Add 1 files
Browse files- 2405/2405.03328.md +148 -0
2405/2405.03328.md
ADDED
|
@@ -0,0 +1,148 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
Title: Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge
|
| 2 |
+
|
| 3 |
+
URL Source: https://arxiv.org/html/2405.03328
|
| 4 |
+
|
| 5 |
+
Published Time: Tue, 07 May 2024 01:05:38 GMT
|
| 6 |
+
|
| 7 |
+
Markdown Content:
|
| 8 |
+
1 1 institutetext: Dept. of Math and Computer Science, University of Catania, Italy 2 2 institutetext: Centre for Medical Image Computing, University College London, UK 3 3 institutetext: School of Physics, Engineering and CS, University of Hertfordshire, UK
|
| 9 |
+
|
| 10 |
+
###### Abstract
|
| 11 |
+
|
| 12 |
+
In this work, we introduce Brain Latent Progression (BrLP), a novel spatiotemporal disease progression model based on latent diffusion. BrLP is designed to predict the evolution of diseases at the individual level on 3D brain MRIs. Existing deep generative models developed for this task are primarily data-driven and face challenges in learning disease progressions. BrLP addresses these challenges by incorporating prior knowledge from disease models to enhance the accuracy of predictions. To implement this, we propose to integrate an auxiliary model that infers volumetric changes in various brain regions. Additionally, we introduce Latent Average Stabilization (LAS), a novel technique to improve spatiotemporal consistency of the predicted progression. BrLP is trained and evaluated on a large dataset comprising 11,730 T1-weighted brain MRIs from 2,805 subjects, collected from three publicly available, longitudinal Alzheimer’s Disease (AD) studies. In our experiments, we compare the MRI scans generated by BrLP with the actual follow-up MRIs available from the subjects, in both cross-sectional and longitudinal settings. BrLP demonstrates significant improvements over existing methods, with an increase of 22% in volumetric accuracy across AD-related brain regions and 43% in image similarity to the ground-truth scans. The ability of BrLP to generate conditioned 3D scans at the subject level, along with the novelty of integrating prior knowledge to enhance accuracy, represents a significant advancement in disease progression modeling, opening new avenues for precision medicine. The code of BrLP is available at the following link: [https://github.com/LemuelPuglisi/BrLP](https://github.com/LemuelPuglisi/BrLP).
|
| 13 |
+
|
| 14 |
+
###### Keywords:
|
| 15 |
+
|
| 16 |
+
Disease Progression Diffusion Models Brain MRI
|
| 17 |
+
|
| 18 |
+
1 Introduction
|
| 19 |
+
--------------
|
| 20 |
+
|
| 21 |
+
Neurodegenerative diseases represent a global health challenge, affecting millions of people and leading to extensive morbidity and mortality. The situation is compounded by an increasingly ageing population, putting more strain on healthcare systems and society as a whole. Additionally, the progression of neurodegenerative diseases is characterized by its heterogeneous nature, with a variety of neuropathological patterns arising from different molecular subtypes[[20](https://arxiv.org/html/2405.03328v1#bib.bib20)]. In particular, these diseases affect brain regions at varying rates and through distinct mechanisms, highlighting the intricate nature of their pathophysiology[[25](https://arxiv.org/html/2405.03328v1#bib.bib25)]. Therefore, we need to develop new methods aimed at better understanding disease development, which will pave the way for more targeted and personalized treatment strategies. Initial approaches used disease progression modeling primarily based on scalar biomarkers[[26](https://arxiv.org/html/2405.03328v1#bib.bib26), [10](https://arxiv.org/html/2405.03328v1#bib.bib10)]. Despite the crude representations of these biomarkers, such approaches have been used to enhance our understanding of diseases[[4](https://arxiv.org/html/2405.03328v1#bib.bib4), [22](https://arxiv.org/html/2405.03328v1#bib.bib22)]. A natural evolution of these efforts is developing spatiotemporal models, which represent disease progression using rich, high-dimensional imaging biomarkers operating directly on medical scans. Unlike scalar biomarkers, these solutions facilitate the visualization and precise localization of complex patterns of structural changes, thereby offering a more detailed understanding of disease dynamics. Recent approaches have leveraged deep generative techniques, such as Variational Autoencoders (VAEs)[[17](https://arxiv.org/html/2405.03328v1#bib.bib17)], Generative Adversarial Networks (GANs)[[15](https://arxiv.org/html/2405.03328v1#bib.bib15), [23](https://arxiv.org/html/2405.03328v1#bib.bib23), [7](https://arxiv.org/html/2405.03328v1#bib.bib7), [28](https://arxiv.org/html/2405.03328v1#bib.bib28), [14](https://arxiv.org/html/2405.03328v1#bib.bib14)], and more recently, diffusion models[[24](https://arxiv.org/html/2405.03328v1#bib.bib24)], to infer the disease progression at the individual level. In particular, DaniNet[[15](https://arxiv.org/html/2405.03328v1#bib.bib15)] is a state-of-the-art model that uses adversarial learning combined with biological constraints to provide individualized predictions of brain MRIs. To mitigate memory requirements, DaniNet generates 2D slices that are then assembled into a 3D volume using a super-resolution module. Another approach is CounterSynth[[14](https://arxiv.org/html/2405.03328v1#bib.bib14)], a GAN-based counterfactual synthesis method that can simulate various conditions within a brain MRI, including ageing and disease progression. Lastly, SADM[[24](https://arxiv.org/html/2405.03328v1#bib.bib24)] is a diffusion model designed to generate longitudinal scans through autoregressive sampling by using a sequence of prior MRI scans.
|
| 22 |
+
|
| 23 |
+
[The](https://arxiv.org/html/2405.03328v1/) primary challenges of these methods are: 1) improving individualization by conditioning on subject-specific metadata; 2) using longitudinal scans when and if available; 3) enhancing spatiotemporal consistency to achieve a smooth progression across spatial and temporal dimensions; 4) managing the high memory demands imposed by the use of high-resolution 3D medical images[[2](https://arxiv.org/html/2405.03328v1#bib.bib2)]. Specifically, DaniNet[[15](https://arxiv.org/html/2405.03328v1#bib.bib15)] and CounterSynth[[14](https://arxiv.org/html/2405.03328v1#bib.bib14)] are not able to directly use longitudinal data if accessible. SADM[[24](https://arxiv.org/html/2405.03328v1#bib.bib24)] is not able to incorporate conditioning on subject-specific metadata and is also memory-intensive. Finally, neither CounterSynth nor SADM offer solutions to enforce spatiotemporal consistency.
|
| 24 |
+
|
| 25 |
+
In response to these challenges, we introduce BrLP, a novel spatiotemporal model, offering several key contributions: i) we propose to combine an LDM[[16](https://arxiv.org/html/2405.03328v1#bib.bib16)] and a ControlNet[[27](https://arxiv.org/html/2405.03328v1#bib.bib27)] to generate individualized brain MRIs conditioned on available subject data – addressing challenge[1;](https://arxiv.org/html/2405.03328v1#C1 "The ‣ 1 Introduction ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")ii) we propose to integrate prior knowledge of disease progression by employing an auxiliary model designed to infer volumetric changes in different brain regions, allowing the use of longitudinal data when available – addressing challenge[2;](https://arxiv.org/html/2405.03328v1#C1 "The ‣ 1 Introduction ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")iii) we propose LAS, a technique to improve spatiotemporal consistency in the predicted progression – addressing challenge[3;](https://arxiv.org/html/2405.03328v1#C1 "The ‣ 1 Introduction ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")and iv) we use latent representations of brain MRIs to limit the memory demands for processing 3D scans – addressing challenge[4.](https://arxiv.org/html/2405.03328v1#C1 "The ‣ 1 Introduction ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")
|
| 26 |
+
|
| 27 |
+
We evaluate BrLP by training it to learn progressive structural changes in the brains of individuals with different cognitive statuses: Cognitively Normal (CN), Mild Cognitive Impairment (MCI), and Alzheimer’s Disease. To do so, we use a large dataset of 11,730 T1-weighted brain MRIs from 2,805 subjects, sourced from three publicly available longitudinal studies on AD. To the best of our knowledge, we are the first to propose a 3D conditional generative model for brain MRI that incorporates prior knowledge of disease progression into the image generation process.
|
| 28 |
+
|
| 29 |
+
2 Methods
|
| 30 |
+
---------
|
| 31 |
+
|
| 32 |
+
### 2.1 Background - Diffusion Models
|
| 33 |
+
|
| 34 |
+
A Denoising Diffusion Probabilistic Model (DDPM)[[5](https://arxiv.org/html/2405.03328v1#bib.bib5)] is a deep generative model with two Markovian processes: forward diffusion and reverse diffusion. In the forward process, Gaussian noise is incrementally added to the original image x 0 subscript 𝑥 0 x_{0}italic_x start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT over T 𝑇 T italic_T steps. At each step t 𝑡 t italic_t, noise is introduced to the current image x t−1 subscript 𝑥 𝑡 1 x_{t-1}italic_x start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT by sampling from a Gaussian transition probability defined as q(x t∣x t−1)≔𝒩(x t;1−β tx t−1,β tI)≔𝑞 conditional subscript 𝑥 𝑡 subscript 𝑥 𝑡 1 𝒩 subscript 𝑥 𝑡 1 subscript 𝛽 𝑡 subscript 𝑥 𝑡 1 subscript 𝛽 𝑡 𝐼 q(x_{t}\mid x_{t-1})\coloneqq\mathcal{N}(x_{t};\sqrt{1-\beta_{t}}x_{t-1},\beta% _{t}I)italic_q ( italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT ∣ italic_x start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT ) ≔ caligraphic_N ( italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT ; square-root start_ARG 1 - italic_β start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT end_ARG italic_x start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT , italic_β start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT italic_I ), where β t subscript 𝛽 𝑡\beta_{t}italic_β start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT follows a variance schedule. If T 𝑇 T italic_T is sufficiently large, x T subscript 𝑥 𝑇 x_{T}italic_x start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT will converge to pure Gaussian noise x T∼𝒩(0,I)similar-to subscript 𝑥 𝑇 𝒩 0 𝐼 x_{T}\sim\mathcal{N}(0,I)italic_x start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT ∼ caligraphic_N ( 0 , italic_I ). The reverse diffusion process aims to revert each diffusion step, allowing the generation of an image from the target distribution starting from pure noise x T subscript 𝑥 𝑇 x_{T}italic_x start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT. The reverse transition probability has a Gaussian closed form, q(x t−1∣x t,x 0)=𝒩(x t−1∣μ~(x 0,x t),β~t)𝑞 conditional subscript 𝑥 𝑡 1 subscript 𝑥 𝑡 subscript 𝑥 0 𝒩 conditional subscript 𝑥 𝑡 1~𝜇 subscript 𝑥 0 subscript 𝑥 𝑡 subscript~𝛽 𝑡 q(x_{t-1}\mid x_{t},x_{0})=\mathcal{N}(x_{t-1}\mid\tilde{\mu}(x_{0},x_{t}),% \tilde{\beta}_{t})italic_q ( italic_x start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT ∣ italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_x start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT ) = caligraphic_N ( italic_x start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT ∣ over~ start_ARG italic_μ end_ARG ( italic_x start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT , italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT ) , over~ start_ARG italic_β end_ARG start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT ), conditioned on the real image x 0 subscript 𝑥 0 x_{0}italic_x start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT. As x 0 subscript 𝑥 0 x_{0}italic_x start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT is not available during generation, a neural network is trained to approximate μ θ(x t,t)≈μ~(x 0,x t)subscript 𝜇 𝜃 subscript 𝑥 𝑡 𝑡~𝜇 subscript 𝑥 0 subscript 𝑥 𝑡\mu_{\theta}(x_{t},t)\approx\tilde{\mu}(x_{0},x_{t})italic_μ start_POSTSUBSCRIPT italic_θ end_POSTSUBSCRIPT ( italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_t ) ≈ over~ start_ARG italic_μ end_ARG ( italic_x start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT , italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT ). Following the work proposed in[[5](https://arxiv.org/html/2405.03328v1#bib.bib5)], it is possible to reparameterise the mean in terms of x t subscript 𝑥 𝑡 x_{t}italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT and a noise term ϵ italic-ϵ\epsilon italic_ϵ, and then use a neural network to predict the noise ϵ θ(x t,t)≈ϵ subscript italic-ϵ 𝜃 subscript 𝑥 𝑡 𝑡 italic-ϵ\epsilon_{\theta}(x_{t},t)\approx\epsilon italic_ϵ start_POSTSUBSCRIPT italic_θ end_POSTSUBSCRIPT ( italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_t ) ≈ italic_ϵ, optimized with the following objective:
|
| 35 |
+
|
| 36 |
+
ℒ ϵ≔𝔼 t,x t,ϵ∼𝒩(0,I)[∥ϵ−ϵ θ(x t,t)∥2].≔subscript ℒ italic-ϵ subscript 𝔼 similar-to 𝑡 subscript 𝑥 𝑡 italic-ϵ 𝒩 0 𝐼 delimited-[]superscript delimited-∥∥italic-ϵ subscript italic-ϵ 𝜃 subscript 𝑥 𝑡 𝑡 2\mathcal{L}_{\epsilon}\coloneqq\mathbb{E}_{t,x_{t},\epsilon\sim\mathcal{N}(0,I% )}\left[\lVert\epsilon-\epsilon_{\theta}(x_{t},t)\rVert^{2}\right].caligraphic_L start_POSTSUBSCRIPT italic_ϵ end_POSTSUBSCRIPT ≔ blackboard_E start_POSTSUBSCRIPT italic_t , italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_ϵ ∼ caligraphic_N ( 0 , italic_I ) end_POSTSUBSCRIPT [ ∥ italic_ϵ - italic_ϵ start_POSTSUBSCRIPT italic_θ end_POSTSUBSCRIPT ( italic_x start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_t ) ∥ start_POSTSUPERSCRIPT 2 end_POSTSUPERSCRIPT ] .(1)
|
| 37 |
+
|
| 38 |
+
An LDM[[16](https://arxiv.org/html/2405.03328v1#bib.bib16)] extends the DDPM by applying the diffusion process to a latent representation z 𝑧 z italic_z of the image x 𝑥 x italic_x, rather than to the image itself. This approach reduces the high memory demand while preserving the quality and flexibility of the models. The latent representation is obtained by training an autoencoder, composed of an encoder ℰ ℰ\mathcal{E}caligraphic_E and a decoder 𝒟,𝒟\mathcal{D},caligraphic_D , such that the encoder maps the sample x 𝑥 x italic_x to the latent space z=ℰ(x),𝑧 ℰ 𝑥 z=\mathcal{E}(x),italic_z = caligraphic_E ( italic_x ) , and the decoder recovers it as x=𝒟(z)𝑥 𝒟 𝑧 x=\mathcal{D}(z)italic_x = caligraphic_D ( italic_z ).
|
| 39 |
+
|
| 40 |
+
|
| 41 |
+
|
| 42 |
+
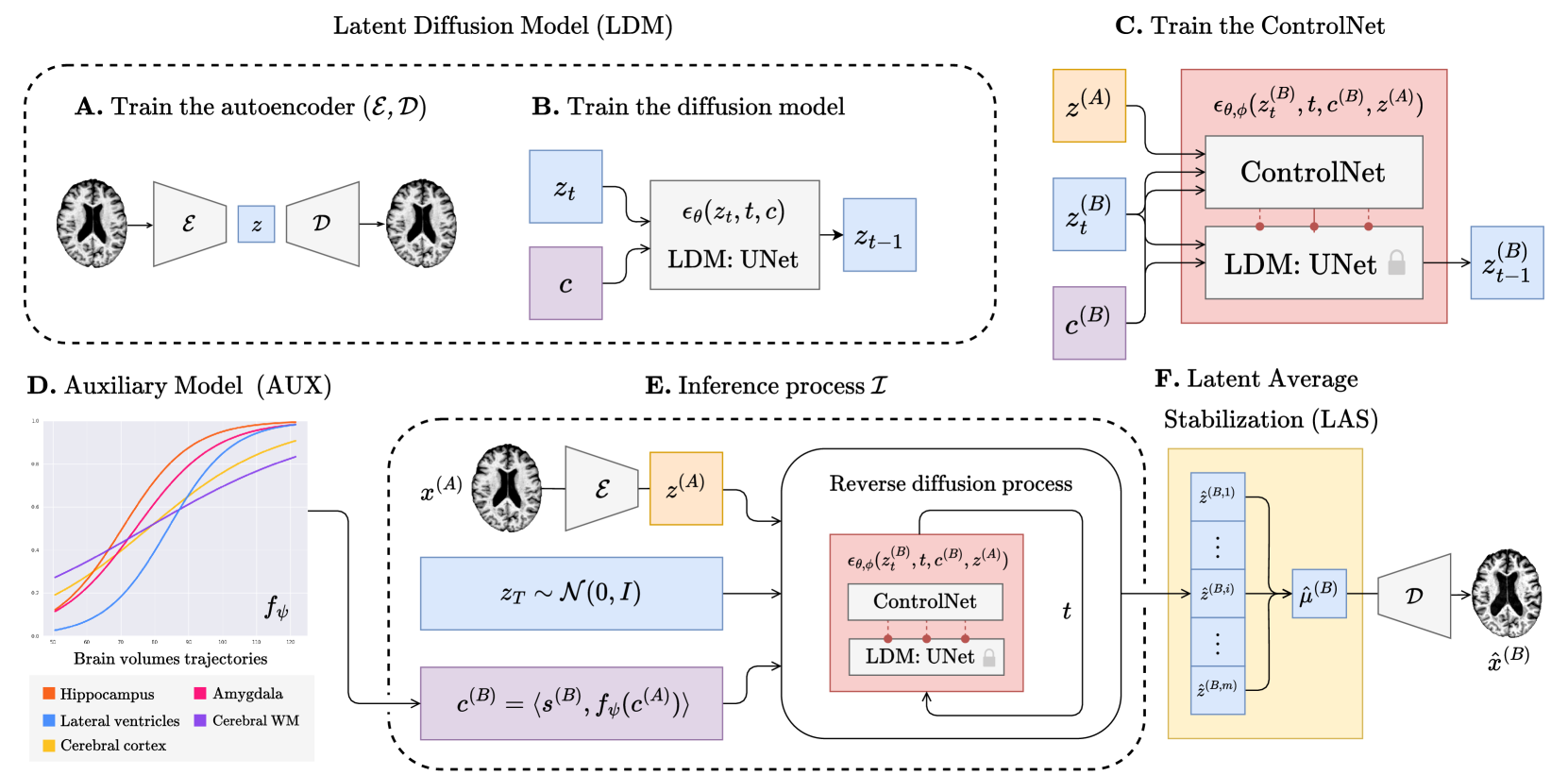
Figure 1: The overview of BrLP training and inference processes.
|
| 43 |
+
|
| 44 |
+
### 2.2 Proposed Pipeline - Brain Latent Progression (BrLP)
|
| 45 |
+
|
| 46 |
+
We now introduce the architecture of BrLP, comprising four key components: an LDM, a ControlNet, an auxiliary model, and a LAS block, each described in successive paragraphs. These four components, summarized in Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge"), collectively address the challenges outlined in the introduction. In particular, the LDM is designed to generate random 3D brain MRIs that conform to specific covariates, while ControlNet aims to specialize these MRI scans to specific anatomical structures of a subject. Additionally, the auxiliary model leverages prior knowledge of disease progression to improve the precision in predicting the volumetric changes of specific brain regions. Finally, the LAS block is used during inference to improve spatiotemporal consistency. Details concerning the training process and hyperparameter settings are provided in Table 1 of the Supplementary Material.
|
| 47 |
+
|
| 48 |
+
#### 2.2.1 LDM - Learning the brain MRIs distribution.
|
| 49 |
+
|
| 50 |
+
Building upon[[12](https://arxiv.org/html/2405.03328v1#bib.bib12)], we train an LDM aimed to generate 3D brain MRIs mirroring specific covariates c=⟨s,v⟩𝑐 𝑠 𝑣 c=\langle s,v\rangle italic_c = ⟨ italic_s , italic_v ⟩, where s 𝑠 s italic_s includes subject-specific metadata (age, sex, and cognitive status) while v 𝑣 v italic_v encompasses progression-related metrics such as volumes of brain regions (hippocampus, cerebral cortex, amygdala, cerebral white matter, and lateral ventricles) linked to AD progression[[13](https://arxiv.org/html/2405.03328v1#bib.bib13)]. The construction of the LDM is a two-phase process. Initially, we train an autoencoder (ℰ,𝒟)ℰ 𝒟(\mathcal{E},\mathcal{D})( caligraphic_E , caligraphic_D ) (block A in Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")) designed to produce a latent representation z=ℰ(x)𝑧 ℰ 𝑥 z=\mathcal{E}(x)italic_z = caligraphic_E ( italic_x ) for each brain MRI x 𝑥 x italic_x within our dataset. Subsequently, we train a conditional UNet (block B in Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")), represented as ϵ θ subscript italic-ϵ 𝜃\epsilon_{\theta}italic_ϵ start_POSTSUBSCRIPT italic_θ end_POSTSUBSCRIPT, with network parameters θ 𝜃\theta italic_θ, aimed to estimate the noise ϵ θ(z t,t,c)subscript italic-ϵ 𝜃 subscript 𝑧 𝑡 𝑡 𝑐\epsilon_{\theta}(z_{t},t,c)italic_ϵ start_POSTSUBSCRIPT italic_θ end_POSTSUBSCRIPT ( italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_t , italic_c ) necessary for reverting from z t subscript 𝑧 𝑡 z_{t}italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT to z t−1 subscript 𝑧 𝑡 1 z_{t-1}italic_z start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT, as mentioned in Section [2.1](https://arxiv.org/html/2405.03328v1#S2.SS1 "2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge"). We train ϵ θ subscript italic-ϵ 𝜃\epsilon_{\theta}italic_ϵ start_POSTSUBSCRIPT italic_θ end_POSTSUBSCRIPT by minimizing the loss ℒ ϵ subscript ℒ italic-ϵ\mathcal{L}_{\epsilon}caligraphic_L start_POSTSUBSCRIPT italic_ϵ end_POSTSUBSCRIPT (Eq.[1](https://arxiv.org/html/2405.03328v1#S2.E1 "In 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")). Covariates c 𝑐 c italic_c are integrated into the network as conditions using a cross-attention mechanism, in line with[[16](https://arxiv.org/html/2405.03328v1#bib.bib16)]. The generation process initiates by sampling random Gaussian noise z T∼𝒩(0,I)similar-to subscript 𝑧 𝑇 𝒩 0 𝐼 z_{T}\sim\mathcal{N}(0,I)italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT ∼ caligraphic_N ( 0 , italic_I ) and then iteratively reverses each diffusion step z t→z t−1→subscript 𝑧 𝑡 subscript 𝑧 𝑡 1 z_{t}\to z_{t-1}italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT → italic_z start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT for t=T,…,1 𝑡 𝑇…1 t=T,\dots,1 italic_t = italic_T , … , 1. Decoding the output z 0 subscript 𝑧 0 z_{0}italic_z start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT from the final step t=1 𝑡 1 t=1 italic_t = 1 yields a synthetic brain MRI x^=𝒟(z 0)^𝑥 𝒟 subscript 𝑧 0\hat{x}=\mathcal{D}(z_{0})over^ start_ARG italic_x end_ARG = caligraphic_D ( italic_z start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT ) that follows the specified covariates c 𝑐 c italic_c.
|
| 51 |
+
|
| 52 |
+
#### 2.2.2 ControlNet - Conditioning on subject brain MRI.
|
| 53 |
+
|
| 54 |
+
The LDM provides only a limited degree of control over the generated brain MRI via the covariates c 𝑐 c italic_c, and it does not allow for conditioning the model on individual anatomical structures. The purpose of this block is to extend the capabilities of the LDM to encompass this additional control. To achieve this, we use ControlNet[[27](https://arxiv.org/html/2405.03328v1#bib.bib27)], (block C in Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")) a neural network designed to work in conjunction with the LDM. We conceptualize ControlNet and LDM as a unified network ϵ θ,ϕ subscript italic-ϵ 𝜃 italic-ϕ\epsilon_{\theta,\phi}italic_ϵ start_POSTSUBSCRIPT italic_θ , italic_ϕ end_POSTSUBSCRIPT, where θ 𝜃\theta italic_θ represents the fixed network’s parameters of the LDM and ϕ italic-ϕ\phi italic_ϕ denotes the trainable network’s parameters of ControlNet. As in the LDM, ϵ θ,ϕ subscript italic-ϵ 𝜃 italic-ϕ\epsilon_{\theta,\phi}italic_ϵ start_POSTSUBSCRIPT italic_θ , italic_ϕ end_POSTSUBSCRIPT is still used to predict the noise ϵ θ,ϕ(z t,t,c,z)subscript italic-ϵ 𝜃 italic-ϕ subscript 𝑧 𝑡 𝑡 𝑐 𝑧\epsilon_{\theta,\phi}(z_{t},t,c,z)italic_ϵ start_POSTSUBSCRIPT italic_θ , italic_ϕ end_POSTSUBSCRIPT ( italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_t , italic_c , italic_z ) in the reverse diffusion step z t→z t−1→subscript 𝑧 𝑡 subscript 𝑧 𝑡 1 z_{t}\to z_{t-1}italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT → italic_z start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT, now incorporating z=ℰ(x)𝑧 ℰ 𝑥 z=\mathcal{E}(x)italic_z = caligraphic_E ( italic_x ) as a condition to encompass the structure of the target brain x 𝑥 x italic_x during the generation process. To train ControlNet, we use the latent representations z(A)superscript 𝑧 𝐴 z^{(A)}italic_z start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT and z(B)superscript 𝑧 𝐵 z^{(B)}italic_z start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT from pairs of brain MRIs of the same patient taken at different ages A 𝐴 A italic_A<B 𝐵 B italic_B. The covariates c(B)superscript 𝑐 𝐵 c^{(B)}italic_c start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT associated with z(B)superscript 𝑧 𝐵 z^{(B)}italic_z start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT are known and used as target covariates. Each training iteration involves: i) sampling t∼U[1,T]similar-to 𝑡 𝑈 1 𝑇 t\sim U[1,T]italic_t ∼ italic_U [ 1 , italic_T ], ii) performing t 𝑡 t italic_t forward diffusion steps z(B)→z t(B)→superscript 𝑧 𝐵 superscript subscript 𝑧 𝑡 𝐵 z^{(B)}\to z_{t}^{(B)}italic_z start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT → italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT, iii) predicting the noise ϵ θ,ϕ(z t(B),t,c(B),z(A))subscript italic-ϵ 𝜃 italic-ϕ superscript subscript 𝑧 𝑡 𝐵 𝑡 superscript 𝑐 𝐵 superscript 𝑧 𝐴\epsilon_{\theta,\phi}(z_{t}^{(B)},t,c^{(B)},z^{(A)})italic_ϵ start_POSTSUBSCRIPT italic_θ , italic_ϕ end_POSTSUBSCRIPT ( italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT , italic_t , italic_c start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT , italic_z start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) to revert z t(B)→z t−1(B)→subscript superscript 𝑧 𝐵 𝑡 subscript superscript 𝑧 𝐵 𝑡 1 z^{(B)}_{t}\to z^{(B)}_{t-1}italic_z start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT → italic_z start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT start_POSTSUBSCRIPT italic_t - 1 end_POSTSUBSCRIPT, and iv) minimizing the loss ℒ ϵ subscript ℒ italic-ϵ\mathcal{L}_{\epsilon}caligraphic_L start_POSTSUBSCRIPT italic_ϵ end_POSTSUBSCRIPT (Eq.[1](https://arxiv.org/html/2405.03328v1#S2.E1 "In 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")).
|
| 55 |
+
|
| 56 |
+
#### 2.2.3 Proposed auxiliary model - Leveraging disease prior knowledge.
|
| 57 |
+
|
| 58 |
+
AD-related regions shrink or expand over time and at different rates[[13](https://arxiv.org/html/2405.03328v1#bib.bib13)]. Deep-learning-based spatiotemporal models strive to learn these progression rates directly from brain MRIs in a black-box manner, which can be very challenging. To aid this process, we propose incorporating prior knowledge of volumetric changes directly into our pipeline. To do so, we exploit an auxiliary model f ψ subscript 𝑓 𝜓 f_{\psi}italic_f start_POSTSUBSCRIPT italic_ψ end_POSTSUBSCRIPT (block D in Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")) able to predict how the volumes of AD-related regions change over time and provide this information to the LDM via the progression-related covariates v 𝑣 v italic_v. The choice of our auxiliary model is tailored to two scenarios, making BrLP flexible for both cross-sectional and longitudinal data. For subjects with a single scan available at age A 𝐴 A italic_A, we employ a regression model to estimate volumetric changes v^(B)=f ψ(c(A))superscript^𝑣 𝐵 subscript 𝑓 𝜓 superscript 𝑐 𝐴\hat{v}^{(B)}=f_{\psi}(c^{(A)})over^ start_ARG italic_v end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = italic_f start_POSTSUBSCRIPT italic_ψ end_POSTSUBSCRIPT ( italic_c start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) at age B 𝐵 B italic_B. For subjects with n 𝑛 n italic_n past visits accessible at ages A 1,…,A n subscript 𝐴 1…subscript 𝐴 𝑛 A_{1},\dots,A_{n}italic_A start_POSTSUBSCRIPT 1 end_POSTSUBSCRIPT , … , italic_A start_POSTSUBSCRIPT italic_n end_POSTSUBSCRIPT, we predict v^(B)=fψ(c(A 1),…,c(A n))superscript^𝑣 𝐵 𝑓 𝜓 superscript 𝑐 subscript 𝐴 1…superscript 𝑐 subscript 𝐴 𝑛\hat{v}^{(B)}=f\psi(c^{(A_{1})},\dots,c^{(A_{n})})over^ start_ARG italic_v end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = italic_f italic_ψ ( italic_c start_POSTSUPERSCRIPT ( italic_A start_POSTSUBSCRIPT 1 end_POSTSUBSCRIPT ) end_POSTSUPERSCRIPT , … , italic_c start_POSTSUPERSCRIPT ( italic_A start_POSTSUBSCRIPT italic_n end_POSTSUBSCRIPT ) end_POSTSUPERSCRIPT ) using Disease Course Mapping (DCM)[[18](https://arxiv.org/html/2405.03328v1#bib.bib18), [8](https://arxiv.org/html/2405.03328v1#bib.bib8)], a model specifically designed for disease progression. DCM is intended to provide a more accurate trajectory in alignment with the subject’s history of volumetric changes available. While we employ DCM as a potential solution, any suitable disease progression model can be used in BrLP.
|
| 59 |
+
|
| 60 |
+
#### 2.2.4 Inference process.
|
| 61 |
+
|
| 62 |
+
Let x(A)superscript 𝑥 𝐴 x^{(A)}italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT be the input brain MRI from a subject at age A 𝐴 A italic_A, with known subject-specific metadata s(A)superscript 𝑠 𝐴 s^{(A)}italic_s start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT and progression-related volumes v(A)superscript 𝑣 𝐴 v^{(A)}italic_v start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT measured from x(A)superscript 𝑥 𝐴 x^{(A)}italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT. As summarized in block E from Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge"), to infer the brain MRI x(B)superscript 𝑥 𝐵 x^{(B)}italic_x start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT at age B>A 𝐵 𝐴 B>A italic_B > italic_A, we perform six steps: i) predict the progression-related volumes v^(B)=f ψ(c(A))superscript^𝑣 𝐵 subscript 𝑓 𝜓 superscript 𝑐 𝐴\hat{v}^{(B)}=f_{\psi}(c^{(A)})over^ start_ARG italic_v end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = italic_f start_POSTSUBSCRIPT italic_ψ end_POSTSUBSCRIPT ( italic_c start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) using the auxiliary model; ii) concatenate this information with the subject-specific metadata s(B)superscript 𝑠 𝐵 s^{(B)}italic_s start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT to form the target covariates c(B)=⟨s(B),v^(B)⟩superscript 𝑐 𝐵 superscript 𝑠 𝐵 superscript^𝑣 𝐵 c^{(B)}=\langle s^{(B)},\hat{v}^{(B)}\rangle italic_c start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = ⟨ italic_s start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT , over^ start_ARG italic_v end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT ⟩; iii) compute the latent z(A)=ℰ(x(A))superscript 𝑧 𝐴 ℰ superscript 𝑥 𝐴 z^{(A)}=\mathcal{E}(x^{(A)})italic_z start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT = caligraphic_E ( italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ); iv) sample random Gaussian noise z T∼𝒩(0,I)similar-to subscript 𝑧 𝑇 𝒩 0 𝐼 z_{T}\sim\mathcal{N}(0,I)italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT ∼ caligraphic_N ( 0 , italic_I ); v) run the reverse diffusion process by predicting the noise ϵ θ,ϕ(z t,t,c(B),z(A))subscript italic-ϵ 𝜃 italic-ϕ subscript 𝑧 𝑡 𝑡 superscript 𝑐 𝐵 superscript 𝑧 𝐴\epsilon_{\theta,\phi}(z_{t},t,c^{(B)},z^{(A)})italic_ϵ start_POSTSUBSCRIPT italic_θ , italic_ϕ end_POSTSUBSCRIPT ( italic_z start_POSTSUBSCRIPT italic_t end_POSTSUBSCRIPT , italic_t , italic_c start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT , italic_z start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) to reverse each diffusion step for t=T,…,1 𝑡 𝑇…1 t=T,\dots,1 italic_t = italic_T , … , 1; and finally vi) employ the decoder 𝒟 𝒟\mathcal{D}caligraphic_D to reconstruct the predicted brain MRI x^(B)=𝒟(z 0)superscript^𝑥 𝐵 𝒟 subscript 𝑧 0\hat{x}^{(B)}=\mathcal{D}(z_{0})over^ start_ARG italic_x end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = caligraphic_D ( italic_z start_POSTSUBSCRIPT 0 end_POSTSUBSCRIPT ) in the imaging domain. This inference process is summarized into a compact notation z^(B)=ℐ(z T,x(A),c(A))superscript^𝑧 𝐵 ℐ subscript 𝑧 𝑇 superscript 𝑥 𝐴 superscript 𝑐 𝐴\hat{z}^{(B)}=\mathcal{I}(z_{T},x^{(A)},c^{(A)})over^ start_ARG italic_z end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = caligraphic_I ( italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT , italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT , italic_c start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) and x^(B)=𝒟(z^(B))superscript^𝑥 𝐵 𝒟 superscript^𝑧 𝐵\hat{x}^{(B)}=\mathcal{D}(\hat{z}^{(B)})over^ start_ARG italic_x end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = caligraphic_D ( over^ start_ARG italic_z end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT ).
|
| 63 |
+
|
| 64 |
+
#### 2.2.5 Enhance inference via proposed Latent Average Stabilization (LAS).
|
| 65 |
+
|
| 66 |
+
Variations in the initial value x T∼𝒩(0,I)similar-to subscript 𝑥 𝑇 𝒩 0 𝐼 x_{T}\sim\mathcal{N}(0,I)italic_x start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT ∼ caligraphic_N ( 0 , italic_I ) can lead to slight discrepancies in the results produced by the inference process. These discrepancies are especially noticeable when making predictions over successive timesteps, manifesting as irregular patterns or non-smooth transitions of progression. Therefore, we introduce LAS (block F in Figure[1](https://arxiv.org/html/2405.03328v1#S2.F1 "Figure 1 ‣ 2.1 Background - Diffusion Models ‣ 2 Methods ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge")), a technique to improve spatiotemporal consistency by averaging different results of the inference process. In particular, LAS is based on the assumption that the predictions z^(B)=ℐ(z T,x(A),c(A))superscript^𝑧 𝐵 ℐ subscript 𝑧 𝑇 superscript 𝑥 𝐴 superscript 𝑐 𝐴\hat{z}^{(B)}=\mathcal{I}(z_{T},x^{(A)},c^{(A)})over^ start_ARG italic_z end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = caligraphic_I ( italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT , italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT , italic_c start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) deviate from a theoretical mean μ(B)=𝔼[z^(B)]superscript 𝜇 𝐵 𝔼 delimited-[]superscript^𝑧 𝐵\mu^{(B)}=\mathbb{E}[\hat{z}^{(B)}]italic_μ start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = blackboard_E [ over^ start_ARG italic_z end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT ]. To estimate the expected value μ(B)superscript 𝜇 𝐵\mu^{(B)}italic_μ start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT, we propose to repeat the inference process m 𝑚 m italic_m times and average the results:
|
| 67 |
+
|
| 68 |
+
μ(B)=𝔼 z T∼𝒩(0,I)[ℐ(z T,x(A),c(A))]≈1 m∑m ℐ(z T,x(A),c(A)).superscript 𝜇 𝐵 subscript 𝔼 similar-to subscript 𝑧 𝑇 𝒩 0 𝐼 delimited-[]ℐ subscript 𝑧 𝑇 superscript 𝑥 𝐴 superscript 𝑐 𝐴 1 𝑚 superscript 𝑚 ℐ subscript 𝑧 𝑇 superscript 𝑥 𝐴 superscript 𝑐 𝐴\mu^{(B)}=\mathop{\mathbb{E}}_{z_{T}\sim\mathcal{N}(0,I)}\bigg{[}\mathcal{I}(z% _{T},x^{(A)},c^{(A)})\bigg{]}\approx\frac{1}{m}\sum^{m}\mathcal{I}(z_{T},x^{(A% )},c^{(A)}).italic_μ start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = blackboard_E start_POSTSUBSCRIPT italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT ∼ caligraphic_N ( 0 , italic_I ) end_POSTSUBSCRIPT [ caligraphic_I ( italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT , italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT , italic_c start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) ] ≈ divide start_ARG 1 end_ARG start_ARG italic_m end_ARG ∑ start_POSTSUPERSCRIPT italic_m end_POSTSUPERSCRIPT caligraphic_I ( italic_z start_POSTSUBSCRIPT italic_T end_POSTSUBSCRIPT , italic_x start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT , italic_c start_POSTSUPERSCRIPT ( italic_A ) end_POSTSUPERSCRIPT ) .(2)
|
| 69 |
+
|
| 70 |
+
Similar to before, we decode the predicted scan as x^(B)=𝒟(μ(B))superscript^𝑥 𝐵 𝒟 superscript 𝜇 𝐵\hat{x}^{(B)}=\mathcal{D}(\mu^{(B)})over^ start_ARG italic_x end_ARG start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT = caligraphic_D ( italic_μ start_POSTSUPERSCRIPT ( italic_B ) end_POSTSUPERSCRIPT ). The entire inference process (with m=4 𝑚 4 m=4 italic_m = 4) requires ∼similar-to\sim∼4.8s per MRI on a consumer GPU.
|
| 71 |
+
|
| 72 |
+
3 Experiments and Results
|
| 73 |
+
-------------------------
|
| 74 |
+
|
| 75 |
+
#### 3.0.1 Data.
|
| 76 |
+
|
| 77 |
+
We collect a large dataset comprising 11,730 T1-weighted brain MRI scans from 2,805 subjects across various publicly available longitudinal studies: ADNI 1/2/3/GO (1,990 subjects)[[11](https://arxiv.org/html/2405.03328v1#bib.bib11)], OASIS-3 (573 subjects)[[9](https://arxiv.org/html/2405.03328v1#bib.bib9)], and AIBL (242 subjects)[[3](https://arxiv.org/html/2405.03328v1#bib.bib3)]. Each subject has at least two MRIs, and each scan is acquired during a different visit. Age, sex, and cognitive status were available from all datasets. The average age is 74±7 plus-or-minus 74 7 74\pm 7 74 ± 7 years, and 53% of the subjects are male. Based on the final visit, 43.8% of subjects are classified as CN, 25.7% exhibit or develop MCI, and 30.5% exhibit or develop AD. We randomly split data into a training set (80%), a validation set (5%), and a testing set (15%) with no overlapping subjects. The validation set is used for early stopping during training. Each brain MRI is pre-processed using: N4 bias-field correction[[21](https://arxiv.org/html/2405.03328v1#bib.bib21)], skull stripping[[6](https://arxiv.org/html/2405.03328v1#bib.bib6)], affine registration to the MNI space, intensity normalization[[19](https://arxiv.org/html/2405.03328v1#bib.bib19)] and resampling to 1.5 mm 3. The volumes used as progression-related covariates and for our subsequent evaluation are calculated using SynthSeg 2.0[[1](https://arxiv.org/html/2405.03328v1#bib.bib1)] and are expressed as percentages of the total brain volume to account for individual differences.
|
| 78 |
+
|
| 79 |
+
#### 3.0.2 Evaluation metrics.
|
| 80 |
+
|
| 81 |
+
We evaluate BrLP using image-based and volumetric metrics to compare the predicted brain MRI scans with the subjects’ actual follow-up scans. In particular, the Mean Squared Error (MSE) and the Structural Similarity Index (SSIM) are used to assess image similarity between the scans. Instead, volumetric metrics in AD-related regions (hippocampus, amygdala, lateral ventricles, cerebrospinal fluid (CSF), and thalamus) evaluate the model’s accuracy in tracking disease progression. Specifically, the Mean Absolute Error (MAE) between the volumes of actual follow-up scans and the generated brain MRIs is reported in the results. Notably, CSF and thalamus are excluded from progression-related covariates, enabling the analysis of unconditioned regions in our predictions.
|
| 82 |
+
|
| 83 |
+
#### 3.0.3 Ablation study.
|
| 84 |
+
|
| 85 |
+
We conduct an ablation study to assess the contributions of: i) the auxiliary model (AUX) and ii) the proposed technique for spatiotemporal consistency (LAS). The results are presented at the top of Table[1](https://arxiv.org/html/2405.03328v1#S3.T1 "Table 1 ‣ 3.0.3 Ablation study. ‣ 3 Experiments and Results ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge"). BrLP without AUX and LAS is referred to as “base”. The experiments demonstrate that both LAS and AUX enhance performance, reducing volumetric errors by 5% and 4%, respectively. An example of improvement achieved with LAS is provided in Figure 2 of the Supplementary Material. Employing both AUX and LAS together offers the optimal setup, achieving an average reduction in volumetric error of 7%. This optimal configuration is used for comparisons against other approaches, with the only variation being the type of auxiliary model used in our pipeline.
|
| 86 |
+
|
| 87 |
+
Table 1: Results from the ablation study and comparison with baseline methods. MAE (± SD) in predicted volumes is expressed as a percentage of total brain volume.
|
| 88 |
+
|
| 89 |
+
Config.Image-based metrics MAE (conditional region volumes)MAE (unconditional reg. volumes)
|
| 90 |
+
Method(AUX)MSE ↓↓\downarrow↓SSIM ↑↑\uparrow↑Hippocampus ↓↓\downarrow↓Amygdala ↓↓\downarrow↓Lat. Ventricle ↓↓\downarrow↓Thalamus ↓↓\downarrow↓CSF ↓↓\downarrow↓
|
| 91 |
+
Ablation Base-0.005 ± 0.003 0.89 ± 0.03 0.026 ± 0.023 0.016 ± 0.015 0.279 ± 0.347 0.030 ± 0.023 0.889 ± 0.681
|
| 92 |
+
Base + AUX LM 0.005 ± 0.002 0.90 ± 0.03 0.024 ± 0.021 0.015 ± 0.013 0.279 ± 0.314 0.030 ± 0.024 0.851 ± 0.632
|
| 93 |
+
Base + LAS-0.005 ± 0.002 0.90 ± 0.03 0.025 ± 0.022 0.015 ± 0.014 0.258 ± 0.330 0.029 ± 0.022 0.851 ± 0.659
|
| 94 |
+
Base + LAS + AUX LM 0.004 ± 0.002 0.91 ± 0.03 0.023 ± 0.021 0.015 ± 0.014 0.255 ± 0.303 0.029 ± 0.023 0.829 ± 0.624
|
| 95 |
+
Comparison Study Single-image (Cross-sectional)
|
| 96 |
+
DaniNet[[15](https://arxiv.org/html/2405.03328v1#bib.bib15)]-0.016 ± 0.007 0.62 ± 0.16 0.030 ± 0.030 0.018 ± 0.017 0.257 ± 0.222 0.038 ± 0.030 1.081 ± 0.814
|
| 97 |
+
CounterSynth[[14](https://arxiv.org/html/2405.03328v1#bib.bib14)]-0.010 ± 0.004 0.82 ± 0.05 0.030 ± 0.018 0.014 ± 0.010 0.310 ± 0.311 0.127 ± 0.035 0.881 ± 0.672
|
| 98 |
+
BrLP (Proposed)LM 0.004 ± 0.002 0.91 ± 0.03 0.023 ± 0.021 0.015 ± 0.014 0.255 ± 0.303 0.029 ± 0.023 0.829 ± 0.624
|
| 99 |
+
Sequence-aware (Longitudinal)
|
| 100 |
+
Latent-SADM[[24](https://arxiv.org/html/2405.03328v1#bib.bib24)]-0.008 ± 0.002 0.85 ± 0.02 0.035 ± 0.027 0.018 ± 0.015 0.329 ± 0.328 0.037 ± 0.028 0.924 ± 0.705
|
| 101 |
+
BrLP (Proposed)DCM 0.004 ± 0.002 0.91 ± 0.03 0.020 ± 0.017 0.014 ± 0.013 0.240 ± 0.259 0.031 ± 0.024 0.810 ± 0.631
|
| 102 |
+
|
| 103 |
+

|
| 104 |
+
|
| 105 |
+
Figure 2: A comparison between the real progression of a 70 y.o. subject with MCI over 15 years and the predictions obtained by BrLP and the baseline methods. Each method shows a predicted MRI (left) and its deviation from the subject’s real brain MRI (right).
|
| 106 |
+
|
| 107 |
+
#### 3.0.4 Comparison with baselines.
|
| 108 |
+
|
| 109 |
+
We categorize existing methods into single-image (cross-sectional) and sequence-aware (longitudinal) approaches. Single-image approaches, such as DaniNet[[15](https://arxiv.org/html/2405.03328v1#bib.bib15)] and CounterSynth[[14](https://arxiv.org/html/2405.03328v1#bib.bib14)], predict progression using just one brain MRI as input. Sequence-aware methods, like SADM[[24](https://arxiv.org/html/2405.03328v1#bib.bib24)], leverage a series of prior brain MRIs as input. Due to the large memory demands of SADM, we have re-implemented it using an LDM, allowing the comparisons in our experiments. We refer to it as Latent-SADM. To evaluate all these methods, we conduct two separate experiments. In single-image methods, we predict all subsequent MRIs for a subject based on their initial scan. For sequence-aware methods, we use the first half of a subject’s MRI visits to predict all subsequent MRIs in the latter half. In single-image settings, our approach uses a Linear Model (LM) as the auxiliary model. In contrast, for sequence-aware experiments, we employ the last available MRI in the sequence as the input for BrLP and fit a logistic DCM on the first half of the subject’s visits as the auxiliary model.
|
| 110 |
+
|
| 111 |
+
Results from our experiments are presented in Table 1. We observe an average decrease of 62% (SD = 10%) in MSE and an average increase of 43% (SD = 18%) in SSIM compared to other baselines. In terms of volumetric measurements across various brain regions, our method shows improvements of 17.55% (SD = 8.79%) over DaniNet, 23.40% (SD = 28.85%) over CounterSynth, and 24.14% (SD = 10.63%) over Latent-SADM. We did not observe any particular differences in the improvement obtained in conditioned and non-conditioned regions. Additionally, paired t-tests (p< 0.001) verified the statistical significance of the observed improvements. Finally, Figure[2](https://arxiv.org/html/2405.03328v1#S3.F2 "Figure 2 ‣ 3.0.3 Ablation study. ‣ 3 Experiments and Results ‣ Enhancing Spatiotemporal Disease Progression Models via Latent Diffusion and Prior Knowledge") presents a visual comparison between the actual progression of a 70-year-old subject over 15 years and the predictions obtained by BrLP and the baseline methods. The results from Latent-SADM and DaniNet exhibit a spatiotemporal mismatch in predicting the lateral ventricles’ enlargement, whereas CounterSynth fails to capture the structural changes observed in the real progression. On the other hand, BrLP shows the most accurate prediction of the brain’s anatomical changes, confirming the previous quantitative findings. It is worth noting that we observed limitations in LAS performance in underrepresented conditions, such as ages over 90 years, resulting in slightly non-monotonic progression (see Case Study 4 in our supplementary video).
|
| 112 |
+
|
| 113 |
+
4 Conclusion
|
| 114 |
+
------------
|
| 115 |
+
|
| 116 |
+
In this work, we propose BrLP, a 3D spatiotemporal model that accurately captures the progression patterns of neurodegenerative diseases by forecasting the evolution of 3D brain MRIs at the individual level. While we have showcased the application of our pipeline on brain MRIs, BrLP holds potential for use with other imaging modalities and to model different progressive diseases. Importantly, our framework can be easily extended to integrate additional covariates, such as genetic data, providing further personalized insights into our predictions.
|
| 117 |
+
|
| 118 |
+
References
|
| 119 |
+
----------
|
| 120 |
+
|
| 121 |
+
* [1] Billot, B., Greve, D.N., Puonti, O., Thielscher, A., Van Leemput, K., Fischl, B., Dalca, A.V., Iglesias, J.E., et al.: Synthseg: Segmentation of brain mri scans of any contrast and resolution without retraining. Medical image analysis 86, 102789 (2023)
|
| 122 |
+
* [2] Blumberg, S.B., Tanno, R., Kokkinos, I., Alexander, D.C.: Deeper image quality transfer: Training low-memory neural networks for 3d images. In: Medical Image Computing and Computer Assisted Intervention–MICCAI 2018: 21st International Conference, Granada, Spain, September 16-20, 2018, Proceedings, Part I. pp. 118–125. Springer (2018)
|
| 123 |
+
* [3] Ellis, K.A., Bush, A.I., Darby, D., De Fazio, D., Foster, J., Hudson, P., Lautenschlager, N.T., Lenzo, N., Martins, R.N., Maruff, P., et al.: The australian imaging, biomarkers and lifestyle (aibl) study of aging: methodology and baseline characteristics of 1112 individuals recruited for a longitudinal study of alzheimer’s disease. International psychogeriatrics 21(4), 672–687 (2009)
|
| 124 |
+
* [4] Eshaghi, A., Young, A.L., Wijeratne, P.A., Prados, F., Arnold, D.L., Narayanan, S., Guttmann, C.R., Barkhof, F., Alexander, D.C., Thompson, A.J., et al.: Identifying multiple sclerosis subtypes using unsupervised machine learning and mri data. Nature communications 12(1), 2078 (2021)
|
| 125 |
+
* [5] Ho, J., Jain, A., Abbeel, P.: Denoising diffusion probabilistic models. Advances in neural information processing systems 33, 6840–6851 (2020)
|
| 126 |
+
* [6] Hoopes, A., Mora, J.S., Dalca, A.V., Fischl, B., Hoffmann, M.: Synthstrip: Skull-stripping for any brain image. NeuroImage 260, 119474 (2022)
|
| 127 |
+
* [7] Jung, E., Luna, M., Park, S.H.: Conditional gan with an attention-based generator and a 3d discriminator for 3d medical image generation. In: Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part VI 24. pp. 318–328. Springer (2021)
|
| 128 |
+
* [8] Koval, I., Bône, A., Louis, M., Lartigue, T., Bottani, S., Marcoux, A., Samper-Gonzalez, J., Burgos, N., Charlier, B., Bertrand, A., et al.: Ad course map charts alzheimer’s disease progression. Scientific Reports 11(1), 8020 (2021)
|
| 129 |
+
* [9] LaMontagne, P.J., Benzinger, T.L., Morris, J.C., Keefe, S., Hornbeck, R., Xiong, C., Grant, E., Hassenstab, J., Moulder, K., Vlassenko, A.G., et al.: Oasis-3: longitudinal neuroimaging, clinical, and cognitive dataset for normal aging and alzheimer disease. MedRxiv pp. 2019–12 (2019)
|
| 130 |
+
* [10] Oxtoby, N.P., Alexander, D.C.: Imaging plus x: multimodal models of neurodegenerative disease. Current opinion in neurology 30(4), 371 (2017)
|
| 131 |
+
* [11] Petersen, R.C., Aisen, P.S., Beckett, L.A., Donohue, M.C., Gamst, A.C., Harvey, D.J., Jack, C.R., Jagust, W.J., Shaw, L.M., Toga, A.W., et al.: Alzheimer’s disease neuroimaging initiative (adni): clinical characterization. Neurology 74(3), 201–209 (2010)
|
| 132 |
+
* [12] Pinaya, W.H., Tudosiu, P.D., Dafflon, J., Da Costa, P.F., Fernandez, V., Nachev, P., Ourselin, S., Cardoso, M.J.: Brain imaging generation with latent diffusion models. In: MICCAI Workshop on Deep Generative Models. pp. 117–126. Springer (2022)
|
| 133 |
+
* [13] Pini, L., Pievani, M., Bocchetta, M., Altomare, D., Bosco, P., Cavedo, E., Galluzzi, S., Marizzoni, M., Frisoni, G.B.: Brain atrophy in alzheimer’s disease and aging. Ageing research reviews 30, 25–48 (2016)
|
| 134 |
+
* [14] Pombo, G., Gray, R., Cardoso, M.J., Ourselin, S., Rees, G., Ashburner, J., Nachev, P.: Equitable modelling of brain imaging by counterfactual augmentation with morphologically constrained 3d deep generative models. Medical Image Analysis 84, 102723 (2023)
|
| 135 |
+
* [15] Ravi, D., Blumberg, S.B., Ingala, S., Barkhof, F., Alexander, D.C., Oxtoby, N.P., Initiative, A.D.N., et al.: Degenerative adversarial neuroimage nets for brain scan simulations: Application in ageing and dementia. Medical Image Analysis 75, 102257 (2022)
|
| 136 |
+
* [16] Rombach, R., Blattmann, A., Lorenz, D., Esser, P., Ommer, B.: High-resolution image synthesis with latent diffusion models. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. pp. 10684–10695 (2022)
|
| 137 |
+
* [17] Sauty, B., Durrleman, S.: Progression models for imaging data with longitudinal variational auto encoders. In: International Conference on Medical Image Computing and Computer-Assisted Intervention. pp. 3–13. Springer (2022)
|
| 138 |
+
* [18] Schiratti, J.B., Allassonnière, S., Colliot, O., Durrleman, S.: A bayesian mixed-effects model to learn trajectories of changes from repeated manifold-valued observations. The Journal of Machine Learning Research 18(1), 4840–4872 (2017)
|
| 139 |
+
* [19] Shinohara, R.T., Sweeney, E.M., Goldsmith, J., Shiee, N., Mateen, F.J., Calabresi, P.A., Jarso, S., Pham, D.L., Reich, D.S., Crainiceanu, C.M., et al.: Statistical normalization techniques for magnetic resonance imaging. NeuroImage: Clinical 6, 9–19 (2014)
|
| 140 |
+
* [20] Tijms, B.M., Vromen, E.M., Mjaavatten, O., Holstege, H., Reus, L.M., van der Lee, S., Wesenhagen, K.E., Lorenzini, L., Vermunt, L., Venkatraghavan, V., et al.: Cerebrospinal fluid proteomics in patients with alzheimer’s disease reveals five molecular subtypes with distinct genetic risk profiles. Nature Aging pp. 1–15 (2024)
|
| 141 |
+
* [21] Tustison, N.J., Avants, B.B., Cook, P.A., Zheng, Y., Egan, A., Yushkevich, P.A., Gee, J.C.: N4itk: improved n3 bias correction. IEEE transactions on medical imaging 29(6), 1310–1320 (2010)
|
| 142 |
+
* [22] Vogel, J.W., Young, A.L., Oxtoby, N.P., Smith, R., Ossenkoppele, R., Strandberg, O.T., La Joie, R., Aksman, L.M., Grothe, M.J., Iturria-Medina, Y., et al.: Four distinct trajectories of tau deposition identified in alzheimer’s disease. Nature medicine 27(5), 871–881 (2021)
|
| 143 |
+
* [23] Xia, T., Chartsias, A., Wang, C., Tsaftaris, S.A., Initiative, A.D.N., et al.: Learning to synthesise the ageing brain without longitudinal data. Medical Image Analysis 73, 102169 (2021)
|
| 144 |
+
* [24] Yoon, J.S., Zhang, C., Suk, H.I., Guo, J., Li, X.: Sadm: Sequence-aware diffusion model for longitudinal medical image generation. In: International Conference on Information Processing in Medical Imaging. pp. 388–400. Springer (2023)
|
| 145 |
+
* [25] Young, A.L., Marinescu, R.V., Oxtoby, N.P., Bocchetta, M., Yong, K., Firth, N.C., Cash, D.M., Thomas, D.L., Dick, K.M., Cardoso, J., et al.: Uncovering the heterogeneity and temporal complexity of neurodegenerative diseases with subtype and stage inference. Nature communications 9(1), 4273 (2018)
|
| 146 |
+
* [26] Young, A.L., Oxtoby, N.P., Garbarino, S., Fox, N.C., Barkhof, F., Schott, J.M., Alexander, D.C.: Data-driven modelling of neurodegenerative disease progression: thinking outside the black box. Nature Reviews Neuroscience pp. 1–20 (2024)
|
| 147 |
+
* [27] Zhang, L., Rao, A., Agrawala, M.: Adding conditional control to text-to-image diffusion models. In: Proceedings of the IEEE/CVF International Conference on Computer Vision. pp. 3836–3847 (2023)
|
| 148 |
+
* [28] Zhao, Y., Ma, B., Jiang, P., Zeng, D., Wang, X., Li, S.: Prediction of alzheimer’s disease progression with multi-information generative adversarial network. IEEE Journal of Biomedical and Health Informatics 25(3), 711–719 (2020)
|