---
title: Brain Tumor Classification With InceptionV3-Grad-CAM
emoji: 🧠
colorFrom: pink
colorTo: red
sdk: gradio
sdk_version: 6.0.0
app_file: app.py
pinned: false
license: mit
short_description: InceptionV3-based brain tumor detection with Grad-CAM.
models:
- AIOmarRehan/Brain_Tumor_Classification_with_Grad-CAM
datasets:
- AIOmarRehan/Brain_Tumor_MRI_Dataset
---
# **Brain Tumor Classification Using InceptionV3 and Grad-CAM**
A complete deep learning pipeline for **brain tumor classification** using MRI scans.
This project demonstrates:
* **End-to-end data preprocessing**
* **Augmentation & dataset balancing**
* **Efficient tf.data pipelines**
* **Transfer learning with InceptionV3**
* **Deep model evaluation**
* **Grad-CAM interpretability**
* **LaTeX mathematical explanations**
---
## **1. Dataset Exploration & Inspection**
We begin by recursively scanning all MRI images and creating a structured DataFrame:
```python
from pathlib import Path
import pandas as pd
image_extensions = {'.jpg', '.jpeg', '.png'}
paths = [
(path.parts[-2], path.name, str(path))
for path in Path("/content/my_data").rglob('*.*')
if path.suffix.lower() in image_extensions
]
df = pd.DataFrame(paths, columns=['class', 'image', 'full_path'])
df = df.sort_values('class').reset_index(drop=True)
df.head()
```
Count images per class:
```python
class_count = df['class'].value_counts()
print(class_count)
```
### **Visualizations**
```python
import matplotlib.pyplot as plt
plt.figure(figsize=(32,16))
class_count.plot(kind='bar', edgecolor='black')
plt.title('Number of Images per Class')
plt.show()
```
### **Insights**
* Classes are **imbalanced**
* Images have **variable resolution**
* Some outliers require **cleaning**
---
## **2. Data Cleaning & Quality Checks**
### **Duplicate removal using MD5 hashes**
```python
import hashlib
def get_hash(file_path):
with open(file_path, 'rb') as f:
return hashlib.md5(f.read()).hexdigest()
df['file_hash'] = df['full_path'].apply(get_hash)
df_unique = df.drop_duplicates(subset='file_hash', keep='first')
```
### **Additional checks**
* Corrupted image detection
* Resolution anomalies
* Brightness/contrast outliers
Cleaning ensures a **robust dataset** with minimal noise.
---
## **3. Data Augmentation & Class Balancing**
Target ~200 images per class using heavy augmentation:
```python
from tensorflow.keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
rotation_range=20,
width_shift_range=0.1,
height_shift_range=0.1,
shear_range=0.1,
zoom_range=0.1,
horizontal_flip=True,
fill_mode='nearest'
)
```
Used for minority class upsampling and preventing overfitting.
---
## **4. Image Preprocessing Pipeline**
```python
import tensorflow as tf
def preprocess_image(path, target_size=(512, 512), augment=True):
img = tf.io.read_file(path)
img = tf.image.decode_image(img, channels=3)
img = tf.image.resize(img, target_size)
img = tf.cast(img, tf.float32) / 255.0
if augment:
img = tf.image.random_flip_left_right(img)
img = tf.image.random_flip_up_down(img)
img = tf.image.random_brightness(img, max_delta=0.1)
img = tf.image.random_contrast(img, 0.9, 1.1)
return tf.clip_by_value(img, 0.0, 1.0)
```
* **Train set:** augmentation enabled
* **Validation/Test sets:** kept clean
---
## **5. Dataset Preparation with `tf.data`**
```python
AUTOTUNE = tf.data.AUTOTUNE
batch_size = 32
train_ds = tf.data.Dataset.from_tensor_slices((train_paths, train_labels))
train_ds = train_ds.shuffle(len(train_paths))
train_ds = train_ds.map(
lambda x, y: (preprocess_image(x, augment=True), y),
num_parallel_calls=AUTOTUNE
)
train_ds = train_ds.batch(batch_size).prefetch(AUTOTUNE)
```
Benefits:
* Parallel loading
* Smart prefetching
* GPU utilization maximized
---
## **6. Model Architecture: InceptionV3**
Transfer learning from ImageNet:
```python
from tensorflow.keras.applications.inception_v3 import InceptionV3
from tensorflow.keras.layers import GlobalAveragePooling2D, Dense, Dropout
from tensorflow.keras.models import Model
inception = InceptionV3(input_shape=input_shape, weights='imagenet', include_top=False)
for layer in inception.layers:
layer.trainable = False
x = GlobalAveragePooling2D()(inception.output)
x = Dense(512, activation='relu')(x)
x = Dropout(0.5)(x)
prediction = Dense(len(le.classes_), activation='softmax')(x)
model = Model(inputs=inception.input, outputs=prediction)
```
### Why InceptionV3?
* Factorized convolutions
* Multi-scale feature extraction
* Lightweight and fast
* Strong performance in medical imaging
---
## **7. Training & Callbacks**
```python
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
model.compile(
loss='sparse_categorical_crossentropy',
optimizer='adam',
metrics=['accuracy']
)
callbacks = [
EarlyStopping(monitor='val_loss', patience=40, restore_best_weights=True),
ModelCheckpoint("best_model.h5", save_best_only=True, monitor='val_loss'),
ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=10, min_lr=1e-5)
]
```
Training:
```python
history = model.fit(train_ds, validation_data=val_ds, epochs=50, callbacks=callbacks)
```
---
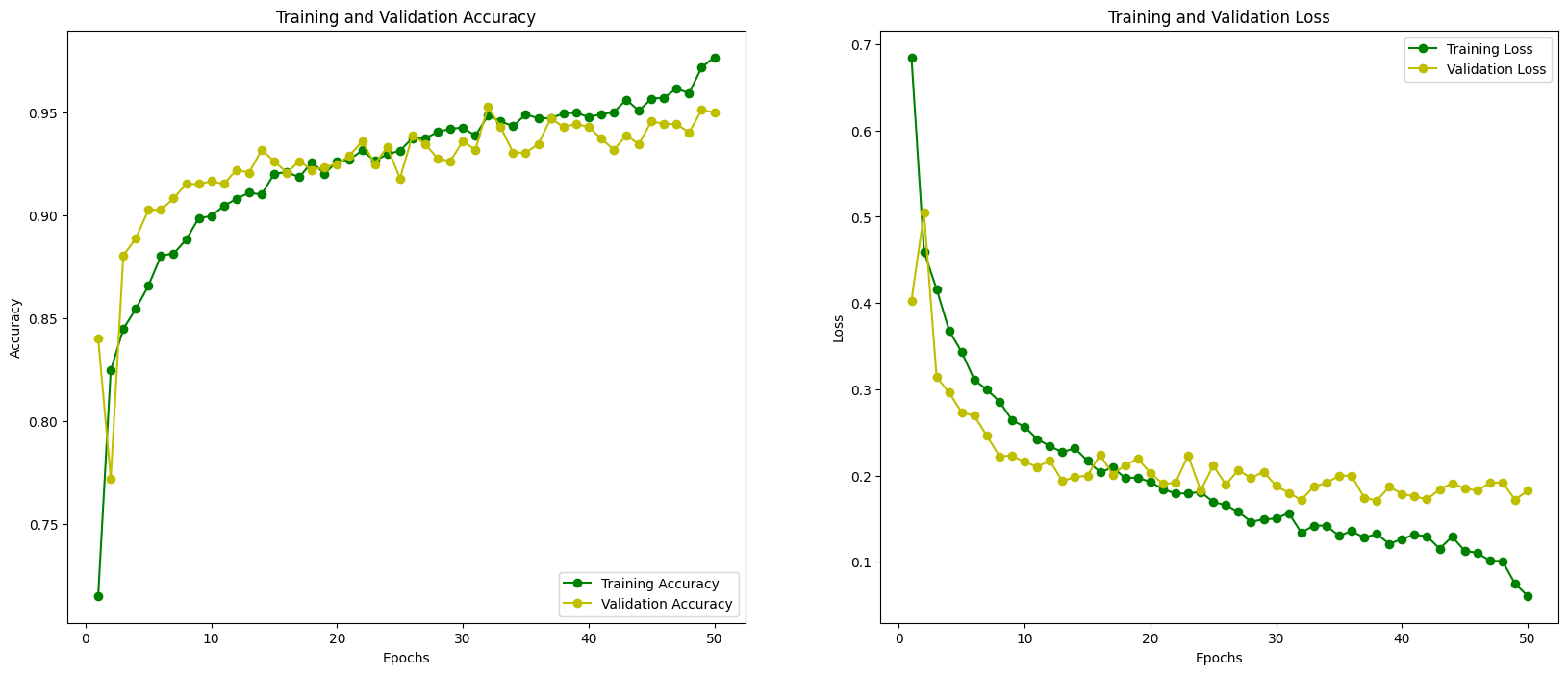
## **8. Training Curves**
```python
import matplotlib.pyplot as plt
plt.plot(history.history['accuracy'], label='Train Accuracy')
plt.plot(history.history['val_accuracy'], label='Val Accuracy')
plt.title('Training vs Validation Accuracy')
plt.legend()
plt.show()
```
* Curves indicate **smooth convergence**
* Small train/val gap → **limited overfitting**
---
## **9. Performance Metrics**
### Confusion Matrix
```python
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
cm = confusion_matrix(y_true, y_pred)
ConfusionMatrixDisplay(cm, display_labels=le.classes_).plot(cmap='Blues')
```
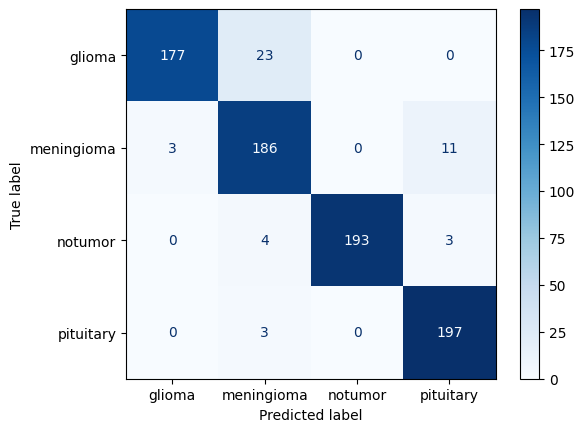
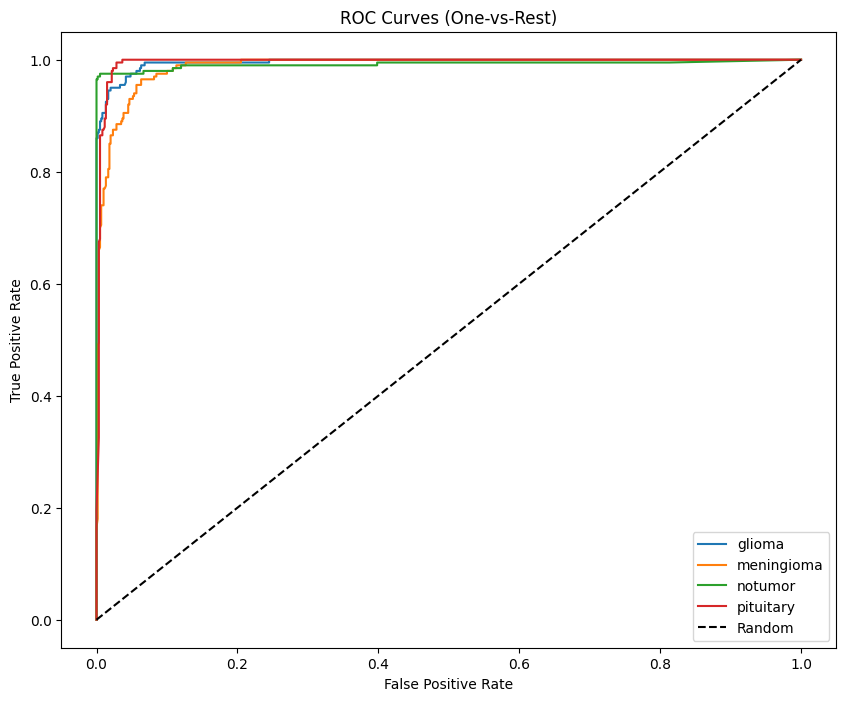
### Multi-class AUC (One-vs-Rest)
**Macro AUC formula:**
 ```python
from sklearn.preprocessing import label_binarize
from sklearn.metrics import roc_curve, auc
y_true_bin = label_binarize(y_true, classes=np.arange(len(le.classes_)))
```
```python
from sklearn.preprocessing import label_binarize
from sklearn.metrics import roc_curve, auc
y_true_bin = label_binarize(y_true, classes=np.arange(len(le.classes_)))
```
---
## **10. Grad-CAM: Interpretability**
Grad-CAM highlights regions the model uses for classification.
### Grad-CAM heatmap:
) Where:
Where:
 Python implementation:
```python
def gradcam(model, img, cls=None):
# last conv
lc = next(l for l in reversed(model.layers) if "conv" in l.name.lower())
gm = tf.keras.Model(model.input, [lc.output, model.output])
with tf.GradientTape() as t:
conv, pred = gm(img[None])
cls = tf.argmax(pred[0]) if cls is None else cls
loss = pred[:, cls]
g = t.gradient(loss, conv)
w = tf.reduce_mean(g, axis=(0,1,2))
cam = tf.reduce_sum(w * conv[0], -1)
cam = tf.nn.relu(cam)
cam /= tf.reduce_max(cam) + 1e-8
return cam.numpy()
```
Visualization example:
```python
plt.figure(figsize=(20,10))
for i, img in enumerate(sample_images):
overlay, info = VizGradCAM(model, img)
plt.subplot(2, 5, i+1)
plt.imshow(overlay)
plt.axis("off")
plt.title(f"True Label: {le.classes_[sample_labels[i]]}")
plt.show()
```
Python implementation:
```python
def gradcam(model, img, cls=None):
# last conv
lc = next(l for l in reversed(model.layers) if "conv" in l.name.lower())
gm = tf.keras.Model(model.input, [lc.output, model.output])
with tf.GradientTape() as t:
conv, pred = gm(img[None])
cls = tf.argmax(pred[0]) if cls is None else cls
loss = pred[:, cls]
g = t.gradient(loss, conv)
w = tf.reduce_mean(g, axis=(0,1,2))
cam = tf.reduce_sum(w * conv[0], -1)
cam = tf.nn.relu(cam)
cam /= tf.reduce_max(cam) + 1e-8
return cam.numpy()
```
Visualization example:
```python
plt.figure(figsize=(20,10))
for i, img in enumerate(sample_images):
overlay, info = VizGradCAM(model, img)
plt.subplot(2, 5, i+1)
plt.imshow(overlay)
plt.axis("off")
plt.title(f"True Label: {le.classes_[sample_labels[i]]}")
plt.show()
```
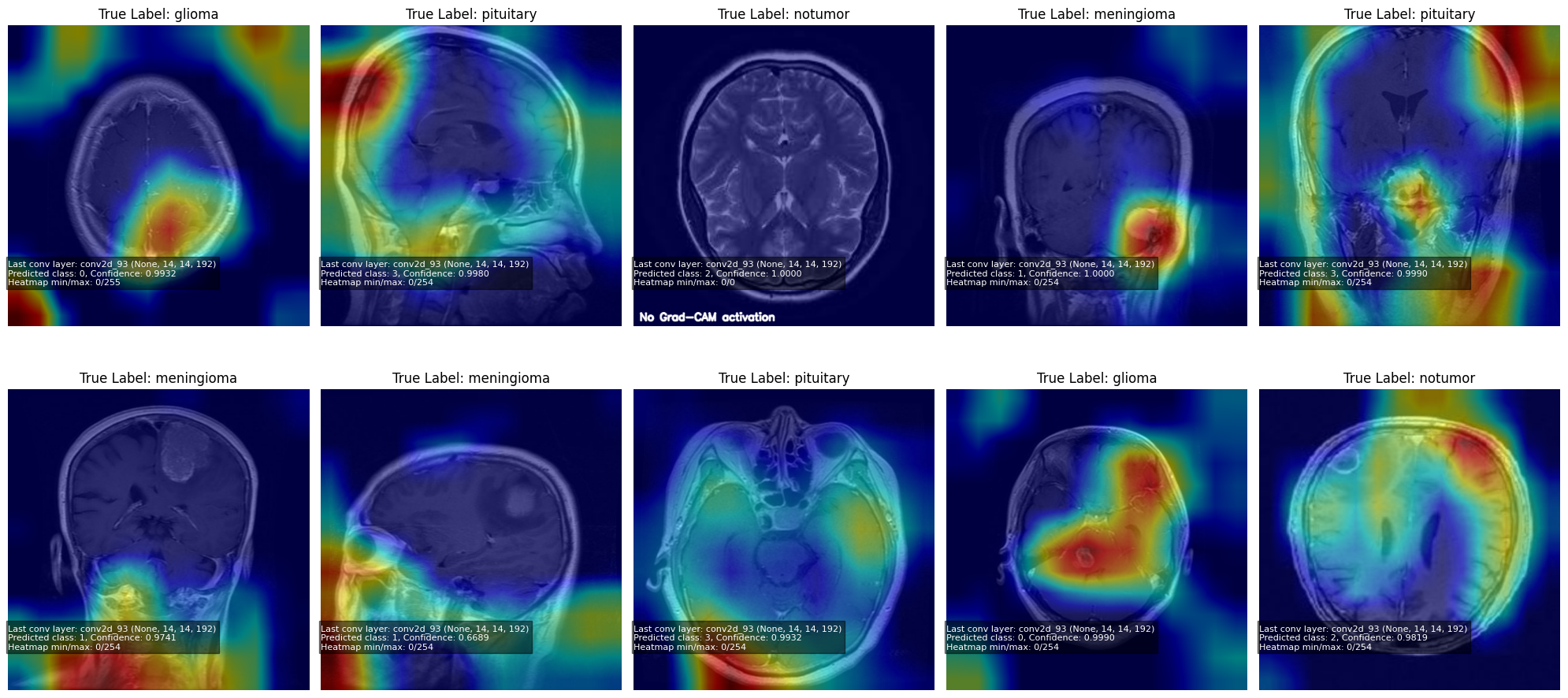
> **Note:** When the model is highly confident in a prediction, the Grad-CAM gradients become near-zero, producing little to no heatmap activation.
---
## **11. Technical LaTeX Notes**
### Sparse Categorical Crossentropy
) ### Global Average Pooling
### Global Average Pooling
 ---
## **12. Model Saving**
```python
model.save("InceptionV3_Brain_Tumor_MRI.h5")
```
---
## **13. Results**
> **Note:** Click the image below to view the video showcasing the project’s results.
---
## **12. Model Saving**
```python
model.save("InceptionV3_Brain_Tumor_MRI.h5")
```
---
## **13. Results**
> **Note:** Click the image below to view the video showcasing the project’s results.

> **Note:** If the video above is not working, you can access it directly via the link below.
[Watch Demo Video](Results/InceptionV3_Brain_Tumor_MRI.mp4)
---
## **Key Takeaways**
* Strong data cleaning = reliable model
* Heavy augmentation reduces bias
* InceptionV3 provides excellent feature extraction
* Evaluation metrics reveal clinical reliability
* Grad-CAM adds essential interpretability




