Updating README.md with metadata and consistency improvements (#1)
Browse files- Updating README.md with metadata and consistency improvements (fd3e5c93a7ca6bf8daed5e5d476855a76cf6647a)
Co-authored-by: Michael Zephyr <nv-mzephyr@users.noreply.huggingface.co>
README.md
CHANGED
|
@@ -1,13 +1,29 @@
|
|
| 1 |
-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 2 |
|
| 3 |
## Description:
|
| 4 |
-
NV-Segment-CT
|
| 5 |
-
This model is a hugging face refactored version of the [MONAI VISTA3D](https://github.com/Project-MONAI/model-zoo/tree/dev/models/vista3d) bundle. A pipeline with transformer library interfaces is provided by this model. For more details about the original model, please visit the [MONAI model zoo](https://github.com/Project-MONAI/model-zoo).
|
| 6 |
|
| 7 |
This model is for research purposes and not for clinical usage.
|
| 8 |
|
|
|
|
| 9 |
|
| 10 |
-
Core to
|
| 11 |
|
| 12 |
- **Segment everything**: Enables whole body exploration, crucial for understanding complex diseases affecting multiple organs and for holistic treatment planning.
|
| 13 |
- **Segment using class**: Provides detailed sectional views based on specific classes, essential for targeted disease analysis or organ mapping, such as tumor identification in critical organs.
|
|
@@ -17,7 +33,7 @@ Core to VISTA-3D are three workflows:
|
|
| 17 |
https://github.com/NVIDIA-Medtech/NV-Segment-CTMR
|
| 18 |
|
| 19 |
## Run pipeline:
|
| 20 |
-
For running the pipeline,
|
| 21 |
|
| 22 |
Here is a code snippet to showcase how to execute inference with this model.
|
| 23 |
```python
|
|
@@ -69,8 +85,6 @@ list(set([i+1 for i in range(132)]) - set([2,16,18,20,21,23,24,25,26,27,128,129,
|
|
| 69 |
- To specify a new class for zero-shot segmentation, set the `label_prompt` to a value between 133 and 254. Ensure that `points` and `point_labels` are also provided; otherwise, the inference result will be a tensor of zeros.
|
| 70 |
|
| 71 |
|
| 72 |
-
|
| 73 |
-
|
| 74 |
## Model Architecture:
|
| 75 |
**Architecture Type:** Transformer <br>
|
| 76 |
**Network Architecture:** SAM-like<br>
|
|
@@ -97,45 +111,6 @@ MONAI Core v.1.3 <br>
|
|
| 97 |
**[Preferred/Supported] Operating System(s):** <br>
|
| 98 |
* Linux <br>
|
| 99 |
|
| 100 |
-
## Model Version(s):
|
| 101 |
-
Internal ONLY
|
| 102 |
-
0.1.9 <br>
|
| 103 |
-
Version changelog: https://gitlab-master.nvidia.com/dlmed/vista3d_bundle/-/blob/main/configs/metadata.json
|
| 104 |
-
|
| 105 |
-
# Training & Evaluation:
|
| 106 |
-
## Training Dataset:
|
| 107 |
-
Internal ONLY
|
| 108 |
-
15 Datasets
|
| 109 |
-
Name, JIRA/SWIPAT, Commercial, and # of Data Tracked
|
| 110 |
-
"VISTA" Sheet: https://docs.google.com/spreadsheets/d/14frhzELquSF_-tF7yGFDBHmSdnp-9-5pmbONQx8iQWk/edit?usp=sharing
|
| 111 |
-
|
| 112 |
-
## Evaluation Dataset:
|
| 113 |
-
Internal ONLY
|
| 114 |
-
15 Datasets
|
| 115 |
-
Name, JIRA/SWIPAT, Commercial, and # of Data Tracked
|
| 116 |
-
"VISTA" Sheet: https://docs.google.com/spreadsheets/d/14frhzELquSF_-tF7yGFDBHmSdnp-9-5pmbONQx8iQWk/edit?usp=sharing
|
| 117 |
-
https://docs.google.com/spreadsheets/d/1hmv-O-f6tdgndsRnoqCgcunR2uQ9IySDhZWmjsXwgbM/edit?usp=sharing
|
| 118 |
-
|
| 119 |
-
** Data Collection Method by dataset <br>
|
| 120 |
-
* [Hybrid: Human, Automatic/Sensors] <br>
|
| 121 |
-
|
| 122 |
-
** Labeling Method by dataset <br>
|
| 123 |
-
* [Hybrid: Human, Automatic/Sensors] <br>
|
| 124 |
-
|
| 125 |
-
**Properties:** Custom internal and public dataset of organs from multiple scanner types. <br>
|
| 126 |
-
|
| 127 |
-
|
| 128 |
-
## Evaluation Dataset:
|
| 129 |
-
|
| 130 |
-
** Data Collection Method by dataset <br>
|
| 131 |
-
* [Hybrid: Human, Automatic/Sensors] <br>
|
| 132 |
-
|
| 133 |
-
** Labeling Method by dataset <br>
|
| 134 |
-
* [Hybrid: Human, Automatic/Sensors] <br>
|
| 135 |
-
|
| 136 |
-
**Properties:** Custom internal and public dataset of organs from multiple scanner types. <br>
|
| 137 |
-
|
| 138 |
-
|
| 139 |
## Inference:
|
| 140 |
**Engine:** Triton <br>
|
| 141 |
**Test Hardware:**
|
|
@@ -144,10 +119,10 @@ H100<br>
|
|
| 144 |
L40<br>
|
| 145 |
|
| 146 |
## Ethical Considerations:
|
| 147 |
-
NVIDIA believes Trustworthy AI is a shared responsibility and we have established policies and practices to enable development for a wide array of AI applications. When downloaded or used in accordance with our terms of service, developers should work with their internal model team to ensure this model meets requirements for the relevant industry and use case and addresses unforeseen product misuse.
|
| 148 |
|
| 149 |
## Additional Information:
|
| 150 |
-
The current list of classes available
|
| 151 |
|
| 152 |
"0": "background",
|
| 153 |
"1": "liver",
|
|
@@ -283,6 +258,12 @@ The current list of classes available within VISTA-3D:
|
|
| 283 |
"131": "vertebrae L6",
|
| 284 |
"132": "airway"
|
| 285 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 286 |
# License
|
| 287 |
|
| 288 |
## Code License
|
|
|
|
| 1 |
+
---
|
| 2 |
+
license: other
|
| 3 |
+
license_name: nvidia-open-model-license-agreement
|
| 4 |
+
license_link: https://www.nvidia.com/en-us/agreements/enterprise-software/nvidia-open-model-license/
|
| 5 |
+
pipeline_tag: image-segmentation
|
| 6 |
+
library_name: monai
|
| 7 |
+
tags:
|
| 8 |
+
- nvidia
|
| 9 |
+
- medical-imaging
|
| 10 |
+
- ct
|
| 11 |
+
- segmentation
|
| 12 |
+
- vista3d
|
| 13 |
+
---
|
| 14 |
+
|
| 15 |
+
# NV-Segment-CT
|
| 16 |
+
|
| 17 |
+
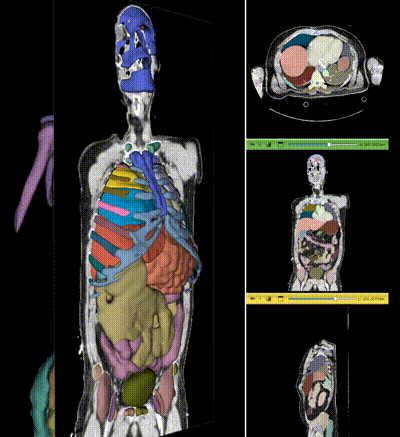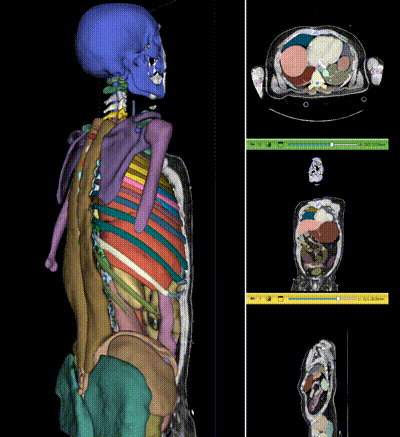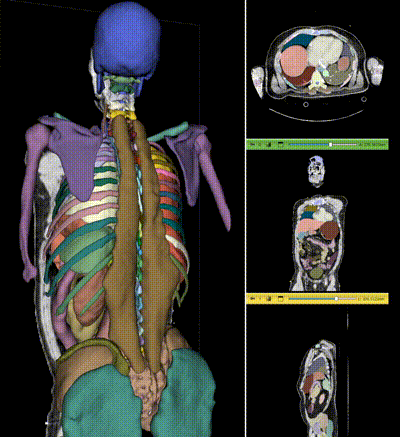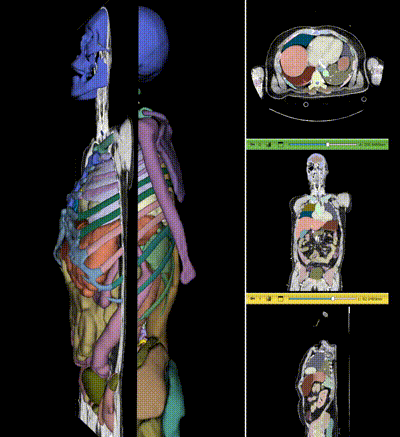
|
| 18 |
|
| 19 |
## Description:
|
| 20 |
+
NV-Segment-CT is a specialized interactive foundation model for 3D medical imaging. It excels in providing accurate and adaptable segmentation analysis across anatomies and modalities. Utilizing a multi-head architecture, NV-Segment-CT adapts to varying conditions and anatomical areas, helping guide users' annotation workflow.
|
|
|
|
| 21 |
|
| 22 |
This model is for research purposes and not for clinical usage.
|
| 23 |
|
| 24 |
+
**Training & Fine-tuning**: Visit [GitHub](https://github.com/NVIDIA-Medtech/NV-Segment-CTMR) for training code, fine-tuning guides, continual learning examples, and comprehensive development documentation.
|
| 25 |
|
| 26 |
+
Core to NV-Segment-CT are three workflows:
|
| 27 |
|
| 28 |
- **Segment everything**: Enables whole body exploration, crucial for understanding complex diseases affecting multiple organs and for holistic treatment planning.
|
| 29 |
- **Segment using class**: Provides detailed sectional views based on specific classes, essential for targeted disease analysis or organ mapping, such as tumor identification in critical organs.
|
|
|
|
| 33 |
https://github.com/NVIDIA-Medtech/NV-Segment-CTMR
|
| 34 |
|
| 35 |
## Run pipeline:
|
| 36 |
+
For running the pipeline, NV-Segment-CT requires at least one prompt for segmentation. It supports label prompt, which is the index of the class for automatic segmentation. It also supports point-click prompts for binary interactive segmentation. Users can provide both prompts at the same time.
|
| 37 |
|
| 38 |
Here is a code snippet to showcase how to execute inference with this model.
|
| 39 |
```python
|
|
|
|
| 85 |
- To specify a new class for zero-shot segmentation, set the `label_prompt` to a value between 133 and 254. Ensure that `points` and `point_labels` are also provided; otherwise, the inference result will be a tensor of zeros.
|
| 86 |
|
| 87 |
|
|
|
|
|
|
|
| 88 |
## Model Architecture:
|
| 89 |
**Architecture Type:** Transformer <br>
|
| 90 |
**Network Architecture:** SAM-like<br>
|
|
|
|
| 111 |
**[Preferred/Supported] Operating System(s):** <br>
|
| 112 |
* Linux <br>
|
| 113 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 114 |
## Inference:
|
| 115 |
**Engine:** Triton <br>
|
| 116 |
**Test Hardware:**
|
|
|
|
| 119 |
L40<br>
|
| 120 |
|
| 121 |
## Ethical Considerations:
|
| 122 |
+
NVIDIA believes Trustworthy AI is a shared responsibility and we have established policies and practices to enable development for a wide array of AI applications. When downloaded or used in accordance with our terms of service, developers should work with their internal model team to ensure this model meets requirements for the relevant industry and use case and addresses unforeseen product misuse. Please report security vulnerabilities or NVIDIA AI Concerns [here](https://www.nvidia.com/en-us/support/submit-security-vulnerability/).
|
| 123 |
|
| 124 |
## Additional Information:
|
| 125 |
+
The current list of classes available:
|
| 126 |
|
| 127 |
"0": "background",
|
| 128 |
"1": "liver",
|
|
|
|
| 258 |
"131": "vertebrae L6",
|
| 259 |
"132": "airway"
|
| 260 |
|
| 261 |
+
## Resources
|
| 262 |
+
|
| 263 |
+
- **Training & Fine-tuning**: [GitHub Repository](https://github.com/NVIDIA-Medtech/NV-Segment-CTMR) - Comprehensive training guides, fine-tuning examples, and development documentation
|
| 264 |
+
- **Sister Model**: [NV-Segment-CTMR](https://huggingface.co/nvidia/NV-Segment-CTMR) - Non-commercial model with CT+MRI support (345+ classes)
|
| 265 |
+
- **Clara Medical Collection**: [View all NVIDIA medical AI models](https://huggingface.co/collections/nvidia/clara-medical)
|
| 266 |
+
|
| 267 |
# License
|
| 268 |
|
| 269 |
## Code License
|