spines['right'].set_color('none') # right, top, left, bottom\nax.spines['top'].set_color('none')\nax.xaxis.set_ticks_position('bottom') # 将xaxis, x坐标轴设定成底边的bottom spines\nax.yaxis.set_ticks_position('left')\nax.spines['bottom'].set_position(('data', 0.5)) # 将位置设定在y轴的0.5上\nax.spines['left'].set_position(('data', 0.5)) # 将位置设定在x轴的0.5刻度上\nplt.show()\n\n# 添加标签\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\nplt.plot(x, c, color='blue', linewidth=2.5, linestyle='-', label='cosline') # label属性制定之后的legend的内容\nplt.plot(x, s, color='red', linewidth=2.5, linestyle='-', label='sine')\nplt.legend(loc='upper left') # 将lengend图标显示在左上角,不加的话,默认将其添加在最合适的位置上\n\n# +\n# 注释 annotate\nax = plt.gca() # 获取当前的坐标轴\n# 单个运行下面的语句,将右边的边界置空\nax.spines['right'].set_color('none') # right, top, left, bottom\nax.spines['top'].set_color('none')\nax.xaxis.set_ticks_position('bottom') # 将xaxis, x坐标轴设定成底边的bottom spines\nax.yaxis.set_ticks_position('left')\nax.spines['bottom'].set_position(('data', 0)) # 将位置设定在y轴的0.5上\nax.spines['left'].set_position(('data', 0)) # 将位置设定在x轴的0.5刻度上\n\n# 为了可以让我们的刻度显示出来不被线条遮盖\nfor label in ax.get_xticklabels() + ax.get_yticklabels():\n # 将刻度的字体大小调整\n label.set_fontsize(16)\n # 字体的矩形显示框\n label.set_bbox(dict(facecolor='white', edgecolor='None', alpha=0.65))\n\n# 注释的代码\nx = np.linspace(-np.pi, np.pi, 256, endpoint=True)\nc, s = np.cos(x), np.sin(x)\nplt.plot(x, c, color='blue', linewidth=10, linestyle='-', label='cosline', alpha=0.5) # label属性制定之后的legend的内容\nplt.plot(x, s, color='red', linewidth=2.5, linestyle='-', label='sine')\nplt.legend(loc='upper left') # 将lengend图标显示在左上角,不加的话,默认将其添加在最合适的位置上\n\nt = 2 * np.pi / 3\nplt.plot([t, t], [0, np.cos(t)], color='blue', linewidth=2.5, linestyle='--') # 显示直线(t,0)->(t,cos(t))\nplt.scatter([t, ], [np.cos(t), ], 50, color='blue') # 将(t,np.cos(t))处打上点,显示\n\n# 第一个参数是注释的文本Latex嵌入,注释的目标点是(t, cos(t)), 注释的文本坐标是(-90, -100), 距离使用offset point相对与终点的\n# arrowprops箭头的属性 , arrowstyle是箭头的格式,还有连接的方式\nplt.annotate(r'$cos(\\frac{2\\pi}{3})=-\\frac{1}{2}$', \\\n xy=(t, np.cos(t)), xycoords='data', xytext=(-90, -100), \\\n textcoords='offset points', fontsize=16, arrowprops=dict(arrowstyle='->', connectionstyle='arc3,rad=.2'))\n\nt = 2 * np.pi / 3\nplt.plot([t, t], [0, np.sin(t)], color='blue', linewidth=2.5, linestyle='--') # 显示直线(t,0)->(t,cos(t))\nplt.scatter([t, ], [np.sin(t), ], 50, color='blue') # 将(t,np.cos(t))处打上点,显示\n\nplt.annotate(r'$sin(\\frac{2\\pi}{3})=-\\frac{\\sqrt{3}}{2}$', \\\n xy=(t, np.sin(t)), xycoords='data', xytext=(-90, -100), \\\n textcoords='offset points', fontsize=16, arrowprops=dict(arrowstyle='->', connectionstyle='arc3,rad=.2'))\n# -\n\n# * 数字,子图,轴的技巧使用\n# * figure : 是一个窗口\n# * subplots : 是一个窗口的子图,默认使用figure创建的是subplots(111)\n\n# %matplotlib inline\n# num是图的编号,不指定默认自增,figsize是图片的边框大小,facecolor是前景色,edgecolor是后景色,frameon是是否绘制图框\nplt.figure(num=1, figsize=(80, 80), dpi=100, facecolor='green', edgecolor='blue', frameon=True)\nplt.show()\nplt.close()\n\n# 子图命令\n# 使用子图可以在常规网格中排雷图,并按照指示制定行数和列数以及绘制图\nplt.subplot(2, 1, 1)\nplt.subplot(2, 1, 2)\n\n# Axes 很类似子图但是语序将图放置在图的任何位置\nplt.axes([0, 0, 0.5, 0.5]) # x, y, xs, ys (x, y)坐标,xs, ys是长宽\nplt.axes([0.1, 0.1, 0.5, 0.5])\nplt.axes([0.2, 0.2, 0.5, 0.5])\n\n# +\n# 颜色填充\nn = 256\nX = np.linspace(-np.pi, np.pi, n, endpoint=True)\nY= np.sin(2 * X)\n\nplt.axes([0.025, 0.025, 0.95, 0.95])\n\nplt.plot(X, Y + 1, color='blue', alpha=1.00)\nplt.fill_between(X, 1, Y + 1, color='blue', alpha=0.25) # 1 - Y + 1之间使用蓝色填充\n\nplt.plot(X, Y - 1, color='blue', alpha=1.00)\nplt.fill_between(X, -1, Y - 1, (Y - 1) > -1, color='blue', alpha=0.25) # where数组选择性填充\nplt.fill_between(X, -1, Y - 1, (Y - 1) < -1, color='red', alpha=0.25)\n\n# +\n# 散点图\nn = 1024\nX = np.random.normal(0, 1, n) # 高斯分布,均值,标准差,规模\nY = np.random.normal(0, 1, n)\nT = np.arctan2(Y, X)\n\nplt.axes([0.25, 2.5, 0.95, 0.95])\nplt.scatter(X, Y, s=75, c=T, alpha=0.5) # size大小,c颜色,alpha是透明度\n\n# +\n# 条形图\nn = 12\nx = np.arange(n)\ny1 = (1 - x / float(n)) * np.random.uniform(0.5, 1.0, n) # 均匀分布中随机采样,[low, high),size是规模\ny2 = (1 - x / float(n)) * np.random.uniform(0.5, 1.0, n)\n\nplt.bar(x, +y1, facecolor='#9999ff', edgecolor='white') # 条形图\nplt.bar(x, -y2, facecolor='#ff9999', edgecolor='white')\n\nfor x1, y in zip(x, y1):\n plt.text(x1, y + 0.05, '%.2f' % y) # x,y是文本注释的坐标,第三个参数显示的内容,alpha是透明度,family设置字体\nfor x1, y in zip(x, y2):\n plt.text(x1 + 0.4, - y - 0.05, '%.2f' % y, ha='center', va='top')\n\n\n# +\n# 等高线\n# np.meshgrid函数用来用两个一维数组(x, y)生成两个二维网格(X, Y)\n# 其中X是x的简单复制,Y是y的简单复制,两个矩阵的规模都是n * m(x - m, y - n)\ndef f(x, y):\n return (1 - x / 2 + x ** 5 + y ** 3) * np.exp(- x ** 2 - y ** 2)\n\nn = 256\nx = np.linspace(-3, 3, n)\ny = np.linspace(-3, 3, n)\nX, Y = np.meshgrid(x, y)\nZ = f(X, Y)\n\nplt.axes([0.025, 0.025, 0.95, 0.95])\nplt.imshow(Z, interpolation='nearest', cmap='bone', origin='lower')\nplt.colorbar(shrink=0.92)\n\n# +\n# 饼图\nn = 20\nZ = np.ones(n)\nplt.axes([0.025, 0.025, 0.95, 0.95])\n\nplt.pie(Z, explode=Z*0.05, colors=['%f' % (i / float(n)) for i in range(n)])\nplt.axis('equal') # 刻度对齐,保证是标准圆形\nplt.xticks()\nplt.yticks()\n\nplt.show()\n","repo_name":"gmftbyGMFTBY/Study","sub_path":"Python/AllForPython/Python-Scipy/.ipynb_checkpoints/Matplotlib-checkpoint.ipynb","file_name":"Matplotlib-checkpoint.ipynb","file_ext":"py","file_size_in_byte":7933,"program_lang":"python","lang":"en","doc_type":"code","stars":10,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"35935036017","text":"# # Author: Yoonhyuck WOO / JBNU_Industrial Information system Engineering\n# # Date; 3. 7. 2022 - 3. 17. 2022\n# # Title: Korean_NER\n# # Professor: Seung-Hoon Na\n\nimport os\nimport json\nimport numpy as np\nfrom functools import partial\nimport torch\nimport torch.nn as nn\nfrom torch.utils.data import DataLoader\nfrom torch.optim import AdamW\nfrom transformers import BertTokenizer, BertModel, FlaubertModel\n# from transformers import BertModel\nfrom sklearn.metrics import classification_report\n\nfrom DataPreprocessing import Data_NER, ner_collate_fn\nfrom modeling_ner import Bert_NER\n\nfrom ner_collate_fn import collate_fn_custom\n\nPRETAINED_MODEL_NAME = 'bert-base-multilingual-cased'\ntokenizer = BertTokenizer.from_pretrained(PRETAINED_MODEL_NAME)\nbert = BertModel.from_pretrained(PRETAINED_MODEL_NAME)\n# flaubert = FlaubertModel.from_pretrained(language_model_dir, output_loading_info=True)\n\n# # BertTokenizer\n# - return tensors = \"pt\" : Finally, you want the tokenizer to return the actual tensors that are fed to the model.\n# \n# # BertModel\n# - Attention_mask : 1 where you care and 0 where you don't care.\n# - Input_ids : the IDs of the sentence morpheme.\n# - Token_type_ids : for the question problem, but it's enough to set it to zero now.\n#\n# reference: https://huggingface.co/docs/transformers/model_doc/bert\n#\n# # Example\n\n# + active=\"\"\n# tokenizer = BertTokenizer.from_pretrained(\"bert-base-uncased\")\n# model = BertModel.from_pretrained(\"bert-base-uncased\")\n#\n# inputs = tokenizer(\"Hello, my dog is cute\", return_tensors=\"pt\")\n# outputs = model(**inputs)\n#\n# last_hidden_states = outputs.last_hidden_state\n#\n# print(inputs)\n#\n# => {'input_ids': tensor([[ 101, 7592, 1010, 2026, 3899, 2003, 10140, 102]]), 'token_type_ids': tensor([[0, 0, 0, 0, 0, 0, 0, 0]]), 'attention_mask': tensor([[1, 1, 1, 1, 1, 1, 1, 1]])}\n# -\n\n# # Loading data\n\nPATH_dir = 'C:\\\\Users\\\\LG\\\\Desktop\\\\github\\\\JBNU-2022-SPRING\\\\English world class tagging & Korean_Named Entity Recognition\\\\Ko_En_NER_POStag_data\\Ko_NER_POS'\nPATH_ko_train = os.path.join(PATH_dir, 'prepro_train.json')\nPATH_ko_test = os.path.join(PATH_dir, 'prepro_test.json')\nPATH_ko_dev = os.path.join(PATH_dir, 'prepro_dev.json')\ntotal_tag = os.path.join(PATH_dir, 'total_tag.json')\n\nwith open(PATH_ko_train) as f: \n dataset_train = json.load(f)\nwith open(PATH_ko_test) as f: \n dataset_test = json.load(f)\nwith open(PATH_ko_dev) as f: \n dataset_dev = json.load(f)\nwith open(total_tag, 'r') as f:\n tag_converter = json.load(f)\n\nprint('train', len(dataset_train))\nprint('test', len(dataset_test))\nprint('dev', len(dataset_dev))\n# tag_converter.id_to_tag\n\n# # collate_fn\n# - Tie it in batches.\n# - If the data set is variable length, it cannot be tied right away and causes an error, so you have to make a 'collate_fn' and hand it over.\n# - partial_collate_fn: when collating, with padding\n# \n# \n# - reference: https://hulk89.github.io/pytorch/2019/09/30/pytorch_dataset/\n\n# # collate_fn_custom\n\n# + jupyter={\"source_hidden\": true}\ndef collate_fn_custom(data):\n \n input_sent = [sample[0] for sample in data]\n labels = [sample[2] for sample in data]\n \n padded_inputs = torch.nn.utils.rnn.pad_sequence(input_sent, batch_first = True)\n \n return {'input': padded_inputs.contiguous(),\n 'label': torch.stack(labels).contiguous()}\n\n\n# -\n\n# # 0309\n\nfrom ner_collate_fn import make_same_len\n\ntag_to_id ={'PAD': 0, 'B-<휠': 1, 'B-OG': 2, 'I-조선': 3, 'I-PS': 4, 'B-LC': 5, 'B-1': 6, 'I-<휠': 7, 'I-LC': 8, 'B-PS': 9, 'I-TI': 10, \n 'B-목소': 11, 'O': 12, 'I-목소': 13, 'I-1': 14, 'I-': 15, 'B-조선': 16, 'I-OG': 17, 'B-': 18, 'I-DT': 19, 'B-TI': 20, 'B-DT': 21}\n\n\n# + active=\"\"\n# convert_to = []\n# def make_same_len(batch):\n# each_len_list = [len(sample) for sample in batch]\n# max_len = max(each_len_list)\n# \n# padded_batch = []\n# pad_id = 0\n# special_token = 'PAD'\n#\n# for sample in batch:\n# padded_batch.append(['CLS'] + sample+ ['PAD'] * (max_len - len(sample)) + ['SEP'])\n# \n# return padded_batch\n#\n# -\n\ndef tag_to_id2(batch):\n total = []\n convert_to = []\n for sample in batch:\n for j in sample:\n j = tag_to_id.get(j)\n convert_to.append(j)\n total.append(convert_to)\n convert_to = []\n \n return total\n\n\n\n# + active=\"\"\n# def make_same_len(batch):\n# each_len_list = [len(sample) for sample in batch]\n# max_len = max(each_len_list)\n# total = []\n# convert_to = []\n# print(max_len)\n# padded_batch = []\n# pad_id = 0\n# special_token = 'PAD'\n#\n# for sample in batch:\n# for j in sample:\n# j = tag_to_id.get(j)\n# convert_to.append(j)\n# total.append(convert_to)\n# convert_to = []\n# sample = total\n# print(len(total))\n# padded_batch.append(['PAD'] + sample + ['PAD'] * (max_len - 2) + ['PAD'])\n# total = []\n# return padded_batch\n#\n# -\n\na = [['PAD'],['B-OG','I-조선'],['I-PS','PAD','I-OG','O']]\ntag_to_id2(a)\n\nprint(make_same_len(a))\ntag_to_id2(make_same_len(a))\n\n\n\nconvert_to = []\ndef tag_to_id(data):\n for j in data:\n j = tag_to_id.get(j)\n convert_to.append(j)\n convert_to = torch.tensor(convert_to)\n \n \n return convert_to\n\n\ndef make_same_len(batch):\n each_len_list = [len(sample) for sample in batch]\n max_len = max(each_len_list)\n \n padded_batch = []\n pad_id = 0\n special_token = 'PAD'\n\n for sample in batch:\n padded_batch.append(['PAD'] + sample + ['PAD'] * (max_len - len(sample)) + ['PAD'])\n \n return padded_batch\n\n\n\nb = [[1],[2,2],[3,3,3],[4,4,4,4],[5,5,5,5,5],[6,6,6,6,6,6]]\nmake_same_len(b)\n\n# +\nconvert_to = []\ndef make_same_len(batch):\n each_len_list = [len(sample) for sample in batch]\n max_len = max(each_len_list)\n \n padded_batch = []\n pad_id = 0\n special_token = 'PAD'\n\n for sample in batch:\n padded_batch.append(['CLS'] + sample+ ['PAD'] * (max_len - len(sample)) + ['SEP'])\n \n for tag in padded_batch:\n print(tag)\n for j in tag:\n j = tag_to_id.get(j)\n convert_to.append(j)\n \n padded_batch = convert_to\n# print('before',padded_batch)\n# for tag in padded_batch:\n# print(tag)\n# print('='*90)\n# for j in tag:\n# # print(j)\n# j = tag_to_id.get(j)\n# convert_to.append(j)\n \n# padded_batch = convert_to\n# # conver_to = []\n# print('aftre')\n return padded_batch\n\n\n\n# -\n\na = [['PAD'],['B-OG','I-조선'],[2,2,2]]\nmake_same_len(a)\n\n\ndef ner_fn_custom(tokenizer, data):\n \n input_sent = [sample[0] for sample in data]\n labels = [sample[2] for sample in data]\n\n batch_inputs = tokenizer(input_sent, padding = True, return_tensors = \"pt\")\n batch_labels = make_same_len(labels)\n batch_labels = tag_to_id2(batch_labels)\n batch_labels = torch.tensor(batch_labels)\n \n return batch_inputs, batch_labels\n# return {'input': padded_inputs.contiguous(),\n# 'label': torch.stack(labels).contiguous()}\n\n\ndef ner_fn_custom(tokenizer, data):\n \n input_sent = [sample[0] for sample in data]\n labels = [sample[2] for sample in data]\n\n batch_inputs = tokenizer(input_sent, padding = True, return_tensors = \"pt\")\n batch_labels = make_same_len(labels)\n \n \n return batch_inputs, batch_labels\n# return {'input': padded_inputs.contiguous(),\n# 'label': torch.stack(labels).contiguous()}\n\nbatch_size = 10\n\n# +\nbatch_size = 1\n# partial (func, /, *args, **keywords) positional argument: args, keyword argument: keywords\n\n# partial_collate_fn = partial(collate_fn_custom, tokenizer, tag_converter)\n# partial_collate_fn = collate_fn_custom\n\npartial_collate_fn = partial(ner_fn_custom, tokenizer)\n\n'''\npartial_collate_fn = partial(collate_fn_custom, tokenizer, tag_converter)\n'''\n# ner_collate_fn: padding & making batch?\n'''\ndef ner_collate_fn(tokenizer, tag_converter):\n \n'''\n\n# +\ndataloader_train = DataLoader(\n dataset_train,\n batch_size = batch_size,\n shuffle = True,\n collate_fn = partial_collate_fn)\n\ndataloader_test = DataLoader(\n dataset_test,\n batch_size = batch_size,\n shuffle = False,\n collate_fn = partial_collate_fn)\n\ndataloader_dev = DataLoader(\n dataset_dev,\n batch_size = batch_size,\n shuffle = False,\n collate_fn = partial_collate_fn)\n\n\n# -\n\n# # BERT_NER\n# - Dropout: method to solve overfitting, one of the problems in deep learning learning. In summary, how to prevent some units of the hidden layer from being overfitted by inoperative
\n# ->the outputs are scaled by a factor of (1 / 1-p) during training\n\n# # *args, **kargs\n# - *args(non-keworded arguments): Tuple type of argument without limitation.\n# - **kargs(keworded arguments): Dictionary type of argument without limitation.\n\nclass Bert_NER(nn.Module):\n \n def __init__(self, bert):\n super().__init__() \n self.bert = bert\n self.dorpout = nn.Dropout(p = 0.1)\n self.linear = nn.Linear(768, 22)\n self.softmax = nn.Softmax(dim = 2) # A dimension along which Softmax will be computed\n \n \n def forward(self, **kwargs):\n emb = self.bert(**kwargs)\n e = self.dropout(emb['last_hidden_state'])\n w = self.linear(e)\n \n return w\n\n\n\nlen(tag_to_id)\n\ntag_num = len(tag_to_id) # 22\nmodel = Bert_NER(22)\n\nCELoss = nn.CrossEntropyLoss(ignore_index = 0)\noptimizer = AdamW(model.parameters(), lr = 1.0e-3)\n\n# +\n# model.cuda(6)\n# device = model.bert.device\n# print(device)\n\n# + jupyter={\"outputs_hidden\": true}\nfor iteration, batch in enumerate(dataloader_test):\n batch_inputs = {k: v for k, v in list (batch[0].items())}\n batch_labels = batch[1]\n \n print('before')\n print(batch[1])\n for i in batch[1]:\n for j in i:\n j = tag_to_id.get(j)\n convert_to.append(j)\n\n batch_labels = convert_to\n convert_to = []\n# batch_labels = tag_to_id.get(batch_labels)\n \n print('='*33,'batch_inputs','='*33)\n print(batch_inputs)\n print('='*33,'after_labels','='*33)\n print(batch_labels)\n# -\n\n# # START!~\n#\n# # Sample\n\n# + jupyter={\"outputs_hidden\": true}\nfor epoch in range(3): # 숫자 ->train_epoch\n model.train()\n \n for iteration, batch in enumerate(dataloader_train):\n batch_inputs = {k: v for k, v in list (batch[0].items())}\n batch_labels = batch[1]\n\n output = model(**batch_inputs)\n print(output)\n loss = CELoss(output.view(-1, output.size(-1)), batch_labels.view(-1))\n\n# optimizer.zero_grad()\n# loss.backward()\n\n# optimizer.step()\n\n# if (iteration + 1) % 10 == 0:\n# print(f'{iteration:3} - loss: {loss.item()}')\n\n# # todo 매 에포크가 끝나면 dev 데이터로 성능 비교하기\n# # Early Stopping 적용하기\n# -\n\n# # 0309_1600\n\n# +\nfor epoch in range(3): # 숫자 ->train_epoch\n# model.train()\n \n for iteration, batch in enumerate(dataloader_train):\n batch_inputs = {k: v for k, v in list (batch[0].items())}\n batch_labels = batch[1] \n \n emb = bert(**batch_inputs)\n e = nn.Dropout(0.1)\n r = e(emb['last_hidden_state'])\n w = nn.Linear(768, 22)\n output = w(r)\n \n# for i in batch[1]:\n# for j in i:\n# j = tag_to_id.get(j)\n# convert_to.append(j)\n\n# batch_labels = convert_to\n# convert_to = []\n# batch_labels = torch.tensor(batch_labels)\n\n# batch_labels.view(-1)\n# q = nn.Softmax(dim = 2) # A dimension along which Softmax will be computed\n \n loss = CELoss(output.view(-1, output.size(-1)), batch_labels.view(-1))\n\n optimizer.zero_grad()\n loss.backward()\n\n optimizer.step()\n\n if (iteration + 1) % 10 == 0:\n print(f'{iteration:3} - loss: {loss.item()}')\n\n # todo 매 에포크가 끝나면 dev 데이터로 성능 비교하기\n # Early Stopping 적용하기\n\n# +\nfor epoch in range(5): # 숫자 ->train_epoch\n model.train()\n\n for data in dataloader_train:\n print(data['input'], data['label'])\n# loss = CELoss(data['input'].view(-1, output.size(-1)), data['label'].view(-1))\n\n# optimizer.zero_grad()\n# loss.backward()\n\n# optimizer.step()\n\n# if (iteration + 1) % 10 == 0:\n# print(f'{iteration:3} - loss: {loss.item()}')\n\n # todo 매 에포크가 끝나면 dev 데이터로 성능 비교하기\n # Early Stopping 적용하기\n# -\n\n# # Mine\n\nfor epoch in range(10): # 숫자 ->train_epoch\n model.train()\n\n for iteration, batch in enumerate(dataloader_train):\n batch_inputs = {k: v.cuda(device) for k, v in list(batch[0].items())}\n batch_labels = batch[1].cuda(device)\n\n output = model(**batch_inputs)\n loss = CELoss(output.view(-1, output.size(-1)), batch_labels.view(-1))\n\n optimizer.zero_grad()\n loss.backward()\n\n optimizer.step()\n\n if (iteration + 1) % 10 == 0:\n print(f'{iteration:3} - loss: {loss.item()}')\n\n # todo 매 에포크가 끝나면 dev 데이터로 성능 비교하기\n # Early Stopping 적용하기\n\n# # Sample\n\n# +\nmodel.eval()\n\ngold_list = []\npred_list = []\n\nwith torch.no_grad():\n for iteration, batch in enumerate(dataloader_test):\n batch_inputs = {k: v.cuda(device) for k, v in list(batch[0].items())}\n batch_labels = batch[1].cuda(device)\n \n output = model(**batch_inputs)\n loss = CELoss(output.view(-1, output.size(-1)), batch_labels.view(-1))\n \n print('loss:', loss.item())\n pred_ids = torch.argmax(output, dim=-1)\n \n for g, p in zip(batch_labels, pred_ids):\n gold_mask = g != tag_converter.pad_id\n \n gold = tag_converter.convert_ids_to_tags(g[gold_mask].tolist())\n pred = tag_converter.convert_ids_to_tags(p[gold_mask].tolist())\n gold_list.append(gold)\n pred_list.append(pred)\n \n print(gold)\n print(pred)\n# -\n\n# # Mine\n\n# +\nmodel.eval()\n\ngold_list = []\npred_list = []\n\nwith torch.no_grad():\n for iteration, batch in enumerate(dataloader_test):\n batch_inputs = {k: v for k, v in list(batch[0].items())}\n batch_labels = batch[1]\n \n output = model(**batch_inputs)\n loss = CELoss(output.view(-1, output.size(-1)), batch_labels.view(-1))\n \n print('loss:', loss.item())\n pred_ids = torch.argmax(output, dim=-1)\n \n for g, p in zip(batch_labels, pred_ids):\n gold_mask = g != tag_converter.pad_id\n \n gold = tag_converter.convert_ids_to_tags(g[gold_mask].tolist())\n pred = tag_converter.convert_ids_to_tags(p[gold_mask].tolist())\n gold_list.append(gold)\n pred_list.append(pred)\n \n print(gold)\n print(pred)\n# -\n\ngold_list_flat = []\npred_list_flat = []\nfor g, p in zip(gold_list, pred_list):\n gold_list_flat += g\n pred_list_flat += p\n\nprint(classification_report(gold_list_flat, pred_list_flat, digits=5, labels=list(tag_converter.tag_to_id.keys())[1:]))\n\n\n\ndef get_chunk_type(tag_name):\n tag_class = tag_name.split('-')[0]\n tag_type = tag_name.split('-')[-1]\n return tag_class, tag_type\n\n\ndef get_chunks(seq):\n default = \"O\"\n\n chunks = []\n chunk_type, chunk_start = None, None\n for i, tok in enumerate(seq):\n # End of a chunk 1\n if tok == default and chunk_type is not None:\n # Add a chunk.\n chunk = (chunk_type, chunk_start, i)\n chunks.append(chunk)\n chunk_type, chunk_start = None, None\n\n # End of a chunk + start of a chunk!\n elif tok != default:\n tok_chunk_class, tok_chunk_type = get_chunk_type(tok)\n if chunk_type is None:\n chunk_type, chunk_start = tok_chunk_type, i\n elif tok_chunk_type != chunk_type or tok_chunk_class == \"B\":\n chunk = (chunk_type, chunk_start, i)\n chunks.append(chunk)\n chunk_type, chunk_start = tok_chunk_type, i\n else:\n pass\n\n # end condition\n if chunk_type is not None:\n chunk = (chunk_type, chunk_start, len(seq))\n chunks.append(chunk)\n\n return chunks\n\n\ndef evaluate_ner_F1(total_answers, total_preds):\n num_match = num_preds = num_answers = 0\n\n for answers, preds in zip(total_answers, total_preds):\n\n answer_seg_result = set(get_chunks(answers))\n pred_seg_result = set(get_chunks(preds))\n\n num_match += len(answer_seg_result & pred_seg_result)\n num_answers += len(answer_seg_result)\n num_preds += len(pred_seg_result)\n\n precision = 100.0 * num_match / num_preds\n recall = 100.0 * num_match / num_answers\n F1 = 2 * precision * recall / (precision + recall)\n\n return precision, recall, F1\n\n\nevaluate_ner_F1(gold_list, pred_list)\n\n\n","repo_name":"YHWOO13/JBNU-2022-SPRING","sub_path":"English world class tagging & Korean_Named Entity Recognition/Korean_Named Entity Recognition/3_Training.ipynb","file_name":"3_Training.ipynb","file_ext":"py","file_size_in_byte":17122,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"9588264018","text":"# # 1. Even or Odd Numbers\n\nnum=int(input(\"Enter a number: \"))\nif num%2==0:\n print(\"This is a even number\")\nelse:\n print(\"this is an odd number\")\n\n# # 2. Reversing Numbers\n\nn=int(input('enter a number'))\nrev=0\nwhile n!=0:\n rem=n%10\n rev=(rev*10)+rem\n n=n//10\n print(rev)\n\n# # 3. Number reversal by converting into string\n\nnumerical=str(input(\"Enter the value: \"))\nreverse=numerical[::-1]\nprint(reverse)\n\n# # 4. Prime Numbers\n\n# Method - I\n\nfor Number in range (1, 100):\n count = 0\n for i in range(2, (Number//2 + 1)):\n if(Number % i == 0):\n count = count + 1\n break\n\n if (count == 0 and Number != 1):\n print(\" %d\" %Number)\n\n# Method - II\n\n# +\nlower=int(input(\"Enter the lower value: \"))\nupper=int(input(\"Enter the upper value: \"))\n\nfor num in range(lower,upper+1):\n if num>1:\n for i in range(2,num):\n if (num%i)==0:\n break\n else:\n print(num)\n# -\n\n# # 5. Fibonacci series\n\nfn=0\nsn=1\nprint(fn)\nprint(sn)\nfor num in range(15):\n new=fn+sn\n print(new)\n fn=sn\n sn=new\n\nn=int(input(\"Enter the series: \"))\nfn=0\nsn=1\nprint(fn)\nprint(sn)\nfor i in range(n):\n new=fn+sn\n print(new)\n fn=sn\n sn=new\n\n# # 6. Armstrong Number\n\nn=int(input(\"Enter a number: \"))\ntemp=n\narm=0\nwhile n!=0:\n rem=n%10\n arm=arm+(rem*rem*rem)\n n=n//10\nif temp==arm:\n print(\"This number is an armstrong number\")\nelse:\n print(\"this number is not an armstrong number\")\n","repo_name":"MuhilanPandian/Data-Science-with-Python","sub_path":"Tasks.ipynb","file_name":"Tasks.ipynb","file_ext":"py","file_size_in_byte":1481,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"12453445075","text":"# ### resonator fitting, to get internal quality factor and external factor, etc\n# two packages are used here, one is scresonators, https://github.com/Boulder-Cryogenic-Quantum-Testbed/scresonators, one is resonator, https://github.com/danielflanigan/resonator\n# ---Zhenhai 20230227\n\nimport fit_resonator.resonator as scres\nfrom resonator import shunt, see\nimport Labber\nimport numpy as np\nimport matplotlib.pyplot as plt\n\n# ### fitting with scresonators\n\n# +\nlogfile = Labber.LogFile(r'C:\\Users\\T2-1\\Dropbox\\My PC (T2-1)\\Desktop\\Zhenhai\\data_processing\\generalCode\\Res_powerscan_2.hdf5') #load the hdf5 file\nrawData = logfile.getTraceXY(entry=20) # get the x data and y data for one specific entry\ncolumn0, column1, column2 = rawData[0]/1e9, 20*np.log10(np.abs(rawData[1])), np.angle(rawData[1]) # prepare the raw data for the fitting, 1st one for frequency in GHz, 2nd one is the S21 magnitute in dB, 3rd one is the phase in radians\nrawData4fit = np.column_stack((column0, column1, column2)) # data shape [column0, column1, column2]\n\nmy_resonator = scres.Resonator()\nmy_resonator.from_columns(rawData4fit)\n\nfit_type = 'DCM'\nMC_iteration = 20\nMC_rounds = 1e3\nMC_fix = ['w1']\nmanual_init = None\n\nmy_resonator.fit_method(fit_type, MC_iteration, MC_rounds=MC_rounds, MC_fix=MC_fix, manual_init=manual_init,\n MC_step_const=0.3)\nparams1 = my_resonator.fit()\nparams1\n# -\n\n# ### fit with Danielflanigan resonator\n\n# for using this code, you need to pieces of packages, one is called Labber, one is called Dainelflanigan resonator which you can find it on github\nimport lmfit\nimport scipy.io as sio\nimport resonator\nfrom resonator import background, shunt, see\nimport matplotlib.patches as mpatches\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport Labber\n\n# +\nfilename = r\"C:\\Users\\T2-1\\Dropbox\\My PC (T2-1)\\Desktop\\Zhenhai\\data_processing\\generalCode\\Res8_powerscan_10dB_2.hdf5\" # the file name and the path\nQi = []\nLfile = Labber.LogFile(filename)\nfreq, _ = Lfile.getTraceXY(entry=2) # get the frequency span used to measure S21, entry can be random number \nS21 = Lfile.getData()\nfor i in range(np.shape(S21)[0]): \n r = shunt.LinearShuntFitter(frequency=freq, \n data=S21[i, :],\n background_model=background.MagnitudeSlopeOffsetPhaseDelay())\n Qi.append(r.Q_i)\n\nplt.figure(figsize=(7, 5))\npower = np.linspace(10, -60, 36) # the power range used to measure resonator spectroscopy power scan\nQi = np.array(Qi)\nplt.plot(power, Qi*1e-6, 'o', color = 'C0', markersize = 6, linewidth = 10)\nplt.ylabel(\"$Q_i$ (1e6)\", fontsize=15)\nplt.xlabel(\"VNA power ($dBm$)\", fontsize=15)\nplt.xticks(fontsize=15)\nplt.grid(None)\nplt.ylim([0, 0.1e6])\nplt.legend(frameon=False,fontsize=15)\nplt.show()\n# -\n\n\n\n\n","repo_name":"zheezit/Fluxonium_vrs.3","sub_path":"Designs_scripts/Modules/resonatorFitting.ipynb","file_name":"resonatorFitting.ipynb","file_ext":"py","file_size_in_byte":2778,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"749795580","text":"# status: Draft\n#\n#\n\n# %load_ext autoreload\n\n# +\n# %autoreload 2\n# %matplotlib inline\n\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nimport WP19_analysis as wpa\n# -\n\n# create zero-filled data in memory\ndata_zf = {}\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name']\n energy_data = wpa.load_timeseries_file(vname + '-clean.csv')\n messages = wpa.load_message_file(vname + '-messages.csv')\n # zero filled data\n data_zf[vname] = wpa.insert_zeros_kVA(energy_data, messages)\n\n# Here are the CDFs for the kVA reports with nulls removed.\n#\n# TODO: is this zerofilled?\n\n# +\nfig, ax = plt.subplots(1, 2)\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name']\n if vname == 'kensio' or vname == 'atamali':\n axes = ax[0]\n else:\n axes = ax[1]\n\n sns.distplot(data_zf[vname]['kVA sliding window Demand'].dropna(), \n hist_kws={'cumulative':True,\n 'histtype':'step',\n 'linewidth':2},\n kde=False,\n norm_hist=True,\n bins=100,\n ax=axes,\n label=vname.title())\n\nax[0].set_xlabel('Reported kVA')\nax[1].set_xlabel('Reported kVA')\nax[0].set_ylabel('Cumulative Fraction of Observations')\nax[0].legend(loc='upper left')\nax[1].legend(loc='upper left')\nax[0].set_ylim((0.7, 1.0))\nax[1].set_ylim((0.7, 1.0))\nplt.savefig('../plots/power-CDF.png')\n# -\n\nfig, ax = plt.subplots()\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name']\n kVA = data_zf[vname]['kVA sliding window Demand']\n operating_kVA = kVA[kVA > 0]\n sns.distplot(operating_kVA.dropna(), \n hist_kws={'cumulative':True,\n 'histtype':'step',\n 'linewidth':2},\n kde=False,\n norm_hist=True,\n bins=100,\n ax=ax,\n label=vname)\nax.set_xlabel('Reported kVA')\nax.set_ylabel('Cumulative Fraction of Observations')\nax.legend()\nplt.savefig('../plots/power-CDF-no-zeros.png')\n\n\n# +\ndef wpa_averaged_power_profile(energy_data, messages):\n # create power column from current and voltage or use existing kVA fields?\n # print(energy_data.columns)\n # group by and aggregate by time\n # return series with averaged 24 hour power profile\n resampled_data = wpa.insert_zeros_kVA(energy_data, messages)\n kVA = resampled_data['kVA sliding window Demand']\n return kVA.groupby(kVA.index.time).mean()\n \nfig, ax = plt.subplots()\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name']\n energy_data = wpa.load_timeseries_file(vname + '-clean.csv')\n messages = wpa.load_message_file(vname + '-messages.csv')\n wpa_averaged_power_profile(energy_data, messages).plot(marker='', \n linestyle='-',\n label=vname)\n \nax.set_xlabel('Time of Day')\nax.set_ylabel('Mean kVA')\nax.legend()\nplt.savefig('../plots/hourly_kVA.png')\n\n# +\ndata = []\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name']\n energy_data = wpa.load_timeseries_file(vname + '-clean.csv')\n kVA = energy_data['kVA sliding window Demand']\n data.append({'village': vname, \n 'mean kVA': kVA.mean(), \n 'max kVA': kVA.max(), \n 'kVA factor': kVA.mean()/kVA.max(), \n 'HH': rfd['connections'], \n 'max kVA per HH': kVA.max()/rfd['connections'],\n 'mean kVA per HH': kVA.mean()/rfd['connections']})\n\nkVA_data = pd.DataFrame(data)\n\nimport tabulate\nprint(tabulate.tabulate(kVA_data, tablefmt='pipe', headers=kVA_data.columns))\n\n\n# +\n# load duration curve\n\n# FIXME: we need to use a zero-filled data set\n# FIXME: check that these are in WP19\n\ndef insert_power_gap_zeros(energy_data, messages):\n # this puts zeros in the recorded power gaps and leaves data gaps untouched\n power_down = messages[messages['message']=='Power Down'].index.values\n power_up = messages[messages['message']=='Power Up'].index.values\n # this is diffing all columns and possibly slowing down function\n diffed = energy_data[['kWh export']].resample('1T').asfreq().diff().shift(-1)\n for i in diffed.index.values:\n # if either boundary of the interval is in a power gap, we want to insert a zero\n # if the insertion point of the index is one greater for the power_down time, you are in a gap\n if np.searchsorted(power_down, i) == np.searchsorted(power_up, i) + 1:\n diffed.loc[i]['kWh export'] = 0\n end_interval = i + np.timedelta64(1,'m')\n if np.searchsorted(power_down, end_interval) == np.searchsorted(power_up, end_interval) + 1:\n diffed.loc[i]['kWh export'] = 0\n return diffed['kWh export']\n\ndef wpa_insert_zeros_kVA(energy_data, message_data):\n power_down = messages[messages['message']=='Power Down'].index.values\n power_up = messages[messages['message']=='Power Up'].index.values\n energy_data_rs = energy_data.resample('1T').asfreq()\n for i in energy_data_rs.index.values:\n if np.searchsorted(power_down, i) == np.searchsorted(power_up, i) + 1:\n energy_data_rs.loc[i]['kVA sliding window Demand'] = 0\n return energy_data_rs\n \n \ndef wpa_load_duration_curve(energy_data):\n # sort and order from largest to smallest\n # x-axis should be percent of time but we'll do raw first\n data = energy_data['kVA sliding window Demand'].sort_values(ascending=False).dropna().values\n index = np.linspace(0, 1, len(data))\n return pd.Series(index=index, data=data)\n\nvname = 'ajau'\nvname = 'asei'\nvname = 'ayapo'\nvname = 'kensio'\nvname = 'atamali'\n\nfig, ax = plt.subplots()\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name']\n energy_data = wpa.load_timeseries_file(vname + '-clean.csv')\n messages = wpa.load_message_file(vname + '-messages.csv')\n energy_data_zf = wpa_insert_zeros_kVA(energy_data, messages)\n plt.plot(wpa_load_duration_curve(energy_data_zf),\n label=vname)\n\nax.set_xlabel('Fraction of Valid Observations')\nax.set_ylabel('Apparent Power Consumed (kVA)')\nax.legend()\nplt.savefig('../plots/load_duration_curve.png')\n\n# +\ntmp_dict = {}\nfor rfd in wpa.raw_file_data[2:]:\n vname = rfd['village_name'].title()\n energy_data = wpa.load_timeseries_file(vname + '-clean.csv')\n kVA = energy_data['kVA sliding window Demand']\n try:\n rating = rfd['genset_rating_kVA']\n except:\n rating = np.nan\n tmp_dict[vname] = {'Mean load (kVA)' : kVA.mean(),\n 'max' : kVA.max(),\n 'Fraction Rated Load': kVA.mean()/kVA.max(),\n 'Rating (kVA)':rating,\n 'grid_type': rfd['grid_type']}\n\npower_stats = pd.DataFrame(tmp_dict).T\npower_stats['Fraction Generator Load'] = power_stats['Mean load (kVA)'] / power_stats['Rating (kVA)']\npower_stats\n\n# +\nimport tabulate\noutput_table = power_stats[['Rating (kVA)', 'Mean load (kVA)', 'Fraction Generator Load']]\n\ntable_string = tabulate.tabulate(output_table, tablefmt='pipe', floatfmt='.3g', headers=output_table.columns)\n\nwith open('../tables/genset_utilization.md', 'w') as f:\n f.write(table_string)\n\nprint(table_string)\noutput_table\n# -\n\n\n\n\n","repo_name":"dsoto/WP19","sub_path":"analysis/SI-power-analysis.ipynb","file_name":"SI-power-analysis.ipynb","file_ext":"py","file_size_in_byte":7332,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"70410376781","text":"# +\nimport sys\ninput = sys.stdin.readline\n\nn = int(input())\narr = sorted(list(map(int, input().split())))\n\nstart = 0\nend = n-1\nczero = abs(arr[start]+arr[end])\ncstart = start\ncend = end\n\nwhile start 0:\n end -= 1\n else:\n start += 1\n\nprint(arr[cstart], arr[cend])\n\n \n# -\n\n\n","repo_name":"youngjun0627/Rgorithm","sub_path":"두 용액.ipynb","file_name":"두 용액.ipynb","file_ext":"py","file_size_in_byte":469,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"8095897770","text":"#\n# # Using ANN model for longawave estimation of cloud fraction \n\n#\n# ## Step 1 - Import and init\n\n# +\nimport tensorflow as tf\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport pandas as pd\nimport os,sys\nimport time\nimport keras_tuner as kt \n\nfrom tensorflow import keras\nfrom keras.callbacks import EarlyStopping\nfrom numpy import loadtxt\nfrom keras.models import load_model\nfrom pathlib import Path\nfrom keras.callbacks import EarlyStopping\nfrom keras_tuner import HyperParameters \n\nsys.path.append('..')\npath = '/home/monitor/IGE_DOC/GIT_REPO/LongwaveEstimationOfCloudFraction-With-XGBoost-And-ANNmodel/Data/'\n\n# -\n\n# ## Step 2 - Retrieve data\n\ndata = pd.read_csv(path + 'DataForML_5min.csv', header=0)\ndel data['timestamp']\ndata\n\n# ## Step 3 - Preparing the data\n# ### 3.1 - Split data\n# We will use 90% of the data for training and 10% for validation. \n# The dataset is **shuffled** and shared between **learning** and **testing**. \n# x will be input data and y the expected output\n\n# +\ndata_pred = data[-10000:]\n\ny_real = data_pred['CF']\nx_pred = data_pred.drop('CF', axis=1)\n\ndata_pred = data_pred.reset_index()\n\ndata_pred = data_pred.drop('index', axis=1)\ndata_pred\n\n# +\n# ---- Shuffle and Split => train, test\n#\ndata = data.drop(data_pred.index)\ndata = data.sample(frac=1., axis=0)\ndata_train = data.sample(frac=0.95, axis=0)\ndata_test = data.drop(data_train.index)\n\n# ---- Split => x,y (medv is price)\n#\nx_train = data_train.drop('CF', axis=1)\ny_train = data_train['CF']\nx_test = data_test.drop('CF', axis=1)\ny_test = data_test['CF']\n\nprint('Original data shape was : ',data.shape)\nprint('x_train : ',x_train.shape, 'y_train : ',y_train.shape)\nprint('x_test : ',x_test.shape, 'y_test : ',y_test.shape)\nprint('x_pred : ',x_pred.shape, 'y_real : ',y_real.shape)\n# -\n\nx_train.shape, x_test.shape\n\n# ### 3.2 - Data normalization\n# **Note :** \n# - All input data must be normalized, train and test. \n# - To do this we will **subtract the mean** and **divide by the standard deviation**. \n# - But test data should not be used in any way, even for normalization. \n# - The mean and the standard deviation will therefore only be calculated with the train data.\n\n# +\ndisplay(x_train.describe().style.format(\"{0:.2f}\").set_caption(\"Before normalization :\"))\n\nmean = x_train.mean()\nstd = x_train.std()\nx_train = (x_train - mean) / std\nx_test = (x_test - mean) / std\n\ndisplay(x_train.describe().style.format(\"{0:.2f}\").set_caption(\"After normalization :\"))\ndisplay(x_train.head(5).style.format(\"{0:.2f}\").set_caption(\"Few lines of the dataset :\"))\n\nx_train, y_train = np.array(x_train), np.array(y_train)\nx_test, y_test = np.array(x_test), np.array(y_test)\n\n\n# +\ndisplay(x_pred.describe().style.format(\"{0:.2f}\").set_caption(\"Before normalization :\"))\n# for pred \nx_pred = (x_pred - mean) / std\n\ndisplay(x_pred.describe().style.format(\"{0:.2f}\").set_caption(\"After normalization :\"))\ndisplay(x_pred.head(5).style.format(\"{0:.2f}\").set_caption(\"Few lines of the dataset :\"))\n\nx_pred, y_real = np.array(x_pred), np.array(y_real)\n\n\n# -\n\n# ## Step 4 - Build a model\n# About informations about : \n# - [Optimizer](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers)\n# - [Activation](https://www.tensorflow.org/api_docs/python/tf/keras/activations)\n# - [Loss](https://www.tensorflow.org/api_docs/python/tf/keras/losses)\n# - [Metrics](https://www.tensorflow.org/api_docs/python/tf/keras/metrics)\n# \n# The range of layers and neuron have been chosen according to the data and the number of features.\n\ndef build_model(hp):\n \n model = keras.models.Sequential()\n \n model.add(keras.layers.Input(7, name=\"InputLayer\"))\n \n model.add(keras.layers.Dense(hp.Choice('units', [8, 10, 12]), activation='relu', name='Dense_n1'))\n \n model.add(keras.layers.Dense(hp.Choice('units', [8, 10, 12]), activation='relu', name='Dense_n2'))\n \n model.add(keras.layers.Dense(hp.Choice('units', [8, 10, 12]), activation='relu', name='Dense_n3'))\n \n model.add(keras.layers.Dense(1, name='Output'))\n \n learning_rate = hp.Float(\"lr\", min_value=1e-3, max_value=1e-2, sampling=\"log\")\n \n model.compile(optimizer= keras.optimizers.Adam(learning_rate=learning_rate),\n loss = 'mse',\n metrics = ['mae', 'mse'])\n \n return model\n\n\n# Here we use keras tuner to tune the model. There are many package that we can use for optimizing or tuning the hyperparameters. We chose keras tuner beacause it's easy to use. \n\n# Model to optimize :\n\n# +\nhp = HyperParameters()\n\nshape = 7 \n\nbuild_model(kt.HyperParameters())\n# -\n\n# RandomSearch method for Tuning hyperparameters \n\ntuner = kt.RandomSearch(hypermodel=build_model, objective=\"val_loss\",\n \n max_trials=3,executions_per_trial=2,overwrite=True,\n \n directory=\"D:/TUTO/data_science/formation_fedel/fidle-master/BHPD/Optimization_results\", project_name=\"Optim_RandomForest\",)\n\n# Summary\n\ntuner.search_space_summary()\n\n#\n#\n# Runing keras tuner and search the best hyperparameters\n#\n#\n\n# +\n\ntuner.search(x_train, y_train, epochs=50, validation_data=(x_test, y_test)) # , callbacks=[stop_early]\n\n# -\n\n# Here we take the best model \n\n# +\n# Get the top 2 models.\n# models = tuner.get_best_models(num_models=1)\n\nbest_model_1 = tuner.get_best_models()[0]\n\nbest_hps=tuner.get_best_hyperparameters(num_trials=1)[0]\n\nbest_model_1 , best_hps\n# -\n\n# ## Step 5 - Train the model\n# ### 5.1 - Get it\n\nbest_model = tuner.hypermodel.build(best_hps)\ntuner.results_summary()\n\n# ### 5.2 - Train it\n\n# We need to retrain the best model \n\n# + jupyter={\"outputs_hidden\": true}\nes = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=200)\n\ncheckpoint_cb = keras.callbacks.ModelCheckpoint(\"check_point_my_keras_model.h5\",save_best_only=True)\n\nbest_model = tuner.hypermodel.build(best_hps)\n\nhistory = best_model.fit(x_train,\n y_train,\n epochs = 50, #60 init \n batch_size = 10, #10\n verbose = fit_verbosity,\n callbacks =[es,checkpoint_cb],\n validation_data = (x_test, y_test))\n# -\n\n# #### Recovery of the model in the event of a training stoppage \n\nmodel = keras.models.load_model(\"check_point_my_keras_model.h5\") \n\nkeras.utils.plot_model(best_model)\n\n# #### Best epoch for training \n\n# +\nval_loss_per_epoch = history.history['val_loss']\n\nbest_epoch = val_loss_per_epoch.index(max(val_loss_per_epoch)) + 1\n\nprint('Best epoch: %d' % (best_epoch,))\n# -\n\n# We retrain with the best epoch \n\n# + jupyter={\"outputs_hidden\": true}\nbest_model_epoch_ = tuner.hypermodel.build(best_hps)\n\nhistory_2 = best_model_epoch_.fit(x_train,\n y_train,\n epochs = 50, #60 init \n batch_size = 10, #10\n verbose = fit_verbosity,\n callbacks =[es],\n validation_data = (x_test, y_test))\n# -\n\neval_result = best_model.evaluate(x_test, y_test)\nprint(\"[test loss, test score]:\", eval_result)\n\n# ## Step 6 - Evaluate\n# ### 6.1 - Model evaluation\n# MAE = Mean Absolute Error (between the labels and predictions) \n\n# +\nscore = best_model.evaluate(x_test, y_test, verbose=0)\n\nprint('x_test / loss : {:5.4f}'.format(score[0]))\nprint('x_test / mae : {:5.4f}'.format(score[1]))\nprint('x_test / mse : {:5.4f}'.format(score[2]))\n\nfor i in range(3):\n print(\"%s: %.2f%%\" % (best_model.metrics_names[i], score[i]*100))\n# -\n\nName_to_save = \"50_3L_epoch_Best_model_ANN\"\nnp.mean(score[0]*100)\n\n# ### Save history in a csv file\n\n# +\ndf=pd.DataFrame(data=history_2.history)\n\nfilepath = Path('D:\\TUTO\\data science\\formation_fedel\\fidle-datasets\\model.csv') \n\ndf.to_csv(Name_to_save+'.csv') \n# -\n\n# ### Save model \n\n# +\nNAME = Name_to_save+\"_{}\".format(int(time.time())) \n\n# save model and architecture to single file\nbest_model.save(NAME+\".h5\")\n\nprint(\"Saved model to disk\")\n# -\n\n# ### Load model \n\n# +\n# load and evaluate a saved model\n\n# load model\nmodel = load_model(NAME+\".h5\")\n# model = load_model(\"50_3L_epoch_Best_model_ANN_1645999529.h5\")\n\n# summarize model.\nmodel.summary()\n\n# evaluate the model\nscore = model.evaluate(x_test, y_test, verbose=0)\nfor i in range(3):\n print(\"%s: %.2f%%\" % (model.metrics_names[i], score[i]*100))\n# -\n\n# figure, axis = plt.subplots(1, 3, figsize=(25,8))\nfor i in range(3):\n \n plt.plot(range(50), df[cols[i]], 'r')\n \n plt.plot(range(50), df[cols[i+3]], 'b')\n \n plt.title(cols[i])\n \n plt.show()\n\n\nprint(\"min( val_mae ) : {:.4f}\".format( min(history_2.history[\"val_mae\"]) ) )\n\n\n# ## Step 7 - Make a prediction\n# The data must be normalized with the parameters (mean, std) previously used.\n\n# ### Plot outcome \n\n# +\ndef plot_pred(start,end, y_pred, y): \n \n # Setup plot (2 rows, 1 column)\n fig, comb = plt.subplots(nrows=1,ncols=1, sharex=True, figsize=(20, 10))\n \n # PLOT sup\n comb.plot(range(len(y_pred[start:end])),y_pred[start:end], color='r', label = \"y_pred\") \n \n comb.plot(range(len(y[start:end])),y[start:end], color='b', label = \"y_real\")\n \n comb.set_title(\"Prediction\", fontsize=18)\n \n comb.legend(prop={'size':15})\n \n comb.set_ylabel('Cloud fraction', fontsize=18)\n \n comb.set_xlabel('Times', fontsize=18)\n \n comb.tick_params(labelsize=14)\n \n return \n\ndata_comp_resutl = pd.DataFrame({'y_pred': y_pred_cln, 'y_real': y_real[:].tolist()})\n\nstart = 0\n\nfor i in range(1):\n \n end = start + 800\n \n plot_pred(start,end,data_comp_resutl['y_pred'], data_comp_resutl['y_real'])\n \n start = start + end\n \n# -\n\n# ### Score \n#\n# * r2_score\n\nround(r2_score(y_real[:], predictions[0][:]),4)*100\n","repo_name":"TinaAndriantsalama/Estimation-of-cloud-fraction-by-longwave-radiation-using-machine-learning","sub_path":"ANN/DNN_CF_Optimization.ipynb","file_name":"DNN_CF_Optimization.ipynb","file_ext":"py","file_size_in_byte":9868,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"39421278728","text":"import numpy as np\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n# %matplotlib inline\n# %config InlineBackend.figure_format = 'retina'\nfrom sklearn import decomposition # for PCA\nfrom sklearn import datasets\nfrom mpl_toolkits.mplot3d import Axes3D\n\ndf = datasets.load_iris() # Load the dataset\n\n\n# +\nX = df.data\nY = df.target\nfig = plt.figure(1,figsize=(4,4))\nplt.clf()\nax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134) # Create a 3D plot\n\nplt.cla()\nfor name, label in [('Setosa', 0), ('Versicolour', 1), ('Virginica', 2)]:\n ax.text3D(X[Y == label, 0].mean(),\n X[Y == label, 1].mean() + 1.5,\n X[Y == label, 2].mean(), name,\n horizontalalignment='center',\n bbox=dict(alpha=.5, edgecolor='w', facecolor='w'))\n \n \n \n# Change the order of labels, so that they match\ny_clr = np.choose(Y, [1, 2, 0]).astype(np.float)\nax.scatter(X[:, 0], X[:, 1], X[:, 2], c=y_clr, \n cmap=plt.cm.nipy_spectral)\n\nax.w_xaxis.set_ticklabels([])\nax.w_yaxis.set_ticklabels([])\nax.w_zaxis.set_ticklabels([]);\n\n\n# +\nfrom sklearn.tree import DecisionTreeClassifier\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import accuracy_score, roc_auc_score\n\nX_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=.3, \n stratify=Y, \n random_state=42)\n# divide data into train and test\nclf = DecisionTreeClassifier(max_depth=2, random_state=42)\nclf.fit(X_train, y_train)\npreds = clf.predict_proba(X_test)\nprint('Accuracy: {:.5f}'.format(accuracy_score(y_test, \n preds.argmax(axis=1))))\n\n\n# +\npca = decomposition.PCA(n_components=3) # Keep the number of axes as three\npca\nX_centered = X - X.mean(axis=0)\npca.fit(X_centered)\nX_pca = pca.transform(X_centered)\n\n# Plotting the results of PCA\nplt.plot(X_pca[Y == 0, 0], X_pca[Y == 0, 1], 'bo', label='Setosa')\nplt.plot(X_pca[Y == 1, 0], X_pca[Y == 1, 1], 'go', label='Versicolour')\nplt.plot(X_pca[Y == 2, 0], X_pca[Y == 2, 1], 'ro', label='Virginica')\nplt.legend(loc=0);\n\n# +\nX_train, X_test, y_train, y_test = train_test_split(X_pca, Y, test_size=.3, \n stratify=Y, \n random_state=42)\n\nclf = DecisionTreeClassifier(max_depth=2, random_state=42)\nclf.fit(X_train, y_train)\npreds = clf.predict_proba(X_test)\nprint('Accuracy: {:.5f}'.format(accuracy_score(y_test, \n preds.argmax(axis=1))))\n\n# Increase in accuray after performing PCA\n\n# +\n# Apply k_Means for a random dataset\nX = np.zeros((300,2))\nnp.random.seed(seed=0) # Produce same number everytime\n\nX[:100, 0] = np.random.normal(loc=0.0, scale=.3, size=100) #The NumPy random normal function enables to create a NumPy array that contains normally distributed data.\nX[:100, 1] = np.random.normal(loc=0.0, scale=.3, size=100)\n\nX[100:200, 0] = np.random.normal(loc=2.0, scale=.5, size=100)\nX[100:200, 1] = np.random.normal(loc=-1.0, scale=.2, size=100)\n\nX[200:300, 0] = np.random.normal(loc=-1.0, scale=.2, size=100)\nX[200:300, 1] = np.random.normal(loc=2.0, scale=.5, size=100)\n\nplt.figure(figsize=(5, 7))\nplt.plot(X[:, 0], X[:, 1], 'bo');\n\n\n# +\nfrom scipy.spatial.distance import cdist\n\n# Randomly allocate the 3 centroids \nnp.random.seed(seed=42)\ncentroids = np.random.normal(loc=0.0, scale=1., size=6)\ncentroids = centroids.reshape((3, 2))\n\ncent_history = []\ncent_history.append(centroids)\n\nfor i in range(3):\n # Calculating the distance from a point to a centroid\n distances = cdist(X, centroids)\n # Checking what's the closest centroid for the point\n labels = distances.argmin(axis=1)\n \n #distance of each point\n centroids = centroids.copy()\n centroids[0, :] = np.mean(X[labels == 0, :], axis=0)\n centroids[1, :] = np.mean(X[labels == 1, :], axis=0)\n centroids[2, :] = np.mean(X[labels == 2, :], axis=0)\n \n cent_history.append(centroids)\n# -\n\nplt.figure(figsize=(8, 8))\nfor i in range(4):\n distances = cdist(X, cent_history[i])\n labels = distances.argmin(axis=1)\n \n plt.subplot(2, 2, i + 1)\n plt.plot(X[labels == 0, 0], X[labels == 0, 1], 'bo', label='cluster #1')\n plt.plot(X[labels == 1, 0], X[labels == 1, 1], 'co', label='cluster #2')\n plt.plot(X[labels == 2, 0], X[labels == 2, 1], 'mo', label='cluster #3')\n plt.plot(cent_history[i][:, 0], cent_history[i][:, 1], 'rX')\n plt.legend(loc=0)\n plt.title('Step {:}'.format(i + 1));\n\nfrom sklearn.cluster import KMeans\ninertia = []\nfor k in range(1, 8):\n kmeans = KMeans(n_clusters=k, random_state=1).fit(X)\n inertia.append(np.sqrt(kmeans.inertia_))\n\nplt.plot(range(1, 8), inertia, marker='s');\nplt.xlabel('number of clusters')\nplt.ylabel('intra SSE');\n\n# +\n# Heirarchial clustering \n# Agglomerative clustering\n\nfrom scipy.cluster import hierarchy\nfrom scipy.spatial.distance import pdist\n\nX = np.zeros((150, 2))\n\nnp.random.seed(seed=42)\nX[:50, 0] = np.random.normal(loc=0.0, scale=.3, size=50)\nX[:50, 1] = np.random.normal(loc=0.0, scale=.3, size=50)\n\nX[50:100, 0] = np.random.normal(loc=2.0, scale=.5, size=50)\nX[50:100, 1] = np.random.normal(loc=-1.0, scale=.2, size=50)\n\nX[100:150, 0] = np.random.normal(loc=-1.0, scale=.2, size=50)\nX[100:150, 1] = np.random.normal(loc=2.0, scale=.5, size=50)\n\n\n# pdist will calculate the upper triangle of the pairwise distance matrix\ndistance_mat = pdist(X) \n# linkage — is an implementation if agglomerative algorithm\nZ = hierarchy.linkage(distance_mat, 'single')\nplt.figure(figsize=(10, 5))\ndn = hierarchy.dendrogram(Z, color_threshold=0.5)\n# -\n\n\n","repo_name":"krutikabapat/Data_Mining-and-ML-Algorithms","sub_path":"Unsupervised clustering and PCA.ipynb","file_name":"Unsupervised clustering and PCA.ipynb","file_ext":"py","file_size_in_byte":5697,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"15538366504","text":"#importing modules\n# %matplotlib inline\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport sympy as sym\nfrom sympy import *\n\n\n# +\n# func - the function you would like to approximate.\n# start, the beginning of your interval.\n# end, the end of your interval.\n# degree, the last integer where you will be truncating your series.\n# fixed_c, the point you will be expanding around.\n\ndef TaylorSeriesApprox (func,start,end,degree,fixed_c):\n x1 = np.linspace(start,end,100) # independent x vaules\n tsa=0 \n for i in range(degree): #taylor series expanding for the given specification\n tsa += (diff(func,x,i).subs(x,fixed_c))/(factorial(i))*(x-fixed_c)**i\n \n #substituting the x values to find y \n y1 = [tsa.subs(x,j) for j in x1[:]]\n return y1\n\n\n# +\n#x taken as variable\nx=sym.symbols(\"x\")\n\n#the function you would like to approximate.\nfunc = x*sin(x)**2+cos(x)\n\n# independent x vaules\nx1 = np.linspace(-10,10,100)\n\n#Approximated y vlaues\ny1 = TaylorSeriesApprox (func,-10,10,99,0)\n\n#subsituted y values\ny2 = [func.subs(x,j) for j in x1[:]]\n\n#plotting the grpah\nplt.plot(x1,y1,'ro')\nplt.plot(x1,y2,'k-')\nplt.xlabel('$x$',fontsize=12)\nplt.ylabel('$xsin^2(x)+cos(x)$',fontsize=12)\nplt.legend(['Taylor Approximation','Actual'],loc=2)\nplt.show()\n# -\n\n\n","repo_name":"revanth3003/Taylor-Series-Approximation","sub_path":"taylor.ipynb","file_name":"taylor.ipynb","file_ext":"py","file_size_in_byte":1275,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"36570911603","text":"# # Cylindrical Nanowire Hamiltonian Spectrum\n# In this notebook, I will investigate the Hamiltonian of a cylindrical nanowire SNS junction. Spin-orbit coupling and Zeeman term will be assumed to be present. \n#\n# Consider a nanowire placed longitudinally along the z-axis. A magnetic field $B_z$ is also assumed to be present along the z-axis.\n#\n# $$ H_e = \\frac{p_z^2 + p_\\phi^2 + p_r^2}{2m} - \\mu + \\alpha (\\vec{\\sigma} \\times \\vec{p}) \\cdot \\hat{r} + \\frac{1}{2} g \\mu_B \\hbar B_z \\sigma_z $$\n#\n# $\\textbf{Update 4th May:\n# In the intial version of code in this document, the spin-orbit and Zeeman terms will be dropped.}$\n\n# +\nimport numpy as np\nimport matplotlib.pyplot as plt\n# %matplotlib inline\n\nimport itertools\nimport scipy.special\nimport scipy.integrate\n\n\n# -\n\n# %load_ext line_profiler\n\n# +\n# create the Hamiltonian matrix\n\ndef calc_hamiltonian(params):\n '''\n Returns the Hamiltonian in the real space basis.\n params is a dict with the following information:\n N_z : number of points along z\n N_phi : number of points along azimuthal direction\n t_z : h^2/(2 m a^2), where a is lattice spacing\n t_phi : h^2/(2 m R^2), where a is lattice spacing\n mu : chemical potential\n '''\n N_z = params['N_z']\n N_phi = params['N_phi']\n t_z = params['t_z']\n t_phi = params['t_phi']\n mu = params['mu']\n flux = params['flux']\n Delta = params['Delta']\n \n def calc_matrix_element(x,y):\n '''\n Returns the matrix element between two real space points x and y\n '''\n (z1,phi1) = x\n (z2,phi2) = y\n # onsite element\n if z1 == z2 and phi1 == phi2:\n #diag_ele = 2*t_z + np.abs(t_phi)*(2 + (2*np.pi*flux/N_phi)**2) - mu\n diag_ele = 2*t_z + np.abs(t_phi)*(2 + flux**2) - mu\n #if N_phi != 1:\n # diag_ele = 2*t_z + np.abs(t_phi)*(2 + (2*np.pi*flux/N_phi)**2) - mu\n #else:\n # # diagonal elemenet for N_phi = 1 does not make sense\n # diag_ele = 2*t_z - mu\n return np.array([[diag_ele,Delta],[np.conj(Delta),-diag_ele ]])\n # z hopping\n elif abs(z1-z2) == 1 and phi1 == phi2:\n return np.array([[-t_z,0],[0,t_z]])\n # phi hopping\n elif (phi1-phi2 == 1 or phi1-phi2 == N_phi-1)and z1 == z2:\n return np.array([[-t_phi,0],[0,np.conj(t_phi)]])\n elif (phi1-phi2 == -1 or phi1-phi2 == -N_phi+1)and z1 == z2:\n return np.conj(np.array([[-t_phi,0],[0,np.conj(t_phi)]])).T\n else:\n return np.array([[0,0],[0,0]])\n \n basis = list(itertools.product(range(N_z),range(N_phi)))\n H_list = [calc_matrix_element(x,y) for x in basis for y in basis]\n N = N_phi*N_z\n H = np.array(H_list).reshape((N,N,2,2))\n \n # magic to flatten the Hamiltonian\n # Are you wathing closely?\n H = np.array([H[x,:,y,:].flatten() for x in range(H.shape[0]) for y in range(H.shape[2])])\\\n .flatten().reshape(2*N,2*N)\n \n return H\n \n\n\n# +\n# test of the Hamiltonian function\nparams = {\n \"N_z\" : 2,\n \"N_phi\" : 5,\n \"flux\" : 0e0,\n \"t_z\" : 5e-3,\n \"Delta\" : 1e-3,\n \"mu\" : 0e-3,\n}\n\nparams[\"t_phi\"] = 1e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\n\n# #%lprun -f calc_hamiltonian H = calc_hamiltonian(params)\nH = calc_hamiltonian(params)\nprint(H.shape)\n\n\n# +\n# surface Green's funciton routines\n# ordinary iteration schemes will be used\n\ndef calc_surface_g(E,alpha,beta,eta,eps = 1e-8,max_iter=100000,kappa=0.5):\n '''\n Return the surface Green's function at energy E \n \n eta is used to calculated the Green's function\n eps controls the accuracy\n \n Returns the surface Green's function as a solution of the recursive relation \n g = ((E + 1j*eta) - alpha - beta'*g*beta)^-1\n '''\n # conv here to avoid problems later\n E = np.array(E)\n \n \n def func_g(g,E,alpha,beta,eta):\n return np.linalg.inv((E + 1j*eta).reshape(-1,1,1)*np.eye(alpha.shape[0])- alpha - beta @ g @ np.conj(beta).T)\n \n g0 = np.zeros((len(E),alpha.shape[0],alpha.shape[1]))\n g = np.zeros((len(E),alpha.shape[0],alpha.shape[1]))\n \n err_vec = []\n for i in range(max_iter):\n g = func_g(g,E,alpha,beta,eta)\n g = ((1-kappa)*g + kappa*g0)\n err = np.linalg.norm(g - g0)/len(E)\n err_vec.append(err)\n g0 = np.copy(g)\n if err < eps:\n break\n return g\n\n# +\n# testing the surface_g code\n# test of the Hamiltonian function\nparams = {\n \"N_z\" : 2,\n \"N_phi\" : 5,\n \"flux\" : 0e0,\n \"t_z\" : 100e-3,\n \"Delta\" : 1e-3,\n \"mu\" : 10e-3,\n}\n\nparams[\"t_phi\"] = 1e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\n\n\nH = calc_hamiltonian(params)\nsurface_dof = 2*params['N_phi']\nalpha = H[:surface_dof,:surface_dof]\nbeta = H[:surface_dof,surface_dof:2*surface_dof]\n\nE = np.linspace(-2e-3,2e-3,100)\neta = 0.1*np.abs(E[1]-E[0])\n\n#g = calc_surface_g(E,alpha,beta,eta)\n# %lprun -f calc_surface_g g = calc_surface_g(E,alpha,beta,eta)\n\na = 1j*(g - np.conj(np.transpose(g,(0,2,1))))\n\ndos = np.real(np.trace(a,axis1=-1,axis2=-2))\n\nplt.plot(E,dos)\nplt.xlabel(\"Energy\")\nplt.ylabel(\"Surface Density of States\")\n\n\n# -\n\ndef calc_sns_negf(E,params):\n '''\n Returns the G,A,Sigma matrices for SNS device. The chemical potential will be assumed to same everywhere and\n no bias is present across the device.\n params dict takes in these parameters:\n N_z : number of points along z in the normal region\n N_phi : number of points along azimuthal direction\n t_z : h^2/(2 m a^2), where a is lattice spacing\n t_phi : h^2/(2 m R^2), where a is lattice spacing\n mu : chemical potential\n Delta_1 : superconducting parameter 1\n Delta_2 : superconducting parameter 2\n eta : imaginary 0+ added for convergence\n '''\n N_z = params['N_z']\n N_phi = params['N_phi']\n t_z = params['t_z']\n t_phi = params['t_phi']\n Delta_1 = params['Delta_1']\n Delta_2 = params['Delta_2']\n mu = params['mu']\n eta = params['eta']\n flux = params['flux']\n kT = params['kT']\n \n # create alpha/beta hamiltonians for S regions\n # calc surface Green's function\n \n params_S1 = {'t_z' : t_z,'t_phi' : t_phi,'N_z' : N_z,\"N_phi\" : N_phi, \"Delta\" : Delta_1, \"mu\" : mu,\"flux\" : flux}\n H_S1 = calc_hamiltonian(params_S1) \n surface_dof = 2*N_phi\n alpha_1 = H_S1[:surface_dof,:surface_dof]\n beta_1 = H_S1[:surface_dof,surface_dof:2*surface_dof]\n g_1 = calc_surface_g(E,alpha_1,beta_1,eta)\n \n params_S2 = {'t_z' : t_z,'t_phi' : t_phi,'N_z' : N_z,\"N_phi\" : N_phi, \"Delta\" : Delta_2, \"mu\" : mu,\"flux\" : flux}\n H_S2 = calc_hamiltonian(params_S2) \n surface_dof = 2*N_phi\n alpha_2 = H_S2[:surface_dof,:surface_dof]\n beta_2 = H_S2[:surface_dof,surface_dof:2*surface_dof]\n g_2 = calc_surface_g(E,alpha_2,beta_2,eta)\n \n \n # create Hamiltonian for N region\n params_N = {'t_z' : t_z,'t_phi' : t_phi,'N_z' : N_z,\"N_phi\" : N_phi, \"Delta\" : 0, \"mu\" : mu,\"flux\" : flux}\n H_mat = calc_hamiltonian(params_N)\n \n #number of dof in a layer\n N_dof_lat = N_phi*2\n # the hopping element between layers\n beta_layer = H_mat[:N_dof_lat,N_dof_lat:2*N_dof_lat]\n \n # the only non-zero elements in sigma\n sigma_mini_1 = np.conj(np.transpose(beta_layer[np.newaxis,:,:],(0,2,1)))@g_1@beta_layer\n sigma_mini_2 = np.conj(np.transpose(beta_layer[np.newaxis,:,:],(0,2,1)))@g_2@beta_layer\n \n sigma_1 = np.zeros((len(E),H_mat.shape[0],H_mat.shape[1]),dtype=np.complex64)\n sigma_1[:,:N_dof_lat,:N_dof_lat] = sigma_mini_1\n gamma_1 = 1j*(sigma_1 - np.transpose(np.conj(sigma_1),(0,2,1)))\n \n sigma_2 = np.zeros((len(E),H_mat.shape[0],H_mat.shape[1]),dtype=np.complex64)\n sigma_2[:,-N_dof_lat:,-N_dof_lat:] = sigma_mini_2\n gamma_2 = 1j*(sigma_2 - np.transpose(np.conj(sigma_2),(0,2,1)))\n \n # NEGF matrices\n G = np.linalg.inv((E + 1j*eta).reshape(-1,1,1)*np.eye(H_mat.shape[0]) - H_mat - sigma_1 - sigma_2)\n \n A = 1j*(G - np.transpose(np.conj(G),(0,2,1))) \n dos = np.real(np.trace(A,axis1=-2,axis2=-1))\n \n def fermi(E,kT):\n return scipy.special.expit(-E/kT)\n \n def generate_fermi_matrix(E,mu,kT):\n return np.array([[[fermi(ele - mu,kT),0],[0,fermi(ele + mu,kT)]] for ele in E])\n \n F1 = np.kron(np.eye(N_phi*N_z),generate_fermi_matrix(E,mu-mu,kT))\n F2 = np.kron(np.eye(N_phi*N_z),generate_fermi_matrix(E,mu-mu,kT))\n sigma_in = np.matmul(gamma_1,F1) + np.matmul(gamma_2,F2)\n \n G_n = np.matmul(G,np.matmul(sigma_in,np.transpose(np.conj(G),(0,2,1))))\n \n #I_mat = 1j*(np.matmul(G_n[:N_dof_lat,N_dof_lat:2*N_dof_lat],beta_layer) \\\n # - np.matmul(G_n[N_dof_lat:2*N_dof_lat,:N_dof_lat],beta_layer))\n I_mat = 1j*(np.matmul(H_mat,G_n) - np.matmul(G_n,H_mat))\n # current = electron current - hole current\n I = np.real(np.trace(I_mat[:,0:2*N_phi:2,0:2*N_phi:2]-I_mat[:,1:2*N_phi:2,1:2*N_phi:2],axis1=-1,axis2=-2))\n \n return G,A,dos,I\n\n# +\nparams = {\n \"N_z\" : 2,\n \"N_phi\" : 5,\n \"flux\" : 1e-1,\n \"t_z\" : 100e-3,\n \"Delta_1\" : 1e-3,\n \"Delta_2\" : 1e-3*np.exp(1j*np.pi/2),\n \"mu\" : 10e-3,\n \"kT\" : 1e-5,\n}\n\nparams[\"t_phi\"] = 1e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\nE_vec = np.linspace(-2e-3,2e-3,100)\nparams['eta'] = np.abs(E_vec[1] - E_vec[0])\n\n#G,A,dos,I = calc_sns_negf(E_vec,params)\n# %lprun -f calc_sns_negf G,A,dos,I = calc_sns_negf(E_vec,params)\n# -\n\nplt.plot(E_vec,dos)\n\nplt.plot(E_vec,I)\n\n# ## Dos vs phase map\n\n# +\n# testing the dos code\nparams = {\n \"N_z\" : 20,\n \"N_phi\" : 3,\n \"flux\" : 2,\n \"t_z\" : 100e-3,\n \"Delta_1\" : 1e-3,\n \"Delta_2\" : 1e-3,\n \"mu\" : 20e-3,\n \"kT\" : 1e-5,\n}\nparams[\"t_phi\"] = 2e-3*np.exp(1j*params[\"flux\"]/params[\"N_phi\"])\nE_vec = np.linspace(-2e-3,2e-3,100)\nparams['eta'] = np.abs(E_vec[1] - E_vec[0])\n\ndos_arr = []\nI_arr = []\nphi_vec = np.linspace(0,2*np.pi,25)\nfor phi in phi_vec:\n params['Delta_2'] = params['Delta_1']*np.exp(1j*phi)\n G,A,dos,I = calc_sns_negf(E_vec,params)\n dos_arr.append(dos)\n I_arr.append(I)\n# -\n\ndos_arr = np.array(dos_arr).T\nXX,YY = np.meshgrid(phi_vec,E_vec)\nplt.pcolor(XX,YY,dos_arr)\n\n# DOS vs current map\nI_arr = np.array(I_arr).T\nXX,YY = np.meshgrid(phi_vec,E_vec)\nplt.pcolor(XX,YY,I_arr)\n\n\n# ## Current phase\n\n# +\ndef integrate_current(E_vec,I_vec):\n return scipy.integrate.simps(I_vec,E_vec) \n\ndef calc_I_phi(params,E_vec,phi_vec):\n I_phi = []\n for phi in phi_vec:\n params['Delta_2'] = params['Delta_1']*np.exp(1j*phi)\n G,A,dos,I = calc_sns_negf(E_vec,params)\n I_phi.append(integrate_current(E_vec,I))\n I_phi = np.array(I_phi)\n return I_phi\n\n\n# +\nparams = {\n \"N_z\" : 2,\n \"N_phi\" : 5,\n \"flux\" : 1e-1,\n \"t_z\" : 100e-3,\n \"Delta_1\" : 1e-3,\n \"Delta_2\" : 1e-3,\n \"mu\" : 10e-3,\n \"kT\" : 1e-5,\n}\nparams[\"t_phi\"] = 1e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\nE_vec = np.linspace(-2e-3,2e-3,100)\nphi_vec = np.linspace(0,2*np.pi,100)\nparams['eta'] = np.abs(E_vec[1] - E_vec[0])\n\nI_phi = calc_I_phi(params,E_vec,phi_vec)\n# #%lprun -f calc_I_phi I_phi = calc_I_phi(params,E_vec,phi_vec)\n# -\n\nplt.plot(phi_vec,I_phi)\n\n# ## Study of current phase relation with changing chemical potential\n\n# +\nparams = {\n \"N_z\" : 2,\n \"N_phi\" : 1,\n \"flux\" : 0e0,\n \"t_z\" : 1000e-3,\n \"Delta_1\" : 1e-3,\n \"Delta_2\" : 1e-3,\n \"mu\" : 100e-3,\n \"kT\" : 1e-5,\n}\nparams[\"t_phi\"] = 5e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\nE_vec = np.linspace(-2e-3,2e-3,100)\nphi_vec = np.linspace(0,2*np.pi,100)\nparams['eta'] = np.abs(E_vec[1] - E_vec[0])\n\nmu_vec = np.linspace(10e-3,100e-3,100)\nI_arr = []\nfor mu in mu_vec:\n params['mu'] = mu\n I_arr.append(calc_I_phi(params,E_vec,phi_vec))\nI_arr = np.array(I_arr)\n# -\n\nXX,YY = np.meshgrid(phi_vec,mu_vec)\nplt.pcolor(XX,YY,I_arr)\nplt.xlabel(r\"$\\phi$\")\nplt.ylabel(r\"$\\mu$\")\nplt.title(\"Current-Phase vs $\\mu$\")\n\nplt.plot(np.sqrt(100e-3 - mu_vec),np.max(I_arr,axis=1))\n\n# ## Study of I-phi with changing flux\n\n# +\nparams = {\n \"N_z\" : 2,\n \"N_phi\" : 5,\n \"flux\" : 0e0,\n \"t_z\" : 100e-3,\n \"Delta_1\" : 1e-3,\n \"Delta_2\" : 1e-3,\n \"mu\" : 10e-3,\n \"kT\" : 1e-5,\n}\nparams[\"t_phi\"] = 1e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\nE_vec = np.linspace(-2e-3,2e-3,100)\nphi_vec = np.linspace(0,2*np.pi,100)\nparams['eta'] = np.abs(E_vec[1] - E_vec[0])\n\nflux_vec = np.linspace(0,4,100)\nI_arr = []\nfor flux in flux_vec:\n print(flux)\n params['flux'] = flux\n params[\"t_phi\"] = 1e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\n I_arr.append(calc_I_phi(params,E_vec,phi_vec))\nI_arr = np.array(I_arr)\n# -\n\nXX,YY = np.meshgrid(phi_vec,flux_vec)\nplt.pcolor(XX,YY,I_arr)\nplt.xlabel(r\"$\\phi$\")\nplt.ylabel(r\"$flux$\")\nplt.title(\"Current-Phase vs flux$\")\n\nplt.plot(phi_vec,I_arr[10])\n\nplt.plot(flux_vec,np.max(I_arr,axis=1))\nplt.ylabel(\"Critical Current\")\nplt.xlabel(\"flux\")\n\n# # Notes\n# - I am unsure how to incorporate the flux term in the diagonal element.\n# - The spectrum is too slow to compute with finite-cross section, this looks like a design flaw\n# - Are $l \\neq 0$ states different in energies from $l = 0$ states in the absence of a B field?\n# - How do l states evolve as a function of $\\mu$? More importantly, what the right chemical potential so that only $l = 0,1$ are populated?\n# - The current phase relationship does not change as a function of $\\mu$. Since adding a B-field to a 1D nanowire only chnages $\\mu$, I do not expect to see any change as a function of B.\n# \n# **Observation** : Low $\\mu$ is hard to converge. I don't know why?\n\n# # Intersting Asides\n# 1. Visualisation of surface density of states in $\\mu$-$\\Delta$ space. I see that for $t \\gg \\mu \\gg \\Delta$, the dos outside the gap is almost flat. \n\n# +\nparams = {\n \"N_z\" : 10,\n \"N_phi\" : 1,\n \"flux\" : 0e0,\n \"t_z\" : 1000e-3,\n \"Delta\" : 1e-3,\n \"mu\" : 100e-3,\n}\n\nparams[\"t_phi\"] = 0e-3*np.exp(1j*2*np.pi*params[\"flux\"]/params[\"N_phi\"])\n\nE = np.linspace(-10e-3,10e-3,1000)\neta = 0.1*np.abs(E[1]-E[0])\n\nmu_vec = np.linspace(0,100e-3,25) \ndata = []\nfor mu in mu_vec:\n params['mu'] = mu\n \n H = calc_hamiltonian(params)\n surface_dof = 2*params['N_phi']\n alpha = H[:surface_dof,:surface_dof]\n beta = H[:surface_dof,surface_dof:2*surface_dof]\n \n g = calc_surface_g(E,alpha,beta,eta)\n a = 1j*(g - np.conj(np.transpose(g,(0,2,1))))\n dos = np.real(np.trace(a,axis1=-1,axis2=-2))\n\n data.append(dos)\ndata = np.array(data)\n# -\n\nXX,YY = np.meshgrid(E,mu_vec)\nplt.pcolor(XX,YY,data)\nplt.xlabel(\"Energy\")\nplt.ylabel(\"Chemical potential\")\n","repo_name":"sandeshkalantre/orbital-josephson-negf","sub_path":"code/Cylindrical Nanowire Hamiltonian Spectrum.ipynb","file_name":"Cylindrical Nanowire Hamiltonian Spectrum.ipynb","file_ext":"py","file_size_in_byte":14558,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"39972034625","text":"# # Step 3. Importing libraries and dataframes\n\n# +\n#Importing libraries\n\nimport pandas as pd\nimport numpy as np\nimport os\n\n# +\n#Import the dataframe 'customers'\n#Create a path\n\npath = r'/Users/alexbor/Desktop/Data Analytics/Immersion/Submissions/Instacart Basket Analysis'\n\n#Import the df \n\ncust_df = pd.read_csv(os.path.join(path, '02 Data ', 'Original data', 'customers.csv'))\n# -\n\n# # Step 4. Exploring and wrangling the data\n\n# # 4.1. Exploring the data\n#\n\n# +\n#Wrangling the data\n#Check the head\n\ncust_df.head()\n\n# +\n#Check the tail\n\ncust_df.tail()\n\n# +\n#Check the shape \n\ncust_df.shape\n\n# +\n#Check the info\n\ncust_df.info()\n# -\n\n# **Note: I can already see here that there are problems with the column 'First Name'. I should keep it in mind for the next steps**\n\n# +\n#Describe the numerical data \n\ncust_df.describe()\n# -\n\n# **Note: I can already see here that the income data values have a very wide scope. I should keep it in mind for now**\n\n# # 4.2. Wrangling data\n\n# +\n#I decided to rename the 'Surnam' column to 'Last name' and the column 'n_dependance' to 'Number of dependants'\n\ncust_df.rename(columns = {'Surnam':'Last Name'}, inplace = True)\n# -\n\ncust_df.rename(columns = {'n_dependants':'Number of dependants'}, inplace = True)\n\n#Checking the head\ncust_df.head()\n\n# # Step 5. Missing values, duplicates and data types\n\n# # 5.1. Missing values\n\n# +\n#Check for missing values\n\ncust_df.isnull().sum()\n# -\n\n#Investigate the column in detail\ncust_df['First Name'].value_counts(dropna = False)\n\n# **There are 11259 missing first names, but I don't think it's relevant to do something about it. It also consitutes only 5% of the data**\n\n# # 5.2. Data types\n\n# +\n#Checking for data types\n\ncust_df.dtypes\n\n# +\n#I decided to change the data type of user_id to string. \n\ncust_df['user_id'] = cust_df['user_id'].astype('str')\n\n# +\n#Checking the result\n\ncust_df['user_id'].dtypes\n# -\n\n# # 5.3. Mixed-type data\n\n# +\n#Checking for mixed-type data \n\nfor col in cust_df.columns.tolist():\n weird = (cust_df[[col]].applymap(type) != cust_df[[col]].iloc[0].apply(type)).any(axis=1)\n if len (cust_df[weird]) > 0:\n print (col)\n# -\n\n# **I suppose the 'First Name' column has mixed-type data because there are NaNs there. I decided to change the data type to string**\n\n# +\n#Change the data type of the column 'First Name'\n\ncust_df['First Name'] = cust_df['First Name'].astype('str')\n\n# +\n#Checking the result\n\ncust_df['First Name'].dtypes\n\n# +\n#Checking again for mixed-type data \n\nfor col in cust_df.columns.tolist():\n weird = (cust_df[[col]].applymap(type) != cust_df[[col]].iloc[0].apply(type)).any(axis=1)\n if len (cust_df[weird]) > 0:\n print (col)\n# -\n\n# # 5.4. Duplicates\n\n# +\n#Finding duplicates \n\ndf_dupl = cust_df[cust_df.duplicated()]\n# -\n\ndf_dupl.shape\n\n# **There are no duplicates**\n\n# # Step 6. Merging the 'cust_df' df with the df from 4.8.\n\n# +\n#Importing the df 'ords_prods_merged'\n\nords_prods_merge = pd.read_pickle(os.path.join(path, '02 Data ', 'Prepared data', 'orders_products_merged_finall_with flags_4_8.pkl'))\n\n# +\n#Checking the data type of 'user_id' to make sure I can merge this df with 'cust_df'\n\nords_prods_merge['user_id'].dtypes\n\n# +\n#Changing the data type of 'user_id' in 'ords_prods_merge'\n\nords_prods_merge['user_id'] = ords_prods_merge['user_id'].astype('str')\n\n# -\n\nords_prods_merge['user_id'].dtypes\n\n# +\n#Drop the '_merge' column from 'ords_prods_merge'\n\nords_prods_merge = ords_prods_merge.drop(columns = ['_merge'])\n\n# +\n#Merging two dataframes \n\nords_prods_cust_merge = cust_df.merge(ords_prods_merge, on = 'user_id', indicator=True)\n\n# +\n#Check the new df info\n\nords_prods_cust_merge.info()\n# -\n\n# # Step 8. Exporting the new df \n\nords_prods_cust_merge.to_pickle(os.path.join(path, '02 Data ', 'Prepared data', 'ords_prods_cust_merge.pkl'))\n","repo_name":"Shurnaja/PYTHON_INSTACART_PROJECT","sub_path":"Scripts Instacart/Visualizations Part 1.ipynb","file_name":"Visualizations Part 1.ipynb","file_ext":"py","file_size_in_byte":3801,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"37343789036","text":"# +\nfrom IPython.core.display import display, HTML\ndisplay(HTML(\"\"))\n# %matplotlib inline\nimport matplotlib\nimport matplotlib.pyplot as plt\nimport matplotlib.gridspec as gridspec\nimport seaborn as sns\nimport os\n\nimport numpy as np\nimport pandas as pd\nimport tensorflow as tf\n\nimport atecml.data\n\nfrom contextlib import contextmanager\nfrom tqdm import tqdm\nfrom sklearn.neighbors import KNeighborsClassifier\nfrom sklearn.preprocessing import StandardScaler, MinMaxScaler\nfrom imblearn.over_sampling import SMOTE, ADASYN\nfrom sklearn.externals import joblib\n\nplt.style.use('ggplot')\n\n# +\n#Loading Data....\ndata = atecml.data.load_train()\ntrain_df = atecml.data.filter_date(data,start_date='2017-09-05',end_date='2017-10-15')\nval_df = atecml.data.filter_date(data,start_date='2017-10-16',end_date='2018-10-15')\n\npredictors = [x for x in train_df.columns if x not in atecml.data.NOT_FEATURE_SUM]\n\n#Loading Data....\ndata = atecml.data.load_train()\npredictors = [x for x in data.columns if x not in atecml.data.NOT_FEATURE_SUM]\n\n\nfeature_tree_num = 20\ntrain_model =[]\ntarget_list = ['Normal','Fraud']\n#target_list = ['Normal']\n#target_list = ['Fraud']\n\nfor idx in range(0,1):\n for item in ['dart','gbdt']:\n for feature_grp_idx in range(0,feature_tree_num):\n for target in target_list:\n train_id = item + '__'+target +'__'+str(idx) +'__' + str(feature_grp_idx)\n train_model.append(train_id)\n\n \ndef model_load(model_name):\n model_cache_name = './'+model_name+'.model'\n if (os.path.exists(model_cache_name)):\n clf = joblib.load(model_cache_name)\n else:\n print('ERROR...............')\n return clf\n\n\nif (os.path.exists('./feature_list.dat')):\n print('Load Feature List from persistant store...')\n feature_list = joblib.load('./feature_list.dat')\nelse:\n print('Error: Feature Cache File missing...')\n\n\ntrained_model_list =[]\nwith atecml.data.timer('Classification: Model Training'):\n for train_id in tqdm(range(len(train_model))):\n fit_model = model_load(train_model[train_id])\n trained_model_list.append(fit_model)\n\n\n# -\n\ndef predict_proba_matrix(trained_model_list,df):\n low_band = 0.4\n high_band = 0.9\n data = df.reset_index(drop=True)\n verify_df =pd.DataFrame()\n with atecml.data.timer('Validation.....'):\n for idx in tqdm(range(len(trained_model_list))):\n clf = trained_model_list[idx]\n model_params = train_model[idx].split('__')\n f_idx = int(model_params[3])\n select_feature = feature_list[f_idx]\n y_predict = clf.predict(data[select_feature])\n if (model_params[1] == 'Fraud'):\n verify_df['n'+str(idx)] = abs(y_predict)\n else:\n verify_df['n'+str(idx)] = abs(1- y_predict)\n verify_df['mean']=verify_df.T.mean()\n result = pd.concat([verify_df,data['id']],axis=1)\n filter_id_list = list(result[(result['mean'] > low_band) & (result['mean'] < high_band)]['id'])\n residual_df = data[data['id'].isin(filter_id_list)].reset_index(drop=True)\n return result,residual_df\n\n\nverify_df = atecml.data.load_train()\nverify_data = atecml.data.filter_date(verify_df,start_date='2017-10-16',end_date='2018-10-15')\nresult,res = predict_proba_matrix(trained_model_list,verify_df)\n\na,b,c=atecml.data.accuracy_validation(verify_df['Fraud'],result['mean'])\n\n# +\nlow_band=0.5\nhigh_band = 0.9\n\nfilter_id_list = list(result[(result['mean'] > low_band) & (result['mean'] < high_band)]['id'])\nprint(len(filter_id_list))\n\n\ntemp_df = verify_df[~verify_df['id'].isin(filter_id_list)].reset_index(drop=True)\ntemp_df_proba = result[~result['id'].isin(filter_id_list)].reset_index(drop=True)\n\nprint(len(temp_df[temp_df['Fraud']==1]['id']))\na,b,c=atecml.data.accuracy_validation(temp_df['Fraud'],temp_df_proba['mean'])\n# -\n\nres_df = verify_df[verify_df['id'].isin(filter_id_list)].reset_index(drop=True)\nres_df_proba = result[result['id'].isin(filter_id_list)].reset_index(drop=True)\nd,e,f=atecml.data.accuracy_validation(res_df['Fraud'],res_df_proba['mean'])\n\npd.concat([temp_df,res_df],axis=1)\n\n\n\n\n\n\n\n\n\n\n\n# +\n\n\nres_train = pd.concat([result[result['id'].isin(list(res['id']))].reset_index(drop=True),res],axis=1)\nres_train.to_pickle('./res.dat')\n# -\n\nresult.columns\n\n\n\n\n","repo_name":"zartbot/anti_fraud","sub_path":"model/model_backup/Residual_Tree/Result_Analytics.ipynb","file_name":"Result_Analytics.ipynb","file_ext":"py","file_size_in_byte":4353,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"23640022427","text":"# # azimuth venues\n\n# +\nimport os\nimport supabase\nimport pandas as pd\nfrom postgrest import APIError\nfrom supabase import create_client, Client\n\nsupabase_url: str = os.getenv(\"SUPABASE_URL\")\nsupabase_key: str = os.getenv(\"SUPABASE_KEY\")\nsupabase_client: Client = create_client(SUPABASE_URL, SUPABASE_KEY)\n# -\n\n# ---------------------------------------------------------------\n# venues from seatgeek database\n\nresponse = supabase_client.table('seatgeek_data').select(\"*\").execute()\ndef seatGeek(response):\n # Create a DataFrame from the API response data\n df = pd.DataFrame(response.data)\n\n # Reset the index\n df = df.reset_index(drop=True)\n\n # Remove duplicates based on 'venue.name' column\n venue = df[~df['venue.name'].duplicated()]\n\n # Get the columns of interest\n cols = list(venue.columns)\n data = venue[cols[27:-12]]\n\n return data\n\n\n# +\n# seatGeek(response)\n# -\n\n# ----------------------------------------------------------\n# _venues from ticketmaster database\n\nresponse2 = supabase_client.table('ticketmasterdata').select(\"*\").execute()\ndef ticketMaster(response):\n # Create a DataFrame from the API response data\n tkMaster_df = pd.DataFrame(response.data)\n\n # Reset the index\n tkMaster_df = tkMaster_df.reset_index(drop=True)\n\n # Select the columns of interest\n tkMaster_df = tkMaster_df.iloc[:, 17:-7]\n\n # Remove duplicates based on 'Venue Name' column\n tkMaster_df = tkMaster_df[~tkMaster_df['Venue Name'].duplicated()]\n\n return tkMaster_df\n\n\n\n# +\n# ticketMaster(response2)\n# -\n\n# ---------------------------------------------------------------------\n# Venues.bson file from crankDB\n\ndef processVenuesData(filename):\n # Read the CSV file into a DataFrame\n venues = pd.read_csv(filename)\n\n # Remove duplicates based on 'Name' column\n venues = venues[~venues['Name'].duplicated()]\n\n # Drop unnecessary columns\n venues.drop(['MetroArea._id', '_id'], axis=1, inplace=True)\n\n return venues\n\n\n\n# +\n# processVenuesData('venues.csv')\n# -\n\n# ----------------------------------------------------\n# chceking if there exist similarity between the unique values in tkmaster and crankdb\n# \n# --------------------------------------------\n#\n# checking if the values in ticketmaster db exists in seatgeek\n\n# +\ncommon_vanues = processVenuesData('venues.csv')['Name'].isin(seatGeek(response)['venue.name'])\ncrankdb_nique = processVenuesData('venues.csv')[~common_vanues]\nprint(crankdb_nique.shape)\n\ncomon = ticketMaster(response2)['Venue Name'].isin(crankdb_unique['name'])\ntkMaster_unique = ticketMaster(response2)[comon]\nprint(tkMaster_unique.shape)\n# -\n\ncrankdb_unique.shape, seatGeek(response).shape, tkMaster_unique.shape\n\nseatGeek = seatGeek(response)\n\n# +\nimport pandas as pd\n\ndef processSeatGeekData(seatGeek):\n # Rename columns\n cols = list(seatGeek.columns)\n seatGeek.columns = [i.replace('venue.', '').lower() for i in cols]\n\n # Select desired columns\n seatGeek = seatGeek[['name', 'state', 'country', 'postal_code', 'city', 'capacity', 'popularity', 'address', 'url', 'timezone', 'location.lat', 'location.lon']]\n\n # Rename columns\n seatGeek.columns = ['name', 'state', 'country', 'postalCode', 'city', 'capacity', 'popularity', 'address', 'url', 'timezone', 'longitude', 'latitude']\n\n # Add new columns\n seatGeek['description'] = None\n seatGeek['phone'] = ''\n seatGeek['metroareaName'] = None\n\n return seatGeek\n\nseatGeek = processSeatGeekData(seatGeek)\n# -\n\ncrankdb_unique['postal_code'] = None\ncrankdb_unique['timezone'] = None\ncrankdb_unique.columns\n\n# ---------------------------\n#\n# joining the datasets\n# ---------------------\n\nazimuthVenues = pd.concat((seatGeek, crankdb_unique), axis = 0)\n\nazimuthVenues.shape\n\n\n# ----------------------------------------------------------\n#\n# inserting the following data into our database table\n# ----------------------------------------------\n\n# +\ndef loadToDb(azimuthVenues, supabase_client):\n # Replace NaN values with None\n azimuthVenues.replace({pd.np.nan: None}, inplace=True)\n\n # Convert DataFrame to JSON records\n json_data = azimuthVenues.to_dict(orient='records')\n\n # Insert records into the database\n for record in json_data:\n try:\n supabase_client.table('azimuth_venues').insert(record).execute()\n except APIError as e:\n print(\"Error occurred while inserting the record:\")\n print(record)\n print(\"Error message:\", e.message)\n print()\n\nloadToDb(azimuthVenues, supabase_client)\n\n\n# -\n\n# --------------------------------------\n#\n# removing duplicated rows\n# -------------------------------\n\n# +\ndef filterDups(supabase_client):\n response = supabase_client.table('azimuth_venues').select('*').execute()\n data = response.data\n\n # Create a set to store unique IDs\n unique_ids = set()\n\n # List to store rows to be deleted\n rows_to_delete = []\n\n # Iterate through each row and filter duplicates based on 'name'\n for row in data:\n row_name = row['name']\n if row_name not in unique_ids:\n unique_ids.add(row_name)\n else:\n rows_to_delete.append(row_name)\n\n # Delete the duplicate rows from the table\n for row_name in rows_to_delete:\n # Retrieve the first occurrence of the duplicate row\n first_occurrence = next(row for row in data if row['name'] == row_name)\n # Delete all other occurrences of the duplicate row\n supabase_client.table('azimuth_venues').delete().match({'name': row_name}).execute()\n # Insert the first occurrence back into the table\n supabase_client.table('azimuth_venues').insert(first_occurrence).execute()\n\n print(\"Duplicate rows retained and others deleted successfully.\")\n\nfilterDups(supabase_client)\n","repo_name":"C4LEB-ai/Azimuth","sub_path":"azimuth_venues.ipynb","file_name":"azimuth_venues.ipynb","file_ext":"py","file_size_in_byte":5825,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"46"}
+{"seq_id":"74123462871","text":"import logging\nlogging.basicConfig()\nlog = logging.getLogger()\nlog.setLevel(logging.DEBUG)\n\n# +\n# from pymodbus.client.sync import ModbusTcpClient\nimport datetime\n\nclient = ModbusTcpClient('192.168.11.150', port=502, retries=3, timeout=3)\nclient.connect()\ntime = datetime.datetime.now()\n\ntry:\n rr = client.read_holding_registers(address=0,count=103,unit=1)\n #rr = client.read_holding_registers(0x0,10,unit=1)\n time = datetime.datetime.now()\nexcept ConnectionException:\n print('connection error')\nclient.close()\n\n\nprint(time)\nassert(rr.function_code < 0x80), \"register error\"\nprint(rr)\nprint(rr.registers)\n#print(client.read_holding_registers(0x9D41,10,unit=1))\n\n\n# -\n\n\n\nprint(rr)\nprint(rr.registers)\n\n# +\nfrom pymodbus.client.sync import ModbusTcpClient\n\nclient = ModbusTcpClient('192.168.11.51', port=502, retries=3, timeout=3)\nclient.connect()\n\nrr = client.read_holding_registers(address=0,count=10,unit=1)\n\nassert(rr.function_code < 0x80)\nprint(rr)\nprint(rr.registers)\n\nclient.close()\n# -\n\n\n\n# +\nfrom pymodbus.client.sync import ModbusTcpClient\nfrom pymodbus.exceptions import ConnectionException\n\nfrom time import sleep\n\nclient = ModbusTcpClient('192.168.11.150', port=502, retries=3, timeout=3)\nclient.connect()\n\n \naddress = 0\nbulk = 1\nskip = []\nwhile True:\n \n print('address {} - {}'.format(address, address+bulk-1))\n \n try:\n if not address in skip:\n rr = client.read_holding_registers(address=address,count=bulk,unit=1)\n else:\n print('address {} skipped'.format(address))\n continue\n\n \n except AssertionError:\n print('Error: assertion')\n client.close()\n sleep(0.3)\n client.connect()\n except ConnectionException:\n print('Error: connection')\n client.close()\n sleep(0.3)\n client.connect()\n else:\n address += bulk\n assert(rr.function_code < 0x80)\n print('read', rr)\n print(rr.registers)\n \n sleep(0.423)\n# -\n\n0-102\n","repo_name":"ericyeh1995/modbusApp","sub_path":"notebooks/aus test.ipynb","file_name":"aus test.ipynb","file_ext":"py","file_size_in_byte":1992,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"41229714601","text":"# +\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport matlab.engine # to run Cheng's code\nimport os\n\n# add the path to my packages to system paths so they can be imported\nimport sys\nsys.path.append('/home/yasamanparhizkar/Documents/yorku/01_thesis/code/my_packages')\n# sys.path.append('F:\\MAScThesis\\code\\my_packages')\n# sys.path.append('/home/yasamanparhizkar/Documents/thesis/code/my_packages')\n\nimport data_handler_03 as dh\nimport my_simgraph_06 as sg\nimport assess_simgraph_03_cheng as asg\n# -\n\n# # Load spike data\n\n# Spike data shape: (297, 1141, 113) $\\implies$ (movie repeats, frames/time, neurons)\n#
\n# Labels are 1 (= spike) or -1 (= no spike).\n\n# +\n# load all spike data from file\nspikes_dp = '../../../data/original_files/spikes.csv'\nbinned_data = np.loadtxt(spikes_dp, delimiter=',')\nbinned_data = binned_data.reshape(binned_data.shape[0], 1141, 113)\nbinned_data = binned_data * 2 - 1 # turn labels from 0,1 to -1,1\n\nI_order_10 = [54, 35, 10, 60, 74, 9, 61, 56, 91, 104]\n# -\n\n# ## Group all 113 neurons\n#\n# This will create a more balanced dataset which is presumabley easier to solve.\n#
\n# Grouped data shape: (297, 1141, 1) $\\implies$ (movie repeats, frames/time, group)\n\n# group all neurons together\ngrouped_data = np.zeros((297, 1141, 1))\nfor trial in range(297):\n for frame in range(1141):\n grouped_data[trial, frame, :] = 2 * int((binned_data[trial, frame, :] == 1).any()) - 1\n\n# +\n# print some statistics\nprint('grouped_data.shape = ', grouped_data.shape)\n\navg_spike_perc = 0\nprint('trial # | percentage belonging to class 1')\nprint('---------------------------------------------')\nfor trial in range(10):\n pers = dh.class_percentages(grouped_data[trial, :, :].reshape(-1), [-1, 1])\n avg_spike_perc += pers[1]\n print('trial #{:3} | {:.2f} %'.format(trial, pers[1]))\n\navg_spike_perc /= 10\nprint('---------------------------------------------')\nprint('AVERAGE | {:.2f} %'.format(avg_spike_perc))\n\ntotal_perc = np.sum(grouped_data == 1) *100 /(grouped_data.shape[0] * grouped_data.shape[1])\nprint('---------------------------------------------')\nprint('{:.2f} % of the whole data belongs to class 1.'.format(total_perc))\n\n\n# -\n\n# # Assess the model's performance with random tests\n\n# +\ndef transform(fv):\n \"\"\"\n Transform to be applied on feature vectors.\n \n Input: fv\n fv - 1xDf torch tensor representing a feature vector\n \n Output: fvv\n fvv - 1xDf' torch tensor representing the transformed feature vector\n \"\"\"\n \n # for faster run and less memory usage\n fvv = fv[:, ::10]\n \n # for numerical stability during GD\n # fvv = fvv * 10\n \n return fvv\n\n# data retrieval params\ndata_params = {'func': dh.datapoint_torch, 'lbl_func': dh.get_labels, 'features_dp': '../../../data/features/slowfast/slowfast_4732/', \\\n 'spike_data': grouped_data, 'group_id': 0, 'transform': transform, 'ind_min': 1*1141+0, 'ind_max': 2*1141-1}\n\n# graph construction and penalty term parameters\nsg_params = {'mu': 30, 'Dt': None, 'Dv':0, 'Dvt':2000, \\\n 'cnstr_method_tt': 'time', 'cnstr_method_vv': 'time', 'cnstr_method_vt': 'time',\\\n 'train_t': None, 'val_t': None, \\\n 'edges_tt':None, 'edges_vv':None, 'edges_vt':None, }\n\n# gradient descent parameters\nsg_opt_params = { 'epsilon0':1, 'epsilon_decay':0.5, 'epsilon_jump': 2, \\\n 'num_its':16, 'check_freq':1, 'print_checks':False, 'Theta0':None, \\\n 'force_all_its': True, 'threshold': 0.01}\n\n# randomization parameters\nrnd_params = {'train_sizes': [50, 100, 150, 200, 250, 300, 350], 'val_sizes': [30], 'train_its': 5, 'val_its': 10, 'seed': None}\n# rnd_params = {'train_sizes': [400], 'val_sizes': [10], 'train_its': 2, 'val_its': 3, 'seed': None}\n\n# parameters to visualize the optimized M\n# f_sz = 474 # must match data_params\nf_sz = int(np.ceil(4732/10)) # must match data_params\nxloc = np.broadcast_to(np.arange(f_sz), (f_sz, f_sz))\nyloc = xloc.T\nfig_params = {'rmark_th': 30, 'f_sz': f_sz, 'xloc': xloc, 'yloc': yloc}\n\n# start a matlab engine to run Cheng's code\neng = matlab.engine.start_matlab()\neng.cd(r'/home/yasamanparhizkar/Documents/yorku/01_thesis/code/15_cheng', nargout=0)\n\n# path to save the results\nres_path_sg = '../../../data/experiments/comparison/temp/sg/'\nres_path_lgrg = '../../../data/experiments/comparison/temp/lgrg/'\n\n# +\n# import importlib\n# importlib.reload(sg)\n# importlib.reload(asg)\n# -\n\n# times will be measured together and written in res_path_sg\nval_num_res_sg, val_num_err_sg, val_num_res_lgrg, val_num_err_lgrg = asg.assess_sg_model(eng, data_params, sg_params, sg_opt_params, rnd_params, fig_params, res_path_sg, res_path_lgrg)\n\nasg.plot_curves(rnd_params, sg_params, res_path_sg)\n\nasg.plot_curves_without_runtime(rnd_params, sg_params, res_path_lgrg)\n\n# close the matlab engine when you're done\neng.quit()\n\n# # Plot similarity graph and logistic regression validation accuracies together\n\n# +\n# unpack params\ntrain_sizes = rnd_params['train_sizes']\nval_sizes = rnd_params['val_sizes']\ntrain_its = rnd_params['train_its']\nval_its = rnd_params['val_its']\nassess_qs = ['min_acc', 'val_acc', 'missed', 'false_alarm']\n\n# read sg training curves\ncurves_sg = {}\nerrors_sg = {}\nfor i in range(len(val_sizes)):\n curves_i = np.loadtxt(res_path_sg+'curves/train_'+str(i)+'.txt')\n j = 0\n for quantity in assess_qs:\n if i==0:\n curves_sg[quantity] = curves_i[j::8].reshape((1, -1))\n errors_sg[quantity] = curves_i[j+1::8].reshape((1, -1))\n else:\n curves_sg[quantity] = np.concatenate((curves[quantity], [curves_i[j::8]]), axis=0)\n errors_sg[quantity] = np.concatenate((errors[quantity], [curves_i[j+1::8]]), axis=0)\n j += 2\n \n# read lgrg training curves\ncurves_lgrg = {}\nerrors_lgrg = {}\nfor i in range(len(val_sizes)):\n curves_i = np.loadtxt(res_path_lgrg+'curves/train_'+str(i)+'.txt')\n j = 0\n for quantity in assess_qs:\n if i==0:\n curves_lgrg[quantity] = curves_i[j::8].reshape((1, -1))\n errors_lgrg[quantity] = curves_i[j+1::8].reshape((1, -1))\n else:\n curves_lgrg[quantity] = np.concatenate((curves[quantity], [curves_i[j::8]]), axis=0)\n errors_lgrg[quantity] = np.concatenate((errors[quantity], [curves_i[j+1::8]]), axis=0)\n j += 2\n\n# plot training curves\nplt.figure(figsize=(7,4))\nplt.subplots_adjust(left=0.1,bottom=0.1,right=0.9,top=0.9,wspace=0.8,hspace=0.8)\nfor i in range(len(val_sizes)):\n plt.subplot(len(val_sizes), 1, i+1)\n plt.errorbar(train_sizes, curves_sg['val_acc'][i], errors_sg['val_acc'][i])\n plt.errorbar(train_sizes, curves_lgrg['val_acc'][i], errors_lgrg['val_acc'][i])\n plt.legend(['sg', 'lgrg'])\n plt.xlabel('training set size')\n plt.ylabel('{} val repeats x {} train repeats'.format(val_its, train_its))\n _ = plt.title('val. set size = {}, Dt = {}, Dvt = {}, Dv = {}'.format(val_sizes[i], sg_params['Dt'], sg_params['Dvt'], sg_params['Dv']))\n# -\n\n\n\n\n\n\n\n\n","repo_name":"yasamanparhizkar/simgraph","sub_path":"code/07_slowfast/02_slowfast_sg_assess/05_sg_slowfast_cheng_assess.ipynb","file_name":"05_sg_slowfast_cheng_assess.ipynb","file_ext":"py","file_size_in_byte":7003,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"75074544151","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#  \n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"VxwTX1XMulIr\" outputId=\"00a75993-8c88-4f12-b425-a1ba508eb58a\"\n# for / for / if\nfor i in range(3): # 0 1 2\n for j in range(5): # 0 1 2 3 4\n if j >=3: \n print(i+j)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"VS3t2YBSu1Ja\" outputId=\"bba7d2a4-c815-4c89-f09d-d1d07494e12b\"\n# loop list\nIO = ['eb', 'skr', 'hw', 'yn', 'cy', 'cw', 'mj', 'ybk', 'htm', 'yr', 'yj', 'wy']\nKM = [i for i in IO if len(i) == 2]\nKM\n","repo_name":"jh0118/class2021SPRING","sub_path":"20210317.ipynb","file_name":"20210317.ipynb","file_ext":"py","file_size_in_byte":752,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"5822826413","text":"# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# http://seananderson.ca/2013/10/19/reshape.html\n# -\n\n# # Tabular Data Gym 2. Reshaping.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport matplotlib.style\nmatplotlib.style.use('ggplot')\n\n# %matplotlib inline\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# %load_ext rpy2.ipython\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# library(magrittr)\n# library(ggplot2)\n# library(dplyr)\n# library(reshape2)\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# names(airquality) <- tolower(names(airquality))\n# head(airquality)\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# aq_pd = %R airquality\naq_pd.head()\n\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# Since `rpy2` does not treat NA well, we need to change it manually to `np.nan`.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\ndef fix_nans_in_df(colname, df):\n df.ix[df[colname] < 0, colname] = np.NaN \n\nfor colname in aq_pd.columns.values:\n fix_nans_in_df(colname, aq_pd)\n\naq_pd.head()\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# ## Melt/stack: changing wide format into long\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# R function `melt` from `reshape2` package change wide fromat into long. \n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n#\n# aql <- melt(airquality)\n# aql %>% head\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# ### Question 1\n#\n# One can get similar result using pandas' method `melt`. \n# -\n\npd.melt(aq_pd)\n\n# ### Question 2\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# airquality %>%\n# melt(id.vars = c(\"month\", \"day\")) %>%\n# head()\n# -\n\n# Now let's try to get it with pandas.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\npd.melt(aq_pd, id_vars = ['month','day'])\n# -\n\n# ## Question 3 Putting names to columns\n\n# + language=\"R\"\n#\n# aql <-\n# melt(airquality, id.vars = c(\"month\", \"day\"),\n# variable.name = \"climate_variable\", \n# value.name = \"climate_value\") %>%\n# mutate(date=as.Date(paste0(\"2015-\",month,'-',day)))\n# head(aql)\n# -\n\n# Your turn with pandas.\n\npd.melt(aq_pd, id_vars = ['month','day'], var_name = 'climate_variable', value_name= 'climate_value')\n\n# ### Comments about ploting\n#\n# When one uses ggplot it is natural to use long format when ploting.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# aql %>%\n# ggplot() +\n# geom_line(aes(date, climate_value, colour=climate_variable))\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# But with pandas' mathplotlib interface it is easier to use wild format.\n#\n# First let's create dates.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\naq_pd2= aq_pd.copy()\n\naq_pd2['date'] = \\\n [pd.Timestamp(s) \n for s in ('2015-'+aq_pd['month'].astype(int).astype(str)+'-'\n +aq_pd['day'].astype(int).astype(str))]\n# -\n\n# Then selecting columns and adding aproprite index one get: \n\naq_pd2_indexed = aq_pd2[['date', 'ozone', 'solar.r', 'wind', 'temp']]\naq_pd2_indexed.set_index('date')\naq_pd2_indexed.plot()\n\n# ## Long- to wide-format data\n\n# + language=\"R\"\n# aql <- melt(airquality, id.vars = c(\"month\", \"day\"))\n# head(aql)\n# -\n\n# Can you do the same with pandas and then create plot?\n\naq_pd2.plot()\n\n# ### Question 4. Time for dcast\n\n# + language=\"R\"\n# aqw <- dcast(aql, month + day ~ variable)\n# head(aqw)\n# -\n\n# The approprite pandas method seems to be `pivot_table`. Can you try?\n\naq_long = pd.melt(aq_pd, id_vars = ['month','day'])\naq_long.head()\n\naq_wide = pd.pivot_table(aq_long, index = ['month','day'],columns = 'variable')\naq_wide.head()\n\naq_wide.reset_index()\n\n# ### Question 5. Melting with stack\n#\n# There is another pandas' method for maelting. It is called stack. Can you try it with `aq_pd`? What is the type of the result? Can you make it to be DataFrame with comlumns `varaiable` and `value`?\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n\n# -\n\n# ### Question 6. Melt, filter, group, agregate.\n\n# + language=\"R\"\n# aql %>% head\n\n# + language=\"R\"\n# airquality %>%\n# melt(id.vars = c(\"month\", \"day\")) %>%\n# filter(!is.na(value)) %>%\n# group_by(variable) %>%\n# summarize(total= sum(value)) \n\n# + language=\"R\"\n# library(data.table)\n# data.table(airquality) %>%\n# melt(id.vars = c(\"month\", \"day\")) %>%\n# .[!is.na(value), .(total=sum(value)), by=variable]\n# -\n\n# Time for doing this with pands:\n\n\n","repo_name":"DataGym/tabularDataGym2","sub_path":"luca/iteration_01.ipynb","file_name":"iteration_01.ipynb","file_ext":"py","file_size_in_byte":4706,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"14711500924","text":"from keras.datasets import mnist\nfrom keras.models import Sequential, Model, load_model\nfrom keras.utils import to_categorical\nfrom keras.layers import Dense, Flatten, Input, BatchNormalization, Dropout\nfrom keras import optimizers\nfrom keras import regularizers\nfrom keras.callbacks import EarlyStopping, ModelCheckpoint\n\nimport numpy as np\nimport matplotlib.pyplot as plt\n\n# ### Importation de la base de données MNIST (ce sont des images de chiffres manuscrits, hello world du deep learning)\n\n(x_train, y_train), (x_test, y_test) = mnist.load_data()\n\nx_train.shape\n\n# ### Affichage des chiffres\n\n# +\nr = np.random.randint(x_train.shape[0])\n\nplt.imshow(x_train[r],cmap = \"hot\")\n# -\n\n# ### Normalisation des données\n\n# +\nx_train = x_train/255\n\nx_test = x_test/255\n# -\n\n# ### Passage des données (0,1,2,3...) en catégories. Exemple : 3 devient (0,0,0,1,0,0,0,0,0,0)\n\n# +\ny_train_cat = to_categorical(y_train,num_classes = 10)\n\ny_test_cat = to_categorical(y_test,num_classes = 10)\n# -\n\n# ### Construction du réseau\n\nmodel = Sequential()\nmodel.add(Flatten(input_shape = (28,28))) #Passage de tableau 2D à 1D\n\nmodel.add(Dense(128,activation = \"relu\", kernel_initializer = \"he_normal\",kernel_regularizer = regularizers.l2(0.01)))\nmodel.add(Dropout(0.1))\nmodel.add(BatchNormalization())\nmodel.add(Dense(128,activation = \"relu\", kernel_initializer = \"he_normal\",kernel_regularizer = regularizers.l2(0.01)))\nmodel.add(Dropout(0.1))\nmodel.add(BatchNormalization())\nmodel.add(Dense(10,activation = \"softmax\", kernel_initializer = \"he_normal\",kernel_regularizer = regularizers.l2(0.01)))\n\n# ### Optimizer et loss function\n\nopt = optimizers.SGD(lr = 0.01)\n\nmodel.compile(loss = \"binary_crossentropy\",optimizer = opt, metrics = ['categorical_accuracy'])\n\n# ### Affichage de l'architecture du réseau \n\nprint(model.summary())\n\n# ### Mise en place du Early Stopping\n\nearly_stopping = EarlyStopping(monitor = \"val_loss\",patience = 10,restore_best_weights=True)\n\n\n# ### Checkpointer pour sauvegarder le meilleur modèle\n\ncheckpointer = ModelCheckpoint(filepath = \"model_exemple_nist_best.h5\",verbose = 1,save_best_only = True)\n\n# ### Apprentissage\n\nhist = model.fit(x_train,y_train_cat,epochs = 10,batch_size = 128,validation_split = 0.1,callbacks = [early_stopping,checkpointer])\n\n# ### Affichage de l'historique de l'apprentissage\n\n# +\nfigure = plt.figure()\n\nax1 = plt.subplot(121)\nax1.plot(hist.history[\"loss\"],label = \"Training Loss\")\nax1.plot(hist.history[\"val_loss\"],label = \"Validation Loss\")\nplt.xlabel(\"Nombre d'itérations\")\nplt.ylabel(\"Loss function\")\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.plot(hist.history[\"categorical_accuracy\"],label = \"Training Accuracy\")\nax2.plot(hist.history[\"val_categorical_accuracy\"],label = \"Validation Accuracy\")\nplt.xlabel(\"Nombre d'itérations\")\nplt.ylabel(\"Accuracy\")\nplt.legend()\n# -\n\n# ### Accuracy sur la base de test\n\n# +\nprediction_bar = model.predict(x_test)\nprediction = np.argmax(prediction_bar,axis = 1)\n\nprint(\"Accuracy on test set: \" + str(np.round(np.sum(prediction == y_test)/x_test.shape[0]*100,3)) + \" %\")\n# -\n\ny_test.shape\n\n# ### Essai sur la base de test\n\n# +\nfigure = plt.figure()\n\nfigure.clear()\n\nr = np.random.randint(x_test.shape[0])\n\nax1 = plt.subplot(121)\nax1.imshow(x_test[r],cmap = \"hot\")\nplt.title(\"Prédiction du réseau : \" + str(prediction[r]) + \"\\n Vraie valeur : \" + str(y_test[r]))\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.bar(np.arange(10),height = prediction_bar[r],tick_label = np.arange(10))\nplt.xlabel(\"Valeur\")\nplt.ylabel(\"Output du réseau\")\nplt.legend()\n# -\n\n# ### Exemple d'erreur\n\n# +\nfigure = plt.figure()\n\nfigure.clear()\n\nerreurs = np.where(prediction != y_test)[0]\n\nr2 = np.random.randint(erreurs.shape[0])\ne = erreurs[r2]\n\nax1 = plt.subplot(121)\nax1.imshow(x_test[e],cmap = \"hot\")\nplt.title(\"Prédiction du réseau : \" + str(prediction[e]) + \"\\n Vraie valeur : \" + str(y_test[e]))\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.bar(np.arange(10),height = prediction_bar[e],tick_label = np.arange(10))\nplt.xlabel(\"Valeur\")\nplt.ylabel(\"Output du réseau\")\nplt.legend()\n# -\n\n# # TEST SUR LA RECONNAISANCE DIMAGE PERSO\n#\n\n# +\nfrom PIL import Image, ImageFilter\n\n# https://stackoverflow.com/questions/35842274/convert-own-image-to-mnists-image\n# https://www.pluralsight.com/guides/importing-image-data-into-numpy-arrays\n\ndef imageprepare(argv):\n \"\"\"\n This function returns the pixel values.\n The imput is a png file location.\n \"\"\"\n im = Image.open(argv).convert('L')\n width = float(im.size[0])\n height = float(im.size[1])\n newImage = Image.new('L', (28, 28), (255)) # creates white canvas of 28x28 pixels\n\n if width > height: # check which dimension is bigger\n # Width is bigger. Width becomes 20 pixels.\n nheight = int(round((20.0 / width * height), 0)) # resize height according to ratio width\n if (nheight == 0): # rare case but minimum is 1 pixel\n nheight = 1\n # resize and sharpen\n img = im.resize((20, nheight), Image.ANTIALIAS).filter(ImageFilter.SHARPEN)\n wtop = int(round(((28 - nheight) / 2), 0)) # calculate horizontal position\n newImage.paste(img, (4, wtop)) # paste resized image on white canvas\n else:\n # Height is bigger. Heigth becomes 20 pixels.\n nwidth = int(round((20.0 / height * width), 0)) # resize width according to ratio height\n if (nwidth == 0): # rare case but minimum is 1 pixel\n nwidth = 1\n # resize and sharpen\n img = im.resize((nwidth, 20), Image.ANTIALIAS).filter(ImageFilter.SHARPEN)\n wleft = int(round(((28 - nwidth) / 2), 0)) # caculate vertical pozition\n newImage.paste(img, (wleft, 4)) # paste resized image on white canvas\n\n # newImage.save(\"sample.png\n\n tv = list(newImage.getdata()) # get pixel values\n\n # normalize pixels to 0 and 1. 0 is pure white, 1 is pure black.\n tva = [(255 - x) * 1.0 / 255.0 for x in tv]\n print(tva)\n return tva\n\n\n# -\n\n# charger image perso \nx=imageprepare('datasets/mnist_test/test_3.png') #file path here\nprint(len(x))# mnist IMAGES are 28x28=784 pixels\n\n# +\n# TRANSFORM X [784,1] into array(1,28,28)\ncol = 28 # just grab the number of columns here\nxtransfinal = []\nxtransfinal.append([x[i:i+col] for i in range(0, len(x), col)])\nxtransfinal = np.array(xtransfinal)\n\n#VISUALIZATION\nplt.imshow(xtransfinal[0],cmap = \"hot\")\nxtransfinal.shape\n# -\n\n# FEEDING INPUT TO MODEL \npredictionbaralx = model.predict(xtransfinal)\npredictionalx = np.argmax(predictionbaralx,axis = 1)\n\n# +\n# RESULT VIZUALISATION\nfigure = plt.figure()\nfigure.clear()\n\n#r = np.random.randint(x_test.shape[0])\nr=0\n\nax1 = plt.subplot(121)\nax1.imshow(xtransfinal[r],cmap = \"hot\")\nplt.title(\"Prédiction du réseau : \" + str(predictionalx[r]))# + \"\\n Vraie valeur : \" + str(y_test[r]))\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.bar(np.arange(10),height = predictionbaralx[r],tick_label = np.arange(10))\nplt.xlabel(\"Valeur\")\nplt.ylabel(\"Output du réseau\")\nplt.legend()\n\n# +\n#import numpy as np\n#xtrans = np.array(x)\n#shape = (28,28)\n#xtrans.reshape(shape)\n#xtrans\n\n# +\n#col = 28 # just grab the number of columns here\n#xtrans2 = [x[i:i+col] for i in range(0, len(x), col)]\n#xtrans2 = np.array(xtrans2)\n#xtrans2.shape\n","repo_name":"Mathonal/MyMLBasics","sub_path":"3_deep_learning/NNET_Mnist_chiffre.ipynb","file_name":"NNET_Mnist_chiffre.ipynb","file_ext":"py","file_size_in_byte":7211,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"6838024158","text":"# # Addition\n\n1 + 2\n\n# +\n# This program adds two numbers\n\nnum1 = 1.5\nnum2 = 6.3\n\n# Add two numbers\nsum = num1 + num2\n# -\n\nprint(sum)\n\n# ## Another way to print\n\n# Display the sum\nprint('The sum of {0} and {1} is {2}'.format(num1, num2, sum))\n\n# # Subtraction\n\n# +\nnum1 = 128\nnum2 = 256\n\nsub = num1 - num2\nprint('{0} - {1} = {2}'.format(num2,num1,sub))\n# -\n\n# ## Multiplication\n\n# +\nnum1 = 12\nnum2 = 2.5\n\npdt = num1*num2\nprint('{0} * {1} = {2}'.format(num1,num2,pdt))\n# -\n\n# # Division\n\n# +\nnum1 = 12\nnum2 = 2.5\nnum3 = 5\n\ndiv1 = num1/num2\nprint(div1)\n\n# floating division\ndiv4 = num1/num3\nprint(div4)\n\n# integer divions\ndiv5 = num1 // num3 # gives the quotient\nprint(div5)\n\n# -\n\n# # Exponentiation\n\n# +\nnum1 = 1.25\nnum2 = 2\nnum3 = -0.5\n\n# Integer power\nexp1 = num1 ** num2\nprint(exp1)\n\n# negative noninteger power\nexp2 = num2 ** num3\nprint(exp2)\n# -\n\n# # Mod or Remainder Operator\n\n# +\nnum1 = 35\nnum2 = 4\nnum3 = 5\n\nrem1 = num1 % num2\nprint(num1, ' % ',num2, ' = ', rem1) # see how we print \n\nrem2 = num1 % num3\nprint(num1, ' % ',num3, ' = ', rem2)\n# -\n\n# # Other mathematical operators\n\n\n","repo_name":"mcodevb/Python-with-Mainul","sub_path":"Mathematical operations in Python.ipynb","file_name":"Mathematical operations in Python.ipynb","file_ext":"py","file_size_in_byte":1089,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"73207704151","text":"# + id=\"h8se9n3yvcR7\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303943922, \"user_tz\": 180, \"elapsed\": 359, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}}\n#pip install control\n##download da biblioteca de controle\n\n# + id=\"is2Ex_8zKocc\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303944312, \"user_tz\": 180, \"elapsed\": 7, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}}\n#bibliotexas para o projeto\nimport control as ctl\nimport matplotlib.pyplot as plt\nimport numpy as np\n\n# + id=\"aTqN3L9jK95N\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303944312, \"user_tz\": 180, \"elapsed\": 6, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}}\n#variaveis de resistencia e capacitancia\nR = 1.;\nC = 1.;\n\n#vetores que serao os polinomios para a funçao de transferencia\nTsim = 10 #vetor de tempo de simulaçao\nnumerador = [1.]\ndenominador = [R*C, 1.]\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"LO5PJFqBNGAk\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303944312, \"user_tz\": 180, \"elapsed\": 6, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}} outputId=\"3802795f-833e-45d5-8ca0-2c0f18015676\"\n#usa os vetores para criar a funçao de transferencia\nH = ctl.tf(numerador, denominador)\nprint('FT malha aberta:', H)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 279} id=\"8pNM7ArFOWdq\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631304221827, \"user_tz\": 180, \"elapsed\": 434, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}} outputId=\"80c251d6-9ff0-4536-c149-73875085e0b5\"\n#calcula a resposta ao degrau\nT, yout = ctl.step_response(H, Tsim) # retorna dois valores a partir dos parametros de funçao de transferencia e tempo\nplt.plot(T, yout, 'b-')\n\n#calcula um degrau unitario\nT2 = np.linspace(-1., 10., 1000)\ndegrau = np.ones_like(T2)\ndegrau[T2<0] = 0\nplt.plot(T2, degrau, 'r-')\n\n#melhorando o grafico\nplt.ylabel('V (V)')\nplt.xlabel('Tempo (s)')\nplt.legend(['resposta ao degrau', 'degrau unitario'])\nplt.grid()\n","repo_name":"davidolivaa/ufabc-ieee-orion-adcs","sub_path":"Controle/01-introducao/01-circuitoRC.ipynb","file_name":"01-circuitoRC.ipynb","file_ext":"py","file_size_in_byte":2585,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"74010120153","text":"import matplotlib.pyplot as plt \nimport numpy as np \nimport pandas as pd\n\n# +\nphase_no=40\n\narr= np.genfromtxt(f'0.1_list_{phase_no}.csv', delimiter=',')\n# -\n\narr[3][1:]\n\n\ndef unpack_phi(phase_no):\n \n arr= np.genfromtxt(f'0.1_list_{phase_no}.csv', delimiter=',')\n \n phi_list= [i if i < np.pi else i-2*np.pi for i in arr[3][1:]%(2*np.pi)]\n \n phi_array= np.reshape(phi_list, (10,5))\n \n return phi_array\n\n\nunpack_phi(10).T\n\nimport seaborn as sns\n\nsns.heatmap(unpack_phi(0), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(10), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(20), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(30), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(40), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\n# +\n#Takes too long to converge to lateral phase\n\nplt.plot(unpack_phi(40)[0]);\n\n# +\n#Behold a little 'cheating' force a symmetric initial condition on both y-axis and run simulation\n\nphi_list= unpack_phi(40)\n\ndef lamb(x_n, y_n, phi_list):\n \n arr= (np.array([i[::-1] for i in phi_list]) + phi_list)/2\n \n return arr[x_n][y_n]\n\n\n# -\n\nplt.plot(((np.array([i[::-1] for i in phi_list]) + phi_list)/2)[0])\n\n# +\nimport fluid_functions as ff\nimport numpy as np\n\nfrom cilia_test_copy import Cilium\nfrom cilia_test_copy import Cilia_System\nfrom cilia_test_copy import big_phi\n\n\nimport matplotlib.pyplot as plt\nfrom itertools import product\n\nimport matplotlib.cm as cm\n\nl_ambda = 0.1\nr0= 0.5\nlattice= 2\na_radii=0.01\n\nf0 = lambda phi : 0\n\ny_n=5\nx_n=10\n\nCilia_System_1= Cilia_System()\n\nnew_constants= np.array([r0, l_ambda ,10])\n\nCilia_row= [Cilium([i*lattice, j*lattice], [0,r0,lamb(i, j, unpack_phi(40))], constants= np.array([r0, l_ambda ,10]), a= a_radii) for i, j in product(range(x_n), range(y_n))]\n\nCilia_System_1.add(Cilia_row)\n\n# -\n\nwhile Cilia_System_1.phi_list[0] < (20*np.pi):\n \n Cilia_System_1.update(stepsize= 0.02)\n\nCilia_System_1.phi_list\n\n# +\narr= Cilia_System_1.phi_list%(2*np.pi)\n\narr= np.array([i if i < np.pi else i-2*np.pi for i in arr])\n\narr=np.reshape(arr, (10,5))\n\nplt.rcParams[\"figure.figsize\"] = (10,5)\n\nplt.plot(np.mean(arr, axis=0)-np.mean(arr, axis=0)[0]);\nplt.grid()\nplt.title('Average Lateral Phase')\nplt.xlabel('y')\nplt.ylabel('Phase');\n\n\n# +\nphis= np.mean(arr, axis=1)[::-1]-np.mean(arr, axis=1)[::-1][-1]\n\nphis= np.array([i if i < np.pi else i-2*np.pi for i in phis])\n\nplt.plot(phis, np.linspace(0, 9, 10))\n\nplt.rcParams[\"figure.figsize\"] = (8,10)\nplt.grid()\nplt.title('Average Streamwise Phase')\nplt.xlabel('Phase')\nplt.ylabel('x');\n# -\n\n\n","repo_name":"Desmondccw/M4R_Project","sub_path":"Figure 9.ipynb","file_name":"Figure 9.ipynb","file_ext":"py","file_size_in_byte":2611,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"18964869739","text":"import glob\nimport pandas as pd\nimport scipy.constants as scc\nfrom scipy.optimize import curve_fit\nfrom astropy.cosmology import FlatLambdaCDM\nimport astropy.units as u\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom astropy.modeling import models, fitting\nimport scipy\nimport scipy.stats as stats\nfrom astropy.coordinates import SkyCoord\nfrom scipy.integrate import quad\nfrom matplotlib.offsetbox import AnchoredText\nimport os\nimport scipy.special as sc\nimport gc\nimport warnings\nimport eazy\nimport time\nimport grizli\nimport numpy as np\nfrom astropy.table import Table\n\n\n# +\ndef Lv(freq, A, beta, Tdust):\n return A * freq**(3+beta) / (np.exp(h * freq / (k_b * Tdust ) ) - 1 )\n\ndef Lv_rescaled(freq, N, A, beta, Tdust): # Lv\n return N * A * freq**(3+beta) / (np.exp(h * freq / (k_b * Tdust ) ) - 1 )\n\ndef T_dust(z):\n return 32.9 + 4.6*(z-2), 2.4+0.35*(z-2) # mean value and dispersion\n\n\n# -\n\nh = 6.62e-27 # erg/s\nk_b = 1.38e-16 # erg/K\nc = 3e10 # speed of light: cm/s\nsigma = 5.67e-5 # stefan boltzmann constant, erg/cm^2/s/K^4\n\npath='/home/yjm/miniconda/Outputs'\nos.chdir(path)\nos.chdir('./a370')\n\nlamb_rest = (np.arange(8, 1200) * 1e-6 ) # micron meters to meters\nfreq_rest = scc.c / lamb_rest # Hz\nfreq_rest_8_1000 = scc.c / (np.arange(8, 1000) * 1e-6 )\nfreq_sorted = np.sort(freq_rest)\nfreq_8_1000_sorted = np.sort(freq_rest_8_1000)\nindex_1000 = np.where(lamb_rest==np.min(abs(freq_sorted-1)))\nkappa0 = 1.3 # cm^2/g --> m^2/kg\nv0 = 6.66e11 #Hz\nM_16=[]\nM_84=[]\nM_50=[]\nbeta_median=[]\nT_median=[]\nL_16 = []\nL_84 = []\nL_UV = []\nL_50 = []\ndetected_fits = glob.glob('*alma_detected.fits')[0]\ntable = Table.read(detected_fits)\nmagnification = np.array(table['magnification'])\nSv_obs = np.array(table['flux_1.2mm'])/magnification*1e-26 # from micron Jy to erg/sec/cm^2\nredshift = np.array([1., 2., 3., 4., 5., 6., 7., 8., 9., 10., 11., 12.])\nlamb_alma = 1200/(1 + redshift)*1e-6 # --> meter\nfreq_alma = scc.c / lamb_alma\nnum_iteration=1000\nbeta = np.random.normal(1.8, 0.2, size=(num_iteration,))\na=2e-26\ncosmo = FlatLambdaCDM(H0=70 * u.km / u.s / u.Mpc, Om0=0.3)\n\nfor j in range(redshift.size):\n d_L = cosmo.luminosity_distance(redshift[j])\n d_L = d_L.to(u.cm) # from parsec to cm\n low, high = T_dust(redshift[j])\n T = np.random.normal(low, high, size=(num_iteration,))\n Lv_rest = 4*scc.pi*d_L**2*Sv_obs[0]/(1+redshift[j]) # unit: erg/s/Hz\n lamb_obs = lamb_rest * (1 + redshift[j])\n M_dust=[]\n t_peak=[]\n l_IR = []\n for i in range(num_iteration): # loop over random T and beta\n N = Lv_rest.value / Lv(freq_alma[j], 2e-26, beta[i], T[i])\n Lv_rescale = N * Lv(freq_rest, 2e-26, beta[i], T[i]) \n Sv_rescale = Lv_rescale * (1 + redshift[j]) / (4 * np.pi * d_L**2)\n nu = 1.5e11 * (1+redshift[j])\n kappa = kappa0 * (nu / v0)**beta[i] # m^2/kg\n # mistake here: Sv_rest and func() have different units\n L = Lv_rest.value / (Lv(freq_alma[j], a, beta[i], T[i]) ) * Lv(nu, a, beta[i], T[i])\n blackbody = 2 * h / c**2 * nu**3 / (np.exp(h*nu / (k_b*T[i])) - 1)\n mass = L / (4 * np.pi * kappa * blackbody ) # kg\n M_dust = np.append(M_dust, mass)\n #L_integrated = np.sum(Lv_rescale[0:-1]) * dx[-1]\n #sol=quad(Lv_rescaled, freq_rest_8_1000[-1], freq_rest_8_1000[0], args=(N, a, beta[i], T[i]))\n #L_integrated = sol[0]\n lamb_peak = scc.c / freq_rest[np.argmax(Lv_rescale)] * 1e6 # meter --> um\n T_peak = 2898 / lamb_peak\n t_peak = np.append(t_peak, T_peak)\n L_integrated = 4 *np.pi * d_L.value**2 * sigma * T_peak**4\n l_IR = np.append(l_IR, L_integrated/3.82e33 ) # in unit of L_sun\n if j==0:\n plt.hist(t_peak)\n m_16 = np.percentile(M_dust, 16, method='nearest')\n M_16 = np.append(M_16, m_16)\n m_84 = np.percentile(M_dust, 84, method='nearest')\n M_84 = np.append(M_84, m_84)\n l_16 = np.percentile(l_IR, 16, method='nearest')\n L_16 = np.append(L_16, l_16)\n l_50 = np.percentile(l_IR, 50, method='nearest')\n L_50 = np.append(L_50, l_50)\n l_84 = np.percentile(l_IR, 84, method='nearest')\n L_84 = np.append(L_84, l_84)\n m_50 = np.percentile(M_dust, 50, method='nearest')\n M_50 = np.append(M_50, m_50)\n\nprint(Lv_rescale)\n\nprint()\n\n# +\n#Mdust_errors = np.vstack([(M_50 - M_16), (M_84 - M_50)])\n\nplt.scatter(redshift, np.log10(M_50))\nplt.scatter(redshift, np.log10(M_16))\nplt.scatter(redshift, np.log10(M_84))\n#plt.errorbar(redshift, np.log10(M_50), yerr=np.log10(Mdust_errors))\n# -\n\n\n\nplt.scatter(redshift, np.log10(L_50))\nplt.scatter(redshift, np.log10(L_16))\nplt.scatter(redshift, np.log10(L_84))\n\n\n","repo_name":"hunyjm12/Master_Project_ALCS","sub_path":"mbb_test.ipynb","file_name":"mbb_test.ipynb","file_ext":"py","file_size_in_byte":4621,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"73226072151","text":"# +\nplanet = {\n 'name': 'Earth',\n 'moons': 1\n}\n#se puede sacar con get\n\nplanet.update({'name': 'Makemake'})\nplanet['name'] = 'Makemake'\nprint(planet)\n\nplanet.update({'name': 'Jupiter', 'moons': 79})\n\nplanet['orbital period'] = 4333\nprint(planet)\n\nplanet.pop('orbital period')\nprint(planet)\n\nplanet['diameter (km)'] = {\n 'polar': 133709,\n 'equatorial': 142984\n}\nprint(f\"{planet['name']} polar diameter: {planet['diameter (km)']['polar']}\")\n\n\n# +\nrainfall = {\n 'october': 3.5,\n 'november': 4.2,\n 'december': 2.1\n}\nfor key in rainfall.keys():\n print(f'{key}: {rainfall[key]}cm')\n\nif 'december' in rainfall:\n rainfall['december'] = rainfall['december'] + 1\nelse:\n rainfall['december'] = 1\nprint(rainfall)\n\ntotal_rainfall = 0\nfor value in rainfall.values():\n total_rainfall = total_rainfall + value\nprint(f'There was {total_rainfall}cm in the last quarter')\n\n# -\n\n# ## Ejercicio 1: Crear y modificar un diccionario de Python.\n\n# Crea un diccionario llamado planet con los datos propuestos\nplanet = {\n 'name': 'Mars',\n 'moons': 2\n}\n# Muestra el nombre del planeta y el número de lunas que tiene.\nprint(f\"{planet['name']}, {planet['moons']}\")\n# Agrega la clave circunferencia con los datos proporcionados previamente\nplanet['circunferencia km'] = {\n 'polar': 6752,\n 'equatorial': 6792\n}\nprint(f\"{planet['name']} has a polar circunference of {planet['circunferencia km']['polar']} km\")\n\n# ## Ejercicio 2: Programación dinámica con diccionarios\n# ## calculo de valores\n\n# +\n# Planets and moons\n\nplanet_moons = {\n 'mercury': 0,\n 'venus': 0,\n 'earth': 1,\n 'mars': 2,\n 'jupiter': 79,\n 'saturn': 82,\n 'uranus': 27,\n 'neptune': 14,\n 'pluto': 5,\n 'haumea': 2,\n 'makemake': 1,\n 'eris': 1\n}\n# Añade el código para determinar el número de lunas.\n\nmoons = planet_moons.values()\nplanets = len(planet_moons.keys())\n\n# Agrega el código para contar el número de lunas. \ntotal_moons = 0\nfor moon in moons:\n total_moons = total_moons + moon\n\n# Calcula el promedio dividiendo el total_moons por el número de planetas\npromedio = total_moons // planets\nprint(promedio)\n \n","repo_name":"CrisGutierrez05/CursoPython","sub_path":"Kata 8.ipynb","file_name":"Kata 8.ipynb","file_ext":"py","file_size_in_byte":2139,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"17148643519","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#
\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"VxwTX1XMulIr\" outputId=\"00a75993-8c88-4f12-b425-a1ba508eb58a\"\n# for / for / if\nfor i in range(3): # 0 1 2\n for j in range(5): # 0 1 2 3 4\n if j >=3: \n print(i+j)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"VS3t2YBSu1Ja\" outputId=\"bba7d2a4-c815-4c89-f09d-d1d07494e12b\"\n# loop list\nIO = ['eb', 'skr', 'hw', 'yn', 'cy', 'cw', 'mj', 'ybk', 'htm', 'yr', 'yj', 'wy']\nKM = [i for i in IO if len(i) == 2]\nKM\n","repo_name":"jh0118/class2021SPRING","sub_path":"20210317.ipynb","file_name":"20210317.ipynb","file_ext":"py","file_size_in_byte":752,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"5822826413","text":"# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# http://seananderson.ca/2013/10/19/reshape.html\n# -\n\n# # Tabular Data Gym 2. Reshaping.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport matplotlib.style\nmatplotlib.style.use('ggplot')\n\n# %matplotlib inline\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# %load_ext rpy2.ipython\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# library(magrittr)\n# library(ggplot2)\n# library(dplyr)\n# library(reshape2)\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# names(airquality) <- tolower(names(airquality))\n# head(airquality)\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# aq_pd = %R airquality\naq_pd.head()\n\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# Since `rpy2` does not treat NA well, we need to change it manually to `np.nan`.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\ndef fix_nans_in_df(colname, df):\n df.ix[df[colname] < 0, colname] = np.NaN \n\nfor colname in aq_pd.columns.values:\n fix_nans_in_df(colname, aq_pd)\n\naq_pd.head()\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# ## Melt/stack: changing wide format into long\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# R function `melt` from `reshape2` package change wide fromat into long. \n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n#\n# aql <- melt(airquality)\n# aql %>% head\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# ### Question 1\n#\n# One can get similar result using pandas' method `melt`. \n# -\n\npd.melt(aq_pd)\n\n# ### Question 2\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# airquality %>%\n# melt(id.vars = c(\"month\", \"day\")) %>%\n# head()\n# -\n\n# Now let's try to get it with pandas.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\npd.melt(aq_pd, id_vars = ['month','day'])\n# -\n\n# ## Question 3 Putting names to columns\n\n# + language=\"R\"\n#\n# aql <-\n# melt(airquality, id.vars = c(\"month\", \"day\"),\n# variable.name = \"climate_variable\", \n# value.name = \"climate_value\") %>%\n# mutate(date=as.Date(paste0(\"2015-\",month,'-',day)))\n# head(aql)\n# -\n\n# Your turn with pandas.\n\npd.melt(aq_pd, id_vars = ['month','day'], var_name = 'climate_variable', value_name= 'climate_value')\n\n# ### Comments about ploting\n#\n# When one uses ggplot it is natural to use long format when ploting.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"} language=\"R\"\n# aql %>%\n# ggplot() +\n# geom_line(aes(date, climate_value, colour=climate_variable))\n\n# + [markdown] ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n# But with pandas' mathplotlib interface it is easier to use wild format.\n#\n# First let's create dates.\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\naq_pd2= aq_pd.copy()\n\naq_pd2['date'] = \\\n [pd.Timestamp(s) \n for s in ('2015-'+aq_pd['month'].astype(int).astype(str)+'-'\n +aq_pd['day'].astype(int).astype(str))]\n# -\n\n# Then selecting columns and adding aproprite index one get: \n\naq_pd2_indexed = aq_pd2[['date', 'ozone', 'solar.r', 'wind', 'temp']]\naq_pd2_indexed.set_index('date')\naq_pd2_indexed.plot()\n\n# ## Long- to wide-format data\n\n# + language=\"R\"\n# aql <- melt(airquality, id.vars = c(\"month\", \"day\"))\n# head(aql)\n# -\n\n# Can you do the same with pandas and then create plot?\n\naq_pd2.plot()\n\n# ### Question 4. Time for dcast\n\n# + language=\"R\"\n# aqw <- dcast(aql, month + day ~ variable)\n# head(aqw)\n# -\n\n# The approprite pandas method seems to be `pivot_table`. Can you try?\n\naq_long = pd.melt(aq_pd, id_vars = ['month','day'])\naq_long.head()\n\naq_wide = pd.pivot_table(aq_long, index = ['month','day'],columns = 'variable')\naq_wide.head()\n\naq_wide.reset_index()\n\n# ### Question 5. Melting with stack\n#\n# There is another pandas' method for maelting. It is called stack. Can you try it with `aq_pd`? What is the type of the result? Can you make it to be DataFrame with comlumns `varaiable` and `value`?\n\n# + ein.tags=[\"worksheet-0\"] slideshow={\"slide_type\": \"-\"}\n\n# -\n\n# ### Question 6. Melt, filter, group, agregate.\n\n# + language=\"R\"\n# aql %>% head\n\n# + language=\"R\"\n# airquality %>%\n# melt(id.vars = c(\"month\", \"day\")) %>%\n# filter(!is.na(value)) %>%\n# group_by(variable) %>%\n# summarize(total= sum(value)) \n\n# + language=\"R\"\n# library(data.table)\n# data.table(airquality) %>%\n# melt(id.vars = c(\"month\", \"day\")) %>%\n# .[!is.na(value), .(total=sum(value)), by=variable]\n# -\n\n# Time for doing this with pands:\n\n\n","repo_name":"DataGym/tabularDataGym2","sub_path":"luca/iteration_01.ipynb","file_name":"iteration_01.ipynb","file_ext":"py","file_size_in_byte":4706,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"14711500924","text":"from keras.datasets import mnist\nfrom keras.models import Sequential, Model, load_model\nfrom keras.utils import to_categorical\nfrom keras.layers import Dense, Flatten, Input, BatchNormalization, Dropout\nfrom keras import optimizers\nfrom keras import regularizers\nfrom keras.callbacks import EarlyStopping, ModelCheckpoint\n\nimport numpy as np\nimport matplotlib.pyplot as plt\n\n# ### Importation de la base de données MNIST (ce sont des images de chiffres manuscrits, hello world du deep learning)\n\n(x_train, y_train), (x_test, y_test) = mnist.load_data()\n\nx_train.shape\n\n# ### Affichage des chiffres\n\n# +\nr = np.random.randint(x_train.shape[0])\n\nplt.imshow(x_train[r],cmap = \"hot\")\n# -\n\n# ### Normalisation des données\n\n# +\nx_train = x_train/255\n\nx_test = x_test/255\n# -\n\n# ### Passage des données (0,1,2,3...) en catégories. Exemple : 3 devient (0,0,0,1,0,0,0,0,0,0)\n\n# +\ny_train_cat = to_categorical(y_train,num_classes = 10)\n\ny_test_cat = to_categorical(y_test,num_classes = 10)\n# -\n\n# ### Construction du réseau\n\nmodel = Sequential()\nmodel.add(Flatten(input_shape = (28,28))) #Passage de tableau 2D à 1D\n\nmodel.add(Dense(128,activation = \"relu\", kernel_initializer = \"he_normal\",kernel_regularizer = regularizers.l2(0.01)))\nmodel.add(Dropout(0.1))\nmodel.add(BatchNormalization())\nmodel.add(Dense(128,activation = \"relu\", kernel_initializer = \"he_normal\",kernel_regularizer = regularizers.l2(0.01)))\nmodel.add(Dropout(0.1))\nmodel.add(BatchNormalization())\nmodel.add(Dense(10,activation = \"softmax\", kernel_initializer = \"he_normal\",kernel_regularizer = regularizers.l2(0.01)))\n\n# ### Optimizer et loss function\n\nopt = optimizers.SGD(lr = 0.01)\n\nmodel.compile(loss = \"binary_crossentropy\",optimizer = opt, metrics = ['categorical_accuracy'])\n\n# ### Affichage de l'architecture du réseau \n\nprint(model.summary())\n\n# ### Mise en place du Early Stopping\n\nearly_stopping = EarlyStopping(monitor = \"val_loss\",patience = 10,restore_best_weights=True)\n\n\n# ### Checkpointer pour sauvegarder le meilleur modèle\n\ncheckpointer = ModelCheckpoint(filepath = \"model_exemple_nist_best.h5\",verbose = 1,save_best_only = True)\n\n# ### Apprentissage\n\nhist = model.fit(x_train,y_train_cat,epochs = 10,batch_size = 128,validation_split = 0.1,callbacks = [early_stopping,checkpointer])\n\n# ### Affichage de l'historique de l'apprentissage\n\n# +\nfigure = plt.figure()\n\nax1 = plt.subplot(121)\nax1.plot(hist.history[\"loss\"],label = \"Training Loss\")\nax1.plot(hist.history[\"val_loss\"],label = \"Validation Loss\")\nplt.xlabel(\"Nombre d'itérations\")\nplt.ylabel(\"Loss function\")\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.plot(hist.history[\"categorical_accuracy\"],label = \"Training Accuracy\")\nax2.plot(hist.history[\"val_categorical_accuracy\"],label = \"Validation Accuracy\")\nplt.xlabel(\"Nombre d'itérations\")\nplt.ylabel(\"Accuracy\")\nplt.legend()\n# -\n\n# ### Accuracy sur la base de test\n\n# +\nprediction_bar = model.predict(x_test)\nprediction = np.argmax(prediction_bar,axis = 1)\n\nprint(\"Accuracy on test set: \" + str(np.round(np.sum(prediction == y_test)/x_test.shape[0]*100,3)) + \" %\")\n# -\n\ny_test.shape\n\n# ### Essai sur la base de test\n\n# +\nfigure = plt.figure()\n\nfigure.clear()\n\nr = np.random.randint(x_test.shape[0])\n\nax1 = plt.subplot(121)\nax1.imshow(x_test[r],cmap = \"hot\")\nplt.title(\"Prédiction du réseau : \" + str(prediction[r]) + \"\\n Vraie valeur : \" + str(y_test[r]))\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.bar(np.arange(10),height = prediction_bar[r],tick_label = np.arange(10))\nplt.xlabel(\"Valeur\")\nplt.ylabel(\"Output du réseau\")\nplt.legend()\n# -\n\n# ### Exemple d'erreur\n\n# +\nfigure = plt.figure()\n\nfigure.clear()\n\nerreurs = np.where(prediction != y_test)[0]\n\nr2 = np.random.randint(erreurs.shape[0])\ne = erreurs[r2]\n\nax1 = plt.subplot(121)\nax1.imshow(x_test[e],cmap = \"hot\")\nplt.title(\"Prédiction du réseau : \" + str(prediction[e]) + \"\\n Vraie valeur : \" + str(y_test[e]))\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.bar(np.arange(10),height = prediction_bar[e],tick_label = np.arange(10))\nplt.xlabel(\"Valeur\")\nplt.ylabel(\"Output du réseau\")\nplt.legend()\n# -\n\n# # TEST SUR LA RECONNAISANCE DIMAGE PERSO\n#\n\n# +\nfrom PIL import Image, ImageFilter\n\n# https://stackoverflow.com/questions/35842274/convert-own-image-to-mnists-image\n# https://www.pluralsight.com/guides/importing-image-data-into-numpy-arrays\n\ndef imageprepare(argv):\n \"\"\"\n This function returns the pixel values.\n The imput is a png file location.\n \"\"\"\n im = Image.open(argv).convert('L')\n width = float(im.size[0])\n height = float(im.size[1])\n newImage = Image.new('L', (28, 28), (255)) # creates white canvas of 28x28 pixels\n\n if width > height: # check which dimension is bigger\n # Width is bigger. Width becomes 20 pixels.\n nheight = int(round((20.0 / width * height), 0)) # resize height according to ratio width\n if (nheight == 0): # rare case but minimum is 1 pixel\n nheight = 1\n # resize and sharpen\n img = im.resize((20, nheight), Image.ANTIALIAS).filter(ImageFilter.SHARPEN)\n wtop = int(round(((28 - nheight) / 2), 0)) # calculate horizontal position\n newImage.paste(img, (4, wtop)) # paste resized image on white canvas\n else:\n # Height is bigger. Heigth becomes 20 pixels.\n nwidth = int(round((20.0 / height * width), 0)) # resize width according to ratio height\n if (nwidth == 0): # rare case but minimum is 1 pixel\n nwidth = 1\n # resize and sharpen\n img = im.resize((nwidth, 20), Image.ANTIALIAS).filter(ImageFilter.SHARPEN)\n wleft = int(round(((28 - nwidth) / 2), 0)) # caculate vertical pozition\n newImage.paste(img, (wleft, 4)) # paste resized image on white canvas\n\n # newImage.save(\"sample.png\n\n tv = list(newImage.getdata()) # get pixel values\n\n # normalize pixels to 0 and 1. 0 is pure white, 1 is pure black.\n tva = [(255 - x) * 1.0 / 255.0 for x in tv]\n print(tva)\n return tva\n\n\n# -\n\n# charger image perso \nx=imageprepare('datasets/mnist_test/test_3.png') #file path here\nprint(len(x))# mnist IMAGES are 28x28=784 pixels\n\n# +\n# TRANSFORM X [784,1] into array(1,28,28)\ncol = 28 # just grab the number of columns here\nxtransfinal = []\nxtransfinal.append([x[i:i+col] for i in range(0, len(x), col)])\nxtransfinal = np.array(xtransfinal)\n\n#VISUALIZATION\nplt.imshow(xtransfinal[0],cmap = \"hot\")\nxtransfinal.shape\n# -\n\n# FEEDING INPUT TO MODEL \npredictionbaralx = model.predict(xtransfinal)\npredictionalx = np.argmax(predictionbaralx,axis = 1)\n\n# +\n# RESULT VIZUALISATION\nfigure = plt.figure()\nfigure.clear()\n\n#r = np.random.randint(x_test.shape[0])\nr=0\n\nax1 = plt.subplot(121)\nax1.imshow(xtransfinal[r],cmap = \"hot\")\nplt.title(\"Prédiction du réseau : \" + str(predictionalx[r]))# + \"\\n Vraie valeur : \" + str(y_test[r]))\nplt.legend()\n\nax2 = plt.subplot(122)\nax2.bar(np.arange(10),height = predictionbaralx[r],tick_label = np.arange(10))\nplt.xlabel(\"Valeur\")\nplt.ylabel(\"Output du réseau\")\nplt.legend()\n\n# +\n#import numpy as np\n#xtrans = np.array(x)\n#shape = (28,28)\n#xtrans.reshape(shape)\n#xtrans\n\n# +\n#col = 28 # just grab the number of columns here\n#xtrans2 = [x[i:i+col] for i in range(0, len(x), col)]\n#xtrans2 = np.array(xtrans2)\n#xtrans2.shape\n","repo_name":"Mathonal/MyMLBasics","sub_path":"3_deep_learning/NNET_Mnist_chiffre.ipynb","file_name":"NNET_Mnist_chiffre.ipynb","file_ext":"py","file_size_in_byte":7211,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"6838024158","text":"# # Addition\n\n1 + 2\n\n# +\n# This program adds two numbers\n\nnum1 = 1.5\nnum2 = 6.3\n\n# Add two numbers\nsum = num1 + num2\n# -\n\nprint(sum)\n\n# ## Another way to print\n\n# Display the sum\nprint('The sum of {0} and {1} is {2}'.format(num1, num2, sum))\n\n# # Subtraction\n\n# +\nnum1 = 128\nnum2 = 256\n\nsub = num1 - num2\nprint('{0} - {1} = {2}'.format(num2,num1,sub))\n# -\n\n# ## Multiplication\n\n# +\nnum1 = 12\nnum2 = 2.5\n\npdt = num1*num2\nprint('{0} * {1} = {2}'.format(num1,num2,pdt))\n# -\n\n# # Division\n\n# +\nnum1 = 12\nnum2 = 2.5\nnum3 = 5\n\ndiv1 = num1/num2\nprint(div1)\n\n# floating division\ndiv4 = num1/num3\nprint(div4)\n\n# integer divions\ndiv5 = num1 // num3 # gives the quotient\nprint(div5)\n\n# -\n\n# # Exponentiation\n\n# +\nnum1 = 1.25\nnum2 = 2\nnum3 = -0.5\n\n# Integer power\nexp1 = num1 ** num2\nprint(exp1)\n\n# negative noninteger power\nexp2 = num2 ** num3\nprint(exp2)\n# -\n\n# # Mod or Remainder Operator\n\n# +\nnum1 = 35\nnum2 = 4\nnum3 = 5\n\nrem1 = num1 % num2\nprint(num1, ' % ',num2, ' = ', rem1) # see how we print \n\nrem2 = num1 % num3\nprint(num1, ' % ',num3, ' = ', rem2)\n# -\n\n# # Other mathematical operators\n\n\n","repo_name":"mcodevb/Python-with-Mainul","sub_path":"Mathematical operations in Python.ipynb","file_name":"Mathematical operations in Python.ipynb","file_ext":"py","file_size_in_byte":1089,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"73207704151","text":"# + id=\"h8se9n3yvcR7\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303943922, \"user_tz\": 180, \"elapsed\": 359, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}}\n#pip install control\n##download da biblioteca de controle\n\n# + id=\"is2Ex_8zKocc\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303944312, \"user_tz\": 180, \"elapsed\": 7, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}}\n#bibliotexas para o projeto\nimport control as ctl\nimport matplotlib.pyplot as plt\nimport numpy as np\n\n# + id=\"aTqN3L9jK95N\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303944312, \"user_tz\": 180, \"elapsed\": 6, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}}\n#variaveis de resistencia e capacitancia\nR = 1.;\nC = 1.;\n\n#vetores que serao os polinomios para a funçao de transferencia\nTsim = 10 #vetor de tempo de simulaçao\nnumerador = [1.]\ndenominador = [R*C, 1.]\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"LO5PJFqBNGAk\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631303944312, \"user_tz\": 180, \"elapsed\": 6, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}} outputId=\"3802795f-833e-45d5-8ca0-2c0f18015676\"\n#usa os vetores para criar a funçao de transferencia\nH = ctl.tf(numerador, denominador)\nprint('FT malha aberta:', H)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 279} id=\"8pNM7ArFOWdq\" executionInfo={\"status\": \"ok\", \"timestamp\": 1631304221827, \"user_tz\": 180, \"elapsed\": 434, \"user\": {\"displayName\": \"David Oliveira\", \"photoUrl\": \"https://lh3.googleusercontent.com/a-/AOh14Gi82wE6qIe5omAkHJKMK6mqmW_ja_uu5IjOde1-Hw=s64\", \"userId\": \"13272824796631453103\"}} outputId=\"80c251d6-9ff0-4536-c149-73875085e0b5\"\n#calcula a resposta ao degrau\nT, yout = ctl.step_response(H, Tsim) # retorna dois valores a partir dos parametros de funçao de transferencia e tempo\nplt.plot(T, yout, 'b-')\n\n#calcula um degrau unitario\nT2 = np.linspace(-1., 10., 1000)\ndegrau = np.ones_like(T2)\ndegrau[T2<0] = 0\nplt.plot(T2, degrau, 'r-')\n\n#melhorando o grafico\nplt.ylabel('V (V)')\nplt.xlabel('Tempo (s)')\nplt.legend(['resposta ao degrau', 'degrau unitario'])\nplt.grid()\n","repo_name":"davidolivaa/ufabc-ieee-orion-adcs","sub_path":"Controle/01-introducao/01-circuitoRC.ipynb","file_name":"01-circuitoRC.ipynb","file_ext":"py","file_size_in_byte":2585,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"74010120153","text":"import matplotlib.pyplot as plt \nimport numpy as np \nimport pandas as pd\n\n# +\nphase_no=40\n\narr= np.genfromtxt(f'0.1_list_{phase_no}.csv', delimiter=',')\n# -\n\narr[3][1:]\n\n\ndef unpack_phi(phase_no):\n \n arr= np.genfromtxt(f'0.1_list_{phase_no}.csv', delimiter=',')\n \n phi_list= [i if i < np.pi else i-2*np.pi for i in arr[3][1:]%(2*np.pi)]\n \n phi_array= np.reshape(phi_list, (10,5))\n \n return phi_array\n\n\nunpack_phi(10).T\n\nimport seaborn as sns\n\nsns.heatmap(unpack_phi(0), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(10), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(20), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(30), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\nsns.heatmap(unpack_phi(40), vmin= -np.pi, vmax= np.pi, cmap= 'hsv');\n\n# +\n#Takes too long to converge to lateral phase\n\nplt.plot(unpack_phi(40)[0]);\n\n# +\n#Behold a little 'cheating' force a symmetric initial condition on both y-axis and run simulation\n\nphi_list= unpack_phi(40)\n\ndef lamb(x_n, y_n, phi_list):\n \n arr= (np.array([i[::-1] for i in phi_list]) + phi_list)/2\n \n return arr[x_n][y_n]\n\n\n# -\n\nplt.plot(((np.array([i[::-1] for i in phi_list]) + phi_list)/2)[0])\n\n# +\nimport fluid_functions as ff\nimport numpy as np\n\nfrom cilia_test_copy import Cilium\nfrom cilia_test_copy import Cilia_System\nfrom cilia_test_copy import big_phi\n\n\nimport matplotlib.pyplot as plt\nfrom itertools import product\n\nimport matplotlib.cm as cm\n\nl_ambda = 0.1\nr0= 0.5\nlattice= 2\na_radii=0.01\n\nf0 = lambda phi : 0\n\ny_n=5\nx_n=10\n\nCilia_System_1= Cilia_System()\n\nnew_constants= np.array([r0, l_ambda ,10])\n\nCilia_row= [Cilium([i*lattice, j*lattice], [0,r0,lamb(i, j, unpack_phi(40))], constants= np.array([r0, l_ambda ,10]), a= a_radii) for i, j in product(range(x_n), range(y_n))]\n\nCilia_System_1.add(Cilia_row)\n\n# -\n\nwhile Cilia_System_1.phi_list[0] < (20*np.pi):\n \n Cilia_System_1.update(stepsize= 0.02)\n\nCilia_System_1.phi_list\n\n# +\narr= Cilia_System_1.phi_list%(2*np.pi)\n\narr= np.array([i if i < np.pi else i-2*np.pi for i in arr])\n\narr=np.reshape(arr, (10,5))\n\nplt.rcParams[\"figure.figsize\"] = (10,5)\n\nplt.plot(np.mean(arr, axis=0)-np.mean(arr, axis=0)[0]);\nplt.grid()\nplt.title('Average Lateral Phase')\nplt.xlabel('y')\nplt.ylabel('Phase');\n\n\n# +\nphis= np.mean(arr, axis=1)[::-1]-np.mean(arr, axis=1)[::-1][-1]\n\nphis= np.array([i if i < np.pi else i-2*np.pi for i in phis])\n\nplt.plot(phis, np.linspace(0, 9, 10))\n\nplt.rcParams[\"figure.figsize\"] = (8,10)\nplt.grid()\nplt.title('Average Streamwise Phase')\nplt.xlabel('Phase')\nplt.ylabel('x');\n# -\n\n\n","repo_name":"Desmondccw/M4R_Project","sub_path":"Figure 9.ipynb","file_name":"Figure 9.ipynb","file_ext":"py","file_size_in_byte":2611,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"18964869739","text":"import glob\nimport pandas as pd\nimport scipy.constants as scc\nfrom scipy.optimize import curve_fit\nfrom astropy.cosmology import FlatLambdaCDM\nimport astropy.units as u\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom astropy.modeling import models, fitting\nimport scipy\nimport scipy.stats as stats\nfrom astropy.coordinates import SkyCoord\nfrom scipy.integrate import quad\nfrom matplotlib.offsetbox import AnchoredText\nimport os\nimport scipy.special as sc\nimport gc\nimport warnings\nimport eazy\nimport time\nimport grizli\nimport numpy as np\nfrom astropy.table import Table\n\n\n# +\ndef Lv(freq, A, beta, Tdust):\n return A * freq**(3+beta) / (np.exp(h * freq / (k_b * Tdust ) ) - 1 )\n\ndef Lv_rescaled(freq, N, A, beta, Tdust): # Lv\n return N * A * freq**(3+beta) / (np.exp(h * freq / (k_b * Tdust ) ) - 1 )\n\ndef T_dust(z):\n return 32.9 + 4.6*(z-2), 2.4+0.35*(z-2) # mean value and dispersion\n\n\n# -\n\nh = 6.62e-27 # erg/s\nk_b = 1.38e-16 # erg/K\nc = 3e10 # speed of light: cm/s\nsigma = 5.67e-5 # stefan boltzmann constant, erg/cm^2/s/K^4\n\npath='/home/yjm/miniconda/Outputs'\nos.chdir(path)\nos.chdir('./a370')\n\nlamb_rest = (np.arange(8, 1200) * 1e-6 ) # micron meters to meters\nfreq_rest = scc.c / lamb_rest # Hz\nfreq_rest_8_1000 = scc.c / (np.arange(8, 1000) * 1e-6 )\nfreq_sorted = np.sort(freq_rest)\nfreq_8_1000_sorted = np.sort(freq_rest_8_1000)\nindex_1000 = np.where(lamb_rest==np.min(abs(freq_sorted-1)))\nkappa0 = 1.3 # cm^2/g --> m^2/kg\nv0 = 6.66e11 #Hz\nM_16=[]\nM_84=[]\nM_50=[]\nbeta_median=[]\nT_median=[]\nL_16 = []\nL_84 = []\nL_UV = []\nL_50 = []\ndetected_fits = glob.glob('*alma_detected.fits')[0]\ntable = Table.read(detected_fits)\nmagnification = np.array(table['magnification'])\nSv_obs = np.array(table['flux_1.2mm'])/magnification*1e-26 # from micron Jy to erg/sec/cm^2\nredshift = np.array([1., 2., 3., 4., 5., 6., 7., 8., 9., 10., 11., 12.])\nlamb_alma = 1200/(1 + redshift)*1e-6 # --> meter\nfreq_alma = scc.c / lamb_alma\nnum_iteration=1000\nbeta = np.random.normal(1.8, 0.2, size=(num_iteration,))\na=2e-26\ncosmo = FlatLambdaCDM(H0=70 * u.km / u.s / u.Mpc, Om0=0.3)\n\nfor j in range(redshift.size):\n d_L = cosmo.luminosity_distance(redshift[j])\n d_L = d_L.to(u.cm) # from parsec to cm\n low, high = T_dust(redshift[j])\n T = np.random.normal(low, high, size=(num_iteration,))\n Lv_rest = 4*scc.pi*d_L**2*Sv_obs[0]/(1+redshift[j]) # unit: erg/s/Hz\n lamb_obs = lamb_rest * (1 + redshift[j])\n M_dust=[]\n t_peak=[]\n l_IR = []\n for i in range(num_iteration): # loop over random T and beta\n N = Lv_rest.value / Lv(freq_alma[j], 2e-26, beta[i], T[i])\n Lv_rescale = N * Lv(freq_rest, 2e-26, beta[i], T[i]) \n Sv_rescale = Lv_rescale * (1 + redshift[j]) / (4 * np.pi * d_L**2)\n nu = 1.5e11 * (1+redshift[j])\n kappa = kappa0 * (nu / v0)**beta[i] # m^2/kg\n # mistake here: Sv_rest and func() have different units\n L = Lv_rest.value / (Lv(freq_alma[j], a, beta[i], T[i]) ) * Lv(nu, a, beta[i], T[i])\n blackbody = 2 * h / c**2 * nu**3 / (np.exp(h*nu / (k_b*T[i])) - 1)\n mass = L / (4 * np.pi * kappa * blackbody ) # kg\n M_dust = np.append(M_dust, mass)\n #L_integrated = np.sum(Lv_rescale[0:-1]) * dx[-1]\n #sol=quad(Lv_rescaled, freq_rest_8_1000[-1], freq_rest_8_1000[0], args=(N, a, beta[i], T[i]))\n #L_integrated = sol[0]\n lamb_peak = scc.c / freq_rest[np.argmax(Lv_rescale)] * 1e6 # meter --> um\n T_peak = 2898 / lamb_peak\n t_peak = np.append(t_peak, T_peak)\n L_integrated = 4 *np.pi * d_L.value**2 * sigma * T_peak**4\n l_IR = np.append(l_IR, L_integrated/3.82e33 ) # in unit of L_sun\n if j==0:\n plt.hist(t_peak)\n m_16 = np.percentile(M_dust, 16, method='nearest')\n M_16 = np.append(M_16, m_16)\n m_84 = np.percentile(M_dust, 84, method='nearest')\n M_84 = np.append(M_84, m_84)\n l_16 = np.percentile(l_IR, 16, method='nearest')\n L_16 = np.append(L_16, l_16)\n l_50 = np.percentile(l_IR, 50, method='nearest')\n L_50 = np.append(L_50, l_50)\n l_84 = np.percentile(l_IR, 84, method='nearest')\n L_84 = np.append(L_84, l_84)\n m_50 = np.percentile(M_dust, 50, method='nearest')\n M_50 = np.append(M_50, m_50)\n\nprint(Lv_rescale)\n\nprint()\n\n# +\n#Mdust_errors = np.vstack([(M_50 - M_16), (M_84 - M_50)])\n\nplt.scatter(redshift, np.log10(M_50))\nplt.scatter(redshift, np.log10(M_16))\nplt.scatter(redshift, np.log10(M_84))\n#plt.errorbar(redshift, np.log10(M_50), yerr=np.log10(Mdust_errors))\n# -\n\n\n\nplt.scatter(redshift, np.log10(L_50))\nplt.scatter(redshift, np.log10(L_16))\nplt.scatter(redshift, np.log10(L_84))\n\n\n","repo_name":"hunyjm12/Master_Project_ALCS","sub_path":"mbb_test.ipynb","file_name":"mbb_test.ipynb","file_ext":"py","file_size_in_byte":4621,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"73226072151","text":"# +\nplanet = {\n 'name': 'Earth',\n 'moons': 1\n}\n#se puede sacar con get\n\nplanet.update({'name': 'Makemake'})\nplanet['name'] = 'Makemake'\nprint(planet)\n\nplanet.update({'name': 'Jupiter', 'moons': 79})\n\nplanet['orbital period'] = 4333\nprint(planet)\n\nplanet.pop('orbital period')\nprint(planet)\n\nplanet['diameter (km)'] = {\n 'polar': 133709,\n 'equatorial': 142984\n}\nprint(f\"{planet['name']} polar diameter: {planet['diameter (km)']['polar']}\")\n\n\n# +\nrainfall = {\n 'october': 3.5,\n 'november': 4.2,\n 'december': 2.1\n}\nfor key in rainfall.keys():\n print(f'{key}: {rainfall[key]}cm')\n\nif 'december' in rainfall:\n rainfall['december'] = rainfall['december'] + 1\nelse:\n rainfall['december'] = 1\nprint(rainfall)\n\ntotal_rainfall = 0\nfor value in rainfall.values():\n total_rainfall = total_rainfall + value\nprint(f'There was {total_rainfall}cm in the last quarter')\n\n# -\n\n# ## Ejercicio 1: Crear y modificar un diccionario de Python.\n\n# Crea un diccionario llamado planet con los datos propuestos\nplanet = {\n 'name': 'Mars',\n 'moons': 2\n}\n# Muestra el nombre del planeta y el número de lunas que tiene.\nprint(f\"{planet['name']}, {planet['moons']}\")\n# Agrega la clave circunferencia con los datos proporcionados previamente\nplanet['circunferencia km'] = {\n 'polar': 6752,\n 'equatorial': 6792\n}\nprint(f\"{planet['name']} has a polar circunference of {planet['circunferencia km']['polar']} km\")\n\n# ## Ejercicio 2: Programación dinámica con diccionarios\n# ## calculo de valores\n\n# +\n# Planets and moons\n\nplanet_moons = {\n 'mercury': 0,\n 'venus': 0,\n 'earth': 1,\n 'mars': 2,\n 'jupiter': 79,\n 'saturn': 82,\n 'uranus': 27,\n 'neptune': 14,\n 'pluto': 5,\n 'haumea': 2,\n 'makemake': 1,\n 'eris': 1\n}\n# Añade el código para determinar el número de lunas.\n\nmoons = planet_moons.values()\nplanets = len(planet_moons.keys())\n\n# Agrega el código para contar el número de lunas. \ntotal_moons = 0\nfor moon in moons:\n total_moons = total_moons + moon\n\n# Calcula el promedio dividiendo el total_moons por el número de planetas\npromedio = total_moons // planets\nprint(promedio)\n \n","repo_name":"CrisGutierrez05/CursoPython","sub_path":"Kata 8.ipynb","file_name":"Kata 8.ipynb","file_ext":"py","file_size_in_byte":2139,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"17148643519","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#  \n\n# + id=\"S60eHj8Eecoz\"\nfrom tensorflow.keras.layers import Dense,SimpleRNN,Embedding,Flatten\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.datasets import imdb\n\n# + id=\"XKaFH9BOeuPy\"\n(xtrain,ytrain),(xtest,ytest) = imdb.load_data()\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"zT8GmbsZe718\" outputId=\"a11d66d8-cbe5-42b2-d9da-2b807e73b57b\"\nxtrain\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"WhvEZczsfGgw\" outputId=\"1b9abeff-3544-40ad-bfd5-d907bae5cac5\"\nxtrain.shape\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"WHED7k0DfYSQ\" outputId=\"e3792a15-09f9-4417-bd56-730adb1d41b3\"\nxtest.shape\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"2ISo0IZsfavd\" outputId=\"ac94ea57-dd7e-49a1-cc0f-cb86a1d94f35\"\nxtrain[100]\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"kWmSJcFjfnWo\" outputId=\"63920c58-3ea2-4ec4-dc43-b992da9b5f81\"\nlen(xtrain[0])\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"z5FRZt7Ufxc8\" outputId=\"880b5280-6c12-43bf-cb8c-fa65f84fc67f\"\nlen(xtrain[10])\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"zADUWbbxfzOA\" outputId=\"471633ad-e794-49fb-fd57-cdc3091f6e9d\"\nlen(xtrain[25])\n\n# + id=\"A10lBHq3f3Ue\"\n# same sequence\nfrom tensorflow.keras.utils import pad_sequences\nxtrain = pad_sequences(xtrain,padding='post',maxlen=50)\nxtest = pad_sequences(xtest,padding='post',maxlen=50)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"VNcuTAFXgnKX\" outputId=\"16768bef-af33-4591-a278-9631a1203117\"\nxtrain.shape\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"dHP_3LLqgunv\" outputId=\"3e05961e-adb5-4d80-f0ef-21eee26288e3\"\nxtrain[0]\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"Yp7R49ZLgy5B\" outputId=\"79ed4a10-3763-450b-a1de-142cb6222240\"\nmodel = Sequential()\n\nmodel.add(SimpleRNN(32,activation='tanh',input_shape=(50,1),return_sequences=False))\n\nmodel.add(Dense(1,activation='sigmoid'))\n\nmodel.summary()\n\n# + id=\"Xstha28NhXho\"\nmodel.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"lSv7bOSoioCP\" outputId=\"a3f8e9d8-7cde-4057-a664-1a8a85ab4a2e\"\nhistory = model.fit(xtrain,ytrain, validation_data=(xtest,ytest),epochs=1)\n\n# + id=\"s_MCOwzoiyz-\"\n\n","repo_name":"adityadhumal06/RNN-imdb-Project","sub_path":"RNN_imdb_dataset.ipynb","file_name":"RNN_imdb_dataset.ipynb","file_ext":"py","file_size_in_byte":2504,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"74073385752","text":"# 1. 데이터 벡터 다루기\n#\n# [예제] A 회사의 2010년 월별매출액 데이터가 다음과 같이 주어졌다고 하자.\n# \n# 100, 120, 130, 124, 150, 167, 170, 163, 160, 155, 145, 157\n# \n# 지수(index)를 이용하여 데이터벡터를 다루어 보기로 한다.\n\nimport numpy as np\nx=np.array([100, 120, 130, 124, 150, 167, 170, 163, 160, 155, 145, 157])\nx\n\nx[1]\n\nx[1:3]\n\nx[1:4]\n\nx[-1]\n\nx[-3]\n\nx[1:-3]\n\nx[7:]\n\nx[:3]\n\nx[::2]\n\nx[1:4:2]\n\nx>2\n\nx2=np.array([[1,2,3,4],[5,6,7,8],[9,10,11,12]])\n\nx2\n\nx2[0]\n\nx2[0,]\n\nx2[0,:]\n\nx2[0,1]\n\nx2[:2,1]\n\nx2[:2,]\n\nx2>2\n\nx2[x2>0]\n\n# + active=\"\"\n# 1. 순서함수 활용\n# -\n\n#데이터 프레임 생성\nimport pandas as pd\ndf=pd.DataFrame({\n 'name':['Kim','Lee','Park','Bae','Kang'],\n 'age': [24,32,43,24,30],\n 'height':[160,170,165,173,158]\n})\ndf\n\n# +\n#나이를 기준으로 순위를 주어. rank_average 변수 생성 -> rank() 사용\n\ndf['rank_average']=df['age'].rank(method='min', ascending=False) #method='min': 동점일 경우 최솟값을 줌\ndf #ascending=False: 내림차순으로 순위를 줌 (값이 클수록 높은 순위)\n# -\n\ndf['age'].rank(method='max',ascending=False) #method='max' 동점일 경우 최댓값을 줌\n\ndf['age'].rank(method='average',ascending=False) #method='average' 동점일 경우 평균을 줌 default\n\ndf['age'].rank(ascending=False)\n\n# +\n#결측값 포함한 데이터 프레임인 경우\n\ndf2=pd.DataFrame({\n 'name':['Kim','Lee','Park','Bae','Kang'],\n 'age':[24,32,43,24,np.nan],\n 'height':[160,170,165,173,158]\n})\ndf2\n\n# +\n#결측값 처리 위한 옵션 na_option 추가\n\ndf2['rank_average']=df2['age'].rank(method='min', ascending=False, na_option='bottom')\n#na_option='bottom': 결측값에 후순위를 주고자 할 경우\n#na_option='top': 결측값에 선순위를 주고자 할 경우\n#na_option='keep': 결측값을 NaN으로 두고자 할 경우 default\ndf2\n\n# +\n#데이터 프레임 df에서 name변수 기준으로 오름차순 정렬하기\n\ndf_sort=df.sort_values(by='name')\ndf_sort\n# -\n\n#데이터 프레임 df에서 age 변수 기준으로 오름차순 정렬하기\ndf_sort=df.sort_values(by='age')\ndf_sort\n\n# +\n#데이터 프레임 df에서 height 변수 기준으로 내림차순으로 정렬하기\n\ndf_sort=df.sort_values(by='height', ascending=False)\ndf_sort\n# -\n\n#데이터 프레임 df에서 두 변수 age, height 변수 기준으로 오름차순으로 정렬하기\ndf_sort=df.sort_values(by=['age','height'])\ndf_sort\n\n# + active=\"\"\n# 1. 결측값\n\n# +\n#데이터 프레임 df2에 결측값 포함 여부 확인하기\n\ndf2.isnull().values.any() #True:결측값 포함\n\n# +\n#데이터 프레임 df2 결측값에 0으로 대체하기\n\ndf2_0=df2.fillna(0)\ndf2_0\n# -\n\n#결측값을 변수별 평균으로 대체\ndf_mean=df2.fillna(df2.mean(numeric_only=True))\ndf_mean\n\n# +\n#순서상 결측값의 바로 앞의 값으로 대체하기\n\ndf2_f=df2.fillna(method='ffill')\ndf2_f\n\n# +\n#순서상 결측값의 바로 뒤의 값으로 대체하기\n\ndf2_f=df2.fillna(method='backfill')\ndf2_f\n# -\n\ndf3=df2.sort_values(by='height')\ndf3\n\ndf3_f=df3.fillna(method='backfill')\ndf3_f\n\ndf3_f2=df2.sort_values(by='height').fillna(method='backfill')\ndf3_f2\n\n\n\n\n\ndf2.sort_values(by='height').fillna(method='backfill')[['name','age']]\n\n\n","repo_name":"jpark630/stat-software","sub_path":"Statistitcal Software Week 13 practice.ipynb","file_name":"Statistitcal Software Week 13 practice.ipynb","file_ext":"py","file_size_in_byte":3335,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"18226709526","text":"# +\nimport numpy as np \nimport pandas as pd\nimport matplotlib.pyplot as plt \nimport export as export \nimport eli5\nimport warnings\nwarnings.filterwarnings('ignore')\n\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.preprocessing import OrdinalEncoder\nfrom sksurv.datasets import load_gbsg2\nfrom sksurv.preprocessing import OneHotEncoder\nfrom sksurv.ensemble import RandomSurvivalForest\nfrom sksurv.datasets import load_whas500\nfrom sksurv.tree import SurvivalTree\nfrom sksurv.util import Surv\n# -\n\nX, y = load_whas500()\nX = X.astype(float)\nestimator = SurvivalTree().fit(X, y)\n\nexport.plot_tree(estimator, feature_names = estimator.feature_names_in_, impurity = False,)\n\n# +\npredict_outcomes = estimator.predict(X)\nsurv_funcs = estimator.predict_survival_function(X.iloc[:5])\n\nfor fn in surv_funcs:\n plt.step(fn.x, fn(fn.x), where=\"post\")\nplt.ylim(0,1)\nplt.show()\n\n# +\nchf_funcs = estimator.predict_cumulative_hazard_function(X.iloc[:5])\n\nfor fn in chf_funcs:\n plt.step(fn.x, fn(fn.x), where=\"post\")\nplt.ylim(0,1)\nplt.show()\n\n\n# +\nX,y = load_gbsg2()\ngrade_str = X.loc[:, \"tgrade\"].astype(object).values[:,np.newaxis]\ngrade_num = OrdinalEncoder(categories=[[\"I\", \"II\", \"III\"]]).fit_transform(grade_str)\nX_no_grade = X.drop(\"tgrade\",axis=1)\nXt = OneHotEncoder().fit_transform(X_no_grade)\nXt.loc[:, \"tgrade\"] = grade_num\n\n\nrandom_state = 20\nX_train, X_test,y_train,y_test = train_test_split(Xt, y, test_size=0.25, random_state=random_state)\n\nrsf = RandomSurvivalForest(n_estimators = 1000, \n min_samples_split = 10,\n min_samples_leaf = 15,\n max_features = \"sqrt\",\n n_jobs = -1,\n random_state = random_state)\n\nrsf.fit(X_train,y_train)\nrsf.score(X_test, y_test)\n# -\n\nX_test_sorted = X_test.sort_values(by=[\"pnodes\", \"age\"])\nX_test_sel = pd.concat((X_test_sorted.head(3), X_test_sorted.tail(3)))\nX_test_sel\n\npd.Series(rsf.predict(X_test_sel))\n\n\nsurv = rsf.predict_survival_function(X_test_sel, return_array=True)\nfor i,s in enumerate(surv):\n plt.step(rsf.event_times_,s, where=\"post\", label=str(i))\nplt.ylabel(\"Survivalprobability\")\nplt.xlabel(\"Timeindays\")\nplt.legend()\nplt.grid(True)\n\nsurv = rsf.predict_cumulative_hazard_function(X_test_sel, return_array=True)\nfor i,s in enumerate(surv):\n plt.step(rsf.event_times_,s, where=\"post\", label=str(i))\nplt.ylabel(\"Cumulativehazard\")\nplt.xlabel(\"Timeindays\")\nplt.legend()\nplt.grid(True)\n\n# +\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nfrom sksurv.datasets import load_whas500\n\nX, y = load_whas500()\n# -\n\nX.head(10).style.hide_index()\n\ny[:10]\n\ndf_y = pd.DataFrame(data={'time': [y[i][1] for i in range(len(y))], 'event': [y[i][0] for i in range(len(y))]})\ndf_y[:10].style.hide_index().highlight_min('event', color='lightgreen')\n\n# One of the main challenges of survival analysis is right censoring, i.e., by the end of the study, the event of interest (for example, in medicine 'death of a patient' or in this dataset 'recurrence of cancer') has only occurred for a subset of the observations.\n#\n# The right censoring in this dataset is given by the column named 'event' and it's a variable which can take value 'True' if the patient had a recurrence of cancer or 'False' if the patient is recurrence free at the indicated time (right-censored samples).\n#\n# Let's see how many right-censored samples do we have.\n\nprint(f'Number of samples: {len(df_y)}')\nprint(f'Number of right censored samples: {len(df_y.query(\"event == False\"))}')\nprint(f'Percentage of right censored samples: {100*len(df_y.query(\"event == False\"))/len(df_y):.1f}%')\n\n# There are 387 patients (56.4%) who were right censored (recurrence free) at the end of the study.\n#\n# Let's divide our dataset in training and test sets.\n\n# +\nfrom sklearn.model_selection import train_test_split\n\nX_trn, X_test, y_trn, y_test = train_test_split(X, y, test_size = 0.30, random_state = 42)\n\nprint(f'Number of training samples: {len(y_trn)}')\nprint(f'Number of test samples: {len(y_test)}')\n# -\n\n# We divide the features/covariates into continuous and categorical.\n\nscaling_cols = [c for c in X.columns if X[c].dtype.kind in ['i', 'f']]\ncat_cols = [c for c in X.columns if X[c].dtype.kind not in [\"i\", \"f\"]]\n\n# We use ordinal encoding for categorical features/covariates and standard scaling for continuous features/covariates.\n\n# +\nfrom sklearn.compose import ColumnTransformer\nfrom sklearn.preprocessing import OrdinalEncoder\nfrom sklearn.preprocessing import StandardScaler\n\npreprocessor = ColumnTransformer(\n [('cat-preprocessor', OrdinalEncoder(), cat_cols),\n ('standard-scaler', StandardScaler(), scaling_cols)],\n remainder='passthrough', sparse_threshold=0)\n\n# +\nfrom sksurv.ensemble import RandomSurvivalForest\nfrom sklearn.pipeline import make_pipeline\nfrom sksurv.linear_model import CoxPHSurvivalAnalysis\nfrom sksurv.metrics import concordance_index_censored\nfrom scipy.stats import reciprocal\nfrom sklearn.model_selection import RandomizedSearchCV\n\nrsf = make_pipeline(preprocessor, RandomSurvivalForest(random_state=42))\nrsf.fit(X_trn, y_trn)\nci_rsf = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], rsf.predict(X_test))\nprint(f'The c-index of Random Survival Forest is given by {ci_rsf[0]:.3f}')\n\n\n# +\nclass reciprocal_int:\n def __init__(self, a, b):\n self._distribution = reciprocal(a, b)\n\n def rvs(self, *args, **kwargs):\n return self._distribution.rvs(*args, **kwargs).astype(int)\n\nparam_distributions1 = {\n 'randomsurvivalforest__max_features': reciprocal_int(3, 8),\n 'randomsurvivalforest__max_depth': reciprocal_int(5, 50),\n 'randomsurvivalforest__min_samples_leaf': reciprocal_int(1, 40),\n}\n\nmodel_random_search = RandomizedSearchCV(\n rsf, param_distributions=param_distributions1, n_iter=50, n_jobs=-1, cv=3, random_state=0)\nmodel_random_search.fit(X_trn, y_trn)\n\nprint(\n f\"The c-index of random survival forest using a {model_random_search.__class__.__name__} is \"\n f\"{model_random_search.score(X_test, y_test):.3f}\")\nprint(\n f\"The best set of parameters is: {model_random_search.best_params_}\"\n)\n# -\n\nmax_depth = model_random_search.best_params_['randomsurvivalforest__max_depth']\nmax_features = model_random_search.best_params_['randomsurvivalforest__max_features']\nmin_samples_leaf = model_random_search.best_params_['randomsurvivalforest__min_samples_leaf']\n\nrsf_best = make_pipeline(preprocessor, RandomSurvivalForest(random_state=42, max_depth=max_depth, max_features=max_features, min_samples_leaf=min_samples_leaf))\nrsf_best.fit(X_trn, y_trn)\nci_rsf = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], rsf_best.predict(X_test))\nprint(f'The c-index of Random Survival Forest is given by {ci_rsf[0]:.3f}')\n\n# +\nfrom eli5.sklearn import PermutationImportance\nperm = PermutationImportance(rsf.steps[-1][1], n_iter=100).fit(preprocessor.fit_transform(X_trn),y_trn)\n\ndata = perm.results_\ndata = pd.DataFrame(data, columns=X_trn.columns)\nmeds = data.median()\nmeds = meds.sort_values(ascending=False)\ndata = data[meds.index]\n# -\n\nfig, ax = plt.subplots(figsize=(10,7))\ndata.boxplot(ax=ax)\nax.set_title('Feature Importances')\nplt.show()\n\neli5.show_weights(perm, feature_names = X.columns.tolist())\n\n# +\ncox = make_pipeline(preprocessor, CoxPHSurvivalAnalysis())\ncox.fit(X_trn, y_trn)\n\nci_cox = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], cox.predict(X_test))\nprint(f'The c-index of Cox is given by {ci_cox[0]:.3f}')\n\n# +\nnp.random.seed(0)\nseeds = np.random.permutation(1000)[:25]\n\nparam_distributions = {\n 'coxphsurvivalanalysis__alpha': reciprocal(0.1, 100),\n}\n\n# alpha = model_random_search.best_params_['coxphsurvivalanalysis__alpha']\ncox_scores = []\nrsf_scores = []\nfor _seed in seeds:\n X_trn, X_test, y_trn, y_test = train_test_split(X, y, random_state=_seed)\n \n cox_best = make_pipeline(preprocessor, CoxPHSurvivalAnalysis())\n cox_best.fit(X_trn, y_trn)\n ci_cox = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], cox_best.predict(X_test))\n cox_scores.append(ci_cox[0])\n\n rsf = make_pipeline(preprocessor, RandomSurvivalForest(n_estimators=1000, \n min_samples_split=10,\n min_samples_leaf=15,\n max_features=\"sqrt\",\n n_jobs=-1,\n random_state=42))\n rsf.fit(X_trn, y_trn)\n ci_rsf = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], rsf.predict(X_test))\n rsf_scores.append(ci_rsf[0])\n# -\n\nprint(f'Cox proportional hazards model: Mean c-index: {np.mean(cox_scores):.3f}, standard deviation: {np.std(cox_scores):.3f}')\nprint(f'Random Survival Forest model: Mean c-index: {np.mean(rsf_scores):.3f}, standard deviation: {np.std(rsf_scores):.3f}')\n\n\nrsf = RandomSurvivalForest(n_estimators=1000,\n min_samples_split=10,\n min_samples_leaf=15,\n max_features=\"sqrt\",\n n_jobs=-1,\n random_state=random_state)\nrsf.fit(X_train,y_train)\n\npd.Series(rsf_best.predict(X_test))\n\nrsf_best.score(X_test, y_test)\n\nX_test\n\nX_test_sorted = X_test.sort_values(by=[\"age\", \"diasbp\"])\nX_test_sel = pd.concat((X_test_sorted.head(3), X_test_sorted.tail(3)))\nX_test_sel\n\npd.Series(rsf_best.predict(X_test_sel))\n\n# +\nfrom mpl_toolkits.mplot3d import Axes3D\nfrom matplotlib import pyplot as plt\nimport numpy as np\nfrom skimage import measure\n\n\n# Set up mesh\nn = 100 \n\nx = np.linspace(-3,3,n)\ny = np.linspace(-3,3,n)\nz = np.linspace(-3,3,n)\nX, Y, Z = np.meshgrid(x, y, z)\n\n# Create cardioid function \ndef f_heart(x,y,z):\n F = 320 * ((-x**2 * z**3 -9*y**2 * z**3/80) +\n (x**2 + 9*y**2/4 + z**2-1)**3)\n return F\n\n# Obtain value to at every point in mesh\nvol = f_heart(X,Y,Z) \n\n# Extract a 2D surface mesh from a 3D volume (F=0)\nverts, faces, normals, values = measure.marching_cubes_lewiner(vol, 0, spacing=(0.1, 0.1, 0.1))\n\n\n# Create a 3D figure\nfig = plt.figure(figsize=(12,8))\nax = fig.add_subplot(111, projection='3d')\n\n# Plot the surface\nax.plot_trisurf(verts[:, 0], verts[:,1], faces, verts[:, 2],\n cmap='Spectral', lw=1)\n\n# Change the angle of view and title\nax.view_init(15, -15)\n\n# ax.set_title(u\"Made with ❤ (and Python)\", fontsize=15) # if you have Python 3\nax.set_title(\"Made with <3 (and Python)\", fontsize=15)\n\n# Show me some love ^^\nplt.show()\n","repo_name":"rrwabina/RADI605","sub_path":"07 Random Survival Forests/scripts/rsf.ipynb","file_name":"rsf.ipynb","file_ext":"py","file_size_in_byte":10643,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"70487119511","text":"import numpy as np\nimport matplotlib.pyplot as plt\nimport plotutils as pu\nfrom sympy.mpmath import nsum, inf, mp\nfrom sympy import *\ninit_printing()\n# %matplotlib inline\n\n# The **Leibniz formula** states that:\n#\n# $$\n# 1 - \\frac{1}{3} + \\frac{1}{5} - \\frac{1}{7} + \\frac{1}{9} - \\ldots = \\frac{\\pi}{4}\n# $$\n#\n# So in other words:\n#\n# $$\n# \\sum^{\\infty}_{n = 0}\\frac{(-1)^n}{2n + 1} = \\frac{\\pi}{4}\n# $$\n#\n# Let's see if we can understand the sum formula first. What we'll do is just take the top part of the fraction $(-1)^n$ and the bottom part $2n + 1$ and plot them seperately for some values of $n$:\n\nwith plt.xkcd():\n x = np.arange(0, 10, 1)\n fig, axes = plt.subplots(1, 2, figsize=(10, 4))\n for ax in axes: pu.setup_axes(ax)\n axes[0].plot(x, (-1)**x, 'bo', zorder=10)\n axes[0].set_xlim(0, 9)\n axes[0].set_ylim(-5, 5)\n axes[1].plot(x, (2*x) + 1)\n axes[1].set_xlim(0, 9)\n axes[1].set_ylim(0, 10)\n\n# So as $n$ gets bigger and bigger we have two things. One flips between $1$ and $-1$ and the other is just a linear value $y = 2n + 1$ that just keeps on getting bigger and bigger. Now the equation above tells us that if we take an near infinite sum of these values we will get closer and closer to the value of $\\frac{\\pi}{4}$ so let's see if that's true.\n#\n# Below are two lines, one line represents $y = \\frac{(-1)^n}{2n + 1}$ and the other line is the sum of all the values of that equation for $y$ at $n = 0, 1, 2, \\ldots, n$. You can see that it (slowly) converges to some value, namely the value $\\frac{4}{\\pi}$.\n\nn = np.arange(0, 10, 1)\nf = lambda x: ((-1)**x) / (2*x + 1)\nwith plt.xkcd():\n fig, axes = plt.subplots(1, figsize=(8, 8))\n pu.setup_axes(axes, xlim=(-1, 9), ylim=(-0.5, 1.2), yticks=[1], yticklabels=[1], xticks=[1,2,3,4,5,6,7,8])\n plt.plot(n, f(n), zorder=10, label='THE INFINITE SERIES')\n plt.plot(n, [nsum(f, [0, n]) for n in n], label='SUMMATION OF THE SERIES')\n plt.annotate('THE LEIBNIZ FORMULA FOR PI', (1, 1))\n axes.set_aspect(4.0)\n axes.legend(loc=4)\n\n# Now if we sum up all the terms of that line above for $x = 0, 1, 2, 3, \\ldots, n$ we'll get closer and closer to $\\frac{4}{\\pi}$. Using `mpmath` we can calculate $\\pi$ with pretty good detail using the `mp.dps` setting to control the precision.\n\nleibniz = lambda n: ((-1)**n) / (2 * n + 1)\nmp.dps = 50\nnsum(leibniz, [0, inf]) * 4\n\n# Of course we can compute it symbolically as well. These fractions get pretty crazy real quickly.\n\nleibniz = S('((-1)^n)/(2*n+1)')\nn = S('n')\nsum([leibniz.subs(n, i) for i in range(100)])\n\n\n","repo_name":"basp/notes","sub_path":"leibniz_formula.ipynb","file_name":"leibniz_formula.ipynb","file_ext":"py","file_size_in_byte":2555,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"24682123021","text":"# +\nfrom sklearn import metrics\nfrom sklearn.linear_model import LogisticRegression\n\ndef rocplot(truth, pred, model):\n \"\"\"This function will roc curve of a given model and return its auc. \n This function is a slighly augmented copy of the plotUnivariateROC from Lab_FeatureRanking_AUC\"\"\"\n fpr, tpr, thresholds = metrics.roc_curve(truth, pred)\n roc_auc = metrics.auc(fpr, tpr)\n \n if roc_auc < 0.5:\n fpr, tpr, thresholds = metrics.roc_curve(truth, -1 * pred)\n roc_auc = metrics.auc(fpr, tpr)\n \n c = (np.random.rand(), np.random.rand(), np.random.rand())\n plt.plot(fpr, tpr, color = c, label = model + ' (AUC = %0.3f)' % roc_auc)\n \n\n plt.plot([0, 1], [0, 1], 'k--')\n plt.xlim([0.0, 1.0])\n plt.ylim([0.0, 1.0])\n plt.xlabel('FPR')\n plt.ylabel('TPR')\n plt.title('ROC')\n plt.legend(loc=\"lower right\")\n \n return roc_auc\n\n\n\n\n# +\nimport pandas as pd\nimport numpy as np\nimport scipy as sp\nimport matplotlib.pyplot as plt\nimport math\nfrom sklearn.tree import DecisionTreeClassifier\nfrom sklearn.tree import export_graphviz\nfrom sklearn.metrics import confusion_matrix,roc_auc_score\n# %matplotlib inline\n\ntrain = pd.read_csv('data/cleaned_mostRecStat_train.csv',header=0,encoding='iso-8859-1')\ntrain_y = train['is_placed']\ntrain = train.drop(\"is_placed\",1)\ntrain = train.drop(\"in_process\",1)\n\n\n\ntest = pd.read_csv('data/cleaned_mostRecStat_test.csv',header=0,encoding='iso-8859-1')\ntest_y = test['is_placed']\ntest = test.drop(\"is_placed\",1)\ntest = test.drop(\"in_process\",1)\n\nlogreg = LogisticRegression(solver='liblinear')\nlogreg = logreg.fit(train, train_y)\npred = logreg.predict_proba(test)[:, 1]\nrocplot(test_y, pred, \"Logistic Regression\")\ntrain.columns\nscore = logreg.score(train, train_y)\n\nprint(score)\n\n# +\nfrom sklearn.feature_selection import SelectFromModel\nfeature_sel = SelectFromModel(logreg, prefit = True)\ntrain_new = feature_sel.transform(train)\ntest_new = feature_sel.transform(test)\nnew_logreg = LogisticRegression(C = 1)\nnew_logreg = new_logreg.fit(train_new, train_y)\nnew_pred = new_logreg.predict_proba(test_new)[:, 1]\nrocplot(test_y, new_pred, \"Logistic Regression\")\n\nfeatures = train.columns.values\nselected_features = features[feature_sel.get_support()]\nselected_features\n# -\n\nAUCs = []\nfig = plt.figure(figsize = (12, 6))\nax = plt.subplot(111)\nfor feature in selected_features:\n auc = rocplot(train_y, train[feature], feature)\n AUCs.append((feature, auc))\nbox = ax.get_position()\nax.set_position([box.x0, box.y0 + box.height * 0.0 , box.width, box.height * 1])\nlgd = ax.legend(loc = 'upper center', bbox_to_anchor = (0.5, -0.15), fancybox = True,\n shadow = True, ncol = 4, prop = {'size':10})\n\nAUCs.sort(key=lambda tup: tup[1])\nAUCs[-10:]\n\nlogreg.score(test, test_y)\n\n# +\n\nlogreg2 = LogisticRegression(solver='liblinear', class_weight = 'balanced')\nlogreg2 = logreg2.fit(train, train_y)\npred = logreg2.predict_proba(test)[:, 1]\nrocplot(test_y, pred, \"Logistic Regression\")\ntrain.columns\nscore = logreg2.score(test, test_y)\n\nprint(score)\n\n# +\nfrom sklearn.feature_selection import SelectFromModel\nfeature_sel = SelectFromModel(logreg2, prefit = True)\ntrain_new = feature_sel.transform(train)\ntest_new = feature_sel.transform(test)\nnew_logreg2 = LogisticRegression(solver='liblinear', class_weight = 'balanced')\nnew_logreg2 = new_logreg2.fit(train_new, train_y)\nnew_pred = new_logreg2.predict_proba(test_new)[:, 1]\nrocplot(test_y, new_pred, \"Logistic Regression\")\n\nfeatures = train.columns.values\nselected_features = features[feature_sel.get_support()]\nselected_features\n\n# +\nAUCs = []\nfig = plt.figure(figsize = (12, 6))\nax = plt.subplot(111)\nfor feature in selected_features:\n auc = rocplot(train_y, train[feature], feature)\n AUCs.append((feature, auc))\nbox = ax.get_position()\nax.set_position([box.x0, box.y0 + box.height * 0.0 , box.width, box.height * 1])\nlgd = ax.legend(loc = 'upper center', bbox_to_anchor = (0.5, -0.15), fancybox = True,\n shadow = True, ncol = 4, prop = {'size':10})\n\nAUCs.sort(key=lambda tup: tup[1])\nAUCs[-10:]\n# -\n\n\n","repo_name":"jls731/infrep17project","sub_path":"Feature Importance.ipynb","file_name":"Feature Importance.ipynb","file_ext":"py","file_size_in_byte":4075,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"26789673648","text":"# # (Bayesian) Naive Bayes algorithm\n\nimport pymc3 as pm\nimport theano as tt\nimport numpy as np\nfrom sklearn import datasets\nfrom sklearn.model_selection import train_test_split\n# %matplotlib inline\n\n# ## Definition\n\n# The Naive Bayes algorithm is a classification algorithm that is often used in topic classification for text data. Imagine a situation where we have $n_{features}$ documents, each represented by a vector $\\vec{x}$ where $x_i$ is the number of times that the word $i$ appears in the document (bag-of-word representation). The quantity we want to estimate is the probability that the document $i$ belongs to a class $c$ given the data $\\mathcal{D}$:\n#\n# $$P( y = c | \\mathbf{x}, \\mathcal{D})$$\n#\n# Following `Bayes` rule we can write:\n#\n# $$P(y = c | \\mathbf{x}, \\mathcal{D}) \\sim P(\\mathbf{x}| y = c, \\mathcal{D})\\;P(y=c|\\mathcal{D})$$\n#\n# The `Naive` part of the algorithm is to write that the random variables describing each component of the vector $\\mathbf{x}$ are independant:\n#\n# $$P(\\mathbf{x}|y=c) = \\prod_{j=1}^{n_{dims}} P(x_j | y=c)$$\n#\n# This, of course, is a coarse approximation: it is--for instance--more likely that the word `computer` will occur often in a document that contains the word `email` often than the word `leek`. There are correlations between the different components. But, surprisingly, this simplified model is very effective.\n\n# ## Generative Model\n#\n# ### Description\n#\n# Statistically, for continuous data, the Naive Bayes model can be defined as: \n#\n# $$\\mathbf{\\alpha} = [1, \\dots, 1]$$\n# $$\\mathbf{\\pi} \\sim Dirichlet\\left(\\mathbf{\\alpha}\\right)$$\n# $$y \\sim Cat(\\pi)$$\n# $$x_{jc} \\sim Normal\\left(\\mu_{jc}, \\sigma_{jc}\\right)$$\n#\n# This is the statistical translation of the following *generative* story: to get the feature $j$ of a vector $\\mathbf{x}$ (the number of times the word $j$ appears in document $\\mathbf{x}$), we choose the category (topic) $c$ to which $\\mathbf{x}$ belongs with probability $\\pi_c$ and then sample $x_j$ from a normal distribution of mean $\\mu_{jc}$ and standard deviation $\\sigma_{jc}$.\n#\n# The model can be summarized by the following graphical representation:\n#\n# \n#\n# It is difficult to read in the previous figure tha\n\n# ### Implementation\n#\n# To get a sense of how the implementation of `GaussianNaiveBayes` works in PyMC3-models, let us show the implementation in PyMC3. We use the full iris dataset available in `scikit-learn` to train the model.\n\niris = datasets.load_iris()\n\n# +\n# Define as theano shared variables so the value can be changed later on\nX = tt.shared(iris.data)\ny = tt.shared(iris.target)\n\nn_dims = iris.data.shape[1]\nn_classes = len(set(iris.target))\nn_features = iris.data.shape[0]\n# -\n\nwith pm.Model() as model:\n # Priors\n alpha = np.ones(n_classes)\n pi = pm.Dirichlet('pi', alpha, shape=n_classes)\n mu = pm.Normal('mu', 0, 100, shape=(n_classes, n_dims))\n sigma = pm.HalfNormal('sigma', 100, shape=(n_classes, n_dims))\n \n # Assign class to data points\n z = pm.Categorical('z', pi, shape=n_features, observed=y)\n \n # The components are independent and normal-distributed\n a = pm.Normal('a', mu[z], sigma[z], observed=X)\n\n# %timeit\nwith model:\n trace = pm.sample(5000)\n\n_ = pm.traceplot(trace)\n\n# ### Prediction time!\n\n# Prediction for Bayesian models are slightly different from their frequentist counterpart, they require to evaluate many integrals over the parameters. Given a data point $\\mathbf{x} = {x_j}$, we would like to estimate the probability that it belongs to any category $c$. Following the notations in [1], the probability reads:\n#\n# $$\n# P(y=c | \\mathbf{x}, \\mathcal{D}) = P(y=c|\\mathcal{D}) \\prod_{j=1}^{dims} P(x_j | y=c, \\mathcal{D})\n# $$\n#\n# The data $\\mathcal{D}$ helped us estimate the distribution of the parameter $\\mathbf{\\mu}$, $\\mathbf{\\pi}$ and $\\mathbf{\\sigma}$. We now need to integrate over the values of these parameters:\n#\n# $$\n# P(y=c | \\mathbf{x}, \\mathbf{D}) = \\left[\\int Cat(y=c|\\pi) P(\\pi|\\mathcal{D}) \\mathrm{d}\\mathbf{\\pi}\\right] \\prod_{j=1}^{dims} \\int Normal(x_j | \\mu_{jc}, \\sigma_{jc}) P(\\mu_{jc}|\\mathcal{D}) P(\\sigma_{jc} | \\mathcal{D}) \\mathrm{d} \\mu_{jc} \\mathrm{d} \\sigma_{jc}\n# $$\n#\n# which can be evaluated from the samples we previously computed (show).\n\n# +\nimport itertools as it\nimport operator\nimport functools as ft\nimport scipy.stats\n\ndef normalize(array):\n \"\"\" Normalize the number in an array to get probabilities.\n \"\"\"\n return array/np.sum(array)\n \n\ndef predict(trace, X):\n \"\"\" Predict the probabilities that the data points belong to a class. \"\"\"\n n_dims = X.shape[1]\n \n predictions = np.array([])\n for x in X:\n normal_probs = scipy.stats.norm(trace['mu'], trace['sigma']).pdf(x)\n per_dim = [np.sum(normal_probs[:,:,i], axis=0)/len(trace['mu']) for i in range(n_dims)]\n a = normalize(ft.reduce(lambda x,y: x*y, per_dim))\n a[np.abs(a) < 0.00001] = 0\n if len(predictions) == 0:\n predictions = a\n else:\n predictions = np.vstack((predictions,a))\n return predictions\n\npred = predict(trace, iris.data) # Probability the the input belongs to each class\npred_class = np.argmax(pred, axis=1) # Class to which the input is predicted to belong (zero-one loss)\naccuracy = sum(np.equal(pred_class, iris.target))/len(iris.data) # Accuracy of the predictions\n# -\n\n# ## Now with pymc3_models\n#\n# ### Features\n#\n# Let us now re-do the analysis with the `GaussianNaiveBayes` class, and show its features.\n\n# +\n# Import the library\nimport os\nimport sys\n\nmodule_path = os.path.abspath(os.path.join('..'))\nif module_path not in sys.path:\n sys.path.append(module_path)\nfrom pymc3_models.models.NaiveBayes import GaussianNaiveBayes\n# -\n\n# First create the model:\n\nN = GaussianNaiveBayes()\n\n# Then fit (still using the Iris data). `pymc3_models` uses ADVI inference by default, but you can switch to NUTS by passing `inference_type='nuts'` as a parameter to the `fit` function.\n\n# +\n# Load the data and split in train and test set\niris = datasets.load_iris()\nX = iris.data\ny = iris.target\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)\n\n# Fit with the training data\nN.fit(X_train, y_train)\n# -\n\n# Now you can predict the probabilities that data points belong to each category using the zero-one cost function:\n\npred_prob = N.predict_proba(X_test)\npred = N.predict(X_test)\n\n# And finally, you can compute the accuracy score:\n\nscore = N.score(X_test, y_test)\nprint(score)\n","repo_name":"parsing-science/pymc3_models","sub_path":"notebooks/NaiveBayes.ipynb","file_name":"NaiveBayes.ipynb","file_ext":"py","file_size_in_byte":6549,"program_lang":"python","lang":"en","doc_type":"code","stars":156,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"35506960086","text":"# # Developing an AI application\n#\n# Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications. \n#\n# In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below. \n#\n#
\n\n# + id=\"S60eHj8Eecoz\"\nfrom tensorflow.keras.layers import Dense,SimpleRNN,Embedding,Flatten\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.datasets import imdb\n\n# + id=\"XKaFH9BOeuPy\"\n(xtrain,ytrain),(xtest,ytest) = imdb.load_data()\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"zT8GmbsZe718\" outputId=\"a11d66d8-cbe5-42b2-d9da-2b807e73b57b\"\nxtrain\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"WhvEZczsfGgw\" outputId=\"1b9abeff-3544-40ad-bfd5-d907bae5cac5\"\nxtrain.shape\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"WHED7k0DfYSQ\" outputId=\"e3792a15-09f9-4417-bd56-730adb1d41b3\"\nxtest.shape\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"2ISo0IZsfavd\" outputId=\"ac94ea57-dd7e-49a1-cc0f-cb86a1d94f35\"\nxtrain[100]\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"kWmSJcFjfnWo\" outputId=\"63920c58-3ea2-4ec4-dc43-b992da9b5f81\"\nlen(xtrain[0])\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"z5FRZt7Ufxc8\" outputId=\"880b5280-6c12-43bf-cb8c-fa65f84fc67f\"\nlen(xtrain[10])\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"zADUWbbxfzOA\" outputId=\"471633ad-e794-49fb-fd57-cdc3091f6e9d\"\nlen(xtrain[25])\n\n# + id=\"A10lBHq3f3Ue\"\n# same sequence\nfrom tensorflow.keras.utils import pad_sequences\nxtrain = pad_sequences(xtrain,padding='post',maxlen=50)\nxtest = pad_sequences(xtest,padding='post',maxlen=50)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"VNcuTAFXgnKX\" outputId=\"16768bef-af33-4591-a278-9631a1203117\"\nxtrain.shape\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"dHP_3LLqgunv\" outputId=\"3e05961e-adb5-4d80-f0ef-21eee26288e3\"\nxtrain[0]\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"Yp7R49ZLgy5B\" outputId=\"79ed4a10-3763-450b-a1de-142cb6222240\"\nmodel = Sequential()\n\nmodel.add(SimpleRNN(32,activation='tanh',input_shape=(50,1),return_sequences=False))\n\nmodel.add(Dense(1,activation='sigmoid'))\n\nmodel.summary()\n\n# + id=\"Xstha28NhXho\"\nmodel.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"lSv7bOSoioCP\" outputId=\"a3f8e9d8-7cde-4057-a664-1a8a85ab4a2e\"\nhistory = model.fit(xtrain,ytrain, validation_data=(xtest,ytest),epochs=1)\n\n# + id=\"s_MCOwzoiyz-\"\n\n","repo_name":"adityadhumal06/RNN-imdb-Project","sub_path":"RNN_imdb_dataset.ipynb","file_name":"RNN_imdb_dataset.ipynb","file_ext":"py","file_size_in_byte":2504,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"74073385752","text":"# 1. 데이터 벡터 다루기\n#\n# [예제] A 회사의 2010년 월별매출액 데이터가 다음과 같이 주어졌다고 하자.\n# \n# 100, 120, 130, 124, 150, 167, 170, 163, 160, 155, 145, 157\n# \n# 지수(index)를 이용하여 데이터벡터를 다루어 보기로 한다.\n\nimport numpy as np\nx=np.array([100, 120, 130, 124, 150, 167, 170, 163, 160, 155, 145, 157])\nx\n\nx[1]\n\nx[1:3]\n\nx[1:4]\n\nx[-1]\n\nx[-3]\n\nx[1:-3]\n\nx[7:]\n\nx[:3]\n\nx[::2]\n\nx[1:4:2]\n\nx>2\n\nx2=np.array([[1,2,3,4],[5,6,7,8],[9,10,11,12]])\n\nx2\n\nx2[0]\n\nx2[0,]\n\nx2[0,:]\n\nx2[0,1]\n\nx2[:2,1]\n\nx2[:2,]\n\nx2>2\n\nx2[x2>0]\n\n# + active=\"\"\n# 1. 순서함수 활용\n# -\n\n#데이터 프레임 생성\nimport pandas as pd\ndf=pd.DataFrame({\n 'name':['Kim','Lee','Park','Bae','Kang'],\n 'age': [24,32,43,24,30],\n 'height':[160,170,165,173,158]\n})\ndf\n\n# +\n#나이를 기준으로 순위를 주어. rank_average 변수 생성 -> rank() 사용\n\ndf['rank_average']=df['age'].rank(method='min', ascending=False) #method='min': 동점일 경우 최솟값을 줌\ndf #ascending=False: 내림차순으로 순위를 줌 (값이 클수록 높은 순위)\n# -\n\ndf['age'].rank(method='max',ascending=False) #method='max' 동점일 경우 최댓값을 줌\n\ndf['age'].rank(method='average',ascending=False) #method='average' 동점일 경우 평균을 줌 default\n\ndf['age'].rank(ascending=False)\n\n# +\n#결측값 포함한 데이터 프레임인 경우\n\ndf2=pd.DataFrame({\n 'name':['Kim','Lee','Park','Bae','Kang'],\n 'age':[24,32,43,24,np.nan],\n 'height':[160,170,165,173,158]\n})\ndf2\n\n# +\n#결측값 처리 위한 옵션 na_option 추가\n\ndf2['rank_average']=df2['age'].rank(method='min', ascending=False, na_option='bottom')\n#na_option='bottom': 결측값에 후순위를 주고자 할 경우\n#na_option='top': 결측값에 선순위를 주고자 할 경우\n#na_option='keep': 결측값을 NaN으로 두고자 할 경우 default\ndf2\n\n# +\n#데이터 프레임 df에서 name변수 기준으로 오름차순 정렬하기\n\ndf_sort=df.sort_values(by='name')\ndf_sort\n# -\n\n#데이터 프레임 df에서 age 변수 기준으로 오름차순 정렬하기\ndf_sort=df.sort_values(by='age')\ndf_sort\n\n# +\n#데이터 프레임 df에서 height 변수 기준으로 내림차순으로 정렬하기\n\ndf_sort=df.sort_values(by='height', ascending=False)\ndf_sort\n# -\n\n#데이터 프레임 df에서 두 변수 age, height 변수 기준으로 오름차순으로 정렬하기\ndf_sort=df.sort_values(by=['age','height'])\ndf_sort\n\n# + active=\"\"\n# 1. 결측값\n\n# +\n#데이터 프레임 df2에 결측값 포함 여부 확인하기\n\ndf2.isnull().values.any() #True:결측값 포함\n\n# +\n#데이터 프레임 df2 결측값에 0으로 대체하기\n\ndf2_0=df2.fillna(0)\ndf2_0\n# -\n\n#결측값을 변수별 평균으로 대체\ndf_mean=df2.fillna(df2.mean(numeric_only=True))\ndf_mean\n\n# +\n#순서상 결측값의 바로 앞의 값으로 대체하기\n\ndf2_f=df2.fillna(method='ffill')\ndf2_f\n\n# +\n#순서상 결측값의 바로 뒤의 값으로 대체하기\n\ndf2_f=df2.fillna(method='backfill')\ndf2_f\n# -\n\ndf3=df2.sort_values(by='height')\ndf3\n\ndf3_f=df3.fillna(method='backfill')\ndf3_f\n\ndf3_f2=df2.sort_values(by='height').fillna(method='backfill')\ndf3_f2\n\n\n\n\n\ndf2.sort_values(by='height').fillna(method='backfill')[['name','age']]\n\n\n","repo_name":"jpark630/stat-software","sub_path":"Statistitcal Software Week 13 practice.ipynb","file_name":"Statistitcal Software Week 13 practice.ipynb","file_ext":"py","file_size_in_byte":3335,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"18226709526","text":"# +\nimport numpy as np \nimport pandas as pd\nimport matplotlib.pyplot as plt \nimport export as export \nimport eli5\nimport warnings\nwarnings.filterwarnings('ignore')\n\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.preprocessing import OrdinalEncoder\nfrom sksurv.datasets import load_gbsg2\nfrom sksurv.preprocessing import OneHotEncoder\nfrom sksurv.ensemble import RandomSurvivalForest\nfrom sksurv.datasets import load_whas500\nfrom sksurv.tree import SurvivalTree\nfrom sksurv.util import Surv\n# -\n\nX, y = load_whas500()\nX = X.astype(float)\nestimator = SurvivalTree().fit(X, y)\n\nexport.plot_tree(estimator, feature_names = estimator.feature_names_in_, impurity = False,)\n\n# +\npredict_outcomes = estimator.predict(X)\nsurv_funcs = estimator.predict_survival_function(X.iloc[:5])\n\nfor fn in surv_funcs:\n plt.step(fn.x, fn(fn.x), where=\"post\")\nplt.ylim(0,1)\nplt.show()\n\n# +\nchf_funcs = estimator.predict_cumulative_hazard_function(X.iloc[:5])\n\nfor fn in chf_funcs:\n plt.step(fn.x, fn(fn.x), where=\"post\")\nplt.ylim(0,1)\nplt.show()\n\n\n# +\nX,y = load_gbsg2()\ngrade_str = X.loc[:, \"tgrade\"].astype(object).values[:,np.newaxis]\ngrade_num = OrdinalEncoder(categories=[[\"I\", \"II\", \"III\"]]).fit_transform(grade_str)\nX_no_grade = X.drop(\"tgrade\",axis=1)\nXt = OneHotEncoder().fit_transform(X_no_grade)\nXt.loc[:, \"tgrade\"] = grade_num\n\n\nrandom_state = 20\nX_train, X_test,y_train,y_test = train_test_split(Xt, y, test_size=0.25, random_state=random_state)\n\nrsf = RandomSurvivalForest(n_estimators = 1000, \n min_samples_split = 10,\n min_samples_leaf = 15,\n max_features = \"sqrt\",\n n_jobs = -1,\n random_state = random_state)\n\nrsf.fit(X_train,y_train)\nrsf.score(X_test, y_test)\n# -\n\nX_test_sorted = X_test.sort_values(by=[\"pnodes\", \"age\"])\nX_test_sel = pd.concat((X_test_sorted.head(3), X_test_sorted.tail(3)))\nX_test_sel\n\npd.Series(rsf.predict(X_test_sel))\n\n\nsurv = rsf.predict_survival_function(X_test_sel, return_array=True)\nfor i,s in enumerate(surv):\n plt.step(rsf.event_times_,s, where=\"post\", label=str(i))\nplt.ylabel(\"Survivalprobability\")\nplt.xlabel(\"Timeindays\")\nplt.legend()\nplt.grid(True)\n\nsurv = rsf.predict_cumulative_hazard_function(X_test_sel, return_array=True)\nfor i,s in enumerate(surv):\n plt.step(rsf.event_times_,s, where=\"post\", label=str(i))\nplt.ylabel(\"Cumulativehazard\")\nplt.xlabel(\"Timeindays\")\nplt.legend()\nplt.grid(True)\n\n# +\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nfrom sksurv.datasets import load_whas500\n\nX, y = load_whas500()\n# -\n\nX.head(10).style.hide_index()\n\ny[:10]\n\ndf_y = pd.DataFrame(data={'time': [y[i][1] for i in range(len(y))], 'event': [y[i][0] for i in range(len(y))]})\ndf_y[:10].style.hide_index().highlight_min('event', color='lightgreen')\n\n# One of the main challenges of survival analysis is right censoring, i.e., by the end of the study, the event of interest (for example, in medicine 'death of a patient' or in this dataset 'recurrence of cancer') has only occurred for a subset of the observations.\n#\n# The right censoring in this dataset is given by the column named 'event' and it's a variable which can take value 'True' if the patient had a recurrence of cancer or 'False' if the patient is recurrence free at the indicated time (right-censored samples).\n#\n# Let's see how many right-censored samples do we have.\n\nprint(f'Number of samples: {len(df_y)}')\nprint(f'Number of right censored samples: {len(df_y.query(\"event == False\"))}')\nprint(f'Percentage of right censored samples: {100*len(df_y.query(\"event == False\"))/len(df_y):.1f}%')\n\n# There are 387 patients (56.4%) who were right censored (recurrence free) at the end of the study.\n#\n# Let's divide our dataset in training and test sets.\n\n# +\nfrom sklearn.model_selection import train_test_split\n\nX_trn, X_test, y_trn, y_test = train_test_split(X, y, test_size = 0.30, random_state = 42)\n\nprint(f'Number of training samples: {len(y_trn)}')\nprint(f'Number of test samples: {len(y_test)}')\n# -\n\n# We divide the features/covariates into continuous and categorical.\n\nscaling_cols = [c for c in X.columns if X[c].dtype.kind in ['i', 'f']]\ncat_cols = [c for c in X.columns if X[c].dtype.kind not in [\"i\", \"f\"]]\n\n# We use ordinal encoding for categorical features/covariates and standard scaling for continuous features/covariates.\n\n# +\nfrom sklearn.compose import ColumnTransformer\nfrom sklearn.preprocessing import OrdinalEncoder\nfrom sklearn.preprocessing import StandardScaler\n\npreprocessor = ColumnTransformer(\n [('cat-preprocessor', OrdinalEncoder(), cat_cols),\n ('standard-scaler', StandardScaler(), scaling_cols)],\n remainder='passthrough', sparse_threshold=0)\n\n# +\nfrom sksurv.ensemble import RandomSurvivalForest\nfrom sklearn.pipeline import make_pipeline\nfrom sksurv.linear_model import CoxPHSurvivalAnalysis\nfrom sksurv.metrics import concordance_index_censored\nfrom scipy.stats import reciprocal\nfrom sklearn.model_selection import RandomizedSearchCV\n\nrsf = make_pipeline(preprocessor, RandomSurvivalForest(random_state=42))\nrsf.fit(X_trn, y_trn)\nci_rsf = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], rsf.predict(X_test))\nprint(f'The c-index of Random Survival Forest is given by {ci_rsf[0]:.3f}')\n\n\n# +\nclass reciprocal_int:\n def __init__(self, a, b):\n self._distribution = reciprocal(a, b)\n\n def rvs(self, *args, **kwargs):\n return self._distribution.rvs(*args, **kwargs).astype(int)\n\nparam_distributions1 = {\n 'randomsurvivalforest__max_features': reciprocal_int(3, 8),\n 'randomsurvivalforest__max_depth': reciprocal_int(5, 50),\n 'randomsurvivalforest__min_samples_leaf': reciprocal_int(1, 40),\n}\n\nmodel_random_search = RandomizedSearchCV(\n rsf, param_distributions=param_distributions1, n_iter=50, n_jobs=-1, cv=3, random_state=0)\nmodel_random_search.fit(X_trn, y_trn)\n\nprint(\n f\"The c-index of random survival forest using a {model_random_search.__class__.__name__} is \"\n f\"{model_random_search.score(X_test, y_test):.3f}\")\nprint(\n f\"The best set of parameters is: {model_random_search.best_params_}\"\n)\n# -\n\nmax_depth = model_random_search.best_params_['randomsurvivalforest__max_depth']\nmax_features = model_random_search.best_params_['randomsurvivalforest__max_features']\nmin_samples_leaf = model_random_search.best_params_['randomsurvivalforest__min_samples_leaf']\n\nrsf_best = make_pipeline(preprocessor, RandomSurvivalForest(random_state=42, max_depth=max_depth, max_features=max_features, min_samples_leaf=min_samples_leaf))\nrsf_best.fit(X_trn, y_trn)\nci_rsf = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], rsf_best.predict(X_test))\nprint(f'The c-index of Random Survival Forest is given by {ci_rsf[0]:.3f}')\n\n# +\nfrom eli5.sklearn import PermutationImportance\nperm = PermutationImportance(rsf.steps[-1][1], n_iter=100).fit(preprocessor.fit_transform(X_trn),y_trn)\n\ndata = perm.results_\ndata = pd.DataFrame(data, columns=X_trn.columns)\nmeds = data.median()\nmeds = meds.sort_values(ascending=False)\ndata = data[meds.index]\n# -\n\nfig, ax = plt.subplots(figsize=(10,7))\ndata.boxplot(ax=ax)\nax.set_title('Feature Importances')\nplt.show()\n\neli5.show_weights(perm, feature_names = X.columns.tolist())\n\n# +\ncox = make_pipeline(preprocessor, CoxPHSurvivalAnalysis())\ncox.fit(X_trn, y_trn)\n\nci_cox = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], cox.predict(X_test))\nprint(f'The c-index of Cox is given by {ci_cox[0]:.3f}')\n\n# +\nnp.random.seed(0)\nseeds = np.random.permutation(1000)[:25]\n\nparam_distributions = {\n 'coxphsurvivalanalysis__alpha': reciprocal(0.1, 100),\n}\n\n# alpha = model_random_search.best_params_['coxphsurvivalanalysis__alpha']\ncox_scores = []\nrsf_scores = []\nfor _seed in seeds:\n X_trn, X_test, y_trn, y_test = train_test_split(X, y, random_state=_seed)\n \n cox_best = make_pipeline(preprocessor, CoxPHSurvivalAnalysis())\n cox_best.fit(X_trn, y_trn)\n ci_cox = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], cox_best.predict(X_test))\n cox_scores.append(ci_cox[0])\n\n rsf = make_pipeline(preprocessor, RandomSurvivalForest(n_estimators=1000, \n min_samples_split=10,\n min_samples_leaf=15,\n max_features=\"sqrt\",\n n_jobs=-1,\n random_state=42))\n rsf.fit(X_trn, y_trn)\n ci_rsf = concordance_index_censored(y_test[\"fstat\"], y_test[\"lenfol\"], rsf.predict(X_test))\n rsf_scores.append(ci_rsf[0])\n# -\n\nprint(f'Cox proportional hazards model: Mean c-index: {np.mean(cox_scores):.3f}, standard deviation: {np.std(cox_scores):.3f}')\nprint(f'Random Survival Forest model: Mean c-index: {np.mean(rsf_scores):.3f}, standard deviation: {np.std(rsf_scores):.3f}')\n\n\nrsf = RandomSurvivalForest(n_estimators=1000,\n min_samples_split=10,\n min_samples_leaf=15,\n max_features=\"sqrt\",\n n_jobs=-1,\n random_state=random_state)\nrsf.fit(X_train,y_train)\n\npd.Series(rsf_best.predict(X_test))\n\nrsf_best.score(X_test, y_test)\n\nX_test\n\nX_test_sorted = X_test.sort_values(by=[\"age\", \"diasbp\"])\nX_test_sel = pd.concat((X_test_sorted.head(3), X_test_sorted.tail(3)))\nX_test_sel\n\npd.Series(rsf_best.predict(X_test_sel))\n\n# +\nfrom mpl_toolkits.mplot3d import Axes3D\nfrom matplotlib import pyplot as plt\nimport numpy as np\nfrom skimage import measure\n\n\n# Set up mesh\nn = 100 \n\nx = np.linspace(-3,3,n)\ny = np.linspace(-3,3,n)\nz = np.linspace(-3,3,n)\nX, Y, Z = np.meshgrid(x, y, z)\n\n# Create cardioid function \ndef f_heart(x,y,z):\n F = 320 * ((-x**2 * z**3 -9*y**2 * z**3/80) +\n (x**2 + 9*y**2/4 + z**2-1)**3)\n return F\n\n# Obtain value to at every point in mesh\nvol = f_heart(X,Y,Z) \n\n# Extract a 2D surface mesh from a 3D volume (F=0)\nverts, faces, normals, values = measure.marching_cubes_lewiner(vol, 0, spacing=(0.1, 0.1, 0.1))\n\n\n# Create a 3D figure\nfig = plt.figure(figsize=(12,8))\nax = fig.add_subplot(111, projection='3d')\n\n# Plot the surface\nax.plot_trisurf(verts[:, 0], verts[:,1], faces, verts[:, 2],\n cmap='Spectral', lw=1)\n\n# Change the angle of view and title\nax.view_init(15, -15)\n\n# ax.set_title(u\"Made with ❤ (and Python)\", fontsize=15) # if you have Python 3\nax.set_title(\"Made with <3 (and Python)\", fontsize=15)\n\n# Show me some love ^^\nplt.show()\n","repo_name":"rrwabina/RADI605","sub_path":"07 Random Survival Forests/scripts/rsf.ipynb","file_name":"rsf.ipynb","file_ext":"py","file_size_in_byte":10643,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"70487119511","text":"import numpy as np\nimport matplotlib.pyplot as plt\nimport plotutils as pu\nfrom sympy.mpmath import nsum, inf, mp\nfrom sympy import *\ninit_printing()\n# %matplotlib inline\n\n# The **Leibniz formula** states that:\n#\n# $$\n# 1 - \\frac{1}{3} + \\frac{1}{5} - \\frac{1}{7} + \\frac{1}{9} - \\ldots = \\frac{\\pi}{4}\n# $$\n#\n# So in other words:\n#\n# $$\n# \\sum^{\\infty}_{n = 0}\\frac{(-1)^n}{2n + 1} = \\frac{\\pi}{4}\n# $$\n#\n# Let's see if we can understand the sum formula first. What we'll do is just take the top part of the fraction $(-1)^n$ and the bottom part $2n + 1$ and plot them seperately for some values of $n$:\n\nwith plt.xkcd():\n x = np.arange(0, 10, 1)\n fig, axes = plt.subplots(1, 2, figsize=(10, 4))\n for ax in axes: pu.setup_axes(ax)\n axes[0].plot(x, (-1)**x, 'bo', zorder=10)\n axes[0].set_xlim(0, 9)\n axes[0].set_ylim(-5, 5)\n axes[1].plot(x, (2*x) + 1)\n axes[1].set_xlim(0, 9)\n axes[1].set_ylim(0, 10)\n\n# So as $n$ gets bigger and bigger we have two things. One flips between $1$ and $-1$ and the other is just a linear value $y = 2n + 1$ that just keeps on getting bigger and bigger. Now the equation above tells us that if we take an near infinite sum of these values we will get closer and closer to the value of $\\frac{\\pi}{4}$ so let's see if that's true.\n#\n# Below are two lines, one line represents $y = \\frac{(-1)^n}{2n + 1}$ and the other line is the sum of all the values of that equation for $y$ at $n = 0, 1, 2, \\ldots, n$. You can see that it (slowly) converges to some value, namely the value $\\frac{4}{\\pi}$.\n\nn = np.arange(0, 10, 1)\nf = lambda x: ((-1)**x) / (2*x + 1)\nwith plt.xkcd():\n fig, axes = plt.subplots(1, figsize=(8, 8))\n pu.setup_axes(axes, xlim=(-1, 9), ylim=(-0.5, 1.2), yticks=[1], yticklabels=[1], xticks=[1,2,3,4,5,6,7,8])\n plt.plot(n, f(n), zorder=10, label='THE INFINITE SERIES')\n plt.plot(n, [nsum(f, [0, n]) for n in n], label='SUMMATION OF THE SERIES')\n plt.annotate('THE LEIBNIZ FORMULA FOR PI', (1, 1))\n axes.set_aspect(4.0)\n axes.legend(loc=4)\n\n# Now if we sum up all the terms of that line above for $x = 0, 1, 2, 3, \\ldots, n$ we'll get closer and closer to $\\frac{4}{\\pi}$. Using `mpmath` we can calculate $\\pi$ with pretty good detail using the `mp.dps` setting to control the precision.\n\nleibniz = lambda n: ((-1)**n) / (2 * n + 1)\nmp.dps = 50\nnsum(leibniz, [0, inf]) * 4\n\n# Of course we can compute it symbolically as well. These fractions get pretty crazy real quickly.\n\nleibniz = S('((-1)^n)/(2*n+1)')\nn = S('n')\nsum([leibniz.subs(n, i) for i in range(100)])\n\n\n","repo_name":"basp/notes","sub_path":"leibniz_formula.ipynb","file_name":"leibniz_formula.ipynb","file_ext":"py","file_size_in_byte":2555,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"24682123021","text":"# +\nfrom sklearn import metrics\nfrom sklearn.linear_model import LogisticRegression\n\ndef rocplot(truth, pred, model):\n \"\"\"This function will roc curve of a given model and return its auc. \n This function is a slighly augmented copy of the plotUnivariateROC from Lab_FeatureRanking_AUC\"\"\"\n fpr, tpr, thresholds = metrics.roc_curve(truth, pred)\n roc_auc = metrics.auc(fpr, tpr)\n \n if roc_auc < 0.5:\n fpr, tpr, thresholds = metrics.roc_curve(truth, -1 * pred)\n roc_auc = metrics.auc(fpr, tpr)\n \n c = (np.random.rand(), np.random.rand(), np.random.rand())\n plt.plot(fpr, tpr, color = c, label = model + ' (AUC = %0.3f)' % roc_auc)\n \n\n plt.plot([0, 1], [0, 1], 'k--')\n plt.xlim([0.0, 1.0])\n plt.ylim([0.0, 1.0])\n plt.xlabel('FPR')\n plt.ylabel('TPR')\n plt.title('ROC')\n plt.legend(loc=\"lower right\")\n \n return roc_auc\n\n\n\n\n# +\nimport pandas as pd\nimport numpy as np\nimport scipy as sp\nimport matplotlib.pyplot as plt\nimport math\nfrom sklearn.tree import DecisionTreeClassifier\nfrom sklearn.tree import export_graphviz\nfrom sklearn.metrics import confusion_matrix,roc_auc_score\n# %matplotlib inline\n\ntrain = pd.read_csv('data/cleaned_mostRecStat_train.csv',header=0,encoding='iso-8859-1')\ntrain_y = train['is_placed']\ntrain = train.drop(\"is_placed\",1)\ntrain = train.drop(\"in_process\",1)\n\n\n\ntest = pd.read_csv('data/cleaned_mostRecStat_test.csv',header=0,encoding='iso-8859-1')\ntest_y = test['is_placed']\ntest = test.drop(\"is_placed\",1)\ntest = test.drop(\"in_process\",1)\n\nlogreg = LogisticRegression(solver='liblinear')\nlogreg = logreg.fit(train, train_y)\npred = logreg.predict_proba(test)[:, 1]\nrocplot(test_y, pred, \"Logistic Regression\")\ntrain.columns\nscore = logreg.score(train, train_y)\n\nprint(score)\n\n# +\nfrom sklearn.feature_selection import SelectFromModel\nfeature_sel = SelectFromModel(logreg, prefit = True)\ntrain_new = feature_sel.transform(train)\ntest_new = feature_sel.transform(test)\nnew_logreg = LogisticRegression(C = 1)\nnew_logreg = new_logreg.fit(train_new, train_y)\nnew_pred = new_logreg.predict_proba(test_new)[:, 1]\nrocplot(test_y, new_pred, \"Logistic Regression\")\n\nfeatures = train.columns.values\nselected_features = features[feature_sel.get_support()]\nselected_features\n# -\n\nAUCs = []\nfig = plt.figure(figsize = (12, 6))\nax = plt.subplot(111)\nfor feature in selected_features:\n auc = rocplot(train_y, train[feature], feature)\n AUCs.append((feature, auc))\nbox = ax.get_position()\nax.set_position([box.x0, box.y0 + box.height * 0.0 , box.width, box.height * 1])\nlgd = ax.legend(loc = 'upper center', bbox_to_anchor = (0.5, -0.15), fancybox = True,\n shadow = True, ncol = 4, prop = {'size':10})\n\nAUCs.sort(key=lambda tup: tup[1])\nAUCs[-10:]\n\nlogreg.score(test, test_y)\n\n# +\n\nlogreg2 = LogisticRegression(solver='liblinear', class_weight = 'balanced')\nlogreg2 = logreg2.fit(train, train_y)\npred = logreg2.predict_proba(test)[:, 1]\nrocplot(test_y, pred, \"Logistic Regression\")\ntrain.columns\nscore = logreg2.score(test, test_y)\n\nprint(score)\n\n# +\nfrom sklearn.feature_selection import SelectFromModel\nfeature_sel = SelectFromModel(logreg2, prefit = True)\ntrain_new = feature_sel.transform(train)\ntest_new = feature_sel.transform(test)\nnew_logreg2 = LogisticRegression(solver='liblinear', class_weight = 'balanced')\nnew_logreg2 = new_logreg2.fit(train_new, train_y)\nnew_pred = new_logreg2.predict_proba(test_new)[:, 1]\nrocplot(test_y, new_pred, \"Logistic Regression\")\n\nfeatures = train.columns.values\nselected_features = features[feature_sel.get_support()]\nselected_features\n\n# +\nAUCs = []\nfig = plt.figure(figsize = (12, 6))\nax = plt.subplot(111)\nfor feature in selected_features:\n auc = rocplot(train_y, train[feature], feature)\n AUCs.append((feature, auc))\nbox = ax.get_position()\nax.set_position([box.x0, box.y0 + box.height * 0.0 , box.width, box.height * 1])\nlgd = ax.legend(loc = 'upper center', bbox_to_anchor = (0.5, -0.15), fancybox = True,\n shadow = True, ncol = 4, prop = {'size':10})\n\nAUCs.sort(key=lambda tup: tup[1])\nAUCs[-10:]\n# -\n\n\n","repo_name":"jls731/infrep17project","sub_path":"Feature Importance.ipynb","file_name":"Feature Importance.ipynb","file_ext":"py","file_size_in_byte":4075,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"26789673648","text":"# # (Bayesian) Naive Bayes algorithm\n\nimport pymc3 as pm\nimport theano as tt\nimport numpy as np\nfrom sklearn import datasets\nfrom sklearn.model_selection import train_test_split\n# %matplotlib inline\n\n# ## Definition\n\n# The Naive Bayes algorithm is a classification algorithm that is often used in topic classification for text data. Imagine a situation where we have $n_{features}$ documents, each represented by a vector $\\vec{x}$ where $x_i$ is the number of times that the word $i$ appears in the document (bag-of-word representation). The quantity we want to estimate is the probability that the document $i$ belongs to a class $c$ given the data $\\mathcal{D}$:\n#\n# $$P( y = c | \\mathbf{x}, \\mathcal{D})$$\n#\n# Following `Bayes` rule we can write:\n#\n# $$P(y = c | \\mathbf{x}, \\mathcal{D}) \\sim P(\\mathbf{x}| y = c, \\mathcal{D})\\;P(y=c|\\mathcal{D})$$\n#\n# The `Naive` part of the algorithm is to write that the random variables describing each component of the vector $\\mathbf{x}$ are independant:\n#\n# $$P(\\mathbf{x}|y=c) = \\prod_{j=1}^{n_{dims}} P(x_j | y=c)$$\n#\n# This, of course, is a coarse approximation: it is--for instance--more likely that the word `computer` will occur often in a document that contains the word `email` often than the word `leek`. There are correlations between the different components. But, surprisingly, this simplified model is very effective.\n\n# ## Generative Model\n#\n# ### Description\n#\n# Statistically, for continuous data, the Naive Bayes model can be defined as: \n#\n# $$\\mathbf{\\alpha} = [1, \\dots, 1]$$\n# $$\\mathbf{\\pi} \\sim Dirichlet\\left(\\mathbf{\\alpha}\\right)$$\n# $$y \\sim Cat(\\pi)$$\n# $$x_{jc} \\sim Normal\\left(\\mu_{jc}, \\sigma_{jc}\\right)$$\n#\n# This is the statistical translation of the following *generative* story: to get the feature $j$ of a vector $\\mathbf{x}$ (the number of times the word $j$ appears in document $\\mathbf{x}$), we choose the category (topic) $c$ to which $\\mathbf{x}$ belongs with probability $\\pi_c$ and then sample $x_j$ from a normal distribution of mean $\\mu_{jc}$ and standard deviation $\\sigma_{jc}$.\n#\n# The model can be summarized by the following graphical representation:\n#\n# \n#\n# It is difficult to read in the previous figure tha\n\n# ### Implementation\n#\n# To get a sense of how the implementation of `GaussianNaiveBayes` works in PyMC3-models, let us show the implementation in PyMC3. We use the full iris dataset available in `scikit-learn` to train the model.\n\niris = datasets.load_iris()\n\n# +\n# Define as theano shared variables so the value can be changed later on\nX = tt.shared(iris.data)\ny = tt.shared(iris.target)\n\nn_dims = iris.data.shape[1]\nn_classes = len(set(iris.target))\nn_features = iris.data.shape[0]\n# -\n\nwith pm.Model() as model:\n # Priors\n alpha = np.ones(n_classes)\n pi = pm.Dirichlet('pi', alpha, shape=n_classes)\n mu = pm.Normal('mu', 0, 100, shape=(n_classes, n_dims))\n sigma = pm.HalfNormal('sigma', 100, shape=(n_classes, n_dims))\n \n # Assign class to data points\n z = pm.Categorical('z', pi, shape=n_features, observed=y)\n \n # The components are independent and normal-distributed\n a = pm.Normal('a', mu[z], sigma[z], observed=X)\n\n# %timeit\nwith model:\n trace = pm.sample(5000)\n\n_ = pm.traceplot(trace)\n\n# ### Prediction time!\n\n# Prediction for Bayesian models are slightly different from their frequentist counterpart, they require to evaluate many integrals over the parameters. Given a data point $\\mathbf{x} = {x_j}$, we would like to estimate the probability that it belongs to any category $c$. Following the notations in [1], the probability reads:\n#\n# $$\n# P(y=c | \\mathbf{x}, \\mathcal{D}) = P(y=c|\\mathcal{D}) \\prod_{j=1}^{dims} P(x_j | y=c, \\mathcal{D})\n# $$\n#\n# The data $\\mathcal{D}$ helped us estimate the distribution of the parameter $\\mathbf{\\mu}$, $\\mathbf{\\pi}$ and $\\mathbf{\\sigma}$. We now need to integrate over the values of these parameters:\n#\n# $$\n# P(y=c | \\mathbf{x}, \\mathbf{D}) = \\left[\\int Cat(y=c|\\pi) P(\\pi|\\mathcal{D}) \\mathrm{d}\\mathbf{\\pi}\\right] \\prod_{j=1}^{dims} \\int Normal(x_j | \\mu_{jc}, \\sigma_{jc}) P(\\mu_{jc}|\\mathcal{D}) P(\\sigma_{jc} | \\mathcal{D}) \\mathrm{d} \\mu_{jc} \\mathrm{d} \\sigma_{jc}\n# $$\n#\n# which can be evaluated from the samples we previously computed (show).\n\n# +\nimport itertools as it\nimport operator\nimport functools as ft\nimport scipy.stats\n\ndef normalize(array):\n \"\"\" Normalize the number in an array to get probabilities.\n \"\"\"\n return array/np.sum(array)\n \n\ndef predict(trace, X):\n \"\"\" Predict the probabilities that the data points belong to a class. \"\"\"\n n_dims = X.shape[1]\n \n predictions = np.array([])\n for x in X:\n normal_probs = scipy.stats.norm(trace['mu'], trace['sigma']).pdf(x)\n per_dim = [np.sum(normal_probs[:,:,i], axis=0)/len(trace['mu']) for i in range(n_dims)]\n a = normalize(ft.reduce(lambda x,y: x*y, per_dim))\n a[np.abs(a) < 0.00001] = 0\n if len(predictions) == 0:\n predictions = a\n else:\n predictions = np.vstack((predictions,a))\n return predictions\n\npred = predict(trace, iris.data) # Probability the the input belongs to each class\npred_class = np.argmax(pred, axis=1) # Class to which the input is predicted to belong (zero-one loss)\naccuracy = sum(np.equal(pred_class, iris.target))/len(iris.data) # Accuracy of the predictions\n# -\n\n# ## Now with pymc3_models\n#\n# ### Features\n#\n# Let us now re-do the analysis with the `GaussianNaiveBayes` class, and show its features.\n\n# +\n# Import the library\nimport os\nimport sys\n\nmodule_path = os.path.abspath(os.path.join('..'))\nif module_path not in sys.path:\n sys.path.append(module_path)\nfrom pymc3_models.models.NaiveBayes import GaussianNaiveBayes\n# -\n\n# First create the model:\n\nN = GaussianNaiveBayes()\n\n# Then fit (still using the Iris data). `pymc3_models` uses ADVI inference by default, but you can switch to NUTS by passing `inference_type='nuts'` as a parameter to the `fit` function.\n\n# +\n# Load the data and split in train and test set\niris = datasets.load_iris()\nX = iris.data\ny = iris.target\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)\n\n# Fit with the training data\nN.fit(X_train, y_train)\n# -\n\n# Now you can predict the probabilities that data points belong to each category using the zero-one cost function:\n\npred_prob = N.predict_proba(X_test)\npred = N.predict(X_test)\n\n# And finally, you can compute the accuracy score:\n\nscore = N.score(X_test, y_test)\nprint(score)\n","repo_name":"parsing-science/pymc3_models","sub_path":"notebooks/NaiveBayes.ipynb","file_name":"NaiveBayes.ipynb","file_ext":"py","file_size_in_byte":6549,"program_lang":"python","lang":"en","doc_type":"code","stars":156,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"35506960086","text":"# # Developing an AI application\n#\n# Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications. \n#\n# In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below. \n#\n#  \n#\n# The project is broken down into multiple steps:\n#\n# * Load and preprocess the image dataset\n# * Train the image classifier on your dataset\n# * Use the trained classifier to predict image content\n#\n# We'll lead you through each part which you'll implement in Python.\n#\n# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.\n#\n# First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.\n\n# +\n# Imports here\nimport numpy as np\nimport matplotlib.pyplot as plt\n\nimport torch\nimport torchvision\nfrom torchvision import datasets, transforms, models\nfrom torch import nn\nfrom torch import optim\n#import torch.nn.functional as F\nfrom torch.optim import lr_scheduler\nfrom torch.autograd import Variable\n\nimport copy\nimport time\nimport json\n\nfrom collections import OrderedDict\n\nfrom PIL import Image\nimport random\n# -\n\n# Set seed\nrandom.seed(2)\nnp.random.seed(2)\ntorch.manual_seed(2)\ntorch.cuda.manual_seed_all(2)\ntorch.backends.cudnn.deterministic=True\n\n# ## Load the data\n#\n# Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.\n#\n# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.\n#\n# The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.\n# \n\ndata_dir = 'flowers'\ntrain_dir = data_dir + '/train'\nvalid_dir = data_dir + '/valid'\ntest_dir = data_dir + '/test'\n\n# +\n# TODO: Define your transforms for the training, validation, and testing sets\ndata_transforms = {'train' : transforms.Compose([transforms.RandomRotation(30),\n transforms.RandomResizedCrop(224),\n transforms.RandomHorizontalFlip(),\n transforms.ToTensor(),\n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'valid' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'test' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])])}\n\n# TODO: Load the datasets with ImageFolder\nimage_datasets = {'train' : datasets.ImageFolder(train_dir, transform=data_transforms['train']),\n 'valid' : datasets.ImageFolder(valid_dir, transform=data_transforms['valid']),\n 'test' : datasets.ImageFolder(test_dir, transform=data_transforms['test'])}\n\n# TODO: Using the image datasets and the trainforms, define the dataloaders\ndataloaders = {'train' : torch.utils.data.DataLoader(image_datasets['train'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'valid' : torch.utils.data.DataLoader(image_datasets['valid'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'test' : torch.utils.data.DataLoader(image_datasets['test'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2))}\n\ndataset_sizes = {'train': len(image_datasets['train']),\n 'valid': len(image_datasets['valid']),\n 'test': len(image_datasets['test'])} \n# -\n\n# ### Label mapping\n#\n# You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.\n\nwith open('cat_to_name.json', 'r') as f:\n cat_to_name = json.load(f)\n\n# # Building and training the classifier\n#\n# Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.\n#\n# We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students! You can also ask questions on the forums or join the instructors in office hours.\n#\n# Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:\n#\n# * Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)\n# * Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout\n# * Train the classifier layers using backpropagation using the pre-trained network to get the features\n# * Track the loss and accuracy on the validation set to determine the best hyperparameters\n#\n# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!\n#\n# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.\n\n# +\n# TODO: Build and train your network\n\n# +\n# default device for all functions with device argument: the fastest device available\n# all functions move models to device at the start and all returned models are moved to cpu at the end: \n# to avoid having problems with loading checkpoint when cuda is not available \ndevice = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\n\n# Defaults for functions to follow: architecture and parameters of the best_model\narchitecture = 'vgg16'\nlr = 0.0001\nhu = 4096\nstep_size = 5\n# Best num_epochs was 8 with lr=0.0001 but 10 with lr=0.001. \n# Keeping value at 10 is ok as train_model picks best number of epochs anyway.\n# We keep default num_epochs=10 in train.py as well. \nnum_epochs = 10 \n\n\n# -\n\ndef pick_arch(architecture):\n \"\"\"Pick from the following model architectures: vgg11, vgg13, vgg16, vgg19.\"\"\"\n if architecture == 'vgg11':\n model = models.vgg11(pretrained=True)\n elif architecture == 'vgg13':\n model = models.vgg13(pretrained=True)\n elif architecture == 'vgg16':\n model = models.vgg16(pretrained=True)\n elif architecture == 'vgg19':\n model = models.vgg19(pretrained=True)\n else:\n model = None\n print('Wrong model architecture. Pick vgg11, vgg13, vgg16 or vgg19.')\n \n return model\n\n\ndef build_model(architecture=architecture, hu=hu):\n \"\"\"Build a model from given architecture and replace classifier with a custom one.\"\"\"\n model = pick_arch(architecture)\n\n for param in model.parameters():\n param.requires_grad = False\n\n input_size = list(model.classifier.parameters())[0].size()[1]\n output_size = len(image_datasets['train'].class_to_idx)\n\n classifier = nn.Sequential(OrderedDict([\n ('fc1', nn.Linear(input_size, hu)),\n ('relu', nn.ReLU()),\n ('dropout', nn.Dropout(p=0.5)),\n ('fc2', nn.Linear(hu, output_size)),\n ('output', nn.LogSoftmax(dim=1))\n ]))\n\n model.classifier = classifier\n \n return model\n\n\ndef train_model(model=None, criterion=None, optimizer=None, scheduler=None,\n num_epochs=num_epochs, device=device):\n since = time.time()\n \n if model == None:\n model = build_model()\n \n if criterion == None:\n criterion = nn.NLLLoss()\n \n if optimizer == None:\n optimizer = optim.Adam(model.classifier.parameters(), lr)\n \n if scheduler == None:\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) \n \n model = model.to(device)\n \n best_model_wts = copy.deepcopy(model.state_dict())\n best_acc = 0.0\n best_num_epochs = 0\n\n for epoch in range(num_epochs):\n print('Epoch {}/{}'.format(epoch, num_epochs - 1))\n print('-' * 10)\n\n # Each epoch has a training and validation phase\n for phase in ['train', 'valid']:\n if phase == 'train':\n scheduler.step()\n model.train() # Set model to training mode\n else:\n model.eval() # Set model to evaluate mode\n\n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n # zero the parameter gradients\n optimizer.zero_grad()\n\n # forward\n # track history if only in train\n with torch.set_grad_enabled(phase == 'train'):\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # backward + optimize only if in training phase\n if phase == 'train':\n loss.backward()\n optimizer.step()\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n epoch_loss = running_loss / dataset_sizes[phase]\n epoch_acc = running_corrects.double() / dataset_sizes[phase]\n\n print('{} Loss: {:.4f} Acc: {:.4f}'.format(\n phase, epoch_loss, epoch_acc))\n\n # deep copy the model\n if phase == 'valid' and epoch_acc > best_acc:\n best_acc = epoch_acc\n best_model_wts = copy.deepcopy(model.state_dict())\n best_num_epochs = epoch + 1\n\n print('-' * 10)\n time_elapsed = time.time() - since\n print('Training complete in {:.0f}m {:.0f}s'.format(\n time_elapsed // 60, time_elapsed % 60))\n print('Best validation Accuracy of {:.4f} for {} epochs'.format(best_acc, best_num_epochs))\n print()\n\n # load best model weights\n model.load_state_dict(best_model_wts)\n model.to('cpu')\n return model, best_acc, best_num_epochs\n\n\n# The trial_model below was trained when default lr was set to 0.001. \n\ntrial_model, trial_acc, trial_num_epochs = train_model(num_epochs=2)\n\n\ndef tune_hyperparams(architecture=architecture, num_epochs=num_epochs,\n lr_list=[lr], hu_list=[hu], step_size_list=[step_size], device=device):\n param_triples = [(lr, hu, step_size) for lr in lr_list for hu in hu_list for step_size in step_size_list]\n \n best_model = pick_arch(architecture)\n best_acc = 0.0\n best_num_epochs = 0\n best_lr = 0.0\n best_hu = 0\n best_step_size = 0\n \n for lr, hu, step_size in param_triples:\n print('Training with')\n print('Learning rate: {} Number of hidden units: {} Step size: {}'.format(lr, hu, step_size))\n print('-' * 10)\n\n model = build_model(architecture=architecture, hu=hu)\n model = model.to(device)\n \n criterion = nn.NLLLoss()\n optimizer = optim.Adam(model.classifier.parameters(), lr=lr)\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) #add these to hyperparams\n \n model_trained, acc_trained, num_epochs_trained = train_model(model, criterion, optimizer, scheduler, num_epochs=num_epochs, device=device)\n \n # deep copy the model\n if acc_trained > best_acc:\n best_acc = acc_trained\n best_model = copy.deepcopy(model_trained)\n best_num_epochs = num_epochs_trained\n best_lr = lr\n best_hu = hu\n best_step_size = step_size\n \n print()\n print('Best results:')\n print('Learning Rate: {:.4f} Number of Hidden Units: {:.4f} Step size: {:.4f}'.format(best_lr, best_hu, best_step_size))\n print('Number of Epochs: {:.4f} Validation Acc: {:.4f}'.format(best_num_epochs, best_acc))\n best_model.to('cpu')\n return best_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size\n\n\ntune_hyperparams(num_epochs=2)\n\n# I tried learning rate of 0.01, number of hidden units 2048 and step size 7 but they did not do as well as lr=0.001, hu=4096 and step_size=5 and I decided to not spend more time training combinations with the above values. The last step in deciding on parameters is to see if learning rates of 0.0001 does better than 0.001. The train_model function already chooses the best number of epochs between 1 and 10. \n\nbest_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size = tune_hyperparams(lr_list=[0.0001, 0.001], hu_list=[4096], step_size_list=[5])\n\n\n# ## Testing your network\n#\n# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.\n\n# TODO: Do validation on the test set\ndef do_validation(model=None, criterion=None, phase='test', device=device):\n if model == None:\n model = build_model()\n if criterion == None:\n criterion = nn.NLLLoss()\n \n model.to(device)\n model.eval()\n \n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n\n # forward\n # don't track history \n with torch.no_grad():\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n total_loss = running_loss / dataset_sizes[phase]\n total_acc = running_corrects.double() / dataset_sizes[phase]\n \n model.to('cpu')\n \n print('Loss: {:.4f} Acc: {:.4f}'.format(\n total_loss, total_acc))\n return total_loss, total_acc.item()\n\ntotal_loss, total_acc = do_validation(model = best_model) \n\n# +\n#next(best_model.parameters()).is_cuda\n# -\n\n# ## Save the checkpoint\n#\n# Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.\n#\n# ```model.class_to_idx = image_datasets['train'].class_to_idx```\n#\n# Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.\n\n# TODO: Save the checkpoint \ncheckpoint = {'state_dict': best_model.state_dict(),\n 'hu': best_hu,\n 'architecture': architecture,\n 'num_epochs': best_num_epochs,\n 'lr': best_lr,\n 'step_size': best_step_size,\n 'class_to_idx': image_datasets['train'].class_to_idx}\n\ntorch.save(checkpoint, 'checkpoint_best.pth')\n\nbest_model.class_to_idx = image_datasets['train'].class_to_idx\n\n\n# ## Loading the checkpoint\n#\n# At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.\n\n# TODO: Write a function that loads a checkpoint and rebuilds the model\ndef load_checkpoint(filepath):\n checkpoint = torch.load(filepath)\n \n model = build_model(architecture=checkpoint['architecture'], hu=checkpoint['hu'])\n\n model.load_state_dict(checkpoint['state_dict'])\n model.class_to_idx = checkpoint['class_to_idx']\n \n return model, {'num_epochs':checkpoint['num_epochs'], \n 'lr': checkpoint['lr'], \n 'step_size': checkpoint['step_size'], \n 'hu': checkpoint['hu'], \n 'architecture': checkpoint['architecture']}\n\n\nreloaded_best_model, reloaded_best_params_dict = load_checkpoint('checkpoint_best.pth') \n\ndo_validation(model = reloaded_best_model)\n\n\n# # Inference for classification\n#\n# Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like \n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n#\n# First you'll need to handle processing the input image such that it can be used in your network. \n#\n# ## Image Preprocessing\n#\n# You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training. \n#\n# First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.\n#\n# Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.\n#\n# As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation. \n#\n# And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.\n\ndef process_image(image):\n ''' Scales, crops, and normalizes a PIL image for a PyTorch model,\n returns an Numpy array\n '''\n # TODO: Process a PIL image for use in a PyTorch model\n pil_image = Image.open(image)\n width, height = pil_image.size\n l = int(256 * max(width, height) / min(width, height))\n pil_image.thumbnail(size = (l,l))\n width, height = pil_image.size \n pil_image = pil_image.crop(box = (width / 2 - 112, height / 2 - 112, width / 2 + 112, height / 2 + 112))\n \n np_image = np.array(pil_image)\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n np_image = (np_image / 255.0 - mean) / std\n \n np_image = np_image.transpose((2, 0, 1))\n \n return np_image\n\n\n# To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).\n\ndef imshow(image, ax=None, title=None):\n if ax is None:\n fig, ax = plt.subplots()\n \n # PyTorch tensors assume the color channel is the first dimension\n # but matplotlib assumes is the third dimension\n image = image.transpose((1, 2, 0))\n \n # Undo preprocessing\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n image = std * image + mean\n \n # Image needs to be clipped between 0 and 1 or it looks like noise when displayed\n image = np.clip(image, 0, 1)\n \n ax.imshow(image)\n \n return ax\n\n\nimshow(process_image('flowers/test/1/image_06743.jpg'))\n\n\n# ## Class Prediction\n#\n# Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.\n#\n# To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.\n#\n# Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.\n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n\n# +\ndef predict(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\ndef predict2(image_path, checkpoint, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model from checkpoint.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model, _ = load_checkpoint(checkpoint)\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\n\n\n# small change in processing image: use transforms instead of process_image function\ndef predict3(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n transfs = data_transforms['test']\n\n img_pil = Image.open(image_path)\n img_tensor = transfs(img_pil)\n img_tensor.unsqueeze_(0)\n img_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(img_variable)\n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes \n\n\n# -\n\nprobs, classes = predict('flowers/test/1/image_06743.jpg', reloaded_best_model)\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\nprobs, classes = predict2('flowers/test/1/image_06743.jpg', 'checkpoint_best.pth')\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\n\n# ## Sanity Checking\n#\n# Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:\n#\n#
\n#\n# The project is broken down into multiple steps:\n#\n# * Load and preprocess the image dataset\n# * Train the image classifier on your dataset\n# * Use the trained classifier to predict image content\n#\n# We'll lead you through each part which you'll implement in Python.\n#\n# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.\n#\n# First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.\n\n# +\n# Imports here\nimport numpy as np\nimport matplotlib.pyplot as plt\n\nimport torch\nimport torchvision\nfrom torchvision import datasets, transforms, models\nfrom torch import nn\nfrom torch import optim\n#import torch.nn.functional as F\nfrom torch.optim import lr_scheduler\nfrom torch.autograd import Variable\n\nimport copy\nimport time\nimport json\n\nfrom collections import OrderedDict\n\nfrom PIL import Image\nimport random\n# -\n\n# Set seed\nrandom.seed(2)\nnp.random.seed(2)\ntorch.manual_seed(2)\ntorch.cuda.manual_seed_all(2)\ntorch.backends.cudnn.deterministic=True\n\n# ## Load the data\n#\n# Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.\n#\n# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.\n#\n# The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.\n# \n\ndata_dir = 'flowers'\ntrain_dir = data_dir + '/train'\nvalid_dir = data_dir + '/valid'\ntest_dir = data_dir + '/test'\n\n# +\n# TODO: Define your transforms for the training, validation, and testing sets\ndata_transforms = {'train' : transforms.Compose([transforms.RandomRotation(30),\n transforms.RandomResizedCrop(224),\n transforms.RandomHorizontalFlip(),\n transforms.ToTensor(),\n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'valid' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'test' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])])}\n\n# TODO: Load the datasets with ImageFolder\nimage_datasets = {'train' : datasets.ImageFolder(train_dir, transform=data_transforms['train']),\n 'valid' : datasets.ImageFolder(valid_dir, transform=data_transforms['valid']),\n 'test' : datasets.ImageFolder(test_dir, transform=data_transforms['test'])}\n\n# TODO: Using the image datasets and the trainforms, define the dataloaders\ndataloaders = {'train' : torch.utils.data.DataLoader(image_datasets['train'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'valid' : torch.utils.data.DataLoader(image_datasets['valid'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'test' : torch.utils.data.DataLoader(image_datasets['test'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2))}\n\ndataset_sizes = {'train': len(image_datasets['train']),\n 'valid': len(image_datasets['valid']),\n 'test': len(image_datasets['test'])} \n# -\n\n# ### Label mapping\n#\n# You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.\n\nwith open('cat_to_name.json', 'r') as f:\n cat_to_name = json.load(f)\n\n# # Building and training the classifier\n#\n# Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.\n#\n# We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students! You can also ask questions on the forums or join the instructors in office hours.\n#\n# Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:\n#\n# * Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)\n# * Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout\n# * Train the classifier layers using backpropagation using the pre-trained network to get the features\n# * Track the loss and accuracy on the validation set to determine the best hyperparameters\n#\n# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!\n#\n# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.\n\n# +\n# TODO: Build and train your network\n\n# +\n# default device for all functions with device argument: the fastest device available\n# all functions move models to device at the start and all returned models are moved to cpu at the end: \n# to avoid having problems with loading checkpoint when cuda is not available \ndevice = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\n\n# Defaults for functions to follow: architecture and parameters of the best_model\narchitecture = 'vgg16'\nlr = 0.0001\nhu = 4096\nstep_size = 5\n# Best num_epochs was 8 with lr=0.0001 but 10 with lr=0.001. \n# Keeping value at 10 is ok as train_model picks best number of epochs anyway.\n# We keep default num_epochs=10 in train.py as well. \nnum_epochs = 10 \n\n\n# -\n\ndef pick_arch(architecture):\n \"\"\"Pick from the following model architectures: vgg11, vgg13, vgg16, vgg19.\"\"\"\n if architecture == 'vgg11':\n model = models.vgg11(pretrained=True)\n elif architecture == 'vgg13':\n model = models.vgg13(pretrained=True)\n elif architecture == 'vgg16':\n model = models.vgg16(pretrained=True)\n elif architecture == 'vgg19':\n model = models.vgg19(pretrained=True)\n else:\n model = None\n print('Wrong model architecture. Pick vgg11, vgg13, vgg16 or vgg19.')\n \n return model\n\n\ndef build_model(architecture=architecture, hu=hu):\n \"\"\"Build a model from given architecture and replace classifier with a custom one.\"\"\"\n model = pick_arch(architecture)\n\n for param in model.parameters():\n param.requires_grad = False\n\n input_size = list(model.classifier.parameters())[0].size()[1]\n output_size = len(image_datasets['train'].class_to_idx)\n\n classifier = nn.Sequential(OrderedDict([\n ('fc1', nn.Linear(input_size, hu)),\n ('relu', nn.ReLU()),\n ('dropout', nn.Dropout(p=0.5)),\n ('fc2', nn.Linear(hu, output_size)),\n ('output', nn.LogSoftmax(dim=1))\n ]))\n\n model.classifier = classifier\n \n return model\n\n\ndef train_model(model=None, criterion=None, optimizer=None, scheduler=None,\n num_epochs=num_epochs, device=device):\n since = time.time()\n \n if model == None:\n model = build_model()\n \n if criterion == None:\n criterion = nn.NLLLoss()\n \n if optimizer == None:\n optimizer = optim.Adam(model.classifier.parameters(), lr)\n \n if scheduler == None:\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) \n \n model = model.to(device)\n \n best_model_wts = copy.deepcopy(model.state_dict())\n best_acc = 0.0\n best_num_epochs = 0\n\n for epoch in range(num_epochs):\n print('Epoch {}/{}'.format(epoch, num_epochs - 1))\n print('-' * 10)\n\n # Each epoch has a training and validation phase\n for phase in ['train', 'valid']:\n if phase == 'train':\n scheduler.step()\n model.train() # Set model to training mode\n else:\n model.eval() # Set model to evaluate mode\n\n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n # zero the parameter gradients\n optimizer.zero_grad()\n\n # forward\n # track history if only in train\n with torch.set_grad_enabled(phase == 'train'):\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # backward + optimize only if in training phase\n if phase == 'train':\n loss.backward()\n optimizer.step()\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n epoch_loss = running_loss / dataset_sizes[phase]\n epoch_acc = running_corrects.double() / dataset_sizes[phase]\n\n print('{} Loss: {:.4f} Acc: {:.4f}'.format(\n phase, epoch_loss, epoch_acc))\n\n # deep copy the model\n if phase == 'valid' and epoch_acc > best_acc:\n best_acc = epoch_acc\n best_model_wts = copy.deepcopy(model.state_dict())\n best_num_epochs = epoch + 1\n\n print('-' * 10)\n time_elapsed = time.time() - since\n print('Training complete in {:.0f}m {:.0f}s'.format(\n time_elapsed // 60, time_elapsed % 60))\n print('Best validation Accuracy of {:.4f} for {} epochs'.format(best_acc, best_num_epochs))\n print()\n\n # load best model weights\n model.load_state_dict(best_model_wts)\n model.to('cpu')\n return model, best_acc, best_num_epochs\n\n\n# The trial_model below was trained when default lr was set to 0.001. \n\ntrial_model, trial_acc, trial_num_epochs = train_model(num_epochs=2)\n\n\ndef tune_hyperparams(architecture=architecture, num_epochs=num_epochs,\n lr_list=[lr], hu_list=[hu], step_size_list=[step_size], device=device):\n param_triples = [(lr, hu, step_size) for lr in lr_list for hu in hu_list for step_size in step_size_list]\n \n best_model = pick_arch(architecture)\n best_acc = 0.0\n best_num_epochs = 0\n best_lr = 0.0\n best_hu = 0\n best_step_size = 0\n \n for lr, hu, step_size in param_triples:\n print('Training with')\n print('Learning rate: {} Number of hidden units: {} Step size: {}'.format(lr, hu, step_size))\n print('-' * 10)\n\n model = build_model(architecture=architecture, hu=hu)\n model = model.to(device)\n \n criterion = nn.NLLLoss()\n optimizer = optim.Adam(model.classifier.parameters(), lr=lr)\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) #add these to hyperparams\n \n model_trained, acc_trained, num_epochs_trained = train_model(model, criterion, optimizer, scheduler, num_epochs=num_epochs, device=device)\n \n # deep copy the model\n if acc_trained > best_acc:\n best_acc = acc_trained\n best_model = copy.deepcopy(model_trained)\n best_num_epochs = num_epochs_trained\n best_lr = lr\n best_hu = hu\n best_step_size = step_size\n \n print()\n print('Best results:')\n print('Learning Rate: {:.4f} Number of Hidden Units: {:.4f} Step size: {:.4f}'.format(best_lr, best_hu, best_step_size))\n print('Number of Epochs: {:.4f} Validation Acc: {:.4f}'.format(best_num_epochs, best_acc))\n best_model.to('cpu')\n return best_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size\n\n\ntune_hyperparams(num_epochs=2)\n\n# I tried learning rate of 0.01, number of hidden units 2048 and step size 7 but they did not do as well as lr=0.001, hu=4096 and step_size=5 and I decided to not spend more time training combinations with the above values. The last step in deciding on parameters is to see if learning rates of 0.0001 does better than 0.001. The train_model function already chooses the best number of epochs between 1 and 10. \n\nbest_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size = tune_hyperparams(lr_list=[0.0001, 0.001], hu_list=[4096], step_size_list=[5])\n\n\n# ## Testing your network\n#\n# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.\n\n# TODO: Do validation on the test set\ndef do_validation(model=None, criterion=None, phase='test', device=device):\n if model == None:\n model = build_model()\n if criterion == None:\n criterion = nn.NLLLoss()\n \n model.to(device)\n model.eval()\n \n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n\n # forward\n # don't track history \n with torch.no_grad():\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n total_loss = running_loss / dataset_sizes[phase]\n total_acc = running_corrects.double() / dataset_sizes[phase]\n \n model.to('cpu')\n \n print('Loss: {:.4f} Acc: {:.4f}'.format(\n total_loss, total_acc))\n return total_loss, total_acc.item()\n\ntotal_loss, total_acc = do_validation(model = best_model) \n\n# +\n#next(best_model.parameters()).is_cuda\n# -\n\n# ## Save the checkpoint\n#\n# Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.\n#\n# ```model.class_to_idx = image_datasets['train'].class_to_idx```\n#\n# Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.\n\n# TODO: Save the checkpoint \ncheckpoint = {'state_dict': best_model.state_dict(),\n 'hu': best_hu,\n 'architecture': architecture,\n 'num_epochs': best_num_epochs,\n 'lr': best_lr,\n 'step_size': best_step_size,\n 'class_to_idx': image_datasets['train'].class_to_idx}\n\ntorch.save(checkpoint, 'checkpoint_best.pth')\n\nbest_model.class_to_idx = image_datasets['train'].class_to_idx\n\n\n# ## Loading the checkpoint\n#\n# At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.\n\n# TODO: Write a function that loads a checkpoint and rebuilds the model\ndef load_checkpoint(filepath):\n checkpoint = torch.load(filepath)\n \n model = build_model(architecture=checkpoint['architecture'], hu=checkpoint['hu'])\n\n model.load_state_dict(checkpoint['state_dict'])\n model.class_to_idx = checkpoint['class_to_idx']\n \n return model, {'num_epochs':checkpoint['num_epochs'], \n 'lr': checkpoint['lr'], \n 'step_size': checkpoint['step_size'], \n 'hu': checkpoint['hu'], \n 'architecture': checkpoint['architecture']}\n\n\nreloaded_best_model, reloaded_best_params_dict = load_checkpoint('checkpoint_best.pth') \n\ndo_validation(model = reloaded_best_model)\n\n\n# # Inference for classification\n#\n# Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like \n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n#\n# First you'll need to handle processing the input image such that it can be used in your network. \n#\n# ## Image Preprocessing\n#\n# You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training. \n#\n# First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.\n#\n# Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.\n#\n# As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation. \n#\n# And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.\n\ndef process_image(image):\n ''' Scales, crops, and normalizes a PIL image for a PyTorch model,\n returns an Numpy array\n '''\n # TODO: Process a PIL image for use in a PyTorch model\n pil_image = Image.open(image)\n width, height = pil_image.size\n l = int(256 * max(width, height) / min(width, height))\n pil_image.thumbnail(size = (l,l))\n width, height = pil_image.size \n pil_image = pil_image.crop(box = (width / 2 - 112, height / 2 - 112, width / 2 + 112, height / 2 + 112))\n \n np_image = np.array(pil_image)\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n np_image = (np_image / 255.0 - mean) / std\n \n np_image = np_image.transpose((2, 0, 1))\n \n return np_image\n\n\n# To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).\n\ndef imshow(image, ax=None, title=None):\n if ax is None:\n fig, ax = plt.subplots()\n \n # PyTorch tensors assume the color channel is the first dimension\n # but matplotlib assumes is the third dimension\n image = image.transpose((1, 2, 0))\n \n # Undo preprocessing\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n image = std * image + mean\n \n # Image needs to be clipped between 0 and 1 or it looks like noise when displayed\n image = np.clip(image, 0, 1)\n \n ax.imshow(image)\n \n return ax\n\n\nimshow(process_image('flowers/test/1/image_06743.jpg'))\n\n\n# ## Class Prediction\n#\n# Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.\n#\n# To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.\n#\n# Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.\n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n\n# +\ndef predict(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\ndef predict2(image_path, checkpoint, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model from checkpoint.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model, _ = load_checkpoint(checkpoint)\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\n\n\n# small change in processing image: use transforms instead of process_image function\ndef predict3(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n transfs = data_transforms['test']\n\n img_pil = Image.open(image_path)\n img_tensor = transfs(img_pil)\n img_tensor.unsqueeze_(0)\n img_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(img_variable)\n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes \n\n\n# -\n\nprobs, classes = predict('flowers/test/1/image_06743.jpg', reloaded_best_model)\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\nprobs, classes = predict2('flowers/test/1/image_06743.jpg', 'checkpoint_best.pth')\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\n\n# ## Sanity Checking\n#\n# Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:\n#\n#  \n#\n# You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.\n\n# +\n# TODO: Display an image along with the top 5 classes\ndef plot_im_probs_classes(image_path, model, device=device):\n probs, classes = predict(image_path, model, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout()\n \ndef plot_im_probs_classes2(image_path, checkpoint, device=device):\n probs, classes = predict2(image_path, checkpoint, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout() \n\n\n# -\n\nplot_im_probs_classes('flowers/test/1/image_06760.jpg', model=reloaded_best_model, device=device)\n\nplot_im_probs_classes2('flowers/test/1/image_06760.jpg', 'checkpoint_best.pth', device=device)\n\n# Resources used for this project:\n# * Material from the first term of Udacity's Data Scientist Nanodegree program. \n# * [Transfer Learning tutorial](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html). `train_model` function was taken from this tutorial with small changes as well as other hints such as shuffling the datasets and using a scheduler. \n# * [PyTorch documentation](https://pytorch.org/docs/stable/index.html).\n# * [Argparse tutorial](https://pymotw.com/3/argparse/#argument-types) and [documentation](https://docs.python.org/dev/library/argparse.html#argparse.Namespace).\n","repo_name":"anamamatelashvili/deep_learning_project","sub_path":"Image Classifier Project.ipynb","file_name":"Image Classifier Project.ipynb","file_ext":"py","file_size_in_byte":31574,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"581944992","text":"# + [markdown] id=\"E-aBmrp8931c\"\n# # Proyecto empresa de viajes compartidos\n\n# + [markdown] id=\"uEJABzxG931c\"\n# ### Descripción del proyecto\n#\n# Durante este proyecto analizaré datos para una nueva empresa de viajes compartidos que se esta lanzando en Chicago. El objetivo será encontrar patrones en los datos y entender las preferencias de los pasajeros y el impacto de los factores externos en los viajes.\n#\n# #### Descripción de los datos\n#\n# Tabla 1: contiene datos sobre la cantidad de viajes que realizó cada compañía.\n# - company_name: nombre de la empresa de taxis\n# - trips_amount: el número de viajes de cada compañía de taxis el 15 y 16 de noviembre de 2017.\n#\n# Tabla 2: contiene datos sobre la finalización de los viajes \n# - dropoff_location_name: barrios de Chicago donde finalizaron los viajes\n# - average_trips: el promedio de viajes que terminaron en cada barrio en noviembre de 2017.\n#\n# Tabla 3:\n# Contiene datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. \n# - start_ts: fecha y hora de la recogida\n# - weather_conditions: condiciones climáticas en el momento en el que comenzó el viaje\n# - duration_seconds: duración del viaje en segundos\n\n# + [markdown] id=\"1G-R9FNe931c\"\n# ### Análisis exploratorio de datos\n\n# + id=\"PZCA0wo3931c\"\n# Importación de librerías a utilizar\n\nimport pandas as pd\nfrom scipy import stats as st\nfrom scipy.stats import levene\nimport matplotlib.pyplot as plt\nfrom seaborn import boxplot\n\n# + id=\"r3wEulH9931d\" outputId=\"1375e687-549f-4792-8c56-1ebfe3ff3c0f\"\n# Importación de archivo 1: número de viajes por compañia\n\ncompanies_trips= pd.read_csv('/datasets/project_sql_result_01.csv')\ncompanies_trips.info()\ncompanies_trips.head()\n\n\n# + id=\"MA1CVDmo931d\" outputId=\"0d2dabbc-9c46-4fb2-e35a-d36d79ab3335\"\n# Chequeo de valores ausentes\n\ncompanies_trips.isna().sum()\n\n# + id=\"UVq55dyy931d\" outputId=\"9735fac2-c40f-4ee0-8c3e-73d4238f734d\"\n# Chequeo de valores duplicados\n\ncompanies_trips.duplicated().sum()\n\n# + id=\"Bio8N7b7931d\" outputId=\"01191f13-de81-4e3f-bc49-83e4a9496fa8\"\n# Importación de archivo 2: promedio de viajes por barrio \n\ndropoff_location= pd.read_csv('/datasets/project_sql_result_04.csv')\ndropoff_location.info()\ndropoff_location.head()\n\n# + id=\"VfGtzMP-931e\" outputId=\"c6356ec7-0873-41da-c118-50a4d1e55f63\"\n# Chequeo de valores ausentes \n\ndropoff_location.isna().sum()\n\n# + id=\"bYD1uV96931e\" outputId=\"1d75298f-3319-4eb2-818a-4e3f4d0a2a96\"\n# Chequeo de valores duplicados\n\ndropoff_location.duplicated().sum()\n\n# + id=\"8aHq5XT-931e\" outputId=\"384087f9-8cea-4f88-e3a1-f5c338d36c98\"\n# Identificar los 10 principales barrios en términos de finalización del recorrido\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10)\n\n# + id=\"LdV71Unp931e\" outputId=\"5da0eaf9-c0e7-4ef7-f5c7-d51006a0a7ba\"\n#Gráfico: empresas de taxis y número de viajes\n\ncompanies_trips.plot(kind='bar',x='company_name', y='trips_amount', ylabel='trips_amount', figsize=(20,10), title='Viajes por empresa de taxi')\n\n# + [markdown] id=\"shnqBODc931e\"\n# Como podemos observar en el gráfico de arriba, la empresa de taxis más usada los días 15 y 16 de noviembre fue Flash Cab. Ésta realizó casi el doble de viajes que la empresa que le sigue: Taxi Affiliation Services. \n#\n# A su vez, dejando de lado Flash Cab que fue sin dudas la más poplar, podemos observar que hay pequeños grupos de compañías en los cuales no difiere tanto la cantidad de viajes realizados. Por ejemplo, las otras 5 copañías principales de taxis realizaron entre 9180 a 11400 viajes.\n#\n# Por otro lado, podemos observar que la mitad de las compañías de taxis tuvieron muy pocos viajes esos días. \n\n# + id=\"bzjPAp8R931e\" outputId=\"4031eafb-cf1d-4fc0-d3f1-73c5e893dc32\"\n# Gráfico 10 barrios principales por número de finalizaciones\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10).plot(kind='bar',x='dropoff_location_name', y='average_trips', ylabel='Trips amount', figsize=(20,10), title='Principales barrios')\n\n# + [markdown] id=\"Hl0_zDu_931f\"\n# Como podemos observar en el gráfico de arriba, se muestran los 10 barrios principales hacia los cuales se dirigieron los taxis (destino final). El barrio con más viajes fue Loop con un promedio de 10727 viajes en el mes de noviembre seguido por River North con un promedio de 9523 viajes.\n#\n# Después se encuentran Streeterville\ty West Loop con un promedio de viajes de 6664 y 5163 respectivamente y por último un tercer grupo en la tabla con barrios con un promedio de viajes entre 1260 a 2420 en el mes de noviembre. \n\n# + [markdown] id=\"ibJP9mHu931f\"\n# ### Prueba de hipótesis\n\n# + [markdown] id=\"9qceopHk931f\"\n# A continuación voy a importar el archivo con los datos que usare para hacer la prueba de hipótesis.\n\n# + id=\"Mj53hprj931f\" outputId=\"216d861a-a20e-40c7-bfca-172a9615cafb\"\n# Importación de archivo para trabajar sobre la hipótesis\n\narchivo_hipotesis = pd.read_csv('/datasets/project_sql_result_07.csv')\narchivo_hipotesis.info()\narchivo_hipotesis.head()\n\n# + id=\"nT7L8FIk931f\" outputId=\"04025974-4c5f-4a5f-84e1-3c46cf2c3852\"\n# Chequeo de valores ausentes\n\narchivo_hipotesis.isna().sum()\n\n# + id=\"ZqMuqgQS931f\" outputId=\"9b37c186-3558-4c3a-b90c-d3e53abca6f5\"\n# Chequeo de valores duplicados\n\narchivo_hipotesis.duplicated().sum()\n\n# + id=\"2gIJlz-9931f\" outputId=\"7e2315d2-3169-427a-c706-8443f22436e4\"\narchivo_hipotesis.value_counts()\n\n# + [markdown] id=\"aygBH6hT931f\"\n# Como podemos observar, en el archivo que nos dieron para hacer la prueba de hipótesis tenemos valores duplicados. Estos pueden deberse a que el archivo contiene los datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. Puede que haya determinadas paradas en Loop con destino hacia el aeropuerto. Al ser un punto fijo es posible que en condiciones normales, el recorrido dure siempre lo mismo; todos los los viajes que salgan de ahi a una determnada hora van a tardar lo mismo generando que haya varios datos con os mismos tiempos.\n\n# + id=\"fb9UZ9-Q931f\" outputId=\"128662bf-d6f4-403b-9c56-d506f9680a7d\"\nboxplot(data= archivo_hipotesis, x=\"duration_seconds\", y=\"weather_conditions\")\n\n# + [markdown] id=\"OxVmL-Y2931g\"\n# Voy a probar la hipótesis de que la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare cambia los sábados lluviosos.\n#\n# Hipótesis nula: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos. \n#\n# Hipótesis alternativa: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare difiere los sábados lluviosos.\n#\n# Previo a la prueba de hipótesis voy a realizar la prueba de levene para ver si las varianzas son iguales. Usaré como hipótesis nula: las varianzas son iguales y como hipótesis alternativa las varianzas no son iguales.\n\n# + id=\"3sY6DdYD931g\" outputId=\"a465661a-30aa-4dd3-92e3-eb7b9dee7791\"\n# Prueba de hipotesis nula levene: varianzas son iguales \n\ndatos_lluvia= archivo_hipotesis.query('weather_conditions == \"Bad\" and duration_seconds >0')\ndatos_good= archivo_hipotesis.query('weather_conditions ==\"Good\"and duration_seconds >0')\n\nlluvia= datos_lluvia['duration_seconds']\ngood= datos_good['duration_seconds']\n\nalpha= 0.05\n\nstat,results= levene(lluvia, good)\nprint('P_value:', results)\n\nif results < alpha:\n print('Rechazo la hipótesis nula')\nelse:\n print('No rechazo la hipótesis nula')\n\n# + [markdown] id=\"huB3ekYd931g\"\n# Como podemos observar de la prueba de Levene realizada arriba, no rechazamos la hipótesis nula que indica que las varianzas son iguales. \n#\n# A continuación realizare la prueba de hipótesis.\n\n# + id=\"ABv0Xklx931g\" outputId=\"7198fbf1-68b8-4240-bb38-b4164b0464f5\"\n# Ho: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos.\n\nalpha= 0.05\nresults= st.ttest_ind(lluvia,good, equal_var= True)\nprint('P_value:', results.pvalue)\n\nif results.pvalue < alpha:\n print('Rechazo la hipótesis nula')\nelse: \n print('No rechazo hipótesis nula')\n\n# + [markdown] id=\"Ns2N_ZTw931g\"\n# De acuerdo con los datos con los que contamos, rechazo la hipótesis nula. La duración promedio de los viajes desde el Loop hasta el aeropuerto de O'Hare cambia los sábados lluviosos.\n\n# + [markdown] id=\"dtnLXksI931g\"\n# ## Conclusión \n#\n# Para concluir, podemos decir que la empresa de taxis más utilizada es Flash Cab seguida por Taxi Affiliation Services, teniendo la primera casi el doble de viajes que la segunda. Esta diferencia puede deberse a que Flash Cab sea una empresa mucho más grande que las otras; al tener una flota de autos más grande en la calle le permite tener el doble de viajes que su competidor más cercano. \n#\n# Con respecto a los detinos finales de los viajes, Loop resulto el más concurrido seguido de cerca por River North.\n#\n# Por último la duración promedio de los viajes difiere un día lluvioso en comparación a un día de buen tiempo. Esta diferencia puede deberse principalmente al factor climático. Los días de mal tiempo puede que haya más tránsito porque la gente decide salir con sus autos y se genera más congestión. También la gente anda un poco más despacio impactando en la duración pormedio de los viajes. \n","repo_name":"CarmenPujato/practicum","sub_path":"empresa de viajes compartidos/Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_name":"Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_ext":"py","file_size_in_byte":9307,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43095479709","text":"# # Kaggle class competition\n# # Deep Learning and Neural Network Models to predict traffic volume\n# # Models: LSTM, GRUs\n# # Library used: Tensorflow, Keras\n\n# ### Set Up Libraries\n\n# +\n# ns - as typically happens, outdated components like as keywords, functions, and classes trigger a warning.\nimport warnings\nwarnings.filterwarnings('ignore')\n# ns - below are the neccessary packages utilized in this project\nimport os\nimport datetime\n\nimport IPython\nimport IPython.display\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns\nimport tensorflow as tf\n\nmpl.rcParams['figure.figsize'] = (8, 6)\nmpl.rcParams['axes.grid'] = False\n# -\n\n# ### Import data\n\n# ns - use pandas package to read in data from file located within folder \ndata = pd.read_csv('Metro_Interstate_Traffic_Volume.csv', infer_datetime_format=True)\ndata.head()\n\n# ### Inspect and Clean up\n\n# ns - investigate the data info\ndata.info()\n\n# #### It could be seen that there are both str and num types of data as categorized below\n# categorical features: holiday, weather_main, weather_description\n#\n# Continious features: temp, rain_1h, show_1h, clouds_all\n#\n# Target: traffic_volume\n\n# ### Convert date_time column to datetime type\n#\n# It is usable if we don't convert string type to datetime type in this project. It is due to the fact that time series will absolutely need datetime. So below is the demonstration.\n\n# ns - datetime type conversion\ndate_time = pd.to_datetime(data.pop('date_time'), format='%Y-%m-%d %H:%M:%S')\n\n# ### Convert to a usable signal \n#\n# In this form of signal, we would be able to utilize its capacity in the ultimate way as it reveals more insignhts as illjustrated below. \n\ntimestamp_s = date_time.map(datetime.datetime.timestamp)\n\n# +\nday = 24*60*60\nyear = (365.2425)*day\n\ndata['Day sin'] = np.sin(timestamp_s * (2 * np.pi / day))\ndata['Day cos'] = np.cos(timestamp_s * (2 * np.pi / day))\ndata['Year sin'] = np.sin(timestamp_s * (2 * np.pi / year))\ndata['Year cos'] = np.cos(timestamp_s * (2 * np.pi / year))\n# -\n\nplt.plot(np.array(data['Day sin'])[:25])\nplt.plot(np.array(data['Day cos'])[:25])\nplt.xlabel('Time [h]')\nplt.title('Time of day signal')\n\n# The best way to identify which frequencies are most essential is to use a \"fft\". As a preliminary test, below is the traffic volume rfft experiment. While we're discussing common frequencies, it's interesting to note the highs at 1/year and 1/day frequencies:\n\n# +\nfft = tf.signal.rfft(data['temp'])\nf_per_dataset = np.arange(0, len(fft))\n\nn_samples_h = len(data['temp'])\nhours_per_year = 24*365.2524\nyears_per_dataset = n_samples_h/(hours_per_year)\n\nf_per_year = f_per_dataset/years_per_dataset\nplt.step(f_per_year, np.abs(fft))\nplt.xscale('log')\nplt.ylim(0, 400000)\nplt.xlim([0.1, max(plt.xlim())])\nplt.xticks([1, 365.2524], labels=['1/Year', '1/day'])\n_ = plt.xlabel('Frequency (log scale)')\n# -\n\n# ### Drop variable \"weather_description\" and convert \"holiday\" and \"weather_main\" to dummies variables\n#\n# It is assumed that the weather description might be more irrelevant to this project or its interpretation could benefit less. Thus, I have decided to drop it. While both weather main and the holiday are deemed significant.\n\n# ns - considered to drop some variable and generate dummies variables for the rest of obj type\ndata.drop('weather_description', axis=1, inplace=True)\ndata = pd.get_dummies(data, columns = ['holiday', 'weather_main'], drop_first=True)\ndata.head()\n\n# ### Split the data\n\n# +\n# ns - as indicated in the instruction, the test set should be excluded with the size of 5000 ea.\n# ns - the slicing method is used in this data split\ncolumn_indices = {name: i for i, name in enumerate(data.columns)}\n\nn = len(data)\nprint(\"There are \", n, \"records and divisions are as below\")\n\ntest_df = data[-5000:]\nprint(\"Test set: \", test_df.shape)\n\nval_df = data[-13000:-5000]\nprint(\"Validation set: \", val_df.shape)\n\ntrain_df = data[0:-13000]\nprint(\"Training set: \", train_df.shape)\n\nnum_features = data.shape[1]\n# -\n\n# ### Data Normalization\n\n# +\ntrain_mean = train_df.mean()\ntrain_std = train_df.std()\n\ntrain_df = (train_df - train_mean) / train_std\nval_df = (val_df - train_mean) / train_std\ntest_df = (test_df - train_mean) / train_std\n\n\n# -\n\n# ### Data windowing\n#\n# The models in this project will make a set of predictions based on a window of consecutive samples from the data and below is the useful windowgenerator class for segmenting data as required for modelling.\n\nclass WindowGenerator():\n def __init__(self, input_width, label_width, shift,\n train_df=train_df, val_df=val_df, test_df=test_df,\n label_columns=None):\n # Store the raw data.\n self.train_df = train_df\n self.val_df = val_df\n self.test_df = test_df\n\n # Work out the label column indices.\n self.label_columns = label_columns\n if label_columns is not None:\n self.label_columns_indices = {name: i for i, name in\n enumerate(label_columns)}\n self.column_indices = {name: i for i, name in\n enumerate(train_df.columns)}\n\n # Work out the window parameters.\n self.input_width = input_width\n self.label_width = label_width\n self.shift = shift\n\n self.total_window_size = input_width + shift\n\n self.input_slice = slice(0, input_width)\n self.input_indices = np.arange(self.total_window_size)[self.input_slice]\n\n self.label_start = self.total_window_size - self.label_width\n self.labels_slice = slice(self.label_start, None)\n self.label_indices = np.arange(self.total_window_size)[self.labels_slice]\n\n def __repr__(self):\n return '\\n'.join([\n f'Total window size: {self.total_window_size}',\n f'Input indices: {self.input_indices}',\n f'Label indices: {self.label_indices}',\n f'Label column name(s): {self.label_columns}'])\n\n\n# As per required, the windowgenerator here will provide 6 width with 2 future prediction and 1 label.\n\nw2 = WindowGenerator(input_width=6, label_width=1, shift=2,\n label_columns=['traffic_volume'])\nw2\n\n\n# +\ndef split_window(self, features):\n inputs = features[:, self.input_slice, :]\n labels = features[:, self.labels_slice, :]\n if self.label_columns is not None:\n labels = tf.stack(\n [labels[:, :, self.column_indices[name]] for name in self.label_columns],\n axis=-1)\n\n # Slicing doesn't preserve static shape information, so set the shapes\n # manually. This way the `tf.data.Datasets` are easier to inspect.\n inputs.set_shape([None, self.input_width, None])\n labels.set_shape([None, self.label_width, None])\n\n return inputs, labels\n\nWindowGenerator.split_window = split_window\n# -\n\n# ### Note: \n#\n# Typically data in TensorFlow is packed into arrays where the outermost index is across examples (the \"batch\" dimension). The middle indices are the \"time\" or \"space\" (width, height) dimension(s). The innermost indices are the features.\n\n# +\n# ns - I tried with other stack but it failed to corporate with the model\n# Stack three slices, the length of the total window:\nexample_window = tf.stack([np.array(train_df[:w2.total_window_size]),\n np.array(train_df[100:100+w2.total_window_size]),\n np.array(train_df[200:200+w2.total_window_size])])\n\n\nexample_inputs, example_labels = w2.split_window(example_window)\n\nprint('All shapes are: (batch, time, features)')\nprint(f'Window shape: {example_window.shape}')\nprint(f'Inputs shape: {example_inputs.shape}')\nprint(f'labels shape: {example_labels.shape}')\n# -\n\n# ### Plot \n#\n# #### Here is a plot method that allows a simple visualization of the split window\n\nw2.example = example_inputs, example_labels\n\n\n# +\ndef plot(self, model=None, plot_col='traffic_volume', max_subplots=3):\n inputs, labels = self.example\n plt.figure(figsize=(12, 8))\n plot_col_index = self.column_indices[plot_col]\n max_n = min(max_subplots, len(inputs))\n for n in range(max_n):\n plt.subplot(max_n, 1, n+1)\n plt.ylabel(f'{plot_col} [normed]')\n plt.plot(self.input_indices, inputs[n, :, plot_col_index],\n label='Inputs', marker='.', zorder=-10)\n\n if self.label_columns:\n label_col_index = self.label_columns_indices.get(plot_col, None)\n else:\n label_col_index = plot_col_index\n\n if label_col_index is None:\n continue\n\n plt.scatter(self.label_indices, labels[n, :, label_col_index],\n edgecolors='k', label='Labels', c='#2ca02c', s=64)\n if model is not None:\n predictions = model(inputs)\n plt.scatter(self.label_indices, predictions[n, :, label_col_index],\n marker='X', edgecolors='k', label='Predictions',\n c='#ff7f0e', s=64)\n\n if n == 0:\n plt.legend()\n\n plt.xlabel('Time [h]')\n\nWindowGenerator.plot = plot\n# -\n\nw2.plot()\n\n\n# The plots above illustrate the 6 inputs and 1 label as we defined them ealier. \n\n# ### Create tf.data.Datasets\n\n# +\ndef make_dataset(self, data):\n data = np.array(data, dtype=np.float32)\n ds = tf.keras.preprocessing.timeseries_dataset_from_array(\n data=data,\n targets=None,\n sequence_length=self.total_window_size,\n sequence_stride=1,\n shuffle=False,\n batch_size=32,)\n\n ds = ds.map(self.split_window)\n\n return ds\n\nWindowGenerator.make_dataset = make_dataset\n\n\n# -\n\n# ### Note:\n#\n# This make_dataset method will take a time series DataFrame and convert it to a tf.data.Dataset of (input_window, label_window) pairs using the preprocessing.timeseries_dataset_from_array function. The WindowGenerator object holds training, validation and test data. Add properties for accessing them as tf.data.Datasets using the above make_dataset method.\n\n# +\n@property\ndef train(self):\n return self.make_dataset(self.train_df)\n\n@property\ndef val(self):\n return self.make_dataset(self.val_df)\n\n@property\ndef test(self):\n return self.make_dataset(self.test_df)\n\n@property\ndef example(self):\n \"\"\"Get and cache an example batch of `inputs, labels` for plotting.\"\"\"\n result = getattr(self, '_example', None)\n if result is None:\n # No example batch was found, so get one from the `.train` dataset\n result = next(iter(self.train))\n # And cache it for next time\n self._example = result\n return result\n\nWindowGenerator.train = train\nWindowGenerator.val = val\nWindowGenerator.test = test\nWindowGenerator.example = example\n# -\n\n# Each element is an (inputs, label) pair\nw2.train.element_spec\n\n# #### Iterating over a Dataset yields concrete batches:\n\nfor example_inputs, example_labels in w2.train.take(1):\n print(f'Inputs shape (batch, time, features): {example_inputs.shape}')\n print(f'Labels shape (batch, time, features): {example_labels.shape}')\n\n# #### This project trains many models, so package the training procedure into a function\n\n# +\nMAX_EPOCHS = 100\n\ndef compile_and_fit(model, window, patience=2):\n early_stopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss',\n patience=patience,\n mode='min')\n\n model.compile(loss=tf.losses.MeanSquaredError(),\n optimizer=tf.optimizers.Adam(),\n metrics=[tf.metrics.MeanAbsoluteError()])\n\n history = model.fit(window.train, epochs=MAX_EPOCHS,\n validation_data=window.val,\n callbacks=[early_stopping])\n return history\n\n\n# -\n\n# For the multi-step model, the training data consists of hourly samples. However, here, the models will learn to predict 2h of the future, given 6h of the past. Here is a Window object that generates these slices from the dataset.\n\n# +\n# ns - indicating the 2 hours future output from the 6 hours history input\nOUT_STEPS = 2\nmulti_window = WindowGenerator(input_width=6,\n label_width=OUT_STEPS,\n shift=OUT_STEPS)\n\nmulti_window.plot()\nmulti_window\n# -\n\n# Illustration here displays the 6 inputs with 2 hours prediction in the future.\n\n# ### Model Construction\n# #### (1) The Base LSTM model\n#\n# This model is expected to be used as the baseline model in this project. The error is aimed to be minimized in further steps. The architecture of this base model consists of LSTM layer with 32 unit, Dense layer and then reshape respectively.\n\n# +\nmulti_val_performance = {}\nmulti_performance = {}\n\nmulti_lstm_model = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.val)\nmulti_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model)\n# -\n\n# #### (2) The Base LSTM with parameter Recurrent Dropout (0.1) and dropout layer (0.1)\n#\n# In this model, the recurrent Dropout with value 0.1 is added into the base model. It is expected to improve the mean absolute error. Recurrent Dropout is a regularization method for recurrent neural networks. Dropout is applied to the updates to LSTM memory cells (or GRU states), i.e. it drops out the input/update gate in LSTM/GRU.\n\n# +\n# ns - model with first implementation based on the base \nmulti_lstm_model2 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model2, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model2)\n# -\n\n# #### (3) The Base LSTM with parameter Recurrent Dropout (0.1) + unit_forget_bias, and dropout layer (0.1)\n#\n# In this model the unit forget bias is added for the experiment. It is hoped that If True, it will add 1 to the bias of the forget gate at initialization. Setting it to true will also force bias_initializer=\"zeros\". The result looks better than the prev ious models. Nevertheless, it is so varying in each run but it seems to capture the minimal MAE.\n\n# +\n# ns - model with second implementation based on the previous model \nmulti_lstm_model3 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model3, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model3)\n# -\n\n# #### Additional Model\n# This is an additional model which I tried to tune after I have completed my report. The result is very impressive, so I keep it on this notebook so that I can make the improvement later when I have time. I tried to add the use_bias parameter to the Dense layer. It enables bias vector in this experiment. I want to utilize Gridsearch to this LSTM but time is running out. Therefore, I end up with this best model in terms of lowest MAE of 0.1724.\n\n# +\n# ns - this is a last minute trial model but it appears efficient\nmulti_lstm_model9 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1), \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model9, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model9)\n# -\n\n# #### (4) The GRU Base Model with recurrent_dropout=0.1\n#\n# GRU is a recurrent neural network (RNN) in the most recent generation, and it is also equivalent to LSTM. In all likelihood, the newer GRU model will outperform the older LSTM model. I started with a simple 32-unit network and a 0.1 recurrence dropout rate.\n\n# +\n# ns - this model is assigned as the GRU base model\nmulti_lstm_model4 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.1, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model4, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model4)\n# -\n\n# #### (5) The GRU Base Model with recurrent_dropout=0.3\n\n# +\n# ns - model with first implementation based on the GRU base \nmulti_lstm_model5 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model5, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model5)\n# -\n\n# #### (6) The GRU Base Model with recurrent_dropout=0.3 and dropout layer (0.1)\n#\n# In addition to the recurrent dropout parameter, this experiment includes the dropout layer with 0.1 figure which previously demonstrated the better version among dropout varying. The outcome below is quite as anticipated. the MAE becomes better. And again the results are very varying in each run.\n\n# +\n# ns - model with second implementation based on the GRU base \nmulti_lstm_model6 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model6, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model6)\n# -\n\n# #### (7) The Bidirectional Model with LSTM\n#\n# I proceed with the selection of Bidirectional technique which I wanted to experimenrt with both LSTM and GRU models. It is started with the Bidirectional LSTM in this trial. I believe that allowing the LSTM model to learn the input sequence both forward and backward, and then concatenating both interpretations, can be useful on some sequence prediction challenges. the result looks great but it is not good as I predicted. In each run, it shows unsatisfied value of 1.7\n\n# +\n# ns - this model is built to try on Bidirectional with LSTM layer\nfrom keras.layers import Bidirectional\nfrom keras.layers import LSTM\n\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, \n return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### (8) The Bidirectional Model with GRU\n#\n# The best model of GRU is selected to be the layer of this Bidirectional method. As mentioned ealier, this result also exibits some fluctuation and not high enough as expected.\n\n# +\n# ns - this model is built to try on Bidirectional with GRU layer\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.GRU(32, activation='tanh', \n recurrent_activation='hard_sigmoid', \n recurrent_dropout=0.1, return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### Additional stacked Bidirectional model\n# This implementation is to show that the more layer I have added into the model the more complex the model would be. This complication diminishes the ability of model. As a result, this experiment proved to be quite weak in comparison with the value of MAE.\n\n# +\n# ns - this model is built to try on Bidirectional with stacked layer\nmulti_lstm_model10 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, return_sequences=True)),\n tf.keras.layers.Dropout(0.1), \n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(8)),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model10, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model10)\n# -\n\n# ### Model Prediction and Selection\n\n# ns - The best MAE is 0.1724 which is from model#9 \n# ns - Note that every time the kernel is reran, the MAE is varied\nprediction = multi_lstm_model9.predict(multi_window.test)\n# ns - The shape of prediction\nprediction.shape\n\n# ns - Extract the traffic volum column which is number 4\npredictedValue = prediction[:,1,4]\npredictedValue\n\n# #### Denormalization\n\n# ns - Extracting the std and mean values\nstd = train_std[4]\nmean = train_mean[4]\nprint(\"std: \", std, \"mean: \", mean)\n\n# ns - Performing data conversion to denormalized values\ndenorm = []\nfor i in predictedValue:\n val = (i*std)+mean\n denorm.append(val)\n\n# #### Export data to CSV file\n\n# ns - dataframe to export raw output to csv file\nimport pandas as pd\ndf = pd.DataFrame(denorm, columns = ['prediction'])\ndf.insert(0, 'id', range(1,4994))\ndf.to_csv(\"model9.csv\", index = False)\n\n# # Thank you!\n\n\n","repo_name":"iamnatapong55/Deep-Learning-Neural-Network","sub_path":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_name":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_ext":"py","file_size_in_byte":26193,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"44313882936","text":"# +\nimport numpy as np\nimport pandas as pd\nfrom matplotlib import pyplot as plt\n\nnp.set_printoptions(suppress=True, precision=3)\n\n\n# + [markdown] heading_collapsed=true\n# # Show Graph\n\n# + hidden=true\ndef ShowGraph(a, b, X, Y, line=\"a+bx\"):\n line_x = np.arange(np.amin(X), np.amax(X), 0.01)\n if line == \"a+bx\":\n line_y = a + b * line_x\n elif line == \"ax^b\":\n line_y = a * line_x ** b\n elif line == \"ae^(bx)\":\n line_y = a * np.exp(b*line_x)\n\n plt.clf()\n plt.plot(line_x, line_y, '-r')\n plt.plot(X, Y, 'ob')\n plt.grid()\n plt.show()\n\n\n# + [markdown] heading_collapsed=true\n# # Error\n\n# + [markdown] heading_collapsed=true hidden=true\n# ## Least Square Error\n\n# + hidden=true\ndef LeastSquareError(a, b, xi, yi):\n Dt = np.sum((yi - np.average(yi)) ** 2)\n D = np.sum((yi - a - b*xi) ** 2)\n \n r = ((Dt-D)/Dt) ** 0.5\n \n return r\n\n\n# + [markdown] hidden=true\n# ## Correlation Coeficient\n\n# + hidden=true\ndef CorrelationCoef(X, Y):\n difX = X - np.average(X)\n difY = Y - np.average(Y)\n \n r = (np.sum(difX*difY))/((np.sum((difX)**2)*np.sum((difY)**2))**(0.5))\n \n return r;\n\n\n# + [markdown] heading_collapsed=true\n# # Linear Regression → y = a + bx\n\n# + hidden=true\ndef LinearRegression(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'xiyi': X_data*Y_data,\n 'xi2': X_data*X_data\n })\n\n x_bar = np.average(data['xi'])\n y_bar = np.average(data['yi'])\n\n b = (n * np.sum(data['xiyi']) - np.sum(data['xi']) *\n np.sum(data['yi']))/(n * np.sum(data['xi2']) - np.sum(data['xi'])**2)\n a = y_bar - b * x_bar\n\n if showGraph:\n ShowGraph(a, b, data['xi'], data['yi'])\n\n if showData:\n print(\"linear regression data\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([4, 6, 8, 10, 14, 16, 20, 22, 24, 28])\nY_data = np.array([30, 18, 22, 28, 14, 22, 16, 8, 20, 8])\n\na, b = LinearRegression(X_data, Y_data, showGraph=True, showData=False)\nprint(\"y = a + bx\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] heading_collapsed=true\n# # Non-Linear Regression\n\n# + [markdown] hidden=true\n# ## y = ax^b\n\n# + hidden=true\ndef NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': np.log10(X_data),\n 'qi': np.log10(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = 10 ** a\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ax^b\")\n\n if showData:\n print(\"non-linear regression data (y = ax^b)\")\n print(\"pi = log(xi)\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ax^b\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] hidden=true\n# ## y = ae^(bx)\n\n# + hidden=true\ndef NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': X_data,\n 'qi': np.log(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = np.exp(a)\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ae^(bx)\")\n\n if showData:\n print(\"non-linear regression data (y = ae^(bx))\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ae^(bx)\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# -\n\n# # Polinomial Regression\n\ndef GaussJordan(A, B, printStep=False):\n size = B.size\n matrix = A\n matrix = np.concatenate((A, B.reshape(size, 1)), axis=1)\n \n for i in range(0, size):\n if printStep:\n print(\"step\", i+1, \"\\n\")\n \n matrix[i] /= matrix[i, i]\n\n for j in range(0, size):\n if i == j:\n continue\n \n matrix[j] -= matrix[i] * matrix[j, i]\n if printStep:\n print(matrix, \"\\n\")\n \n return matrix\n\n\n# ## Ordo 2\n\ndef CorrelationCoefPoli2(x, y, a0, a1, a2):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt((Dt-D)/Dt)\n \n table = pd.DataFrame({\n 'x': x,\n 'y': y,\n 'x2': x**2,\n \"y'\": ya,\n \"y - y'\": y - ya,\n \"(y - y')^2\": (y - ya)**2,\n 'y - ybar': y - np.average(y),\n '(y - ybar)^2': (y - np.average(y))**2\n })\n \n print(\"Dt (y-ybar)^2 :\", Dt)\n print(\"D (y-ya)^2 :\", D)\n print(table)\n \n return r\n\n\ndef ShowGraphPoli2(X, Y, a0, a1, a2):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([2, 4.25, 5.25, 7.81, 9.2, 10.6], dtype=\"float32\")\nY = np.array([7.2, 7.1, 6, 5, 3.5, 5], dtype=\"float32\")\n\n# orde 2\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y\n})\n\nprint(\"y avg\", np.average(Y))\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, y, xy, x2y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2],\n [x, x2, x3],\n [x2, x3, x4]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2 = result\nprint(\"\\ny = a0 + a1x + a2x^2\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli2(X, Y, a0, a1, a2))\n\nShowGraphPoli2(X, Y, a0, a1, a2)\n\n\n# -\n\n# ## Ordo 3\n\ndef CorrelationCoefPoli3(x, y, a0, a1, a2, a3):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2) + a3*(x**3)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt(((Dt-D)/Dt))\n \n return r\n\n\ndef ShowGraphPoli3(X, Y, a0, a1, a2, a3):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) + a3*(x**3) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10,\n 11, 12, 13, 14, 15, 16, 17, 18, 19, 20,\n 21, 22, 23, 24, 25], dtype=\"float32\")\nY = np.array([0.90, 1.42, 1.30, 1.55, 1.63, 1.32, 1.35,\n 1.47, 1.95, 1.66, 1.96, 1.47, 1.92, 1.35,\n 1.05, 1.85, 1.74, 1.65, 1.78, 1.71, 2.29,\n 1.82, 2.06, 2.14, 1.27], dtype=\"float32\")\n\n# orde 3\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx3y = np.sum((X**3)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\nx5 = np.sum(X**5)\nx6 = np.sum(X**6)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'x5': X**5,\n 'x6': X**6,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y,\n 'x3y': (X**3)*Y\n})\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, x5, x6, y, xy, x2y, x3y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2, x3],\n [x, x2, x3, x4],\n [x2, x3, x4, x5],\n [x3, x4, x5, x6]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y, x3y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2, a3 = result\nprint(\"\\ny = a0 + a1x + a2x^2 + a3x^3\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"a3 =\", a3)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli3(X, Y, a0, a1, a2, a3))\n\nShowGraphPoli3(X, Y, a0, a1, a2, a3)\n# -\n\n\n","repo_name":"lutfiandri/numerical-methods-notebook","sub_path":"03 - Regression.ipynb","file_name":"03 - Regression.ipynb","file_ext":"py","file_size_in_byte":9564,"program_lang":"python","lang":"en","doc_type":"code","stars":21,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"37435824621","text":"# +\nfrom math import trunc\nfrom multiprocessing.sharedctypes import Value\nimport xml.etree.ElementTree as ET\nimport xml.dom.minidom\nimport os\nimport cv2\nfrom tqdm import tqdm, trange\nimport numpy as np\nimport pandas as pd\nimport time\nimport warnings\nwarnings.filterwarnings(\"ignore\")\n\n\nfrom data_aug.data_aug import *\nfrom data_aug.bbox_util import *\n\nimport matplotlib.pyplot as plt \nimport pickle as pkl\n# %matplotlib inline\n\n\n# -\n\nimport os\nif not os.path.exists('jittering_data'):\n os.mkdir('jittering_data')\n\n# +\norigin_path = os.path.join('.', 'jittering_data', 'origin')\norigin_Image_Path = os.path.join(origin_path, 'JPEGImages')\norigin_Annotation_Path = os.path.join(origin_path, 'Annotations_with_ID')\n\noutput_path = os.path.join('.', 'jittering_data', 'output')\noutput_Image_Path = os.path.join(output_path, 'JPEGImages')\noutput_Annotation_Path = os.path.join(output_path, 'Annotations_with_ID')\nif not os.path.exists(output_Image_Path):\n os.mkdir(output_Image_Path)\nif not os.path.exists(output_Annotation_Path):\n os.mkdir(output_Annotation_Path)\n\n# -\n\ntarget_list = [i.split('.')[0] for i in os.listdir(origin_Annotation_Path)[:]]\n\n\n# +\ndef process(target):\n origin_img_path = os.path.join(origin_Image_Path, target+'.jpg')\n origin_xml_path = os.path.join(origin_Annotation_Path, target+'.xml')\n origin_boxes = xml_to_boxes(origin_xml_path)\n origin_img = cv2.imread(origin_img_path)\n #origin_img = 255*np.ones(origin_img.shape, origin_img.dtype)\n #plt.imshow(origin_img[:,:,::-1])\n #plt.show()\n #origin_img2 = cv2.imread(origin_img_path)\n # Demo the data\n #print(origin_boxes)\n #plotted_img = draw_rect(origin_img, origin_boxes)\n #cv2.imwrite('nb.jpg', origin_img)\n #cv2.imwrite('nbt.jpg', origin_img2)\n #cv2.imshow('t',origin_img)\n #cv2.waitKey()\n #plt.show()\n # just copy\n #seq = Sequence([RandomHSV(0, 0, 100),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(50, 100, 100)\n #img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n #plt.imshow(img_[:,:,::-1])\n #plt.show()\n #return\n output_img_path = os.path.join(output_Image_Path, target+str(0)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(0)+'.xml')\n cv2.imwrite(output_img_path, origin_img)\n for i in range(1,5):\n output_img_path = os.path.join(output_Image_Path, target+str(i)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(i)+'.xml')\n \n seq = Sequence([RandomHSV(40, 50, 50),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(0, 0, 10)\n #ad = origin_img.copy()\n #print(ad.shape, ad.dtype)\n #cv2.imshow('title',origin_img)\n #cv2.waitKey()\n #cv2.imshow('title',ad)\n #cv2.waitKey()\n try:\n img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n cv2.imwrite(output_img_path, img_)\n tree = gen_tree(origin_xml_path, img_.shape,bboxes_)\n tree.write(output_xml_path)\n except:\n print(\"STOP\")\n \n #plotted_img = draw_rect(img_, bboxes_)\n #plt.imshow(plotted_img[:,:,::-1])\n #plt.show()\n \n\n \n# -\n\ndef gen_tree(xml_path, img_shape, bboxes):\n tree = ET.parse(xml_path)\n root = tree.getroot()\n root.find('size/width').text = str(img_shape[0])\n root.find('size/height').text = str(img_shape[1])\n boxes = [i for i in root.iter('object')]\n for box in boxes:\n root.remove(box)\n for box in bboxes:\n new_box = ET.Element('object')\n ET.SubElement(new_box, 'ID').text = str(int(box[4]))\n bndbox = ET.SubElement(new_box, 'bndbox')\n ET.SubElement(bndbox, 'xmin').text = str(int(box[0]))\n ET.SubElement(bndbox, 'ymin').text = str(int(box[1]))\n ET.SubElement(bndbox, 'xmax').text = str(int(box[2]))\n ET.SubElement(bndbox, 'ymax').text = str(int(box[3]))\n root.append(new_box)\n ET.indent(root)\n return tree\n\n\n# +\n\ndef xml_to_boxes(path):\n tree = ET.parse(path)\n root = tree.getroot()\n list_with_all_boxes = []\n for box in root.iter('object'):\n xmin = int(box.find(\"bndbox/xmin\").text)\n ymin = int(box.find(\"bndbox/ymin\").text)\n xmax = int(box.find(\"bndbox/xmax\").text)\n ymax = int(box.find(\"bndbox/ymax\").text)\n Id = int(box.find('ID').text)\n list_with_all_boxes.append([xmin, ymin, xmax, ymax, Id])\n return np.array(list_with_all_boxes, dtype='float64')\n\n\n\n# -\n\n#process('000007')\nfor target in tqdm(target_list):\n process(target)\n\n#\n","repo_name":"Pxter7777/DLcomp2","sub_path":"jittering.ipynb","file_name":"jittering.ipynb","file_ext":"py","file_size_in_byte":4665,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"28041736766","text":"# # Project: TMDB Data Analysis \n#\n# ## Table of Contents\n# \n\n# \n# ## Introduction\n#\n# TMDB (The Movie DataBase) is a dataset that contains data about movies. These data were collected from the community. Each movie has some interesting attributes such as:\n#
\n#\n# You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.\n\n# +\n# TODO: Display an image along with the top 5 classes\ndef plot_im_probs_classes(image_path, model, device=device):\n probs, classes = predict(image_path, model, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout()\n \ndef plot_im_probs_classes2(image_path, checkpoint, device=device):\n probs, classes = predict2(image_path, checkpoint, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout() \n\n\n# -\n\nplot_im_probs_classes('flowers/test/1/image_06760.jpg', model=reloaded_best_model, device=device)\n\nplot_im_probs_classes2('flowers/test/1/image_06760.jpg', 'checkpoint_best.pth', device=device)\n\n# Resources used for this project:\n# * Material from the first term of Udacity's Data Scientist Nanodegree program. \n# * [Transfer Learning tutorial](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html). `train_model` function was taken from this tutorial with small changes as well as other hints such as shuffling the datasets and using a scheduler. \n# * [PyTorch documentation](https://pytorch.org/docs/stable/index.html).\n# * [Argparse tutorial](https://pymotw.com/3/argparse/#argument-types) and [documentation](https://docs.python.org/dev/library/argparse.html#argparse.Namespace).\n","repo_name":"anamamatelashvili/deep_learning_project","sub_path":"Image Classifier Project.ipynb","file_name":"Image Classifier Project.ipynb","file_ext":"py","file_size_in_byte":31574,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"581944992","text":"# + [markdown] id=\"E-aBmrp8931c\"\n# # Proyecto empresa de viajes compartidos\n\n# + [markdown] id=\"uEJABzxG931c\"\n# ### Descripción del proyecto\n#\n# Durante este proyecto analizaré datos para una nueva empresa de viajes compartidos que se esta lanzando en Chicago. El objetivo será encontrar patrones en los datos y entender las preferencias de los pasajeros y el impacto de los factores externos en los viajes.\n#\n# #### Descripción de los datos\n#\n# Tabla 1: contiene datos sobre la cantidad de viajes que realizó cada compañía.\n# - company_name: nombre de la empresa de taxis\n# - trips_amount: el número de viajes de cada compañía de taxis el 15 y 16 de noviembre de 2017.\n#\n# Tabla 2: contiene datos sobre la finalización de los viajes \n# - dropoff_location_name: barrios de Chicago donde finalizaron los viajes\n# - average_trips: el promedio de viajes que terminaron en cada barrio en noviembre de 2017.\n#\n# Tabla 3:\n# Contiene datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. \n# - start_ts: fecha y hora de la recogida\n# - weather_conditions: condiciones climáticas en el momento en el que comenzó el viaje\n# - duration_seconds: duración del viaje en segundos\n\n# + [markdown] id=\"1G-R9FNe931c\"\n# ### Análisis exploratorio de datos\n\n# + id=\"PZCA0wo3931c\"\n# Importación de librerías a utilizar\n\nimport pandas as pd\nfrom scipy import stats as st\nfrom scipy.stats import levene\nimport matplotlib.pyplot as plt\nfrom seaborn import boxplot\n\n# + id=\"r3wEulH9931d\" outputId=\"1375e687-549f-4792-8c56-1ebfe3ff3c0f\"\n# Importación de archivo 1: número de viajes por compañia\n\ncompanies_trips= pd.read_csv('/datasets/project_sql_result_01.csv')\ncompanies_trips.info()\ncompanies_trips.head()\n\n\n# + id=\"MA1CVDmo931d\" outputId=\"0d2dabbc-9c46-4fb2-e35a-d36d79ab3335\"\n# Chequeo de valores ausentes\n\ncompanies_trips.isna().sum()\n\n# + id=\"UVq55dyy931d\" outputId=\"9735fac2-c40f-4ee0-8c3e-73d4238f734d\"\n# Chequeo de valores duplicados\n\ncompanies_trips.duplicated().sum()\n\n# + id=\"Bio8N7b7931d\" outputId=\"01191f13-de81-4e3f-bc49-83e4a9496fa8\"\n# Importación de archivo 2: promedio de viajes por barrio \n\ndropoff_location= pd.read_csv('/datasets/project_sql_result_04.csv')\ndropoff_location.info()\ndropoff_location.head()\n\n# + id=\"VfGtzMP-931e\" outputId=\"c6356ec7-0873-41da-c118-50a4d1e55f63\"\n# Chequeo de valores ausentes \n\ndropoff_location.isna().sum()\n\n# + id=\"bYD1uV96931e\" outputId=\"1d75298f-3319-4eb2-818a-4e3f4d0a2a96\"\n# Chequeo de valores duplicados\n\ndropoff_location.duplicated().sum()\n\n# + id=\"8aHq5XT-931e\" outputId=\"384087f9-8cea-4f88-e3a1-f5c338d36c98\"\n# Identificar los 10 principales barrios en términos de finalización del recorrido\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10)\n\n# + id=\"LdV71Unp931e\" outputId=\"5da0eaf9-c0e7-4ef7-f5c7-d51006a0a7ba\"\n#Gráfico: empresas de taxis y número de viajes\n\ncompanies_trips.plot(kind='bar',x='company_name', y='trips_amount', ylabel='trips_amount', figsize=(20,10), title='Viajes por empresa de taxi')\n\n# + [markdown] id=\"shnqBODc931e\"\n# Como podemos observar en el gráfico de arriba, la empresa de taxis más usada los días 15 y 16 de noviembre fue Flash Cab. Ésta realizó casi el doble de viajes que la empresa que le sigue: Taxi Affiliation Services. \n#\n# A su vez, dejando de lado Flash Cab que fue sin dudas la más poplar, podemos observar que hay pequeños grupos de compañías en los cuales no difiere tanto la cantidad de viajes realizados. Por ejemplo, las otras 5 copañías principales de taxis realizaron entre 9180 a 11400 viajes.\n#\n# Por otro lado, podemos observar que la mitad de las compañías de taxis tuvieron muy pocos viajes esos días. \n\n# + id=\"bzjPAp8R931e\" outputId=\"4031eafb-cf1d-4fc0-d3f1-73c5e893dc32\"\n# Gráfico 10 barrios principales por número de finalizaciones\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10).plot(kind='bar',x='dropoff_location_name', y='average_trips', ylabel='Trips amount', figsize=(20,10), title='Principales barrios')\n\n# + [markdown] id=\"Hl0_zDu_931f\"\n# Como podemos observar en el gráfico de arriba, se muestran los 10 barrios principales hacia los cuales se dirigieron los taxis (destino final). El barrio con más viajes fue Loop con un promedio de 10727 viajes en el mes de noviembre seguido por River North con un promedio de 9523 viajes.\n#\n# Después se encuentran Streeterville\ty West Loop con un promedio de viajes de 6664 y 5163 respectivamente y por último un tercer grupo en la tabla con barrios con un promedio de viajes entre 1260 a 2420 en el mes de noviembre. \n\n# + [markdown] id=\"ibJP9mHu931f\"\n# ### Prueba de hipótesis\n\n# + [markdown] id=\"9qceopHk931f\"\n# A continuación voy a importar el archivo con los datos que usare para hacer la prueba de hipótesis.\n\n# + id=\"Mj53hprj931f\" outputId=\"216d861a-a20e-40c7-bfca-172a9615cafb\"\n# Importación de archivo para trabajar sobre la hipótesis\n\narchivo_hipotesis = pd.read_csv('/datasets/project_sql_result_07.csv')\narchivo_hipotesis.info()\narchivo_hipotesis.head()\n\n# + id=\"nT7L8FIk931f\" outputId=\"04025974-4c5f-4a5f-84e1-3c46cf2c3852\"\n# Chequeo de valores ausentes\n\narchivo_hipotesis.isna().sum()\n\n# + id=\"ZqMuqgQS931f\" outputId=\"9b37c186-3558-4c3a-b90c-d3e53abca6f5\"\n# Chequeo de valores duplicados\n\narchivo_hipotesis.duplicated().sum()\n\n# + id=\"2gIJlz-9931f\" outputId=\"7e2315d2-3169-427a-c706-8443f22436e4\"\narchivo_hipotesis.value_counts()\n\n# + [markdown] id=\"aygBH6hT931f\"\n# Como podemos observar, en el archivo que nos dieron para hacer la prueba de hipótesis tenemos valores duplicados. Estos pueden deberse a que el archivo contiene los datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. Puede que haya determinadas paradas en Loop con destino hacia el aeropuerto. Al ser un punto fijo es posible que en condiciones normales, el recorrido dure siempre lo mismo; todos los los viajes que salgan de ahi a una determnada hora van a tardar lo mismo generando que haya varios datos con os mismos tiempos.\n\n# + id=\"fb9UZ9-Q931f\" outputId=\"128662bf-d6f4-403b-9c56-d506f9680a7d\"\nboxplot(data= archivo_hipotesis, x=\"duration_seconds\", y=\"weather_conditions\")\n\n# + [markdown] id=\"OxVmL-Y2931g\"\n# Voy a probar la hipótesis de que la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare cambia los sábados lluviosos.\n#\n# Hipótesis nula: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos. \n#\n# Hipótesis alternativa: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare difiere los sábados lluviosos.\n#\n# Previo a la prueba de hipótesis voy a realizar la prueba de levene para ver si las varianzas son iguales. Usaré como hipótesis nula: las varianzas son iguales y como hipótesis alternativa las varianzas no son iguales.\n\n# + id=\"3sY6DdYD931g\" outputId=\"a465661a-30aa-4dd3-92e3-eb7b9dee7791\"\n# Prueba de hipotesis nula levene: varianzas son iguales \n\ndatos_lluvia= archivo_hipotesis.query('weather_conditions == \"Bad\" and duration_seconds >0')\ndatos_good= archivo_hipotesis.query('weather_conditions ==\"Good\"and duration_seconds >0')\n\nlluvia= datos_lluvia['duration_seconds']\ngood= datos_good['duration_seconds']\n\nalpha= 0.05\n\nstat,results= levene(lluvia, good)\nprint('P_value:', results)\n\nif results < alpha:\n print('Rechazo la hipótesis nula')\nelse:\n print('No rechazo la hipótesis nula')\n\n# + [markdown] id=\"huB3ekYd931g\"\n# Como podemos observar de la prueba de Levene realizada arriba, no rechazamos la hipótesis nula que indica que las varianzas son iguales. \n#\n# A continuación realizare la prueba de hipótesis.\n\n# + id=\"ABv0Xklx931g\" outputId=\"7198fbf1-68b8-4240-bb38-b4164b0464f5\"\n# Ho: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos.\n\nalpha= 0.05\nresults= st.ttest_ind(lluvia,good, equal_var= True)\nprint('P_value:', results.pvalue)\n\nif results.pvalue < alpha:\n print('Rechazo la hipótesis nula')\nelse: \n print('No rechazo hipótesis nula')\n\n# + [markdown] id=\"Ns2N_ZTw931g\"\n# De acuerdo con los datos con los que contamos, rechazo la hipótesis nula. La duración promedio de los viajes desde el Loop hasta el aeropuerto de O'Hare cambia los sábados lluviosos.\n\n# + [markdown] id=\"dtnLXksI931g\"\n# ## Conclusión \n#\n# Para concluir, podemos decir que la empresa de taxis más utilizada es Flash Cab seguida por Taxi Affiliation Services, teniendo la primera casi el doble de viajes que la segunda. Esta diferencia puede deberse a que Flash Cab sea una empresa mucho más grande que las otras; al tener una flota de autos más grande en la calle le permite tener el doble de viajes que su competidor más cercano. \n#\n# Con respecto a los detinos finales de los viajes, Loop resulto el más concurrido seguido de cerca por River North.\n#\n# Por último la duración promedio de los viajes difiere un día lluvioso en comparación a un día de buen tiempo. Esta diferencia puede deberse principalmente al factor climático. Los días de mal tiempo puede que haya más tránsito porque la gente decide salir con sus autos y se genera más congestión. También la gente anda un poco más despacio impactando en la duración pormedio de los viajes. \n","repo_name":"CarmenPujato/practicum","sub_path":"empresa de viajes compartidos/Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_name":"Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_ext":"py","file_size_in_byte":9307,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43095479709","text":"# # Kaggle class competition\n# # Deep Learning and Neural Network Models to predict traffic volume\n# # Models: LSTM, GRUs\n# # Library used: Tensorflow, Keras\n\n# ### Set Up Libraries\n\n# +\n# ns - as typically happens, outdated components like as keywords, functions, and classes trigger a warning.\nimport warnings\nwarnings.filterwarnings('ignore')\n# ns - below are the neccessary packages utilized in this project\nimport os\nimport datetime\n\nimport IPython\nimport IPython.display\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns\nimport tensorflow as tf\n\nmpl.rcParams['figure.figsize'] = (8, 6)\nmpl.rcParams['axes.grid'] = False\n# -\n\n# ### Import data\n\n# ns - use pandas package to read in data from file located within folder \ndata = pd.read_csv('Metro_Interstate_Traffic_Volume.csv', infer_datetime_format=True)\ndata.head()\n\n# ### Inspect and Clean up\n\n# ns - investigate the data info\ndata.info()\n\n# #### It could be seen that there are both str and num types of data as categorized below\n# categorical features: holiday, weather_main, weather_description\n#\n# Continious features: temp, rain_1h, show_1h, clouds_all\n#\n# Target: traffic_volume\n\n# ### Convert date_time column to datetime type\n#\n# It is usable if we don't convert string type to datetime type in this project. It is due to the fact that time series will absolutely need datetime. So below is the demonstration.\n\n# ns - datetime type conversion\ndate_time = pd.to_datetime(data.pop('date_time'), format='%Y-%m-%d %H:%M:%S')\n\n# ### Convert to a usable signal \n#\n# In this form of signal, we would be able to utilize its capacity in the ultimate way as it reveals more insignhts as illjustrated below. \n\ntimestamp_s = date_time.map(datetime.datetime.timestamp)\n\n# +\nday = 24*60*60\nyear = (365.2425)*day\n\ndata['Day sin'] = np.sin(timestamp_s * (2 * np.pi / day))\ndata['Day cos'] = np.cos(timestamp_s * (2 * np.pi / day))\ndata['Year sin'] = np.sin(timestamp_s * (2 * np.pi / year))\ndata['Year cos'] = np.cos(timestamp_s * (2 * np.pi / year))\n# -\n\nplt.plot(np.array(data['Day sin'])[:25])\nplt.plot(np.array(data['Day cos'])[:25])\nplt.xlabel('Time [h]')\nplt.title('Time of day signal')\n\n# The best way to identify which frequencies are most essential is to use a \"fft\". As a preliminary test, below is the traffic volume rfft experiment. While we're discussing common frequencies, it's interesting to note the highs at 1/year and 1/day frequencies:\n\n# +\nfft = tf.signal.rfft(data['temp'])\nf_per_dataset = np.arange(0, len(fft))\n\nn_samples_h = len(data['temp'])\nhours_per_year = 24*365.2524\nyears_per_dataset = n_samples_h/(hours_per_year)\n\nf_per_year = f_per_dataset/years_per_dataset\nplt.step(f_per_year, np.abs(fft))\nplt.xscale('log')\nplt.ylim(0, 400000)\nplt.xlim([0.1, max(plt.xlim())])\nplt.xticks([1, 365.2524], labels=['1/Year', '1/day'])\n_ = plt.xlabel('Frequency (log scale)')\n# -\n\n# ### Drop variable \"weather_description\" and convert \"holiday\" and \"weather_main\" to dummies variables\n#\n# It is assumed that the weather description might be more irrelevant to this project or its interpretation could benefit less. Thus, I have decided to drop it. While both weather main and the holiday are deemed significant.\n\n# ns - considered to drop some variable and generate dummies variables for the rest of obj type\ndata.drop('weather_description', axis=1, inplace=True)\ndata = pd.get_dummies(data, columns = ['holiday', 'weather_main'], drop_first=True)\ndata.head()\n\n# ### Split the data\n\n# +\n# ns - as indicated in the instruction, the test set should be excluded with the size of 5000 ea.\n# ns - the slicing method is used in this data split\ncolumn_indices = {name: i for i, name in enumerate(data.columns)}\n\nn = len(data)\nprint(\"There are \", n, \"records and divisions are as below\")\n\ntest_df = data[-5000:]\nprint(\"Test set: \", test_df.shape)\n\nval_df = data[-13000:-5000]\nprint(\"Validation set: \", val_df.shape)\n\ntrain_df = data[0:-13000]\nprint(\"Training set: \", train_df.shape)\n\nnum_features = data.shape[1]\n# -\n\n# ### Data Normalization\n\n# +\ntrain_mean = train_df.mean()\ntrain_std = train_df.std()\n\ntrain_df = (train_df - train_mean) / train_std\nval_df = (val_df - train_mean) / train_std\ntest_df = (test_df - train_mean) / train_std\n\n\n# -\n\n# ### Data windowing\n#\n# The models in this project will make a set of predictions based on a window of consecutive samples from the data and below is the useful windowgenerator class for segmenting data as required for modelling.\n\nclass WindowGenerator():\n def __init__(self, input_width, label_width, shift,\n train_df=train_df, val_df=val_df, test_df=test_df,\n label_columns=None):\n # Store the raw data.\n self.train_df = train_df\n self.val_df = val_df\n self.test_df = test_df\n\n # Work out the label column indices.\n self.label_columns = label_columns\n if label_columns is not None:\n self.label_columns_indices = {name: i for i, name in\n enumerate(label_columns)}\n self.column_indices = {name: i for i, name in\n enumerate(train_df.columns)}\n\n # Work out the window parameters.\n self.input_width = input_width\n self.label_width = label_width\n self.shift = shift\n\n self.total_window_size = input_width + shift\n\n self.input_slice = slice(0, input_width)\n self.input_indices = np.arange(self.total_window_size)[self.input_slice]\n\n self.label_start = self.total_window_size - self.label_width\n self.labels_slice = slice(self.label_start, None)\n self.label_indices = np.arange(self.total_window_size)[self.labels_slice]\n\n def __repr__(self):\n return '\\n'.join([\n f'Total window size: {self.total_window_size}',\n f'Input indices: {self.input_indices}',\n f'Label indices: {self.label_indices}',\n f'Label column name(s): {self.label_columns}'])\n\n\n# As per required, the windowgenerator here will provide 6 width with 2 future prediction and 1 label.\n\nw2 = WindowGenerator(input_width=6, label_width=1, shift=2,\n label_columns=['traffic_volume'])\nw2\n\n\n# +\ndef split_window(self, features):\n inputs = features[:, self.input_slice, :]\n labels = features[:, self.labels_slice, :]\n if self.label_columns is not None:\n labels = tf.stack(\n [labels[:, :, self.column_indices[name]] for name in self.label_columns],\n axis=-1)\n\n # Slicing doesn't preserve static shape information, so set the shapes\n # manually. This way the `tf.data.Datasets` are easier to inspect.\n inputs.set_shape([None, self.input_width, None])\n labels.set_shape([None, self.label_width, None])\n\n return inputs, labels\n\nWindowGenerator.split_window = split_window\n# -\n\n# ### Note: \n#\n# Typically data in TensorFlow is packed into arrays where the outermost index is across examples (the \"batch\" dimension). The middle indices are the \"time\" or \"space\" (width, height) dimension(s). The innermost indices are the features.\n\n# +\n# ns - I tried with other stack but it failed to corporate with the model\n# Stack three slices, the length of the total window:\nexample_window = tf.stack([np.array(train_df[:w2.total_window_size]),\n np.array(train_df[100:100+w2.total_window_size]),\n np.array(train_df[200:200+w2.total_window_size])])\n\n\nexample_inputs, example_labels = w2.split_window(example_window)\n\nprint('All shapes are: (batch, time, features)')\nprint(f'Window shape: {example_window.shape}')\nprint(f'Inputs shape: {example_inputs.shape}')\nprint(f'labels shape: {example_labels.shape}')\n# -\n\n# ### Plot \n#\n# #### Here is a plot method that allows a simple visualization of the split window\n\nw2.example = example_inputs, example_labels\n\n\n# +\ndef plot(self, model=None, plot_col='traffic_volume', max_subplots=3):\n inputs, labels = self.example\n plt.figure(figsize=(12, 8))\n plot_col_index = self.column_indices[plot_col]\n max_n = min(max_subplots, len(inputs))\n for n in range(max_n):\n plt.subplot(max_n, 1, n+1)\n plt.ylabel(f'{plot_col} [normed]')\n plt.plot(self.input_indices, inputs[n, :, plot_col_index],\n label='Inputs', marker='.', zorder=-10)\n\n if self.label_columns:\n label_col_index = self.label_columns_indices.get(plot_col, None)\n else:\n label_col_index = plot_col_index\n\n if label_col_index is None:\n continue\n\n plt.scatter(self.label_indices, labels[n, :, label_col_index],\n edgecolors='k', label='Labels', c='#2ca02c', s=64)\n if model is not None:\n predictions = model(inputs)\n plt.scatter(self.label_indices, predictions[n, :, label_col_index],\n marker='X', edgecolors='k', label='Predictions',\n c='#ff7f0e', s=64)\n\n if n == 0:\n plt.legend()\n\n plt.xlabel('Time [h]')\n\nWindowGenerator.plot = plot\n# -\n\nw2.plot()\n\n\n# The plots above illustrate the 6 inputs and 1 label as we defined them ealier. \n\n# ### Create tf.data.Datasets\n\n# +\ndef make_dataset(self, data):\n data = np.array(data, dtype=np.float32)\n ds = tf.keras.preprocessing.timeseries_dataset_from_array(\n data=data,\n targets=None,\n sequence_length=self.total_window_size,\n sequence_stride=1,\n shuffle=False,\n batch_size=32,)\n\n ds = ds.map(self.split_window)\n\n return ds\n\nWindowGenerator.make_dataset = make_dataset\n\n\n# -\n\n# ### Note:\n#\n# This make_dataset method will take a time series DataFrame and convert it to a tf.data.Dataset of (input_window, label_window) pairs using the preprocessing.timeseries_dataset_from_array function. The WindowGenerator object holds training, validation and test data. Add properties for accessing them as tf.data.Datasets using the above make_dataset method.\n\n# +\n@property\ndef train(self):\n return self.make_dataset(self.train_df)\n\n@property\ndef val(self):\n return self.make_dataset(self.val_df)\n\n@property\ndef test(self):\n return self.make_dataset(self.test_df)\n\n@property\ndef example(self):\n \"\"\"Get and cache an example batch of `inputs, labels` for plotting.\"\"\"\n result = getattr(self, '_example', None)\n if result is None:\n # No example batch was found, so get one from the `.train` dataset\n result = next(iter(self.train))\n # And cache it for next time\n self._example = result\n return result\n\nWindowGenerator.train = train\nWindowGenerator.val = val\nWindowGenerator.test = test\nWindowGenerator.example = example\n# -\n\n# Each element is an (inputs, label) pair\nw2.train.element_spec\n\n# #### Iterating over a Dataset yields concrete batches:\n\nfor example_inputs, example_labels in w2.train.take(1):\n print(f'Inputs shape (batch, time, features): {example_inputs.shape}')\n print(f'Labels shape (batch, time, features): {example_labels.shape}')\n\n# #### This project trains many models, so package the training procedure into a function\n\n# +\nMAX_EPOCHS = 100\n\ndef compile_and_fit(model, window, patience=2):\n early_stopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss',\n patience=patience,\n mode='min')\n\n model.compile(loss=tf.losses.MeanSquaredError(),\n optimizer=tf.optimizers.Adam(),\n metrics=[tf.metrics.MeanAbsoluteError()])\n\n history = model.fit(window.train, epochs=MAX_EPOCHS,\n validation_data=window.val,\n callbacks=[early_stopping])\n return history\n\n\n# -\n\n# For the multi-step model, the training data consists of hourly samples. However, here, the models will learn to predict 2h of the future, given 6h of the past. Here is a Window object that generates these slices from the dataset.\n\n# +\n# ns - indicating the 2 hours future output from the 6 hours history input\nOUT_STEPS = 2\nmulti_window = WindowGenerator(input_width=6,\n label_width=OUT_STEPS,\n shift=OUT_STEPS)\n\nmulti_window.plot()\nmulti_window\n# -\n\n# Illustration here displays the 6 inputs with 2 hours prediction in the future.\n\n# ### Model Construction\n# #### (1) The Base LSTM model\n#\n# This model is expected to be used as the baseline model in this project. The error is aimed to be minimized in further steps. The architecture of this base model consists of LSTM layer with 32 unit, Dense layer and then reshape respectively.\n\n# +\nmulti_val_performance = {}\nmulti_performance = {}\n\nmulti_lstm_model = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.val)\nmulti_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model)\n# -\n\n# #### (2) The Base LSTM with parameter Recurrent Dropout (0.1) and dropout layer (0.1)\n#\n# In this model, the recurrent Dropout with value 0.1 is added into the base model. It is expected to improve the mean absolute error. Recurrent Dropout is a regularization method for recurrent neural networks. Dropout is applied to the updates to LSTM memory cells (or GRU states), i.e. it drops out the input/update gate in LSTM/GRU.\n\n# +\n# ns - model with first implementation based on the base \nmulti_lstm_model2 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model2, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model2)\n# -\n\n# #### (3) The Base LSTM with parameter Recurrent Dropout (0.1) + unit_forget_bias, and dropout layer (0.1)\n#\n# In this model the unit forget bias is added for the experiment. It is hoped that If True, it will add 1 to the bias of the forget gate at initialization. Setting it to true will also force bias_initializer=\"zeros\". The result looks better than the prev ious models. Nevertheless, it is so varying in each run but it seems to capture the minimal MAE.\n\n# +\n# ns - model with second implementation based on the previous model \nmulti_lstm_model3 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model3, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model3)\n# -\n\n# #### Additional Model\n# This is an additional model which I tried to tune after I have completed my report. The result is very impressive, so I keep it on this notebook so that I can make the improvement later when I have time. I tried to add the use_bias parameter to the Dense layer. It enables bias vector in this experiment. I want to utilize Gridsearch to this LSTM but time is running out. Therefore, I end up with this best model in terms of lowest MAE of 0.1724.\n\n# +\n# ns - this is a last minute trial model but it appears efficient\nmulti_lstm_model9 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1), \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model9, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model9)\n# -\n\n# #### (4) The GRU Base Model with recurrent_dropout=0.1\n#\n# GRU is a recurrent neural network (RNN) in the most recent generation, and it is also equivalent to LSTM. In all likelihood, the newer GRU model will outperform the older LSTM model. I started with a simple 32-unit network and a 0.1 recurrence dropout rate.\n\n# +\n# ns - this model is assigned as the GRU base model\nmulti_lstm_model4 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.1, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model4, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model4)\n# -\n\n# #### (5) The GRU Base Model with recurrent_dropout=0.3\n\n# +\n# ns - model with first implementation based on the GRU base \nmulti_lstm_model5 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model5, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model5)\n# -\n\n# #### (6) The GRU Base Model with recurrent_dropout=0.3 and dropout layer (0.1)\n#\n# In addition to the recurrent dropout parameter, this experiment includes the dropout layer with 0.1 figure which previously demonstrated the better version among dropout varying. The outcome below is quite as anticipated. the MAE becomes better. And again the results are very varying in each run.\n\n# +\n# ns - model with second implementation based on the GRU base \nmulti_lstm_model6 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model6, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model6)\n# -\n\n# #### (7) The Bidirectional Model with LSTM\n#\n# I proceed with the selection of Bidirectional technique which I wanted to experimenrt with both LSTM and GRU models. It is started with the Bidirectional LSTM in this trial. I believe that allowing the LSTM model to learn the input sequence both forward and backward, and then concatenating both interpretations, can be useful on some sequence prediction challenges. the result looks great but it is not good as I predicted. In each run, it shows unsatisfied value of 1.7\n\n# +\n# ns - this model is built to try on Bidirectional with LSTM layer\nfrom keras.layers import Bidirectional\nfrom keras.layers import LSTM\n\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, \n return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### (8) The Bidirectional Model with GRU\n#\n# The best model of GRU is selected to be the layer of this Bidirectional method. As mentioned ealier, this result also exibits some fluctuation and not high enough as expected.\n\n# +\n# ns - this model is built to try on Bidirectional with GRU layer\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.GRU(32, activation='tanh', \n recurrent_activation='hard_sigmoid', \n recurrent_dropout=0.1, return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### Additional stacked Bidirectional model\n# This implementation is to show that the more layer I have added into the model the more complex the model would be. This complication diminishes the ability of model. As a result, this experiment proved to be quite weak in comparison with the value of MAE.\n\n# +\n# ns - this model is built to try on Bidirectional with stacked layer\nmulti_lstm_model10 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, return_sequences=True)),\n tf.keras.layers.Dropout(0.1), \n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(8)),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model10, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model10)\n# -\n\n# ### Model Prediction and Selection\n\n# ns - The best MAE is 0.1724 which is from model#9 \n# ns - Note that every time the kernel is reran, the MAE is varied\nprediction = multi_lstm_model9.predict(multi_window.test)\n# ns - The shape of prediction\nprediction.shape\n\n# ns - Extract the traffic volum column which is number 4\npredictedValue = prediction[:,1,4]\npredictedValue\n\n# #### Denormalization\n\n# ns - Extracting the std and mean values\nstd = train_std[4]\nmean = train_mean[4]\nprint(\"std: \", std, \"mean: \", mean)\n\n# ns - Performing data conversion to denormalized values\ndenorm = []\nfor i in predictedValue:\n val = (i*std)+mean\n denorm.append(val)\n\n# #### Export data to CSV file\n\n# ns - dataframe to export raw output to csv file\nimport pandas as pd\ndf = pd.DataFrame(denorm, columns = ['prediction'])\ndf.insert(0, 'id', range(1,4994))\ndf.to_csv(\"model9.csv\", index = False)\n\n# # Thank you!\n\n\n","repo_name":"iamnatapong55/Deep-Learning-Neural-Network","sub_path":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_name":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_ext":"py","file_size_in_byte":26193,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"44313882936","text":"# +\nimport numpy as np\nimport pandas as pd\nfrom matplotlib import pyplot as plt\n\nnp.set_printoptions(suppress=True, precision=3)\n\n\n# + [markdown] heading_collapsed=true\n# # Show Graph\n\n# + hidden=true\ndef ShowGraph(a, b, X, Y, line=\"a+bx\"):\n line_x = np.arange(np.amin(X), np.amax(X), 0.01)\n if line == \"a+bx\":\n line_y = a + b * line_x\n elif line == \"ax^b\":\n line_y = a * line_x ** b\n elif line == \"ae^(bx)\":\n line_y = a * np.exp(b*line_x)\n\n plt.clf()\n plt.plot(line_x, line_y, '-r')\n plt.plot(X, Y, 'ob')\n plt.grid()\n plt.show()\n\n\n# + [markdown] heading_collapsed=true\n# # Error\n\n# + [markdown] heading_collapsed=true hidden=true\n# ## Least Square Error\n\n# + hidden=true\ndef LeastSquareError(a, b, xi, yi):\n Dt = np.sum((yi - np.average(yi)) ** 2)\n D = np.sum((yi - a - b*xi) ** 2)\n \n r = ((Dt-D)/Dt) ** 0.5\n \n return r\n\n\n# + [markdown] hidden=true\n# ## Correlation Coeficient\n\n# + hidden=true\ndef CorrelationCoef(X, Y):\n difX = X - np.average(X)\n difY = Y - np.average(Y)\n \n r = (np.sum(difX*difY))/((np.sum((difX)**2)*np.sum((difY)**2))**(0.5))\n \n return r;\n\n\n# + [markdown] heading_collapsed=true\n# # Linear Regression → y = a + bx\n\n# + hidden=true\ndef LinearRegression(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'xiyi': X_data*Y_data,\n 'xi2': X_data*X_data\n })\n\n x_bar = np.average(data['xi'])\n y_bar = np.average(data['yi'])\n\n b = (n * np.sum(data['xiyi']) - np.sum(data['xi']) *\n np.sum(data['yi']))/(n * np.sum(data['xi2']) - np.sum(data['xi'])**2)\n a = y_bar - b * x_bar\n\n if showGraph:\n ShowGraph(a, b, data['xi'], data['yi'])\n\n if showData:\n print(\"linear regression data\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([4, 6, 8, 10, 14, 16, 20, 22, 24, 28])\nY_data = np.array([30, 18, 22, 28, 14, 22, 16, 8, 20, 8])\n\na, b = LinearRegression(X_data, Y_data, showGraph=True, showData=False)\nprint(\"y = a + bx\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] heading_collapsed=true\n# # Non-Linear Regression\n\n# + [markdown] hidden=true\n# ## y = ax^b\n\n# + hidden=true\ndef NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': np.log10(X_data),\n 'qi': np.log10(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = 10 ** a\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ax^b\")\n\n if showData:\n print(\"non-linear regression data (y = ax^b)\")\n print(\"pi = log(xi)\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ax^b\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] hidden=true\n# ## y = ae^(bx)\n\n# + hidden=true\ndef NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': X_data,\n 'qi': np.log(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = np.exp(a)\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ae^(bx)\")\n\n if showData:\n print(\"non-linear regression data (y = ae^(bx))\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ae^(bx)\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# -\n\n# # Polinomial Regression\n\ndef GaussJordan(A, B, printStep=False):\n size = B.size\n matrix = A\n matrix = np.concatenate((A, B.reshape(size, 1)), axis=1)\n \n for i in range(0, size):\n if printStep:\n print(\"step\", i+1, \"\\n\")\n \n matrix[i] /= matrix[i, i]\n\n for j in range(0, size):\n if i == j:\n continue\n \n matrix[j] -= matrix[i] * matrix[j, i]\n if printStep:\n print(matrix, \"\\n\")\n \n return matrix\n\n\n# ## Ordo 2\n\ndef CorrelationCoefPoli2(x, y, a0, a1, a2):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt((Dt-D)/Dt)\n \n table = pd.DataFrame({\n 'x': x,\n 'y': y,\n 'x2': x**2,\n \"y'\": ya,\n \"y - y'\": y - ya,\n \"(y - y')^2\": (y - ya)**2,\n 'y - ybar': y - np.average(y),\n '(y - ybar)^2': (y - np.average(y))**2\n })\n \n print(\"Dt (y-ybar)^2 :\", Dt)\n print(\"D (y-ya)^2 :\", D)\n print(table)\n \n return r\n\n\ndef ShowGraphPoli2(X, Y, a0, a1, a2):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([2, 4.25, 5.25, 7.81, 9.2, 10.6], dtype=\"float32\")\nY = np.array([7.2, 7.1, 6, 5, 3.5, 5], dtype=\"float32\")\n\n# orde 2\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y\n})\n\nprint(\"y avg\", np.average(Y))\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, y, xy, x2y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2],\n [x, x2, x3],\n [x2, x3, x4]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2 = result\nprint(\"\\ny = a0 + a1x + a2x^2\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli2(X, Y, a0, a1, a2))\n\nShowGraphPoli2(X, Y, a0, a1, a2)\n\n\n# -\n\n# ## Ordo 3\n\ndef CorrelationCoefPoli3(x, y, a0, a1, a2, a3):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2) + a3*(x**3)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt(((Dt-D)/Dt))\n \n return r\n\n\ndef ShowGraphPoli3(X, Y, a0, a1, a2, a3):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) + a3*(x**3) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10,\n 11, 12, 13, 14, 15, 16, 17, 18, 19, 20,\n 21, 22, 23, 24, 25], dtype=\"float32\")\nY = np.array([0.90, 1.42, 1.30, 1.55, 1.63, 1.32, 1.35,\n 1.47, 1.95, 1.66, 1.96, 1.47, 1.92, 1.35,\n 1.05, 1.85, 1.74, 1.65, 1.78, 1.71, 2.29,\n 1.82, 2.06, 2.14, 1.27], dtype=\"float32\")\n\n# orde 3\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx3y = np.sum((X**3)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\nx5 = np.sum(X**5)\nx6 = np.sum(X**6)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'x5': X**5,\n 'x6': X**6,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y,\n 'x3y': (X**3)*Y\n})\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, x5, x6, y, xy, x2y, x3y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2, x3],\n [x, x2, x3, x4],\n [x2, x3, x4, x5],\n [x3, x4, x5, x6]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y, x3y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2, a3 = result\nprint(\"\\ny = a0 + a1x + a2x^2 + a3x^3\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"a3 =\", a3)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli3(X, Y, a0, a1, a2, a3))\n\nShowGraphPoli3(X, Y, a0, a1, a2, a3)\n# -\n\n\n","repo_name":"lutfiandri/numerical-methods-notebook","sub_path":"03 - Regression.ipynb","file_name":"03 - Regression.ipynb","file_ext":"py","file_size_in_byte":9564,"program_lang":"python","lang":"en","doc_type":"code","stars":21,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"37435824621","text":"# +\nfrom math import trunc\nfrom multiprocessing.sharedctypes import Value\nimport xml.etree.ElementTree as ET\nimport xml.dom.minidom\nimport os\nimport cv2\nfrom tqdm import tqdm, trange\nimport numpy as np\nimport pandas as pd\nimport time\nimport warnings\nwarnings.filterwarnings(\"ignore\")\n\n\nfrom data_aug.data_aug import *\nfrom data_aug.bbox_util import *\n\nimport matplotlib.pyplot as plt \nimport pickle as pkl\n# %matplotlib inline\n\n\n# -\n\nimport os\nif not os.path.exists('jittering_data'):\n os.mkdir('jittering_data')\n\n# +\norigin_path = os.path.join('.', 'jittering_data', 'origin')\norigin_Image_Path = os.path.join(origin_path, 'JPEGImages')\norigin_Annotation_Path = os.path.join(origin_path, 'Annotations_with_ID')\n\noutput_path = os.path.join('.', 'jittering_data', 'output')\noutput_Image_Path = os.path.join(output_path, 'JPEGImages')\noutput_Annotation_Path = os.path.join(output_path, 'Annotations_with_ID')\nif not os.path.exists(output_Image_Path):\n os.mkdir(output_Image_Path)\nif not os.path.exists(output_Annotation_Path):\n os.mkdir(output_Annotation_Path)\n\n# -\n\ntarget_list = [i.split('.')[0] for i in os.listdir(origin_Annotation_Path)[:]]\n\n\n# +\ndef process(target):\n origin_img_path = os.path.join(origin_Image_Path, target+'.jpg')\n origin_xml_path = os.path.join(origin_Annotation_Path, target+'.xml')\n origin_boxes = xml_to_boxes(origin_xml_path)\n origin_img = cv2.imread(origin_img_path)\n #origin_img = 255*np.ones(origin_img.shape, origin_img.dtype)\n #plt.imshow(origin_img[:,:,::-1])\n #plt.show()\n #origin_img2 = cv2.imread(origin_img_path)\n # Demo the data\n #print(origin_boxes)\n #plotted_img = draw_rect(origin_img, origin_boxes)\n #cv2.imwrite('nb.jpg', origin_img)\n #cv2.imwrite('nbt.jpg', origin_img2)\n #cv2.imshow('t',origin_img)\n #cv2.waitKey()\n #plt.show()\n # just copy\n #seq = Sequence([RandomHSV(0, 0, 100),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(50, 100, 100)\n #img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n #plt.imshow(img_[:,:,::-1])\n #plt.show()\n #return\n output_img_path = os.path.join(output_Image_Path, target+str(0)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(0)+'.xml')\n cv2.imwrite(output_img_path, origin_img)\n for i in range(1,5):\n output_img_path = os.path.join(output_Image_Path, target+str(i)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(i)+'.xml')\n \n seq = Sequence([RandomHSV(40, 50, 50),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(0, 0, 10)\n #ad = origin_img.copy()\n #print(ad.shape, ad.dtype)\n #cv2.imshow('title',origin_img)\n #cv2.waitKey()\n #cv2.imshow('title',ad)\n #cv2.waitKey()\n try:\n img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n cv2.imwrite(output_img_path, img_)\n tree = gen_tree(origin_xml_path, img_.shape,bboxes_)\n tree.write(output_xml_path)\n except:\n print(\"STOP\")\n \n #plotted_img = draw_rect(img_, bboxes_)\n #plt.imshow(plotted_img[:,:,::-1])\n #plt.show()\n \n\n \n# -\n\ndef gen_tree(xml_path, img_shape, bboxes):\n tree = ET.parse(xml_path)\n root = tree.getroot()\n root.find('size/width').text = str(img_shape[0])\n root.find('size/height').text = str(img_shape[1])\n boxes = [i for i in root.iter('object')]\n for box in boxes:\n root.remove(box)\n for box in bboxes:\n new_box = ET.Element('object')\n ET.SubElement(new_box, 'ID').text = str(int(box[4]))\n bndbox = ET.SubElement(new_box, 'bndbox')\n ET.SubElement(bndbox, 'xmin').text = str(int(box[0]))\n ET.SubElement(bndbox, 'ymin').text = str(int(box[1]))\n ET.SubElement(bndbox, 'xmax').text = str(int(box[2]))\n ET.SubElement(bndbox, 'ymax').text = str(int(box[3]))\n root.append(new_box)\n ET.indent(root)\n return tree\n\n\n# +\n\ndef xml_to_boxes(path):\n tree = ET.parse(path)\n root = tree.getroot()\n list_with_all_boxes = []\n for box in root.iter('object'):\n xmin = int(box.find(\"bndbox/xmin\").text)\n ymin = int(box.find(\"bndbox/ymin\").text)\n xmax = int(box.find(\"bndbox/xmax\").text)\n ymax = int(box.find(\"bndbox/ymax\").text)\n Id = int(box.find('ID').text)\n list_with_all_boxes.append([xmin, ymin, xmax, ymax, Id])\n return np.array(list_with_all_boxes, dtype='float64')\n\n\n\n# -\n\n#process('000007')\nfor target in tqdm(target_list):\n process(target)\n\n#\n","repo_name":"Pxter7777/DLcomp2","sub_path":"jittering.ipynb","file_name":"jittering.ipynb","file_ext":"py","file_size_in_byte":4665,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"28041736766","text":"# # Project: TMDB Data Analysis \n#\n# ## Table of Contents\n# \n\n# \n# ## Introduction\n#\n# TMDB (The Movie DataBase) is a dataset that contains data about movies. These data were collected from the community. Each movie has some interesting attributes such as:\n# \n# - original title
\n# - genres
\n# - cast
\n# - director
\n# - release year
\n# - budget
\n# - revenue
\n# - rating\n#
\n# To get an insight on the different data patterns, I'll try to breakdown the dataset and answer some question, such as:\n# \n# - Which genres are most popular from year to year?\n#
- Most popular actor from year to year?\n#
- Did movies with higher vote count received a better rating?\n#
- What kinds of properties are associated with movies that have high revenues?\n#
- How did the number of film production changed over time?\n#
\n\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\n# %matplotlib inline\nimport seaborn as sns\n\n# \n# ## Data Wrangling\n#\n# In this section of the report, I will load in the data, check for cleanliness, and then trim and clean the dataset for analysis.\n#\n# ### General Properties\n\n# load the dataset\ndf = pd.read_csv('tmdb-movies.csv')\ndf.head()\n\n# more info about data types of columns, missing values.\ndf.info()\n\n# Investigate the shape of dataset\ndf.shape\n\n# check for duplicates in data set, and count them if exists\nsum(df.duplicated())\n\n#To understand the statistics of the dataset\ndf.describe()\n\n#\n# ### Initial Impression of Data\n#\n# The dataset contains some columns that aren't useful for my investigation. To simplify the analysis, I'll remove these columns:\n# 1. imdb_id\n# 2. homepage\n# 3. tagline\n# 4. release_date\n# 5. overview\n# 6. keywords\n# \n#\n#\n# Upon a deeper look into the dataset, there're movies with some important values such as budget, and revenue set to 0. This is unrealistic and can't be analyized. Thus, any movie with 0 value in one, or more of these columns (\"budget_adj\", \"revenue_adj\", \"budget\", \"revenue\", \"runtime\") will be removed.\n#\n# Additionally, columns (\"cast\", \"directors\", \"genres\", \"production_companies\") have enteries seperated by \"|\". These could be problematic when trying to extract the different entities within one entery. Thus, the entities in one entery will be seperated into different columns to ease the data exploration process.\n\n# ## Data Cleaning\n\n# Removing the irrelevant columns\ndf.drop(['imdb_id','homepage','tagline','release_date','keywords','overview'], axis = 1, inplace = True)\ndf.info()\n\n#Removing the duplicate entery, and check again\ndf.drop_duplicates(inplace=True)\nsum(df.duplicated())\n\n#Replace all the films with 0 values in the specified columns to NaN values, then remove all NaN enteries from the dataset\ndf['revenue_adj'].replace(0, np.NAN, inplace=True)\ndf['revenue'].replace(0, np.NAN, inplace=True)\ndf['budget_adj'].replace(0, np.NAN, inplace=True)\ndf['budget'].replace(0, np.NAN, inplace=True)\ndf['runtime'].replace(0, np.NAN, inplace=True)\ndf.dropna(axis=0, inplace=True)\n\n# +\n#Split the colums into seperate ones (split by '|')\n#reference: https://apassionatechie.wordpress.com/2018/02/24/how-do-i-split-a-string-into-several-columns-in-a-dataframe-with-pandas-python/\n\ncast_df = (df['cast'].str.split('|', expand=True).rename(columns=lambda x: f\"cast{x+1}\"))\ndir_df = (df['director'].str.split('|', expand=True).rename(columns=lambda x: f\"director{x+1}\"))\ngen_df = (df['genres'].str.split('|', expand=True).rename(columns=lambda x: f\"genre{x+1}\"))\nprod_df = (df['production_companies'].str.split('|', expand=True).rename(columns=lambda x: f\"production_company{x+1}\"))\n# -\n\n#Updating the dataset\ndf = df.join([cast_df, gen_df, prod_df, dir_df])\ndf = df.drop(['cast', 'director', 'genres', 'production_companies'], axis=1)\n\n#Check the new shape of the dataset\ndf.info()\ndf.shape\n\n# ## Exploratory Data Analysis\n\n# ### Question 1: Which genres are most popular from year to year?\n\n# +\n#Determine most popular genres each year\n#create a dataframe containing all generes columns\ngen_year_df = df.loc[:, 'genre1': 'genre5']\n#add the release_date to the dataframe\ngen_year_df = gen_year_df.join(df['release_year'])\n#identify the unique release years\nyears = gen_year_df['release_year'].unique()\n#empty typles list to carry the popular genre for every year\npop_gen = []\nfor year in years:\n #create a new df with only films where their release year is equal to the iterating year \n year_df = gen_year_df[gen_year_df['release_year']==year]\n #remove column release_year\n year_df = year_df.drop(['release_year'], axis = 1)\n #find the most recurring genre entry\n most_pop = year_df.stack().value_counts().idxmax() \n \n pop_gen.append((year, most_pop))\n \n#Create a dataframe to show the sorted \"year - popular genre\" tuples\npop_df = pd.DataFrame(pop_gen, columns=['Year','Most popular genre']).sort_values('Year').reset_index().drop(['index'], axis=1)\npop_df\n# -\n\n# 'Drama' seems to be predominantly the most popular genre throughout the years. \n\n# ### Question 2: Most popular actor from year to year?\n\n# +\n#Same as the previous process\ncast_year_df = df.loc[:, 'cast1': 'cast5']\ncast_year_df = cast_year_df.join(df['release_year'])\nyears = cast_year_df['release_year'].unique()\npop_cast = []\nfor year in years:\n year_df = cast_year_df[cast_year_df['release_year']==year] \n year_df = year_df.drop(['release_year'], axis = 1)\n most_pop = year_df.stack().value_counts().idxmax() \n \n pop_cast.append((year, most_pop))\n \npop_df = pd.DataFrame(pop_cast, columns=['Year','Most popular actor']).sort_values('Year').reset_index().drop(['index'], axis=1)\npop_df\n# -\n\n# The table shows some of the biggest names in Hollywood, and when were they most popular.\n\n# ### Question 3: Did movies with higher vote count received a better rating?\n\n# +\ndf_vote = df.loc[:, 'vote_count' : 'vote_average']\n\ndf_vote.plot(x='vote_count', y='vote_average', kind='scatter')\nplt.xlabel('Vote Count')\nplt.ylabel('Vote Average')\n# -\n\n# It's hard to deduce a relationship from this graph, we can check the correlation value\n\ndf_vote.corr()\n\n# The corrolation is 0.39 which is a \"moderate to low\" correlation. It seems like vote count doesn't hold a strong effect on the vote average.\n\n# ### Question 4: What kinds of properties are associated with movies that have high revenues?\n\n#Define movies with high revenues as : movies with revenue higher than the mean of revenues and higher than its own budget \nrev_df = df\nrev_df['revenue_adj'].mean()\n\nrev_df = df.query('revenue_adj >= 138715934.34610373' and 'revenue_adj >=budget_adj')\n\n# +\n#For the high revenue films\n#Find the most frequent director\ndirector = rev_df.loc[:, 'director1': 'director5']\npop_dir = director.stack().value_counts().idxmax()\n\n#find the most frequent cast member\ncast = rev_df.loc[:, 'cast1': 'cast5']\npop_cast = cast.stack().value_counts().idxmax()\n\n#find the most frequent genre\ngen = rev_df.loc[:, 'genre1': 'genre5']\npop_gen = gen.stack().value_counts().idxmax()\n\n#find the most frequent production company\nprod = rev_df.loc[:, 'production_company1': 'production_company5']\npop_prod = prod.stack().value_counts().idxmax()\n\n#find the most frequent release year\nrelease_year = rev_df['release_year'].value_counts().idxmax()\n\n#find the avg budget\navg_budget = rev_df['budget_adj'].mean()\n\n#find the avg vote\navg_vote = rev_df['vote_average'].mean()\n\n#find the avg revenue\navg_rev = rev_df['revenue_adj'].mean()\n# -\n\nproperties = {'Popular director' : pop_dir, \n 'Popular cast' : pop_cast,\n 'Popular genre' : pop_gen,\n 'Production company' : pop_prod,\n 'Release year' : release_year,\n 'Average budget' : avg_budget,\n 'Average Revenue' : avg_rev,\n 'Average vote' : avg_vote}\nproperties\n\n# ### Question 5: How did the number of film production changed over time?\n\n# +\nyears = df['release_year'].unique()\nprod = []\nfor year in years:\n year_df = df[df['release_year']==year] \n count = len(year_df.index) \n \n prod.append((year, count))\n \nprod_df = pd.DataFrame(prod, columns=['Year','Number of movies']).sort_values('Year').reset_index().drop(['index'], axis=1)\nprod_df\n# -\n\nprod_df.plot(x='Year', y='Number of movies', kind='bar', figsize=(20,10))\nplt.xlabel('Year')\nplt.ylabel('Number of movies produced')\n\n# There's a constant increase with the number of movies produced annually. This makes sense, the industry is expected to grow as more people now can access movies easier than older years.\n\n# ## Conclusions\n\n# Afetr cleaning the data, I was able to answer the research questions:\n#\n# #### Question 1: Which genres are most popular from year to year?\n# Throughout the different years, 'Drama' seems to be the most popular genre.\n#\n# #### Question 2: Most popular actor from year to year?\n# The table showed some of the biggest names in Hollywood, and when was their peak.\n#\n# #### Question 3: Did movies with higher vote count received a better rating?\n# Not quite true. The corralation doesn't indicate a strong relationship between vote counts and higher vote average.\n#\n# #### Question 4: What kinds of properties are associated with movies that have high revenues?\n# \n# \n# | Popular director | \n# Steven Spielberg | \n#
\n# \n# | Popular actor | \n# Robert De Niro | \n#
\n# \n# | Popular genre | \n# Drama | \n#
\n# \n# | Production Company | \n# Universal Pictures | \n#
\n# \n# | Release year | \n# 2011 | \n#
\n# \n# | Average budget | \n# 47,896,614 | \n#
\n# \n# | Average Revenue | \n# 184,138,833 | \n#
\n# \n# | Average vote | \n# 6.29 | \n#
\n#
\n#\n# #### Question 5: How did the number of film production changed over time?\n# There's a significant increase in the number of movies produced each year.\n\n# ### Limitations\n# In the cleaning stage, many movies were discarded due to insuffecient data given. The original dataset contained 10,866 movie entry, and the cleaned data set contains 3,805; this means almost 65% of the enteries were discarded. Of course this will have an effect on all the answers found, for instance the number of movies produced each year may not accuratly represent the actual case, in case more movies from certain years were discarded than the other years.\n#\n# In the revenue data, there was no normalization, exchange rate, or currency conversion considered during the nalysis. The analysis was limited to the given numerical values.\n","repo_name":"Yomna521/Investigate-a-dataset","sub_path":"Investigate-a-dataset.ipynb","file_name":"Investigate-a-dataset.ipynb","file_ext":"py","file_size_in_byte":11104,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"366680518","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#  \n\n# + id=\"0iDzG5n8eBm_\" colab_type=\"code\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34} outputId=\"fcb93288-5194-40c8-c624-7fff7f2ac118\"\nsome_list = [5, 10, 20, 30, 40, 50]\n\ndef only_ends(the_list):\n ends_list = []\n ends_list.append(the_list[0])\n ends_list.append(the_list[-1])\n return ends_list\n\nonly_ends(some_list)\n","repo_name":"tastiz/python_challenges","sub_path":"onlyListEnds.ipynb","file_name":"onlyListEnds.ipynb","file_ext":"py","file_size_in_byte":607,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"28612213132","text":"import pkgutil\nimport inspect\nimport importlib\nimport datetime\n\nbuiltin_modules = dir(__builtin__)\n\n# +\n# pkgutil.iter_modules()\n# -\n\nmodule_list = []\nfor module_name in pkgutil.iter_modules():\n [file_finder, module_name, ispkg] = module_name\n if not ispkg:\n continue\n # print(module_name)\n try:\n module = importlib.import_module(module_name)\n module_list.append([file_finder, module_name, ispkg])\n except:\n pass\n # print(module_name['ispkg'])\n # print(module_name[2])\n # for key in module_name:\n # print('\\t', key)\n\n# +\nmodule_name_list = []\ndef get_module_name(module_obj):\n str_module_obj = str(module_obj)\n add_module_obj = str_module_obj.split(\" -1:\n return_bool = True\n elif str(obj_name)[:2] == '__' and str(obj_name)[-2:] == '__':\n return_bool = True\n return return_bool\n \ndef get_modules(module_obj):\n for obj_name, obj in inspect.getmembers(module_obj, inspect.ismodule):\n try:\n if check_builtin_function(obj_name, obj):\n continue\n if get_module_name(obj):\n continue\n get_modules(obj)\n get_classes(obj)\n except:\n pass\n \ndef get_classes(module_obj):\n for obj_name, obj in inspect.getmembers(module_obj, inspect.isclass):\n try:\n if check_builtin_function(obj_name, obj):\n continue\n if get_class_name(obj):\n continue\n # get_modules(obj)\n get_classes(obj)\n # get_functions(obj)\n except:\n pass\n \ndef get_functions(module_obj, module_name_list=[]):\n for obj_name, obj in inspect.getmembers(module_obj, inspect.isfunction):\n try:\n if check_builtin_function(obj_name, obj):\n continue\n module_name_list.append(obj_name)\n # print('\\t\\t\\t', obj_name, obj)\n except:\n pass\n return module_name_list\n\n\n# -\n\nstart_time = datetime.datetime.now()\nprint('start_time', start_time)\nfor module_atr in module_list:\n # print('*'*50)\n # print(module_atr)\n [file_finder, module_name, ispkg] = module_atr\n # print(file_finder, module_name, ispkg)\n module = importlib.import_module(module_name)\n # print('\\t', module)\n # print(module)\n # print('')\n # print('\\t', dir(module))\n \n get_modules(module)\n get_classes(module)\n # get_functions(module)\n # break\nend_time = datetime.datetime.now()\nprint('end_time', end_time)\nprint(end_time - start_time)\n\nfor idx, x in enumerate(class_name_list):\n print(idx, '\\t', x)\n if idx > 10:\n break\n\nfor idx, x in enumerate(module_name_list):\n print(idx, '\\t', x)\n if idx > 10:\n break\n\nidx = 1\nclass_name = class_name_list[idx]\nprint(class_name)\nmodule_name = class_name.split('.')\nprint(module_name, len(module_name))\n# print(module_name[:-1])\n# print('.'.join(module_name[:-1]))\nif len(mdoule_name) > 1:\n module_name = '.'.join(module_name[:-1])\nprint(module_name)\nclass_obj = importlib.import_module(module_name)\n\n# dir(class_obj)\n# print(inspect.getmembers(class_obj))\n# function_name_list = []\n# for obj in inspect.getmembers(class_obj, inspect.isfunction):\n# # for obj in inspect.getmembers(class_obj):\n# print(obj)\nfunction_name_list = get_functions(class_obj)\n\nfor idx in range(0, len(function_name_list)):\n print('*'*50)\n # print(idx, '\\t', function_name_list[idx])\n # function_obj = getattr(class_obj, function_name_list[idx])\n # print('\\t', function_obj)\n # print(inspect.getsource(function_obj))\n print(inspect.getsource(getattr(class_obj, function_name_list[idx])))\n\nprint(inspect.getsource(class_obj))\n\n\n","repo_name":"chibadai/JupyterLab","sub_path":"notebooks/pkgutil-to-source-code-20200308-1.ipynb","file_name":"pkgutil-to-source-code-20200308-1.ipynb","file_ext":"py","file_size_in_byte":4494,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43231152486","text":"# + [markdown] pycharm={\"name\": \"#%% md\\n\"}\n# ### 从零实现线性回归\n#\n# #### 基础操作定义\n\n# +\nimport random\nimport torch\nfrom matplotlib import pyplot as plt\nimport os\n\n# 避免当下版本matplotlib与pytorch的冲突\nos.environ['KMP_DUPLICATE_LIB_OK'] = 'True'\n\n\n# + pycharm={\"name\": \"#%%\\n\"}\ndef synthetic_data(w, b, num_examples): #@save\n \"\"\"生成 y = Xw + b + 噪声\"\"\"\n X = torch.normal(0, 1, (num_examples, len(w)))\n y = torch.matmul(X, w) + b\n y += torch.normal(0, 0.01, y.shape)\n return X, y.reshape((-1, 1))\n\n\ntrue_w = torch.tensor([2, -3.4])\ntrue_b = 4.2\nfeatures, labels = synthetic_data(true_w, true_b, 1000)\n\nplt.scatter(features[:, 1].detach().numpy(), labels.detach().numpy(), 1)\nplt.show()\n\n\n# + pycharm={\"name\": \"#%%\\n\"}\ndef data_iter(batch_size, features, labels):\n \"\"\"\n 随机采样\n :param batch_size: 随机样本大小\n :param features: 数据集\n :param labels: 数据集对应的标签\n :return: 随机样本\n \"\"\"\n num_examples = len(features)\n indices = list(range(num_examples))\n # 这些样本是随机读取的,没有特定的顺序\n random.shuffle(indices)\n for i in range(0, num_examples, batch_size):\n batch_indices = torch.tensor(indices[i: min(i + batch_size, num_examples)])\n yield features[batch_indices], labels[batch_indices]\n\n\n# + [markdown] pycharm={\"name\": \"#%% md\\n\"}\n# #### 模型定义\n\n# + pycharm={\"name\": \"#%%\\n\"}\n# 初始化模型参数\n# 从均值为0、标准差为0.01的正态分布中采样随机数来初始化权重,并将偏置初始化为0\nw = torch.normal(0, 0.01, size=(2, 1), requires_grad=True)\nb = torch.zeros(1, requires_grad=True)\n\n\n# 定义线性回归模型\ndef lin_reg(X, w, b):\n return torch.matmul(X, w) + b\n\n\n# 定义损失函数\ndef squared_loss(y_hat, y):\n return (y_hat - y.reshape(y_hat.shape)) ** 2 / 2\n\n\n# 定义优化算法: 小批量随机梯度下降\ndef sgd(params, lr, batch_size):\n with torch.no_grad():\n for param in params:\n param -= lr * param.grad / batch_size\n param.grad.zero_()\n\n\n# -\n\n# #### 训练模型\n\n# + pycharm={\"name\": \"#%%\\n\"}\ndef train(lr, num_epochs, batch_size, net, loss):\n for epoch in range(num_epochs):\n for X, y in data_iter(batch_size, features, labels):\n # X和y的小批量损失\n l = loss(net(X, w, b), y)\n\n # l的形状是(batch_size, 1),而不是一个标量\n # l中的所有元素被加到一起,并以此计算关于[`w`, `b`]的梯度\n l.sum().backward()\n\n # 使用参数的梯度更新参数\n sgd([w, b], lr, batch_size)\n # 接下来的计算不需要梯度,所以使用语句with torch.no_grad():\n with torch.no_grad():\n train_l = loss(net(features, w, b), labels)\n print(f'epoch {epoch + 1}, loss {float(train_l.mean()):f}')\n\n\ntrain(0.2, 10, 10, lin_reg, squared_loss)\nprint(f'w的估计误差: {true_w - w.reshape(true_w.shape)}')\nprint(f'b的估计误差: {true_b - b}')\n# -\n\n# ### 简洁实现\n\n# + pycharm={\"name\": \"#%%\\n\"}\nimport numpy as np\nfrom torch.utils import data\nfrom torch import nn\n\n# 生成数据集\ntrue_w = torch.tensor([2, -3.4])\ntrue_b = 4.2\nfeatures, labels = synthetic_data(true_w, true_b, 1000)\n\n\n# 加载数据集\ndef load_array(data_arrays, batch_size, is_train=True): #@save\n # 构造一个PyTorch数据迭代器\n dataset = data.TensorDataset(*data_arrays)\n # 从数据集dataset中抽取batch_size个数据,shuffle指示是否随机抽取\n return data.DataLoader(dataset, batch_size, shuffle=is_train)\n\n\n\"\"\"\n创建模型\n 利用Sequential容器建立神经网络\n 利用Linear生成全连接层,也就是本次需要的线性回归函数,两个参数分别为输入个数与输出个数\n\"\"\"\nnet = nn.Sequential(nn.Linear(2, 1))\n# 初始化权重与偏差\n# 利用平均值为0,方差为0.01的高斯分布初始化模型权重\nnet[0].weight.data.normal_(0, 0.01)\nnet[0].bias.data.fill_(0)\n\n# 损失函数与优化算法\nloss = nn.MSELoss()\nopt = torch.optim.SGD(net.parameters(), lr=0.3)\n\n# 训练\nbatch_size = 10\ndata_iter = load_array((features, labels), batch_size)\nnum_epochs = 3\nfor i in range(num_epochs):\n for X, y in data_iter:\n # 计算损失函数\n l = loss(net(X), y)\n # 清零梯度\n opt.zero_grad()\n # 计算梯度\n l.backward()\n # 更新模型\n opt.step()\n l = loss(net(features), labels)\n print(f'epoch {i + 1}, loss {l:f}')\n","repo_name":"MikuAndRabbit/note-DiveIntoDeepLeaning","sub_path":"03-linear_neural_network/01-linear_regression.ipynb","file_name":"01-linear_regression.ipynb","file_ext":"py","file_size_in_byte":4515,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"18281383662","text":"import pandas as pd\nimport requests\nfrom pycoingecko import CoinGeckoAPI\nfrom pprint import pprint\nfrom binance import Client\nfrom datetime import datetime\nimport yfinance as yf\n\ncg = CoinGeckoAPI()\nclient = Client()\n\n# Se decide hacer un análisis sobre la categoría Capa 1 (L1) de criptomonedas por su capitalización de mercado.\n\n# Categorías de Interes\n\nCategorias=['Layer 1 (L1)']\n\n#Lista de todas las categorías\ncategoria=pd.DataFrame(cg.get_coins_categories_list())\ncategoria[categoria['name'].isin(Categorias)]\n\n#Lista de todas las categorías con datos de mercado\ndatos_merc_ct=pd.DataFrame(cg.get_coins_categories(order='market_cap_asc'))\ndatos_merc_ct=datos_merc_ct.sort_values(by='market_cap',ascending=False).dropna(subset='market_cap')\ndatos_merc_ct.head()\n\n\ndef get_coin_list():\n return cg.get_coins_list()\n\n\ndf1=get_coin_list()\ndf1=pd.DataFrame(df1)\n\n#crypto_symbol=['btc','eth','bnb','ada','sol','usdt','usdc','dai','busd','tusd']\ndf1[df1['name'].str.contains('TrueUS', case=False)]\n\n\n# Criptomonedas seleccionadas para el análisis.\n\ncriptos_order=['Bitcoin','Ethereum','BNB','Cardano','Solana','Polkadot','Bitcoin Cash','Avalanche',\n 'Cosmos Hub','Hedera']\ncriptos = df1[df1['name'].isin(criptos_order)]\ncriptos\n\ncriptos['symbol']\n\n\n# ### Generando Datos para las diferentes Criptos Seleccionadas:\n\n# Obtenga datos de mercado históricos que incluyen precio, capitalización de mercado y volumen de 24 horas dentro de un rango de marca de tiempo (granularidad automática)\n\ndef get_coin_market_chart_range_by_id(id:str,markert_chart:str,to:str,From:str):\n ''' Ejem: id = Bitcoin , markert_chart = usd, to = 1660539600, from = 1692079200 '''\n return cg.get_coin_market_chart_range_by_id(id,markert_chart,to,From)\n\n\n# +\nfrom datetime import datetime\n#1660971600 ----> 2022-08-20\n#1692424800 ----> 2023-08-19\n\ntimestamp = 1692424800\n\n# Convertir el timestamp a una fecha legible\nfecha_legible = datetime.utcfromtimestamp(timestamp).strftime('%Y-%m-%d %H:%M:%S')\n\nprint(fecha_legible)\n\n\n# -\n\ndef extract_unix_timestamp(item):\n fecha_ilegible= item[0] // 1000 # Dividir por 1000 para obtener 10 dígitos\n fecha_legible = datetime.utcfromtimestamp(fecha_ilegible).strftime('%Y-%m-%d')\n return fecha_legible\n\n\ndef remove_unix_timestamp(item):\n return item[1]\n\n\n# +\n#Está lista se ocupara para que todos los DataFrames tengan las mismas fechas.\nfecha_inicio = '2022-08-20'\nfecha_fin = '2023-08-19'\n\nrango_fechas = pd.date_range(start=fecha_inicio, end=fecha_fin, freq='D')\n\n# Crear un DataFrame con la columna de fechas\ndf_Fechas = pd.DataFrame({'fechas': rango_fechas})\n\n\n# -\n\ndef df_market(id:str): \n #Creación DataFrame\n gcmcrange=get_coin_market_chart_range_by_id(id,'usd','1660971600','1692424800')\n df=pd.DataFrame(gcmcrange)\n #Extracción Fechas:\n df['fechas'] = df['prices'].apply(extract_unix_timestamp)\n #Remove Unix Time\n for i in df:\n df[i+'2'] = df[i].apply(remove_unix_timestamp)\n #Formato DataTime para posterior merge:\n df['fechas'] = pd.to_datetime(df['fechas'])\n\n columnas_a_eliminar = ['prices', 'market_caps','total_volumes','fechas2']\n df=df.drop(columns= columnas_a_eliminar)\n\n df_union = pd.merge(df_Fechas, df, on='fechas', how='outer')\n\n #Agregando Columna con ID \n df_union.insert(0, 'Cripto', id)\n \n\n return df_union\n\n# #### Categoría: Layer 1 (L1)\n\ncriptos\n\ndf_Bitcoin_market=df_market('bitcoin')\ndf_Ethereum_market=df_market('ethereum')\ndf_BNB_market=df_market('binancecoin')\ndf_Cardano_market=df_market('cardano')\ndf_Solana_market=df_market('solana')\ndf_polkadot_market=df_market('polkadot')\ndf_bitcoin_cash_market=df_market('bitcoin-cash')\ndf_Avalanche_market=df_market('avalanche-2')\ndf_cosmos_market=df_market('cosmos')\ndf_hedera_hashgraph_market=df_market('hedera-hashgraph')\n\n# +\n# Lista de DataFrames a agregar\ndataframes = [df_Bitcoin_market, df_Ethereum_market,df_BNB_market,df_Cardano_market,\n df_Solana_market,df_polkadot_market,df_bitcoin_cash_market,df_Avalanche_market\n ,df_cosmos_market,df_hedera_hashgraph_market]\n\ndf_concatenated = pd.concat(dataframes, ignore_index=False)\n# -\n\ndf_concatenated.shape\n\n\n# ### Extracción de precios de apertura, mayor precio, menor y cierrre.\n\ndef Candles_Cripto(id):\n\n df = pd.DataFrame(client.get_historical_klines(id,'1d','1660971600','1692424800'))\n df = df.iloc[:,:5]\n df.columns = ['fechas','Open','High','Low','Close',]\n df['fechas'] = pd.to_datetime(df.fechas,unit='ms')\n df = df.set_index('fechas')\n df = df.astype(float)\n return df\n\n\nSymbol=(criptos['symbol'].tolist())\n\nSymbol\nfor i in range(len(Symbol)):\n Symbol[i] = Symbol[i] + 'usdt'\nSymbol = [item.upper() for item in Symbol]\n\ncriptos\n\nSymbol\n\ndf_Candles_Bitcoin=Candles_Cripto('BTCUSDT')\ndf_Candles_Ethereum=Candles_Cripto('ETHUSDT')\ndf_Candles_BNB=Candles_Cripto('BNBUSDT')\ndf_Candles_Cardano=Candles_Cripto('ADAUSDT')\ndf_Candles_Solana=Candles_Cripto('SOLUSDT')\ndf_Candles_polkadot=Candles_Cripto('DOTUSDT')\ndf_Candles_bitcoin_cash=Candles_Cripto('BCHUSDT')\ndf_Candles_Avalanche=Candles_Cripto('AVAXUSDT')\ndf_Candles_cosmos=Candles_Cripto('ATOMUSDT')\ndf_Candles_hedera_hashgraph=Candles_Cripto('HBARUSDT')\n\n\ndef merge_data(df1,df2):\n df1.set_index('fechas', inplace=True)\n df=df1.merge(df2,left_index=True,right_index=True)\n return df\n\n#merged_df = df_Bitcoin_market.merge(df_Candles_Bitcoin, left_index=True, right_index=True)\n\n\ndf_Bitcoin=merge_data(df_Bitcoin_market,df_Candles_Bitcoin)\n\n\ndf_Ethereum=merge_data(df_Ethereum_market,df_Candles_Ethereum)\ndf_BNB=merge_data(df_BNB_market,df_Candles_BNB)\ndf_Cardano=merge_data(df_Cardano_market,df_Candles_Cardano)\ndf_Solana=merge_data(df_Solana_market,df_Candles_Solana)\ndf_Polkadot=merge_data(df_polkadot_market,df_Candles_polkadot)\ndf_Avalanche=merge_data(df_Avalanche_market,df_Candles_Avalanche)\ndf_Cosmos=merge_data(df_cosmos_market,df_Candles_cosmos)\ndf_Hedera=merge_data(df_hedera_hashgraph_market,df_Candles_hedera_hashgraph)\ndf_Bitcoin_Cash=merge_data(df_bitcoin_cash_market,df_Candles_bitcoin_cash)\n\n\ndf_Bitcoin_Cash.shape\n\n# +\nLayer_1=[df_Bitcoin,df_Ethereum,df_BNB,df_Cardano,df_Solana,df_Polkadot,df_Avalanche\n ,df_Cosmos,df_Hedera,df_Bitcoin_Cash]\n\nLayer_1=pd.concat(Layer_1,ignore_index=False)\n# -\n\n#Asignar Ranking de acuerdo al MarketCap\nlatest_data = Layer_1.groupby('Cripto').apply(lambda group: group[group.index == group.index.max()])\nlatest_data = latest_data.sort_values(by='market_caps2', ascending=False)\nlatest_data['Ranking'] = range(1, len(latest_data) + 1)\nLayer_1['Ranking'] = Layer_1['Cripto'].map(latest_data.set_index('Cripto')['Ranking'])\n\n# Agregar Body Ratio\n\n# +\n# Calcular el Body Ratio\nLayer_1['Body_Ratio'] = abs(Layer_1['Close'] - Layer_1['Open']) / (Layer_1['High'] - Layer_1['Low'])\n\n# Clasificar las velas en tres categorías basadas en el Body Ratio y dirección de cierre\ndef classify_vela(row):\n if row['Body_Ratio'] < 0.3:\n return 'Indecisión'\n elif row['Body_Ratio'] >= 0.3 and row['Body_Ratio'] <= 0.7:\n return 'Fuerte Volatilidad'\n else:\n if row['Close'] > row['Open']:\n return 'Fuerte Dirección Alcista'\n else:\n return 'Fuerte Dirección Bajista'\n\nLayer_1['Clasificación'] = Layer_1.apply(classify_vela, axis=1)\n# -\n\n# Agregar Retorno Diario\n\nLayer_1['Daily_Return'] = Layer_1.groupby('Cripto')['Close'].pct_change()\n\nLayer_1['Daily_Return'].fillna(0, inplace=True)\n\n# +\n#Layer_1.to_csv('DataCriptos.csv',index=True)\n# -\n\n# Obtención de datos del Top 3 compañias en el sector de Tecnología\n#\n# Fuente Yahoo.\n\n# AAPL\tApple Inc.\n#\n# MSFT\tMicrosoft Corporation\n\nstart_date = \"2022-08-20\"\nend_date = \"2023-08-19\"\ncolumns=['Dividends','Stock Splits']\n\nAAPL = yf.Ticker(\"AAPL\")\ndf_AAPL = AAPL.history(period=\"1d\", start=start_date, end=end_date)\ndf_AAPL = AAPL.history(period=\"1d\", start=start_date, end=end_date)\ndf_AAPL.index = df_AAPL.index.date\ndf_AAPL=df_AAPL.drop(columns=columns)\nstart_date = df_AAPL.index.min()\nend_date = df_AAPL.index.max()\ndate_range = pd.date_range(start=start_date, end=end_date)\ndf_AAPL = df_AAPL.reindex(date_range)\ndf_AAPL.insert(0, 'Company', 'Apple Inc.')\ndf_AAPL=df_AAPL.interpolate()\n\nMSFT = yf.Ticker(\"MSFT\")\ndf_MSFT = MSFT.history(period=\"1d\", start=start_date, end=end_date)\ndf_MSFT = MSFT.history(period=\"1d\", start=start_date, end=end_date)\ndf_MSFT.index = df_MSFT.index.date\ndf_MSFT=df_MSFT.drop(columns=columns)\nstart_date = df_MSFT.index.min()\nend_date = df_MSFT.index.max()\ndate_range = pd.date_range(start=start_date, end=end_date)\ndf_MSFT = df_MSFT.reindex(date_range)\ndf_MSFT.insert(0, 'Company', 'Microsoft Corporation')\ndf_MSFT=df_MSFT.interpolate()\n\n\n# +\nDataTec=[df_AAPL,df_MSFT]\n\nDataTec=pd.concat(DataTec,ignore_index=False)\n\n# +\n# Calcular el Body Ratio\nDataTec['Body_Ratio'] = abs(DataTec['Close'] - DataTec['Open']) / (DataTec['High'] - DataTec['Low'])\n\nDataTec['Clasificación'] = DataTec.apply(classify_vela, axis=1)\n# -\n\n# Retorno Diario\n\nDataTec['Daily_Return'] = DataTec.groupby('Company')['Close'].pct_change()\n\nDataTec['Daily_Return'].fillna(0, inplace=True)\n\n# +\n#DataTec.to_csv('DataTec.csv',index=True)\n# -\n\nLayer_1['Cripto'].unique()\n\nDataTec[DataTec.Company=='Apple Inc.'].info()\n\nDataTec.info()\n","repo_name":"Benjaminmxxx/DataAnalytics","sub_path":"ETL.ipynb","file_name":"ETL.ipynb","file_ext":"py","file_size_in_byte":9255,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"18703646438","text":"# +\n#tweetlerin veritabanından alınması\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport nltk\nimport zemberek\nimport jpype,os\nfrom nltk.tokenize import word_tokenize\nfrom nltk.tokenize import TweetTokenizer\nfrom nltk.corpus import stopwords \ndurakwords= set(stopwords.words(\"turkish\"))\nimport string\nimport re\nfrom typing import List\nfrom jpype import JClass, JString, getDefaultJVMPath, shutdownJVM, startJVM, java\nfrom scipy import stats\nimport seaborn as sns\nfrom collections import Counter\nfrom keras.models import Sequential\nfrom keras.layers import Embedding,SpatialDropout1D,LSTM,Dense\nfrom keras.callbacks import ModelCheckpoint, EarlyStopping\n# %matplotlib inline\ndf=pd.DataFrame()\ndf=pd.read_csv(\"C:/Users/user/tweets/tweets2.csv\")\n\n\n# -\n\n#Tweetlerin küçük harfe dönüştürülmesi\ndf= df.rename(columns = {'hash,content,class': 'tweets'})\ndf['tweets']=df['tweets'].str.lower()\n\n#tweetlerin son değerlerine göre yeni bir dataframe e yerleştirilmesi\n#1 tek,2 spor,eko,pol,mag,sağ,kült\n#df[df.tweets.str.endswith('2')]\ndf['icerik']='icerik'\ndf['cleantext']='cleantext'\ndf.loc[df['tweets'].str.endswith('1'),'icerik']=\"teknoloji\"\ndf.loc[df['tweets'].str.endswith('2'),'icerik']=\"spor\"\ndf.loc[df['tweets'].str.endswith('3'),'icerik']=\"ekonomi\"\ndf.loc[df['tweets'].str.endswith('4'),'icerik']=\"politika\"\ndf.loc[df['tweets'].str.endswith('5'),'icerik']=\"magazin\"\ndf.loc[df['tweets'].str.endswith('8'),'icerik']=\"kultur\"\ndf.loc[df['tweets'].str.endswith('7'),'icerik']=\"saglik\"\ndf.loc[df['tweets'].str.endswith('1'),'label']=\"1\"\ndf.loc[df['tweets'].str.endswith('2'),'label']=\"2\"\ndf.loc[df['tweets'].str.endswith('3'),'label']=\"3\"\ndf.loc[df['tweets'].str.endswith('4'),'label']=\"4\"\ndf.loc[df['tweets'].str.endswith('5'),'label']=\"5\"\ndf.loc[df['tweets'].str.endswith('8'),'label']=\"8\"\ndf.loc[df['tweets'].str.endswith('7'),'label']=\"7\"\npd.to_numeric(df.label)\n\n#Zemberek kütüphanesinin kullanılabilmesi için Jpype ile JVM oluşturulması\nZEMBEREK_PATH = r'C:\\Users\\user\\Desktop\\dersler\\tweets\\zemberek-full.jar'\nstartJVM(getDefaultJVMPath(), '-ea', '-Djava.class.path=%s' % (ZEMBEREK_PATH))\n#Morphology kütüphanesini kullanacağımızdan burada oluşturuyoruz\nTurkishMorphology = JClass('zemberek.morphology.TurkishMorphology')\nmorphology = TurkishMorphology.createWithDefaults()\n\n\ndef mentionRemove(text):\n text = re.sub(r'@\\w+', '', text)#mentionların çıkartırılması\n return text\n\n\ndef siteRemove(text):\n text = re.sub(r'http.?://[^\\s]+[\\s]?', '', text)#bağlantıların çıkarılması\n return text\n\n\ndef puncRemove(text):\n text = re.sub('[^a-zığüşiöç\\s]', '', text)#özel karakterlerin ve sayıların çıkarılması\n text = re.sub(r\"Â\", \"A\", text)#umlautlu karakterlerinn değiştirilmesi\n text = re.sub(r\"â\", \"a\", text)\n text = re.sub(r\"Î\", \"I\", text)\n text = re.sub(r\"î\", \"ı\", text)\n text = re.sub(r\"Û\", \"U\", text)\n text = re.sub(r\"û\", \"u\", text)\n\n return text\n\n\n\ndef removeSw(text):\n text=[w for w in tweet if w.lower() not in durakwords]#stopwordlerin çıkarılması\n return text\n\n\ndef rootFinder(text):\n pos=[]\n textlist=text.split(\" \")\n analysis: java.util.ArrayList = ( morphology.analyzeAndDisambiguate(text).bestAnalysis() )\n for i, analysis in enumerate(analysis, start=1):\n f'\\nAnalysis {i}: {analysis}',\n f'\\nPrimary POS {i}: {analysis.getPos()}' \n if (str(analysis.getLemmas()[0]) == \"UNK\"):\n pos.append(textlist[i-1])\n else: \n pos.append(f'{str(analysis.getLemmas()[0])}')\n return pos\n\n\n\ndef emojiRemove(tweet):\n # Smile -- :), : ), :-), (:, ( :, (-:, :')\n tweet = re.sub(r'(:\\s?\\)|:-\\)|\\(\\s?:|\\(-:|:\\'\\))', ' EMO_POS ', tweet)\n # Laugh -- :D, : D, :-D, xD, x-D, XD, X-D\n tweet = re.sub(r'(:\\s?D|:-D|x-?D|X-?D)', ' EMO_POS ', tweet)\n # Love -- <3, :*\n tweet = re.sub(r'(<3|:\\*)', ' EMO_POS ', tweet)\n # Wink -- ;-), ;), ;-D, ;D, (;, (-;\n tweet = re.sub(r'(;-?\\)|;-?D|\\(-?;)', ' EMO_POS ', tweet)\n # Sad -- :-(, : (, :(, ):, )-:\n tweet = re.sub(r'(:\\s?\\(|:-\\(|\\)\\s?:|\\)-:)', ' EMO_NEG ', tweet)\n # Cry -- :,(, :'(, :\"(\n tweet = re.sub(r'(:,\\(|:\\'\\(|:\"\\()', ' EMO_NEG ', tweet)\n return tweet\n\n\n# +\n#hash kısmının ve sondaki içerik belirteci ve tweet bağlantısının kaldırılması\n#utf-16 karakterlerin elenmesi\n#kelimelerin ayrıştırılması\n#tweet içerisindeki bağlantıların çıkarılması\n#tweet içindeki mentionların çıkarılması\np=set(string.punctuation)\n\ndef cleaner(text):\n text=text\n text=mentionRemove(text)#mentionların çıkartırılması\n text=siteRemove(text)#bağlantıların çıkarılması\n text=puncRemove(text)#özel karakterlerin ve sayıların çıkarılması\n text=emojiRemove(text)\n text=rootFinder(text)#kelime köklerinin bulunması\n text=removeSw(text)#durak kelimelerin elenmesi\n return text\nfor x in range(0,len(df)):\n #tweet=\"ve ya da için baba veya\"\n tweet=df.iloc[x,0]\n tweet=mentionRemove(tweet)#mentionların çıkartırılması\n tweet=siteRemove(tweet)#bağlantıların çıkarılması\n #tweet.replace(\"aracılığıyla\",\"a\")\n tweet=puncRemove(tweet)#özel karakterlerin ve sayıların çıkarılması\n tweet=rootFinder(tweet)#kelime köklerinin bulunması\n tweet=removeSw(tweet)#durak kelimelerin elenmesi\n tweet=' '.join(tweet)\n #tweet=cleaner(tweet)\n df.iloc[x,2]=tweet\n pos=[]\n\n\n# -\n\nimport matplotlib.pyplot as plt\nimport os\nimport re\nimport shutil\nimport string\nimport tensorflow as tf\nfrom tensorflow.keras import regularizers\nfrom tensorflow.keras import layers\nfrom tensorflow.keras import losses\nfrom collections import Counter\nfrom sklearn.preprocessing import LabelEncoder\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import classification_report\nfrom tensorflow.keras.preprocessing.text import Tokenizer\nfrom tensorflow.keras.preprocessing.sequence import pad_sequences\nfrom sklearn.metrics import accuracy_score,confusion_matrix\n\nprint(tf.__version__)\nif tf.test.gpu_device_name():\n print(\"gpu device :{}\".format(tf.test.gpu_device_name()))\nelse:\n print(\"please install gpu version of tf\")\n\n\ndef counter_words(df):\n count = Counter()\n for i in df:\n for w in i.split():\n count[w] += 1\n return count\n\n\nc=counter_words(df.cleantext)\nnum_words = len(c)\nmax_seq_lenght = 12\n\ntweets=df.cleantext\nlabels=df.icerik\n\n# +\ntokenizer=Tokenizer(num_words=num_words)\ntokenizer.fit_on_texts(tweets.tolist())\n\nprint(str(tokenizer.texts_to_sequences([\"fenerbahçe ben sevgili\"])))\n# -\n\nX_train, X_test, Y_train, Y_test = train_test_split(tweets.tolist(),labels.tolist(), test_size = 0.3,stratify=labels.tolist(),\n random_state = 44)\nprint('Train data len: '+ str(len(X_train)))\nprint('labels: ', Counter(Y_train))\nprint('test data len: '+ str(len(X_test)))\nprint('labels: ' ,Counter(Y_test))\n\nX_train=np.array(tokenizer.texts_to_sequences(X_train))\nX_test=np.array(tokenizer.texts_to_sequences(X_test))\n#Y_train=np.array(tokenizer.texts_to_sequences(Y_train))\nX_train=pad_sequences(X_train, padding = 'post',maxlen=max_seq_lenght,truncating=\"post\")\nX_test=pad_sequences(X_test, padding = 'post',maxlen=max_seq_lenght,truncating=\"post\")\n\nprint(X_train[1])\nprint(Y_train[:10])\n\nle = LabelEncoder()\ntrain_labels=le.fit_transform(Y_train)\nprint(train_labels)\ntrain_labels=np.asarray(tf.keras.utils.to_categorical(train_labels))\nprint(train_labels)\ntest_labels=le.fit_transform(Y_test)\ntest_labels=np.asarray(tf.keras.utils.to_categorical(test_labels))\nlist(le.classes_)\n\ntrain_ds=tf.data.Dataset.from_tensor_slices((X_train,train_labels))\ntest_ds=tf.data.Dataset.from_tensor_slices((X_test,test_labels))\n\ntrain_labels=np.asarray(tf.keras.utils.to_categorical(le.fit_transform(Y_train)))\ntrain_labels\n\nnum_words = len(c)\n# Max number of words in each complaint.\nmax_seq_lenght = 12\n# This is fixed.\ndim = 100\nprint(len(c))\nmodel = tf.keras.Sequential()\nmodel.add(tf.keras.layers.Embedding(num_words+1,dim,input_length=max_seq_lenght,embeddings_regularizer=regularizers.l2(0.0001)))\nmodel.add(tf.keras.layers.Conv1D(128,7,activation='relu',kernel_regularizer=regularizers.l2(0.0001),bias_regularizer=regularizers.l2(0.0001)))\nmodel.add(tf.keras.layers.GlobalMaxPooling1D())\nmodel.add(tf.keras.layers.Dropout(0.2))\nmodel.add(tf.keras.layers.Dense(7,activation='sigmoid',kernel_regularizer=regularizers.l2(0.001),bias_regularizer=regularizers.l2(0.001)))\nmodel.summary()\n\nmodel.compile(loss=tf.keras.losses.CategoricalCrossentropy(),optimizer='Adam',metrics=[\"CategoricalAccuracy\"])\n\nepochs=10\nhistory=model.fit(train_ds.batch(128),epochs=epochs,validation_data=test_ds.batch(128),verbose=1)\n\nnew_complaint = ['erdoğan hastane millet millet oy']\nseq = tokenizer.texts_to_sequences(new_complaint)\npadded = pad_sequences(seq, maxlen=max_seq_lenght)\npred = model.predict(padded)\nlabels = ['ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji']\nprint(pred, labels[np.argmax(pred)])\n\npreds=model.predict(X_test)\npreds=preds.argmax(axis=1)\nactual=le.fit_transform(Y_test)\nprint(\"accuracy score is\",accuracy_score(preds,actual))\n\nprint(classification_report(actual, preds,target_names=['ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji']))\n\nconf_mat = confusion_matrix(actual,preds)\nfig, ax = plt.subplots(figsize=(15,15))\nsns.heatmap(conf_mat, annot=True, cmap=\"Blues\", fmt='d',\n xticklabels={'ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji'},\n yticklabels={'ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji'})\nplt.ylabel('Actual')\nplt.xlabel('Predicted')\nplt.title(\"CONFUSION MATRIX\\n\", size=16);\n\npred=model.predict()\n\n\n","repo_name":"alprbykn/tweet_classification","sub_path":"tweets_neuralNetworks.ipynb","file_name":"tweets_neuralNetworks.ipynb","file_ext":"py","file_size_in_byte":9783,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"71874153753","text":"# ## Taylor Sin(x) Serisi by OVO Analytics\n# Made by Eren BOZARIK\n\nimport numpy as np\nimport time\n\n\ndef taylor_sin(x):\n start_time = time.time()\n x1=x*(np.pi/180)\n val=0\n for n in range(0, 25):\n add=(((-1) ** n) * (x1 ** (2*n+1)) / np.math.factorial(2*n + 1))\n val+=add\n end_time = time.time()\n print(\"{:.2f} surede hesaplandi\" .format(float(end_time - start_time)))\n return val\n\n\na=int(input(\"Hesaplanacak Dereceyi Giriniz :\"))\nprint(\"Sin({}) = {:.2f}\".format(a,taylor_sin(a)))\n","repo_name":"erenbozarik/Statistical-Scripts","sub_path":"Taylor_Series.ipynb","file_name":"Taylor_Series.ipynb","file_ext":"py","file_size_in_byte":517,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"70983907353","text":"# # Project: Writing a Data Scientist Blog Post\n\n# ## Introduction and Question\n\n# - Ford GoBike System Data: This data set includes information about individual rides made in a bike-sharing system covering the greater San Francisco Bay area.\n#\n# - The data including information of 183.412 ford gobike trips in San Francisco area with 16 variables (duration_sec, start_time, end_time, start_station_id, start_station_name, start_station_latitude, start_station_longitude, end_station_id, end_station_name, end_station_latitude ,end_station_longitude, bike_id, user_type, member_birth_year, member_gender, bike_share_for_all_trip). But some data have wrong so it needs to be clean them.\n#\n# - Question: How location distribution, user type, user age and also gender afftect to the trip?\n\n# ## Data Cleaning and Accessing\n\n# ### Data Cleaning\n\n# +\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nimport warnings\n\n# %matplotlib inline\nwarnings.simplefilter(\"ignore\")\n# -\n\nford_df = pd.read_csv(\"201902-fordgobike-tripdata.csv\")\nford_df.head()\n\nford_df.info()\n\nford_df.columns\n\n# #### Some columns have wrong datatype. So I need to convert them.\n#\n# 1. start_time and end_time shoud be datetime instend object\n#\n# 2. bike_share_for_all_trip should be converted to boolean instead\n\nford_df['start_time'] = ford_df['start_time'].astype('datetime64[ns]')\nford_df['end_time'] = ford_df['end_time'].astype('datetime64[ns]')\n\nford_df.bike_share_for_all_trip = ford_df.bike_share_for_all_trip.replace({\"Yes\": True, \"No\":False}).astype(bool)\n\nford_df.info()\n\n# ### Data Accessing\n\n# #### What are percentage of User Type and Member Gender?\n\nuser_type = ford_df['user_type'].value_counts()\nax = user_type.plot(kind='pie', autopct= '%1.1f%%',figsize=(7, 7));\n# set the title\nax.set_title(\"User Type\" , fontsize=20, fontweight='bold');\n\n# There's huge number of people that have subscribed.\n\nmember_gender = ford_df['member_gender'].value_counts()\nax = member_gender.plot(kind='pie', autopct= '%1.1f%%',figsize=(7, 7));\n# set the title\nax.set_title(\"Member Gender\" , fontsize=20, fontweight='bold');\n\n\n# Member gender male is more than female. Female is more than Other\n\n# #### Is there any relationship between duration second and user type?\n\ndef barplot(col, title, xlabel, ylabel, rotation):\n \"\"\"\n This function provide the counts for the each series in the column and plot it in bar plot.\n Input: column, title, xlabel, ylabel, rotation of xticks.\n \n Output: plt bar plot of value counts\n \"\"\"\n fig, ax = plt.subplots(figsize = (6,6), dpi = 120);\n col.value_counts().plot(kind='bar');\n plt.title(title, fontweight='bold');\n plt.ylabel(xlabel);\n plt.xlabel(ylabel);\n plt.xticks(rotation=rotation);\n\n\nbarplot(ford_df['user_type'], \"User Type by mean trip duration\", \"User Type\", \"Trip duration in second\", 0)\n\n# Although the number of Subscriber user is more than number of Customer, Customer trip duration is more than Subscriber\n\n# #### Is there any relationship between duration second, week day and user type?\n\nx = ford_df['start_time'].dt.day_name()\ny = 'duration_sec'\norders = [\"Monday\", \"Tuesday\",\"Wednesday\", \"Thursday\", \"Friday\",\"Saturday\",\"Sunday\"]\n\nfig, ax = plt.subplots(figsize = (12,5), dpi = 150);\nsns.boxplot(data=ford_df, x=x, y=y, order=orders,hue='user_type');\nplt.ylim([0,2500]);\nplt.title(\"Duration second by day of week\", fontweight='bold');\nplt.xlabel(\"Day of Week\");\nplt.ylabel(\"Duration second\");\nplt.legend(loc=1);\n\n# It seems that more trip in weekend than day week for Customer and for Subscriber it the same for all week.\n\n# #### What is top 10 Start and End Station have most rider?\n\naverage_line = ford_df['bike_id'].count() / ford_df.groupby(['start_station_name'])['start_station_name'].count()\naverage_line = average_line.nunique()\n\n# +\n# Separate the genre based on the pipe (|).\nford_df['start_station_name'] = ford_df['start_station_name'].astype(str)\nstationList = ford_df['start_station_name'].str.split('|')\n\n# Create list of station\nstation = []\nfor i in stationList:\n station.extend(i)\n \n# Plot based on the value counts\nplt.subplots(figsize=(12,10));\npd.Series(station).value_counts()[:10].sort_values(ascending=False).plot.barh(width=0.9,color=sns.color_palette('coolwarm',10));\nplt.title('Top 10 Start Station have most rider and Average line', fontweight='bold');\nplt.xlabel('Count');\nplt.ylabel('Station Name');\nplt.vlines(x=average_line, ymin=-2, ymax=20, color='r');\nplt.show();\n# -\n\n# Based on the chart above, it demonstrates that Market St at 10th St has most rider when they start riding\n\n# +\n# Separate the genre based on the pipe (|).\nford_df['end_station_name'] = ford_df['end_station_name'].astype(str)\nstationList = ford_df['end_station_name'].str.split('|')\n\n# Create list of station\nstation = []\nfor i in stationList:\n station.extend(i)\n \n# Plot based on the value counts\nplt.subplots(figsize=(12,8));\npd.Series(station).value_counts()[:10].sort_values(ascending=False).plot.bar(width=0.9,color=sns.color_palette('coolwarm',10));\nplt.title('Top 10 End Station have most rider', fontweight='bold');\nplt.xlabel('Count');\nplt.ylabel('Station Name');\nplt.show();\n# -\n\n# Based on the chart above, it demonstrates that Market St at 10th St has most rider when they start riding. But the end station is not this one. It's a San Francisco Caltrain Staion 2 because maybe around this station have a lot the beautiful place or famous restaurant,...\n\n# ## Conclusions\n\n# In conclusion, Subcribers have a huge ratio, but customers spend more time on riding than another. As a result, we need to have a strategy to convert them to our customers. Besides, Almost riders usually ride bicycle at weekend. That is great if we orginize some gameshow, marketing programs at that time to attract more customers.\n","repo_name":"tranhoang030998/Writing-a-Data-Scientist-Blog-Post","sub_path":"Udacity_BlogPost.ipynb","file_name":"Udacity_BlogPost.ipynb","file_ext":"py","file_size_in_byte":5848,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"10957102887","text":"# # # Kata 8\n#\n# Módulo 8 - Administrar Datos con Diccionarios\n#\n# Ejercicio 1\n\nplanet = {\n 'name' : 'Mars',\n 'moons' : 2\n}\n\nprint(f'{planet[\"name\"]} has {planet[\"moons\"]} moons.')\n\nplanet['circunference (km)'] = {\n 'polar': 6752,\n 'equatorial': 6792\n}\n\nprint(f'{planet[\"name\"]} polar circunference of: {planet[\"circunference (km)\"][\"polar\"]} km. And a equatorial circunference of: {planet[\"circunference (km)\"][\"equatorial\"]} km.')\n\n\n# Ejercicio 2\n\nplanet_moons = {\n 'mercury': 0,\n 'venus': 0,\n 'earth': 1,\n 'mars': 2,\n 'jupiter': 79,\n 'saturn': 82,\n 'uranus': 27,\n 'neptune': 14,\n 'pluto': 5,\n 'haumea': 2,\n 'makemake': 1,\n 'eris': 1\n}\n\n# +\ntotal_moons = 0\n\nfor value in planet_moons.values():\n total_moons = total_moons + value\nprint(f'There is a total of {total_moons} in the solar system.')\nprint(f'That is an average of {total_moons/len(planet_moons.keys())}')\n","repo_name":"ErickViveros/KatasLaunchX","sub_path":"Kata_08.ipynb","file_name":"Kata_08.ipynb","file_ext":"py","file_size_in_byte":919,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"4007700494","text":"# # Stage 4: Advanced Analysis\n# - Sentiment Analysis\n# - Text Classification\n\n# ### Importing Libraries\n\n# +\nimport pandas as pd\nfrom tqdm import tqdm\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\nfrom transformers import pipeline\n# -\n\n# ### Files Import\n\n# +\ndata = pd.read_pickle('data_clean_summarized.pkl')\ndata.reset_index(drop=True, inplace=True)\n\ndata\n\n# +\n# from Amazon website (https://www.amazon.com/AmazonBasics-Microwave-Small-Works-Alexa/dp/B07894S727)\nspecs = pd.read_csv(\"specs.csv\")\nspecifications = list(specs[\"Specifications\"]) # extract specifications headers from specs.csv\n\n# from Mr Right Ideas website (https://www.mrright.in/ideas/appliances/microwave/microwave-oven-components-and-their-functions/)\ncomps = pd.read_csv(\"comps.csv\")\ncomponents = list(dict.fromkeys(comps[\"Components\"])) # extract component headers from comps.csv\ncomps_renamed = list(dict.fromkeys(comps[\"Comps_renamed\"])) # extract renamed components from comps.csv\n\n# from wiki_scraper.py\nwiki_adjusted = pd.read_csv(\"wiki_adjusted.csv\")\nwiki_adjusted = list(wiki_adjusted['Keywords']) # extract sustainability keywords from wiki_adjusted.csv\n\nothers = ['Packaging', 'Delivery']\nspecifications += others\n\ndata['comps_mention'] = 'na'\n# -\n\n# ### Components Mentioned in Reviews\n# - With Sentiment Analysis\n\n# +\nfor i in range(data.shape[0]):\n for j in comps_renamed:\n if j in data['text'][i]:\n data['comps_mention'][i] = j\n\ndata\n\n# +\n# Create quick lambda functions to find the polarity and subjectivity of each routine\nfrom textblob import TextBlob\n\ncount_result = []\nfor comps in comps_renamed:\n count = [data[data.comps_mention == comps].shape[0]]\n count_result += count\n\npol = lambda x: TextBlob(x).sentiment.polarity\ndata['polarity'] = data['summary'].apply(pol)\n\npol_count = lambda x: 1 if x > 0 else -1\ndata['pol_count'] = data['polarity'].apply(pol_count)\ndisplay(data)\n\nsentiment_list = []\nfor comps in comps_renamed:\n data_count = data[data.comps_mention == comps]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': comps_renamed,\n 'values': count_result,\n 'sentiments': sentiment_list}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n\n# +\n# Data check\n\ndata_selected = data[data.comps_mention == df_chart.labels[2]] # check number accordingly\ndata_selected.reset_index(drop=True, inplace=True)\n\ndata_selected\n# -\n\n# ### Sustainability Mentioned in Reviews\n# - With Sentiment Analysis\n\n# +\ndata['sus_mention'] = 'na'\nwiki_adjusted = ['waste', 'eneryg', 'environmental', 'material']\nfor i in range(data.shape[0]):\n for j in wiki_adjusted:\n if j in data['text'][i]:\n data['sus_mention'][i] = j\n\ndata\n\n# +\n# Create quick lambda functions to find the polarity and subjectivity of each routine\nfrom textblob import TextBlob\n\ncount_result = []\nfor sus in wiki_adjusted:\n count = [data[data.sus_mention == sus].shape[0]]\n count_result += count\n\npol = lambda x: TextBlob(x).sentiment.polarity\ndata['polarity'] = data['summary'].apply(pol)\n\npol_count = lambda x: 1 if x > 0 else -1\ndata['pol_count'] = data['polarity'].apply(pol_count)\ndisplay(data)\n\nsentiment_list = []\nfor sus in wiki_adjusted:\n data_count = data[data.sus_mention == sus]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': wiki_adjusted,\n 'values': count_result,\n 'sentiments': sentiment_list}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n\n# +\n# Data check\n\ndata_selected = data[data.comps_mention == df_chart.labels[2]] # check number accordingly\ndata_selected.reset_index(drop=True, inplace=True)\n\ndata_selected\n# -\n\n# ## Zero-Shot-Classification\n\nclassifier = pipeline(\"zero-shot-classification\", device=0)\n\n# ### Specification Categorization\n# - With Sentiment Analysis \n\n# +\ndisplay(specifications)\n#Applying it into the dataframe base on original review\ndata['category'] = data['text'].apply(lambda x: classifier(x, candidate_labels=specifications))\n\ndata\n# -\n\n#Replacing the category with the top matched category\nfor i in tqdm(range(data.shape[0])):\n if data['category'][i] not in specifications:\n data['category'][i] = data['category'][i].get('labels')[0]\n\ndata_cat = data.copy()\n\n# +\ncandidate_results = list(0 for i in range(len(specifications)))\n\n\ndef count(category):\n for i in range(len(specifications)):\n if category == specifications[i]:\n candidate_results[i] = candidate_results[i] + 1\n\n\n# +\n# Run the counting function\ndata_cat['category'].apply(lambda x: count(x))\n\ncandidate_results\n\n\n# +\nsentiment_list_func = []\nfor func in specifications:\n data_count = data[data.category == func]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list_func += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': specifications,\n 'values': candidate_results,\n 'sentiments': sentiment_list_func}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n# -\n\n# ### Sustainability Categorization \n# - Sentiment Analysis included\n\n# +\n#Categorying in terms of sustainability\n\n#Applying it into the dataframe base on original review\ndata['sus_category'] = data['text'].apply(lambda x: classifier(x, candidate_labels=wiki_adjusted))\n\ndata\n# -\n\n#Replacing the category with the top matched category\nfor i in tqdm(range(data.shape[0])):\n if data['sus_category'][i] not in wiki_adjusted:\n data['sus_category'][i] = data['sus_category'][i].get('labels')[0]\n\n# +\ndata_cat = data.copy()\n\ncandidate_results = list(0 for i in range(len(wiki_adjusted)))\n\n\ndef count(category):\n for i in range(len(wiki_adjusted)):\n if category == wiki_adjusted[i]:\n candidate_results[i] = candidate_results[i] + 1\n\n\n# +\n# Run the counting function\ndata_cat['sus_category'].apply(lambda x: count(x))\n\ncandidate_results\n\n\n# +\nsentiment_list_func = []\nfor func in wiki_adjusted:\n data_count = data[data.category == func]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list_func += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': wiki_adjusted,\n 'values': candidate_results,\n 'sentiments': sentiment_list_func}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n\n# +\ndata.to_pickle('data_analysed.pkl')\n\ndata\n# -\n\n\n","repo_name":"jiasenn/text-analysis","sub_path":"4.Advanced-Analysis.ipynb","file_name":"4.Advanced-Analysis.ipynb","file_ext":"py","file_size_in_byte":7104,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"32876589411","text":"# \n\n# # Final Project \n\n# ## Задание 1. A/B–тестирование\n\n# Одной из основных задач аналитика в нашей команде является корректное проведение экспериментов. Для этого мы применяем метод A/B–тестирования. В ходе тестирования одной гипотезы целевой группе была предложена новая механика оплаты услуг на сайте, у контрольной группы оставалась базовая механика. В качестве задания Вам необходимо проанализировать итоги эксперимента и сделать вывод, стоит ли запускать новую механику оплаты на всех пользователей.\n#\n# Предлагаем Вам ответить на следующие вопросы:\n#\n# - На какие метрики Вы смотрите в ходе анализа и почему?\n# - Имеются ли различия в показателях и с чем они могут быть связаны?\n# - Являются ли эти различия статистически значимыми?\n# - Стоит ли запускать новую механику на всех пользователей?\n#\n\n# ## В качестве входных данных Вы имеете 4 csv-файла:\n#\n# - groups.csv - файл с информацией о принадлежности пользователя к контрольной или экспериментальной группе (А – контроль, B – целевая группа) \n# - groups_add.csv - дополнительный файл с пользователями, который вам прислали спустя 2 дня после передачи данных\n# - active_studs.csv - файл с информацией о пользователях, которые зашли на платформу в дни проведения эксперимента. \n# - checks.csv - файл с информацией об оплатах пользователей в дни проведения эксперимента. \n\n# ____\n\n# +\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nfrom scipy.stats import chi2_contingency, ttest_ind, levene, normaltest, mannwhitneyu, norm\nfrom tqdm.auto import tqdm\nimport pandahouse as ph\n\nsns.set(\n font_scale=1.5,\n style=\"whitegrid\",\n rc={'figure.figsize':(16,8)}\n )\n# -\n\n# #### Загрузка и первичный анализ данных:\n\ngroups = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_groups.csv', sep=';')\ngroups.head()\n\nprint(f'Кол-во строк: {groups.shape[0]}')\nprint(f'Кол-во дубликатов: {groups.duplicated().sum()}')\nprint(f'Кол-во уникальных id: {groups.id.nunique()}')\n\ngroups.isna().sum() # проверка на пропущенные значения \n\ngroups_add = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_group_add.csv', sep=',')\ngroups_add.head()\n\nprint(f'Кол-во строк: {groups_add.shape[0]}')\nprint(f'Кол-во дубликатов: {groups_add.duplicated().sum()}')\nprint(f'Кол-во уникальных id: {groups_add.id.nunique()}')\n\n# Объединим данные в один фрейм:\n\ngroups_total = pd.concat([groups,groups_add], sort=False,ignore_index=True)\ngroups_total\n\ngroups_total.duplicated().sum() # прверяем что не появилось дублей \n\n# поменяем назваие стольбца для удобства дальнейшего объединения таблиц\ngroups_total.rename(columns={'id':'student_id'}, inplace=True) \ngroups_total.head()\n\n# Посмотрим на распределение пользователей по группам:\n\ngroups_total.grp.value_counts()\n\nsns.countplot(data=groups_total, x='grp');\n\n# Распределение оказалось крайне не равномерным, что само по себе не является чем-то неправильным или критичным для A/B теста.\n# Дело в том, что обычно тестовая группа бывает меньше, в виду того, что катить новую фичу на большое кол-во пользователей рисковано и дорого, а в наших данных тестовая группа более чем в 3 раза больше.\n\n# ____\n\n# пользователи зашедшие на платфому в дни проведения эксперимента\nactive_studs = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_active_studs.csv')\nactive_studs.head()\n\nprint(f'Кол-во дубликатов:{active_studs.duplicated().sum()}')\n\nprint(f'Кол-во пользователей зашедших на платформу во время эксперимента:{active_studs.shape[0]}')\n\n# Кажется логичным добавить колонку, которая будет как-то выделять пользователей которые заходили на платформу/сайт во время эксперимента, при дальнейшем объединении таблиц.Так и поступим. И судя по колнке student_id мы имеем дело с образовательной платформой.\n\nactive_studs['active'] = 1\nactive_studs.head()\n\n# данные об оплатах в дни проведеня эксперимента\nchecks = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_checks.csv',sep=';')\nchecks.head()\n\nchecks.isna().sum() # проверка на пропущенные значения\n\nprint(f'Кол-во строк: {checks.shape[0]}')\nprint(f'Кол-во дубликатов: {checks.duplicated().sum()}')\nprint(f'Кол-во уникальных id: {checks.student_id.nunique()}')\n\nchecks.rev.describe()# посмортим немного описательной статистки по платежам \n\n# - Максимальный платеж - 4650\n# - Минимальный платеж - 199\n# - Медианный - 840 \n\nchecks.rev.value_counts().head(10) # распределение по платежам \n\nchecks.rev.plot(kind='hist', bins=25);\n\n# Распределение платежей не равномерное, имеется 3 моды(платежи по 1900, 290, 690).\n# Но это без привязки к группам, а нас интересуют различия как раз между группми.\n\n# ____\n\n# Предварительно можно сделать следующие выводы:\n# - данные очень \"чистые\", без пропусков и нужного типа\n# - таблица с активными пользователми насчитывает 8341 записи, тогда как общая таблица 74576 записи.\n# Исходя из условия задания, если пользователь не заходил на платформу/сайт во время эксперимента, то и увидеть/ощутить \n# изменения механики оплат на себе он не мог. Из этого следует, что анализировать и сранивать нужно только активн��х пользователей.\n\n# Соединим наши таблицы в одну.\n# Кажется логичным соединять таблицу активных пользователей с общей с помощью inner джойна, чтобы остались толкько те, кто нам нужен, но сделаем все обьединения через left чтобы пока что не потерять пользоватлей и покрутить данные еще, отфильтровать нужное всегда успеем.\n\n# делаем мердж и заполняем пропуски 0, так у нас 0 помечены те, кто не заходил во время эксперимента\nfinal_df = groups_total.merge(active_studs, how='left', on = 'student_id').fillna(0)\nfinal_df.head()\n\n# ____\n\n# Теперь присоединим таблицу с оплатами, но перед этим тоже создадим столбец для того чтобы выделить в общем фрейме тех, кто платил.\n\nchecks['has_rev'] = 1\nchecks.head()\n\nfinal_df = final_df.merge(checks, how='left', on='student_id').fillna(0)\nfinal_df.head()\n\n# Покрутим/проанализируем получившийся датафрейм\n\nprint(f'Число активных пользователей: {final_df.query(\"active == 1\").shape[0]}')\nprint(f'Число активных и сделавших покупку пользователей: {final_df.query(\"active == 1 & has_rev == 1\").shape[0]}')\nprint(f'Число активных и не сделавших покупку пользователей: {final_df.query(\"active == 1 & has_rev == 0\").shape[0]}')\nprint(f'Число неактивных и сделавших покупку пользователей: {final_df.query(\"active == 0 & has_rev == 1\").shape[0]}')\nprint(f'Число неактивных и не сделавших покупку пользователей: {final_df.query(\"active == 0 & has_rev == 0\").shape[0]}')\n\n# ____\n\n# Стоить прокомментировать 149 пользователей заплативших не заходя на сайт/платформу:\n# - можно проедположить что имеют место автоплатежи\n# - ошибка системы сплитования \n# - ошибки в логфайлах \n#\n# Утверждать с уверенностью, не имея дополнительных данных о том, как долго продолжался А/В тест,данные о времени захода пользователя и.т.д, мы не можем.\n# Следуя условию задачи, далее будем продаолжать анализ только с теми пользователми на которых проводился тест.\n\nexp_data = final_df.query(\"active == 1\")\nexp_data.head()\n\nexp_data.grp.value_counts()\n\n# Cреди активных пользователей так же наблюдаем дисбаланс по группам.\n\n# Посмотрим на распределение платежей:\n\nsns.histplot(data = exp_data, x = 'rev', hue = 'grp', bins= 30);\n\n# График оказался не очень информативным в силу того, что подавляющее большинство пользователей заходивших во время эксперимента не сделало покупку.\n#\n# Построим распредление трат только тех пользователей чей revenue > 0:\n\nsns.histplot(data = exp_data.query('rev > 0'), x = 'rev', hue = 'grp', bins= 25);\n\n# Глядя на график можно заметить, что наши распределения отличаются только пиком в группе В в районе rev = ~1900.\n#\n# Проверим цифрами:\n\nexp_data.query(\"rev > 0\")\\\n .groupby(['grp','rev'],as_index=False)\\\n .agg({'student_id':'count'})\\\n .rename(columns={'student_id':'num_checks'})\\\n .pivot(columns='grp', values='num_checks', index='rev')\\\n .sort_values(by='B', ascending=False)\\\n .fillna(0)\\\n .head(10)\n\n# В группе В 128 пользователей совершили платеж на 1900, тогда как в группе А таких не оказалось вовсе.\n\n# _____\n\n# #### Так как мы тестировали влияние новой механики оплаты, и исходя из имеющихся данных, нашими метриками будут:\n# - конверсия(CR) \n# - средний доход с пользователя (ARPU) \n# - средний доход с платящего пользователя (ARPPU).\n\ngrpA = exp_data.loc[exp_data.grp=='A']\ngrpA\n\ngrpB = exp_data.loc[exp_data.grp=='B']\ngrpB\n\n# #### Conversion Rate\n\ncr = exp_data.groupby('grp', as_index=False)\\\n .agg({'has_rev':'mean'})\ncr\n\n# В тестовой группе конверсия уменьшилась. Необходимо удостовериться в том, является ли это различие стат. значимо.\n# Для этого используем критерий согласия Пирсона или иначе хи квадрат.\n#\n# Сформулируем гипотезы:\n# \n# H0 - конверсии в двух группах не отличаются\n# \n# H1 - конверсии в двух группах отличаются\n\n# +\nstat, p, dof, expected = chi2_contingency(pd.crosstab(exp_data.grp, exp_data.has_rev))\n\nprint(p)\nif p > 0.05:\n print('Оснований для отклонения нулевой гипотезы у нас нет.')\nelif p < 0.05:\n print('Отклоняем нулевую гипотезу.')\nelse:\n print('Необходимо перестмотерть p-value')\n# -\n\n# ____\n\n# #### ARPU - Avarage Revenue Per User\n\nnp.mean(grpA.rev), np.mean(grpB.rev)\n\nlen(grpB.rev), len(grpA.rev) # размер выборок\n\n# Cредний доход с пользователя в двух группах отличается.\n# Необходимо подтвердить или отвергнуть это с помощью стат. тестов.\n#\n# Н0 - средний доход с пользователя в двух группах не отличается\n#\n# Н1 - средний доход с пользователя в двух группах отличается\n#\n# В нашем распоряжении критерий Стьюдента, его непараметрический аналог - критерий Манна-Уитни и бустрап.\n#\n# Критерий Стьюдента имеет ряд необходимых допущений: случайность выборки, ее репрезентативность, и нормалььное распределение, а так же гомогенность дисперсий. Последние два являются не строгими. При не нермальном распределении необходимо чтобы выборки были достаточно большими, а при различии в дисперсиях можно воспользоваться поправкой Уэлча при использовании t-критерия.\n# Так же можно прологорифмировать наши выбрки, но данная операция может навредить при принятии решений бизнесса, так как отличия в логорифмированных данных не то же самое что при сравнении обычный средних.И в наших данных при подсчете ARPU это невозможно так как есть большое количество записей с 0 оплатой, а как известно логорифм от 0 это бесконечность).\n#\n#\n# Критерий Манна-Уитни хорош когда распределение признака не нормально, но он плохо интерпритируем и так же чувсителен при наличии в данных большого кол-ва повторяющихся знаяений, а это как раз наш случай. \n#\n# Бустрап же менее чувствителен ко всем вышеизложенным требованиям но является 'тяжелым'.\n#\n# Я буду использовать кртерий стьюдента и бутсрап.\n\n# Из графиков выше было очевидно, что распределение не нормальное, но следует подтвердить это:\n\nprint(normaltest(grpA.rev))\nprint(normaltest(grpB.rev))\n\n# С помощью теста Левена, проверим различаются ли дисперсии:\n\nlevene(grpB.rev,grpA.rev)\n\n# p-value равный 0.22 не дает нам оснований отклонить гипотезу о равенстве дисперсий.\n\nttest_ind(grpA.rev, grpB.rev)\n\n# t-test на наших данных показал p-value > 0.05 , что не дает нам оснований отклонить нулевую гипотезу.\n\nmannwhitneyu(grpA.rev, grpB.rev)\n\n# Манн-Уитни так же показал уровень значимости выше 0.05.\n\n# Но давайте посмотрим что произойдет, если мы будем многократно извлекать средние, складывать их в массив, затем посторим распределение этих средних и применим стат тесты:\n\nmean_rev_grpA_1000 = []\nfor i in range(1000):\n mean_rev_grpA_1000.append(\n np.mean(grpA.rev.sample(frac=1,replace=True))\n )\n\nplt.hist(mean_rev_grpA_1000);\n\nmean_rev_grpB_1000 = []\nfor i in range(1000):\n mean_rev_grpB_1000.append(\n np.mean(grpB.rev.sample(frac=1,replace=True))\n )\n\nplt.hist(mean_rev_grpB_1000);\n\n# ЦПТ во всей красе!)\n\nttest_ind(mean_rev_grpA_1000,mean_rev_grpB_1000)\n\n\n# А вот средние из многократных выборок средних уже отличаются стат значимо!\n\n# Теперь напишем функцию бутсрапа, котрая будет извлекать из наших групп подвыборки, сравнивать разницу средних в этих выборках и класть эту разницу в массив и затем строить распределение разницы средних:\n\ndef get_bootstrap(\n data_column_1, # числовые значения первой выборки\n data_column_2, # числовые значения второй выборки\n boot_it = 1000, # количество бутстрэп-подвыборок\n statistic = np.mean, # интересующая нас статистика\n bootstrap_conf_level = 0.95 # уровень значимости\n):\n boot_len = max([len(data_column_1), len(data_column_2)])\n boot_data = []\n for i in tqdm(range(boot_it)): # извлекаем подвыборки\n samples_1 = data_column_1.sample(\n boot_len, \n replace = True # параметр возвращения\n ).values\n \n samples_2 = data_column_2.sample(\n boot_len, # чтобы сохранить дисперсию, берем такой же размер выборки\n replace = True\n ).values\n \n boot_data.append(statistic(samples_1-samples_2)) \n pd_boot_data = pd.DataFrame(boot_data)\n \n left_quant = (1 - bootstrap_conf_level)/2\n right_quant = 1 - (1 - bootstrap_conf_level) / 2\n quants = pd_boot_data.quantile([left_quant, right_quant])\n \n p_1 = norm.cdf(\n x = 0, \n loc = np.mean(boot_data), \n scale = np.std(boot_data)\n )\n p_2 = norm.cdf(\n x = 0, \n loc = -np.mean(boot_data), \n scale = np.std(boot_data)\n )\n p_value = min(p_1, p_2) * 2\n \n # Визуализация\n _, _, bars = plt.hist(pd_boot_data[0], bins = 50)\n for bar in bars:\n if abs(bar.get_x()) <= quants.iloc[0][0] or abs(bar.get_x()) >= quants.iloc[1][0]:\n bar.set_facecolor('red')\n else: \n bar.set_facecolor('grey')\n bar.set_edgecolor('black')\n \n plt.style.use('ggplot')\n plt.vlines(quants,ymin=0,ymax=50,linestyle='--')\n plt.xlabel('boot_data')\n plt.ylabel('frequency')\n plt.title(\"Histogram of boot_data\")\n plt.show()\n \n return {#\"boot_data\": boot_data, \n \"quants\": quants, \n \"p_value\": p_value}\n\n\nget_bootstrap(grpB.rev, grpA.rev)\n\n# Результат приминения бустрапа так же говорит нам о стат значимом различии среднего дохода с пользователя в двух группах.\n# На графике видно, что ноль не входит в доверительный интервал и p-value < 0.05. Хотя стоит отметить что p-value очень близка к 0.05. \n\n# _____\n\n# #### ARPPU - Avarage Revenue Per Paying User\n\n# Теперь анализируем только платящих пользователей.\n#\n# Данных еще меньше.\n# Гипотезы все те же:\n#\n# Н0 - средний доход с платящего пользователя в группах не различается\n#\n# Н1 - средний доход с платящего пользователя в группах различается\n#\n# Так как данных еще меньше, то выбор бустрапа будет еще более оправдан, но посмотрим на другие критерии тоже.\n\nlen(grpA.query('rev>0'))\n\nlen(grpB.query('rev>0'))\n\nttest_ind(grpA.query('rev>0').rev, grpB.query('rev>0').rev)\n\n# t-test дает нам основания для отклонения нулевой гипотезы.\n\nmannwhitneyu(grpA.query('rev>0').rev, grpB.query('rev>0').rev)\n\nget_bootstrap(grpA.query('rev>0').rev, grpB.query('rev>0').rev)\n\n# Все три теста гвоврят нам о том что мы должны отклонить нулевую гипотезу.\n\n# ### Выводы:\n#\n# 1. Конверсия в тестовой группе снизилась, но статистически это изменение оказалось не значимым.\n# Однако сам факт этого негативен, так как мы оценивали влияние влияние новой механики оплаты и по идее она должны была делать процесс покупки легче/удобнее и следовательно вести к увеличению конверсии.\n# 2. Средний доход с пользователя вырос, однако стат тесты показали разные результаты:\n# т-тест и манн-уитни показали что различия не значимы, тогда как бустрап показал значимые различия, но с р-value близким к пороговому значению принятия решения. Я думаю что дело в том, что в данных подавляющее большинство записей где сумма равна нулю. \n# 3. Средний доход с платящих пользователей так же вырос в тестовой группе и все тесты показали, что различия значимы.\n# 4. Так же хочется отметить что по сути единственное отличие в распределениях оплат в двух группах заключается в том, что в тестовой группе имеется пик оплат на сумму в 1900. Тут надо подумать как изменения механики оплаты может вести к тому, что пользователи наснут тратить определенную сумму. Самый плохой вариант это если во время проведения эксперимента проводилась еще какая нибудь акция и при сплитовании пользователей получилось так, что пользователи в тестовой группе видели не только изменение в механике оплаты, но и акцию, в таком случае наш А/В тест полностью не валиден. \n# 5. Однако с другой стороны, так как у нас образовательная платформа, и если представить что раньше у нас поекпались все курсы по отдельности, а теперь появилась возможность обьединять курсы в специальсности и брать их вместе платя определенную сумму, и если это считать изменением механики оплаты, то результаты теста показыват что результат положительный.\n#\n# Подводя итог скажу что катить новую механику оплаты(чтобы под этим не подразумевалось) на всех, я бы пока не стал.\n# Собрал бы еще данные, проверил правильно сплитуются пользователи на группы, продлил бы время проведения А/В теста и убедился что мы тестируем влияние какого то одного эффекта, а не новой мехаики и акций и прочего.\n#\n#\n\n# ____\n\n# ## Задание 2. SQL\n\n# Все данные находятся в табличном виде в Clickhouse.\n\n# #### 2.1 Очень усердные ученики.\n#\n# 2.1.1 Условие\n#\n# Образовательные курсы состоят из различных уроков, каждый из которых состоит из нескольких маленьких заданий. Каждое такое маленькое задание называется \"горошиной\".\n#\n# Назовём очень усердным учеником того пользователя, который хотя бы раз за текущий месяц правильно решил 20 горошин.\n\n# Подключаемся к базе данных:\n\nconnection = {'host': 'http://clickhouse.beslan.pro:8080',\n 'database':'default',\n 'user':'student', \n 'password':'dpo_python_2020'\n }\n\n# Посмотрим на данные хранящиеся в таблице:\n\nq_1 = '''\n SELECT\n st_id,\n toDateTime(timest) as timest,\n correct, \n subject\n \n FROM \n default.peas\n LIMIT 10\n \n '''\nquery_1 = ph.read_clickhouse(query=q_1, connection=connection)\nquery_1\n\n# Cмотрим за какой период времени данные в тадлице:\n\nq_2 = '''\n SELECT\n min(toDateTime(timest)) as first_date,\n max(toDateTime(timest)) as last_date\n FROM \n default.peas\n \n '''\nquery_2 = ph.read_clickhouse(query=q_2, connection=connection)\nquery_2\n\n# Считаем усердных учеников:\n\nq_3 = '''\n SELECT\n COUNT(DISTINCT st_id) as good_students\n FROM (\n SELECT\n st_id,\n SUM(correct) as corr_tasks\n FROM \n default.peas\n GROUP BY\n st_id\n HAVING\n corr_tasks >= 20\n ) \n '''\nquery_3 = ph.read_clickhouse(query=q_3, connection=connection)\nquery_3\n\n# Ну очевидно, что если в условии задачи под текущим месяцем подразумевается ноябрь 2022, то таких студентов в таблице не обнаружится.\n\n# ____\n\n# #### 2.2 Оптимизация воронки\n#\n# 2.2.1 Условие\n#\n# Образовательная платформа предлагает пройти студентам курсы по модели trial: студент может решить бесплатно лишь 30 горошин в день. Для неограниченного количества заданий в определенной дисциплине студенту необходимо приобрести полный доступ. Команда провела эксперимент, где был протестирован новый экран оплаты.\n\n# \n#\n\n# Необходимо в одном запросе выгрузить следующую информацию о группах пользователей:\n#\n# - ARPU \n# - ARPAU \n# - CR в покупку \n# - СR активного пользователя в покупку \n# - CR пользователя из активности по математике (subject = ’math’) в покупку курса по математике\n# - ARPU считается относительно всех пользователей, попавших в группы.\n#\n# Активным считается пользователь, за все время решивший больше 10 задач правильно в любых дисциплинах.\n#\n# Активным по математике считается пользователь, за все время решивший 2 или больше задач правильно по математике.\n\nq_4 = '''\n\n SELECT \n test_grp as grp,\n round(SUM(fcp.money)/uniqExact(s.st_id),2) as ARPU,\n round(SUMIf(fcp.money, query_in.act > 10)/uniqIf(s.st_id, query_in.act > 10),2) as ARPAU,\n round(uniqIf(s.st_id, fcp.money > 0)/uniqExact(s.st_id)*100, 2) as CR,\n round(uniqIf(s.st_id, query_in.act > 10 and fcp.money > 0)/uniqIf(s.st_id, query_in.act > 10)*100,2) as CR_active,\n round(uniqIf(s.st_id, query_in.math_active >= 2 and fcp.subject = 'Math' and fcp.money > 0)*100 /\n uniqIf(s.st_id, query_in.math_active >= 2), 2) as CR_math\n FROM \n studs s\n LEFT JOIN final_project_check fcp\n ON s.st_id = fcp.st_id\n LEFT JOIN (\n SELECT \n st_id, \n SUM(correct) as act, \n SUMIf(correct, subject = 'Math') as math_active\n FROM \n peas\n GROUP BY \n st_id) as query_in \n ON s.st_id = query_in.st_id\n\n GROUP BY test_grp\n\n'''\nquery_4 = ph.read_clickhouse(query=q_4, connection=connection)\nquery_4\n\n\n# ____\n\n# ## Задание 3. Python\n#\n# #### 3.1 Задача\n#\n# - Реализуйте функцию, которая будет автоматически подгружать информацию из дополнительного файла groups_add.csv (заголовки могут отличаться) и на основании дополнительных параметров пересчитывать метрики.\n#\n# - Реализуйте функцию, которая будет строить графики по получаемым метрикам.\n\ndef get_data(df1: 'DataFrame', df2: 'DataFrame', df3: 'DataFrame', df4: 'DataFrame') -> 'DataFrame':\n '''\n Функция get_data принимает в качестве атрибутов \n 4 датафрейма соединяет их рассчитавет метрики\n и подает на выход датафрейм с метриками.\n \n '''\n df1.columns = df2.columns\n all_groups = pd.concat([df1, df2])\n all_groups.rename(columns={'id':'student_id'}, inplace=True)\n df3['active'] = 1\n df4['has_rev'] = 1\n final_df = all_groups.merge(df3, how='left', on='student_id').fillna(0)\n final_df = final_df.merge(df4, how='left', on='student_id').fillna(0)\n final_df = final_df.query('active == 1')\n metrics_df = final_df.groupby('grp', as_index=False)\\\n .agg({'student_id':'count','rev':['sum','mean'],'has_rev':['sum','mean'], })\n \n metrics_df.columns = ['grp','active_users','revenue','ARPU','paying_users','CR']\n #metrics_df = metrics_df.assign(paying_users = final_df.query('rev > 0').groupby('grp').agg({'rev': 'count'})) \n #metrics_df['ARPAU'] = (metrics_df.revenue / metrics_df.active_users)\n metrics_df['ARPPU'] = round(metrics_df.revenue / metrics_df.paying_users,2)\n metrics_df['ARPU'] = round(metrics_df['ARPU'],2)\n metrics_df['CR'] = round(metrics_df.CR*100,2)\n metrics_df['revenue'] = round(metrics_df.revenue,2)\n \n return metrics_df\n\n\nmetrics = get_data(groups,groups_add,active_studs,checks)\nmetrics\n\n\ndef show_metrics(metrics: 'DataFrame') -> 'graphs':\n '''\n Функция show_metrics принимает на вход один атрибут-датафрейм с метриками\n и возвращает графики данных метрик.\n \n '''\n \n plt.figure(figsize=(10, 6))\n splot=sns.barplot(x=metrics.grp,y=\"CR\",data=metrics)\n for p in splot.patches:\n splot.annotate(format(p.get_height(), '.1f'), \n (p.get_x() + p.get_width() / 2., p.get_height()), \n ha = 'center', va = 'center', \n xytext = (0, 9), \n textcoords = 'offset points')\n \n plt.figure(figsize=(10, 6))\n splot=sns.barplot(x=metrics.grp,y=\"ARPU\",data=metrics)\n for p in splot.patches:\n splot.annotate(format(p.get_height(), '.1f'), \n (p.get_x() + p.get_width() / 2., p.get_height()), \n ha = 'center', va = 'center', \n xytext = (0, 9), \n textcoords = 'offset points')\n\n plt.figure(figsize=(10, 6))\n splot=sns.barplot(x=metrics.grp,y=\"ARPPU\",data=metrics)\n for p in splot.patches:\n splot.annotate(format(p.get_height(), '.1f'), \n (p.get_x() + p.get_width() / 2., p.get_height()), \n ha = 'center', va = 'center', \n xytext = (0, 9), \n textcoords = 'offset points')\n \n\n \n return plt.show()\n\nshow_metrics(metrics)\n","repo_name":"AlexKuchiev/Karpov.Courses_projects","sub_path":"final.ipynb","file_name":"final.ipynb","file_ext":"py","file_size_in_byte":35093,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"31667184147","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#
\n\n# + id=\"0iDzG5n8eBm_\" colab_type=\"code\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34} outputId=\"fcb93288-5194-40c8-c624-7fff7f2ac118\"\nsome_list = [5, 10, 20, 30, 40, 50]\n\ndef only_ends(the_list):\n ends_list = []\n ends_list.append(the_list[0])\n ends_list.append(the_list[-1])\n return ends_list\n\nonly_ends(some_list)\n","repo_name":"tastiz/python_challenges","sub_path":"onlyListEnds.ipynb","file_name":"onlyListEnds.ipynb","file_ext":"py","file_size_in_byte":607,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"28612213132","text":"import pkgutil\nimport inspect\nimport importlib\nimport datetime\n\nbuiltin_modules = dir(__builtin__)\n\n# +\n# pkgutil.iter_modules()\n# -\n\nmodule_list = []\nfor module_name in pkgutil.iter_modules():\n [file_finder, module_name, ispkg] = module_name\n if not ispkg:\n continue\n # print(module_name)\n try:\n module = importlib.import_module(module_name)\n module_list.append([file_finder, module_name, ispkg])\n except:\n pass\n # print(module_name['ispkg'])\n # print(module_name[2])\n # for key in module_name:\n # print('\\t', key)\n\n# +\nmodule_name_list = []\ndef get_module_name(module_obj):\n str_module_obj = str(module_obj)\n add_module_obj = str_module_obj.split(\" -1:\n return_bool = True\n elif str(obj_name)[:2] == '__' and str(obj_name)[-2:] == '__':\n return_bool = True\n return return_bool\n \ndef get_modules(module_obj):\n for obj_name, obj in inspect.getmembers(module_obj, inspect.ismodule):\n try:\n if check_builtin_function(obj_name, obj):\n continue\n if get_module_name(obj):\n continue\n get_modules(obj)\n get_classes(obj)\n except:\n pass\n \ndef get_classes(module_obj):\n for obj_name, obj in inspect.getmembers(module_obj, inspect.isclass):\n try:\n if check_builtin_function(obj_name, obj):\n continue\n if get_class_name(obj):\n continue\n # get_modules(obj)\n get_classes(obj)\n # get_functions(obj)\n except:\n pass\n \ndef get_functions(module_obj, module_name_list=[]):\n for obj_name, obj in inspect.getmembers(module_obj, inspect.isfunction):\n try:\n if check_builtin_function(obj_name, obj):\n continue\n module_name_list.append(obj_name)\n # print('\\t\\t\\t', obj_name, obj)\n except:\n pass\n return module_name_list\n\n\n# -\n\nstart_time = datetime.datetime.now()\nprint('start_time', start_time)\nfor module_atr in module_list:\n # print('*'*50)\n # print(module_atr)\n [file_finder, module_name, ispkg] = module_atr\n # print(file_finder, module_name, ispkg)\n module = importlib.import_module(module_name)\n # print('\\t', module)\n # print(module)\n # print('')\n # print('\\t', dir(module))\n \n get_modules(module)\n get_classes(module)\n # get_functions(module)\n # break\nend_time = datetime.datetime.now()\nprint('end_time', end_time)\nprint(end_time - start_time)\n\nfor idx, x in enumerate(class_name_list):\n print(idx, '\\t', x)\n if idx > 10:\n break\n\nfor idx, x in enumerate(module_name_list):\n print(idx, '\\t', x)\n if idx > 10:\n break\n\nidx = 1\nclass_name = class_name_list[idx]\nprint(class_name)\nmodule_name = class_name.split('.')\nprint(module_name, len(module_name))\n# print(module_name[:-1])\n# print('.'.join(module_name[:-1]))\nif len(mdoule_name) > 1:\n module_name = '.'.join(module_name[:-1])\nprint(module_name)\nclass_obj = importlib.import_module(module_name)\n\n# dir(class_obj)\n# print(inspect.getmembers(class_obj))\n# function_name_list = []\n# for obj in inspect.getmembers(class_obj, inspect.isfunction):\n# # for obj in inspect.getmembers(class_obj):\n# print(obj)\nfunction_name_list = get_functions(class_obj)\n\nfor idx in range(0, len(function_name_list)):\n print('*'*50)\n # print(idx, '\\t', function_name_list[idx])\n # function_obj = getattr(class_obj, function_name_list[idx])\n # print('\\t', function_obj)\n # print(inspect.getsource(function_obj))\n print(inspect.getsource(getattr(class_obj, function_name_list[idx])))\n\nprint(inspect.getsource(class_obj))\n\n\n","repo_name":"chibadai/JupyterLab","sub_path":"notebooks/pkgutil-to-source-code-20200308-1.ipynb","file_name":"pkgutil-to-source-code-20200308-1.ipynb","file_ext":"py","file_size_in_byte":4494,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43231152486","text":"# + [markdown] pycharm={\"name\": \"#%% md\\n\"}\n# ### 从零实现线性回归\n#\n# #### 基础操作定义\n\n# +\nimport random\nimport torch\nfrom matplotlib import pyplot as plt\nimport os\n\n# 避免当下版本matplotlib与pytorch的冲突\nos.environ['KMP_DUPLICATE_LIB_OK'] = 'True'\n\n\n# + pycharm={\"name\": \"#%%\\n\"}\ndef synthetic_data(w, b, num_examples): #@save\n \"\"\"生成 y = Xw + b + 噪声\"\"\"\n X = torch.normal(0, 1, (num_examples, len(w)))\n y = torch.matmul(X, w) + b\n y += torch.normal(0, 0.01, y.shape)\n return X, y.reshape((-1, 1))\n\n\ntrue_w = torch.tensor([2, -3.4])\ntrue_b = 4.2\nfeatures, labels = synthetic_data(true_w, true_b, 1000)\n\nplt.scatter(features[:, 1].detach().numpy(), labels.detach().numpy(), 1)\nplt.show()\n\n\n# + pycharm={\"name\": \"#%%\\n\"}\ndef data_iter(batch_size, features, labels):\n \"\"\"\n 随机采样\n :param batch_size: 随机样本大小\n :param features: 数据集\n :param labels: 数据集对应的标签\n :return: 随机样本\n \"\"\"\n num_examples = len(features)\n indices = list(range(num_examples))\n # 这些样本是随机读取的,没有特定的顺序\n random.shuffle(indices)\n for i in range(0, num_examples, batch_size):\n batch_indices = torch.tensor(indices[i: min(i + batch_size, num_examples)])\n yield features[batch_indices], labels[batch_indices]\n\n\n# + [markdown] pycharm={\"name\": \"#%% md\\n\"}\n# #### 模型定义\n\n# + pycharm={\"name\": \"#%%\\n\"}\n# 初始化模型参数\n# 从均值为0、标准差为0.01的正态分布中采样随机数来初始化权重,并将偏置初始化为0\nw = torch.normal(0, 0.01, size=(2, 1), requires_grad=True)\nb = torch.zeros(1, requires_grad=True)\n\n\n# 定义线性回归模型\ndef lin_reg(X, w, b):\n return torch.matmul(X, w) + b\n\n\n# 定义损失函数\ndef squared_loss(y_hat, y):\n return (y_hat - y.reshape(y_hat.shape)) ** 2 / 2\n\n\n# 定义优化算法: 小批量随机梯度下降\ndef sgd(params, lr, batch_size):\n with torch.no_grad():\n for param in params:\n param -= lr * param.grad / batch_size\n param.grad.zero_()\n\n\n# -\n\n# #### 训练模型\n\n# + pycharm={\"name\": \"#%%\\n\"}\ndef train(lr, num_epochs, batch_size, net, loss):\n for epoch in range(num_epochs):\n for X, y in data_iter(batch_size, features, labels):\n # X和y的小批量损失\n l = loss(net(X, w, b), y)\n\n # l的形状是(batch_size, 1),而不是一个标量\n # l中的所有元素被加到一起,并以此计算关于[`w`, `b`]的梯度\n l.sum().backward()\n\n # 使用参数的梯度更新参数\n sgd([w, b], lr, batch_size)\n # 接下来的计算不需要梯度,所以使用语句with torch.no_grad():\n with torch.no_grad():\n train_l = loss(net(features, w, b), labels)\n print(f'epoch {epoch + 1}, loss {float(train_l.mean()):f}')\n\n\ntrain(0.2, 10, 10, lin_reg, squared_loss)\nprint(f'w的估计误差: {true_w - w.reshape(true_w.shape)}')\nprint(f'b的估计误差: {true_b - b}')\n# -\n\n# ### 简洁实现\n\n# + pycharm={\"name\": \"#%%\\n\"}\nimport numpy as np\nfrom torch.utils import data\nfrom torch import nn\n\n# 生成数据集\ntrue_w = torch.tensor([2, -3.4])\ntrue_b = 4.2\nfeatures, labels = synthetic_data(true_w, true_b, 1000)\n\n\n# 加载数据集\ndef load_array(data_arrays, batch_size, is_train=True): #@save\n # 构造一个PyTorch数据迭代器\n dataset = data.TensorDataset(*data_arrays)\n # 从数据集dataset中抽取batch_size个数据,shuffle指示是否随机抽取\n return data.DataLoader(dataset, batch_size, shuffle=is_train)\n\n\n\"\"\"\n创建模型\n 利用Sequential容器建立神经网络\n 利用Linear生成全连接层,也就是本次需要的线性回归函数,两个参数分别为输入个数与输出个数\n\"\"\"\nnet = nn.Sequential(nn.Linear(2, 1))\n# 初始化权重与偏差\n# 利用平均值为0,方差为0.01的高斯分布初始化模型权重\nnet[0].weight.data.normal_(0, 0.01)\nnet[0].bias.data.fill_(0)\n\n# 损失函数与优化算法\nloss = nn.MSELoss()\nopt = torch.optim.SGD(net.parameters(), lr=0.3)\n\n# 训练\nbatch_size = 10\ndata_iter = load_array((features, labels), batch_size)\nnum_epochs = 3\nfor i in range(num_epochs):\n for X, y in data_iter:\n # 计算损失函数\n l = loss(net(X), y)\n # 清零梯度\n opt.zero_grad()\n # 计算梯度\n l.backward()\n # 更新模型\n opt.step()\n l = loss(net(features), labels)\n print(f'epoch {i + 1}, loss {l:f}')\n","repo_name":"MikuAndRabbit/note-DiveIntoDeepLeaning","sub_path":"03-linear_neural_network/01-linear_regression.ipynb","file_name":"01-linear_regression.ipynb","file_ext":"py","file_size_in_byte":4515,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"18281383662","text":"import pandas as pd\nimport requests\nfrom pycoingecko import CoinGeckoAPI\nfrom pprint import pprint\nfrom binance import Client\nfrom datetime import datetime\nimport yfinance as yf\n\ncg = CoinGeckoAPI()\nclient = Client()\n\n# Se decide hacer un análisis sobre la categoría Capa 1 (L1) de criptomonedas por su capitalización de mercado.\n\n# Categorías de Interes\n\nCategorias=['Layer 1 (L1)']\n\n#Lista de todas las categorías\ncategoria=pd.DataFrame(cg.get_coins_categories_list())\ncategoria[categoria['name'].isin(Categorias)]\n\n#Lista de todas las categorías con datos de mercado\ndatos_merc_ct=pd.DataFrame(cg.get_coins_categories(order='market_cap_asc'))\ndatos_merc_ct=datos_merc_ct.sort_values(by='market_cap',ascending=False).dropna(subset='market_cap')\ndatos_merc_ct.head()\n\n\ndef get_coin_list():\n return cg.get_coins_list()\n\n\ndf1=get_coin_list()\ndf1=pd.DataFrame(df1)\n\n#crypto_symbol=['btc','eth','bnb','ada','sol','usdt','usdc','dai','busd','tusd']\ndf1[df1['name'].str.contains('TrueUS', case=False)]\n\n\n# Criptomonedas seleccionadas para el análisis.\n\ncriptos_order=['Bitcoin','Ethereum','BNB','Cardano','Solana','Polkadot','Bitcoin Cash','Avalanche',\n 'Cosmos Hub','Hedera']\ncriptos = df1[df1['name'].isin(criptos_order)]\ncriptos\n\ncriptos['symbol']\n\n\n# ### Generando Datos para las diferentes Criptos Seleccionadas:\n\n# Obtenga datos de mercado históricos que incluyen precio, capitalización de mercado y volumen de 24 horas dentro de un rango de marca de tiempo (granularidad automática)\n\ndef get_coin_market_chart_range_by_id(id:str,markert_chart:str,to:str,From:str):\n ''' Ejem: id = Bitcoin , markert_chart = usd, to = 1660539600, from = 1692079200 '''\n return cg.get_coin_market_chart_range_by_id(id,markert_chart,to,From)\n\n\n# +\nfrom datetime import datetime\n#1660971600 ----> 2022-08-20\n#1692424800 ----> 2023-08-19\n\ntimestamp = 1692424800\n\n# Convertir el timestamp a una fecha legible\nfecha_legible = datetime.utcfromtimestamp(timestamp).strftime('%Y-%m-%d %H:%M:%S')\n\nprint(fecha_legible)\n\n\n# -\n\ndef extract_unix_timestamp(item):\n fecha_ilegible= item[0] // 1000 # Dividir por 1000 para obtener 10 dígitos\n fecha_legible = datetime.utcfromtimestamp(fecha_ilegible).strftime('%Y-%m-%d')\n return fecha_legible\n\n\ndef remove_unix_timestamp(item):\n return item[1]\n\n\n# +\n#Está lista se ocupara para que todos los DataFrames tengan las mismas fechas.\nfecha_inicio = '2022-08-20'\nfecha_fin = '2023-08-19'\n\nrango_fechas = pd.date_range(start=fecha_inicio, end=fecha_fin, freq='D')\n\n# Crear un DataFrame con la columna de fechas\ndf_Fechas = pd.DataFrame({'fechas': rango_fechas})\n\n\n# -\n\ndef df_market(id:str): \n #Creación DataFrame\n gcmcrange=get_coin_market_chart_range_by_id(id,'usd','1660971600','1692424800')\n df=pd.DataFrame(gcmcrange)\n #Extracción Fechas:\n df['fechas'] = df['prices'].apply(extract_unix_timestamp)\n #Remove Unix Time\n for i in df:\n df[i+'2'] = df[i].apply(remove_unix_timestamp)\n #Formato DataTime para posterior merge:\n df['fechas'] = pd.to_datetime(df['fechas'])\n\n columnas_a_eliminar = ['prices', 'market_caps','total_volumes','fechas2']\n df=df.drop(columns= columnas_a_eliminar)\n\n df_union = pd.merge(df_Fechas, df, on='fechas', how='outer')\n\n #Agregando Columna con ID \n df_union.insert(0, 'Cripto', id)\n \n\n return df_union\n\n# #### Categoría: Layer 1 (L1)\n\ncriptos\n\ndf_Bitcoin_market=df_market('bitcoin')\ndf_Ethereum_market=df_market('ethereum')\ndf_BNB_market=df_market('binancecoin')\ndf_Cardano_market=df_market('cardano')\ndf_Solana_market=df_market('solana')\ndf_polkadot_market=df_market('polkadot')\ndf_bitcoin_cash_market=df_market('bitcoin-cash')\ndf_Avalanche_market=df_market('avalanche-2')\ndf_cosmos_market=df_market('cosmos')\ndf_hedera_hashgraph_market=df_market('hedera-hashgraph')\n\n# +\n# Lista de DataFrames a agregar\ndataframes = [df_Bitcoin_market, df_Ethereum_market,df_BNB_market,df_Cardano_market,\n df_Solana_market,df_polkadot_market,df_bitcoin_cash_market,df_Avalanche_market\n ,df_cosmos_market,df_hedera_hashgraph_market]\n\ndf_concatenated = pd.concat(dataframes, ignore_index=False)\n# -\n\ndf_concatenated.shape\n\n\n# ### Extracción de precios de apertura, mayor precio, menor y cierrre.\n\ndef Candles_Cripto(id):\n\n df = pd.DataFrame(client.get_historical_klines(id,'1d','1660971600','1692424800'))\n df = df.iloc[:,:5]\n df.columns = ['fechas','Open','High','Low','Close',]\n df['fechas'] = pd.to_datetime(df.fechas,unit='ms')\n df = df.set_index('fechas')\n df = df.astype(float)\n return df\n\n\nSymbol=(criptos['symbol'].tolist())\n\nSymbol\nfor i in range(len(Symbol)):\n Symbol[i] = Symbol[i] + 'usdt'\nSymbol = [item.upper() for item in Symbol]\n\ncriptos\n\nSymbol\n\ndf_Candles_Bitcoin=Candles_Cripto('BTCUSDT')\ndf_Candles_Ethereum=Candles_Cripto('ETHUSDT')\ndf_Candles_BNB=Candles_Cripto('BNBUSDT')\ndf_Candles_Cardano=Candles_Cripto('ADAUSDT')\ndf_Candles_Solana=Candles_Cripto('SOLUSDT')\ndf_Candles_polkadot=Candles_Cripto('DOTUSDT')\ndf_Candles_bitcoin_cash=Candles_Cripto('BCHUSDT')\ndf_Candles_Avalanche=Candles_Cripto('AVAXUSDT')\ndf_Candles_cosmos=Candles_Cripto('ATOMUSDT')\ndf_Candles_hedera_hashgraph=Candles_Cripto('HBARUSDT')\n\n\ndef merge_data(df1,df2):\n df1.set_index('fechas', inplace=True)\n df=df1.merge(df2,left_index=True,right_index=True)\n return df\n\n#merged_df = df_Bitcoin_market.merge(df_Candles_Bitcoin, left_index=True, right_index=True)\n\n\ndf_Bitcoin=merge_data(df_Bitcoin_market,df_Candles_Bitcoin)\n\n\ndf_Ethereum=merge_data(df_Ethereum_market,df_Candles_Ethereum)\ndf_BNB=merge_data(df_BNB_market,df_Candles_BNB)\ndf_Cardano=merge_data(df_Cardano_market,df_Candles_Cardano)\ndf_Solana=merge_data(df_Solana_market,df_Candles_Solana)\ndf_Polkadot=merge_data(df_polkadot_market,df_Candles_polkadot)\ndf_Avalanche=merge_data(df_Avalanche_market,df_Candles_Avalanche)\ndf_Cosmos=merge_data(df_cosmos_market,df_Candles_cosmos)\ndf_Hedera=merge_data(df_hedera_hashgraph_market,df_Candles_hedera_hashgraph)\ndf_Bitcoin_Cash=merge_data(df_bitcoin_cash_market,df_Candles_bitcoin_cash)\n\n\ndf_Bitcoin_Cash.shape\n\n# +\nLayer_1=[df_Bitcoin,df_Ethereum,df_BNB,df_Cardano,df_Solana,df_Polkadot,df_Avalanche\n ,df_Cosmos,df_Hedera,df_Bitcoin_Cash]\n\nLayer_1=pd.concat(Layer_1,ignore_index=False)\n# -\n\n#Asignar Ranking de acuerdo al MarketCap\nlatest_data = Layer_1.groupby('Cripto').apply(lambda group: group[group.index == group.index.max()])\nlatest_data = latest_data.sort_values(by='market_caps2', ascending=False)\nlatest_data['Ranking'] = range(1, len(latest_data) + 1)\nLayer_1['Ranking'] = Layer_1['Cripto'].map(latest_data.set_index('Cripto')['Ranking'])\n\n# Agregar Body Ratio\n\n# +\n# Calcular el Body Ratio\nLayer_1['Body_Ratio'] = abs(Layer_1['Close'] - Layer_1['Open']) / (Layer_1['High'] - Layer_1['Low'])\n\n# Clasificar las velas en tres categorías basadas en el Body Ratio y dirección de cierre\ndef classify_vela(row):\n if row['Body_Ratio'] < 0.3:\n return 'Indecisión'\n elif row['Body_Ratio'] >= 0.3 and row['Body_Ratio'] <= 0.7:\n return 'Fuerte Volatilidad'\n else:\n if row['Close'] > row['Open']:\n return 'Fuerte Dirección Alcista'\n else:\n return 'Fuerte Dirección Bajista'\n\nLayer_1['Clasificación'] = Layer_1.apply(classify_vela, axis=1)\n# -\n\n# Agregar Retorno Diario\n\nLayer_1['Daily_Return'] = Layer_1.groupby('Cripto')['Close'].pct_change()\n\nLayer_1['Daily_Return'].fillna(0, inplace=True)\n\n# +\n#Layer_1.to_csv('DataCriptos.csv',index=True)\n# -\n\n# Obtención de datos del Top 3 compañias en el sector de Tecnología\n#\n# Fuente Yahoo.\n\n# AAPL\tApple Inc.\n#\n# MSFT\tMicrosoft Corporation\n\nstart_date = \"2022-08-20\"\nend_date = \"2023-08-19\"\ncolumns=['Dividends','Stock Splits']\n\nAAPL = yf.Ticker(\"AAPL\")\ndf_AAPL = AAPL.history(period=\"1d\", start=start_date, end=end_date)\ndf_AAPL = AAPL.history(period=\"1d\", start=start_date, end=end_date)\ndf_AAPL.index = df_AAPL.index.date\ndf_AAPL=df_AAPL.drop(columns=columns)\nstart_date = df_AAPL.index.min()\nend_date = df_AAPL.index.max()\ndate_range = pd.date_range(start=start_date, end=end_date)\ndf_AAPL = df_AAPL.reindex(date_range)\ndf_AAPL.insert(0, 'Company', 'Apple Inc.')\ndf_AAPL=df_AAPL.interpolate()\n\nMSFT = yf.Ticker(\"MSFT\")\ndf_MSFT = MSFT.history(period=\"1d\", start=start_date, end=end_date)\ndf_MSFT = MSFT.history(period=\"1d\", start=start_date, end=end_date)\ndf_MSFT.index = df_MSFT.index.date\ndf_MSFT=df_MSFT.drop(columns=columns)\nstart_date = df_MSFT.index.min()\nend_date = df_MSFT.index.max()\ndate_range = pd.date_range(start=start_date, end=end_date)\ndf_MSFT = df_MSFT.reindex(date_range)\ndf_MSFT.insert(0, 'Company', 'Microsoft Corporation')\ndf_MSFT=df_MSFT.interpolate()\n\n\n# +\nDataTec=[df_AAPL,df_MSFT]\n\nDataTec=pd.concat(DataTec,ignore_index=False)\n\n# +\n# Calcular el Body Ratio\nDataTec['Body_Ratio'] = abs(DataTec['Close'] - DataTec['Open']) / (DataTec['High'] - DataTec['Low'])\n\nDataTec['Clasificación'] = DataTec.apply(classify_vela, axis=1)\n# -\n\n# Retorno Diario\n\nDataTec['Daily_Return'] = DataTec.groupby('Company')['Close'].pct_change()\n\nDataTec['Daily_Return'].fillna(0, inplace=True)\n\n# +\n#DataTec.to_csv('DataTec.csv',index=True)\n# -\n\nLayer_1['Cripto'].unique()\n\nDataTec[DataTec.Company=='Apple Inc.'].info()\n\nDataTec.info()\n","repo_name":"Benjaminmxxx/DataAnalytics","sub_path":"ETL.ipynb","file_name":"ETL.ipynb","file_ext":"py","file_size_in_byte":9255,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"18703646438","text":"# +\n#tweetlerin veritabanından alınması\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport nltk\nimport zemberek\nimport jpype,os\nfrom nltk.tokenize import word_tokenize\nfrom nltk.tokenize import TweetTokenizer\nfrom nltk.corpus import stopwords \ndurakwords= set(stopwords.words(\"turkish\"))\nimport string\nimport re\nfrom typing import List\nfrom jpype import JClass, JString, getDefaultJVMPath, shutdownJVM, startJVM, java\nfrom scipy import stats\nimport seaborn as sns\nfrom collections import Counter\nfrom keras.models import Sequential\nfrom keras.layers import Embedding,SpatialDropout1D,LSTM,Dense\nfrom keras.callbacks import ModelCheckpoint, EarlyStopping\n# %matplotlib inline\ndf=pd.DataFrame()\ndf=pd.read_csv(\"C:/Users/user/tweets/tweets2.csv\")\n\n\n# -\n\n#Tweetlerin küçük harfe dönüştürülmesi\ndf= df.rename(columns = {'hash,content,class': 'tweets'})\ndf['tweets']=df['tweets'].str.lower()\n\n#tweetlerin son değerlerine göre yeni bir dataframe e yerleştirilmesi\n#1 tek,2 spor,eko,pol,mag,sağ,kült\n#df[df.tweets.str.endswith('2')]\ndf['icerik']='icerik'\ndf['cleantext']='cleantext'\ndf.loc[df['tweets'].str.endswith('1'),'icerik']=\"teknoloji\"\ndf.loc[df['tweets'].str.endswith('2'),'icerik']=\"spor\"\ndf.loc[df['tweets'].str.endswith('3'),'icerik']=\"ekonomi\"\ndf.loc[df['tweets'].str.endswith('4'),'icerik']=\"politika\"\ndf.loc[df['tweets'].str.endswith('5'),'icerik']=\"magazin\"\ndf.loc[df['tweets'].str.endswith('8'),'icerik']=\"kultur\"\ndf.loc[df['tweets'].str.endswith('7'),'icerik']=\"saglik\"\ndf.loc[df['tweets'].str.endswith('1'),'label']=\"1\"\ndf.loc[df['tweets'].str.endswith('2'),'label']=\"2\"\ndf.loc[df['tweets'].str.endswith('3'),'label']=\"3\"\ndf.loc[df['tweets'].str.endswith('4'),'label']=\"4\"\ndf.loc[df['tweets'].str.endswith('5'),'label']=\"5\"\ndf.loc[df['tweets'].str.endswith('8'),'label']=\"8\"\ndf.loc[df['tweets'].str.endswith('7'),'label']=\"7\"\npd.to_numeric(df.label)\n\n#Zemberek kütüphanesinin kullanılabilmesi için Jpype ile JVM oluşturulması\nZEMBEREK_PATH = r'C:\\Users\\user\\Desktop\\dersler\\tweets\\zemberek-full.jar'\nstartJVM(getDefaultJVMPath(), '-ea', '-Djava.class.path=%s' % (ZEMBEREK_PATH))\n#Morphology kütüphanesini kullanacağımızdan burada oluşturuyoruz\nTurkishMorphology = JClass('zemberek.morphology.TurkishMorphology')\nmorphology = TurkishMorphology.createWithDefaults()\n\n\ndef mentionRemove(text):\n text = re.sub(r'@\\w+', '', text)#mentionların çıkartırılması\n return text\n\n\ndef siteRemove(text):\n text = re.sub(r'http.?://[^\\s]+[\\s]?', '', text)#bağlantıların çıkarılması\n return text\n\n\ndef puncRemove(text):\n text = re.sub('[^a-zığüşiöç\\s]', '', text)#özel karakterlerin ve sayıların çıkarılması\n text = re.sub(r\"Â\", \"A\", text)#umlautlu karakterlerinn değiştirilmesi\n text = re.sub(r\"â\", \"a\", text)\n text = re.sub(r\"Î\", \"I\", text)\n text = re.sub(r\"î\", \"ı\", text)\n text = re.sub(r\"Û\", \"U\", text)\n text = re.sub(r\"û\", \"u\", text)\n\n return text\n\n\n\ndef removeSw(text):\n text=[w for w in tweet if w.lower() not in durakwords]#stopwordlerin çıkarılması\n return text\n\n\ndef rootFinder(text):\n pos=[]\n textlist=text.split(\" \")\n analysis: java.util.ArrayList = ( morphology.analyzeAndDisambiguate(text).bestAnalysis() )\n for i, analysis in enumerate(analysis, start=1):\n f'\\nAnalysis {i}: {analysis}',\n f'\\nPrimary POS {i}: {analysis.getPos()}' \n if (str(analysis.getLemmas()[0]) == \"UNK\"):\n pos.append(textlist[i-1])\n else: \n pos.append(f'{str(analysis.getLemmas()[0])}')\n return pos\n\n\n\ndef emojiRemove(tweet):\n # Smile -- :), : ), :-), (:, ( :, (-:, :')\n tweet = re.sub(r'(:\\s?\\)|:-\\)|\\(\\s?:|\\(-:|:\\'\\))', ' EMO_POS ', tweet)\n # Laugh -- :D, : D, :-D, xD, x-D, XD, X-D\n tweet = re.sub(r'(:\\s?D|:-D|x-?D|X-?D)', ' EMO_POS ', tweet)\n # Love -- <3, :*\n tweet = re.sub(r'(<3|:\\*)', ' EMO_POS ', tweet)\n # Wink -- ;-), ;), ;-D, ;D, (;, (-;\n tweet = re.sub(r'(;-?\\)|;-?D|\\(-?;)', ' EMO_POS ', tweet)\n # Sad -- :-(, : (, :(, ):, )-:\n tweet = re.sub(r'(:\\s?\\(|:-\\(|\\)\\s?:|\\)-:)', ' EMO_NEG ', tweet)\n # Cry -- :,(, :'(, :\"(\n tweet = re.sub(r'(:,\\(|:\\'\\(|:\"\\()', ' EMO_NEG ', tweet)\n return tweet\n\n\n# +\n#hash kısmının ve sondaki içerik belirteci ve tweet bağlantısının kaldırılması\n#utf-16 karakterlerin elenmesi\n#kelimelerin ayrıştırılması\n#tweet içerisindeki bağlantıların çıkarılması\n#tweet içindeki mentionların çıkarılması\np=set(string.punctuation)\n\ndef cleaner(text):\n text=text\n text=mentionRemove(text)#mentionların çıkartırılması\n text=siteRemove(text)#bağlantıların çıkarılması\n text=puncRemove(text)#özel karakterlerin ve sayıların çıkarılması\n text=emojiRemove(text)\n text=rootFinder(text)#kelime köklerinin bulunması\n text=removeSw(text)#durak kelimelerin elenmesi\n return text\nfor x in range(0,len(df)):\n #tweet=\"ve ya da için baba veya\"\n tweet=df.iloc[x,0]\n tweet=mentionRemove(tweet)#mentionların çıkartırılması\n tweet=siteRemove(tweet)#bağlantıların çıkarılması\n #tweet.replace(\"aracılığıyla\",\"a\")\n tweet=puncRemove(tweet)#özel karakterlerin ve sayıların çıkarılması\n tweet=rootFinder(tweet)#kelime köklerinin bulunması\n tweet=removeSw(tweet)#durak kelimelerin elenmesi\n tweet=' '.join(tweet)\n #tweet=cleaner(tweet)\n df.iloc[x,2]=tweet\n pos=[]\n\n\n# -\n\nimport matplotlib.pyplot as plt\nimport os\nimport re\nimport shutil\nimport string\nimport tensorflow as tf\nfrom tensorflow.keras import regularizers\nfrom tensorflow.keras import layers\nfrom tensorflow.keras import losses\nfrom collections import Counter\nfrom sklearn.preprocessing import LabelEncoder\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import classification_report\nfrom tensorflow.keras.preprocessing.text import Tokenizer\nfrom tensorflow.keras.preprocessing.sequence import pad_sequences\nfrom sklearn.metrics import accuracy_score,confusion_matrix\n\nprint(tf.__version__)\nif tf.test.gpu_device_name():\n print(\"gpu device :{}\".format(tf.test.gpu_device_name()))\nelse:\n print(\"please install gpu version of tf\")\n\n\ndef counter_words(df):\n count = Counter()\n for i in df:\n for w in i.split():\n count[w] += 1\n return count\n\n\nc=counter_words(df.cleantext)\nnum_words = len(c)\nmax_seq_lenght = 12\n\ntweets=df.cleantext\nlabels=df.icerik\n\n# +\ntokenizer=Tokenizer(num_words=num_words)\ntokenizer.fit_on_texts(tweets.tolist())\n\nprint(str(tokenizer.texts_to_sequences([\"fenerbahçe ben sevgili\"])))\n# -\n\nX_train, X_test, Y_train, Y_test = train_test_split(tweets.tolist(),labels.tolist(), test_size = 0.3,stratify=labels.tolist(),\n random_state = 44)\nprint('Train data len: '+ str(len(X_train)))\nprint('labels: ', Counter(Y_train))\nprint('test data len: '+ str(len(X_test)))\nprint('labels: ' ,Counter(Y_test))\n\nX_train=np.array(tokenizer.texts_to_sequences(X_train))\nX_test=np.array(tokenizer.texts_to_sequences(X_test))\n#Y_train=np.array(tokenizer.texts_to_sequences(Y_train))\nX_train=pad_sequences(X_train, padding = 'post',maxlen=max_seq_lenght,truncating=\"post\")\nX_test=pad_sequences(X_test, padding = 'post',maxlen=max_seq_lenght,truncating=\"post\")\n\nprint(X_train[1])\nprint(Y_train[:10])\n\nle = LabelEncoder()\ntrain_labels=le.fit_transform(Y_train)\nprint(train_labels)\ntrain_labels=np.asarray(tf.keras.utils.to_categorical(train_labels))\nprint(train_labels)\ntest_labels=le.fit_transform(Y_test)\ntest_labels=np.asarray(tf.keras.utils.to_categorical(test_labels))\nlist(le.classes_)\n\ntrain_ds=tf.data.Dataset.from_tensor_slices((X_train,train_labels))\ntest_ds=tf.data.Dataset.from_tensor_slices((X_test,test_labels))\n\ntrain_labels=np.asarray(tf.keras.utils.to_categorical(le.fit_transform(Y_train)))\ntrain_labels\n\nnum_words = len(c)\n# Max number of words in each complaint.\nmax_seq_lenght = 12\n# This is fixed.\ndim = 100\nprint(len(c))\nmodel = tf.keras.Sequential()\nmodel.add(tf.keras.layers.Embedding(num_words+1,dim,input_length=max_seq_lenght,embeddings_regularizer=regularizers.l2(0.0001)))\nmodel.add(tf.keras.layers.Conv1D(128,7,activation='relu',kernel_regularizer=regularizers.l2(0.0001),bias_regularizer=regularizers.l2(0.0001)))\nmodel.add(tf.keras.layers.GlobalMaxPooling1D())\nmodel.add(tf.keras.layers.Dropout(0.2))\nmodel.add(tf.keras.layers.Dense(7,activation='sigmoid',kernel_regularizer=regularizers.l2(0.001),bias_regularizer=regularizers.l2(0.001)))\nmodel.summary()\n\nmodel.compile(loss=tf.keras.losses.CategoricalCrossentropy(),optimizer='Adam',metrics=[\"CategoricalAccuracy\"])\n\nepochs=10\nhistory=model.fit(train_ds.batch(128),epochs=epochs,validation_data=test_ds.batch(128),verbose=1)\n\nnew_complaint = ['erdoğan hastane millet millet oy']\nseq = tokenizer.texts_to_sequences(new_complaint)\npadded = pad_sequences(seq, maxlen=max_seq_lenght)\npred = model.predict(padded)\nlabels = ['ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji']\nprint(pred, labels[np.argmax(pred)])\n\npreds=model.predict(X_test)\npreds=preds.argmax(axis=1)\nactual=le.fit_transform(Y_test)\nprint(\"accuracy score is\",accuracy_score(preds,actual))\n\nprint(classification_report(actual, preds,target_names=['ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji']))\n\nconf_mat = confusion_matrix(actual,preds)\nfig, ax = plt.subplots(figsize=(15,15))\nsns.heatmap(conf_mat, annot=True, cmap=\"Blues\", fmt='d',\n xticklabels={'ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji'},\n yticklabels={'ekonomi', 'kultur', 'magazin', 'politika', 'saglik', 'spor', 'teknoloji'})\nplt.ylabel('Actual')\nplt.xlabel('Predicted')\nplt.title(\"CONFUSION MATRIX\\n\", size=16);\n\npred=model.predict()\n\n\n","repo_name":"alprbykn/tweet_classification","sub_path":"tweets_neuralNetworks.ipynb","file_name":"tweets_neuralNetworks.ipynb","file_ext":"py","file_size_in_byte":9783,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"71874153753","text":"# ## Taylor Sin(x) Serisi by OVO Analytics\n# Made by Eren BOZARIK\n\nimport numpy as np\nimport time\n\n\ndef taylor_sin(x):\n start_time = time.time()\n x1=x*(np.pi/180)\n val=0\n for n in range(0, 25):\n add=(((-1) ** n) * (x1 ** (2*n+1)) / np.math.factorial(2*n + 1))\n val+=add\n end_time = time.time()\n print(\"{:.2f} surede hesaplandi\" .format(float(end_time - start_time)))\n return val\n\n\na=int(input(\"Hesaplanacak Dereceyi Giriniz :\"))\nprint(\"Sin({}) = {:.2f}\".format(a,taylor_sin(a)))\n","repo_name":"erenbozarik/Statistical-Scripts","sub_path":"Taylor_Series.ipynb","file_name":"Taylor_Series.ipynb","file_ext":"py","file_size_in_byte":517,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"70983907353","text":"# # Project: Writing a Data Scientist Blog Post\n\n# ## Introduction and Question\n\n# - Ford GoBike System Data: This data set includes information about individual rides made in a bike-sharing system covering the greater San Francisco Bay area.\n#\n# - The data including information of 183.412 ford gobike trips in San Francisco area with 16 variables (duration_sec, start_time, end_time, start_station_id, start_station_name, start_station_latitude, start_station_longitude, end_station_id, end_station_name, end_station_latitude ,end_station_longitude, bike_id, user_type, member_birth_year, member_gender, bike_share_for_all_trip). But some data have wrong so it needs to be clean them.\n#\n# - Question: How location distribution, user type, user age and also gender afftect to the trip?\n\n# ## Data Cleaning and Accessing\n\n# ### Data Cleaning\n\n# +\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nimport warnings\n\n# %matplotlib inline\nwarnings.simplefilter(\"ignore\")\n# -\n\nford_df = pd.read_csv(\"201902-fordgobike-tripdata.csv\")\nford_df.head()\n\nford_df.info()\n\nford_df.columns\n\n# #### Some columns have wrong datatype. So I need to convert them.\n#\n# 1. start_time and end_time shoud be datetime instend object\n#\n# 2. bike_share_for_all_trip should be converted to boolean instead\n\nford_df['start_time'] = ford_df['start_time'].astype('datetime64[ns]')\nford_df['end_time'] = ford_df['end_time'].astype('datetime64[ns]')\n\nford_df.bike_share_for_all_trip = ford_df.bike_share_for_all_trip.replace({\"Yes\": True, \"No\":False}).astype(bool)\n\nford_df.info()\n\n# ### Data Accessing\n\n# #### What are percentage of User Type and Member Gender?\n\nuser_type = ford_df['user_type'].value_counts()\nax = user_type.plot(kind='pie', autopct= '%1.1f%%',figsize=(7, 7));\n# set the title\nax.set_title(\"User Type\" , fontsize=20, fontweight='bold');\n\n# There's huge number of people that have subscribed.\n\nmember_gender = ford_df['member_gender'].value_counts()\nax = member_gender.plot(kind='pie', autopct= '%1.1f%%',figsize=(7, 7));\n# set the title\nax.set_title(\"Member Gender\" , fontsize=20, fontweight='bold');\n\n\n# Member gender male is more than female. Female is more than Other\n\n# #### Is there any relationship between duration second and user type?\n\ndef barplot(col, title, xlabel, ylabel, rotation):\n \"\"\"\n This function provide the counts for the each series in the column and plot it in bar plot.\n Input: column, title, xlabel, ylabel, rotation of xticks.\n \n Output: plt bar plot of value counts\n \"\"\"\n fig, ax = plt.subplots(figsize = (6,6), dpi = 120);\n col.value_counts().plot(kind='bar');\n plt.title(title, fontweight='bold');\n plt.ylabel(xlabel);\n plt.xlabel(ylabel);\n plt.xticks(rotation=rotation);\n\n\nbarplot(ford_df['user_type'], \"User Type by mean trip duration\", \"User Type\", \"Trip duration in second\", 0)\n\n# Although the number of Subscriber user is more than number of Customer, Customer trip duration is more than Subscriber\n\n# #### Is there any relationship between duration second, week day and user type?\n\nx = ford_df['start_time'].dt.day_name()\ny = 'duration_sec'\norders = [\"Monday\", \"Tuesday\",\"Wednesday\", \"Thursday\", \"Friday\",\"Saturday\",\"Sunday\"]\n\nfig, ax = plt.subplots(figsize = (12,5), dpi = 150);\nsns.boxplot(data=ford_df, x=x, y=y, order=orders,hue='user_type');\nplt.ylim([0,2500]);\nplt.title(\"Duration second by day of week\", fontweight='bold');\nplt.xlabel(\"Day of Week\");\nplt.ylabel(\"Duration second\");\nplt.legend(loc=1);\n\n# It seems that more trip in weekend than day week for Customer and for Subscriber it the same for all week.\n\n# #### What is top 10 Start and End Station have most rider?\n\naverage_line = ford_df['bike_id'].count() / ford_df.groupby(['start_station_name'])['start_station_name'].count()\naverage_line = average_line.nunique()\n\n# +\n# Separate the genre based on the pipe (|).\nford_df['start_station_name'] = ford_df['start_station_name'].astype(str)\nstationList = ford_df['start_station_name'].str.split('|')\n\n# Create list of station\nstation = []\nfor i in stationList:\n station.extend(i)\n \n# Plot based on the value counts\nplt.subplots(figsize=(12,10));\npd.Series(station).value_counts()[:10].sort_values(ascending=False).plot.barh(width=0.9,color=sns.color_palette('coolwarm',10));\nplt.title('Top 10 Start Station have most rider and Average line', fontweight='bold');\nplt.xlabel('Count');\nplt.ylabel('Station Name');\nplt.vlines(x=average_line, ymin=-2, ymax=20, color='r');\nplt.show();\n# -\n\n# Based on the chart above, it demonstrates that Market St at 10th St has most rider when they start riding\n\n# +\n# Separate the genre based on the pipe (|).\nford_df['end_station_name'] = ford_df['end_station_name'].astype(str)\nstationList = ford_df['end_station_name'].str.split('|')\n\n# Create list of station\nstation = []\nfor i in stationList:\n station.extend(i)\n \n# Plot based on the value counts\nplt.subplots(figsize=(12,8));\npd.Series(station).value_counts()[:10].sort_values(ascending=False).plot.bar(width=0.9,color=sns.color_palette('coolwarm',10));\nplt.title('Top 10 End Station have most rider', fontweight='bold');\nplt.xlabel('Count');\nplt.ylabel('Station Name');\nplt.show();\n# -\n\n# Based on the chart above, it demonstrates that Market St at 10th St has most rider when they start riding. But the end station is not this one. It's a San Francisco Caltrain Staion 2 because maybe around this station have a lot the beautiful place or famous restaurant,...\n\n# ## Conclusions\n\n# In conclusion, Subcribers have a huge ratio, but customers spend more time on riding than another. As a result, we need to have a strategy to convert them to our customers. Besides, Almost riders usually ride bicycle at weekend. That is great if we orginize some gameshow, marketing programs at that time to attract more customers.\n","repo_name":"tranhoang030998/Writing-a-Data-Scientist-Blog-Post","sub_path":"Udacity_BlogPost.ipynb","file_name":"Udacity_BlogPost.ipynb","file_ext":"py","file_size_in_byte":5848,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"10957102887","text":"# # # Kata 8\n#\n# Módulo 8 - Administrar Datos con Diccionarios\n#\n# Ejercicio 1\n\nplanet = {\n 'name' : 'Mars',\n 'moons' : 2\n}\n\nprint(f'{planet[\"name\"]} has {planet[\"moons\"]} moons.')\n\nplanet['circunference (km)'] = {\n 'polar': 6752,\n 'equatorial': 6792\n}\n\nprint(f'{planet[\"name\"]} polar circunference of: {planet[\"circunference (km)\"][\"polar\"]} km. And a equatorial circunference of: {planet[\"circunference (km)\"][\"equatorial\"]} km.')\n\n\n# Ejercicio 2\n\nplanet_moons = {\n 'mercury': 0,\n 'venus': 0,\n 'earth': 1,\n 'mars': 2,\n 'jupiter': 79,\n 'saturn': 82,\n 'uranus': 27,\n 'neptune': 14,\n 'pluto': 5,\n 'haumea': 2,\n 'makemake': 1,\n 'eris': 1\n}\n\n# +\ntotal_moons = 0\n\nfor value in planet_moons.values():\n total_moons = total_moons + value\nprint(f'There is a total of {total_moons} in the solar system.')\nprint(f'That is an average of {total_moons/len(planet_moons.keys())}')\n","repo_name":"ErickViveros/KatasLaunchX","sub_path":"Kata_08.ipynb","file_name":"Kata_08.ipynb","file_ext":"py","file_size_in_byte":919,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"4007700494","text":"# # Stage 4: Advanced Analysis\n# - Sentiment Analysis\n# - Text Classification\n\n# ### Importing Libraries\n\n# +\nimport pandas as pd\nfrom tqdm import tqdm\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\nfrom transformers import pipeline\n# -\n\n# ### Files Import\n\n# +\ndata = pd.read_pickle('data_clean_summarized.pkl')\ndata.reset_index(drop=True, inplace=True)\n\ndata\n\n# +\n# from Amazon website (https://www.amazon.com/AmazonBasics-Microwave-Small-Works-Alexa/dp/B07894S727)\nspecs = pd.read_csv(\"specs.csv\")\nspecifications = list(specs[\"Specifications\"]) # extract specifications headers from specs.csv\n\n# from Mr Right Ideas website (https://www.mrright.in/ideas/appliances/microwave/microwave-oven-components-and-their-functions/)\ncomps = pd.read_csv(\"comps.csv\")\ncomponents = list(dict.fromkeys(comps[\"Components\"])) # extract component headers from comps.csv\ncomps_renamed = list(dict.fromkeys(comps[\"Comps_renamed\"])) # extract renamed components from comps.csv\n\n# from wiki_scraper.py\nwiki_adjusted = pd.read_csv(\"wiki_adjusted.csv\")\nwiki_adjusted = list(wiki_adjusted['Keywords']) # extract sustainability keywords from wiki_adjusted.csv\n\nothers = ['Packaging', 'Delivery']\nspecifications += others\n\ndata['comps_mention'] = 'na'\n# -\n\n# ### Components Mentioned in Reviews\n# - With Sentiment Analysis\n\n# +\nfor i in range(data.shape[0]):\n for j in comps_renamed:\n if j in data['text'][i]:\n data['comps_mention'][i] = j\n\ndata\n\n# +\n# Create quick lambda functions to find the polarity and subjectivity of each routine\nfrom textblob import TextBlob\n\ncount_result = []\nfor comps in comps_renamed:\n count = [data[data.comps_mention == comps].shape[0]]\n count_result += count\n\npol = lambda x: TextBlob(x).sentiment.polarity\ndata['polarity'] = data['summary'].apply(pol)\n\npol_count = lambda x: 1 if x > 0 else -1\ndata['pol_count'] = data['polarity'].apply(pol_count)\ndisplay(data)\n\nsentiment_list = []\nfor comps in comps_renamed:\n data_count = data[data.comps_mention == comps]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': comps_renamed,\n 'values': count_result,\n 'sentiments': sentiment_list}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n\n# +\n# Data check\n\ndata_selected = data[data.comps_mention == df_chart.labels[2]] # check number accordingly\ndata_selected.reset_index(drop=True, inplace=True)\n\ndata_selected\n# -\n\n# ### Sustainability Mentioned in Reviews\n# - With Sentiment Analysis\n\n# +\ndata['sus_mention'] = 'na'\nwiki_adjusted = ['waste', 'eneryg', 'environmental', 'material']\nfor i in range(data.shape[0]):\n for j in wiki_adjusted:\n if j in data['text'][i]:\n data['sus_mention'][i] = j\n\ndata\n\n# +\n# Create quick lambda functions to find the polarity and subjectivity of each routine\nfrom textblob import TextBlob\n\ncount_result = []\nfor sus in wiki_adjusted:\n count = [data[data.sus_mention == sus].shape[0]]\n count_result += count\n\npol = lambda x: TextBlob(x).sentiment.polarity\ndata['polarity'] = data['summary'].apply(pol)\n\npol_count = lambda x: 1 if x > 0 else -1\ndata['pol_count'] = data['polarity'].apply(pol_count)\ndisplay(data)\n\nsentiment_list = []\nfor sus in wiki_adjusted:\n data_count = data[data.sus_mention == sus]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': wiki_adjusted,\n 'values': count_result,\n 'sentiments': sentiment_list}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n\n# +\n# Data check\n\ndata_selected = data[data.comps_mention == df_chart.labels[2]] # check number accordingly\ndata_selected.reset_index(drop=True, inplace=True)\n\ndata_selected\n# -\n\n# ## Zero-Shot-Classification\n\nclassifier = pipeline(\"zero-shot-classification\", device=0)\n\n# ### Specification Categorization\n# - With Sentiment Analysis \n\n# +\ndisplay(specifications)\n#Applying it into the dataframe base on original review\ndata['category'] = data['text'].apply(lambda x: classifier(x, candidate_labels=specifications))\n\ndata\n# -\n\n#Replacing the category with the top matched category\nfor i in tqdm(range(data.shape[0])):\n if data['category'][i] not in specifications:\n data['category'][i] = data['category'][i].get('labels')[0]\n\ndata_cat = data.copy()\n\n# +\ncandidate_results = list(0 for i in range(len(specifications)))\n\n\ndef count(category):\n for i in range(len(specifications)):\n if category == specifications[i]:\n candidate_results[i] = candidate_results[i] + 1\n\n\n# +\n# Run the counting function\ndata_cat['category'].apply(lambda x: count(x))\n\ncandidate_results\n\n\n# +\nsentiment_list_func = []\nfor func in specifications:\n data_count = data[data.category == func]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list_func += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': specifications,\n 'values': candidate_results,\n 'sentiments': sentiment_list_func}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n# -\n\n# ### Sustainability Categorization \n# - Sentiment Analysis included\n\n# +\n#Categorying in terms of sustainability\n\n#Applying it into the dataframe base on original review\ndata['sus_category'] = data['text'].apply(lambda x: classifier(x, candidate_labels=wiki_adjusted))\n\ndata\n# -\n\n#Replacing the category with the top matched category\nfor i in tqdm(range(data.shape[0])):\n if data['sus_category'][i] not in wiki_adjusted:\n data['sus_category'][i] = data['sus_category'][i].get('labels')[0]\n\n# +\ndata_cat = data.copy()\n\ncandidate_results = list(0 for i in range(len(wiki_adjusted)))\n\n\ndef count(category):\n for i in range(len(wiki_adjusted)):\n if category == wiki_adjusted[i]:\n candidate_results[i] = candidate_results[i] + 1\n\n\n# +\n# Run the counting function\ndata_cat['sus_category'].apply(lambda x: count(x))\n\ncandidate_results\n\n\n# +\nsentiment_list_func = []\nfor func in wiki_adjusted:\n data_count = data[data.category == func]\n sen_ana = data_count['pol_count'].sum()\n if sen_ana > 0:\n sen_ana = 'positive'\n else:\n sen_ana = 'negative'\n sentiment_list_func += [sen_ana]\n # df_chart['sentiments'] = [df_chart['labels'] == comps]\n\n# data['pol_count'].sum()\n\nchart = {'labels': wiki_adjusted,\n 'values': candidate_results,\n 'sentiments': sentiment_list_func}\n\ndf_chart = pd.DataFrame(chart, columns=['labels','values', 'sentiments'])\n\ndf_chart\n\n# +\ndata.to_pickle('data_analysed.pkl')\n\ndata\n# -\n\n\n","repo_name":"jiasenn/text-analysis","sub_path":"4.Advanced-Analysis.ipynb","file_name":"4.Advanced-Analysis.ipynb","file_ext":"py","file_size_in_byte":7104,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"32876589411","text":"# \n\n# # Final Project \n\n# ## Задание 1. A/B–тестирование\n\n# Одной из основных задач аналитика в нашей команде является корректное проведение экспериментов. Для этого мы применяем метод A/B–тестирования. В ходе тестирования одной гипотезы целевой группе была предложена новая механика оплаты услуг на сайте, у контрольной группы оставалась базовая механика. В качестве задания Вам необходимо проанализировать итоги эксперимента и сделать вывод, стоит ли запускать новую механику оплаты на всех пользователей.\n#\n# Предлагаем Вам ответить на следующие вопросы:\n#\n# - На какие метрики Вы смотрите в ходе анализа и почему?\n# - Имеются ли различия в показателях и с чем они могут быть связаны?\n# - Являются ли эти различия статистически значимыми?\n# - Стоит ли запускать новую механику на всех пользователей?\n#\n\n# ## В качестве входных данных Вы имеете 4 csv-файла:\n#\n# - groups.csv - файл с информацией о принадлежности пользователя к контрольной или экспериментальной группе (А – контроль, B – целевая группа) \n# - groups_add.csv - дополнительный файл с пользователями, который вам прислали спустя 2 дня после передачи данных\n# - active_studs.csv - файл с информацией о пользователях, которые зашли на платформу в дни проведения эксперимента. \n# - checks.csv - файл с информацией об оплатах пользователей в дни проведения эксперимента. \n\n# ____\n\n# +\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nfrom scipy.stats import chi2_contingency, ttest_ind, levene, normaltest, mannwhitneyu, norm\nfrom tqdm.auto import tqdm\nimport pandahouse as ph\n\nsns.set(\n font_scale=1.5,\n style=\"whitegrid\",\n rc={'figure.figsize':(16,8)}\n )\n# -\n\n# #### Загрузка и первичный анализ данных:\n\ngroups = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_groups.csv', sep=';')\ngroups.head()\n\nprint(f'Кол-во строк: {groups.shape[0]}')\nprint(f'Кол-во дубликатов: {groups.duplicated().sum()}')\nprint(f'Кол-во уникальных id: {groups.id.nunique()}')\n\ngroups.isna().sum() # проверка на пропущенные значения \n\ngroups_add = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_group_add.csv', sep=',')\ngroups_add.head()\n\nprint(f'Кол-во строк: {groups_add.shape[0]}')\nprint(f'Кол-во дубликатов: {groups_add.duplicated().sum()}')\nprint(f'Кол-во уникальных id: {groups_add.id.nunique()}')\n\n# Объединим данные в один фрейм:\n\ngroups_total = pd.concat([groups,groups_add], sort=False,ignore_index=True)\ngroups_total\n\ngroups_total.duplicated().sum() # прверяем что не появилось дублей \n\n# поменяем назваие стольбца для удобства дальнейшего объединения таблиц\ngroups_total.rename(columns={'id':'student_id'}, inplace=True) \ngroups_total.head()\n\n# Посмотрим на распределение пользователей по группам:\n\ngroups_total.grp.value_counts()\n\nsns.countplot(data=groups_total, x='grp');\n\n# Распределение оказалось крайне не равномерным, что само по себе не является чем-то неправильным или критичным для A/B теста.\n# Дело в том, что обычно тестовая группа бывает меньше, в виду того, что катить новую фичу на большое кол-во пользователей рисковано и дорого, а в наших данных тестовая группа более чем в 3 раза больше.\n\n# ____\n\n# пользователи зашедшие на платфому в дни проведения эксперимента\nactive_studs = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_active_studs.csv')\nactive_studs.head()\n\nprint(f'Кол-во дубликатов:{active_studs.duplicated().sum()}')\n\nprint(f'Кол-во пользователей зашедших на платформу во время эксперимента:{active_studs.shape[0]}')\n\n# Кажется логичным добавить колонку, которая будет как-то выделять пользователей которые заходили на платформу/сайт во время эксперимента, при дальнейшем объединении таблиц.Так и поступим. И судя по колнке student_id мы имеем дело с образовательной платформой.\n\nactive_studs['active'] = 1\nactive_studs.head()\n\n# данные об оплатах в дни проведеня эксперимента\nchecks = pd.read_csv('/mnt/HC_Volume_18315164/home-jupyter/jupyter-a-kuchiev/F/Проект_2_checks.csv',sep=';')\nchecks.head()\n\nchecks.isna().sum() # проверка на пропущенные значения\n\nprint(f'Кол-во строк: {checks.shape[0]}')\nprint(f'Кол-во дубликатов: {checks.duplicated().sum()}')\nprint(f'Кол-во уникальных id: {checks.student_id.nunique()}')\n\nchecks.rev.describe()# посмортим немного описательной статистки по платежам \n\n# - Максимальный платеж - 4650\n# - Минимальный платеж - 199\n# - Медианный - 840 \n\nchecks.rev.value_counts().head(10) # распределение по платежам \n\nchecks.rev.plot(kind='hist', bins=25);\n\n# Распределение платежей не равномерное, имеется 3 моды(платежи по 1900, 290, 690).\n# Но это без привязки к группам, а нас интересуют различия как раз между группми.\n\n# ____\n\n# Предварительно можно сделать следующие выводы:\n# - данные очень \"чистые\", без пропусков и нужного типа\n# - таблица с активными пользователми насчитывает 8341 записи, тогда как общая таблица 74576 записи.\n# Исходя из условия задания, если пользователь не заходил на платформу/сайт во время эксперимента, то и увидеть/ощутить \n# изменения механики оплат на себе он не мог. Из этого следует, что анализировать и сранивать нужно только активн��х пользователей.\n\n# Соединим наши таблицы в одну.\n# Кажется логичным соединять таблицу активных пользователей с общей с помощью inner джойна, чтобы остались толкько те, кто нам нужен, но сделаем все обьединения через left чтобы пока что не потерять пользоватлей и покрутить данные еще, отфильтровать нужное всегда успеем.\n\n# делаем мердж и заполняем пропуски 0, так у нас 0 помечены те, кто не заходил во время эксперимента\nfinal_df = groups_total.merge(active_studs, how='left', on = 'student_id').fillna(0)\nfinal_df.head()\n\n# ____\n\n# Теперь присоединим таблицу с оплатами, но перед этим тоже создадим столбец для того чтобы выделить в общем фрейме тех, кто платил.\n\nchecks['has_rev'] = 1\nchecks.head()\n\nfinal_df = final_df.merge(checks, how='left', on='student_id').fillna(0)\nfinal_df.head()\n\n# Покрутим/проанализируем получившийся датафрейм\n\nprint(f'Число активных пользователей: {final_df.query(\"active == 1\").shape[0]}')\nprint(f'Число активных и сделавших покупку пользователей: {final_df.query(\"active == 1 & has_rev == 1\").shape[0]}')\nprint(f'Число активных и не сделавших покупку пользователей: {final_df.query(\"active == 1 & has_rev == 0\").shape[0]}')\nprint(f'Число неактивных и сделавших покупку пользователей: {final_df.query(\"active == 0 & has_rev == 1\").shape[0]}')\nprint(f'Число неактивных и не сделавших покупку пользователей: {final_df.query(\"active == 0 & has_rev == 0\").shape[0]}')\n\n# ____\n\n# Стоить прокомментировать 149 пользователей заплативших не заходя на сайт/платформу:\n# - можно проедположить что имеют место автоплатежи\n# - ошибка системы сплитования \n# - ошибки в логфайлах \n#\n# Утверждать с уверенностью, не имея дополнительных данных о том, как долго продолжался А/В тест,данные о времени захода пользователя и.т.д, мы не можем.\n# Следуя условию задачи, далее будем продаолжать анализ только с теми пользователми на которых проводился тест.\n\nexp_data = final_df.query(\"active == 1\")\nexp_data.head()\n\nexp_data.grp.value_counts()\n\n# Cреди активных пользователей так же наблюдаем дисбаланс по группам.\n\n# Посмотрим на распределение платежей:\n\nsns.histplot(data = exp_data, x = 'rev', hue = 'grp', bins= 30);\n\n# График оказался не очень информативным в силу того, что подавляющее большинство пользователей заходивших во время эксперимента не сделало покупку.\n#\n# Построим распредление трат только тех пользователей чей revenue > 0:\n\nsns.histplot(data = exp_data.query('rev > 0'), x = 'rev', hue = 'grp', bins= 25);\n\n# Глядя на график можно заметить, что наши распределения отличаются только пиком в группе В в районе rev = ~1900.\n#\n# Проверим цифрами:\n\nexp_data.query(\"rev > 0\")\\\n .groupby(['grp','rev'],as_index=False)\\\n .agg({'student_id':'count'})\\\n .rename(columns={'student_id':'num_checks'})\\\n .pivot(columns='grp', values='num_checks', index='rev')\\\n .sort_values(by='B', ascending=False)\\\n .fillna(0)\\\n .head(10)\n\n# В группе В 128 пользователей совершили платеж на 1900, тогда как в группе А таких не оказалось вовсе.\n\n# _____\n\n# #### Так как мы тестировали влияние новой механики оплаты, и исходя из имеющихся данных, нашими метриками будут:\n# - конверсия(CR) \n# - средний доход с пользователя (ARPU) \n# - средний доход с платящего пользователя (ARPPU).\n\ngrpA = exp_data.loc[exp_data.grp=='A']\ngrpA\n\ngrpB = exp_data.loc[exp_data.grp=='B']\ngrpB\n\n# #### Conversion Rate\n\ncr = exp_data.groupby('grp', as_index=False)\\\n .agg({'has_rev':'mean'})\ncr\n\n# В тестовой группе конверсия уменьшилась. Необходимо удостовериться в том, является ли это различие стат. значимо.\n# Для этого используем критерий согласия Пирсона или иначе хи квадрат.\n#\n# Сформулируем гипотезы:\n# \n# H0 - конверсии в двух группах не отличаются\n# \n# H1 - конверсии в двух группах отличаются\n\n# +\nstat, p, dof, expected = chi2_contingency(pd.crosstab(exp_data.grp, exp_data.has_rev))\n\nprint(p)\nif p > 0.05:\n print('Оснований для отклонения нулевой гипотезы у нас нет.')\nelif p < 0.05:\n print('Отклоняем нулевую гипотезу.')\nelse:\n print('Необходимо перестмотерть p-value')\n# -\n\n# ____\n\n# #### ARPU - Avarage Revenue Per User\n\nnp.mean(grpA.rev), np.mean(grpB.rev)\n\nlen(grpB.rev), len(grpA.rev) # размер выборок\n\n# Cредний доход с пользователя в двух группах отличается.\n# Необходимо подтвердить или отвергнуть это с помощью стат. тестов.\n#\n# Н0 - средний доход с пользователя в двух группах не отличается\n#\n# Н1 - средний доход с пользователя в двух группах отличается\n#\n# В нашем распоряжении критерий Стьюдента, его непараметрический аналог - критерий Манна-Уитни и бустрап.\n#\n# Критерий Стьюдента имеет ряд необходимых допущений: случайность выборки, ее репрезентативность, и нормалььное распределение, а так же гомогенность дисперсий. Последние два являются не строгими. При не нермальном распределении необходимо чтобы выборки были достаточно большими, а при различии в дисперсиях можно воспользоваться поправкой Уэлча при использовании t-критерия.\n# Так же можно прологорифмировать наши выбрки, но данная операция может навредить при принятии решений бизнесса, так как отличия в логорифмированных данных не то же самое что при сравнении обычный средних.И в наших данных при подсчете ARPU это невозможно так как есть большое количество записей с 0 оплатой, а как известно логорифм от 0 это бесконечность).\n#\n#\n# Критерий Манна-Уитни хорош когда распределение признака не нормально, но он плохо интерпритируем и так же чувсителен при наличии в данных большого кол-ва повторяющихся знаяений, а это как раз наш случай. \n#\n# Бустрап же менее чувствителен ко всем вышеизложенным требованиям но является 'тяжелым'.\n#\n# Я буду использовать кртерий стьюдента и бутсрап.\n\n# Из графиков выше было очевидно, что распределение не нормальное, но следует подтвердить это:\n\nprint(normaltest(grpA.rev))\nprint(normaltest(grpB.rev))\n\n# С помощью теста Левена, проверим различаются ли дисперсии:\n\nlevene(grpB.rev,grpA.rev)\n\n# p-value равный 0.22 не дает нам оснований отклонить гипотезу о равенстве дисперсий.\n\nttest_ind(grpA.rev, grpB.rev)\n\n# t-test на наших данных показал p-value > 0.05 , что не дает нам оснований отклонить нулевую гипотезу.\n\nmannwhitneyu(grpA.rev, grpB.rev)\n\n# Манн-Уитни так же показал уровень значимости выше 0.05.\n\n# Но давайте посмотрим что произойдет, если мы будем многократно извлекать средние, складывать их в массив, затем посторим распределение этих средних и применим стат тесты:\n\nmean_rev_grpA_1000 = []\nfor i in range(1000):\n mean_rev_grpA_1000.append(\n np.mean(grpA.rev.sample(frac=1,replace=True))\n )\n\nplt.hist(mean_rev_grpA_1000);\n\nmean_rev_grpB_1000 = []\nfor i in range(1000):\n mean_rev_grpB_1000.append(\n np.mean(grpB.rev.sample(frac=1,replace=True))\n )\n\nplt.hist(mean_rev_grpB_1000);\n\n# ЦПТ во всей красе!)\n\nttest_ind(mean_rev_grpA_1000,mean_rev_grpB_1000)\n\n\n# А вот средние из многократных выборок средних уже отличаются стат значимо!\n\n# Теперь напишем функцию бутсрапа, котрая будет извлекать из наших групп подвыборки, сравнивать разницу средних в этих выборках и класть эту разницу в массив и затем строить распределение разницы средних:\n\ndef get_bootstrap(\n data_column_1, # числовые значения первой выборки\n data_column_2, # числовые значения второй выборки\n boot_it = 1000, # количество бутстрэп-подвыборок\n statistic = np.mean, # интересующая нас статистика\n bootstrap_conf_level = 0.95 # уровень значимости\n):\n boot_len = max([len(data_column_1), len(data_column_2)])\n boot_data = []\n for i in tqdm(range(boot_it)): # извлекаем подвыборки\n samples_1 = data_column_1.sample(\n boot_len, \n replace = True # параметр возвращения\n ).values\n \n samples_2 = data_column_2.sample(\n boot_len, # чтобы сохранить дисперсию, берем такой же размер выборки\n replace = True\n ).values\n \n boot_data.append(statistic(samples_1-samples_2)) \n pd_boot_data = pd.DataFrame(boot_data)\n \n left_quant = (1 - bootstrap_conf_level)/2\n right_quant = 1 - (1 - bootstrap_conf_level) / 2\n quants = pd_boot_data.quantile([left_quant, right_quant])\n \n p_1 = norm.cdf(\n x = 0, \n loc = np.mean(boot_data), \n scale = np.std(boot_data)\n )\n p_2 = norm.cdf(\n x = 0, \n loc = -np.mean(boot_data), \n scale = np.std(boot_data)\n )\n p_value = min(p_1, p_2) * 2\n \n # Визуализация\n _, _, bars = plt.hist(pd_boot_data[0], bins = 50)\n for bar in bars:\n if abs(bar.get_x()) <= quants.iloc[0][0] or abs(bar.get_x()) >= quants.iloc[1][0]:\n bar.set_facecolor('red')\n else: \n bar.set_facecolor('grey')\n bar.set_edgecolor('black')\n \n plt.style.use('ggplot')\n plt.vlines(quants,ymin=0,ymax=50,linestyle='--')\n plt.xlabel('boot_data')\n plt.ylabel('frequency')\n plt.title(\"Histogram of boot_data\")\n plt.show()\n \n return {#\"boot_data\": boot_data, \n \"quants\": quants, \n \"p_value\": p_value}\n\n\nget_bootstrap(grpB.rev, grpA.rev)\n\n# Результат приминения бустрапа так же говорит нам о стат значимом различии среднего дохода с пользователя в двух группах.\n# На графике видно, что ноль не входит в доверительный интервал и p-value < 0.05. Хотя стоит отметить что p-value очень близка к 0.05. \n\n# _____\n\n# #### ARPPU - Avarage Revenue Per Paying User\n\n# Теперь анализируем только платящих пользователей.\n#\n# Данных еще меньше.\n# Гипотезы все те же:\n#\n# Н0 - средний доход с платящего пользователя в группах не различается\n#\n# Н1 - средний доход с платящего пользователя в группах различается\n#\n# Так как данных еще меньше, то выбор бустрапа будет еще более оправдан, но посмотрим на другие критерии тоже.\n\nlen(grpA.query('rev>0'))\n\nlen(grpB.query('rev>0'))\n\nttest_ind(grpA.query('rev>0').rev, grpB.query('rev>0').rev)\n\n# t-test дает нам основания для отклонения нулевой гипотезы.\n\nmannwhitneyu(grpA.query('rev>0').rev, grpB.query('rev>0').rev)\n\nget_bootstrap(grpA.query('rev>0').rev, grpB.query('rev>0').rev)\n\n# Все три теста гвоврят нам о том что мы должны отклонить нулевую гипотезу.\n\n# ### Выводы:\n#\n# 1. Конверсия в тестовой группе снизилась, но статистически это изменение оказалось не значимым.\n# Однако сам факт этого негативен, так как мы оценивали влияние влияние новой механики оплаты и по идее она должны была делать процесс покупки легче/удобнее и следовательно вести к увеличению конверсии.\n# 2. Средний доход с пользователя вырос, однако стат тесты показали разные результаты:\n# т-тест и манн-уитни показали что различия не значимы, тогда как бустрап показал значимые различия, но с р-value близким к пороговому значению принятия решения. Я думаю что дело в том, что в данных подавляющее большинство записей где сумма равна нулю. \n# 3. Средний доход с платящих пользователей так же вырос в тестовой группе и все тесты показали, что различия значимы.\n# 4. Так же хочется отметить что по сути единственное отличие в распределениях оплат в двух группах заключается в том, что в тестовой группе имеется пик оплат на сумму в 1900. Тут надо подумать как изменения механики оплаты может вести к тому, что пользователи наснут тратить определенную сумму. Самый плохой вариант это если во время проведения эксперимента проводилась еще какая нибудь акция и при сплитовании пользователей получилось так, что пользователи в тестовой группе видели не только изменение в механике оплаты, но и акцию, в таком случае наш А/В тест полностью не валиден. \n# 5. Однако с другой стороны, так как у нас образовательная платформа, и если представить что раньше у нас поекпались все курсы по отдельности, а теперь появилась возможность обьединять курсы в специальсности и брать их вместе платя определенную сумму, и если это считать изменением механики оплаты, то результаты теста показыват что результат положительный.\n#\n# Подводя итог скажу что катить новую механику оплаты(чтобы под этим не подразумевалось) на всех, я бы пока не стал.\n# Собрал бы еще данные, проверил правильно сплитуются пользователи на группы, продлил бы время проведения А/В теста и убедился что мы тестируем влияние какого то одного эффекта, а не новой мехаики и акций и прочего.\n#\n#\n\n# ____\n\n# ## Задание 2. SQL\n\n# Все данные находятся в табличном виде в Clickhouse.\n\n# #### 2.1 Очень усердные ученики.\n#\n# 2.1.1 Условие\n#\n# Образовательные курсы состоят из различных уроков, каждый из которых состоит из нескольких маленьких заданий. Каждое такое маленькое задание называется \"горошиной\".\n#\n# Назовём очень усердным учеником того пользователя, который хотя бы раз за текущий месяц правильно решил 20 горошин.\n\n# Подключаемся к базе данных:\n\nconnection = {'host': 'http://clickhouse.beslan.pro:8080',\n 'database':'default',\n 'user':'student', \n 'password':'dpo_python_2020'\n }\n\n# Посмотрим на данные хранящиеся в таблице:\n\nq_1 = '''\n SELECT\n st_id,\n toDateTime(timest) as timest,\n correct, \n subject\n \n FROM \n default.peas\n LIMIT 10\n \n '''\nquery_1 = ph.read_clickhouse(query=q_1, connection=connection)\nquery_1\n\n# Cмотрим за какой период времени данные в тадлице:\n\nq_2 = '''\n SELECT\n min(toDateTime(timest)) as first_date,\n max(toDateTime(timest)) as last_date\n FROM \n default.peas\n \n '''\nquery_2 = ph.read_clickhouse(query=q_2, connection=connection)\nquery_2\n\n# Считаем усердных учеников:\n\nq_3 = '''\n SELECT\n COUNT(DISTINCT st_id) as good_students\n FROM (\n SELECT\n st_id,\n SUM(correct) as corr_tasks\n FROM \n default.peas\n GROUP BY\n st_id\n HAVING\n corr_tasks >= 20\n ) \n '''\nquery_3 = ph.read_clickhouse(query=q_3, connection=connection)\nquery_3\n\n# Ну очевидно, что если в условии задачи под текущим месяцем подразумевается ноябрь 2022, то таких студентов в таблице не обнаружится.\n\n# ____\n\n# #### 2.2 Оптимизация воронки\n#\n# 2.2.1 Условие\n#\n# Образовательная платформа предлагает пройти студентам курсы по модели trial: студент может решить бесплатно лишь 30 горошин в день. Для неограниченного количества заданий в определенной дисциплине студенту необходимо приобрести полный доступ. Команда провела эксперимент, где был протестирован новый экран оплаты.\n\n# \n#\n\n# Необходимо в одном запросе выгрузить следующую информацию о группах пользователей:\n#\n# - ARPU \n# - ARPAU \n# - CR в покупку \n# - СR активного пользователя в покупку \n# - CR пользователя из активности по математике (subject = ’math’) в покупку курса по математике\n# - ARPU считается относительно всех пользователей, попавших в группы.\n#\n# Активным считается пользователь, за все время решивший больше 10 задач правильно в любых дисциплинах.\n#\n# Активным по математике считается пользователь, за все время решивший 2 или больше задач правильно по математике.\n\nq_4 = '''\n\n SELECT \n test_grp as grp,\n round(SUM(fcp.money)/uniqExact(s.st_id),2) as ARPU,\n round(SUMIf(fcp.money, query_in.act > 10)/uniqIf(s.st_id, query_in.act > 10),2) as ARPAU,\n round(uniqIf(s.st_id, fcp.money > 0)/uniqExact(s.st_id)*100, 2) as CR,\n round(uniqIf(s.st_id, query_in.act > 10 and fcp.money > 0)/uniqIf(s.st_id, query_in.act > 10)*100,2) as CR_active,\n round(uniqIf(s.st_id, query_in.math_active >= 2 and fcp.subject = 'Math' and fcp.money > 0)*100 /\n uniqIf(s.st_id, query_in.math_active >= 2), 2) as CR_math\n FROM \n studs s\n LEFT JOIN final_project_check fcp\n ON s.st_id = fcp.st_id\n LEFT JOIN (\n SELECT \n st_id, \n SUM(correct) as act, \n SUMIf(correct, subject = 'Math') as math_active\n FROM \n peas\n GROUP BY \n st_id) as query_in \n ON s.st_id = query_in.st_id\n\n GROUP BY test_grp\n\n'''\nquery_4 = ph.read_clickhouse(query=q_4, connection=connection)\nquery_4\n\n\n# ____\n\n# ## Задание 3. Python\n#\n# #### 3.1 Задача\n#\n# - Реализуйте функцию, которая будет автоматически подгружать информацию из дополнительного файла groups_add.csv (заголовки могут отличаться) и на основании дополнительных параметров пересчитывать метрики.\n#\n# - Реализуйте функцию, которая будет строить графики по получаемым метрикам.\n\ndef get_data(df1: 'DataFrame', df2: 'DataFrame', df3: 'DataFrame', df4: 'DataFrame') -> 'DataFrame':\n '''\n Функция get_data принимает в качестве атрибутов \n 4 датафрейма соединяет их рассчитавет метрики\n и подает на выход датафрейм с метриками.\n \n '''\n df1.columns = df2.columns\n all_groups = pd.concat([df1, df2])\n all_groups.rename(columns={'id':'student_id'}, inplace=True)\n df3['active'] = 1\n df4['has_rev'] = 1\n final_df = all_groups.merge(df3, how='left', on='student_id').fillna(0)\n final_df = final_df.merge(df4, how='left', on='student_id').fillna(0)\n final_df = final_df.query('active == 1')\n metrics_df = final_df.groupby('grp', as_index=False)\\\n .agg({'student_id':'count','rev':['sum','mean'],'has_rev':['sum','mean'], })\n \n metrics_df.columns = ['grp','active_users','revenue','ARPU','paying_users','CR']\n #metrics_df = metrics_df.assign(paying_users = final_df.query('rev > 0').groupby('grp').agg({'rev': 'count'})) \n #metrics_df['ARPAU'] = (metrics_df.revenue / metrics_df.active_users)\n metrics_df['ARPPU'] = round(metrics_df.revenue / metrics_df.paying_users,2)\n metrics_df['ARPU'] = round(metrics_df['ARPU'],2)\n metrics_df['CR'] = round(metrics_df.CR*100,2)\n metrics_df['revenue'] = round(metrics_df.revenue,2)\n \n return metrics_df\n\n\nmetrics = get_data(groups,groups_add,active_studs,checks)\nmetrics\n\n\ndef show_metrics(metrics: 'DataFrame') -> 'graphs':\n '''\n Функция show_metrics принимает на вход один атрибут-датафрейм с метриками\n и возвращает графики данных метрик.\n \n '''\n \n plt.figure(figsize=(10, 6))\n splot=sns.barplot(x=metrics.grp,y=\"CR\",data=metrics)\n for p in splot.patches:\n splot.annotate(format(p.get_height(), '.1f'), \n (p.get_x() + p.get_width() / 2., p.get_height()), \n ha = 'center', va = 'center', \n xytext = (0, 9), \n textcoords = 'offset points')\n \n plt.figure(figsize=(10, 6))\n splot=sns.barplot(x=metrics.grp,y=\"ARPU\",data=metrics)\n for p in splot.patches:\n splot.annotate(format(p.get_height(), '.1f'), \n (p.get_x() + p.get_width() / 2., p.get_height()), \n ha = 'center', va = 'center', \n xytext = (0, 9), \n textcoords = 'offset points')\n\n plt.figure(figsize=(10, 6))\n splot=sns.barplot(x=metrics.grp,y=\"ARPPU\",data=metrics)\n for p in splot.patches:\n splot.annotate(format(p.get_height(), '.1f'), \n (p.get_x() + p.get_width() / 2., p.get_height()), \n ha = 'center', va = 'center', \n xytext = (0, 9), \n textcoords = 'offset points')\n \n\n \n return plt.show()\n\nshow_metrics(metrics)\n","repo_name":"AlexKuchiev/Karpov.Courses_projects","sub_path":"final.ipynb","file_name":"final.ipynb","file_ext":"py","file_size_in_byte":35093,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"31667184147","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#  \n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"V5Behma77t2H\" outputId=\"65cd96aa-a98f-4e61-aa3a-f10f3e00aa8b\"\ndef sol(N):\n S = list(map(int,input().split()))\n dp = S.copy()\n for i in range(1,N):\n Max = [0]\n for j in range(i):\n if S[i] > S[j]:\n Max.append(dp[j])\n dp[i] += max(Max)\n return max(dp)\n\nprint(sol(int(input())))\n","repo_name":"Kang-Yoonah/BaekJoon","sub_path":"Silver/11055.ipynb","file_name":"11055.ipynb","file_ext":"py","file_size_in_byte":656,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"18780268394","text":"# + id=\"5s_uheqcdmU5\" colab_type=\"code\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1788} outputId=\"e961f0cc-ff66-4e88-c487-ed99e5f760cb\" executionInfo={\"status\": \"ok\", \"timestamp\": 1538983182426, \"user_tz\": -480, \"elapsed\": 35829, \"user\": {\"displayName\": \"Justin Wu\", \"photoUrl\": \"\", \"userId\": \"12321085329230288327\"}}\nfrom bs4 import BeautifulSoup\nimport requests\nimport re\nimport os\nimport json\n\n\ndef get_soup(url,header):\n return BeautifulSoup(requests.get(url, headers=header).content,'html.parser')\n\nquery = input(\"query: \")# you can change the query for the image here\nimage_type=\"ActiOn\"\nquery= query.split()\nquery='+'.join(query)\nurl=\"https://www.google.co.in/search?q=\"+query+\"&source=lnms&tbm=isch\"\nprint('query_url: {}'.format(url))\n\n#add the directory for your image here\nDIR=\"Images\"\nheader={'User-Agent':\"Mozilla/5.0 (Windows NT 6.1; WOW64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/43.0.2357.134 Safari/537.36\"\n}\nsoup = get_soup(url,header)\n\n\nActualImages=[]# contains the link for Large original images, type of image\nfor a in soup.find_all(\"div\",{\"class\":\"rg_meta\"}):\n link , Type =json.loads(a.text)[\"ou\"] ,json.loads(a.text)[\"ity\"]\n ActualImages.append((link,Type))\n\nprint (\"Downloading total {} images\".format(len(ActualImages)))\n\nif not os.path.exists(DIR):\n os.mkdir(DIR)\nDIR = os.path.join(DIR, query.split()[0])\n\nif not os.path.exists(DIR):\n os.mkdir(DIR)\n \n###print images\nfor i , (img , Type) in enumerate( ActualImages):\n try:\n\n img_data = requests.get(img, stream=True).content\n cntr = len([i for i in os.listdir(DIR) if image_type in i]) + 1\n print (cntr, img)\n if len(Type)==0:\n f = open(os.path.join(DIR , image_type + \"_\"+ str(cntr)+\".jpg\"), 'wb')\n else :\n f = open(os.path.join(DIR , image_type + \"_\"+ str(cntr)+\".\"+Type), 'wb')\n\n\n f.write(img_data)\n f.close()\n except Exception as e:\n print (\"could not load : \"+img)\n print (e)\n","repo_name":"CloudMile/allianz_courses","sub_path":"Intro_to_ML_NLP_workshop/google _crawler.ipynb","file_name":"google _crawler.ipynb","file_ext":"py","file_size_in_byte":2007,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"1666058090","text":"# ### Entrenando un algoritmo de machine learning\n#\n# Para el presente entregable seleccioné un dataset obtenido del Instituto Nacional de Estadistica de Uruguay (INE).\n#\n# El mismo corresponde a la encuesta continua de hogares realizada en el año 2017.\n#\n# El objetivo del analisis sera trabajar con predicciones del ingreso en relación de dependencia de los encuestados.\n#\n# Archivos utilizados:\n#\n# P_2017_Terceros.sav -> el dataset que contiene los datos recopilados en la encuesta continua de hogares.\n# Dicccionario de Variables ECH 2017 -> funciona como anexo para determinar a que corresponde la codificación de cada columna del dataset.\n\nimport numpy as np\nimport pandas as pd\nfrom sklearn.model_selection import train_test_split\nfrom sklearn import linear_model\nfrom sklearn.metrics import mean_squared_error\nfrom sklearn.compose import make_column_transformer\nfrom sklearn.preprocessing import OneHotEncoder, StandardScaler\n\ndfP2017 = pd.read_spss(r'..\\\\data\\\\P_2017_Terceros.sav')\n\ncolumns = dfP2017.columns.values.tolist()\nprint(columns)\n\ndfP2017.head(2)\n\n# A continuación se definen las variables a utilizar en el analisis y se genera un dataset independiente para realizar modificaciones sin afectar al dataset original\n\n# +\ndf_ysos = pd.DataFrame()\n\n# variable de respuesta\ndf_ysos['ysos_dep'] = dfP2017['g126_1'] + dfP2017['g126_2'] + dfP2017['g126_3'] + dfP2017['g126_4'] \\\n + dfP2017['g134_1'] + dfP2017['g134_2'] + dfP2017['g134_3'] + dfP2017['g134_4']\n\n# caracteristicas personales\ndf_ysos['edad'] = dfP2017['e27']\ndf_ysos['sexo'] = dfP2017['e26']\ndf_ysos['estrato'] = dfP2017['estred13']\ndf_ysos['ascendencia_negra'] = dfP2017['e29_1']\ndf_ysos['estado_civil'] = dfP2017['e36']\ndf_ysos['asiste_comedor']=dfP2017['e559']\ndf_ysos['seguro_privado']=dfP2017['e45_3']\ndf_ysos['pobre']=dfP2017['pobre06']\ndf_ysos['indigente']=dfP2017['indigente06']\n\n## educación\ndf_ysos['anios_educ'] = dfP2017['e51_2'] + dfP2017['e51_3'] + dfP2017['e51_4'] + dfP2017['e51_5'] + dfP2017['e51_6'] \\\n + dfP2017['e51_7'] + dfP2017['e51_8'] + dfP2017['e51_9'] + dfP2017['e51_10'] + dfP2017['e51_11']\n\n## relación de empleo\ndf_ysos['antiguedad'] = dfP2017['f88_2']\ndf_ysos['categoria_ocup'] = dfP2017['f73']\ndf_ysos['dimension_empresa']=dfP2017['f77']\n\ndf_ysos.info()\n# -\n\n# ##### DETALLE DE VARIABLES A UTILIZAR\n#\n# ysos_dep -> ingresos dependientes de cada persona\n#\n# edad -> edad de la persona\n#\n# sexo -> sexo declarado de la persona\n#\n# estrato -> estrato social al que se asocia al encuestado\n#\n# ascendencia_negra -> dato booleano que indica si el encuestado tiene ascendencia \n# negra\n#\n# estado_civil -> estado civil declarado del encuestado\n#\n# asiste_comedor -> dato booleano que indica si el encuestado consume alguna de sus \n# comidas diarias en comedor público\n#\n# seguro_privado -> dato booleano que indica si el encuestado tiene asistencia medica \n# privada\n#\n# pobre -> dato que indica si el encuestado esta por debajo de la franja de pobreza establecida por el INE\n#\n# indigente -> dato que indica si el encuestado se clasifica como indigente\n#\n# anios_educ -> años de educación formal que tiene el encuestado\n#\n# antiguedad -> antigüedad laboral del encuestado\n#\n# categoria_ocup -> categoria ocupacional del encuestado\n#\n# dimension_empresa -> cantidad de trabajadores activos de la empresa a la cual pertenence el encuestado\n\n# genero archivo con datos a utilizar para alojar en Github, ya que el archivo .sav es muy pesado y no se puede guardar incluir al hacer commit a github.\nexport = '../data/ECH2017.xlsx'\ndf_ysos.to_excel(export, index=False)\n\ndf_ysos['dimension_empresa'].dtype\n\ndf_ysos.head(2)\n\n#se eliminan del dataset todos los ingresos menores a 0\ndf_ysos = df_ysos[df_ysos['ysos_dep'] > 0]\n\ndf_ysos = df_ysos.loc[(df_ysos['categoria_ocup'] == \"Asalariado/a privado/a\") | (df_ysos['categoria_ocup'] == \"Asalariado/a público/a\")]\n\n#transformación logaritmica de la variable ingresos\ndf_ysos['l_ysos_dep'] = np.log(df_ysos['ysos_dep'])\ndf_l_ysos = df_ysos.drop(columns=['ysos_dep'])\ndf_l_ysos.head(2)\n\n\n# A continuacón se realizan transformaciones a ciertas columnas para contar con la información agrupada de una forma que permita realizar analisis de forma más eficiente.\n\n# +\ndef map_estrato(estrato):\n if estrato in ['Montevideo Alto']:\n return 'Mdeo - Alto'\n elif estrato in ['Montevideo Medio', 'Montevideo Medio Bajo', 'Montevideo Medio Alto']:\n return 'Mdeo - Medio'\n elif estrato in ['Montevideo Bajo']:\n return 'Mdeo - Bajo'\n elif estrato in ['Zona Metropolitana']:\n return 'Metropolitana'\n elif estrato in ['Centro Norte', 'Centro Sur', 'Costa Este', 'Litoral Norte', 'Litoral Sur','Norte']:\n return 'Interior'\n\n# se crea nueva columna que contenga los nombres de los grupos\ndf_l_ysos['estrato'] = df_l_ysos['estrato'].apply(map_estrato)\n\n\n# +\ndef map_dimenisonempresa(dimension):\n if dimension in ['Una persona', 'De 2 a 4 personas', 'De 5 a 9 personas', 'De 10 a 19 personas']:\n return 'Empresa_chica'\n elif dimension in ['De 20 a 49 personas']:\n return 'Empresa_mediana'\n elif dimension =='De 50 personas o más':\n return 'Empresa_grande'\n elif dimension ==0.0:\n return 'N/A'\n\n# se crea nueva columna que contenga los nombres de los grupos\ndf_l_ysos['dimension_empresa'] = df_l_ysos['dimension_empresa'].apply(map_dimenisonempresa)\n# -\n\ndf_l_ysos.head(2)\n\n# +\n#Se agrupan las variables separandolas en categoricas y numericas.\n\ncategorical_columns = ['sexo', 'categoria_ocup','asiste_comedor','ascendencia_negra','estrato','seguro_privado', 'dimension_empresa']\nnumerical_columns = ['edad', 'anios_educ', 'antiguedad']\n\n# Se genera la función transformadora\ncolumn_transform = make_column_transformer(\n (OneHotEncoder(drop='if_binary'), categorical_columns), # cuando son dos categorías, elimina una\n (StandardScaler(), numerical_columns), # comentado mientras no se decida estandarizar\n remainder='passthrough', # deja el resto de las columnas sin transformar\n verbose_feature_names_out=False)\n\n# +\ndf_l_ysos_transf = pd.DataFrame(\n column_transform.fit_transform(df_l_ysos), \n columns=column_transform.get_feature_names_out()\n)\n\ndf_l_ysos_transf.head()\n# -\n\npredictoras = df_l_ysos_transf.drop('l_ysos_dep', axis=1)\n\ndf_l_ysos.head(2)\n\nX_train, X_test, y_train, y_test = train_test_split(predictoras,\n df_l_ysos_transf['l_ysos_dep'], \n test_size=0.20, \n random_state=1234)\n\n# #### Regresión Lineal\n#\n# A continuación se plantea el modelo propiamente dicho, para una primera instancia se trabaja solamente con la variable que toma los años de educación del encuestado.\n\nreg = linear_model.LinearRegression()\n\n# +\nmodelo_X_train = X_train[['anios_educ']]\nmodelo_X_test = X_test[['anios_educ']]\n\nmodelo = reg.fit(modelo_X_train, y_train)\n\n# +\n# predicción en train\nmodelo_y_train_pred = modelo.predict(modelo_X_train)\n\n# predicción en test\nmodelo_y_test_pred = modelo.predict(modelo_X_test)\n# -\n\nmodelo_mse_train = mean_squared_error(y_train, modelo_y_train_pred)\nprint(f\"Error cuadratico medio del modelo en train: \", modelo_mse_train)\n\nmodelo_mse_test = mean_squared_error(y_test, modelo_y_test_pred)\nprint(f\"Error cuadratico medio del modelo en test: \", modelo_mse_test)\n\n# ##### Actualización 31/05/2023\n#\n# Se agregan las variables 'edad' y 'antiguedad' con el objetivo de reducir los valores de error cuadratico medio del modelo tanto en train como en test.\n\n# +\n#variables_adicionales = ['edad']+['sexo']\n\nmodelo_X_train2 = X_train[['anios_educ', 'edad', 'antiguedad']]\nmodelo_X_test2 = X_test[['anios_educ', 'edad', 'antiguedad']]\n\nmodelo = reg.fit(modelo_X_train2, y_train)\n\n# +\n# predicción en train\nmodelo_y_train_pred2 = modelo.predict(modelo_X_train2)\n\n# predicción en test\nmodelo_y_test_pred2 = modelo.predict(modelo_X_test2)\n\n# -\n\nmodelo_mse_train2 = mean_squared_error(y_train, modelo_y_train_pred2)\nprint(f\"Error cuadratico medio del modelo en train: \", modelo_mse_train2)\n\nmodelo_mse_test2 = mean_squared_error(y_test, modelo_y_test_pred2)\nprint(f\"Error cuadratico medio del modelo en test: \", modelo_mse_test2)\n\n# Efectivamente, agregando las variables 'edad' y 'sexo' se generan mejoras en los resultados del modelo.\n","repo_name":"MartinMF88/Tarea-machine-learning","sub_path":"src/Desafío_EvaluaciónML_Martin_ Martinez.ipynb","file_name":"Desafío_EvaluaciónML_Martin_ Martinez.ipynb","file_ext":"py","file_size_in_byte":8406,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"15007352664","text":"# +\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport more_itertools\n\nfrom keras.models import Sequential\nfrom keras.layers import LSTM, Dense, Flatten, ConvLSTM2D\n\nfrom sklearn.metrics import mean_squared_error\n\n# -\n\n# # Preparação dos dados\n\n# Dividir uma sequência univariada em amostras\n\n# +\nX = np.linspace(0, 5*np.pi, 100, endpoint=True)\nC = np.cos(X)\n\n# transform numpy array to list\nC = C.tolist()\n\nplt.plot(X, C)\nplt.show()\n# -\n\nprint(C)\n\n\n# Podemos dividir a sequência em vários padrões de entrada / saída chamados amostras, onde três etapas de tempo são usadas como entrada e uma etapa de tempo é usada como saída para a previsão de uma etapa que está sendo aprendida.\n\n# +\n# Função para dividir uma sequência univariada em amostras para o método de janelas\n# X - amostras de entrada; y - variável que será predita\n\ndef split_sequence(sequence, n_steps):\n\tX, y = list(), list()\n\tfor i in range(len(sequence)):\n\t\tend_ix = i + n_steps\n\t\tif end_ix > len(sequence)-1:\n\t\t\tbreak\n\t\tseq_x, seq_y = sequence[i:end_ix], sequence[end_ix]\n\t\tX.append(seq_x)\n\t\ty.append(seq_y)\n\treturn np.array(X), np.array(y)\n\n# +\n# Escolha do numero de passos no tempo\nn_steps = 4\n\nX, y = split_sequence(C, n_steps)\n\nfor i in range(len(X)):\n\tprint(X[i], y[i])\n# -\n\n# ## Reshape\n\n# Primeiro, devemos remodelar o conjunto de dados de treinamento na estrutura esperada de [samples, timesteps] para [samples, timesteps, rows, cols, channels].\n\nX.shape\n\nn_features = 1 # série temporal univariada (uma variável de predição)\nn_seq = 2\nn_steps = 2\nX = X.reshape((X.shape[0], n_seq, 1, n_steps, n_features))\n\nX.shape\n\n# ## Divisão da série temporal em treino e teste\n\n# As 60 primeiras linhas como treino\nX_train = X[0:60]\n\n# O restante das linhas como teste\nX_test = X[60:]\n\n# Analogamente para a variável preditora\n\ny_train = y[0:60]\n\ny_test = y[60:]\n\n# # Modelos LSTM para séries temporais univariadas\n\n# Nesse problema de previsão de cosseno, tratamos de um caso de série temporal univariada. Sendo assim, podemos aplicar os seguintes modelos LSTM:\n#\n# * Vanilla LSTM\n# * Stacked LSTM\n# * Bidirectional LSTM\n# * CNN LSTM\n# * ConvLSTM\n\n# ## Convolutional LSTM - ConvLSTM\n\n# Uma extensão adicional da abordagem CNN-LSTM é realizar as convoluções do CNN (por exemplo, como o CNN lê os dados da sequência de entrada) como parte do LSTM para cada etapa de tempo.\n#\n# Essa combinação é chamada de LSTM convolucional ou, abreviadamente, ConvLSTM e, como o CNN-LSTM, também é usada para dados espaço-temporais.\n#\n# Ao contrário de um LSTM que lê os dados diretamente para calcular o estado interno e as transições de estado, e ao contrário do CNN-LSTM que está interpretando a saída dos modelos CNN, o ConvLSTM está usando convoluções diretamente como parte da leitura de entrada nas próprias unidades LSTM .\n#\n# A biblioteca Keras fornece a classe ConvLSTM2D que suporta o modelo ConvLSTM para dados 2D.\n\n# The ConvLSTM2D class, by default, expects input data to have the shape:\n\n# [samples, timesteps, rows, cols, features]\n\n# Onde cada etapa de tempo de dados é definida como uma imagem de pontos de dados (linhas * colunas).\n\n# ### Definindo o modelo ConvLSTM\n\n# Definindo o modelo\nmodel = Sequential()\nmodel.add(ConvLSTM2D(filters=64, kernel_size=(1,2), activation='relu', input_shape=(n_seq, 1, n_steps, n_features)))\nmodel.add(Flatten())\nmodel.add(Dense(1))\nmodel.compile(optimizer='adam', loss='mse')\n\n# fit model\nmodel_history = model.fit(X_train, y_train, epochs=1000, verbose=0)\n\nplt.plot(model_history.history['loss'])\nplt.title('Convolutional LSTM loss')\nplt.ylabel('loss')\nplt.xlabel('epoch')\nplt.legend(['train'], loc='upper right')\nplt.yscale('log')\nplt.show()\n\n# model prediction\nyhat = model.predict(X_test, verbose=0)\n\n#model evaluation\nmean_squared_error(y_test,yhat)\n\nplt.plot(y_test, label='test')\nplt.plot(yhat, label='Conv2D LSTM prediction')\nplt.legend()\n\n# # Observações\n#\n# - Imports: LSTM, Dense, Flatten, ConvLSTM2D\n# - Número de passos no tempo: 4\n# - Reshape a série temporal de [samples, timesteps] para [samples, timesteps, rows, columns, features]\n# - Treino realizado com 1000 epochs foi rápido, e o erro foi bem baixo, como pode ser visto na figura acima\n","repo_name":"bruna-capistrano/TrabalhoPPA","sub_path":"lstm/conv_lstm.ipynb","file_name":"conv_lstm.ipynb","file_ext":"py","file_size_in_byte":4247,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"32231656518","text":"import numpy as np\nfrom sklearn.datasets import load_iris\nfrom sklearn import tree\niris = load_iris()\n\n\nfor i in range(len(iris.target_names)):\n print(\"Target:\\t\", iris.target_names[i], \"\\tValue:\\t\", i)\n\nprint(\"Features: \", \", \".join(e for e in iris.feature_names))\n\nfor i in range(len(iris.target)):\n print(i, iris.data[i], iris.target[i])\n\ntest_idx = [0, 50, 100] #keep some testing data around\n\n#training data\ntrain_target = np.delete(iris.target, test_idx)\ntrain_data = np.delete(iris.data, test_idx, axis=0)\n\n#testing data\ntest_target = iris.target[test_idx]\ntest_data = iris.data[test_idx]\n\nclf = tree.DecisionTreeClassifier()\nclf.fit(train_data, train_target)\n\nprint(test_target)\nprint(clf.predict(test_data))\n\n#visualize here\nfrom sklearn.externals.six import StringIO\nfile = open('tree.dot', 'w+')\ntree.export_graphviz(clf,\n out_file=file,\n feature_names=iris.feature_names,\n class_names=iris.target_names,\n filled=True, rounded=True,\n impurity=False)\nfile.close()\n\n# # Visualize The Data\n# `$ xdot tree.dot`\n\n\n","repo_name":"Stuart4/Machine-Learning-Journey","sub_path":"lab2/Iris.ipynb","file_name":"Iris.ipynb","file_ext":"py","file_size_in_byte":1121,"program_lang":"python","lang":"en","doc_type":"code","stars":3,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"7672155754","text":"# !pip install --quiet --no-cache-dir h3ronpy\n\nimport h3\nfrom h3ronpy import raster\nimport rasterio as rio\nimport rasterio.plot\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom rasterstats import gen_zonal_stats, gen_point_query\nfrom rasterstats import zonal_stats\nimport pandas as pd\nimport geopandas as gpd\nimport h3\nimport json\nimport os\nfrom shapely.geometry import shape, mapping, box, Point, LinearRing, Polygon\n\n\n# +\ntest_raster = '../../data/raw/crop_data/default_pasture/CroplandPastureArea2000_Geotiff/Pasture2000_5m.tif'\n\ntest_area = (-10,40,0,50)\n\n# +\nwith rio.open(test_raster) as src:\n window = rio.windows.from_bounds(*test_area, src.transform)\n transform = rio.windows.transform(window, src.transform)\n print(src.profile)\n rio.plot.show(src.read(window=window, masked=True))\n gdf = raster.raster_to_geodataframe(src.read(1, window=window), transform, h3_resolution=4, nodata_value=src.profile['nodata'], compacted=False)\n\ngdf.plot('value')\n#gdf['h3index'] = gdf['h3index'].apply(hex)\ngdf.head()\n\n# +\ngeom = box(*test_area)\nidx = [h for h in h3.polyfill(mapping(geom), 4, geo_json_conformant=True)]\npts = [Point(h3.h3_to_geo(h)[::-1]) for h in idx]\nidx_int = [int(h, 16) for h in idx]\ngeoms = h3.h3_set_to_multi_polygon(idx, geo_json=True)\n\ndf = pd.DataFrame({'h3index':idx_int, 'value':gen_point_query(pts, test_raster, interpolate='nearest')})\ndf = df.dropna()\ndf.plot('value')\ndf.head()\n# -\n\nj = gdf.set_index('h3index').join(df.set_index('h3index'), rsuffix='t')\nj\n\n# %%timeit\npd.DataFrame({'h3index':idx_int, 'value':gen_point_query(pts, test_raster, interpolate='nearest')})\n\n# %%timeit\nwith rio.open(test_raster) as src:\n raster.to_dataframe(src.read(1, window=window), transform, h3_resolution=4, nodata_value=src.profile['nodata'], compacted=False)\n\n# +\ndfs = []\nwith rio.open(test_raster) as src:\n for ji, window in src.block_windows():\n transform = rio.windows.transform(window, src.transform)\n arr = src.read(1, window=window)\n \n df = h3ronpy.raster.raster_to_geodataframe(arr, transform, 4, nodata_value=src.profile['nodata'], compacted=False)\n dfs.append(df)\n\n\n# -\n\nl = [i for df in dfs for i in df['h3index']]\nprint(len(l))\nprint(len(set(l)))\n\npd.concat(dfs).plot('value')\n\n# +\nfrom math import ceil\nBLOCKSIZE = 512\ndef gen_raster_h3(raster_list, h3_res):\n \"\"\"Convert a list of identically formatted rasters to H3\n \n A function for efficiently turning a set of rasters into an H3 table.\n \n Takes a list of 1-band rasters with identical projection/transform.\n Reads each raster in blocks, and converts to h3 (nearest to centroid).\n Yields a dataframe with an h3index and one column for each raster's value.\n\n Args:\n raster_list: list of paths to rasters\n h3_res: h3 resolution to use for resampling\n \n Yields:\n A Pandas dataframe for each raster block (usu. 512x512) with an \n h3index and one column for each raster's value.\n \"\"\"\n readers = [rio.open(r) for r in raster_list]\n names = [os.path.splitext(os.path.basename(r))[0].lower() for r in raster_list]\n \n base = readers[0]\n for j in range(ceil(base.height/BLOCKSIZE)):\n for i in range(ceil(base.width/BLOCKSIZE)):\n window = rio.windows.Window(i*BLOCKSIZE, j*BLOCKSIZE, BLOCKSIZE, BLOCKSIZE)\n w_transform = rio.windows.transform(window, base.transform)\n dfs = []\n for src in readers:\n if src.transform != base.transform:\n raise ValueError(\"Transforms do not match\")\n arr = src.read(1, window=window)\n _df = raster.raster_to_dataframe(arr, w_transform, h3_res, nodata_value=src.profile['nodata'], compacted=False)\n dfs.append(_df.set_index('h3index')['value'])\n df = pd.concat(dfs, axis=1)\n print(f'Reading block {j}, {i}: h3index count {len(df)}')\n if len(df):\n df.columns = names\n # cast h3index from int64 to hex string\n yield df\n for src in readers:\n src.close()\n\n\n\n# +\ntest_list = [\n '../../data/raw/crop_data/default_pasture/CroplandPastureArea2000_Geotiff/Pasture2000_5m.tif',\n '../../data/raw/crop_data/default_pasture/CroplandPastureArea2000_Geotiff/Cropland2000_5m.tif'\n]\ntest_dir = '../../data/seed/mapspam/spam2017v2r1_ssa_prod'\ntest_list2 = [os.path.join(test_dir, f) for f in os.listdir(test_dir)]\n\nh3grid = pd.concat(list(gen_raster_h3(test_list2, 4)))\nh3grid = gpd.GeoDataFrame(h3grid, geometry=[Polygon(h3.h3_to_geo_boundary(h, geo_json=True)) for h in h3grid.index])\nh3grid.plot()\n# -\n\n\n","repo_name":"Vizzuality/landgriffon","sub_path":"data/notebooks/Lab/0_6_test_h3ronpy_fg.ipynb","file_name":"0_6_test_h3ronpy_fg.ipynb","file_ext":"py","file_size_in_byte":4650,"program_lang":"python","lang":"en","doc_type":"code","stars":14,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"33577372063","text":"def func(Manufacturer,target_period,reference_period):\n from tkinter.filedialog import askopenfilename\n filename = askopenfilename()\n \n import sys\n import pandas as pd\n df = pd.read_excel(filename,sheet_name = 'input_data')\n del filename\n \n print('Validating availability of Manufacturer in data')\n if Manufacturer in df['Manufacturer'].unique():\n print('Manufacturer available in data')\n else:\n print('Manufacturer not available in the data')\n print('Ensure input from below mentioned list')\n print(df['Manufacturer'].unique())\n sys.exit('Please input the correct Manufacturer and Retry!')\n \n #Converting month to a format as per input\n df.month = pd.PeriodIndex(df.month, freq='M').strftime('%b%Y')\n \n print('Validating availability of target period in data')\n if target_period in df['month'].unique():\n print('Target period input available in data')\n else:\n print('Target Period not available in the data')\n print('Ensure input from below mentioned list')\n print(df['month'].unique())\n sys.exit('Please input the correct date format and Retry!')\n\n print('Validating availability of reference_period in data')\n if reference_period in df['month'].unique():\n print('Reference Period input available in data')\n else:\n print('Reference Period not available in the data')\n print('Ensure input from below mentioned list')\n print(df['month'].unique())\n sys.exit('Please input the correct date format and Retry!')\n \n \n #Filtering Data based on Date\n df = df.loc[(df.month == target_period) | (df.month == reference_period),:]\n\n #Filtering Data based on Manufacturer\n df = df.loc[df['Manufacturer'] == Manufacturer]\n\n #Pivoting Data to Convert Data from long to wide format\n df = (df.pivot_table(index=['Zone', 'Region', 'Manufacturer', 'Brand', 'Subbrand', 'Item','PackSize', 'Packtype'],\n columns='month',\n values='Value Offtake(000 Rs)').reset_index())\n\n #Data is now in a workable format let us start with Analysis\n #Step 1: Lets Check if this Manufacturer had a drop\n\n BY_MANUFACTURER = df.groupby('Manufacturer')\n Manu_Table = BY_MANUFACTURER.sum()\n\n\n if (Manu_Table[target_period] > Manu_Table[reference_period])[0]:\n sys.exit(\"There is no drop in the sales of {0} for the period {1}\".format(Manufacturer,target_period))\n\n print(\"There has been a drop in the sales of {0} for the period {1}\".format(Manufacturer,target_period))\n print(\".\")\n print(\"Initiating deep dive\")\n \n #Making Table for Analysis\n\n \n\n Desired_Levels = ['Zone', 'Region','Manufacturer', 'Brand', 'Subbrand', 'Item']\n Output_Table = pd.DataFrame()\n for levels in Desired_Levels:\n df_1 = df.groupby([levels]).sum()\n df_1 = df_1.reset_index()\n df_1 = df_1.rename(columns={levels:'Focus_Area'},)\n df_1['Growth_Rate'] = round(((df_1[target_period] - df_1[reference_period])/df_1[reference_period])*100)\n df_1['Contribution'] = round(((df_1[target_period] + df_1[reference_period])/(sum(df_1[target_period])+sum(df_1[reference_period]))*100))\n df_1['Product'] = round(((df_1['Growth_Rate']/100) * (df_1['Contribution']/100)),2)\n df_1['Manufacturer'] = Manufacturer\n del df_1[target_period]\n del df_1[reference_period]\n df_1 = df_1[['Manufacturer','Focus_Area','Growth_Rate','Contribution','Product']]\n Output_Table = Output_Table.append(df_1, ignore_index=True)\n\n Output_Table = Output_Table.sort_values(by='Product',ascending=True)\n Output_Table = Output_Table.reset_index()\n del Output_Table['index']\n\n return(Output_Table)\nfunc(Manufacturer='AMUL',target_period='May2019',reference_period='Apr2019')\n","repo_name":"amisha-74/coding","sub_path":"Final Deep Dive Analysis-checkpoint.ipynb","file_name":"Final Deep Dive Analysis-checkpoint.ipynb","file_ext":"py","file_size_in_byte":3849,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"36471005240","text":"# ### Use trained models to classify collection of tweets\n\nimport pandas as pd\nfrom itertools import combinations\nfrom torchtext import vocab\nimport os\nimport sys\nimport torch\nfrom torch.nn import functional as F\nimport numpy as np\nfrom torchtext import data\nfrom torchtext import datasets\nfrom torchtext.vocab import Vectors, GloVe\nfrom torchtext.legacy.data import Field, LabelField, TabularDataset, BucketIterator, Iterator\nimport torch.optim as optim\nfrom sklearn.metrics import accuracy_score, classification_report, confusion_matrix\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\ndf_test = pd.read_csv(\"data/Twitter/clean_twitter_info_annotated.csv\")\ndf_test = df_test[df_test['clean_tweet'].notna()]\n# assign dummy label to work more easily with prediction code\ndf_test['label'] = 0\n\n# +\ndf_train = pd.read_csv(\"data/Twitter/hate_twitter/hate_upsampled_train.csv\")\ndf_val = pd.read_csv(\"data/Twitter/hate_twitter/hate_upsampled_val.csv\")\n\ndf_train = df_train[df_train['clean_tweet'].notna()]\ndf_val = df_val[df_val['clean_tweet'].notna()]\n\ndf_train[[\"clean_tweet\", \"label\"]].to_csv(\"data/Twitter/hate_twitter/train.csv\")\ndf_val[[\"clean_tweet\", \"label\"]].to_csv(\"data/Twitter/hate_twitter/val.csv\")\ndf_test[[\"clean_tweet\", \"label\"]].to_csv(\"data/Twitter/hate_twitter/test.csv\")\n\n# +\n#device = torch.device('cuda' if torch.cuda.is_available() else 'cpu') \ndevice = 'cpu'\n\nif device == 'cpu':\n VECTORS_CACHE_DIR = '.vector_cache/'\n # Please change above to your cache\nelse:\n VECTORS_CACHE_DIR = '.vector_cache/'\n \nBATCH_SIZE = 64\n\n\n# +\nimport torch.nn as nn\nfrom torch.nn.utils.rnn import pack_padded_sequence, pad_packed_sequence\nimport torch.nn.functional as F\n\nclass classifier(nn.Module):\n \n #define all the layers used in model\n def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, text, n_layers, \n bidirectional, dropout):\n \n #Constructor\n super().__init__() \n self.embedding_dim = embedding_dim\n #embedding layer\n self.embedding = nn.Embedding(vocab_size, embedding_dim)\n self.embedding.weight.data = text.vectors\n self.dropout = nn.Dropout(dropout)\n\n #lstm layer\n self.lstm = nn.LSTM(embedding_dim, \n hidden_dim, \n num_layers=n_layers, \n bidirectional=True, \n # dropout=dropout,\n batch_first=True)\n \n #dense layer\n self.fc = nn.Linear(hidden_dim * 2, output_dim)\n \n # #activation function\n self.act = nn.Sigmoid()\n \n def forward(self, text, text_lengths):\n \n #text = [batch size,sent_length]\n embedded = self.embedding(text)\n #embedded = [batch size, sent_len, emb dim]\n\n #packed sequence\n packed_embedded = pack_padded_sequence(embedded, text_lengths,batch_first=True, \n enforce_sorted=False)\n \n packed_output, (hidden, cell) = self.lstm(packed_embedded)\n output, _ = pad_packed_sequence(packed_output, batch_first=True)\n\n out_forward = output[range(len(output)), text_lengths - 1, :self.embedding_dim]\n out_reverse = output[:, 0, self.embedding_dim:]\n out_reduced = torch.cat((out_forward, out_reverse), 1)\n dense_outputs = self.dropout(out_reduced)\n dense_outputs=self.fc(dense_outputs)\n\n dense_outputs = torch.squeeze(dense_outputs, 1)\n outputs = torch.sigmoid(dense_outputs)\n return outputs.to(device)\n\n\n# -\n\nTEXT = Field(tokenize='spacy',batch_first=True,include_lengths=True)\nLABEL = LabelField(sequential=False, use_vocab=False, dtype = torch.float,batch_first=True)\nfields = [(None, None), ('clean_tweet',TEXT),('label', LABEL)]\ntrain, val, test = TabularDataset.splits(path = \"data/Twitter/hate_twitter/\", \n train='train.csv',validation='val.csv',\n test='test.csv', format='csv',\n fields= fields, skip_header=True)\nTEXT.build_vocab(train,min_freq=3,vectors = \"glove.twitter.27B.100d\")\ntrain_iterator, valid_iterator, test_iterator = BucketIterator.splits(\n (train, val, test), \n batch_size = BATCH_SIZE,\n sort_key = lambda x: len(x.clean_tweet),\n sort_within_batch=True,\n device = device)\nsize_of_vocab = len(TEXT.vocab)\nembedding_dim = 100\nnum_hidden_nodes = 50\nnum_output_nodes = 1\nnum_layers = 2\ndropout = 0.2\n\n\ndef load_checkpoint(load_path, model, optimizer, device):\n '''\n Used in evaluate() to load the current best model.\n \n Inputs:\n - load_path: string, where to save model specs.\n - model: a PyTorch model object.\n - optimizer: a PyTorch Optimizer to be used.\n - device: string, 'cpu' or 'cuda' if using google colab.\n \n Returns: float, the models last validation loss.\n \n Other Effects:\n - Loads the saved state at load_path\n into the current model object\n '''\n if load_path==None:\n return\n\n state_dict = torch.load(load_path, map_location=device)\n print(f'Model loaded from <== {load_path}')\n \n model.load_state_dict(state_dict['model_state_dict'])\n optimizer.load_state_dict(state_dict['optimizer_state_dict'])\n \n return model\n\n\n\ndef classify_unlabeled(model, test_loader, device):\n '''\n Accepts the current best model and evaluates\n the test dataset. Printing test accuracy and\n an sklearn confusion matrix report.\n \n Inputs:\n - model: PyTorch model object, the current best model.\n - test_loader: an iterator with test data.\n - device: string, 'cpu' or 'cuda' if using google colab.\n \n Returns: None.\n \n Other Effects:\n Prints test accuracy.\n Prints an Accuracy / F1 Report (sklearn)\n Prints a Confusion Matrix (sklearn & matplotlib)\n '''\n y_pred = []\n\n model.eval()\n with torch.no_grad():\n for ((text, text_len),_) in test_loader: \n \n text = text.to(device)\n text_len = text_len.to(device)\n output = model(text, text_len) \n output = torch.round(output)\n\n y_pred.extend(output.tolist())\n\n return y_pred\n\n\n# +\n# Evaluate\nbest_model = classifier(size_of_vocab, embedding_dim, num_hidden_nodes,num_output_nodes, TEXT.vocab, num_layers, \n bidirectional = True, dropout = dropout)\n\noptimizer = optim.Adam(best_model.parameters(), lr=0.00001)\n\nbest_model = load_checkpoint('model.pt', best_model, optimizer, device)\n\ny_pred = classify_unlabeled(best_model, test_iterator, device)\n# -\n\n# confirm y pred and df test have same size\nlen(y_pred)\n\n# confirm y pred and df test have same size\ndf_test.shape\n\ndf_test['label_pred'] = y_pred\ndf_test['date_dt'] = pd.to_datetime(df_test['Datetime'])\ndf_test.drop(columns=['Unnamed: 0', 'Unnamed: 0.1', 'label'], inplace=True)\n\ndf_test['date_day'] = df_test['date_dt'].dt.date\n\n# show number of hate tweets in labeled data\ndf_test.groupby(['tag'])['tweet_id'].count()\n\ndf_test.groupby(['tag', 'label_pred'])['tweet_id'].count()\n\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].head(70)\n\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].iloc[10].Text\n\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].iloc[15].Text\n\n# examine single tweet example of non-hate, sexist tweets\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].iloc[68].Text\n\ndf_test[(df_test['label_pred'] > 0) & (df_test['tag'] == 'none')& (df_test['date_day'] > pd.to_datetime('2015-01-15'))].head(70)\n\n# examine individual tweet from the above set of hateful, 'none' overlap\ndf_test[(df_test['label_pred'] > 0) & (df_test['tag'] == 'none')& (df_test['date_day'] > pd.to_datetime('2015-01-15'))].iloc[66].Text\n\n# ### Visualize predicted hate comments over time\n\ntweet_preds_day = df_test.groupby(['label_pred', 'date_day'])['tweet_id'].count().reset_index()\ntweet_preds_day.rename(columns={'tweet_id':'num_tweets'},inplace=True)\n\ntweet_preds_day.head()\n\n# +\n# View predicted hate comments over time\nf, ax = plt.subplots(1,1,figsize=(15,10))\n\nsns.lineplot('date_day', 'num_tweets',hue='label_pred', \\\n data=tweet_preds_day[tweet_preds_day['label_pred']>0])\n\nax.set_xlabel(\"Date\")\nax.set_ylabel(\"Number of Hateful Tweets\")\nax.set_title(\"Tweets Classified as Hateful by LSTM Model\")\n# -\n\n# ### Compare with Google Trends\n\n# +\nimport pytrends\nimport pandas as pd\nfrom pytrends.request import TrendReq\nimport matplotlib\nimport csv\nfrom datetime import date\nimport time \nimport requests\nimport warnings\nwarnings.filterwarnings('ignore')\n\npytrends = TrendReq(hl='en-US', tz=360)\n\n# +\nkeyword = [\"nigger\", \"niggers\", \"kike\", \"chink\", \"faggot\"]\n\npytrends.build_payload(keyword, cat=0, timeframe='2013-09-01 2015-04-01', geo='US', gprop='')\ninterest_over_time_df = pytrends.interest_over_time()\n\n# +\nimport matplotlib.pyplot as plt\nimport numpy as np\n\nf, (ax1, ax2) = plt.subplots(2, 1, figsize=(15,10))\n\n\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"nigger\"], label = \"Anti-Black word\")\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"kike\"], label = \"Anti-Semitic word\")\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"chink\"], label = \"Anti-Asian word\")\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"faggot\"], label = \"Anti-LGBT word\")\nax1.set_title(\"Google Trends Hate Speech\")\nax2.plot('date_day', 'num_tweets', data=tweet_preds_day[tweet_preds_day['label_pred']>0])\nax2.set_title(\"Classified Hate Speech\")\nplt.plot()\nplt.legend()\nplt.show()\n\n# -\n\n\n","repo_name":"keiraou/life_cycle_of_hate_speech","sub_path":"classify_with_trained_models.ipynb","file_name":"classify_with_trained_models.ipynb","file_ext":"py","file_size_in_byte":9716,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"42823273365","text":"import pandas as pd\nimport numpy as np\n\nimport sklearn as sk\nfrom sklearn import tree\n\nratings = pd.read_csv(\"ratings.csv\")\nmean_ratings = ratings.groupby(\"movieId\").rating.mean()\n\nmovies = pd.read_csv(\"movies.csv\")\nmovies = movies.set_index(\"movieId\")\nmovies['rating'] = mean_ratings\nmovies.head()\n\nmovies['year'] = movies.title.str.extract(\"\\((\\d{4})\\)\", expand=False)\nmovies.year.astype(float)\nmovies.head()\n\ngenres = [\"Drama\", \"Comedy\", \"Thrilles\", \"Action\" ,\"Adventure\", \"Crime\"]\nfor genre in genres:\n movies[genre] = movies.genres.str.contains(genre)\nmovies['good'] = np.where(movies.rating>3.5, \"Good\", \"Bad\")\nmovies.head(5)\n\nfrom sklearn import tree\nclf = tree.DecisionTreeClassifier(max_depth=2)\nm = clf.fit(movies[genres], movies.good)\n\npredictions = m.predict(movies[genres])\npd.crosstab(movies.good, predictions)\n\nfrom sklearn.metrics import confusion_matrix\nprint(confusion_matrix(movies.good, predictions))\n\ncorrect = movies.good == predictions\ncorrect.mean()\n\nimport pydotplus\nfrom IPython.display import Image\ndot_data = tree.export_graphviz(m,\n feature_names=genres, class_names=['bad', 'good'], \n filled=True, rounded=True) \ngraph = pydotplus.graph_from_dot_data(dot_data) \n\n\n","repo_name":"peiweihe/analyse_amazon_reviews","sub_path":"trees.ipynb","file_name":"trees.ipynb","file_ext":"py","file_size_in_byte":1201,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"28043068283","text":"# +\nrecords = [\n ('foo',1,2),\n ('bar','hello'),\n ('foo',3,4),\n]\ndef do_foo(x,y):\n print('foo',x,y)\n\ndef do_bar(s):\n print('bar',s)\n\nfor tag, *args in records:\n if tag == 'foo':\n do_foo(*args)\n elif tag == 'bar':\n do_bar(*args)\n# -\n\nimport heapq\nportfolio = [\n {'name': 'ibm', 'shares':100, 'price': 91.1},\n {'name': 'appl', 'shares':50, 'price': 543.22},\n {'name': 'fb', 'shares':200, 'price': 21.09},\n {'name': 'yhoo', 'shares':45, 'price': 16.35},\n]\ncheap = heapq.nsmallest(3, portfolio, key=lambda s : s['price'])\nexpensive = heapq.nlargest(3, portfolio, key=lambda s : s['price'])\n\ncheap\n\nexpensive\n\nnums = [1,3,2,5,4.5, 77,23,9,6, 54,96,0,12,11,2.3]\n\nlen(nums)\n\nimport time\n\nt1 = time.time()\nnew_lis = sorted(nums)[:5]\nt2 = time.time() - t1\n\nnew_lis\n\nt2\n\n\nt3 = time.time()\nnew_hlis= heapq.nsmallest(5,nums)\nt4 = time.time()- t3\n\nnew_hlis\n\nt4\n\n\n# # 下划线起名的规则\n# + _xx:前置单下划线,私有化属性或方法,一般来讲,变量名_xx被看作是“私有 的”,在模块或类外不可以使用。当变量是私有的时候,用_xx 来表示变量是很好的习惯。类对象和子类可以访问,这并不能完全做到真正的私有,只是约定俗成的而已,这样写表示不希望这个变量在外部被直接调用\n# + \\__xx:前置双下划线,私有化属性或方法,无法在外部直接访问(名字重整所以访问不到,只能是允许这个类本身进行访问了。连子类也不可以)\n# + \\__xx\\__:前后双下划线,系统定义名字(这就是在python中强大的魔法方法),因为变量名__xxx__对Python 来说有特殊含义,对于普通的变量应当避免这种命名风格。\n# + xx_:后置单下划线,用于避免与Python关键词的冲突\n# 如以下例子所示,我在test类中定义了x,_x和 __x三个属性,并创建了test的类对象t,对这三个属性进行访问,__x不能被访问到\n\nclass test(object):\n def __init__(self):\n self.x = 10\n self._x = 20\n self.__x = 30\nt = test()\nprint(t.x) # 10\nprint(t._x) # 20\nprint(t.__x)\n# print(t.__x) # AttributeError: 'test' object has no attribute '__x'\n\n# 构建一个优先级队列\nclass PriorityQueue:\n def __init__(self):\n self._queue= []\n self._index = 0\n \n def push(self, item, priority):\n heapq.heappush(self._queue, (-priority, self._index,item))\n self._index += 1\n \n def pop(self):\n return heapq.heappop(self._queue)[-1]\n\n\nclass Item:\n def __init__(self, name):\n self.name = name\n \n def __repr__(self):\n return f\"Item('{self.name}')\"\n\n\nq = PriorityQueue()\nq.push(Item('foo'), 1)\nq.push(Item('bar'), 2)\nq.push(Item('spam'), 3)\nq.push(Item('grok'), 5)\n\nq.pop()\n\nq.pop()\n\nq.push(Item('test'), 1)\n\nq.pop()\n\nq.pop()\n\nq.pop()\n\nq.pop()\n\n# # 字典中的键映射多个值(multidict)\n\n# +\nfrom collections import defaultdict\n\nd = defaultdict(list)\nd['a'].append(1)\nd['a'].append(2)\nd['b'].append(4)\n\n# -\n\nd\n\nd_prime = defaultdict(set)\nd_prime['a'].add(1)\nd_prime['a'].add(2)\nd_prime['b'].add(4)\n\nd_prime\n\n\n# # 字典排序\n\n# +\nfrom collections import OrderedDict\ndef ordered_dict():\n d = OrderedDict()\n d['foo']=1\n d['bar']=2\n d['spam']=3\n d['grok']=4\n \n for key in d:\n print(key, d[key])\n\nc= ordered_dict()\n \n\n# -\n\n# # 字典的运算\n\nprices = {\n 'acme': 45.23,\n 'aapl': 612.78,\n 'ibm':205.55,\n 'hpq': 37.20,\n 'fb': 10.75 \n}\n\nmin_price = min(zip(prices.values(), prices.keys()))\n\nmin_price\n\nmax_price = max(zip(prices.values(), prices.keys()))\n\nmax_price\n\nsorted_prices = sorted(zip(prices.values(), prices.keys()))\n\nsorted_prices\n\nprices_and_names = zip(prices.values(), prices.keys())\nprint(min(prices_and_names)) # ok\nprint(max(prices_and_names)) # ValueError: max() arg is an empty set\n\nprint(max(prices))\nprint(min(prices))\n\nprint(min(prices.values()))\nprint(max(prices.values()))\n\nprint(min(prices, key= lambda k: prices[k]))\nprint(max(prices, key= lambda k: prices[k]))\n\nmin_value = prices[min(prices, key= lambda k: prices[k])]\n\nmin_value\n\n# ## `总之,用zip函数达到字典键值反转的效果`\n\n# # 查找两字典的相同点\n\na = {\n 'x': 1,\n 'y': 2,\n 'z': 3,\n}\nb= {\n 'p':233,\n 'y': 11,\n 'x': 1\n}\n\n# find keys in common\na.keys() & b.keys()\n\n# find keys in a that not in b\na.keys() - b.keys()\n\n# find (key, value) pairs in common\na.items() & b.items()\n\n# make a new dict with certain keys removed\nc = {key: a[key] for key in a.keys() - {'z', 'w'}}\n\nc\n\n\n# # 删除序列相同元素并保持顺序\n\n# When hashable\ndef dedupe(items):\n seen = set()\n for item in items:\n if item not in seen:\n yield item\n seen.add(item)\n\n\na = [1,5,2,1,9,1,5,10]\nlist(dedupe(a))\n\n# ## About 'Hashale'\n# > An object is hashable if it has a hash value which never changes during its lifetime (it needs a __hash__() method), and can be compared to other objects (it needs an __eq__() or __cmp__() method). Hashable objects which compare equal must have the same hash value.\n#\n# > Hashability makes an object usable as a dictionary key and a set member, because these data structures use the hash value internally.\n#\n# > All of Python’s immutable built-in objects are hashable, while no mutable containers (such as lists or dictionaries) are. Objects which are instances of user-defined classes are hashable by default; they all compare unequal, and their hash value is their id().\n\ntype(a)\n\n\n# when it's not hashable\ndef dedupe(items, key= None):\n seen = set()\n for item in items:\n val = item if key is None else key(item)\n if val not in seen:\n yield item \n seen.add(val)\n\n\na = [{'x': 1, 'y':2}, {'x':1,'y':3}, {'x':2,'y':4}, {'x': 1, 'y':2}]\n\nlist(dedupe(a, key=lambda d: (d['x'], d['y'])))\n\nlist(dedupe(a, key=lambda d: (d['x'])))\n\n# 当然了,最简单的还是set(),但是不能保证顺序\n\na = [1,5,2,1,9,1,5,10]\n\nset(a)\n\nrecord = '....................100 .......513.25 ..........'\ncost = int(record[20:23]) * float(record[31:37])\n\nprint(cost)\n\nSHARES = slice(20, 23)\nPRICE = slice(31, 37)\ncost = int(record[SHARES]) * float(record[PRICE])\n\ncost\n\nitems = [0, 1, 2, 3, 4, 5, 6]\na= slice(2,4)\nitems[a]\n\nitems[a]=[10,11]\n\nitems\n\ndel items[a]\n\nitems\n\ns= slice(5,50,2)\n\n\ns.start\n\ns.stop\n\ns.step\n\ns = 'HelloWorld'\n\na.indices(len(s))\n\nfor i in range(*a.indices(len(s))):\n print(s[i])\n \n\n# # 找出序列中出现次数最多的元素\n\nwords = [\n'look', 'into', 'my', 'eyes', 'look', 'into', 'my', 'eyes',\n'the', 'eyes', 'the', 'eyes', 'the', 'eyes', 'not', 'around', 'the',\n'eyes', \"don't\", 'look', 'around', 'the', 'eyes', 'look', 'into',\n'my', 'eyes', \"you're\", 'under'\n]\n\nfrom collections import Counter\nword_counts = Counter(words)\ntop_three = word_counts.most_common(3)\n\ntop_three\n\n# Counter 对象可以接受任意的hashable 序列对象\n\nword_counts['eyes']\n\nword_counts['not']\n\n# 如果你想手动增加计数,可以简单的用加法:\n\nmorewords = ['why','are','you','not','looking','in','my','eyes']\n\nfor item in morewords:\n print(item)\n\ntype(word_counts)\n\nword_counts\n\nfor word in morewords:\n word_counts['word']+=1\n \n\nword_counts['eyes']\n\nword_counts\n\n# 上面写的手动修改方法已经失效\n\nword_counts.update(morewords)\n\nword_counts\n\nword_counts['not']\n\nword_counts['eyes']\n\n# Counter 实例一个鲜为人知的特性是它们可以很容易的跟数学运算操作相结合\n\n# +\na = Counter(words)\nb = Counter(morewords)\n\n\n# -\n\na\n\nb\n\na + b\n\na - b\n\na & b\n\n# # 通过某个关键字排序一个字典列表\n\nrows = [\n{'fname': 'Brian', 'lname': 'Jones', 'uid': 1003},\n{'fname': 'David', 'lname': 'Beazley', 'uid': 1002},\n{'fname': 'John', 'lname': 'Cleese', 'uid': 1001},\n{'fname': 'Big', 'lname': 'Jones', 'uid': 1004}\n]\n\nfrom operator import itemgetter\n\nrows_by_fname = sorted(rows, key=itemgetter('fname'))\nrows_by_uid = sorted(rows, key=itemgetter('uid'))\n\nrows_by_fname\n\nrows_by_uid\n\n# 支持多个keys\nrows_by_lfname = sorted(rows, key=itemgetter('lname','fname'))\n\nrows_by_lfname\n\n# itemgetter() 有时候也可以用lambda 表达式代替\nrows_by_fname = sorted(rows, key= lambda r: r['fname'])\n\n# 当然还是itemgetter效率更快\nrows_by_fname\n\nmin(rows, key=itemgetter('uid'))\n\n\n# # 排序不支持原生比较的对象\n\nclass User:\n def __init__(self, user_id):\n self.user_id = user_id\n \n def __repr__(self):\n return f'User({self.user_id})'\n\n\ndef sort_notcompare():\n users = [User(233), User(64), User(89)]\n print(users)\n print(sorted(users, key= lambda u: u.user_id))\n\n\nsort_notcompare()\n\nusers = [User(233), User(64), User(89)]\n\n# 使用operator.attrgetter\nfrom operator import attrgetter\nsorted(users, key=attrgetter('user_id'))\n\nmin(users, key=attrgetter('user_id'))\n\n# ### 不管是itemgetter还是attrgetter, 效果基本都和lambda一样,前者效率更高, 且能多个字段比较\n\n# # 通过某个字段将记录分组\n\n# 你有一个字典或者实例的序列,然后你想根据某个特定的字段比如date 来分组迭代访问。\n\nrows = [\n{'address': '5412 N CLARK', 'date': '07/01/2012'},\n{'address': '5148 N CLARK', 'date': '07/04/2012'},\n{'address': '5800 E 58TH', 'date': '07/02/2012'},\n{'address': '2122 N CLARK', 'date': '07/03/2012'},\n{'address': '5645 N RAVENSWOOD', 'date': '07/02/2012'},\n{'address': '1060 W ADDISON', 'date': '07/02/2012'},\n{'address': '4801 N BROADWAY', 'date': '07/01/2012'},\n{'address': '1039 W GRANVILLE', 'date': '07/04/2012'},\n]\n\nfrom operator import itemgetter\nfrom itertools import groupby\n\nrows.sort(key=itemgetter('date'))\n\nrows\n\nfor date, items in groupby(rows, key=itemgetter('date')):\n print(date)\n for i in items:\n print('+', i)\n\n# ### groupby需要事先排序,假如说只是想将一些字段分组到更大的数据结构中,并且允许随机访问,还是使用defaultdict为佳\n\nrows_by_date = defaultdict(list)\nfor row in rows:\n rows_by_date[row['date']].append(row)\n\nfor r in rows_by_date['07/01/2012']:\n print(r)\n\nrows_by_date\n\n# # 过滤序列元素\n\n# ### 最简单的过滤序列元素的方法就是使用列表推导\n\nmylist = [1, 4, -5, 10, -7, 2, 3, -1]\n\n[n for n in mylist if n > 0]\n\n# ### 使用列表推导的一个潜在缺陷就是如果输入非常大的时候会产生一个非常大的结果集,占用大量内存\n\npos = (n for n in mylist if n > 0)\n\npos\n\nfor i in pos:\n print(i)\n\n# ### 当过滤规比较复杂时,需要写一个函数,使用内建filter(返回的是迭代器)\n\nvalues = ['1', '2', '-3', '-', '4', 'N/A', '5']\n\n\ndef is_int(val):\n try: \n x = int(val)\n return True\n except ValueError:\n return False\nivals = list(filter(is_int, values))\n\nivals\n\n# +\n# filter()\n#Return an iterator yielding those items of iterable for which function(item)\n# -\n\n# ### 列表推导和生成器表达式通常情况下是过滤数据最简单的方式, 其实它们还能在过滤的时候转换数据\n\n# +\nmylist = [1, 4, -5, 10, -7, 2, 3, -1]\nimport math\n\n[math.sqrt(n) for n in mylist if n > 0]\n# -\n\n# ### 过滤操作的一个变种就是将不符合条件的值用新的值代替,而不是丢弃它们\n\nclip_neg = [n if n > 0 else 0 for n in mylist]\n\nclip_neg\n\naddresses = [\n'5412 N CLARK',\n'5148 N CLARK',\n'5800 E 58TH',\n'2122 N CLARK',\n'5645 N RAVENSWOOD',\n'1060 W ADDISON',\n'4801 N BROADWAY',\n'1039 W GRANVILLE',\n]\ncounts = [ 0, 3, 10, 4, 1, 7, 6, 1]\n\nlen(addresses)\n\nlen(counts)\n\n# ### itertools.compress() ,它以一个iterable对象和一个相对应的Boolean 选择器序列作为输入参数。然后输出iterable 对象中对应选择器为True 的元素。当你需要用另外一个相关联的序列来过滤某个序列的时候,这个函数是非常有用的。\n\n# +\nfrom itertools import compress\n\nmore5 = [n>5 for n in counts]\n# -\n\nmore5\n\nlist(compress(addresses, more5))\n\n# # 从字典中提取子集\n\n# +\nprices = {\n'ACME': 45.23,\n'AAPL': 612.78,\n'IBM': 205.55,\n'HPQ': 37.20,\n'FB': 10.75\n}\n\n# 字典推导\n#Make a dictionary of all prices over 200\np1 = {key: value for key, value in prices.items() if value > 200}\n# Make a dictionary of tech stocks\ntech_names = {'AAPL', 'IBM', 'HPQ', 'MSFT'}\np2 = {key: value for key, value in prices.items() if key in tech_names}\n# -\n\np1\n\np2\n\n# 比较慢\np1 = dict((key, value) for key, value in prices.items() if value > 200)\n\np1\n\ntech_names = { 'AAPL', 'IBM', 'HPQ', 'MSFT' }\np2 = { key:prices[key] for key in prices.keys() & tech_names }\n\n# 同理,也比较慢\np2\n\n# # 映射名称到序列元素\n\nfrom collections import namedtuple\nSubscriber = namedtuple('Subscriber', ['addr', 'joined'])\n\nsub = Subscriber('joney@example.com', '2021-3-15')\n\nsub.addr\n\nsub.joined\n\nlen(sub)\n\n# ### 尽管namedtuple 的实例看起来像一个普通的类实例,但是它跟元组类型是可交换的,支持所有的普通元组操作,比如索引和解压。\n\naddr, joined = sub\n\naddr\n\njoined\n\nsub\n\n\ndef compute_cost(records):\n total = 0.0\n for rec in records:\n total += rec[1] * rec[2]\n return total\n\n\nStock = namedtuple('Stock', ['name', 'shares', 'price'])\ndef compute_cost(records):\n total = 0.0\n \n total += s.shares * s.price\n return total\n\n\ns = Stock('ACME', 100, 123.45)\n\ns\n\ns.shares\n\ns.price\n\ncompute_cost(s)\n\ns.shares = 75\n\ns = s._replace(shares = 75)\n\ns\n\nStock = namedtuple('Stock', ['name', 'shares', 'price', 'date', 'time'])\n# Create a prototype instance\nstock_prototype = Stock('', 0, 0.0, None, None)\n# Function to convert a dictionary to a Stock\ndef dict_to_stock(s):\n return stock_prototype._replace(**s)\n\n\na = {'name': 'ACME', 'shares': 100, 'price': 123.45}\n\ndict_to_stock(a)\n\nstock_prototype\n\n# ### 如果你的目标是定义一个需要更新很多实例属性的高效数据结构,那么命名元组并不是你的最佳选择。这时候你应该考虑定义一个包含slots 方法的类\n\n# # 转换并同时计算数据\n\nnums = [1, 2, 3, 4, 5]\ns = sum(x * x for x in nums)\n\ns\n\nimport os\n# Determine if any .py files exist in a directory\nfiles = os.listdir('/Users/joriri/Code')\nif any(name.endswith('.py') for name in files):\n print('There be python!')\nelse:\n print('Sorry, no python.')\n\n# Output a tuple as CSV\ns = ('ACME', 50, 123.45)\nprint(','.join(str(x) for x in s))\n# Data reduction across fields of a data structure\nportfolio = [\n{'name':'GOOG', 'shares': 50},\n{'name':'YHOO', 'shares': 75},\n{'name':'AOL', 'shares': 20},\n{'name':'SCOX', 'shares': 65}\n]\nmin_shares = min(s['shares'] for s in portfolio)\n\nmin_shares\n\n# ### 在使用一些聚集函数比如min() 和max() 的时候你可能更加倾向于使用生成器版本,它们接受的一个key 关键字参数或许对你很有帮助。\n\n# Original: Returns 20\nmin_shares = min(s['shares'] for s in portfolio)\n# Alternative: Returns {'name': 'AOL', 'shares': 20}\nmin_shares = min(portfolio, key=lambda s: s['shares'])\n\nmin_shares\n\n# # 合并多个字典或映射\n\na = {'x': 1, 'z': 3 }\nb = {'y': 2, 'z': 4 , 'p': 5}\n\n# +\nfrom collections import ChainMap\nc = ChainMap(a,b)\n\nprint(c['x'])\nprint(c['z'])\nprint(c['p'])\n# -\n\ndel c['y']\n\ndel c['x']\n\nc['x']\n\na\n\nc['w']= 6\n\na\n\n# ### 对chainmap的字典(列表)来说, 更新删除影响的是列表中的第一个字典\n\nvalues = ChainMap()\nvalues['x'] =1 \n\nvalues=values.new_child()\nvalues['x'] = 2\nvalues=values.new_child()\nvalues['x']=3\n\n\nvalues\n\nvalues['x']\n\n# discard last mapping\nvalues = values.parents\nvalues['x']\n\nvalues = values.parents\nvalues['x']\n\nvalues\n\n# ### 当然也可以用updata来代替chainmap\n\na = {'x': 1, 'z': 3 }\nb = {'y': 2, 'z': 4 }\n\nmerged = dict(b)\nmerged.update(a)\n\nmerged\n\na['x']= 233\n\nmerged\n\n\n# ### 这样也能行得通,但是它需要你创建一个完全不同的字典对象(或者是破坏现有字典结构)。同时,如果原字典做了更新,这种改变不会反应到新的合并字典中去\n\n# ### ChianMap 使用原来的字典,它自己不创建新的字典。所以它并不会产生上面所说的结果\n\na = {'x': 1, 'z': 3 }\nb = {'y': 2, 'z': 4 }\nmerged = ChainMap(a,b)\n\nmerged\n\na['x']= 233\n\nmerged\n\n# # 第二章 字符串和文本\n# # 使用多个界定符分割字符串\n\n# ### string 对象的split() 方法只适应于非常简单的字符串分割情形,它并不允许有多个分隔符或者是分隔符周围不确定的空格。当你需要更加灵活的切割字符串的时候,最好使用re.split() 方法:\n\n# +\nline = 'asdf fjdk; afed, fjek,asdf, foo'\n\nimport re\nre.split(r'[;,\\s]\\s*', line)\n# -\n\n# ### 当你使用re.split() 函数时候,需要特别注意的是正则表达式中是否包含一个括号捕获分组。如果使用了捕获分组,那么被匹配的文本也将出现在结果列表中\n\n# 分隔符被捕获了\nfields = re.split(r'(,|;|\\s)\\s*', line)\nfields\n\nvalues = fields[::2]\n\nvalues\n\ndelimeters = fields[1::2]\n\ndelimeters\n\nlen(values)\n\nlen(delimeters)\n\nlen(fields)\n\ndelimeters += ['']\n\ndelimeters\n\nlen(delimeters)\n\n# reform the line using the same delimiters \n''.join(v+d for v, d in zip(values, delimeters))\n\n# ### 如果你不想保留分割字符串到结果列表中去,但仍然需要使用到括号来分组正则表达式的话,确保你的分组是非捕获分组,形如(?:...)\n\nre.split(r'(?:,|;|\\s)\\s*', line)\n\n# # 字符串开头或结尾匹配\n\n# ### 直接使用str.startswith & str.endswith\n\nfilename = 'spam.txt'\n\nfilename.startswith('spam')\n\nfilename.endswith('ttt')\n\nurl = 'http://www.baidu.com'\n\nurl.startswith('https')\n\nurl.endswith('com')\n\n# ### 匹配多种可能,先成一个`元组`\n\nimport os\nfilenames = os.listdir('.')\n\nfilenames\n\n[name for name in filenames if name.endswith(('.ipynb', 'csv'))]\n\nany(name.endswith('.py') for name in filenames)\n\n# +\nfrom urllib.request import urlopen\n\ndef read_data(name):\n if name.startswith('http', 'https', 'ftp'):\n return urlopen(name).read()\n else:\n with open(name) as f:\n return f.read()\n\n\n# -\n\n# 必须是元组\nchoices = ['http:', 'ftp:']\nurl = 'http://www.python.org'\nurl.startswith(choices)\n\nurl.startswith(tuple(choices))\n\n# ### 这种匹配也可以用正则或者切片,但是startswith是最简单且快速的。 并且,这种方法和其他普通数据聚合方法相结合(any…)也十分不错\n\n# # 用Shell 通配符匹配字符串\n\n# ## 你想使用Unix Shell 中常用的通配符(比如*.py , Dat[0-9]*.csv 等) 去匹配文本字符串\n\nfrom fnmatch import fnmatch, fnmatchcase\n\nfnmatch('foo.txt', '*.txt')\n\nfnmatch('foo.txt', '?oo.txt')\n\nfnmatch('Date45.cvs', 'Date[0-9]*')\n\nnames = ['Dat1.csv', 'Dat2.csv', 'config.ini', 'foo.py']\n\n[name for name in names if fnmatch(name, 'Dat*.csv')]\n\n# 大小写敏感 \n# False on Mac, True on Windows\nfnmatch('foo.txt', '*.TXT')\n\n# 如果要把大小写考虑进去,用fnmatchcase\nfnmatchcase('foo.txt', '*.TXT')\n\n# ### 这两个函数通常会被忽略的一个特性是在处理非文件名的字符串时候它们也是很有用的\n\naddresses = [\n'5412 N CLARK ST',\n'1060 W ADDISON ST',\n'1039 W GRANVILLE AVE',\n'2122 N CLARK ST',\n'4802 N BROADWAY',\n]\n\n[addr for addr in addresses if fnmatchcase(addr, '* ST')]\n\n[addr for addr in addresses if fnmatchcase(addr, '54[0-9][0-9] *CLARK*')]\n\n# ### fnmatch() 函数匹配能力介于简单的字符串方法和强大的正则表达式之间。如果在数据处理操作中只需要简单的通配符就能完成的时候,这通常是一个比较合理的方案。如果你的代码需要做文件名的匹配,最好使用glob 模块\n\n# # 字符串匹配和搜索\n\n# ### startswith endswith find\n\ntext = 'yeah, but no, but yeah, but no, but yeah'\n\ntext.find('no')\n\n# 使用正则\ntext1 = '11/27/2012'\ntext2 = 'Nov 27, 2012'\n\nif re.match(r'\\d+/\\d+/\\d+', text1):\n print('yes')\nelse:\n print('no')\n\n# ### 如果你想使用同一个模式去做多次匹配,你应该先将模式字符串预编译为模式对象\n\ndatepat = re.compile(r'\\d+/\\d+/\\d+')\n\nif datepat.match(text1):\n print('yes')\nelse:\n print('no')\n\n# ### match() 总是从字符串开始去匹配,如果你想查找字符串任意部分的模式出现位置,使用findall() 方法去代替。\n\ntext = 'Today is 11/27/2012. PyCon starts 3/13/2013.'\ndatepat.findall(text)\n\n# 在定义正则式的时候,通常会利用括号去捕获分组\ndatepat = re.compile(r'(\\d+)/(\\d+)/(\\d+)')\n\nm = datepat.match('11/27/2021')\n\n\nm\n\n# Return subgroup(s) of the match by indices or names.\n# For 0 returns the entire match.\nm.group(0)\n\nm.group(1)\n\nm.group(2)\n\nm.groups()\n\nm.group()\n\ntext\n\ndatepat.findall(text)\n\nfor month, day, year in datepat.findall(text):\n print(f'{year}-{month}-{day}')\n\n# # 字符串搜索和替换\n\ntext = 'yeah, but no, but yeah, but no, but yeah'\ntext.replace('yeah', 'yep')\n\ntext = 'Today is 11/27/2012. PyCon starts 3/13/2013.'\n\n# 注意这里匹配的模式有括号,意味着有group, 后面的\\3 \\1 \\2对应的是组号\nre.sub(r'(\\d+)/(\\d+)/(\\d+)', r'\\3-\\1-\\2', text)\n\n# ### 对于更加复杂的替换,可以传递一个替换回调函数来代替\n\nfrom calendar import month_abbr\ndef change_date(m):\n month_name = month_abbr[int(m.group(1))]\n return f'{m.group(2)} {month_name} {m.group(1)}'\n\n\ndatepat.sub(change_date, text)\n\ndatepat\n\nnewtext, n = datepat.subn(r'\\3-\\1-\\2', text)\n\nnewtext\n\nn\n\n# # 字符串忽略大小写的搜索替换\n\ntext = 'UPPER PYTHON, lower python, Mixed Python'\n\nre.findall('python', text, flags=re.IGNORECASE)\n\nre.sub('python', 'snake', text, flags=re.IGNORECASE)\n\n\n# 如果要使替换的单词和原有的单词 大小写一致\ndef matchcase(word):\n def replace(m):\n text = m.group()\n if text.isupper():\n return word.upper()\n elif text.islower():\n return word.lower()\n elif text[0].isupper():\n return word.capitalize()\n else:\n return word\n return replace\n\n\nre.sub('python', matchcase('snake'), text, flags=re.IGNORECASE)\n\n# # 最短匹配模式\n\nstr_pat = re.compile(r'\\\"(.*)\\\"')\n\ntext1= 'Computer says \"no.\"'\n\nstr_pat.findall(text1)\n\ntext2 = 'Computer says \"no.\" Phone says \"yes.\"'\nstr_pat.findall(text2)\n\nstr_pat = re.compile(r'\\\"(.*?)\\\"')\nstr_pat.findall(text2)\n\n# # 多行匹配模式\n\ncomment = re.compile(r'/\\*(.*?)\\*/')\ntext1= '/*this is a comment.*/'\ntext2 = '''/*this is a \nmultiline comment.*/'''\n\ntext1\n\ntext2\n\ncomment.findall(text1)\n\ncomment.findall(text2)\n\ncomment = re.compile(r'/\\*((?:.|\\n)*?)\\*/')\n\ncomment.findall(text2)\n\nnew_comment = re.compile(r'/\\*([\\s\\S]*?)\\*/')\n\nnew_comment.findall(text2)\n\n# re.compile的 re.DOTALL\ncomment = re.compile(r'/\\*(.*?)\\*/', re.DOTALL)\n\ncomment.findall(text2)\n\n# ### 对于简单的情况使用re.DOTALL 标记参数工作的很好,但是如果模式非常复杂或者是为了构造字符串令牌而将多个模式合并起来(2.18 节有详细描述),这时候使用个标记参数就可能出现一些问题。如果让你选择的话,最好还是定义自己的正则表达���模式,这样它可以在不需要额外的标记参数下也能工作的很好。\n#\n#\n\n# # 将Unicode 文本标准化\n\ns1 = 'Spicy Jalape\\u00f1o'\ns2 = 'Spicy Jalapen\\u0303o'\n\ns1\n\ns2\n\ns1 == s2\n\nlen(s1)\n\nlen(s2)\n\nimport unicodedata\nt1 = unicodedata.normalize('NFC', s1)\nt2 = unicodedata.normalize('NFC', s2)\n\nt1\n\nt2\n\nt1 == t2\n\nlen(t1)\n\nt1 = unicodedata.normalize('NFD', s1)\nt2 = unicodedata.normalize('NFD', s2)\n\nt1\n\nlen(t1)\n\ns = '\\ufb01' # A single character\n\ns\n\nunicodedata.normalize('NFC', s)\n\nunicodedata.normalize('NFD', s)\n\nunicodedata.normalize('NFKD', s)\n\nunicodedata.normalize('NFKC', s)\n\nt1\n\n''.join(c for c in t1 if not unicodedata.combining(c))\n\n# # 在正则式中使用Unicode\n\n# +\n# return a match Object or None\n# re.match()\n# -\n\nnum= re.compile('\\d+')\nnum.match('212')\n\nnum.match('\\u0661\\u0662\\u0663')\n\narabic = re.compile('[\\u0600-\\u06ff\\u0750-\\u077f\\u08a0-\\u08ff]+')\n\npat = re.compile('stra\\u00dfe', re.IGNORECASE)\n\n\ns = 'straße'\npat.match(s)\n\n# does't match\npat.match(s.upper())\n\ns.upper()\n\n# ### 混合使用Unicode 和正则表达式通常会让你抓狂。如果你真的打算这样做的话,最好考虑下安装第三方正则式库,它们会为Unicode 的大小写转换和其他大量有趣特性提供全面的支持,包括模糊匹配。\n\n# # 删除字符串中不需要的字符\n\ns = ' hello world \\n'\n\ns.strip()\n\ns.lstrip()\n\ns.rstrip()\n\nt = '-----hello====='\n\nt.lstrip('-')\n\nt.rstrip('=')\n\nt.strip('-=')\n\ns = ' hello world \\n'\n\nre.sub('\\s+', ' ', s)\n\n# # 审查清理文本字符串\n\n# ### 文本清理问题会涉及到包括文本解析与数据处理等一系列问题。在非常简单的情形下,你可能会选择使用字符串函数(比如str.upper() 和str.lower() ) 将文本转为标准格式。使用str.replace() 或者re.sub() 的简单替换操作能删除或者改变指定的字符序列。你同样还可以使用2.9 小节的nicodedata.normalize() 函数将unicode文本标准化。\n# ### 有时候你可能还想在清理操作上更进一步。比如,你可能想消除整个区间上的字符或者去除变音符。为了这样做,你可以使用经常会被忽视的str.translate()方法。\n\ns = 'pýtĥöñ\\fis\\tawesome\\r\\n'\n\ns\n\n# +\nremap = {\n ord('\\t'): ' ',\n ord('\\f'): ' ',\n ord('\\r'): None,\n}\n\na = s.translate(remap)\n# -\n\na\n\n# +\nimport sys\n\ncmb_chrs = dict.fromkeys(c for c in range(sys.maxunicode) \n if unicodedata.combining(chr(c)))\n# -\n\nb = unicodedata.normalize('NFD', a)\n\nb\n\nb.translate(cmb_chrs)\n\n# +\n# 构造一个将所有Unicode 数字字符映射到对应的ASCII 字\n# 符上的表格\ndigitmap = {c: ord('0')+ unicodedata.digit(chr(c))\n for c in range(sys.maxunicode)\n if unicodedata.category(chr(c)) =='Nd'}\n\nlen(digitmap)\n# -\n\ndigitmap = { c: ord('0') + unicodedata.digit(chr(c))\n for c in range(sys.maxunicode)\n if unicodedata.category(chr(c)) == 'Nd' }\n\nlen(digitmap)\n\nx = '\\u0661\\u0662\\u0663'\n\nx.translate(digitmap)\n\n# ### 另一种清理文本的技术涉及到I/O 解码与编码函数。这里的思路是先对文本做一些初步的清理,然后再结合encode() 或者decode() 操作来清除或修改它\n\na\n\n\nb = unicodedata.normalize('NFD',a)\n\nb\n\nb.encode('ascii', 'ignore').decode('ascii')\n\n\n# ### 这里的标准化操作将原来的文本分解为单独的和音符。接下来的ASCII 编码/解码只是简单的一下子丢弃掉那些字符。当然,这种方法仅仅只在最后的目标就是获取到文本对应ACSII 表示的时候生效。\n\n# ### 文本字符清理一个最主要的问题应该是运行的性能。一般来讲,代码越简单运行越快。对于简单的替换操作, str.replace() 方法通常是最快的,甚至在你需要多次调用的时候\n#\n# ### 另一方面,如果你需要执行任何复杂字符对字符的重新映射或者删除操作的话,tanslate() 方法会非常的快。\n\ndef clean_spaces(s):\n s = s.replace('\\r', '')\n s = s.replace('\\t', ' ')\n s = s.replace('\\f', ' ')\n return s\n\n\n# # 字符串的对齐\n\ntext = 'Hello World'\n\ntext\n\ntext.ljust(20)\n\ntext.rjust(20)\n\ntext.center(20)\n\ntext.rjust(20,'=')\n\ntext.center(20,'*')\n\nformat(text, '>20')\n\nformat(text, '<20')\n\nformat(text, '^20')\n\nformat(text, '=>20s')\n\nformat(text, '*^20s')\n\n'{:>10s} {:>10s}'.format('Hello', 'World')\n\n# ### format() 函数的一个好处是它不仅适用于字符串。它可以用来格式化任何值,使得它非常的通用。\n\nx = 1.2345\n\nformat(x, '>10')\n\nformat(x,'<10.2f')\n\n# # 合并拼接字符串\n\nparts = ['Is', 'Chicago', 'Not', 'Chicago?']\n\n' '.join(parts)\n\n','.join(parts)\n\n# # 字符串中插入变量\n\ns = '{name} has {n} messages'\ns.format(name='Guido', n=37)\n\nname = 'tami'\nn = 520\ns.format_map(vars())\n\n\n# +\n# vars() 也是适用于对象实例\nclass Info:\n def __init__(self, name, n):\n self.name = name\n self.n = n\n\na= Info('kotoshi', 2)\ns.format_map(vars(a))\n\n\n# +\n# 处理缺省值\nclass safesub(dict):\n def __missing__(self, key):\n return '{' + key + '}'\n\ndel n\ns.format_map(safesub(vars()))\n# -\n\nimport sys\ndef sub(text):\n return text.format_map(safesub(sys._getframe(1).f_locals))\n\n\nname = 'guido'\nn =37\nprint(sub('hello {name}'))\n\nprint(sub('you have {n} messages'))\n\nprint(sub('your favorite color is {color}'))\n\ncolor = 'yellow'\n\nprint(sub('your favorite color is {color}'))\n\n# ### 比较老的一些使用方法\n\n'%(name) has %(n) messages.' % vars()\n\nname\n\nn\n\nimport string\ns= string.Template('$name has $n messages')\ns.substitute(vars())\n\n# # 以指定列宽格式化字符串\n\ns = \"Look into my eyes, look into my eyes, the eyes, the eyes, \\\nthe eyes, not around the eyes, don't look around the eyes, \\\nlook into my eyes, you're under.\"\n\nimport textwrap\nprint(textwrap.fill(s, 70))\n\nprint(textwrap.fill(s, 40))\n\nprint(textwrap.fill(s, 40, initial_indent = ' '))\n\nimport os\nos.get_terminal_size()\n# 在vscode里可以, 可能jupyter不适用\n\n# # 在字符串里处理html xml\n\ns = 'Elements are written as \"text\".'\nimport html\nprint(s)\n\nprint(html.escape(s))\n\nprint(html.escape(s, quote=False))\n\ns = 'Spicy Jalapeño'\n\ns.encode('ascii', errors='xmlcharrefreplace')\n\ns = 'Spicy "Jalapeño".'\nfrom html.parser import HTMLParser\np = HTMLParser()\np.unescape(s)\n\n\nt = 'The prompt is >>>'\nfrom xml.sax.saxutils import unescape\nunescape(t)\n\n# # 字符串令牌解析\n\ntext = 'foo = 23 + 42 * 10'\n\nimport re\nNAME = r'(?P[a-zA-Z_][a-zA-Z_0-9]*)'\nNUM = r'(?P\\d+)'\nPLUS = r'(?P\\+)'\nTIMES = r'(?P\\*)'\nEQ = r'(?P=)'\nWS = r'(?P\\s+)'\nmaster_pat = re.compile('|'.join([NAME, NUM, PLUS, TIMES, EQ, WS]))\n\nscanner = master_pat.scanner('foo = 42')\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n# _.lastgroup, _.group()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\nfrom collections import namedtuple\ndef generate_tokens(pat, text):\n Token = namedtuple('Token', ['type', 'value'])\n scanner = pat.scanner(text)\n for m in iter(scanner.match, None):\n yield Token(m.lastgroup, m.group())\n# Example use\nfor tok in generate_tokens(master_pat, 'foo = 42'):\n print(tok)\n\ntokens = (tok for tok in generate_tokens(master_pat, text)\n if tok.type != 'WS')\nfor tok in tokens:\n print(tok)\n\n# +\nPRINT = r'(Pprint)'\nNAME = r'(P[a-zA-Z][a-zA-z0-9]*)'\nmaster_pat = re.compile('|'.join([PRINT, NAME]))\n\nfor test in generate_tokens(master_pat, 'printer'):\n print(test)\n# -\n\n# # 实现一个简单的递归下降分析器\n\n# +\n# #!/usr/bin/env python\n# -*- encoding: utf-8 -*-\n\"\"\"\nTopic: 下降解析器\nDesc: \n\"\"\"\nimport re\nimport collections\n\n# Token specification\nNUM = r'(?P\\d+)'\nPLUS = r'(?P\\+)'\nMINUS = r'(?P-)'\nTIMES = r'(?P\\*)'\nDIVIDE = r'(?P/)'\nLPAREN = r'(?P\\()'\nRPAREN = r'(?P\\))'\nWS = r'(?P\\s+)'\n\nmaster_pat = re.compile('|'.join([NUM, PLUS, MINUS, TIMES, \n DIVIDE, LPAREN, RPAREN, WS]))\n\n# tokenizer\nToken = collections.namedtuple('Token', ['type', 'value'])\n\ndef generate_tokens(text):\n scanner = master_pat.scanner(text)\n for m in iter(scanner.match, None):\n tok = Token(m.lastgroup, m.group())\n if tok.type !='WS':\n yield tok\n\n\n# Parser\nclass ExpressionEvaluator:\n '''\n Implementation of a recursive descent parser. Each method\n implements a single grammar rule. Use the ._accept() method\n to test and accept the current lookahead token. Use the ._expect()\n method to exactly match and discard the next token on on the input\n (or raise a SyntaxError if it doesn't match).\n'''\n\n def parse(self, text):\n self.tokens = generate_tokens(text)\n self.tok = None # Last symbol consumed\n self.nexttok = None # Next symbol tokenized\n self._advance() # Load first lookahead token\n return self.expr()\n \n def _advance(self):\n 'Advance one token ahead'\n self.tok, self.nexttok = self.nexttok, next(self.tokens, None)\n\n def _accept(self, toktype):\n 'Test and consume the next token if it matches toktype'\n if self.nexttok and self.nexttok.type ==toktype:\n self._advance()\n return True\n else: \n return False\n \n def _expect(self, toktype):\n 'Consume next token if it matches toktype or raise SyntaxError'\n if not self._accept(toktype):\n raise SyntaxError('Ex[ected ' + toktype)\n \n # Grammer rules follow\n def expr(self):\n \"expression ::= term {('+'|'-') term }*\"\n exprval = self.term()\n while self._accept('PLUS') or self._accept('MINUS'):\n op = self.tok.type\n right = self.term()\n if op == 'PLUS':\n exprval += right\n elif op =='MINUS':\n exprval -= right\n return exprval\n \n def term(self):\n \"term ::= factor { ('*'|'/') factor }* \"\n termval = self.factor()\n while self._accept('TIMES') or self._accept('DIVIDE'):\n op = self.tok.type\n right = self.factor()\n if op == 'TIMES':\n termval *= right\n elif op == 'DIVIDE':\n termval /= right\n return termval\n \n def factor(self):\n \" factor ::= NUM | ( expr )\"\n if self._accept('NUM'):\n return int(self.tok.value)\n elif self._accept('LPAREN'):\n exprval = self.expr()\n self._expect('RPAREN')\n return exprval\n else:\n raise SyntaxError('Expected NUMBER or LPAREN')\n\n\ndef descent_parser():\n e = ExpressionEvaluator()\n print(e.parse('2'))\n print(e.parse('2+3'))\n print(e.parse('2+3*4'))\n print(e.parse('2+(3+4)*5'))\n\n\nif __name__ == '__main__':\n descent_parser()\n# -\n\n# ### 下面的代码不能运行在ipython上, 必须存为一个.py文件\n\n# +\n# plyexample.py\n#\n# Example of parsing with PLY\n\nfrom ply.lex import lex\nfrom ply.yacc import yacc\n\n# Token list\ntokens = [ 'NUM', 'PLUS', 'MINUS', 'TIMES', 'DIVIDE', 'LPAREN', 'RPAREN' ]\n\n# Ignored characters\n\nt_ignore = ' \\t\\n'\n\n# Token specifications (as regexs)\nt_PLUS = r'\\+'\nt_MINUS = r'-'\nt_TIMES = r'\\*'\nt_DIVIDE = r'/'\nt_LPAREN = r'\\('\nt_RPAREN = r'\\)'\n\n# Token processing functions\ndef t_NUM(t):\n r'\\d+'\n t.value = int(t.value)\n return t\n\n# Error handler\ndef t_error(t):\n print('Bad character: {!r}'.format(t.value[0]))\n t.skip(1)\n\n# Build the lexer\nlexer = lex()\n\n# Grammar rules and handler functions\ndef p_expr(p):\n '''\n expr : expr PLUS term\n | expr MINUS term\n '''\n if p[2] == '+':\n p[0] = p[1] + p[3]\n elif p[2] == '-':\n p[0] = p[1] - p[3]\n\ndef p_expr_term(p):\n '''\n expr : term\n '''\n p[0] = p[1]\n\ndef p_term(p):\n '''\n term : term TIMES factor\n | term DIVIDE factor\n '''\n if p[2] == '*':\n p[0] = p[1] * p[3]\n elif p[2] == '/':\n p[0] = p[1] / p[3]\n\ndef p_term_factor(p):\n '''\n term : factor\n '''\n p[0] = p[1]\n\ndef p_factor(p):\n '''\n factor : NUM\n '''\n p[0] = p[1]\n\ndef p_factor_group(p):\n '''\n factor : LPAREN expr RPAREN\n '''\n p[0] = p[2]\n\ndef p_error(p):\n print('Syntax error')\n\nparser = yacc()\n\nif __name__ == '__main__':\n print(parser.parse('2'))\n print(parser.parse('2+3'))\n print(parser.parse('2+(3+4)*5'))\n# -\n\n# # 字节字符串上的操作\n\ndata = bytearray(b'Hello World')\n\ndata[:5]\n\ndata.replace(b'Hello', b'Hello Cruel')\n\ndata = b'FOO:BAR,SPAM'\nimport re\nre.split('[:,]', data)\n\nre.split(b'[:,]', data)\n\n# ### some differences with standard string \n\na = 'Hello World'\na[0]\n\nb = b'Hello World'\nb[0]\n\nprint(a)\n\nprint(b) # Observe b'...'\n\n# +\n# formatting ? \nb'%10s %10d %10.2f' % (b'ACME', 100, 490.1)\n\n# It said there's no formatting for bytestring, but somehow formatting works = =;\n\n# +\nwith open ('halape\\xf10.txt', 'w') as f:\n f.write('spicy')\n \nimport os\nos.listdir('.')\n# -\n\nos.listdir(b'.')\n\n# # 第三章\n# # 数字日期和时间\n\n# ## 数字的四舍五入\n\nround(1.23,1)\n\nround(1.27,1)\n\nround(1.32455, 3)\n\n# ## 执行精确的浮点数运算\n\na = 4.2\nb = 2.1\na + b \n\n(a+b) == 6.3\n\nfrom decimal import Decimal\na = Decimal('4.2')\nb = Decimal('2.1')\na + b \n\n(a + b ) == Decimal('6.3')\n\n# Decimal can allow you to control the comoputation, like digit and round\na = Decimal('1.3')\nb = Decimal('1.7')\nprint(a/b)\n\nfrom decimal import localcontext\nwith localcontext() as ctx:\n ctx.prec = 3\n print(a/b)\n\nnums = [1.23e+18, 1, -1.23e+18]\nsum(nums)\n\nimport math\nmath.fsum(nums)\n\n# # 2 8 16 进制\n\nx = 1234\n\n\nbin(x)\n\nhex(x)\n\noct(x)\n\nformat(x, 'b')\n\nformat(x, 'x')\n\nformat(x, 'o')\n\n# ## 字节到大整数的打包与解包\n\n# ## 复数的数学运算\n\na = complex(2,4)\nb =3 - 5j\n\na\n\nb\n\na.real\n\na.imag\n\na + b\n\nc = 1 + 1j\n\nc\n\n\nabs(c)\n\nimport cmath\ncmath.sqrt(-1)\n\n# ## 无穷大与NaN\n\na = float('inf')\n\nb = float('-inf')\n\nc = float('nan')\n\na \n\nb \n\nc \n\nmath.isnan(c)\n\nmath.isinf(b)\n\n# ## 分数运算\n\nfrom fractions import Fraction\na = Fraction(2,3)\nb = Fraction(1,6)\n\nprint(a+b)\n\na.numerator\n\na.denominator\n\nc = 0.667\nFraction(c).limit_denominator(10)\n\nx = 3.75\n# y = Fraction()\nx.as_integer_ratio()\n\ny = Fraction(*x.as_integer_ratio())\n\ny\n\n# ## 大型数组运算 & 矩阵雨线性代数\n\n# ### 用numpy\n\n# # 随机选择\n\nimport random\nvalues = [1,2,3,4,5,6]\n\nrandom.choice(values)\n\nrandom.choice(values)\n\nrandom.sample(values,3)\n\nrandom.shuffle(values)\n\nvalues\n\nrandom.randint(0,10)\n\n# 生成0到1 范围内均匀分布的浮点数, 用random.random()\nrandom.random()\n\n# 获得 N 位随机二进制的整数\nrandom.getrandbits(3)\n\n\nrandom.seed(233)\nrandom.randint(1,100)\n\nrandom.randint(1,100)\n\n# +\n# random.uniform() random.gauss()\n# -\n\n# ### 在random 模块中的函数不应该用在和密码学相关的程序中。如果你确实需要类似的功能,可以使用ssl 模块中相应的函数。\n\n# # 基本的日期与时间转换\n\n# 表示一个时间段 用timedelta\nfrom datetime import timedelta\na = timedelta(days=2, hours=3)\nb = timedelta(hours=6.5)\na + b \n\n(a + b).seconds / 3600\n\n(a + b).hours\n\n(a + b).total_seconds() / 3600\n\nfrom datetime import datetime\na = datetime(2008, 8, 8)\n\na + timedelta(days=10)\n\nb = datetime(2023, 7, 4)\nb - a \n\nprint(datetime.today())\n\nprint(datetime.today() + timedelta(minutes= 10))\n\n# ### 如果要处理更加复杂的日期, 用dateutil\n\na \n\na + timedelta(months=1)\n\nfrom dateutil.relativedelta import relativedelta\na + relativedelta(months=+10)\n\nb\n\nb - a \n\nd = relativedelta(b,a)\n\nd \n\n# # 查找最后一个周五的日期\n\n# #!/usr/bin/env python\n# -*- encoding: utf-8 -*-\n\"\"\"\nTopic: 最后的周五\nDesc :\n\"\"\"\nfrom datetime import datetime, timedelta\nweekdays = ['Monday', 'Tuesday', 'Wednesday', 'Thursday',\n'Friday', 'Saturday', 'Sunday']\ndef get_previous_byday(dayname, start_date=None):\n if start_date is None:\n start_date = datetime.today()\n day_num = start_date.weekday()\n day_num_target = weekdays.index(dayname)\n days_ago = (7 + day_num - day_num_target) % 7\n if days_ago == 0:\n days_ago = 7\n target_date = start_date - timedelta(days=days_ago)\n return target_date\n\n\ndatetime.today()\n\nget_previous_byday('Friday')\n\nday_num = datetime.today().weekday()\n\nday_num\n\nday_num_target = 4\n\n# 4 is the index of Friday\ndays_ago = (7 + day_num - day_num_target) % 7\n\ndays_ago\n\ndatetime.today() - timedelta(days= days_ago)\n\nget_previous_byday('Tuesday', datetime(2023,7,1))\n\n# ### 如果你要像这样执行大量的日期计算的话,你最好安装第三方包python-dateutil来代替。\n\nfrom datetime import datetime\nfrom dateutil.relativedelta import relativedelta\nfrom dateutil.rrule import *\nd = datetime.now()\n\n\nd \n\n# next Friday\nprint(d + relativedelta(weekday=FR))\n\n# previous 2 Friday\nprint(d + relativedelta(weekday=FR(-2)))\n\n# # 计算当前月份的日期范围\n\nfrom datetime import datetime, date, timedelta\nimport calendar\ndef get_month_range(start_date=None):\n if start_date is None:\n start_date = date.today().replace(day=1)\n _, days_in_month = calendar.monthrange(start_date.year, start_date.month)\n end_date = start_date + timedelta(days=days_in_month)\n return (start_date, end_date)\n\n\na_day = timedelta(days=1)\nfirst_day, last_day = get_month_range()\nwhile first_day < last_day:\n print(first_day)\n first_day += a_day\n\ndate.today()\n\ndate.today().replace(day=1)\n\n\ndef date_range(start, stop, step):\n while start < stop:\n yield start\n start += step\n\n\nfor d in date_range(datetime(2023, 7, 1), datetime(2023, 8, 1), timedelta(hours=6)):\n print(d)\n\n# # 字符串转换为日期\n\nfrom datetime import datetime\ntext = '2012-9-23'\ny = datetime.strptime(text, '%Y-%m-%d')\nz = datetime.now()\ndiff = z - y\n\ndiff\n\nz\n\n# 性能差,可以自己写一个日期的解析\nnice_z = datetime.strftime(z, '%A %B %d, %Y')\n\nnice_z\n\n# # 结合时区的日期操作\n\nfrom datetime import datetime\nfrom pytz import timezone\nd = datetime(2012, 12, 21, 9, 30, 0)\nprint(d)\n\n# localize the date for Chicago\ncentral = timezone('US/Central')\nloc_d = central.localize(d)\n\nloc_d\n\nbang_d = loc_d.astimezone(timezone('Asia/Kolkata'))\n\nbang_d\n\n\n# ### 注意美国夏时令的转换 & pytz不建议使用了\n\n# # 第四章 迭代器与生成器\n\n# ## 手动遍历迭代器\n\ndef manual_iter():\n with open('./lines.txt') as f:\n try:\n while True:\n line = next(f)\n print(line, end='')\n except StopIteration:\n pass\n\n\nmanual_iter()\n\n\ndef manual_iter():\n with open('./lines.txt') as f:\n while True:\n line = next(f)\n if line is None:\n break\n print(line, end='')\n\n\nmanual_iter()\n\nitems = [1, 2, 3]\nit = iter(items)\n\nnext(it)\n\nnext(it)\n\nnext(it)\n\nnext(it)\n\n\n# ## 代理迭代\n\n# +\nclass Node:\n def __init__(self, value):\n self._value = value\n self._children = []\n \n def __repr__(self):\n return f'Node({self._value})'\n \n def add_child(self, node):\n self._children.append(node)\n \n def __iter__(self):\n return iter(self._children)\n\n\nif __name__ == '__main__':\n root = Node(0)\n child1 = Node(1)\n child2 = Node(2)\n root.add_child(child1)\n root.add_child(child2)\n \n for ch in root:\n print(ch)\n\n\n# -\n\n# ## 使用生成器创建新的迭代模式\n\n# +\ndef frange(start, stop, increment):\n x = start\n while x < stop:\n yield x\n x += increment\n \nfor n in frange(0,4,0.5):\n print(n)\n# -\n\nfrange(2,3,0.1)\n\n\n# ## 实现迭代器协议\n\n# +\nclass Node:\n def __init__(self, value):\n self._value = value\n self._children = []\n \n def __repr__(self):\n return f'Node({self._value})'\n \n def add_child(self, node):\n self._children.append(node)\n \n def __iter__(self):\n return iter(self._children)\n \n def depth_first(self):\n yield self\n for c in self:\n yield from c.depth_first()\n \n\nif __name__ == '__main__': \n root = Node(0)\n child1 = Node(1)\n child2 = Node(2)\n root.add_child(child1)\n root.add_child(child2)\n child1.add_child(Node(3))\n child1.add_child(Node(4))\n child2.add_child(Node(5))\n \n for ch in root.depth_first():\n print(ch)\n# -\n\n# # 反向迭代\n\na = [1,3,5,7]\nfor x in reversed(a):\n print(x)\n\nf = open('./lines.txt')\nfor line in reversed(list(f)):\n print(line, end='')\n\nf.close()\n\n\n# +\nclass Countdown:\n def __init__(self, start):\n self.start = start\n \n # forward iterator\n def __iter__(self):\n n = self.start\n while n > 0:\n yield n\n n -= 1\n \n # reverse iterator\n def __reversed__(self):\n n = 1\n while n <= self.start:\n yield n\n n += 1\n \n\nfor rr in reversed(Countdown(30)):\n print(rr)\n \nprint('*'*20) \n \nfor rr in Countdown(30):\n print(rr)\n \n# -\n\n# # 带有外部状态的生成器函数\n\n# ### 如果你想让你的生成器暴露外部状态给用户,别忘了你可以简单的将它实现为一个类,然后把生成器函数放到iter () 方法中过去。\n\nfrom collections import deque\nclass linehistory:\n def __init__(self, lines, histlen=3):\n self.lines = lines\n self.history = deque(maxlen=histlen)\n \n def __iter__(self):\n for lineno, line in enumerate(self.lines, 1):\n self.history.append((lineno, line))\n yield line\n\n def clear(self):\n self.history.clear()\n\n\nwith open('somefile.txt') as f:\n lines = linehistory(f)\n for line in lines:\n if 'python' in line:\n for lineno, hline in lines.history:\n print('*{}:{}'.format(lineno, hline), end='')\n\n\n# ### 没有看懂上面的结果\n\n# # 迭代器切片\n\n# +\ndef count(n):\n while True:\n yield n\n n += 1\n \nc = count(0)\nc[10:20]\n# -\n\nimport itertools\nfor x in itertools.islice(c, 10, 20):\n print(x)\n\n# ### islice 用过就废,如需再次访问,先放到一个列表里\n\n# # 跳过可迭代对象的开始部分\n\nfrom itertools import dropwhile\nwith open('/etc/passwd') as f:\n for line in dropwhile(lambda line: line.startswith('#'), f):\n print(line, end='')\n\nfrom itertools import islice\nitems = ['a', 'b', 'c', 1, 3, 5, 9]\nfor x in islice(items, 3, None):\n print(x)\n\n# # 排列组合的迭代\n\nitems = ['a', 'b', 'c']\nfrom itertools import permutations\nfor p in permutations(items):\n print(p)\n\nfor p in permutations(items, 2):\n print(p)\n\nfrom itertools import combinations\nfor c in combinations(items, 2):\n print(c)\n\nfrom itertools import combinations_with_replacement\nfor c in combinations_with_replacement(items, 3):\n print(c)\n\n# # 序列上索引值迭代\n\nmy_list = ['a', 'b', 'c']\nfor idx, val in enumerate(my_list):\n print(idx, val)\n\nfor idx, val in enumerate(my_list, 1):\n print(idx, val)\n\n\ndef parse_data(filename):\n with open(filename, 'rt') as f:\n for lineno, line in enumerate(f, 1):\n fields = line.split()\n try:\n count = int(fields[1])\n ...\n except ValueError as e:\n print('Line {}: Parse error: {}'.format(lineno, e))\n\n\n# +\nfrom collections import defaultdict\nword_summary = defaultdict(list)\nwith open('somefile.txt', 'r') as f:\n lines = f.readlines()\n\nfor idx, line in enumerate(lines):\n # Create a list of words in current line\n# print(line)\n# print(line.split())\n words = [w.strip().lower() for w in line.split()]\n for word in words:\n word_summary[word].append(idx)\n# -\n\nword_summary\n\nss = 'this '\nss.strip()\n\n# # 同时迭代多个序列\n\na = [1, 2, 3]\nb = ['w', 'x', 'y', 'z']\nfor i in zip(a,b):\n print(i)\n\nfrom itertools import zip_longest\nfor i in zip_longest(a,b):\n print(i)\n\nfor i in zip_longest(a, b, fillvalue=0):\n print(i)\n\nheaders = ['name', 'shares', 'price']\nvalues = ['ACME', 100, 490.1]\ns = dict(zip(headers, values))\n\ns\n\n# ### zip可以接受多于两个的序列的参数 & 返回的是一个迭代器\n\na = [1, 2, 3]\nb = [10, 11, 12]\nc = ['x','y','z']\nfor i in zip(a, b, c):\n print(i)\n\n# # 不同集合上元素的迭代\n\n# ### 多个容器, 但不想写多次循环\n\nfrom itertools import chain\na = [1,2,3,4]\nb = ['x', 'y', 'z']\nfor x in chain(a,b):\n print(x)\n\nfor x in a+b:\n ...\n# 这种要比上面的低效\n\n# # 创建数据处理管道\n\n# +\nimport os\nimport fnmatch\nimport gzip\nimport bz2\nimport re\n\ndef gen_find(filepat, top):\n '''\n Find all filenames in a directory tree that match a shell wildcard pattern \n '''\n for path, dirlist, filelist in os.walk(top):\n for name in fnmatch.filter(filelist, filepat):\n yield os.path.join(path, name)\n \n\ndef gen_opener(filenames):\n '''\n Open a ssequence of filenames one at a time producing a file object.\n The fle is closed immediately when proceeding t othe next iteration.\n '''\n for filename in filenames:\n if filename.endswith('.gz'):\n f = gzip.open(filename, 'rt')\n elif filename.endswith('.bz2'):\n f = bz2.open(filenamee, 'rt')\n else:\n f = open(filename, 'rt')\n yield f\n f.close()\n \ndef gen_concatenate(iterators):\n '''\n Chain a sequence of iterators together into a single sequence\n '''\n for it in iterators:\n yield from it\n \ndef gen_grep(pattern, lines):\n '''\n Look for a regex pattern in a sequence of lines\n '''\n pat = re.compile(pattern)\n for line in lines:\n if pat.search(line):\n yield line\n \n \n# -\n\nlognames = gen_find('access-log*', 'www')\nfiles = gen_opener(lognames)\nlines = gen_concatenate(files)\npylines = gen_grep('(?i)python', lines)\nfor line in pylines:\n print(line)\n\nlognames = gen_find('access-log*', 'www')\nfiles = gen_opener(lognames)\nlines = gen_concatenate(files)\npylines = gen_grep('(?i)python', lines)\nbytecolumn = (line.rsplit(None,1)[1] for line in pylines)\nbytes = (int(x) for x in bytecolumn if x != '-')\nprint('Total', sum(bytes))\n\n# # 展开嵌套的序列\n\n# +\n# 在vscode里可行\nfrom collections import Iterable\n\ndef flatten(items, ignore_types=(str, bytes)):\n for x in items:\n if isinstance(x, Iterable) and not isinstance(x, ignore_types):\n yield from flatten(x)\n else:\n yield x\n\nitems = [1,2,[3,4, [5,6], 7], 8]\nfor x in flatten(items):\n print(x)\n# -\n\nitems_2 = ['Dave', 'Paula', ['Thomas', 'Lewis']] \nfor x in flatten(items_2):\n print(x)\n\n# # 顺序迭代合并后的排序迭代对象\n\nimport heapq\nx = [1,4,7,10]\ny= [2,5,6,11]\nfor z in heapq.merge(x,y):\n print(z)\n\na = ['a', 'c', 'z']\nb = ['j', 'k' ,'b']\nfor c in heapq.merge(a,b):\n print(c)\n\nwith open('sorted_file_1', 'rt') as file1, \\\n open('sorted_file_2', 'rt') as file2, \\\n open('merged_file', 'wt') as outf:\n for line in heapq.merge(file1, file2):\n outf.write(line)\n\n# # 迭代器代替while 无限循环\n\nimport sys\nf = open('/etc/passwd')\nfor chunk in iter(lambda: f.read(10), ''):\n n = sys.stdout.write(chunk)\n\n\n","repo_name":"LightRoastKenyaPeaberry/ipython_notes","sub_path":"python_cookbook/python_cookbook_1_to_4.ipynb","file_name":"python_cookbook_1_to_4.ipynb","file_ext":"py","file_size_in_byte":50199,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"34151305078","text":"# # [AI 이노베이션 스퀘어] 11기 기본 저녁반\n# - github: https://github.com/hysKim1/AI_Fundamental\n# - [파이썬 공식문서]https://docs.python.org/ko/3/\n# - [Numpy 공식문서]https://numpy.org/devdocs/\n# - [ML]https://developers.google.com/machine-learning/crash-course/ml-intro?hl=ko\n# - [DL]https://www.tensorflow.org/\n# ---\n#\n# * [1. layers API ](#layers )
\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"V5Behma77t2H\" outputId=\"65cd96aa-a98f-4e61-aa3a-f10f3e00aa8b\"\ndef sol(N):\n S = list(map(int,input().split()))\n dp = S.copy()\n for i in range(1,N):\n Max = [0]\n for j in range(i):\n if S[i] > S[j]:\n Max.append(dp[j])\n dp[i] += max(Max)\n return max(dp)\n\nprint(sol(int(input())))\n","repo_name":"Kang-Yoonah/BaekJoon","sub_path":"Silver/11055.ipynb","file_name":"11055.ipynb","file_ext":"py","file_size_in_byte":656,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"18780268394","text":"# + id=\"5s_uheqcdmU5\" colab_type=\"code\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1788} outputId=\"e961f0cc-ff66-4e88-c487-ed99e5f760cb\" executionInfo={\"status\": \"ok\", \"timestamp\": 1538983182426, \"user_tz\": -480, \"elapsed\": 35829, \"user\": {\"displayName\": \"Justin Wu\", \"photoUrl\": \"\", \"userId\": \"12321085329230288327\"}}\nfrom bs4 import BeautifulSoup\nimport requests\nimport re\nimport os\nimport json\n\n\ndef get_soup(url,header):\n return BeautifulSoup(requests.get(url, headers=header).content,'html.parser')\n\nquery = input(\"query: \")# you can change the query for the image here\nimage_type=\"ActiOn\"\nquery= query.split()\nquery='+'.join(query)\nurl=\"https://www.google.co.in/search?q=\"+query+\"&source=lnms&tbm=isch\"\nprint('query_url: {}'.format(url))\n\n#add the directory for your image here\nDIR=\"Images\"\nheader={'User-Agent':\"Mozilla/5.0 (Windows NT 6.1; WOW64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/43.0.2357.134 Safari/537.36\"\n}\nsoup = get_soup(url,header)\n\n\nActualImages=[]# contains the link for Large original images, type of image\nfor a in soup.find_all(\"div\",{\"class\":\"rg_meta\"}):\n link , Type =json.loads(a.text)[\"ou\"] ,json.loads(a.text)[\"ity\"]\n ActualImages.append((link,Type))\n\nprint (\"Downloading total {} images\".format(len(ActualImages)))\n\nif not os.path.exists(DIR):\n os.mkdir(DIR)\nDIR = os.path.join(DIR, query.split()[0])\n\nif not os.path.exists(DIR):\n os.mkdir(DIR)\n \n###print images\nfor i , (img , Type) in enumerate( ActualImages):\n try:\n\n img_data = requests.get(img, stream=True).content\n cntr = len([i for i in os.listdir(DIR) if image_type in i]) + 1\n print (cntr, img)\n if len(Type)==0:\n f = open(os.path.join(DIR , image_type + \"_\"+ str(cntr)+\".jpg\"), 'wb')\n else :\n f = open(os.path.join(DIR , image_type + \"_\"+ str(cntr)+\".\"+Type), 'wb')\n\n\n f.write(img_data)\n f.close()\n except Exception as e:\n print (\"could not load : \"+img)\n print (e)\n","repo_name":"CloudMile/allianz_courses","sub_path":"Intro_to_ML_NLP_workshop/google _crawler.ipynb","file_name":"google _crawler.ipynb","file_ext":"py","file_size_in_byte":2007,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"1666058090","text":"# ### Entrenando un algoritmo de machine learning\n#\n# Para el presente entregable seleccioné un dataset obtenido del Instituto Nacional de Estadistica de Uruguay (INE).\n#\n# El mismo corresponde a la encuesta continua de hogares realizada en el año 2017.\n#\n# El objetivo del analisis sera trabajar con predicciones del ingreso en relación de dependencia de los encuestados.\n#\n# Archivos utilizados:\n#\n# P_2017_Terceros.sav -> el dataset que contiene los datos recopilados en la encuesta continua de hogares.\n# Dicccionario de Variables ECH 2017 -> funciona como anexo para determinar a que corresponde la codificación de cada columna del dataset.\n\nimport numpy as np\nimport pandas as pd\nfrom sklearn.model_selection import train_test_split\nfrom sklearn import linear_model\nfrom sklearn.metrics import mean_squared_error\nfrom sklearn.compose import make_column_transformer\nfrom sklearn.preprocessing import OneHotEncoder, StandardScaler\n\ndfP2017 = pd.read_spss(r'..\\\\data\\\\P_2017_Terceros.sav')\n\ncolumns = dfP2017.columns.values.tolist()\nprint(columns)\n\ndfP2017.head(2)\n\n# A continuación se definen las variables a utilizar en el analisis y se genera un dataset independiente para realizar modificaciones sin afectar al dataset original\n\n# +\ndf_ysos = pd.DataFrame()\n\n# variable de respuesta\ndf_ysos['ysos_dep'] = dfP2017['g126_1'] + dfP2017['g126_2'] + dfP2017['g126_3'] + dfP2017['g126_4'] \\\n + dfP2017['g134_1'] + dfP2017['g134_2'] + dfP2017['g134_3'] + dfP2017['g134_4']\n\n# caracteristicas personales\ndf_ysos['edad'] = dfP2017['e27']\ndf_ysos['sexo'] = dfP2017['e26']\ndf_ysos['estrato'] = dfP2017['estred13']\ndf_ysos['ascendencia_negra'] = dfP2017['e29_1']\ndf_ysos['estado_civil'] = dfP2017['e36']\ndf_ysos['asiste_comedor']=dfP2017['e559']\ndf_ysos['seguro_privado']=dfP2017['e45_3']\ndf_ysos['pobre']=dfP2017['pobre06']\ndf_ysos['indigente']=dfP2017['indigente06']\n\n## educación\ndf_ysos['anios_educ'] = dfP2017['e51_2'] + dfP2017['e51_3'] + dfP2017['e51_4'] + dfP2017['e51_5'] + dfP2017['e51_6'] \\\n + dfP2017['e51_7'] + dfP2017['e51_8'] + dfP2017['e51_9'] + dfP2017['e51_10'] + dfP2017['e51_11']\n\n## relación de empleo\ndf_ysos['antiguedad'] = dfP2017['f88_2']\ndf_ysos['categoria_ocup'] = dfP2017['f73']\ndf_ysos['dimension_empresa']=dfP2017['f77']\n\ndf_ysos.info()\n# -\n\n# ##### DETALLE DE VARIABLES A UTILIZAR\n#\n# ysos_dep -> ingresos dependientes de cada persona\n#\n# edad -> edad de la persona\n#\n# sexo -> sexo declarado de la persona\n#\n# estrato -> estrato social al que se asocia al encuestado\n#\n# ascendencia_negra -> dato booleano que indica si el encuestado tiene ascendencia \n# negra\n#\n# estado_civil -> estado civil declarado del encuestado\n#\n# asiste_comedor -> dato booleano que indica si el encuestado consume alguna de sus \n# comidas diarias en comedor público\n#\n# seguro_privado -> dato booleano que indica si el encuestado tiene asistencia medica \n# privada\n#\n# pobre -> dato que indica si el encuestado esta por debajo de la franja de pobreza establecida por el INE\n#\n# indigente -> dato que indica si el encuestado se clasifica como indigente\n#\n# anios_educ -> años de educación formal que tiene el encuestado\n#\n# antiguedad -> antigüedad laboral del encuestado\n#\n# categoria_ocup -> categoria ocupacional del encuestado\n#\n# dimension_empresa -> cantidad de trabajadores activos de la empresa a la cual pertenence el encuestado\n\n# genero archivo con datos a utilizar para alojar en Github, ya que el archivo .sav es muy pesado y no se puede guardar incluir al hacer commit a github.\nexport = '../data/ECH2017.xlsx'\ndf_ysos.to_excel(export, index=False)\n\ndf_ysos['dimension_empresa'].dtype\n\ndf_ysos.head(2)\n\n#se eliminan del dataset todos los ingresos menores a 0\ndf_ysos = df_ysos[df_ysos['ysos_dep'] > 0]\n\ndf_ysos = df_ysos.loc[(df_ysos['categoria_ocup'] == \"Asalariado/a privado/a\") | (df_ysos['categoria_ocup'] == \"Asalariado/a público/a\")]\n\n#transformación logaritmica de la variable ingresos\ndf_ysos['l_ysos_dep'] = np.log(df_ysos['ysos_dep'])\ndf_l_ysos = df_ysos.drop(columns=['ysos_dep'])\ndf_l_ysos.head(2)\n\n\n# A continuacón se realizan transformaciones a ciertas columnas para contar con la información agrupada de una forma que permita realizar analisis de forma más eficiente.\n\n# +\ndef map_estrato(estrato):\n if estrato in ['Montevideo Alto']:\n return 'Mdeo - Alto'\n elif estrato in ['Montevideo Medio', 'Montevideo Medio Bajo', 'Montevideo Medio Alto']:\n return 'Mdeo - Medio'\n elif estrato in ['Montevideo Bajo']:\n return 'Mdeo - Bajo'\n elif estrato in ['Zona Metropolitana']:\n return 'Metropolitana'\n elif estrato in ['Centro Norte', 'Centro Sur', 'Costa Este', 'Litoral Norte', 'Litoral Sur','Norte']:\n return 'Interior'\n\n# se crea nueva columna que contenga los nombres de los grupos\ndf_l_ysos['estrato'] = df_l_ysos['estrato'].apply(map_estrato)\n\n\n# +\ndef map_dimenisonempresa(dimension):\n if dimension in ['Una persona', 'De 2 a 4 personas', 'De 5 a 9 personas', 'De 10 a 19 personas']:\n return 'Empresa_chica'\n elif dimension in ['De 20 a 49 personas']:\n return 'Empresa_mediana'\n elif dimension =='De 50 personas o más':\n return 'Empresa_grande'\n elif dimension ==0.0:\n return 'N/A'\n\n# se crea nueva columna que contenga los nombres de los grupos\ndf_l_ysos['dimension_empresa'] = df_l_ysos['dimension_empresa'].apply(map_dimenisonempresa)\n# -\n\ndf_l_ysos.head(2)\n\n# +\n#Se agrupan las variables separandolas en categoricas y numericas.\n\ncategorical_columns = ['sexo', 'categoria_ocup','asiste_comedor','ascendencia_negra','estrato','seguro_privado', 'dimension_empresa']\nnumerical_columns = ['edad', 'anios_educ', 'antiguedad']\n\n# Se genera la función transformadora\ncolumn_transform = make_column_transformer(\n (OneHotEncoder(drop='if_binary'), categorical_columns), # cuando son dos categorías, elimina una\n (StandardScaler(), numerical_columns), # comentado mientras no se decida estandarizar\n remainder='passthrough', # deja el resto de las columnas sin transformar\n verbose_feature_names_out=False)\n\n# +\ndf_l_ysos_transf = pd.DataFrame(\n column_transform.fit_transform(df_l_ysos), \n columns=column_transform.get_feature_names_out()\n)\n\ndf_l_ysos_transf.head()\n# -\n\npredictoras = df_l_ysos_transf.drop('l_ysos_dep', axis=1)\n\ndf_l_ysos.head(2)\n\nX_train, X_test, y_train, y_test = train_test_split(predictoras,\n df_l_ysos_transf['l_ysos_dep'], \n test_size=0.20, \n random_state=1234)\n\n# #### Regresión Lineal\n#\n# A continuación se plantea el modelo propiamente dicho, para una primera instancia se trabaja solamente con la variable que toma los años de educación del encuestado.\n\nreg = linear_model.LinearRegression()\n\n# +\nmodelo_X_train = X_train[['anios_educ']]\nmodelo_X_test = X_test[['anios_educ']]\n\nmodelo = reg.fit(modelo_X_train, y_train)\n\n# +\n# predicción en train\nmodelo_y_train_pred = modelo.predict(modelo_X_train)\n\n# predicción en test\nmodelo_y_test_pred = modelo.predict(modelo_X_test)\n# -\n\nmodelo_mse_train = mean_squared_error(y_train, modelo_y_train_pred)\nprint(f\"Error cuadratico medio del modelo en train: \", modelo_mse_train)\n\nmodelo_mse_test = mean_squared_error(y_test, modelo_y_test_pred)\nprint(f\"Error cuadratico medio del modelo en test: \", modelo_mse_test)\n\n# ##### Actualización 31/05/2023\n#\n# Se agregan las variables 'edad' y 'antiguedad' con el objetivo de reducir los valores de error cuadratico medio del modelo tanto en train como en test.\n\n# +\n#variables_adicionales = ['edad']+['sexo']\n\nmodelo_X_train2 = X_train[['anios_educ', 'edad', 'antiguedad']]\nmodelo_X_test2 = X_test[['anios_educ', 'edad', 'antiguedad']]\n\nmodelo = reg.fit(modelo_X_train2, y_train)\n\n# +\n# predicción en train\nmodelo_y_train_pred2 = modelo.predict(modelo_X_train2)\n\n# predicción en test\nmodelo_y_test_pred2 = modelo.predict(modelo_X_test2)\n\n# -\n\nmodelo_mse_train2 = mean_squared_error(y_train, modelo_y_train_pred2)\nprint(f\"Error cuadratico medio del modelo en train: \", modelo_mse_train2)\n\nmodelo_mse_test2 = mean_squared_error(y_test, modelo_y_test_pred2)\nprint(f\"Error cuadratico medio del modelo en test: \", modelo_mse_test2)\n\n# Efectivamente, agregando las variables 'edad' y 'sexo' se generan mejoras en los resultados del modelo.\n","repo_name":"MartinMF88/Tarea-machine-learning","sub_path":"src/Desafío_EvaluaciónML_Martin_ Martinez.ipynb","file_name":"Desafío_EvaluaciónML_Martin_ Martinez.ipynb","file_ext":"py","file_size_in_byte":8406,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"15007352664","text":"# +\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport more_itertools\n\nfrom keras.models import Sequential\nfrom keras.layers import LSTM, Dense, Flatten, ConvLSTM2D\n\nfrom sklearn.metrics import mean_squared_error\n\n# -\n\n# # Preparação dos dados\n\n# Dividir uma sequência univariada em amostras\n\n# +\nX = np.linspace(0, 5*np.pi, 100, endpoint=True)\nC = np.cos(X)\n\n# transform numpy array to list\nC = C.tolist()\n\nplt.plot(X, C)\nplt.show()\n# -\n\nprint(C)\n\n\n# Podemos dividir a sequência em vários padrões de entrada / saída chamados amostras, onde três etapas de tempo são usadas como entrada e uma etapa de tempo é usada como saída para a previsão de uma etapa que está sendo aprendida.\n\n# +\n# Função para dividir uma sequência univariada em amostras para o método de janelas\n# X - amostras de entrada; y - variável que será predita\n\ndef split_sequence(sequence, n_steps):\n\tX, y = list(), list()\n\tfor i in range(len(sequence)):\n\t\tend_ix = i + n_steps\n\t\tif end_ix > len(sequence)-1:\n\t\t\tbreak\n\t\tseq_x, seq_y = sequence[i:end_ix], sequence[end_ix]\n\t\tX.append(seq_x)\n\t\ty.append(seq_y)\n\treturn np.array(X), np.array(y)\n\n# +\n# Escolha do numero de passos no tempo\nn_steps = 4\n\nX, y = split_sequence(C, n_steps)\n\nfor i in range(len(X)):\n\tprint(X[i], y[i])\n# -\n\n# ## Reshape\n\n# Primeiro, devemos remodelar o conjunto de dados de treinamento na estrutura esperada de [samples, timesteps] para [samples, timesteps, rows, cols, channels].\n\nX.shape\n\nn_features = 1 # série temporal univariada (uma variável de predição)\nn_seq = 2\nn_steps = 2\nX = X.reshape((X.shape[0], n_seq, 1, n_steps, n_features))\n\nX.shape\n\n# ## Divisão da série temporal em treino e teste\n\n# As 60 primeiras linhas como treino\nX_train = X[0:60]\n\n# O restante das linhas como teste\nX_test = X[60:]\n\n# Analogamente para a variável preditora\n\ny_train = y[0:60]\n\ny_test = y[60:]\n\n# # Modelos LSTM para séries temporais univariadas\n\n# Nesse problema de previsão de cosseno, tratamos de um caso de série temporal univariada. Sendo assim, podemos aplicar os seguintes modelos LSTM:\n#\n# * Vanilla LSTM\n# * Stacked LSTM\n# * Bidirectional LSTM\n# * CNN LSTM\n# * ConvLSTM\n\n# ## Convolutional LSTM - ConvLSTM\n\n# Uma extensão adicional da abordagem CNN-LSTM é realizar as convoluções do CNN (por exemplo, como o CNN lê os dados da sequência de entrada) como parte do LSTM para cada etapa de tempo.\n#\n# Essa combinação é chamada de LSTM convolucional ou, abreviadamente, ConvLSTM e, como o CNN-LSTM, também é usada para dados espaço-temporais.\n#\n# Ao contrário de um LSTM que lê os dados diretamente para calcular o estado interno e as transições de estado, e ao contrário do CNN-LSTM que está interpretando a saída dos modelos CNN, o ConvLSTM está usando convoluções diretamente como parte da leitura de entrada nas próprias unidades LSTM .\n#\n# A biblioteca Keras fornece a classe ConvLSTM2D que suporta o modelo ConvLSTM para dados 2D.\n\n# The ConvLSTM2D class, by default, expects input data to have the shape:\n\n# [samples, timesteps, rows, cols, features]\n\n# Onde cada etapa de tempo de dados é definida como uma imagem de pontos de dados (linhas * colunas).\n\n# ### Definindo o modelo ConvLSTM\n\n# Definindo o modelo\nmodel = Sequential()\nmodel.add(ConvLSTM2D(filters=64, kernel_size=(1,2), activation='relu', input_shape=(n_seq, 1, n_steps, n_features)))\nmodel.add(Flatten())\nmodel.add(Dense(1))\nmodel.compile(optimizer='adam', loss='mse')\n\n# fit model\nmodel_history = model.fit(X_train, y_train, epochs=1000, verbose=0)\n\nplt.plot(model_history.history['loss'])\nplt.title('Convolutional LSTM loss')\nplt.ylabel('loss')\nplt.xlabel('epoch')\nplt.legend(['train'], loc='upper right')\nplt.yscale('log')\nplt.show()\n\n# model prediction\nyhat = model.predict(X_test, verbose=0)\n\n#model evaluation\nmean_squared_error(y_test,yhat)\n\nplt.plot(y_test, label='test')\nplt.plot(yhat, label='Conv2D LSTM prediction')\nplt.legend()\n\n# # Observações\n#\n# - Imports: LSTM, Dense, Flatten, ConvLSTM2D\n# - Número de passos no tempo: 4\n# - Reshape a série temporal de [samples, timesteps] para [samples, timesteps, rows, columns, features]\n# - Treino realizado com 1000 epochs foi rápido, e o erro foi bem baixo, como pode ser visto na figura acima\n","repo_name":"bruna-capistrano/TrabalhoPPA","sub_path":"lstm/conv_lstm.ipynb","file_name":"conv_lstm.ipynb","file_ext":"py","file_size_in_byte":4247,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"32231656518","text":"import numpy as np\nfrom sklearn.datasets import load_iris\nfrom sklearn import tree\niris = load_iris()\n\n\nfor i in range(len(iris.target_names)):\n print(\"Target:\\t\", iris.target_names[i], \"\\tValue:\\t\", i)\n\nprint(\"Features: \", \", \".join(e for e in iris.feature_names))\n\nfor i in range(len(iris.target)):\n print(i, iris.data[i], iris.target[i])\n\ntest_idx = [0, 50, 100] #keep some testing data around\n\n#training data\ntrain_target = np.delete(iris.target, test_idx)\ntrain_data = np.delete(iris.data, test_idx, axis=0)\n\n#testing data\ntest_target = iris.target[test_idx]\ntest_data = iris.data[test_idx]\n\nclf = tree.DecisionTreeClassifier()\nclf.fit(train_data, train_target)\n\nprint(test_target)\nprint(clf.predict(test_data))\n\n#visualize here\nfrom sklearn.externals.six import StringIO\nfile = open('tree.dot', 'w+')\ntree.export_graphviz(clf,\n out_file=file,\n feature_names=iris.feature_names,\n class_names=iris.target_names,\n filled=True, rounded=True,\n impurity=False)\nfile.close()\n\n# # Visualize The Data\n# `$ xdot tree.dot`\n\n\n","repo_name":"Stuart4/Machine-Learning-Journey","sub_path":"lab2/Iris.ipynb","file_name":"Iris.ipynb","file_ext":"py","file_size_in_byte":1121,"program_lang":"python","lang":"en","doc_type":"code","stars":3,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"7672155754","text":"# !pip install --quiet --no-cache-dir h3ronpy\n\nimport h3\nfrom h3ronpy import raster\nimport rasterio as rio\nimport rasterio.plot\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom rasterstats import gen_zonal_stats, gen_point_query\nfrom rasterstats import zonal_stats\nimport pandas as pd\nimport geopandas as gpd\nimport h3\nimport json\nimport os\nfrom shapely.geometry import shape, mapping, box, Point, LinearRing, Polygon\n\n\n# +\ntest_raster = '../../data/raw/crop_data/default_pasture/CroplandPastureArea2000_Geotiff/Pasture2000_5m.tif'\n\ntest_area = (-10,40,0,50)\n\n# +\nwith rio.open(test_raster) as src:\n window = rio.windows.from_bounds(*test_area, src.transform)\n transform = rio.windows.transform(window, src.transform)\n print(src.profile)\n rio.plot.show(src.read(window=window, masked=True))\n gdf = raster.raster_to_geodataframe(src.read(1, window=window), transform, h3_resolution=4, nodata_value=src.profile['nodata'], compacted=False)\n\ngdf.plot('value')\n#gdf['h3index'] = gdf['h3index'].apply(hex)\ngdf.head()\n\n# +\ngeom = box(*test_area)\nidx = [h for h in h3.polyfill(mapping(geom), 4, geo_json_conformant=True)]\npts = [Point(h3.h3_to_geo(h)[::-1]) for h in idx]\nidx_int = [int(h, 16) for h in idx]\ngeoms = h3.h3_set_to_multi_polygon(idx, geo_json=True)\n\ndf = pd.DataFrame({'h3index':idx_int, 'value':gen_point_query(pts, test_raster, interpolate='nearest')})\ndf = df.dropna()\ndf.plot('value')\ndf.head()\n# -\n\nj = gdf.set_index('h3index').join(df.set_index('h3index'), rsuffix='t')\nj\n\n# %%timeit\npd.DataFrame({'h3index':idx_int, 'value':gen_point_query(pts, test_raster, interpolate='nearest')})\n\n# %%timeit\nwith rio.open(test_raster) as src:\n raster.to_dataframe(src.read(1, window=window), transform, h3_resolution=4, nodata_value=src.profile['nodata'], compacted=False)\n\n# +\ndfs = []\nwith rio.open(test_raster) as src:\n for ji, window in src.block_windows():\n transform = rio.windows.transform(window, src.transform)\n arr = src.read(1, window=window)\n \n df = h3ronpy.raster.raster_to_geodataframe(arr, transform, 4, nodata_value=src.profile['nodata'], compacted=False)\n dfs.append(df)\n\n\n# -\n\nl = [i for df in dfs for i in df['h3index']]\nprint(len(l))\nprint(len(set(l)))\n\npd.concat(dfs).plot('value')\n\n# +\nfrom math import ceil\nBLOCKSIZE = 512\ndef gen_raster_h3(raster_list, h3_res):\n \"\"\"Convert a list of identically formatted rasters to H3\n \n A function for efficiently turning a set of rasters into an H3 table.\n \n Takes a list of 1-band rasters with identical projection/transform.\n Reads each raster in blocks, and converts to h3 (nearest to centroid).\n Yields a dataframe with an h3index and one column for each raster's value.\n\n Args:\n raster_list: list of paths to rasters\n h3_res: h3 resolution to use for resampling\n \n Yields:\n A Pandas dataframe for each raster block (usu. 512x512) with an \n h3index and one column for each raster's value.\n \"\"\"\n readers = [rio.open(r) for r in raster_list]\n names = [os.path.splitext(os.path.basename(r))[0].lower() for r in raster_list]\n \n base = readers[0]\n for j in range(ceil(base.height/BLOCKSIZE)):\n for i in range(ceil(base.width/BLOCKSIZE)):\n window = rio.windows.Window(i*BLOCKSIZE, j*BLOCKSIZE, BLOCKSIZE, BLOCKSIZE)\n w_transform = rio.windows.transform(window, base.transform)\n dfs = []\n for src in readers:\n if src.transform != base.transform:\n raise ValueError(\"Transforms do not match\")\n arr = src.read(1, window=window)\n _df = raster.raster_to_dataframe(arr, w_transform, h3_res, nodata_value=src.profile['nodata'], compacted=False)\n dfs.append(_df.set_index('h3index')['value'])\n df = pd.concat(dfs, axis=1)\n print(f'Reading block {j}, {i}: h3index count {len(df)}')\n if len(df):\n df.columns = names\n # cast h3index from int64 to hex string\n yield df\n for src in readers:\n src.close()\n\n\n\n# +\ntest_list = [\n '../../data/raw/crop_data/default_pasture/CroplandPastureArea2000_Geotiff/Pasture2000_5m.tif',\n '../../data/raw/crop_data/default_pasture/CroplandPastureArea2000_Geotiff/Cropland2000_5m.tif'\n]\ntest_dir = '../../data/seed/mapspam/spam2017v2r1_ssa_prod'\ntest_list2 = [os.path.join(test_dir, f) for f in os.listdir(test_dir)]\n\nh3grid = pd.concat(list(gen_raster_h3(test_list2, 4)))\nh3grid = gpd.GeoDataFrame(h3grid, geometry=[Polygon(h3.h3_to_geo_boundary(h, geo_json=True)) for h in h3grid.index])\nh3grid.plot()\n# -\n\n\n","repo_name":"Vizzuality/landgriffon","sub_path":"data/notebooks/Lab/0_6_test_h3ronpy_fg.ipynb","file_name":"0_6_test_h3ronpy_fg.ipynb","file_ext":"py","file_size_in_byte":4650,"program_lang":"python","lang":"en","doc_type":"code","stars":14,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"33577372063","text":"def func(Manufacturer,target_period,reference_period):\n from tkinter.filedialog import askopenfilename\n filename = askopenfilename()\n \n import sys\n import pandas as pd\n df = pd.read_excel(filename,sheet_name = 'input_data')\n del filename\n \n print('Validating availability of Manufacturer in data')\n if Manufacturer in df['Manufacturer'].unique():\n print('Manufacturer available in data')\n else:\n print('Manufacturer not available in the data')\n print('Ensure input from below mentioned list')\n print(df['Manufacturer'].unique())\n sys.exit('Please input the correct Manufacturer and Retry!')\n \n #Converting month to a format as per input\n df.month = pd.PeriodIndex(df.month, freq='M').strftime('%b%Y')\n \n print('Validating availability of target period in data')\n if target_period in df['month'].unique():\n print('Target period input available in data')\n else:\n print('Target Period not available in the data')\n print('Ensure input from below mentioned list')\n print(df['month'].unique())\n sys.exit('Please input the correct date format and Retry!')\n\n print('Validating availability of reference_period in data')\n if reference_period in df['month'].unique():\n print('Reference Period input available in data')\n else:\n print('Reference Period not available in the data')\n print('Ensure input from below mentioned list')\n print(df['month'].unique())\n sys.exit('Please input the correct date format and Retry!')\n \n \n #Filtering Data based on Date\n df = df.loc[(df.month == target_period) | (df.month == reference_period),:]\n\n #Filtering Data based on Manufacturer\n df = df.loc[df['Manufacturer'] == Manufacturer]\n\n #Pivoting Data to Convert Data from long to wide format\n df = (df.pivot_table(index=['Zone', 'Region', 'Manufacturer', 'Brand', 'Subbrand', 'Item','PackSize', 'Packtype'],\n columns='month',\n values='Value Offtake(000 Rs)').reset_index())\n\n #Data is now in a workable format let us start with Analysis\n #Step 1: Lets Check if this Manufacturer had a drop\n\n BY_MANUFACTURER = df.groupby('Manufacturer')\n Manu_Table = BY_MANUFACTURER.sum()\n\n\n if (Manu_Table[target_period] > Manu_Table[reference_period])[0]:\n sys.exit(\"There is no drop in the sales of {0} for the period {1}\".format(Manufacturer,target_period))\n\n print(\"There has been a drop in the sales of {0} for the period {1}\".format(Manufacturer,target_period))\n print(\".\")\n print(\"Initiating deep dive\")\n \n #Making Table for Analysis\n\n \n\n Desired_Levels = ['Zone', 'Region','Manufacturer', 'Brand', 'Subbrand', 'Item']\n Output_Table = pd.DataFrame()\n for levels in Desired_Levels:\n df_1 = df.groupby([levels]).sum()\n df_1 = df_1.reset_index()\n df_1 = df_1.rename(columns={levels:'Focus_Area'},)\n df_1['Growth_Rate'] = round(((df_1[target_period] - df_1[reference_period])/df_1[reference_period])*100)\n df_1['Contribution'] = round(((df_1[target_period] + df_1[reference_period])/(sum(df_1[target_period])+sum(df_1[reference_period]))*100))\n df_1['Product'] = round(((df_1['Growth_Rate']/100) * (df_1['Contribution']/100)),2)\n df_1['Manufacturer'] = Manufacturer\n del df_1[target_period]\n del df_1[reference_period]\n df_1 = df_1[['Manufacturer','Focus_Area','Growth_Rate','Contribution','Product']]\n Output_Table = Output_Table.append(df_1, ignore_index=True)\n\n Output_Table = Output_Table.sort_values(by='Product',ascending=True)\n Output_Table = Output_Table.reset_index()\n del Output_Table['index']\n\n return(Output_Table)\nfunc(Manufacturer='AMUL',target_period='May2019',reference_period='Apr2019')\n","repo_name":"amisha-74/coding","sub_path":"Final Deep Dive Analysis-checkpoint.ipynb","file_name":"Final Deep Dive Analysis-checkpoint.ipynb","file_ext":"py","file_size_in_byte":3849,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"36471005240","text":"# ### Use trained models to classify collection of tweets\n\nimport pandas as pd\nfrom itertools import combinations\nfrom torchtext import vocab\nimport os\nimport sys\nimport torch\nfrom torch.nn import functional as F\nimport numpy as np\nfrom torchtext import data\nfrom torchtext import datasets\nfrom torchtext.vocab import Vectors, GloVe\nfrom torchtext.legacy.data import Field, LabelField, TabularDataset, BucketIterator, Iterator\nimport torch.optim as optim\nfrom sklearn.metrics import accuracy_score, classification_report, confusion_matrix\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\ndf_test = pd.read_csv(\"data/Twitter/clean_twitter_info_annotated.csv\")\ndf_test = df_test[df_test['clean_tweet'].notna()]\n# assign dummy label to work more easily with prediction code\ndf_test['label'] = 0\n\n# +\ndf_train = pd.read_csv(\"data/Twitter/hate_twitter/hate_upsampled_train.csv\")\ndf_val = pd.read_csv(\"data/Twitter/hate_twitter/hate_upsampled_val.csv\")\n\ndf_train = df_train[df_train['clean_tweet'].notna()]\ndf_val = df_val[df_val['clean_tweet'].notna()]\n\ndf_train[[\"clean_tweet\", \"label\"]].to_csv(\"data/Twitter/hate_twitter/train.csv\")\ndf_val[[\"clean_tweet\", \"label\"]].to_csv(\"data/Twitter/hate_twitter/val.csv\")\ndf_test[[\"clean_tweet\", \"label\"]].to_csv(\"data/Twitter/hate_twitter/test.csv\")\n\n# +\n#device = torch.device('cuda' if torch.cuda.is_available() else 'cpu') \ndevice = 'cpu'\n\nif device == 'cpu':\n VECTORS_CACHE_DIR = '.vector_cache/'\n # Please change above to your cache\nelse:\n VECTORS_CACHE_DIR = '.vector_cache/'\n \nBATCH_SIZE = 64\n\n\n# +\nimport torch.nn as nn\nfrom torch.nn.utils.rnn import pack_padded_sequence, pad_packed_sequence\nimport torch.nn.functional as F\n\nclass classifier(nn.Module):\n \n #define all the layers used in model\n def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, text, n_layers, \n bidirectional, dropout):\n \n #Constructor\n super().__init__() \n self.embedding_dim = embedding_dim\n #embedding layer\n self.embedding = nn.Embedding(vocab_size, embedding_dim)\n self.embedding.weight.data = text.vectors\n self.dropout = nn.Dropout(dropout)\n\n #lstm layer\n self.lstm = nn.LSTM(embedding_dim, \n hidden_dim, \n num_layers=n_layers, \n bidirectional=True, \n # dropout=dropout,\n batch_first=True)\n \n #dense layer\n self.fc = nn.Linear(hidden_dim * 2, output_dim)\n \n # #activation function\n self.act = nn.Sigmoid()\n \n def forward(self, text, text_lengths):\n \n #text = [batch size,sent_length]\n embedded = self.embedding(text)\n #embedded = [batch size, sent_len, emb dim]\n\n #packed sequence\n packed_embedded = pack_padded_sequence(embedded, text_lengths,batch_first=True, \n enforce_sorted=False)\n \n packed_output, (hidden, cell) = self.lstm(packed_embedded)\n output, _ = pad_packed_sequence(packed_output, batch_first=True)\n\n out_forward = output[range(len(output)), text_lengths - 1, :self.embedding_dim]\n out_reverse = output[:, 0, self.embedding_dim:]\n out_reduced = torch.cat((out_forward, out_reverse), 1)\n dense_outputs = self.dropout(out_reduced)\n dense_outputs=self.fc(dense_outputs)\n\n dense_outputs = torch.squeeze(dense_outputs, 1)\n outputs = torch.sigmoid(dense_outputs)\n return outputs.to(device)\n\n\n# -\n\nTEXT = Field(tokenize='spacy',batch_first=True,include_lengths=True)\nLABEL = LabelField(sequential=False, use_vocab=False, dtype = torch.float,batch_first=True)\nfields = [(None, None), ('clean_tweet',TEXT),('label', LABEL)]\ntrain, val, test = TabularDataset.splits(path = \"data/Twitter/hate_twitter/\", \n train='train.csv',validation='val.csv',\n test='test.csv', format='csv',\n fields= fields, skip_header=True)\nTEXT.build_vocab(train,min_freq=3,vectors = \"glove.twitter.27B.100d\")\ntrain_iterator, valid_iterator, test_iterator = BucketIterator.splits(\n (train, val, test), \n batch_size = BATCH_SIZE,\n sort_key = lambda x: len(x.clean_tweet),\n sort_within_batch=True,\n device = device)\nsize_of_vocab = len(TEXT.vocab)\nembedding_dim = 100\nnum_hidden_nodes = 50\nnum_output_nodes = 1\nnum_layers = 2\ndropout = 0.2\n\n\ndef load_checkpoint(load_path, model, optimizer, device):\n '''\n Used in evaluate() to load the current best model.\n \n Inputs:\n - load_path: string, where to save model specs.\n - model: a PyTorch model object.\n - optimizer: a PyTorch Optimizer to be used.\n - device: string, 'cpu' or 'cuda' if using google colab.\n \n Returns: float, the models last validation loss.\n \n Other Effects:\n - Loads the saved state at load_path\n into the current model object\n '''\n if load_path==None:\n return\n\n state_dict = torch.load(load_path, map_location=device)\n print(f'Model loaded from <== {load_path}')\n \n model.load_state_dict(state_dict['model_state_dict'])\n optimizer.load_state_dict(state_dict['optimizer_state_dict'])\n \n return model\n\n\n\ndef classify_unlabeled(model, test_loader, device):\n '''\n Accepts the current best model and evaluates\n the test dataset. Printing test accuracy and\n an sklearn confusion matrix report.\n \n Inputs:\n - model: PyTorch model object, the current best model.\n - test_loader: an iterator with test data.\n - device: string, 'cpu' or 'cuda' if using google colab.\n \n Returns: None.\n \n Other Effects:\n Prints test accuracy.\n Prints an Accuracy / F1 Report (sklearn)\n Prints a Confusion Matrix (sklearn & matplotlib)\n '''\n y_pred = []\n\n model.eval()\n with torch.no_grad():\n for ((text, text_len),_) in test_loader: \n \n text = text.to(device)\n text_len = text_len.to(device)\n output = model(text, text_len) \n output = torch.round(output)\n\n y_pred.extend(output.tolist())\n\n return y_pred\n\n\n# +\n# Evaluate\nbest_model = classifier(size_of_vocab, embedding_dim, num_hidden_nodes,num_output_nodes, TEXT.vocab, num_layers, \n bidirectional = True, dropout = dropout)\n\noptimizer = optim.Adam(best_model.parameters(), lr=0.00001)\n\nbest_model = load_checkpoint('model.pt', best_model, optimizer, device)\n\ny_pred = classify_unlabeled(best_model, test_iterator, device)\n# -\n\n# confirm y pred and df test have same size\nlen(y_pred)\n\n# confirm y pred and df test have same size\ndf_test.shape\n\ndf_test['label_pred'] = y_pred\ndf_test['date_dt'] = pd.to_datetime(df_test['Datetime'])\ndf_test.drop(columns=['Unnamed: 0', 'Unnamed: 0.1', 'label'], inplace=True)\n\ndf_test['date_day'] = df_test['date_dt'].dt.date\n\n# show number of hate tweets in labeled data\ndf_test.groupby(['tag'])['tweet_id'].count()\n\ndf_test.groupby(['tag', 'label_pred'])['tweet_id'].count()\n\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].head(70)\n\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].iloc[10].Text\n\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].iloc[15].Text\n\n# examine single tweet example of non-hate, sexist tweets\ndf_test[(df_test['label_pred'] == 0) & (df_test['tag'] == 'sexism')].iloc[68].Text\n\ndf_test[(df_test['label_pred'] > 0) & (df_test['tag'] == 'none')& (df_test['date_day'] > pd.to_datetime('2015-01-15'))].head(70)\n\n# examine individual tweet from the above set of hateful, 'none' overlap\ndf_test[(df_test['label_pred'] > 0) & (df_test['tag'] == 'none')& (df_test['date_day'] > pd.to_datetime('2015-01-15'))].iloc[66].Text\n\n# ### Visualize predicted hate comments over time\n\ntweet_preds_day = df_test.groupby(['label_pred', 'date_day'])['tweet_id'].count().reset_index()\ntweet_preds_day.rename(columns={'tweet_id':'num_tweets'},inplace=True)\n\ntweet_preds_day.head()\n\n# +\n# View predicted hate comments over time\nf, ax = plt.subplots(1,1,figsize=(15,10))\n\nsns.lineplot('date_day', 'num_tweets',hue='label_pred', \\\n data=tweet_preds_day[tweet_preds_day['label_pred']>0])\n\nax.set_xlabel(\"Date\")\nax.set_ylabel(\"Number of Hateful Tweets\")\nax.set_title(\"Tweets Classified as Hateful by LSTM Model\")\n# -\n\n# ### Compare with Google Trends\n\n# +\nimport pytrends\nimport pandas as pd\nfrom pytrends.request import TrendReq\nimport matplotlib\nimport csv\nfrom datetime import date\nimport time \nimport requests\nimport warnings\nwarnings.filterwarnings('ignore')\n\npytrends = TrendReq(hl='en-US', tz=360)\n\n# +\nkeyword = [\"nigger\", \"niggers\", \"kike\", \"chink\", \"faggot\"]\n\npytrends.build_payload(keyword, cat=0, timeframe='2013-09-01 2015-04-01', geo='US', gprop='')\ninterest_over_time_df = pytrends.interest_over_time()\n\n# +\nimport matplotlib.pyplot as plt\nimport numpy as np\n\nf, (ax1, ax2) = plt.subplots(2, 1, figsize=(15,10))\n\n\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"nigger\"], label = \"Anti-Black word\")\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"kike\"], label = \"Anti-Semitic word\")\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"chink\"], label = \"Anti-Asian word\")\nax1.plot(interest_over_time_df.index, interest_over_time_df[\"faggot\"], label = \"Anti-LGBT word\")\nax1.set_title(\"Google Trends Hate Speech\")\nax2.plot('date_day', 'num_tweets', data=tweet_preds_day[tweet_preds_day['label_pred']>0])\nax2.set_title(\"Classified Hate Speech\")\nplt.plot()\nplt.legend()\nplt.show()\n\n# -\n\n\n","repo_name":"keiraou/life_cycle_of_hate_speech","sub_path":"classify_with_trained_models.ipynb","file_name":"classify_with_trained_models.ipynb","file_ext":"py","file_size_in_byte":9716,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"42823273365","text":"import pandas as pd\nimport numpy as np\n\nimport sklearn as sk\nfrom sklearn import tree\n\nratings = pd.read_csv(\"ratings.csv\")\nmean_ratings = ratings.groupby(\"movieId\").rating.mean()\n\nmovies = pd.read_csv(\"movies.csv\")\nmovies = movies.set_index(\"movieId\")\nmovies['rating'] = mean_ratings\nmovies.head()\n\nmovies['year'] = movies.title.str.extract(\"\\((\\d{4})\\)\", expand=False)\nmovies.year.astype(float)\nmovies.head()\n\ngenres = [\"Drama\", \"Comedy\", \"Thrilles\", \"Action\" ,\"Adventure\", \"Crime\"]\nfor genre in genres:\n movies[genre] = movies.genres.str.contains(genre)\nmovies['good'] = np.where(movies.rating>3.5, \"Good\", \"Bad\")\nmovies.head(5)\n\nfrom sklearn import tree\nclf = tree.DecisionTreeClassifier(max_depth=2)\nm = clf.fit(movies[genres], movies.good)\n\npredictions = m.predict(movies[genres])\npd.crosstab(movies.good, predictions)\n\nfrom sklearn.metrics import confusion_matrix\nprint(confusion_matrix(movies.good, predictions))\n\ncorrect = movies.good == predictions\ncorrect.mean()\n\nimport pydotplus\nfrom IPython.display import Image\ndot_data = tree.export_graphviz(m,\n feature_names=genres, class_names=['bad', 'good'], \n filled=True, rounded=True) \ngraph = pydotplus.graph_from_dot_data(dot_data) \n\n\n","repo_name":"peiweihe/analyse_amazon_reviews","sub_path":"trees.ipynb","file_name":"trees.ipynb","file_ext":"py","file_size_in_byte":1201,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"28043068283","text":"# +\nrecords = [\n ('foo',1,2),\n ('bar','hello'),\n ('foo',3,4),\n]\ndef do_foo(x,y):\n print('foo',x,y)\n\ndef do_bar(s):\n print('bar',s)\n\nfor tag, *args in records:\n if tag == 'foo':\n do_foo(*args)\n elif tag == 'bar':\n do_bar(*args)\n# -\n\nimport heapq\nportfolio = [\n {'name': 'ibm', 'shares':100, 'price': 91.1},\n {'name': 'appl', 'shares':50, 'price': 543.22},\n {'name': 'fb', 'shares':200, 'price': 21.09},\n {'name': 'yhoo', 'shares':45, 'price': 16.35},\n]\ncheap = heapq.nsmallest(3, portfolio, key=lambda s : s['price'])\nexpensive = heapq.nlargest(3, portfolio, key=lambda s : s['price'])\n\ncheap\n\nexpensive\n\nnums = [1,3,2,5,4.5, 77,23,9,6, 54,96,0,12,11,2.3]\n\nlen(nums)\n\nimport time\n\nt1 = time.time()\nnew_lis = sorted(nums)[:5]\nt2 = time.time() - t1\n\nnew_lis\n\nt2\n\n\nt3 = time.time()\nnew_hlis= heapq.nsmallest(5,nums)\nt4 = time.time()- t3\n\nnew_hlis\n\nt4\n\n\n# # 下划线起名的规则\n# + _xx:前置单下划线,私有化属性或方法,一般来讲,变量名_xx被看作是“私有 的”,在模块或类外不可以使用。当变量是私有的时候,用_xx 来表示变量是很好的习惯。类对象和子类可以访问,这并不能完全做到真正的私有,只是约定俗成的而已,这样写表示不希望这个变量在外部被直接调用\n# + \\__xx:前置双下划线,私有化属性或方法,无法在外部直接访问(名字重整所以访问不到,只能是允许这个类本身进行访问了。连子类也不可以)\n# + \\__xx\\__:前后双下划线,系统定义名字(这就是在python中强大的魔法方法),因为变量名__xxx__对Python 来说有特殊含义,对于普通的变量应当避免这种命名风格。\n# + xx_:后置单下划线,用于避免与Python关键词的冲突\n# 如以下例子所示,我在test类中定义了x,_x和 __x三个属性,并创建了test的类对象t,对这三个属性进行访问,__x不能被访问到\n\nclass test(object):\n def __init__(self):\n self.x = 10\n self._x = 20\n self.__x = 30\nt = test()\nprint(t.x) # 10\nprint(t._x) # 20\nprint(t.__x)\n# print(t.__x) # AttributeError: 'test' object has no attribute '__x'\n\n# 构建一个优先级队列\nclass PriorityQueue:\n def __init__(self):\n self._queue= []\n self._index = 0\n \n def push(self, item, priority):\n heapq.heappush(self._queue, (-priority, self._index,item))\n self._index += 1\n \n def pop(self):\n return heapq.heappop(self._queue)[-1]\n\n\nclass Item:\n def __init__(self, name):\n self.name = name\n \n def __repr__(self):\n return f\"Item('{self.name}')\"\n\n\nq = PriorityQueue()\nq.push(Item('foo'), 1)\nq.push(Item('bar'), 2)\nq.push(Item('spam'), 3)\nq.push(Item('grok'), 5)\n\nq.pop()\n\nq.pop()\n\nq.push(Item('test'), 1)\n\nq.pop()\n\nq.pop()\n\nq.pop()\n\nq.pop()\n\n# # 字典中的键映射多个值(multidict)\n\n# +\nfrom collections import defaultdict\n\nd = defaultdict(list)\nd['a'].append(1)\nd['a'].append(2)\nd['b'].append(4)\n\n# -\n\nd\n\nd_prime = defaultdict(set)\nd_prime['a'].add(1)\nd_prime['a'].add(2)\nd_prime['b'].add(4)\n\nd_prime\n\n\n# # 字典排序\n\n# +\nfrom collections import OrderedDict\ndef ordered_dict():\n d = OrderedDict()\n d['foo']=1\n d['bar']=2\n d['spam']=3\n d['grok']=4\n \n for key in d:\n print(key, d[key])\n\nc= ordered_dict()\n \n\n# -\n\n# # 字典的运算\n\nprices = {\n 'acme': 45.23,\n 'aapl': 612.78,\n 'ibm':205.55,\n 'hpq': 37.20,\n 'fb': 10.75 \n}\n\nmin_price = min(zip(prices.values(), prices.keys()))\n\nmin_price\n\nmax_price = max(zip(prices.values(), prices.keys()))\n\nmax_price\n\nsorted_prices = sorted(zip(prices.values(), prices.keys()))\n\nsorted_prices\n\nprices_and_names = zip(prices.values(), prices.keys())\nprint(min(prices_and_names)) # ok\nprint(max(prices_and_names)) # ValueError: max() arg is an empty set\n\nprint(max(prices))\nprint(min(prices))\n\nprint(min(prices.values()))\nprint(max(prices.values()))\n\nprint(min(prices, key= lambda k: prices[k]))\nprint(max(prices, key= lambda k: prices[k]))\n\nmin_value = prices[min(prices, key= lambda k: prices[k])]\n\nmin_value\n\n# ## `总之,用zip函数达到字典键值反转的效果`\n\n# # 查找两字典的相同点\n\na = {\n 'x': 1,\n 'y': 2,\n 'z': 3,\n}\nb= {\n 'p':233,\n 'y': 11,\n 'x': 1\n}\n\n# find keys in common\na.keys() & b.keys()\n\n# find keys in a that not in b\na.keys() - b.keys()\n\n# find (key, value) pairs in common\na.items() & b.items()\n\n# make a new dict with certain keys removed\nc = {key: a[key] for key in a.keys() - {'z', 'w'}}\n\nc\n\n\n# # 删除序列相同元素并保持顺序\n\n# When hashable\ndef dedupe(items):\n seen = set()\n for item in items:\n if item not in seen:\n yield item\n seen.add(item)\n\n\na = [1,5,2,1,9,1,5,10]\nlist(dedupe(a))\n\n# ## About 'Hashale'\n# > An object is hashable if it has a hash value which never changes during its lifetime (it needs a __hash__() method), and can be compared to other objects (it needs an __eq__() or __cmp__() method). Hashable objects which compare equal must have the same hash value.\n#\n# > Hashability makes an object usable as a dictionary key and a set member, because these data structures use the hash value internally.\n#\n# > All of Python’s immutable built-in objects are hashable, while no mutable containers (such as lists or dictionaries) are. Objects which are instances of user-defined classes are hashable by default; they all compare unequal, and their hash value is their id().\n\ntype(a)\n\n\n# when it's not hashable\ndef dedupe(items, key= None):\n seen = set()\n for item in items:\n val = item if key is None else key(item)\n if val not in seen:\n yield item \n seen.add(val)\n\n\na = [{'x': 1, 'y':2}, {'x':1,'y':3}, {'x':2,'y':4}, {'x': 1, 'y':2}]\n\nlist(dedupe(a, key=lambda d: (d['x'], d['y'])))\n\nlist(dedupe(a, key=lambda d: (d['x'])))\n\n# 当然了,最简单的还是set(),但是不能保证顺序\n\na = [1,5,2,1,9,1,5,10]\n\nset(a)\n\nrecord = '....................100 .......513.25 ..........'\ncost = int(record[20:23]) * float(record[31:37])\n\nprint(cost)\n\nSHARES = slice(20, 23)\nPRICE = slice(31, 37)\ncost = int(record[SHARES]) * float(record[PRICE])\n\ncost\n\nitems = [0, 1, 2, 3, 4, 5, 6]\na= slice(2,4)\nitems[a]\n\nitems[a]=[10,11]\n\nitems\n\ndel items[a]\n\nitems\n\ns= slice(5,50,2)\n\n\ns.start\n\ns.stop\n\ns.step\n\ns = 'HelloWorld'\n\na.indices(len(s))\n\nfor i in range(*a.indices(len(s))):\n print(s[i])\n \n\n# # 找出序列中出现次数最多的元素\n\nwords = [\n'look', 'into', 'my', 'eyes', 'look', 'into', 'my', 'eyes',\n'the', 'eyes', 'the', 'eyes', 'the', 'eyes', 'not', 'around', 'the',\n'eyes', \"don't\", 'look', 'around', 'the', 'eyes', 'look', 'into',\n'my', 'eyes', \"you're\", 'under'\n]\n\nfrom collections import Counter\nword_counts = Counter(words)\ntop_three = word_counts.most_common(3)\n\ntop_three\n\n# Counter 对象可以接受任意的hashable 序列对象\n\nword_counts['eyes']\n\nword_counts['not']\n\n# 如果你想手动增加计数,可以简单的用加法:\n\nmorewords = ['why','are','you','not','looking','in','my','eyes']\n\nfor item in morewords:\n print(item)\n\ntype(word_counts)\n\nword_counts\n\nfor word in morewords:\n word_counts['word']+=1\n \n\nword_counts['eyes']\n\nword_counts\n\n# 上面写的手动修改方法已经失效\n\nword_counts.update(morewords)\n\nword_counts\n\nword_counts['not']\n\nword_counts['eyes']\n\n# Counter 实例一个鲜为人知的特性是它们可以很容易的跟数学运算操作相结合\n\n# +\na = Counter(words)\nb = Counter(morewords)\n\n\n# -\n\na\n\nb\n\na + b\n\na - b\n\na & b\n\n# # 通过某个关键字排序一个字典列表\n\nrows = [\n{'fname': 'Brian', 'lname': 'Jones', 'uid': 1003},\n{'fname': 'David', 'lname': 'Beazley', 'uid': 1002},\n{'fname': 'John', 'lname': 'Cleese', 'uid': 1001},\n{'fname': 'Big', 'lname': 'Jones', 'uid': 1004}\n]\n\nfrom operator import itemgetter\n\nrows_by_fname = sorted(rows, key=itemgetter('fname'))\nrows_by_uid = sorted(rows, key=itemgetter('uid'))\n\nrows_by_fname\n\nrows_by_uid\n\n# 支持多个keys\nrows_by_lfname = sorted(rows, key=itemgetter('lname','fname'))\n\nrows_by_lfname\n\n# itemgetter() 有时候也可以用lambda 表达式代替\nrows_by_fname = sorted(rows, key= lambda r: r['fname'])\n\n# 当然还是itemgetter效率更快\nrows_by_fname\n\nmin(rows, key=itemgetter('uid'))\n\n\n# # 排序不支持原生比较的对象\n\nclass User:\n def __init__(self, user_id):\n self.user_id = user_id\n \n def __repr__(self):\n return f'User({self.user_id})'\n\n\ndef sort_notcompare():\n users = [User(233), User(64), User(89)]\n print(users)\n print(sorted(users, key= lambda u: u.user_id))\n\n\nsort_notcompare()\n\nusers = [User(233), User(64), User(89)]\n\n# 使用operator.attrgetter\nfrom operator import attrgetter\nsorted(users, key=attrgetter('user_id'))\n\nmin(users, key=attrgetter('user_id'))\n\n# ### 不管是itemgetter还是attrgetter, 效果基本都和lambda一样,前者效率更高, 且能多个字段比较\n\n# # 通过某个字段将记录分组\n\n# 你有一个字典或者实例的序列,然后你想根据某个特定的字段比如date 来分组迭代访问。\n\nrows = [\n{'address': '5412 N CLARK', 'date': '07/01/2012'},\n{'address': '5148 N CLARK', 'date': '07/04/2012'},\n{'address': '5800 E 58TH', 'date': '07/02/2012'},\n{'address': '2122 N CLARK', 'date': '07/03/2012'},\n{'address': '5645 N RAVENSWOOD', 'date': '07/02/2012'},\n{'address': '1060 W ADDISON', 'date': '07/02/2012'},\n{'address': '4801 N BROADWAY', 'date': '07/01/2012'},\n{'address': '1039 W GRANVILLE', 'date': '07/04/2012'},\n]\n\nfrom operator import itemgetter\nfrom itertools import groupby\n\nrows.sort(key=itemgetter('date'))\n\nrows\n\nfor date, items in groupby(rows, key=itemgetter('date')):\n print(date)\n for i in items:\n print('+', i)\n\n# ### groupby需要事先排序,假如说只是想将一些字段分组到更大的数据结构中,并且允许随机访问,还是使用defaultdict为佳\n\nrows_by_date = defaultdict(list)\nfor row in rows:\n rows_by_date[row['date']].append(row)\n\nfor r in rows_by_date['07/01/2012']:\n print(r)\n\nrows_by_date\n\n# # 过滤序列元素\n\n# ### 最简单的过滤序列元素的方法就是使用列表推导\n\nmylist = [1, 4, -5, 10, -7, 2, 3, -1]\n\n[n for n in mylist if n > 0]\n\n# ### 使用列表推导的一个潜在缺陷就是如果输入非常大的时候会产生一个非常大的结果集,占用大量内存\n\npos = (n for n in mylist if n > 0)\n\npos\n\nfor i in pos:\n print(i)\n\n# ### 当过滤规比较复杂时,需要写一个函数,使用内建filter(返回的是迭代器)\n\nvalues = ['1', '2', '-3', '-', '4', 'N/A', '5']\n\n\ndef is_int(val):\n try: \n x = int(val)\n return True\n except ValueError:\n return False\nivals = list(filter(is_int, values))\n\nivals\n\n# +\n# filter()\n#Return an iterator yielding those items of iterable for which function(item)\n# -\n\n# ### 列表推导和生成器表达式通常情况下是过滤数据最简单的方式, 其实它们还能在过滤的时候转换数据\n\n# +\nmylist = [1, 4, -5, 10, -7, 2, 3, -1]\nimport math\n\n[math.sqrt(n) for n in mylist if n > 0]\n# -\n\n# ### 过滤操作的一个变种就是将不符合条件的值用新的值代替,而不是丢弃它们\n\nclip_neg = [n if n > 0 else 0 for n in mylist]\n\nclip_neg\n\naddresses = [\n'5412 N CLARK',\n'5148 N CLARK',\n'5800 E 58TH',\n'2122 N CLARK',\n'5645 N RAVENSWOOD',\n'1060 W ADDISON',\n'4801 N BROADWAY',\n'1039 W GRANVILLE',\n]\ncounts = [ 0, 3, 10, 4, 1, 7, 6, 1]\n\nlen(addresses)\n\nlen(counts)\n\n# ### itertools.compress() ,它以一个iterable对象和一个相对应的Boolean 选择器序列作为输入参数。然后输出iterable 对象中对应选择器为True 的元素。当你需要用另外一个相关联的序列来过滤某个序列的时候,这个函数是非常有用的。\n\n# +\nfrom itertools import compress\n\nmore5 = [n>5 for n in counts]\n# -\n\nmore5\n\nlist(compress(addresses, more5))\n\n# # 从字典中提取子集\n\n# +\nprices = {\n'ACME': 45.23,\n'AAPL': 612.78,\n'IBM': 205.55,\n'HPQ': 37.20,\n'FB': 10.75\n}\n\n# 字典推导\n#Make a dictionary of all prices over 200\np1 = {key: value for key, value in prices.items() if value > 200}\n# Make a dictionary of tech stocks\ntech_names = {'AAPL', 'IBM', 'HPQ', 'MSFT'}\np2 = {key: value for key, value in prices.items() if key in tech_names}\n# -\n\np1\n\np2\n\n# 比较慢\np1 = dict((key, value) for key, value in prices.items() if value > 200)\n\np1\n\ntech_names = { 'AAPL', 'IBM', 'HPQ', 'MSFT' }\np2 = { key:prices[key] for key in prices.keys() & tech_names }\n\n# 同理,也比较慢\np2\n\n# # 映射名称到序列元素\n\nfrom collections import namedtuple\nSubscriber = namedtuple('Subscriber', ['addr', 'joined'])\n\nsub = Subscriber('joney@example.com', '2021-3-15')\n\nsub.addr\n\nsub.joined\n\nlen(sub)\n\n# ### 尽管namedtuple 的实例看起来像一个普通的类实例,但是它跟元组类型是可交换的,支持所有的普通元组操作,比如索引和解压。\n\naddr, joined = sub\n\naddr\n\njoined\n\nsub\n\n\ndef compute_cost(records):\n total = 0.0\n for rec in records:\n total += rec[1] * rec[2]\n return total\n\n\nStock = namedtuple('Stock', ['name', 'shares', 'price'])\ndef compute_cost(records):\n total = 0.0\n \n total += s.shares * s.price\n return total\n\n\ns = Stock('ACME', 100, 123.45)\n\ns\n\ns.shares\n\ns.price\n\ncompute_cost(s)\n\ns.shares = 75\n\ns = s._replace(shares = 75)\n\ns\n\nStock = namedtuple('Stock', ['name', 'shares', 'price', 'date', 'time'])\n# Create a prototype instance\nstock_prototype = Stock('', 0, 0.0, None, None)\n# Function to convert a dictionary to a Stock\ndef dict_to_stock(s):\n return stock_prototype._replace(**s)\n\n\na = {'name': 'ACME', 'shares': 100, 'price': 123.45}\n\ndict_to_stock(a)\n\nstock_prototype\n\n# ### 如果你的目标是定义一个需要更新很多实例属性的高效数据结构,那么命名元组并不是你的最佳选择。这时候你应该考虑定义一个包含slots 方法的类\n\n# # 转换并同时计算数据\n\nnums = [1, 2, 3, 4, 5]\ns = sum(x * x for x in nums)\n\ns\n\nimport os\n# Determine if any .py files exist in a directory\nfiles = os.listdir('/Users/joriri/Code')\nif any(name.endswith('.py') for name in files):\n print('There be python!')\nelse:\n print('Sorry, no python.')\n\n# Output a tuple as CSV\ns = ('ACME', 50, 123.45)\nprint(','.join(str(x) for x in s))\n# Data reduction across fields of a data structure\nportfolio = [\n{'name':'GOOG', 'shares': 50},\n{'name':'YHOO', 'shares': 75},\n{'name':'AOL', 'shares': 20},\n{'name':'SCOX', 'shares': 65}\n]\nmin_shares = min(s['shares'] for s in portfolio)\n\nmin_shares\n\n# ### 在使用一些聚集函数比如min() 和max() 的时候你可能更加倾向于使用生成器版本,它们接受的一个key 关键字参数或许对你很有帮助。\n\n# Original: Returns 20\nmin_shares = min(s['shares'] for s in portfolio)\n# Alternative: Returns {'name': 'AOL', 'shares': 20}\nmin_shares = min(portfolio, key=lambda s: s['shares'])\n\nmin_shares\n\n# # 合并多个字典或映射\n\na = {'x': 1, 'z': 3 }\nb = {'y': 2, 'z': 4 , 'p': 5}\n\n# +\nfrom collections import ChainMap\nc = ChainMap(a,b)\n\nprint(c['x'])\nprint(c['z'])\nprint(c['p'])\n# -\n\ndel c['y']\n\ndel c['x']\n\nc['x']\n\na\n\nc['w']= 6\n\na\n\n# ### 对chainmap的字典(列表)来说, 更新删除影响的是列表中的第一个字典\n\nvalues = ChainMap()\nvalues['x'] =1 \n\nvalues=values.new_child()\nvalues['x'] = 2\nvalues=values.new_child()\nvalues['x']=3\n\n\nvalues\n\nvalues['x']\n\n# discard last mapping\nvalues = values.parents\nvalues['x']\n\nvalues = values.parents\nvalues['x']\n\nvalues\n\n# ### 当然也可以用updata来代替chainmap\n\na = {'x': 1, 'z': 3 }\nb = {'y': 2, 'z': 4 }\n\nmerged = dict(b)\nmerged.update(a)\n\nmerged\n\na['x']= 233\n\nmerged\n\n\n# ### 这样也能行得通,但是它需要你创建一个完全不同的字典对象(或者是破坏现有字典结构)。同时,如果原字典做了更新,这种改变不会反应到新的合并字典中去\n\n# ### ChianMap 使用原来的字典,它自己不创建新的字典。所以它并不会产生上面所说的结果\n\na = {'x': 1, 'z': 3 }\nb = {'y': 2, 'z': 4 }\nmerged = ChainMap(a,b)\n\nmerged\n\na['x']= 233\n\nmerged\n\n# # 第二章 字符串和文本\n# # 使用多个界定符分割字符串\n\n# ### string 对象的split() 方法只适应于非常简单的字符串分割情形,它并不允许有多个分隔符或者是分隔符周围不确定的空格。当你需要更加灵活的切割字符串的时候,最好使用re.split() 方法:\n\n# +\nline = 'asdf fjdk; afed, fjek,asdf, foo'\n\nimport re\nre.split(r'[;,\\s]\\s*', line)\n# -\n\n# ### 当你使用re.split() 函数时候,需要特别注意的是正则表达式中是否包含一个括号捕获分组。如果使用了捕获分组,那么被匹配的文本也将出现在结果列表中\n\n# 分隔符被捕获了\nfields = re.split(r'(,|;|\\s)\\s*', line)\nfields\n\nvalues = fields[::2]\n\nvalues\n\ndelimeters = fields[1::2]\n\ndelimeters\n\nlen(values)\n\nlen(delimeters)\n\nlen(fields)\n\ndelimeters += ['']\n\ndelimeters\n\nlen(delimeters)\n\n# reform the line using the same delimiters \n''.join(v+d for v, d in zip(values, delimeters))\n\n# ### 如果你不想保留分割字符串到结果列表中去,但仍然需要使用到括号来分组正则表达式的话,确保你的分组是非捕获分组,形如(?:...)\n\nre.split(r'(?:,|;|\\s)\\s*', line)\n\n# # 字符串开头或结尾匹配\n\n# ### 直接使用str.startswith & str.endswith\n\nfilename = 'spam.txt'\n\nfilename.startswith('spam')\n\nfilename.endswith('ttt')\n\nurl = 'http://www.baidu.com'\n\nurl.startswith('https')\n\nurl.endswith('com')\n\n# ### 匹配多种可能,先成一个`元组`\n\nimport os\nfilenames = os.listdir('.')\n\nfilenames\n\n[name for name in filenames if name.endswith(('.ipynb', 'csv'))]\n\nany(name.endswith('.py') for name in filenames)\n\n# +\nfrom urllib.request import urlopen\n\ndef read_data(name):\n if name.startswith('http', 'https', 'ftp'):\n return urlopen(name).read()\n else:\n with open(name) as f:\n return f.read()\n\n\n# -\n\n# 必须是元组\nchoices = ['http:', 'ftp:']\nurl = 'http://www.python.org'\nurl.startswith(choices)\n\nurl.startswith(tuple(choices))\n\n# ### 这种匹配也可以用正则或者切片,但是startswith是最简单且快速的。 并且,这种方法和其他普通数据聚合方法相结合(any…)也十分不错\n\n# # 用Shell 通配符匹配字符串\n\n# ## 你想使用Unix Shell 中常用的通配符(比如*.py , Dat[0-9]*.csv 等) 去匹配文本字符串\n\nfrom fnmatch import fnmatch, fnmatchcase\n\nfnmatch('foo.txt', '*.txt')\n\nfnmatch('foo.txt', '?oo.txt')\n\nfnmatch('Date45.cvs', 'Date[0-9]*')\n\nnames = ['Dat1.csv', 'Dat2.csv', 'config.ini', 'foo.py']\n\n[name for name in names if fnmatch(name, 'Dat*.csv')]\n\n# 大小写敏感 \n# False on Mac, True on Windows\nfnmatch('foo.txt', '*.TXT')\n\n# 如果要把大小写考虑进去,用fnmatchcase\nfnmatchcase('foo.txt', '*.TXT')\n\n# ### 这两个函数通常会被忽略的一个特性是在处理非文件名的字符串时候它们也是很有用的\n\naddresses = [\n'5412 N CLARK ST',\n'1060 W ADDISON ST',\n'1039 W GRANVILLE AVE',\n'2122 N CLARK ST',\n'4802 N BROADWAY',\n]\n\n[addr for addr in addresses if fnmatchcase(addr, '* ST')]\n\n[addr for addr in addresses if fnmatchcase(addr, '54[0-9][0-9] *CLARK*')]\n\n# ### fnmatch() 函数匹配能力介于简单的字符串方法和强大的正则表达式之间。如果在数据处理操作中只需要简单的通配符就能完成的时候,这通常是一个比较合理的方案。如果你的代码需要做文件名的匹配,最好使用glob 模块\n\n# # 字符串匹配和搜索\n\n# ### startswith endswith find\n\ntext = 'yeah, but no, but yeah, but no, but yeah'\n\ntext.find('no')\n\n# 使用正则\ntext1 = '11/27/2012'\ntext2 = 'Nov 27, 2012'\n\nif re.match(r'\\d+/\\d+/\\d+', text1):\n print('yes')\nelse:\n print('no')\n\n# ### 如果你想使用同一个模式去做多次匹配,你应该先将模式字符串预编译为模式对象\n\ndatepat = re.compile(r'\\d+/\\d+/\\d+')\n\nif datepat.match(text1):\n print('yes')\nelse:\n print('no')\n\n# ### match() 总是从字符串开始去匹配,如果你想查找字符串任意部分的模式出现位置,使用findall() 方法去代替。\n\ntext = 'Today is 11/27/2012. PyCon starts 3/13/2013.'\ndatepat.findall(text)\n\n# 在定义正则式的时候,通常会利用括号去捕获分组\ndatepat = re.compile(r'(\\d+)/(\\d+)/(\\d+)')\n\nm = datepat.match('11/27/2021')\n\n\nm\n\n# Return subgroup(s) of the match by indices or names.\n# For 0 returns the entire match.\nm.group(0)\n\nm.group(1)\n\nm.group(2)\n\nm.groups()\n\nm.group()\n\ntext\n\ndatepat.findall(text)\n\nfor month, day, year in datepat.findall(text):\n print(f'{year}-{month}-{day}')\n\n# # 字符串搜索和替换\n\ntext = 'yeah, but no, but yeah, but no, but yeah'\ntext.replace('yeah', 'yep')\n\ntext = 'Today is 11/27/2012. PyCon starts 3/13/2013.'\n\n# 注意这里匹配的模式有括号,意味着有group, 后面的\\3 \\1 \\2对应的是组号\nre.sub(r'(\\d+)/(\\d+)/(\\d+)', r'\\3-\\1-\\2', text)\n\n# ### 对于更加复杂的替换,可以传递一个替换回调函数来代替\n\nfrom calendar import month_abbr\ndef change_date(m):\n month_name = month_abbr[int(m.group(1))]\n return f'{m.group(2)} {month_name} {m.group(1)}'\n\n\ndatepat.sub(change_date, text)\n\ndatepat\n\nnewtext, n = datepat.subn(r'\\3-\\1-\\2', text)\n\nnewtext\n\nn\n\n# # 字符串忽略大小写的搜索替换\n\ntext = 'UPPER PYTHON, lower python, Mixed Python'\n\nre.findall('python', text, flags=re.IGNORECASE)\n\nre.sub('python', 'snake', text, flags=re.IGNORECASE)\n\n\n# 如果要使替换的单词和原有的单词 大小写一致\ndef matchcase(word):\n def replace(m):\n text = m.group()\n if text.isupper():\n return word.upper()\n elif text.islower():\n return word.lower()\n elif text[0].isupper():\n return word.capitalize()\n else:\n return word\n return replace\n\n\nre.sub('python', matchcase('snake'), text, flags=re.IGNORECASE)\n\n# # 最短匹配模式\n\nstr_pat = re.compile(r'\\\"(.*)\\\"')\n\ntext1= 'Computer says \"no.\"'\n\nstr_pat.findall(text1)\n\ntext2 = 'Computer says \"no.\" Phone says \"yes.\"'\nstr_pat.findall(text2)\n\nstr_pat = re.compile(r'\\\"(.*?)\\\"')\nstr_pat.findall(text2)\n\n# # 多行匹配模式\n\ncomment = re.compile(r'/\\*(.*?)\\*/')\ntext1= '/*this is a comment.*/'\ntext2 = '''/*this is a \nmultiline comment.*/'''\n\ntext1\n\ntext2\n\ncomment.findall(text1)\n\ncomment.findall(text2)\n\ncomment = re.compile(r'/\\*((?:.|\\n)*?)\\*/')\n\ncomment.findall(text2)\n\nnew_comment = re.compile(r'/\\*([\\s\\S]*?)\\*/')\n\nnew_comment.findall(text2)\n\n# re.compile的 re.DOTALL\ncomment = re.compile(r'/\\*(.*?)\\*/', re.DOTALL)\n\ncomment.findall(text2)\n\n# ### 对于简单的情况使用re.DOTALL 标记参数工作的很好,但是如果模式非常复杂或者是为了构造字符串令牌而将多个模式合并起来(2.18 节有详细描述),这时候使用个标记参数就可能出现一些问题。如果让你选择的话,最好还是定义自己的正则表达���模式,这样它可以在不需要额外的标记参数下也能工作的很好。\n#\n#\n\n# # 将Unicode 文本标准化\n\ns1 = 'Spicy Jalape\\u00f1o'\ns2 = 'Spicy Jalapen\\u0303o'\n\ns1\n\ns2\n\ns1 == s2\n\nlen(s1)\n\nlen(s2)\n\nimport unicodedata\nt1 = unicodedata.normalize('NFC', s1)\nt2 = unicodedata.normalize('NFC', s2)\n\nt1\n\nt2\n\nt1 == t2\n\nlen(t1)\n\nt1 = unicodedata.normalize('NFD', s1)\nt2 = unicodedata.normalize('NFD', s2)\n\nt1\n\nlen(t1)\n\ns = '\\ufb01' # A single character\n\ns\n\nunicodedata.normalize('NFC', s)\n\nunicodedata.normalize('NFD', s)\n\nunicodedata.normalize('NFKD', s)\n\nunicodedata.normalize('NFKC', s)\n\nt1\n\n''.join(c for c in t1 if not unicodedata.combining(c))\n\n# # 在正则式中使用Unicode\n\n# +\n# return a match Object or None\n# re.match()\n# -\n\nnum= re.compile('\\d+')\nnum.match('212')\n\nnum.match('\\u0661\\u0662\\u0663')\n\narabic = re.compile('[\\u0600-\\u06ff\\u0750-\\u077f\\u08a0-\\u08ff]+')\n\npat = re.compile('stra\\u00dfe', re.IGNORECASE)\n\n\ns = 'straße'\npat.match(s)\n\n# does't match\npat.match(s.upper())\n\ns.upper()\n\n# ### 混合使用Unicode 和正则表达式通常会让你抓狂。如果你真的打算这样做的话,最好考虑下安装第三方正则式库,它们会为Unicode 的大小写转换和其他大量有趣特性提供全面的支持,包括模糊匹配。\n\n# # 删除字符串中不需要的字符\n\ns = ' hello world \\n'\n\ns.strip()\n\ns.lstrip()\n\ns.rstrip()\n\nt = '-----hello====='\n\nt.lstrip('-')\n\nt.rstrip('=')\n\nt.strip('-=')\n\ns = ' hello world \\n'\n\nre.sub('\\s+', ' ', s)\n\n# # 审查清理文本字符串\n\n# ### 文本清理问题会涉及到包括文本解析与数据处理等一系列问题。在非常简单的情形下,你可能会选择使用字符串函数(比如str.upper() 和str.lower() ) 将文本转为标准格式。使用str.replace() 或者re.sub() 的简单替换操作能删除或者改变指定的字符序列。你同样还可以使用2.9 小节的nicodedata.normalize() 函数将unicode文本标准化。\n# ### 有时候你可能还想在清理操作上更进一步。比如,你可能想消除整个区间上的字符或者去除变音符。为了这样做,你可以使用经常会被忽视的str.translate()方法。\n\ns = 'pýtĥöñ\\fis\\tawesome\\r\\n'\n\ns\n\n# +\nremap = {\n ord('\\t'): ' ',\n ord('\\f'): ' ',\n ord('\\r'): None,\n}\n\na = s.translate(remap)\n# -\n\na\n\n# +\nimport sys\n\ncmb_chrs = dict.fromkeys(c for c in range(sys.maxunicode) \n if unicodedata.combining(chr(c)))\n# -\n\nb = unicodedata.normalize('NFD', a)\n\nb\n\nb.translate(cmb_chrs)\n\n# +\n# 构造一个将所有Unicode 数字字符映射到对应的ASCII 字\n# 符上的表格\ndigitmap = {c: ord('0')+ unicodedata.digit(chr(c))\n for c in range(sys.maxunicode)\n if unicodedata.category(chr(c)) =='Nd'}\n\nlen(digitmap)\n# -\n\ndigitmap = { c: ord('0') + unicodedata.digit(chr(c))\n for c in range(sys.maxunicode)\n if unicodedata.category(chr(c)) == 'Nd' }\n\nlen(digitmap)\n\nx = '\\u0661\\u0662\\u0663'\n\nx.translate(digitmap)\n\n# ### 另一种清理文本的技术涉及到I/O 解码与编码函数。这里的思路是先对文本做一些初步的清理,然后再结合encode() 或者decode() 操作来清除或修改它\n\na\n\n\nb = unicodedata.normalize('NFD',a)\n\nb\n\nb.encode('ascii', 'ignore').decode('ascii')\n\n\n# ### 这里的标准化操作将原来的文本分解为单独的和音符。接下来的ASCII 编码/解码只是简单的一下子丢弃掉那些字符。当然,这种方法仅仅只在最后的目标就是获取到文本对应ACSII 表示的时候生效。\n\n# ### 文本字符清理一个最主要的问题应该是运行的性能。一般来讲,代码越简单运行越快。对于简单的替换操作, str.replace() 方法通常是最快的,甚至在你需要多次调用的时候\n#\n# ### 另一方面,如果你需要执行任何复杂字符对字符的重新映射或者删除操作的话,tanslate() 方法会非常的快。\n\ndef clean_spaces(s):\n s = s.replace('\\r', '')\n s = s.replace('\\t', ' ')\n s = s.replace('\\f', ' ')\n return s\n\n\n# # 字符串的对齐\n\ntext = 'Hello World'\n\ntext\n\ntext.ljust(20)\n\ntext.rjust(20)\n\ntext.center(20)\n\ntext.rjust(20,'=')\n\ntext.center(20,'*')\n\nformat(text, '>20')\n\nformat(text, '<20')\n\nformat(text, '^20')\n\nformat(text, '=>20s')\n\nformat(text, '*^20s')\n\n'{:>10s} {:>10s}'.format('Hello', 'World')\n\n# ### format() 函数的一个好处是它不仅适用于字符串。它可以用来格式化任何值,使得它非常的通用。\n\nx = 1.2345\n\nformat(x, '>10')\n\nformat(x,'<10.2f')\n\n# # 合并拼接字符串\n\nparts = ['Is', 'Chicago', 'Not', 'Chicago?']\n\n' '.join(parts)\n\n','.join(parts)\n\n# # 字符串中插入变量\n\ns = '{name} has {n} messages'\ns.format(name='Guido', n=37)\n\nname = 'tami'\nn = 520\ns.format_map(vars())\n\n\n# +\n# vars() 也是适用于对象实例\nclass Info:\n def __init__(self, name, n):\n self.name = name\n self.n = n\n\na= Info('kotoshi', 2)\ns.format_map(vars(a))\n\n\n# +\n# 处理缺省值\nclass safesub(dict):\n def __missing__(self, key):\n return '{' + key + '}'\n\ndel n\ns.format_map(safesub(vars()))\n# -\n\nimport sys\ndef sub(text):\n return text.format_map(safesub(sys._getframe(1).f_locals))\n\n\nname = 'guido'\nn =37\nprint(sub('hello {name}'))\n\nprint(sub('you have {n} messages'))\n\nprint(sub('your favorite color is {color}'))\n\ncolor = 'yellow'\n\nprint(sub('your favorite color is {color}'))\n\n# ### 比较老的一些使用方法\n\n'%(name) has %(n) messages.' % vars()\n\nname\n\nn\n\nimport string\ns= string.Template('$name has $n messages')\ns.substitute(vars())\n\n# # 以指定列宽格式化字符串\n\ns = \"Look into my eyes, look into my eyes, the eyes, the eyes, \\\nthe eyes, not around the eyes, don't look around the eyes, \\\nlook into my eyes, you're under.\"\n\nimport textwrap\nprint(textwrap.fill(s, 70))\n\nprint(textwrap.fill(s, 40))\n\nprint(textwrap.fill(s, 40, initial_indent = ' '))\n\nimport os\nos.get_terminal_size()\n# 在vscode里可以, 可能jupyter不适用\n\n# # 在字符串里处理html xml\n\ns = 'Elements are written as \"text\".'\nimport html\nprint(s)\n\nprint(html.escape(s))\n\nprint(html.escape(s, quote=False))\n\ns = 'Spicy Jalapeño'\n\ns.encode('ascii', errors='xmlcharrefreplace')\n\ns = 'Spicy "Jalapeño".'\nfrom html.parser import HTMLParser\np = HTMLParser()\np.unescape(s)\n\n\nt = 'The prompt is >>>'\nfrom xml.sax.saxutils import unescape\nunescape(t)\n\n# # 字符串令牌解析\n\ntext = 'foo = 23 + 42 * 10'\n\nimport re\nNAME = r'(?P[a-zA-Z_][a-zA-Z_0-9]*)'\nNUM = r'(?P\\d+)'\nPLUS = r'(?P\\+)'\nTIMES = r'(?P\\*)'\nEQ = r'(?P=)'\nWS = r'(?P\\s+)'\nmaster_pat = re.compile('|'.join([NAME, NUM, PLUS, TIMES, EQ, WS]))\n\nscanner = master_pat.scanner('foo = 42')\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n# _.lastgroup, _.group()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\n_.lastgroup, _.group()\n\nscanner.match()\n\nfrom collections import namedtuple\ndef generate_tokens(pat, text):\n Token = namedtuple('Token', ['type', 'value'])\n scanner = pat.scanner(text)\n for m in iter(scanner.match, None):\n yield Token(m.lastgroup, m.group())\n# Example use\nfor tok in generate_tokens(master_pat, 'foo = 42'):\n print(tok)\n\ntokens = (tok for tok in generate_tokens(master_pat, text)\n if tok.type != 'WS')\nfor tok in tokens:\n print(tok)\n\n# +\nPRINT = r'(Pprint)'\nNAME = r'(P[a-zA-Z][a-zA-z0-9]*)'\nmaster_pat = re.compile('|'.join([PRINT, NAME]))\n\nfor test in generate_tokens(master_pat, 'printer'):\n print(test)\n# -\n\n# # 实现一个简单的递归下降分析器\n\n# +\n# #!/usr/bin/env python\n# -*- encoding: utf-8 -*-\n\"\"\"\nTopic: 下降解析器\nDesc: \n\"\"\"\nimport re\nimport collections\n\n# Token specification\nNUM = r'(?P\\d+)'\nPLUS = r'(?P\\+)'\nMINUS = r'(?P-)'\nTIMES = r'(?P\\*)'\nDIVIDE = r'(?P/)'\nLPAREN = r'(?P\\()'\nRPAREN = r'(?P\\))'\nWS = r'(?P\\s+)'\n\nmaster_pat = re.compile('|'.join([NUM, PLUS, MINUS, TIMES, \n DIVIDE, LPAREN, RPAREN, WS]))\n\n# tokenizer\nToken = collections.namedtuple('Token', ['type', 'value'])\n\ndef generate_tokens(text):\n scanner = master_pat.scanner(text)\n for m in iter(scanner.match, None):\n tok = Token(m.lastgroup, m.group())\n if tok.type !='WS':\n yield tok\n\n\n# Parser\nclass ExpressionEvaluator:\n '''\n Implementation of a recursive descent parser. Each method\n implements a single grammar rule. Use the ._accept() method\n to test and accept the current lookahead token. Use the ._expect()\n method to exactly match and discard the next token on on the input\n (or raise a SyntaxError if it doesn't match).\n'''\n\n def parse(self, text):\n self.tokens = generate_tokens(text)\n self.tok = None # Last symbol consumed\n self.nexttok = None # Next symbol tokenized\n self._advance() # Load first lookahead token\n return self.expr()\n \n def _advance(self):\n 'Advance one token ahead'\n self.tok, self.nexttok = self.nexttok, next(self.tokens, None)\n\n def _accept(self, toktype):\n 'Test and consume the next token if it matches toktype'\n if self.nexttok and self.nexttok.type ==toktype:\n self._advance()\n return True\n else: \n return False\n \n def _expect(self, toktype):\n 'Consume next token if it matches toktype or raise SyntaxError'\n if not self._accept(toktype):\n raise SyntaxError('Ex[ected ' + toktype)\n \n # Grammer rules follow\n def expr(self):\n \"expression ::= term {('+'|'-') term }*\"\n exprval = self.term()\n while self._accept('PLUS') or self._accept('MINUS'):\n op = self.tok.type\n right = self.term()\n if op == 'PLUS':\n exprval += right\n elif op =='MINUS':\n exprval -= right\n return exprval\n \n def term(self):\n \"term ::= factor { ('*'|'/') factor }* \"\n termval = self.factor()\n while self._accept('TIMES') or self._accept('DIVIDE'):\n op = self.tok.type\n right = self.factor()\n if op == 'TIMES':\n termval *= right\n elif op == 'DIVIDE':\n termval /= right\n return termval\n \n def factor(self):\n \" factor ::= NUM | ( expr )\"\n if self._accept('NUM'):\n return int(self.tok.value)\n elif self._accept('LPAREN'):\n exprval = self.expr()\n self._expect('RPAREN')\n return exprval\n else:\n raise SyntaxError('Expected NUMBER or LPAREN')\n\n\ndef descent_parser():\n e = ExpressionEvaluator()\n print(e.parse('2'))\n print(e.parse('2+3'))\n print(e.parse('2+3*4'))\n print(e.parse('2+(3+4)*5'))\n\n\nif __name__ == '__main__':\n descent_parser()\n# -\n\n# ### 下面的代码不能运行在ipython上, 必须存为一个.py文件\n\n# +\n# plyexample.py\n#\n# Example of parsing with PLY\n\nfrom ply.lex import lex\nfrom ply.yacc import yacc\n\n# Token list\ntokens = [ 'NUM', 'PLUS', 'MINUS', 'TIMES', 'DIVIDE', 'LPAREN', 'RPAREN' ]\n\n# Ignored characters\n\nt_ignore = ' \\t\\n'\n\n# Token specifications (as regexs)\nt_PLUS = r'\\+'\nt_MINUS = r'-'\nt_TIMES = r'\\*'\nt_DIVIDE = r'/'\nt_LPAREN = r'\\('\nt_RPAREN = r'\\)'\n\n# Token processing functions\ndef t_NUM(t):\n r'\\d+'\n t.value = int(t.value)\n return t\n\n# Error handler\ndef t_error(t):\n print('Bad character: {!r}'.format(t.value[0]))\n t.skip(1)\n\n# Build the lexer\nlexer = lex()\n\n# Grammar rules and handler functions\ndef p_expr(p):\n '''\n expr : expr PLUS term\n | expr MINUS term\n '''\n if p[2] == '+':\n p[0] = p[1] + p[3]\n elif p[2] == '-':\n p[0] = p[1] - p[3]\n\ndef p_expr_term(p):\n '''\n expr : term\n '''\n p[0] = p[1]\n\ndef p_term(p):\n '''\n term : term TIMES factor\n | term DIVIDE factor\n '''\n if p[2] == '*':\n p[0] = p[1] * p[3]\n elif p[2] == '/':\n p[0] = p[1] / p[3]\n\ndef p_term_factor(p):\n '''\n term : factor\n '''\n p[0] = p[1]\n\ndef p_factor(p):\n '''\n factor : NUM\n '''\n p[0] = p[1]\n\ndef p_factor_group(p):\n '''\n factor : LPAREN expr RPAREN\n '''\n p[0] = p[2]\n\ndef p_error(p):\n print('Syntax error')\n\nparser = yacc()\n\nif __name__ == '__main__':\n print(parser.parse('2'))\n print(parser.parse('2+3'))\n print(parser.parse('2+(3+4)*5'))\n# -\n\n# # 字节字符串上的操作\n\ndata = bytearray(b'Hello World')\n\ndata[:5]\n\ndata.replace(b'Hello', b'Hello Cruel')\n\ndata = b'FOO:BAR,SPAM'\nimport re\nre.split('[:,]', data)\n\nre.split(b'[:,]', data)\n\n# ### some differences with standard string \n\na = 'Hello World'\na[0]\n\nb = b'Hello World'\nb[0]\n\nprint(a)\n\nprint(b) # Observe b'...'\n\n# +\n# formatting ? \nb'%10s %10d %10.2f' % (b'ACME', 100, 490.1)\n\n# It said there's no formatting for bytestring, but somehow formatting works = =;\n\n# +\nwith open ('halape\\xf10.txt', 'w') as f:\n f.write('spicy')\n \nimport os\nos.listdir('.')\n# -\n\nos.listdir(b'.')\n\n# # 第三章\n# # 数字日期和时间\n\n# ## 数字的四舍五入\n\nround(1.23,1)\n\nround(1.27,1)\n\nround(1.32455, 3)\n\n# ## 执行精确的浮点数运算\n\na = 4.2\nb = 2.1\na + b \n\n(a+b) == 6.3\n\nfrom decimal import Decimal\na = Decimal('4.2')\nb = Decimal('2.1')\na + b \n\n(a + b ) == Decimal('6.3')\n\n# Decimal can allow you to control the comoputation, like digit and round\na = Decimal('1.3')\nb = Decimal('1.7')\nprint(a/b)\n\nfrom decimal import localcontext\nwith localcontext() as ctx:\n ctx.prec = 3\n print(a/b)\n\nnums = [1.23e+18, 1, -1.23e+18]\nsum(nums)\n\nimport math\nmath.fsum(nums)\n\n# # 2 8 16 进制\n\nx = 1234\n\n\nbin(x)\n\nhex(x)\n\noct(x)\n\nformat(x, 'b')\n\nformat(x, 'x')\n\nformat(x, 'o')\n\n# ## 字节到大整数的打包与解包\n\n# ## 复数的数学运算\n\na = complex(2,4)\nb =3 - 5j\n\na\n\nb\n\na.real\n\na.imag\n\na + b\n\nc = 1 + 1j\n\nc\n\n\nabs(c)\n\nimport cmath\ncmath.sqrt(-1)\n\n# ## 无穷大与NaN\n\na = float('inf')\n\nb = float('-inf')\n\nc = float('nan')\n\na \n\nb \n\nc \n\nmath.isnan(c)\n\nmath.isinf(b)\n\n# ## 分数运算\n\nfrom fractions import Fraction\na = Fraction(2,3)\nb = Fraction(1,6)\n\nprint(a+b)\n\na.numerator\n\na.denominator\n\nc = 0.667\nFraction(c).limit_denominator(10)\n\nx = 3.75\n# y = Fraction()\nx.as_integer_ratio()\n\ny = Fraction(*x.as_integer_ratio())\n\ny\n\n# ## 大型数组运算 & 矩阵雨线性代数\n\n# ### 用numpy\n\n# # 随机选择\n\nimport random\nvalues = [1,2,3,4,5,6]\n\nrandom.choice(values)\n\nrandom.choice(values)\n\nrandom.sample(values,3)\n\nrandom.shuffle(values)\n\nvalues\n\nrandom.randint(0,10)\n\n# 生成0到1 范围内均匀分布的浮点数, 用random.random()\nrandom.random()\n\n# 获得 N 位随机二进制的整数\nrandom.getrandbits(3)\n\n\nrandom.seed(233)\nrandom.randint(1,100)\n\nrandom.randint(1,100)\n\n# +\n# random.uniform() random.gauss()\n# -\n\n# ### 在random 模块中的函数不应该用在和密码学相关的程序中。如果你确实需要类似的功能,可以使用ssl 模块中相应的函数。\n\n# # 基本的日期与时间转换\n\n# 表示一个时间段 用timedelta\nfrom datetime import timedelta\na = timedelta(days=2, hours=3)\nb = timedelta(hours=6.5)\na + b \n\n(a + b).seconds / 3600\n\n(a + b).hours\n\n(a + b).total_seconds() / 3600\n\nfrom datetime import datetime\na = datetime(2008, 8, 8)\n\na + timedelta(days=10)\n\nb = datetime(2023, 7, 4)\nb - a \n\nprint(datetime.today())\n\nprint(datetime.today() + timedelta(minutes= 10))\n\n# ### 如果要处理更加复杂的日期, 用dateutil\n\na \n\na + timedelta(months=1)\n\nfrom dateutil.relativedelta import relativedelta\na + relativedelta(months=+10)\n\nb\n\nb - a \n\nd = relativedelta(b,a)\n\nd \n\n# # 查找最后一个周五的日期\n\n# #!/usr/bin/env python\n# -*- encoding: utf-8 -*-\n\"\"\"\nTopic: 最后的周五\nDesc :\n\"\"\"\nfrom datetime import datetime, timedelta\nweekdays = ['Monday', 'Tuesday', 'Wednesday', 'Thursday',\n'Friday', 'Saturday', 'Sunday']\ndef get_previous_byday(dayname, start_date=None):\n if start_date is None:\n start_date = datetime.today()\n day_num = start_date.weekday()\n day_num_target = weekdays.index(dayname)\n days_ago = (7 + day_num - day_num_target) % 7\n if days_ago == 0:\n days_ago = 7\n target_date = start_date - timedelta(days=days_ago)\n return target_date\n\n\ndatetime.today()\n\nget_previous_byday('Friday')\n\nday_num = datetime.today().weekday()\n\nday_num\n\nday_num_target = 4\n\n# 4 is the index of Friday\ndays_ago = (7 + day_num - day_num_target) % 7\n\ndays_ago\n\ndatetime.today() - timedelta(days= days_ago)\n\nget_previous_byday('Tuesday', datetime(2023,7,1))\n\n# ### 如果你要像这样执行大量的日期计算的话,你最好安装第三方包python-dateutil来代替。\n\nfrom datetime import datetime\nfrom dateutil.relativedelta import relativedelta\nfrom dateutil.rrule import *\nd = datetime.now()\n\n\nd \n\n# next Friday\nprint(d + relativedelta(weekday=FR))\n\n# previous 2 Friday\nprint(d + relativedelta(weekday=FR(-2)))\n\n# # 计算当前月份的日期范围\n\nfrom datetime import datetime, date, timedelta\nimport calendar\ndef get_month_range(start_date=None):\n if start_date is None:\n start_date = date.today().replace(day=1)\n _, days_in_month = calendar.monthrange(start_date.year, start_date.month)\n end_date = start_date + timedelta(days=days_in_month)\n return (start_date, end_date)\n\n\na_day = timedelta(days=1)\nfirst_day, last_day = get_month_range()\nwhile first_day < last_day:\n print(first_day)\n first_day += a_day\n\ndate.today()\n\ndate.today().replace(day=1)\n\n\ndef date_range(start, stop, step):\n while start < stop:\n yield start\n start += step\n\n\nfor d in date_range(datetime(2023, 7, 1), datetime(2023, 8, 1), timedelta(hours=6)):\n print(d)\n\n# # 字符串转换为日期\n\nfrom datetime import datetime\ntext = '2012-9-23'\ny = datetime.strptime(text, '%Y-%m-%d')\nz = datetime.now()\ndiff = z - y\n\ndiff\n\nz\n\n# 性能差,可以自己写一个日期的解析\nnice_z = datetime.strftime(z, '%A %B %d, %Y')\n\nnice_z\n\n# # 结合时区的日期操作\n\nfrom datetime import datetime\nfrom pytz import timezone\nd = datetime(2012, 12, 21, 9, 30, 0)\nprint(d)\n\n# localize the date for Chicago\ncentral = timezone('US/Central')\nloc_d = central.localize(d)\n\nloc_d\n\nbang_d = loc_d.astimezone(timezone('Asia/Kolkata'))\n\nbang_d\n\n\n# ### 注意美国夏时令的转换 & pytz不建议使用了\n\n# # 第四章 迭代器与生成器\n\n# ## 手动遍历迭代器\n\ndef manual_iter():\n with open('./lines.txt') as f:\n try:\n while True:\n line = next(f)\n print(line, end='')\n except StopIteration:\n pass\n\n\nmanual_iter()\n\n\ndef manual_iter():\n with open('./lines.txt') as f:\n while True:\n line = next(f)\n if line is None:\n break\n print(line, end='')\n\n\nmanual_iter()\n\nitems = [1, 2, 3]\nit = iter(items)\n\nnext(it)\n\nnext(it)\n\nnext(it)\n\nnext(it)\n\n\n# ## 代理迭代\n\n# +\nclass Node:\n def __init__(self, value):\n self._value = value\n self._children = []\n \n def __repr__(self):\n return f'Node({self._value})'\n \n def add_child(self, node):\n self._children.append(node)\n \n def __iter__(self):\n return iter(self._children)\n\n\nif __name__ == '__main__':\n root = Node(0)\n child1 = Node(1)\n child2 = Node(2)\n root.add_child(child1)\n root.add_child(child2)\n \n for ch in root:\n print(ch)\n\n\n# -\n\n# ## 使用生成器创建新的迭代模式\n\n# +\ndef frange(start, stop, increment):\n x = start\n while x < stop:\n yield x\n x += increment\n \nfor n in frange(0,4,0.5):\n print(n)\n# -\n\nfrange(2,3,0.1)\n\n\n# ## 实现迭代器协议\n\n# +\nclass Node:\n def __init__(self, value):\n self._value = value\n self._children = []\n \n def __repr__(self):\n return f'Node({self._value})'\n \n def add_child(self, node):\n self._children.append(node)\n \n def __iter__(self):\n return iter(self._children)\n \n def depth_first(self):\n yield self\n for c in self:\n yield from c.depth_first()\n \n\nif __name__ == '__main__': \n root = Node(0)\n child1 = Node(1)\n child2 = Node(2)\n root.add_child(child1)\n root.add_child(child2)\n child1.add_child(Node(3))\n child1.add_child(Node(4))\n child2.add_child(Node(5))\n \n for ch in root.depth_first():\n print(ch)\n# -\n\n# # 反向迭代\n\na = [1,3,5,7]\nfor x in reversed(a):\n print(x)\n\nf = open('./lines.txt')\nfor line in reversed(list(f)):\n print(line, end='')\n\nf.close()\n\n\n# +\nclass Countdown:\n def __init__(self, start):\n self.start = start\n \n # forward iterator\n def __iter__(self):\n n = self.start\n while n > 0:\n yield n\n n -= 1\n \n # reverse iterator\n def __reversed__(self):\n n = 1\n while n <= self.start:\n yield n\n n += 1\n \n\nfor rr in reversed(Countdown(30)):\n print(rr)\n \nprint('*'*20) \n \nfor rr in Countdown(30):\n print(rr)\n \n# -\n\n# # 带有外部状态的生成器函数\n\n# ### 如果你想让你的生成器暴露外部状态给用户,别忘了你可以简单的将它实现为一个类,然后把生成器函数放到iter () 方法中过去。\n\nfrom collections import deque\nclass linehistory:\n def __init__(self, lines, histlen=3):\n self.lines = lines\n self.history = deque(maxlen=histlen)\n \n def __iter__(self):\n for lineno, line in enumerate(self.lines, 1):\n self.history.append((lineno, line))\n yield line\n\n def clear(self):\n self.history.clear()\n\n\nwith open('somefile.txt') as f:\n lines = linehistory(f)\n for line in lines:\n if 'python' in line:\n for lineno, hline in lines.history:\n print('*{}:{}'.format(lineno, hline), end='')\n\n\n# ### 没有看懂上面的结果\n\n# # 迭代器切片\n\n# +\ndef count(n):\n while True:\n yield n\n n += 1\n \nc = count(0)\nc[10:20]\n# -\n\nimport itertools\nfor x in itertools.islice(c, 10, 20):\n print(x)\n\n# ### islice 用过就废,如需再次访问,先放到一个列表里\n\n# # 跳过可迭代对象的开始部分\n\nfrom itertools import dropwhile\nwith open('/etc/passwd') as f:\n for line in dropwhile(lambda line: line.startswith('#'), f):\n print(line, end='')\n\nfrom itertools import islice\nitems = ['a', 'b', 'c', 1, 3, 5, 9]\nfor x in islice(items, 3, None):\n print(x)\n\n# # 排列组合的迭代\n\nitems = ['a', 'b', 'c']\nfrom itertools import permutations\nfor p in permutations(items):\n print(p)\n\nfor p in permutations(items, 2):\n print(p)\n\nfrom itertools import combinations\nfor c in combinations(items, 2):\n print(c)\n\nfrom itertools import combinations_with_replacement\nfor c in combinations_with_replacement(items, 3):\n print(c)\n\n# # 序列上索引值迭代\n\nmy_list = ['a', 'b', 'c']\nfor idx, val in enumerate(my_list):\n print(idx, val)\n\nfor idx, val in enumerate(my_list, 1):\n print(idx, val)\n\n\ndef parse_data(filename):\n with open(filename, 'rt') as f:\n for lineno, line in enumerate(f, 1):\n fields = line.split()\n try:\n count = int(fields[1])\n ...\n except ValueError as e:\n print('Line {}: Parse error: {}'.format(lineno, e))\n\n\n# +\nfrom collections import defaultdict\nword_summary = defaultdict(list)\nwith open('somefile.txt', 'r') as f:\n lines = f.readlines()\n\nfor idx, line in enumerate(lines):\n # Create a list of words in current line\n# print(line)\n# print(line.split())\n words = [w.strip().lower() for w in line.split()]\n for word in words:\n word_summary[word].append(idx)\n# -\n\nword_summary\n\nss = 'this '\nss.strip()\n\n# # 同时迭代多个序列\n\na = [1, 2, 3]\nb = ['w', 'x', 'y', 'z']\nfor i in zip(a,b):\n print(i)\n\nfrom itertools import zip_longest\nfor i in zip_longest(a,b):\n print(i)\n\nfor i in zip_longest(a, b, fillvalue=0):\n print(i)\n\nheaders = ['name', 'shares', 'price']\nvalues = ['ACME', 100, 490.1]\ns = dict(zip(headers, values))\n\ns\n\n# ### zip可以接受多于两个的序列的参数 & 返回的是一个迭代器\n\na = [1, 2, 3]\nb = [10, 11, 12]\nc = ['x','y','z']\nfor i in zip(a, b, c):\n print(i)\n\n# # 不同集合上元素的迭代\n\n# ### 多个容器, 但不想写多次循环\n\nfrom itertools import chain\na = [1,2,3,4]\nb = ['x', 'y', 'z']\nfor x in chain(a,b):\n print(x)\n\nfor x in a+b:\n ...\n# 这种要比上面的低效\n\n# # 创建数据处理管道\n\n# +\nimport os\nimport fnmatch\nimport gzip\nimport bz2\nimport re\n\ndef gen_find(filepat, top):\n '''\n Find all filenames in a directory tree that match a shell wildcard pattern \n '''\n for path, dirlist, filelist in os.walk(top):\n for name in fnmatch.filter(filelist, filepat):\n yield os.path.join(path, name)\n \n\ndef gen_opener(filenames):\n '''\n Open a ssequence of filenames one at a time producing a file object.\n The fle is closed immediately when proceeding t othe next iteration.\n '''\n for filename in filenames:\n if filename.endswith('.gz'):\n f = gzip.open(filename, 'rt')\n elif filename.endswith('.bz2'):\n f = bz2.open(filenamee, 'rt')\n else:\n f = open(filename, 'rt')\n yield f\n f.close()\n \ndef gen_concatenate(iterators):\n '''\n Chain a sequence of iterators together into a single sequence\n '''\n for it in iterators:\n yield from it\n \ndef gen_grep(pattern, lines):\n '''\n Look for a regex pattern in a sequence of lines\n '''\n pat = re.compile(pattern)\n for line in lines:\n if pat.search(line):\n yield line\n \n \n# -\n\nlognames = gen_find('access-log*', 'www')\nfiles = gen_opener(lognames)\nlines = gen_concatenate(files)\npylines = gen_grep('(?i)python', lines)\nfor line in pylines:\n print(line)\n\nlognames = gen_find('access-log*', 'www')\nfiles = gen_opener(lognames)\nlines = gen_concatenate(files)\npylines = gen_grep('(?i)python', lines)\nbytecolumn = (line.rsplit(None,1)[1] for line in pylines)\nbytes = (int(x) for x in bytecolumn if x != '-')\nprint('Total', sum(bytes))\n\n# # 展开嵌套的序列\n\n# +\n# 在vscode里可行\nfrom collections import Iterable\n\ndef flatten(items, ignore_types=(str, bytes)):\n for x in items:\n if isinstance(x, Iterable) and not isinstance(x, ignore_types):\n yield from flatten(x)\n else:\n yield x\n\nitems = [1,2,[3,4, [5,6], 7], 8]\nfor x in flatten(items):\n print(x)\n# -\n\nitems_2 = ['Dave', 'Paula', ['Thomas', 'Lewis']] \nfor x in flatten(items_2):\n print(x)\n\n# # 顺序迭代合并后的排序迭代对象\n\nimport heapq\nx = [1,4,7,10]\ny= [2,5,6,11]\nfor z in heapq.merge(x,y):\n print(z)\n\na = ['a', 'c', 'z']\nb = ['j', 'k' ,'b']\nfor c in heapq.merge(a,b):\n print(c)\n\nwith open('sorted_file_1', 'rt') as file1, \\\n open('sorted_file_2', 'rt') as file2, \\\n open('merged_file', 'wt') as outf:\n for line in heapq.merge(file1, file2):\n outf.write(line)\n\n# # 迭代器代替while 无限循环\n\nimport sys\nf = open('/etc/passwd')\nfor chunk in iter(lambda: f.read(10), ''):\n n = sys.stdout.write(chunk)\n\n\n","repo_name":"LightRoastKenyaPeaberry/ipython_notes","sub_path":"python_cookbook/python_cookbook_1_to_4.ipynb","file_name":"python_cookbook_1_to_4.ipynb","file_ext":"py","file_size_in_byte":50199,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"34151305078","text":"# # [AI 이노베이션 스퀘어] 11기 기본 저녁반\n# - github: https://github.com/hysKim1/AI_Fundamental\n# - [파이썬 공식문서]https://docs.python.org/ko/3/\n# - [Numpy 공식문서]https://numpy.org/devdocs/\n# - [ML]https://developers.google.com/machine-learning/crash-course/ml-intro?hl=ko\n# - [DL]https://www.tensorflow.org/\n# ---\n#\n# * [1. layers API ](#layers )
\n# * [ weights](#weights)\n# * [variables](#variables)\n# * [ Layer Class 구성 ](#Layer-Class-구성)\n# * [\\_\\_init__](#\\_\\_init__)\n# * [\\_\\_call__](#\\_\\_call__)\n# * [build()](#build)\n# * [call()](#call)\n# * [built](#call)\n# * [build 따로 호츌한 경우](#build-따로-호츌한-경우)\n# * [build 없는 경우](#build-없는-경우)\n#\n# * [응용 : Sequential model, Model 상속](#응용-:-Sequential-model)\n#\n# * [2. 학습 방법 4가지 ](#학습-방법-4가지 )
\n# * [2.1.train](#1.train)\n# * [2.2.fit](#2.fit)\n# * [batch](#batch)\n# * [epochs](#epochs)\n# * [verbose](#verbose)\n# * [callbacks](#callbacks)\n# * [validation_split](#validation_split)\n# * [class_weight](#class_weight)\n# * [initial_epoch](#initial_epoch)\n# * [steps_per_epoch](#steps_per_epoch)\n# * [workers](#workers)\n# * [trainable](#trainable)\n# * [2.3.train_on_batch](#3.train_on_batch)\n# * [train_step](#train_step)\n# * [2.4.GradientTape](#4.GradientTape)\n# ---\n\n# ## 모뎅 생성 방법 \n#\n# 1. tensor 이용 처음부터 끝까지 함 (low level)\n# - 처음부터 끝까지 다 만듦\n# - 지원하지 않는 기능을 만들어 사용할 때 씀\n# - tensor 와 nn package\n# - `tf.nn.relu`\n# 2. estimator\n# - 이미 만들어진 estimator 가져다가 씀\n#\n# keras 에서도 저수준으로 생성가능하여 권고함\n# \n# 3. Keras: `Sequential`\n# - 단순,단방향\n# - 편리\n# 4. Keras: `Model`\n# 1. functional paradigm\n# - 이론,실제 간극을 줄임\n# 2. multi-input,multi-output 지원\n# - 복잡한 모델을 만들 수 있음\n# 3. 미리 만들어진 기능 사용\n# 4. 유지 보수 쉬움\n#\n# 5. Keras: `Model` 상속(하위 클래스 sublass)\n# - 만들어진 모델에 기능 추가\n# - 상속해서 사용자 맞춤 레이어, 모델 사용 가능 \n# - 예.`Callback` 상속\n# - 동적 사용 가능(if, while 이용 조건에 따라서 바꿈)\n#\n#\n# - 혼재 사용되어 모든 것의 이해가 필요\n# - numpy array programmming , python 의 functional paradigm을 알아야 쉽게 이해 할 수 있음 \n# - 생성 모델마다 학습 방법이 조금씩 다름\n\nimport tensorflow as tf\n\n\n# #### Callback\n# - `Callbacks`: 모델 훈련 중 특정 지점에서 호출된 유틸리티\n# - Callback을 상속받아 메소드를 overriding하면 customizing 가능\n\nclass MyCallback(tf.keras.callbacks.Callback): #상속 예시 기억나시나요?\n pass \n\n\n# ## layers\n#\n# - Keras layers API.\n#\n#\n# - `Model`, `Sequential`은 Input(데이터 크기), output layer(예측 갯수) 모양만 맞춰주면 알아서 자동으로 맞춰줌\n#\n# - `Model`, `Sequential` 사용하지 않기 때문에 `input_shape` 입력 없이도 에러 안나고 계산됨\n\n# functional paradigm\nlayer= tf.keras.layers.Dense(100, input_shape=(3,)) #input layer\nlayer(tf.constant([[1,2,3]])) #\n\nx=tf.constant([[1,2,3]]) #위와 동일\nlayer(x) #3개의 input 이 100개의 node에 각 곱해지게됨\n\nlayer \n\ndir(layer) #weigths, variables, bias, kernel...\n\n# ### weights\n# layer 의 weigths 를 리스트 형태로 보여줌\n# keras의 `weight/variable` :bias, kernel\n#\n\nlayer.weights\n\n# 각 kernel, bias array를 unapcking 하여 나눠 저장\n\nlen(layer.weights) #2개 list\n\nkernel,bias =layer.weights #unpacking technique\n\nkernel #가중치 (3,100)\n\n# ### variables\n# - `weights`가 이론에서 가리키는 가중치와 혼동되어 사용하는 별칭(alias)\n\nlayer.variables == layer.weights #둘이 동일 \n\nlayer.variables\n\n\n# ### Layer Class 구성\n#\n# > 실행순서\n# 0. init\n# - layer에 필요한 매개변수를 입력\n# 1. __call__\n# - build, call() 순서대로 호출\n# 2. build\n# - 입력의 크기를 받고 나머지 초기화\n# - 명시적으로 크기 지정할 필요없이 입력 바탕으로 자동으로 kernel 크기 맞춰 생성\n# - __call__로 부터 한 번만 \n# 3. call()\n# - forward compuation ( 정방향 연산) : 행렬 곱으로 feed forward함\n#\n#\n# ### \\_\\_init__\n# - class 안에 정의하는 special/magic method `__ __`\n# - 인스턴스 생성하여 초기화하는 생성자\n\nclass X:\n def __init__(self):\n print('A')\n\n\nX\n\nX()\n\n\n# `super`:자식 클래스가 부모 클래스로부터 상속받은 멤버를 참조할 때 사용하는 참조 변수\n\n#\n# tf 공식문서 맞춤 레이어 참고: https://www.tensorflow.org/tutorials/customization/custom_layers?hl=ko\n# \n# \n# - tf.keras.Layer 클래스를 상속하고 하단처럼 구현\n\nclass MyDenseLayer(tf.keras.layers.Layer):\n def __init__(self, num_outputs):\n super().__init__()\n print('__init__')\n self.num_outputs = num_outputs\n\n def build(self, input_shape):\n print('build')\n print(input_shape)\n self.kernel = self.add_weight(\"kernel\",\n shape=[int(input_shape[-1]),\n self.num_outputs])\n print(self.kernel.shape)\n\n \n def call(self, input):\n print('call')\n return tf.matmul(input, self.kernel)\n\n# 4개의 output이며 아직 총 몇개의 데이터가 입력 받을지 모름\n# ( None,4)\n\ny=MyDenseLayer(4) #num_outputs =4\n\n# ### \\_\\_call__\n# - 객체에 `()` 붙일 수 있음\n# - **클래스의 인스턴스를 callale하게 만들어 내부적으로 함수처럼 호출**\n# - `build()` 존재하면 호출(optional)\n# - self.kernel 초기화\n# - 자동으로 input, output에 따라서 variables(kernel,bias) 크기 맞춰 줌\n# - `call()` 호출\n\n# build를 정확히 언제 호출하는지 알아야함\n# ### build\n# - optional\n# - 자동으로 input, output에 따라서 variables(kernel,bias) 크기 맞춰 줌\n# - 행렬 곱 예: (l, m)@ (m, n) @ (n, y) @ (y a) = (l,a)\n# \n# > 응용: Sequentials\n# - 입력 레이어: `input_shape=(x,a)` 지정 해주면 알아서 맞춰 줌\n#\n# > 응용:` Model ( inputs= ,ouputs= )`지정 해주면 알아서 맞춰 줌\n#\n\ny=MyDenseLayer(4) #num_outputs =4\n\n# 1) build 호출 , input_shape=( 2,3)\n# - self.kernel =(3,4) 자동으로 맞춰줌\n# 2) call 호출\n# - 행렬곱 : input @ self.kernel = (2,3)@(3,4)=(2,4) \n\n# + active=\"\"\n# input @ weights= outputs\n# (2,3) @ (3,4) = (2,4)\n# -\n\ny(tf.zeros((2,3))) #input_shape=(2,3)\n\ny.kernel #self.kernel (3,4) 생성됨\n\n# ### call\n# - `call()`을 직접적으로 부르면 `build()` 호출 안함\n#\n# 기존의 self.kernel = (3,4 )을 가져다 씀\n# - input의 (행,열)에서 열을 3으로 맞추니 가능\n# - 행렬 곱 : (1,3)@(3,4)=(1,4)\n\ny.call(tf.zeros((1,3))) # 직접 call하면 build 안부르지만 kernel 크기에 맞춰 바꾸면 가능\n\n# InvalidArgumentError: Matrix size-incompatible: In[0]: [2,4], In[1]: [3,4] [Op:MatMul]\n# - build를 부르지 않기 때문에 위에서 저장된 값 기반으로 실행\n# - input=(2,4)와 kernel에서 나온 self.kernel (3,4) 행렬곱을 크기가 맞지 않아 할 수가 없음.\n\ny.call(tf.zeros((2,4))) # 직접 call하면 build 안 부른 경우 기존의 kenel 크기 안맞으면 에러 \n\n# AttributeError: 'MyDenseLayer' object has no attribute 'kernel'\n# - build 부르지 않기 때문에 self.kernel 생성되지 않아 에러 남\n\ny=MyDenseLayer(4) #1) __init__ outputs=4\ny.call(tf.zeros((2,4))) #2) call()\n\ny.kernel #self.kernel 없기 때문에 없음\n\n# #### built\n# - `build()` 호축 되었는지 안 되었는지 여부를 boolean으로 반환값으로 확인\n# - build 호출 됨: x.built=True\n\nx=MyDenseLayer(4) #1) __init__ outputs=4\nx.call(tf.zeros((2,4))) #2) call() input_shape ( 2,4) ->build 호출 안 함\n\nx.built # build 호출 안됨 = False\n\nx(tf.zeros((2,4))) #build 호출 \n\n# `build` 는 인스턴스 한 다음 최초 한번만 호출됨\n# - 코드 출력값에 에 `build` 없음\n\nx(tf.zeros((2,4))) #build 호출 더이상 없음\n\nx.built # build 호출 = True\n\nx.built=False\n\nx(tf.zeros((2,4))) #build 다시 재호출 \n\n# #### build 따로 호츌한 경우\n# - build에 입력된 값과 call 입력된값 일치시켜야 함\n\ndd=MyDenseLayer(4) #num_outputs=4\n\n# + active=\"\"\n# input @ weights= outputs\n# (3,2) @ (2,4) = (3,4)\n# -\n\ndd.build((3,2)) #build 부르면\n\ndd.call(tf.ones((3,2))) #기존에 생성된 커널과 동일해야만 작동\n\ndd.kernel\n\n# + active=\"\"\n# input @ weights= outputs\n# (1,9) @ (2,4) = (1,4)\n# -\n\ndd(tf.ones((1,9))) #build 재호출 되어 문제 없음\n\n\n# #### build 없는 경우\n# - 가능 하긴 하나 기본적으로 `Layer`, `Model` 클래스에서는 대부분 있으므로 상속해서 overriding 하려면 알아야함\n\nclass MyDenseLayer(tf.keras.layers.Layer):\n def __init__(self, num_outputs):\n super().__init__()\n print('__init__')\n self.num_outputs = num_outputs\n\n def call(self, input_):\n print('call')\n self.kernel = self.add_weight(\"kernel\",\n shape=[int(input_.shape[-1]),\n self.num_outputs])\n print(self.kernel)\n return tf.matmul(input_, self.kernel)\n\n\n# + active=\"\"\n# input @ kenel = output\n# (2,6) @ (6,3) = (2,3)\n# -\n\nb=MyDenseLayer(2) \n\nb.built #build 호출 없음\n\nb.call(tf.ones((4,3))) \n\nb.kernel #kernel 생성됨\n\nb(tf.ones((3,4))) # call : self.kernel에서 (6,3)생성 됨\n\n\nclass MyDenseLayer(tf.keras.layers.Layer):\n def __init__(self, num_outputs):\n super().__init__()\n print('__init__')\n self.num_outputs = num_outputs\n\n def call(self, input_):\n print('call')\n print(input_)\n return tf.matmul(input_, tf.ones((input_.shape[-1],self.num_outputs) ))\n# 하단 부분 변경 \n\n\nb=MyDenseLayer(2) #build 없는 경우 \n\nb.built\n\nb(tf.ones((4,3))) #build 없어도 가능\n\nb.call(tf.ones((4,3))) #상동\n\n# ### 응용 : Sequential model\n# - functional paradigm 가능\n\n#instance 방식\nmodel= tf.keras.Sequential([ #input_shape없음 - summary 불가 \n \n tf.keras.layers.Dense(2,activation='relu', name='layer1'),\n tf.keras.layers.Dense(3,activation='relu', name='layer2'),\n tf.keras.layers.Dense(4 ,name='layer3')\n])\n\nmodel.built #build 안 불림\n\n# `dir` build가 있는 것은 입력 크기 자동으로 맞춰주는 기능 갖춰주어 input,output 크기만 맞추면 됨\n\ndir(model) # __call__ ->callble->functional paradigm , build-> 자동 크기 맞춰줌\n\nx=tf.ones((3,5)) #input_shape= (3,5)\ny=model(x)\n\nmodel.built #build 불림\n\n# 코드를 기반으로 tensor 크기를 하단과 같이 알 수 있음\n# - input=(3,5)\n# - layer1 node 2 -> (5,2 )\n# - layer2 node 2 -> (2,2)\n# - layer3 node 4 -> output ( 2 , 4)\n\n# + active=\"\"\n# 행렬 곱 : (3,5)@(5,2)@(2,2)@(2,4)=(3,4)\n# -\n\nmodel(tf.ones((3,5))) #내부��으로 build를 부름\n\n# + active=\"\"\n# 1. (3,5)@(5,2)=(3,2)\n# 2. (3,2)@(2,2)=(3,2)\n# 3. (3,2)@(2,4)=(3,4) <- 최종\n# -\n\nmodel.summary() #위와 같은 방식으로 자동으로 맞춰주어 Output Shape 나옴\n\n# ### 응용: 상속\n#\n# #### 레이어 상속\n\n# - 단순하게 layer 묶은 것이기 때문애 모델안에 모델 들어갈 수 있음 \n# - layer과 동일한 접근\n\n# +\n#instance model\nmodel= tf.keras.Sequential([ \n \n tf.keras.layers.Dense(2,activation='relu', name='layer1'),\n tf.keras.layers.Dense(2,activation='relu', name='layer2'),\n tf.keras.layers.Dense(4 ,name='layer3')\n])\n\nmodel1= tf.keras.Sequential([\n model, # 위에거 가져옴\n tf.keras.layers.Dense(2,activation='relu', name='layer1')\n\n])\n# -\n\nmodel1.layers[0].layers # 단순히 묶음\n\n\n# #### Model 상속\n# - 상황에 따라 동적 프로그래밍 가능하여 복잡한 모델 생성 가능\n# - `Model` class 상속하는 고급 기법\n# - \\_\\_init__ \n# - flatten & dense 레이어 2개 \n# - call\n# \n# 전문가를 위한 빠른 시작 공식 문서 참고:https://www.tensorflow.org/tutorials/quickstart/advanced?hl=ko\n\n# +\n#https://www.tensorflow.org/tutorials/quickstart/advanced?hl=ko\n\nclass MyModel(tf.keras.models.Model):\n def __init__(self):\n super(MyModel, self).__init__()\n self.flatten = tf.keras.layers.Flatten()\n self.d1 = tf.keras.layers.Dense(128, activation='relu')\n self.d2 = tf.keras.layers.Dense(10, activation='softmax')\n\n def call(self, x):\n if x.shape[-1]>3 : #조건문에 따른 동적 프로그래밍 가능!\n x = self.flatten(x)\n x = self.d1(x)\n return self.d2(x)\n else:\n return self.d2(x)\n\n\n# -\n\nmodel=MyModel() #안스턴스화\n\ndir(model) # __call__ ->callble->f\n\nmodel(tf.ones(10,5)) # _call_ -> build(),call()\n\n# ---\n# 초보자를 위한 시작 https://www.tensorflow.org/tutorials/quickstart/beginner?hl=ko\n\n# +\n#초보자를 위한 시작 https://www.tensorflow.org/tutorials/quickstart/beginner?hl=ko\n\nmnist = tf.keras.datasets.mnist\n\n(x_train, y_train), (x_test, y_test) = mnist.load_data()\nx_train, x_test = x_train / 255.0, x_test / 255.0\n# -\n\n#Sequential\nmodel = tf.keras.models.Sequential([\n tf.keras.layers.Flatten(input_shape=(28, 28)),\n tf.keras.layers.Dense(128, activation='relu'),\n tf.keras.layers.Dropout(0.2),\n tf.keras.layers.Dense(10, activation='softmax')\n])\n\n# +\n# Model - 상동\ninputs=tf.keras.Input(shape=(28,28))\nflatten=tf.keras.layers.Flatten()(inputs)\n\nlayer1=tf.keras.layers.Dense(128,activation='relu')(flatten)\ndropout= tf.keras.layers.Dropout(0.2)(layer1)\noutput= tf.keras.layers.Dense(10, activation='softmax')(dropout)\n\nmodel1=tf.keras.models.Model(inputs=inputs,outputs=output)\n# -\n\nmodel.built\n\nmodel.summary() #모델 생성 완료\n\n# #### add_ method\n#\n# #### add_loss\n# - compile( loss=) 에서 제공하는 것 외에 loss function 을 사용하고 싶을때 사용\n# - subclassed layer or model의 상속을 받는 하위 클래스 내부에 'call' 함수에서 사용 가능\n#\n# stackoverflow 참고 :https://stackoverflow.com/questions/50063613/what-is-the-purpose-of-the-add-loss-function-in-keras\n\n# + active=\"\"\n# #https://keras.io/api/layers/base_layer/#add_loss-method\n# #call 에 사용한 예시\n# class MyLayer(tf.keras.layers.Layer):\n# def call(self, inputs):\n# self.add_loss(tf.abs(tf.reduce_mean(inputs)))\n# return inputs\n# -\n\n# - `SparseCategoricalCrossentropy`: label encoding 다중 분류문제에 사용\n\nmodel.add_loss(tf.keras.losses.SparseCategoricalCrossentropy())\n\n# add_loss loss가 2개 돌아가기 때문에 한 번 정하면 compile 해도 변경이 안됨\n# - 사용자 정의 학습 할때 사용\n# - compile -> fit\n\nmodel.compile(loss=tf.keras.losses.SparseCategoricalCrossentropy())\n\nmodel.fit(x_train,y_train,epochs=5)\n\n# +\n#한 가지 예시로 \ninputs = tf.keras.Input(shape=(28,28))\nx = tf.keras.layers.Dense(10)(inputs)\noutputs = tf.keras.layers.Dense(1)(x)\nmodell = tf.keras.Model(inputs, outputs)\n\n# Activity regularization- 종속성이 다른 경우\n\nmodell.add_loss(tf.abs(tf.reduce_mean(x)))\n# -\n\nmodell.compile(loss='sparse_categorical_crossentropy', \n optimizer='adam')#- fit 사용하기 위해서 compile 해야함`\n\nmodell.fit(x_train,y_train,epochs=5) #activity regularization 으로 가능\n\n# - 보통 `compile (loss, metrics)`에서 제공되지 않는 경우\n# - 활성화만 요구할 수 있으며(이 경우에는 대상이 없음), 이 활성화는 모델 출력이 아닐 수 있다.\n#\n# - ActivityRegularizationLayer 경우에는 activation of a layer만 필요함(target 불필요) \n# - 커스텀한 layer의 call method 로 self.add_loss(loss_value)를 호출 가능.\n# - 이러한 방식으로 추가된 손실을 학습 동안에 `compile`로 보내어지는 loss에 추가된다. \n#\n# TF 공식문서 학습과 평가 참고 : https://www.tensorflow.org/guide/keras/train_and_evaluate\n\n# + active=\"\"\n# # A layer that creates an activity regularization loss 클래스 예시\n# class ActivityRegularizationLayer(tf.keras.layers.Layer):\n# def call(self, inputs):\n# self.add_loss(tf.reduce_sum(inputs) * 0.1) #호출\n# return inputs # Pass-through layer.\n#\n# #functional paradigm\n# # activity regularization 를 레이어 처럼 사용\n# inputs = tf.keras.Input(shape=(28,28))\n# x = tf.keras.layers.Dense(64, activation=\"relu\", name=\"dense_1\")(inputs)\n# x = ActivityRegularizationLayer()(x) \n#\n# x = tf.keras.layers.Dense(64, activation=\"relu\", name=\"dense_2\")(x)\n# outputs = tf.keras.layers.Dense(10, name=\"pred\")(x)\n#\n# model = tf.keras.Model(inputs=inputs, outputs=outputs)\n# model.compile(\n# optimizer=tf.keras.optimizers.RMSprop(learning_rate=1e-3),\n# loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),\n# )\n#\n#\n# model.fit(x_train, y_train, batch_size=64, epochs=1)\n# -\n\n# ## 학습 방법 4가지\n# 상황에따라서 학습 방법이 다름\n# - 모델 만드는법 5가지 : tensor , estimator, keras(Sequentials, Model 함수형 패러다임, Model subclass)\n# - 학습 방법 4가지: train,fit, train_on_batch, GradientTape\n#\n# 혼용 주의\n#\n# ## 1.train\n# - estimator는 scikit-learn이랑 유사.\n# - compile 없이 바로 학습 \n# - 둘 다 한번에 layer,node 갯수 정함\n# - 상세한 변경은 불가하여 맞춤 학습이 어려움\n\n# + active=\"\"\n# #scikit-learn\n# from sklearn.neural_network import MLPClassifier\n# mlp = MLPClassifier(hidden_layer_sizes=(30,102,30))\n# mlp.fit(x_train, y_train)\n# -\n\n# Init signature:\n# MLPClassifier(\n# hidden_layer_sizes=(100,),\n# activation='relu',\n# solver='adam',\n# alpha=0.0001,\n# batch_size='auto',\n# learning_rate='constant',\n# learning_rate_init=0.001,\n# power_t=0.5,\n# max_iter=200,\n# shuffle=True,\n# random_state=None,\n# tol=0.0001,\n# verbose=False,\n# warm_start=False,\n# momentum=0.9,\n# nesterovs_momentum=True,\n# early_stopping=False,\n# validation_fraction=0.1,\n# beta_1=0.9,\n# beta_2=0.999,\n# epsilon=1e-08,\n# n_iter_no_change=10,\n# max_fun=15000,\n# )\n\n# + active=\"\"\n# # estimator\n# # Build a DNN with 2 hidden layers with 30 and 10 hidden nodes each.\n#\n# classifier = tf.estimator.DNNClassifier(\n# feature_columns=my_feature_columns,\n# # Two hidden layers of 30 and 10 nodes respectively.\n# hidden_units=[30, 10],\n# # The model must choose between 3 classes.\n# n_classes=3)\n#\n#\n# estimator.train(\n# input_fn=(x_train,y_train), hooks=None, steps=None, max_steps=None, saving_listeners=None\n# )\n# -\n\n# ## 2.fit\n# - 상황에 맞게 조절하여 사용 \n# \n# ### batch\n# - 학습 시 몇 개마다 업데이트 할 것인지\n# - 크기가 클수록 빠르나 정확도가 떨어짐 \n# - 최소 1 ~ 최대 전체 샘플 갯수\n# - 비유:수학 문제를 다 풀고 답을 풀것인지 나눠서 확인하고 다 풀것인지..\n#\n# ### epochs \n# - 데이터 전체를 몇번 학습 시킬지\n\ndir(model1) #fit, train_ ...\n\n# ### verbose \n# - 설명 양 \n# - verbose: Integer. 0, 1, or 2(- 숫자카 커질수록 설명 양이 적어짐)\n# > Verbosity mode. \n# - 0 = silent, \n# - 1 = progress bar, \n# - 2 = one line per epoch\n# - 3 >= epoch\n\nmodel1.compile(loss='sparse_categorical_crossentropy', \n optimizer='adam') \nmodel1.fit(x_train,y_train,epochs=3, verbose=0)\n\nmodel1.fit(x_train,y_train,epochs=3, verbose=1)\n\nmodel1.fit(x_train,y_train,epochs=3, verbose=2)\n\nmodel1.fit(x_train,y_train,epochs=3, verbose=4)\n\n\n# ### callbacks\n# - 한 번 실행하고 나서 epoch 실행되게 함\n# - 학습과 검증 과정에서 적용할 keras.callbacks.Callback 인스턴스의 리스트.\n# - 학습 과정의 특정 단계에서 적용할 함수의 세트\n# - keras.callbacks.Callback :확작해서 커스텀 콜백 생성 가능\n# - keras.callbacks.BaseLogger\n# - keras.callbacks.TerminateOnNaN\n# - keras.callbacks.ProgbarLogger\n# - keras.callbacks.History\n# - keras.callbacks.ModelCheckpoint\n# - keras.callbacks.RemoteMonitor\n# - LearningRateScheduler\n# - keras.callbacks.TensorBoard\n# - keras.callbacks.ReduceLROnPlateau\n# - keras.callbacks.CSVLogger\n#\n# keras 공식 문서 참고 : https://keras.io/ko/callbacks/#_1\n#\n\n# +\n# 각 batch에 대한 loss 내역 기록\nclass LossHistory(keras.callbacks.Callback): #상속 -> self.model 접근 가능\n def on_train_begin(self, logs={}):\n self.losses = []\n\n def on_batch_end(self, batch, logs={}):\n self.losses.append(logs.get('loss'))\n\n \nmodel = Sequential()\nmodel.add(Dense(10, input_dim=784, kernel_initializer='uniform'))\nmodel.add(Activation('softmax'))\nmodel.compile(loss='categorical_crossentropy', optimizer='rmsprop')\n\nhistory = LossHistory() #인스턴스화 한 후 fit에 사용\nmodel.fit(x_train, y_train, batch_size=128, epochs=20, verbose=0, callbacks=[history])\n\nprint(history.losses)\n\n# +\n# epoch마다 모델 저장\nfrom keras.callbacks import ModelCheckpoint\n\nmodel = Sequential()\nmodel.add(Dense(10, input_dim=784, kernel_initializer='uniform'))\nmodel.add(Activation('softmax'))\nmodel.compile(loss='categorical_crossentropy', optimizer='rmsprop')\n\n'''\n만약 검증에 대하 loss가 감소하다면 각 epoch 후의 모델의 가중치를 저장함\n'''\ncheckpointer = ModelCheckpoint(filepath='/tmp/weights.hdf5', verbose=1, save_best_only=True)\nmodel.fit(x_train, y_train, batch_size=128, epochs=20, verbose=0, validation_data=(X_test, Y_test), callbacks=[checkpointer])\n# -\n\n# ### validation_split\n# - 내부적으로 hold out으로 기본 0.2는 전체의 80%는 학습 나머지 20%은 성능검정에 사용해서 매 epoch 마다 val_loss 출력\n\nmodel1.fit(x_train,y_train,epochs=3, validation_split=0.2) #val_loss 생김\n\n# predict 단순히 input 을 넣어서 예측값 넣음\n# 차이를 줄이는것이 loss 함수이며 줄여가는 과정을 학습\n# - back propagation \n# - 뒤로가면서 loss가 가장 작은 weight값 찾음\n\n# ### class_weight\n# - imbalanced dataset : 데이터가 불균형한 경우 특정 데이터들은 잘 학습이 안되기 때문에 데이터들에 가중치를 더 주어 집중 학습\n# - [1,1,1,1,1,1,1,1,0,0] 데이터에서 0은 수가 절대적으로 적어서 학습이 잘 안됨\n#\n# ### sample_weight\n# - 전문가,초보자가 수집한 데이터중 신뢰성이 다르기 때문에 데이터 별로 가중치를 신뢰도에 따라 다르게 줌\n\n# ### initial_epoch\n# - 학습 시작시 몇번째 epcoch부터 시작할지 정하여 건너 뛰고 학습\n# - 학습시킬때마다 누적되기 때문에 학습을 시작하는 epcoh (이전의 학습 과정을 재개하는데 유용)\n\nmodel1.fit(x_train,y_train,epochs=3, initial_epoch=1)\n\nmodel1.fit(x_train,y_train,epochs=3, initial_epoch=2)\n\n# ### steps_per_epoch\n# - step: Weight와 Bias를 1회 업데이트\n# $steps_per_epoch=\\frac {no samples X epochs}{batch_size}$\n# - 전체 데이터 하나(1 epoch)에 대해서 weight,Bias를 업데이트 하는 횟수(총 step횟수=batches of samples)\n\nmodel1.fit(x_train,y_train,epochs=3, steps_per_epoch=3)\n\nmodel1.fit(x_train,y_train,epochs=3,batch_size=60000) #상동\n\n# 계산이 안 맞는경우 다음과같이 됨.\n# - `steps_per_epoch * epochs` batches 갯수까지 가능한데 초과 하므로 학습 경고메시지 생김\n# - steps_per_epoch 먼저 맞춤 \n# - 4 *3 =12 개의 batches 가능하나 최대 batch_size= 60,000/12=5,000개\n#\n\nmodel1.fit(x_train,y_train,epochs=3,batch_size=60000,steps_per_epoch=4) #경고\n\nmodel1.fit(x_train,y_train,epochs=3,batch_size=5000,steps_per_epoch=4) #잘 됨\n\n# ### workers\n# - scikit-learn의 `n_jobs`: 동시에 core 몇개 사용할지\n# - 가속을 위한 프로세스의 최대 갯수를 정수형으로 입력 \n\nmodel1.fit(x_train,y_train,epochs=3, workers=5)\n\ndir(model1) #train_step\n\n# ### trainable\n# - 독립적으로 학습 시킬지 아닐지 여부를 정함\n# - 'freezing' 이라 불리우며 `fit`학습 시킬때 weights 업데이트 시키지 않고 그대로 가져오기 때문\n#\n# 참고: https://www.tensorflow.org/guide/keras/transfer_learning\n\n# +\ninputs=tf.keras.Input(shape=(28,28))\nflatten=tf.keras.layers.Flatten()(inputs)\nlayer1=tf.keras.layers.Dense(128,activation='relu')(flatten)\n\nlayer1.trainable=False # 추가 frozen layer1\n\ndropout= tf.keras.layers.Dropout(0.2)(layer1)\noutput= tf.keras.layers.Dense(10, activation='softmax')(dropout)\nmodel1=tf.keras.models.Model(inputs=inputs,outputs=output)\n\n\nmodel1.compile(loss='sparse_categorical_crossentropy')\n# -\n\n# - 육안으로 변화 없음 trainable에대한 summary 버그라는 글 찾음: https://github.com/keras-team/keras/issues/4674\n#\n#\n# - 학습시킬수 있는 파라미터수 : Trainable params: 101,770\n#\n\nmodel1.summary() \n\nmodel1.layers # 모든 레이어 속함\n\nmodel1.layers[-1].weights #ㅐoutput layer \n\n# Non-trainable params: 0 에서 1,290 이 됨.\n\nmodel1.layers[-1].trainable=False #freezing output layer \nmodel1.summary()\n\nmodel1.layers[-1].weights #안변함\n\n# 모델에 적용하면 해당 모델에 속한 레이어 전체에 적용됨\n# - Non-trainable params: 101,770 전체\n\nmodel1.trainable=False #모든 레이어 \nmodel1.summary() \n\n# Runs a single gradient update on a single batch of data.\n# 1의 batch 데이터에서 업데이트\n\nmodel1.fit(x_train,y_train,epochs=3)\n\nmodel.weights #초기 가중치 가져옴\n\n# ## 3.train_on_batch\n# - 1 batch 당 1번의 gradient업데이트로 얻은 training loss \n# - fit의 저수준 버전\n\nmodel1.train_on_batch(x_train,y_train) #fit과 비슷\n\n# - fit은 하단을 train_on_batch for 이용하여 array연산 \n# - 4 levels : train > fit > **train_on_batch** > GradientTape\n\nfor i in range(3):\n print(model1.train_on_batch(x_train,y_train))\n\n# #### train_step \n# - `fit()`에서 반복적 호출( tf.version 2.2 이상 부터 )하여 사용자가 정의한 학습 loop을 조정 가능함\n# https://www.tensorflow.org/api_docs/python/tf/keras/Model\n\n# ## 자동 미분\n# - 학습: gradient descent 통해 loss function 최소화하는 파라미터 찾음\n# - loss function : 입력 & 실체값 차이를 함수로 표현\n# - gradient descent : 현재 위치와 learning rate곱하여 이동하면서 loss/cost function 최소값(미분값=0)을 주는 파라미터를 찾음\n# - gradient : 특정점의 기울기\n# - learning rate $\\alpha$: 현재 그래디언트에서 다음 위치까지 얼마만큼 가야하는지\n# - 미분 : 하나의 변수 \n# - 편미분: 변수가 여러개 인 경우 각각에 대해 미분)\n# - 자동 미분 :auto differentiation\n# - tensorflow 지원 4가지: `train`,`fit`, `train_on_batch`, `GradientTape`\n#\n# ## 4.GradientTape\n# - 1970년대에 인기 저장 매체로 tape 사용하여 영상물 기록\n# - 주어진 입력에 대한 연산의 gradient에서 최소점을 찾기위해 학습한 것이 어떻겨 변화하는지 편미분값 저장\n# - 가장 저주순\n# - 미분하기위해서 미분하는 함수를 GradientTape에 넣으면 자동미분 할 수 있도록 구조체로 변경\n# - 변수가 여러개 이므로 편미분을 통해서함 \n#\n# tf 공식문서 자동미분과 그래디어트 테이프 참고 : https://www.tensorflow.org/tutorials/customization/autodiff?hl=ko\n# - 자세한것은 다음 시간에 이어서\n\n# + active=\"\"\n# # 데이터의 batches를 반복하여 사용\n#\n# for x_batch_train, y_batch_train in train_dataset:\n# with tf.GradientTape() as tape:\n# logits = layer(x_batch_train) # Logits for this minibatch\n# # Loss value for this minibatch\n# loss_value = loss_fn(y_batch_train, logits)\n# # Add extra losses created during this forward pass:\n# loss_value += sum(model.losses)\n#\n# grads = tape.gradient(loss_value, model.trainable_weights)\n# optimizer.apply_gradients(zip(grads, model.trainable_weights))\n","repo_name":"hysKim1/AI_Fundamental","sub_path":"200820_11기_기본저녁반_필기.ipynb","file_name":"200820_11기_기본저녁반_필기.ipynb","file_ext":"py","file_size_in_byte":28573,"program_lang":"python","lang":"en","doc_type":"code","stars":2,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"4408696176","text":"# + id=\"OLqeQ2zISoxH\" colab_type=\"code\" colab={}\nimport gdown\nimport os\nfrom pandas_profiling import ProfileReport\n\n# https://drive.google.com/file/d/1l_J0P9A_AD8d_rzZHJ5Fg8F4y1nGP_x3/view?usp=sharing\n\nurl = f'https://drive.google.com/uc?id=1l_J0P9A_AD8d_rzZHJ5Fg8F4y1nGP_x3'\nfilename = 'dataset.csv'\nif not os.path.exists(filename):\n gdown.download(url, filename, quiet=True)\n\n# + id=\"O8lpxHGCTRJ0\" colab_type=\"code\" colab={}\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\n# + id=\"FvhCE49JTUA1\" colab_type=\"code\" colab={}\ncolumns = ['emotion', 'text']\n\ndf = pd.read_csv(filename, names=columns)\n\n# + id=\"gSSP9JxBTg9X\" colab_type=\"code\" colab={}\nXraw = df['text'].values\nyraw = df['emotion'].values\n\n# + [markdown] id=\"ujQ3AdDST8Ow\" colab_type=\"text\"\n# ### Preprocessing\n\n# + id=\"iEIB57-UU8K4\" colab_type=\"code\" outputId=\"7ae49867-ef47-4942-f2fa-ed3285f7c940\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 104}\nimport nltk\n\nnltk.download(\"punkt\")\nnltk.download('stopwords')\n\n# + id=\"mna_HRCvZThv\" colab_type=\"code\" outputId=\"2e25b4b2-2ad7-46e6-c3c2-e10ccb1dee40\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34}\n# !pip install autocorrect\n\n# + id=\"9jUUMijzTul2\" colab_type=\"code\" colab={}\nfrom nltk.tokenize import word_tokenize\nfrom nltk.corpus import stopwords\nfrom autocorrect import Speller\n\nspell = Speller(lang='en')\n\nimport re\n\ndef process_text(text):\n text = text if type(text) == str else ''\n\n # clean the words, remove symbols special chars\n pattern = r'[^a-zA-z0-9\\s]'\n text = re.sub(pattern, '', text)\n\n # convert to lowercase\n text = text.lower()\n\n # first tokenize the text\n word_tokenized = word_tokenize(text)\n\n # let's remove the stop words\n en_stopwords = stopwords.words('english')\n words_swords_removed = [spell(word) for word in word_tokenized if word not in en_stopwords]\n\n return words_swords_removed\n\n\n# + id=\"CFtBXFT1VHuo\" colab_type=\"code\" colab={}\nfrom sklearn.feature_extraction.text import CountVectorizer\n\n# + id=\"KruKnL3oVdxQ\" colab_type=\"code\" colab={}\ncvector = CountVectorizer(tokenizer=process_text)\n\n# + id=\"RqqxOeqwV-3m\" colab_type=\"code\" colab={}\nX_tr = cvector.fit_transform(Xraw)\n\n# + id=\"1MNBe5SfWTBR\" colab_type=\"code\" outputId=\"0b745028-5561-430a-80cc-485b5c4e9e14\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 52}\nX_tr\n\n# + id=\"SsAx6FIgZuro\" colab_type=\"code\" outputId=\"9ee1fc4b-2d0b-4182-b005-19644c11dd36\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 1000}\nhelp(X_tr)\n\n# + id=\"9ljbyaI9aWJ_\" colab_type=\"code\" colab={}\nfrom sklearn.preprocessing import LabelEncoder\n\n# + id=\"zBzuXHu-bW4c\" colab_type=\"code\" colab={}\nlabel_encoder = LabelEncoder()\n\n# + id=\"Yldq3yGIba_3\" colab_type=\"code\" outputId=\"52f51f1a-5e84-4400-8694-b16257021393\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34}\nlabel_encoder.fit(yraw)\n\n# + id=\"-OKIBgWsbdDw\" colab_type=\"code\" outputId=\"50e55445-eeeb-4640-c81d-18a6649c0241\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 52}\nlabel_encoder.classes_\n\n# + id=\"S5eFQTSqblAE\" colab_type=\"code\" outputId=\"b4cee2ea-8771-4a25-9956-bb6d8739640f\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34}\ny_tr = label_encoder.transform(yraw)\ny_tr\n\n# + id=\"T6w3XIVQbrz_\" colab_type=\"code\" outputId=\"c534769c-afde-440b-df1e-9a259ec5002c\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34}\ny_tr.shape\n\n# + [markdown] id=\"slPdEoaLcv9a\" colab_type=\"text\"\n# ### Support vector machine classifier\n\n# + id=\"-F4VMMdHb2FI\" colab_type=\"code\" colab={}\nfrom sklearn.svm import SVC\n\n# + id=\"GxiXIknyfgRz\" colab_type=\"code\" colab={}\nfrom sklearn.model_selection import train_test_split\n\n# + id=\"LtpTJ1SbfqpB\" colab_type=\"code\" outputId=\"0f3c06c4-2515-442a-a9e0-4530829800b8\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 34}\nX_train, X_test, y_train, y_test = train_test_split(X_tr, y_tr, test_size=0.3, shuffle=True)\n\nprint(f'X_train: {X_train.shape}, X_test: {X_test.shape}, y_train: {y_train.shape}, y_test: {y_test.shape}')\n\n# + id=\"KbyFw6j9gqyr\" colab_type=\"code\" outputId=\"f13980be-796a-4640-f312-ae48cb98475b\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 425}\nplt.figure(figsize=(10, 6))\n\nplt.subplot(1, 2, 1)\nsns.distplot(y_train)\nplt.title('Training set')\n\nplt.subplot(1, 2, 2)\nsns.distplot(y_test)\nplt.title('Test set')\nprint(np.bincount(y_train))\nprint(np.bincount(y_test))\n\n# + [markdown] id=\"8UUd-mrxh4up\" colab_type=\"text\"\n# The distribution of training set and test set seems similar.\n\n# + id=\"SURoqpBHdAVe\" colab_type=\"code\" outputId=\"9163c56c-1e68-47da-85c2-ff7cba82b256\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 86}\nmodel = SVC(kernel='poly', degree=2)\nmodel.fit(X_train, y_train)\n\n# + id=\"X_3MjIevdPFw\" colab_type=\"code\" colab={}\ny_train_pred = model.predict(X_train)\ny_test_pred = model.predict(X_test)\n\n# + id=\"fcYsJ-lEdS0V\" colab_type=\"code\" outputId=\"e8e39238-6cec-47d4-d54b-913215f949b4\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 538}\nfrom sklearn.metrics import classification_report\n\nprint('-' * 10 + ' Training Set ' + '-' * 10)\nprint(classification_report(y_train, y_train_pred))\n\nprint('-' * 10 + ' Testing Set ' + '-' * 10)\nprint(classification_report(y_test, y_test_pred))\n\n# + [markdown] id=\"uNvKZdNYdj97\" colab_type=\"text\"\n# With a simple support vector machine classifier, we have got an accuracy of $81\\%$ on training set but just $56\\%$ accuracy on the test set. We have to try other models or improve this for better accuracy for using baseline model.\n","repo_name":"nbnbhattarai/emotion-detection-system","sub_path":"notebooks/3_0_base_model_experimentation_svm.ipynb","file_name":"3_0_base_model_experimentation_svm.ipynb","file_ext":"py","file_size_in_byte":5543,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"72488829604","text":"# + id=\"pHnIE8IhjNKQ\"\nfrom PIL import Image\n\n# Buka gambar yang ingin diubah menjadi grayscale\ninput_image_path = \"Upin.jpeg\"\noutput_image_path = \"Upin abu.jpg\"\nimage = Image.open(input_image_path)\n\n# Konversi gambar ke skala abu-abu\ngrayscale_image = image.convert(\"L\")\n\n# Simpan gambar skala abu-abu\ngrayscale_image.save(output_image_path)\n\n# Tampilkan gambar skala abu-abu (opsional)\ngrayscale_image.show()\n\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 449} id=\"gxgUC96FkKg4\" outputId=\"4dc74bf9-177f-4f7a-844b-49aada64b53c\"\nfrom PIL import Image\nfrom IPython.display import display\n\n# Load the grayscale image\ngrayscale_image = Image.open(\"Upin abu.jpg\")\n\n# Display the grayscale image\ndisplay(grayscale_image)\n\n\n# + [markdown] id=\"WMZPrPLjpIsN\"\n# Resizing\n#\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"Ba6Z_1KipKVU\" outputId=\"7f08810a-60e4-4001-d51d-2341f75e7fb8\"\nfrom PIL import Image\n\n# Load the image\ninput_image_path = \"Upin.jpeg\"\nimage = Image.open(input_image_path)\n\n# Dapatkan lebar dan tinggi gambar dalam piksel\nwidth, height = image.size\n\n# Cetak lebar dan tinggi gambar\nprint(f\"Lebar gambar: {width} piksel\")\nprint(f\"Tinggi gambar: {height} piksel\")\n\n\n# + id=\"6o264HyApnTc\"\n# Define the new dimensions (width x height) in pixels\nnew_width = 400\nnew_height = 500\n\n# Resize the image\nresized_image = image.resize((new_width, new_height))\n\n# Save the resized image\noutput_image_path = \"resized_Upin.jpg\"\nresized_image.save(output_image_path)\n\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 252} id=\"B-nuc5H5qOYG\" outputId=\"6a50d21c-ebe3-4e7f-f89f-903911f16b18\"\nfrom PIL import Image\nfrom IPython.display import display\n\n# Load the grayscale image\nresize_image = Image.open(\"resized_Upin.jpg\")\n\n# Display the grayscale image\ndisplay(resize_image)\n\n# Dapatkan lebar dan tinggi gambar dalam piksel\nwidthresize, heightresize = resize_image.size\n\n# Cetak lebar dan tinggi gambar\nprint(f\"Lebar gambar: {widthresize} piksel\")\nprint(f\"Tinggi gambar: {heightresize} piksel\")\n\n# + [markdown] id=\"v6ec1xCyvLMM\"\n# Blurring Image\n#\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 881} id=\"QP683B0KvM7U\" outputId=\"1916dda2-5e53-4cf5-d832-4926aefdcb5c\"\nimport cv2\nimport numpy as np\nimport matplotlib.pyplot as plt\n# Load the image\ninput_image_path = \"Upin.jpeg\"\nimage = cv2.imread(input_image_path)\n\n# Apply Gaussian blur\nkernel_size = (5, 5) # Adjust the kernel size as needed\nblurred_image = cv2.GaussianBlur(image, kernel_size, 0)\n\n# Save the blurred image\noutput_image_path = \"blurred_Upin.jpg\"\ncv2.imwrite(output_image_path, blurred_image)\n\n# Display the original and blurred images (optional)\n\nplt.subplot(121), plt.imshow(cv2.cvtColor(image, cv2.COLOR_BGR2RGB), cmap='gray'), plt.title('Original Image')\nplt.xticks([]), plt.yticks([])\nplt.subplot(122), plt.imshow(cv2.cvtColor(blurred_image, cv2.COLOR_BGR2RGB), cmap='gray'), plt.title('Blurred Image')\nplt.xticks([]), plt.yticks([])\nplt.show()\n\n\n# + [markdown] id=\"VzqOQpg70Ly0\"\n# Reducing Noise dengan LPF\n#\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 467} id=\"jHIRGZO1qpnz\" outputId=\"30e848e0-1565-4072-a141-f3ef4ab2c130\"\nimport cv2\nimport numpy as np\n\n# Baca citra dengan noise (contoh: citra berumur)\nimage_with_noise = cv2.imread('Noise.jpg') # Ganti 'citra_noise.jpg' dengan nama file citra Anda\n\n# Terapkan filter Gaussian untuk mengurangi noise\nkernal_size = (7,7) # Ukuran kernel (biasanya ganjil)\nsigma_x = 0 # Nilai sigma dalam arah X (0 untuk otomatis menghitung berdasarkan ukuran kernel)\nsigma_y = 0 # Nilai sigma dalam arah Y (0 untuk otomatis menghitung berdasarkan ukuran kernel)\n\n# Terapkan filter rata-rata (LPF) untuk mengurangi noise\nimage_without_noise = cv2.GaussianBlur(image_with_noise, kernal_size, sigma_x, sigma_y)\n\n# Tampilkan citra asli dan citra tanpa noise\ncv2.imshow('Original Image', image_with_noise)\ncv2.imshow('Image Without Noise', image_without_noise)\n# Save the image\noutput_image_path = \"reduce_Noise.jpg\"\ncv2.imwrite(output_image_path,image_without_noise)\n\n# Tunggu sampai pengguna menekan tombol apapun\ncv2.waitKey(0)\ncv2.destroyAllWindows()\n\n\n# + [markdown] id=\"-ir_6-SI3hGC\"\n# Reducing Noise dengan Median Filter\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 484} id=\"CyFFskvi6GyK\" outputId=\"ad6b2fe1-13fb-4781-b972-66ad7cf41a31\"\nimport cv2\nimport numpy as np\n\n\n# Baca gambar dari Google Drive (ganti dengan path yang sesuai)\nimage_path = 'Noise.jpg' # Ganti dengan path gambar Anda\nimage = cv2.imread(image_path)\n\n# Konversi gambar ke grayscale (opsional, tergantung pada jenis gambar Anda)\ngray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)\n\n# Terapkan Median Filter\nkernel_size = 3 # Cobalah berbagai ukuran kernel\nimage_filtered = cv2.medianBlur(gray, kernel_size)\n\n# Tampilkan gambar asli dan hasil Median Filter\ncv2.imshow('Original Image', gray)\ncv2.imshow('Filtered Image', image_filtered)\n# Tunggu sampai pengguna menekan tombol apapun dan kemudian tutup jendela\ncv2.waitKey(0)\ncv2.destroyAllWindows()\n\n# Simpan gambar hasil Median Filter (jika diperlukan)\ncv2.imwrite('hasil_filtered.jpg', image_filtered) # Ganti dengan path penyimpanan yang sesuai\n\n\n# + [markdown] id=\"ISM3jnLS1JdQ\"\n# Cropping Objek\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 73} id=\"przNbF6l4bCI\" outputId=\"8eea6ed7-03f0-43a2-c1e8-9e4dc7f607ce\"\nimport cv2\nfrom matplotlib import pyplot as plt\n\n# Tentukan nama file gambar yang diunggah\nimage_filename = 'Upin.jpeg'\n\n# Muat gambar menggunakan OpenCV\nimage = cv2.imread(image_filename)\n\n\n# + id=\"036Zym9q1XLf\"\nx1, y1 = 100, 100 # Koordinat kiri atas\nx2, y2 = 300, 300 # Koordinat kanan bawah\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 406} id=\"YcRG26Af1gfF\" outputId=\"7d1ff232-091c-4fe0-b4ad-d4d53a05826e\"\ncropped_image = image[y1:y2, x1:x2]\n\n# Konversi warna gambar dari BGR ke RGB (jika diperlukan)\ncropped_image_rgb = cv2.cvtColor(cropped_image, cv2.COLOR_BGR2RGB)\n\n# Save the image\noutput_image_path = \"Upin_potong.jpg\"\ncv2.imwrite(output_image_path,cropped_image_rgb)\n\n# Tampilkan gambar\nplt.imshow(cropped_image_rgb)\nplt.axis('off')\nplt.show()\n\n","repo_name":"Holylight5/CV_Sesi5","sub_path":"CV5_ImageProcessing_Dirga.ipynb","file_name":"CV5_ImageProcessing_Dirga.ipynb","file_ext":"py","file_size_in_byte":6108,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"3274707759","text":"# # Chapter 7. 앙상블 학습과 랜덤 포레스트\n#\n# 무작위로 선택된 수천명의 사람에게 복잡한 질문을 하고 대답을 모은다고 가정합시다. 많은 경우 이렇게 모은 답이 전문가의 답보다 낫습니다. 이를 *대중의 지혜*라고 합니다.\n#\n# 이와 비슷하게 일련의 예측기로부터 예측을 수집하면 가장 좋은 모델 하나보다 더 좋은 예측을 얻을 수 있을 것입니다. 일련의 예측기를 **앙상블**이라고 부르기 때문에 이를 **앙상블 학습**(ensemble learning)이라고 하며, 앙상블 학습 알고리즘을 앙상블 방법이라고 합니다. \n#\n# 예를 들어 훈련 세트로부터 무작위로 각기 다른 서브셋을 만들어 일련의 결정 트리 분류기를 훈련시킬 수 있습니다. 예측을 하려면 모든 개별 트리의 예측을 구하면 됩니다. 그런 다음 가장 많은 선택을 받은 클래스를 예측으로 삼습니다. 결정 트리의 앙상블을 **랜덤 포레스트**(random forest)라고 합니다. 가장 간단한 방법이지만, 오늘날 가장 강력한 머신러닝 알고리즘 중 하나입니다.\n#\n#\n# 이 장에서는 **배깅, 부스팅, 스태킹** 등 가장 인기있는 앙상블 방법을 설명하겠습니다.\n#\n#\n\n# 시작하기 전에 먼저 필요한 모듈 import!\nimport matplotlib.pyplot as plt\nimport matplotlib\nmatplotlib.rc('font', family='AppleGothic')\nplt.rcParams['axes.unicode_minus'] = False\n\n# ## 1. 투표 기반 분류기\n#\n# 정확도가 80%인 분류기 여러개를 훈련시켰다고 가정합시다. 아마도 로지스틱 회귀 분류기, SVM 분류기, 랜덤 포레스트 분류기, k-최근접 이웃 분류기 등을 가지고 있을 수 있습니다.\n#\n# 더 좋은 분류기를 만드는 매우 간단한 방법은 각 분류기의 예측을 모아서 가장 많이 선택된 클래스를 예측하는 것입니다. 이렇게 다수결 투표로 정해지는 분류기를 **직접 투표**(hard voting) 분류기라고 합니다.\n#\n# 조금 놀랍게도, 이 다수결 투표 분류기가 앙상블에 포함된 개별 분류기 중 가장 뛰어난 것보다도 정확도가 높을 경우가 많습니다. 사실 각 분류기가 약한 학습기일지라도, 많고 다양하다면 앙상블은 강한 학습기가 될 수 있습니다.\n#\n# 다음은 여러 분류기를 조합하여 사이킷런의 **투표 기반 분류기**(`VotingClassifier`)를 만들고 훈련시키는 코드입니다.\n#\n\n# +\n# moons 데이터셋을 읽어와 학습시켜봅시다!\nfrom sklearn.datasets import make_moons\nfrom sklearn.model_selection import train_test_split\n\nX, y = make_moons(n_samples=500, noise=0.30, random_state=42)\nX_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)\nprint(X_train.shape, X_test.shape)\n\n# +\nfrom sklearn.ensemble import RandomForestClassifier\nfrom sklearn.ensemble import VotingClassifier\nfrom sklearn.linear_model import LogisticRegression\nfrom sklearn.svm import SVC\n\nlog_clf = LogisticRegression(solver='liblinear')\nrnd_clf = RandomForestClassifier(n_estimators=10)\nsvm_clf = SVC(gamma='auto')\n\nvoting_clf = VotingClassifier(\n estimators=[('lr', log_clf), ('rf', rnd_clf), ('svc', svm_clf)],\n voting='hard'\n)\n\nvoting_clf.fit(X_train, y_train)\n# -\n\n# 각 분류기의 테스트셋 정확도를 확인해봅시다.\n\n# +\nfrom sklearn.metrics import accuracy_score\n\nfor clf in (log_clf, rnd_clf, svm_clf, voting_clf):\n clf.fit(X_train, y_train)\n y_pred = clf.predict(X_test)\n print(clf.__class__.__name__, accuracy_score(y_test, y_pred))\n# -\n\n# VoingClassifier가 꽤 좋은 성적을 보이고 있는 것 같죠? \n#\n# 모든 분류기가 클래스의 확률을 예측할 수 있으면 (즉, `predict_proba()`메서드가 있으면), 개별 분류기의 예측의 평균을 내어 확률이 가장 높은 클래스를 예측할 수 있습니다. 이를 **간접 투표**(soft voting)이라고 합니다. 이 방식은 확률이 높은 투표에 비중을 두기 때문에 직접 투표 방식보다 성능이 높습니다. 이 방식을 사용하기 위해서는 `voting='hard'`를 `voting='soft'`로 바꾸고 모든 분류기가 클래스의 확률을 추정할 수 있으면 됩니다. `SVC`는 기본값에서는 클래스 확률을 제공하지 않으므로 `probability` 매개변수를 `True`로 지정해야 합니다. (이렇게하면 클래스 확률을 추정하기 위해 교차검증을 사용하므로 훈련 속도가 느려지지만 `SVC`에서 `predict_proba()`를 사용할 수 있습니다.) 앞의 코드를 간접 투표 방식을 사용하도록 변경하면 91%의 정확도를 얻을 것입니다.\n\n# ## 2. 배깅과 페이스팅\n#\n# 앞서 말했듯이 다양한 분류기를 만드는 한 가�� 방법은 각기 다른 훈련 알고리즘을 사용하는 것입니다. 또 다른 방법은 같은 알고리즘을 사용하지만 훈련 세트의 서브셋을 무작위로 구성하여 분류기를 각기 다르게 학습시키는 것입니다.\n#\n# 훈련 세트에서 중복을 허용하여 샘플링하는 방식을 **배깅**(bagging)(bootstrap aggregating의 줄임말)이라고 하며, 중복을 허용하지않고 샘플링하는 방식을 **페이스팅**(pasting)이라고 합니다.\n#\n# 다시 말해 배깅과 페이스팅에서는 같은 훈련 샘플을 여러 개의 예측기에 걸쳐 사용할 수 있습니다. 하지만 배깅만이 한 예측기를 위해 같은 훈련 샘플을 여러 번 샘플링 할 수 있습니다. 이 샘플링과 훈련 과정은 아래와 같습니다.\n#\n# 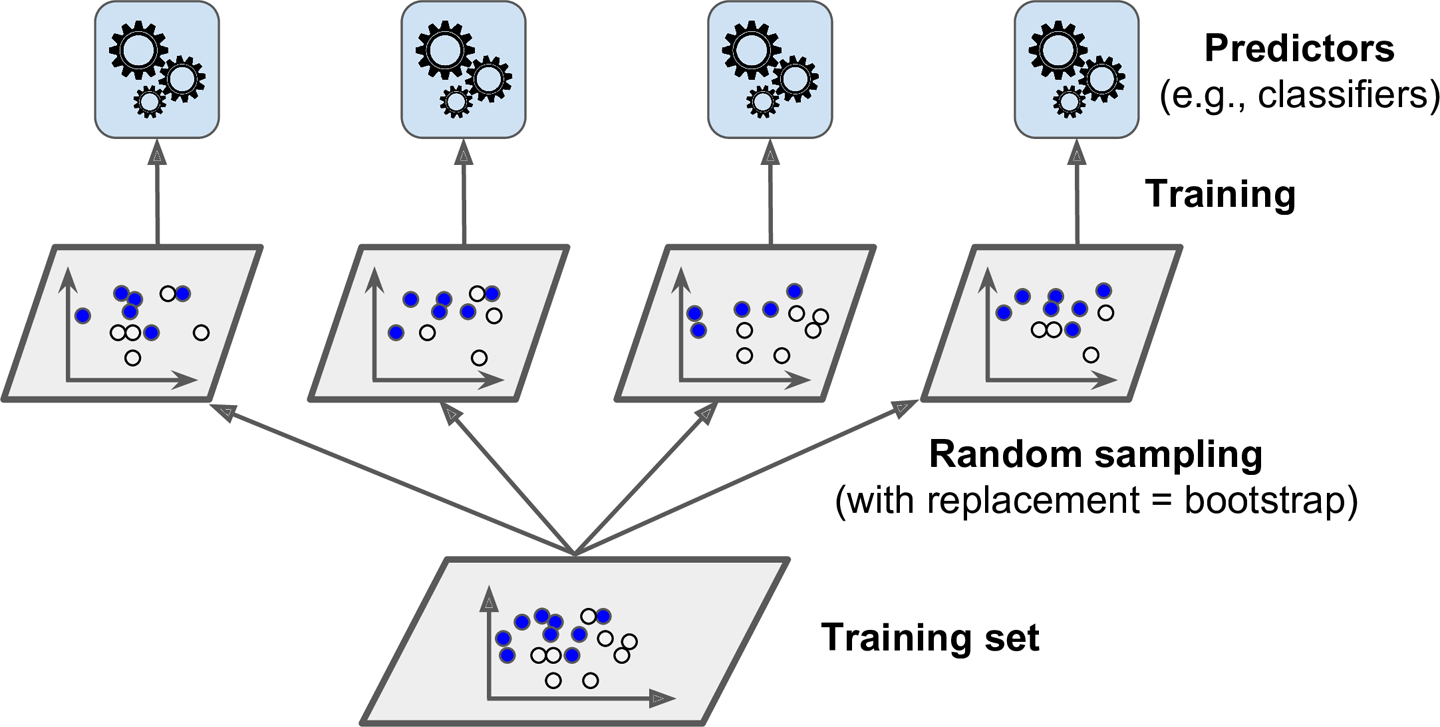\n#\n# 모든 예측기가 훈련을 마치면 앙상블은 모든 예측기의 예측을 모아 새로운 샘플에 대한 예측을 만듭니다. 수집 함수는 전형적으로 분류일 때는 **통계적 최빈값**(statistical mode)(즉, 직접 투표 분류기처럼 가장 많은 예측 결과)이고 회귀에 대해서는 평균을 계산합니다. 개별 예측기는 원본 훈련 세트로 훈련시킨 것보다 훨씬 크게 편향되어 있지만 수집 함수를 통과하면 편향과 분산이 모두 감소합니다. \n#\n# 위 그림에서 볼 수 있듯이 *예측기는 동시에 다른 CPU코어나 서버에서 병렬로 학습시킬 수 있습니다.* 이와 유사하게 예측도 병렬로 수행할 수 있습니다. 이런 확장성 덕분에 배깅과 페이스팅의 인기가 높습니다.\n#\n#\n\n# ### 2.1 사이킷런의 배깅과 페이스팅\n#\n# 사이킷런은 배깅과 페이스팅을 위해 간편한 API로 구성된 `BaggingClassifier`(회귀의 경우 `BagginRegressor`)를 제공합니다. 다음은 결정 트리 분류기 500개의 앙상블을 훈련시키는 코드입니다. 각 분류기는 훈련 세트에서 중복을 허용하여 무작위로 선택된 100개의 샘플로 훈련됩니다. 이 경우는 배깅의 경우이고, 페이스팅을 사용하려면 `bootstrap=False`로 지정하면 됩니다. `n_jobs` 매개변수는 사이킷런이 훈련과 예측에 사용할 CPU 코어 수를 지정합니다.\n\n# +\nfrom sklearn.ensemble import BaggingClassifier\nfrom sklearn.tree import DecisionTreeClassifier\n\nbag_clf = BaggingClassifier(\n DecisionTreeClassifier(), n_estimators=500,\n max_samples=100, bootstrap=True, n_jobs=-1\n)\nbag_clf.fit(X_train, y_train)\ny_pred = bag_clf.predict(X_test)\n# -\n\n# `BaggingClassifier`는 기반이 되는 분류기가 결정 트리 분류기처럼 클래스 확률을 추정할 수 있으면 (즉, `predict_proba()` 함수가 있으면) 직접 투표 대신 자동으로 간접 투표 방식을 사용합니다.\n#\n#\n# 아래 그림은 단일 결정 트리의 결정 경계외 500개의 트리를 사용한 배깅 앙상블의 경계를 비교한 것입니다. 둘다 moons 데이터 셋에 훈련시켰습니다. 여기서 볼 수 있듯이, 앙상블의 예측이 결정 트리 하나의 예측보다 일반화가 훨씬 잘 된 것 같습니다. 앙상블은 비슷한 편향에서 더 작은 분산을 만듭니다.\n\n# +\nimport matplotlib.pyplot as plt\nimport numpy as np\nfrom matplotlib.colors import ListedColormap\n\ntree_clf = DecisionTreeClassifier(random_state=42)\ntree_clf.fit(X_train, y_train)\ny_pred_tree = tree_clf.predict(X_test)\nprint(accuracy_score(y_test, y_pred_tree))\n\ndef plot_decision_boundary(clf, X, y, axes=[-1.5, 2.5, -1, 1.5], alpha=0.5, contour=True):\n x1s = np.linspace(axes[0], axes[1], 100)\n x2s = np.linspace(axes[2], axes[3], 100)\n x1, x2 = np.meshgrid(x1s, x2s)\n X_new = np.c_[x1.ravel(), x2.ravel()]\n y_pred = clf.predict(X_new).reshape(x1.shape)\n custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])\n plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=custom_cmap)\n if contour:\n custom_cmap2 = ListedColormap(['#7d7d58','#4c4c7f','#507d50'])\n plt.contour(x1, x2, y_pred, cmap=custom_cmap2, alpha=0.8)\n plt.plot(X[:, 0][y==0], X[:, 1][y==0], \"yo\", alpha=alpha)\n plt.plot(X[:, 0][y==1], X[:, 1][y==1], \"bs\", alpha=alpha)\n plt.axis(axes)\n plt.xlabel(r\"$x_1$\", fontsize=18)\n plt.ylabel(r\"$x_2$\", fontsize=18, rotation=0)\n\nplt.figure(figsize=(11,4))\nplt.subplot(121)\nplot_decision_boundary(tree_clf, X_train, y_train)\nplt.title(\"One Decision Tree\", fontsize=14)\nplt.subplot(122)\nplot_decision_boundary(bag_clf, X_train, y_train)\nplt.title(\"Decision Tree with Bagging\", fontsize=14)\nplt.show()\n# -\n\n# 부트스트래핑은 각 예측기가 학습하는 서브셋에 다양성을 증가시키므로 배깅이 페이스팅보다 편향이 조금 더 높습니다. 하지만 이는 예측기들의 상관관계를 줄이므로 앙상블의 분산을 감소시킵니다. 전반적으로 배깅이 더 나은 모델을 만들기 때문에 일반적으로 더 선호합니다. 그러��� 시간과 CPU 파워에 여유가 있다면 교차 검증으로 배깅과 페이스팅을 모두 평가해서 더 나은 쪽을 선택하는 것이 좋습니다.\n#\n#\n\n# ### 2.2 oob 평가\n#\n# 배깅을 사용하면 어떤 샘플은 한 예측기를 위해 여러 번 샘플링되고 어떤 것은 전혀 선택되지 않을 수 있습니다. `BaggingClassifier`는 기본 값으로 중복을 허용하여(`bootstrap=True`) 훈련 세트 크기만큼 $m$개 샘플을 선택합니다. 이는 평균적으로 각 예측기에 훈련 샘플의 63%만 정도만 샘플링 된다는 것을 의미합니다. 선택되지 않은 훈련 샘플의 나머지 37%를 oob(out-of-bag) 샘플이라고 부릅니다. 예측기마다 남겨진 37%는 모두 다릅니다.\n#\n# 예측기가 훈련되는 동안에는 oob 샘플을 사용하지 않으므로 검증 세트나 교차 검증을 사용하지않고 oob 샘플을 사용해 평가할 수 있습니다. 앙상블의 평가는 각 예측기의 oob 평가를 평균하여 얻습니다.\n#\n# 사이킷런에서 `BaggingClassifier`를 만들 때 `oob_score=True`로 지정하면 훈련이 끝난 후 자동으로 oob 평가를 수행합니다. 다음 코드는 이 과정을 보여줍니다. 평가 점수 결과는 `oob_score_` 변수에 저장되어 있습니다.\n\n# +\nbag_clf = BaggingClassifier(\n DecisionTreeClassifier(), n_estimators=500,\n bootstrap=True, n_jobs=1, oob_score=True\n)\n\nbag_clf.fit(X_train, y_train)\nbag_clf.oob_score_\n# -\n\n# oob 평가 결과를 보면 `BaggingClassifier`는 테스트 세트에서 약 89%의 정확도를 얻을 것으로 보입니다. 확인해봅시다.\n\n# +\nfrom sklearn.metrics import accuracy_score\n\ny_pred = bag_clf.predict(X_test)\naccuracy_score(y_test, y_pred)\n# -\n\n# 테스트 세트에서 90%의 정확도를 얻었습니다. 비슷하네요!\n#\n# oob 샘플에 대한 결정 함수의 값도 `oob_decision_function_` 변수에서 확인할 수 있습니다. 이 경우 결정 함수는 각 훈련의 샘플의 클래스 확률을 반환합니다. (기반이 되는 예측기가 `predict_proba()`를 가지고 있기 때문에) 다음 예를 보면 oob 평가는 첫 번째 훈련 샘플의 양성 클래스에 속할 확률을 59.51%로 추정하고 있습니다.\n#\n\nbag_clf.oob_decision_function_[:5]\n\n# ## 3. 랜덤 패치와 랜덤 서브스페이스\n#\n# `BaggingClassifier`는 특정 샘플링도 지원합니다. `max_features`, `bootstrap_features` 두 매개변수로 조절됩니다. 작동 방식은 `max_samples`, `bootstrap`과 동일하지만 샘플이 아니고 특성에 대한 샘플링입니다. 따라서 예측기는 무작위로 선택한 입력 특성의 일부분으로 훈련 됩니다.\n#\n# 특히 (이미지와 같은) 매우 고차원의 데이터셋을 다룰 때 유용합니다. 훈련 특성과 샘플은 모두 샘플링 하는 것을 **랜덤 패치 방식**(random patches method)라고 합니다. 훈련 샘플을 모두 사용하고, (즉 `bootstrap=False`이고 `max_samples=1.0`) 특성은 샘플링하는(즉, `bootstrap_features=True`이고 `max_features`는 1.0보다 작은) 것을 **랜덤 서브스페이스 방식**(Random subspaces method)이라고 합니다.\n#\n# 특성 샘플링은 더 다양한 예측기를 만들며 편향을 늘리는 대신 분산을 낮춥니다.\n\n# ## 4. 랜덤 포레스트\n#\n# 앞서 언급했듯이 랜덤 포레스트는 일반적으로 배깅 방법(또는 페이스팅)을 적용한 결정 트리의 앙상블입니다. 전형적으로 `max_samples`를 훈련 세트의 크기로 지정합니다. `BaggingClassifier`에 `DecisionTreeClassifier`를 넣어 만드는 대신 결정 트리에 최적화되어 사용하기 편리한 `RandomForestClassifier`를 사용할 수 있습니다. (회귀의 경우 `RandomForestRegressor`) 아래는 최대 16개 리프노드를 가지는 500개의 트리로 이루어진 랜덤 포레스트 분류기를 여러 CPU 코어에서 훈련시키는 코드입니다.\n\n# +\nfrom sklearn.ensemble import RandomForestClassifier\n\nrnd_clf = RandomForestClassifier(n_estimators=500, max_leaf_nodes=16, n_jobs=-1)\nrnd_clf.fit(X_train, y_train)\n\ny_pred_rf = rnd_clf.predict(X_test)\n# -\n\n# `RandomForestClassifier`는 몇가지 예외가 있지만 트리의 성장 조절을 위한 `DecisionTreeClassifier` 매개변수와 앙상블 자체를 제어하는 데 필요한 `BaggingClassifier`의 매개변수를 모두 가지고 있습니다.\n#\n#\n# 랜덤 포레스트 알고리즘은 트리의 노드를 분할할 때 전체 특성 중에서 최선의 특성을 찾는 대신 무작위로 선택한 특성 후보 중에서 최적의 특성을 찾는 식으로 무작위성을 더 주입합니다. 이는 결국 트리를 더욱 다양하게 만들고 편향을 손해보는 대신 분산을 낮추어 전체적으로 더 훌륭한 모델을 만들어 냅니다. 다음은 `BaggingClassifier`를 사용해 앞의 `RandomForestClassifier`와 거의 유사하게 만든 것입니다.\n\nbag_clf = BaggingClassifier(\n DecisionTreeClassifier(splitter='random', max_leaf_nodes=16),\n n_estimators=500, max_samples=1.0, bootstrap=True, n_jobs=-1\n)\n\n# ### 4.1 엑스트라 트리\n#\n# 랜덤 포레스트에서 트리를 만들 때 각 노드는 무작위로 특성의 서브셋을 만들어 분할에 사용합니다. 트리를 더욱 무작위하게 만들기 위해 최적의 임계값을 찾는 대신 후보 특성을 사용해 무작위로 분할한 다음 그 중에서 최상의 분할을 선택합니다.\n#\n# 이와같이 극단적으로 무작위한 트리의 랜덤 포레스트를 **익스트림 랜덤 트리**(Extremely Randomized Tree) 앙상블(또는 줄여서 *엑스트라 트리*(Extra Trees))라고 부릅니다. 여기서도 역시 편향이 늘어나지만 대신 분산을 낮추게 됩니다. 모든 노드에서 특성마다 가장 최적의 임계값을 찾는 것이 트리 알고리즘에서 가장 시간이 많이 소요되는 작업 중 하나이므로 일반적인 랜덤 포레스트보다 엑스트라 트리가 훨씬 빠릅니다.\n#\n# 엑스트라 트리를 만들려면 사이킷런의 `ExtraTreesClassifier`를 사용합니다. 사용법은 `RandomForestClassifier`와 같습니다. 마찬가지로 회귀 문제의 경우 `ExtraTreesRegressor`를 사용합니다. \n\n# ### 4.2 특성 중요도\n#\n# 랜덤 포레스트의 또 다른 장점은 특성의 상대적 중요도를 측정하기 쉽다는 것입니다. 사이킷런은 어떤 특성을 사용한 노드가 (랜덤 포레스트에 있는 모든 트리에 걸쳐서) 평균적으로 불순도를 얼마나 감소시키는지 확인하여 특성의 중요도를 측정합니다. 더 정확히 말하자면 가중치 평귱니며 각 노드의 가중치는 연관된 훈련 샘플 수와 같습니다.\n#\n# 사이킷런은 훈련이 끝난 뒤 특성마다 자동으로 이 점수를 계산하고 중요도의 전체 합이 1이 되도록 결괏값을 정규화합니다. 이 값은 `feature_importances_` 변수에 저장되어 있습니다. 예를 들어 다음 코드는 iris 데이터 셋에 `RandomForestClassifier`를 훈련시키고 각 특성의 중요도를 출력합니다. 가장 중요한 특성은 꽃잎의 길이(44%)와 너비(42%이고 꽃받침의 길이와 너비는 비교적 덜 중요해 보입니다.)\n\n# +\nfrom sklearn.datasets import load_iris\n\niris = load_iris()\nrnd_clf = RandomForestClassifier(n_estimators=500, n_jobs=-1)\nrnd_clf.fit(iris['data'], iris['target'])\n\nfor name, score in zip(iris['feature_names'], rnd_clf.feature_importances_):\n print(name, score)\n# -\n\n# 랜덤 포레스트는 특히 특성을 선택해야할 때 어떤 특성이 중요한지 빠르게 확인할 수 있어 매우 편리합니다.\n\n# ## 5. 부스팅\n#\n# **부스팅**(boosting)은 약한 학습기를 여러개 연결하여 강한 학습기를 만드는 앙상블 방법을 말합니다. 부스팅 방법의 아이디어는 앞의 모델을 보안해나가면서 일련의 예측기를 학습시키는 것입니다. 부스팅 방법은 여러가지가 있지만 가장 있기있는 것은 **아다부스트**(AdaBoost)(Adaptive Boosting의 줄임말)와 **그래디언트 부스팅**(Gradient Boosting)입니다. \n#\n# ### 5.1 아다부스트\n#\n# 이전 예측기를 보완하는 새로운 예측기를 만드는 방법은 이전 모델이 과소적합했던 훈련 샘플의 가중치를 더 높이는 것입니다. 이렇게 하면 훈련 샘플은 학습하기 어려운 샘플에 점점 더 맞춰지게 됩니다. 이것이 아다부스트에서 사용하는 방식입니다.\n#\n# 예를들어 아다부스트 분류기를 만드려면 기반이 되는 첫 번째 분류기 훈련 세트에서 훈련 시키고 예측을 만듭니다. 그 다음에 잘못 분류된 훈련 샘플의 가중치를 상대적으로 높입니다. 두 번째 분류기는 업데이트 된 가중치를 사용해 훈련 세트에서 훈련하고 다시 예측을 만듭니다. 그 다음에 다시 가중치를 업데이트 하는 식으로 계속됩니다. \n\n# +\nm = len(X_train)\n\nplt.figure(figsize=(11, 4))\nfor subplot, learning_rate in ((121, 1), (122, 0.5)):\n sample_weights = np.ones(m)\n plt.subplot(subplot)\n if subplot == 121:\n plt.text(-0.7, -0.65, \"1\", fontsize=14)\n plt.text(-0.6, -0.10, \"2\", fontsize=14)\n plt.text(-0.5, 0.10, \"3\", fontsize=14)\n plt.text(-0.4, 0.55, \"4\", fontsize=14)\n plt.text(-0.3, 0.90, \"5\", fontsize=14) \n for i in range(5):\n svm_clf = SVC(kernel=\"rbf\", C=0.05, gamma='auto', random_state=42)\n svm_clf.fit(X_train, y_train, sample_weight=sample_weights)\n y_pred = svm_clf.predict(X_train)\n sample_weights[y_pred != y_train] *= (1 + learning_rate)\n plot_decision_boundary(svm_clf, X, y, alpha=0.2)\n plt.title(\"learning_rate = {}\".format(learning_rate), fontsize=16)\n\nplt.show()\n# -\n\n# 위 그림은 moons 데이터셋에 훈련시킨 다섯개의 연속된 예측기의 결정 경계입니다. 첫 번째 분류기가 많은 샘플에서 잘못 분류해서 이 샘플들의 가중치가 높아져 두번째 분류기에서는 이 샘플들에 대해 좀 더 정확한 예측을 합니다. 오른쪽 그래프는 학습률을 절반으로 낮춘것을 빼면 똑같은 예측기를 나타냅니다. \n#\n# 그림에서 볼 수 있듯이 이런 연속된 학습 기법은 경사 하강법과 비슷한 면이 있습니다. 경사 하강법은 비용함수를 최소화하기위해 한 예측기의 모델 파라미터를 조정해가는 반면 아다부스트는 점차 좋아지도록 앙상블에 예측기를 추가합니다.\n#\n# 모든 예측기가 훈련을 마치면 이 앙상블은 배깅이나 페이스팅과 비슷한 방식으로 예측을 만듭니다. 하지만 가중치가 적용된 훈련 세트의 전반적인 정확도에 따라 예측기마다 다른 가중치가 적용됩니다.\n#\n#\n\n# +\nfrom sklearn.ensemble import AdaBoostClassifier\n\nada_clf = AdaBoostClassifier(\n DecisionTreeClassifier(max_depth=1), n_estimators=200,\n algorithm='SAMME.R', learning_rate=0.5, random_state=42\n)\nada_clf.fit(X_train, y_train)\nplot_decision_boundary(ada_clf, X, y)\n# -\n\n# 사이킷런은 SAMME라는 아다부스트의 다중 클래스 버전을 사용합니다. 클래스가 두개 뿐일때에는 SAMME가 아다부스트와 동일합니다. 예측기가 클래스의 확률을 추정할 수 있다면, (즉 `predict_proba()` 메서드가 있다면) 사이킷런은 SAMME.R (끝의 R은 'Reat'을 의미합니다.)라는 SAMME의 변종을 사용합니다. 이 알고리즘은 예측값 대신 클래스 확률에 기반하며 일반적으로 성능이 더 좋습니다.\n#\n# 위 코드는 `AdaBoostClassifier`를 사용하여 200개의 아주 얕은 결정트리를 기반으로 하는 아다부스트 분류기를 훈련시킵니다. 마찬가지로 회귀 문제의 경우 `AdaBoostRegressor`를 사용할 수 있습니다.\n\n# ### 5.2 그래디언트 부스팅\n#\n# 인기가 높은 또 하나의 부스팅 알고리즘은 **그래디언트 부스팅**(Gradient Boosting)입니다. 아다부스트처럼 그래디언트 부스팅은 앙상블에 이전까지의 오차를 보정하도록 예측기를 순차적으로 추가합니다. 하지만 아다부스트처럼 반복마다 샘플의 가중치를 수정하는 대신 이전 예측기가 만든 **잔여 오차**(residual error)에 새로운 예측기를 학습시킵니다.\n#\n# 결정 트리 기반 예측기로 사용하는 간단한 회귀 문제를 살펴봅시다. 이를 먼저 `DecisionTreeRegressor`를 훈련 세트에 학습시키고, 이 예측기에서 생긴 잔여 오차에 `DecisionTreeRegressor`를 훈련시킵니다. \n\nnp.random.seed(42)\nX = np.random.rand(100, 1) - 0.5\ny = 3*X[:, 0]**2 + 0.05 * np.random.randn(100)\n\n# +\nfrom sklearn.tree import DecisionTreeRegressor\n\ntree_reg1 = DecisionTreeRegressor(max_depth=2)\ntree_reg1.fit(X, y)\n# -\n\ny2 = y - tree_reg1.predict(X) # 잔여 오차로 학습시킨다!\ntree_reg2 = DecisionTreeRegressor(max_depth=2)\ntree_reg2.fit(X, y2)\n\ny3 = y - tree_reg2.predict(X)\ntree_reg3 = DecisionTreeRegressor(max_depth=2)\ntree_reg3.fit(X, y3)\n\n# 이제 세개의 트리를 포함하는 앙상블 모델이 생겼습니다. 새로운 샘플에 대한 예측을 만들려면 모든 트리의 예측을 더하면 됩니다.\n\nX_new = np.array([[0.8]])\n\ny_pred = sum(tree.predict(X_new) for tree in (tree_reg1, tree_reg2, tree_reg3))\ny_pred\n\n\n\n# +\ndef plot_predictions(regressors, X, y, axes, label=None, style=\"r-\", data_style=\"b.\", data_label=None):\n x1 = np.linspace(axes[0], axes[1], 500)\n y_pred = sum(regressor.predict(x1.reshape(-1, 1)) for regressor in regressors)\n plt.plot(X[:, 0], y, data_style, label=data_label)\n plt.plot(x1, y_pred, style, linewidth=2, label=label)\n if label or data_label:\n plt.legend(loc=\"upper center\", fontsize=16)\n plt.axis(axes)\n\nplt.figure(figsize=(11,11))\n\nplt.subplot(321)\nplot_predictions([tree_reg1], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label=\"$h_1(x_1)$\", style=\"g-\", data_label=\"훈련 세트\")\nplt.ylabel(\"$y$\", fontsize=16, rotation=0)\nplt.title(\"잔여 오차와 트리의 예측\", fontsize=16)\n\nplt.subplot(322)\nplot_predictions([tree_reg1], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label=\"$h(x_1) = h_1(x_1)$\", data_label=\"훈련 세트\")\nplt.ylabel(\"$y$\", fontsize=16, rotation=0)\nplt.title(\"앙상블의 예측\", fontsize=16)\n\nplt.subplot(323)\nplot_predictions([tree_reg2], X, y2, axes=[-0.5, 0.5, -0.5, 0.5], label=\"$h_2(x_1)$\", style=\"g-\", data_style=\"k+\", data_label=\"잔여 오차\")\nplt.ylabel(\"$y - h_1(x_1)$\", fontsize=16)\n\nplt.subplot(324)\nplot_predictions([tree_reg1, tree_reg2], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label=\"$h(x_1) = h_1(x_1) + h_2(x_1)$\")\nplt.ylabel(\"$y$\", fontsize=16, rotation=0)\n\nplt.subplot(325)\nplot_predictions([tree_reg3], X, y3, axes=[-0.5, 0.5, -0.5, 0.5], label=\"$h_3(x_1)$\", style=\"g-\", data_style=\"k+\")\nplt.ylabel(\"$y - h_1(x_1) - h_2(x_1)$\", fontsize=16)\nplt.xlabel(\"$x_1$\", fontsize=16)\n\nplt.subplot(326)\nplot_predictions([tree_reg1, tree_reg2, tree_reg3], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label=\"$h(x_1) = h_1(x_1) + h_2(x_1) + h_3(x_1)$\")\nplt.xlabel(\"$x_1$\", fontsize=16)\nplt.ylabel(\"$y$\", fontsize=16, rotation=0)\n\nplt.show()\n# -\n\n# 위 그림의 왼쪽열은 이 트리의 예측이고 오른쪽 열은 앙상블의 예측입니다. 첫 번째 행에서는 앙상블에 트리가 하나만 있어서 완전히 똑같습니다. 부전째 행부터는 새로운 트리가 첫번째 트리의 잔여 오차에 대해 학습되었습니다. 오른쪽 앙상블의 예측이 두개 트리의 예측의 합과 같은 것을 확인하 수 있습니다. 세번재 행에서는 두번째 트리의 잔여오차에 훈련되었습니다. 트리가 앙상블에 추가될수록 앙상블의 예측이 점차 좋아지는 것을 알 수 있습니다.\n#\n# 사이킷런의 `GradientBoostRegressor`를 사용하면 GBRT 앙상블을 간단하게 훈련시킬 수 있습니다. 트리 수(`n_estimators`)와 같이 앙상블의 훈련을 제어하는 매개변수는 물론 `RandomForestRegressor`와 아주 비슷하게 결정 트리의 성장을 제어하는 매개 변수 (`max_depth`, `min_samples_leaf` 등)를 가지고 있습니다. 다음은 이전에 만든 것과 같은 앙상블을 만드는 코드입니다.\n\n# +\nfrom sklearn.ensemble import GradientBoostingRegressor\n\ngbrt = GradientBoostingRegressor(max_depth=2, n_estimators=3, learning_rate=0.1, random_state=42)\ngbrt.fit(X, y)\n# -\n\n# `learning_rate` 매개변수가 각 트리의 기여 정도를 조절합니다. 이를 0.1처럼 낮게 설정하면 앙상블을 훈련 세트에 학습시키기 위해 많은 트리가 필요하지만 일반적으로 예측의 성능은 좋아집니다. 이는 **축소**(shrinkage)라고 부르는 규제 방법입니다. 아래 그림은 작은 학습률로 훈련시킨 두 개의 GBRT 앙상블을 보여줍니다.\n\ngbrt_slow = GradientBoostingRegressor(max_depth=2, n_estimators=200, learning_rate=0.1, random_state=42)\ngbrt_slow.fit(X, y)\n\n# +\nplt.figure(figsize=(11, 4))\n\nplt.subplot(121)\nplot_predictions([gbrt], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label='앙상블의 예측')\nplt.title(\"learning_rate = {}, n_estimators = {}\".format(gbrt.learning_rate, gbrt.n_estimators))\n\nplt.subplot(122)\nplot_predictions([gbrt_slow], X, y, axes=[-0.5, 0.5, -0.1, 0.8])\nplt.title(\"learning_rate = {}, n_estimators = {}\".format(gbrt_slow.learning_rate, gbrt_slow.n_estimators))\n\nplt.show()\n# -\n\n# 왼쪽은 훈련 세트를 학습시키기에 트리가 충분하지 않은 반면 오른쪽은 트리가 너무 많아 훈련 세트에 과대 적합되었습니다. 최적의 트리 수를 찾기 위해서는 *조기 종료*를 사용할 수 있습니다. 간단하게 구현하려면 `staged_predict()`메서드를 사용합니다. 이 메서드는 훈련의 각 단계(트리 하나, 트리 두개 등)에서 앙상블에 의해 만들어진 예측기를 순회하는 반복자(iterator)를 반환합니다. 다음 코드는 120개읱 트리로 GBRT 앙상블을 훈련시키고 최적의 트리 수를 찾기 위해 각 훈련 단계에서 검증 오차를 측정합니다. 마지막에 최적의 트리 수를 사용해 새로운 GBRT 앙상블을 훈련시킵니다.\n\n# +\nimport numpy as np\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import mean_squared_error\n\nX_train, X_val, y_train, y_val = train_test_split(X, y, random_state=49)\n\ngbrt = GradientBoostingRegressor(max_depth=2, n_estimators=120, random_state=42)\ngbrt.fit(X_train, y_train)\n\nerrors = [mean_squared_error(y_val, y_pred) for y_pred in gbrt.staged_predict(X_val)]\nbst_n_estimators = np.argmin(errors)\n\ngbrt_best = GradientBoostingRegressor(max_depth=2, n_estimators=bst_n_estimators)\ngbrt_best.fit(X_train, y_train)\n# -\n\nmin_error = np.min(errors)\n\n# +\nplt.figure(figsize=(11, 4))\n\nplt.subplot(121)\nplt.plot(errors, \"b.-\")\nplt.plot([bst_n_estimators, bst_n_estimators], [0, min_error], \"k--\")\nplt.plot([0, 120], [min_error, min_error], \"k--\")\nplt.plot(bst_n_estimators, min_error, \"ko\")\nplt.text(bst_n_estimators, min_error*1.2, \"최소\", ha='center')\nplt.axis([0, 120, 0, 0.01])\nplt.xlabel(\"트리 개수\")\nplt.title(\"검증 오차\")\n\nplt.subplot(122)\nplot_predictions([gbrt_best], X, y, axes=[-0.5, 0.5, -0.1, 0.8])\nplt.title(\"최적 모델 (트리 {}개)\".format(bst_n_estimators))\n\nplt.show()\n# -\n\n# 많은 수의 트리를 먼저 훈련시키고 최적의 수를 찾기 위해 살펴보는 대신 실제로 훈련을 중지하는 방법으로 조기종료를 구현할 수 있습니다. `warm_start=True`로 설정하면 사이킷런이 `fit()`메서드가 호출될 때 기존 트리를 유지하고 훈련을 추가할 수 있도록 해줍니다. 다음 코드는 연속해서 다섯 번의 반복동안 검증 오차가 향상되지 않으면 훈련을 멈춥니다.\n\n# +\ngbrt = GradientBoostingRegressor(max_depth=2, warm_start=True)\n\nmin_val_error = float('inf')\nerror_going_up = 0\n\nfor n_estimaotrs in range(1, 120):\n gbrt.n_estimaotrs = n_estimaotrs\n gbrt.fit(X_train, y_train)\n y_pred = gbrt.predict(X_val)\n val_error = mean_squared_error(y_val, y_pred)\n if val_error < min_val_error:\n min_val_error = val_error\n error_going_up = 0\n else:\n error_going_up += 1\n if error_going_up == 5:\n break # 조기 종료 \n# -\n\nprint(gbrt.n_estimaotrs)\nprint(\"최소 검증 MSE : \", min_val_error)\n\n# `GradientBoostingRegressor`는 각 트리가 훈련할 때 사용할 훈련 샘플의 비율을 지정할 수 있는 `subsample` 매개변수도 지원합니다. 예를 들어 `subsample=0.25`라고 하면 각 트리는 무작위로 선택된 25%의 훈련 샘플로 학습됩니다. 아마 추측할 수 있겠지만 편향이 높아지는 대신 분산이 낮아지게 됩니다. 또한 훈련 속도를 상당히 높입니다. 이런 기법을 **확률적 그래디언트 부스팅**(stochastic gradient boosting)이라고 합니다.\n\n# ## 6. 스태킹\n#\n# 이 장에서 이야기할 마지막 앙상블 모델은 **스태킹**(Stacking)(stacked generalization의 줄임말)입니다. 이는 '앙상블에 속한 모든 예측기의 예측을 취합하는 간단한 함수를 사용하는 대신 취합하는 모델 자체를 훈련시킬 수는 없을까?'라는 기본 아이디어에서 출발합니다. 아래 그림은 새로운 샘플에 회귀 작업을 수행하는 앙상블을 보여주고 있습니다. 아래의 세 예측기는 각각 다른 값을 예측하고 마지막 예측기(**블렌더**(blender)또는 **메타 학습기**(meta learner)라고 합니다)가 이 예측을 입력으로 받아 최종 예측을 만듭니다. \n#\n# 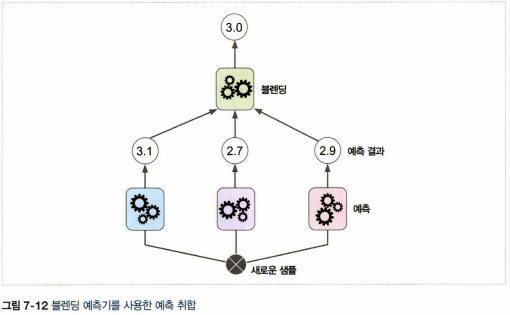\n\n# 블렌더를 학습시키는 일반적인 방법은 홀드 아웃 세트를 사용하는 것입니다. 어떻게 작동하는지 살펴봅시다. 먼저 훈련 세트를 두개의 서브셋으로 나눕니다. 첫 번째 서브셋은 첫 번째 레이어의 예측을 훈련시키기 위해 사용됩니다.\n#\n# 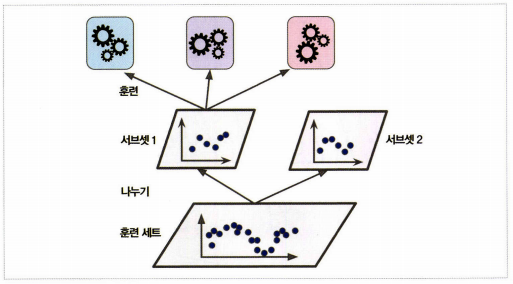\n#\n# 그런 다음 첫 번째 레이어의 예측기를 사용해 두 번째 세트에 대한 예측을 만듭니다. 예측기들이 훈련하는 동안 이 샘플들을 전혀 보지 못했기 때문에 이때 만들어진 예측은 완전히 새로운 것입니다. 이제 홀드 아웃 세트의 각 샘플에 대해 세 개의 예측값이 만들어집니다. 타깃 값은 그대로 쓰고 예측한 값을 입력 특성으로 사용하는 새로운 훈련 세트를 만들 수 있습니다. 블렌더가 새 훈련 세트로 훈련됩니다. 즉, 첫 번째 레이어의 예측을 가지고 타깃값을 예측하도록 학습됩니다. \n#\n# 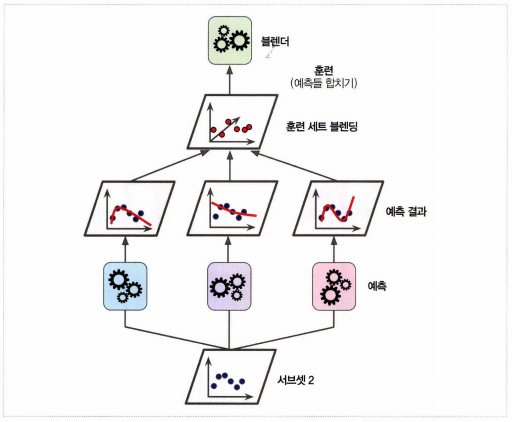\n#\n# 사실 이런 방식의 블렌더를 여러 개 훈련 시키는 것도 가능합니다. 그렇게 한다면 블렌더만의 레이어가 만들어지게 됩니다. 이렇게 하려면 훈련 세트를 세 개의 서브셋으로 나눕니다. 첫 번째 세트는 첫 번째 레이어를 훈련시키는 데 사용되고 두 번째 세트는 두 번째 레이어를 훈련시키기는데 사용되고 두번째 세트는 두 번째 레이어를 훈련시키기 위한 훈련 세트를 만드는 데 사용됩니다. 그리고 세 번재 세트는 세 번째 레이어를 훈련시키기 위한 훈련 세트를 만드는 데 사용됩니다. 작업이 끝나면 아래 그림처럼 각 레이어를 차례대로 실행해서 새로운 샘플에 대한 예측을 만들 수 있습니다. \n#\n# 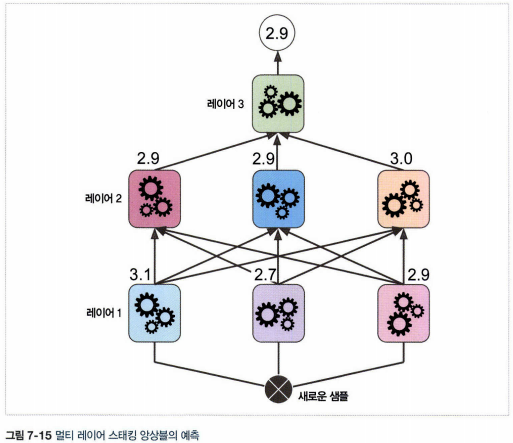\n\n# ---\n#\n# ## 요약 정리하기\n#\n# 1. 앙상블 : 여러 예측기를 훈련시켜 좋은 모델을 만드는 방법.\n# 2. 랜덤 포레스트 : 결정트리의 앙상블\n# 3. 투표 기반 분류기\n# - 직접 투표 : 다수결 투표로 정해지는 분류기\n# - 사이킷런의 `VotingClassifier`에서 매개변수 `voting='hard'`로 설정을 통해 학습 가능\n# - 간접 투표 : 개별 분류기의 예측을 평균내어 확률이 가장 높은 클래스를 예측하는 방식\n# - `VotingClassifier`에서 매개변수 `voting='soft'`로 바꾸면 가능\n# 4. 배깅\n# - 훈련 세트의 **중복을 허용**\n# - 사이킷런에 `BaggingClassifier` 제공하고 있다. 배깅의 경우 `bootstrap=True`(디폴트 값)으로 지정.\n# 5. 페이스팅\n# - 훈련 세트의 **중복ㅇ을 허용하지 않음**\n# - 배깅과 마찬가지로 `BaggingClassifier`를 사용하는데 `bootstrap=False`로 설정하면 된다.\n# 6. oob\n# - 중복을 허용해 샘플링을 하면 평균적으로 각 예측기에 훈련 샘플의 63%정도만 샘플링된다. 선택되지 않은 훈련 샘플의 나머지 37%를 oob(out-of-bag) 샘플이라고 부른다.\n# - 사이킷런에서 `BaggingClassifier`를 만들 때 `oob_score=True`로 지정하면 훈련이 끝난 후 자동으로 oob 평가를 수행한다. 평가 점수 결과는 `oob_score_`에 저장되어 있다. \n# 7. 랜덤 패치\n# - 훈련 특성과 샘플을 모두 샘플링 하는 것.\n# 8. 랜덤 서브스페이스\n# - 훈련 샘플을 모두 사용하고 특성은 샘플링 하는 것. \n# 9. 랜덤 포레스트 \n# - 배깅 방법을 적용한 결정트리의 앙상블\n# - `max_samples`를 통해 훈련 세트의 크기를 지정\n# - 사이킷런에서 `RandomForestClassifier` 제공 \n# 10. 엑스트라 트리\n# - 랜덤 포레스트에서 각 노드는 무작위로 특성의 서브셋을 만들어 분할에 사용하는데, 트리를 더욱 무작위로 만들기 위해 최적의 임계값을 찾는 대신 후보 특성을 사용해 무작위로 분할한 트리의 랜덤 포레스트를 익스트림 랜덤 트리 앙상블, 줄여서 엑스트라 트리라고 부른다.\n# - 사이킷런에서 `ExtraTreesClassifier`를 제공한다\n# 11. 특성 중요도\n# - 특성의 상대적 중요도.\n# - 랜덤포레스트에서는 특성의 사앧적 중요도를 측정할 수 있다. 이 값은 `feature_importances_` 변수에 저장되어 있다.\n# 12. 부스팅\n# - 약한 학습기를 여러개 연결하여 강한 학습기를 만드는 방식. \n# 1. 아다부스트 \n# - 과소적합했던 이전 샘플의 가중치를 높이는 방식\n# - 사이킷런에서 `AdaBoostClassifier`를 제공\n# 2. 그래디언트 부스팅\n# - 이전 예측기가 만든 잔여 오차에 대해 새로운 학습기를 학습시키는 방식. \n# - 결정트리에서 그래디언트 부스팅을 사용한 방식을 그래디언트 부스티드 회귀 트리(GBRT)라고 하는데, 사이킷런의 `GradientBoostingRegressor`통해 구현할 수 있다. \n# 13. 스태킹\n# - 앙상블 취합하는 모델을 훈련시키는 방식.\n# - 사이킷런에서 직접 지원하지는 않고 있다. \n\n\n","repo_name":"jiyeoon/TIL","sub_path":"Hands_On_Machine_Learning/ch7_앙상블학습과랜덤포레스트.ipynb","file_name":"ch7_앙상블학습과랜덤포레스트.ipynb","file_ext":"py","file_size_in_byte":36824,"program_lang":"python","lang":"en","doc_type":"code","stars":8,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"43607327670","text":"# + [markdown] id=\"view-in-github\" colab_type=\"text\"\n#  \n\n# + [markdown] id=\"RzWFEMTRTYD0\"\n# #Imports and Setup\n\n# + id=\"_cajUFlyllOs\"\nimport warnings\nwarnings.filterwarnings('ignore')\n\nimport pandas as pd\nimport numpy as np\nfrom plotnine import *\nimport seaborn as sns\nimport scipy.cluster.hierarchy as shc\n\nfrom sklearn.preprocessing import StandardScaler\n\nfrom sklearn.neighbors import NearestNeighbors\nfrom sklearn.cluster import DBSCAN\nfrom sklearn.cluster import KMeans\nfrom sklearn.cluster import AgglomerativeClustering\nfrom sklearn.mixture import GaussianMixture\n\nfrom sklearn.metrics import silhouette_score\n\nimport pandas as pd\nfrom random import randint\nfrom sklearn import datasets\nimport matplotlib.pyplot as plt\nfrom scipy.spatial import distance\nimport numpy as np\nimport pylab\n\n\n\n# %matplotlib inline\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 206} id=\"C3Q3Vh0GllO6\" outputId=\"72bff194-89b5-450e-e990-d0a52dbfb8e8\"\ndf = pd.read_csv(\"https://raw.githubusercontent.com/cmparlettpelleriti/CPSC392ParlettPelleriti/master/Data/KMEM1.csv\")\n\ndf.head()\n\n\n\n# + [markdown] id=\"JB7hKwymkzT3\"\n# # Prepare data\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 296} id=\"X-WrjCGLfMuc\" outputId=\"469fec00-d835-4ae3-e910-52570ecb6c77\"\nsns.scatterplot(x='x',y='y',data=df)\n\n# + id=\"a0Uhf9EB2-Fj\"\ndf_scaled=StandardScaler().fit_transform(df)\n\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"jS0UDpsN-9Gw\" outputId=\"14757898-4643-4330-ea1a-4e3d7f2281fb\"\ndf_scaled[:,0].shape\n\n# + [markdown] id=\"nH9lDbE8k5DE\"\n# # Elbow method\n\n# + [markdown] id=\"DX92Xrg2lDc6\"\n# - implement elbow method to choose best K value for kmeans clustering\n\n# + id=\"FlU9rud7b5KI\"\ndistortions = []\nK = range(1,10)\nfor k in K:\n kmeanModel = KMeans(n_clusters=k)\n kmeanModel.fit(df)\n distortions.append(kmeanModel.inertia_)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 295} id=\"pII01zCEb5BC\" outputId=\"77e71013-9f32-41ce-c530-ce07e26ab1d6\"\nplt.plot(K, distortions, marker = 'o')\nplt.xlabel('k')\nplt.ylabel('Distortion')\nplt.title('The Elbow Method showing the optimal k')\nplt.show()\n\n# + [markdown] id=\"R8rjzsEflUOJ\"\n# # Dendogram \n\n# + [markdown] id=\"jODcnSidlZaL\"\n# - plot dendrogram visualize the result of a hierarchical clustering if performed on data.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 444} id=\"_mNltcQ9yzrh\" outputId=\"dc842235-ff2b-44f8-ab73-05eea333f522\"\nplt.figure(figsize=(10, 7))\nplt.title(\"Counters Dendograms\")\ndend = shc.dendrogram(shc.linkage(y=df_scaled , method='single',metric='euclidean'))\n\n# + [markdown] id=\"Rbmf8ri5l7t8\"\n# # Clustering\n\n# + [markdown] id=\"1fsF1XjQl0M4\"\n# ## KMeans clustering\n\n# + [markdown] id=\"3b_CnsPDqHdn\"\n# - perform Kmeans clustering\n\n# + id=\"fnL-DNHbmDP0\"\nkm = KMeans(3)\nkm.fit(df_scaled)\ndf[\"km\"] = km.labels_\n\n# + [markdown] id=\"gg8LWKqOqKKe\"\n# - visualize output clusters\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} id=\"u_ru6htvmkuu\" outputId=\"6ae38d9c-b0f5-40f3-d6d9-3197416ab3c0\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(km)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"KMeans\"))\n\n\n# + [markdown] id=\"8ePSYVETqOqw\"\n# - evaluate cluster performance using silhouette score.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"gF8PETyYqGjd\" outputId=\"7594aa61-aecf-45f2-d6f5-7f7b1e381e44\"\npreds = km.predict(df_scaled)\nscore = silhouette_score(df, preds)\nprint(\"silhouette score using kmeans is {}\".format(score))\n\n# + [markdown] id=\"9-E42y5kmH2P\"\n# ## Agglomerative clustering\n\n# + [markdown] id=\"tgCfui1mqck2\"\n# - perform agglomerative clustering.\n\n# + id=\"3KDclawemMWS\"\nag = AgglomerativeClustering(3)\nag.fit(df_scaled)\ndf[\"hac\"] = ag.labels_\n\n# + [markdown] id=\"3yRJ-LkVqiwB\"\n# - visualize output clusters.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} id=\"ma0AwvaXmorz\" outputId=\"95e48459-f32c-4858-a802-b589070c1c74\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(hac)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"Hierarchical Agglomeretive\"))\n\n# + [markdown] id=\"6cf4mgC0qmGO\"\n# - evaluate cluster performance using silhouette score.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"bVhAofOgq6ci\" outputId=\"8070315e-df03-43ee-b7b9-dd91d9af71e2\"\nscore = silhouette_score(df, ag.labels_)\nprint(\"silhouette score using AgglomerativeClustering is {}\".format(score))\n\n# + [markdown] id=\"5ghPqRj_mSQw\"\n# ## Gaussian Mixture clustering\n\n# + [markdown] id=\"CVBpUUq9ruZU\"\n# - perform Gaussian Mixture clustering.\n\n# + id=\"Hhvs-XRpmYZh\"\ngm = GaussianMixture(3)\ngm.fit(df_scaled)\ndf[\"gm\"] = gm.predict(df[[\"x\",\"y\"]])\n\n# + [markdown] id=\"2lxdQyQGrHJJ\"\n# - visualize output clusters.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} id=\"HVUPOXLpmm1b\" outputId=\"4826407b-1917-4655-f9f4-c0214ccdee13\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(gm)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"Gaussian Mixtures\"))\n\n# + [markdown] id=\"AuUsIFZVqv1D\"\n# - evaluate cluster performance using silhouette score.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"WAaNLi1hrPVj\" outputId=\"ff925df2-9de3-4a83-d125-50f04804ac9d\"\npreds = gm.predict(df[[\"x\",\"y\"]])\nscore = silhouette_score(df, preds)\nprint(\"silhouette score using GaussianMixture is {}\".format(score))\n\n# + [markdown] id=\"eQh9GE5qmc84\"\n# ## DBSAN clustering\n\n# + [markdown] id=\"2csU_aLer0DN\"\n# - perform DBSCAN clustering.\n\n# + id=\"97FjNxdahWOD\"\ndbscan = DBSCAN(eps=0.5, min_samples=10)\ndbscan.fit(df_scaled)\ndf[\"db\"] = dbscan.labels_\n\n# + [markdown] id=\"GJdIuOwRrIDk\"\n# - visualize output clusters.\n\n# + id=\"mi33GjH8jEht\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} outputId=\"7446945c-cb90-4487-b2e7-8b687febd967\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(db)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"DBSCAN\"))\n\n\n# + [markdown] id=\"rYKqybQIqw2a\"\n# - evaluate cluster performance using silhouette score.\n\n# + id=\"RExwvq_YjJJx\" colab={\"base_uri\": \"https://localhost:8080/\"} outputId=\"6a49f8f9-6b12-4639-974e-3b6444260245\"\nscore = silhouette_score(df_scaled, dbscan.labels_)\nprint(\"silhouette score using dbscan is {}\".format(score))\n\n# + [markdown] id=\"tEtKxBsbr5RV\"\n# # Conclusion\n\n# + [markdown] id=\"iBR0zUdUr94N\"\n# - kmeans then Agglomerative clustering was the best clustering algorithms to perform having silhouette score of approximatly 6 and 0.57 and that was due to the behavior and distribution while the other clustering algorithms may perform better on other data.\n","repo_name":"ayaamr11/Clustering-Algorithms","sub_path":"ClusteringAlgorithms.ipynb","file_name":"ClusteringAlgorithms.ipynb","file_ext":"py","file_size_in_byte":7454,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"39593347418","text":"import numpy as np\nimport pandas as pd\nimport pyflux as pf\nimport datetime as datetime\nimport matplotlib.pyplot as plt\nimport os\nimport pickle\nimport ForecastModel as fc\n\n# +\nX_fwds = pickle.load(open('../data/forward_rates', 'rb'))\n#X_fwds = df_add_first_diff(X_fwds)\n\n#df_FX = pickle.load( open( \"data/FX_data\", \"rb\" ) )\n# Loading up the federal reserve speech data\nfed_metrics = pickle.load( open( \"../data/final_press_rel_dist\", \"rb\" ) )\n# -\n\n\n\n# +\ncos_last = fed_metrics['cos_last']\ncos_avg_n = fed_metrics['cos_avg_n']\ned_last = fed_metrics['ed_last']\ned_avg_n = fed_metrics['ed_avg_n']\nfed_dates = fed_metrics['dates']\n\n#grouping by date (some dates had multiple speeches)\navgstats = pd.DataFrame({'date':fed_dates,\n 'ed_last': ed_last,\n 'ed_avg_n': ed_avg_n,\n 'cos_last': cos_last,\n 'cos_avg_n': cos_avg_n}).groupby('date').mean()\navgstats.index = pd.to_datetime(avgstats.index)\n# -\n\nthis_min = min(avgstats.index)\nthis_min\n\nX_fwds.info()\n\n# now I need to filter the X_fwds to drop anything before this period!\nX_after = X_fwds[X_fwds.index >= this_min]\n\n# +\nX_fwds = X_fwds[X_fwds.index >= this_min]\nX_fwds = X_fwds.merge(avgstats, how='left', left_index = True, right_index = True)\nX_fwds.fillna(value=0, inplace=True)\n\n# the first row of X_fwds contains zeros for the differenced rates, clear them here\nX_fwds = X_fwds.drop(X_fwds.index[0])\n\n# -\n\nX_fwds.describe()\n\ntemp = X_fwds['ed_last'].copy()\n\ntemp =pd.concat([temp, temp.shift(), temp.shift(2)], axis=1)\n\ntemp.head()\n\nX_fwds['ed_last_l1']= temp.iloc[:,1]\nX_fwds['ed_last_l2']= temp.iloc[:,2]\nX_fwds.head()\n\n\n\n\ntemp2\n\nlen(temp2)\n\nlen(temp)\n\n\n","repo_name":"davidjsmith44/NLP-Fed-Speeches","sub_path":"src/Untitled5.ipynb","file_name":"Untitled5.ipynb","file_ext":"py","file_size_in_byte":1700,"program_lang":"python","lang":"en","doc_type":"code","stars":10,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"33880389887","text":"# # Capstone Project\n# ## Image classifier for the SVHN dataset\n# ### Instructions\n#\n# In this notebook, you will create a neural network that classifies real-world images digits. You will use concepts from throughout this course in building, training, testing, validating and saving your Tensorflow classifier model.\n#\n# This project is peer-assessed. Within this notebook you will find instructions in each section for how to complete the project. Pay close attention to the instructions as the peer review will be carried out according to a grading rubric that checks key parts of the project instructions. Feel free to add extra cells into the notebook as required.\n#\n# ### How to submit\n#\n# When you have completed the Capstone project notebook, you will submit a pdf of the notebook for peer review. First ensure that the notebook has been fully executed from beginning to end, and all of the cell outputs are visible. This is important, as the grading rubric depends on the reviewer being able to view the outputs of your notebook. Save the notebook as a pdf (File -> Download as -> PDF via LaTeX). You should then submit this pdf for review.\n#\n# ### Let's get started!\n#\n# We'll start by running some imports, and loading the dataset. For this project you are free to make further imports throughout the notebook as you wish. \n\nimport tensorflow as tf\nfrom scipy.io import loadmat\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport random\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPooling2D, BatchNormalization\nfrom tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping\n\n# \n# For the capstone project, you will use the [SVHN dataset](http://ufldl.stanford.edu/housenumbers/). This is an image dataset of over 600,000 digit images in all, and is a harder dataset than MNIST as the numbers appear in the context of natural scene images. SVHN is obtained from house numbers in Google Street View images. \n#\n# * Y. Netzer, T. Wang, A. Coates, A. Bissacco, B. Wu and A. Y. Ng. \"Reading Digits in Natural Images with Unsupervised Feature Learning\". NIPS Workshop on Deep Learning and Unsupervised Feature Learning, 2011.\n#\n# Your goal is to develop an end-to-end workflow for building, training, validating, evaluating and saving a neural network that classifies a real-world image into one of ten classes.\n\n# +\n# Run this cell to load the dataset\n\ntrain = loadmat('data/train_32x32.mat')\ntest = loadmat('data/test_32x32.mat')\n\n\n# -\n\n# Both `train` and `test` are dictionaries with keys `X` and `y` for the input images and labels respectively.\n\n# ## 1. Inspect and preprocess the dataset\n# * Extract the training and testing images and labels separately from the train and test dictionaries loaded for you.\n# * Select a random sample of images and corresponding labels from the dataset (at least 10), and display them in a figure.\n# * Convert the training and test images to grayscale by taking the average across all colour channels for each pixel. _Hint: retain the channel dimension, which will now have size 1._\n# * Select a random sample of the grayscale images and corresponding labels from the dataset (at least 10), and display them in a figure.\n\ndef get_train_test_data(train_data, test_data):\n \"\"\"\n This function preprocesses and returns train and test data from received dictionary\n Data is transposed to make batch size dimension first and normalized\n \"\"\"\n x_train, y_train = train_data['X'], train_data['y']\n x_test, y_test = test_data['X'], test_data['y']\n # Set labels between 0 and 9\n y_train -= 1\n y_test -= 1\n # transpose to batch size first\n x_train = np.transpose(x_train, (3, 0, 1, 2))\n x_test = np.transpose(x_test, (3, 0, 1, 2))\n # min - max: 0 - 255\n x_train = x_train / 255.0\n x_test = x_test / 255.0\n return x_train, y_train, x_test, y_test\n\n\nx_train, y_train, x_test, y_test = get_train_test_data(train, test)\nprint(f'x_train shape is: {x_train.shape}')\nprint(f'y_train shape is: {y_train.shape}')\nprint(f'x_test shape is: {x_test.shape}')\nprint(f'y_test shape is: {y_test.shape}')\nprint(f'x_train data: min value is {np.min(x_train)} and max value is {np.max(x_train)}')\nprint(f'x_test data: min value is {np.min(x_test)} and max value is {np.max(x_test)}')\nprint(f'y_train data: min value is {np.min(y_train)} and max value is {np.max(y_train)}')\nprint(f'y_test data: min value is {np.min(y_test)} and max value is {np.max(y_test)}')\n\n# +\nnum_imgs = 10\nfig, ax = plt.subplots(1, num_imgs, figsize=(num_imgs, 1))\nrandom_sample = random.sample(range(len(x_train)), num_imgs)\n\nfor idx, sample in enumerate(random_sample):\n ax[idx].set_axis_off()\n ax[idx].imshow(x_train[sample])\n ax[idx].set_title(y_train[sample][0] + 1)\n# -\n\nx_train = np.mean(x_train, axis = 3)[..., np.newaxis]\nx_test = np.mean(x_test, axis = 3)[..., np.newaxis]\n\nprint(f'x_train shape is: {x_train.shape}')\nprint(f'x_test shape is: {x_test.shape}')\n\nfig, ax = plt.subplots(1, num_imgs, figsize=(num_imgs, 1))\nfor idx, sample in enumerate(random_sample):\n ax[idx].set_axis_off()\n ax[idx].imshow(np.squeeze(x_train[sample]), cmap='gray')\n ax[idx].set_title(y_train[sample][0] + 1)\n\n\n# ## 2. MLP neural network classifier\n# * Build an MLP classifier model using the Sequential API. Your model should use only Flatten and Dense layers, with the final layer having a 10-way softmax output. \n# * You should design and build the model yourself. Feel free to experiment with different MLP architectures. _Hint: to achieve a reasonable accuracy you won't need to use more than 4 or 5 layers._\n# * Print out the model summary (using the summary() method)\n# * Compile and train the model (we recommend a maximum of 30 epochs), making use of both training and validation sets during the training run. \n# * Your model should track at least one appropriate metric, and use at least two callbacks during training, one of which should be a ModelCheckpoint callback.\n# * As a guide, you should aim to achieve a final categorical cross entropy training loss of less than 1.0 (the validation loss might be higher).\n# * Plot the learning curves for loss vs epoch and accuracy vs epoch for both training and validation sets.\n# * Compute and display the loss and accuracy of the trained model on the test set.\n\ndef get_mlp_model(input_shape):\n \"\"\"\n Returns compiled MLP model\n \"\"\"\n model = Sequential([\n Flatten(input_shape = input_shape),\n #Dense(64, activation = 'relu'),\n Dense(256, activation = 'relu'),\n Dense(256, activation = 'relu'),\n Dense(128, activation = 'relu'),\n #BatchNormalization(),\n #Dense(32, activation = 'relu'),\n #Dropout(0.3),\n Dense(10, activation = 'softmax')\n ])\n model.compile(optimizer = 'adam', loss = 'sparse_categorical_crossentropy', metrics = ['accuracy'])\n return model\n\n\nmodel = get_mlp_model(x_train[0].shape)\nmodel.summary()\n\n\ndef get_checkpoints(checkpoint_path):\n \"\"\"\n Returns a list with early stopping and best checkpoint \n \"\"\"\n best_model = ModelCheckpoint(checkpoint_path, \n monitor = 'val_accuracy', save_weights_only = True,\n save_best_only = True)\n early_stopping = EarlyStopping(monitor = 'val_accuracy', patience = 4)\n return [best_model, early_stopping]\n\n\nhistory = model.fit(x_train, y_train, epochs = 30, batch_size = 64, validation_split = 0.15, callbacks = get_checkpoints('mlp/checkpoints_best_only/checkpoint'))\n\nplt.plot(history.history['accuracy'])\nplt.plot(history.history['val_accuracy'])\nplt.title('Accuracy vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='lower right')\nplt.show()\n\nplt.plot(history.history['loss'])\nplt.plot(history.history['val_loss'])\nplt.title('Loss vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='upper right')\nplt.show()\n\nmodel = get_mlp_model(x_train[0].shape)\nmodel.load_weights('mlp/checkpoints_best_only/checkpoint')\ntest_loss, test_acc = model.evaluate(x = x_test, y = y_test, verbose = 0)\nprint('Loss in test set is {:.02f} and Accuracy in test set is {:.02f}'.format(test_loss, test_acc))\n\n\n# ## 3. CNN neural network classifier\n# * Build a CNN classifier model using the Sequential API. Your model should use the Conv2D, MaxPool2D, BatchNormalization, Flatten, Dense and Dropout layers. The final layer should again have a 10-way softmax output. \n# * You should design and build the model yourself. Feel free to experiment with different CNN architectures. _Hint: to achieve a reasonable accuracy you won't need to use more than 2 or 3 convolutional layers and 2 fully connected layers.)_\n# * The CNN model should use fewer trainable parameters than your MLP model.\n# * Compile and train the model (we recommend a maximum of 30 epochs), making use of both training and validation sets during the training run.\n# * Your model should track at least one appropriate metric, and use at least two callbacks during training, one of which should be a ModelCheckpoint callback.\n# * You should aim to beat the MLP model performance with fewer parameters!\n# * Plot the learning curves for loss vs epoch and accuracy vs epoch for both training and validation sets.\n# * Compute and display the loss and accuracy of the trained model on the test set.\n\ndef get_cnn_model(input_shape):\n \"\"\"\n Returns compiled CNN model\n \"\"\"\n model = Sequential([\n Conv2D(16, (3, 3), activation = 'relu', input_shape = input_shape),\n MaxPooling2D((2, 2)),\n Conv2D(32, (3, 3), activation = 'relu'),\n MaxPooling2D((2, 2)),\n BatchNormalization(),\n Flatten(),\n Dense(32, activation = 'relu'),\n Dropout(0.3),\n Dense(10, activation = 'softmax')\n ])\n model.compile(optimizer = 'adam', loss = 'sparse_categorical_crossentropy', metrics = ['accuracy'])\n return model\n\n\nmodel = get_cnn_model(x_train[0].shape)\nmodel.summary()\n\nhistory_cnn = model.fit(x_train, y_train, epochs = 30, batch_size = 64, validation_split = 0.15, callbacks = get_checkpoints('cnn/checkpoints_best_only/checkpoint'))\n\nplt.plot(history_cnn.history['accuracy'])\nplt.plot(history_cnn.history['val_accuracy'])\nplt.title('Accuracy vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='lower right')\nplt.show()\n\nplt.plot(history_cnn.history['loss'])\nplt.plot(history_cnn.history['val_loss'])\nplt.title('Loss vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='upper right')\nplt.show()\n\nmodel = get_cnn_model(x_train[0].shape)\nmodel.load_weights('cnn/checkpoints_best_only/checkpoint')\ntest_loss, test_acc = model.evaluate(x = x_test, y = y_test, verbose = 0)\nprint('Loss in test set is {:.02f} and Accuracy in test set is {:.02f}'.format(test_loss, test_acc))\n\n# ## 4. Get model predictions\n# * Load the best weights for the MLP and CNN models that you saved during the training run.\n# * Randomly select 5 images and corresponding labels from the test set and display the images with their labels.\n# * Alongside the image and label, show each model’s predictive distribution as a bar chart, and the final model prediction given by the label with maximum probability.\n\n# ### 4.1. MLP model\n\nrandom_idx = np.random.choice(len(x_test), 5)\nrandom_test_images = x_test[random_idx]\nrandom_test_labels = y_test[random_idx]\n\nmlp_model = get_mlp_model(x_train[0].shape)\nmlp_model.load_weights('mlp/checkpoints_best_only/checkpoint')\n\n# +\npredictions = mlp_model.predict(random_test_images)\n\nfig, axes = plt.subplots(5, 2, figsize = (16, 12))\nfig.subplots_adjust(hspace = 0.4, wspace = -0.2)\n\nfor i, (prediction, image, label) in enumerate(zip(predictions, random_test_images, random_test_labels)):\n axes[i, 0].set_axis_off()\n axes[i, 0].imshow(np.squeeze(image), cmap = 'gray')\n axes[i, 0].text(10., -1.5, f'Label: {label[0] + 1}')\n axes[i, 1].bar(np.arange(1, len(prediction) + 1), prediction)\n axes[i, 1].set_xticks(np.arange(len(prediction) + 1))\n axes[i, 1].set_title(f\"Model prediction: {np.argmax(prediction) + 1}\")\n \nplt.show()\n# -\n\n# ### 4.2. CNN Model\n\ncnn_model = get_cnn_model(x_train[0].shape)\ncnn_model.load_weights('cnn/checkpoints_best_only/checkpoint')\n\n# +\npredictions = cnn_model.predict(random_test_images)\n\nfig, axes = plt.subplots(5, 2, figsize = (16, 12))\nfig.subplots_adjust(hspace = 0.4, wspace = -0.2)\n\nfor i, (prediction, image, label) in enumerate(zip(predictions, random_test_images, random_test_labels)):\n axes[i, 0].set_axis_off()\n axes[i, 0].imshow(np.squeeze(image), cmap = 'gray')\n axes[i, 0].text(10., -1.5, f'Label: {label[0] + 1}')\n axes[i, 1].bar(np.arange(1, len(prediction) + 1), prediction)\n axes[i, 1].set_xticks(np.arange(len(prediction) + 1))\n axes[i, 1].set_title(f\"Model prediction: {np.argmax(prediction) + 1}\")\n \nplt.show()\n# -\n\n\n","repo_name":"jcgarciaca/getting_started_with_tensorflow2","sub_path":"Capstone Project.ipynb","file_name":"Capstone Project.ipynb","file_ext":"py","file_size_in_byte":13047,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"23535976144","text":"# +\nimport keras\nimport tensorflow as tf\nimport keras.layers as layers\nimport keras.activations as activations\nimport numpy as np\nimport tensorflow_probability as tfp\nimport pylab as pl\nimport matplotlib.pyplot as plt\nimport scipy as sp\nimport scipy.signal \nfrom scipy.io import wavfile\n\nimport sounddevice as sd\n# -\n\n# # 1. Mise en place des données tests\n\n# +\nrate, sls = wavfile.read('Sls.wav')\nsls = sls/np.max(sls)\n\nN = len(sls)\nprint(N/rate)\n\ntime = np.arange(N)/rate\n\nplt.plot(sls)\nplt.title('Signal')\nplt.show()\n\n#sd.play(sls/np.max(sls),rate)\n\n# +\nNs = N\nsigma_carre = 1.2\n\n#compute gaussian signal\nSamples = np.power(1.2,0.5)*np.random.randn(Ns)\nabs = np.arange(Ns)\npl.plot(abs,Samples)\npl.show()\n\n# +\nslsn = sls + Samples/90\n\npl.plot(slsn)\n\npl.show()\n\n#sd.play(slsn[3200000:3200000+3200000]/np.max(sls),rate)\n\n# +\ndef make_extrait(x,fs):\n data = np.zeros((1,fs*2))\n i=0\n data = [x[i:i+fs*2]]\n i+=fs\n while(i+fs*2
\n\n# + [markdown] id=\"RzWFEMTRTYD0\"\n# #Imports and Setup\n\n# + id=\"_cajUFlyllOs\"\nimport warnings\nwarnings.filterwarnings('ignore')\n\nimport pandas as pd\nimport numpy as np\nfrom plotnine import *\nimport seaborn as sns\nimport scipy.cluster.hierarchy as shc\n\nfrom sklearn.preprocessing import StandardScaler\n\nfrom sklearn.neighbors import NearestNeighbors\nfrom sklearn.cluster import DBSCAN\nfrom sklearn.cluster import KMeans\nfrom sklearn.cluster import AgglomerativeClustering\nfrom sklearn.mixture import GaussianMixture\n\nfrom sklearn.metrics import silhouette_score\n\nimport pandas as pd\nfrom random import randint\nfrom sklearn import datasets\nimport matplotlib.pyplot as plt\nfrom scipy.spatial import distance\nimport numpy as np\nimport pylab\n\n\n\n# %matplotlib inline\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 206} id=\"C3Q3Vh0GllO6\" outputId=\"72bff194-89b5-450e-e990-d0a52dbfb8e8\"\ndf = pd.read_csv(\"https://raw.githubusercontent.com/cmparlettpelleriti/CPSC392ParlettPelleriti/master/Data/KMEM1.csv\")\n\ndf.head()\n\n\n\n# + [markdown] id=\"JB7hKwymkzT3\"\n# # Prepare data\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 296} id=\"X-WrjCGLfMuc\" outputId=\"469fec00-d835-4ae3-e910-52570ecb6c77\"\nsns.scatterplot(x='x',y='y',data=df)\n\n# + id=\"a0Uhf9EB2-Fj\"\ndf_scaled=StandardScaler().fit_transform(df)\n\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"jS0UDpsN-9Gw\" outputId=\"14757898-4643-4330-ea1a-4e3d7f2281fb\"\ndf_scaled[:,0].shape\n\n# + [markdown] id=\"nH9lDbE8k5DE\"\n# # Elbow method\n\n# + [markdown] id=\"DX92Xrg2lDc6\"\n# - implement elbow method to choose best K value for kmeans clustering\n\n# + id=\"FlU9rud7b5KI\"\ndistortions = []\nK = range(1,10)\nfor k in K:\n kmeanModel = KMeans(n_clusters=k)\n kmeanModel.fit(df)\n distortions.append(kmeanModel.inertia_)\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 295} id=\"pII01zCEb5BC\" outputId=\"77e71013-9f32-41ce-c530-ce07e26ab1d6\"\nplt.plot(K, distortions, marker = 'o')\nplt.xlabel('k')\nplt.ylabel('Distortion')\nplt.title('The Elbow Method showing the optimal k')\nplt.show()\n\n# + [markdown] id=\"R8rjzsEflUOJ\"\n# # Dendogram \n\n# + [markdown] id=\"jODcnSidlZaL\"\n# - plot dendrogram visualize the result of a hierarchical clustering if performed on data.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 444} id=\"_mNltcQ9yzrh\" outputId=\"dc842235-ff2b-44f8-ab73-05eea333f522\"\nplt.figure(figsize=(10, 7))\nplt.title(\"Counters Dendograms\")\ndend = shc.dendrogram(shc.linkage(y=df_scaled , method='single',metric='euclidean'))\n\n# + [markdown] id=\"Rbmf8ri5l7t8\"\n# # Clustering\n\n# + [markdown] id=\"1fsF1XjQl0M4\"\n# ## KMeans clustering\n\n# + [markdown] id=\"3b_CnsPDqHdn\"\n# - perform Kmeans clustering\n\n# + id=\"fnL-DNHbmDP0\"\nkm = KMeans(3)\nkm.fit(df_scaled)\ndf[\"km\"] = km.labels_\n\n# + [markdown] id=\"gg8LWKqOqKKe\"\n# - visualize output clusters\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} id=\"u_ru6htvmkuu\" outputId=\"6ae38d9c-b0f5-40f3-d6d9-3197416ab3c0\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(km)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"KMeans\"))\n\n\n# + [markdown] id=\"8ePSYVETqOqw\"\n# - evaluate cluster performance using silhouette score.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"gF8PETyYqGjd\" outputId=\"7594aa61-aecf-45f2-d6f5-7f7b1e381e44\"\npreds = km.predict(df_scaled)\nscore = silhouette_score(df, preds)\nprint(\"silhouette score using kmeans is {}\".format(score))\n\n# + [markdown] id=\"9-E42y5kmH2P\"\n# ## Agglomerative clustering\n\n# + [markdown] id=\"tgCfui1mqck2\"\n# - perform agglomerative clustering.\n\n# + id=\"3KDclawemMWS\"\nag = AgglomerativeClustering(3)\nag.fit(df_scaled)\ndf[\"hac\"] = ag.labels_\n\n# + [markdown] id=\"3yRJ-LkVqiwB\"\n# - visualize output clusters.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} id=\"ma0AwvaXmorz\" outputId=\"95e48459-f32c-4858-a802-b589070c1c74\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(hac)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"Hierarchical Agglomeretive\"))\n\n# + [markdown] id=\"6cf4mgC0qmGO\"\n# - evaluate cluster performance using silhouette score.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"bVhAofOgq6ci\" outputId=\"8070315e-df03-43ee-b7b9-dd91d9af71e2\"\nscore = silhouette_score(df, ag.labels_)\nprint(\"silhouette score using AgglomerativeClustering is {}\".format(score))\n\n# + [markdown] id=\"5ghPqRj_mSQw\"\n# ## Gaussian Mixture clustering\n\n# + [markdown] id=\"CVBpUUq9ruZU\"\n# - perform Gaussian Mixture clustering.\n\n# + id=\"Hhvs-XRpmYZh\"\ngm = GaussianMixture(3)\ngm.fit(df_scaled)\ndf[\"gm\"] = gm.predict(df[[\"x\",\"y\"]])\n\n# + [markdown] id=\"2lxdQyQGrHJJ\"\n# - visualize output clusters.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} id=\"HVUPOXLpmm1b\" outputId=\"4826407b-1917-4655-f9f4-c0214ccdee13\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(gm)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"Gaussian Mixtures\"))\n\n# + [markdown] id=\"AuUsIFZVqv1D\"\n# - evaluate cluster performance using silhouette score.\n\n# + colab={\"base_uri\": \"https://localhost:8080/\"} id=\"WAaNLi1hrPVj\" outputId=\"ff925df2-9de3-4a83-d125-50f04804ac9d\"\npreds = gm.predict(df[[\"x\",\"y\"]])\nscore = silhouette_score(df, preds)\nprint(\"silhouette score using GaussianMixture is {}\".format(score))\n\n# + [markdown] id=\"eQh9GE5qmc84\"\n# ## DBSAN clustering\n\n# + [markdown] id=\"2csU_aLer0DN\"\n# - perform DBSCAN clustering.\n\n# + id=\"97FjNxdahWOD\"\ndbscan = DBSCAN(eps=0.5, min_samples=10)\ndbscan.fit(df_scaled)\ndf[\"db\"] = dbscan.labels_\n\n# + [markdown] id=\"GJdIuOwRrIDk\"\n# - visualize output clusters.\n\n# + id=\"mi33GjH8jEht\" colab={\"base_uri\": \"https://localhost:8080/\", \"height\": 503} outputId=\"7446945c-cb90-4487-b2e7-8b687febd967\"\nprint(ggplot(df, aes(x = \"x\", y = \"y\",\n color = \"factor(db)\")) +\n geom_point() +\n theme_minimal() +\n scale_color_discrete(name = \"Cluster Assignment\") +\n theme(panel_grid_major = element_blank()) + \n labs(title = \"DBSCAN\"))\n\n\n# + [markdown] id=\"rYKqybQIqw2a\"\n# - evaluate cluster performance using silhouette score.\n\n# + id=\"RExwvq_YjJJx\" colab={\"base_uri\": \"https://localhost:8080/\"} outputId=\"6a49f8f9-6b12-4639-974e-3b6444260245\"\nscore = silhouette_score(df_scaled, dbscan.labels_)\nprint(\"silhouette score using dbscan is {}\".format(score))\n\n# + [markdown] id=\"tEtKxBsbr5RV\"\n# # Conclusion\n\n# + [markdown] id=\"iBR0zUdUr94N\"\n# - kmeans then Agglomerative clustering was the best clustering algorithms to perform having silhouette score of approximatly 6 and 0.57 and that was due to the behavior and distribution while the other clustering algorithms may perform better on other data.\n","repo_name":"ayaamr11/Clustering-Algorithms","sub_path":"ClusteringAlgorithms.ipynb","file_name":"ClusteringAlgorithms.ipynb","file_ext":"py","file_size_in_byte":7454,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"39593347418","text":"import numpy as np\nimport pandas as pd\nimport pyflux as pf\nimport datetime as datetime\nimport matplotlib.pyplot as plt\nimport os\nimport pickle\nimport ForecastModel as fc\n\n# +\nX_fwds = pickle.load(open('../data/forward_rates', 'rb'))\n#X_fwds = df_add_first_diff(X_fwds)\n\n#df_FX = pickle.load( open( \"data/FX_data\", \"rb\" ) )\n# Loading up the federal reserve speech data\nfed_metrics = pickle.load( open( \"../data/final_press_rel_dist\", \"rb\" ) )\n# -\n\n\n\n# +\ncos_last = fed_metrics['cos_last']\ncos_avg_n = fed_metrics['cos_avg_n']\ned_last = fed_metrics['ed_last']\ned_avg_n = fed_metrics['ed_avg_n']\nfed_dates = fed_metrics['dates']\n\n#grouping by date (some dates had multiple speeches)\navgstats = pd.DataFrame({'date':fed_dates,\n 'ed_last': ed_last,\n 'ed_avg_n': ed_avg_n,\n 'cos_last': cos_last,\n 'cos_avg_n': cos_avg_n}).groupby('date').mean()\navgstats.index = pd.to_datetime(avgstats.index)\n# -\n\nthis_min = min(avgstats.index)\nthis_min\n\nX_fwds.info()\n\n# now I need to filter the X_fwds to drop anything before this period!\nX_after = X_fwds[X_fwds.index >= this_min]\n\n# +\nX_fwds = X_fwds[X_fwds.index >= this_min]\nX_fwds = X_fwds.merge(avgstats, how='left', left_index = True, right_index = True)\nX_fwds.fillna(value=0, inplace=True)\n\n# the first row of X_fwds contains zeros for the differenced rates, clear them here\nX_fwds = X_fwds.drop(X_fwds.index[0])\n\n# -\n\nX_fwds.describe()\n\ntemp = X_fwds['ed_last'].copy()\n\ntemp =pd.concat([temp, temp.shift(), temp.shift(2)], axis=1)\n\ntemp.head()\n\nX_fwds['ed_last_l1']= temp.iloc[:,1]\nX_fwds['ed_last_l2']= temp.iloc[:,2]\nX_fwds.head()\n\n\n\n\ntemp2\n\nlen(temp2)\n\nlen(temp)\n\n\n","repo_name":"davidjsmith44/NLP-Fed-Speeches","sub_path":"src/Untitled5.ipynb","file_name":"Untitled5.ipynb","file_ext":"py","file_size_in_byte":1700,"program_lang":"python","lang":"en","doc_type":"code","stars":10,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"33880389887","text":"# # Capstone Project\n# ## Image classifier for the SVHN dataset\n# ### Instructions\n#\n# In this notebook, you will create a neural network that classifies real-world images digits. You will use concepts from throughout this course in building, training, testing, validating and saving your Tensorflow classifier model.\n#\n# This project is peer-assessed. Within this notebook you will find instructions in each section for how to complete the project. Pay close attention to the instructions as the peer review will be carried out according to a grading rubric that checks key parts of the project instructions. Feel free to add extra cells into the notebook as required.\n#\n# ### How to submit\n#\n# When you have completed the Capstone project notebook, you will submit a pdf of the notebook for peer review. First ensure that the notebook has been fully executed from beginning to end, and all of the cell outputs are visible. This is important, as the grading rubric depends on the reviewer being able to view the outputs of your notebook. Save the notebook as a pdf (File -> Download as -> PDF via LaTeX). You should then submit this pdf for review.\n#\n# ### Let's get started!\n#\n# We'll start by running some imports, and loading the dataset. For this project you are free to make further imports throughout the notebook as you wish. \n\nimport tensorflow as tf\nfrom scipy.io import loadmat\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport random\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPooling2D, BatchNormalization\nfrom tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping\n\n# \n# For the capstone project, you will use the [SVHN dataset](http://ufldl.stanford.edu/housenumbers/). This is an image dataset of over 600,000 digit images in all, and is a harder dataset than MNIST as the numbers appear in the context of natural scene images. SVHN is obtained from house numbers in Google Street View images. \n#\n# * Y. Netzer, T. Wang, A. Coates, A. Bissacco, B. Wu and A. Y. Ng. \"Reading Digits in Natural Images with Unsupervised Feature Learning\". NIPS Workshop on Deep Learning and Unsupervised Feature Learning, 2011.\n#\n# Your goal is to develop an end-to-end workflow for building, training, validating, evaluating and saving a neural network that classifies a real-world image into one of ten classes.\n\n# +\n# Run this cell to load the dataset\n\ntrain = loadmat('data/train_32x32.mat')\ntest = loadmat('data/test_32x32.mat')\n\n\n# -\n\n# Both `train` and `test` are dictionaries with keys `X` and `y` for the input images and labels respectively.\n\n# ## 1. Inspect and preprocess the dataset\n# * Extract the training and testing images and labels separately from the train and test dictionaries loaded for you.\n# * Select a random sample of images and corresponding labels from the dataset (at least 10), and display them in a figure.\n# * Convert the training and test images to grayscale by taking the average across all colour channels for each pixel. _Hint: retain the channel dimension, which will now have size 1._\n# * Select a random sample of the grayscale images and corresponding labels from the dataset (at least 10), and display them in a figure.\n\ndef get_train_test_data(train_data, test_data):\n \"\"\"\n This function preprocesses and returns train and test data from received dictionary\n Data is transposed to make batch size dimension first and normalized\n \"\"\"\n x_train, y_train = train_data['X'], train_data['y']\n x_test, y_test = test_data['X'], test_data['y']\n # Set labels between 0 and 9\n y_train -= 1\n y_test -= 1\n # transpose to batch size first\n x_train = np.transpose(x_train, (3, 0, 1, 2))\n x_test = np.transpose(x_test, (3, 0, 1, 2))\n # min - max: 0 - 255\n x_train = x_train / 255.0\n x_test = x_test / 255.0\n return x_train, y_train, x_test, y_test\n\n\nx_train, y_train, x_test, y_test = get_train_test_data(train, test)\nprint(f'x_train shape is: {x_train.shape}')\nprint(f'y_train shape is: {y_train.shape}')\nprint(f'x_test shape is: {x_test.shape}')\nprint(f'y_test shape is: {y_test.shape}')\nprint(f'x_train data: min value is {np.min(x_train)} and max value is {np.max(x_train)}')\nprint(f'x_test data: min value is {np.min(x_test)} and max value is {np.max(x_test)}')\nprint(f'y_train data: min value is {np.min(y_train)} and max value is {np.max(y_train)}')\nprint(f'y_test data: min value is {np.min(y_test)} and max value is {np.max(y_test)}')\n\n# +\nnum_imgs = 10\nfig, ax = plt.subplots(1, num_imgs, figsize=(num_imgs, 1))\nrandom_sample = random.sample(range(len(x_train)), num_imgs)\n\nfor idx, sample in enumerate(random_sample):\n ax[idx].set_axis_off()\n ax[idx].imshow(x_train[sample])\n ax[idx].set_title(y_train[sample][0] + 1)\n# -\n\nx_train = np.mean(x_train, axis = 3)[..., np.newaxis]\nx_test = np.mean(x_test, axis = 3)[..., np.newaxis]\n\nprint(f'x_train shape is: {x_train.shape}')\nprint(f'x_test shape is: {x_test.shape}')\n\nfig, ax = plt.subplots(1, num_imgs, figsize=(num_imgs, 1))\nfor idx, sample in enumerate(random_sample):\n ax[idx].set_axis_off()\n ax[idx].imshow(np.squeeze(x_train[sample]), cmap='gray')\n ax[idx].set_title(y_train[sample][0] + 1)\n\n\n# ## 2. MLP neural network classifier\n# * Build an MLP classifier model using the Sequential API. Your model should use only Flatten and Dense layers, with the final layer having a 10-way softmax output. \n# * You should design and build the model yourself. Feel free to experiment with different MLP architectures. _Hint: to achieve a reasonable accuracy you won't need to use more than 4 or 5 layers._\n# * Print out the model summary (using the summary() method)\n# * Compile and train the model (we recommend a maximum of 30 epochs), making use of both training and validation sets during the training run. \n# * Your model should track at least one appropriate metric, and use at least two callbacks during training, one of which should be a ModelCheckpoint callback.\n# * As a guide, you should aim to achieve a final categorical cross entropy training loss of less than 1.0 (the validation loss might be higher).\n# * Plot the learning curves for loss vs epoch and accuracy vs epoch for both training and validation sets.\n# * Compute and display the loss and accuracy of the trained model on the test set.\n\ndef get_mlp_model(input_shape):\n \"\"\"\n Returns compiled MLP model\n \"\"\"\n model = Sequential([\n Flatten(input_shape = input_shape),\n #Dense(64, activation = 'relu'),\n Dense(256, activation = 'relu'),\n Dense(256, activation = 'relu'),\n Dense(128, activation = 'relu'),\n #BatchNormalization(),\n #Dense(32, activation = 'relu'),\n #Dropout(0.3),\n Dense(10, activation = 'softmax')\n ])\n model.compile(optimizer = 'adam', loss = 'sparse_categorical_crossentropy', metrics = ['accuracy'])\n return model\n\n\nmodel = get_mlp_model(x_train[0].shape)\nmodel.summary()\n\n\ndef get_checkpoints(checkpoint_path):\n \"\"\"\n Returns a list with early stopping and best checkpoint \n \"\"\"\n best_model = ModelCheckpoint(checkpoint_path, \n monitor = 'val_accuracy', save_weights_only = True,\n save_best_only = True)\n early_stopping = EarlyStopping(monitor = 'val_accuracy', patience = 4)\n return [best_model, early_stopping]\n\n\nhistory = model.fit(x_train, y_train, epochs = 30, batch_size = 64, validation_split = 0.15, callbacks = get_checkpoints('mlp/checkpoints_best_only/checkpoint'))\n\nplt.plot(history.history['accuracy'])\nplt.plot(history.history['val_accuracy'])\nplt.title('Accuracy vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='lower right')\nplt.show()\n\nplt.plot(history.history['loss'])\nplt.plot(history.history['val_loss'])\nplt.title('Loss vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='upper right')\nplt.show()\n\nmodel = get_mlp_model(x_train[0].shape)\nmodel.load_weights('mlp/checkpoints_best_only/checkpoint')\ntest_loss, test_acc = model.evaluate(x = x_test, y = y_test, verbose = 0)\nprint('Loss in test set is {:.02f} and Accuracy in test set is {:.02f}'.format(test_loss, test_acc))\n\n\n# ## 3. CNN neural network classifier\n# * Build a CNN classifier model using the Sequential API. Your model should use the Conv2D, MaxPool2D, BatchNormalization, Flatten, Dense and Dropout layers. The final layer should again have a 10-way softmax output. \n# * You should design and build the model yourself. Feel free to experiment with different CNN architectures. _Hint: to achieve a reasonable accuracy you won't need to use more than 2 or 3 convolutional layers and 2 fully connected layers.)_\n# * The CNN model should use fewer trainable parameters than your MLP model.\n# * Compile and train the model (we recommend a maximum of 30 epochs), making use of both training and validation sets during the training run.\n# * Your model should track at least one appropriate metric, and use at least two callbacks during training, one of which should be a ModelCheckpoint callback.\n# * You should aim to beat the MLP model performance with fewer parameters!\n# * Plot the learning curves for loss vs epoch and accuracy vs epoch for both training and validation sets.\n# * Compute and display the loss and accuracy of the trained model on the test set.\n\ndef get_cnn_model(input_shape):\n \"\"\"\n Returns compiled CNN model\n \"\"\"\n model = Sequential([\n Conv2D(16, (3, 3), activation = 'relu', input_shape = input_shape),\n MaxPooling2D((2, 2)),\n Conv2D(32, (3, 3), activation = 'relu'),\n MaxPooling2D((2, 2)),\n BatchNormalization(),\n Flatten(),\n Dense(32, activation = 'relu'),\n Dropout(0.3),\n Dense(10, activation = 'softmax')\n ])\n model.compile(optimizer = 'adam', loss = 'sparse_categorical_crossentropy', metrics = ['accuracy'])\n return model\n\n\nmodel = get_cnn_model(x_train[0].shape)\nmodel.summary()\n\nhistory_cnn = model.fit(x_train, y_train, epochs = 30, batch_size = 64, validation_split = 0.15, callbacks = get_checkpoints('cnn/checkpoints_best_only/checkpoint'))\n\nplt.plot(history_cnn.history['accuracy'])\nplt.plot(history_cnn.history['val_accuracy'])\nplt.title('Accuracy vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='lower right')\nplt.show()\n\nplt.plot(history_cnn.history['loss'])\nplt.plot(history_cnn.history['val_loss'])\nplt.title('Loss vs. epochs')\nplt.ylabel('Loss')\nplt.xlabel('Epoch')\nplt.legend(['Training', 'Validation'], loc='upper right')\nplt.show()\n\nmodel = get_cnn_model(x_train[0].shape)\nmodel.load_weights('cnn/checkpoints_best_only/checkpoint')\ntest_loss, test_acc = model.evaluate(x = x_test, y = y_test, verbose = 0)\nprint('Loss in test set is {:.02f} and Accuracy in test set is {:.02f}'.format(test_loss, test_acc))\n\n# ## 4. Get model predictions\n# * Load the best weights for the MLP and CNN models that you saved during the training run.\n# * Randomly select 5 images and corresponding labels from the test set and display the images with their labels.\n# * Alongside the image and label, show each model’s predictive distribution as a bar chart, and the final model prediction given by the label with maximum probability.\n\n# ### 4.1. MLP model\n\nrandom_idx = np.random.choice(len(x_test), 5)\nrandom_test_images = x_test[random_idx]\nrandom_test_labels = y_test[random_idx]\n\nmlp_model = get_mlp_model(x_train[0].shape)\nmlp_model.load_weights('mlp/checkpoints_best_only/checkpoint')\n\n# +\npredictions = mlp_model.predict(random_test_images)\n\nfig, axes = plt.subplots(5, 2, figsize = (16, 12))\nfig.subplots_adjust(hspace = 0.4, wspace = -0.2)\n\nfor i, (prediction, image, label) in enumerate(zip(predictions, random_test_images, random_test_labels)):\n axes[i, 0].set_axis_off()\n axes[i, 0].imshow(np.squeeze(image), cmap = 'gray')\n axes[i, 0].text(10., -1.5, f'Label: {label[0] + 1}')\n axes[i, 1].bar(np.arange(1, len(prediction) + 1), prediction)\n axes[i, 1].set_xticks(np.arange(len(prediction) + 1))\n axes[i, 1].set_title(f\"Model prediction: {np.argmax(prediction) + 1}\")\n \nplt.show()\n# -\n\n# ### 4.2. CNN Model\n\ncnn_model = get_cnn_model(x_train[0].shape)\ncnn_model.load_weights('cnn/checkpoints_best_only/checkpoint')\n\n# +\npredictions = cnn_model.predict(random_test_images)\n\nfig, axes = plt.subplots(5, 2, figsize = (16, 12))\nfig.subplots_adjust(hspace = 0.4, wspace = -0.2)\n\nfor i, (prediction, image, label) in enumerate(zip(predictions, random_test_images, random_test_labels)):\n axes[i, 0].set_axis_off()\n axes[i, 0].imshow(np.squeeze(image), cmap = 'gray')\n axes[i, 0].text(10., -1.5, f'Label: {label[0] + 1}')\n axes[i, 1].bar(np.arange(1, len(prediction) + 1), prediction)\n axes[i, 1].set_xticks(np.arange(len(prediction) + 1))\n axes[i, 1].set_title(f\"Model prediction: {np.argmax(prediction) + 1}\")\n \nplt.show()\n# -\n\n\n","repo_name":"jcgarciaca/getting_started_with_tensorflow2","sub_path":"Capstone Project.ipynb","file_name":"Capstone Project.ipynb","file_ext":"py","file_size_in_byte":13047,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"52"}
+{"seq_id":"23535976144","text":"# +\nimport keras\nimport tensorflow as tf\nimport keras.layers as layers\nimport keras.activations as activations\nimport numpy as np\nimport tensorflow_probability as tfp\nimport pylab as pl\nimport matplotlib.pyplot as plt\nimport scipy as sp\nimport scipy.signal \nfrom scipy.io import wavfile\n\nimport sounddevice as sd\n# -\n\n# # 1. Mise en place des données tests\n\n# +\nrate, sls = wavfile.read('Sls.wav')\nsls = sls/np.max(sls)\n\nN = len(sls)\nprint(N/rate)\n\ntime = np.arange(N)/rate\n\nplt.plot(sls)\nplt.title('Signal')\nplt.show()\n\n#sd.play(sls/np.max(sls),rate)\n\n# +\nNs = N\nsigma_carre = 1.2\n\n#compute gaussian signal\nSamples = np.power(1.2,0.5)*np.random.randn(Ns)\nabs = np.arange(Ns)\npl.plot(abs,Samples)\npl.show()\n\n# +\nslsn = sls + Samples/90\n\npl.plot(slsn)\n\npl.show()\n\n#sd.play(slsn[3200000:3200000+3200000]/np.max(sls),rate)\n\n# +\ndef make_extrait(x,fs):\n data = np.zeros((1,fs*2))\n i=0\n data = [x[i:i+fs*2]]\n i+=fs\n while(i+fs*2 \n#\n# The project is broken down into multiple steps:\n#\n# * Load and preprocess the image dataset\n# * Train the image classifier on your dataset\n# * Use the trained classifier to predict image content\n#\n# We'll lead you through each part which you'll implement in Python.\n#\n# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.\n#\n# First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.\n\n# +\n# Imports here\nimport numpy as np\nimport matplotlib.pyplot as plt\n\nimport torch\nimport torchvision\nfrom torchvision import datasets, transforms, models\nfrom torch import nn\nfrom torch import optim\n#import torch.nn.functional as F\nfrom torch.optim import lr_scheduler\nfrom torch.autograd import Variable\n\nimport copy\nimport time\nimport json\n\nfrom collections import OrderedDict\n\nfrom PIL import Image\nimport random\n# -\n\n# Set seed\nrandom.seed(2)\nnp.random.seed(2)\ntorch.manual_seed(2)\ntorch.cuda.manual_seed_all(2)\ntorch.backends.cudnn.deterministic=True\n\n# ## Load the data\n#\n# Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.\n#\n# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.\n#\n# The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.\n# \n\ndata_dir = 'flowers'\ntrain_dir = data_dir + '/train'\nvalid_dir = data_dir + '/valid'\ntest_dir = data_dir + '/test'\n\n# +\n# TODO: Define your transforms for the training, validation, and testing sets\ndata_transforms = {'train' : transforms.Compose([transforms.RandomRotation(30),\n transforms.RandomResizedCrop(224),\n transforms.RandomHorizontalFlip(),\n transforms.ToTensor(),\n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'valid' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'test' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])])}\n\n# TODO: Load the datasets with ImageFolder\nimage_datasets = {'train' : datasets.ImageFolder(train_dir, transform=data_transforms['train']),\n 'valid' : datasets.ImageFolder(valid_dir, transform=data_transforms['valid']),\n 'test' : datasets.ImageFolder(test_dir, transform=data_transforms['test'])}\n\n# TODO: Using the image datasets and the trainforms, define the dataloaders\ndataloaders = {'train' : torch.utils.data.DataLoader(image_datasets['train'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'valid' : torch.utils.data.DataLoader(image_datasets['valid'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'test' : torch.utils.data.DataLoader(image_datasets['test'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2))}\n\ndataset_sizes = {'train': len(image_datasets['train']),\n 'valid': len(image_datasets['valid']),\n 'test': len(image_datasets['test'])} \n# -\n\n# ### Label mapping\n#\n# You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.\n\nwith open('cat_to_name.json', 'r') as f:\n cat_to_name = json.load(f)\n\n# # Building and training the classifier\n#\n# Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.\n#\n# We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students! You can also ask questions on the forums or join the instructors in office hours.\n#\n# Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:\n#\n# * Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)\n# * Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout\n# * Train the classifier layers using backpropagation using the pre-trained network to get the features\n# * Track the loss and accuracy on the validation set to determine the best hyperparameters\n#\n# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!\n#\n# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.\n\n# +\n# TODO: Build and train your network\n\n# +\n# default device for all functions with device argument: the fastest device available\n# all functions move models to device at the start and all returned models are moved to cpu at the end: \n# to avoid having problems with loading checkpoint when cuda is not available \ndevice = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\n\n# Defaults for functions to follow: architecture and parameters of the best_model\narchitecture = 'vgg16'\nlr = 0.0001\nhu = 4096\nstep_size = 5\n# Best num_epochs was 8 with lr=0.0001 but 10 with lr=0.001. \n# Keeping value at 10 is ok as train_model picks best number of epochs anyway.\n# We keep default num_epochs=10 in train.py as well. \nnum_epochs = 10 \n\n\n# -\n\ndef pick_arch(architecture):\n \"\"\"Pick from the following model architectures: vgg11, vgg13, vgg16, vgg19.\"\"\"\n if architecture == 'vgg11':\n model = models.vgg11(pretrained=True)\n elif architecture == 'vgg13':\n model = models.vgg13(pretrained=True)\n elif architecture == 'vgg16':\n model = models.vgg16(pretrained=True)\n elif architecture == 'vgg19':\n model = models.vgg19(pretrained=True)\n else:\n model = None\n print('Wrong model architecture. Pick vgg11, vgg13, vgg16 or vgg19.')\n \n return model\n\n\ndef build_model(architecture=architecture, hu=hu):\n \"\"\"Build a model from given architecture and replace classifier with a custom one.\"\"\"\n model = pick_arch(architecture)\n\n for param in model.parameters():\n param.requires_grad = False\n\n input_size = list(model.classifier.parameters())[0].size()[1]\n output_size = len(image_datasets['train'].class_to_idx)\n\n classifier = nn.Sequential(OrderedDict([\n ('fc1', nn.Linear(input_size, hu)),\n ('relu', nn.ReLU()),\n ('dropout', nn.Dropout(p=0.5)),\n ('fc2', nn.Linear(hu, output_size)),\n ('output', nn.LogSoftmax(dim=1))\n ]))\n\n model.classifier = classifier\n \n return model\n\n\ndef train_model(model=None, criterion=None, optimizer=None, scheduler=None,\n num_epochs=num_epochs, device=device):\n since = time.time()\n \n if model == None:\n model = build_model()\n \n if criterion == None:\n criterion = nn.NLLLoss()\n \n if optimizer == None:\n optimizer = optim.Adam(model.classifier.parameters(), lr)\n \n if scheduler == None:\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) \n \n model = model.to(device)\n \n best_model_wts = copy.deepcopy(model.state_dict())\n best_acc = 0.0\n best_num_epochs = 0\n\n for epoch in range(num_epochs):\n print('Epoch {}/{}'.format(epoch, num_epochs - 1))\n print('-' * 10)\n\n # Each epoch has a training and validation phase\n for phase in ['train', 'valid']:\n if phase == 'train':\n scheduler.step()\n model.train() # Set model to training mode\n else:\n model.eval() # Set model to evaluate mode\n\n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n # zero the parameter gradients\n optimizer.zero_grad()\n\n # forward\n # track history if only in train\n with torch.set_grad_enabled(phase == 'train'):\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # backward + optimize only if in training phase\n if phase == 'train':\n loss.backward()\n optimizer.step()\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n epoch_loss = running_loss / dataset_sizes[phase]\n epoch_acc = running_corrects.double() / dataset_sizes[phase]\n\n print('{} Loss: {:.4f} Acc: {:.4f}'.format(\n phase, epoch_loss, epoch_acc))\n\n # deep copy the model\n if phase == 'valid' and epoch_acc > best_acc:\n best_acc = epoch_acc\n best_model_wts = copy.deepcopy(model.state_dict())\n best_num_epochs = epoch + 1\n\n print('-' * 10)\n time_elapsed = time.time() - since\n print('Training complete in {:.0f}m {:.0f}s'.format(\n time_elapsed // 60, time_elapsed % 60))\n print('Best validation Accuracy of {:.4f} for {} epochs'.format(best_acc, best_num_epochs))\n print()\n\n # load best model weights\n model.load_state_dict(best_model_wts)\n model.to('cpu')\n return model, best_acc, best_num_epochs\n\n\n# The trial_model below was trained when default lr was set to 0.001. \n\ntrial_model, trial_acc, trial_num_epochs = train_model(num_epochs=2)\n\n\ndef tune_hyperparams(architecture=architecture, num_epochs=num_epochs,\n lr_list=[lr], hu_list=[hu], step_size_list=[step_size], device=device):\n param_triples = [(lr, hu, step_size) for lr in lr_list for hu in hu_list for step_size in step_size_list]\n \n best_model = pick_arch(architecture)\n best_acc = 0.0\n best_num_epochs = 0\n best_lr = 0.0\n best_hu = 0\n best_step_size = 0\n \n for lr, hu, step_size in param_triples:\n print('Training with')\n print('Learning rate: {} Number of hidden units: {} Step size: {}'.format(lr, hu, step_size))\n print('-' * 10)\n\n model = build_model(architecture=architecture, hu=hu)\n model = model.to(device)\n \n criterion = nn.NLLLoss()\n optimizer = optim.Adam(model.classifier.parameters(), lr=lr)\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) #add these to hyperparams\n \n model_trained, acc_trained, num_epochs_trained = train_model(model, criterion, optimizer, scheduler, num_epochs=num_epochs, device=device)\n \n # deep copy the model\n if acc_trained > best_acc:\n best_acc = acc_trained\n best_model = copy.deepcopy(model_trained)\n best_num_epochs = num_epochs_trained\n best_lr = lr\n best_hu = hu\n best_step_size = step_size\n \n print()\n print('Best results:')\n print('Learning Rate: {:.4f} Number of Hidden Units: {:.4f} Step size: {:.4f}'.format(best_lr, best_hu, best_step_size))\n print('Number of Epochs: {:.4f} Validation Acc: {:.4f}'.format(best_num_epochs, best_acc))\n best_model.to('cpu')\n return best_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size\n\n\ntune_hyperparams(num_epochs=2)\n\n# I tried learning rate of 0.01, number of hidden units 2048 and step size 7 but they did not do as well as lr=0.001, hu=4096 and step_size=5 and I decided to not spend more time training combinations with the above values. The last step in deciding on parameters is to see if learning rates of 0.0001 does better than 0.001. The train_model function already chooses the best number of epochs between 1 and 10. \n\nbest_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size = tune_hyperparams(lr_list=[0.0001, 0.001], hu_list=[4096], step_size_list=[5])\n\n\n# ## Testing your network\n#\n# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.\n\n# TODO: Do validation on the test set\ndef do_validation(model=None, criterion=None, phase='test', device=device):\n if model == None:\n model = build_model()\n if criterion == None:\n criterion = nn.NLLLoss()\n \n model.to(device)\n model.eval()\n \n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n\n # forward\n # don't track history \n with torch.no_grad():\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n total_loss = running_loss / dataset_sizes[phase]\n total_acc = running_corrects.double() / dataset_sizes[phase]\n \n model.to('cpu')\n \n print('Loss: {:.4f} Acc: {:.4f}'.format(\n total_loss, total_acc))\n return total_loss, total_acc.item()\n\ntotal_loss, total_acc = do_validation(model = best_model) \n\n# +\n#next(best_model.parameters()).is_cuda\n# -\n\n# ## Save the checkpoint\n#\n# Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.\n#\n# ```model.class_to_idx = image_datasets['train'].class_to_idx```\n#\n# Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.\n\n# TODO: Save the checkpoint \ncheckpoint = {'state_dict': best_model.state_dict(),\n 'hu': best_hu,\n 'architecture': architecture,\n 'num_epochs': best_num_epochs,\n 'lr': best_lr,\n 'step_size': best_step_size,\n 'class_to_idx': image_datasets['train'].class_to_idx}\n\ntorch.save(checkpoint, 'checkpoint_best.pth')\n\nbest_model.class_to_idx = image_datasets['train'].class_to_idx\n\n\n# ## Loading the checkpoint\n#\n# At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.\n\n# TODO: Write a function that loads a checkpoint and rebuilds the model\ndef load_checkpoint(filepath):\n checkpoint = torch.load(filepath)\n \n model = build_model(architecture=checkpoint['architecture'], hu=checkpoint['hu'])\n\n model.load_state_dict(checkpoint['state_dict'])\n model.class_to_idx = checkpoint['class_to_idx']\n \n return model, {'num_epochs':checkpoint['num_epochs'], \n 'lr': checkpoint['lr'], \n 'step_size': checkpoint['step_size'], \n 'hu': checkpoint['hu'], \n 'architecture': checkpoint['architecture']}\n\n\nreloaded_best_model, reloaded_best_params_dict = load_checkpoint('checkpoint_best.pth') \n\ndo_validation(model = reloaded_best_model)\n\n\n# # Inference for classification\n#\n# Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like \n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n#\n# First you'll need to handle processing the input image such that it can be used in your network. \n#\n# ## Image Preprocessing\n#\n# You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training. \n#\n# First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.\n#\n# Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.\n#\n# As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation. \n#\n# And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.\n\ndef process_image(image):\n ''' Scales, crops, and normalizes a PIL image for a PyTorch model,\n returns an Numpy array\n '''\n # TODO: Process a PIL image for use in a PyTorch model\n pil_image = Image.open(image)\n width, height = pil_image.size\n l = int(256 * max(width, height) / min(width, height))\n pil_image.thumbnail(size = (l,l))\n width, height = pil_image.size \n pil_image = pil_image.crop(box = (width / 2 - 112, height / 2 - 112, width / 2 + 112, height / 2 + 112))\n \n np_image = np.array(pil_image)\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n np_image = (np_image / 255.0 - mean) / std\n \n np_image = np_image.transpose((2, 0, 1))\n \n return np_image\n\n\n# To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).\n\ndef imshow(image, ax=None, title=None):\n if ax is None:\n fig, ax = plt.subplots()\n \n # PyTorch tensors assume the color channel is the first dimension\n # but matplotlib assumes is the third dimension\n image = image.transpose((1, 2, 0))\n \n # Undo preprocessing\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n image = std * image + mean\n \n # Image needs to be clipped between 0 and 1 or it looks like noise when displayed\n image = np.clip(image, 0, 1)\n \n ax.imshow(image)\n \n return ax\n\n\nimshow(process_image('flowers/test/1/image_06743.jpg'))\n\n\n# ## Class Prediction\n#\n# Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.\n#\n# To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.\n#\n# Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.\n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n\n# +\ndef predict(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\ndef predict2(image_path, checkpoint, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model from checkpoint.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model, _ = load_checkpoint(checkpoint)\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\n\n\n# small change in processing image: use transforms instead of process_image function\ndef predict3(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n transfs = data_transforms['test']\n\n img_pil = Image.open(image_path)\n img_tensor = transfs(img_pil)\n img_tensor.unsqueeze_(0)\n img_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(img_variable)\n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes \n\n\n# -\n\nprobs, classes = predict('flowers/test/1/image_06743.jpg', reloaded_best_model)\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\nprobs, classes = predict2('flowers/test/1/image_06743.jpg', 'checkpoint_best.pth')\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\n\n# ## Sanity Checking\n#\n# Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:\n#\n#
\n#\n# The project is broken down into multiple steps:\n#\n# * Load and preprocess the image dataset\n# * Train the image classifier on your dataset\n# * Use the trained classifier to predict image content\n#\n# We'll lead you through each part which you'll implement in Python.\n#\n# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.\n#\n# First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.\n\n# +\n# Imports here\nimport numpy as np\nimport matplotlib.pyplot as plt\n\nimport torch\nimport torchvision\nfrom torchvision import datasets, transforms, models\nfrom torch import nn\nfrom torch import optim\n#import torch.nn.functional as F\nfrom torch.optim import lr_scheduler\nfrom torch.autograd import Variable\n\nimport copy\nimport time\nimport json\n\nfrom collections import OrderedDict\n\nfrom PIL import Image\nimport random\n# -\n\n# Set seed\nrandom.seed(2)\nnp.random.seed(2)\ntorch.manual_seed(2)\ntorch.cuda.manual_seed_all(2)\ntorch.backends.cudnn.deterministic=True\n\n# ## Load the data\n#\n# Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.\n#\n# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.\n#\n# The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.\n# \n\ndata_dir = 'flowers'\ntrain_dir = data_dir + '/train'\nvalid_dir = data_dir + '/valid'\ntest_dir = data_dir + '/test'\n\n# +\n# TODO: Define your transforms for the training, validation, and testing sets\ndata_transforms = {'train' : transforms.Compose([transforms.RandomRotation(30),\n transforms.RandomResizedCrop(224),\n transforms.RandomHorizontalFlip(),\n transforms.ToTensor(),\n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'valid' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])]),\n 'test' : transforms.Compose([transforms.Resize(256),\n transforms.CenterCrop(224),\n transforms.ToTensor(), \n transforms.Normalize([0.485, 0.456, 0.406], \n [0.229, 0.224, 0.225])])}\n\n# TODO: Load the datasets with ImageFolder\nimage_datasets = {'train' : datasets.ImageFolder(train_dir, transform=data_transforms['train']),\n 'valid' : datasets.ImageFolder(valid_dir, transform=data_transforms['valid']),\n 'test' : datasets.ImageFolder(test_dir, transform=data_transforms['test'])}\n\n# TODO: Using the image datasets and the trainforms, define the dataloaders\ndataloaders = {'train' : torch.utils.data.DataLoader(image_datasets['train'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'valid' : torch.utils.data.DataLoader(image_datasets['valid'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2)),\n 'test' : torch.utils.data.DataLoader(image_datasets['test'], batch_size=32, shuffle=True, num_workers=0, worker_init_fn=np.random.seed(2))}\n\ndataset_sizes = {'train': len(image_datasets['train']),\n 'valid': len(image_datasets['valid']),\n 'test': len(image_datasets['test'])} \n# -\n\n# ### Label mapping\n#\n# You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.\n\nwith open('cat_to_name.json', 'r') as f:\n cat_to_name = json.load(f)\n\n# # Building and training the classifier\n#\n# Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.\n#\n# We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students! You can also ask questions on the forums or join the instructors in office hours.\n#\n# Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:\n#\n# * Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)\n# * Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout\n# * Train the classifier layers using backpropagation using the pre-trained network to get the features\n# * Track the loss and accuracy on the validation set to determine the best hyperparameters\n#\n# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!\n#\n# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.\n\n# +\n# TODO: Build and train your network\n\n# +\n# default device for all functions with device argument: the fastest device available\n# all functions move models to device at the start and all returned models are moved to cpu at the end: \n# to avoid having problems with loading checkpoint when cuda is not available \ndevice = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\n\n# Defaults for functions to follow: architecture and parameters of the best_model\narchitecture = 'vgg16'\nlr = 0.0001\nhu = 4096\nstep_size = 5\n# Best num_epochs was 8 with lr=0.0001 but 10 with lr=0.001. \n# Keeping value at 10 is ok as train_model picks best number of epochs anyway.\n# We keep default num_epochs=10 in train.py as well. \nnum_epochs = 10 \n\n\n# -\n\ndef pick_arch(architecture):\n \"\"\"Pick from the following model architectures: vgg11, vgg13, vgg16, vgg19.\"\"\"\n if architecture == 'vgg11':\n model = models.vgg11(pretrained=True)\n elif architecture == 'vgg13':\n model = models.vgg13(pretrained=True)\n elif architecture == 'vgg16':\n model = models.vgg16(pretrained=True)\n elif architecture == 'vgg19':\n model = models.vgg19(pretrained=True)\n else:\n model = None\n print('Wrong model architecture. Pick vgg11, vgg13, vgg16 or vgg19.')\n \n return model\n\n\ndef build_model(architecture=architecture, hu=hu):\n \"\"\"Build a model from given architecture and replace classifier with a custom one.\"\"\"\n model = pick_arch(architecture)\n\n for param in model.parameters():\n param.requires_grad = False\n\n input_size = list(model.classifier.parameters())[0].size()[1]\n output_size = len(image_datasets['train'].class_to_idx)\n\n classifier = nn.Sequential(OrderedDict([\n ('fc1', nn.Linear(input_size, hu)),\n ('relu', nn.ReLU()),\n ('dropout', nn.Dropout(p=0.5)),\n ('fc2', nn.Linear(hu, output_size)),\n ('output', nn.LogSoftmax(dim=1))\n ]))\n\n model.classifier = classifier\n \n return model\n\n\ndef train_model(model=None, criterion=None, optimizer=None, scheduler=None,\n num_epochs=num_epochs, device=device):\n since = time.time()\n \n if model == None:\n model = build_model()\n \n if criterion == None:\n criterion = nn.NLLLoss()\n \n if optimizer == None:\n optimizer = optim.Adam(model.classifier.parameters(), lr)\n \n if scheduler == None:\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) \n \n model = model.to(device)\n \n best_model_wts = copy.deepcopy(model.state_dict())\n best_acc = 0.0\n best_num_epochs = 0\n\n for epoch in range(num_epochs):\n print('Epoch {}/{}'.format(epoch, num_epochs - 1))\n print('-' * 10)\n\n # Each epoch has a training and validation phase\n for phase in ['train', 'valid']:\n if phase == 'train':\n scheduler.step()\n model.train() # Set model to training mode\n else:\n model.eval() # Set model to evaluate mode\n\n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n # zero the parameter gradients\n optimizer.zero_grad()\n\n # forward\n # track history if only in train\n with torch.set_grad_enabled(phase == 'train'):\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # backward + optimize only if in training phase\n if phase == 'train':\n loss.backward()\n optimizer.step()\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n epoch_loss = running_loss / dataset_sizes[phase]\n epoch_acc = running_corrects.double() / dataset_sizes[phase]\n\n print('{} Loss: {:.4f} Acc: {:.4f}'.format(\n phase, epoch_loss, epoch_acc))\n\n # deep copy the model\n if phase == 'valid' and epoch_acc > best_acc:\n best_acc = epoch_acc\n best_model_wts = copy.deepcopy(model.state_dict())\n best_num_epochs = epoch + 1\n\n print('-' * 10)\n time_elapsed = time.time() - since\n print('Training complete in {:.0f}m {:.0f}s'.format(\n time_elapsed // 60, time_elapsed % 60))\n print('Best validation Accuracy of {:.4f} for {} epochs'.format(best_acc, best_num_epochs))\n print()\n\n # load best model weights\n model.load_state_dict(best_model_wts)\n model.to('cpu')\n return model, best_acc, best_num_epochs\n\n\n# The trial_model below was trained when default lr was set to 0.001. \n\ntrial_model, trial_acc, trial_num_epochs = train_model(num_epochs=2)\n\n\ndef tune_hyperparams(architecture=architecture, num_epochs=num_epochs,\n lr_list=[lr], hu_list=[hu], step_size_list=[step_size], device=device):\n param_triples = [(lr, hu, step_size) for lr in lr_list for hu in hu_list for step_size in step_size_list]\n \n best_model = pick_arch(architecture)\n best_acc = 0.0\n best_num_epochs = 0\n best_lr = 0.0\n best_hu = 0\n best_step_size = 0\n \n for lr, hu, step_size in param_triples:\n print('Training with')\n print('Learning rate: {} Number of hidden units: {} Step size: {}'.format(lr, hu, step_size))\n print('-' * 10)\n\n model = build_model(architecture=architecture, hu=hu)\n model = model.to(device)\n \n criterion = nn.NLLLoss()\n optimizer = optim.Adam(model.classifier.parameters(), lr=lr)\n scheduler = lr_scheduler.StepLR(optimizer, step_size, gamma=0.1) #add these to hyperparams\n \n model_trained, acc_trained, num_epochs_trained = train_model(model, criterion, optimizer, scheduler, num_epochs=num_epochs, device=device)\n \n # deep copy the model\n if acc_trained > best_acc:\n best_acc = acc_trained\n best_model = copy.deepcopy(model_trained)\n best_num_epochs = num_epochs_trained\n best_lr = lr\n best_hu = hu\n best_step_size = step_size\n \n print()\n print('Best results:')\n print('Learning Rate: {:.4f} Number of Hidden Units: {:.4f} Step size: {:.4f}'.format(best_lr, best_hu, best_step_size))\n print('Number of Epochs: {:.4f} Validation Acc: {:.4f}'.format(best_num_epochs, best_acc))\n best_model.to('cpu')\n return best_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size\n\n\ntune_hyperparams(num_epochs=2)\n\n# I tried learning rate of 0.01, number of hidden units 2048 and step size 7 but they did not do as well as lr=0.001, hu=4096 and step_size=5 and I decided to not spend more time training combinations with the above values. The last step in deciding on parameters is to see if learning rates of 0.0001 does better than 0.001. The train_model function already chooses the best number of epochs between 1 and 10. \n\nbest_model, best_acc, best_num_epochs, best_lr, best_hu, best_step_size = tune_hyperparams(lr_list=[0.0001, 0.001], hu_list=[4096], step_size_list=[5])\n\n\n# ## Testing your network\n#\n# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.\n\n# TODO: Do validation on the test set\ndef do_validation(model=None, criterion=None, phase='test', device=device):\n if model == None:\n model = build_model()\n if criterion == None:\n criterion = nn.NLLLoss()\n \n model.to(device)\n model.eval()\n \n running_loss = 0.0\n running_corrects = 0\n\n # Iterate over data.\n for inputs, labels in dataloaders[phase]:\n inputs = inputs.to(device)\n labels = labels.to(device)\n\n\n # forward\n # don't track history \n with torch.no_grad():\n outputs = model(inputs)\n _, preds = torch.max(outputs, 1)\n loss = criterion(outputs, labels)\n\n # statistics\n running_loss += loss.item() * inputs.size(0)\n running_corrects += torch.sum(preds == labels.data)\n\n total_loss = running_loss / dataset_sizes[phase]\n total_acc = running_corrects.double() / dataset_sizes[phase]\n \n model.to('cpu')\n \n print('Loss: {:.4f} Acc: {:.4f}'.format(\n total_loss, total_acc))\n return total_loss, total_acc.item()\n\ntotal_loss, total_acc = do_validation(model = best_model) \n\n# +\n#next(best_model.parameters()).is_cuda\n# -\n\n# ## Save the checkpoint\n#\n# Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.\n#\n# ```model.class_to_idx = image_datasets['train'].class_to_idx```\n#\n# Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.\n\n# TODO: Save the checkpoint \ncheckpoint = {'state_dict': best_model.state_dict(),\n 'hu': best_hu,\n 'architecture': architecture,\n 'num_epochs': best_num_epochs,\n 'lr': best_lr,\n 'step_size': best_step_size,\n 'class_to_idx': image_datasets['train'].class_to_idx}\n\ntorch.save(checkpoint, 'checkpoint_best.pth')\n\nbest_model.class_to_idx = image_datasets['train'].class_to_idx\n\n\n# ## Loading the checkpoint\n#\n# At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.\n\n# TODO: Write a function that loads a checkpoint and rebuilds the model\ndef load_checkpoint(filepath):\n checkpoint = torch.load(filepath)\n \n model = build_model(architecture=checkpoint['architecture'], hu=checkpoint['hu'])\n\n model.load_state_dict(checkpoint['state_dict'])\n model.class_to_idx = checkpoint['class_to_idx']\n \n return model, {'num_epochs':checkpoint['num_epochs'], \n 'lr': checkpoint['lr'], \n 'step_size': checkpoint['step_size'], \n 'hu': checkpoint['hu'], \n 'architecture': checkpoint['architecture']}\n\n\nreloaded_best_model, reloaded_best_params_dict = load_checkpoint('checkpoint_best.pth') \n\ndo_validation(model = reloaded_best_model)\n\n\n# # Inference for classification\n#\n# Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like \n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n#\n# First you'll need to handle processing the input image such that it can be used in your network. \n#\n# ## Image Preprocessing\n#\n# You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training. \n#\n# First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.\n#\n# Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.\n#\n# As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation. \n#\n# And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.\n\ndef process_image(image):\n ''' Scales, crops, and normalizes a PIL image for a PyTorch model,\n returns an Numpy array\n '''\n # TODO: Process a PIL image for use in a PyTorch model\n pil_image = Image.open(image)\n width, height = pil_image.size\n l = int(256 * max(width, height) / min(width, height))\n pil_image.thumbnail(size = (l,l))\n width, height = pil_image.size \n pil_image = pil_image.crop(box = (width / 2 - 112, height / 2 - 112, width / 2 + 112, height / 2 + 112))\n \n np_image = np.array(pil_image)\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n np_image = (np_image / 255.0 - mean) / std\n \n np_image = np_image.transpose((2, 0, 1))\n \n return np_image\n\n\n# To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).\n\ndef imshow(image, ax=None, title=None):\n if ax is None:\n fig, ax = plt.subplots()\n \n # PyTorch tensors assume the color channel is the first dimension\n # but matplotlib assumes is the third dimension\n image = image.transpose((1, 2, 0))\n \n # Undo preprocessing\n mean = np.array([0.485, 0.456, 0.406])\n std = np.array([0.229, 0.224, 0.225])\n image = std * image + mean\n \n # Image needs to be clipped between 0 and 1 or it looks like noise when displayed\n image = np.clip(image, 0, 1)\n \n ax.imshow(image)\n \n return ax\n\n\nimshow(process_image('flowers/test/1/image_06743.jpg'))\n\n\n# ## Class Prediction\n#\n# Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.\n#\n# To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.\n#\n# Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.\n#\n# ```python\n# probs, classes = predict(image_path, model)\n# print(probs)\n# print(classes)\n# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]\n# > ['70', '3', '45', '62', '55']\n# ```\n\n# +\ndef predict(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\ndef predict2(image_path, checkpoint, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model from checkpoint.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model, _ = load_checkpoint(checkpoint)\n model.to(device)\n model.eval()\n image = process_image(image_path)\n img_tensor = torch.from_numpy(image).type(torch.FloatTensor)\n img_tensor.unsqueeze_(0)\n image_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(image_variable) \n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes\n\n\n\n# small change in processing image: use transforms instead of process_image function\ndef predict3(image_path, model, topk=5, device=device):\n ''' Predict the class (or classes) of an image using a trained deep learning model.\n '''\n \n # TODO: Implement the code to predict the class from an image file\n model.to(device)\n model.eval()\n transfs = data_transforms['test']\n\n img_pil = Image.open(image_path)\n img_tensor = transfs(img_pil)\n img_tensor.unsqueeze_(0)\n img_variable = Variable(img_tensor)\n image_variable = image_variable.to(device)\n with torch.no_grad():\n outputs = model(img_variable)\n \n probs, classes = outputs.topk(topk) \n \n probs = list(np.exp(np.asarray(probs).flatten()))\n classes = list(np.asarray(classes).flatten())\n \n class_to_idx = model.class_to_idx\n idx_to_class = {v: k for k, v in class_to_idx.items()}\n \n classes = [idx_to_class[k] for k in classes]\n model.to('cpu')\n return probs, classes \n\n\n# -\n\nprobs, classes = predict('flowers/test/1/image_06743.jpg', reloaded_best_model)\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\nprobs, classes = predict2('flowers/test/1/image_06743.jpg', 'checkpoint_best.pth')\nprint([cat_to_name[k] for k in classes])\nprint(classes)\nprint(probs)\n\n\n# ## Sanity Checking\n#\n# Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:\n#\n#  \n#\n# You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.\n\n# +\n# TODO: Display an image along with the top 5 classes\ndef plot_im_probs_classes(image_path, model, device=device):\n probs, classes = predict(image_path, model, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout()\n \ndef plot_im_probs_classes2(image_path, checkpoint, device=device):\n probs, classes = predict2(image_path, checkpoint, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout() \n\n\n# -\n\nplot_im_probs_classes('flowers/test/1/image_06760.jpg', model=reloaded_best_model, device=device)\n\nplot_im_probs_classes2('flowers/test/1/image_06760.jpg', 'checkpoint_best.pth', device=device)\n\n# Resources used for this project:\n# * Material from the first term of Udacity's Data Scientist Nanodegree program. \n# * [Transfer Learning tutorial](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html). `train_model` function was taken from this tutorial with small changes as well as other hints such as shuffling the datasets and using a scheduler. \n# * [PyTorch documentation](https://pytorch.org/docs/stable/index.html).\n# * [Argparse tutorial](https://pymotw.com/3/argparse/#argument-types) and [documentation](https://docs.python.org/dev/library/argparse.html#argparse.Namespace).\n","repo_name":"anamamatelashvili/deep_learning_project","sub_path":"Image Classifier Project.ipynb","file_name":"Image Classifier Project.ipynb","file_ext":"py","file_size_in_byte":31574,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"581944992","text":"# + [markdown] id=\"E-aBmrp8931c\"\n# # Proyecto empresa de viajes compartidos\n\n# + [markdown] id=\"uEJABzxG931c\"\n# ### Descripción del proyecto\n#\n# Durante este proyecto analizaré datos para una nueva empresa de viajes compartidos que se esta lanzando en Chicago. El objetivo será encontrar patrones en los datos y entender las preferencias de los pasajeros y el impacto de los factores externos en los viajes.\n#\n# #### Descripción de los datos\n#\n# Tabla 1: contiene datos sobre la cantidad de viajes que realizó cada compañía.\n# - company_name: nombre de la empresa de taxis\n# - trips_amount: el número de viajes de cada compañía de taxis el 15 y 16 de noviembre de 2017.\n#\n# Tabla 2: contiene datos sobre la finalización de los viajes \n# - dropoff_location_name: barrios de Chicago donde finalizaron los viajes\n# - average_trips: el promedio de viajes que terminaron en cada barrio en noviembre de 2017.\n#\n# Tabla 3:\n# Contiene datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. \n# - start_ts: fecha y hora de la recogida\n# - weather_conditions: condiciones climáticas en el momento en el que comenzó el viaje\n# - duration_seconds: duración del viaje en segundos\n\n# + [markdown] id=\"1G-R9FNe931c\"\n# ### Análisis exploratorio de datos\n\n# + id=\"PZCA0wo3931c\"\n# Importación de librerías a utilizar\n\nimport pandas as pd\nfrom scipy import stats as st\nfrom scipy.stats import levene\nimport matplotlib.pyplot as plt\nfrom seaborn import boxplot\n\n# + id=\"r3wEulH9931d\" outputId=\"1375e687-549f-4792-8c56-1ebfe3ff3c0f\"\n# Importación de archivo 1: número de viajes por compañia\n\ncompanies_trips= pd.read_csv('/datasets/project_sql_result_01.csv')\ncompanies_trips.info()\ncompanies_trips.head()\n\n\n# + id=\"MA1CVDmo931d\" outputId=\"0d2dabbc-9c46-4fb2-e35a-d36d79ab3335\"\n# Chequeo de valores ausentes\n\ncompanies_trips.isna().sum()\n\n# + id=\"UVq55dyy931d\" outputId=\"9735fac2-c40f-4ee0-8c3e-73d4238f734d\"\n# Chequeo de valores duplicados\n\ncompanies_trips.duplicated().sum()\n\n# + id=\"Bio8N7b7931d\" outputId=\"01191f13-de81-4e3f-bc49-83e4a9496fa8\"\n# Importación de archivo 2: promedio de viajes por barrio \n\ndropoff_location= pd.read_csv('/datasets/project_sql_result_04.csv')\ndropoff_location.info()\ndropoff_location.head()\n\n# + id=\"VfGtzMP-931e\" outputId=\"c6356ec7-0873-41da-c118-50a4d1e55f63\"\n# Chequeo de valores ausentes \n\ndropoff_location.isna().sum()\n\n# + id=\"bYD1uV96931e\" outputId=\"1d75298f-3319-4eb2-818a-4e3f4d0a2a96\"\n# Chequeo de valores duplicados\n\ndropoff_location.duplicated().sum()\n\n# + id=\"8aHq5XT-931e\" outputId=\"384087f9-8cea-4f88-e3a1-f5c338d36c98\"\n# Identificar los 10 principales barrios en términos de finalización del recorrido\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10)\n\n# + id=\"LdV71Unp931e\" outputId=\"5da0eaf9-c0e7-4ef7-f5c7-d51006a0a7ba\"\n#Gráfico: empresas de taxis y número de viajes\n\ncompanies_trips.plot(kind='bar',x='company_name', y='trips_amount', ylabel='trips_amount', figsize=(20,10), title='Viajes por empresa de taxi')\n\n# + [markdown] id=\"shnqBODc931e\"\n# Como podemos observar en el gráfico de arriba, la empresa de taxis más usada los días 15 y 16 de noviembre fue Flash Cab. Ésta realizó casi el doble de viajes que la empresa que le sigue: Taxi Affiliation Services. \n#\n# A su vez, dejando de lado Flash Cab que fue sin dudas la más poplar, podemos observar que hay pequeños grupos de compañías en los cuales no difiere tanto la cantidad de viajes realizados. Por ejemplo, las otras 5 copañías principales de taxis realizaron entre 9180 a 11400 viajes.\n#\n# Por otro lado, podemos observar que la mitad de las compañías de taxis tuvieron muy pocos viajes esos días. \n\n# + id=\"bzjPAp8R931e\" outputId=\"4031eafb-cf1d-4fc0-d3f1-73c5e893dc32\"\n# Gráfico 10 barrios principales por número de finalizaciones\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10).plot(kind='bar',x='dropoff_location_name', y='average_trips', ylabel='Trips amount', figsize=(20,10), title='Principales barrios')\n\n# + [markdown] id=\"Hl0_zDu_931f\"\n# Como podemos observar en el gráfico de arriba, se muestran los 10 barrios principales hacia los cuales se dirigieron los taxis (destino final). El barrio con más viajes fue Loop con un promedio de 10727 viajes en el mes de noviembre seguido por River North con un promedio de 9523 viajes.\n#\n# Después se encuentran Streeterville\ty West Loop con un promedio de viajes de 6664 y 5163 respectivamente y por último un tercer grupo en la tabla con barrios con un promedio de viajes entre 1260 a 2420 en el mes de noviembre. \n\n# + [markdown] id=\"ibJP9mHu931f\"\n# ### Prueba de hipótesis\n\n# + [markdown] id=\"9qceopHk931f\"\n# A continuación voy a importar el archivo con los datos que usare para hacer la prueba de hipótesis.\n\n# + id=\"Mj53hprj931f\" outputId=\"216d861a-a20e-40c7-bfca-172a9615cafb\"\n# Importación de archivo para trabajar sobre la hipótesis\n\narchivo_hipotesis = pd.read_csv('/datasets/project_sql_result_07.csv')\narchivo_hipotesis.info()\narchivo_hipotesis.head()\n\n# + id=\"nT7L8FIk931f\" outputId=\"04025974-4c5f-4a5f-84e1-3c46cf2c3852\"\n# Chequeo de valores ausentes\n\narchivo_hipotesis.isna().sum()\n\n# + id=\"ZqMuqgQS931f\" outputId=\"9b37c186-3558-4c3a-b90c-d3e53abca6f5\"\n# Chequeo de valores duplicados\n\narchivo_hipotesis.duplicated().sum()\n\n# + id=\"2gIJlz-9931f\" outputId=\"7e2315d2-3169-427a-c706-8443f22436e4\"\narchivo_hipotesis.value_counts()\n\n# + [markdown] id=\"aygBH6hT931f\"\n# Como podemos observar, en el archivo que nos dieron para hacer la prueba de hipótesis tenemos valores duplicados. Estos pueden deberse a que el archivo contiene los datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. Puede que haya determinadas paradas en Loop con destino hacia el aeropuerto. Al ser un punto fijo es posible que en condiciones normales, el recorrido dure siempre lo mismo; todos los los viajes que salgan de ahi a una determnada hora van a tardar lo mismo generando que haya varios datos con os mismos tiempos.\n\n# + id=\"fb9UZ9-Q931f\" outputId=\"128662bf-d6f4-403b-9c56-d506f9680a7d\"\nboxplot(data= archivo_hipotesis, x=\"duration_seconds\", y=\"weather_conditions\")\n\n# + [markdown] id=\"OxVmL-Y2931g\"\n# Voy a probar la hipótesis de que la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare cambia los sábados lluviosos.\n#\n# Hipótesis nula: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos. \n#\n# Hipótesis alternativa: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare difiere los sábados lluviosos.\n#\n# Previo a la prueba de hipótesis voy a realizar la prueba de levene para ver si las varianzas son iguales. Usaré como hipótesis nula: las varianzas son iguales y como hipótesis alternativa las varianzas no son iguales.\n\n# + id=\"3sY6DdYD931g\" outputId=\"a465661a-30aa-4dd3-92e3-eb7b9dee7791\"\n# Prueba de hipotesis nula levene: varianzas son iguales \n\ndatos_lluvia= archivo_hipotesis.query('weather_conditions == \"Bad\" and duration_seconds >0')\ndatos_good= archivo_hipotesis.query('weather_conditions ==\"Good\"and duration_seconds >0')\n\nlluvia= datos_lluvia['duration_seconds']\ngood= datos_good['duration_seconds']\n\nalpha= 0.05\n\nstat,results= levene(lluvia, good)\nprint('P_value:', results)\n\nif results < alpha:\n print('Rechazo la hipótesis nula')\nelse:\n print('No rechazo la hipótesis nula')\n\n# + [markdown] id=\"huB3ekYd931g\"\n# Como podemos observar de la prueba de Levene realizada arriba, no rechazamos la hipótesis nula que indica que las varianzas son iguales. \n#\n# A continuación realizare la prueba de hipótesis.\n\n# + id=\"ABv0Xklx931g\" outputId=\"7198fbf1-68b8-4240-bb38-b4164b0464f5\"\n# Ho: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos.\n\nalpha= 0.05\nresults= st.ttest_ind(lluvia,good, equal_var= True)\nprint('P_value:', results.pvalue)\n\nif results.pvalue < alpha:\n print('Rechazo la hipótesis nula')\nelse: \n print('No rechazo hipótesis nula')\n\n# + [markdown] id=\"Ns2N_ZTw931g\"\n# De acuerdo con los datos con los que contamos, rechazo la hipótesis nula. La duración promedio de los viajes desde el Loop hasta el aeropuerto de O'Hare cambia los sábados lluviosos.\n\n# + [markdown] id=\"dtnLXksI931g\"\n# ## Conclusión \n#\n# Para concluir, podemos decir que la empresa de taxis más utilizada es Flash Cab seguida por Taxi Affiliation Services, teniendo la primera casi el doble de viajes que la segunda. Esta diferencia puede deberse a que Flash Cab sea una empresa mucho más grande que las otras; al tener una flota de autos más grande en la calle le permite tener el doble de viajes que su competidor más cercano. \n#\n# Con respecto a los detinos finales de los viajes, Loop resulto el más concurrido seguido de cerca por River North.\n#\n# Por último la duración promedio de los viajes difiere un día lluvioso en comparación a un día de buen tiempo. Esta diferencia puede deberse principalmente al factor climático. Los días de mal tiempo puede que haya más tránsito porque la gente decide salir con sus autos y se genera más congestión. También la gente anda un poco más despacio impactando en la duración pormedio de los viajes. \n","repo_name":"CarmenPujato/practicum","sub_path":"empresa de viajes compartidos/Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_name":"Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_ext":"py","file_size_in_byte":9307,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43095479709","text":"# # Kaggle class competition\n# # Deep Learning and Neural Network Models to predict traffic volume\n# # Models: LSTM, GRUs\n# # Library used: Tensorflow, Keras\n\n# ### Set Up Libraries\n\n# +\n# ns - as typically happens, outdated components like as keywords, functions, and classes trigger a warning.\nimport warnings\nwarnings.filterwarnings('ignore')\n# ns - below are the neccessary packages utilized in this project\nimport os\nimport datetime\n\nimport IPython\nimport IPython.display\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns\nimport tensorflow as tf\n\nmpl.rcParams['figure.figsize'] = (8, 6)\nmpl.rcParams['axes.grid'] = False\n# -\n\n# ### Import data\n\n# ns - use pandas package to read in data from file located within folder \ndata = pd.read_csv('Metro_Interstate_Traffic_Volume.csv', infer_datetime_format=True)\ndata.head()\n\n# ### Inspect and Clean up\n\n# ns - investigate the data info\ndata.info()\n\n# #### It could be seen that there are both str and num types of data as categorized below\n# categorical features: holiday, weather_main, weather_description\n#\n# Continious features: temp, rain_1h, show_1h, clouds_all\n#\n# Target: traffic_volume\n\n# ### Convert date_time column to datetime type\n#\n# It is usable if we don't convert string type to datetime type in this project. It is due to the fact that time series will absolutely need datetime. So below is the demonstration.\n\n# ns - datetime type conversion\ndate_time = pd.to_datetime(data.pop('date_time'), format='%Y-%m-%d %H:%M:%S')\n\n# ### Convert to a usable signal \n#\n# In this form of signal, we would be able to utilize its capacity in the ultimate way as it reveals more insignhts as illjustrated below. \n\ntimestamp_s = date_time.map(datetime.datetime.timestamp)\n\n# +\nday = 24*60*60\nyear = (365.2425)*day\n\ndata['Day sin'] = np.sin(timestamp_s * (2 * np.pi / day))\ndata['Day cos'] = np.cos(timestamp_s * (2 * np.pi / day))\ndata['Year sin'] = np.sin(timestamp_s * (2 * np.pi / year))\ndata['Year cos'] = np.cos(timestamp_s * (2 * np.pi / year))\n# -\n\nplt.plot(np.array(data['Day sin'])[:25])\nplt.plot(np.array(data['Day cos'])[:25])\nplt.xlabel('Time [h]')\nplt.title('Time of day signal')\n\n# The best way to identify which frequencies are most essential is to use a \"fft\". As a preliminary test, below is the traffic volume rfft experiment. While we're discussing common frequencies, it's interesting to note the highs at 1/year and 1/day frequencies:\n\n# +\nfft = tf.signal.rfft(data['temp'])\nf_per_dataset = np.arange(0, len(fft))\n\nn_samples_h = len(data['temp'])\nhours_per_year = 24*365.2524\nyears_per_dataset = n_samples_h/(hours_per_year)\n\nf_per_year = f_per_dataset/years_per_dataset\nplt.step(f_per_year, np.abs(fft))\nplt.xscale('log')\nplt.ylim(0, 400000)\nplt.xlim([0.1, max(plt.xlim())])\nplt.xticks([1, 365.2524], labels=['1/Year', '1/day'])\n_ = plt.xlabel('Frequency (log scale)')\n# -\n\n# ### Drop variable \"weather_description\" and convert \"holiday\" and \"weather_main\" to dummies variables\n#\n# It is assumed that the weather description might be more irrelevant to this project or its interpretation could benefit less. Thus, I have decided to drop it. While both weather main and the holiday are deemed significant.\n\n# ns - considered to drop some variable and generate dummies variables for the rest of obj type\ndata.drop('weather_description', axis=1, inplace=True)\ndata = pd.get_dummies(data, columns = ['holiday', 'weather_main'], drop_first=True)\ndata.head()\n\n# ### Split the data\n\n# +\n# ns - as indicated in the instruction, the test set should be excluded with the size of 5000 ea.\n# ns - the slicing method is used in this data split\ncolumn_indices = {name: i for i, name in enumerate(data.columns)}\n\nn = len(data)\nprint(\"There are \", n, \"records and divisions are as below\")\n\ntest_df = data[-5000:]\nprint(\"Test set: \", test_df.shape)\n\nval_df = data[-13000:-5000]\nprint(\"Validation set: \", val_df.shape)\n\ntrain_df = data[0:-13000]\nprint(\"Training set: \", train_df.shape)\n\nnum_features = data.shape[1]\n# -\n\n# ### Data Normalization\n\n# +\ntrain_mean = train_df.mean()\ntrain_std = train_df.std()\n\ntrain_df = (train_df - train_mean) / train_std\nval_df = (val_df - train_mean) / train_std\ntest_df = (test_df - train_mean) / train_std\n\n\n# -\n\n# ### Data windowing\n#\n# The models in this project will make a set of predictions based on a window of consecutive samples from the data and below is the useful windowgenerator class for segmenting data as required for modelling.\n\nclass WindowGenerator():\n def __init__(self, input_width, label_width, shift,\n train_df=train_df, val_df=val_df, test_df=test_df,\n label_columns=None):\n # Store the raw data.\n self.train_df = train_df\n self.val_df = val_df\n self.test_df = test_df\n\n # Work out the label column indices.\n self.label_columns = label_columns\n if label_columns is not None:\n self.label_columns_indices = {name: i for i, name in\n enumerate(label_columns)}\n self.column_indices = {name: i for i, name in\n enumerate(train_df.columns)}\n\n # Work out the window parameters.\n self.input_width = input_width\n self.label_width = label_width\n self.shift = shift\n\n self.total_window_size = input_width + shift\n\n self.input_slice = slice(0, input_width)\n self.input_indices = np.arange(self.total_window_size)[self.input_slice]\n\n self.label_start = self.total_window_size - self.label_width\n self.labels_slice = slice(self.label_start, None)\n self.label_indices = np.arange(self.total_window_size)[self.labels_slice]\n\n def __repr__(self):\n return '\\n'.join([\n f'Total window size: {self.total_window_size}',\n f'Input indices: {self.input_indices}',\n f'Label indices: {self.label_indices}',\n f'Label column name(s): {self.label_columns}'])\n\n\n# As per required, the windowgenerator here will provide 6 width with 2 future prediction and 1 label.\n\nw2 = WindowGenerator(input_width=6, label_width=1, shift=2,\n label_columns=['traffic_volume'])\nw2\n\n\n# +\ndef split_window(self, features):\n inputs = features[:, self.input_slice, :]\n labels = features[:, self.labels_slice, :]\n if self.label_columns is not None:\n labels = tf.stack(\n [labels[:, :, self.column_indices[name]] for name in self.label_columns],\n axis=-1)\n\n # Slicing doesn't preserve static shape information, so set the shapes\n # manually. This way the `tf.data.Datasets` are easier to inspect.\n inputs.set_shape([None, self.input_width, None])\n labels.set_shape([None, self.label_width, None])\n\n return inputs, labels\n\nWindowGenerator.split_window = split_window\n# -\n\n# ### Note: \n#\n# Typically data in TensorFlow is packed into arrays where the outermost index is across examples (the \"batch\" dimension). The middle indices are the \"time\" or \"space\" (width, height) dimension(s). The innermost indices are the features.\n\n# +\n# ns - I tried with other stack but it failed to corporate with the model\n# Stack three slices, the length of the total window:\nexample_window = tf.stack([np.array(train_df[:w2.total_window_size]),\n np.array(train_df[100:100+w2.total_window_size]),\n np.array(train_df[200:200+w2.total_window_size])])\n\n\nexample_inputs, example_labels = w2.split_window(example_window)\n\nprint('All shapes are: (batch, time, features)')\nprint(f'Window shape: {example_window.shape}')\nprint(f'Inputs shape: {example_inputs.shape}')\nprint(f'labels shape: {example_labels.shape}')\n# -\n\n# ### Plot \n#\n# #### Here is a plot method that allows a simple visualization of the split window\n\nw2.example = example_inputs, example_labels\n\n\n# +\ndef plot(self, model=None, plot_col='traffic_volume', max_subplots=3):\n inputs, labels = self.example\n plt.figure(figsize=(12, 8))\n plot_col_index = self.column_indices[plot_col]\n max_n = min(max_subplots, len(inputs))\n for n in range(max_n):\n plt.subplot(max_n, 1, n+1)\n plt.ylabel(f'{plot_col} [normed]')\n plt.plot(self.input_indices, inputs[n, :, plot_col_index],\n label='Inputs', marker='.', zorder=-10)\n\n if self.label_columns:\n label_col_index = self.label_columns_indices.get(plot_col, None)\n else:\n label_col_index = plot_col_index\n\n if label_col_index is None:\n continue\n\n plt.scatter(self.label_indices, labels[n, :, label_col_index],\n edgecolors='k', label='Labels', c='#2ca02c', s=64)\n if model is not None:\n predictions = model(inputs)\n plt.scatter(self.label_indices, predictions[n, :, label_col_index],\n marker='X', edgecolors='k', label='Predictions',\n c='#ff7f0e', s=64)\n\n if n == 0:\n plt.legend()\n\n plt.xlabel('Time [h]')\n\nWindowGenerator.plot = plot\n# -\n\nw2.plot()\n\n\n# The plots above illustrate the 6 inputs and 1 label as we defined them ealier. \n\n# ### Create tf.data.Datasets\n\n# +\ndef make_dataset(self, data):\n data = np.array(data, dtype=np.float32)\n ds = tf.keras.preprocessing.timeseries_dataset_from_array(\n data=data,\n targets=None,\n sequence_length=self.total_window_size,\n sequence_stride=1,\n shuffle=False,\n batch_size=32,)\n\n ds = ds.map(self.split_window)\n\n return ds\n\nWindowGenerator.make_dataset = make_dataset\n\n\n# -\n\n# ### Note:\n#\n# This make_dataset method will take a time series DataFrame and convert it to a tf.data.Dataset of (input_window, label_window) pairs using the preprocessing.timeseries_dataset_from_array function. The WindowGenerator object holds training, validation and test data. Add properties for accessing them as tf.data.Datasets using the above make_dataset method.\n\n# +\n@property\ndef train(self):\n return self.make_dataset(self.train_df)\n\n@property\ndef val(self):\n return self.make_dataset(self.val_df)\n\n@property\ndef test(self):\n return self.make_dataset(self.test_df)\n\n@property\ndef example(self):\n \"\"\"Get and cache an example batch of `inputs, labels` for plotting.\"\"\"\n result = getattr(self, '_example', None)\n if result is None:\n # No example batch was found, so get one from the `.train` dataset\n result = next(iter(self.train))\n # And cache it for next time\n self._example = result\n return result\n\nWindowGenerator.train = train\nWindowGenerator.val = val\nWindowGenerator.test = test\nWindowGenerator.example = example\n# -\n\n# Each element is an (inputs, label) pair\nw2.train.element_spec\n\n# #### Iterating over a Dataset yields concrete batches:\n\nfor example_inputs, example_labels in w2.train.take(1):\n print(f'Inputs shape (batch, time, features): {example_inputs.shape}')\n print(f'Labels shape (batch, time, features): {example_labels.shape}')\n\n# #### This project trains many models, so package the training procedure into a function\n\n# +\nMAX_EPOCHS = 100\n\ndef compile_and_fit(model, window, patience=2):\n early_stopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss',\n patience=patience,\n mode='min')\n\n model.compile(loss=tf.losses.MeanSquaredError(),\n optimizer=tf.optimizers.Adam(),\n metrics=[tf.metrics.MeanAbsoluteError()])\n\n history = model.fit(window.train, epochs=MAX_EPOCHS,\n validation_data=window.val,\n callbacks=[early_stopping])\n return history\n\n\n# -\n\n# For the multi-step model, the training data consists of hourly samples. However, here, the models will learn to predict 2h of the future, given 6h of the past. Here is a Window object that generates these slices from the dataset.\n\n# +\n# ns - indicating the 2 hours future output from the 6 hours history input\nOUT_STEPS = 2\nmulti_window = WindowGenerator(input_width=6,\n label_width=OUT_STEPS,\n shift=OUT_STEPS)\n\nmulti_window.plot()\nmulti_window\n# -\n\n# Illustration here displays the 6 inputs with 2 hours prediction in the future.\n\n# ### Model Construction\n# #### (1) The Base LSTM model\n#\n# This model is expected to be used as the baseline model in this project. The error is aimed to be minimized in further steps. The architecture of this base model consists of LSTM layer with 32 unit, Dense layer and then reshape respectively.\n\n# +\nmulti_val_performance = {}\nmulti_performance = {}\n\nmulti_lstm_model = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.val)\nmulti_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model)\n# -\n\n# #### (2) The Base LSTM with parameter Recurrent Dropout (0.1) and dropout layer (0.1)\n#\n# In this model, the recurrent Dropout with value 0.1 is added into the base model. It is expected to improve the mean absolute error. Recurrent Dropout is a regularization method for recurrent neural networks. Dropout is applied to the updates to LSTM memory cells (or GRU states), i.e. it drops out the input/update gate in LSTM/GRU.\n\n# +\n# ns - model with first implementation based on the base \nmulti_lstm_model2 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model2, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model2)\n# -\n\n# #### (3) The Base LSTM with parameter Recurrent Dropout (0.1) + unit_forget_bias, and dropout layer (0.1)\n#\n# In this model the unit forget bias is added for the experiment. It is hoped that If True, it will add 1 to the bias of the forget gate at initialization. Setting it to true will also force bias_initializer=\"zeros\". The result looks better than the prev ious models. Nevertheless, it is so varying in each run but it seems to capture the minimal MAE.\n\n# +\n# ns - model with second implementation based on the previous model \nmulti_lstm_model3 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model3, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model3)\n# -\n\n# #### Additional Model\n# This is an additional model which I tried to tune after I have completed my report. The result is very impressive, so I keep it on this notebook so that I can make the improvement later when I have time. I tried to add the use_bias parameter to the Dense layer. It enables bias vector in this experiment. I want to utilize Gridsearch to this LSTM but time is running out. Therefore, I end up with this best model in terms of lowest MAE of 0.1724.\n\n# +\n# ns - this is a last minute trial model but it appears efficient\nmulti_lstm_model9 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1), \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model9, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model9)\n# -\n\n# #### (4) The GRU Base Model with recurrent_dropout=0.1\n#\n# GRU is a recurrent neural network (RNN) in the most recent generation, and it is also equivalent to LSTM. In all likelihood, the newer GRU model will outperform the older LSTM model. I started with a simple 32-unit network and a 0.1 recurrence dropout rate.\n\n# +\n# ns - this model is assigned as the GRU base model\nmulti_lstm_model4 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.1, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model4, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model4)\n# -\n\n# #### (5) The GRU Base Model with recurrent_dropout=0.3\n\n# +\n# ns - model with first implementation based on the GRU base \nmulti_lstm_model5 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model5, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model5)\n# -\n\n# #### (6) The GRU Base Model with recurrent_dropout=0.3 and dropout layer (0.1)\n#\n# In addition to the recurrent dropout parameter, this experiment includes the dropout layer with 0.1 figure which previously demonstrated the better version among dropout varying. The outcome below is quite as anticipated. the MAE becomes better. And again the results are very varying in each run.\n\n# +\n# ns - model with second implementation based on the GRU base \nmulti_lstm_model6 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model6, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model6)\n# -\n\n# #### (7) The Bidirectional Model with LSTM\n#\n# I proceed with the selection of Bidirectional technique which I wanted to experimenrt with both LSTM and GRU models. It is started with the Bidirectional LSTM in this trial. I believe that allowing the LSTM model to learn the input sequence both forward and backward, and then concatenating both interpretations, can be useful on some sequence prediction challenges. the result looks great but it is not good as I predicted. In each run, it shows unsatisfied value of 1.7\n\n# +\n# ns - this model is built to try on Bidirectional with LSTM layer\nfrom keras.layers import Bidirectional\nfrom keras.layers import LSTM\n\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, \n return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### (8) The Bidirectional Model with GRU\n#\n# The best model of GRU is selected to be the layer of this Bidirectional method. As mentioned ealier, this result also exibits some fluctuation and not high enough as expected.\n\n# +\n# ns - this model is built to try on Bidirectional with GRU layer\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.GRU(32, activation='tanh', \n recurrent_activation='hard_sigmoid', \n recurrent_dropout=0.1, return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### Additional stacked Bidirectional model\n# This implementation is to show that the more layer I have added into the model the more complex the model would be. This complication diminishes the ability of model. As a result, this experiment proved to be quite weak in comparison with the value of MAE.\n\n# +\n# ns - this model is built to try on Bidirectional with stacked layer\nmulti_lstm_model10 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, return_sequences=True)),\n tf.keras.layers.Dropout(0.1), \n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(8)),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model10, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model10)\n# -\n\n# ### Model Prediction and Selection\n\n# ns - The best MAE is 0.1724 which is from model#9 \n# ns - Note that every time the kernel is reran, the MAE is varied\nprediction = multi_lstm_model9.predict(multi_window.test)\n# ns - The shape of prediction\nprediction.shape\n\n# ns - Extract the traffic volum column which is number 4\npredictedValue = prediction[:,1,4]\npredictedValue\n\n# #### Denormalization\n\n# ns - Extracting the std and mean values\nstd = train_std[4]\nmean = train_mean[4]\nprint(\"std: \", std, \"mean: \", mean)\n\n# ns - Performing data conversion to denormalized values\ndenorm = []\nfor i in predictedValue:\n val = (i*std)+mean\n denorm.append(val)\n\n# #### Export data to CSV file\n\n# ns - dataframe to export raw output to csv file\nimport pandas as pd\ndf = pd.DataFrame(denorm, columns = ['prediction'])\ndf.insert(0, 'id', range(1,4994))\ndf.to_csv(\"model9.csv\", index = False)\n\n# # Thank you!\n\n\n","repo_name":"iamnatapong55/Deep-Learning-Neural-Network","sub_path":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_name":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_ext":"py","file_size_in_byte":26193,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"44313882936","text":"# +\nimport numpy as np\nimport pandas as pd\nfrom matplotlib import pyplot as plt\n\nnp.set_printoptions(suppress=True, precision=3)\n\n\n# + [markdown] heading_collapsed=true\n# # Show Graph\n\n# + hidden=true\ndef ShowGraph(a, b, X, Y, line=\"a+bx\"):\n line_x = np.arange(np.amin(X), np.amax(X), 0.01)\n if line == \"a+bx\":\n line_y = a + b * line_x\n elif line == \"ax^b\":\n line_y = a * line_x ** b\n elif line == \"ae^(bx)\":\n line_y = a * np.exp(b*line_x)\n\n plt.clf()\n plt.plot(line_x, line_y, '-r')\n plt.plot(X, Y, 'ob')\n plt.grid()\n plt.show()\n\n\n# + [markdown] heading_collapsed=true\n# # Error\n\n# + [markdown] heading_collapsed=true hidden=true\n# ## Least Square Error\n\n# + hidden=true\ndef LeastSquareError(a, b, xi, yi):\n Dt = np.sum((yi - np.average(yi)) ** 2)\n D = np.sum((yi - a - b*xi) ** 2)\n \n r = ((Dt-D)/Dt) ** 0.5\n \n return r\n\n\n# + [markdown] hidden=true\n# ## Correlation Coeficient\n\n# + hidden=true\ndef CorrelationCoef(X, Y):\n difX = X - np.average(X)\n difY = Y - np.average(Y)\n \n r = (np.sum(difX*difY))/((np.sum((difX)**2)*np.sum((difY)**2))**(0.5))\n \n return r;\n\n\n# + [markdown] heading_collapsed=true\n# # Linear Regression → y = a + bx\n\n# + hidden=true\ndef LinearRegression(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'xiyi': X_data*Y_data,\n 'xi2': X_data*X_data\n })\n\n x_bar = np.average(data['xi'])\n y_bar = np.average(data['yi'])\n\n b = (n * np.sum(data['xiyi']) - np.sum(data['xi']) *\n np.sum(data['yi']))/(n * np.sum(data['xi2']) - np.sum(data['xi'])**2)\n a = y_bar - b * x_bar\n\n if showGraph:\n ShowGraph(a, b, data['xi'], data['yi'])\n\n if showData:\n print(\"linear regression data\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([4, 6, 8, 10, 14, 16, 20, 22, 24, 28])\nY_data = np.array([30, 18, 22, 28, 14, 22, 16, 8, 20, 8])\n\na, b = LinearRegression(X_data, Y_data, showGraph=True, showData=False)\nprint(\"y = a + bx\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] heading_collapsed=true\n# # Non-Linear Regression\n\n# + [markdown] hidden=true\n# ## y = ax^b\n\n# + hidden=true\ndef NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': np.log10(X_data),\n 'qi': np.log10(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = 10 ** a\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ax^b\")\n\n if showData:\n print(\"non-linear regression data (y = ax^b)\")\n print(\"pi = log(xi)\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ax^b\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] hidden=true\n# ## y = ae^(bx)\n\n# + hidden=true\ndef NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': X_data,\n 'qi': np.log(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = np.exp(a)\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ae^(bx)\")\n\n if showData:\n print(\"non-linear regression data (y = ae^(bx))\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ae^(bx)\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# -\n\n# # Polinomial Regression\n\ndef GaussJordan(A, B, printStep=False):\n size = B.size\n matrix = A\n matrix = np.concatenate((A, B.reshape(size, 1)), axis=1)\n \n for i in range(0, size):\n if printStep:\n print(\"step\", i+1, \"\\n\")\n \n matrix[i] /= matrix[i, i]\n\n for j in range(0, size):\n if i == j:\n continue\n \n matrix[j] -= matrix[i] * matrix[j, i]\n if printStep:\n print(matrix, \"\\n\")\n \n return matrix\n\n\n# ## Ordo 2\n\ndef CorrelationCoefPoli2(x, y, a0, a1, a2):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt((Dt-D)/Dt)\n \n table = pd.DataFrame({\n 'x': x,\n 'y': y,\n 'x2': x**2,\n \"y'\": ya,\n \"y - y'\": y - ya,\n \"(y - y')^2\": (y - ya)**2,\n 'y - ybar': y - np.average(y),\n '(y - ybar)^2': (y - np.average(y))**2\n })\n \n print(\"Dt (y-ybar)^2 :\", Dt)\n print(\"D (y-ya)^2 :\", D)\n print(table)\n \n return r\n\n\ndef ShowGraphPoli2(X, Y, a0, a1, a2):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([2, 4.25, 5.25, 7.81, 9.2, 10.6], dtype=\"float32\")\nY = np.array([7.2, 7.1, 6, 5, 3.5, 5], dtype=\"float32\")\n\n# orde 2\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y\n})\n\nprint(\"y avg\", np.average(Y))\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, y, xy, x2y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2],\n [x, x2, x3],\n [x2, x3, x4]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2 = result\nprint(\"\\ny = a0 + a1x + a2x^2\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli2(X, Y, a0, a1, a2))\n\nShowGraphPoli2(X, Y, a0, a1, a2)\n\n\n# -\n\n# ## Ordo 3\n\ndef CorrelationCoefPoli3(x, y, a0, a1, a2, a3):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2) + a3*(x**3)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt(((Dt-D)/Dt))\n \n return r\n\n\ndef ShowGraphPoli3(X, Y, a0, a1, a2, a3):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) + a3*(x**3) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10,\n 11, 12, 13, 14, 15, 16, 17, 18, 19, 20,\n 21, 22, 23, 24, 25], dtype=\"float32\")\nY = np.array([0.90, 1.42, 1.30, 1.55, 1.63, 1.32, 1.35,\n 1.47, 1.95, 1.66, 1.96, 1.47, 1.92, 1.35,\n 1.05, 1.85, 1.74, 1.65, 1.78, 1.71, 2.29,\n 1.82, 2.06, 2.14, 1.27], dtype=\"float32\")\n\n# orde 3\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx3y = np.sum((X**3)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\nx5 = np.sum(X**5)\nx6 = np.sum(X**6)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'x5': X**5,\n 'x6': X**6,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y,\n 'x3y': (X**3)*Y\n})\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, x5, x6, y, xy, x2y, x3y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2, x3],\n [x, x2, x3, x4],\n [x2, x3, x4, x5],\n [x3, x4, x5, x6]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y, x3y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2, a3 = result\nprint(\"\\ny = a0 + a1x + a2x^2 + a3x^3\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"a3 =\", a3)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli3(X, Y, a0, a1, a2, a3))\n\nShowGraphPoli3(X, Y, a0, a1, a2, a3)\n# -\n\n\n","repo_name":"lutfiandri/numerical-methods-notebook","sub_path":"03 - Regression.ipynb","file_name":"03 - Regression.ipynb","file_ext":"py","file_size_in_byte":9564,"program_lang":"python","lang":"en","doc_type":"code","stars":21,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"37435824621","text":"# +\nfrom math import trunc\nfrom multiprocessing.sharedctypes import Value\nimport xml.etree.ElementTree as ET\nimport xml.dom.minidom\nimport os\nimport cv2\nfrom tqdm import tqdm, trange\nimport numpy as np\nimport pandas as pd\nimport time\nimport warnings\nwarnings.filterwarnings(\"ignore\")\n\n\nfrom data_aug.data_aug import *\nfrom data_aug.bbox_util import *\n\nimport matplotlib.pyplot as plt \nimport pickle as pkl\n# %matplotlib inline\n\n\n# -\n\nimport os\nif not os.path.exists('jittering_data'):\n os.mkdir('jittering_data')\n\n# +\norigin_path = os.path.join('.', 'jittering_data', 'origin')\norigin_Image_Path = os.path.join(origin_path, 'JPEGImages')\norigin_Annotation_Path = os.path.join(origin_path, 'Annotations_with_ID')\n\noutput_path = os.path.join('.', 'jittering_data', 'output')\noutput_Image_Path = os.path.join(output_path, 'JPEGImages')\noutput_Annotation_Path = os.path.join(output_path, 'Annotations_with_ID')\nif not os.path.exists(output_Image_Path):\n os.mkdir(output_Image_Path)\nif not os.path.exists(output_Annotation_Path):\n os.mkdir(output_Annotation_Path)\n\n# -\n\ntarget_list = [i.split('.')[0] for i in os.listdir(origin_Annotation_Path)[:]]\n\n\n# +\ndef process(target):\n origin_img_path = os.path.join(origin_Image_Path, target+'.jpg')\n origin_xml_path = os.path.join(origin_Annotation_Path, target+'.xml')\n origin_boxes = xml_to_boxes(origin_xml_path)\n origin_img = cv2.imread(origin_img_path)\n #origin_img = 255*np.ones(origin_img.shape, origin_img.dtype)\n #plt.imshow(origin_img[:,:,::-1])\n #plt.show()\n #origin_img2 = cv2.imread(origin_img_path)\n # Demo the data\n #print(origin_boxes)\n #plotted_img = draw_rect(origin_img, origin_boxes)\n #cv2.imwrite('nb.jpg', origin_img)\n #cv2.imwrite('nbt.jpg', origin_img2)\n #cv2.imshow('t',origin_img)\n #cv2.waitKey()\n #plt.show()\n # just copy\n #seq = Sequence([RandomHSV(0, 0, 100),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(50, 100, 100)\n #img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n #plt.imshow(img_[:,:,::-1])\n #plt.show()\n #return\n output_img_path = os.path.join(output_Image_Path, target+str(0)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(0)+'.xml')\n cv2.imwrite(output_img_path, origin_img)\n for i in range(1,5):\n output_img_path = os.path.join(output_Image_Path, target+str(i)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(i)+'.xml')\n \n seq = Sequence([RandomHSV(40, 50, 50),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(0, 0, 10)\n #ad = origin_img.copy()\n #print(ad.shape, ad.dtype)\n #cv2.imshow('title',origin_img)\n #cv2.waitKey()\n #cv2.imshow('title',ad)\n #cv2.waitKey()\n try:\n img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n cv2.imwrite(output_img_path, img_)\n tree = gen_tree(origin_xml_path, img_.shape,bboxes_)\n tree.write(output_xml_path)\n except:\n print(\"STOP\")\n \n #plotted_img = draw_rect(img_, bboxes_)\n #plt.imshow(plotted_img[:,:,::-1])\n #plt.show()\n \n\n \n# -\n\ndef gen_tree(xml_path, img_shape, bboxes):\n tree = ET.parse(xml_path)\n root = tree.getroot()\n root.find('size/width').text = str(img_shape[0])\n root.find('size/height').text = str(img_shape[1])\n boxes = [i for i in root.iter('object')]\n for box in boxes:\n root.remove(box)\n for box in bboxes:\n new_box = ET.Element('object')\n ET.SubElement(new_box, 'ID').text = str(int(box[4]))\n bndbox = ET.SubElement(new_box, 'bndbox')\n ET.SubElement(bndbox, 'xmin').text = str(int(box[0]))\n ET.SubElement(bndbox, 'ymin').text = str(int(box[1]))\n ET.SubElement(bndbox, 'xmax').text = str(int(box[2]))\n ET.SubElement(bndbox, 'ymax').text = str(int(box[3]))\n root.append(new_box)\n ET.indent(root)\n return tree\n\n\n# +\n\ndef xml_to_boxes(path):\n tree = ET.parse(path)\n root = tree.getroot()\n list_with_all_boxes = []\n for box in root.iter('object'):\n xmin = int(box.find(\"bndbox/xmin\").text)\n ymin = int(box.find(\"bndbox/ymin\").text)\n xmax = int(box.find(\"bndbox/xmax\").text)\n ymax = int(box.find(\"bndbox/ymax\").text)\n Id = int(box.find('ID').text)\n list_with_all_boxes.append([xmin, ymin, xmax, ymax, Id])\n return np.array(list_with_all_boxes, dtype='float64')\n\n\n\n# -\n\n#process('000007')\nfor target in tqdm(target_list):\n process(target)\n\n#\n","repo_name":"Pxter7777/DLcomp2","sub_path":"jittering.ipynb","file_name":"jittering.ipynb","file_ext":"py","file_size_in_byte":4665,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"28041736766","text":"# # Project: TMDB Data Analysis \n#\n# ## Table of Contents\n#
\n#\n# You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.\n\n# +\n# TODO: Display an image along with the top 5 classes\ndef plot_im_probs_classes(image_path, model, device=device):\n probs, classes = predict(image_path, model, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout()\n \ndef plot_im_probs_classes2(image_path, checkpoint, device=device):\n probs, classes = predict2(image_path, checkpoint, topk=5, device=device)\n probs.reverse()\n classes.reverse()\n classes = [cat_to_name[k] for k in classes]\n \n image = Image.open(image_path)\n tranfs = transforms.Compose([transforms.Resize(256), transforms.CenterCrop(224)])\n \n fig, (ax1, ax2) = plt.subplots(figsize=(6,9), nrows=2)\n ax1.imshow(tranfs(image))\n ax1.axis('off')\n ax1.set_title(classes[-1], size=20)\n ax2.barh(np.arange(5), probs)\n ax2.set_aspect(0.19 * probs[-1] * 1.05, adjustable='box')\n ax2.set_xlim(0, probs[-1] * 1.05)\n ax2.set_yticks(np.arange(5))\n ax2.set_yticklabels(classes, size='large');\n \n plt.tight_layout() \n\n\n# -\n\nplot_im_probs_classes('flowers/test/1/image_06760.jpg', model=reloaded_best_model, device=device)\n\nplot_im_probs_classes2('flowers/test/1/image_06760.jpg', 'checkpoint_best.pth', device=device)\n\n# Resources used for this project:\n# * Material from the first term of Udacity's Data Scientist Nanodegree program. \n# * [Transfer Learning tutorial](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html). `train_model` function was taken from this tutorial with small changes as well as other hints such as shuffling the datasets and using a scheduler. \n# * [PyTorch documentation](https://pytorch.org/docs/stable/index.html).\n# * [Argparse tutorial](https://pymotw.com/3/argparse/#argument-types) and [documentation](https://docs.python.org/dev/library/argparse.html#argparse.Namespace).\n","repo_name":"anamamatelashvili/deep_learning_project","sub_path":"Image Classifier Project.ipynb","file_name":"Image Classifier Project.ipynb","file_ext":"py","file_size_in_byte":31574,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"5"}
+{"seq_id":"581944992","text":"# + [markdown] id=\"E-aBmrp8931c\"\n# # Proyecto empresa de viajes compartidos\n\n# + [markdown] id=\"uEJABzxG931c\"\n# ### Descripción del proyecto\n#\n# Durante este proyecto analizaré datos para una nueva empresa de viajes compartidos que se esta lanzando en Chicago. El objetivo será encontrar patrones en los datos y entender las preferencias de los pasajeros y el impacto de los factores externos en los viajes.\n#\n# #### Descripción de los datos\n#\n# Tabla 1: contiene datos sobre la cantidad de viajes que realizó cada compañía.\n# - company_name: nombre de la empresa de taxis\n# - trips_amount: el número de viajes de cada compañía de taxis el 15 y 16 de noviembre de 2017.\n#\n# Tabla 2: contiene datos sobre la finalización de los viajes \n# - dropoff_location_name: barrios de Chicago donde finalizaron los viajes\n# - average_trips: el promedio de viajes que terminaron en cada barrio en noviembre de 2017.\n#\n# Tabla 3:\n# Contiene datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. \n# - start_ts: fecha y hora de la recogida\n# - weather_conditions: condiciones climáticas en el momento en el que comenzó el viaje\n# - duration_seconds: duración del viaje en segundos\n\n# + [markdown] id=\"1G-R9FNe931c\"\n# ### Análisis exploratorio de datos\n\n# + id=\"PZCA0wo3931c\"\n# Importación de librerías a utilizar\n\nimport pandas as pd\nfrom scipy import stats as st\nfrom scipy.stats import levene\nimport matplotlib.pyplot as plt\nfrom seaborn import boxplot\n\n# + id=\"r3wEulH9931d\" outputId=\"1375e687-549f-4792-8c56-1ebfe3ff3c0f\"\n# Importación de archivo 1: número de viajes por compañia\n\ncompanies_trips= pd.read_csv('/datasets/project_sql_result_01.csv')\ncompanies_trips.info()\ncompanies_trips.head()\n\n\n# + id=\"MA1CVDmo931d\" outputId=\"0d2dabbc-9c46-4fb2-e35a-d36d79ab3335\"\n# Chequeo de valores ausentes\n\ncompanies_trips.isna().sum()\n\n# + id=\"UVq55dyy931d\" outputId=\"9735fac2-c40f-4ee0-8c3e-73d4238f734d\"\n# Chequeo de valores duplicados\n\ncompanies_trips.duplicated().sum()\n\n# + id=\"Bio8N7b7931d\" outputId=\"01191f13-de81-4e3f-bc49-83e4a9496fa8\"\n# Importación de archivo 2: promedio de viajes por barrio \n\ndropoff_location= pd.read_csv('/datasets/project_sql_result_04.csv')\ndropoff_location.info()\ndropoff_location.head()\n\n# + id=\"VfGtzMP-931e\" outputId=\"c6356ec7-0873-41da-c118-50a4d1e55f63\"\n# Chequeo de valores ausentes \n\ndropoff_location.isna().sum()\n\n# + id=\"bYD1uV96931e\" outputId=\"1d75298f-3319-4eb2-818a-4e3f4d0a2a96\"\n# Chequeo de valores duplicados\n\ndropoff_location.duplicated().sum()\n\n# + id=\"8aHq5XT-931e\" outputId=\"384087f9-8cea-4f88-e3a1-f5c338d36c98\"\n# Identificar los 10 principales barrios en términos de finalización del recorrido\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10)\n\n# + id=\"LdV71Unp931e\" outputId=\"5da0eaf9-c0e7-4ef7-f5c7-d51006a0a7ba\"\n#Gráfico: empresas de taxis y número de viajes\n\ncompanies_trips.plot(kind='bar',x='company_name', y='trips_amount', ylabel='trips_amount', figsize=(20,10), title='Viajes por empresa de taxi')\n\n# + [markdown] id=\"shnqBODc931e\"\n# Como podemos observar en el gráfico de arriba, la empresa de taxis más usada los días 15 y 16 de noviembre fue Flash Cab. Ésta realizó casi el doble de viajes que la empresa que le sigue: Taxi Affiliation Services. \n#\n# A su vez, dejando de lado Flash Cab que fue sin dudas la más poplar, podemos observar que hay pequeños grupos de compañías en los cuales no difiere tanto la cantidad de viajes realizados. Por ejemplo, las otras 5 copañías principales de taxis realizaron entre 9180 a 11400 viajes.\n#\n# Por otro lado, podemos observar que la mitad de las compañías de taxis tuvieron muy pocos viajes esos días. \n\n# + id=\"bzjPAp8R931e\" outputId=\"4031eafb-cf1d-4fc0-d3f1-73c5e893dc32\"\n# Gráfico 10 barrios principales por número de finalizaciones\n\ndropoff_location.sort_values(by= 'average_trips',ascending= False).head(10).plot(kind='bar',x='dropoff_location_name', y='average_trips', ylabel='Trips amount', figsize=(20,10), title='Principales barrios')\n\n# + [markdown] id=\"Hl0_zDu_931f\"\n# Como podemos observar en el gráfico de arriba, se muestran los 10 barrios principales hacia los cuales se dirigieron los taxis (destino final). El barrio con más viajes fue Loop con un promedio de 10727 viajes en el mes de noviembre seguido por River North con un promedio de 9523 viajes.\n#\n# Después se encuentran Streeterville\ty West Loop con un promedio de viajes de 6664 y 5163 respectivamente y por último un tercer grupo en la tabla con barrios con un promedio de viajes entre 1260 a 2420 en el mes de noviembre. \n\n# + [markdown] id=\"ibJP9mHu931f\"\n# ### Prueba de hipótesis\n\n# + [markdown] id=\"9qceopHk931f\"\n# A continuación voy a importar el archivo con los datos que usare para hacer la prueba de hipótesis.\n\n# + id=\"Mj53hprj931f\" outputId=\"216d861a-a20e-40c7-bfca-172a9615cafb\"\n# Importación de archivo para trabajar sobre la hipótesis\n\narchivo_hipotesis = pd.read_csv('/datasets/project_sql_result_07.csv')\narchivo_hipotesis.info()\narchivo_hipotesis.head()\n\n# + id=\"nT7L8FIk931f\" outputId=\"04025974-4c5f-4a5f-84e1-3c46cf2c3852\"\n# Chequeo de valores ausentes\n\narchivo_hipotesis.isna().sum()\n\n# + id=\"ZqMuqgQS931f\" outputId=\"9b37c186-3558-4c3a-b90c-d3e53abca6f5\"\n# Chequeo de valores duplicados\n\narchivo_hipotesis.duplicated().sum()\n\n# + id=\"2gIJlz-9931f\" outputId=\"7e2315d2-3169-427a-c706-8443f22436e4\"\narchivo_hipotesis.value_counts()\n\n# + [markdown] id=\"aygBH6hT931f\"\n# Como podemos observar, en el archivo que nos dieron para hacer la prueba de hipótesis tenemos valores duplicados. Estos pueden deberse a que el archivo contiene los datos sobre viajes desde el Loop hasta el Aeropuerto Internacional O'Hare. Puede que haya determinadas paradas en Loop con destino hacia el aeropuerto. Al ser un punto fijo es posible que en condiciones normales, el recorrido dure siempre lo mismo; todos los los viajes que salgan de ahi a una determnada hora van a tardar lo mismo generando que haya varios datos con os mismos tiempos.\n\n# + id=\"fb9UZ9-Q931f\" outputId=\"128662bf-d6f4-403b-9c56-d506f9680a7d\"\nboxplot(data= archivo_hipotesis, x=\"duration_seconds\", y=\"weather_conditions\")\n\n# + [markdown] id=\"OxVmL-Y2931g\"\n# Voy a probar la hipótesis de que la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare cambia los sábados lluviosos.\n#\n# Hipótesis nula: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos. \n#\n# Hipótesis alternativa: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional O'Hare difiere los sábados lluviosos.\n#\n# Previo a la prueba de hipótesis voy a realizar la prueba de levene para ver si las varianzas son iguales. Usaré como hipótesis nula: las varianzas son iguales y como hipótesis alternativa las varianzas no son iguales.\n\n# + id=\"3sY6DdYD931g\" outputId=\"a465661a-30aa-4dd3-92e3-eb7b9dee7791\"\n# Prueba de hipotesis nula levene: varianzas son iguales \n\ndatos_lluvia= archivo_hipotesis.query('weather_conditions == \"Bad\" and duration_seconds >0')\ndatos_good= archivo_hipotesis.query('weather_conditions ==\"Good\"and duration_seconds >0')\n\nlluvia= datos_lluvia['duration_seconds']\ngood= datos_good['duration_seconds']\n\nalpha= 0.05\n\nstat,results= levene(lluvia, good)\nprint('P_value:', results)\n\nif results < alpha:\n print('Rechazo la hipótesis nula')\nelse:\n print('No rechazo la hipótesis nula')\n\n# + [markdown] id=\"huB3ekYd931g\"\n# Como podemos observar de la prueba de Levene realizada arriba, no rechazamos la hipótesis nula que indica que las varianzas son iguales. \n#\n# A continuación realizare la prueba de hipótesis.\n\n# + id=\"ABv0Xklx931g\" outputId=\"7198fbf1-68b8-4240-bb38-b4164b0464f5\"\n# Ho: la duración promedio de los viajes desde el Loop hasta el Aeropuerto Internacional de O'Hare no difiere los sábados lluviosos.\n\nalpha= 0.05\nresults= st.ttest_ind(lluvia,good, equal_var= True)\nprint('P_value:', results.pvalue)\n\nif results.pvalue < alpha:\n print('Rechazo la hipótesis nula')\nelse: \n print('No rechazo hipótesis nula')\n\n# + [markdown] id=\"Ns2N_ZTw931g\"\n# De acuerdo con los datos con los que contamos, rechazo la hipótesis nula. La duración promedio de los viajes desde el Loop hasta el aeropuerto de O'Hare cambia los sábados lluviosos.\n\n# + [markdown] id=\"dtnLXksI931g\"\n# ## Conclusión \n#\n# Para concluir, podemos decir que la empresa de taxis más utilizada es Flash Cab seguida por Taxi Affiliation Services, teniendo la primera casi el doble de viajes que la segunda. Esta diferencia puede deberse a que Flash Cab sea una empresa mucho más grande que las otras; al tener una flota de autos más grande en la calle le permite tener el doble de viajes que su competidor más cercano. \n#\n# Con respecto a los detinos finales de los viajes, Loop resulto el más concurrido seguido de cerca por River North.\n#\n# Por último la duración promedio de los viajes difiere un día lluvioso en comparación a un día de buen tiempo. Esta diferencia puede deberse principalmente al factor climático. Los días de mal tiempo puede que haya más tránsito porque la gente decide salir con sus autos y se genera más congestión. También la gente anda un poco más despacio impactando en la duración pormedio de los viajes. \n","repo_name":"CarmenPujato/practicum","sub_path":"empresa de viajes compartidos/Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_name":"Proyecto6_Empresa_de_viajes_compartidos.ipynb","file_ext":"py","file_size_in_byte":9307,"program_lang":"python","lang":"en","doc_type":"code","stars":1,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"43095479709","text":"# # Kaggle class competition\n# # Deep Learning and Neural Network Models to predict traffic volume\n# # Models: LSTM, GRUs\n# # Library used: Tensorflow, Keras\n\n# ### Set Up Libraries\n\n# +\n# ns - as typically happens, outdated components like as keywords, functions, and classes trigger a warning.\nimport warnings\nwarnings.filterwarnings('ignore')\n# ns - below are the neccessary packages utilized in this project\nimport os\nimport datetime\n\nimport IPython\nimport IPython.display\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns\nimport tensorflow as tf\n\nmpl.rcParams['figure.figsize'] = (8, 6)\nmpl.rcParams['axes.grid'] = False\n# -\n\n# ### Import data\n\n# ns - use pandas package to read in data from file located within folder \ndata = pd.read_csv('Metro_Interstate_Traffic_Volume.csv', infer_datetime_format=True)\ndata.head()\n\n# ### Inspect and Clean up\n\n# ns - investigate the data info\ndata.info()\n\n# #### It could be seen that there are both str and num types of data as categorized below\n# categorical features: holiday, weather_main, weather_description\n#\n# Continious features: temp, rain_1h, show_1h, clouds_all\n#\n# Target: traffic_volume\n\n# ### Convert date_time column to datetime type\n#\n# It is usable if we don't convert string type to datetime type in this project. It is due to the fact that time series will absolutely need datetime. So below is the demonstration.\n\n# ns - datetime type conversion\ndate_time = pd.to_datetime(data.pop('date_time'), format='%Y-%m-%d %H:%M:%S')\n\n# ### Convert to a usable signal \n#\n# In this form of signal, we would be able to utilize its capacity in the ultimate way as it reveals more insignhts as illjustrated below. \n\ntimestamp_s = date_time.map(datetime.datetime.timestamp)\n\n# +\nday = 24*60*60\nyear = (365.2425)*day\n\ndata['Day sin'] = np.sin(timestamp_s * (2 * np.pi / day))\ndata['Day cos'] = np.cos(timestamp_s * (2 * np.pi / day))\ndata['Year sin'] = np.sin(timestamp_s * (2 * np.pi / year))\ndata['Year cos'] = np.cos(timestamp_s * (2 * np.pi / year))\n# -\n\nplt.plot(np.array(data['Day sin'])[:25])\nplt.plot(np.array(data['Day cos'])[:25])\nplt.xlabel('Time [h]')\nplt.title('Time of day signal')\n\n# The best way to identify which frequencies are most essential is to use a \"fft\". As a preliminary test, below is the traffic volume rfft experiment. While we're discussing common frequencies, it's interesting to note the highs at 1/year and 1/day frequencies:\n\n# +\nfft = tf.signal.rfft(data['temp'])\nf_per_dataset = np.arange(0, len(fft))\n\nn_samples_h = len(data['temp'])\nhours_per_year = 24*365.2524\nyears_per_dataset = n_samples_h/(hours_per_year)\n\nf_per_year = f_per_dataset/years_per_dataset\nplt.step(f_per_year, np.abs(fft))\nplt.xscale('log')\nplt.ylim(0, 400000)\nplt.xlim([0.1, max(plt.xlim())])\nplt.xticks([1, 365.2524], labels=['1/Year', '1/day'])\n_ = plt.xlabel('Frequency (log scale)')\n# -\n\n# ### Drop variable \"weather_description\" and convert \"holiday\" and \"weather_main\" to dummies variables\n#\n# It is assumed that the weather description might be more irrelevant to this project or its interpretation could benefit less. Thus, I have decided to drop it. While both weather main and the holiday are deemed significant.\n\n# ns - considered to drop some variable and generate dummies variables for the rest of obj type\ndata.drop('weather_description', axis=1, inplace=True)\ndata = pd.get_dummies(data, columns = ['holiday', 'weather_main'], drop_first=True)\ndata.head()\n\n# ### Split the data\n\n# +\n# ns - as indicated in the instruction, the test set should be excluded with the size of 5000 ea.\n# ns - the slicing method is used in this data split\ncolumn_indices = {name: i for i, name in enumerate(data.columns)}\n\nn = len(data)\nprint(\"There are \", n, \"records and divisions are as below\")\n\ntest_df = data[-5000:]\nprint(\"Test set: \", test_df.shape)\n\nval_df = data[-13000:-5000]\nprint(\"Validation set: \", val_df.shape)\n\ntrain_df = data[0:-13000]\nprint(\"Training set: \", train_df.shape)\n\nnum_features = data.shape[1]\n# -\n\n# ### Data Normalization\n\n# +\ntrain_mean = train_df.mean()\ntrain_std = train_df.std()\n\ntrain_df = (train_df - train_mean) / train_std\nval_df = (val_df - train_mean) / train_std\ntest_df = (test_df - train_mean) / train_std\n\n\n# -\n\n# ### Data windowing\n#\n# The models in this project will make a set of predictions based on a window of consecutive samples from the data and below is the useful windowgenerator class for segmenting data as required for modelling.\n\nclass WindowGenerator():\n def __init__(self, input_width, label_width, shift,\n train_df=train_df, val_df=val_df, test_df=test_df,\n label_columns=None):\n # Store the raw data.\n self.train_df = train_df\n self.val_df = val_df\n self.test_df = test_df\n\n # Work out the label column indices.\n self.label_columns = label_columns\n if label_columns is not None:\n self.label_columns_indices = {name: i for i, name in\n enumerate(label_columns)}\n self.column_indices = {name: i for i, name in\n enumerate(train_df.columns)}\n\n # Work out the window parameters.\n self.input_width = input_width\n self.label_width = label_width\n self.shift = shift\n\n self.total_window_size = input_width + shift\n\n self.input_slice = slice(0, input_width)\n self.input_indices = np.arange(self.total_window_size)[self.input_slice]\n\n self.label_start = self.total_window_size - self.label_width\n self.labels_slice = slice(self.label_start, None)\n self.label_indices = np.arange(self.total_window_size)[self.labels_slice]\n\n def __repr__(self):\n return '\\n'.join([\n f'Total window size: {self.total_window_size}',\n f'Input indices: {self.input_indices}',\n f'Label indices: {self.label_indices}',\n f'Label column name(s): {self.label_columns}'])\n\n\n# As per required, the windowgenerator here will provide 6 width with 2 future prediction and 1 label.\n\nw2 = WindowGenerator(input_width=6, label_width=1, shift=2,\n label_columns=['traffic_volume'])\nw2\n\n\n# +\ndef split_window(self, features):\n inputs = features[:, self.input_slice, :]\n labels = features[:, self.labels_slice, :]\n if self.label_columns is not None:\n labels = tf.stack(\n [labels[:, :, self.column_indices[name]] for name in self.label_columns],\n axis=-1)\n\n # Slicing doesn't preserve static shape information, so set the shapes\n # manually. This way the `tf.data.Datasets` are easier to inspect.\n inputs.set_shape([None, self.input_width, None])\n labels.set_shape([None, self.label_width, None])\n\n return inputs, labels\n\nWindowGenerator.split_window = split_window\n# -\n\n# ### Note: \n#\n# Typically data in TensorFlow is packed into arrays where the outermost index is across examples (the \"batch\" dimension). The middle indices are the \"time\" or \"space\" (width, height) dimension(s). The innermost indices are the features.\n\n# +\n# ns - I tried with other stack but it failed to corporate with the model\n# Stack three slices, the length of the total window:\nexample_window = tf.stack([np.array(train_df[:w2.total_window_size]),\n np.array(train_df[100:100+w2.total_window_size]),\n np.array(train_df[200:200+w2.total_window_size])])\n\n\nexample_inputs, example_labels = w2.split_window(example_window)\n\nprint('All shapes are: (batch, time, features)')\nprint(f'Window shape: {example_window.shape}')\nprint(f'Inputs shape: {example_inputs.shape}')\nprint(f'labels shape: {example_labels.shape}')\n# -\n\n# ### Plot \n#\n# #### Here is a plot method that allows a simple visualization of the split window\n\nw2.example = example_inputs, example_labels\n\n\n# +\ndef plot(self, model=None, plot_col='traffic_volume', max_subplots=3):\n inputs, labels = self.example\n plt.figure(figsize=(12, 8))\n plot_col_index = self.column_indices[plot_col]\n max_n = min(max_subplots, len(inputs))\n for n in range(max_n):\n plt.subplot(max_n, 1, n+1)\n plt.ylabel(f'{plot_col} [normed]')\n plt.plot(self.input_indices, inputs[n, :, plot_col_index],\n label='Inputs', marker='.', zorder=-10)\n\n if self.label_columns:\n label_col_index = self.label_columns_indices.get(plot_col, None)\n else:\n label_col_index = plot_col_index\n\n if label_col_index is None:\n continue\n\n plt.scatter(self.label_indices, labels[n, :, label_col_index],\n edgecolors='k', label='Labels', c='#2ca02c', s=64)\n if model is not None:\n predictions = model(inputs)\n plt.scatter(self.label_indices, predictions[n, :, label_col_index],\n marker='X', edgecolors='k', label='Predictions',\n c='#ff7f0e', s=64)\n\n if n == 0:\n plt.legend()\n\n plt.xlabel('Time [h]')\n\nWindowGenerator.plot = plot\n# -\n\nw2.plot()\n\n\n# The plots above illustrate the 6 inputs and 1 label as we defined them ealier. \n\n# ### Create tf.data.Datasets\n\n# +\ndef make_dataset(self, data):\n data = np.array(data, dtype=np.float32)\n ds = tf.keras.preprocessing.timeseries_dataset_from_array(\n data=data,\n targets=None,\n sequence_length=self.total_window_size,\n sequence_stride=1,\n shuffle=False,\n batch_size=32,)\n\n ds = ds.map(self.split_window)\n\n return ds\n\nWindowGenerator.make_dataset = make_dataset\n\n\n# -\n\n# ### Note:\n#\n# This make_dataset method will take a time series DataFrame and convert it to a tf.data.Dataset of (input_window, label_window) pairs using the preprocessing.timeseries_dataset_from_array function. The WindowGenerator object holds training, validation and test data. Add properties for accessing them as tf.data.Datasets using the above make_dataset method.\n\n# +\n@property\ndef train(self):\n return self.make_dataset(self.train_df)\n\n@property\ndef val(self):\n return self.make_dataset(self.val_df)\n\n@property\ndef test(self):\n return self.make_dataset(self.test_df)\n\n@property\ndef example(self):\n \"\"\"Get and cache an example batch of `inputs, labels` for plotting.\"\"\"\n result = getattr(self, '_example', None)\n if result is None:\n # No example batch was found, so get one from the `.train` dataset\n result = next(iter(self.train))\n # And cache it for next time\n self._example = result\n return result\n\nWindowGenerator.train = train\nWindowGenerator.val = val\nWindowGenerator.test = test\nWindowGenerator.example = example\n# -\n\n# Each element is an (inputs, label) pair\nw2.train.element_spec\n\n# #### Iterating over a Dataset yields concrete batches:\n\nfor example_inputs, example_labels in w2.train.take(1):\n print(f'Inputs shape (batch, time, features): {example_inputs.shape}')\n print(f'Labels shape (batch, time, features): {example_labels.shape}')\n\n# #### This project trains many models, so package the training procedure into a function\n\n# +\nMAX_EPOCHS = 100\n\ndef compile_and_fit(model, window, patience=2):\n early_stopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss',\n patience=patience,\n mode='min')\n\n model.compile(loss=tf.losses.MeanSquaredError(),\n optimizer=tf.optimizers.Adam(),\n metrics=[tf.metrics.MeanAbsoluteError()])\n\n history = model.fit(window.train, epochs=MAX_EPOCHS,\n validation_data=window.val,\n callbacks=[early_stopping])\n return history\n\n\n# -\n\n# For the multi-step model, the training data consists of hourly samples. However, here, the models will learn to predict 2h of the future, given 6h of the past. Here is a Window object that generates these slices from the dataset.\n\n# +\n# ns - indicating the 2 hours future output from the 6 hours history input\nOUT_STEPS = 2\nmulti_window = WindowGenerator(input_width=6,\n label_width=OUT_STEPS,\n shift=OUT_STEPS)\n\nmulti_window.plot()\nmulti_window\n# -\n\n# Illustration here displays the 6 inputs with 2 hours prediction in the future.\n\n# ### Model Construction\n# #### (1) The Base LSTM model\n#\n# This model is expected to be used as the baseline model in this project. The error is aimed to be minimized in further steps. The architecture of this base model consists of LSTM layer with 32 unit, Dense layer and then reshape respectively.\n\n# +\nmulti_val_performance = {}\nmulti_performance = {}\n\nmulti_lstm_model = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.val)\nmulti_performance['LSTM_Base'] = multi_lstm_model.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model)\n# -\n\n# #### (2) The Base LSTM with parameter Recurrent Dropout (0.1) and dropout layer (0.1)\n#\n# In this model, the recurrent Dropout with value 0.1 is added into the base model. It is expected to improve the mean absolute error. Recurrent Dropout is a regularization method for recurrent neural networks. Dropout is applied to the updates to LSTM memory cells (or GRU states), i.e. it drops out the input/update gate in LSTM/GRU.\n\n# +\n# ns - model with first implementation based on the base \nmulti_lstm_model2 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model2, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout'] = multi_lstm_model2.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model2)\n# -\n\n# #### (3) The Base LSTM with parameter Recurrent Dropout (0.1) + unit_forget_bias, and dropout layer (0.1)\n#\n# In this model the unit forget bias is added for the experiment. It is hoped that If True, it will add 1 to the bias of the forget gate at initialization. Setting it to true will also force bias_initializer=\"zeros\". The result looks better than the prev ious models. Nevertheless, it is so varying in each run but it seems to capture the minimal MAE.\n\n# +\n# ns - model with second implementation based on the previous model \nmulti_lstm_model3 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model3, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout2'] = multi_lstm_model3.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model3)\n# -\n\n# #### Additional Model\n# This is an additional model which I tried to tune after I have completed my report. The result is very impressive, so I keep it on this notebook so that I can make the improvement later when I have time. I tried to add the use_bias parameter to the Dense layer. It enables bias vector in this experiment. I want to utilize Gridsearch to this LSTM but time is running out. Therefore, I end up with this best model in terms of lowest MAE of 0.1724.\n\n# +\n# ns - this is a last minute trial model but it appears efficient\nmulti_lstm_model9 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.LSTM(32, unit_forget_bias=True, recurrent_dropout=0.1, return_sequences=False),\n tf.keras.layers.Dropout(0.1), \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model9, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout3'] = multi_lstm_model9.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model9)\n# -\n\n# #### (4) The GRU Base Model with recurrent_dropout=0.1\n#\n# GRU is a recurrent neural network (RNN) in the most recent generation, and it is also equivalent to LSTM. In all likelihood, the newer GRU model will outperform the older LSTM model. I started with a simple 32-unit network and a 0.1 recurrence dropout rate.\n\n# +\n# ns - this model is assigned as the GRU base model\nmulti_lstm_model4 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.1, return_sequences=False),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model4, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU'] = multi_lstm_model4.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model4)\n# -\n\n# #### (5) The GRU Base Model with recurrent_dropout=0.3\n\n# +\n# ns - model with first implementation based on the GRU base \nmulti_lstm_model5 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model5, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU2'] = multi_lstm_model5.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model5)\n# -\n\n# #### (6) The GRU Base Model with recurrent_dropout=0.3 and dropout layer (0.1)\n#\n# In addition to the recurrent dropout parameter, this experiment includes the dropout layer with 0.1 figure which previously demonstrated the better version among dropout varying. The outcome below is quite as anticipated. the MAE becomes better. And again the results are very varying in each run.\n\n# +\n# ns - model with second implementation based on the GRU base \nmulti_lstm_model6 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n tf.keras.layers.GRU(32, activation='tanh', recurrent_activation='hard_sigmoid', recurrent_dropout=0.3),\n tf.keras.layers.Dropout(0.1),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model6, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.val)\nmulti_performance['LSTM_GRU3'] = multi_lstm_model6.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model6)\n# -\n\n# #### (7) The Bidirectional Model with LSTM\n#\n# I proceed with the selection of Bidirectional technique which I wanted to experimenrt with both LSTM and GRU models. It is started with the Bidirectional LSTM in this trial. I believe that allowing the LSTM model to learn the input sequence both forward and backward, and then concatenating both interpretations, can be useful on some sequence prediction challenges. the result looks great but it is not good as I predicted. In each run, it shows unsatisfied value of 1.7\n\n# +\n# ns - this model is built to try on Bidirectional with LSTM layer\nfrom keras.layers import Bidirectional\nfrom keras.layers import LSTM\n\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, \n return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi1'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### (8) The Bidirectional Model with GRU\n#\n# The best model of GRU is selected to be the layer of this Bidirectional method. As mentioned ealier, this result also exibits some fluctuation and not high enough as expected.\n\n# +\n# ns - this model is built to try on Bidirectional with GRU layer\nmulti_lstm_model7 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.GRU(32, activation='tanh', \n recurrent_activation='hard_sigmoid', \n recurrent_dropout=0.1, return_sequences=False)),\n tf.keras.layers.Dropout(0.1),\n \n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros()),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model7, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.val)\nmulti_performance['LSTM_Bi2'] = multi_lstm_model7.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model7)\n# -\n\n# #### Additional stacked Bidirectional model\n# This implementation is to show that the more layer I have added into the model the more complex the model would be. This complication diminishes the ability of model. As a result, this experiment proved to be quite weak in comparison with the value of MAE.\n\n# +\n# ns - this model is built to try on Bidirectional with stacked layer\nmulti_lstm_model10 = tf.keras.Sequential([\n # Shape [batch, time, features] => [batch, lstm_units]\n # Adding more `lstm_units` just overfits more quickly.\n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, unit_forget_bias=True, \n recurrent_dropout=0.1, return_sequences=True)),\n tf.keras.layers.Dropout(0.1), \n tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(8)),\n # Shape => [batch, out_steps*features]\n tf.keras.layers.Dense(OUT_STEPS*num_features,\n kernel_initializer=tf.initializers.zeros(), use_bias=True),\n # Shape => [batch, out_steps, features]\n tf.keras.layers.Reshape([OUT_STEPS, num_features])\n])\n\nhistory = compile_and_fit(multi_lstm_model10, multi_window)\n\nIPython.display.clear_output()\n\nmulti_val_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.val)\nmulti_performance['LSTM_ReDropout10'] = multi_lstm_model10.evaluate(multi_window.test, verbose=0)\nmulti_window.plot(multi_lstm_model10)\n# -\n\n# ### Model Prediction and Selection\n\n# ns - The best MAE is 0.1724 which is from model#9 \n# ns - Note that every time the kernel is reran, the MAE is varied\nprediction = multi_lstm_model9.predict(multi_window.test)\n# ns - The shape of prediction\nprediction.shape\n\n# ns - Extract the traffic volum column which is number 4\npredictedValue = prediction[:,1,4]\npredictedValue\n\n# #### Denormalization\n\n# ns - Extracting the std and mean values\nstd = train_std[4]\nmean = train_mean[4]\nprint(\"std: \", std, \"mean: \", mean)\n\n# ns - Performing data conversion to denormalized values\ndenorm = []\nfor i in predictedValue:\n val = (i*std)+mean\n denorm.append(val)\n\n# #### Export data to CSV file\n\n# ns - dataframe to export raw output to csv file\nimport pandas as pd\ndf = pd.DataFrame(denorm, columns = ['prediction'])\ndf.insert(0, 'id', range(1,4994))\ndf.to_csv(\"model9.csv\", index = False)\n\n# # Thank you!\n\n\n","repo_name":"iamnatapong55/Deep-Learning-Neural-Network","sub_path":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_name":"Deep Learning and Neural Networks - LSTM and GRUs Models.ipynb","file_ext":"py","file_size_in_byte":26193,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"44313882936","text":"# +\nimport numpy as np\nimport pandas as pd\nfrom matplotlib import pyplot as plt\n\nnp.set_printoptions(suppress=True, precision=3)\n\n\n# + [markdown] heading_collapsed=true\n# # Show Graph\n\n# + hidden=true\ndef ShowGraph(a, b, X, Y, line=\"a+bx\"):\n line_x = np.arange(np.amin(X), np.amax(X), 0.01)\n if line == \"a+bx\":\n line_y = a + b * line_x\n elif line == \"ax^b\":\n line_y = a * line_x ** b\n elif line == \"ae^(bx)\":\n line_y = a * np.exp(b*line_x)\n\n plt.clf()\n plt.plot(line_x, line_y, '-r')\n plt.plot(X, Y, 'ob')\n plt.grid()\n plt.show()\n\n\n# + [markdown] heading_collapsed=true\n# # Error\n\n# + [markdown] heading_collapsed=true hidden=true\n# ## Least Square Error\n\n# + hidden=true\ndef LeastSquareError(a, b, xi, yi):\n Dt = np.sum((yi - np.average(yi)) ** 2)\n D = np.sum((yi - a - b*xi) ** 2)\n \n r = ((Dt-D)/Dt) ** 0.5\n \n return r\n\n\n# + [markdown] hidden=true\n# ## Correlation Coeficient\n\n# + hidden=true\ndef CorrelationCoef(X, Y):\n difX = X - np.average(X)\n difY = Y - np.average(Y)\n \n r = (np.sum(difX*difY))/((np.sum((difX)**2)*np.sum((difY)**2))**(0.5))\n \n return r;\n\n\n# + [markdown] heading_collapsed=true\n# # Linear Regression → y = a + bx\n\n# + hidden=true\ndef LinearRegression(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'xiyi': X_data*Y_data,\n 'xi2': X_data*X_data\n })\n\n x_bar = np.average(data['xi'])\n y_bar = np.average(data['yi'])\n\n b = (n * np.sum(data['xiyi']) - np.sum(data['xi']) *\n np.sum(data['yi']))/(n * np.sum(data['xi2']) - np.sum(data['xi'])**2)\n a = y_bar - b * x_bar\n\n if showGraph:\n ShowGraph(a, b, data['xi'], data['yi'])\n\n if showData:\n print(\"linear regression data\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([4, 6, 8, 10, 14, 16, 20, 22, 24, 28])\nY_data = np.array([30, 18, 22, 28, 14, 22, 16, 8, 20, 8])\n\na, b = LinearRegression(X_data, Y_data, showGraph=True, showData=False)\nprint(\"y = a + bx\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] heading_collapsed=true\n# # Non-Linear Regression\n\n# + [markdown] hidden=true\n# ## y = ax^b\n\n# + hidden=true\ndef NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': np.log10(X_data),\n 'qi': np.log10(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = 10 ** a\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ax^b\")\n\n if showData:\n print(\"non-linear regression data (y = ax^b)\")\n print(\"pi = log(xi)\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression1(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ax^b\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# + [markdown] hidden=true\n# ## y = ae^(bx)\n\n# + hidden=true\ndef NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True):\n if X_data.size != Y_data.size:\n print(\"banyak data X dan Y tidak sama!\")\n exit()\n\n n = X_data.size\n data = pd.DataFrame({\n 'xi': X_data,\n 'yi': Y_data,\n 'pi': X_data,\n 'qi': np.log(Y_data)\n })\n\n a, b = LinearRegression(data['pi'], data['qi'],\n showGraph=False, showData=showData)\n a = np.exp(a)\n\n if showGraph:\n ShowGraph(a, b, X_data, Y_data, line=\"ae^(bx)\")\n\n if showData:\n print(\"non-linear regression data (y = ae^(bx))\")\n print(\"qi = log(yi)\")\n print(data)\n print()\n\n return [a, b]\n\n\nX_data = np.array([1, 2, 3, 4, 5])\nY_data = np.array([0.5, 1.7, 3.4, 5.7, 8.4])\n\na, b = NonLinearRegression2(X_data, Y_data, showGraph=True, showData=True)\nprint(\"y = ae^(bx)\\n\")\nprint(\"a:\", a)\nprint(\"b:\", b)\nprint(\"least suqare error:\", LeastSquareError(a, b, X_data, Y_data))\n\n\n# -\n\n# # Polinomial Regression\n\ndef GaussJordan(A, B, printStep=False):\n size = B.size\n matrix = A\n matrix = np.concatenate((A, B.reshape(size, 1)), axis=1)\n \n for i in range(0, size):\n if printStep:\n print(\"step\", i+1, \"\\n\")\n \n matrix[i] /= matrix[i, i]\n\n for j in range(0, size):\n if i == j:\n continue\n \n matrix[j] -= matrix[i] * matrix[j, i]\n if printStep:\n print(matrix, \"\\n\")\n \n return matrix\n\n\n# ## Ordo 2\n\ndef CorrelationCoefPoli2(x, y, a0, a1, a2):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt((Dt-D)/Dt)\n \n table = pd.DataFrame({\n 'x': x,\n 'y': y,\n 'x2': x**2,\n \"y'\": ya,\n \"y - y'\": y - ya,\n \"(y - y')^2\": (y - ya)**2,\n 'y - ybar': y - np.average(y),\n '(y - ybar)^2': (y - np.average(y))**2\n })\n \n print(\"Dt (y-ybar)^2 :\", Dt)\n print(\"D (y-ya)^2 :\", D)\n print(table)\n \n return r\n\n\ndef ShowGraphPoli2(X, Y, a0, a1, a2):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([2, 4.25, 5.25, 7.81, 9.2, 10.6], dtype=\"float32\")\nY = np.array([7.2, 7.1, 6, 5, 3.5, 5], dtype=\"float32\")\n\n# orde 2\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y\n})\n\nprint(\"y avg\", np.average(Y))\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, y, xy, x2y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2],\n [x, x2, x3],\n [x2, x3, x4]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2 = result\nprint(\"\\ny = a0 + a1x + a2x^2\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli2(X, Y, a0, a1, a2))\n\nShowGraphPoli2(X, Y, a0, a1, a2)\n\n\n# -\n\n# ## Ordo 3\n\ndef CorrelationCoefPoli3(x, y, a0, a1, a2, a3):\n # x y adalah data awal\n ya = a0 + a1*x + a2*(x**2) + a3*(x**3)\n \n D = np.sum((y - ya)**2)\n Dt = np.sum((y - np.average(y))**2)\n r = np.sqrt(((Dt-D)/Dt))\n \n return r\n\n\ndef ShowGraphPoli3(X, Y, a0, a1, a2, a3):\n # X Y (kapital) itu data awalnya yakk\n \n x = np.arange(np.amin(X), np.amax(X), 0.01) # ini tulis aja, susah jelasinnya :v\n y = a0 + a1*x + a2*(x**2) + a3*(x**3) # ini garis f(x) nya\n\n plt.clf() # ini buat clear plt yang sebelumnya, dihapus aja gpp kan cuma 1\n plt.plot(x, y, '-r') # ini nampilin garis f(x) tadi. '-r' => - artine garis; r artine red => garis merah\n plt.plot(X, Y, 'ob') # ini nampilin titik-titik data. 'ob' => o artine bulet2 (titik); b artine blue => titik2 biru\n plt.grid() # ini biar ada background gridnya\n plt.show() # ini buat menampilkan yang dibuat tadi\n\n\n# +\nX = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10,\n 11, 12, 13, 14, 15, 16, 17, 18, 19, 20,\n 21, 22, 23, 24, 25], dtype=\"float32\")\nY = np.array([0.90, 1.42, 1.30, 1.55, 1.63, 1.32, 1.35,\n 1.47, 1.95, 1.66, 1.96, 1.47, 1.92, 1.35,\n 1.05, 1.85, 1.74, 1.65, 1.78, 1.71, 2.29,\n 1.82, 2.06, 2.14, 1.27], dtype=\"float32\")\n\n# orde 3\nn = X.size\n\nx = np.sum(X)\ny = np.sum(Y)\nxy = np.sum(X*Y)\nx2y = np.sum((X**2)*Y)\nx3y = np.sum((X**3)*Y)\nx2 = np.sum(X**2)\nx3 = np.sum(X**3)\nx4 = np.sum(X**4)\nx5 = np.sum(X**5)\nx6 = np.sum(X**6)\n\ntable = pd.DataFrame({\n 'x': X,\n 'x2': X**2,\n 'x3': X**3,\n 'x4': X**4,\n 'x5': X**5,\n 'x6': X**6,\n 'y': Y,\n 'xy': X*Y,\n 'x2y': (X**2)*Y,\n 'x3y': (X**3)*Y\n})\n\nprint(table, '\\n')\nprint(n, x, x2, x3, x4, x5, x6, y, xy, x2y, x3y, '\\n', sep=\" | \")\n\n\nA = np.array([[n, x, x2, x3],\n [x, x2, x3, x4],\n [x2, x3, x4, x5],\n [x3, x4, x5, x6]], dtype=\"float32\")\n\nB = np.array([y, xy, x2y, x3y], dtype=\"float32\")\n\nresultMatrix = GaussJordan(A, B, printStep=True)\nresult = resultMatrix[:, -1]\na0, a1, a2, a3 = result\nprint(\"\\ny = a0 + a1x + a2x^2 + a3x^3\")\nprint(\"a0 =\", a0)\nprint(\"a1 =\", a1)\nprint(\"a2 =\", a2)\nprint(\"a3 =\", a3)\nprint(\"\\nKoefisien korelasi =\", CorrelationCoefPoli3(X, Y, a0, a1, a2, a3))\n\nShowGraphPoli3(X, Y, a0, a1, a2, a3)\n# -\n\n\n","repo_name":"lutfiandri/numerical-methods-notebook","sub_path":"03 - Regression.ipynb","file_name":"03 - Regression.ipynb","file_ext":"py","file_size_in_byte":9564,"program_lang":"python","lang":"en","doc_type":"code","stars":21,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"37435824621","text":"# +\nfrom math import trunc\nfrom multiprocessing.sharedctypes import Value\nimport xml.etree.ElementTree as ET\nimport xml.dom.minidom\nimport os\nimport cv2\nfrom tqdm import tqdm, trange\nimport numpy as np\nimport pandas as pd\nimport time\nimport warnings\nwarnings.filterwarnings(\"ignore\")\n\n\nfrom data_aug.data_aug import *\nfrom data_aug.bbox_util import *\n\nimport matplotlib.pyplot as plt \nimport pickle as pkl\n# %matplotlib inline\n\n\n# -\n\nimport os\nif not os.path.exists('jittering_data'):\n os.mkdir('jittering_data')\n\n# +\norigin_path = os.path.join('.', 'jittering_data', 'origin')\norigin_Image_Path = os.path.join(origin_path, 'JPEGImages')\norigin_Annotation_Path = os.path.join(origin_path, 'Annotations_with_ID')\n\noutput_path = os.path.join('.', 'jittering_data', 'output')\noutput_Image_Path = os.path.join(output_path, 'JPEGImages')\noutput_Annotation_Path = os.path.join(output_path, 'Annotations_with_ID')\nif not os.path.exists(output_Image_Path):\n os.mkdir(output_Image_Path)\nif not os.path.exists(output_Annotation_Path):\n os.mkdir(output_Annotation_Path)\n\n# -\n\ntarget_list = [i.split('.')[0] for i in os.listdir(origin_Annotation_Path)[:]]\n\n\n# +\ndef process(target):\n origin_img_path = os.path.join(origin_Image_Path, target+'.jpg')\n origin_xml_path = os.path.join(origin_Annotation_Path, target+'.xml')\n origin_boxes = xml_to_boxes(origin_xml_path)\n origin_img = cv2.imread(origin_img_path)\n #origin_img = 255*np.ones(origin_img.shape, origin_img.dtype)\n #plt.imshow(origin_img[:,:,::-1])\n #plt.show()\n #origin_img2 = cv2.imread(origin_img_path)\n # Demo the data\n #print(origin_boxes)\n #plotted_img = draw_rect(origin_img, origin_boxes)\n #cv2.imwrite('nb.jpg', origin_img)\n #cv2.imwrite('nbt.jpg', origin_img2)\n #cv2.imshow('t',origin_img)\n #cv2.waitKey()\n #plt.show()\n # just copy\n #seq = Sequence([RandomHSV(0, 0, 100),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(50, 100, 100)\n #img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n #plt.imshow(img_[:,:,::-1])\n #plt.show()\n #return\n output_img_path = os.path.join(output_Image_Path, target+str(0)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(0)+'.xml')\n cv2.imwrite(output_img_path, origin_img)\n for i in range(1,5):\n output_img_path = os.path.join(output_Image_Path, target+str(i)+'.jpg')\n output_xml_path = os.path.join(output_Annotation_Path, target+str(i)+'.xml')\n \n seq = Sequence([RandomHSV(40, 50, 50),RandomHorizontalFlip(), RandomScale(0.2), RandomRotate(20), RandomShear(0.2)])\n #seq = RandomHSV(0, 0, 10)\n #ad = origin_img.copy()\n #print(ad.shape, ad.dtype)\n #cv2.imshow('title',origin_img)\n #cv2.waitKey()\n #cv2.imshow('title',ad)\n #cv2.waitKey()\n try:\n img_, bboxes_ = seq(origin_img.copy(), origin_boxes.copy())\n cv2.imwrite(output_img_path, img_)\n tree = gen_tree(origin_xml_path, img_.shape,bboxes_)\n tree.write(output_xml_path)\n except:\n print(\"STOP\")\n \n #plotted_img = draw_rect(img_, bboxes_)\n #plt.imshow(plotted_img[:,:,::-1])\n #plt.show()\n \n\n \n# -\n\ndef gen_tree(xml_path, img_shape, bboxes):\n tree = ET.parse(xml_path)\n root = tree.getroot()\n root.find('size/width').text = str(img_shape[0])\n root.find('size/height').text = str(img_shape[1])\n boxes = [i for i in root.iter('object')]\n for box in boxes:\n root.remove(box)\n for box in bboxes:\n new_box = ET.Element('object')\n ET.SubElement(new_box, 'ID').text = str(int(box[4]))\n bndbox = ET.SubElement(new_box, 'bndbox')\n ET.SubElement(bndbox, 'xmin').text = str(int(box[0]))\n ET.SubElement(bndbox, 'ymin').text = str(int(box[1]))\n ET.SubElement(bndbox, 'xmax').text = str(int(box[2]))\n ET.SubElement(bndbox, 'ymax').text = str(int(box[3]))\n root.append(new_box)\n ET.indent(root)\n return tree\n\n\n# +\n\ndef xml_to_boxes(path):\n tree = ET.parse(path)\n root = tree.getroot()\n list_with_all_boxes = []\n for box in root.iter('object'):\n xmin = int(box.find(\"bndbox/xmin\").text)\n ymin = int(box.find(\"bndbox/ymin\").text)\n xmax = int(box.find(\"bndbox/xmax\").text)\n ymax = int(box.find(\"bndbox/ymax\").text)\n Id = int(box.find('ID').text)\n list_with_all_boxes.append([xmin, ymin, xmax, ymax, Id])\n return np.array(list_with_all_boxes, dtype='float64')\n\n\n\n# -\n\n#process('000007')\nfor target in tqdm(target_list):\n process(target)\n\n#\n","repo_name":"Pxter7777/DLcomp2","sub_path":"jittering.ipynb","file_name":"jittering.ipynb","file_ext":"py","file_size_in_byte":4665,"program_lang":"python","lang":"en","doc_type":"code","stars":0,"dataset":"github-jupyter-script","pt":"47"}
+{"seq_id":"28041736766","text":"# # Project: TMDB Data Analysis \n#\n# ## Table of Contents\n#