add workflow, train loss and validation accuracy figures
Browse files- README.md +12 -17
- configs/metadata.json +2 -1
- docs/README.md +12 -17
README.md
CHANGED
|
@@ -12,6 +12,8 @@ A pre-trained model for the endoscopic inbody classification task.
|
|
| 12 |
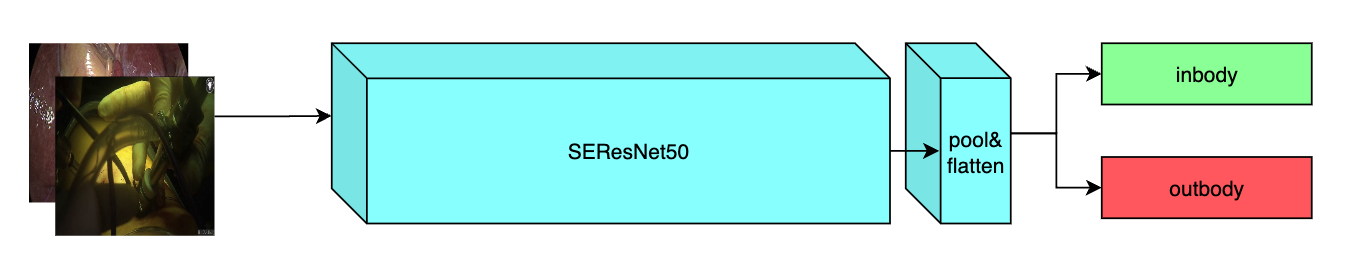
This model is trained using the SEResNet50 structure, whose details can be found in [1]. All datasets are from private samples of [Activ Surgical](https://www.activsurgical.com/). Samples in training and validation dataset are from the same 4 videos, while test samples are from different two videos.
|
| 13 |
The [pytorch model](https://drive.google.com/file/d/14CS-s1uv2q6WedYQGeFbZeEWIkoyNa-x/view?usp=sharing) and [torchscript model](https://drive.google.com/file/d/1fOoJ4n5DWKHrt9QXTZ2sXwr9C-YvVGCM/view?usp=sharing) are shared in google drive. Modify the `bundle_root` parameter specified in `configs/train.json` and `configs/inference.json` to reflect where models are downloaded. Expected directory path to place downloaded models is `models/` under `bundle_root`.
|
| 14 |
|
|
|
|
|
|
|
| 15 |
## Data
|
| 16 |
Datasets used in this work were provided by [Activ Surgical](https://www.activsurgical.com/). Here is a [link](https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/inbody_outbody_samples.zip) of 20 samples (10 in-body and 10 out-body) to show what this dataset looks like. After downloading this dataset, python script in `scripts` folder naming `data_process` can be used to get label json files by running the command below and replacing datapath and outpath parameters.
|
| 17 |
```
|
|
@@ -59,6 +61,16 @@ This model achieves the following accuracy score on the test dataset:
|
|
| 59 |
|
| 60 |
Accuracy = 0.98
|
| 61 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 62 |
## commands example
|
| 63 |
Execute training:
|
| 64 |
|
|
@@ -109,23 +121,6 @@ python -m monai.bundle ckpt_export network_def \
|
|
| 109 |
--config_file configs/inference.json
|
| 110 |
```
|
| 111 |
|
| 112 |
-
Export checkpoint to onnx file, which has been tested on pytorch 1.12.0:
|
| 113 |
-
|
| 114 |
-
```
|
| 115 |
-
python scripts/export_to_onnx.py --model models/model.pt --outpath models/model.onnx
|
| 116 |
-
```
|
| 117 |
-
|
| 118 |
-
Export TensorRT float16 model from the onnx model:
|
| 119 |
-
|
| 120 |
-
```
|
| 121 |
-
trtexec --onnx=models/model.onnx --saveEngine=models/model.trt --fp16 \
|
| 122 |
-
--minShapes=INPUT__0:1x3x256x256 \
|
| 123 |
-
--optShapes=INPUT__0:16x3x256x256 \
|
| 124 |
-
--maxShapes=INPUT__0:32x3x256x256 \
|
| 125 |
-
--shapes=INPUT__0:8x3x256x256
|
| 126 |
-
```
|
| 127 |
-
This command need TensorRT with correct CUDA installed in the environment. For the detail of installing TensorRT, please refer to [this link](https://docs.nvidia.com/deeplearning/tensorrt/install-guide/index.html).
|
| 128 |
-
|
| 129 |
# References
|
| 130 |
[1] J. Hu, L. Shen and G. Sun, Squeeze-and-Excitation Networks, 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2018, pp. 7132-7141. https://arxiv.org/pdf/1709.01507.pdf
|
| 131 |
|
|
|
|
| 12 |
This model is trained using the SEResNet50 structure, whose details can be found in [1]. All datasets are from private samples of [Activ Surgical](https://www.activsurgical.com/). Samples in training and validation dataset are from the same 4 videos, while test samples are from different two videos.
|
| 13 |
The [pytorch model](https://drive.google.com/file/d/14CS-s1uv2q6WedYQGeFbZeEWIkoyNa-x/view?usp=sharing) and [torchscript model](https://drive.google.com/file/d/1fOoJ4n5DWKHrt9QXTZ2sXwr9C-YvVGCM/view?usp=sharing) are shared in google drive. Modify the `bundle_root` parameter specified in `configs/train.json` and `configs/inference.json` to reflect where models are downloaded. Expected directory path to place downloaded models is `models/` under `bundle_root`.
|
| 14 |
|
| 15 |
+
|
| 16 |
+
|
| 17 |
## Data
|
| 18 |
Datasets used in this work were provided by [Activ Surgical](https://www.activsurgical.com/). Here is a [link](https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/inbody_outbody_samples.zip) of 20 samples (10 in-body and 10 out-body) to show what this dataset looks like. After downloading this dataset, python script in `scripts` folder naming `data_process` can be used to get label json files by running the command below and replacing datapath and outpath parameters.
|
| 19 |
```
|
|
|
|
| 61 |
|
| 62 |
Accuracy = 0.98
|
| 63 |
|
| 64 |
+
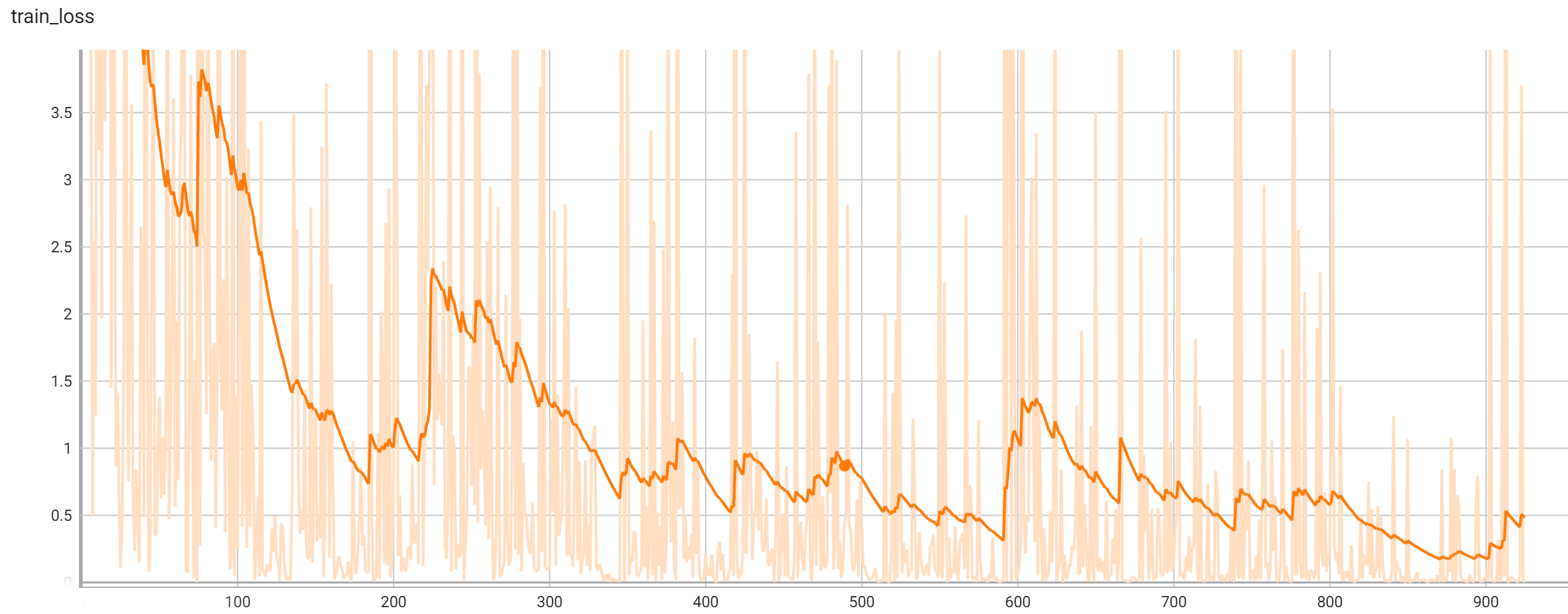
## Training Performance
|
| 65 |
+
A graph showing the training loss over 25 epochs.
|
| 66 |
+
|
| 67 |
+
 <br>
|
| 68 |
+
|
| 69 |
+
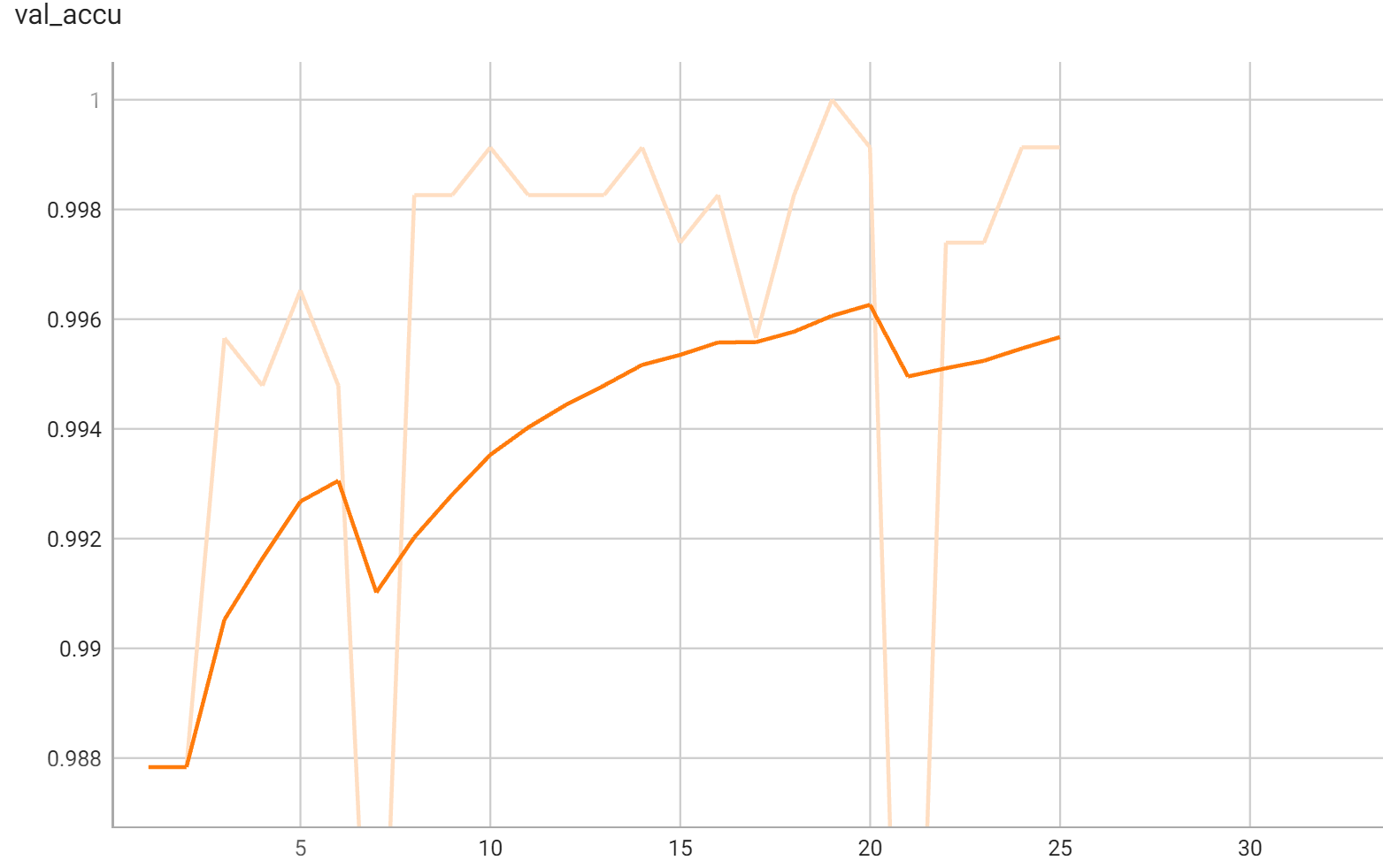
## Validation Performance
|
| 70 |
+
A graph showing the validation accuracy over 25 epochs.
|
| 71 |
+
|
| 72 |
+
 <br>
|
| 73 |
+
|
| 74 |
## commands example
|
| 75 |
Execute training:
|
| 76 |
|
|
|
|
| 121 |
--config_file configs/inference.json
|
| 122 |
```
|
| 123 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 124 |
# References
|
| 125 |
[1] J. Hu, L. Shen and G. Sun, Squeeze-and-Excitation Networks, 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2018, pp. 7132-7141. https://arxiv.org/pdf/1709.01507.pdf
|
| 126 |
|
configs/metadata.json
CHANGED
|
@@ -1,7 +1,8 @@
|
|
| 1 |
{
|
| 2 |
"schema": "https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/meta_schema_20220324.json",
|
| 3 |
-
"version": "0.3.
|
| 4 |
"changelog": {
|
|
|
|
| 5 |
"0.3.0": "update dataset processing",
|
| 6 |
"0.2.2": "update to use monai 1.0.1",
|
| 7 |
"0.2.1": "enhance readme on commands example",
|
|
|
|
| 1 |
{
|
| 2 |
"schema": "https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/meta_schema_20220324.json",
|
| 3 |
+
"version": "0.3.1",
|
| 4 |
"changelog": {
|
| 5 |
+
"0.3.1": "add workflow, train loss and validation accuracy figures",
|
| 6 |
"0.3.0": "update dataset processing",
|
| 7 |
"0.2.2": "update to use monai 1.0.1",
|
| 8 |
"0.2.1": "enhance readme on commands example",
|
docs/README.md
CHANGED
|
@@ -5,6 +5,8 @@ A pre-trained model for the endoscopic inbody classification task.
|
|
| 5 |
This model is trained using the SEResNet50 structure, whose details can be found in [1]. All datasets are from private samples of [Activ Surgical](https://www.activsurgical.com/). Samples in training and validation dataset are from the same 4 videos, while test samples are from different two videos.
|
| 6 |
The [pytorch model](https://drive.google.com/file/d/14CS-s1uv2q6WedYQGeFbZeEWIkoyNa-x/view?usp=sharing) and [torchscript model](https://drive.google.com/file/d/1fOoJ4n5DWKHrt9QXTZ2sXwr9C-YvVGCM/view?usp=sharing) are shared in google drive. Modify the `bundle_root` parameter specified in `configs/train.json` and `configs/inference.json` to reflect where models are downloaded. Expected directory path to place downloaded models is `models/` under `bundle_root`.
|
| 7 |
|
|
|
|
|
|
|
| 8 |
## Data
|
| 9 |
Datasets used in this work were provided by [Activ Surgical](https://www.activsurgical.com/). Here is a [link](https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/inbody_outbody_samples.zip) of 20 samples (10 in-body and 10 out-body) to show what this dataset looks like. After downloading this dataset, python script in `scripts` folder naming `data_process` can be used to get label json files by running the command below and replacing datapath and outpath parameters.
|
| 10 |
```
|
|
@@ -52,6 +54,16 @@ This model achieves the following accuracy score on the test dataset:
|
|
| 52 |
|
| 53 |
Accuracy = 0.98
|
| 54 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 55 |
## commands example
|
| 56 |
Execute training:
|
| 57 |
|
|
@@ -102,23 +114,6 @@ python -m monai.bundle ckpt_export network_def \
|
|
| 102 |
--config_file configs/inference.json
|
| 103 |
```
|
| 104 |
|
| 105 |
-
Export checkpoint to onnx file, which has been tested on pytorch 1.12.0:
|
| 106 |
-
|
| 107 |
-
```
|
| 108 |
-
python scripts/export_to_onnx.py --model models/model.pt --outpath models/model.onnx
|
| 109 |
-
```
|
| 110 |
-
|
| 111 |
-
Export TensorRT float16 model from the onnx model:
|
| 112 |
-
|
| 113 |
-
```
|
| 114 |
-
trtexec --onnx=models/model.onnx --saveEngine=models/model.trt --fp16 \
|
| 115 |
-
--minShapes=INPUT__0:1x3x256x256 \
|
| 116 |
-
--optShapes=INPUT__0:16x3x256x256 \
|
| 117 |
-
--maxShapes=INPUT__0:32x3x256x256 \
|
| 118 |
-
--shapes=INPUT__0:8x3x256x256
|
| 119 |
-
```
|
| 120 |
-
This command need TensorRT with correct CUDA installed in the environment. For the detail of installing TensorRT, please refer to [this link](https://docs.nvidia.com/deeplearning/tensorrt/install-guide/index.html).
|
| 121 |
-
|
| 122 |
# References
|
| 123 |
[1] J. Hu, L. Shen and G. Sun, Squeeze-and-Excitation Networks, 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2018, pp. 7132-7141. https://arxiv.org/pdf/1709.01507.pdf
|
| 124 |
|
|
|
|
| 5 |
This model is trained using the SEResNet50 structure, whose details can be found in [1]. All datasets are from private samples of [Activ Surgical](https://www.activsurgical.com/). Samples in training and validation dataset are from the same 4 videos, while test samples are from different two videos.
|
| 6 |
The [pytorch model](https://drive.google.com/file/d/14CS-s1uv2q6WedYQGeFbZeEWIkoyNa-x/view?usp=sharing) and [torchscript model](https://drive.google.com/file/d/1fOoJ4n5DWKHrt9QXTZ2sXwr9C-YvVGCM/view?usp=sharing) are shared in google drive. Modify the `bundle_root` parameter specified in `configs/train.json` and `configs/inference.json` to reflect where models are downloaded. Expected directory path to place downloaded models is `models/` under `bundle_root`.
|
| 7 |
|
| 8 |
+

|
| 9 |
+
|
| 10 |
## Data
|
| 11 |
Datasets used in this work were provided by [Activ Surgical](https://www.activsurgical.com/). Here is a [link](https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/inbody_outbody_samples.zip) of 20 samples (10 in-body and 10 out-body) to show what this dataset looks like. After downloading this dataset, python script in `scripts` folder naming `data_process` can be used to get label json files by running the command below and replacing datapath and outpath parameters.
|
| 12 |
```
|
|
|
|
| 54 |
|
| 55 |
Accuracy = 0.98
|
| 56 |
|
| 57 |
+
## Training Performance
|
| 58 |
+
A graph showing the training loss over 25 epochs.
|
| 59 |
+
|
| 60 |
+
 <br>
|
| 61 |
+
|
| 62 |
+
## Validation Performance
|
| 63 |
+
A graph showing the validation accuracy over 25 epochs.
|
| 64 |
+
|
| 65 |
+
 <br>
|
| 66 |
+
|
| 67 |
## commands example
|
| 68 |
Execute training:
|
| 69 |
|
|
|
|
| 114 |
--config_file configs/inference.json
|
| 115 |
```
|
| 116 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 117 |
# References
|
| 118 |
[1] J. Hu, L. Shen and G. Sun, Squeeze-and-Excitation Networks, 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2018, pp. 7132-7141. https://arxiv.org/pdf/1709.01507.pdf
|
| 119 |
|