Update README.md
Browse files
README.md
CHANGED
|
@@ -3,15 +3,15 @@ license: apache-2.0
|
|
| 3 |
tags:
|
| 4 |
- biology
|
| 5 |
- chemistry
|
| 6 |
-
- IntFold
|
| 7 |
- biomolecular-structure-prediction
|
|
|
|
| 8 |
---
|
| 9 |
|
| 10 |
-
](https://huggingface.co/GAGABIG/CNN)
|
| 14 |
-
[](LICENSE)
|
| 16 |
[](#contact-us)
|
| 17 |
|
|
@@ -22,38 +22,38 @@ tags:
|
|
| 22 |
</div>
|
| 23 |
|
| 24 |
|
| 25 |
-
.
|
| 64 |
|
| 65 |
-
 for the most accurate, complete, and convenient biomolecular structure predictions.** It requires no installation and provides an intuitive web interface to submit your sequences and visualize results directly in your browser. The server runs the **full, optimized, latest** IntelliFold implementation for optimal performance.
|
| 71 |
|
| 72 |
-
 and [FastFold](https://github.com/hpcaitech/FastFold), following [Protenix](https://github.com/bytedance/Protenix)'s usage.
|
| 94 |
-
- Many components in `
|
| 95 |
- This repository, the implementation of **Inference Data Pipeline**(Data/Feature Processing and MSA generation tasks) referred to [Boltz-1](https://github.com/jwohlwend/boltz), and modify some codes to adapt to the input of our model.
|
| 96 |
|
| 97 |
|
| 98 |
|
| 99 |
## ⚖️ License
|
| 100 |
|
| 101 |
-
The
|
| 102 |
|
| 103 |
## 📬 Contact Us
|
| 104 |
|
|
|
|
| 3 |
tags:
|
| 4 |
- biology
|
| 5 |
- chemistry
|
|
|
|
| 6 |
- biomolecular-structure-prediction
|
| 7 |
+
- IntelliFold
|
| 8 |
---
|
| 9 |
|
| 10 |
+

|
| 11 |
|
| 12 |
+
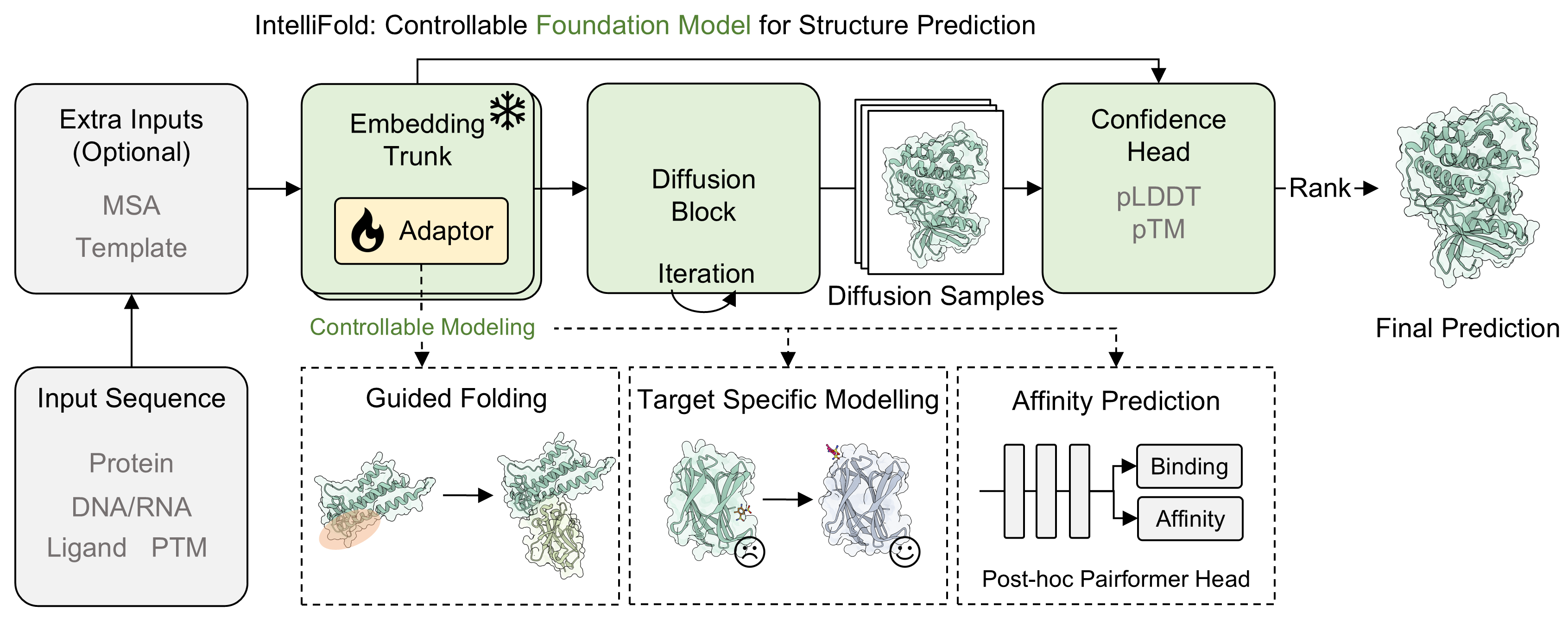
# IntelliFold: A Controllable Foundation Model for General and Specialized Biomolecular Structure Prediction.
|
| 13 |
[](https://huggingface.co/GAGABIG/CNN)
|
| 14 |
+
[](https://pypi.org/project/intellifold/)
|
| 15 |
[](LICENSE)
|
| 16 |
[](#contact-us)
|
| 17 |
|
|
|
|
| 22 |
</div>
|
| 23 |
|
| 24 |
|
| 25 |
+
|
| 26 |
|
| 27 |
|
| 28 |
## 🚀 Quick Start
|
| 29 |
|
| 30 |
To quickly get started with IntelliFold, you can use the following commands:
|
| 31 |
```bash
|
| 32 |
+
# Install IntelliFold from PyPI
|
| 33 |
+
pip install intellifold
|
| 34 |
# Run inference with an example YAML file
|
| 35 |
+
intellifold predict ./examples/5S8I_A.yaml --out_dir ./output
|
| 36 |
```
|
| 37 |
|
| 38 |
## ⚙️ Installation
|
| 39 |
|
| 40 |
+
To more complete installation instructions and usage, please refer to the [Installation Guide](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/installation.md).
|
| 41 |
|
| 42 |
|
| 43 |
## 🔍 Inference
|
| 44 |
|
| 45 |
+
1. **Prepare Input File**: Create a YAML file with your sequences following our [input format specification](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/input_yaml_format.md)
|
| 46 |
|
| 47 |
2. **Run Prediction**:
|
| 48 |
```bash
|
| 49 |
+
intellifold predict your_input.yaml --out_dir ./results
|
| 50 |
```
|
| 51 |
|
| 52 |
+
3. **Check Results**: Find predicted structures and confidence scores in the output directory, you can also check the section of **output format** in [output documentation](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/input_yaml_format.md#output-format).
|
| 53 |
|
| 54 |
+
4. **Optional Optimization**: Enable [custom kernels](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/kernels.md) for faster inference and reduced memory usage
|
| 55 |
|
| 56 |
+
For comprehensive usage instructions and examples, refer to the [Usage Guide](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/usage.md).
|
| 57 |
|
| 58 |
|
| 59 |
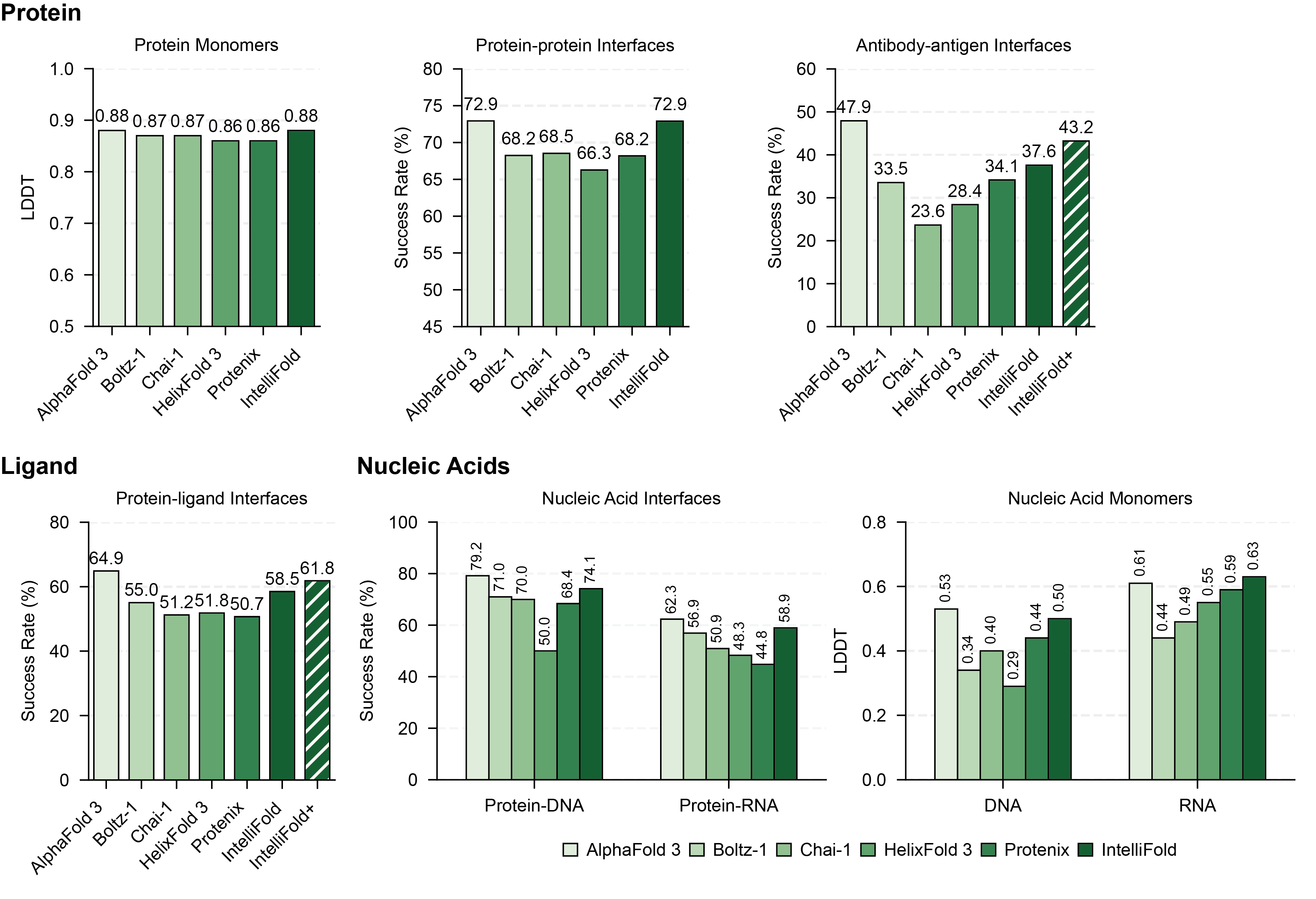
## 📊 Benchmarking
|
|
|
|
| 62 |
|
| 63 |
For more details on the benchmarking process and results, please refer to our [Technical Report](https://arxiv.org/abs/2507.02025).
|
| 64 |
|
| 65 |
+
|
| 66 |
|
| 67 |
|
| 68 |
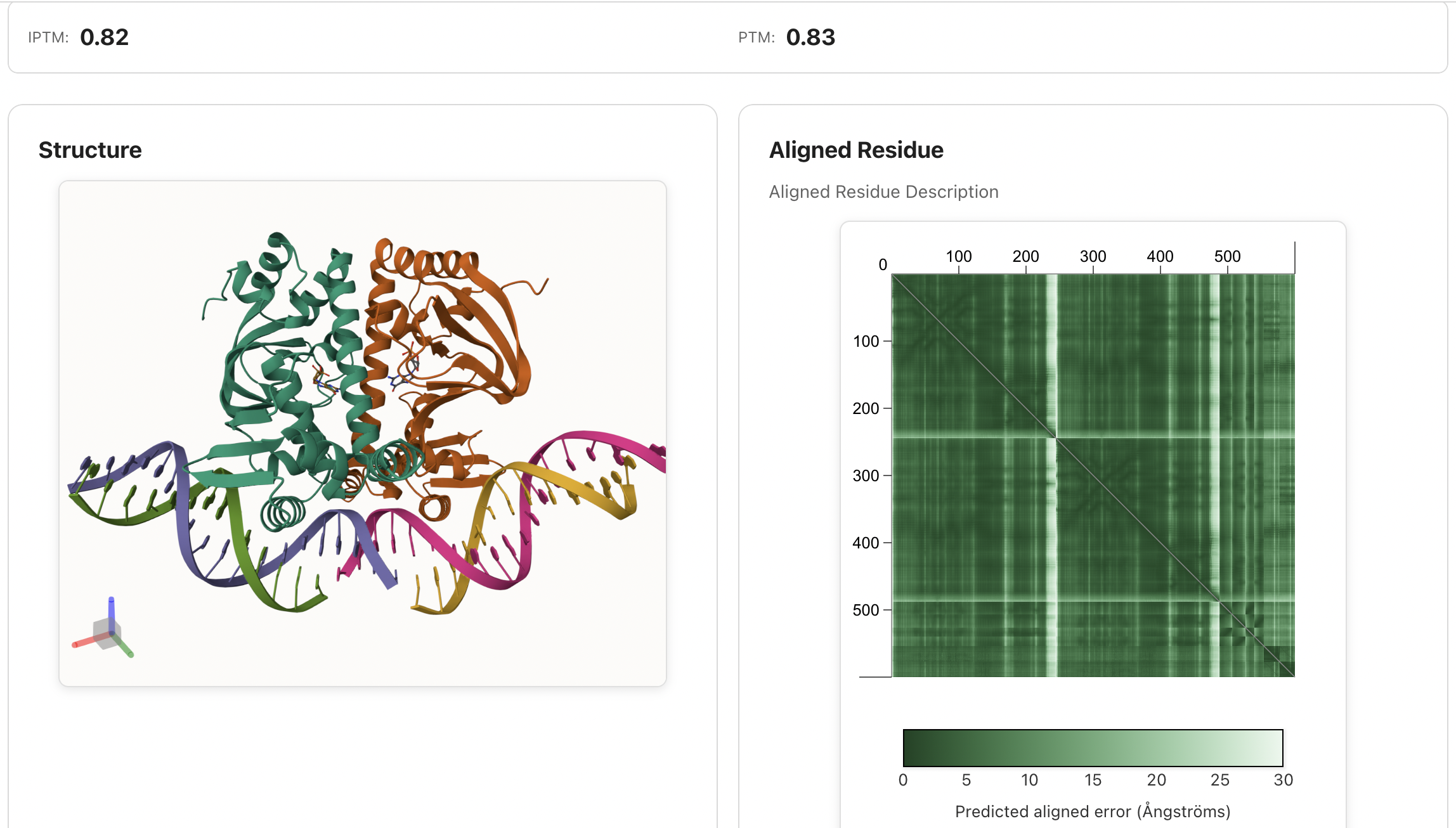
## 🌐 IntelliFold Server
|
| 69 |
|
| 70 |
**We highly recommend using the [IntelliFold Server](https://server.intfold.com) for the most accurate, complete, and convenient biomolecular structure predictions.** It requires no installation and provides an intuitive web interface to submit your sequences and visualize results directly in your browser. The server runs the **full, optimized, latest** IntelliFold implementation for optimal performance.
|
| 71 |
|
| 72 |
+
|
| 73 |
|
| 74 |
|
| 75 |
## 📜 Citation
|
|
|
|
| 91 |
## 🔗 Acknowledgements
|
| 92 |
|
| 93 |
- The implementation of **fast layernorm operators** is inspired by [OneFlow](https://github.com/Oneflow-Inc/oneflow) and [FastFold](https://github.com/hpcaitech/FastFold), following [Protenix](https://github.com/bytedance/Protenix)'s usage.
|
| 94 |
+
- Many components in `intellifold/openfold/` are adapted from [OpenFold](https://github.com/aqlaboratory/openfold), with substantial modifications and improvements by our team (except for the `LayerNorm` part).
|
| 95 |
- This repository, the implementation of **Inference Data Pipeline**(Data/Feature Processing and MSA generation tasks) referred to [Boltz-1](https://github.com/jwohlwend/boltz), and modify some codes to adapt to the input of our model.
|
| 96 |
|
| 97 |
|
| 98 |
|
| 99 |
## ⚖️ License
|
| 100 |
|
| 101 |
+
The IntelliFold project, including code and model parameters, is made available under the [Apache 2.0 License](https://github.com/IntelliGen-AI/IntelliFold/blob/main/LICENSE), it is free for both academic research and commercial use.
|
| 102 |
|
| 103 |
## 📬 Contact Us
|
| 104 |
|