File size: 10,498 Bytes
fd3e5c9 8696833 fd3e5c9 8696833 fd3e5c9 8696833 fd3e5c9 8696833 fd3e5c9 8696833 fd3e5c9 8696833 fd3e5c9 8696833 fd3e5c9 8696833 |
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 |
---
license: other
license_name: nvidia-open-model-license-agreement
license_link: https://www.nvidia.com/en-us/agreements/enterprise-software/nvidia-open-model-license/
pipeline_tag: image-segmentation
library_name: monai
tags:
- nvidia
- medical-imaging
- ct
- segmentation
- vista3d
---
# NV-Segment-CT
## Description:
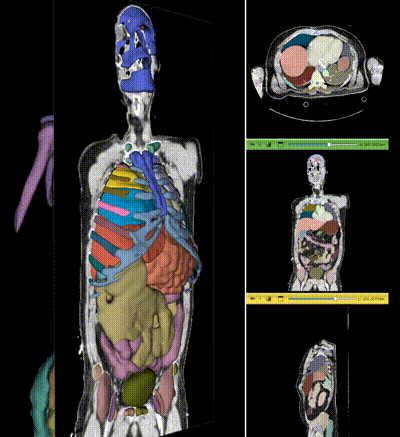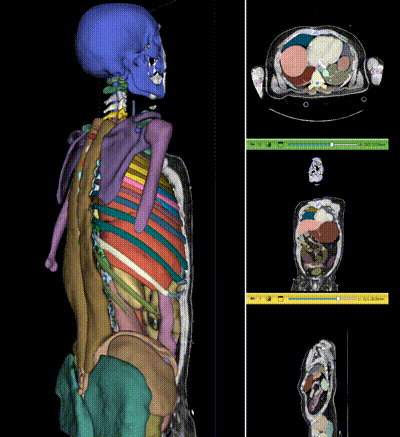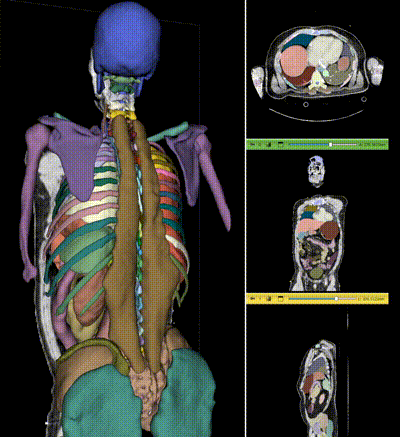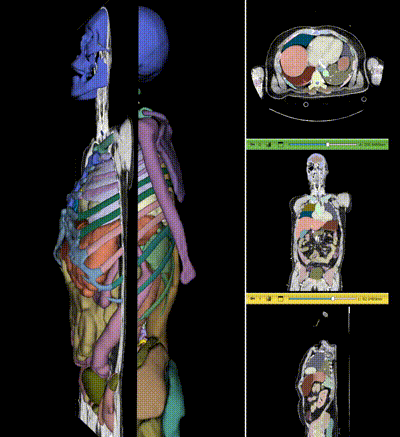
NV-Segment-CT is a specialized interactive foundation model for 3D medical imaging. It excels in providing accurate and adaptable segmentation analysis across anatomies and modalities. Utilizing a multi-head architecture, NV-Segment-CT adapts to varying conditions and anatomical areas, helping guide users' annotation workflow.
This model is for research purposes and not for clinical usage.
**Training & Fine-tuning**: Visit [GitHub](https://github.com/NVIDIA-Medtech/NV-Segment-CTMR) for training code, fine-tuning guides, continual learning examples, and comprehensive development documentation.
Core to NV-Segment-CT are three workflows:
- **Segment everything**: Enables whole body exploration, crucial for understanding complex diseases affecting multiple organs and for holistic treatment planning.
- **Segment using class**: Provides detailed sectional views based on specific classes, essential for targeted disease analysis or organ mapping, such as tumor identification in critical organs.
- **Segment point prompts**: Enhances segmentation precision through user-directed, click-based selection. This interactive approach accelerates the creation of accurate ground-truth data, essential in medical imaging analysis.
## Run pipeline:
For running the pipeline, NV-Segment-CT requires at least one prompt for segmentation. It supports label prompt, which is the index of the class for automatic segmentation. It also supports point-click prompts for binary interactive segmentation. Users can provide both prompts at the same time.
Here is a code snippet to showcase how to execute inference with this model.
```python
import os
import tempfile
import torch
from hugging_face_pipeline import HuggingFacePipelineHelper
FILE_PATH = os.path.dirname(__file__)
with tempfile.TemporaryDirectory() as tmp_dir:
output_dir = os.path.join(tmp_dir, "output_dir")
pipeline_helper = HuggingFacePipelineHelper("vista3d")
pipeline = pipeline_helper.init_pipeline(
os.path.join(FILE_PATH, "vista3d_pretrained_model"),
device=torch.device("cuda:0"),
)
inputs = [
{
"image": "/data/Task09_Spleen/imagesTs/spleen_1.nii.gz",
"label_prompt": [3],
},
{
"image": "/data/Task09_Spleen/imagesTs/spleen_11.nii.gz",
"label_prompt": [3],
},
]
pipeline(inputs, output_dir=output_dir)
```
The inputs defines the image to segment and the prompt for segmentation.
```python
inputs = {'image': '/data/Task09_Spleen/imagesTs/spleen_15.nii.gz', 'label_prompt':[1]}
inputs = {'image': '/data/Task09_Spleen/imagesTs/spleen_15.nii.gz', 'points':[[138,245,18], [271,343,27]], 'point_labels':[1,0]}
```
- The inputs must include the key `image` which contain the absolute path to the nii image file, and includes prompt keys of `label_prompt`, `points` and `point_labels`.
- The `label_prompt` is a list of length `B`, which can perform `B` foreground objects segmentation, e.g. `[2,3,4,5]`. If `B>1`, Point prompts must NOT be provided.
- The `points` is of shape `[N, 3]` like `[[x1,y1,z1],[x2,y2,z2],...[xN,yN,zN]]`, representing `N` point coordinates **IN THE ORIGINAL IMAGE SPACE** of a single foreground object. `point_labels` is a list of length [N] like [1,1,0,-1,...], which
matches the `points`. 0 means background, 1 means foreground, -1 means ignoring this point. `points` and `point_labels` must pe provided together and match length.
- **B must be 1 if label_prompt and points are provided together**. The inferer only supports SINGLE OBJECT point click segmentatation.
- If no prompt is provided, the model will use `everything_labels` to segment 117 classes:
```Python
list(set([i+1 for i in range(132)]) - set([2,16,18,20,21,23,24,25,26,27,128,129,130,131,132]))
```
- The `points` together with `label_prompts` for "Kidney", "Lung", "Bone" (class index [2, 20, 21]) are not allowed since those prompts will be divided into sub-categories (e.g. left kidney and right kidney). Use `points` for the sub-categories as defined in the `inference.json`.
- To specify a new class for zero-shot segmentation, set the `label_prompt` to a value between 133 and 254. Ensure that `points` and `point_labels` are also provided; otherwise, the inference result will be a tensor of zeros.
## Model Architecture:
**Architecture Type:** Transformer <br>
**Network Architecture:** SAM-like<br>
## Input:
**Input Type(s):** Computed Tomography (CT) Image<br>
**Input Format(s):** (Neuroimaging Informatics Technology Initiative) NIfTI <br>
**Input Parameters:** Three-Dimensional (3D) <br>
**Other Properties Related to Input:** Array of Class/Point Information
## Output:
**Output Type(s):** Image <br>
**Output Format:** NIfTI <br>
**Output Parameters:** 3D <br>
## Software Integration:
**Runtime Engine(s):**
MONAI Core v.1.3 <br>
**Supported Hardware Microarchitecture Compatibility:** <br>
* Ampere <br>
* Hopper <br>
**[Preferred/Supported] Operating System(s):** <br>
* Linux <br>
## Inference:
**Engine:** Triton <br>
**Test Hardware:**
A100<br>
H100<br>
L40<br>
## Ethical Considerations:
NVIDIA believes Trustworthy AI is a shared responsibility and we have established policies and practices to enable development for a wide array of AI applications. When downloaded or used in accordance with our terms of service, developers should work with their internal model team to ensure this model meets requirements for the relevant industry and use case and addresses unforeseen product misuse. Please report security vulnerabilities or NVIDIA AI Concerns [here](https://www.nvidia.com/en-us/support/submit-security-vulnerability/).
## Additional Information:
The current list of classes available:
"0": "background",
"1": "liver",
"2": "kidney",
"3": "spleen",
"4": "pancreas",
"5": "right kidney",
"6": "aorta",
"7": "inferior vena cava",
"8": "right adrenal gland",
"9": "left adrenal gland",
"10": "gallbladder",
"11": "esophagus",
"12": "stomach",
"13": "duodenum",
"14": "left kidney",
"15": "bladder",
"16": "prostate or uterus",
"17": "portal vein and splenic vein",
"18": "rectum",
"19": "small bowel",
"20": "lung",
"21": "bone",
"22": "brain",
"23": "lung tumor",
"24": "pancreatic tumor",
"25": "hepatic vessel",
"26": "hepatic tumor",
"27": "colon cancer primaries",
"28": "left lung upper lobe",
"29": "left lung lower lobe",
"30": "right lung upper lobe",
"31": "right lung middle lobe",
"32": "right lung lower lobe",
"33": "vertebrae L5",
"34": "vertebrae L4",
"35": "vertebrae L3",
"36": "vertebrae L2",
"37": "vertebrae L1",
"38": "vertebrae T12",
"39": "vertebrae T11",
"40": "vertebrae T10",
"41": "vertebrae T9",
"42": "vertebrae T8",
"43": "vertebrae T7",
"44": "vertebrae T6",
"45": "vertebrae T5",
"46": "vertebrae T4",
"47": "vertebrae T3",
"48": "vertebrae T2",
"49": "vertebrae T1",
"50": "vertebrae C7",
"51": "vertebrae C6",
"52": "vertebrae C5",
"53": "vertebrae C4",
"54": "vertebrae C3",
"55": "vertebrae C2",
"56": "vertebrae C1",
"57": "trachea",
"58": "left iliac artery",
"59": "right iliac artery",
"60": "left iliac vena",
"61": "right iliac vena",
"62": "colon",
"63": "left rib 1",
"64": "left rib 2",
"65": "left rib 3",
"66": "left rib 4",
"67": "left rib 5",
"68": "left rib 6",
"69": "left rib 7",
"70": "left rib 8",
"71": "left rib 9",
"72": "left rib 10",
"73": "left rib 11",
"74": "left rib 12",
"75": "right rib 1",
"76": "right rib 2",
"77": "right rib 3",
"78": "right rib 4",
"79": "right rib 5",
"80": "right rib 6",
"81": "right rib 7",
"82": "right rib 8",
"83": "right rib 9",
"84": "right rib 10",
"85": "right rib 11",
"86": "right rib 12",
"87": "left humerus",
"88": "right humerus",
"89": "left scapula",
"90": "right scapula",
"91": "left clavicula",
"92": "right clavicula",
"93": "left femur",
"94": "right femur",
"95": "left hip",
"96": "right hip",
"97": "sacrum",
"98": "left gluteus maximus",
"99": "right gluteus maximus",
"100": "left gluteus medius",
"101": "right gluteus medius",
"102": "left gluteus minimus",
"103": "right gluteus minimus",
"104": "left autochthon",
"105": "right autochthon",
"106": "left iliopsoas",
"107": "right iliopsoas",
"108": "left atrial appendage",
"109": "brachiocephalic trunk",
"110": "left brachiocephalic vein",
"111": "right brachiocephalic vein",
"112": "left common carotid artery",
"113": "right common carotid artery",
"114": "costal cartilages",
"115": "heart",
"116": "left kidney cyst",
"117": "right kidney cyst",
"118": "prostate",
"119": "pulmonary vein",
"120": "skull",
"121": "spinal cord",
"122": "sternum",
"123": "left subclavian artery",
"124": "right subclavian artery",
"125": "superior vena cava",
"126": "thyroid gland",
"127": "vertebrae S1",
"128": "bone lesion",
"129": "kidney mass",
"130": "liver tumor",
"131": "vertebrae L6",
"132": "airway"
## Resources
- **Training & Fine-tuning**: [GitHub Repository](https://github.com/NVIDIA-Medtech/NV-Segment-CTMR) - Comprehensive training guides, fine-tuning examples, and development documentation
- **Sister Model**: [NV-Segment-CTMR](https://huggingface.co/nvidia/NV-Segment-CTMR) - Non-commercial model with CT+MRI support (345+ classes)
- **Clara Medical Collection**: [View all NVIDIA medical AI models](https://huggingface.co/collections/nvidia/clara-medical)
# License
## Code License
This project includes code licensed under the Apache License 2.0.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
## Model Weights License
The model weights included in this project are licensed under the NCLS v1 License.
Both licenses' full texts have been combined into a single `LICENSE` file. Please refer to this `LICENSE` file for more details about the terms and conditions of both licenses.
# References
- Antonelli, M., Reinke, A., Bakas, S. et al. The Medical Segmentation Decathlon. Nat Commun 13, 4128 (2022). https://doi.org/10.1038/s41467-022-30695-9
- He, Yufan, et al. VISTA3D: A unified segmentation foundation model for 3D medical imaging. CVPR 2025. https://arxiv.org/abs/2406.05285
|