diff --git a/.dockerignore b/.dockerignore
new file mode 100644
index 0000000000000000000000000000000000000000..8771911bd9b6ef9f6390b603a5ce8a5242958c4b
--- /dev/null
+++ b/.dockerignore
@@ -0,0 +1,6 @@
+.git
+.env
+.claude
+__pycache__
+*.pyc
+outputs/
diff --git a/.env.example b/.env.example
new file mode 100644
index 0000000000000000000000000000000000000000..632ee4515aefc80e66060d18d7bf26f8a80bdead
--- /dev/null
+++ b/.env.example
@@ -0,0 +1,10 @@
+# Required
+GEMINI_API_KEY=your_gemini_api_key
+SUPABASE_URL=your_supabase_project_url
+SUPABASE_ANON_KEY=your_supabase_anon_key
+
+# Optional (auto-detected on Linux, only needed for custom installs)
+# TESSERACT_CMD=/usr/bin/tesseract
+# POPPLER_PATH=/usr/bin
+# PORT=7860
+# FLASK_DEBUG=false
diff --git a/.gitattributes b/.gitattributes
index a6344aac8c09253b3b630fb776ae94478aa0275b..6b61c5fd0906fef9908d8e77e7b3094b94ce67d9 100644
--- a/.gitattributes
+++ b/.gitattributes
@@ -1,35 +1,44 @@
-*.7z filter=lfs diff=lfs merge=lfs -text
-*.arrow filter=lfs diff=lfs merge=lfs -text
-*.bin filter=lfs diff=lfs merge=lfs -text
-*.bz2 filter=lfs diff=lfs merge=lfs -text
-*.ckpt filter=lfs diff=lfs merge=lfs -text
-*.ftz filter=lfs diff=lfs merge=lfs -text
-*.gz filter=lfs diff=lfs merge=lfs -text
-*.h5 filter=lfs diff=lfs merge=lfs -text
-*.joblib filter=lfs diff=lfs merge=lfs -text
-*.lfs.* filter=lfs diff=lfs merge=lfs -text
-*.mlmodel filter=lfs diff=lfs merge=lfs -text
-*.model filter=lfs diff=lfs merge=lfs -text
-*.msgpack filter=lfs diff=lfs merge=lfs -text
-*.npy filter=lfs diff=lfs merge=lfs -text
-*.npz filter=lfs diff=lfs merge=lfs -text
-*.onnx filter=lfs diff=lfs merge=lfs -text
-*.ot filter=lfs diff=lfs merge=lfs -text
-*.parquet filter=lfs diff=lfs merge=lfs -text
-*.pb filter=lfs diff=lfs merge=lfs -text
-*.pickle filter=lfs diff=lfs merge=lfs -text
-*.pkl filter=lfs diff=lfs merge=lfs -text
-*.pt filter=lfs diff=lfs merge=lfs -text
-*.pth filter=lfs diff=lfs merge=lfs -text
-*.rar filter=lfs diff=lfs merge=lfs -text
-*.safetensors filter=lfs diff=lfs merge=lfs -text
-saved_model/**/* filter=lfs diff=lfs merge=lfs -text
-*.tar.* filter=lfs diff=lfs merge=lfs -text
-*.tar filter=lfs diff=lfs merge=lfs -text
-*.tflite filter=lfs diff=lfs merge=lfs -text
-*.tgz filter=lfs diff=lfs merge=lfs -text
-*.wasm filter=lfs diff=lfs merge=lfs -text
-*.xz filter=lfs diff=lfs merge=lfs -text
-*.zip filter=lfs diff=lfs merge=lfs -text
-*.zst filter=lfs diff=lfs merge=lfs -text
-*tfevents* filter=lfs diff=lfs merge=lfs -text
+# Auto detect text files and perform LF normalization
+* text=auto
+Images/OCRSheet.jpg filter=lfs diff=lfs merge=lfs -text
+Images/OcrSheetMarked.jpg filter=lfs diff=lfs merge=lfs -text
+Images/omr_answer_key.pdf filter=lfs diff=lfs merge=lfs -text
+Images/omr_answer_key.png filter=lfs diff=lfs merge=lfs -text
+Images/OMRSheet.jpg filter=lfs diff=lfs merge=lfs -text
+Images/OMRSheet2.jpg filter=lfs diff=lfs merge=lfs -text
+Images/OMRSheet3.jpg filter=lfs diff=lfs merge=lfs -text
+Images/OMRTest.jpg filter=lfs diff=lfs merge=lfs -text
+Images/question_paper.png filter=lfs diff=lfs merge=lfs -text
+Images/test.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/inputs/OMRImage.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/outputs/CheckedOMRs/OMRImage.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/Sandeep-1507/omr-1.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/Sandeep-1507/omr-2.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/Sandeep-1507/omr-3.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/Shamanth/omr_sheet_01.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/UPSC-mock/answer_key.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample1/MobileCamera/sheet1.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample2/AdrianSample/adrian_omr.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample4/IMG_20201116_143512.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample4/IMG_20201116_150717658.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample4/IMG_20201116_150750830.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg filter=lfs diff=lfs merge=lfs -text
+OMRChecker/samples/sample6/reference.png filter=lfs diff=lfs merge=lfs -text
+OMRChecker/src/tests/test_samples/sample2/sample.jpg filter=lfs diff=lfs merge=lfs -text
+output/test_ocr_res_img.jpg filter=lfs diff=lfs merge=lfs -text
+output/test_preprocessed_img.jpg filter=lfs diff=lfs merge=lfs -text
diff --git a/.gitignore b/.gitignore
new file mode 100644
index 0000000000000000000000000000000000000000..c0d9bca9ec94a13bea3f3ab28d790fae28fab60f
--- /dev/null
+++ b/.gitignore
@@ -0,0 +1,188 @@
+# Byte-compiled / optimized / DLL files
+__pycache__/
+*.py[cod]
+*$py.class
+
+# C extensions
+*.so
+
+# Distribution / packaging
+.Python
+build/
+develop-eggs/
+dist/
+downloads/
+eggs/
+.eggs/
+lib/
+lib64/
+parts/
+sdist/
+var/
+wheels/
+share/python-wheels/
+*.egg-info/
+.installed.cfg
+*.egg
+MANIFEST
+
+# PyInstaller
+# Usually these files are written by a python script from a template
+# before PyInstaller builds the exe, so as to inject date/other infos into it.
+*.manifest
+*.spec
+
+# Installer logs
+pip-log.txt
+pip-delete-this-directory.txt
+
+# Unit test / coverage reports
+htmlcov/
+.tox/
+.nox/
+.coverage
+.coverage.*
+.cache
+nosetests.xml
+coverage.xml
+*.cover
+*.py,cover
+.hypothesis/
+.pytest_cache/
+cover/
+
+# Translations
+*.mo
+*.pot
+
+# Django stuff:
+*.log
+local_settings.py
+db.sqlite3
+db.sqlite3-journal
+
+# Flask stuff:
+instance/
+.webassets-cache
+
+# Scrapy stuff:
+.scrapy
+
+# Sphinx documentation
+docs/_build/
+
+# PyBuilder
+.pybuilder/
+target/
+
+# Jupyter Notebook
+.ipynb_checkpoints
+
+# IPython
+profile_default/
+ipython_config.py
+
+# pyenv
+# For a library or package, you might want to ignore these files since the code is
+# intended to run in multiple environments; otherwise, check them in:
+# .python-version
+
+# pipenv
+# According to pypa/pipenv#598, it is recommended to include Pipfile.lock in version control.
+# However, in case of collaboration, if having platform-specific dependencies or dependencies
+# having no cross-platform support, pipenv may install dependencies that don't work, or not
+# install all needed dependencies.
+#Pipfile.lock
+
+# UV
+# Similar to Pipfile.lock, it is generally recommended to include uv.lock in version control.
+# This is especially recommended for binary packages to ensure reproducibility, and is more
+# commonly ignored for libraries.
+#uv.lock
+
+# poetry
+# Similar to Pipfile.lock, it is generally recommended to include poetry.lock in version control.
+# This is especially recommended for binary packages to ensure reproducibility, and is more
+# commonly ignored for libraries.
+# https://python-poetry.org/docs/basic-usage/#commit-your-poetrylock-file-to-version-control
+#poetry.lock
+
+# pdm
+# Similar to Pipfile.lock, it is generally recommended to include pdm.lock in version control.
+#pdm.lock
+# pdm stores project-wide configurations in .pdm.toml, but it is recommended to not include it
+# in version control.
+# https://pdm.fming.dev/latest/usage/project/#working-with-version-control
+.pdm.toml
+.pdm-python
+.pdm-build/
+
+# PEP 582; used by e.g. github.com/David-OConnor/pyflow and github.com/pdm-project/pdm
+__pypackages__/
+
+# Celery stuff
+celerybeat-schedule
+celerybeat.pid
+
+# SageMath parsed files
+*.sage.py
+
+# Environments
+.env
+.venv
+env/
+venv/
+ENV/
+env.bak/
+venv.bak/
+
+# Spyder project settings
+.spyderproject
+.spyproject
+
+# Rope project settings
+.ropeproject
+
+# mkdocs documentation
+/site
+
+# mypy
+.mypy_cache/
+.dmypy.json
+dmypy.json
+
+# Pyre type checker
+.pyre/
+
+# pytype static type analyzer
+.pytype/
+
+# Cython debug symbols
+cython_debug/
+
+# PyCharm
+# JetBrains specific template is maintained in a separate JetBrains.gitignore that can
+# be found at https://github.com/github/gitignore/blob/main/Global/JetBrains.gitignore
+# and can be added to the global gitignore or merged into this file. For a more nuclear
+# option (not recommended) you can uncomment the following to ignore the entire idea folder.
+#.idea/
+
+# App outputs
+outputs/
+.claude/
+
+# Temp/debug images
+Images/debug_*.png
+
+# Ruff stuff:
+.ruff_cache/
+
+# PyPI configuration file
+.pypirc
+
+# Cursor
+# Cursor is an AI-powered code editor.`.cursorignore` specifies files/directories to
+# exclude from AI features like autocomplete and code analysis. Recommended for sensitive data
+# refer to https://docs.cursor.com/context/ignore-files
+.cursorignore
+.cursorindexingignore
\ No newline at end of file
diff --git a/Dockerfile b/Dockerfile
new file mode 100644
index 0000000000000000000000000000000000000000..5161358c714a9d7a803d0d1e061582f88f339358
--- /dev/null
+++ b/Dockerfile
@@ -0,0 +1,30 @@
+FROM python:3.11-slim
+
+# System deps: tesseract, poppler, opencv libs
+RUN apt-get update && apt-get install -y --no-install-recommends \
+ tesseract-ocr \
+ poppler-utils \
+ libgl1 \
+ libglib2.0-0 \
+ && rm -rf /var/lib/apt/lists/*
+
+WORKDIR /app
+
+# Install CPU-only PyTorch first (avoids ~4GB of NVIDIA CUDA libs)
+RUN pip install --no-cache-dir torch torchvision --index-url https://download.pytorch.org/whl/cpu
+
+# Install remaining Python deps
+COPY requirements.txt .
+RUN pip install --no-cache-dir -r requirements.txt
+
+# Copy application code
+COPY . .
+
+# Create required directories
+RUN mkdir -p OMRChecker/inputs outputs/Results outputs/CheckedOMRs outputs/Manual outputs/Evaluation Images
+
+# HF Spaces expects port 7860
+ENV PORT=7860
+EXPOSE 7860
+
+CMD ["python", "app.py"]
diff --git a/Images/GK_Question_Paper_A4.pdf b/Images/GK_Question_Paper_A4.pdf
new file mode 100644
index 0000000000000000000000000000000000000000..12edfc5cdbfb2cfbef01e6624510cfea5ac342da
Binary files /dev/null and b/Images/GK_Question_Paper_A4.pdf differ
diff --git a/Images/OCR2.jpg b/Images/OCR2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..d235818d3997a061dde34a148b53b56947acd61b
Binary files /dev/null and b/Images/OCR2.jpg differ
diff --git a/Images/OCRSheet.jpg b/Images/OCRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c0f7e5ce9292cf397fa5dd921dc926968529c444
--- /dev/null
+++ b/Images/OCRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:76d3a863bc910ffa2de222ead17eb7021999e594c9fd5a6a0116dc3ed7149fff
+size 112779
diff --git a/Images/OCRTest.pdf b/Images/OCRTest.pdf
new file mode 100644
index 0000000000000000000000000000000000000000..5e464937d24356e93f2457e463659deec9b21605
Binary files /dev/null and b/Images/OCRTest.pdf differ
diff --git a/Images/OMRSheet.jpg b/Images/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c9e3dcccfc8af7c60becb3a4cc25f650861db98e
--- /dev/null
+++ b/Images/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4439b6a479cb9fea92de9656b900a528052616ae8e8962a6c33b8e81ed7f327a
+size 267398
diff --git a/Images/OMRSheet2.jpg b/Images/OMRSheet2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..42fcf6eb88cc072090a6296e2faa9e335963715c
--- /dev/null
+++ b/Images/OMRSheet2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4c6661dce53a4ff8292ee218a2acf5f0aa921c3f337cc0117bda6c71f487735b
+size 232847
diff --git a/Images/OMRSheet3.jpg b/Images/OMRSheet3.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa0600c4643d69ea5302f406486f5aaef764a7b2
--- /dev/null
+++ b/Images/OMRSheet3.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:2072f72607f9e9363cba21b700c86e5dffd3fce0cf2d7e2d1288c23b5066b257
+size 237950
diff --git a/Images/OMRTest.jpg b/Images/OMRTest.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..97e9257fa2472bf70f02de39c51f999c39f5ec8a
--- /dev/null
+++ b/Images/OMRTest.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:daaa90fa7b3fe40fbf1dd0fee2941de0c0d138290f1d32523979b95bcd253907
+size 273119
diff --git a/Images/OcrSheetMarked.jpg b/Images/OcrSheetMarked.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..87ec203468051d0ce456111cd85b5d117e141f10
--- /dev/null
+++ b/Images/OcrSheetMarked.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:328a515ffb5367799509a284fc85bca8114508337db76049e86c9c8415b96221
+size 221191
diff --git a/Images/omr_answer_key.pdf b/Images/omr_answer_key.pdf
new file mode 100644
index 0000000000000000000000000000000000000000..e769a97a4d0fd07a11655997a915b9892c3085f2
--- /dev/null
+++ b/Images/omr_answer_key.pdf
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:879c8f7aa369a6f0cbc65f9a5279a18ae42a35d541ba53e70b140178e83acf3f
+size 105714
diff --git a/Images/omr_answer_key.png b/Images/omr_answer_key.png
new file mode 100644
index 0000000000000000000000000000000000000000..c9e3dcccfc8af7c60becb3a4cc25f650861db98e
--- /dev/null
+++ b/Images/omr_answer_key.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4439b6a479cb9fea92de9656b900a528052616ae8e8962a6c33b8e81ed7f327a
+size 267398
diff --git a/Images/question.png b/Images/question.png
new file mode 100644
index 0000000000000000000000000000000000000000..4d10d743f3d7f5fb533a86ff4e0c1a287a8e75c0
Binary files /dev/null and b/Images/question.png differ
diff --git a/Images/question_paper.pdf b/Images/question_paper.pdf
new file mode 100644
index 0000000000000000000000000000000000000000..5e464937d24356e93f2457e463659deec9b21605
Binary files /dev/null and b/Images/question_paper.pdf differ
diff --git a/Images/question_paper.png b/Images/question_paper.png
new file mode 100644
index 0000000000000000000000000000000000000000..a5a2f9cf04c0bb5f790a542f3480199bf24670c3
--- /dev/null
+++ b/Images/question_paper.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:ebf1ff49a9b76cf2259890ec30c7c369f60c900331d06bc81d096e2d3efbe134
+size 199992
diff --git a/Images/test.jpg b/Images/test.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3344b5f30842cb22944fede561995c526e3bc3d7
--- /dev/null
+++ b/Images/test.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:dc40095b38d1a4e8248d464ba1e14d943e6ce2eaf5a0173316c4ce3bcbfabf5e
+size 331992
diff --git a/OMRChecker/.pre-commit-config.yaml b/OMRChecker/.pre-commit-config.yaml
new file mode 100644
index 0000000000000000000000000000000000000000..1fe9847c20f354b81701b994f9589c0e37488464
--- /dev/null
+++ b/OMRChecker/.pre-commit-config.yaml
@@ -0,0 +1,59 @@
+exclude: "__snapshots__/.*$"
+default_install_hook_types: [pre-commit, pre-push]
+repos:
+ - repo: https://github.com/pre-commit/pre-commit-hooks
+ rev: v4.4.0
+ hooks:
+ - id: check-yaml
+ stages: [commit]
+ - id: check-added-large-files
+ args: ['--maxkb=300']
+ fail_fast: false
+ stages: [commit]
+ - id: pretty-format-json
+ args: ['--autofix', '--no-sort-keys']
+ - id: end-of-file-fixer
+ exclude_types: ["csv", "json"]
+ stages: [commit]
+ - id: trailing-whitespace
+ stages: [commit]
+ - repo: https://github.com/pycqa/isort
+ rev: 5.12.0
+ hooks:
+ - id: isort
+ args: ["--profile", "black"]
+ stages: [commit]
+ - repo: https://github.com/psf/black
+ rev: 23.3.0
+ hooks:
+ - id: black
+ fail_fast: true
+ stages: [commit]
+ - repo: https://github.com/pycqa/flake8
+ rev: 6.0.0
+ hooks:
+ - id: flake8
+ args:
+ - "--ignore=E501,W503,E203,E741,F541" # Line too long, Line break occurred before a binary operator, Whitespace before ':'
+ fail_fast: true
+ stages: [commit]
+ - repo: local
+ hooks:
+ - id: pytest-on-commit
+ name: Running single sample test
+ entry: python3 -m pytest -rfpsxEX --disable-warnings --verbose -k sample1
+ language: system
+ pass_filenames: false
+ always_run: true
+ fail_fast: true
+ stages: [commit]
+ - repo: local
+ hooks:
+ - id: pytest-on-push
+ name: Running all tests before push...
+ entry: python3 -m pytest -rfpsxEX --disable-warnings --verbose --durations=3
+ language: system
+ pass_filenames: false
+ always_run: true
+ fail_fast: true
+ stages: [push]
diff --git a/OMRChecker/.pylintrc b/OMRChecker/.pylintrc
new file mode 100644
index 0000000000000000000000000000000000000000..369568f5560bc52faf695ad91c6df93c568195f4
--- /dev/null
+++ b/OMRChecker/.pylintrc
@@ -0,0 +1,43 @@
+[BASIC]
+# Regular expression matching correct variable names. Overrides variable-naming-style.
+# snake_case with single letter regex -
+variable-rgx=[a-z0-9_]{1,30}$
+
+# Good variable names which should always be accepted, separated by a comma.
+good-names=x,y,pt
+
+[MESSAGES CONTROL]
+
+# Disable the message, report, category or checker with the given id(s). You
+# can either give multiple identifiers separated by comma (,) or put this
+# option multiple times (only on the command line, not in the configuration
+# file where it should appear only once). You can also use "--disable=all" to
+# disable everything first and then reenable specific checks. For example, if
+# you want to run only the similarities checker, you can use "--disable=all
+# --enable=similarities". If you want to run only the classes checker, but have
+# no Warning level messages displayed, use "--disable=all --enable=classes
+# --disable=W".
+disable=import-error,
+ unresolved-import,
+ too-few-public-methods,
+ missing-docstring,
+ relative-beyond-top-level,
+ too-many-instance-attributes,
+ bad-continuation,
+ no-member
+
+# Note: bad-continuation is a false positive showing bug in pylint
+# https://github.com/psf/black/issues/48
+
+
+[REPORTS]
+# Set the output format. Available formats are text, parseable, colorized, json
+# and msvs (visual studio). You can also give a reporter class, e.g.
+# mypackage.mymodule.MyReporterClass.
+output-format=text
+
+# Tells whether to display a full report or only the messages.
+reports=no
+
+# Activate the evaluation score.
+score=yes
diff --git a/OMRChecker/CODE_OF_CONDUCT.md b/OMRChecker/CODE_OF_CONDUCT.md
new file mode 100644
index 0000000000000000000000000000000000000000..4a84b4a0ddccf4a14ef9b5c0b65f6eafdb5549cc
--- /dev/null
+++ b/OMRChecker/CODE_OF_CONDUCT.md
@@ -0,0 +1,133 @@
+
+# Contributor Covenant Code of Conduct
+
+## Our Pledge
+
+We as members, contributors, and leaders pledge to make participation in our
+community a harassment-free experience for everyone, regardless of age, body
+size, visible or invisible disability, ethnicity, sex characteristics, gender
+identity and expression, level of experience, education, socio-economic status,
+nationality, personal appearance, race, caste, color, religion, or sexual
+identity and orientation.
+
+We pledge to act and interact in ways that contribute to an open, welcoming,
+diverse, inclusive, and healthy community.
+
+## Our Standards
+
+Examples of behavior that contributes to a positive environment for our
+community include:
+
+* Demonstrating empathy and kindness toward other people
+* Being respectful of differing opinions, viewpoints, and experiences
+* Giving and gracefully accepting constructive feedback
+* Accepting responsibility and apologizing to those affected by our mistakes,
+ and learning from the experience
+* Focusing on what is best not just for us as individuals, but for the overall
+ community
+
+Examples of unacceptable behavior include:
+
+* The use of sexualized language or imagery, and sexual attention or advances of
+ any kind
+* Trolling, insulting or derogatory comments, and personal or political attacks
+* Public or private harassment
+* Publishing others' private information, such as a physical or email address,
+ without their explicit permission
+* Other conduct which could reasonably be considered inappropriate in a
+ professional setting
+
+## Enforcement Responsibilities
+
+Community leaders are responsible for clarifying and enforcing our standards of
+acceptable behavior and will take appropriate and fair corrective action in
+response to any behavior that they deem inappropriate, threatening, offensive,
+or harmful.
+
+Community leaders have the right and responsibility to remove, edit, or reject
+comments, commits, code, wiki edits, issues, and other contributions that are
+not aligned to this Code of Conduct, and will communicate reasons for moderation
+decisions when appropriate.
+
+## Scope
+
+This Code of Conduct applies within all community spaces, and also applies when
+an individual is officially representing the community in public spaces.
+Examples of representing our community include using an official email address,
+posting via an official social media account, or acting as an appointed
+representative at an online or offline event.
+
+## Enforcement
+
+Instances of abusive, harassing, or otherwise unacceptable behavior may be
+reported to the community leaders responsible for enforcement at
+[INSERT CONTACT METHOD].
+All complaints will be reviewed and investigated promptly and fairly.
+
+All community leaders are obligated to respect the privacy and security of the
+reporter of any incident.
+
+## Enforcement Guidelines
+
+Community leaders will follow these Community Impact Guidelines in determining
+the consequences for any action they deem in violation of this Code of Conduct:
+
+### 1. Correction
+
+**Community Impact**: Use of inappropriate language or other behavior deemed
+unprofessional or unwelcome in the community.
+
+**Consequence**: A private, written warning from community leaders, providing
+clarity around the nature of the violation and an explanation of why the
+behavior was inappropriate. A public apology may be requested.
+
+### 2. Warning
+
+**Community Impact**: A violation through a single incident or series of
+actions.
+
+**Consequence**: A warning with consequences for continued behavior. No
+interaction with the people involved, including unsolicited interaction with
+those enforcing the Code of Conduct, for a specified period of time. This
+includes avoiding interactions in community spaces as well as external channels
+like social media. Violating these terms may lead to a temporary or permanent
+ban.
+
+### 3. Temporary Ban
+
+**Community Impact**: A serious violation of community standards, including
+sustained inappropriate behavior.
+
+**Consequence**: A temporary ban from any sort of interaction or public
+communication with the community for a specified period of time. No public or
+private interaction with the people involved, including unsolicited interaction
+with those enforcing the Code of Conduct, is allowed during this period.
+Violating these terms may lead to a permanent ban.
+
+### 4. Permanent Ban
+
+**Community Impact**: Demonstrating a pattern of violation of community
+standards, including sustained inappropriate behavior, harassment of an
+individual, or aggression toward or disparagement of classes of individuals.
+
+**Consequence**: A permanent ban from any sort of public interaction within the
+community.
+
+## Attribution
+
+This Code of Conduct is adapted from the [Contributor Covenant][homepage],
+version 2.1, available at
+[https://www.contributor-covenant.org/version/2/1/code_of_conduct.html][v2.1].
+
+Community Impact Guidelines were inspired by
+[Mozilla's code of conduct enforcement ladder][Mozilla CoC].
+
+For answers to common questions about this code of conduct, see the FAQ at
+[https://www.contributor-covenant.org/faq][FAQ]. Translations are available at
+[https://www.contributor-covenant.org/translations][translations].
+
+[homepage]: https://www.contributor-covenant.org
+[v2.1]: https://www.contributor-covenant.org/version/2/1/code_of_conduct.html
+[Mozilla CoC]: https://github.com/mozilla/diversity
+[FAQ]: https://www.contributor-covenant.org/faq
+[translations]: https://www.contributor-covenant.org/translations
diff --git a/OMRChecker/CONTRIBUTING.md b/OMRChecker/CONTRIBUTING.md
new file mode 100644
index 0000000000000000000000000000000000000000..c649a048362ecfe9ac4e260feac8b0f3f8447a94
--- /dev/null
+++ b/OMRChecker/CONTRIBUTING.md
@@ -0,0 +1,32 @@
+# How to contribute
+So you want to write code and get it landed in the official OMRChecker repository?
+First, fork our repository into your own GitHub account, and create a local clone of it as described in the installation instructions.
+The latter will be used to get new features implemented or bugs fixed.
+
+Once done and you have the code locally on the disk, you can get started. We advise you to not work directly on the master branch,
+but to create a separate branch for each issue you are working on. That way you can easily switch between different work,
+and you can update each one for the latest changes on the upstream master individually.
+
+
+# Writing Code
+For writing the code just follow the [Pep8 Python style](https://peps.python.org/pep-0008/) guide, If there is something unclear about the style, just look at existing code which might help you to understand it better.
+
+Also, try to use commits with [conventional messages](https://www.conventionalcommits.org/en/v1.0.0/#summary).
+
+
+# Code Formatting
+Before committing your code, make sure to run the following command to format your code according to the PEP8 style guide:
+```.sh
+pip install -r requirements.dev.txt && pre-commit install
+```
+
+Run `pre-commit` before committing your changes:
+```.sh
+git add .
+pre-commit run -a
+```
+
+# Where to contribute from
+
+- You can pickup any open [issues](https://github.com/Udayraj123/OMRChecker/issues) to solve.
+- You can also check out the [ideas list](https://github.com/users/Udayraj123/projects/2/views/1)
diff --git a/OMRChecker/Contributors.md b/OMRChecker/Contributors.md
new file mode 100644
index 0000000000000000000000000000000000000000..b7a428a77223e4e227c9b74c3e54eb43444f9017
--- /dev/null
+++ b/OMRChecker/Contributors.md
@@ -0,0 +1,22 @@
+# Contributors
+
+- [Udayraj123](https://github.com/Udayraj123)
+- [leongwaikay](https://github.com/leongwaikay)
+- [deepakgouda](https://github.com/deepakgouda)
+- [apurva91](https://github.com/apurva91)
+- [sparsh2706](https://github.com/sparsh2706)
+- [namit2saxena](https://github.com/namit2saxena)
+- [Harsh-Kapoorr](https://github.com/Harsh-Kapoorr)
+- [Sandeep-1507](https://github.com/Sandeep-1507)
+- [SpyzzVVarun](https://github.com/SpyzzVVarun)
+- [asc249](https://github.com/asc249)
+- [05Alston](https://github.com/05Alston)
+- [Antibodyy](https://github.com/Antibodyy)
+- [infinity1729](https://github.com/infinity1729)
+- [Rohan-G](https://github.com/Rohan-G)
+- [UjjwalMahar](https://github.com/UjjwalMahar)
+- [Kurtsley](https://github.com/Kurtsley)
+- [gaursagar21](https://github.com/gaursagar21)
+- [aayushibansal2001](https://github.com/aayushibansal2001)
+- [ShamanthVallem](https://github.com/ShamanthVallem)
+- [rudrapsc](https://github.com/rudrapsc)
diff --git a/OMRChecker/LICENSE b/OMRChecker/LICENSE
new file mode 100644
index 0000000000000000000000000000000000000000..33cdce9d8e013750ea0e519ffc9a03b2d14d4d0c
--- /dev/null
+++ b/OMRChecker/LICENSE
@@ -0,0 +1,22 @@
+MIT License
+
+Copyright (c) 2024-present Udayraj Deshmukh and other contributors
+
+Permission is hereby granted, free of charge, to any person obtaining
+a copy of this software and associated documentation files (the
+"Software"), to deal in the Software without restriction, including
+without limitation the rights to use, copy, modify, merge, publish,
+distribute, sublicense, and/or sell copies of the Software, and to
+permit persons to whom the Software is furnished to do so, subject to
+the following conditions:
+
+The above copyright notice and this permission notice shall be
+included in all copies or substantial portions of the Software.
+
+THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND,
+EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF
+MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND
+NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE
+LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION
+OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION
+WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
diff --git a/OMRChecker/README.md b/OMRChecker/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..fba176b81f7766e0aacd394b79d0db5573015512
--- /dev/null
+++ b/OMRChecker/README.md
@@ -0,0 +1,359 @@
+# OMR Checker
+
+Read OMR sheets fast and accurately using a scanner 🖨 or your phone 🤳.
+
+## What is OMR?
+
+OMR stands for Optical Mark Recognition, used to detect and interpret human-marked data on documents. OMR refers to the process of reading and evaluating OMR sheets, commonly used in exams, surveys, and other forms.
+
+#### **Quick Links**
+
+- [Installation](#getting-started)
+- [User Guide](https://github.com/Udayraj123/OMRChecker/wiki)
+- [Contributor Guide](https://github.com/Udayraj123/OMRChecker/blob/master/CONTRIBUTING.md)
+- [Project Ideas List](https://github.com/users/Udayraj123/projects/2/views/1)
+
+
+
+[](https://github.com/Udayraj123/OMRChecker/pull/new/master)
+[](https://github.com/Udayraj123/OMRChecker/pulls?q=is%3Aclosed)
+[](https://GitHub.com/Udayraj123/OMRChecker/issues?q=is%3Aissue+is%3Aclosed)
+[](https://github.com/Udayraj123/OMRChecker/issues/5)
+
+
+
+[](https://GitHub.com/Udayraj123/OMRChecker/stargazers/)
+[](https://discord.gg/qFv2Vqf)
+
+
+
+## 🎯 Features
+
+A full-fledged OMR checking software that can read and evaluate OMR sheets scanned at any angle and having any color.
+
+| Specs ![]() |  
|   ![]() |
+| :--------------------- | :--------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| 💯 **Accurate** | Currently nearly 100% accurate on good quality document scans; and about 90% accurate on mobile images. |
+| 💪🏿 **Robust** | Supports low resolution, xeroxed sheets. See [**Robustness**](https://github.com/Udayraj123/OMRChecker/wiki/Robustness) for more. |
+| ⏩ **Fast** | Current processing speed without any optimization is 200 OMRs/minute. |
+| ✅ **Customizable** | [Easily apply](https://github.com/Udayraj123/OMRChecker/wiki/User-Guide) to custom OMR layouts, surveys, etc. |
+| 📊 **Visually Rich** | [Get insights](https://github.com/Udayraj123/OMRChecker/wiki/Rich-Visuals) to configure and debug easily. |
+| 🎈 **Lightweight** | Very minimal core code size. |
+| 🏫 **Large Scale** | Tested on a large scale at [Technothlon](https://en.wikipedia.org/wiki/Technothlon). |
+| 👩🏿💻 **Dev Friendly** | [Pylinted](http://pylint.pycqa.org/) and [Black formatted](https://github.com/psf/black) code. Also has a [developer community](https://discord.gg/qFv2Vqf) on discord. |
+
+Note: For solving interesting challenges, developers can check out [**TODOs**](https://github.com/Udayraj123/OMRChecker/wiki/TODOs).
+
+See the complete guide and details at [Project Wiki](https://github.com/Udayraj123/OMRChecker/wiki/).
+
+
+
+## 💡 What can OMRChecker do for me?
+
+Once you configure the OMR layout, just throw images of the sheets at the software; and you'll get back the marked responses in an excel sheet!
+
+Images can be taken from various angles as shown below-
+
+
|
+| :--------------------- | :--------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| 💯 **Accurate** | Currently nearly 100% accurate on good quality document scans; and about 90% accurate on mobile images. |
+| 💪🏿 **Robust** | Supports low resolution, xeroxed sheets. See [**Robustness**](https://github.com/Udayraj123/OMRChecker/wiki/Robustness) for more. |
+| ⏩ **Fast** | Current processing speed without any optimization is 200 OMRs/minute. |
+| ✅ **Customizable** | [Easily apply](https://github.com/Udayraj123/OMRChecker/wiki/User-Guide) to custom OMR layouts, surveys, etc. |
+| 📊 **Visually Rich** | [Get insights](https://github.com/Udayraj123/OMRChecker/wiki/Rich-Visuals) to configure and debug easily. |
+| 🎈 **Lightweight** | Very minimal core code size. |
+| 🏫 **Large Scale** | Tested on a large scale at [Technothlon](https://en.wikipedia.org/wiki/Technothlon). |
+| 👩🏿💻 **Dev Friendly** | [Pylinted](http://pylint.pycqa.org/) and [Black formatted](https://github.com/psf/black) code. Also has a [developer community](https://discord.gg/qFv2Vqf) on discord. |
+
+Note: For solving interesting challenges, developers can check out [**TODOs**](https://github.com/Udayraj123/OMRChecker/wiki/TODOs).
+
+See the complete guide and details at [Project Wiki](https://github.com/Udayraj123/OMRChecker/wiki/).
+
+
+
+## 💡 What can OMRChecker do for me?
+
+Once you configure the OMR layout, just throw images of the sheets at the software; and you'll get back the marked responses in an excel sheet!
+
+Images can be taken from various angles as shown below-
+
+
+  +
+
+
+### Code in action on images taken by scanner:
+
+
+  +
+
+
+
+
+### Code in action on images taken by a mobile phone:
+
+
+  +
+
+
+## Visuals
+
+### Processing steps
+
+See step-by-step processing of any OMR sheet:
+
+
+
+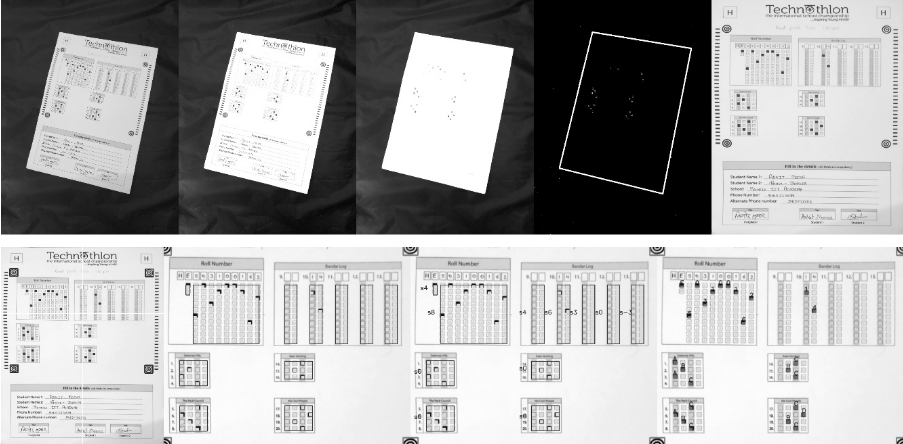  +
+
+
+
+ *Note: This image is generated by the code itself!*
+
+
+### Output
+
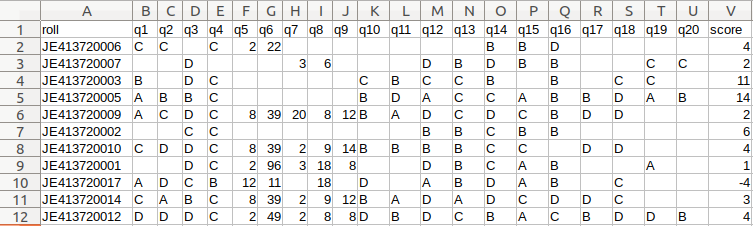
+Get a CSV sheet containing the detected responses and evaluated scores:
+
+
+
+  +
+
+
+
+
+We now support [colored outputs](https://github.com/Udayraj123/OMRChecker/wiki/%5Bv2%5D-About-Evaluation) as well. Here's a sample output on another image -
+
+
+  +
+
+
+
+
+#### There are many more visuals in the wiki. Check them out [here!](https://github.com/Udayraj123/OMRChecker/wiki/Rich-Visuals)
+
+## Getting started
+
+
+
+**Operating system:** OSX or Linux is recommended although Windows is also supported.
+
+### 1. Install global dependencies
+
+ 
+
+To check if python3 and pip is already installed:
+
+```bash
+python3 --version
+python3 -m pip --version
+```
+
+
+ Install Python3
+
+To install python3 follow instructions [here](https://www.python.org/downloads/)
+
+To install pip - follow instructions [here](https://pip.pypa.io/en/stable/installation/)
+
+
+
+Install OpenCV
+
+**Any installation method is fine.**
+
+Recommended:
+
+```bash
+python3 -m pip install --user --upgrade pip
+python3 -m pip install --user opencv-python
+python3 -m pip install --user opencv-contrib-python
+```
+
+More details on pip install openCV [here](https://www.pyimagesearch.com/2018/09/19/pip-install-opencv/).
+
+
+
+
+
+Extra steps(for Linux users only)
+
+Installing missing libraries(if any):
+
+On a fresh computer, some of the libraries may get missing in event after a successful pip install. Install them using following commands[(ref)](https://www.pyimagesearch.com/2018/05/28/ubuntu-18-04-how-to-install-opencv/):
+
+```bash
+sudo apt-get install -y build-essential cmake unzip pkg-config
+sudo apt-get install -y libjpeg-dev libpng-dev libtiff-dev
+sudo apt-get install -y libavcodec-dev libavformat-dev libswscale-dev libv4l-dev
+sudo apt-get install -y libatlas-base-dev gfortran
+```
+
+
+
+### 2. Install project dependencies
+
+Clone the repo
+
+```bash
+git clone https://github.com/Udayraj123/OMRChecker
+cd OMRChecker/
+```
+
+Install pip requirements
+
+```bash
+python3 -m pip install --user -r requirements.txt
+```
+
+_**Note:** If you face a distutils error in pip, use `--ignore-installed` flag in above command._
+
+
+
+### 3. Run the code
+
+1. First copy and examine the sample data to know how to structure your inputs:
+ ```bash
+ cp -r ./samples/sample1 inputs/
+ # Note: you may remove previous inputs (if any) with `mv inputs/* ~/.trash`
+ # Change the number N in sampleN to see more examples
+ ```
+2. Run OMRChecker:
+ ```bash
+ python3 main.py
+ ```
+
+Alternatively you can also use `python3 main.py -i ./samples/sample1`.
+
+Each example in the samples folder demonstrates different ways in which OMRChecker can be used.
+
+### Common Issues
+
+
+
+ 1. [Windows] ERROR: Could not open requirements file
+
+Command: python3 -m pip install --user -r requirements.txt
+
+ Link to Solution: #54
+
+
+
+2. [Linux] ERROR: No module named pip
+
+Command: python3 -m pip install --user --upgrade pip
+
+ Link to Solution: #70
+
+
+## OMRChecker for custom OMR Sheets
+
+1. First, [create your own template.json](https://github.com/Udayraj123/OMRChecker/wiki/User-Guide).
+2. Configure the tuning parameters.
+3. Run OMRChecker with appropriate arguments (See full usage).
+
+
+## Full Usage
+
+```
+python3 main.py [--setLayout] [--inputDir dir1] [--outputDir dir1]
+```
+
+Explanation for the arguments:
+
+`--setLayout`: Set up OMR template layout - modify your json file and run again until the template is set.
+
+`--inputDir`: Specify an input directory.
+
+`--outputDir`: Specify an output directory.
+
+
+
+ Deprecation logs
+
+
+- The old `--noCropping` flag has been replaced with the 'CropPage' plugin in "preProcessors" of the template.json(see [samples](https://github.com/Udayraj123/OMRChecker/tree/master/samples)).
+- The `--autoAlign` flag is deprecated due to low performance on a generic OMR sheet
+- The `--template` flag is deprecated and instead it's recommended to keep the template file at the parent folder containing folders of different images
+
+
+
+
+## FAQ
+
+
+
+Why is this software free?
+
+
+This project was born out of a student-led organization called as [Technothlon](https://technothlon.techniche.org.in). It is a logic-based international school championship organized by students of IIT Guwahati. Being a non-profit organization, and after seeing it work fabulously at such a large scale we decided to share this tool with the world. The OMR checking processes still involves so much tediousness which we aim to reduce dramatically.
+
+We believe in the power of open source! Currently, OMRChecker is in an intermediate stage where only developers can use it. We hope to see it become more user-friendly as well as robust from exposure to different inputs from you all!
+
+[](https://github.com/ellerbrock/open-source-badges/)
+
+
+
+
+
+Can I use this code in my (public) work?
+
+
+OMRChecker can be forked and modified. You are encouraged to play with it and we would love to see your own projects in action!
+
+It is published under the [MIT license](https://github.com/Udayraj123/OMRChecker/blob/master/LICENSE).
+
+
+
+
+
+What are the ways to contribute?
+
+
+
+
+- Join the developer community on [Discord](https://discord.gg/qFv2Vqf) to fix [issues](https://github.com/Udayraj123/OMRChecker/issues) with OMRChecker.
+
+- If this project saved you large costs on OMR Software licenses, or saved efforts to make one. Consider donating an amount of your choice(donate section).
+
+
+
+
+
+
+
+## Credits
+
+_A Huge thanks to:_
+_**Adrian Rosebrock** for his exemplary blog:_ https://pyimagesearch.com
+
+_**Harrison Kinsley** aka sentdex for his [video tutorials](https://www.youtube.com/watch?v=Z78zbnLlPUA&list=PLQVvvaa0QuDdttJXlLtAJxJetJcqmqlQq) and many other resources._
+
+_**Satya Mallic** for his resourceful blog:_ https://www.learnopencv.com
+
+_And to other amazing people from all over the globe who've made significant improvements in this project._
+
+_Thank you!_
+
+
+
+## Related Projects
+
+Here's a snapshot of the [Android OMR Helper App (archived)](https://github.com/Udayraj123/AndroidOMRHelper):
+
+
+
+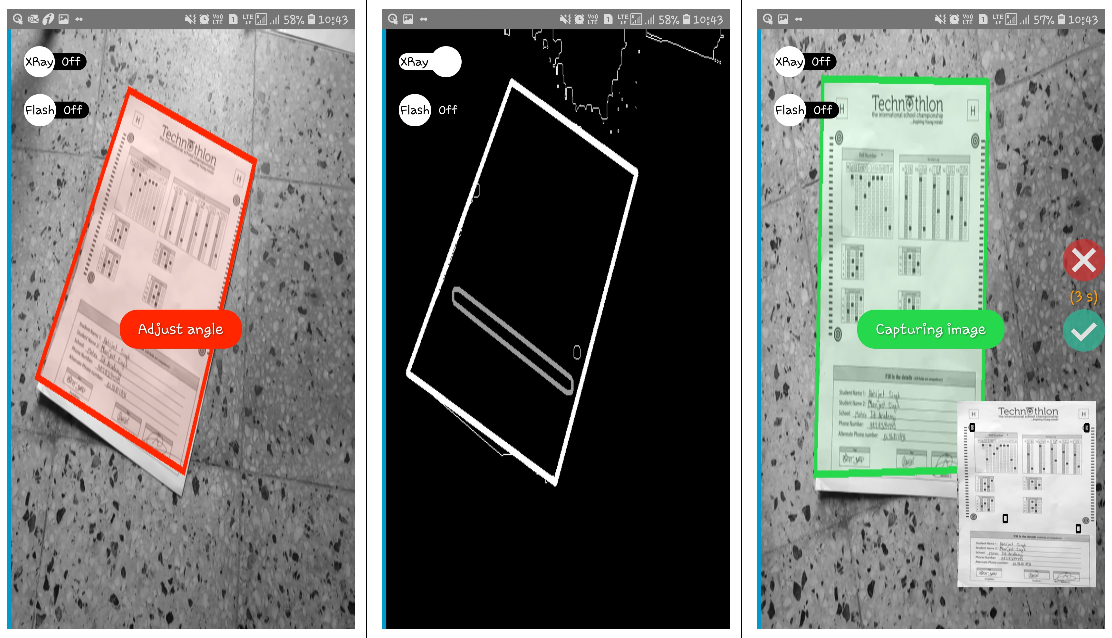  +
+
+
+
+
+## Stargazers over time
+
+[](https://starchart.cc/Udayraj123/OMRChecker)
+
+---
+
+Made with ❤️ by Awesome Contributors
+
+
+  +
+
+---
+
+### License
+
+[](https://github.com/Udayraj123/OMRChecker/blob/master/LICENSE)
+
+For more details see [LICENSE](https://github.com/Udayraj123/OMRChecker/blob/master/LICENSE).
+
+### Donate
+
+
+
+
+---
+
+### License
+
+[](https://github.com/Udayraj123/OMRChecker/blob/master/LICENSE)
+
+For more details see [LICENSE](https://github.com/Udayraj123/OMRChecker/blob/master/LICENSE).
+
+### Donate
+
+ [](https://www.paypal.me/Udayraj123/500)
+
+_Find OMRChecker on_ [**_Product Hunt_**](https://www.producthunt.com/posts/omr-checker/) **|** [**_Reddit_**](https://www.reddit.com/r/computervision/comments/ccbj6f/omrchecker_grade_exams_using_python_and_opencv/) **|** [**Discord**](https://discord.gg/qFv2Vqf) **|** [**Linkedin**](https://www.linkedin.com/pulse/open-source-talks-udayraj-udayraj-deshmukh/) **|** [**goodfirstissue.dev**](https://goodfirstissue.dev/language/python) **|** [**codepeak.tech**](https://www.codepeak.tech/) **|** [**fossoverflow.dev**](https://fossoverflow.dev/projects) **|** [**Interview on Console by CodeSee**](https://console.substack.com/p/console-140) **|** [**Open Source Hub**](https://opensourcehub.io/udayraj123/omrchecker)
+
+
+
diff --git a/OMRChecker/docs/assets/colored_output.jpg b/OMRChecker/docs/assets/colored_output.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3cafa473b9a1134f845858be4cf0e98f3bd3bb7c
Binary files /dev/null and b/OMRChecker/docs/assets/colored_output.jpg differ
diff --git a/OMRChecker/inputs/OMRImage.jpg b/OMRChecker/inputs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c9e3dcccfc8af7c60becb3a4cc25f650861db98e
--- /dev/null
+++ b/OMRChecker/inputs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4439b6a479cb9fea92de9656b900a528052616ae8e8962a6c33b8e81ed7f327a
+size 267398
diff --git a/OMRChecker/inputs/template.json b/OMRChecker/inputs/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..823fe2e58a4182ef04d1fff99ecf146b5ec53aae
--- /dev/null
+++ b/OMRChecker/inputs/template.json
@@ -0,0 +1,37 @@
+{
+ "pageDimensions": [
+ 1122,
+ 1600
+ ],
+ "bubbleDimensions": [
+ 48,
+ 50
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 100,
+ 175
+ ],
+ "bubblesGap": 55,
+ "labelsGap": 67,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q02block": {
+ "origin": [
+ 100,
+ 845
+ ],
+ "bubblesGap": 58,
+ "labelsGap": 70,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ }
+
+}
\ No newline at end of file
diff --git a/OMRChecker/main.py b/OMRChecker/main.py
new file mode 100644
index 0000000000000000000000000000000000000000..19e3ebd43d4be2a096cf709b04817247f9952506
--- /dev/null
+++ b/OMRChecker/main.py
@@ -0,0 +1,99 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+
+import argparse
+import sys
+from pathlib import Path
+
+from src.entry import entry_point
+from src.logger import logger
+
+
+def parse_args():
+ # construct the argument parse and parse the arguments
+ argparser = argparse.ArgumentParser()
+
+ argparser.add_argument(
+ "-i",
+ "--inputDir",
+ default=["inputs"],
+ # https://docs.python.org/3/library/argparse.html#nargs
+ nargs="*",
+ required=False,
+ type=str,
+ dest="input_paths",
+ help="Specify an input directory.",
+ )
+
+ argparser.add_argument(
+ "-d",
+ "--debug",
+ required=False,
+ dest="debug",
+ action="store_false",
+ help="Enables debugging mode for showing detailed errors",
+ )
+
+ argparser.add_argument(
+ "-o",
+ "--outputDir",
+ default="outputs",
+ required=False,
+ dest="output_dir",
+ help="Specify an output directory.",
+ )
+
+ argparser.add_argument(
+ "-a",
+ "--autoAlign",
+ required=False,
+ dest="autoAlign",
+ action="store_true",
+ help="(experimental) Enables automatic template alignment - \
+ use if the scans show slight misalignments.",
+ )
+
+ argparser.add_argument(
+ "-l",
+ "--setLayout",
+ required=False,
+ dest="setLayout",
+ action="store_true",
+ help="Set up OMR template layout - modify your json file and \
+ run again until the template is set.",
+ )
+
+ (
+ args,
+ unknown,
+ ) = argparser.parse_known_args()
+
+ args = vars(args)
+
+ if len(unknown) > 0:
+ logger.warning(f"\nError: Unknown arguments: {unknown}", unknown)
+ argparser.print_help()
+ exit(11)
+ return args
+
+
+def entry_point_for_args(args):
+ if args["debug"] is True:
+ # Disable tracebacks
+ sys.tracebacklimit = 0
+ for root in args["input_paths"]:
+ entry_point(
+ Path(root),
+ args,
+ )
+
+
+if __name__ == "__main__":
+ args = parse_args()
+ entry_point_for_args(args)
diff --git a/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..d8d1a2325170014745edf3503b40e7c7bda160dc
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:f16c45dda1d02bf06e33a6d219c8282c363bf3943b94960eede1efe2bc6204d9
+size 408744
diff --git a/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3a968368fa191241dda21ea0c41f5d2d840f2e52
--- /dev/null
+++ b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:7e9678555042535c83ab920d0b616d4f99f9f08017629d46a15eb64d7c3abd3c
+size 407394
diff --git a/OMRChecker/outputs/Images/Manual/ErrorFiles.csv b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Results/Results_10PM.csv b/OMRChecker/outputs/Images/Results/Results_10PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..f3b83ec4b12f9e002ca0824ee43fc41f1f9ae30a
--- /dev/null
+++ b/OMRChecker/outputs/Images/Results/Results_10PM.csv
@@ -0,0 +1,2 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\Images\OMRSheet.jpg","outputs\Images\CheckedOMRs\OMRSheet.jpg","0","C","D","A","C","B","C","C","D","D","B","B","B","D","C","C","ABCD","B","ABCD","C","ABCD"
diff --git a/OMRChecker/outputs/Manual/ErrorFiles.csv b/OMRChecker/outputs/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..a9bb85c601677eefe49c6a75b39d5bd404506322
--- /dev/null
+++ b/OMRChecker/outputs/Manual/ErrorFiles.csv
@@ -0,0 +1,3 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
diff --git a/OMRChecker/outputs/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/pyproject.toml b/OMRChecker/pyproject.toml
new file mode 100644
index 0000000000000000000000000000000000000000..efa55e8fc8bf1623d8569abd406fef4d81270291
--- /dev/null
+++ b/OMRChecker/pyproject.toml
@@ -0,0 +1,18 @@
+[tool.black]
+exclude = '''
+(
+ /(
+ \.eggs # exclude a few common directories in the
+ | \.git # root of the project
+ | \.venv
+ | _build
+ | build
+ | dist
+ )/
+ | foo.py # also separately exclude a file named foo.py in
+ # the root of the project
+)
+'''
+include = '\.pyi?$'
+line-length = 88
+target-version = ['py37']
diff --git a/OMRChecker/pytest.ini b/OMRChecker/pytest.ini
new file mode 100644
index 0000000000000000000000000000000000000000..417c4a2dd810294aa9afaa9b4f73464fc9808c96
--- /dev/null
+++ b/OMRChecker/pytest.ini
@@ -0,0 +1,6 @@
+# pytest.ini
+[pytest]
+minversion = 7.0
+addopts = -qq --capture=no
+testpaths =
+ src/tests
diff --git a/OMRChecker/requirements.dev.txt b/OMRChecker/requirements.dev.txt
new file mode 100644
index 0000000000000000000000000000000000000000..fb31b5f80558481a3f2a5edbb741b363a1bd1089
--- /dev/null
+++ b/OMRChecker/requirements.dev.txt
@@ -0,0 +1,7 @@
+-r requirements.txt
+flake8>=6.0.0
+freezegun>=1.2.2
+pre-commit>=3.3.3
+pytest-mock>=3.11.1
+pytest>=7.4.0
+syrupy>=4.0.4
diff --git a/OMRChecker/requirements.txt b/OMRChecker/requirements.txt
new file mode 100644
index 0000000000000000000000000000000000000000..d0c3af875835b492a48ef689c3dc9335eb7cdca7
--- /dev/null
+++ b/OMRChecker/requirements.txt
@@ -0,0 +1,8 @@
+deepmerge>=1.1.0
+dotmap>=1.3.30
+jsonschema>=4.17.3
+matplotlib>=3.7.1
+numpy>=1.25.0
+pandas>=2.0.2
+rich>=13.4.2
+screeninfo>=0.8.1
diff --git a/OMRChecker/samples/answer-key/using-csv/adrian_omr.png b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png differ
diff --git a/OMRChecker/samples/answer-key/using-csv/answer_key.csv b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..acf09be611a66dfaeb7127e7249eadfbba6b0881
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
@@ -0,0 +1,5 @@
+q1,C
+q2,E
+q3,A
+q4,B
+q5,B
\ No newline at end of file
diff --git a/OMRChecker/samples/answer-key/using-csv/evaluation.json b/OMRChecker/samples/answer-key/using-csv/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/using-csv/template.json b/OMRChecker/samples/answer-key/using-csv/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/evaluation.json b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..09508b18e3490efcd68d6cb0b57a03d6e203f2ca
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
@@ -0,0 +1,35 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..5"
+ ],
+ "answers_in_order": [
+ "C",
+ "E",
+ [
+ "A",
+ "C"
+ ],
+ [
+ [
+ "B",
+ 2
+ ],
+ [
+ "C",
+ "3/2"
+ ]
+ ],
+ "C"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/answer-key/weighted-answers/template.json b/OMRChecker/samples/answer-key/weighted-answers/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..661d5f4fa73d7eea57e73837cf3862a25b9efcc3
Binary files /dev/null and b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg differ
diff --git a/OMRChecker/samples/community/Antibodyy/template.json b/OMRChecker/samples/community/Antibodyy/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..55e016e8eb402156d62f6553067b68abb37377ee
--- /dev/null
+++ b/OMRChecker/samples/community/Antibodyy/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 299,
+ 398
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 79
+ ],
+ "bubblesGap": 43,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "q1..6"
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-1.png b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
new file mode 100644
index 0000000000000000000000000000000000000000..5ed00d194a806d58c3db1dab7831796df7d4aaef
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:ed3024e5b709fbbdb1bcbef35dcfa50203ddb811f74b26e657d3f1b86c7e8c94
+size 381247
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-2.png b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d250db2c94980defb7ab78f9114ca56c1f223024
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:571109a06a09b792694a0ecac329283da199ac4495b1f10ffb7e5925c76a75c9
+size 130973
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-3.png b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
new file mode 100644
index 0000000000000000000000000000000000000000..c414be1c6fa93808d3419715e2e610884e233926
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:3976c705b45d889b06acc4bf82b27f129ad55c4875ee2aa6eb01aecf5a0250f6
+size 144032
diff --git a/OMRChecker/samples/community/Sandeep-1507/template.json b/OMRChecker/samples/community/Sandeep-1507/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bdb929ccd1a61c3ae54df4cdd6a8cf4281369733
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/template.json
@@ -0,0 +1,234 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 15,
+ 15
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Booklet_No": [
+ "b1..7"
+ ]
+ },
+ "fieldBlocks": {
+ "Booklet_No": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 112,
+ 530
+ ],
+ "fieldLabels": [
+ "b1..7"
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 28,
+ "labelsGap": 26.5
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 100
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 370
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q21..35"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 638
+ ]
+ },
+ "MCQBlock2a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 100
+ ]
+ },
+ "MCQBlock2a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 370
+ ]
+ },
+ "MCQBlock2a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q71..85"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 638
+ ]
+ },
+ "MCQBlock3a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q101..110"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 100
+ ]
+ },
+ "MCQBlock3a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q111..120"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 370
+ ]
+ },
+ "MCQBlock3a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q121..135"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 638
+ ]
+ },
+ "MCQBlock4a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q151..160"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 100
+ ]
+ },
+ "MCQBlock4a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q161..170"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 370
+ ]
+ },
+ "MCQBlock4a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q171..185"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 638
+ ]
+ },
+ "MCQBlock1a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q36..50"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 480,
+ 1061
+ ]
+ },
+ "MCQBlock2a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q86..100"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 648,
+ 1061
+ ]
+ },
+ "MCQBlock3a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q136..150"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 1061
+ ]
+ },
+ "MCQBlock4a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q186..200"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.6,
+ "origin": [
+ 986,
+ 1061
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Shamanth/omr_sheet_01.png b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
new file mode 100644
index 0000000000000000000000000000000000000000..7c317db90f80fdd8d3dcfbbadcb1f9b764e619e3
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:9dd72b86ce9c6544f4a628187b4021df5746e9579222e7598c4d88846f2fbeb3
+size 107647
diff --git a/OMRChecker/samples/community/Shamanth/template.json b/OMRChecker/samples/community/Shamanth/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..7d8c34d5e4b7af5ca94415b5096c82300156bcbc
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/template.json
@@ -0,0 +1,25 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 78,
+ 41
+ ],
+ "fieldLabels": [
+ "q21..28"
+ ],
+ "bubblesGap": 56,
+ "labelsGap": 46
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/answer_key.jpg b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..9d39b263e8fe5c23e391ad39bd0905efbf976038
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:13bb31818680ca8497ee4c640a15d4acab2048cf11c973d758c8c75fd71c73be
+size 291737
diff --git a/OMRChecker/samples/community/UPSC-mock/config.json b/OMRChecker/samples/community/UPSC-mock/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..388060a33b04db4b550d0efecafd32aa277355d5
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 1800,
+ "display_width": 2400,
+ "processing_height": 2400,
+ "processing_width": 1800
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/evaluation.json b/OMRChecker/samples/community/UPSC-mock/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..3271cbd5e39161d428d765145f34d811701bdc64
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/evaluation.json
@@ -0,0 +1,18 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "answer_key_image_path": "answer_key.jpg",
+ "questions_in_order": [
+ "q1..100"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "2",
+ "incorrect": "-2/3",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..5dd9ae6adbd327c7995bfcd5f53989002e240cf2
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:020705f4618fd55a72dbc35719565307b844dbd28838c790fe858c3ebe5fb36e
+size 160882
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c696b2e2d39118dec97d4f4c7479bfbc2dc6fc66
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1bb28daa10d276768d4fe3fd3f162ff8d3556cb805c4cf3f362e84d00d808095
+size 182385
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..edb7bbb600db8175e5beaa707c58d5a95972ec1e
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:695e6cc5a497bcf114a101533ee0464a223ecb8f914bc28a8d356533bcf6e341
+size 179112
diff --git a/OMRChecker/samples/community/UPSC-mock/template.json b/OMRChecker/samples/community/UPSC-mock/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..2fadab79ca7f7f9adb03a85cde30f1ba2ccca588
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/template.json
@@ -0,0 +1,195 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 30,
+ 25
+ ],
+ "customLabels": {
+ "Subject Code": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "Roll": [
+ "roll1..10"
+ ]
+ },
+ "fieldBlocks": {
+ "bookletNo": {
+ "origin": [
+ 595,
+ 545
+ ],
+ "bubblesGap": 68,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "bookletNo"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "C",
+ "D"
+ ],
+ "direction": "vertical"
+ },
+ "subjectCode": {
+ "origin": [
+ 912,
+ 512
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.5,
+ "fieldLabels": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "roll": {
+ "origin": [
+ 1200,
+ 510
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.8,
+ "fieldLabels": [
+ "roll1..10"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "q01block": {
+ "origin": [
+ 500,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 500,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 495,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1125,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1125,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..ffd67e704a23baa49bec476e04ba6b1796ff9b52
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
@@ -0,0 +1,200 @@
+q1,C
+q2,C
+q3,"D,E"
+q4,"A,AB"
+q5,"[['A', '1'], ['B', '2']]"
+q6,"['A', 'B']"
+q7,C
+q8,D
+q9,B
+q10,B
+q11,A
+q12,A
+q13,C
+q14,B
+q15,D
+q16,B
+q17,C
+q18,A
+q19,B
+q20,D
+q21,D
+q22,C
+q23,A
+q24,C
+q25,D
+q26,C
+q27,C
+q28,B
+q29,A
+q30,D
+q31,C
+q32,B
+q33,B
+q34,C
+q35,A
+q36,D
+q37,C
+q38,B
+q39,C
+q40,A
+q41,A
+q42,C
+q43,D
+q44,D
+q45,B
+q46,C
+q47,C
+q48,A
+q49,C
+q50,B
+q51,B
+q52,C
+q53,D
+q54,C
+q55,B
+q56,B
+q57,A
+q58,A
+q59,D
+q60,C
+q61,C
+q62,B
+q63,A
+q64,C
+q65,D
+q66,C
+q67,B
+q68,A
+q69,B
+q70,B
+q71,C
+q72,B
+q73,C
+q74,A
+q75,A
+q76,C
+q77,D
+q78,D
+q79,B
+q80,A
+q81,B
+q82,C
+q83,D
+q84,C
+q85,A
+q86,C
+q87,D
+q88,B
+q89,C
+q90,B
+q91,B
+q92,A
+q93,C
+q94,D
+q95,C
+q96,B
+q97,B
+q98,A
+q99,A
+q100,A
+q101,A
+q102,B
+q103,C
+q104,C
+q105,A
+q106,D
+q107,B
+q108,A
+q109,C
+q110,B
+q111,B
+q112,C
+q113,C
+q114,B
+q115,D
+q116,B
+q117,A
+q118,C
+q119,D
+q120,C
+q121,C
+q122,A
+q123,B
+q124,C
+q125,D
+q126,C
+q127,C
+q128,D
+q129,D
+q130,A
+q131,A
+q132,C
+q133,B
+q134,C
+q135,D
+q136,B
+q137,C
+q138,A
+q139,B
+q140,D
+q141,D
+q142,C
+q143,D
+q144,A
+q145,A
+q146,C
+q147,A
+q148,D
+q149,D
+q150,B
+q151,A
+q152,B
+q153,B
+q154,D
+q155,D
+q156,B
+q157,A
+q158,B
+q159,A
+q160,C
+q161,D
+q162,C
+q163,A
+q164,B
+q165,D
+q166,D
+q167,C
+q168,C
+q169,C
+q170,D
+q171,A
+q172,A
+q173,C
+q174,C
+q175,B
+q176,D
+q177,A
+q178,B
+q179,B
+q180,C
+q181,D
+q182,C
+q183,B
+q184,B
+q185,C
+q186,D
+q187,D
+q188,A
+q189,A
+q190,B
+q191,C
+q192,B
+q193,D
+q194,C
+q195,B
+q196,B
+q197,A
+q198,B
+q199,B
+q200,A
diff --git a/OMRChecker/samples/community/UmarFarootAPS/config.json b/OMRChecker/samples/community/UmarFarootAPS/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..5a9d75731b7f5ae8c6fd803a7426b29ac39c6bfb
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 960,
+ "display_width": 1280,
+ "processing_height": 1640,
+ "processing_width": 1332
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/evaluation.json b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg differ
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..73d60b9caa9860aa6a9bb75956fd3430b1aee44e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:49e1a14e0c1be809282b6e43c913e17598380a33f607a5c46723764c32e17b59
+size 304722
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..25d75107cf4885f2974c52cf42003255be285a3e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18b85df487c0bc98eb9139949c3fbca2bad57e8f224fe48b495e0951660f65d
+size 237392
diff --git a/OMRChecker/samples/community/UmarFarootAPS/template.json b/OMRChecker/samples/community/UmarFarootAPS/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..f303edabdf3bc64c41c50152ae72d09f5a0de913
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 2550,
+ 3300
+ ],
+ "bubbleDimensions": [
+ 32,
+ 32
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll_no": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ]
+ },
+ "fieldBlocks": {
+ "Roll_no": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 2169,
+ 180
+ ],
+ "fieldLabels": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ],
+ "bubblesGap": 61,
+ "labelsGap": 93
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q1..17"
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q18..34"
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q35..50"
+ ]
+ },
+ "MCQBlock1a4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q51..67"
+ ]
+ },
+ "MCQBlock1a5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q68..84"
+ ]
+ },
+ "MCQBlock1a6": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q85..100"
+ ]
+ },
+ "MCQBlock1a7": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q101..117"
+ ]
+ },
+ "MCQBlock1a8": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q118..134"
+ ]
+ },
+ "MCQBlock1a9": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q135..150"
+ ]
+ },
+ "MCQBlock1a10": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q151..167"
+ ]
+ },
+ "MCQBlock1a11": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q168..184"
+ ]
+ },
+ "MCQBlock1a12": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q185..200"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..b9d70c6be1aa35398e2ac125f3e7bea0ff924fd0
Binary files /dev/null and b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg differ
diff --git a/OMRChecker/samples/community/dxuian/template.json b/OMRChecker/samples/community/dxuian/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e055f557537fa531a0546c2155146763c8d937a6
--- /dev/null
+++ b/OMRChecker/samples/community/dxuian/template.json
@@ -0,0 +1,48 @@
+{
+ "pageDimensions": [707, 484],
+ "bubbleDimensions": [15, 10],
+ "fieldBlocks": {
+ "Column1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [82, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q1", "Q2", "Q3", "Q4", "Q5", "Q6", "Q7", "Q8", "Q9", "Q10", "Q11", "Q12", "Q13", "Q14", "Q15", "Q16", "Q17", "Q18", "Q19", "Q20"]
+ },
+ "Column2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [205, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q21", "Q22", "Q23", "Q24", "Q25", "Q26", "Q27", "Q28", "Q29", "Q30", "Q31", "Q32", "Q33", "Q34", "Q35", "Q36", "Q37", "Q38", "Q39", "Q40"]
+ },
+ "Column3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [327, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q41", "Q42", "Q43", "Q44", "Q45", "Q46", "Q47", "Q48", "Q49", "Q50", "Q51", "Q52", "Q53", "Q54", "Q55", "Q56", "Q57", "Q58", "Q59", "Q60"]
+ },
+ "Column4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [450, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q61", "Q62", "Q63", "Q64", "Q65", "Q66", "Q67", "Q68", "Q69", "Q70", "Q71", "Q72", "Q73", "Q74", "Q75", "Q76", "Q77", "Q78", "Q79", "Q80"]
+ },
+ "Column5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [573, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q81", "Q82", "Q83", "Q84", "Q85", "Q86", "Q87", "Q88", "Q89", "Q90", "Q91", "Q92", "Q93", "Q94", "Q95", "Q96", "Q97", "Q98", "Q99", "Q100"]
+ }
+ },
+
+ "emptyValue": "-"
+}
\ No newline at end of file
diff --git a/OMRChecker/samples/community/ibrahimkilic/template.json b/OMRChecker/samples/community/ibrahimkilic/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..3064ee19192fc7474370c0f518a51bea9a30b639
--- /dev/null
+++ b/OMRChecker/samples/community/ibrahimkilic/template.json
@@ -0,0 +1,30 @@
+{
+ "pageDimensions": [
+ 299,
+ 328
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "emptyValue": "no",
+ "fieldBlocks": {
+ "YesNoBlock1": {
+ "direction": "horizontal",
+ "bubbleValues": [
+ "yes"
+ ],
+ "origin": [
+ 15,
+ 55
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 48,
+ "labelsGap": 48,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..e9436f41ee8db1269e2255ee902b73de2525d502
Binary files /dev/null and b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg differ
diff --git a/OMRChecker/samples/community/samuelIkoli/template.json b/OMRChecker/samples/community/samuelIkoli/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..19c46f7c96bb07cc0a5a1731bbb8febeba9686e9
--- /dev/null
+++ b/OMRChecker/samples/community/samuelIkoli/template.json
@@ -0,0 +1,28 @@
+{
+ "pageDimensions": [630, 404],
+ "bubbleDimensions": [20, 15],
+ "customLabels": {},
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [33, 6],
+ "fieldLabels": ["q1", "q2", "q3", "q4", "q5", "q6", "q7", "q8", "q9", "q10", "q11", "q12", "q13", "q14", "q15", "q16", "q17", "q18", "q19", "q20"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock2": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [248, 6],
+ "fieldLabels": ["q21", "q22", "q23", "q24", "q25", "q26", "q27", "q28", "q29", "q30", "q31", "q32", "q33", "q34", "q35", "q36", "q37", "q38", "q39", "q40"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock3": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [465, 6],
+ "fieldLabels": ["q41", "q42", "q43", "q44", "q45", "q46", "q47", "q48", "q49", "q50", "q51", "q52", "q53", "q54", "q55", "q56", "q57", "q58", "q59", "q60"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg
new file mode 100644
index 0000000000000000000000000000000000000000..fcd61e808e4a2f33095ea0dbef0ec27741450465
Binary files /dev/null and b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg differ
diff --git a/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6626eb476d38cb16156dc15a48d30bcd41ebaac1
--- /dev/null
+++ b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1be7b0fc432e811fd07b07fe325d9ca8d0d97782eea9f95ed62ff0e47076e1cd
+size 162012
diff --git a/OMRChecker/samples/sample1/config.json b/OMRChecker/samples/sample1/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample1/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample1/omr_marker.jpg b/OMRChecker/samples/sample1/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample1/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample1/template.json b/OMRChecker/samples/sample1/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e4795ad56c919618354f18d4de7a14f14642df07
--- /dev/null
+++ b/OMRChecker/samples/sample1/template.json
@@ -0,0 +1,197 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q5": [
+ "q5_1",
+ "q5_2"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubblesGap": 41,
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "fieldLabels": [
+ "Medium"
+ ],
+ "labelsGap": 0,
+ "origin": [
+ 170,
+ 282
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "roll1..9"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 58,
+ "origin": [
+ 225,
+ 282
+ ]
+ },
+ "Int_Block_Q5": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q5_1",
+ "q5_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 903,
+ 282
+ ]
+ },
+ "Int_Block_Q6": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1077,
+ 282
+ ]
+ },
+ "Int_Block_Q7": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1240,
+ 282
+ ]
+ },
+ "Int_Block_Q8": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1410,
+ 282
+ ]
+ },
+ "Int_Block_Q9": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1580,
+ 282
+ ]
+ },
+ "MCQ_Block_Q1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..4"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 860
+ ]
+ },
+ "MCQ_Block_Q10": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q10..13"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 1195
+ ]
+ },
+ "MCQ_Block_Q14": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q14..16"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 860
+ ]
+ },
+ "MCQ_Block_Q17": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q17..20"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 1195
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ },
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/sample2/config.json b/OMRChecker/samples/sample2/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample2/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample2/template.json b/OMRChecker/samples/sample2/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/sample2/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample3/README.md b/OMRChecker/samples/sample3/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..4b1e325dfa284cf676a1f3d9e37d426f5d20ebb6
--- /dev/null
+++ b/OMRChecker/samples/sample3/README.md
@@ -0,0 +1,31 @@
+## Observation
+The OMR layout is slightly different on colored thick papers vs on xeroxed thin papers. The shift becomes noticible in case of OMR with large number of questions.
+
+We overlapped a colored OMR sheet with a xerox copy of the same OMR sheet(both printed on A4 papers) and observed that there is a great amount of layout sheet as we reach the bottom of the OMR.
+
+Link to an explainer with a real life example: [Google Drive](https://drive.google.com/drive/folders/1GpZTmpEhEjSALJEMjHwDafzWKgEoCTOI?usp=sharing)
+
+## Reasons for shifts in Template layout:
+Listing out a few reasons for the above observation:
+### Printer margin setting
+The margin settings for different printers may be different for the same OMR layout. Thus causing the print to become elongated either horizontally, vertically or both ways.
+
+### The Fan-out effect
+The fan-out effect is usually observed in a sheet fed offset press. Depending on how the papers are made, their dimensions have a tendency to change when they are exposed to moisture or water.
+
+The standard office papers(80 gsm) can easily capture moisture and change shape e.g. in case they get stored for multiple days in a place where the weather is highly humid.
+
+Below are some examples of the GSM ranges:
+
+- 74gsm to 90gsm – This is the basic standard office paper, used in your laser printers.
+- 100gsm to 120gsm – This is stationary paper used for standard letterheads, complimentary slips.
+- 130 to 170gsm – Mostly used for leaflets, posters, single-sided flyers, and brochures.
+
+## Solution
+
+It is recommended to scan each types of prints into different folders and use a separate template.json layout for each of the folders. The same is presented in this sample folder.
+
+## References
+
+- [Paper dimensional stability in sheet-fed offset printing](https://www.diva-portal.org/smash/get/diva2:517895/FULLTEXT01.pdf)
+- An analysis of a few ["Interesting" bubble sheets](https://imgur.com/a/10qwL)
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..4877db1feb1c758bc35b41f543dc9bb14d6ca62f
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18eba01e5d0c466b4b750aa20e710e7354489ed9bef91966c2349007fee349d
+size 137681
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/template.json b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ab35b30f1e3ec35d1c4698cc68b76c4a47d6eb37
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 504,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 504,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 500,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1120,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1120,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample3/config.json b/OMRChecker/samples/sample3/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample3/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..2be38bbfb8e8075f0da00d5fb56d4689d2e72e26
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:eab4df0efac2614b2deddcadc0b754aa72c668170e5d9af49ad77c877198556c
+size 187367
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd499ade7e54d5061817985e40effb13fce8f012
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 492,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 492,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 492,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 487,
+ 1920
+ ],
+ "bubblesGap": 61.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 807,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 803,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 803,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 803,
+ 1920
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1115,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1115,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample4/IMG_20201116_143512.jpg b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6c817d4f620b6bc728eeb410bd9c1da8d27a5cfe
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0b1f0e0afd8a2c40deb7f7c992c9fb6046eb1b28a5f9a674e80ce3418c7fc307
+size 421479
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..023f66bc9eb9100a8500d84b82f7725141962082
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8c3f847e6c37c96562ee17b3d69cb919bb52743f8b95d02679ed62e20dbd5096
+size 386960
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..21869f5c27011e067f1fd8cb3a4e6ecb0e4c7e11
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0bdc5eca48c13d19f228ef7252b5e855b5c62926ba1e9e3297fc6a46e074b1be
+size 307198
diff --git a/OMRChecker/samples/sample4/config.json b/OMRChecker/samples/sample4/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..6c82782a3f6486145d6367eabe6910cca5268214
--- /dev/null
+++ b/OMRChecker/samples/sample4/config.json
@@ -0,0 +1,13 @@
+{
+ "dimensions": {
+ "display_width": 1189,
+ "display_height": 1682
+ },
+ "threshold_params": {
+ "MIN_JUMP": 30
+ },
+ "outputs": {
+ "filter_out_multimarked_files": false,
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample4/evaluation.json b/OMRChecker/samples/sample4/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..8ea935e70ded2a9c53c13bf22b5904f1ef3fa923
--- /dev/null
+++ b/OMRChecker/samples/sample4/evaluation.json
@@ -0,0 +1,34 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..11"
+ ],
+ "answers_in_order": [
+ "B",
+ "D",
+ "C",
+ "B",
+ "D",
+ "C",
+ [
+ "B",
+ "C",
+ "BC"
+ ],
+ "A",
+ "C",
+ "D",
+ "C"
+ ],
+ "should_explain_scoring": true,
+ "enable_evaluation_table_to_csv": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample4/template.json b/OMRChecker/samples/sample4/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..d1bef2985939f8ea9c01bfdec28e0d5b6bab11a3
--- /dev/null
+++ b/OMRChecker/samples/sample4/template.json
@@ -0,0 +1,45 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 30,
+ 30
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ },
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 134,
+ 684
+ ],
+ "fieldLabels": [
+ "q1..11"
+ ],
+ "bubblesGap": 79,
+ "labelsGap": 62
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/README.md b/OMRChecker/samples/sample5/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..b70cedd6cbf22de260649c08e855d929d5e8ae87
--- /dev/null
+++ b/OMRChecker/samples/sample5/README.md
@@ -0,0 +1,6 @@
+### OMRChecker Sample
+
+This sample demonstrates multiple things, namely -
+- Running OMRChecker on images scanned using popular document scanning apps
+- Using a common template.json file for sub-folders (e.g. multiple scan batches)
+- Using evaluation.json file with custom marking (without streak-based marking)
diff --git a/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..72b450e07dd7d4c4a18287183128ad7edbf5607a
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:08bb2e9fecf23be6850485cb7776a604aa2efe9800855ea763dee7ade5362323
+size 127651
diff --git a/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..46fb2ac36c0d5c2366fdf9a7ed56eb933578a60f
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0da09483868ad499925fefa284153e9544822894335e16cea859057740c3c063
+size 127863
diff --git a/OMRChecker/samples/sample5/config.json b/OMRChecker/samples/sample5/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample5/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample5/evaluation.json b/OMRChecker/samples/sample5/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..706e49ee9d9e7d445ca98ea87a1a140ce53da54d
--- /dev/null
+++ b/OMRChecker/samples/sample5/evaluation.json
@@ -0,0 +1,93 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..22"
+ ],
+ "answers_in_order": [
+ "C",
+ "C",
+ "B",
+ "C",
+ "C",
+ [
+ "1",
+ "01"
+ ],
+ "19",
+ "10",
+ "10",
+ "18",
+ "D",
+ "A",
+ "D",
+ "D",
+ "D",
+ "C",
+ "C",
+ "C",
+ "C",
+ "D",
+ "B",
+ "A"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ },
+ "BOOMERANG_1": {
+ "questions": [
+ "q1..5"
+ ],
+ "marking": {
+ "correct": 4,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "PROXIMITY_1": {
+ "questions": [
+ "q6..10"
+ ],
+ "marking": {
+ "correct": 3,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_1": {
+ "questions": [
+ "q11..14"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "POWER_SECTION_1": {
+ "questions": [
+ "q15..18"
+ ],
+ "marking": {
+ "correct": 1,
+ "incorrect": 0,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_2": {
+ "questions": [
+ "q19..22"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/omr_marker.jpg b/OMRChecker/samples/sample5/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample5/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample5/template.json b/OMRChecker/samples/sample5/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ac2436c02a8a19024e8050ff7c02dca9aa7ea016
--- /dev/null
+++ b/OMRChecker/samples/sample5/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ],
+ "q10": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "origin": [
+ 200,
+ 215
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "Medium"
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 261,
+ 210
+ ],
+ "bubblesGap": 46.5,
+ "labelsGap": 58,
+ "fieldLabels": [
+ "roll1..9"
+ ]
+ },
+ "Int1": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 935,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ]
+ },
+ "Int2": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1100,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ]
+ },
+ "Int3": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1275,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ]
+ },
+ "Int4": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1449,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "Int5": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1620,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "Mcq1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 198,
+ 826
+ ],
+ "bubblesGap": 93,
+ "labelsGap": 62,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ },
+ "Mcq2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 830
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q11..14"
+ ]
+ },
+ "Mcq3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 1270
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q15..18"
+ ]
+ },
+ "Mcq4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1481,
+ 830
+ ],
+ "bubblesGap": 73,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q19..22"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/config.json b/OMRChecker/samples/sample6/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..2ac30e718a44f7336aafda1b0f57aec1320fc72c
--- /dev/null
+++ b/OMRChecker/samples/sample6/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_width": 2480,
+ "display_height": 3508,
+ "processing_width": 1653,
+ "processing_height": 2339
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..fdaa0f4112e15037a31bc90da204b7c19114820b
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:de8a5ed5b7360a71db83b49a3d0d6e1af1482339d1116286a064d285be42e37e
+size 246590
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..bf56d121c1061302fc4ce87fca61bb8584ea9e4a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:aca51de61ba3ee76e00e22a666d8cd2b72190af674fa051c9cf70346e9ee4d0a
+size 244049
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..22bfa55de7a687729b733a615ad78dad64ac467a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:950dbf86fc152f464be30ebaeb518c61067b0c494ab5fd70f05de10b5dfabb12
+size 244321
diff --git a/OMRChecker/samples/sample6/readme.md b/OMRChecker/samples/sample6/readme.md
new file mode 100644
index 0000000000000000000000000000000000000000..d8491b2a049eb4e721e007e8e8bc4fe5ca536802
--- /dev/null
+++ b/OMRChecker/samples/sample6/readme.md
@@ -0,0 +1,21 @@
+# Demo for feature-based alignment
+
+## Background
+OMR is used to match student roll on final exam scripts. Scripts are scanned using a document scanner and the cover pages are extracted for OMR. Even though a document scanner does not produce any warpped perspective, the alignment is not perfect, causing some rotation and translation in the scans.
+
+The scripts in this sample were specifically selected incorrectly marked scripts to demonstrate how feature-based alignment can correct transformation errors using a reference image. In the actual batch. 156 out of 532 scripts were incorrectly marked. With feature-based alignment, all scripts were correctly marked.
+
+## Usage
+Two template files are given in the sample folder, one with feature-based alignment (template_fb_align), the other without (template_no_fb_align).
+
+## Additional Notes
+
+### Reference Image
+When using a reference image for feature-based alignment, it is better not to have many repeated patterns as it is causes ambiguity when trying to match similar feature points. The bubbles in an OMR form are identical and should not be used for feature-extraction.
+
+Thus, the reference image should be cleared of any bubbles. Forms with lots of text as in this example would be effective.
+
+Note the reference image in this sample was generated from a vector pdf, and not from a scanned blank, producing in a perfectly aligned reference.
+
+### Level adjustment
+The bubbles on the scripts were not shaded dark enough. Thus, a level adjustment was done to bring the black point to 70% to darken the light shading. White point was brought down to 80% to remove the light-grey background in the columns.
diff --git a/OMRChecker/samples/sample6/reference.png b/OMRChecker/samples/sample6/reference.png
new file mode 100644
index 0000000000000000000000000000000000000000..e2e692f4638e9c7fa2d7d159e044c778b18acdc9
--- /dev/null
+++ b/OMRChecker/samples/sample6/reference.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8379ba055209c065def041fc1fb1cea444b913d835c1cfeffa40091446e5e3e5
+size 178069
diff --git a/OMRChecker/samples/sample6/template.json b/OMRChecker/samples/sample6/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_fb_align.json b/OMRChecker/samples/sample6/template_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd45b9c8fd3ba024b951ff4e7965c98bb56211ee
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_fb_align.json
@@ -0,0 +1,118 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "FeatureBasedAlignment",
+ "options": {
+ "reference": "reference.png",
+ "maxFeatures": 1000,
+ "2d": true
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_no_fb_align.json b/OMRChecker/samples/sample6/template_no_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_no_fb_align.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/src/__init__.py b/OMRChecker/src/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..5c1baacaa53bb1b4219d33eee566c6a613ab88d6
--- /dev/null
+++ b/OMRChecker/src/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+from src.logger import logger
+
+# It takes a few seconds for the imports
+logger.info(f"Loading OMRChecker modules...")
diff --git a/OMRChecker/src/constants.py b/OMRChecker/src/constants.py
new file mode 100644
index 0000000000000000000000000000000000000000..628eb75ac892ddc184cc99429036aa9196889c83
--- /dev/null
+++ b/OMRChecker/src/constants.py
@@ -0,0 +1,54 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+from dotmap import DotMap
+
+# Filenames
+TEMPLATE_FILENAME = "template.json"
+EVALUATION_FILENAME = "evaluation.json"
+CONFIG_FILENAME = "config.json"
+
+FIELD_LABEL_NUMBER_REGEX = r"([^\d]+)(\d*)"
+#
+ERROR_CODES = DotMap(
+ {
+ "MULTI_BUBBLE_WARN": 1,
+ "NO_MARKER_ERR": 2,
+ },
+ _dynamic=False,
+)
+
+FIELD_TYPES = {
+ "QTYPE_INT": {
+ "bubbleValues": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"],
+ "direction": "vertical",
+ },
+ "QTYPE_INT_FROM_1": {
+ "bubbleValues": ["1", "2", "3", "4", "5", "6", "7", "8", "9", "0"],
+ "direction": "vertical",
+ },
+ "QTYPE_MCQ4": {"bubbleValues": ["A", "B", "C", "D"], "direction": "horizontal"},
+ "QTYPE_MCQ5": {
+ "bubbleValues": ["A", "B", "C", "D", "E"],
+ "direction": "horizontal",
+ },
+ #
+ # You can create and append custom field types here-
+ #
+}
+
+# TODO: move to interaction.py
+TEXT_SIZE = 0.95
+CLR_BLACK = (50, 150, 150)
+CLR_WHITE = (250, 250, 250)
+CLR_GRAY = (130, 130, 130)
+CLR_DARK_GRAY = (100, 100, 100)
+
+# TODO: move to config.json
+GLOBAL_PAGE_THRESHOLD_WHITE = 200
+GLOBAL_PAGE_THRESHOLD_BLACK = 100
diff --git a/OMRChecker/src/core.py b/OMRChecker/src/core.py
new file mode 100644
index 0000000000000000000000000000000000000000..15279cf49fb9fb12c88611bcc69611ea524344dc
--- /dev/null
+++ b/OMRChecker/src/core.py
@@ -0,0 +1,721 @@
+import os
+from collections import defaultdict
+from typing import Any
+
+import cv2
+import matplotlib.pyplot as plt
+import numpy as np
+
+import src.constants as constants
+from src.logger import logger
+from src.utils.image import CLAHE_HELPER, ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class ImageInstanceOps:
+ """Class to hold fine-tuned utilities for a group of images. One instance for each processing directory."""
+
+ save_img_list: Any = defaultdict(list)
+
+ def __init__(self, tuning_config):
+ super().__init__()
+ self.tuning_config = tuning_config
+ self.save_image_level = tuning_config.outputs.save_image_level
+
+ def apply_preprocessors(self, file_path, in_omr, template):
+ tuning_config = self.tuning_config
+ # resize to conform to template
+ in_omr = ImageUtils.resize_util(
+ in_omr,
+ tuning_config.dimensions.processing_width,
+ tuning_config.dimensions.processing_height,
+ )
+
+ # run pre_processors in sequence
+ for pre_processor in template.pre_processors:
+ in_omr = pre_processor.apply_filter(in_omr, file_path)
+ return in_omr
+
+ def read_omr_response(self, template, image, name, save_dir=None):
+ config = self.tuning_config
+ auto_align = config.alignment_params.auto_align
+ try:
+ img = image.copy()
+ # origDim = img.shape[:2]
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ if img.max() > img.min():
+ img = ImageUtils.normalize_util(img)
+ # Processing copies
+ transp_layer = img.copy()
+ final_marked = img.copy()
+
+ morph = img.copy()
+ self.append_save_img(3, morph)
+
+ if auto_align:
+ # Note: clahe is good for morphology, bad for thresholding
+ morph = CLAHE_HELPER.apply(morph)
+ self.append_save_img(3, morph)
+ # Remove shadows further, make columns/boxes darker (less gamma)
+ morph = ImageUtils.adjust_gamma(
+ morph, config.threshold_params.GAMMA_LOW
+ )
+ # TODO: all numbers should come from either constants or config
+ _, morph = cv2.threshold(morph, 220, 220, cv2.THRESH_TRUNC)
+ morph = ImageUtils.normalize_util(morph)
+ self.append_save_img(3, morph)
+ if config.outputs.show_image_level >= 4:
+ InteractionUtils.show("morph1", morph, 0, 1, config)
+
+ # Move them to data class if needed
+ # Overlay Transparencies
+ alpha = 0.65
+ omr_response = {}
+ multi_marked, multi_roll = 0, 0
+
+ # TODO Make this part useful for visualizing status checks
+ # blackVals=[0]
+ # whiteVals=[255]
+
+ if config.outputs.show_image_level >= 5:
+ all_c_box_vals = {"int": [], "mcq": []}
+ # TODO: simplify this logic
+ q_nums = {"int": [], "mcq": []}
+
+ # Find Shifts for the field_blocks --> Before calculating threshold!
+ if auto_align:
+ # print("Begin Alignment")
+ # Open : erode then dilate
+ v_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (2, 10))
+ morph_v = cv2.morphologyEx(
+ morph, cv2.MORPH_OPEN, v_kernel, iterations=3
+ )
+ _, morph_v = cv2.threshold(morph_v, 200, 200, cv2.THRESH_TRUNC)
+ morph_v = 255 - ImageUtils.normalize_util(morph_v)
+
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morphed_vertical", morph_v, 0, 1, config=config
+ )
+
+ # InteractionUtils.show("morph1",morph,0,1,config=config)
+ # InteractionUtils.show("morphed_vertical",morph_v,0,1,config=config)
+
+ self.append_save_img(3, morph_v)
+
+ morph_thr = 60 # for Mobile images, 40 for scanned Images
+ _, morph_v = cv2.threshold(morph_v, morph_thr, 255, cv2.THRESH_BINARY)
+ # kernel best tuned to 5x5 now
+ morph_v = cv2.erode(morph_v, np.ones((5, 5), np.uint8), iterations=2)
+
+ self.append_save_img(3, morph_v)
+ # h_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (10, 2))
+ # morph_h = cv2.morphologyEx(morph, cv2.MORPH_OPEN, h_kernel, iterations=3)
+ # ret, morph_h = cv2.threshold(morph_h,200,200,cv2.THRESH_TRUNC)
+ # morph_h = 255 - normalize_util(morph_h)
+ # InteractionUtils.show("morph_h",morph_h,0,1,config=config)
+ # _, morph_h = cv2.threshold(morph_h,morph_thr,255,cv2.THRESH_BINARY)
+ # morph_h = cv2.erode(morph_h, np.ones((5,5),np.uint8), iterations = 2)
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morph_thr_eroded", morph_v, 0, 1, config=config
+ )
+
+ self.append_save_img(6, morph_v)
+
+ # template relative alignment code
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+
+ match_col, max_steps, align_stride, thk = map(
+ config.alignment_params.get,
+ [
+ "match_col",
+ "max_steps",
+ "stride",
+ "thickness",
+ ],
+ )
+ shift, steps = 0, 0
+ while steps < max_steps:
+ left_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0] + shift - thk : -thk + s[0] + shift + match_col,
+ ]
+ )
+ right_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0]
+ + shift
+ - match_col
+ + d[0]
+ + thk : thk
+ + s[0]
+ + shift
+ + d[0],
+ ]
+ )
+
+ # For demonstration purposes-
+ # if(field_block.name == "int1"):
+ # ret = morph_v.copy()
+ # cv2.rectangle(ret,
+ # (s[0]+shift-thk,s[1]),
+ # (s[0]+shift+thk+d[0],s[1]+d[1]),
+ # constants.CLR_WHITE,
+ # 3)
+ # appendSaveImg(6,ret)
+ # print(shift, left_mean, right_mean)
+ left_shift, right_shift = left_mean > 100, right_mean > 100
+ if left_shift:
+ if right_shift:
+ break
+ else:
+ shift -= align_stride
+ else:
+ if right_shift:
+ shift += align_stride
+ else:
+ break
+ steps += 1
+
+ field_block.shift = shift
+ # print("Aligned field_block: ",field_block.name,"Corrected Shift:",
+ # field_block.shift,", dimensions:", field_block.dimensions,
+ # "origin:", field_block.origin,'\n')
+ # print("End Alignment")
+
+ final_align = None
+ if config.outputs.show_image_level >= 2:
+ initial_align = self.draw_template_layout(img, template, shifted=False)
+ final_align = self.draw_template_layout(
+ img, template, shifted=True, draw_qvals=True
+ )
+ # appendSaveImg(4,mean_vals)
+ self.append_save_img(2, initial_align)
+ self.append_save_img(2, final_align)
+
+ if auto_align:
+ final_align = np.hstack((initial_align, final_align))
+ self.append_save_img(5, img)
+
+ # Get mean bubbleValues n other stats
+ all_q_vals, all_q_strip_arrs, all_q_std_vals = [], [], []
+ total_q_strip_no = 0
+ for field_block in template.field_blocks:
+ box_w, box_h = field_block.bubble_dimensions
+ q_std_vals = []
+ for field_block_bubbles in field_block.traverse_bubbles:
+ q_strip_vals = []
+ for pt in field_block_bubbles:
+ # shifted
+ x, y = (pt.x + field_block.shift, pt.y)
+ rect = [y, y + box_h, x, x + box_w]
+ q_strip_vals.append(
+ cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0]
+ # detectCross(img, rect) ? 100 : 0
+ )
+ q_std_vals.append(round(np.std(q_strip_vals), 2))
+ all_q_strip_arrs.append(q_strip_vals)
+ # _, _, _ = get_global_threshold(q_strip_vals, "QStrip Plot",
+ # plot_show=False, sort_in_plot=True)
+ # hist = getPlotImg()
+ # InteractionUtils.show("QStrip "+field_block_bubbles[0].field_label, hist, 0, 1,config=config)
+ all_q_vals.extend(q_strip_vals)
+ # print(total_q_strip_no, field_block_bubbles[0].field_label, q_std_vals[len(q_std_vals)-1])
+ total_q_strip_no += 1
+ all_q_std_vals.extend(q_std_vals)
+
+ global_std_thresh, _, _ = self.get_global_threshold(
+ all_q_std_vals
+ ) # , "Q-wise Std-dev Plot", plot_show=True, sort_in_plot=True)
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # Note: Plotting takes Significant times here --> Change Plotting args
+ # to support show_image_level
+ # , "Mean Intensity Histogram",plot_show=True, sort_in_plot=True)
+ global_thr, _, _ = self.get_global_threshold(all_q_vals, looseness=4)
+
+ logger.info(
+ f"Thresholding: \tglobal_thr: {round(global_thr, 2)} \tglobal_std_THR: {round(global_std_thresh, 2)}\t{'(Looks like a Xeroxed OMR)' if (global_thr == 255) else ''}"
+ )
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # if(config.outputs.show_image_level>=1):
+ # hist = getPlotImg()
+ # InteractionUtils.show("Hist", hist, 0, 1,config=config)
+ # appendSaveImg(4,hist)
+ # appendSaveImg(5,hist)
+ # appendSaveImg(2,hist)
+
+ per_omr_threshold_avg, total_q_strip_no, total_q_box_no = 0, 0, 0
+ for field_block in template.field_blocks:
+ block_q_strip_no = 1
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ s, d = field_block.origin, field_block.dimensions
+ key = field_block.name[:3]
+ # cv2.rectangle(final_marked,(s[0]+shift,s[1]),(s[0]+shift+d[0],
+ # s[1]+d[1]),CLR_BLACK,3)
+ for field_block_bubbles in field_block.traverse_bubbles:
+ # All Black or All White case
+ no_outliers = all_q_std_vals[total_q_strip_no] < global_std_thresh
+ # print(total_q_strip_no, field_block_bubbles[0].field_label,
+ # all_q_std_vals[total_q_strip_no], "no_outliers:", no_outliers)
+ per_q_strip_threshold = self.get_local_threshold(
+ all_q_strip_arrs[total_q_strip_no],
+ global_thr,
+ no_outliers,
+ f"Mean Intensity Histogram for {key}.{field_block_bubbles[0].field_label}.{block_q_strip_no}",
+ config.outputs.show_image_level >= 6,
+ )
+ # print(field_block_bubbles[0].field_label,key,block_q_strip_no, "THR: ",
+ # round(per_q_strip_threshold,2))
+ per_omr_threshold_avg += per_q_strip_threshold
+
+ # Note: Little debugging visualization - view the particular Qstrip
+ # if(
+ # 0
+ # # or "q17" in (field_block_bubbles[0].field_label)
+ # # or (field_block_bubbles[0].field_label+str(block_q_strip_no))=="q15"
+ # ):
+ # st, end = qStrip
+ # InteractionUtils.show("QStrip: "+key+"-"+str(block_q_strip_no),
+ # img[st[1] : end[1], st[0]+shift : end[0]+shift],0,config=config)
+
+ # TODO: get rid of total_q_box_no
+ detected_bubbles = []
+ for bubble in field_block_bubbles:
+ bubble_is_marked = (
+ per_q_strip_threshold > all_q_vals[total_q_box_no]
+ )
+ total_q_box_no += 1
+ if bubble_is_marked:
+ detected_bubbles.append(bubble)
+ x, y, field_value = (
+ bubble.x + field_block.shift,
+ bubble.y,
+ bubble.field_value,
+ )
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 12), int(y + box_h / 12)),
+ (
+ int(x + box_w - box_w / 12),
+ int(y + box_h - box_h / 12),
+ ),
+ constants.CLR_DARK_GRAY,
+ 3,
+ )
+
+ cv2.putText(
+ final_marked,
+ str(field_value),
+ (x, y),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ (20, 20, 10),
+ int(1 + 3.5 * constants.TEXT_SIZE),
+ )
+ else:
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (
+ int(x + box_w - box_w / 10),
+ int(y + box_h - box_h / 10),
+ ),
+ constants.CLR_GRAY,
+ -1,
+ )
+
+ for bubble in detected_bubbles:
+ field_label, field_value = (
+ bubble.field_label,
+ bubble.field_value,
+ )
+ # Only send rolls multi-marked in the directory
+ multi_marked_local = field_label in omr_response
+ omr_response[field_label] = (
+ (omr_response[field_label] + field_value)
+ if multi_marked_local
+ else field_value
+ )
+ # TODO: generalize this into identifier
+ # multi_roll = multi_marked_local and "Roll" in str(q)
+ multi_marked = multi_marked or multi_marked_local
+
+ if len(detected_bubbles) == 0:
+ field_label = field_block_bubbles[0].field_label
+ omr_response[field_label] = field_block.empty_val
+
+ if config.outputs.show_image_level >= 5:
+ if key in all_c_box_vals:
+ q_nums[key].append(f"{key[:2]}_c{str(block_q_strip_no)}")
+ all_c_box_vals[key].append(
+ all_q_strip_arrs[total_q_strip_no]
+ )
+
+ block_q_strip_no += 1
+ total_q_strip_no += 1
+ # /for field_block
+
+ per_omr_threshold_avg /= total_q_strip_no
+ per_omr_threshold_avg = round(per_omr_threshold_avg, 2)
+ # Translucent
+ cv2.addWeighted(
+ final_marked, alpha, transp_layer, 1 - alpha, 0, final_marked
+ )
+ # Box types
+ if config.outputs.show_image_level >= 6:
+ # plt.draw()
+ f, axes = plt.subplots(len(all_c_box_vals), sharey=True)
+ f.canvas.manager.set_window_title(name)
+ ctr = 0
+ type_name = {
+ "int": "Integer",
+ "mcq": "MCQ",
+ "med": "MED",
+ "rol": "Roll",
+ }
+ for k, boxvals in all_c_box_vals.items():
+ axes[ctr].title.set_text(type_name[k] + " Type")
+ axes[ctr].boxplot(boxvals)
+ # thrline=axes[ctr].axhline(per_omr_threshold_avg,color='red',ls='--')
+ # thrline.set_label("Average THR")
+ axes[ctr].set_ylabel("Intensity")
+ axes[ctr].set_xticklabels(q_nums[k])
+ # axes[ctr].legend()
+ ctr += 1
+ # imshow will do the waiting
+ plt.tight_layout(pad=0.5)
+ plt.show()
+
+ if config.outputs.show_image_level >= 3 and final_align is not None:
+ final_align = ImageUtils.resize_util_h(
+ final_align, int(config.dimensions.display_height)
+ )
+ # [final_align.shape[1],0])
+ InteractionUtils.show(
+ "Template Alignment Adjustment", final_align, 0, 0, config=config
+ )
+
+ if config.outputs.save_detections and save_dir is not None:
+ if multi_roll:
+ save_dir = save_dir.joinpath("_MULTI_")
+ image_path = str(save_dir.joinpath(name))
+ ImageUtils.save_img(image_path, final_marked)
+
+ self.append_save_img(2, final_marked)
+
+ if save_dir is not None:
+ for i in range(config.outputs.save_image_level):
+ self.save_image_stacks(i + 1, name, save_dir)
+
+ return omr_response, final_marked, multi_marked, multi_roll
+
+ except Exception as e:
+ raise e
+
+ @staticmethod
+ def draw_template_layout(img, template, shifted=True, draw_qvals=False, border=-1):
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ final_align = img.copy()
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ if shifted:
+ cv2.rectangle(
+ final_align,
+ (s[0] + shift, s[1]),
+ (s[0] + shift + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ else:
+ cv2.rectangle(
+ final_align,
+ (s[0], s[1]),
+ (s[0] + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ for field_block_bubbles in field_block.traverse_bubbles:
+ for pt in field_block_bubbles:
+ x, y = (pt.x + field_block.shift, pt.y) if shifted else (pt.x, pt.y)
+ cv2.rectangle(
+ final_align,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (int(x + box_w - box_w / 10), int(y + box_h - box_h / 10)),
+ constants.CLR_GRAY,
+ border,
+ )
+ if draw_qvals:
+ rect = [y, y + box_h, x, x + box_w]
+ cv2.putText(
+ final_align,
+ f"{int(cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0])}",
+ (rect[2] + 2, rect[0] + (box_h * 2) // 3),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ 0.6,
+ constants.CLR_BLACK,
+ 2,
+ )
+ if shifted:
+ text_in_px = cv2.getTextSize(
+ field_block.name, cv2.FONT_HERSHEY_SIMPLEX, constants.TEXT_SIZE, 4
+ )
+ cv2.putText(
+ final_align,
+ field_block.name,
+ (int(s[0] + d[0] - text_in_px[0][0]), int(s[1] - text_in_px[0][1])),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ constants.CLR_BLACK,
+ 4,
+ )
+ return final_align
+
+ def get_global_threshold(
+ self,
+ q_vals_orig,
+ plot_title=None,
+ plot_show=True,
+ sort_in_plot=True,
+ looseness=1,
+ ):
+ """
+ Note: Cannot assume qStrip has only-gray or only-white bg
+ (in which case there is only one jump).
+ So there will be either 1 or 2 jumps.
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ 2 Jumps :
+ ......
+ |||||| <-- wrong THR
+ ....||||||
+ |||||||||| <-- safe THR
+ ..||||||||||
+ ||||||||||||
+
+ The abstract "First LARGE GAP" is perfect for this.
+ Current code is considering ONLY TOP 2 jumps(>= MIN_GAP) to be big,
+ gives the smaller one
+
+ """
+ config = self.tuning_config
+ PAGE_TYPE_FOR_THRESHOLD, MIN_JUMP, JUMP_DELTA = map(
+ config.threshold_params.get,
+ [
+ "PAGE_TYPE_FOR_THRESHOLD",
+ "MIN_JUMP",
+ "JUMP_DELTA",
+ ],
+ )
+
+ global_default_threshold = (
+ constants.GLOBAL_PAGE_THRESHOLD_WHITE
+ if PAGE_TYPE_FOR_THRESHOLD == "white"
+ else constants.GLOBAL_PAGE_THRESHOLD_BLACK
+ )
+
+ # Sort the Q bubbleValues
+ # TODO: Change var name of q_vals
+ q_vals = sorted(q_vals_orig)
+ # Find the FIRST LARGE GAP and set it as threshold:
+ ls = (looseness + 1) // 2
+ l = len(q_vals) - ls
+ max1, thr1 = MIN_JUMP, global_default_threshold
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - ls] + jump / 2
+
+ # NOTE: thr2 is deprecated, thus is JUMP_DELTA
+ # Make use of the fact that the JUMP_DELTA(Vertical gap ofc) between
+ # values at detected jumps would be atleast 20
+ max2, thr2 = MIN_JUMP, global_default_threshold
+ # Requires atleast 1 gray box to be present (Roll field will ensure this)
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ new_thr = q_vals[i - ls] + jump / 2
+ if jump > max2 and abs(thr1 - new_thr) > JUMP_DELTA:
+ max2 = jump
+ thr2 = new_thr
+ # global_thr = min(thr1,thr2)
+ global_thr, j_low, j_high = thr1, thr1 - max1 // 2, thr1 + max1 // 2
+
+ # # For normal images
+ # thresholdRead = 116
+ # if(thr1 > thr2 and thr2 > thresholdRead):
+ # print("Note: taking safer thr line.")
+ # global_thr, j_low, j_high = thr2, thr2 - max2//2, thr2 + max2//2
+
+ if plot_title:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals_orig)), q_vals if sort_in_plot else q_vals_orig)
+ ax.set_title(plot_title)
+ thrline = ax.axhline(global_thr, color="green", ls="--", linewidth=5)
+ thrline.set_label("Global Threshold")
+ thrline = ax.axhline(thr2, color="red", ls=":", linewidth=3)
+ thrline.set_label("THR2 Line")
+ # thrline=ax.axhline(j_low,color='red',ls='-.', linewidth=3)
+ # thrline=ax.axhline(j_high,color='red',ls='-.', linewidth=3)
+ # thrline.set_label("Boundary Line")
+ # ax.set_ylabel("Mean Intensity")
+ ax.set_ylabel("Values")
+ ax.set_xlabel("Position")
+ ax.legend()
+ if plot_show:
+ plt.title(plot_title)
+ plt.show()
+
+ return global_thr, j_low, j_high
+
+ def get_local_threshold(
+ self, q_vals, global_thr, no_outliers, plot_title=None, plot_show=True
+ ):
+ """
+ TODO: Update this documentation too-
+ //No more - Assumption : Colwise background color is uniformly gray or white,
+ but not alternating. In this case there is atmost one jump.
+
+ 0 Jump :
+ <-- safe THR?
+ .......
+ ...|||||||
+ |||||||||| <-- safe THR?
+ // How to decide given range is above or below gray?
+ -> global q_vals shall absolutely help here. Just run same function
+ on total q_vals instead of colwise _//
+ How to decide it is this case of 0 jumps
+
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ """
+ config = self.tuning_config
+ # Sort the Q bubbleValues
+ q_vals = sorted(q_vals)
+
+ # Small no of pts cases:
+ # base case: 1 or 2 pts
+ if len(q_vals) < 3:
+ thr1 = (
+ global_thr
+ if np.max(q_vals) - np.min(q_vals) < config.threshold_params.MIN_GAP
+ else np.mean(q_vals)
+ )
+ else:
+ # qmin, qmax, qmean, qstd = round(np.min(q_vals),2), round(np.max(q_vals),2),
+ # round(np.mean(q_vals),2), round(np.std(q_vals),2)
+ # GVals = [round(abs(q-qmean),2) for q in q_vals]
+ # gmean, gstd = round(np.mean(GVals),2), round(np.std(GVals),2)
+ # # DISCRETION: Pretty critical factor in reading response
+ # # Doesn't work well for small number of values.
+ # DISCRETION = 2.7 # 2.59 was closest hit, 3.0 is too far
+ # L2MaxGap = round(max([abs(g-gmean) for g in GVals]),2)
+ # if(L2MaxGap > DISCRETION*gstd):
+ # no_outliers = False
+
+ # # ^Stackoverflow method
+ # print(field_label, no_outliers,"qstd",round(np.std(q_vals),2), "gstd", gstd,
+ # "Gaps in gvals",sorted([round(abs(g-gmean),2) for g in GVals],reverse=True),
+ # '\t',round(DISCRETION*gstd,2), L2MaxGap)
+
+ # else:
+ # Find the LARGEST GAP and set it as threshold: //(FIRST LARGE GAP)
+ l = len(q_vals) - 1
+ max1, thr1 = config.threshold_params.MIN_JUMP, 255
+ for i in range(1, l):
+ jump = q_vals[i + 1] - q_vals[i - 1]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - 1] + jump / 2
+ # print(field_label,q_vals,max1)
+
+ confident_jump = (
+ config.threshold_params.MIN_JUMP
+ + config.threshold_params.CONFIDENT_SURPLUS
+ )
+ # If not confident, then only take help of global_thr
+ if max1 < confident_jump:
+ if no_outliers:
+ # All Black or All White case
+ thr1 = global_thr
+ else:
+ # TODO: Low confidence parameters here
+ pass
+
+ # if(thr1 == 255):
+ # print("Warning: threshold is unexpectedly 255! (Outlier Delta issue?)",plot_title)
+
+ # Make a common plot function to show local and global thresholds
+ if plot_show and plot_title is not None:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals)), q_vals)
+ thrline = ax.axhline(thr1, color="green", ls=("-."), linewidth=3)
+ thrline.set_label("Local Threshold")
+ thrline = ax.axhline(global_thr, color="red", ls=":", linewidth=5)
+ thrline.set_label("Global Threshold")
+ ax.set_title(plot_title)
+ ax.set_ylabel("Bubble Mean Intensity")
+ ax.set_xlabel("Bubble Number(sorted)")
+ ax.legend()
+ # TODO append QStrip to this plot-
+ # appendSaveImg(6,getPlotImg())
+ if plot_show:
+ plt.show()
+ return thr1
+
+ def append_save_img(self, key, img):
+ if self.save_image_level >= int(key):
+ self.save_img_list[key].append(img.copy())
+
+ def save_image_stacks(self, key, filename, save_dir):
+ config = self.tuning_config
+ if self.save_image_level >= int(key) and self.save_img_list[key] != []:
+ name = os.path.splitext(filename)[0]
+ result = np.hstack(
+ tuple(
+ [
+ ImageUtils.resize_util_h(img, config.dimensions.display_height)
+ for img in self.save_img_list[key]
+ ]
+ )
+ )
+ result = ImageUtils.resize_util(
+ result,
+ min(
+ len(self.save_img_list[key]) * config.dimensions.display_width // 3,
+ int(config.dimensions.display_width * 2.5),
+ ),
+ )
+ ImageUtils.save_img(f"{save_dir}stack/{name}_{str(key)}_stack.jpg", result)
+
+ def reset_all_save_img(self):
+ for i in range(self.save_image_level):
+ self.save_img_list[i + 1] = []
diff --git a/OMRChecker/src/defaults/__init__.py b/OMRChecker/src/defaults/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..db3d42e87b3e47d0cbee08032e65e87ac1bd8165
--- /dev/null
+++ b/OMRChecker/src/defaults/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+# Use all imports relative to root directory
+# (https://chrisyeh96.github.io/2017/08/08/definitive-guide-python-imports.html)
+from src.defaults.config import * # NOQA
+from src.defaults.template import * # NOQA
diff --git a/OMRChecker/src/defaults/config.py b/OMRChecker/src/defaults/config.py
new file mode 100644
index 0000000000000000000000000000000000000000..6d1315fb45b0e736ad7daedcea5f3b23e3a9078d
--- /dev/null
+++ b/OMRChecker/src/defaults/config.py
@@ -0,0 +1,35 @@
+from dotmap import DotMap
+
+CONFIG_DEFAULTS = DotMap(
+ {
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666,
+ },
+ "threshold_params": {
+ "GAMMA_LOW": 0.7,
+ "MIN_GAP": 30,
+ "MIN_JUMP": 25,
+ "CONFIDENT_SURPLUS": 5,
+ "JUMP_DELTA": 30,
+ "PAGE_TYPE_FOR_THRESHOLD": "white",
+ },
+ "alignment_params": {
+ # Note: 'auto_align' enables automatic template alignment, use if the scans show slight misalignments.
+ "auto_align": False,
+ "match_col": 5,
+ "max_steps": 20,
+ "stride": 1,
+ "thickness": 3,
+ },
+ "outputs": {
+ "show_image_level": 0,
+ "save_image_level": 0,
+ "save_detections": True,
+ "filter_out_multimarked_files": False,
+ },
+ },
+ _dynamic=False,
+)
diff --git a/OMRChecker/src/defaults/template.py b/OMRChecker/src/defaults/template.py
new file mode 100644
index 0000000000000000000000000000000000000000..efd75420f01cf49f72a2e285734c0d79cafe4837
--- /dev/null
+++ b/OMRChecker/src/defaults/template.py
@@ -0,0 +1,6 @@
+TEMPLATE_DEFAULTS = {
+ "preProcessors": [],
+ "emptyValue": "",
+ "customLabels": {},
+ "outputColumns": [],
+}
diff --git a/OMRChecker/src/entry.py b/OMRChecker/src/entry.py
new file mode 100644
index 0000000000000000000000000000000000000000..94d3137c3ac99502399cbb40d6d378cd8249d8de
--- /dev/null
+++ b/OMRChecker/src/entry.py
@@ -0,0 +1,371 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import os
+from csv import QUOTE_NONNUMERIC
+from pathlib import Path
+from time import time
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src import constants
+from src.defaults import CONFIG_DEFAULTS
+from src.evaluation import EvaluationConfig, evaluate_concatenated_response
+from src.logger import console, logger
+from src.template import Template
+from src.utils.file import Paths, setup_dirs_for_paths, setup_outputs_for_template
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils, Stats
+from src.utils.parsing import get_concatenated_response, open_config_with_defaults
+
+# Load processors
+STATS = Stats()
+
+
+def entry_point(input_dir, args):
+ if not os.path.exists(input_dir):
+ raise Exception(f"Given input directory does not exist: '{input_dir}'")
+ curr_dir = input_dir
+ return process_dir(input_dir, curr_dir, args)
+
+
+def print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+):
+ logger.info("")
+ table = Table(title="Current Configurations", show_header=False, show_lines=False)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Value", style="magenta")
+ table.add_row("Directory Path", f"{curr_dir}")
+ table.add_row("Count of Images", f"{len(omr_files)}")
+ table.add_row("Set Layout Mode ", "ON" if args["setLayout"] else "OFF")
+ pre_processor_names = [pp.__class__.__name__ for pp in template.pre_processors]
+ table.add_row(
+ "Markers Detection",
+ "ON" if "CropOnMarkers" in pre_processor_names else "OFF",
+ )
+ table.add_row("Auto Alignment", f"{tuning_config.alignment_params.auto_align}")
+ table.add_row("Detected Template Path", f"{template}")
+ if local_config_path:
+ table.add_row("Detected Local Config", f"{local_config_path}")
+ if evaluation_config:
+ table.add_row("Detected Evaluation Config", f"{evaluation_config}")
+
+ table.add_row(
+ "Detected pre-processors",
+ ", ".join(pre_processor_names),
+ )
+ console.print(table, justify="center")
+
+
+def process_dir(
+ root_dir,
+ curr_dir,
+ args,
+ template=None,
+ tuning_config=CONFIG_DEFAULTS,
+ evaluation_config=None,
+):
+ # Update local tuning_config (in current recursion stack)
+ local_config_path = curr_dir.joinpath(constants.CONFIG_FILENAME)
+ if os.path.exists(local_config_path):
+ tuning_config = open_config_with_defaults(local_config_path)
+
+ # Update local template (in current recursion stack)
+ local_template_path = curr_dir.joinpath(constants.TEMPLATE_FILENAME)
+ local_template_exists = os.path.exists(local_template_path)
+ if local_template_exists:
+ template = Template(
+ local_template_path,
+ tuning_config,
+ )
+ # Look for subdirectories for processing
+ subdirs = [d for d in curr_dir.iterdir() if d.is_dir()]
+
+ output_dir = Path(args["output_dir"], curr_dir.relative_to(root_dir))
+ paths = Paths(output_dir)
+
+ # look for images in current dir to process
+ exts = ("*.[pP][nN][gG]", "*.[jJ][pP][gG]", "*.[jJ][pP][eE][gG]")
+ omr_files = sorted([f for ext in exts for f in curr_dir.glob(ext)])
+
+ # Exclude images (take union over all pre_processors)
+ excluded_files = []
+ if template:
+ for pp in template.pre_processors:
+ excluded_files.extend(Path(p) for p in pp.exclude_files())
+
+ local_evaluation_path = curr_dir.joinpath(constants.EVALUATION_FILENAME)
+ if not args["setLayout"] and os.path.exists(local_evaluation_path):
+ if not local_template_exists:
+ logger.warning(
+ f"Found an evaluation file without a parent template file: {local_evaluation_path}"
+ )
+ evaluation_config = EvaluationConfig(
+ curr_dir,
+ local_evaluation_path,
+ template,
+ tuning_config,
+ )
+
+ excluded_files.extend(
+ Path(exclude_file) for exclude_file in evaluation_config.get_exclude_files()
+ )
+
+ omr_files = [f for f in omr_files if f not in excluded_files]
+
+ if omr_files:
+ if not template:
+ logger.error(
+ f"Found images, but no template in the directory tree \
+ of '{curr_dir}'. \nPlace {constants.TEMPLATE_FILENAME} in the \
+ appropriate directory."
+ )
+ raise Exception(
+ f"No template file found in the directory tree of {curr_dir}"
+ )
+
+ setup_dirs_for_paths(paths)
+ outputs_namespace = setup_outputs_for_template(paths, template)
+
+ print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+ )
+ if args["setLayout"]:
+ show_template_layouts(omr_files, template, tuning_config)
+ else:
+ process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+ )
+
+ elif not subdirs:
+ # Each subdirectory should have images or should be non-leaf
+ logger.info(
+ f"No valid images or sub-folders found in {curr_dir}.\
+ Empty directories not allowed."
+ )
+
+ # recursively process sub-folders
+ for d in subdirs:
+ process_dir(
+ root_dir,
+ d,
+ args,
+ template,
+ tuning_config,
+ evaluation_config,
+ )
+
+
+def show_template_layouts(omr_files, template, tuning_config):
+ for file_path in omr_files:
+ file_name = file_path.name
+ file_path = str(file_path)
+ in_omr = cv2.imread(file_path, cv2.IMREAD_GRAYSCALE)
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+ template_layout = template.image_instance_ops.draw_template_layout(
+ in_omr, template, shifted=False, border=2
+ )
+ InteractionUtils.show(
+ f"Template Layout: {file_name}", template_layout, 1, 1, config=tuning_config
+ )
+
+
+def process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+):
+ start_time = int(time())
+ files_counter = 0
+ STATS.files_not_moved = 0
+
+ for file_path in omr_files:
+ files_counter += 1
+ file_name = file_path.name
+
+ in_omr = cv2.imread(str(file_path), cv2.IMREAD_GRAYSCALE)
+
+ logger.info("")
+ logger.info(
+ f"({files_counter}) Opening image: \t'{file_path}'\tResolution: {in_omr.shape}"
+ )
+
+ template.image_instance_ops.reset_all_save_img()
+
+ template.image_instance_ops.append_save_img(1, in_omr)
+
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+
+ if in_omr is None:
+ # Error OMR case
+ new_file_path = outputs_namespace.paths.errors_dir.joinpath(file_name)
+ outputs_namespace.OUTPUT_SET.append(
+ [file_name] + outputs_namespace.empty_resp
+ )
+ if check_and_move(
+ constants.ERROR_CODES.NO_MARKER_ERR, file_path, new_file_path
+ ):
+ err_line = [
+ file_name,
+ file_path,
+ new_file_path,
+ "NA",
+ ] + outputs_namespace.empty_resp
+ pd.DataFrame(err_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Errors"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ continue
+
+ # uniquify
+ file_id = str(file_name)
+ save_dir = outputs_namespace.paths.save_marked_dir
+ (
+ response_dict,
+ final_marked,
+ multi_marked,
+ _,
+ ) = template.image_instance_ops.read_omr_response(
+ template, image=in_omr, name=file_id, save_dir=save_dir
+ )
+
+ # TODO: move inner try catch here
+ # concatenate roll nos, set unmarked responses, etc
+ omr_response = get_concatenated_response(response_dict, template)
+
+ if (
+ evaluation_config is None
+ or not evaluation_config.get_should_explain_scoring()
+ ):
+ logger.info(f"Read Response: \n{omr_response}")
+
+ score = 0
+ if evaluation_config is not None:
+ score = evaluate_concatenated_response(
+ omr_response, evaluation_config, file_path, outputs_namespace.paths.evaluation_dir
+ )
+ logger.info(
+ f"(/{files_counter}) Graded with score: {round(score, 2)}\t for file: '{file_id}'"
+ )
+ else:
+ logger.info(f"(/{files_counter}) Processed file: '{file_id}'")
+
+ if tuning_config.outputs.show_image_level >= 2:
+ InteractionUtils.show(
+ f"Final Marked Bubbles : '{file_id}'",
+ ImageUtils.resize_util_h(
+ final_marked, int(tuning_config.dimensions.display_height * 1.3)
+ ),
+ 1,
+ 1,
+ config=tuning_config,
+ )
+
+ resp_array = []
+ for k in template.output_columns:
+ resp_array.append(omr_response[k])
+
+ outputs_namespace.OUTPUT_SET.append([file_name] + resp_array)
+
+ if multi_marked == 0 or not tuning_config.outputs.filter_out_multimarked_files:
+ STATS.files_not_moved += 1
+ new_file_path = save_dir.joinpath(file_id)
+ # Enter into Results sheet-
+ results_line = [file_name, file_path, new_file_path, score] + resp_array
+ # Write/Append to results_line file(opened in append mode)
+ pd.DataFrame(results_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Results"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ else:
+ # multi_marked file
+ logger.info(f"[{files_counter}] Found multi-marked file: '{file_id}'")
+ new_file_path = outputs_namespace.paths.multi_marked_dir.joinpath(file_name)
+ if check_and_move(
+ constants.ERROR_CODES.MULTI_BUBBLE_WARN, file_path, new_file_path
+ ):
+ mm_line = [file_name, file_path, new_file_path, "NA"] + resp_array
+ pd.DataFrame(mm_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["MultiMarked"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ # else:
+ # TODO: Add appropriate record handling here
+ # pass
+
+ print_stats(start_time, files_counter, tuning_config)
+
+
+def check_and_move(error_code, file_path, filepath2):
+ # TODO: fix file movement into error/multimarked/invalid etc again
+ STATS.files_not_moved += 1
+ return True
+
+
+def print_stats(start_time, files_counter, tuning_config):
+ time_checking = max(1, round(time() - start_time, 2))
+ log = logger.info
+ log("")
+ log(f"{'Total file(s) moved': <27}: {STATS.files_moved}")
+ log(f"{'Total file(s) not moved': <27}: {STATS.files_not_moved}")
+ log("--------------------------------")
+ log(
+ f"{'Total file(s) processed': <27}: {files_counter} ({'Sum Tallied!' if files_counter == (STATS.files_moved + STATS.files_not_moved) else 'Not Tallying!'})"
+ )
+
+ if tuning_config.outputs.show_image_level <= 0:
+ log(
+ f"\nFinished Checking {files_counter} file(s) in {round(time_checking, 1)} seconds i.e. ~{round(time_checking / 60, 1)} minute(s)."
+ )
+ log(
+ f"{'OMR Processing Rate': <27}: \t ~ {round(time_checking / files_counter, 2)} seconds/OMR"
+ )
+ log(
+ f"{'OMR Processing Speed': <27}: \t ~ {round((files_counter * 60) / time_checking, 2)} OMRs/minute"
+ )
+ else:
+ log(f"\n{'Total script time': <27}: {time_checking} seconds")
+
+ if tuning_config.outputs.show_image_level <= 1:
+ log(
+ "\nTip: To see some awesome visuals, open config.json and increase 'show_image_level'"
+ )
diff --git a/OMRChecker/src/evaluation.py b/OMRChecker/src/evaluation.py
new file mode 100644
index 0000000000000000000000000000000000000000..55c2455a2967063e7604d0f0c904c09720e1064d
--- /dev/null
+++ b/OMRChecker/src/evaluation.py
@@ -0,0 +1,546 @@
+import ast
+import os
+import re
+from copy import deepcopy
+from csv import QUOTE_NONNUMERIC
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src.logger import console, logger
+from src.schemas.constants import (
+ BONUS_SECTION_PREFIX,
+ DEFAULT_SECTION_KEY,
+ MARKING_VERDICT_TYPES,
+)
+from src.utils.parsing import (
+ get_concatenated_response,
+ open_evaluation_with_validation,
+ parse_fields,
+ parse_float_or_fraction,
+)
+
+
+class AnswerMatcher:
+ def __init__(self, answer_item, section_marking_scheme):
+ self.section_marking_scheme = section_marking_scheme
+ self.answer_item = answer_item
+ self.answer_type = self.validate_and_get_answer_type(answer_item)
+ self.set_defaults_from_scheme(section_marking_scheme)
+
+ @staticmethod
+ def is_a_marking_score(answer_element):
+ # Note: strict type checking is already done at schema validation level,
+ # Here we focus on overall struct type
+ return type(answer_element) == str or type(answer_element) == int
+
+ @staticmethod
+ def is_standard_answer(answer_element):
+ return type(answer_element) == str and len(answer_element) >= 1
+
+ def validate_and_get_answer_type(self, answer_item):
+ if self.is_standard_answer(answer_item):
+ return "standard"
+ elif type(answer_item) == list:
+ if (
+ # Array of answer elements: ['A', 'B', 'AB']
+ len(answer_item) >= 2
+ and all(
+ self.is_standard_answer(answers_or_score)
+ for answers_or_score in answer_item
+ )
+ ):
+ return "multiple-correct"
+ elif (
+ # Array of two-tuples: [['A', 1], ['B', 1], ['C', 3], ['AB', 2]]
+ len(answer_item) >= 1
+ and all(
+ type(answer_and_score) == list and len(answer_and_score) == 2
+ for answer_and_score in answer_item
+ )
+ and all(
+ self.is_standard_answer(allowed_answer)
+ and self.is_a_marking_score(answer_score)
+ for allowed_answer, answer_score in answer_item
+ )
+ ):
+ return "multiple-correct-weighted"
+
+ logger.critical(
+ f"Unable to determine answer type for answer item: {answer_item}"
+ )
+ raise Exception("Unable to determine answer type")
+
+ def set_defaults_from_scheme(self, section_marking_scheme):
+ answer_type = self.answer_type
+ self.empty_val = section_marking_scheme.empty_val
+ answer_item = self.answer_item
+ self.marking = deepcopy(section_marking_scheme.marking)
+ # TODO: reuse part of parse_scheme_marking here -
+ if answer_type == "standard":
+ # no local overrides
+ pass
+ elif answer_type == "multiple-correct":
+ # override marking scheme scores for each allowed answer
+ for allowed_answer in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = self.marking["correct"]
+ elif answer_type == "multiple-correct-weighted":
+ # Note: No override using marking scheme as answer scores are provided in answer_item
+ for allowed_answer, answer_score in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = parse_float_or_fraction(
+ answer_score
+ )
+
+ def get_marking_scheme(self):
+ return self.section_marking_scheme
+
+ def get_section_explanation(self):
+ answer_type = self.answer_type
+ if answer_type in ["standard", "multiple-correct"]:
+ return self.section_marking_scheme.section_key
+ elif answer_type == "multiple-correct-weighted":
+ return f"Custom: {self.marking}"
+
+ def get_verdict_marking(self, marked_answer):
+ answer_type = self.answer_type
+ question_verdict = "incorrect"
+ if answer_type == "standard":
+ question_verdict = self.get_standard_verdict(marked_answer)
+ elif answer_type == "multiple-correct":
+ question_verdict = self.get_multiple_correct_verdict(marked_answer)
+ elif answer_type == "multiple-correct-weighted":
+ question_verdict = self.get_multiple_correct_weighted_verdict(marked_answer)
+ return question_verdict, self.marking[question_verdict]
+
+ def get_standard_verdict(self, marked_answer):
+ allowed_answer = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer == allowed_answer:
+ return "correct"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_verdict(self, marked_answer):
+ allowed_answers = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_weighted_verdict(self, marked_answer):
+ allowed_answers = [
+ allowed_answer for allowed_answer, _answer_score in self.answer_item
+ ]
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def __str__(self):
+ return f"{self.answer_item}"
+
+
+class SectionMarkingScheme:
+ def __init__(self, section_key, section_scheme, empty_val):
+ # TODO: get local empty_val from qblock
+ self.empty_val = empty_val
+ self.section_key = section_key
+ # DEFAULT marking scheme follows a shorthand
+ if section_key == DEFAULT_SECTION_KEY:
+ self.questions = None
+ self.marking = self.parse_scheme_marking(section_scheme)
+ else:
+ self.questions = parse_fields(section_key, section_scheme["questions"])
+ self.marking = self.parse_scheme_marking(section_scheme["marking"])
+
+ def __str__(self):
+ return self.section_key
+
+ def parse_scheme_marking(self, marking):
+ parsed_marking = {}
+ for verdict_type in MARKING_VERDICT_TYPES:
+ verdict_marking = parse_float_or_fraction(marking[verdict_type])
+ if (
+ verdict_marking > 0
+ and verdict_type == "incorrect"
+ and not self.section_key.startswith(BONUS_SECTION_PREFIX)
+ ):
+ logger.warning(
+ f"Found positive marks({round(verdict_marking, 2)}) for incorrect answer in the schema '{self.section_key}'. For Bonus sections, add a prefix 'BONUS_' to them."
+ )
+ parsed_marking[verdict_type] = verdict_marking
+
+ return parsed_marking
+
+ def match_answer(self, marked_answer, answer_matcher):
+ question_verdict, verdict_marking = answer_matcher.get_verdict_marking(
+ marked_answer
+ )
+
+ return verdict_marking, question_verdict
+
+
+class EvaluationConfig:
+ """Note: this instance will be reused for multiple omr sheets"""
+
+ def __init__(self, curr_dir, evaluation_path, template, tuning_config):
+ self.path = evaluation_path
+ evaluation_json = open_evaluation_with_validation(evaluation_path)
+ options, marking_schemes, source_type = map(
+ evaluation_json.get, ["options", "marking_schemes", "source_type"]
+ )
+ self.should_explain_scoring = options.get("should_explain_scoring", False)
+ self.has_non_default_section = False
+ self.exclude_files = []
+ self.enable_evaluation_table_to_csv = options.get(
+ "enable_evaluation_table_to_csv", False
+ )
+
+ if source_type == "csv":
+ csv_path = curr_dir.joinpath(options["answer_key_csv_path"])
+ if not os.path.exists(csv_path):
+ logger.warning(f"Answer key csv does not exist at: '{csv_path}'.")
+
+ answer_key_image_path = options.get("answer_key_image_path", None)
+ if os.path.exists(csv_path):
+ # TODO: CSV parsing/validation for each row with a (qNo, ) pair
+ answer_key = pd.read_csv(
+ csv_path,
+ header=None,
+ names=["question", "answer"],
+ converters={"question": str, "answer": self.parse_answer_column},
+ )
+
+ self.questions_in_order = answer_key["question"].to_list()
+ answers_in_order = answer_key["answer"].to_list()
+ elif not answer_key_image_path:
+ raise Exception(f"Answer key csv not found at '{csv_path}'")
+ else:
+ image_path = str(curr_dir.joinpath(answer_key_image_path))
+ if not os.path.exists(image_path):
+ raise Exception(f"Answer key image not found at '{image_path}'")
+
+ # self.exclude_files.append(image_path)
+
+ logger.debug(
+ f"Attempting to generate answer key from image: '{image_path}'"
+ )
+ # TODO: use a common function for below changes?
+ in_omr = cv2.imread(image_path, cv2.IMREAD_GRAYSCALE)
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ image_path, in_omr, template
+ )
+ if in_omr is None:
+ raise Exception(
+ f"Could not read answer key from image {image_path}"
+ )
+ (
+ response_dict,
+ _final_marked,
+ _multi_marked,
+ _multi_roll,
+ ) = template.image_instance_ops.read_omr_response(
+ template,
+ image=in_omr,
+ name=image_path,
+ save_dir=None,
+ )
+ omr_response = get_concatenated_response(response_dict, template)
+
+ empty_val = template.global_empty_val
+ empty_answer_regex = (
+ rf"{re.escape(empty_val)}+" if empty_val != "" else r"^$"
+ )
+
+ if "questions_in_order" in options:
+ self.questions_in_order = self.parse_questions_in_order(
+ options["questions_in_order"]
+ )
+ empty_answered_questions = [
+ question
+ for question in self.questions_in_order
+ if re.search(empty_answer_regex, omr_response[question])
+ ]
+ if len(empty_answered_questions) > 0:
+ logger.error(
+ f"Found empty answers for questions: {empty_answered_questions}, empty value used: '{empty_val}'"
+ )
+ raise Exception(
+ f"Found empty answers in file '{image_path}'. Please check your template again in the --setLayout mode."
+ )
+ else:
+ logger.warning(
+ f"questions_in_order not provided, proceeding to use non-empty values as answer key"
+ )
+ self.questions_in_order = sorted(
+ question
+ for (question, answer) in omr_response.items()
+ if not re.search(empty_answer_regex, answer)
+ )
+ answers_in_order = [
+ omr_response[question] for question in self.questions_in_order
+ ]
+ # TODO: save the CSV
+ else:
+ self.questions_in_order = self.parse_questions_in_order(
+ options["questions_in_order"]
+ )
+ answers_in_order = options["answers_in_order"]
+
+ self.validate_questions(answers_in_order)
+
+ self.section_marking_schemes, self.question_to_scheme = {}, {}
+ for section_key, section_scheme in marking_schemes.items():
+ section_marking_scheme = SectionMarkingScheme(
+ section_key, section_scheme, template.global_empty_val
+ )
+ if section_key != DEFAULT_SECTION_KEY:
+ self.section_marking_schemes[section_key] = section_marking_scheme
+ for q in section_marking_scheme.questions:
+ # TODO: check the answer key for custom scheme here?
+ self.question_to_scheme[q] = section_marking_scheme
+ self.has_non_default_section = True
+ else:
+ self.default_marking_scheme = section_marking_scheme
+
+ self.validate_marking_schemes()
+
+ self.question_to_answer_matcher = self.parse_answers_and_map_questions(
+ answers_in_order
+ )
+ self.validate_answers(answers_in_order, tuning_config)
+
+ def __str__(self):
+ return str(self.path)
+
+ # Externally called methods have higher abstraction level.
+ def prepare_and_validate_omr_response(self, omr_response):
+ self.reset_explanation_table()
+
+ omr_response_questions = set(omr_response.keys())
+ all_questions = set(self.questions_in_order)
+ missing_questions = sorted(all_questions.difference(omr_response_questions))
+ if len(missing_questions) > 0:
+ logger.critical(f"Missing OMR response for: {missing_questions}")
+ raise Exception(
+ f"Some questions are missing in the OMR response for the given answer key"
+ )
+
+ prefixed_omr_response_questions = set(
+ [k for k in omr_response.keys() if k.startswith("q")]
+ )
+ missing_prefixed_questions = sorted(
+ prefixed_omr_response_questions.difference(all_questions)
+ )
+ if len(missing_prefixed_questions) > 0:
+ logger.warning(
+ f"No answer given for potential questions in OMR response: {missing_prefixed_questions}"
+ )
+
+ def match_answer_for_question(self, current_score, question, marked_answer):
+ answer_matcher = self.question_to_answer_matcher[question]
+ question_verdict, delta = answer_matcher.get_verdict_marking(marked_answer)
+ self.conditionally_add_explanation(
+ answer_matcher,
+ delta,
+ marked_answer,
+ question_verdict,
+ question,
+ current_score,
+ )
+ return delta
+
+ def conditionally_print_explanation(self):
+ if self.should_explain_scoring:
+ console.print(self.explanation_table, justify="center")
+
+ # Explanation Table to CSV
+ def conditionally_save_explanation_csv(self, file_path, evaluation_output_dir):
+ if self.enable_evaluation_table_to_csv:
+ data = {col.header: col._cells for col in self.explanation_table.columns}
+
+ output_path = os.path.join(
+ evaluation_output_dir,
+ f"{file_path.stem}_evaluation.csv",
+ )
+
+ pd.DataFrame(data, dtype=str).to_csv(
+ output_path,
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ index=False,
+ )
+
+ def get_should_explain_scoring(self):
+ return self.should_explain_scoring
+
+ def get_exclude_files(self):
+ return self.exclude_files
+
+ @staticmethod
+ def parse_answer_column(answer_column):
+ if answer_column[0] == "[":
+ # multiple-correct-weighted or multiple-correct
+ parsed_answer = ast.literal_eval(answer_column)
+ elif "," in answer_column:
+ # multiple-correct
+ parsed_answer = answer_column.split(",")
+ else:
+ # single-correct
+ parsed_answer = answer_column
+ return parsed_answer
+
+ def parse_questions_in_order(self, questions_in_order):
+ return parse_fields("questions_in_order", questions_in_order)
+
+ def validate_answers(self, answers_in_order, tuning_config):
+ answer_matcher_map = self.question_to_answer_matcher
+ if tuning_config.outputs.filter_out_multimarked_files:
+ multi_marked_answer = False
+ for question, answer_item in zip(self.questions_in_order, answers_in_order):
+ answer_type = answer_matcher_map[question].answer_type
+ if answer_type == "standard":
+ if len(answer_item) > 1:
+ multi_marked_answer = True
+ if answer_type == "multiple-correct":
+ for single_answer in answer_item:
+ if len(single_answer) > 1:
+ multi_marked_answer = True
+ break
+ if answer_type == "multiple-correct-weighted":
+ for single_answer, _answer_score in answer_item:
+ if len(single_answer) > 1:
+ multi_marked_answer = True
+
+ if multi_marked_answer:
+ raise Exception(
+ f"Provided answer key contains multiple correct answer(s), but config.filter_out_multimarked_files is True. Scoring will get skipped."
+ )
+
+ def validate_questions(self, answers_in_order):
+ questions_in_order = self.questions_in_order
+ len_questions_in_order, len_answers_in_order = len(questions_in_order), len(
+ answers_in_order
+ )
+ if len_questions_in_order != len_answers_in_order:
+ logger.critical(
+ f"questions_in_order({len_questions_in_order}): {questions_in_order}\nanswers_in_order({len_answers_in_order}): {answers_in_order}"
+ )
+ raise Exception(
+ f"Unequal lengths for questions_in_order and answers_in_order ({len_questions_in_order} != {len_answers_in_order})"
+ )
+
+ def validate_marking_schemes(self):
+ section_marking_schemes = self.section_marking_schemes
+ section_questions = set()
+ for section_key, section_scheme in section_marking_schemes.items():
+ if section_key == DEFAULT_SECTION_KEY:
+ continue
+ current_set = set(section_scheme.questions)
+ if not section_questions.isdisjoint(current_set):
+ raise Exception(
+ f"Section '{section_key}' has overlapping question(s) with other sections"
+ )
+ section_questions = section_questions.union(current_set)
+
+ all_questions = set(self.questions_in_order)
+ missing_questions = sorted(section_questions.difference(all_questions))
+ if len(missing_questions) > 0:
+ logger.critical(f"Missing answer key for: {missing_questions}")
+ raise Exception(
+ f"Some questions are missing in the answer key for the given marking scheme"
+ )
+
+ def parse_answers_and_map_questions(self, answers_in_order):
+ question_to_answer_matcher = {}
+ for question, answer_item in zip(self.questions_in_order, answers_in_order):
+ section_marking_scheme = self.get_marking_scheme_for_question(question)
+ answer_matcher = AnswerMatcher(answer_item, section_marking_scheme)
+ question_to_answer_matcher[question] = answer_matcher
+ if (
+ answer_matcher.answer_type == "multiple-correct-weighted"
+ and section_marking_scheme.section_key != DEFAULT_SECTION_KEY
+ ):
+ logger.warning(
+ f"The custom scheme '{section_marking_scheme}' will not apply to question '{question}' as it will use the given answer weights f{answer_item}"
+ )
+ return question_to_answer_matcher
+
+ # Then unfolding lower abstraction levels
+ def reset_explanation_table(self):
+ self.explanation_table = None
+ self.prepare_explanation_table()
+
+ def prepare_explanation_table(self):
+ # TODO: provide a way to export this as csv/pdf
+ if not self.should_explain_scoring:
+ return
+ table = Table(title="Evaluation Explanation Table", show_lines=True)
+ table.add_column("Question")
+ table.add_column("Marked")
+ table.add_column("Answer(s)")
+ table.add_column("Verdict")
+ table.add_column("Delta")
+ table.add_column("Score")
+ # TODO: Add max and min score in explanation (row-wise and total)
+ if self.has_non_default_section:
+ table.add_column("Section")
+ self.explanation_table = table
+
+ def get_marking_scheme_for_question(self, question):
+ return self.question_to_scheme.get(question, self.default_marking_scheme)
+
+ def conditionally_add_explanation(
+ self,
+ answer_matcher,
+ delta,
+ marked_answer,
+ question_verdict,
+ question,
+ current_score,
+ ):
+ if self.should_explain_scoring:
+ next_score = current_score + delta
+ # Conditionally add cells
+ row = [
+ item
+ for item in [
+ question,
+ marked_answer,
+ str(answer_matcher),
+ str.title(question_verdict),
+ str(round(delta, 2)),
+ str(round(next_score, 2)),
+ (
+ answer_matcher.get_section_explanation()
+ if self.has_non_default_section
+ else None
+ ),
+ ]
+ if item is not None
+ ]
+ self.explanation_table.add_row(*row)
+
+
+def evaluate_concatenated_response(
+ concatenated_response, evaluation_config, file_path, evaluation_output_dir
+):
+ evaluation_config.prepare_and_validate_omr_response(concatenated_response)
+ current_score = 0.0
+ for question in evaluation_config.questions_in_order:
+ marked_answer = concatenated_response[question]
+ delta = evaluation_config.match_answer_for_question(
+ current_score, question, marked_answer
+ )
+ current_score += delta
+
+ evaluation_config.conditionally_print_explanation()
+ evaluation_config.conditionally_save_explanation_csv(file_path, evaluation_output_dir)
+
+ return current_score
diff --git a/OMRChecker/src/logger.py b/OMRChecker/src/logger.py
new file mode 100644
index 0000000000000000000000000000000000000000..d78988b8e8b6dc5d123c58fd0068c13bf9e67f42
--- /dev/null
+++ b/OMRChecker/src/logger.py
@@ -0,0 +1,68 @@
+import logging
+from typing import Union
+
+from rich.console import Console
+from rich.logging import RichHandler
+
+FORMAT = "%(message)s"
+
+# TODO: set logging level from config.json dynamically
+logging.basicConfig(
+ level=logging.INFO,
+ format="%(message)s",
+ datefmt="[%X]",
+ handlers=[RichHandler(rich_tracebacks=True)],
+)
+
+
+class Logger:
+ def __init__(
+ self,
+ name,
+ level: Union[int, str] = logging.NOTSET,
+ message_format="%(message)s",
+ date_format="[%X]",
+ ):
+ self.log = logging.getLogger(name)
+ self.log.setLevel(level)
+ self.log.__format__ = message_format
+ self.log.__date_format__ = date_format
+
+ def debug(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("debug", *msg, sep=sep)
+
+ def info(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("info", *msg, sep=sep)
+
+ def warning(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("warning", *msg, sep=sep)
+
+ def error(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("error", *msg, sep=sep)
+
+ def critical(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("critical", *msg, sep=sep)
+
+ def stringify(func):
+ def inner(self, method_type: str, *msg: object, sep=" "):
+ nmsg = []
+ for v in msg:
+ if not isinstance(v, str):
+ v = str(v)
+ nmsg.append(v)
+ return func(self, method_type, *nmsg, sep=sep)
+
+ return inner
+
+ # set stack level to 3 so that the caller of this function is logged, not this function itself.
+ # stack-frame - self.log.debug - logutil - stringify - log method - caller
+ @stringify
+ def logutil(self, method_type: str, *msg: object, sep=" ") -> None:
+ func = getattr(self.log, method_type, None)
+ if not func:
+ raise AttributeError(f"Logger has no method {method_type}")
+ return func(sep.join(msg), stacklevel=4)
+
+
+logger = Logger(__name__)
+console = Console()
diff --git a/OMRChecker/src/processors/CropOnMarkers.py b/OMRChecker/src/processors/CropOnMarkers.py
new file mode 100644
index 0000000000000000000000000000000000000000..3801d88af313c83dd514ab33ab9bde1cf2afd16d
--- /dev/null
+++ b/OMRChecker/src/processors/CropOnMarkers.py
@@ -0,0 +1,233 @@
+import os
+
+import cv2
+import numpy as np
+
+from src.logger import logger
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class CropOnMarkers(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ config = self.tuning_config
+ marker_ops = self.options
+ self.threshold_circles = []
+ # img_utils = ImageUtils()
+
+ # options with defaults
+ self.marker_path = os.path.join(
+ self.relative_dir, marker_ops.get("relativePath", "omr_marker.jpg")
+ )
+ self.min_matching_threshold = marker_ops.get("min_matching_threshold", 0.3)
+ self.max_matching_variation = marker_ops.get("max_matching_variation", 0.41)
+ self.marker_rescale_range = tuple(
+ int(r) for r in marker_ops.get("marker_rescale_range", (35, 100))
+ )
+ self.marker_rescale_steps = int(marker_ops.get("marker_rescale_steps", 10))
+ self.apply_erode_subtract = marker_ops.get("apply_erode_subtract", True)
+ self.marker = self.load_marker(marker_ops, config)
+
+ def __str__(self):
+ return self.marker_path
+
+ def exclude_files(self):
+ return [self.marker_path]
+
+ def apply_filter(self, image, file_path):
+ config = self.tuning_config
+ image_instance_ops = self.image_instance_ops
+ image_eroded_sub = ImageUtils.normalize_util(
+ image
+ if self.apply_erode_subtract
+ else (image - cv2.erode(image, kernel=np.ones((5, 5)), iterations=5))
+ )
+ # Quads on warped image
+ quads = {}
+ h1, w1 = image_eroded_sub.shape[:2]
+ midh, midw = h1 // 3, w1 // 2
+ origins = [[0, 0], [midw, 0], [0, midh], [midw, midh]]
+ quads[0] = image_eroded_sub[0:midh, 0:midw]
+ quads[1] = image_eroded_sub[0:midh, midw:w1]
+ quads[2] = image_eroded_sub[midh:h1, 0:midw]
+ quads[3] = image_eroded_sub[midh:h1, midw:w1]
+
+ # Draw Quadlines
+ image_eroded_sub[:, midw : midw + 2] = 255
+ image_eroded_sub[midh : midh + 2, :] = 255
+
+ best_scale, all_max_t = self.getBestMatch(image_eroded_sub)
+ if best_scale is None:
+ if config.outputs.show_image_level >= 1:
+ InteractionUtils.show("Quads", image_eroded_sub, config=config)
+ return None
+
+ optimal_marker = ImageUtils.resize_util_h(
+ self.marker, u_height=int(self.marker.shape[0] * best_scale)
+ )
+ _h, w = optimal_marker.shape[:2]
+ centres = []
+ sum_t, max_t = 0, 0
+ quarter_match_log = "Matching Marker: "
+ for k in range(0, 4):
+ res = cv2.matchTemplate(quads[k], optimal_marker, cv2.TM_CCOEFF_NORMED)
+ max_t = res.max()
+ quarter_match_log += f"Quarter{str(k + 1)}: {str(round(max_t, 3))}\t"
+ if (
+ max_t < self.min_matching_threshold
+ or abs(all_max_t - max_t) >= self.max_matching_variation
+ ):
+ logger.error(
+ file_path,
+ "\nError: No circle found in Quad",
+ k + 1,
+ "\n\t min_matching_threshold",
+ self.min_matching_threshold,
+ "\t max_matching_variation",
+ self.max_matching_variation,
+ "\t max_t",
+ max_t,
+ "\t all_max_t",
+ all_max_t,
+ )
+ if config.outputs.show_image_level >= 1:
+ InteractionUtils.show(
+ f"No markers: {file_path}",
+ image_eroded_sub,
+ 0,
+ config=config,
+ )
+ InteractionUtils.show(
+ f"res_Q{str(k + 1)} ({str(max_t)})",
+ res,
+ 1,
+ config=config,
+ )
+ return None
+
+ pt = np.argwhere(res == max_t)[0]
+ pt = [pt[1], pt[0]]
+ pt[0] += origins[k][0]
+ pt[1] += origins[k][1]
+ # print(">>",pt)
+ image = cv2.rectangle(
+ image, tuple(pt), (pt[0] + w, pt[1] + _h), (150, 150, 150), 2
+ )
+ # display:
+ image_eroded_sub = cv2.rectangle(
+ image_eroded_sub,
+ tuple(pt),
+ (pt[0] + w, pt[1] + _h),
+ (50, 50, 50) if self.apply_erode_subtract else (155, 155, 155),
+ 4,
+ )
+ centres.append([pt[0] + w / 2, pt[1] + _h / 2])
+ sum_t += max_t
+
+ logger.info(quarter_match_log)
+ logger.info(f"Optimal Scale: {best_scale}")
+ # analysis data
+ self.threshold_circles.append(sum_t / 4)
+
+ image = ImageUtils.four_point_transform(image, np.array(centres))
+ # appendSaveImg(1,image_eroded_sub)
+ # appendSaveImg(1,image_norm)
+
+ image_instance_ops.append_save_img(2, image_eroded_sub)
+ # Debugging image -
+ # res = cv2.matchTemplate(image_eroded_sub,optimal_marker,cv2.TM_CCOEFF_NORMED)
+ # res[ : , midw:midw+2] = 255
+ # res[ midh:midh+2, : ] = 255
+ # show("Markers Matching",res)
+ if config.outputs.show_image_level >= 2 and config.outputs.show_image_level < 4:
+ image_eroded_sub = ImageUtils.resize_util_h(
+ image_eroded_sub, image.shape[0]
+ )
+ image_eroded_sub[:, -5:] = 0
+ h_stack = np.hstack((image_eroded_sub, image))
+ InteractionUtils.show(
+ f"Warped: {file_path}",
+ ImageUtils.resize_util(
+ h_stack, int(config.dimensions.display_width * 1.6)
+ ),
+ 0,
+ 0,
+ [0, 0],
+ config=config,
+ )
+ # iterations : Tuned to 2.
+ # image_eroded_sub = image_norm - cv2.erode(image_norm, kernel=np.ones((5,5)),iterations=2)
+ return image
+
+ def load_marker(self, marker_ops, config):
+ if not os.path.exists(self.marker_path):
+ logger.error(
+ "Marker not found at path provided in template:",
+ self.marker_path,
+ )
+ exit(31)
+
+ marker = cv2.imread(self.marker_path, cv2.IMREAD_GRAYSCALE)
+
+ if "sheetToMarkerWidthRatio" in marker_ops:
+ marker = ImageUtils.resize_util(
+ marker,
+ config.dimensions.processing_width
+ / int(marker_ops["sheetToMarkerWidthRatio"]),
+ )
+ marker = cv2.GaussianBlur(marker, (5, 5), 0)
+ marker = cv2.normalize(
+ marker, None, alpha=0, beta=255, norm_type=cv2.NORM_MINMAX
+ )
+
+ if self.apply_erode_subtract:
+ marker -= cv2.erode(marker, kernel=np.ones((5, 5)), iterations=5)
+
+ return marker
+
+ # Resizing the marker within scaleRange at rate of descent_per_step to
+ # find the best match.
+ def getBestMatch(self, image_eroded_sub):
+ config = self.tuning_config
+ descent_per_step = (
+ self.marker_rescale_range[1] - self.marker_rescale_range[0]
+ ) // self.marker_rescale_steps
+ _h, _w = self.marker.shape[:2]
+ res, best_scale = None, None
+ all_max_t = 0
+
+ for r0 in np.arange(
+ self.marker_rescale_range[1],
+ self.marker_rescale_range[0],
+ -1 * descent_per_step,
+ ): # reverse order
+ s = float(r0 * 1 / 100)
+ if s == 0.0:
+ continue
+ rescaled_marker = ImageUtils.resize_util_h(
+ self.marker, u_height=int(_h * s)
+ )
+ # res is the black image with white dots
+ res = cv2.matchTemplate(
+ image_eroded_sub, rescaled_marker, cv2.TM_CCOEFF_NORMED
+ )
+
+ max_t = res.max()
+ if all_max_t < max_t:
+ # print('Scale: '+str(s)+', Circle Match: '+str(round(max_t*100,2))+'%')
+ best_scale, all_max_t = s, max_t
+
+ if all_max_t < self.min_matching_threshold:
+ logger.warning(
+ "\tTemplate matching too low! Consider rechecking preProcessors applied before this."
+ )
+ if config.outputs.show_image_level >= 1:
+ InteractionUtils.show("res", res, 1, 0, config=config)
+
+ if best_scale is None:
+ logger.warning(
+ "No matchings for given scaleRange:", self.marker_rescale_range
+ )
+ return best_scale, all_max_t
diff --git a/OMRChecker/src/processors/CropPage.py b/OMRChecker/src/processors/CropPage.py
new file mode 100644
index 0000000000000000000000000000000000000000..49a6782597aaeae6ee00de28eedef5d40f1fc5a6
--- /dev/null
+++ b/OMRChecker/src/processors/CropPage.py
@@ -0,0 +1,112 @@
+"""
+https://www.pyimagesearch.com/2015/04/06/zero-parameter-automatic-canny-edge-detection-with-python-and-opencv/
+"""
+import cv2
+import numpy as np
+
+from src.logger import logger
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils
+
+MIN_PAGE_AREA = 80000
+
+
+def normalize(image):
+ return cv2.normalize(image, 0, 255, norm_type=cv2.NORM_MINMAX)
+
+
+def check_max_cosine(approx):
+ # assumes 4 pts present
+ max_cosine = 0
+ min_cosine = 1.5
+ for i in range(2, 5):
+ cosine = abs(angle(approx[i % 4], approx[i - 2], approx[i - 1]))
+ max_cosine = max(cosine, max_cosine)
+ min_cosine = min(cosine, min_cosine)
+
+ if max_cosine >= 0.35:
+ logger.warning("Quadrilateral is not a rectangle.")
+ return False
+ return True
+
+
+def validate_rect(approx):
+ return len(approx) == 4 and check_max_cosine(approx.reshape(4, 2))
+
+
+def angle(p_1, p_2, p_0):
+ dx1 = float(p_1[0] - p_0[0])
+ dy1 = float(p_1[1] - p_0[1])
+ dx2 = float(p_2[0] - p_0[0])
+ dy2 = float(p_2[1] - p_0[1])
+ return (dx1 * dx2 + dy1 * dy2) / np.sqrt(
+ (dx1 * dx1 + dy1 * dy1) * (dx2 * dx2 + dy2 * dy2) + 1e-10
+ )
+
+
+class CropPage(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ cropping_ops = self.options
+ self.morph_kernel = tuple(
+ int(x) for x in cropping_ops.get("morphKernel", [10, 10])
+ )
+
+ def apply_filter(self, image, file_path):
+ image = normalize(cv2.GaussianBlur(image, (3, 3), 0))
+
+ # Resize should be done with another preprocessor is needed
+ sheet = self.find_page(image, file_path)
+ if len(sheet) == 0:
+ logger.error(
+ f"\tError: Paper boundary not found for: '{file_path}'\nHave you accidentally included CropPage preprocessor?"
+ )
+ return None
+
+ logger.info(f"Found page corners: \t {sheet.tolist()}")
+
+ # Warp layer 1
+ image = ImageUtils.four_point_transform(image, sheet)
+
+ # Return preprocessed image
+ return image
+
+ def find_page(self, image, file_path):
+ config = self.tuning_config
+
+ image = normalize(image)
+
+ _ret, image = cv2.threshold(image, 200, 255, cv2.THRESH_TRUNC)
+ image = normalize(image)
+
+ kernel = cv2.getStructuringElement(cv2.MORPH_RECT, self.morph_kernel)
+
+ # Close the small holes, i.e. Complete the edges on canny image
+ closed = cv2.morphologyEx(image, cv2.MORPH_CLOSE, kernel)
+
+ edge = cv2.Canny(closed, 185, 55)
+
+ if config.outputs.show_image_level >= 5:
+ InteractionUtils.show("edge", edge, config=config)
+
+ # findContours returns outer boundaries in CW and inner ones, ACW.
+ cnts = ImageUtils.grab_contours(
+ cv2.findContours(edge, cv2.RETR_LIST, cv2.CHAIN_APPROX_SIMPLE)
+ )
+ # convexHull to resolve disordered curves due to noise
+ cnts = [cv2.convexHull(c) for c in cnts]
+ cnts = sorted(cnts, key=cv2.contourArea, reverse=True)[:5]
+ sheet = []
+ for c in cnts:
+ if cv2.contourArea(c) < MIN_PAGE_AREA:
+ continue
+ peri = cv2.arcLength(c, True)
+ approx = cv2.approxPolyDP(c, epsilon=0.025 * peri, closed=True)
+ if validate_rect(approx):
+ sheet = np.reshape(approx, (4, -1))
+ cv2.drawContours(image, [approx], -1, (0, 255, 0), 2)
+ cv2.drawContours(edge, [approx], -1, (255, 255, 255), 10)
+ break
+
+ return sheet
diff --git a/OMRChecker/src/processors/FeatureBasedAlignment.py b/OMRChecker/src/processors/FeatureBasedAlignment.py
new file mode 100644
index 0000000000000000000000000000000000000000..c7b0fb26cc0f3efd4eb27fa8b1e3461b91bced0a
--- /dev/null
+++ b/OMRChecker/src/processors/FeatureBasedAlignment.py
@@ -0,0 +1,94 @@
+"""
+Image based feature alignment
+Credits: https://www.learnopencv.com/image-alignment-feature-based-using-opencv-c-python/
+"""
+import cv2
+import numpy as np
+
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class FeatureBasedAlignment(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+ config = self.tuning_config
+
+ # process reference image
+ self.ref_path = self.relative_dir.joinpath(options["reference"])
+ ref_img = cv2.imread(str(self.ref_path), cv2.IMREAD_GRAYSCALE)
+ self.ref_img = ImageUtils.resize_util(
+ ref_img,
+ config.dimensions.processing_width,
+ config.dimensions.processing_height,
+ )
+ # get options with defaults
+ self.max_features = int(options.get("maxFeatures", 500))
+ self.good_match_percent = options.get("goodMatchPercent", 0.15)
+ self.transform_2_d = options.get("2d", False)
+ # Extract keypoints and description of source image
+ self.orb = cv2.ORB_create(self.max_features)
+ self.to_keypoints, self.to_descriptors = self.orb.detectAndCompute(
+ self.ref_img, None
+ )
+
+ def __str__(self):
+ return self.ref_path.name
+
+ def exclude_files(self):
+ return [self.ref_path]
+
+ def apply_filter(self, image, _file_path):
+ config = self.tuning_config
+ # Convert images to grayscale
+ # im1Gray = cv2.cvtColor(im1, cv2.COLOR_BGR2GRAY)
+ # im2Gray = cv2.cvtColor(im2, cv2.COLOR_BGR2GRAY)
+
+ image = cv2.normalize(image, 0, 255, norm_type=cv2.NORM_MINMAX)
+
+ # Detect ORB features and compute descriptors.
+ from_keypoints, from_descriptors = self.orb.detectAndCompute(image, None)
+
+ # Match features.
+ matcher = cv2.DescriptorMatcher_create(
+ cv2.DESCRIPTOR_MATCHER_BRUTEFORCE_HAMMING
+ )
+
+ # create BFMatcher object (alternate matcher)
+ # matcher = cv2.BFMatcher(cv2.NORM_HAMMING, crossCheck=True)
+
+ matches = np.array(matcher.match(from_descriptors, self.to_descriptors, None))
+
+ # Sort matches by score
+ matches = sorted(matches, key=lambda x: x.distance, reverse=False)
+
+ # Remove not so good matches
+ num_good_matches = int(len(matches) * self.good_match_percent)
+ matches = matches[:num_good_matches]
+
+ # Draw top matches
+ if config.outputs.show_image_level > 2:
+ im_matches = cv2.drawMatches(
+ image, from_keypoints, self.ref_img, self.to_keypoints, matches, None
+ )
+ InteractionUtils.show("Aligning", im_matches, resize=True, config=config)
+
+ # Extract location of good matches
+ points1 = np.zeros((len(matches), 2), dtype=np.float32)
+ points2 = np.zeros((len(matches), 2), dtype=np.float32)
+
+ for i, match in enumerate(matches):
+ points1[i, :] = from_keypoints[match.queryIdx].pt
+ points2[i, :] = self.to_keypoints[match.trainIdx].pt
+
+ # Find homography
+ height, width = self.ref_img.shape
+ if self.transform_2_d:
+ m, _inliers = cv2.estimateAffine2D(points1, points2)
+ return cv2.warpAffine(image, m, (width, height))
+
+ # Use homography
+ h, _mask = cv2.findHomography(points1, points2, cv2.RANSAC)
+ return cv2.warpPerspective(image, h, (width, height))
diff --git a/OMRChecker/src/processors/builtins.py b/OMRChecker/src/processors/builtins.py
new file mode 100644
index 0000000000000000000000000000000000000000..3e3f54683ee104c67bdd422c0fa3cdacfcc4fd8d
--- /dev/null
+++ b/OMRChecker/src/processors/builtins.py
@@ -0,0 +1,54 @@
+import cv2
+import numpy as np
+
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+
+
+class Levels(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+
+ def output_level(value, low, high, gamma):
+ if value <= low:
+ return 0
+ if value >= high:
+ return 255
+ inv_gamma = 1.0 / gamma
+ return (((value - low) / (high - low)) ** inv_gamma) * 255
+
+ self.gamma = np.array(
+ [
+ output_level(
+ i,
+ int(255 * options.get("low", 0)),
+ int(255 * options.get("high", 1)),
+ options.get("gamma", 1.0),
+ )
+ for i in np.arange(0, 256)
+ ]
+ ).astype("uint8")
+
+ def apply_filter(self, image, _file_path):
+ return cv2.LUT(image, self.gamma)
+
+
+class MedianBlur(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+ self.kSize = int(options.get("kSize", 5))
+
+ def apply_filter(self, image, _file_path):
+ return cv2.medianBlur(image, self.kSize)
+
+
+class GaussianBlur(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+ self.kSize = tuple(int(x) for x in options.get("kSize", (3, 3)))
+ self.sigmaX = int(options.get("sigmaX", 0))
+
+ def apply_filter(self, image, _file_path):
+ return cv2.GaussianBlur(image, self.kSize, self.sigmaX)
diff --git a/OMRChecker/src/processors/interfaces/ImagePreprocessor.py b/OMRChecker/src/processors/interfaces/ImagePreprocessor.py
new file mode 100644
index 0000000000000000000000000000000000000000..619a8ad711f48affc0ebaad0c6a2bf34ee2f184d
--- /dev/null
+++ b/OMRChecker/src/processors/interfaces/ImagePreprocessor.py
@@ -0,0 +1,18 @@
+# Use all imports relative to root directory
+from src.processors.manager import Processor
+
+
+class ImagePreprocessor(Processor):
+ """Base class for an extension that applies some preprocessing to the input image"""
+
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+
+ def apply_filter(self, image, filename):
+ """Apply filter to the image and returns modified image"""
+ raise NotImplementedError
+
+ @staticmethod
+ def exclude_files():
+ """Returns a list of file paths that should be excluded from processing"""
+ return []
diff --git a/OMRChecker/src/processors/manager.py b/OMRChecker/src/processors/manager.py
new file mode 100644
index 0000000000000000000000000000000000000000..18f4346c736513fea4d141d2489e37bf920003d4
--- /dev/null
+++ b/OMRChecker/src/processors/manager.py
@@ -0,0 +1,80 @@
+"""
+Processor/Extension framework
+Adapated from https://github.com/gdiepen/python_processor_example
+"""
+import inspect
+import pkgutil
+
+from src.logger import logger
+
+
+class Processor:
+ """Base class that each processor must inherit from."""
+
+ def __init__(
+ self,
+ options=None,
+ relative_dir=None,
+ image_instance_ops=None,
+ ):
+ self.options = options
+ self.relative_dir = relative_dir
+ self.image_instance_ops = image_instance_ops
+ self.tuning_config = image_instance_ops.tuning_config
+ self.description = "UNKNOWN"
+
+
+class ProcessorManager:
+ """Upon creation, this class will read the processors package for modules
+ that contain a class definition that is inheriting from the Processor class
+ """
+
+ def __init__(self, processors_dir="src.processors"):
+ """Constructor that initiates the reading of all available processors
+ when an instance of the ProcessorCollection object is created
+ """
+ self.processors_dir = processors_dir
+ self.reload_processors()
+
+ @staticmethod
+ def get_name_filter(processor_name):
+ def filter_function(member):
+ return inspect.isclass(member) and member.__module__ == processor_name
+
+ return filter_function
+
+ def reload_processors(self):
+ """Reset the list of all processors and initiate the walk over the main
+ provided processor package to load all available processors
+ """
+ self.processors = {}
+ self.seen_paths = []
+
+ logger.info(f'Loading processors from "{self.processors_dir}"...')
+ self.walk_package(self.processors_dir)
+
+ def walk_package(self, package):
+ """walk the supplied package to retrieve all processors"""
+ imported_package = __import__(package, fromlist=["blah"])
+ loaded_packages = []
+ for _, processor_name, ispkg in pkgutil.walk_packages(
+ imported_package.__path__, imported_package.__name__ + "."
+ ):
+ if not ispkg and processor_name != __name__:
+ processor_module = __import__(processor_name, fromlist=["blah"])
+ # https://stackoverflow.com/a/46206754/6242649
+ clsmembers = inspect.getmembers(
+ processor_module,
+ ProcessorManager.get_name_filter(processor_name),
+ )
+ for _, c in clsmembers:
+ # Only add classes that are a sub class of Processor, but NOT Processor itself
+ if issubclass(c, Processor) & (c is not Processor):
+ self.processors[c.__name__] = c
+ loaded_packages.append(c.__name__)
+
+ logger.info(f"Loaded processors: {loaded_packages}")
+
+
+# Singleton export
+PROCESSOR_MANAGER = ProcessorManager()
diff --git a/OMRChecker/src/schemas/__init__.py b/OMRChecker/src/schemas/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..e325daf6b22ed9b0e1c37a4d251cc58e040b431d
--- /dev/null
+++ b/OMRChecker/src/schemas/__init__.py
@@ -0,0 +1,18 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+from jsonschema import Draft202012Validator
+
+from src.schemas.config_schema import CONFIG_SCHEMA
+from src.schemas.evaluation_schema import EVALUATION_SCHEMA
+from src.schemas.template_schema import TEMPLATE_SCHEMA
+
+SCHEMA_JSONS = {
+ "config": CONFIG_SCHEMA,
+ "evaluation": EVALUATION_SCHEMA,
+ "template": TEMPLATE_SCHEMA,
+}
+
+SCHEMA_VALIDATORS = {
+ "config": Draft202012Validator(CONFIG_SCHEMA),
+ "evaluation": Draft202012Validator(EVALUATION_SCHEMA),
+ "template": Draft202012Validator(TEMPLATE_SCHEMA),
+}
diff --git a/OMRChecker/src/schemas/config_schema.py b/OMRChecker/src/schemas/config_schema.py
new file mode 100644
index 0000000000000000000000000000000000000000..ffb41f45df1373c5fd0ab3bbe8132e74ef77fc73
--- /dev/null
+++ b/OMRChecker/src/schemas/config_schema.py
@@ -0,0 +1,57 @@
+CONFIG_SCHEMA = {
+ "$schema": "https://json-schema.org/draft/2020-12/schema",
+ "$id": "https://github.com/Udayraj123/OMRChecker/tree/master/src/schemas/config-schema.json",
+ "title": "Config Schema",
+ "description": "OMRChecker config schema for custom tuning",
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "dimensions": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "display_height": {"type": "integer"},
+ "display_width": {"type": "integer"},
+ "processing_height": {"type": "integer"},
+ "processing_width": {"type": "integer"},
+ },
+ },
+ "threshold_params": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "GAMMA_LOW": {"type": "number", "minimum": 0, "maximum": 1},
+ "MIN_GAP": {"type": "integer", "minimum": 10, "maximum": 100},
+ "MIN_JUMP": {"type": "integer", "minimum": 10, "maximum": 100},
+ "CONFIDENT_SURPLUS": {"type": "integer", "minimum": 0, "maximum": 20},
+ "JUMP_DELTA": {"type": "integer", "minimum": 10, "maximum": 100},
+ "PAGE_TYPE_FOR_THRESHOLD": {
+ "enum": ["white", "black"],
+ "type": "string",
+ },
+ },
+ },
+ "alignment_params": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "auto_align": {"type": "boolean"},
+ "match_col": {"type": "integer", "minimum": 0, "maximum": 10},
+ "max_steps": {"type": "integer", "minimum": 1, "maximum": 100},
+ "stride": {"type": "integer", "minimum": 1, "maximum": 10},
+ "thickness": {"type": "integer", "minimum": 1, "maximum": 10},
+ },
+ },
+ "outputs": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "show_image_level": {"type": "integer", "minimum": 0, "maximum": 6},
+ "save_image_level": {"type": "integer", "minimum": 0, "maximum": 6},
+ "save_detections": {"type": "boolean"},
+ # This option moves multimarked files into a separate folder for manual checking, skipping evaluation
+ "filter_out_multimarked_files": {"type": "boolean"},
+ },
+ },
+ },
+}
diff --git a/OMRChecker/src/schemas/constants.py b/OMRChecker/src/schemas/constants.py
new file mode 100644
index 0000000000000000000000000000000000000000..34b9b24d3fac2f60ecbba2d20d8f039d729db11c
--- /dev/null
+++ b/OMRChecker/src/schemas/constants.py
@@ -0,0 +1,17 @@
+DEFAULT_SECTION_KEY = "DEFAULT"
+
+BONUS_SECTION_PREFIX = "BONUS"
+
+MARKING_VERDICT_TYPES = ["correct", "incorrect", "unmarked"]
+
+ARRAY_OF_STRINGS = {
+ "type": "array",
+ "items": {"type": "string"},
+}
+
+FIELD_STRING_TYPE = {
+ "type": "string",
+ "pattern": "^([^\\.]+|[^\\.\\d]+\\d+\\.{2,3}\\d+)$",
+}
+
+FIELD_STRING_REGEX_GROUPS = r"([^\.\d]+)(\d+)\.{2,3}(\d+)"
diff --git a/OMRChecker/src/schemas/evaluation_schema.py b/OMRChecker/src/schemas/evaluation_schema.py
new file mode 100644
index 0000000000000000000000000000000000000000..c51af58b849eaa3202815ca1c9358ec34839975b
--- /dev/null
+++ b/OMRChecker/src/schemas/evaluation_schema.py
@@ -0,0 +1,151 @@
+from src.schemas.constants import (
+ ARRAY_OF_STRINGS,
+ DEFAULT_SECTION_KEY,
+ FIELD_STRING_TYPE,
+)
+
+marking_score_regex = "-?(\\d+)(/(\\d+))?"
+
+marking_score = {
+ "oneOf": [
+ {"type": "string", "pattern": marking_score_regex},
+ {"type": "number"},
+ ]
+}
+
+marking_object_properties = {
+ "additionalProperties": False,
+ "required": ["correct", "incorrect", "unmarked"],
+ "type": "object",
+ "properties": {
+ # TODO: can support streak marking if we allow array of marking_scores here
+ "correct": marking_score,
+ "incorrect": marking_score,
+ "unmarked": marking_score,
+ },
+}
+
+EVALUATION_SCHEMA = {
+ "$schema": "https://json-schema.org/draft/2020-12/schema",
+ "$id": "https://github.com/Udayraj123/OMRChecker/tree/master/src/schemas/evaluation-schema.json",
+ "title": "Evaluation Schema",
+ "description": "OMRChecker evaluation schema i.e. the marking scheme",
+ "type": "object",
+ "additionalProperties": True,
+ "required": ["source_type", "options", "marking_schemes"],
+ "properties": {
+ "additionalProperties": False,
+ "source_type": {"type": "string", "enum": ["csv", "custom"]},
+ "options": {"type": "object"},
+ "marking_schemes": {
+ "type": "object",
+ "required": [DEFAULT_SECTION_KEY],
+ "patternProperties": {
+ f"^{DEFAULT_SECTION_KEY}$": marking_object_properties,
+ f"^(?!{DEFAULT_SECTION_KEY}$).*": {
+ "additionalProperties": False,
+ "required": ["marking", "questions"],
+ "type": "object",
+ "properties": {
+ "questions": {
+ "oneOf": [
+ FIELD_STRING_TYPE,
+ {
+ "type": "array",
+ "items": FIELD_STRING_TYPE,
+ },
+ ]
+ },
+ "marking": marking_object_properties,
+ },
+ },
+ },
+ },
+ },
+ "allOf": [
+ {
+ "if": {"properties": {"source_type": {"const": "csv"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "additionalProperties": False,
+ "required": ["answer_key_csv_path"],
+ "dependentRequired": {
+ "answer_key_image_path": [
+ "answer_key_csv_path",
+ "questions_in_order",
+ ]
+ },
+ "type": "object",
+ "properties": {
+ "should_explain_scoring": {"type": "boolean"},
+ "answer_key_csv_path": {"type": "string"},
+ "answer_key_image_path": {"type": "string"},
+ "questions_in_order": ARRAY_OF_STRINGS,
+ },
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"source_type": {"const": "custom"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "additionalProperties": False,
+ "required": ["answers_in_order", "questions_in_order"],
+ "type": "object",
+ "properties": {
+ "should_explain_scoring": {"type": "boolean"},
+ "answers_in_order": {
+ "oneOf": [
+ {
+ "type": "array",
+ "items": {
+ "oneOf": [
+ # "standard": single correct, multi-marked single-correct
+ # Example: "q1" --> '67'
+ {"type": "string"},
+ # "multiple-correct": multiple-correct (for ambiguous/bonus questions)
+ # Example: "q1" --> [ 'A', 'B' ]
+ {
+ "type": "array",
+ "items": {"type": "string"},
+ "minItems": 2,
+ },
+ # "multiple-correct-weighted": array of answer-wise weights (marking scheme not applicable)
+ # Example 1: "q1" --> [['A', 1], ['B', 2], ['C', 3]] or
+ # Example 2: "q2" --> [['A', 1], ['B', 1], ['AB', 2]]
+ {
+ "type": "array",
+ "items": {
+ "type": "array",
+ "items": False,
+ "minItems": 2,
+ "maxItems": 2,
+ "prefixItems": [
+ {"type": "string"},
+ marking_score,
+ ],
+ },
+ },
+ # Multiple-correct with custom marking scheme
+ # ["A", ["1", "2", "3"]],
+ # [["A", "B", "AB"], ["1", "2", "3"]]
+ ],
+ },
+ },
+ ]
+ },
+ "questions_in_order": ARRAY_OF_STRINGS,
+ "enable_evaluation_table_to_csv": {
+ "type": "boolean",
+ "default": False,
+ },
+ },
+ }
+ }
+ },
+ },
+ ],
+}
diff --git a/OMRChecker/src/schemas/template_schema.py b/OMRChecker/src/schemas/template_schema.py
new file mode 100644
index 0000000000000000000000000000000000000000..12b7ffdd8577054c03db25f12815d91813026c40
--- /dev/null
+++ b/OMRChecker/src/schemas/template_schema.py
@@ -0,0 +1,226 @@
+from src.constants import FIELD_TYPES
+from src.schemas.constants import ARRAY_OF_STRINGS, FIELD_STRING_TYPE
+
+positive_number = {"type": "number", "minimum": 0}
+positive_integer = {"type": "integer", "minimum": 0}
+two_positive_integers = {
+ "type": "array",
+ "prefixItems": [
+ positive_integer,
+ positive_integer,
+ ],
+ "maxItems": 2,
+ "minItems": 2,
+}
+two_positive_numbers = {
+ "type": "array",
+ "prefixItems": [
+ positive_number,
+ positive_number,
+ ],
+ "maxItems": 2,
+ "minItems": 2,
+}
+zero_to_one_number = {
+ "type": "number",
+ "minimum": 0,
+ "maximum": 1,
+}
+
+TEMPLATE_SCHEMA = {
+ "$schema": "https://json-schema.org/draft/2020-12/schema",
+ "$id": "https://github.com/Udayraj123/OMRChecker/tree/master/src/schemas/template-schema.json",
+ "title": "Template Validation Schema",
+ "description": "OMRChecker input template schema",
+ "type": "object",
+ "required": [
+ "bubbleDimensions",
+ "pageDimensions",
+ "preProcessors",
+ "fieldBlocks",
+ ],
+ "additionalProperties": False,
+ "properties": {
+ "bubbleDimensions": {
+ **two_positive_integers,
+ "description": "The dimensions of the overlay bubble area: [width, height]",
+ },
+ "customLabels": {
+ "description": "The customLabels contain fields that need to be joined together before generating the results sheet",
+ "type": "object",
+ "patternProperties": {
+ "^.*$": {"type": "array", "items": FIELD_STRING_TYPE}
+ },
+ },
+ "outputColumns": {
+ "type": "array",
+ "items": FIELD_STRING_TYPE,
+ "description": "The ordered list of columns to be contained in the output csv(default order: alphabetical)",
+ },
+ "pageDimensions": {
+ **two_positive_integers,
+ "description": "The dimensions(width, height) to which the page will be resized to before applying template",
+ },
+ "preProcessors": {
+ "description": "Custom configuration values to use in the template's directory",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "name": {
+ "type": "string",
+ "enum": [
+ "CropOnMarkers",
+ "CropPage",
+ "FeatureBasedAlignment",
+ "GaussianBlur",
+ "Levels",
+ "MedianBlur",
+ ],
+ },
+ },
+ "required": ["name", "options"],
+ "allOf": [
+ {
+ "if": {"properties": {"name": {"const": "CropOnMarkers"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "apply_erode_subtract": {"type": "boolean"},
+ "marker_rescale_range": two_positive_numbers,
+ "marker_rescale_steps": {"type": "number"},
+ "max_matching_variation": {"type": "number"},
+ "min_matching_threshold": {"type": "number"},
+ "relativePath": {"type": "string"},
+ "sheetToMarkerWidthRatio": {"type": "number"},
+ },
+ "required": ["relativePath"],
+ }
+ }
+ },
+ },
+ {
+ "if": {
+ "properties": {"name": {"const": "FeatureBasedAlignment"}}
+ },
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "2d": {"type": "boolean"},
+ "goodMatchPercent": {"type": "number"},
+ "maxFeatures": {"type": "integer"},
+ "reference": {"type": "string"},
+ },
+ "required": ["reference"],
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "Levels"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "gamma": zero_to_one_number,
+ "high": zero_to_one_number,
+ "low": zero_to_one_number,
+ },
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "MedianBlur"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {"kSize": {"type": "integer"}},
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "GaussianBlur"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "kSize": two_positive_integers,
+ "sigmaX": {"type": "number"},
+ },
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "CropPage"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "morphKernel": two_positive_integers
+ },
+ }
+ }
+ },
+ },
+ ],
+ },
+ },
+ "fieldBlocks": {
+ "description": "The fieldBlocks denote small groups of adjacent fields",
+ "type": "object",
+ "patternProperties": {
+ "^.*$": {
+ "type": "object",
+ "required": [
+ "origin",
+ "bubblesGap",
+ "labelsGap",
+ "fieldLabels",
+ ],
+ "oneOf": [
+ {"required": ["fieldType"]},
+ {"required": ["bubbleValues", "direction"]},
+ ],
+ "properties": {
+ "bubbleDimensions": two_positive_numbers,
+ "bubblesGap": positive_number,
+ "bubbleValues": ARRAY_OF_STRINGS,
+ "direction": {
+ "type": "string",
+ "enum": ["horizontal", "vertical"],
+ },
+ "emptyValue": {"type": "string"},
+ "fieldLabels": {"type": "array", "items": FIELD_STRING_TYPE},
+ "labelsGap": positive_number,
+ "origin": two_positive_integers,
+ "fieldType": {
+ "type": "string",
+ "enum": list(FIELD_TYPES.keys()),
+ },
+ },
+ }
+ },
+ },
+ "emptyValue": {
+ "description": "The value to be used in case of empty bubble detected at global level.",
+ "type": "string",
+ },
+ },
+}
diff --git a/OMRChecker/src/template.py b/OMRChecker/src/template.py
new file mode 100644
index 0000000000000000000000000000000000000000..1ab6c9173bb9bf237d8902eaef004835ed8a1446
--- /dev/null
+++ b/OMRChecker/src/template.py
@@ -0,0 +1,327 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+from src.constants import FIELD_TYPES
+from src.core import ImageInstanceOps
+from src.logger import logger
+from src.processors.manager import PROCESSOR_MANAGER
+from src.utils.parsing import (
+ custom_sort_output_columns,
+ open_template_with_defaults,
+ parse_fields,
+)
+
+
+class Template:
+ def __init__(self, template_path, tuning_config):
+ self.path = template_path
+ self.image_instance_ops = ImageInstanceOps(tuning_config)
+
+ json_object = open_template_with_defaults(template_path)
+ (
+ custom_labels_object,
+ field_blocks_object,
+ output_columns_array,
+ pre_processors_object,
+ self.bubble_dimensions,
+ self.global_empty_val,
+ self.options,
+ self.page_dimensions,
+ ) = map(
+ json_object.get,
+ [
+ "customLabels",
+ "fieldBlocks",
+ "outputColumns",
+ "preProcessors",
+ "bubbleDimensions",
+ "emptyValue",
+ "options",
+ "pageDimensions",
+ ],
+ )
+
+ self.parse_output_columns(output_columns_array)
+ self.setup_pre_processors(pre_processors_object, template_path.parent)
+ self.setup_field_blocks(field_blocks_object)
+ self.parse_custom_labels(custom_labels_object)
+
+ non_custom_columns, all_custom_columns = (
+ list(self.non_custom_labels),
+ list(custom_labels_object.keys()),
+ )
+
+ if len(self.output_columns) == 0:
+ self.fill_output_columns(non_custom_columns, all_custom_columns)
+
+ self.validate_template_columns(non_custom_columns, all_custom_columns)
+
+ def parse_output_columns(self, output_columns_array):
+ self.output_columns = parse_fields(f"Output Columns", output_columns_array)
+
+ def setup_pre_processors(self, pre_processors_object, relative_dir):
+ # load image pre_processors
+ self.pre_processors = []
+ for pre_processor in pre_processors_object:
+ ProcessorClass = PROCESSOR_MANAGER.processors[pre_processor["name"]]
+ pre_processor_instance = ProcessorClass(
+ options=pre_processor["options"],
+ relative_dir=relative_dir,
+ image_instance_ops=self.image_instance_ops,
+ )
+ self.pre_processors.append(pre_processor_instance)
+
+ def setup_field_blocks(self, field_blocks_object):
+ # Add field_blocks
+ self.field_blocks = []
+ self.all_parsed_labels = set()
+ for block_name, field_block_object in field_blocks_object.items():
+ self.parse_and_add_field_block(block_name, field_block_object)
+
+ def parse_custom_labels(self, custom_labels_object):
+ all_parsed_custom_labels = set()
+ self.custom_labels = {}
+ for custom_label, label_strings in custom_labels_object.items():
+ parsed_labels = parse_fields(f"Custom Label: {custom_label}", label_strings)
+ parsed_labels_set = set(parsed_labels)
+ self.custom_labels[custom_label] = parsed_labels
+
+ missing_custom_labels = sorted(
+ parsed_labels_set.difference(self.all_parsed_labels)
+ )
+ if len(missing_custom_labels) > 0:
+ logger.critical(
+ f"For '{custom_label}', Missing labels - {missing_custom_labels}"
+ )
+ raise Exception(
+ f"Missing field block label(s) in the given template for {missing_custom_labels} from '{custom_label}'"
+ )
+
+ if not all_parsed_custom_labels.isdisjoint(parsed_labels_set):
+ # Note: this can be made a warning, but it's a choice
+ logger.critical(
+ f"field strings overlap for labels: {label_strings} and existing custom labels: {all_parsed_custom_labels}"
+ )
+ raise Exception(
+ f"The field strings for custom label '{custom_label}' overlap with other existing custom labels"
+ )
+
+ all_parsed_custom_labels.update(parsed_labels)
+
+ self.non_custom_labels = self.all_parsed_labels.difference(
+ all_parsed_custom_labels
+ )
+
+ def fill_output_columns(self, non_custom_columns, all_custom_columns):
+ all_template_columns = non_custom_columns + all_custom_columns
+ # Typical case: sort alpha-numerical (natural sort)
+ self.output_columns = sorted(
+ all_template_columns, key=custom_sort_output_columns
+ )
+
+ def validate_template_columns(self, non_custom_columns, all_custom_columns):
+ output_columns_set = set(self.output_columns)
+ all_custom_columns_set = set(all_custom_columns)
+
+ missing_output_columns = sorted(
+ output_columns_set.difference(all_custom_columns_set).difference(
+ self.all_parsed_labels
+ )
+ )
+ if len(missing_output_columns) > 0:
+ logger.critical(f"Missing output columns: {missing_output_columns}")
+ raise Exception(
+ f"Some columns are missing in the field blocks for the given output columns"
+ )
+
+ all_template_columns_set = set(non_custom_columns + all_custom_columns)
+ missing_label_columns = sorted(
+ all_template_columns_set.difference(output_columns_set)
+ )
+ if len(missing_label_columns) > 0:
+ logger.warning(
+ f"Some label columns are not covered in the given output columns: {missing_label_columns}"
+ )
+
+ def parse_and_add_field_block(self, block_name, field_block_object):
+ field_block_object = self.pre_fill_field_block(field_block_object)
+ block_instance = FieldBlock(block_name, field_block_object)
+ self.field_blocks.append(block_instance)
+ self.validate_parsed_labels(field_block_object["fieldLabels"], block_instance)
+
+ def pre_fill_field_block(self, field_block_object):
+ if "fieldType" in field_block_object:
+ field_block_object = {
+ **field_block_object,
+ **FIELD_TYPES[field_block_object["fieldType"]],
+ }
+ else:
+ field_block_object = {**field_block_object, "fieldType": "__CUSTOM__"}
+
+ return {
+ "direction": "vertical",
+ "emptyValue": self.global_empty_val,
+ "bubbleDimensions": self.bubble_dimensions,
+ **field_block_object,
+ }
+
+ def validate_parsed_labels(self, field_labels, block_instance):
+ parsed_field_labels, block_name = (
+ block_instance.parsed_field_labels,
+ block_instance.name,
+ )
+ field_labels_set = set(parsed_field_labels)
+ if not self.all_parsed_labels.isdisjoint(field_labels_set):
+ # Note: in case of two fields pointing to same column, use a custom column instead of same field labels.
+ logger.critical(
+ f"An overlap found between field string: {field_labels} in block '{block_name}' and existing labels: {self.all_parsed_labels}"
+ )
+ raise Exception(
+ f"The field strings for field block {block_name} overlap with other existing fields"
+ )
+ self.all_parsed_labels.update(field_labels_set)
+
+ page_width, page_height = self.page_dimensions
+ block_width, block_height = block_instance.dimensions
+ [block_start_x, block_start_y] = block_instance.origin
+
+ block_end_x, block_end_y = (
+ block_start_x + block_width,
+ block_start_y + block_height,
+ )
+
+ if (
+ block_end_x >= page_width
+ or block_end_y >= page_height
+ or block_start_x < 0
+ or block_start_y < 0
+ ):
+ raise Exception(
+ f"Overflowing field block '{block_name}' with origin {block_instance.origin} and dimensions {block_instance.dimensions} in template with dimensions {self.page_dimensions}"
+ )
+
+ def __str__(self):
+ return str(self.path)
+
+
+class FieldBlock:
+ def __init__(self, block_name, field_block_object):
+ self.name = block_name
+ self.shift = 0
+ self.setup_field_block(field_block_object)
+
+ def setup_field_block(self, field_block_object):
+ # case mapping
+ (
+ bubble_dimensions,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ field_labels,
+ field_type,
+ labels_gap,
+ origin,
+ self.empty_val,
+ ) = map(
+ field_block_object.get,
+ [
+ "bubbleDimensions",
+ "bubbleValues",
+ "bubblesGap",
+ "direction",
+ "fieldLabels",
+ "fieldType",
+ "labelsGap",
+ "origin",
+ "emptyValue",
+ ],
+ )
+ self.parsed_field_labels = parse_fields(
+ f"Field Block Labels: {self.name}", field_labels
+ )
+ self.origin = origin
+ self.bubble_dimensions = bubble_dimensions
+ self.calculate_block_dimensions(
+ bubble_dimensions,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ labels_gap,
+ )
+ self.generate_bubble_grid(
+ bubble_values,
+ bubbles_gap,
+ direction,
+ field_type,
+ labels_gap,
+ )
+
+ def calculate_block_dimensions(
+ self,
+ bubble_dimensions,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ labels_gap,
+ ):
+ _h, _v = (1, 0) if (direction == "vertical") else (0, 1)
+
+ values_dimension = int(
+ bubbles_gap * (len(bubble_values) - 1) + bubble_dimensions[_h]
+ )
+ fields_dimension = int(
+ labels_gap * (len(self.parsed_field_labels) - 1) + bubble_dimensions[_v]
+ )
+ self.dimensions = (
+ [fields_dimension, values_dimension]
+ if (direction == "vertical")
+ else [values_dimension, fields_dimension]
+ )
+
+ def generate_bubble_grid(
+ self,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ field_type,
+ labels_gap,
+ ):
+ _h, _v = (1, 0) if (direction == "vertical") else (0, 1)
+ self.traverse_bubbles = []
+ # Generate the bubble grid
+ lead_point = [float(self.origin[0]), float(self.origin[1])]
+ for field_label in self.parsed_field_labels:
+ bubble_point = lead_point.copy()
+ field_bubbles = []
+ for bubble_value in bubble_values:
+ field_bubbles.append(
+ Bubble(bubble_point.copy(), field_label, field_type, bubble_value)
+ )
+ bubble_point[_h] += bubbles_gap
+ self.traverse_bubbles.append(field_bubbles)
+ lead_point[_v] += labels_gap
+
+
+class Bubble:
+ """
+ Container for a Point Box on the OMR
+
+ field_label is the point's property- field to which this point belongs to
+ It can be used as a roll number column as well. (eg roll1)
+ It can also correspond to a single digit of integer type Q (eg q5d1)
+ """
+
+ def __init__(self, pt, field_label, field_type, field_value):
+ self.x = round(pt[0])
+ self.y = round(pt[1])
+ self.field_label = field_label
+ self.field_type = field_type
+ self.field_value = field_value
+
+ def __str__(self):
+ return str([self.x, self.y])
diff --git a/OMRChecker/src/tests/__init__.py b/OMRChecker/src/tests/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..28a0acd8198648687d60beb6d8dded390f95a29a
--- /dev/null
+++ b/OMRChecker/src/tests/__init__.py
@@ -0,0 +1 @@
+# https://stackoverflow.com/a/50169991/6242649
diff --git a/OMRChecker/src/tests/__snapshots__/test_all_samples.ambr b/OMRChecker/src/tests/__snapshots__/test_all_samples.ambr
new file mode 100644
index 0000000000000000000000000000000000000000..45df1372201a0679ded211fbc424716db37f7632
--- /dev/null
+++ b/OMRChecker/src/tests/__snapshots__/test_all_samples.ambr
@@ -0,0 +1,302 @@
+# serializer version: 1
+# name: test_run_answer_key_using_csv
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "adrian_omr.png","samples/answer-key/using-csv/adrian_omr.png","outputs/answer-key/using-csv/CheckedOMRs/adrian_omr.png","5.0","C","E","A","B","B"
+
+ ''',
+ })
+# ---
+# name: test_run_answer_key_weighted_answers
+ dict({
+ 'images/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'images/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'images/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "adrian_omr.png","samples/answer-key/weighted-answers/images/adrian_omr.png","outputs/answer-key/weighted-answers/images/CheckedOMRs/adrian_omr.png","5.5","B","E","A","C","B"
+ "adrian_omr_2.png","samples/answer-key/weighted-answers/images/adrian_omr_2.png","outputs/answer-key/weighted-answers/images/CheckedOMRs/adrian_omr_2.png","10.0","C","E","A","B","B"
+
+ ''',
+ })
+# ---
+# name: test_run_community_Antibodyy
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6"
+ "simple_omr_sheet.jpg","samples/community/Antibodyy/simple_omr_sheet.jpg","outputs/community/Antibodyy/CheckedOMRs/simple_omr_sheet.jpg","0","A","C","B","D","E","B"
+
+ ''',
+ })
+# ---
+# name: test_run_community_Sandeep_1507
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Booklet_No","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Booklet_No","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Booklet_No","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+ "omr-1.png","samples/community/Sandeep-1507/omr-1.png","outputs/community/Sandeep-1507/CheckedOMRs/omr-1.png","0","0190880","D","C","B","A","A","B","C","D","D","C","B","A","D","A","B","C","D","","B","D","C","A","C","C","B","A","D","A","AC","C","B","D","C","B","A","B","B","D","D","A","C","B","D","A","C","B","D","B","D","A","A","B","C","D","C","B","A","D","D","A","B","C","D","C","B","A","B","C","D","A","B","C","D","B","A","C","D","C","B","A","D","B","D","A","A","B","A","C","B","D","C","D","B","A","C","C","B","D","B","C","B","A","D","C","B","A","B","C","D","A","A","A","B","B","A","B","C","D","A","A","D","C","B","A","","A","B","C","D","D","D","B","B","C","C","D","C","C","D","D","C","C","B","B","A","A","D","D","B","A","D","C","B","A","A","D","D","B","B","A","A","B","C","D","D","C","B","A","B","D","A","C","C","C","A","A","B","B","D","D","A","A","B","C","D","B","D","A","B","C","D","AD","C","D","B","C","A","B","C","D"
+ "omr-2.png","samples/community/Sandeep-1507/omr-2.png","outputs/community/Sandeep-1507/CheckedOMRs/omr-2.png","0","0no22nonono","A","B","B","A","D","C","B","D","C","D","D","D","B","B","D","D","D","B","C","C","A","A","B","A","D","A","A","B","A","C","A","C","D","D","D","","","C","C","B","B","B","","D","","C","D","","D","B","A","D","B","A","C","A","C","A","C","B","A","D","C","B","C","B","C","D","B","B","D","C","C","D","D","A","D","A","D","C","B","D","C","A","C","","C","B","B","","A","A","D","","B","A","","C","A","D","D","C","C","A","C","A","C","D","A","A","A","D","D","B","C","B","B","B","D","A","C","D","D","A","A","A","C","D","C","C","B","D","A","A","C","B","","D","A","C","C","C","","","","A","C","","D","A","B","A","A","C","A","D","B","B","A","D","A","B","C","A","C","D","D","D","C","A","C","A","C","D","A","A","A","D","A","B","A","B","C","B","A","","B","C","D","D","","","D","C","C","C","","C","A",""
+ "omr-3.png","samples/community/Sandeep-1507/omr-3.png","outputs/community/Sandeep-1507/CheckedOMRs/omr-3.png","0","0nononono73","B","A","C","D","A","D","D","A","C","A","A","B","C","A","A","C","A","B","A","D","C","C","A","D","D","C","C","C","A","C","C","B","B","D","D","C","","","C","B","","","D","A","A","A","A","","A","C","C","C","D","C","","A","B","C","D","B","C","C","C","D","A","B","B","B","D","D","B","B","C","D","B","D","A","B","A","B","C","A","C","A","C","D","","","A","B","","B","C","D","A","D","D","","","C","D","B","B","A","A","D","D","B","A","B","B","C","C","D","D","C","A","D","C","D","C","C","B","C","D","C","D","A","B","D","C","B","D","B","B","","D","","B","D","B","B","C","A","D","","C","","C","","B","C","A","B","B","D","D","D","B","A","D","D","A","D","D","C","B","B","D","C","B","A","C","D","A","D","D","A","C","A","B","D","C","C","C","A","D","","","B","B","","C","C","B","B","C","","","B"
+
+ ''',
+ })
+# ---
+# name: test_run_community_Shamanth
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q21","q22","q23","q24","q25","q26","q27","q28"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q21","q22","q23","q24","q25","q26","q27","q28"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q21","q22","q23","q24","q25","q26","q27","q28"
+ "omr_sheet_01.png","samples/community/Shamanth/omr_sheet_01.png","outputs/community/Shamanth/CheckedOMRs/omr_sheet_01.png","0","A","B","C","D","A","C","C","D"
+
+ ''',
+ })
+# ---
+# name: test_run_community_UPSC_mock
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "answer_key.jpg","samples/community/UPSC-mock/answer_key.jpg","outputs/community/UPSC-mock/CheckedOMRs/answer_key.jpg","200.0","","","","C","D","A","C","C","C","B","A","C","C","B","D","B","D","C","C","B","D","B","D","C","C","C","B","D","D","D","B","A","D","D","C","A","B","C","A","D","A","A","A","D","D","B","A","B","C","B","A","C","D","C","D","A","B","C","A","C","C","C","D","B","C","C","C","C","A","D","A","D","A","D","C","C","D","C","D","A","A","C","B","C","D","C","A","B","C","B","D","A","A","C","A","B","D","C","D","A","C","B","A"
+
+ ''',
+ 'scan-angles/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'scan-angles/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'scan-angles/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "angle-1.jpg","samples/community/UPSC-mock/scan-angles/angle-1.jpg","outputs/community/UPSC-mock/scan-angles/CheckedOMRs/angle-1.jpg","70.66666666666669","","","","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+ "angle-2.jpg","samples/community/UPSC-mock/scan-angles/angle-2.jpg","outputs/community/UPSC-mock/scan-angles/CheckedOMRs/angle-2.jpg","70.66666666666669","","","","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+ "angle-3.jpg","samples/community/UPSC-mock/scan-angles/angle-3.jpg","outputs/community/UPSC-mock/scan-angles/CheckedOMRs/angle-3.jpg","70.66666666666669","","","","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+
+ ''',
+ })
+# ---
+# name: test_run_community_UmarFarootAPS
+ dict({
+ 'scans/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll_no","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'scans/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll_no","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'scans/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll_no","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+ "scan-type-1.jpg","samples/community/UmarFarootAPS/scans/scan-type-1.jpg","outputs/community/UmarFarootAPS/scans/CheckedOMRs/scan-type-1.jpg","49.0","2468","A","C","B","C","A","D","B","C","B","D","C","A","C","D","B","C","A","B","C","A","C","B","D","C","A","B","D","C","A","C","B","D","B","A","C","D","B","C","A","C","D","A","C","D","A","B","D","C","A","C","D","B","C","A","C","D","B","C","D","A","B","C","B","C","D","B","D","A","C","B","D","A","B","C","B","A","C","D","B","A","C","B","C","B","A","D","B","A","C","D","B","D","B","C","B","D","A","C","B","C","B","C","D","B","C","A","B","C","A","D","C","B","D","B","A","B","C","D","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","B","A","C","B","A","C","A","B","C","B","C","B","A","C","A","C","B","B","C","B","A","C","A","B","A","B","A","B","C","D","B","C","A","C","D","C","A","C","B","A","C","A","B","C","B","D","A","B","C","D","C","B","B","C","A","B","C","B"
+ "scan-type-2.jpg","samples/community/UmarFarootAPS/scans/scan-type-2.jpg","outputs/community/UmarFarootAPS/scans/CheckedOMRs/scan-type-2.jpg","20.0","0234","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","A","D","","","AD","","","","A","D","","","","","","","D","A","","D","","A","","D","","","","A","","","C","","","D","","","A","","","","D","","C","","A","","C","","D","B","B","","","A","","D","","","","D","","","","","A","D","","","B","","","D","","","A","","","D","","","","","","D","","","","A","D","","","A","","B","","D","","","","C","C","D","D","A","","D","","A","D","","","D","","B","D","","","D","","D","B","","","","D","","A","","","","D","","B","","","","","","D","","","A","","","A","","D","","","D"
+
+ ''',
+ })
+# ---
+# name: test_run_community_ibrahimkilic
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "yes_no_questionnarie.jpg","samples/community/ibrahimkilic/yes_no_questionnarie.jpg","outputs/community/ibrahimkilic/CheckedOMRs/yes_no_questionnarie.jpg","0","no","no","no","no","no"
+
+ ''',
+ })
+# ---
+# name: test_run_sample1
+ dict({
+ 'MobileCamera/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+
+ ''',
+ 'MobileCamera/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+
+ ''',
+ 'MobileCamera/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+ "sheet1.jpg","samples/sample1/MobileCamera/sheet1.jpg","outputs/sample1/MobileCamera/CheckedOMRs/sheet1.jpg","0","E503110026","B","","D","B","6","11","20","7","16","B","D","C","D","A","D","B","A","C","C","D"
+
+ ''',
+ })
+# ---
+# name: test_run_sample2
+ dict({
+ 'AdrianSample/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'AdrianSample/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'AdrianSample/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "adrian_omr.png","samples/sample2/AdrianSample/adrian_omr.png","outputs/sample2/AdrianSample/CheckedOMRs/adrian_omr.png","0","B","E","A","C","B"
+ "adrian_omr_2.png","samples/sample2/AdrianSample/adrian_omr_2.png","outputs/sample2/AdrianSample/CheckedOMRs/adrian_omr_2.png","0","C","E","A","B","B"
+
+ ''',
+ })
+# ---
+# name: test_run_sample3
+ dict({
+ 'colored-thick-sheet/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'colored-thick-sheet/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'colored-thick-sheet/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "rgb-100-gsm.jpg","samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg","outputs/sample3/colored-thick-sheet/CheckedOMRs/rgb-100-gsm.jpg","0","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+
+ ''',
+ 'xeroxed-thin-sheet/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'xeroxed-thin-sheet/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'xeroxed-thin-sheet/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "grayscale-80-gsm.jpg","samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg","outputs/sample3/xeroxed-thin-sheet/CheckedOMRs/grayscale-80-gsm.jpg","0","C","D","A","C","C","C","B","A","C","C","B","D","B","D","C","C","B","D","B","D","C","C","C","B","D","D","D","B","A","D","D","C","A","B","C","A","D","A","A","A","D","D","B","A","B","C","B","A","C","D","C","D","A","B","C","A","C","C","C","D","B","C","C","C","C","A","D","A","D","A","D","C","C","D","C","D","A","A","C","B","C","D","C","A","B","C","B","D","A","A","C","A","B","D","C","D","A","C","B","A"
+
+ ''',
+ })
+# ---
+# name: test_run_sample4
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11"
+ "IMG_20201116_143512.jpg","samples/sample4/IMG_20201116_143512.jpg","outputs/sample4/CheckedOMRs/IMG_20201116_143512.jpg","33.0","B","D","C","B","D","C","BC","A","C","D","C"
+ "IMG_20201116_150717658.jpg","samples/sample4/IMG_20201116_150717658.jpg","outputs/sample4/CheckedOMRs/IMG_20201116_150717658.jpg","33.0","B","D","C","B","D","C","BC","A","C","D","C"
+ "IMG_20201116_150750830.jpg","samples/sample4/IMG_20201116_150750830.jpg","outputs/sample4/CheckedOMRs/IMG_20201116_150750830.jpg","-2.0","A","","D","C","AC","A","D","B","C","D","D"
+
+ ''',
+ })
+# ---
+# name: test_run_sample5
+ dict({
+ 'ScanBatch1/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch1/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch1/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+ "camscanner-1.jpg","samples/sample5/ScanBatch1/camscanner-1.jpg","outputs/sample5/ScanBatch1/CheckedOMRs/camscanner-1.jpg","-4.0","E204420102","D","C","A","C","B","08","52","21","85","36","B","C","A","A","D","C","C","AD","A","A","D",""
+
+ ''',
+ 'ScanBatch2/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch2/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch2/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+ "camscanner-2.jpg","samples/sample5/ScanBatch2/camscanner-2.jpg","outputs/sample5/ScanBatch2/CheckedOMRs/camscanner-2.jpg","55.0","E204420109","C","C","B","C","C","01","19","10","10","18","D","A","D","D","D","C","C","C","C","D","B","A"
+
+ ''',
+ })
+# ---
+# name: test_run_sample6
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+ "reference.png","samples/sample6/reference.png","outputs/sample6/CheckedOMRs/reference.png","0","A"
+
+ ''',
+ 'doc-scans/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'doc-scans/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'doc-scans/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+ "sample_roll_01.jpg","samples/sample6/doc-scans/sample_roll_01.jpg","outputs/sample6/doc-scans/CheckedOMRs/sample_roll_01.jpg","0","A0188877Y"
+ "sample_roll_02.jpg","samples/sample6/doc-scans/sample_roll_02.jpg","outputs/sample6/doc-scans/CheckedOMRs/sample_roll_02.jpg","0","A0203959W"
+ "sample_roll_03.jpg","samples/sample6/doc-scans/sample_roll_03.jpg","outputs/sample6/doc-scans/CheckedOMRs/sample_roll_03.jpg","0","A0204729A"
+
+ ''',
+ })
+# ---
diff --git a/OMRChecker/src/tests/test_all_samples.py b/OMRChecker/src/tests/test_all_samples.py
new file mode 100644
index 0000000000000000000000000000000000000000..79095bf5e96da15680d667ae8216bb801864943e
--- /dev/null
+++ b/OMRChecker/src/tests/test_all_samples.py
@@ -0,0 +1,112 @@
+import os
+import shutil
+from glob import glob
+
+from src.tests.utils import run_entry_point, setup_mocker_patches
+
+
+def read_file(path):
+ with open(path) as file:
+ return file.read()
+
+
+def run_sample(mocker, sample_path):
+ setup_mocker_patches(mocker)
+
+ input_path = os.path.join("samples", sample_path)
+ output_dir = os.path.join("outputs", sample_path)
+ if os.path.exists(output_dir):
+ print(
+ f"Warning: output directory already exists: {output_dir}. This may affect the test execution."
+ )
+
+ run_entry_point(input_path, output_dir)
+
+ sample_outputs = extract_sample_outputs(output_dir)
+
+ print(f"Note: removing output directory: {output_dir}")
+ shutil.rmtree(output_dir)
+
+ return sample_outputs
+
+
+EXT = "*.csv"
+
+
+def extract_sample_outputs(output_dir):
+ sample_outputs = {}
+ for _dir, _subdir, _files in os.walk(output_dir):
+ for file in glob(os.path.join(_dir, EXT)):
+ relative_path = os.path.relpath(file, output_dir)
+ sample_outputs[relative_path] = read_file(file)
+ return sample_outputs
+
+
+def test_run_answer_key_using_csv(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "answer-key/using-csv")
+ assert snapshot == sample_outputs
+
+
+def test_run_answer_key_weighted_answers(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "answer-key/weighted-answers")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample1(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample1")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample2(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample2")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample3(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample3")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample4(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample4")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample5(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample5")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample6(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample6")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_Antibodyy(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/Antibodyy")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_ibrahimkilic(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/ibrahimkilic")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_Sandeep_1507(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/Sandeep-1507")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_Shamanth(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/Shamanth")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_UmarFarootAPS(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/UmarFarootAPS")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_UPSC_mock(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/UPSC-mock")
+ assert snapshot == sample_outputs
diff --git a/OMRChecker/src/tests/test_edge_cases.py b/OMRChecker/src/tests/test_edge_cases.py
new file mode 100644
index 0000000000000000000000000000000000000000..155d37223006680928dbbbd75054fcbfcc2fb33a
--- /dev/null
+++ b/OMRChecker/src/tests/test_edge_cases.py
@@ -0,0 +1,95 @@
+import os
+from pathlib import Path
+
+import pandas as pd
+
+from src.tests.test_samples.sample2.boilerplate import (
+ CONFIG_BOILERPLATE,
+ TEMPLATE_BOILERPLATE,
+)
+from src.tests.utils import (
+ generate_write_jsons_and_run,
+ remove_file,
+ run_entry_point,
+ setup_mocker_patches,
+)
+
+FROZEN_TIMESTAMP = "1970-01-01"
+CURRENT_DIR = Path("src/tests")
+BASE_SAMPLE_PATH = CURRENT_DIR.joinpath("test_samples", "sample2")
+BASE_RESULTS_CSV_PATH = os.path.join(
+ "outputs", BASE_SAMPLE_PATH, "Results", "Results_05AM.csv"
+)
+BASE_MULTIMARKED_CSV_PATH = os.path.join(
+ "outputs", BASE_SAMPLE_PATH, "Manual", "MultiMarkedFiles.csv"
+)
+
+
+def run_sample(mocker, input_path):
+ setup_mocker_patches(mocker)
+ output_dir = os.path.join("outputs", input_path)
+ run_entry_point(input_path, output_dir)
+
+
+def extract_output_data(path):
+ output_data = pd.read_csv(path, keep_default_na=False)
+ return output_data
+
+
+write_jsons_and_run = generate_write_jsons_and_run(
+ run_sample,
+ sample_path=BASE_SAMPLE_PATH,
+ template_boilerplate=TEMPLATE_BOILERPLATE,
+ config_boilerplate=CONFIG_BOILERPLATE,
+)
+
+
+def test_config_low_dimensions(mocker):
+ def modify_config(config):
+ config["dimensions"]["processing_height"] = 1000
+ config["dimensions"]["processing_width"] = 1000
+
+ exception = write_jsons_and_run(mocker, modify_config=modify_config)
+
+ assert str(exception) == "No Error"
+
+
+def test_different_bubble_dimensions(mocker):
+ # Prevent appending to output csv:
+ remove_file(BASE_RESULTS_CSV_PATH)
+ remove_file(BASE_MULTIMARKED_CSV_PATH)
+
+ exception = write_jsons_and_run(mocker)
+ assert str(exception) == "No Error"
+ original_output_data = extract_output_data(BASE_RESULTS_CSV_PATH)
+
+ def modify_template(template):
+ # Incorrect global bubble size
+ template["bubbleDimensions"] = [5, 5]
+ # Correct bubble size for MCQBlock1a1
+ template["fieldBlocks"]["MCQBlock1a1"]["bubbleDimensions"] = [32, 32]
+ # Incorrect bubble size for MCQBlock1a11
+ template["fieldBlocks"]["MCQBlock1a11"]["bubbleDimensions"] = [10, 10]
+
+ remove_file(BASE_RESULTS_CSV_PATH)
+ remove_file(BASE_MULTIMARKED_CSV_PATH)
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == "No Error"
+
+ results_output_data = extract_output_data(BASE_RESULTS_CSV_PATH)
+
+ assert results_output_data.empty
+
+ output_data = extract_output_data(BASE_MULTIMARKED_CSV_PATH)
+
+ equal_columns = [f"q{i}" for i in range(1, 18)]
+ assert (
+ output_data[equal_columns].iloc[0].to_list()
+ == original_output_data[equal_columns].iloc[0].to_list()
+ )
+
+ unequal_columns = [f"q{i}" for i in range(168, 185)]
+ assert not (
+ output_data[unequal_columns].iloc[0].to_list()
+ == original_output_data[unequal_columns].iloc[0].to_list()
+ )
diff --git a/OMRChecker/src/tests/test_samples/sample1/boilerplate.py b/OMRChecker/src/tests/test_samples/sample1/boilerplate.py
new file mode 100644
index 0000000000000000000000000000000000000000..f35c77c1f7081925edbbf8a207b933d45916e7f0
--- /dev/null
+++ b/OMRChecker/src/tests/test_samples/sample1/boilerplate.py
@@ -0,0 +1,14 @@
+TEMPLATE_BOILERPLATE = {
+ "pageDimensions": [300, 400],
+ "bubbleDimensions": [25, 25],
+ "preProcessors": [{"name": "CropPage", "options": {"morphKernel": [10, 10]}}],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [65, 60],
+ "fieldLabels": ["q1..5"],
+ "labelsGap": 52,
+ "bubblesGap": 41,
+ }
+ },
+}
diff --git a/OMRChecker/src/tests/test_samples/sample1/sample.png b/OMRChecker/src/tests/test_samples/sample1/sample.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/src/tests/test_samples/sample1/sample.png differ
diff --git a/OMRChecker/src/tests/test_samples/sample2/boilerplate.py b/OMRChecker/src/tests/test_samples/sample2/boilerplate.py
new file mode 100644
index 0000000000000000000000000000000000000000..22ddfb3a061a96191533c436f0f626401dc7fdfc
--- /dev/null
+++ b/OMRChecker/src/tests/test_samples/sample2/boilerplate.py
@@ -0,0 +1,39 @@
+TEMPLATE_BOILERPLATE = {
+ "pageDimensions": [2550, 3300],
+ "bubbleDimensions": [32, 32],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17,
+ },
+ }
+ ],
+ "fieldBlocks": {
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [197, 300],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": ["q1..17"],
+ },
+ "MCQBlock1a11": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [1770, 1310],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": ["q168..184"],
+ },
+ },
+}
+
+CONFIG_BOILERPLATE = {
+ "dimensions": {
+ "display_height": 960,
+ "display_width": 1280,
+ "processing_height": 1640,
+ "processing_width": 1332,
+ },
+ "outputs": {"show_image_level": 0, "filter_out_multimarked_files": True},
+}
diff --git a/OMRChecker/src/tests/test_samples/sample2/omr_marker.jpg b/OMRChecker/src/tests/test_samples/sample2/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/src/tests/test_samples/sample2/omr_marker.jpg differ
diff --git a/OMRChecker/src/tests/test_samples/sample2/sample.jpg b/OMRChecker/src/tests/test_samples/sample2/sample.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..25d75107cf4885f2974c52cf42003255be285a3e
--- /dev/null
+++ b/OMRChecker/src/tests/test_samples/sample2/sample.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18b85df487c0bc98eb9139949c3fbca2bad57e8f224fe48b495e0951660f65d
+size 237392
diff --git a/OMRChecker/src/tests/test_template_validations.py b/OMRChecker/src/tests/test_template_validations.py
new file mode 100644
index 0000000000000000000000000000000000000000..6a4a84f52469b89a3ac0e1f5f24b9944040ede5f
--- /dev/null
+++ b/OMRChecker/src/tests/test_template_validations.py
@@ -0,0 +1,159 @@
+import os
+from pathlib import Path
+
+from src.tests.test_samples.sample1.boilerplate import TEMPLATE_BOILERPLATE
+from src.tests.utils import (
+ generate_write_jsons_and_run,
+ run_entry_point,
+ setup_mocker_patches,
+)
+
+FROZEN_TIMESTAMP = "1970-01-01"
+CURRENT_DIR = Path("src/tests")
+BASE_SAMPLE_PATH = CURRENT_DIR.joinpath("test_samples", "sample1")
+BASE_SAMPLE_TEMPLATE_PATH = BASE_SAMPLE_PATH.joinpath("template.json")
+
+
+def run_sample(mocker, input_path):
+ setup_mocker_patches(mocker)
+ output_dir = os.path.join("outputs", input_path)
+ run_entry_point(input_path, output_dir)
+
+
+write_jsons_and_run = generate_write_jsons_and_run(
+ run_sample,
+ sample_path=BASE_SAMPLE_PATH,
+ template_boilerplate=TEMPLATE_BOILERPLATE,
+)
+
+
+def test_no_input_dir(mocker):
+ try:
+ run_sample(mocker, "X")
+ except Exception as e:
+ assert str(e) == "Given input directory does not exist: 'X'"
+
+
+def test_no_template(mocker):
+ if os.path.exists(BASE_SAMPLE_TEMPLATE_PATH):
+ os.remove(BASE_SAMPLE_TEMPLATE_PATH)
+ try:
+ run_sample(mocker, BASE_SAMPLE_PATH)
+ except Exception as e:
+ assert (
+ str(e)
+ == "No template file found in the directory tree of src/tests/test_samples/sample1"
+ )
+
+
+def test_empty_template(mocker):
+ def modify_template(_):
+ return {}
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == f"Provided Template JSON is Invalid: '{BASE_SAMPLE_TEMPLATE_PATH}'"
+ )
+
+
+def test_invalid_field_type(mocker):
+ def modify_template(template):
+ template["fieldBlocks"]["MCQ_Block_1"]["fieldType"] = "X"
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == f"Provided Template JSON is Invalid: '{BASE_SAMPLE_TEMPLATE_PATH}'"
+ )
+
+
+def test_overflow_labels(mocker):
+ def modify_template(template):
+ template["fieldBlocks"]["MCQ_Block_1"]["fieldLabels"] = ["q1..100"]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "Overflowing field block 'MCQ_Block_1' with origin [65, 60] and dimensions [189, 5173] in template with dimensions [300, 400]"
+ )
+
+
+def test_overflow_safe_dimensions(mocker):
+ def modify_template(template):
+ template["pageDimensions"] = [255, 400]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == "No Error"
+
+
+def test_field_strings_overlap(mocker):
+ def modify_template(template):
+ template["fieldBlocks"] = {
+ **template["fieldBlocks"],
+ "New_Block": {
+ **template["fieldBlocks"]["MCQ_Block_1"],
+ "fieldLabels": ["q5"],
+ },
+ }
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == (
+ "The field strings for field block New_Block overlap with other existing fields"
+ )
+
+
+def test_custom_label_strings_overlap_single(mocker):
+ def modify_template(template):
+ template["customLabels"] = {
+ "label1": ["q1..2", "q2..3"],
+ }
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "Given field string 'q2..3' has overlapping field(s) with other fields in 'Custom Label: label1': ['q1..2', 'q2..3']"
+ )
+
+
+def test_custom_label_strings_overlap_multiple(mocker):
+ def modify_template(template):
+ template["customLabels"] = {
+ "label1": ["q1..2"],
+ "label2": ["q2..3"],
+ }
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "The field strings for custom label 'label2' overlap with other existing custom labels"
+ )
+
+
+def test_missing_field_block_labels(mocker):
+ def modify_template(template):
+ template["customLabels"] = {"Combined": ["qX", "qY"]}
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "Missing field block label(s) in the given template for ['qX', 'qY'] from 'Combined'"
+ )
+
+
+def test_missing_output_columns(mocker):
+ def modify_template(template):
+ template["outputColumns"] = ["qX", "q1..5"]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == (
+ "Some columns are missing in the field blocks for the given output columns"
+ )
+
+
+def test_safe_missing_label_columns(mocker):
+ def modify_template(template):
+ template["outputColumns"] = ["q1..4"]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == "No Error"
diff --git a/OMRChecker/src/tests/utils.py b/OMRChecker/src/tests/utils.py
new file mode 100644
index 0000000000000000000000000000000000000000..2b24215e319a6c35463662e1951982deb89cb910
--- /dev/null
+++ b/OMRChecker/src/tests/utils.py
@@ -0,0 +1,97 @@
+import json
+import os
+from copy import deepcopy
+
+from freezegun import freeze_time
+
+from main import entry_point_for_args
+
+FROZEN_TIMESTAMP = "1970-01-01"
+
+
+def setup_mocker_patches(mocker):
+ mock_imshow = mocker.patch("cv2.imshow")
+ mock_imshow.return_value = True
+
+ mock_destroy_all_windows = mocker.patch("cv2.destroyAllWindows")
+ mock_destroy_all_windows.return_value = True
+
+ mock_wait_key = mocker.patch("cv2.waitKey")
+ mock_wait_key.return_value = ord("q")
+
+
+def run_entry_point(input_path, output_dir):
+ args = {
+ "autoAlign": False,
+ "debug": False,
+ "input_paths": [input_path],
+ "output_dir": output_dir,
+ "setLayout": False,
+ "silent": True,
+ }
+ with freeze_time(FROZEN_TIMESTAMP):
+ entry_point_for_args(args)
+
+
+def write_modified(modify_content, boilerplate, sample_json_path):
+ if boilerplate is None:
+ return
+
+ content = deepcopy(boilerplate)
+
+ if modify_content is not None:
+ returned_value = modify_content(content)
+ if returned_value is not None:
+ content = returned_value
+
+ with open(sample_json_path, "w") as f:
+ json.dump(content, f)
+
+
+def remove_file(path):
+ if os.path.exists(path):
+ os.remove(path)
+
+
+def generate_write_jsons_and_run(
+ run_sample,
+ sample_path,
+ template_boilerplate=None,
+ config_boilerplate=None,
+ evaluation_boilerplate=None,
+):
+ if (template_boilerplate or config_boilerplate or evaluation_boilerplate) is None:
+ raise Exception(
+ f"No boilerplates found. Provide atleast one boilerplate to write json."
+ )
+
+ def write_jsons_and_run(
+ mocker,
+ modify_template=None,
+ modify_config=None,
+ modify_evaluation=None,
+ ):
+ sample_template_path, sample_config_path, sample_evaluation_path = (
+ sample_path.joinpath("template.json"),
+ sample_path.joinpath("config.json"),
+ sample_path.joinpath("evaluation.json"),
+ )
+ write_modified(modify_template, template_boilerplate, sample_template_path)
+ write_modified(modify_config, config_boilerplate, sample_config_path)
+ write_modified(
+ modify_evaluation, evaluation_boilerplate, sample_evaluation_path
+ )
+
+ exception = "No Error"
+ try:
+ run_sample(mocker, sample_path)
+ except Exception as e:
+ exception = e
+
+ remove_file(sample_template_path)
+ remove_file(sample_config_path)
+ remove_file(sample_evaluation_path)
+
+ return exception
+
+ return write_jsons_and_run
diff --git a/OMRChecker/src/utils/__init__.py b/OMRChecker/src/utils/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..e69de29bb2d1d6434b8b29ae775ad8c2e48c5391
diff --git a/OMRChecker/src/utils/file.py b/OMRChecker/src/utils/file.py
new file mode 100644
index 0000000000000000000000000000000000000000..89d827b49c62d78c66510547f1f312a0263e5496
--- /dev/null
+++ b/OMRChecker/src/utils/file.py
@@ -0,0 +1,95 @@
+import argparse
+import json
+import os
+from csv import QUOTE_NONNUMERIC
+from time import localtime, strftime
+
+import pandas as pd
+
+from src.logger import logger
+
+
+def load_json(path, **rest):
+ try:
+ with open(path, "r") as f:
+ loaded = json.load(f, **rest)
+ except json.decoder.JSONDecodeError as error:
+ logger.critical(f"Error when loading json file at: '{path}'\n{error}")
+ exit(1)
+ return loaded
+
+
+class Paths:
+ def __init__(self, output_dir):
+ self.output_dir = output_dir
+ self.save_marked_dir = output_dir.joinpath("CheckedOMRs")
+ self.results_dir = output_dir.joinpath("Results")
+ self.manual_dir = output_dir.joinpath("Manual")
+ self.evaluation_dir = output_dir.joinpath("Evaluation")
+ self.errors_dir = self.manual_dir.joinpath("ErrorFiles")
+ self.multi_marked_dir = self.manual_dir.joinpath("MultiMarkedFiles")
+
+
+def setup_dirs_for_paths(paths):
+ logger.info("Checking Directories...")
+ for save_output_dir in [paths.save_marked_dir]:
+ if not os.path.exists(save_output_dir):
+ logger.info(f"Created : {save_output_dir}")
+ os.makedirs(save_output_dir)
+ os.mkdir(save_output_dir.joinpath("stack"))
+ os.mkdir(save_output_dir.joinpath("_MULTI_"))
+ os.mkdir(save_output_dir.joinpath("_MULTI_", "stack"))
+
+ for save_output_dir in [paths.manual_dir, paths.results_dir, paths.evaluation_dir]:
+ if not os.path.exists(save_output_dir):
+ logger.info(f"Created : {save_output_dir}")
+ os.makedirs(save_output_dir)
+
+ for save_output_dir in [paths.multi_marked_dir, paths.errors_dir]:
+ if not os.path.exists(save_output_dir):
+ logger.info(f"Created : {save_output_dir}")
+ os.makedirs(save_output_dir)
+
+
+def setup_outputs_for_template(paths, template):
+ # TODO: consider moving this into a class instance
+ ns = argparse.Namespace()
+ logger.info("Checking Files...")
+
+ # Include current output paths
+ ns.paths = paths
+
+ ns.empty_resp = [""] * len(template.output_columns)
+ ns.sheetCols = [
+ "file_id",
+ "input_path",
+ "output_path",
+ "score",
+ ] + template.output_columns
+ ns.OUTPUT_SET = []
+ ns.files_obj = {}
+ TIME_NOW_HRS = strftime("%I%p", localtime())
+ ns.filesMap = {
+ "Results": os.path.join(paths.results_dir, f"Results_{TIME_NOW_HRS}.csv"),
+ "MultiMarked": os.path.join(paths.manual_dir, "MultiMarkedFiles.csv"),
+ "Errors": os.path.join(paths.manual_dir, "ErrorFiles.csv"),
+ }
+
+ for file_key, file_name in ns.filesMap.items():
+ if not os.path.exists(file_name):
+ logger.info(f"Created new file: '{file_name}'")
+ # moved handling of files to pandas csv writer
+ ns.files_obj[file_key] = file_name
+ # Create Header Columns
+ pd.DataFrame([ns.sheetCols], dtype=str).to_csv(
+ ns.files_obj[file_key],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ else:
+ logger.info(f"Present : appending to '{file_name}'")
+ ns.files_obj[file_key] = open(file_name, "a")
+
+ return ns
diff --git a/OMRChecker/src/utils/image.py b/OMRChecker/src/utils/image.py
new file mode 100644
index 0000000000000000000000000000000000000000..cb546b0151225306fa2804958e515b6017687255
--- /dev/null
+++ b/OMRChecker/src/utils/image.py
@@ -0,0 +1,155 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import cv2
+import matplotlib.pyplot as plt
+import numpy as np
+
+from src.logger import logger
+
+plt.rcParams["figure.figsize"] = (10.0, 8.0)
+CLAHE_HELPER = cv2.createCLAHE(clipLimit=5.0, tileGridSize=(8, 8))
+
+
+class ImageUtils:
+ """A Static-only Class to hold common image processing utilities & wrappers over OpenCV functions"""
+
+ @staticmethod
+ def save_img(path, final_marked):
+ logger.info(f"Saving Image to '{path}'")
+ cv2.imwrite(path, final_marked)
+
+ @staticmethod
+ def resize_util(img, u_width, u_height=None):
+ if u_height is None:
+ h, w = img.shape[:2]
+ u_height = int(h * u_width / w)
+ return cv2.resize(img, (int(u_width), int(u_height)))
+
+ @staticmethod
+ def resize_util_h(img, u_height, u_width=None):
+ if u_width is None:
+ h, w = img.shape[:2]
+ u_width = int(w * u_height / h)
+ return cv2.resize(img, (int(u_width), int(u_height)))
+
+ @staticmethod
+ def grab_contours(cnts):
+ # source: imutils package
+
+ # if the length the contours tuple returned by cv2.findContours
+ # is '2' then we are using either OpenCV v2.4, v4-beta, or
+ # v4-official
+ if len(cnts) == 2:
+ cnts = cnts[0]
+
+ # if the length of the contours tuple is '3' then we are using
+ # either OpenCV v3, v4-pre, or v4-alpha
+ elif len(cnts) == 3:
+ cnts = cnts[1]
+
+ # otherwise OpenCV has changed their cv2.findContours return
+ # signature yet again and I have no idea WTH is going on
+ else:
+ raise Exception(
+ (
+ "Contours tuple must have length 2 or 3, "
+ "otherwise OpenCV changed their cv2.findContours return "
+ "signature yet again. Refer to OpenCV's documentation "
+ "in that case"
+ )
+ )
+
+ # return the actual contours array
+ return cnts
+
+ @staticmethod
+ def normalize_util(img, alpha=0, beta=255):
+ return cv2.normalize(img, alpha, beta, norm_type=cv2.NORM_MINMAX)
+
+ @staticmethod
+ def auto_canny(image, sigma=0.93):
+ # compute the median of the single channel pixel intensities
+ v = np.median(image)
+
+ # apply automatic Canny edge detection using the computed median
+ lower = int(max(0, (1.0 - sigma) * v))
+ upper = int(min(255, (1.0 + sigma) * v))
+ edged = cv2.Canny(image, lower, upper)
+
+ # return the edged image
+ return edged
+
+ @staticmethod
+ def adjust_gamma(image, gamma=1.0):
+ # build a lookup table mapping the pixel values [0, 255] to
+ # their adjusted gamma values
+ inv_gamma = 1.0 / gamma
+ table = np.array(
+ [((i / 255.0) ** inv_gamma) * 255 for i in np.arange(0, 256)]
+ ).astype("uint8")
+
+ # apply gamma correction using the lookup table
+ return cv2.LUT(image, table)
+
+ @staticmethod
+ def four_point_transform(image, pts):
+ # obtain a consistent order of the points and unpack them
+ # individually
+ rect = ImageUtils.order_points(pts)
+ (tl, tr, br, bl) = rect
+
+ # compute the width of the new image, which will be the
+ width_a = np.sqrt(((br[0] - bl[0]) ** 2) + ((br[1] - bl[1]) ** 2))
+ width_b = np.sqrt(((tr[0] - tl[0]) ** 2) + ((tr[1] - tl[1]) ** 2))
+
+ max_width = max(int(width_a), int(width_b))
+ # max_width = max(int(np.linalg.norm(br-bl)), int(np.linalg.norm(tr-tl)))
+
+ # compute the height of the new image, which will be the
+ height_a = np.sqrt(((tr[0] - br[0]) ** 2) + ((tr[1] - br[1]) ** 2))
+ height_b = np.sqrt(((tl[0] - bl[0]) ** 2) + ((tl[1] - bl[1]) ** 2))
+ max_height = max(int(height_a), int(height_b))
+ # max_height = max(int(np.linalg.norm(tr-br)), int(np.linalg.norm(tl-br)))
+
+ # now that we have the dimensions of the new image, construct
+ # the set of destination points to obtain a "birds eye view",
+ # (i.e. top-down view) of the image, again specifying points
+ # in the top-left, top-right, bottom-right, and bottom-left
+ # order
+ dst = np.array(
+ [
+ [0, 0],
+ [max_width - 1, 0],
+ [max_width - 1, max_height - 1],
+ [0, max_height - 1],
+ ],
+ dtype="float32",
+ )
+
+ transform_matrix = cv2.getPerspectiveTransform(rect, dst)
+ warped = cv2.warpPerspective(image, transform_matrix, (max_width, max_height))
+
+ # return the warped image
+ return warped
+
+ @staticmethod
+ def order_points(pts):
+ rect = np.zeros((4, 2), dtype="float32")
+
+ # the top-left point will have the smallest sum, whereas
+ # the bottom-right point will have the largest sum
+ s = pts.sum(axis=1)
+ rect[0] = pts[np.argmin(s)]
+ rect[2] = pts[np.argmax(s)]
+ diff = np.diff(pts, axis=1)
+ rect[1] = pts[np.argmin(diff)]
+ rect[3] = pts[np.argmax(diff)]
+
+ # return the ordered coordinates
+ return rect
diff --git a/OMRChecker/src/utils/interaction.py b/OMRChecker/src/utils/interaction.py
new file mode 100644
index 0000000000000000000000000000000000000000..72968ec3951bd5bc21fabe545b0f87c0ef3fe896
--- /dev/null
+++ b/OMRChecker/src/utils/interaction.py
@@ -0,0 +1,110 @@
+from dataclasses import dataclass
+
+import cv2
+try:
+ from screeninfo import get_monitors
+ monitor_window = get_monitors()[0]
+ _monitor_width, _monitor_height = monitor_window.width, monitor_window.height
+except Exception:
+ _monitor_width, _monitor_height = 1920, 1080
+
+from src.logger import logger
+from src.utils.image import ImageUtils
+
+
+@dataclass
+class ImageMetrics:
+ # TODO: Move TEXT_SIZE, etc here and find a better class name
+ window_width, window_height = _monitor_width, _monitor_height
+ # for positioning image windows
+ window_x, window_y = 0, 0
+ reset_pos = [0, 0]
+
+
+class InteractionUtils:
+ """Perform primary functions such as displaying images and reading responses"""
+
+ image_metrics = ImageMetrics()
+
+ @staticmethod
+ def show(name, origin, pause=1, resize=False, reset_pos=None, config=None):
+ image_metrics = InteractionUtils.image_metrics
+ if origin is None:
+ logger.info(f"'{name}' - NoneType image to show!")
+ if pause:
+ cv2.destroyAllWindows()
+ return
+ if resize:
+ if not config:
+ raise Exception("config not provided for resizing the image to show")
+ img = ImageUtils.resize_util(origin, config.dimensions.display_width)
+ else:
+ img = origin
+
+ if not is_window_available(name):
+ cv2.namedWindow(name)
+
+ cv2.imshow(name, img)
+
+ if reset_pos:
+ image_metrics.window_x = reset_pos[0]
+ image_metrics.window_y = reset_pos[1]
+
+ cv2.moveWindow(
+ name,
+ image_metrics.window_x,
+ image_metrics.window_y,
+ )
+
+ h, w = img.shape[:2]
+
+ # Set next window position
+ margin = 25
+ w += margin
+ h += margin
+
+ w, h = w // 2, h // 2
+ if image_metrics.window_x + w > image_metrics.window_width:
+ image_metrics.window_x = 0
+ if image_metrics.window_y + h > image_metrics.window_height:
+ image_metrics.window_y = 0
+ else:
+ image_metrics.window_y += h
+ else:
+ image_metrics.window_x += w
+
+ if pause:
+ logger.info(
+ f"Showing '{name}'\n\t Press Q on image to continue. Press Ctrl + C in terminal to exit"
+ )
+
+ wait_q()
+ InteractionUtils.image_metrics.window_x = 0
+ InteractionUtils.image_metrics.window_y = 0
+
+
+@dataclass
+class Stats:
+ # TODO Fill these for stats
+ # Move qbox_vals here?
+ # badThresholds = []
+ # veryBadPoints = []
+ files_moved = 0
+ files_not_moved = 0
+
+
+def wait_q():
+ esc_key = 27
+ while cv2.waitKey(1) & 0xFF not in [ord("q"), esc_key]:
+ pass
+ cv2.destroyAllWindows()
+
+
+def is_window_available(name: str) -> bool:
+ """Checks if a window is available"""
+ try:
+ cv2.getWindowProperty(name, cv2.WND_PROP_VISIBLE)
+ return True
+ except Exception as e:
+ print(e)
+ return False
diff --git a/OMRChecker/src/utils/parsing.py b/OMRChecker/src/utils/parsing.py
new file mode 100644
index 0000000000000000000000000000000000000000..b500271ca038fc0654fba6d823ea517b70719fa1
--- /dev/null
+++ b/OMRChecker/src/utils/parsing.py
@@ -0,0 +1,113 @@
+import re
+from copy import deepcopy
+from fractions import Fraction
+
+from deepmerge import Merger
+from dotmap import DotMap
+
+from src.constants import FIELD_LABEL_NUMBER_REGEX
+from src.defaults import CONFIG_DEFAULTS, TEMPLATE_DEFAULTS
+from src.schemas.constants import FIELD_STRING_REGEX_GROUPS
+from src.utils.file import load_json
+from src.utils.validations import (
+ validate_config_json,
+ validate_evaluation_json,
+ validate_template_json,
+)
+
+OVERRIDE_MERGER = Merger(
+ # pass in a list of tuples,with the
+ # strategies you are looking to apply
+ # to each type.
+ [
+ # (list, ["prepend"]),
+ (dict, ["merge"])
+ ],
+ # next, choose the fallback strategies,
+ # applied to all other types:
+ ["override"],
+ # finally, choose the strategies in
+ # the case where the types conflict:
+ ["override"],
+)
+
+
+def get_concatenated_response(omr_response, template):
+ # Multi-column/multi-row questions which need to be concatenated
+ concatenated_response = {}
+ for field_label, concatenate_keys in template.custom_labels.items():
+ custom_label = "".join([omr_response[k] for k in concatenate_keys])
+ concatenated_response[field_label] = custom_label
+
+ for field_label in template.non_custom_labels:
+ concatenated_response[field_label] = omr_response[field_label]
+
+ return concatenated_response
+
+
+def open_config_with_defaults(config_path):
+ user_tuning_config = load_json(config_path)
+ user_tuning_config = OVERRIDE_MERGER.merge(
+ deepcopy(CONFIG_DEFAULTS), user_tuning_config
+ )
+ validate_config_json(user_tuning_config, config_path)
+ # https://github.com/drgrib/dotmap/issues/74
+ return DotMap(user_tuning_config, _dynamic=False)
+
+
+def open_template_with_defaults(template_path):
+ user_template = load_json(template_path)
+ user_template = OVERRIDE_MERGER.merge(deepcopy(TEMPLATE_DEFAULTS), user_template)
+ validate_template_json(user_template, template_path)
+ return user_template
+
+
+def open_evaluation_with_validation(evaluation_path):
+ user_evaluation_config = load_json(evaluation_path)
+ validate_evaluation_json(user_evaluation_config, evaluation_path)
+ return user_evaluation_config
+
+
+def parse_fields(key, fields):
+ parsed_fields = []
+ fields_set = set()
+ for field_string in fields:
+ fields_array = parse_field_string(field_string)
+ current_set = set(fields_array)
+ if not fields_set.isdisjoint(current_set):
+ raise Exception(
+ f"Given field string '{field_string}' has overlapping field(s) with other fields in '{key}': {fields}"
+ )
+ fields_set.update(current_set)
+ parsed_fields.extend(fields_array)
+ return parsed_fields
+
+
+def parse_field_string(field_string):
+ if "." in field_string:
+ field_prefix, start, end = re.findall(FIELD_STRING_REGEX_GROUPS, field_string)[
+ 0
+ ]
+ start, end = int(start), int(end)
+ if start >= end:
+ raise Exception(
+ f"Invalid range in fields string: '{field_string}', start: {start} is not less than end: {end}"
+ )
+ return [
+ f"{field_prefix}{field_number}" for field_number in range(start, end + 1)
+ ]
+ else:
+ return [field_string]
+
+
+def custom_sort_output_columns(field_label):
+ label_prefix, label_suffix = re.findall(FIELD_LABEL_NUMBER_REGEX, field_label)[0]
+ return [label_prefix, int(label_suffix) if len(label_suffix) > 0 else 0]
+
+
+def parse_float_or_fraction(result):
+ if type(result) == str and "/" in result:
+ result = float(Fraction(result))
+ else:
+ result = float(result)
+ return result
diff --git a/OMRChecker/src/utils/validations.py b/OMRChecker/src/utils/validations.py
new file mode 100644
index 0000000000000000000000000000000000000000..84332a0bc85cd784bfb9782450c9e510676cc656
--- /dev/null
+++ b/OMRChecker/src/utils/validations.py
@@ -0,0 +1,115 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import re
+
+import jsonschema
+from jsonschema import validate
+from rich.table import Table
+
+from src.logger import console, logger
+from src.schemas import SCHEMA_JSONS, SCHEMA_VALIDATORS
+
+
+def validate_evaluation_json(json_data, evaluation_path):
+ logger.info(f"Loading evaluation.json: {evaluation_path}")
+ try:
+ validate(instance=json_data, schema=SCHEMA_JSONS["evaluation"])
+ except jsonschema.exceptions.ValidationError as _err: # NOQA
+ table = Table(show_lines=True)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Error", style="magenta")
+
+ errors = sorted(
+ SCHEMA_VALIDATORS["evaluation"].iter_errors(json_data),
+ key=lambda e: e.path,
+ )
+ for error in errors:
+ key, validator, msg = parse_validation_error(error)
+ if validator == "required":
+ requiredProperty = re.findall(r"'(.*?)'", msg)[0]
+ table.add_row(
+ f"{key}.{requiredProperty}",
+ msg + ". Make sure the spelling of the key is correct",
+ )
+ else:
+ table.add_row(key, msg)
+ console.print(table, justify="center")
+ raise Exception(
+ f"Provided Evaluation JSON is Invalid: '{evaluation_path}'"
+ ) from None
+
+
+def validate_template_json(json_data, template_path):
+ logger.info(f"Loading template.json: {template_path}")
+ try:
+ validate(instance=json_data, schema=SCHEMA_JSONS["template"])
+ except jsonschema.exceptions.ValidationError as _err: # NOQA
+ table = Table(show_lines=True)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Error", style="magenta")
+
+ errors = sorted(
+ SCHEMA_VALIDATORS["template"].iter_errors(json_data),
+ key=lambda e: e.path,
+ )
+ for error in errors:
+ key, validator, msg = parse_validation_error(error)
+
+ # Print preProcessor name in case of options error
+ if key == "preProcessors":
+ preProcessorName = json_data["preProcessors"][error.path[1]]["name"]
+ preProcessorKey = error.path[2]
+ table.add_row(f"{key}.{preProcessorName}.{preProcessorKey}", msg)
+ elif validator == "required":
+ requiredProperty = re.findall(r"'(.*?)'", msg)[0]
+ table.add_row(
+ f"{key}.{requiredProperty}",
+ f"{msg}. Check for spelling errors and make sure it is in camelCase",
+ )
+ else:
+ table.add_row(key, msg)
+ console.print(table, justify="center")
+ raise Exception(
+ f"Provided Template JSON is Invalid: '{template_path}'"
+ ) from None
+
+
+def validate_config_json(json_data, config_path):
+ logger.info(f"Loading config.json: {config_path}")
+ try:
+ validate(instance=json_data, schema=SCHEMA_JSONS["config"])
+ except jsonschema.exceptions.ValidationError as _err: # NOQA
+ table = Table(show_lines=True)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Error", style="magenta")
+ errors = sorted(
+ SCHEMA_VALIDATORS["config"].iter_errors(json_data),
+ key=lambda e: e.path,
+ )
+ for error in errors:
+ key, validator, msg = parse_validation_error(error)
+
+ if validator == "required":
+ requiredProperty = re.findall(r"'(.*?)'", msg)[0]
+ table.add_row(
+ f"{key}.{requiredProperty}",
+ f"{msg}. Check for spelling errors and make sure it is in camelCase",
+ )
+ else:
+ table.add_row(key, msg)
+ console.print(table, justify="center")
+ raise Exception(f"Provided config JSON is Invalid: '{config_path}'") from None
+
+
+def parse_validation_error(error):
+ return (
+ (error.path[0] if len(error.path) > 0 else "$root"),
+ error.validator,
+ error.message,
+ )
diff --git a/README.md b/README.md
index a49d94a9a4c6b64ee5291b5d4cfa41e385b0c51b..1ff342f984e5b3c4275cf804a348a0460c18a4fe 100644
--- a/README.md
+++ b/README.md
@@ -1,10 +1,90 @@
----
-title: Ocr Omr Backend
-emoji: 📚
-colorFrom: indigo
-colorTo: indigo
-sdk: docker
-pinned: false
----
-
-Check out the configuration reference at https://huggingface.co/docs/hub/spaces-config-reference
+# OCR Backend
+
+Backend API for OCR on handwritten images.
+
+## Setup
+
+1. Create a virtual environment:
+ ```bash
+ python -m venv venv
+ ```
+2. Activate the environment:
+ * Windows:
+ ```bash
+ .\venv\Scripts\activate
+ ```
+3. Install dependencies:
+ ```bash
+ pip install -r requirements.txt
+ ```
+4. **For Tesseract OCR:** Install Tesseract on your system. Download from [Tesseract GitHub](https://github.com/tesseract-ocr/tesseract).
+ * If `pytesseract` can't find Tesseract, you might need to set the path in `app.py`:
+ ```python
+ pytesseract.pytesseract.tesseract_cmd = r'C:\Program Files\Tesseract-OCR\tesseract.exe'
+ ```
+5. **For PDF processing:** Install Poppler. Download from [Poppler for Windows](https://github.com/oschwartz10612/poppler-windows/releases).
+ * You will need to update the `poppler_path` in `app.py` to point to the `bin` directory of your Poppler installation (e.g., `r'C:\Program Files\poppler-0.68.0\bin'`)
+
+## Run
+
+```bash
+python app.py
+```
+
+API will be at `http://127.0.0.1:5000`.
+
+## API Endpoints
+
+### `POST /easyocr`
+
+Uses EasyOCR to extract text from images.
+
+**Request:** `multipart/form-data` with `images` (one or more image files).
+
+**Example (curl):**
+
+```bash
+curl -X POST -F "images=@/path/to/your/image1.png" http://127.0.0.1:5000/easyocr
+```
+
+### `POST /tesseract`
+
+Uses Tesseract OCR to extract text from images.
+
+**Request:** `multipart/form-data` with `images` (one or more image files).
+
+**Example (curl):**
+
+```bash
+curl -X POST -F "images=@/path/to/your/image1.png" http://127.0.0.1:5000/tesseract
+```
+
+### `POST /process_question_paper`
+
+Processes an image or PDF of a question paper to extract questions and answers.
+
+**Request:** `multipart/form-data` with `file` (a single image or PDF file).
+
+**Example (curl for image):**
+
+```bash
+curl -X POST -F "file=@/path/to/your/question_paper.png" http://127.0.0.1:5000/process_question_paper
+```
+
+**Example (curl for PDF):**
+
+```bash
+curl -X POST -F "file=@/path/to/your/question_paper.pdf" http://127.0.0.1:5000/process_question_paper
+```
+
+### `GET /evaluate_answers`
+
+Compares OCR extracted texts with the answers from the last processed question paper.
+
+**Request:** None (GET request).
+
+**Example (curl):**
+
+```bash
+curl -X GET http://127.0.0.1:5000/evaluate_answers
+```
diff --git a/app.py b/app.py
new file mode 100644
index 0000000000000000000000000000000000000000..6706a773d328a3aaf4cdcd993304a233bc9c962a
--- /dev/null
+++ b/app.py
@@ -0,0 +1,2242 @@
+import sys
+import platform
+import easyocr
+from pdf2image import convert_from_path, convert_from_bytes
+from flask import Flask, request, jsonify
+from flask_cors import CORS
+from dataclasses import dataclass
+from typing import List, Tuple, Optional, Dict, Any
+from collections import defaultdict
+import numpy as np
+import cv2
+import pytesseract
+from PIL import Image
+import os
+import tempfile
+import difflib
+import re
+from fuzzywuzzy import fuzz
+from dotenv import load_dotenv
+import google.generativeai as genai
+import asyncio
+import base64
+import io
+import json
+import pandas as pd
+import subprocess
+
+# Import the SupabaseHandler
+import uuid
+from datetime import datetime
+from supabase import create_client, Client
+
+_tesseract_cmd = os.getenv("TESSERACT_CMD")
+if _tesseract_cmd:
+ pytesseract.pytesseract.tesseract_cmd = _tesseract_cmd
+elif platform.system() == "Windows":
+ pytesseract.pytesseract.tesseract_cmd = r"C:\Program Files\Tesseract-OCR\tesseract.exe"
+
+
+def _get_poppler_path():
+ env_path = os.getenv("POPPLER_PATH")
+ if env_path:
+ return env_path
+ if platform.system() == "Windows":
+ # Check common install locations
+ candidates = [
+ r'C:\Program Files\poppler\Library\bin',
+ r'C:\Program Files\poppler\poppler-24.08.0\Library\bin',
+ ]
+ # Also scan for any versioned poppler directory
+ poppler_base = r'C:\Program Files\poppler'
+ if os.path.isdir(poppler_base):
+ for entry in os.listdir(poppler_base):
+ candidate = os.path.join(poppler_base, entry, 'Library', 'bin')
+ if candidate not in candidates:
+ candidates.append(candidate)
+ for path in candidates:
+ if os.path.isdir(path):
+ return path
+ return None
+
+
+load_dotenv()
+GEMINI_API_KEY = os.getenv("GEMINI_API_KEY")
+genai.configure(api_key=GEMINI_API_KEY)
+
+app = Flask(__name__)
+CORS(app)
+reader = easyocr.Reader(['en'])
+
+# Global variables to store processing results
+ocr_extracted_texts = []
+last_processed_question_paper_object = None
+last_processed_omr_key = None # Global variable to store OMR answer key
+last_processed_omr_results = None # Global variable to store OMR processing results
+porcessed_omr_results = []
+OMR_IMAGES = []
+
+class SupabaseHandler:
+ def __init__(self):
+ url: str = os.getenv("SUPABASE_URL")
+ key: str = os.getenv("SUPABASE_ANON_KEY")
+ if not url or not key:
+ raise ValueError("Supabase URL and ANON_KEY must be set in environment variables")
+ self.supabase: Client = create_client(url, key)
+
+ def store_evaluation_result(self, teacher_email, evaluation_data, exam_name=None):
+ """
+ Store evaluation result in Supabase with a unique key and exam name
+ Returns the unique key for retrieval
+ """
+ try:
+ # Generate unique key
+ unique_key = str(uuid.uuid4())
+
+ # Prepare data for storage
+ storage_data = {
+ "unique_key": unique_key,
+ "teacher_email": teacher_email,
+ "evaluation_data": evaluation_data,
+ "exam_name": exam_name, # Add exam name field
+ "created_at": datetime.utcnow().isoformat(),
+ "total_students": evaluation_data.get("total_students", 0)
+ }
+
+ # Insert into Supabase
+ result = self.supabase.table("evaluation_results").insert(storage_data).execute()
+
+ if result.data:
+ print(f"Successfully stored evaluation result with key: {unique_key} for exam: {exam_name}")
+ return unique_key
+ else:
+ print("Failed to store evaluation result")
+ return None
+
+ except Exception as e:
+ print(f"Error storing evaluation result: {str(e)}")
+ return None
+
+ def get_evaluation_result(self, unique_key):
+ """
+ Retrieve evaluation result by unique key
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("*").eq("unique_key", unique_key).execute()
+
+ if result.data and len(result.data) > 0:
+ return result.data[0]
+ else:
+ return None
+
+ except Exception as e:
+ print(f"Error retrieving evaluation result: {str(e)}")
+ return None
+
+ def get_teacher_evaluations(self, teacher_email):
+ """
+ Get all evaluation results for a specific teacher
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("unique_key", "created_at", "total_students", "exam_name").eq("teacher_email", teacher_email).order("created_at", desc=True).execute()
+
+ if result.data:
+ return result.data
+ else:
+ return []
+
+ except Exception as e:
+ print(f"Error retrieving teacher evaluations: {str(e)}")
+ return []
+
+class QuestionPaper:
+ def __init__(self, path=None):
+ self.questions = []
+ self.answers = []
+ self.path = path
+ def clean_answers(self):
+ # Remove unwanted patterns from answers
+ unwanted_patterns = [
+ "Time: 15 MinutesMarks: 20",
+ "Time: 15 Minutes Marks: 20",
+ "GENERAL KNOWLEDGE QUESTION PAPER WITH ANSWERS",
+ "GENERAL KNOWLEDGE QUESTION PAPER",
+ ]
+ # Filter out unwanted answers
+ cleaned_answers = []
+ for answer in self.answers:
+ if answer.strip() and answer.strip() not in unwanted_patterns:
+ # Also check if it doesn't match any unwanted pattern with regex
+ is_unwanted = False
+ for pattern in unwanted_patterns:
+ if pattern and re.search(re.escape(pattern), answer, re.IGNORECASE):
+ is_unwanted = True
+ break
+ if not is_unwanted:
+ cleaned_answers.append(answer.strip())
+
+ self.answers = cleaned_answers
+
+ def add_question(self, question_text):
+ self.questions.append(question_text)
+
+ def add_answer(self, answer_text):
+ self.answers.append(answer_text)
+
+ def to_dict(self):
+ return {
+ 'questions': self.questions,
+ 'answers': self.answers
+ }
+
+class OMRAnswerKey:
+ def __init__(self):
+ self.answers = {} # Dictionary mapping question numbers to correct options
+ self.total_marks = 0
+ self.marks_per_question = 1
+ self.negative_marking = 0
+ self.title = ""
+ self.duration = ""
+ self.total_questions = 0
+ self.path = None
+ self.questions = [] # List to store questions if needed
+ self.question_data = [] # List to store complete question data with options
+
+ def __str__(self):
+ return f"OMR Answer Key: {self.title}\nTotal Questions: {self.total_questions}\nAnswers: {self.answers}"
+
+ def set_answers(self, answers: dict):
+ """Set the answer key with question numbers as keys and correct options (A,B,C,D) as values"""
+ self.answers = {int(k): v.upper() for k, v in answers.items() if v.upper() in ['A', 'B', 'C', 'D']}
+ self.total_questions = len(self.answers)
+
+ def set_marking_scheme(self, marks_per_question: float, negative_marking: float = 0):
+ """Set the marking scheme for the answer key"""
+ self.marks_per_question = marks_per_question
+ self.negative_marking = negative_marking
+ self.total_marks = self.total_questions * marks_per_question
+
+ def set_metadata(self, title: str, duration: str):
+ """Set metadata for the answer key"""
+ self.title = title
+ self.duration = duration
+
+ def set_question_data(self, question_data):
+ """Store complete question data including options"""
+ self.question_data = question_data
+ self.questions = [f"{q['number']}. {q['question']}" for q in question_data]
+ self.answers = {q['number']: q['answer'] for q in question_data if q['answer']}
+ self.total_questions = len(question_data)
+
+ def get_question_details(self, question_number):
+ """Get complete details for a specific question"""
+ for q in self.question_data:
+ print(f"Checking question number: {q['number']} with {question_number}")
+ if str(q['number']) == str(question_number):
+ return q
+ return None
+
+ def to_dict(self):
+ return {
+ 'title': self.title,
+ 'duration': self.duration,
+ 'total_questions': self.total_questions,
+ 'answers': self.answers,
+ 'total_marks': self.total_marks,
+ 'marks_per_question': self.marks_per_question,
+ 'negative_marking': self.negative_marking,
+ 'questions': self.questions,
+ 'question_data': self.question_data # Include complete question data
+ }
+
+
+
+def parse_question_paper_text(text):
+ """
+ Improved parsing function that correctly identifies questions and answers
+ """
+ lines = [line.strip() for line in text.split('\n') if line.strip()]
+
+ questions = []
+ answers = []
+
+ # Patterns to ignore (headers, footers, etc.)
+ ignore_patterns = [
+ r'GENERAL KNOWLEDGE QUESTION PAPER.*',
+ r'Time:\s*\d+\s*Minutes.*Marks:\s*\d+',
+ r'Time:\s*\d+\s*MinutesMarks:\s*\d+',
+ r'^\s*$' # Empty lines
+ ]
+
+ # Filter out unwanted lines
+ filtered_lines = []
+ for line in lines:
+ should_ignore = False
+ for pattern in ignore_patterns:
+ if re.match(pattern, line, re.IGNORECASE):
+ should_ignore = True
+ break
+ if not should_ignore:
+ filtered_lines.append(line)
+
+ # Pattern to identify questions (starts with number followed by dot/parenthesis)
+ question_pattern = r'^\d+\s*[.)]\s*(.+)'
+
+ i = 0
+ while i < len(filtered_lines):
+ current_line = filtered_lines[i].strip()
+
+ # Check if current line is a question
+ question_match = re.match(question_pattern, current_line)
+ if question_match:
+ # This is a question
+ question_text = question_match.group(1).strip()
+ questions.append(f"{current_line}") # Keep the full question with number
+
+ # Look for the answer in the next line
+ if i + 1 < len(filtered_lines):
+ next_line = filtered_lines[i + 1].strip()
+ # If next line is not a question (doesn't start with number), it's likely an answer
+ if not re.match(question_pattern, next_line):
+ answers.append(next_line)
+ i += 2 # Skip both question and answer
+ else:
+ # Next line is also a question, so this question might not have an answer
+ # Or the answer might be embedded in the same line
+ # Try to extract answer from the question line itself if it contains common answer patterns
+ answers.append("") # Placeholder for missing answer
+ i += 1
+ else:
+ # Last line and it's a question without answer
+ answers.append("")
+ i += 1
+ else:
+ # This line doesn't match question pattern, skip it or try to pair it with previous question
+ if len(questions) > len(answers):
+ # We have more questions than answers, this might be an answer
+ answers.append(current_line)
+ i += 1
+
+ # Ensure we have equal number of questions and answers
+ while len(answers) < len(questions):
+ answers.append("")
+ while len(questions) < len(answers):
+ questions.append(f"Question {len(questions) + 1}")
+
+ return questions, answers
+
+def improved_clean_and_parse_ocr_text(ocr_text):
+ """
+ Improved parsing with better answer extraction logic
+ """
+ # Remove special characters but keep important ones
+ cleaned_text = re.sub(r'[|@~¥#$%^&*()_+=\[\]{}\\:";\'<>?,./]', ' ', ocr_text)
+
+ # Split by newlines and filter out empty strings
+ lines = [line.strip() for line in cleaned_text.split('\n') if line.strip()]
+
+ individual_answers = []
+
+ # Try to find numbered patterns first
+ numbered_pattern = re.compile(r'(\d+)\s*[.)]\s*([^0-9]+?)(?=\d+\s*[.)]|$)', re.MULTILINE | re.DOTALL)
+ matches = numbered_pattern.findall(cleaned_text)
+
+ if matches:
+ # If we found numbered patterns, use them
+ for number, answer in matches:
+ answer = answer.strip()
+ if answer and len(answer) > 1:
+ individual_answers.append(answer)
+ else:
+ # Fallback to line-by-line processing
+ for line in lines:
+ # Remove leading numbers and punctuation
+ cleaned_line = re.sub(r'^\d+\s*[.)]\s*', '', line).strip()
+ if cleaned_line and len(cleaned_line) > 1:
+ individual_answers.append(cleaned_line)
+
+ return individual_answers
+
+def find_best_match(student_answer, correct_answers, threshold=0.6):
+ """
+ Find the best matching correct answer for a student answer
+ """
+ best_score = 0
+ best_match = None
+
+ for correct_answer in correct_answers:
+ # Use multiple similarity metrics
+ ratio_score = difflib.SequenceMatcher(None, student_answer.lower(), correct_answer.lower()).ratio()
+ fuzzy_score = fuzz.ratio(student_answer.lower(), correct_answer.lower()) / 100.0
+ partial_score = fuzz.partial_ratio(student_answer.lower(), correct_answer.lower()) / 100.0
+
+ # Take the maximum of all scores
+ combined_score = max(ratio_score, fuzzy_score, partial_score)
+
+ if combined_score > best_score:
+ best_score = combined_score
+ best_match = correct_answer
+
+ # Only return match if it meets the threshold
+ if best_score >= threshold:
+ return best_match, best_score
+ else:
+ return None, best_score
+
+def extract_roll_number(student_answer_path):
+ """
+ Extract roll number from student answer sheet using OCR
+ """
+ try:
+ student_answer_image = Image.open(student_answer_path)
+ text = pytesseract.image_to_string(student_answer_image)
+
+ # Look for common roll number patterns
+ roll_patterns = [
+ r'(?i)roll\s*no\s*[:\-]?\s*(\w+)',
+ r'(?i)roll\s*number\s*[:\-]?\s*(\w+)',
+ r'(?i)roll\s*[:\-]?\s*(\w+)',
+ r'(?i)reg\s*no\s*[:\-]?\s*(\w+)',
+ r'(?i)registration\s*[:\-]?\s*(\w+)'
+ ]
+
+ for pattern in roll_patterns:
+ match = re.search(pattern, text)
+ if match:
+ return match.group(1).strip()
+
+ # If no explicit roll number found, try to find number sequences
+ number_sequences = re.findall(r'\b\d{2,}\b', text)
+ if number_sequences:
+ return number_sequences[0] # Return first significant number sequence
+
+ return "Unknown"
+ except Exception as e:
+ print(f"Error extracting roll number: {str(e)}")
+ return "Unknown"
+
+# OMR Section
+
+
+@dataclass
+class BubbleLocation:
+ """Stores information about each bubble"""
+ question_num: int
+ option: str
+ center: Tuple[int, int]
+ radius: int
+ filled: bool = False
+ fill_ratio: float = 0.0
+
+
+
+class CorrectedOMRReader:
+ def __init__(self, image_path: str = None, image_array: np.ndarray = None):
+ """Initialize the OMR Reader with an image"""
+ if image_array is not None:
+ self.image = image_array
+ self.image_path = None
+ elif image_path is not None:
+ self.image = cv2.imread(image_path)
+ self.image_path = image_path
+ else:
+ raise ValueError("Either image_array or image_path must be provided")
+
+ if self.image is None:
+ raise ValueError("Could not load image")
+
+ self.gray = cv2.cvtColor(self.image, cv2.COLOR_BGR2GRAY)
+ self.height, self.width = self.gray.shape
+ self.bubbles = []
+ self.answers = {}
+
+ # Expected grid parameters
+ self.expected_radius = 15 # Approximate bubble radius
+ self.grid_params = {
+ 'rows': 20, # Maximum rows
+ 'cols': 3, # 3 columns of questions
+ 'options': 4 # 4 options per question (A, B, C, D)
+ }
+
+ def preprocess_for_detection(self):
+ """Preprocess specifically for bubble DETECTION (not fill detection)"""
+ blurred = cv2.GaussianBlur(self.gray, (3, 3), 0)
+ _, thresh = cv2.threshold(blurred, 200, 255, cv2.THRESH_BINARY)
+ self.detection_thresh = cv2.bitwise_not(thresh)
+ return self.detection_thresh
+
+ def find_bubble_grid(self):
+ """Find bubble locations using grid detection"""
+ bubbles = []
+
+ param_sets = [
+ {'dp': 1.0, 'minDist': 20, 'param1': 50, 'param2': 28, 'minRadius': 10, 'maxRadius': 20},
+ {'dp': 1.1, 'minDist': 22, 'param1': 45, 'param2': 25, 'minRadius': 11, 'maxRadius': 19},
+ {'dp': 1.2, 'minDist': 25, 'param1': 40, 'param2': 30, 'minRadius': 9, 'maxRadius': 21},
+ ]
+
+ for params in param_sets:
+ circles = cv2.HoughCircles(
+ self.gray,
+ cv2.HOUGH_GRADIENT,
+ dp=params['dp'],
+ minDist=params['minDist'],
+ param1=params['param1'],
+ param2=params['param2'],
+ minRadius=params['minRadius'],
+ maxRadius=params['maxRadius']
+ )
+
+ if circles is not None:
+ circles = np.round(circles[0, :]).astype("int")
+ for (x, y, r) in circles:
+ is_dup = False
+ for bub in bubbles:
+ if np.sqrt((x - bub[0])**2 + (y - bub[1])**2) < 15:
+ is_dup = True
+ break
+ if not is_dup:
+ bubbles.append((x, y, r))
+
+ print(f" Found {len(bubbles)} bubbles with Hough Circles")
+
+ if len(bubbles) < 180:
+ template_bubbles = self.template_matching_detection()
+ bubbles.extend(template_bubbles)
+ print(f" Added {len(template_bubbles)} bubbles with template matching")
+
+ return bubbles
+
+ def template_matching_detection(self):
+ """Use template matching to find bubble locations"""
+ bubbles = []
+ template_size = 30
+ template = np.zeros((template_size, template_size), dtype=np.uint8)
+ cv2.circle(template, (template_size//2, template_size//2), 12, 255, 2)
+
+ result = cv2.matchTemplate(self.gray, template, cv2.TM_CCOEFF_NORMED)
+ threshold = 0.5
+ locations = np.where(result >= threshold)
+
+ for pt in zip(*locations[::-1]):
+ center_x = pt[0] + template_size // 2
+ center_y = pt[1] + template_size // 2
+ too_close = False
+ for (bx, by, _) in bubbles:
+ if np.sqrt((center_x - bx)**2 + (center_y - by)**2) < 20:
+ too_close = True
+ break
+ if not too_close:
+ bubbles.append((center_x, center_y, 12))
+
+ return bubbles
+
+ def detect_bubbles_by_contours(self):
+ """Detect bubbles using contours - focusing on circular shapes"""
+ bubbles = []
+ edge_params = [(30, 100), (50, 150), (20, 80)]
+
+ for low, high in edge_params:
+ edges = cv2.Canny(self.gray, low, high)
+ contours, _ = cv2.findContours(edges, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
+
+ for contour in contours:
+ area = cv2.contourArea(contour)
+ if 150 < area < 900:
+ (x, y), radius = cv2.minEnclosingCircle(contour)
+ perimeter = cv2.arcLength(contour, True)
+ if perimeter > 0:
+ circularity = 4 * np.pi * area / (perimeter * perimeter)
+ if circularity > 0.6 and 8 < radius < 22:
+ is_dup = False
+ for bub in bubbles:
+ if np.sqrt((x - bub[0])**2 + (y - bub[1])**2) < 15:
+ is_dup = True
+ break
+ if not is_dup:
+ bubbles.append((int(x), int(y), int(radius)))
+
+ return bubbles
+
+ def organize_and_filter_bubbles(self, all_bubbles):
+ if not all_bubbles:
+ return []
+
+ filtered_bubbles = []
+ for bubble in all_bubbles:
+ is_duplicate = False
+ for existing in filtered_bubbles:
+ dist = np.sqrt((bubble[0] - existing[0])**2 + (bubble[1] - existing[1])**2)
+ if dist < 15:
+ is_duplicate = True
+ break
+ if not is_duplicate:
+ filtered_bubbles.append(bubble)
+
+ filtered_bubbles.sort(key=lambda b: (b[1], b[0]))
+
+ rows = []
+ current_row = []
+ row_threshold = 20
+
+ for bubble in filtered_bubbles:
+ if not current_row:
+ current_row.append(bubble)
+ else:
+ avg_y = np.mean([b[1] for b in current_row])
+ if abs(bubble[1] - avg_y) < row_threshold:
+ current_row.append(bubble)
+ else:
+ if len(current_row) >= 4:
+ current_row.sort(key=lambda b: b[0])
+ rows.append(current_row)
+ current_row = [bubble]
+
+ if len(current_row) >= 4:
+ current_row.sort(key=lambda b: b[0])
+ rows.append(current_row)
+
+ return rows
+
+ def map_to_questions(self, bubble_rows):
+ mapped_bubbles = []
+ options = ['A', 'B', 'C', 'D']
+
+ if not bubble_rows:
+ return mapped_bubbles
+
+ col1_max = self.width * 0.35
+ col2_max = self.width * 0.68
+
+ for row_idx, row in enumerate(bubble_rows[:20]):
+ col1 = [b for b in row if b[0] < col1_max]
+ col2 = [b for b in row if col1_max <= b[0] < col2_max]
+ col3 = [b for b in row if b[0] >= col2_max]
+
+ if len(col1) >= 4:
+ col1_sorted = sorted(col1, key=lambda b: b[0])[:4]
+ q_num = row_idx + 1
+ for opt_idx, bubble in enumerate(col1_sorted):
+ mapped_bubbles.append(BubbleLocation(q_num, options[opt_idx], (bubble[0], bubble[1]), bubble[2]))
+
+ if len(col2) >= 4:
+ col2_sorted = sorted(col2, key=lambda b: b[0])[:4]
+ q_num = row_idx + 21
+ for opt_idx, bubble in enumerate(col2_sorted):
+ mapped_bubbles.append(BubbleLocation(q_num, options[opt_idx], (bubble[0], bubble[1]), bubble[2]))
+
+ if row_idx < 10 and len(col3) >= 4:
+ col3_sorted = sorted(col3, key=lambda b: b[0])[:4]
+ q_num = row_idx + 41
+ for opt_idx, bubble in enumerate(col3_sorted):
+ mapped_bubbles.append(BubbleLocation(q_num, options[opt_idx], (bubble[0], bubble[1]), bubble[2]))
+
+ return mapped_bubbles
+
+ def analyze_bubble_fill(self, bubble: BubbleLocation):
+ mask = np.zeros(self.gray.shape, dtype=np.uint8)
+ cv2.circle(mask, bubble.center, max(bubble.radius - 5, 5), 255, -1)
+ mean_val = cv2.mean(self.gray, mask=mask)[0]
+
+ large_ring_mask = np.zeros(self.gray.shape, dtype=np.uint8)
+ cv2.circle(large_ring_mask, bubble.center, bubble.radius + 10, 255, -1)
+ cv2.circle(large_ring_mask, bubble.center, bubble.radius + 5, 0, -1)
+ surrounding_mean = cv2.mean(self.gray, mask=large_ring_mask)[0]
+
+ bubble.darkness_score = surrounding_mean - mean_val
+ darkness_threshold = 50
+ absolute_darkness_threshold = 150 # 150
+ bubble.filled = (bubble.darkness_score > darkness_threshold) and (mean_val < absolute_darkness_threshold)
+
+ pixels = self.gray[mask > 0]
+ if len(pixels) > 0:
+ std_dev = np.std(pixels)
+ if std_dev > 25 and mean_val < 170:
+ bubble.filled = True
+ if mean_val < 120:
+ bubble.filled = True
+
+ return bubble.filled
+
+ def process(self):
+ """Main processing pipeline"""
+ print("Starting corrected OMR processing...")
+ print("Detecting bubble locations...")
+
+ all_bubbles = []
+ circles = self.find_bubble_grid()
+ all_bubbles.extend(circles)
+ contour_bubbles = self.detect_bubbles_by_contours()
+ all_bubbles.extend(contour_bubbles)
+ print(f" Contour bubbles found: {len(contour_bubbles)}")
+ print(f"Total bubbles detected: {len(all_bubbles)}")
+
+ if len(all_bubbles) < 180:
+ print("Not enough bubbles detected, using grid-based approach...")
+ grid_bubbles = self.detect_by_grid_assumption()
+ all_bubbles.extend(grid_bubbles)
+ print(f"Added {len(grid_bubbles)} bubbles from grid assumption")
+
+ print("Organizing bubbles into grid...")
+ bubble_rows = self.organize_and_filter_bubbles(all_bubbles)
+ print(f"Organized into {len(bubble_rows)} rows")
+
+ print("Mapping bubbles to questions...")
+ self.bubbles = self.map_to_questions(bubble_rows)
+ print(f"Mapped {len(self.bubbles)} bubble locations")
+
+ print("Analyzing filled bubbles...")
+ for bubble in self.bubbles:
+ self.analyze_bubble_fill(bubble)
+
+ print("Extracting final answers...")
+ self.extract_answers()
+
+ return self.answers
+
+ def detect_by_grid_assumption(self):
+ bubbles = []
+ col_starts = [60, 360, 660]
+ bubble_spacing_x = 45
+ bubble_spacing_y = 28
+ start_y = 50
+
+ for col_idx, col_x in enumerate(col_starts):
+ num_rows = 20 if col_idx < 2 else 10
+ for row in range(num_rows):
+ y = start_y + row * bubble_spacing_y
+ for opt in range(4):
+ x = col_x + opt * bubble_spacing_x
+ exists = False
+ for existing in bubbles:
+ if np.sqrt((x - existing[0])**2 + (y - existing[1])**2) < 20:
+ exists = True
+ break
+ if not exists:
+ bubbles.append((x, y, 13))
+ return bubbles
+
+ def extract_answers(self):
+ questions = defaultdict(list)
+ for bubble in self.bubbles:
+ questions[bubble.question_num].append(bubble)
+
+ self.answers = {}
+ for q_num in sorted(questions.keys()):
+ q_bubbles = questions[q_num]
+ filled = [b for b in q_bubbles if b.filled]
+
+ if not filled:
+ self.answers[q_num] = "---"
+ elif len(filled) == 1:
+ self.answers[q_num] = filled[0].option
+ else:
+ filled.sort(key=lambda b: b.darkness_score, reverse=True)
+ self.answers[q_num] = filled[0].option
+
+ return self.answers
+
+ def visualize_results(self):
+ result_img = self.image.copy()
+ for bubble in self.bubbles:
+ if bubble.filled:
+ cv2.circle(result_img, bubble.center, bubble.radius, (0, 255, 0), 2)
+ text = f"Q{bubble.question_num}:{bubble.option}"
+ cv2.putText(result_img, text,
+ (bubble.center[0] - 25, bubble.center[1] - bubble.radius - 5),
+ cv2.FONT_HERSHEY_SIMPLEX, 0.3, (0, 0, 255), 1)
+ else:
+ cv2.circle(result_img, bubble.center, bubble.radius, (100, 100, 255), 1)
+ return result_img
+
+ def display_results(self):
+ print("\n" + "="*60)
+ print("DETECTED ANSWERS")
+ print("="*60)
+
+ for i in range(1, 21):
+ row_str = ""
+ ans1 = self.answers.get(i, "---")
+ row_str += f"Q{i:2d}: {ans1:^4} | "
+
+ if i + 20 <= 40:
+ ans2 = self.answers.get(i + 20, "---")
+ row_str += f"Q{i+20:2d}: {ans2:^4} | "
+ else:
+ row_str += " " * 13 + "| "
+
+ if i + 40 <= 50:
+ ans3 = self.answers.get(i + 40, "---")
+ row_str += f"Q{i+40:2d}: {ans3:^4}"
+
+ print(row_str)
+
+ print("\n" + "="*60)
+ print("SUMMARY")
+ print("="*60)
+ answered = sum(1 for v in self.answers.values() if v != "---")
+ print(f"Questions detected: {len(self.answers)}")
+ print(f"Answered: {answered}")
+ print(f"Unanswered: {len(self.answers) - answered}")
+
+
+
+
+def process_single_image(image_data) -> Dict[str, Any]:
+ """Process a single image and return results with fixed indexing"""
+ try:
+ # Convert image data to numpy array
+ if isinstance(image_data, str):
+ # Base64 encoded image
+ image_bytes = base64.b64decode(image_data)
+ image = Image.open(io.BytesIO(image_bytes))
+ image_array = cv2.cvtColor(np.array(image), cv2.COLOR_RGB2BGR)
+ else:
+ # Direct file upload
+ image = Image.open(image_data)
+ image_array = cv2.cvtColor(np.array(image), cv2.COLOR_RGB2BGR)
+
+ # Process the image using the new CorrectedOMRReader
+ reader = CorrectedOMRReader(image_array=image_array)
+ answers = reader.process()
+
+ # No need for indexing fix in the new implementation
+ fixed_answers = answers
+
+ # Calculate CORRECTED statistics for 50 questions total
+ total_questions = 50 # Fixed to always be 50
+ answered = sum(1 for v in fixed_answers.values() if v is not None)
+ unanswered = total_questions - answered
+
+ # Format answers for JSON (convert None to "null" string)
+ formatted_answers = {}
+ for q_num in range(1, total_questions + 1):
+ answer = fixed_answers.get(q_num)
+ formatted_answers[str(q_num)] = answer if answer is not None else "null"
+
+ return {
+ "success": True,
+ "answers": formatted_answers,
+ "summary": {
+ "total_questions": total_questions,
+ "answered": answered,
+ "unanswered": unanswered
+ }
+ }
+
+ except Exception as e:
+ return {
+ "success": False,
+ "error": str(e),
+ "answers": {},
+ "summary": {
+ "total_questions": 50,
+ "answered": 0,
+ "unanswered": 50
+ }
+ }
+
+
+
+
+@app.route('/health', methods=['GET'])
+def health_check():
+ """Health check endpoint"""
+ return jsonify({
+ "status": "healthy",
+ "message": "OMR API is running"
+ })
+
+@app.route('/', methods=['GET'])
+def home():
+ """Home endpoint with API documentation"""
+ return jsonify({
+ "message": "OMR Processing API",
+ "version": "1.0",
+ "endpoints": {
+ "/process_omr": {
+ "method": "POST",
+ "description": "Process OMR answer sheets",
+ "accepts": [
+ "Multipart form data with 'images' field",
+ "JSON with base64 encoded images in 'images' array"
+ ],
+ "returns": "JSON with detected answers and summary"
+ },
+ "/health": {
+ "method": "GET",
+ "description": "Health check endpoint"
+ }
+ },
+ "example_response": {
+ "success": True,
+ "answers": {
+ "1": "A",
+ "2": "B",
+ "3": "null"
+ },
+ "summary": {
+ "total_questions": 50,
+ "answered": 45,
+ "unanswered": 5
+ }
+ }
+ })
+
+
+# <----------------->
+
+@app.route('/easyocr', methods=['POST'])
+def easyocr_image():
+ if 'images' not in request.files:
+ return jsonify({'error': 'No image files provided'}), 400
+
+ images = request.files.getlist('images')
+ extracted_texts = []
+
+ for image_file in images:
+ try:
+ # Save the image to a temporary file
+ with tempfile.NamedTemporaryFile(delete=False, suffix='.png') as temp_image_file:
+ image_file.save(temp_image_file.name)
+ temp_path = temp_image_file.name
+
+ try:
+ image_np = np.frombuffer(open(temp_path, 'rb').read(), np.uint8)
+ image = cv2.imdecode(image_np, cv2.IMREAD_COLOR)
+
+ # Perform OCR
+ result = reader.readtext(image)
+
+ # Extract text from the result
+ text = " ".join([item[1] for item in result])
+ extracted_texts.append(text)
+ ocr_extracted_texts.append(text)
+ finally:
+ # Clean up temp file
+ if os.path.exists(temp_path):
+ os.unlink(temp_path)
+ except Exception as e:
+ extracted_texts.append(f"Error processing image with EasyOCR: {str(e)}")
+
+ return jsonify({'extracted_texts': extracted_texts})
+
+@app.route('/tesseract', methods=['POST'])
+def tesseract_image():
+ if 'images' not in request.files:
+ return jsonify({'error': 'No image files provided'}), 400
+
+ images = request.files.getlist('images')
+ extracted_texts = []
+
+ for image_file in images:
+ try:
+ # Save the image to a temporary file
+ with tempfile.NamedTemporaryFile(delete=False, suffix='.png') as temp_image_file:
+ image_file.save(temp_image_file.name)
+ temp_path = temp_image_file.name
+
+ try:
+ with Image.open(temp_path) as image:
+ # Perform OCR using Tesseract
+ text = pytesseract.image_to_string(image)
+ extracted_texts.append(text.strip())
+ ocr_extracted_texts.append(text.strip())
+ finally:
+ # Clean up the temporary file
+ if os.path.exists(temp_path):
+ os.unlink(temp_path)
+ except Exception as e:
+ extracted_texts.append(f"Error processing image with Tesseract: {str(e)}")
+
+ return jsonify({'extracted_texts': extracted_texts})
+
+@app.route('/process_question_paper', methods=['POST'])
+def process_question_paper():
+ global last_processed_question_paper_object
+
+ if 'file' not in request.files:
+ return jsonify({'error': 'No file provided'}), 400
+
+ file = request.files['file']
+ if file.filename == '':
+ return jsonify({'error': 'No file selected'}), 400
+
+ question_paper = QuestionPaper()
+
+ try:
+ # Create Images directory if it doesn't exist
+ images_dir = os.path.join(app.root_path, 'Images')
+ os.makedirs(images_dir, exist_ok=True)
+
+ if file.filename.lower().endswith('.pdf'):
+ question_paper_filename = "question_paper.pdf"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ # Initialize the global object with the path
+ question_paper.path = question_paper_path
+
+ # For PDF processing
+ images_from_pdf = convert_from_path(question_paper_path, poppler_path=_get_poppler_path())
+
+ all_text = ""
+ for page_image in images_from_pdf:
+ text = pytesseract.image_to_string(page_image)
+ all_text += text + "\n"
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(all_text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ else:
+ # Process as image
+ question_paper_filename = "question_paper.png"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ question_paper.path = question_paper_path
+
+ image = Image.open(question_paper_path)
+ text = pytesseract.image_to_string(image)
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ # Clean the answers (remove any remaining unwanted patterns)
+ question_paper.clean_answers()
+
+ # Store the processed question paper globally
+ last_processed_question_paper_object = question_paper
+
+ return jsonify(question_paper.to_dict())
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+
+def gemini_evaluate_answer_sheet_with_roll(question_paper_path, student_answer_path, questions, correct_answers, paddle_results):
+ """
+ Evaluate entire answer sheet using Gemini and extract roll number
+ """
+ try:
+ model = genai.GenerativeModel('gemini-2.5-flash')
+
+ # Create the expected answers list for the prompt
+ expected_answers_text = "\n".join([f"{i+1}. {answer}" for i, answer in enumerate(correct_answers)])
+
+ prompt_text = f"""You are an OCR Assitant for an evaluvation script.
+ You will be given an image of a question paper and an image of a student's handwritten answers along with traditional OCR evaluvations.
+ Your task is assist the traditional OCR in overcoming its limitation with handwritten text the image may have bad quality handwritten text which the OCR may fail to extract and evaluvate properly, this is where you come in.
+ Your task is to Just do a double check of the OCR results and correct any mistakes or missing answers. and provide the result in a structured way.
+
+ Expected correct answers:
+ {expected_answers_text}
+ Traditional OCR Evaluation Results:
+ {paddle_results}
+
+ Instructions:
+ - First, identify and extract the student's roll number from the answer sheet
+ - Compare the student's handwritten answers with the expected answers above
+ - Small spelling mistakes should be ignored and considered correct
+ - If an answer has been crossed out or strikethrough, consider it incorrect
+ - Be lenient with handwriting recognition issues
+ - Look for answers by question numbers (1, 2, 3, etc.)
+
+ Please evaluate ALL questions and respond in this EXACT JSON format:
+ {{
+ "roll_number": "extracted_roll_number_here",
+ "evaluations": [
+ {{"question_number": 1, "status": "Correct"}},
+ {{"question_number": 2, "status": "Wrong"}},
+ {{"question_number": 3, "status": "Missing"}},
+ ...
+ ]
+ }}
+
+ For roll_number: Look for patterns like "Roll No:", "Roll Number:", "Reg No:", or any number sequence that appears to be a student identifier.
+
+ For each question, use ONLY one of these three status values:
+ - "Correct" - if the student's answer matches the expected answer (allowing for minor spelling)
+ - "Wrong" - if the student's answer is clearly different from the expected answer
+ - "Missing" - if no answer is visible for this question number
+
+ Respond with ONLY the JSON format above, no other text.
+
+ ! Note
+ Ignore texts like `GENERAL KNOWLEDGE QUESTION PAPER WITH ANSWERS` and the final output should only have actual questions.
+
+ """
+
+ # Handle PDF vs Image for question paper
+ if question_paper_path.lower().endswith('.pdf'):
+ # Convert PDF to images
+ pdf_images = convert_from_path(question_paper_path, poppler_path=_get_poppler_path())
+ question_paper_img = pdf_images[0] # Use first page
+ else:
+ question_paper_img = Image.open(question_paper_path)
+
+ # Load student answer image
+ student_answer_img = Image.open(student_answer_path)
+
+ # Create content for the model
+ content = [prompt_text, question_paper_img, student_answer_img]
+
+ response = model.generate_content(content)
+ result_text = response.text.strip()
+
+ print(f"Gemini response: {result_text}")
+
+ # Try to parse JSON response
+ import json
+ try:
+ # Clean the response - sometimes Gemini adds markdown formatting
+ if "```json" in result_text:
+ result_text = result_text.split("```json")[1].split("```")[0].strip()
+ elif "```" in result_text:
+ result_text = result_text.split("```")[1].strip()
+
+ parsed_result = json.loads(result_text)
+ return parsed_result["roll_number"], parsed_result["evaluations"]
+ except (json.JSONDecodeError, KeyError) as e:
+ print(f"Failed to parse JSON response: {e}")
+ print(f"Raw response: {result_text}")
+ # Fallback - extract roll number using OCR and create default "Error" results
+ roll_number = extract_roll_number(student_answer_path)
+ return roll_number, [{"question_number": i+1, "status": "Error"} for i in range(len(correct_answers))]
+
+ except Exception as e:
+ print(f"Error in Gemini evaluation: {str(e)}")
+ # Return error status for all questions with OCR extracted roll number
+ roll_number = extract_roll_number(student_answer_path)
+ return roll_number, [{"question_number": i+1, "status": "Error"} for i in range(len(correct_answers))]
+
+def quick_match(correct_list, messy_student_list, min_score=80):
+ """Quick function to match messy student answers"""
+ from fuzzywuzzy import process
+ import re
+
+ results = []
+ used = set()
+
+ for item in messy_student_list:
+ # Extract content
+ content = re.sub(r'^\d+\.?\s*', '', str(item)).strip()
+ if content and content != '-':
+ # Find best match
+ match = process.extractOne(content, correct_list)
+ if match and match[1] >= min_score:
+ q_num = correct_list.index(match[0]) + 1
+ if q_num not in used:
+ used.add(q_num)
+ results.append((item, q_num, match[0], match[1]))
+
+ return results
+
+def process_with_paddle_ocr(image_path, correct_answers):
+ """
+ Process an image with PaddleOCR and perform similarity matching with correct answers
+ Returns:
+ tuple: (extracted_text, similarity_scores, average_similarity)
+ """
+ try:
+ # Initialize PaddleOCR
+ from paddleocr import PaddleOCR
+ print("Initializing PaddleOCR...")
+ ocr = PaddleOCR(
+ use_doc_orientation_classify=True,
+ use_doc_unwarping=False,
+ use_textline_orientation=False
+ )
+ print("PaddleOCR initialized.")
+ # Read and process the image
+ # result = ocr.ocr(image_path, cls=True)
+ print("Preditcing")
+ result = ocr.predict(image_path)
+ print("PaddleOCR processing completed.")
+ # print(f"PaddleOCR result: {result}")
+ print("Correct Answers are:")
+ print(correct_answers)
+ for res in result:
+ words = res["rec_texts"]
+ print(f"PaddleOCR extracted words: {words}")
+ # words = result["rec_texts"]
+ result = quick_match(correct_answers, words, min_score=85)
+ print(f"PaddleOCR matched results: {result}")
+
+ return result
+
+ except Exception as e:
+ print(f"Error in PaddleOCR processing: {str(e)}")
+ return None, [], 0
+
+# OCR Evaluvation Endpoint
+@app.route('/evaluate_answers', methods=['POST'])
+def evaluate_answers():
+ global ocr_extracted_texts
+ if 'student_answers' not in request.files:
+ return jsonify({"error": "Missing student answers"}), 400
+
+ student_answer_files = request.files.getlist('student_answers')
+
+ # Get teacher email and exam name from the request
+ teacher_email = request.form.get('teacher_email', 'unknown@example.com')
+ exam_name = request.form.get('exam_name', 'Untitled Exam') # Get exam name from form data
+
+ # Retrieve the question paper object
+ question_paper = last_processed_question_paper_object
+
+ if last_processed_question_paper_object is None:
+ return jsonify({'error': 'Question paper not found or processed yet'}), 404
+
+ student_answer_paths = []
+ try:
+ # Save student answer files temporarily
+ for student_answer_file in student_answer_files:
+ with tempfile.NamedTemporaryFile(delete=False, suffix='.png') as temp_ans_file:
+ student_answer_file.save(temp_ans_file.name)
+ student_answer_paths.append(temp_ans_file.name)
+
+ # Process each student's answer sheet
+ all_students_results = []
+
+ if question_paper.path and os.path.exists(question_paper.path):
+ print(f"Starting Gemini evaluation for exam: {exam_name} with {len(student_answer_paths)} students...")
+
+ for idx, student_answer_path in enumerate(student_answer_paths):
+ print(f"Processing answer sheet {idx + 1} with PaddleOCR...")
+
+ # First process with PaddleOCR
+ results = process_with_paddle_ocr(
+ student_answer_path,
+ question_paper.answers
+ )
+
+ roll_number, sheet_evaluations = gemini_evaluate_answer_sheet_with_roll(
+ question_paper.path,
+ student_answer_path,
+ question_paper.questions,
+ question_paper.answers,
+ results
+ )
+
+
+
+
+ # Process the results for this student
+ student_results = []
+ for eval_result in sheet_evaluations:
+ question_num = eval_result["question_number"]
+ if 1 <= question_num <= len(question_paper.questions):
+ student_results.append({
+ 'question_number': question_num,
+ 'question_text': question_paper.questions[question_num - 1],
+ 'correct_answer': question_paper.answers[question_num - 1],
+ 'status': eval_result["status"]
+ })
+
+ # Calculate summary for this student
+ correct_count = sum(1 for result in student_results if result['status'] == 'Correct')
+ total_questions = len(student_results)
+ score_percentage = (correct_count / total_questions) * 100 if total_questions > 0 else 0
+
+ student_summary = {
+ 'roll_number': roll_number,
+ 'total_questions': len(question_paper.answers),
+ 'correct_answers': correct_count,
+ 'wrong_answers': sum(1 for result in student_results if result['status'] == 'Wrong'),
+ 'missing_answers': sum(1 for result in student_results if result['status'] == 'Missing'),
+ 'error_answers': sum(1 for result in student_results if result['status'] == 'Error'),
+ 'score_percentage': round(score_percentage, 2),
+ 'evaluation_results': student_results,
+ 'ocr_results': {
+ 'extracted_text': results,
+ }
+ }
+
+ all_students_results.append(student_summary)
+
+ final_results = {
+ 'exam_name': exam_name, # Include exam name in results
+ 'total_students': len(student_answer_paths),
+ 'students_evaluated': all_students_results
+ }
+
+ # STORE THE RESULTS IN SUPABASE WITH EXAM NAME
+ try:
+ supabase_handler = SupabaseHandler()
+ unique_key = supabase_handler.store_evaluation_result(teacher_email, final_results, exam_name)
+
+ if unique_key:
+ # Add the unique key to the response
+ final_results['unique_key'] = unique_key
+ final_results['storage_success'] = True
+ print(f"Results stored successfully with key: {unique_key} for exam: {exam_name}")
+ else:
+ final_results['storage_success'] = False
+ final_results['storage_error'] = "Failed to store results in database"
+ print("Failed to store results in Supabase")
+
+ except Exception as storage_error:
+ print(f"Error storing results: {str(storage_error)}")
+ final_results['storage_success'] = False
+ final_results['storage_error'] = str(storage_error)
+
+ return jsonify(final_results)
+ else:
+ return jsonify({
+ 'error': 'Question paper file not found for Gemini evaluation.'
+ })
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+ finally:
+ # Clean up temporary student answer files
+ for path in student_answer_paths:
+ try:
+ if os.path.exists(path):
+ os.unlink(path)
+ except PermissionError:
+ pass # File still locked on Windows; OS will clean up temp dir
+
+# Get Evaluation
+@app.route('/get_evaluation_result/', methods=['GET'])
+def get_evaluation_result(unique_key):
+ """
+ Get evaluation result by unique key
+ """
+ try:
+ supabase_handler = SupabaseHandler()
+ result = supabase_handler.get_evaluation_result(unique_key)
+
+ if result:
+ return jsonify({
+ 'success': True,
+ 'data': result
+ })
+ else:
+ return jsonify({
+ 'error': 'Evaluation result not found'
+ }), 404
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+
+# Get Teacher Evaluation
+@app.route('/get_teacher_evaluations/', methods=['GET'])
+def get_teacher_evaluations(teacher_email):
+ """
+ Get all evaluation results for a specific teacher
+ """
+ try:
+ supabase_handler = SupabaseHandler()
+ results = supabase_handler.get_teacher_evaluations(teacher_email)
+
+ return jsonify({
+ 'success': True,
+ 'data': results,
+ 'total_evaluations': len(results)
+ })
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+
+# Get OMR Answer Key
+@app.route('/get_omr_answer_key', methods=['GET'])
+def get_omr_answer_key():
+ """Get the currently stored OMR answer key"""
+ global last_processed_omr_key
+
+ if last_processed_omr_key is None:
+ return jsonify({
+ 'error': 'No answer key has been processed yet'
+ }), 404
+
+ return jsonify({
+ 'success': True,
+ 'answer_key': last_processed_omr_key.to_dict()
+ })
+
+def omr_gemini_process(error_questions, correct_answers, image_file):
+ """
+ Use Gemini to assist in evaluating OMR sheets, especially for error questions
+ """
+ try:
+
+ model = genai.GenerativeModel('gemini-2.5-flash')
+
+ prompt_text = f"""
+ You are an OMR Assistant for an evaluvation script.
+ Your main purpose is to assist in the process.
+
+ Correct Answers to questions sorted by question number: {correct_answers}
+ Error Question numbers: {error_questions}
+ Your task:
+ - From the given image identify the student name and roll number
+ - if for some reason the traditional OMR Processing failed to detect some answers, those question numbers will be provided to you, you should look into those questions form the given image and correct answers.
+ - Only provide answer for the questions that are in the error list.
+ - You can ignore the rest of the question
+ - if Error question is empty, just extract the roll number and name
+
+
+ Please evaluate ALL questions and respond in this EXACT JSON format:
+ {{
+ "roll_number": "extracted_roll_number_here",
+ "evaluations": [
+ {{"question_number": 1, "status": "Correct"}},
+ {{"question_number": 2, "status": "Wrong"}},
+ {{"question_number": 3, "status": "Missing"}},
+ ...
+ ]
+ }}
+
+ """
+ student_answer_img = image_file
+
+ content = [prompt_text, student_answer_img]
+
+ response = model.generate_content(content)
+ result_text = response.text.strip()
+
+ print(f"Gemini response: {result_text}")
+
+ import json
+ try:
+ # Clean the response - sometimes Gemini adds markdown formatting
+ if "```json" in result_text:
+ result_text = result_text.split("```json")[1].split("```")[0].strip()
+ elif "```" in result_text:
+ result_text = result_text.split("```")[1].strip()
+
+ parsed_result = json.loads(result_text)
+ return parsed_result["roll_number"], parsed_result["evaluations"]
+ except (json.JSONDecodeError, KeyError) as e:
+ print(f"Failed to parse JSON response: {e}")
+ print(f"Raw response: {result_text}")
+ # Fallback - extract roll number using OCR and create default "Error" results
+ roll_number = extract_roll_number(os.path.join("OMRChecker", "inputs", "OMRImage.jpg"))
+ return roll_number, [{"question_number": i+1, "status": "Error"} for i in range(len(correct_answers))]
+
+ except Exception as e:
+ print(f"Error in OMR Gemini processing: {str(e)}")
+ return "Unknown", [{"question_number": q, "status": "Error"} for q in error_questions]
+
+@app.route('/evaluate_omr', methods=['POST'])
+def evaluate_omr():
+ """
+ Evaluate OMR answers against stored answer key
+ """
+ global last_processed_omr_key, last_processed_omr_results, porcessed_omr_results, OMR_IMAGES
+
+ # Get teacher email and exam name from the request
+ teacher_email = request.form.get('teacher_email', 'unknown@example.com')
+ exam_name = request.form.get('exam_name', 'Untitled Exam') # Get exam name from form data
+ if not last_processed_omr_key:
+ return jsonify({
+ 'error': 'No answer key has been processed. Please process an answer key first.'
+ }), 400
+
+ if not last_processed_omr_results:
+ return jsonify({
+ 'error': 'No OMR sheet has been processed. Please process an OMR sheet first.'
+ }), 400
+
+ try:
+ # Get the marked answers from the processed OMR
+ if isinstance(last_processed_omr_results, list):
+ omr_data = last_processed_omr_results[0] # Take first sheet if multiple
+ else:
+ omr_data = last_processed_omr_results
+ student_datas = []
+ for idx, omr_data in enumerate(porcessed_omr_results):
+ marked_answers = omr_data
+ image_file = OMR_IMAGES[idx]
+
+ # Get correct answers from answer key (only for questions that exist)
+ correct_answers = last_processed_omr_key.answers
+ total_questions_in_key = len(correct_answers)
+
+ # Evaluate answers only for questions that exist in the answer key
+ evaluation_details = []
+ correct_count = 0
+ wrong_count = 0
+ missing_count = 0
+ error_questions = []
+
+ for q_num in sorted(correct_answers.keys()):
+ print(f"Evaluating Question {q_num}")
+ print(f"Correct Answer: {correct_answers[q_num]} | Marked Answer: {marked_answers.get(str(q_num))}")
+ correct_ans = correct_answers[q_num]
+ marked_ans = marked_answers.get(str(q_num))
+
+ if marked_ans is None or marked_ans == '' or len(str(marked_ans)) > 1 or marked_ans == 'nan':
+ status = 'Missing'
+ error_questions.append(q_num)
+ missing_count += 1
+ elif marked_ans.upper() == correct_ans.upper():
+ status = 'Correct'
+ correct_count += 1
+ else:
+ status = 'Wrong'
+ wrong_count += 1
+
+ evaluation_details.append({
+ 'question_number': q_num,
+ 'question_text': last_processed_omr_key.questions[q_num - 1] if q_num <= len(last_processed_omr_key.questions) else f"Question {q_num}",
+ 'correct_answer': correct_ans,
+ 'marked_answer': marked_ans if marked_ans != 'null' else None,
+ 'status': status
+ })
+
+ roll_no, gemini_result = omr_gemini_process(
+ error_questions,
+ last_processed_omr_key.answers,
+ image_file
+ )
+
+ for err_idx in error_questions:
+ for gemini_eval in gemini_result:
+ if gemini_eval["question_number"] == err_idx:
+ correct_ans = last_processed_omr_key.answers[err_idx]
+ marked_ans = None # Since it was an error question
+ status = gemini_eval["status"]
+
+ if status == "Correct":
+ correct_count += 1
+ # wrong_count -= 1 # Adjust wrong count
+ missing_count -= 1 # Adjust missing count
+ elif status == "Wrong":
+ wrong_count += 1
+ missing_count -= 1 # Adjust missing count
+ elif status == "Missing":
+ missing_count += 1
+
+ # Update the evaluation details
+ for eval_detail in evaluation_details:
+ if eval_detail['question_number'] == err_idx:
+ eval_detail.update({
+ 'marked_answer': marked_ans,
+ 'status': status
+ })
+ break
+ break
+
+ # Calculate score
+ total_score = correct_count * last_processed_omr_key.marks_per_question
+
+ if last_processed_omr_key.negative_marking > 0:
+ total_score -= wrong_count * last_processed_omr_key.negative_marking
+
+ max_score = total_questions_in_key * last_processed_omr_key.marks_per_question
+ student_summary = {
+ 'roll_number': roll_no,
+ 'total_questions': len(last_processed_omr_key.answers),
+ 'correct_answers': correct_count,
+ 'wrong_answers': wrong_count,
+ 'missing_answers': missing_count,
+ 'error_answers': len(error_questions),
+ 'score_percentage': correct_count / len(last_processed_omr_key.answers) * 100 if len(last_processed_omr_key.answers) > 0 else 0,
+ 'evaluation_results': evaluation_details,
+ 'ocr_results': {
+ 'extracted_text': gemini_result,
+ }
+ }
+ student_datas.append(student_summary)
+
+ # Format the data in the required structure for Supabase
+ formatted_evaluation_data = {
+ 'exam_name': exam_name, # Include exam name in results
+ 'total_students': len(student_datas),
+ 'students_evaluated': student_datas
+ }
+
+ # Store results in Supabase (optional — skip if credentials not configured)
+ unique_key = None
+ try:
+ supabase_handler = SupabaseHandler()
+ unique_key = supabase_handler.store_evaluation_result(teacher_email, formatted_evaluation_data, exam_name)
+ except Exception as supa_err:
+ print(f"Supabase storage skipped: {supa_err}")
+
+ # Prepare answer key info
+ answer_key_info = {
+ "title": getattr(last_processed_omr_key, 'title', 'Untitled'),
+ "marks_per_question": last_processed_omr_key.marks_per_question,
+ "negative_marking": last_processed_omr_key.negative_marking
+ }
+
+ # Return response in the same format as stored in Supabase
+ final_result = {
+ "success": True,
+ "unique_key": unique_key,
+ #**formatted_evaluation_data, # Include all the formatted data
+ "additional_info": {
+ "answer_key_info": answer_key_info
+ }
+ }
+
+ return jsonify(final_result)
+
+ except Exception as e:
+ return jsonify({
+ "success": False,
+ "error": f"Evaluation failed: {str(e)}"
+ }), 500
+
+def process_with_gemini(evaluation_details, evaluation_summary, omr_data):
+ """
+ Use Gemini to independently evaluate the OMR sheet and extract student details
+ """
+ global last_processed_omr_key
+
+ try:
+ model = genai.GenerativeModel('gemini-2.5-flash')
+
+ # Prepare the questions and correct answers for Gemini
+ questions_and_answers = ""
+ for i, (q_num, correct_answer) in enumerate(sorted(last_processed_omr_key.answers.items())):
+ question_text = last_processed_omr_key.questions[i] if i < len(last_processed_omr_key.questions) else f"Question {q_num}"
+ questions_and_answers += f"Question {q_num}: {question_text}\nCorrect Answer: {correct_answer}\n\n"
+
+ prompt = f"""
+ You are a teacher grading an OMR answer sheet.
+
+STUDENT INFO: Extract the student's name and roll number from the image.
+
+GRADING TASK: For each question, identify which bubble (A, B, C, or D) is filled/darkened, then compare with the correct answer.
+
+QUESTIONS AND CORRECT ANSWERS:
+{questions_and_answers}
+
+IMPORTANT: Look carefully at each row of bubbles. A filled bubble will be darkened/shaded, while empty bubbles will be white/clear.
+
+Respond in this EXACT JSON format:
+{{
+ "student_info": {{
+ "name": "extracted student name",
+ "roll_no": "extracted roll number"
+ }},
+ "gemini_evaluation": [
+ {{"question": 1, "marked_answer": "C", "correct_answer": "C", "status": "Correct"}},
+ {{"question": 2, "marked_answer": "D", "correct_answer": "D", "status": "Correct"}},
+ // ... continue for all questions
+ ]
+}}
+
+For status: use "Correct", "Wrong", or "Missing" only.
+For marked_answer: use "A", "B", "C", "D", or null if no bubble is clearly filled.
+
+ """
+ # Get the image - we need to retrieve it from the last processed OMR
+ # Since we don't store the image directly, we'll need to work with what we have
+ # For now, let's assume we have access to the image file
+
+ # Check if we have image data stored
+ if 'image_data' in omr_data:
+ # If we have base64 image data
+ image_data = omr_data['image_data']
+ image_bytes = base64.b64decode(image_data)
+ image = Image.open(io.BytesIO(image_bytes))
+ elif 'filename' in omr_data:
+ # Try to find the image file
+ try:
+ # Look for the image in common locations
+ possible_paths = [
+ f"Images/{omr_data['filename']}",
+ f"temp/{omr_data['filename']}",
+ omr_data['filename']
+ ]
+
+ image = None
+ for path in possible_paths:
+ if os.path.exists(path):
+ image = Image.open(path)
+ break
+
+ if image is None:
+ # If we can't find the image, return a fallback result
+ return {
+ "student_info": {
+ "name": "Image not available",
+ "roll_number": "Image not available"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": ["Original image not available for verification"],
+ "notes": "Could not verify due to missing image file"
+ },
+ "gemini_evaluation": []
+ }
+ except Exception as e:
+ print(f"Error loading image: {str(e)}")
+ return {
+ "student_info": {
+ "name": "Error loading image",
+ "roll_number": "Error loading image"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": [f"Error loading image: {str(e)}"],
+ "notes": "Image processing failed"
+ },
+ "gemini_evaluation": []
+ }
+ else:
+ # No image reference available
+ return {
+ "student_info": {
+ "name": "No image data",
+ "roll_number": "No image data"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": ["No image data available"],
+ "notes": "Cannot verify without image"
+ },
+ "gemini_evaluation": []
+ }
+
+ # Generate content with Gemini
+ response = model.generate_content([prompt, image])
+ result_text = response.text.strip()
+
+ print(f"Gemini raw response: {result_text}")
+
+ # Parse the JSON response
+ try:
+ # Clean the response - remove markdown formatting if present
+ if "```json" in result_text:
+ result_text = result_text.split("```json")[1].split("```")[0].strip()
+ elif "```" in result_text:
+ result_text = result_text.split("```")[1].strip()
+
+ parsed_result = json.loads(result_text)
+
+ # Update summary counts and score based on the evaluation
+ if 'gemini_evaluation' in parsed_result:
+ correct_count = sum(1 for item in parsed_result['gemini_evaluation'] if item.get('status') == 'Correct')
+ wrong_count = sum(1 for item in parsed_result['gemini_evaluation'] if item.get('status') == 'Wrong')
+ missing_count = sum(1 for item in parsed_result['gemini_evaluation'] if item.get('status') == 'Missing')
+
+ score = (correct_count * last_processed_omr_key.marks_per_question) - (wrong_count * last_processed_omr_key.negative_marking)
+ max_score = len(last_processed_omr_key.answers) * last_processed_omr_key.marks_per_question
+
+ parsed_result['summary'] = {
+ "total_questions": len(last_processed_omr_key.answers),
+ "correct_count": correct_count,
+ "wrong_count": wrong_count,
+ "missing_count": missing_count,
+ "score": score,
+ "max_score": max_score,
+ "percentage": round((score / max_score) * 100, 2) if max_score > 0 else 0
+ }
+
+ return parsed_result
+
+ except json.JSONDecodeError as e:
+ print(f"Failed to parse Gemini JSON response: {e}")
+ print(f"Raw response: {result_text}")
+
+ # Fallback response with extracted text attempt
+ return {
+ "student_info": {
+ "name": "Parse error",
+ "roll_number": "Parse error"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": ["Failed to parse Gemini response"],
+ "notes": f"JSON parse error: {str(e)}"
+ },
+ "gemini_evaluation": [],
+ "raw_response": result_text # Include raw response for debugging
+ }
+ # print(f"Error in Gemini processing: {str(e)}")
+ except Exception as e:
+ return {
+ "student_info": {
+ "name": "Processing error",
+ "roll_number": "Processing error"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": [f"Gemini processing error: {str(e)}"],
+ "notes": "Failed to process with Gemini"
+ },
+ "gemini_evaluation": []
+ }
+def compare_evaluations(our_evaluation, gemini_evaluation):
+ """
+ Compare our automated evaluation with Gemini's independent evaluation
+ """
+ if not gemini_evaluation:
+ return {
+ "comparison_available": False,
+ "reason": "Gemini evaluation not available"
+ }
+
+ matches = 0
+ differences = []
+ total_compared = 0
+
+ # Create a lookup for our evaluation
+ our_eval_lookup = {detail['question_number']: detail for detail in our_evaluation}
+
+ for gemini_item in gemini_evaluation:
+ q_num = gemini_item.get('question')
+ if q_num in our_eval_lookup:
+ total_compared += 1
+ our_status = our_eval_lookup[q_num]['status']
+ gemini_status = gemini_item.get('status')
+
+ if our_status == gemini_status:
+ matches += 1
+ else:
+ differences.append({
+ "question": q_num,
+ "our_evaluation": {
+ "marked_answer": our_eval_lookup[q_num]['marked_answer'],
+ "status": our_status
+ },
+ "gemini_evaluation": {
+ "marked_answer": gemini_item.get('marked_answer'),
+ "status": gemini_status
+ }
+ })
+
+ agreement_rate = (matches / total_compared) * 100 if total_compared > 0 else 0
+
+ return {
+ "comparison_available": True,
+ "total_questions_compared": total_compared,
+ "agreements": matches,
+ "differences_count": len(differences),
+ "agreement_rate": round(agreement_rate, 2),
+ "differences": differences
+ }
+
+
+
+
+# Also need to modify the process_omr endpoint to store image data for later use
+@app.route('/process_omr', methods=['POST'])
+def process_omr_enhanced():
+ """
+ Enhanced OMR processing that stores image data for later Gemini processing
+ """
+ global last_processed_omr_results
+ global OMR_IMAGES
+ global porcessed_omr_results
+
+ OMR_IMAGES = []
+ porcessed_omr_results = []
+ try:
+ results = []
+ print("Starting OMR processing...")
+ # Check if files were uploaded
+ if 'images' in request.files:
+ files = request.files.getlist('images')
+ results = []
+ for idx, file in enumerate(files):
+ if file.filename == '':
+ continue
+
+ print(f"===================================== Processing file {file.filename} =====================================")
+ name, extension = os.path.splitext(file.filename)
+ filename = os.path.join("OMRChecker", "inputs", "OMRImage" + extension)
+ file.save(filename)
+ OMR_IMAGES.append(Image.open(filename))
+
+ result = subprocess.run([sys.executable, os.path.join('OMRChecker', 'main.py'), '--inputDir=' + os.path.join('OMRChecker', 'inputs')])
+
+ print("OMR Finished Processing Successfully")
+ folder = os.path.join("outputs", "Results")
+ csv_files = [f for f in os.listdir(folder) if f.endswith(".csv")]
+ print("CSV FILES:", csv_files)
+ result_file = os.path.join(folder, csv_files[0])
+ print("Found Result File", result_file)
+ df = pd.read_csv(result_file)
+
+ # Convert to JSON
+ data_json = df.to_json(orient="records")
+ parsed_json = json.loads(data_json)
+
+ columns_dict = df.to_dict(orient="list")
+
+ print(columns_dict)
+ questions_only = {k.replace("q", ""): v[0] for k, v in columns_dict.items() if k.startswith("q")}
+
+ last_processed_omr_results = questions_only
+ porcessed_omr_results.append(questions_only)
+
+ if os.path.exists(result_file):
+ os.remove(result_file)
+ print(f"{result_file} deleted")
+
+ return jsonify(parsed_json)
+
+ else:
+ return jsonify({
+ "success": False,
+ "error": "No images provided. Use 'images' field for file uploads.",
+ "results": []
+ }), 400
+
+ except Exception as e:
+ return jsonify({
+ "success": False,
+ "error": f"Server error: {str(e)}",
+ "results": []
+ }), 500
+
+@app.route('/get_question_details/', methods=['GET'])
+def get_question_details(question_number):
+ """Get detailed information about a specific question"""
+ global last_processed_omr_key
+
+ if last_processed_omr_key is None:
+ return jsonify({
+ 'error': 'No answer key has been processed yet'
+ }), 404
+
+ question_data = last_processed_omr_key.get_question_details(question_number)
+ if question_data is None:
+ return jsonify({
+ 'error': f'Question number {question_number} not found'
+ }), 404
+
+ return jsonify({
+ 'success': True,
+ 'question_data': question_data
+ })
+
+@app.route('/debug_parsing', methods=['GET'])
+def debug_parsing():
+ """
+ Debug endpoint to see how OCR text is being parsed
+ """
+ if not ocr_extracted_texts:
+ return jsonify({'error': 'No OCR extracted texts available.'}), 400
+
+ debug_results = []
+
+ for ocr_text in ocr_extracted_texts:
+ parsed_answers = improved_clean_and_parse_ocr_text(ocr_text)
+ debug_results.append({
+ 'original_ocr_text': ocr_text,
+ 'parsed_answers': parsed_answers
+ })
+
+ return jsonify({'debug_results': debug_results})
+
+def extract_omr_metadata(text: str) -> tuple:
+ """Extract title and duration from the question paper text"""
+ title = ""
+ duration = ""
+
+ # Look for title (usually in first few lines, often in caps)
+ lines = text.split('\n')
+ for line in lines[:5]: # Check first 5 lines
+ if line.strip().upper() == line.strip() and len(line.strip()) > 10:
+ title = line.strip()
+ break
+
+ # Look for duration/time
+ time_pattern = r'Time:\s*(\d+)\s*(minutes|mins|min)'
+ duration_match = re.search(time_pattern, text, re.IGNORECASE)
+ if duration_match:
+ duration = f"{duration_match.group(1)} minutes"
+
+ return title, duration
+
+def extract_omr_answers(text: str) -> dict:
+ """Extract answers from the question paper text"""
+ answers = {}
+ questions = []
+ question_data = []
+ current_question = None
+
+ print("\nStarting answer extraction...")
+
+ # Split text into lines and process line by line
+ lines = [line.strip() for line in text.split('\n') if line.strip()]
+
+ # Skip header lines until we find the first question
+ started = False
+ current_dict = None
+
+ for line in lines:
+ print(f"Processing line: {line}")
+
+ # Skip header or empty lines
+ if not started:
+ if line.startswith('1.'):
+ started = True
+ else:
+ continue
+
+ # Check for new question
+ question_match = re.match(r'^(\d+)[.)](.*?)$', line)
+ if question_match:
+ # Save previous question if exists
+ if current_dict:
+ question_data.append(current_dict)
+
+ # Start new question
+ q_num = int(question_match.group(1))
+ q_text = question_match.group(2).strip()
+ current_dict = {
+ 'number': q_num,
+ 'question': q_text,
+ 'options': {},
+ 'answer': None
+ }
+ continue
+
+ # Check for options
+ option_match = re.match(r'^([A-D])[).](.*?)$', line)
+ if option_match and current_dict is not None:
+ opt_letter = option_match.group(1)
+ opt_text = option_match.group(2).strip()
+ current_dict['options'][opt_letter] = opt_text
+ continue
+
+ # Check for answer
+ answer_match = re.match(r'^\s*Answer[:\s]*([A-D]|.+)$', line, re.IGNORECASE)
+ if answer_match and current_dict is not None:
+ answer = answer_match.group(1).strip()
+ # print(f"For Question: {current_dict['number']}, Options are:")
+ # print(current_dict['options'])
+ for opt_letter, opt_text in current_dict['options'].items():
+ if answer.lower() == opt_text.lower():
+ answer = opt_letter
+ break
+ current_dict['answer'] = answer
+ continue
+
+ # Add last question
+ if current_dict:
+ question_data.append(current_dict)
+
+ print("\nExtracted Question Data:")
+ for q in question_data:
+ print(f"\nQuestion {q['number']}:")
+ print(f"Text: {q['question']}")
+ print(f"Options: {q['options']}")
+ print(f"Answer: {q['answer']}")
+
+ # Add to return format
+ if q['answer']:
+ answers[q['number']] = q['answer']
+ questions.append(f"{q['number']}. {q['question']}")
+
+ print(f"\nExtracted {len(questions)} questions and {len(answers)} answers")
+ print("Questions:", questions)
+ print("Answers:", answers)
+
+ return answers, questions
+
+def debug_text_extraction(text: str):
+ """Helper function to debug text extraction issues"""
+ print("=== Extracted Text ===")
+ print(text)
+ print("\n=== Line by Line Analysis ===")
+ for line in text.split('\n'):
+ if line.strip():
+ print(f"Line: {line.strip()}")
+
+@app.route('/process_omr_answer_key', methods=['POST'])
+def process_omr_answer_key():
+ """
+ Process OMR answer key from either:
+ 1. JSON format with direct answers
+ 2. PDF/Image of question paper with answers marked
+
+ For JSON format:
+ {
+ "answers": {
+ "1": "A",
+ "2": "B",
+ ...
+ },
+ "marks_per_question": 1.0, # optional, defaults to 1
+ "negative_marking": 0.0 # optional, defaults to 0
+ }
+
+ For PDF/Image:
+ multipart/form-data with 'file' field containing the question paper
+ """
+ global last_processed_omr_key
+
+ try:
+ omr_key = OMRAnswerKey()
+
+ # Check if file upload or JSON
+ if 'file' in request.files:
+ file = request.files['file']
+ if file.filename == '':
+ return jsonify({'error': 'No file selected'}), 400
+
+ # Create Images directory if it doesn't exist
+ images_dir = os.path.join(app.root_path, 'Images')
+ os.makedirs(images_dir, exist_ok=True)
+
+ if file.filename.lower().endswith('.pdf'):
+ # Save and process PDF
+ answer_key_path = os.path.join(images_dir, "omr_answer_key.pdf")
+ file.save(answer_key_path)
+ omr_key.path = answer_key_path
+
+ # Convert PDF to images and extract text
+ all_text = ""
+ try:
+ print(f"\nProcessing PDF file: {answer_key_path}")
+ images_from_pdf = convert_from_path(
+ answer_key_path,
+ poppler_path=_get_poppler_path(),
+ dpi=300 # Increase DPI for better quality
+ )
+ print(f"Converted PDF to {len(images_from_pdf)} images")
+
+ for idx, page_image in enumerate(images_from_pdf):
+ print(f"\nProcessing page {idx + 1}")
+
+ # Preprocess the image for better OCR
+ # Convert to numpy array
+ img_np = np.array(page_image)
+
+ # Convert to grayscale
+ gray = cv2.cvtColor(img_np, cv2.COLOR_RGB2GRAY)
+
+ # Apply thresholding to get black and white image
+ _, threshold = cv2.threshold(gray, 0, 255, cv2.THRESH_BINARY + cv2.THRESH_OTSU)
+
+ # Save processed image for debugging
+ debug_image_path = os.path.join(images_dir, f"debug_page_{idx + 1}.png")
+ cv2.imwrite(debug_image_path, threshold)
+ print(f"Saved processed image to {debug_image_path}")
+
+ # Configure Tesseract parameters for better accuracy
+ custom_config = r'--oem 3 --psm 6'
+ text = pytesseract.image_to_string(threshold, config=custom_config)
+ print(f"Extracted text length: {len(text)}")
+ all_text += text + "\n"
+
+ print("\nTotal extracted text length:", len(all_text))
+
+ except Exception as e:
+ print(f"Error during PDF processing: {str(e)}")
+ raise
+
+ # Debug the extracted text
+ print("\nDebugging PDF extraction:")
+ debug_text_extraction(all_text)
+
+ # Extract metadata and answers
+ title, duration = extract_omr_metadata(all_text)
+ answers, questions = extract_omr_answers(all_text)
+
+ print("\nExtracted answers:", answers)
+
+ omr_key.set_metadata(title, duration)
+ omr_key.set_answers(answers)
+ omr_key.questions = questions
+
+ else:
+ # Process as image
+ answer_key_path = os.path.join(images_dir, "omr_answer_key.png")
+ file.save(answer_key_path)
+ omr_key.path = answer_key_path
+
+ image = Image.open(answer_key_path)
+ text = pytesseract.image_to_string(image)
+
+ # Debug the extracted text
+ print("\nDebugging Image extraction:")
+ debug_text_extraction(text)
+
+ # Extract metadata and answers
+ title, duration = extract_omr_metadata(text)
+ answers, questions = extract_omr_answers(text)
+
+ print("\nStructured Extraction Results:")
+ print("Title:", title)
+ print("Duration:", duration)
+ print("\nQuestions found:", len(questions))
+ print("Answers found:", len(answers))
+ print("\nAnswers:", answers)
+
+ omr_key.set_metadata(title, duration)
+ omr_key.set_answers(answers)
+ omr_key.questions = questions
+
+ # Set default marking scheme
+ marks_per_question = float(request.form.get('marks_per_question', 1.0))
+ negative_marking = float(request.form.get('negative_marking', 0.0))
+
+ else:
+ # Process JSON input
+ if not request.is_json:
+ return jsonify({'error': 'Request must be JSON or file upload'}), 400
+
+ data = request.get_json()
+
+ if 'answers' not in data:
+ return jsonify({'error': 'Answer key must be provided'}), 400
+
+ # Validate answer format
+ answer_key = data['answers']
+ for q_num, answer in answer_key.items():
+ try:
+ q_num = int(q_num)
+ if not isinstance(answer, str) or answer.upper() not in ['A', 'B', 'C', 'D']:
+ return jsonify({
+ 'error': f'Invalid answer format for question {q_num}. Must be A, B, C, or D'
+ }), 400
+ except ValueError:
+ return jsonify({
+ 'error': f'Question numbers must be integers, got {q_num}'
+ }), 400
+
+ # Set the answers
+ omr_key.set_answers(answer_key)
+
+ # Set metadata if provided
+ title = data.get('title', '')
+ duration = data.get('duration', '')
+ omr_key.set_metadata(title, duration)
+
+ # Set marking scheme
+ marks_per_question = float(data.get('marks_per_question', 1.0))
+ negative_marking = float(data.get('negative_marking', 0.0))
+
+ # Set marking scheme
+ omr_key.set_marking_scheme(marks_per_question, negative_marking)
+
+ # Store globally
+ last_processed_omr_key = omr_key
+
+ return jsonify({
+ 'success': True,
+ 'message': 'OMR answer key processed successfully',
+ 'answer_key': omr_key.to_dict()
+ })
+
+ except Exception as e:
+ return jsonify({
+ 'error': f'Failed to process answer key: {str(e)}'
+ }), 500
+
+if __name__ == '__main__':
+ app.run(
+ host="0.0.0.0",
+ port=int(os.environ.get("PORT", 5000)),
+ debug=os.environ.get("FLASK_DEBUG", "false").lower() == "true"
+ )
\ No newline at end of file
diff --git a/output/test_ocr_res_img.jpg b/output/test_ocr_res_img.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3a857a483398ec504cad6afd36a4b0d2f6e317b8
--- /dev/null
+++ b/output/test_ocr_res_img.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:31eac088a5ed8769b2837fc5064db5461b8dbb972264a44aa3c5693d81b70675
+size 562726
diff --git a/output/test_preprocessed_img.jpg b/output/test_preprocessed_img.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0d44308ec253f162d92fb2ad3d3141d64fcc2bfc
--- /dev/null
+++ b/output/test_preprocessed_img.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:99af06ba58b1a2ac6ae490f63ba0ce86b4df86dd20625abbdfa70a73c0de590d
+size 1318246
diff --git a/output/test_res.json b/output/test_res.json
new file mode 100644
index 0000000000000000000000000000000000000000..da0d897b478568b9d30fca643fb466695efe18da
--- /dev/null
+++ b/output/test_res.json
@@ -0,0 +1,805 @@
+{
+ "input_path": "Images\\test.jpg",
+ "page_index": null,
+ "model_settings": {
+ "use_doc_preprocessor": true,
+ "use_textline_orientation": false
+ },
+ "doc_preprocessor_res": {
+ "input_path": null,
+ "page_index": null,
+ "model_settings": {
+ "use_doc_orientation_classify": true,
+ "use_doc_unwarping": false
+ },
+ "angle": 0
+ },
+ "dt_polys": [
+ [
+ [
+ 209,
+ 389
+ ],
+ [
+ 426,
+ 386
+ ],
+ [
+ 427,
+ 446
+ ],
+ [
+ 210,
+ 449
+ ]
+ ],
+ [
+ [
+ 68,
+ 421
+ ],
+ [
+ 135,
+ 421
+ ],
+ [
+ 135,
+ 485
+ ],
+ [
+ 68,
+ 485
+ ]
+ ],
+ [
+ [
+ 220,
+ 498
+ ],
+ [
+ 381,
+ 492
+ ],
+ [
+ 383,
+ 567
+ ],
+ [
+ 222,
+ 572
+ ]
+ ],
+ [
+ [
+ 80,
+ 524
+ ],
+ [
+ 151,
+ 524
+ ],
+ [
+ 151,
+ 584
+ ],
+ [
+ 80,
+ 584
+ ]
+ ],
+ [
+ [
+ 88,
+ 630
+ ],
+ [
+ 146,
+ 636
+ ],
+ [
+ 138,
+ 712
+ ],
+ [
+ 80,
+ 706
+ ]
+ ],
+ [
+ [
+ 222,
+ 630
+ ],
+ [
+ 559,
+ 626
+ ],
+ [
+ 560,
+ 702
+ ],
+ [
+ 223,
+ 706
+ ]
+ ],
+ [
+ [
+ 175,
+ 744
+ ],
+ [
+ 520,
+ 738
+ ],
+ [
+ 522,
+ 814
+ ],
+ [
+ 177,
+ 820
+ ]
+ ],
+ [
+ [
+ 83,
+ 772
+ ],
+ [
+ 131,
+ 772
+ ],
+ [
+ 131,
+ 819
+ ],
+ [
+ 83,
+ 819
+ ]
+ ],
+ [
+ [
+ 209,
+ 859
+ ],
+ [
+ 628,
+ 838
+ ],
+ [
+ 632,
+ 914
+ ],
+ [
+ 213,
+ 935
+ ]
+ ],
+ [
+ [
+ 85,
+ 878
+ ],
+ [
+ 140,
+ 878
+ ],
+ [
+ 140,
+ 936
+ ],
+ [
+ 85,
+ 936
+ ]
+ ],
+ [
+ [
+ 202,
+ 997
+ ],
+ [
+ 343,
+ 989
+ ],
+ [
+ 348,
+ 1065
+ ],
+ [
+ 207,
+ 1073
+ ]
+ ],
+ [
+ [
+ 81,
+ 1009
+ ],
+ [
+ 149,
+ 1009
+ ],
+ [
+ 149,
+ 1081
+ ],
+ [
+ 81,
+ 1081
+ ]
+ ],
+ [
+ [
+ 209,
+ 1121
+ ],
+ [
+ 508,
+ 1110
+ ],
+ [
+ 512,
+ 1196
+ ],
+ [
+ 212,
+ 1207
+ ]
+ ],
+ [
+ [
+ 0,
+ 1132
+ ],
+ [
+ 2160,
+ 1052
+ ],
+ [
+ 2160,
+ 2938
+ ],
+ [
+ 0,
+ 2938
+ ]
+ ],
+ [
+ [
+ 77,
+ 1136
+ ],
+ [
+ 157,
+ 1136
+ ],
+ [
+ 157,
+ 1205
+ ],
+ [
+ 77,
+ 1205
+ ]
+ ],
+ [
+ [
+ 86,
+ 1277
+ ],
+ [
+ 501,
+ 1252
+ ],
+ [
+ 504,
+ 1314
+ ],
+ [
+ 90,
+ 1339
+ ]
+ ],
+ [
+ [
+ 80,
+ 1396
+ ],
+ [
+ 465,
+ 1366
+ ],
+ [
+ 470,
+ 1431
+ ],
+ [
+ 85,
+ 1460
+ ]
+ ]
+ ],
+ "text_det_params": {
+ "limit_side_len": 64,
+ "limit_type": "min",
+ "thresh": 0.3,
+ "max_side_limit": 4000,
+ "box_thresh": 0.6,
+ "unclip_ratio": 1.5
+ },
+ "text_type": "general",
+ "textline_orientation_angles": [
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1
+ ],
+ "text_rec_score_thresh": 0.0,
+ "return_word_box": false,
+ "rec_texts": [
+ "Venus",
+ "4",
+ "Beu",
+ "5.",
+ "6",
+ "Graphite",
+ "cheetah",
+ "7.",
+ "Bluewhale",
+ "8.",
+ "Sun",
+ "9.",
+ "Babbage",
+ "-",
+ "10.",
+ "1. sahara",
+ "12.Mount"
+ ],
+ "rec_scores": [
+ 0.9222331047058105,
+ 0.9949488639831543,
+ 0.8761672377586365,
+ 0.769047737121582,
+ 0.9965930581092834,
+ 0.9706618785858154,
+ 0.9598208665847778,
+ 0.9705947637557983,
+ 0.9192090630531311,
+ 0.7382487058639526,
+ 0.9669068455696106,
+ 0.981717586517334,
+ 0.9927512407302856,
+ 0.11772612482309341,
+ 0.9063168168067932,
+ 0.926537811756134,
+ 0.9096950888633728
+ ],
+ "rec_polys": [
+ [
+ [
+ 209,
+ 389
+ ],
+ [
+ 426,
+ 386
+ ],
+ [
+ 427,
+ 446
+ ],
+ [
+ 210,
+ 449
+ ]
+ ],
+ [
+ [
+ 68,
+ 421
+ ],
+ [
+ 135,
+ 421
+ ],
+ [
+ 135,
+ 485
+ ],
+ [
+ 68,
+ 485
+ ]
+ ],
+ [
+ [
+ 220,
+ 498
+ ],
+ [
+ 381,
+ 492
+ ],
+ [
+ 383,
+ 567
+ ],
+ [
+ 222,
+ 572
+ ]
+ ],
+ [
+ [
+ 80,
+ 524
+ ],
+ [
+ 151,
+ 524
+ ],
+ [
+ 151,
+ 584
+ ],
+ [
+ 80,
+ 584
+ ]
+ ],
+ [
+ [
+ 88,
+ 630
+ ],
+ [
+ 146,
+ 636
+ ],
+ [
+ 138,
+ 712
+ ],
+ [
+ 80,
+ 706
+ ]
+ ],
+ [
+ [
+ 222,
+ 630
+ ],
+ [
+ 559,
+ 626
+ ],
+ [
+ 560,
+ 702
+ ],
+ [
+ 223,
+ 706
+ ]
+ ],
+ [
+ [
+ 175,
+ 744
+ ],
+ [
+ 520,
+ 738
+ ],
+ [
+ 522,
+ 814
+ ],
+ [
+ 177,
+ 820
+ ]
+ ],
+ [
+ [
+ 83,
+ 772
+ ],
+ [
+ 131,
+ 772
+ ],
+ [
+ 131,
+ 819
+ ],
+ [
+ 83,
+ 819
+ ]
+ ],
+ [
+ [
+ 209,
+ 859
+ ],
+ [
+ 628,
+ 838
+ ],
+ [
+ 632,
+ 914
+ ],
+ [
+ 213,
+ 935
+ ]
+ ],
+ [
+ [
+ 85,
+ 878
+ ],
+ [
+ 140,
+ 878
+ ],
+ [
+ 140,
+ 936
+ ],
+ [
+ 85,
+ 936
+ ]
+ ],
+ [
+ [
+ 202,
+ 997
+ ],
+ [
+ 343,
+ 989
+ ],
+ [
+ 348,
+ 1065
+ ],
+ [
+ 207,
+ 1073
+ ]
+ ],
+ [
+ [
+ 81,
+ 1009
+ ],
+ [
+ 149,
+ 1009
+ ],
+ [
+ 149,
+ 1081
+ ],
+ [
+ 81,
+ 1081
+ ]
+ ],
+ [
+ [
+ 209,
+ 1121
+ ],
+ [
+ 508,
+ 1110
+ ],
+ [
+ 512,
+ 1196
+ ],
+ [
+ 212,
+ 1207
+ ]
+ ],
+ [
+ [
+ 0,
+ 1132
+ ],
+ [
+ 2160,
+ 1052
+ ],
+ [
+ 2160,
+ 2938
+ ],
+ [
+ 0,
+ 2938
+ ]
+ ],
+ [
+ [
+ 77,
+ 1136
+ ],
+ [
+ 157,
+ 1136
+ ],
+ [
+ 157,
+ 1205
+ ],
+ [
+ 77,
+ 1205
+ ]
+ ],
+ [
+ [
+ 86,
+ 1277
+ ],
+ [
+ 501,
+ 1252
+ ],
+ [
+ 504,
+ 1314
+ ],
+ [
+ 90,
+ 1339
+ ]
+ ],
+ [
+ [
+ 80,
+ 1396
+ ],
+ [
+ 465,
+ 1366
+ ],
+ [
+ 470,
+ 1431
+ ],
+ [
+ 85,
+ 1460
+ ]
+ ]
+ ],
+ "rec_boxes": [
+ [
+ 209,
+ 386,
+ 427,
+ 449
+ ],
+ [
+ 68,
+ 421,
+ 135,
+ 485
+ ],
+ [
+ 220,
+ 492,
+ 383,
+ 572
+ ],
+ [
+ 80,
+ 524,
+ 151,
+ 584
+ ],
+ [
+ 80,
+ 630,
+ 146,
+ 712
+ ],
+ [
+ 222,
+ 626,
+ 560,
+ 706
+ ],
+ [
+ 175,
+ 738,
+ 522,
+ 820
+ ],
+ [
+ 83,
+ 772,
+ 131,
+ 819
+ ],
+ [
+ 209,
+ 838,
+ 632,
+ 935
+ ],
+ [
+ 85,
+ 878,
+ 140,
+ 936
+ ],
+ [
+ 202,
+ 989,
+ 348,
+ 1073
+ ],
+ [
+ 81,
+ 1009,
+ 149,
+ 1081
+ ],
+ [
+ 209,
+ 1110,
+ 512,
+ 1207
+ ],
+ [
+ 0,
+ 1052,
+ 2160,
+ 2938
+ ],
+ [
+ 77,
+ 1136,
+ 157,
+ 1205
+ ],
+ [
+ 86,
+ 1252,
+ 504,
+ 1339
+ ],
+ [
+ 80,
+ 1366,
+ 470,
+ 1460
+ ]
+ ]
+}
\ No newline at end of file
diff --git a/requirements.txt b/requirements.txt
new file mode 100644
index 0000000000000000000000000000000000000000..7a4abd3119741fdb3c41345dd7933e84c4c69aa0
--- /dev/null
+++ b/requirements.txt
@@ -0,0 +1,24 @@
+easyocr
+opencv-python
+flask
+flask-cors
+pytesseract
+pillow
+pdf2image
+fuzzywuzzy
+python-levenshtein
+google-generativeai
+python-dotenv
+supabase
+paddlepaddle
+paddleocr
+openpyxl
+deepmerge>=1.1.0
+dotmap>=1.3.30
+jsonschema>=4.17.3
+matplotlib>=3.7.1
+numpy>=1.25.0
+pandas>=2.0.2
+rich>=13.4.2
+screeninfo>=0.8.1
+requests
diff --git a/server.py b/server.py
new file mode 100644
index 0000000000000000000000000000000000000000..b63634859c02cf1cefd7ce57eb98acc513aa7a34
--- /dev/null
+++ b/server.py
@@ -0,0 +1,182 @@
+class QuestionPaper:
+ def __init__(self, path=None):
+ self.questions = []
+ self.answers = []
+ self.path = path
+
+ def clean_answers(self):
+ # Remove unwanted patterns from answers
+ unwanted_patterns = [
+ "Time: 15 MinutesMarks: 20",
+ "Time: 15 Minutes Marks: 20",
+ "GENERAL KNOWLEDGE QUESTION PAPER WITH ANSWERS",
+ "GENERAL KNOWLEDGE QUESTION PAPER",
+ "" # Empty strings
+ ]
+
+ # Filter out unwanted answers
+ cleaned_answers = []
+ for answer in self.answers:
+ if answer.strip() and answer.strip() not in unwanted_patterns:
+ # Also check if it doesn't match any unwanted pattern with regex
+ is_unwanted = False
+ for pattern in unwanted_patterns:
+ if pattern and re.search(re.escape(pattern), answer, re.IGNORECASE):
+ is_unwanted = True
+ break
+ if not is_unwanted:
+ cleaned_answers.append(answer.strip())
+
+ self.answers = cleaned_answers
+
+ def add_question(self, question_text):
+ self.questions.append(question_text)
+
+ def add_answer(self, answer_text):
+ self.answers.append(answer_text)
+
+ def to_dict(self):
+ return {
+ 'questions': self.questions,
+ 'answers': self.answers
+ }
+
+def parse_question_paper_text(text):
+ """
+ Improved parsing function that correctly identifies questions and answers
+ """
+ lines = [line.strip() for line in text.split('\n') if line.strip()]
+
+ questions = []
+ answers = []
+
+ # Patterns to ignore (headers, footers, etc.)
+ ignore_patterns = [
+ r'GENERAL KNOWLEDGE QUESTION PAPER.*',
+ r'Time:\s*\d+\s*Minutes.*Marks:\s*\d+',
+ r'Time:\s*\d+\s*MinutesMarks:\s*\d+',
+ r'^\s*$' # Empty lines
+ ]
+
+ # Filter out unwanted lines
+ filtered_lines = []
+ for line in lines:
+ should_ignore = False
+ for pattern in ignore_patterns:
+ if re.match(pattern, line, re.IGNORECASE):
+ should_ignore = True
+ break
+ if not should_ignore:
+ filtered_lines.append(line)
+
+ # Pattern to identify questions (starts with number followed by dot/parenthesis)
+ question_pattern = r'^\d+\s*[.)]\s*(.+)'
+
+ i = 0
+ while i < len(filtered_lines):
+ current_line = filtered_lines[i].strip()
+
+ # Check if current line is a question
+ question_match = re.match(question_pattern, current_line)
+ if question_match:
+ # This is a question
+ question_text = question_match.group(1).strip()
+ questions.append(f"{current_line}") # Keep the full question with number
+
+ # Look for the answer in the next line
+ if i + 1 < len(filtered_lines):
+ next_line = filtered_lines[i + 1].strip()
+ # If next line is not a question (doesn't start with number), it's likely an answer
+ if not re.match(question_pattern, next_line):
+ answers.append(next_line)
+ i += 2 # Skip both question and answer
+ else:
+ # Next line is also a question, so this question might not have an answer
+ # Or the answer might be embedded in the same line
+ # Try to extract answer from the question line itself if it contains common answer patterns
+ answers.append("") # Placeholder for missing answer
+ i += 1
+ else:
+ # Last line and it's a question without answer
+ answers.append("")
+ i += 1
+ else:
+ # This line doesn't match question pattern, skip it or try to pair it with previous question
+ if len(questions) > len(answers):
+ # We have more questions than answers, this might be an answer
+ answers.append(current_line)
+ i += 1
+
+ # Ensure we have equal number of questions and answers
+ while len(answers) < len(questions):
+ answers.append("")
+ while len(questions) < len(answers):
+ questions.append(f"Question {len(questions) + 1}")
+
+ return questions, answers
+
+@app.route('/process_question_paper', methods=['POST'])
+def process_question_paper():
+ global last_processed_question_paper_object
+
+ if 'file' not in request.files:
+ return jsonify({'error': 'No file provided'}), 400
+
+ file = request.files['file']
+ if file.filename == '':
+ return jsonify({'error': 'No file selected'}), 400
+
+ question_paper = QuestionPaper()
+
+ try:
+ # Create Images directory if it doesn't exist
+ images_dir = os.path.join(app.root_path, 'Images')
+ os.makedirs(images_dir, exist_ok=True)
+
+ if file.filename.lower().endswith('.pdf'):
+ question_paper_filename = "question_paper.pdf"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ # Initialize the global object with the path
+ question_paper.path = question_paper_path
+
+ # For PDF processing
+ images_from_pdf = convert_from_path(question_paper_path, poppler_path=r'C:\Program Files\poppler\Library\bin')
+
+ all_text = ""
+ for page_image in images_from_pdf:
+ text = pytesseract.image_to_string(page_image)
+ all_text += text + "\n"
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(all_text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ else:
+ # Process as image
+ question_paper_filename = "question_paper.png"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ question_paper.path = question_paper_path
+
+ image = Image.open(question_paper_path)
+ text = pytesseract.image_to_string(image)
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ # Clean the answers (remove any remaining unwanted patterns)
+ question_paper.clean_answers()
+
+ # Store the processed question paper globally
+ last_processed_question_paper_object = question_paper
+
+ return jsonify(question_paper.to_dict())
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
\ No newline at end of file
diff --git a/supabase_handler.py b/supabase_handler.py
new file mode 100644
index 0000000000000000000000000000000000000000..252c666b5bc0798342655da4beefe6f63c30e90a
--- /dev/null
+++ b/supabase_handler.py
@@ -0,0 +1,80 @@
+import os
+import uuid
+from datetime import datetime
+from supabase import create_client, Client
+from dotenv import load_dotenv
+
+load_dotenv()
+
+class SupabaseHandler:
+ def __init__(self):
+ url: str = os.getenv("SUPABASE_URL")
+ key: str = os.getenv("SUPABASE_ANON_KEY")
+ if not url or not key:
+ raise ValueError("SUPABASE_URL and SUPABASE_ANON_KEY must be set in environment variables")
+ self.supabase: Client = create_client(url, key)
+
+ def store_evaluation_result(self, teacher_email, evaluation_data):
+ """
+ Store evaluation result in Supabase with a unique key
+ Returns the unique key for retrieval
+ """
+ try:
+ # Generate unique key
+ unique_key = str(uuid.uuid4())
+
+ # Prepare data for storage
+ storage_data = {
+ "unique_key": unique_key,
+ "teacher_email": teacher_email,
+ "evaluation_data": evaluation_data,
+ "created_at": datetime.utcnow().isoformat(),
+ "total_students": evaluation_data.get("total_students", 0)
+ }
+
+ # Insert into Supabase
+ result = self.supabase.table("evaluation_results").insert(storage_data).execute()
+
+ if result.data:
+ print(f"Successfully stored evaluation result with key: {unique_key}")
+ return unique_key
+ else:
+ print("Failed to store evaluation result")
+ return None
+
+ except Exception as e:
+ print(f"Error storing evaluation result: {str(e)}")
+ return None
+
+ def get_evaluation_result(self, unique_key):
+ """
+ Retrieve evaluation result by unique key
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("*").eq("unique_key", unique_key).execute()
+
+ if result.data and len(result.data) > 0:
+ return result.data[0]
+ else:
+ return None
+
+ except Exception as e:
+ print(f"Error retrieving evaluation result: {str(e)}")
+ return None
+
+ def get_teacher_evaluations(self, teacher_email):
+ """
+ Get all evaluation results for a specific teacher
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("unique_key", "created_at", "total_students").eq("teacher_email", teacher_email).order("created_at", desc=True).execute()
+
+ if result.data:
+ return result.data
+ else:
+ return []
+
+ except Exception as e:
+ print(f"Error retrieving teacher evaluations: {str(e)}")
+ return []
+
diff --git a/test_endpoints.py b/test_endpoints.py
new file mode 100644
index 0000000000000000000000000000000000000000..03ae6de089292ac54acd9c12873773ad35fa8b7f
--- /dev/null
+++ b/test_endpoints.py
@@ -0,0 +1,236 @@
+"""
+Test script for OCR/OMR backend endpoints.
+Usage: python test_endpoints.py [base_url]
+Default base_url: http://localhost:5000
+"""
+
+import sys
+import os
+import requests
+
+BASE_URL = sys.argv[1] if len(sys.argv) > 1 else "http://localhost:5000"
+IMAGES_DIR = os.path.join(os.path.dirname(__file__), "Images")
+
+results = []
+# Track prerequisite passes for dependent tests
+prereqs = {
+ "question_paper": False,
+ "omr_answer_key": False,
+ "process_omr": False,
+}
+
+
+def report(name, passed, detail=""):
+ status = "PASS" if passed else "FAIL"
+ results.append((name, status))
+ msg = f"[{status}] {name}"
+ if detail:
+ msg += f" -- {detail}"
+ print(msg)
+ return passed
+
+
+def skip(name, reason):
+ results.append((name, "SKIP"))
+ print(f"[SKIP] {name} -- {reason}")
+
+
+# ---------------------------------------------------------------------------
+# 1. Health check
+# ---------------------------------------------------------------------------
+def test_health():
+ try:
+ r = requests.get(f"{BASE_URL}/health", timeout=10)
+ return report("/health", r.status_code == 200, f"status={r.status_code}")
+ except Exception as e:
+ return report("/health", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 2. Root endpoint
+# ---------------------------------------------------------------------------
+def test_root():
+ try:
+ r = requests.get(f"{BASE_URL}/", timeout=10)
+ return report("/ (root)", r.status_code == 200, f"status={r.status_code}")
+ except Exception as e:
+ return report("/ (root)", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 3. EasyOCR
+# ---------------------------------------------------------------------------
+def test_easyocr():
+ img = os.path.join(IMAGES_DIR, "test.jpg")
+ if not os.path.exists(img):
+ return report("/easyocr", False, "test.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/easyocr", files={"images": f}, timeout=60)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", keys={list(data.keys()) if isinstance(data, dict) else 'list'}"
+ return report("/easyocr", ok, detail)
+ except Exception as e:
+ return report("/easyocr", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 4. Tesseract
+# ---------------------------------------------------------------------------
+def test_tesseract():
+ img = os.path.join(IMAGES_DIR, "OCRSheet.jpg")
+ if not os.path.exists(img):
+ return report("/tesseract", False, "OCRSheet.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/tesseract", files={"images": f}, timeout=60)
+ ok = r.status_code == 200
+ return report("/tesseract", ok, f"status={r.status_code}")
+ except Exception as e:
+ return report("/tesseract", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 5. Process question paper (PDF)
+# ---------------------------------------------------------------------------
+def test_question_paper_pdf():
+ img = os.path.join(IMAGES_DIR, "question_paper.pdf")
+ if not os.path.exists(img):
+ return report("/process_question_paper (pdf)", False, "question_paper.pdf not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_question_paper", files={"file": f}, timeout=120)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", questions={len(data.get('questions', []))}"
+ prereqs["question_paper"] = True
+ return report("/process_question_paper (pdf)", ok, detail)
+ except Exception as e:
+ return report("/process_question_paper (pdf)", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 6. Process question paper (image)
+# ---------------------------------------------------------------------------
+def test_question_paper_png():
+ img = os.path.join(IMAGES_DIR, "question_paper.png")
+ if not os.path.exists(img):
+ return report("/process_question_paper (png)", False, "question_paper.png not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_question_paper", files={"file": f}, timeout=120)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", questions={len(data.get('questions', []))}"
+ prereqs["question_paper"] = True
+ return report("/process_question_paper (png)", ok, detail)
+ except Exception as e:
+ return report("/process_question_paper (png)", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 7. Process OMR answer key
+# ---------------------------------------------------------------------------
+def test_omr_answer_key():
+ img = os.path.join(IMAGES_DIR, "omr_answer_key.pdf")
+ if not os.path.exists(img):
+ return report("/process_omr_answer_key", False, "omr_answer_key.pdf not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_omr_answer_key", files={"file": f}, timeout=120)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", success={data.get('success')}"
+ prereqs["omr_answer_key"] = True
+ return report("/process_omr_answer_key", ok, detail)
+ except Exception as e:
+ return report("/process_omr_answer_key", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 8. Process OMR sheet
+# ---------------------------------------------------------------------------
+def test_process_omr():
+ img = os.path.join(IMAGES_DIR, "OMRSheet.jpg")
+ if not os.path.exists(img):
+ return report("/process_omr", False, "OMRSheet.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_omr", files={"images": f}, timeout=180)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ prereqs["process_omr"] = True
+ return report("/process_omr", ok, detail)
+ except Exception as e:
+ return report("/process_omr", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 9. Evaluate OMR (depends on 7 + 8)
+# ---------------------------------------------------------------------------
+def test_evaluate_omr():
+ if not prereqs["omr_answer_key"] or not prereqs["process_omr"]:
+ skip("/evaluate_omr", "requires /process_omr_answer_key and /process_omr to pass first")
+ return
+ try:
+ r = requests.post(f"{BASE_URL}/evaluate_omr", timeout=120)
+ ok = r.status_code == 200
+ return report("/evaluate_omr", ok, f"status={r.status_code}")
+ except Exception as e:
+ return report("/evaluate_omr", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 10. Evaluate answers (depends on 5/6)
+# ---------------------------------------------------------------------------
+def test_evaluate_answers():
+ if not prereqs["question_paper"]:
+ skip("/evaluate_answers", "requires /process_question_paper to pass first")
+ return
+ img = os.path.join(IMAGES_DIR, "test.jpg")
+ if not os.path.exists(img):
+ return report("/evaluate_answers", False, "test.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/evaluate_answers", files={"student_answers": f}, timeout=120)
+ ok = r.status_code == 200
+ return report("/evaluate_answers", ok, f"status={r.status_code}")
+ except Exception as e:
+ return report("/evaluate_answers", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# Run all tests in order
+# ---------------------------------------------------------------------------
+if __name__ == "__main__":
+ print(f"\nTesting backend at: {BASE_URL}\n" + "=" * 60)
+
+ test_health()
+ test_root()
+ test_easyocr()
+ test_tesseract()
+ test_question_paper_pdf()
+ test_question_paper_png()
+ test_omr_answer_key()
+ test_process_omr()
+ test_evaluate_omr()
+ test_evaluate_answers()
+
+ # Summary
+ print("\n" + "=" * 60)
+ passed = sum(1 for _, s in results if s == "PASS")
+ failed = sum(1 for _, s in results if s == "FAIL")
+ skipped = sum(1 for _, s in results if s == "SKIP")
+ total = len(results)
+ print(f"Results: {passed}/{total} passed, {failed} failed, {skipped} skipped")
+ sys.exit(0 if failed == 0 else 1)
[](https://www.paypal.me/Udayraj123/500)
+
+_Find OMRChecker on_ [**_Product Hunt_**](https://www.producthunt.com/posts/omr-checker/) **|** [**_Reddit_**](https://www.reddit.com/r/computervision/comments/ccbj6f/omrchecker_grade_exams_using_python_and_opencv/) **|** [**Discord**](https://discord.gg/qFv2Vqf) **|** [**Linkedin**](https://www.linkedin.com/pulse/open-source-talks-udayraj-udayraj-deshmukh/) **|** [**goodfirstissue.dev**](https://goodfirstissue.dev/language/python) **|** [**codepeak.tech**](https://www.codepeak.tech/) **|** [**fossoverflow.dev**](https://fossoverflow.dev/projects) **|** [**Interview on Console by CodeSee**](https://console.substack.com/p/console-140) **|** [**Open Source Hub**](https://opensourcehub.io/udayraj123/omrchecker)
+
+
+
diff --git a/OMRChecker/docs/assets/colored_output.jpg b/OMRChecker/docs/assets/colored_output.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3cafa473b9a1134f845858be4cf0e98f3bd3bb7c
Binary files /dev/null and b/OMRChecker/docs/assets/colored_output.jpg differ
diff --git a/OMRChecker/inputs/OMRImage.jpg b/OMRChecker/inputs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c9e3dcccfc8af7c60becb3a4cc25f650861db98e
--- /dev/null
+++ b/OMRChecker/inputs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4439b6a479cb9fea92de9656b900a528052616ae8e8962a6c33b8e81ed7f327a
+size 267398
diff --git a/OMRChecker/inputs/template.json b/OMRChecker/inputs/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..823fe2e58a4182ef04d1fff99ecf146b5ec53aae
--- /dev/null
+++ b/OMRChecker/inputs/template.json
@@ -0,0 +1,37 @@
+{
+ "pageDimensions": [
+ 1122,
+ 1600
+ ],
+ "bubbleDimensions": [
+ 48,
+ 50
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 100,
+ 175
+ ],
+ "bubblesGap": 55,
+ "labelsGap": 67,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q02block": {
+ "origin": [
+ 100,
+ 845
+ ],
+ "bubblesGap": 58,
+ "labelsGap": 70,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ }
+
+}
\ No newline at end of file
diff --git a/OMRChecker/main.py b/OMRChecker/main.py
new file mode 100644
index 0000000000000000000000000000000000000000..19e3ebd43d4be2a096cf709b04817247f9952506
--- /dev/null
+++ b/OMRChecker/main.py
@@ -0,0 +1,99 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+
+import argparse
+import sys
+from pathlib import Path
+
+from src.entry import entry_point
+from src.logger import logger
+
+
+def parse_args():
+ # construct the argument parse and parse the arguments
+ argparser = argparse.ArgumentParser()
+
+ argparser.add_argument(
+ "-i",
+ "--inputDir",
+ default=["inputs"],
+ # https://docs.python.org/3/library/argparse.html#nargs
+ nargs="*",
+ required=False,
+ type=str,
+ dest="input_paths",
+ help="Specify an input directory.",
+ )
+
+ argparser.add_argument(
+ "-d",
+ "--debug",
+ required=False,
+ dest="debug",
+ action="store_false",
+ help="Enables debugging mode for showing detailed errors",
+ )
+
+ argparser.add_argument(
+ "-o",
+ "--outputDir",
+ default="outputs",
+ required=False,
+ dest="output_dir",
+ help="Specify an output directory.",
+ )
+
+ argparser.add_argument(
+ "-a",
+ "--autoAlign",
+ required=False,
+ dest="autoAlign",
+ action="store_true",
+ help="(experimental) Enables automatic template alignment - \
+ use if the scans show slight misalignments.",
+ )
+
+ argparser.add_argument(
+ "-l",
+ "--setLayout",
+ required=False,
+ dest="setLayout",
+ action="store_true",
+ help="Set up OMR template layout - modify your json file and \
+ run again until the template is set.",
+ )
+
+ (
+ args,
+ unknown,
+ ) = argparser.parse_known_args()
+
+ args = vars(args)
+
+ if len(unknown) > 0:
+ logger.warning(f"\nError: Unknown arguments: {unknown}", unknown)
+ argparser.print_help()
+ exit(11)
+ return args
+
+
+def entry_point_for_args(args):
+ if args["debug"] is True:
+ # Disable tracebacks
+ sys.tracebacklimit = 0
+ for root in args["input_paths"]:
+ entry_point(
+ Path(root),
+ args,
+ )
+
+
+if __name__ == "__main__":
+ args = parse_args()
+ entry_point_for_args(args)
diff --git a/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..d8d1a2325170014745edf3503b40e7c7bda160dc
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:f16c45dda1d02bf06e33a6d219c8282c363bf3943b94960eede1efe2bc6204d9
+size 408744
diff --git a/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3a968368fa191241dda21ea0c41f5d2d840f2e52
--- /dev/null
+++ b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:7e9678555042535c83ab920d0b616d4f99f9f08017629d46a15eb64d7c3abd3c
+size 407394
diff --git a/OMRChecker/outputs/Images/Manual/ErrorFiles.csv b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Results/Results_10PM.csv b/OMRChecker/outputs/Images/Results/Results_10PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..f3b83ec4b12f9e002ca0824ee43fc41f1f9ae30a
--- /dev/null
+++ b/OMRChecker/outputs/Images/Results/Results_10PM.csv
@@ -0,0 +1,2 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\Images\OMRSheet.jpg","outputs\Images\CheckedOMRs\OMRSheet.jpg","0","C","D","A","C","B","C","C","D","D","B","B","B","D","C","C","ABCD","B","ABCD","C","ABCD"
diff --git a/OMRChecker/outputs/Manual/ErrorFiles.csv b/OMRChecker/outputs/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..a9bb85c601677eefe49c6a75b39d5bd404506322
--- /dev/null
+++ b/OMRChecker/outputs/Manual/ErrorFiles.csv
@@ -0,0 +1,3 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
diff --git a/OMRChecker/outputs/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/pyproject.toml b/OMRChecker/pyproject.toml
new file mode 100644
index 0000000000000000000000000000000000000000..efa55e8fc8bf1623d8569abd406fef4d81270291
--- /dev/null
+++ b/OMRChecker/pyproject.toml
@@ -0,0 +1,18 @@
+[tool.black]
+exclude = '''
+(
+ /(
+ \.eggs # exclude a few common directories in the
+ | \.git # root of the project
+ | \.venv
+ | _build
+ | build
+ | dist
+ )/
+ | foo.py # also separately exclude a file named foo.py in
+ # the root of the project
+)
+'''
+include = '\.pyi?$'
+line-length = 88
+target-version = ['py37']
diff --git a/OMRChecker/pytest.ini b/OMRChecker/pytest.ini
new file mode 100644
index 0000000000000000000000000000000000000000..417c4a2dd810294aa9afaa9b4f73464fc9808c96
--- /dev/null
+++ b/OMRChecker/pytest.ini
@@ -0,0 +1,6 @@
+# pytest.ini
+[pytest]
+minversion = 7.0
+addopts = -qq --capture=no
+testpaths =
+ src/tests
diff --git a/OMRChecker/requirements.dev.txt b/OMRChecker/requirements.dev.txt
new file mode 100644
index 0000000000000000000000000000000000000000..fb31b5f80558481a3f2a5edbb741b363a1bd1089
--- /dev/null
+++ b/OMRChecker/requirements.dev.txt
@@ -0,0 +1,7 @@
+-r requirements.txt
+flake8>=6.0.0
+freezegun>=1.2.2
+pre-commit>=3.3.3
+pytest-mock>=3.11.1
+pytest>=7.4.0
+syrupy>=4.0.4
diff --git a/OMRChecker/requirements.txt b/OMRChecker/requirements.txt
new file mode 100644
index 0000000000000000000000000000000000000000..d0c3af875835b492a48ef689c3dc9335eb7cdca7
--- /dev/null
+++ b/OMRChecker/requirements.txt
@@ -0,0 +1,8 @@
+deepmerge>=1.1.0
+dotmap>=1.3.30
+jsonschema>=4.17.3
+matplotlib>=3.7.1
+numpy>=1.25.0
+pandas>=2.0.2
+rich>=13.4.2
+screeninfo>=0.8.1
diff --git a/OMRChecker/samples/answer-key/using-csv/adrian_omr.png b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png differ
diff --git a/OMRChecker/samples/answer-key/using-csv/answer_key.csv b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..acf09be611a66dfaeb7127e7249eadfbba6b0881
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
@@ -0,0 +1,5 @@
+q1,C
+q2,E
+q3,A
+q4,B
+q5,B
\ No newline at end of file
diff --git a/OMRChecker/samples/answer-key/using-csv/evaluation.json b/OMRChecker/samples/answer-key/using-csv/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/using-csv/template.json b/OMRChecker/samples/answer-key/using-csv/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/evaluation.json b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..09508b18e3490efcd68d6cb0b57a03d6e203f2ca
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
@@ -0,0 +1,35 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..5"
+ ],
+ "answers_in_order": [
+ "C",
+ "E",
+ [
+ "A",
+ "C"
+ ],
+ [
+ [
+ "B",
+ 2
+ ],
+ [
+ "C",
+ "3/2"
+ ]
+ ],
+ "C"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/answer-key/weighted-answers/template.json b/OMRChecker/samples/answer-key/weighted-answers/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..661d5f4fa73d7eea57e73837cf3862a25b9efcc3
Binary files /dev/null and b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg differ
diff --git a/OMRChecker/samples/community/Antibodyy/template.json b/OMRChecker/samples/community/Antibodyy/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..55e016e8eb402156d62f6553067b68abb37377ee
--- /dev/null
+++ b/OMRChecker/samples/community/Antibodyy/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 299,
+ 398
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 79
+ ],
+ "bubblesGap": 43,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "q1..6"
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-1.png b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
new file mode 100644
index 0000000000000000000000000000000000000000..5ed00d194a806d58c3db1dab7831796df7d4aaef
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:ed3024e5b709fbbdb1bcbef35dcfa50203ddb811f74b26e657d3f1b86c7e8c94
+size 381247
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-2.png b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d250db2c94980defb7ab78f9114ca56c1f223024
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:571109a06a09b792694a0ecac329283da199ac4495b1f10ffb7e5925c76a75c9
+size 130973
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-3.png b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
new file mode 100644
index 0000000000000000000000000000000000000000..c414be1c6fa93808d3419715e2e610884e233926
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:3976c705b45d889b06acc4bf82b27f129ad55c4875ee2aa6eb01aecf5a0250f6
+size 144032
diff --git a/OMRChecker/samples/community/Sandeep-1507/template.json b/OMRChecker/samples/community/Sandeep-1507/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bdb929ccd1a61c3ae54df4cdd6a8cf4281369733
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/template.json
@@ -0,0 +1,234 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 15,
+ 15
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Booklet_No": [
+ "b1..7"
+ ]
+ },
+ "fieldBlocks": {
+ "Booklet_No": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 112,
+ 530
+ ],
+ "fieldLabels": [
+ "b1..7"
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 28,
+ "labelsGap": 26.5
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 100
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 370
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q21..35"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 638
+ ]
+ },
+ "MCQBlock2a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 100
+ ]
+ },
+ "MCQBlock2a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 370
+ ]
+ },
+ "MCQBlock2a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q71..85"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 638
+ ]
+ },
+ "MCQBlock3a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q101..110"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 100
+ ]
+ },
+ "MCQBlock3a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q111..120"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 370
+ ]
+ },
+ "MCQBlock3a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q121..135"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 638
+ ]
+ },
+ "MCQBlock4a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q151..160"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 100
+ ]
+ },
+ "MCQBlock4a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q161..170"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 370
+ ]
+ },
+ "MCQBlock4a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q171..185"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 638
+ ]
+ },
+ "MCQBlock1a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q36..50"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 480,
+ 1061
+ ]
+ },
+ "MCQBlock2a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q86..100"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 648,
+ 1061
+ ]
+ },
+ "MCQBlock3a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q136..150"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 1061
+ ]
+ },
+ "MCQBlock4a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q186..200"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.6,
+ "origin": [
+ 986,
+ 1061
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Shamanth/omr_sheet_01.png b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
new file mode 100644
index 0000000000000000000000000000000000000000..7c317db90f80fdd8d3dcfbbadcb1f9b764e619e3
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:9dd72b86ce9c6544f4a628187b4021df5746e9579222e7598c4d88846f2fbeb3
+size 107647
diff --git a/OMRChecker/samples/community/Shamanth/template.json b/OMRChecker/samples/community/Shamanth/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..7d8c34d5e4b7af5ca94415b5096c82300156bcbc
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/template.json
@@ -0,0 +1,25 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 78,
+ 41
+ ],
+ "fieldLabels": [
+ "q21..28"
+ ],
+ "bubblesGap": 56,
+ "labelsGap": 46
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/answer_key.jpg b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..9d39b263e8fe5c23e391ad39bd0905efbf976038
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:13bb31818680ca8497ee4c640a15d4acab2048cf11c973d758c8c75fd71c73be
+size 291737
diff --git a/OMRChecker/samples/community/UPSC-mock/config.json b/OMRChecker/samples/community/UPSC-mock/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..388060a33b04db4b550d0efecafd32aa277355d5
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 1800,
+ "display_width": 2400,
+ "processing_height": 2400,
+ "processing_width": 1800
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/evaluation.json b/OMRChecker/samples/community/UPSC-mock/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..3271cbd5e39161d428d765145f34d811701bdc64
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/evaluation.json
@@ -0,0 +1,18 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "answer_key_image_path": "answer_key.jpg",
+ "questions_in_order": [
+ "q1..100"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "2",
+ "incorrect": "-2/3",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..5dd9ae6adbd327c7995bfcd5f53989002e240cf2
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:020705f4618fd55a72dbc35719565307b844dbd28838c790fe858c3ebe5fb36e
+size 160882
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c696b2e2d39118dec97d4f4c7479bfbc2dc6fc66
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1bb28daa10d276768d4fe3fd3f162ff8d3556cb805c4cf3f362e84d00d808095
+size 182385
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..edb7bbb600db8175e5beaa707c58d5a95972ec1e
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:695e6cc5a497bcf114a101533ee0464a223ecb8f914bc28a8d356533bcf6e341
+size 179112
diff --git a/OMRChecker/samples/community/UPSC-mock/template.json b/OMRChecker/samples/community/UPSC-mock/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..2fadab79ca7f7f9adb03a85cde30f1ba2ccca588
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/template.json
@@ -0,0 +1,195 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 30,
+ 25
+ ],
+ "customLabels": {
+ "Subject Code": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "Roll": [
+ "roll1..10"
+ ]
+ },
+ "fieldBlocks": {
+ "bookletNo": {
+ "origin": [
+ 595,
+ 545
+ ],
+ "bubblesGap": 68,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "bookletNo"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "C",
+ "D"
+ ],
+ "direction": "vertical"
+ },
+ "subjectCode": {
+ "origin": [
+ 912,
+ 512
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.5,
+ "fieldLabels": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "roll": {
+ "origin": [
+ 1200,
+ 510
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.8,
+ "fieldLabels": [
+ "roll1..10"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "q01block": {
+ "origin": [
+ 500,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 500,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 495,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1125,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1125,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..ffd67e704a23baa49bec476e04ba6b1796ff9b52
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
@@ -0,0 +1,200 @@
+q1,C
+q2,C
+q3,"D,E"
+q4,"A,AB"
+q5,"[['A', '1'], ['B', '2']]"
+q6,"['A', 'B']"
+q7,C
+q8,D
+q9,B
+q10,B
+q11,A
+q12,A
+q13,C
+q14,B
+q15,D
+q16,B
+q17,C
+q18,A
+q19,B
+q20,D
+q21,D
+q22,C
+q23,A
+q24,C
+q25,D
+q26,C
+q27,C
+q28,B
+q29,A
+q30,D
+q31,C
+q32,B
+q33,B
+q34,C
+q35,A
+q36,D
+q37,C
+q38,B
+q39,C
+q40,A
+q41,A
+q42,C
+q43,D
+q44,D
+q45,B
+q46,C
+q47,C
+q48,A
+q49,C
+q50,B
+q51,B
+q52,C
+q53,D
+q54,C
+q55,B
+q56,B
+q57,A
+q58,A
+q59,D
+q60,C
+q61,C
+q62,B
+q63,A
+q64,C
+q65,D
+q66,C
+q67,B
+q68,A
+q69,B
+q70,B
+q71,C
+q72,B
+q73,C
+q74,A
+q75,A
+q76,C
+q77,D
+q78,D
+q79,B
+q80,A
+q81,B
+q82,C
+q83,D
+q84,C
+q85,A
+q86,C
+q87,D
+q88,B
+q89,C
+q90,B
+q91,B
+q92,A
+q93,C
+q94,D
+q95,C
+q96,B
+q97,B
+q98,A
+q99,A
+q100,A
+q101,A
+q102,B
+q103,C
+q104,C
+q105,A
+q106,D
+q107,B
+q108,A
+q109,C
+q110,B
+q111,B
+q112,C
+q113,C
+q114,B
+q115,D
+q116,B
+q117,A
+q118,C
+q119,D
+q120,C
+q121,C
+q122,A
+q123,B
+q124,C
+q125,D
+q126,C
+q127,C
+q128,D
+q129,D
+q130,A
+q131,A
+q132,C
+q133,B
+q134,C
+q135,D
+q136,B
+q137,C
+q138,A
+q139,B
+q140,D
+q141,D
+q142,C
+q143,D
+q144,A
+q145,A
+q146,C
+q147,A
+q148,D
+q149,D
+q150,B
+q151,A
+q152,B
+q153,B
+q154,D
+q155,D
+q156,B
+q157,A
+q158,B
+q159,A
+q160,C
+q161,D
+q162,C
+q163,A
+q164,B
+q165,D
+q166,D
+q167,C
+q168,C
+q169,C
+q170,D
+q171,A
+q172,A
+q173,C
+q174,C
+q175,B
+q176,D
+q177,A
+q178,B
+q179,B
+q180,C
+q181,D
+q182,C
+q183,B
+q184,B
+q185,C
+q186,D
+q187,D
+q188,A
+q189,A
+q190,B
+q191,C
+q192,B
+q193,D
+q194,C
+q195,B
+q196,B
+q197,A
+q198,B
+q199,B
+q200,A
diff --git a/OMRChecker/samples/community/UmarFarootAPS/config.json b/OMRChecker/samples/community/UmarFarootAPS/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..5a9d75731b7f5ae8c6fd803a7426b29ac39c6bfb
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 960,
+ "display_width": 1280,
+ "processing_height": 1640,
+ "processing_width": 1332
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/evaluation.json b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg differ
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..73d60b9caa9860aa6a9bb75956fd3430b1aee44e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:49e1a14e0c1be809282b6e43c913e17598380a33f607a5c46723764c32e17b59
+size 304722
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..25d75107cf4885f2974c52cf42003255be285a3e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18b85df487c0bc98eb9139949c3fbca2bad57e8f224fe48b495e0951660f65d
+size 237392
diff --git a/OMRChecker/samples/community/UmarFarootAPS/template.json b/OMRChecker/samples/community/UmarFarootAPS/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..f303edabdf3bc64c41c50152ae72d09f5a0de913
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 2550,
+ 3300
+ ],
+ "bubbleDimensions": [
+ 32,
+ 32
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll_no": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ]
+ },
+ "fieldBlocks": {
+ "Roll_no": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 2169,
+ 180
+ ],
+ "fieldLabels": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ],
+ "bubblesGap": 61,
+ "labelsGap": 93
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q1..17"
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q18..34"
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q35..50"
+ ]
+ },
+ "MCQBlock1a4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q51..67"
+ ]
+ },
+ "MCQBlock1a5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q68..84"
+ ]
+ },
+ "MCQBlock1a6": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q85..100"
+ ]
+ },
+ "MCQBlock1a7": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q101..117"
+ ]
+ },
+ "MCQBlock1a8": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q118..134"
+ ]
+ },
+ "MCQBlock1a9": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q135..150"
+ ]
+ },
+ "MCQBlock1a10": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q151..167"
+ ]
+ },
+ "MCQBlock1a11": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q168..184"
+ ]
+ },
+ "MCQBlock1a12": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q185..200"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..b9d70c6be1aa35398e2ac125f3e7bea0ff924fd0
Binary files /dev/null and b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg differ
diff --git a/OMRChecker/samples/community/dxuian/template.json b/OMRChecker/samples/community/dxuian/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e055f557537fa531a0546c2155146763c8d937a6
--- /dev/null
+++ b/OMRChecker/samples/community/dxuian/template.json
@@ -0,0 +1,48 @@
+{
+ "pageDimensions": [707, 484],
+ "bubbleDimensions": [15, 10],
+ "fieldBlocks": {
+ "Column1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [82, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q1", "Q2", "Q3", "Q4", "Q5", "Q6", "Q7", "Q8", "Q9", "Q10", "Q11", "Q12", "Q13", "Q14", "Q15", "Q16", "Q17", "Q18", "Q19", "Q20"]
+ },
+ "Column2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [205, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q21", "Q22", "Q23", "Q24", "Q25", "Q26", "Q27", "Q28", "Q29", "Q30", "Q31", "Q32", "Q33", "Q34", "Q35", "Q36", "Q37", "Q38", "Q39", "Q40"]
+ },
+ "Column3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [327, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q41", "Q42", "Q43", "Q44", "Q45", "Q46", "Q47", "Q48", "Q49", "Q50", "Q51", "Q52", "Q53", "Q54", "Q55", "Q56", "Q57", "Q58", "Q59", "Q60"]
+ },
+ "Column4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [450, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q61", "Q62", "Q63", "Q64", "Q65", "Q66", "Q67", "Q68", "Q69", "Q70", "Q71", "Q72", "Q73", "Q74", "Q75", "Q76", "Q77", "Q78", "Q79", "Q80"]
+ },
+ "Column5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [573, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q81", "Q82", "Q83", "Q84", "Q85", "Q86", "Q87", "Q88", "Q89", "Q90", "Q91", "Q92", "Q93", "Q94", "Q95", "Q96", "Q97", "Q98", "Q99", "Q100"]
+ }
+ },
+
+ "emptyValue": "-"
+}
\ No newline at end of file
diff --git a/OMRChecker/samples/community/ibrahimkilic/template.json b/OMRChecker/samples/community/ibrahimkilic/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..3064ee19192fc7474370c0f518a51bea9a30b639
--- /dev/null
+++ b/OMRChecker/samples/community/ibrahimkilic/template.json
@@ -0,0 +1,30 @@
+{
+ "pageDimensions": [
+ 299,
+ 328
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "emptyValue": "no",
+ "fieldBlocks": {
+ "YesNoBlock1": {
+ "direction": "horizontal",
+ "bubbleValues": [
+ "yes"
+ ],
+ "origin": [
+ 15,
+ 55
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 48,
+ "labelsGap": 48,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..e9436f41ee8db1269e2255ee902b73de2525d502
Binary files /dev/null and b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg differ
diff --git a/OMRChecker/samples/community/samuelIkoli/template.json b/OMRChecker/samples/community/samuelIkoli/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..19c46f7c96bb07cc0a5a1731bbb8febeba9686e9
--- /dev/null
+++ b/OMRChecker/samples/community/samuelIkoli/template.json
@@ -0,0 +1,28 @@
+{
+ "pageDimensions": [630, 404],
+ "bubbleDimensions": [20, 15],
+ "customLabels": {},
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [33, 6],
+ "fieldLabels": ["q1", "q2", "q3", "q4", "q5", "q6", "q7", "q8", "q9", "q10", "q11", "q12", "q13", "q14", "q15", "q16", "q17", "q18", "q19", "q20"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock2": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [248, 6],
+ "fieldLabels": ["q21", "q22", "q23", "q24", "q25", "q26", "q27", "q28", "q29", "q30", "q31", "q32", "q33", "q34", "q35", "q36", "q37", "q38", "q39", "q40"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock3": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [465, 6],
+ "fieldLabels": ["q41", "q42", "q43", "q44", "q45", "q46", "q47", "q48", "q49", "q50", "q51", "q52", "q53", "q54", "q55", "q56", "q57", "q58", "q59", "q60"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg
new file mode 100644
index 0000000000000000000000000000000000000000..fcd61e808e4a2f33095ea0dbef0ec27741450465
Binary files /dev/null and b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg differ
diff --git a/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6626eb476d38cb16156dc15a48d30bcd41ebaac1
--- /dev/null
+++ b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1be7b0fc432e811fd07b07fe325d9ca8d0d97782eea9f95ed62ff0e47076e1cd
+size 162012
diff --git a/OMRChecker/samples/sample1/config.json b/OMRChecker/samples/sample1/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample1/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample1/omr_marker.jpg b/OMRChecker/samples/sample1/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample1/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample1/template.json b/OMRChecker/samples/sample1/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e4795ad56c919618354f18d4de7a14f14642df07
--- /dev/null
+++ b/OMRChecker/samples/sample1/template.json
@@ -0,0 +1,197 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q5": [
+ "q5_1",
+ "q5_2"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubblesGap": 41,
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "fieldLabels": [
+ "Medium"
+ ],
+ "labelsGap": 0,
+ "origin": [
+ 170,
+ 282
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "roll1..9"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 58,
+ "origin": [
+ 225,
+ 282
+ ]
+ },
+ "Int_Block_Q5": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q5_1",
+ "q5_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 903,
+ 282
+ ]
+ },
+ "Int_Block_Q6": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1077,
+ 282
+ ]
+ },
+ "Int_Block_Q7": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1240,
+ 282
+ ]
+ },
+ "Int_Block_Q8": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1410,
+ 282
+ ]
+ },
+ "Int_Block_Q9": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1580,
+ 282
+ ]
+ },
+ "MCQ_Block_Q1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..4"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 860
+ ]
+ },
+ "MCQ_Block_Q10": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q10..13"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 1195
+ ]
+ },
+ "MCQ_Block_Q14": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q14..16"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 860
+ ]
+ },
+ "MCQ_Block_Q17": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q17..20"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 1195
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ },
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/sample2/config.json b/OMRChecker/samples/sample2/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample2/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample2/template.json b/OMRChecker/samples/sample2/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/sample2/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample3/README.md b/OMRChecker/samples/sample3/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..4b1e325dfa284cf676a1f3d9e37d426f5d20ebb6
--- /dev/null
+++ b/OMRChecker/samples/sample3/README.md
@@ -0,0 +1,31 @@
+## Observation
+The OMR layout is slightly different on colored thick papers vs on xeroxed thin papers. The shift becomes noticible in case of OMR with large number of questions.
+
+We overlapped a colored OMR sheet with a xerox copy of the same OMR sheet(both printed on A4 papers) and observed that there is a great amount of layout sheet as we reach the bottom of the OMR.
+
+Link to an explainer with a real life example: [Google Drive](https://drive.google.com/drive/folders/1GpZTmpEhEjSALJEMjHwDafzWKgEoCTOI?usp=sharing)
+
+## Reasons for shifts in Template layout:
+Listing out a few reasons for the above observation:
+### Printer margin setting
+The margin settings for different printers may be different for the same OMR layout. Thus causing the print to become elongated either horizontally, vertically or both ways.
+
+### The Fan-out effect
+The fan-out effect is usually observed in a sheet fed offset press. Depending on how the papers are made, their dimensions have a tendency to change when they are exposed to moisture or water.
+
+The standard office papers(80 gsm) can easily capture moisture and change shape e.g. in case they get stored for multiple days in a place where the weather is highly humid.
+
+Below are some examples of the GSM ranges:
+
+- 74gsm to 90gsm – This is the basic standard office paper, used in your laser printers.
+- 100gsm to 120gsm – This is stationary paper used for standard letterheads, complimentary slips.
+- 130 to 170gsm – Mostly used for leaflets, posters, single-sided flyers, and brochures.
+
+## Solution
+
+It is recommended to scan each types of prints into different folders and use a separate template.json layout for each of the folders. The same is presented in this sample folder.
+
+## References
+
+- [Paper dimensional stability in sheet-fed offset printing](https://www.diva-portal.org/smash/get/diva2:517895/FULLTEXT01.pdf)
+- An analysis of a few ["Interesting" bubble sheets](https://imgur.com/a/10qwL)
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..4877db1feb1c758bc35b41f543dc9bb14d6ca62f
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18eba01e5d0c466b4b750aa20e710e7354489ed9bef91966c2349007fee349d
+size 137681
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/template.json b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ab35b30f1e3ec35d1c4698cc68b76c4a47d6eb37
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 504,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 504,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 500,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1120,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1120,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample3/config.json b/OMRChecker/samples/sample3/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample3/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..2be38bbfb8e8075f0da00d5fb56d4689d2e72e26
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:eab4df0efac2614b2deddcadc0b754aa72c668170e5d9af49ad77c877198556c
+size 187367
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd499ade7e54d5061817985e40effb13fce8f012
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 492,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 492,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 492,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 487,
+ 1920
+ ],
+ "bubblesGap": 61.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 807,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 803,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 803,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 803,
+ 1920
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1115,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1115,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample4/IMG_20201116_143512.jpg b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6c817d4f620b6bc728eeb410bd9c1da8d27a5cfe
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0b1f0e0afd8a2c40deb7f7c992c9fb6046eb1b28a5f9a674e80ce3418c7fc307
+size 421479
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..023f66bc9eb9100a8500d84b82f7725141962082
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8c3f847e6c37c96562ee17b3d69cb919bb52743f8b95d02679ed62e20dbd5096
+size 386960
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..21869f5c27011e067f1fd8cb3a4e6ecb0e4c7e11
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0bdc5eca48c13d19f228ef7252b5e855b5c62926ba1e9e3297fc6a46e074b1be
+size 307198
diff --git a/OMRChecker/samples/sample4/config.json b/OMRChecker/samples/sample4/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..6c82782a3f6486145d6367eabe6910cca5268214
--- /dev/null
+++ b/OMRChecker/samples/sample4/config.json
@@ -0,0 +1,13 @@
+{
+ "dimensions": {
+ "display_width": 1189,
+ "display_height": 1682
+ },
+ "threshold_params": {
+ "MIN_JUMP": 30
+ },
+ "outputs": {
+ "filter_out_multimarked_files": false,
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample4/evaluation.json b/OMRChecker/samples/sample4/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..8ea935e70ded2a9c53c13bf22b5904f1ef3fa923
--- /dev/null
+++ b/OMRChecker/samples/sample4/evaluation.json
@@ -0,0 +1,34 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..11"
+ ],
+ "answers_in_order": [
+ "B",
+ "D",
+ "C",
+ "B",
+ "D",
+ "C",
+ [
+ "B",
+ "C",
+ "BC"
+ ],
+ "A",
+ "C",
+ "D",
+ "C"
+ ],
+ "should_explain_scoring": true,
+ "enable_evaluation_table_to_csv": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample4/template.json b/OMRChecker/samples/sample4/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..d1bef2985939f8ea9c01bfdec28e0d5b6bab11a3
--- /dev/null
+++ b/OMRChecker/samples/sample4/template.json
@@ -0,0 +1,45 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 30,
+ 30
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ },
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 134,
+ 684
+ ],
+ "fieldLabels": [
+ "q1..11"
+ ],
+ "bubblesGap": 79,
+ "labelsGap": 62
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/README.md b/OMRChecker/samples/sample5/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..b70cedd6cbf22de260649c08e855d929d5e8ae87
--- /dev/null
+++ b/OMRChecker/samples/sample5/README.md
@@ -0,0 +1,6 @@
+### OMRChecker Sample
+
+This sample demonstrates multiple things, namely -
+- Running OMRChecker on images scanned using popular document scanning apps
+- Using a common template.json file for sub-folders (e.g. multiple scan batches)
+- Using evaluation.json file with custom marking (without streak-based marking)
diff --git a/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..72b450e07dd7d4c4a18287183128ad7edbf5607a
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:08bb2e9fecf23be6850485cb7776a604aa2efe9800855ea763dee7ade5362323
+size 127651
diff --git a/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..46fb2ac36c0d5c2366fdf9a7ed56eb933578a60f
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0da09483868ad499925fefa284153e9544822894335e16cea859057740c3c063
+size 127863
diff --git a/OMRChecker/samples/sample5/config.json b/OMRChecker/samples/sample5/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample5/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample5/evaluation.json b/OMRChecker/samples/sample5/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..706e49ee9d9e7d445ca98ea87a1a140ce53da54d
--- /dev/null
+++ b/OMRChecker/samples/sample5/evaluation.json
@@ -0,0 +1,93 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..22"
+ ],
+ "answers_in_order": [
+ "C",
+ "C",
+ "B",
+ "C",
+ "C",
+ [
+ "1",
+ "01"
+ ],
+ "19",
+ "10",
+ "10",
+ "18",
+ "D",
+ "A",
+ "D",
+ "D",
+ "D",
+ "C",
+ "C",
+ "C",
+ "C",
+ "D",
+ "B",
+ "A"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ },
+ "BOOMERANG_1": {
+ "questions": [
+ "q1..5"
+ ],
+ "marking": {
+ "correct": 4,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "PROXIMITY_1": {
+ "questions": [
+ "q6..10"
+ ],
+ "marking": {
+ "correct": 3,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_1": {
+ "questions": [
+ "q11..14"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "POWER_SECTION_1": {
+ "questions": [
+ "q15..18"
+ ],
+ "marking": {
+ "correct": 1,
+ "incorrect": 0,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_2": {
+ "questions": [
+ "q19..22"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/omr_marker.jpg b/OMRChecker/samples/sample5/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample5/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample5/template.json b/OMRChecker/samples/sample5/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ac2436c02a8a19024e8050ff7c02dca9aa7ea016
--- /dev/null
+++ b/OMRChecker/samples/sample5/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ],
+ "q10": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "origin": [
+ 200,
+ 215
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "Medium"
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 261,
+ 210
+ ],
+ "bubblesGap": 46.5,
+ "labelsGap": 58,
+ "fieldLabels": [
+ "roll1..9"
+ ]
+ },
+ "Int1": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 935,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ]
+ },
+ "Int2": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1100,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ]
+ },
+ "Int3": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1275,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ]
+ },
+ "Int4": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1449,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "Int5": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1620,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "Mcq1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 198,
+ 826
+ ],
+ "bubblesGap": 93,
+ "labelsGap": 62,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ },
+ "Mcq2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 830
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q11..14"
+ ]
+ },
+ "Mcq3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 1270
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q15..18"
+ ]
+ },
+ "Mcq4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1481,
+ 830
+ ],
+ "bubblesGap": 73,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q19..22"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/config.json b/OMRChecker/samples/sample6/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..2ac30e718a44f7336aafda1b0f57aec1320fc72c
--- /dev/null
+++ b/OMRChecker/samples/sample6/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_width": 2480,
+ "display_height": 3508,
+ "processing_width": 1653,
+ "processing_height": 2339
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..fdaa0f4112e15037a31bc90da204b7c19114820b
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:de8a5ed5b7360a71db83b49a3d0d6e1af1482339d1116286a064d285be42e37e
+size 246590
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..bf56d121c1061302fc4ce87fca61bb8584ea9e4a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:aca51de61ba3ee76e00e22a666d8cd2b72190af674fa051c9cf70346e9ee4d0a
+size 244049
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..22bfa55de7a687729b733a615ad78dad64ac467a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:950dbf86fc152f464be30ebaeb518c61067b0c494ab5fd70f05de10b5dfabb12
+size 244321
diff --git a/OMRChecker/samples/sample6/readme.md b/OMRChecker/samples/sample6/readme.md
new file mode 100644
index 0000000000000000000000000000000000000000..d8491b2a049eb4e721e007e8e8bc4fe5ca536802
--- /dev/null
+++ b/OMRChecker/samples/sample6/readme.md
@@ -0,0 +1,21 @@
+# Demo for feature-based alignment
+
+## Background
+OMR is used to match student roll on final exam scripts. Scripts are scanned using a document scanner and the cover pages are extracted for OMR. Even though a document scanner does not produce any warpped perspective, the alignment is not perfect, causing some rotation and translation in the scans.
+
+The scripts in this sample were specifically selected incorrectly marked scripts to demonstrate how feature-based alignment can correct transformation errors using a reference image. In the actual batch. 156 out of 532 scripts were incorrectly marked. With feature-based alignment, all scripts were correctly marked.
+
+## Usage
+Two template files are given in the sample folder, one with feature-based alignment (template_fb_align), the other without (template_no_fb_align).
+
+## Additional Notes
+
+### Reference Image
+When using a reference image for feature-based alignment, it is better not to have many repeated patterns as it is causes ambiguity when trying to match similar feature points. The bubbles in an OMR form are identical and should not be used for feature-extraction.
+
+Thus, the reference image should be cleared of any bubbles. Forms with lots of text as in this example would be effective.
+
+Note the reference image in this sample was generated from a vector pdf, and not from a scanned blank, producing in a perfectly aligned reference.
+
+### Level adjustment
+The bubbles on the scripts were not shaded dark enough. Thus, a level adjustment was done to bring the black point to 70% to darken the light shading. White point was brought down to 80% to remove the light-grey background in the columns.
diff --git a/OMRChecker/samples/sample6/reference.png b/OMRChecker/samples/sample6/reference.png
new file mode 100644
index 0000000000000000000000000000000000000000..e2e692f4638e9c7fa2d7d159e044c778b18acdc9
--- /dev/null
+++ b/OMRChecker/samples/sample6/reference.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8379ba055209c065def041fc1fb1cea444b913d835c1cfeffa40091446e5e3e5
+size 178069
diff --git a/OMRChecker/samples/sample6/template.json b/OMRChecker/samples/sample6/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_fb_align.json b/OMRChecker/samples/sample6/template_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd45b9c8fd3ba024b951ff4e7965c98bb56211ee
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_fb_align.json
@@ -0,0 +1,118 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "FeatureBasedAlignment",
+ "options": {
+ "reference": "reference.png",
+ "maxFeatures": 1000,
+ "2d": true
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_no_fb_align.json b/OMRChecker/samples/sample6/template_no_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_no_fb_align.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/src/__init__.py b/OMRChecker/src/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..5c1baacaa53bb1b4219d33eee566c6a613ab88d6
--- /dev/null
+++ b/OMRChecker/src/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+from src.logger import logger
+
+# It takes a few seconds for the imports
+logger.info(f"Loading OMRChecker modules...")
diff --git a/OMRChecker/src/constants.py b/OMRChecker/src/constants.py
new file mode 100644
index 0000000000000000000000000000000000000000..628eb75ac892ddc184cc99429036aa9196889c83
--- /dev/null
+++ b/OMRChecker/src/constants.py
@@ -0,0 +1,54 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+from dotmap import DotMap
+
+# Filenames
+TEMPLATE_FILENAME = "template.json"
+EVALUATION_FILENAME = "evaluation.json"
+CONFIG_FILENAME = "config.json"
+
+FIELD_LABEL_NUMBER_REGEX = r"([^\d]+)(\d*)"
+#
+ERROR_CODES = DotMap(
+ {
+ "MULTI_BUBBLE_WARN": 1,
+ "NO_MARKER_ERR": 2,
+ },
+ _dynamic=False,
+)
+
+FIELD_TYPES = {
+ "QTYPE_INT": {
+ "bubbleValues": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"],
+ "direction": "vertical",
+ },
+ "QTYPE_INT_FROM_1": {
+ "bubbleValues": ["1", "2", "3", "4", "5", "6", "7", "8", "9", "0"],
+ "direction": "vertical",
+ },
+ "QTYPE_MCQ4": {"bubbleValues": ["A", "B", "C", "D"], "direction": "horizontal"},
+ "QTYPE_MCQ5": {
+ "bubbleValues": ["A", "B", "C", "D", "E"],
+ "direction": "horizontal",
+ },
+ #
+ # You can create and append custom field types here-
+ #
+}
+
+# TODO: move to interaction.py
+TEXT_SIZE = 0.95
+CLR_BLACK = (50, 150, 150)
+CLR_WHITE = (250, 250, 250)
+CLR_GRAY = (130, 130, 130)
+CLR_DARK_GRAY = (100, 100, 100)
+
+# TODO: move to config.json
+GLOBAL_PAGE_THRESHOLD_WHITE = 200
+GLOBAL_PAGE_THRESHOLD_BLACK = 100
diff --git a/OMRChecker/src/core.py b/OMRChecker/src/core.py
new file mode 100644
index 0000000000000000000000000000000000000000..15279cf49fb9fb12c88611bcc69611ea524344dc
--- /dev/null
+++ b/OMRChecker/src/core.py
@@ -0,0 +1,721 @@
+import os
+from collections import defaultdict
+from typing import Any
+
+import cv2
+import matplotlib.pyplot as plt
+import numpy as np
+
+import src.constants as constants
+from src.logger import logger
+from src.utils.image import CLAHE_HELPER, ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class ImageInstanceOps:
+ """Class to hold fine-tuned utilities for a group of images. One instance for each processing directory."""
+
+ save_img_list: Any = defaultdict(list)
+
+ def __init__(self, tuning_config):
+ super().__init__()
+ self.tuning_config = tuning_config
+ self.save_image_level = tuning_config.outputs.save_image_level
+
+ def apply_preprocessors(self, file_path, in_omr, template):
+ tuning_config = self.tuning_config
+ # resize to conform to template
+ in_omr = ImageUtils.resize_util(
+ in_omr,
+ tuning_config.dimensions.processing_width,
+ tuning_config.dimensions.processing_height,
+ )
+
+ # run pre_processors in sequence
+ for pre_processor in template.pre_processors:
+ in_omr = pre_processor.apply_filter(in_omr, file_path)
+ return in_omr
+
+ def read_omr_response(self, template, image, name, save_dir=None):
+ config = self.tuning_config
+ auto_align = config.alignment_params.auto_align
+ try:
+ img = image.copy()
+ # origDim = img.shape[:2]
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ if img.max() > img.min():
+ img = ImageUtils.normalize_util(img)
+ # Processing copies
+ transp_layer = img.copy()
+ final_marked = img.copy()
+
+ morph = img.copy()
+ self.append_save_img(3, morph)
+
+ if auto_align:
+ # Note: clahe is good for morphology, bad for thresholding
+ morph = CLAHE_HELPER.apply(morph)
+ self.append_save_img(3, morph)
+ # Remove shadows further, make columns/boxes darker (less gamma)
+ morph = ImageUtils.adjust_gamma(
+ morph, config.threshold_params.GAMMA_LOW
+ )
+ # TODO: all numbers should come from either constants or config
+ _, morph = cv2.threshold(morph, 220, 220, cv2.THRESH_TRUNC)
+ morph = ImageUtils.normalize_util(morph)
+ self.append_save_img(3, morph)
+ if config.outputs.show_image_level >= 4:
+ InteractionUtils.show("morph1", morph, 0, 1, config)
+
+ # Move them to data class if needed
+ # Overlay Transparencies
+ alpha = 0.65
+ omr_response = {}
+ multi_marked, multi_roll = 0, 0
+
+ # TODO Make this part useful for visualizing status checks
+ # blackVals=[0]
+ # whiteVals=[255]
+
+ if config.outputs.show_image_level >= 5:
+ all_c_box_vals = {"int": [], "mcq": []}
+ # TODO: simplify this logic
+ q_nums = {"int": [], "mcq": []}
+
+ # Find Shifts for the field_blocks --> Before calculating threshold!
+ if auto_align:
+ # print("Begin Alignment")
+ # Open : erode then dilate
+ v_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (2, 10))
+ morph_v = cv2.morphologyEx(
+ morph, cv2.MORPH_OPEN, v_kernel, iterations=3
+ )
+ _, morph_v = cv2.threshold(morph_v, 200, 200, cv2.THRESH_TRUNC)
+ morph_v = 255 - ImageUtils.normalize_util(morph_v)
+
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morphed_vertical", morph_v, 0, 1, config=config
+ )
+
+ # InteractionUtils.show("morph1",morph,0,1,config=config)
+ # InteractionUtils.show("morphed_vertical",morph_v,0,1,config=config)
+
+ self.append_save_img(3, morph_v)
+
+ morph_thr = 60 # for Mobile images, 40 for scanned Images
+ _, morph_v = cv2.threshold(morph_v, morph_thr, 255, cv2.THRESH_BINARY)
+ # kernel best tuned to 5x5 now
+ morph_v = cv2.erode(morph_v, np.ones((5, 5), np.uint8), iterations=2)
+
+ self.append_save_img(3, morph_v)
+ # h_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (10, 2))
+ # morph_h = cv2.morphologyEx(morph, cv2.MORPH_OPEN, h_kernel, iterations=3)
+ # ret, morph_h = cv2.threshold(morph_h,200,200,cv2.THRESH_TRUNC)
+ # morph_h = 255 - normalize_util(morph_h)
+ # InteractionUtils.show("morph_h",morph_h,0,1,config=config)
+ # _, morph_h = cv2.threshold(morph_h,morph_thr,255,cv2.THRESH_BINARY)
+ # morph_h = cv2.erode(morph_h, np.ones((5,5),np.uint8), iterations = 2)
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morph_thr_eroded", morph_v, 0, 1, config=config
+ )
+
+ self.append_save_img(6, morph_v)
+
+ # template relative alignment code
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+
+ match_col, max_steps, align_stride, thk = map(
+ config.alignment_params.get,
+ [
+ "match_col",
+ "max_steps",
+ "stride",
+ "thickness",
+ ],
+ )
+ shift, steps = 0, 0
+ while steps < max_steps:
+ left_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0] + shift - thk : -thk + s[0] + shift + match_col,
+ ]
+ )
+ right_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0]
+ + shift
+ - match_col
+ + d[0]
+ + thk : thk
+ + s[0]
+ + shift
+ + d[0],
+ ]
+ )
+
+ # For demonstration purposes-
+ # if(field_block.name == "int1"):
+ # ret = morph_v.copy()
+ # cv2.rectangle(ret,
+ # (s[0]+shift-thk,s[1]),
+ # (s[0]+shift+thk+d[0],s[1]+d[1]),
+ # constants.CLR_WHITE,
+ # 3)
+ # appendSaveImg(6,ret)
+ # print(shift, left_mean, right_mean)
+ left_shift, right_shift = left_mean > 100, right_mean > 100
+ if left_shift:
+ if right_shift:
+ break
+ else:
+ shift -= align_stride
+ else:
+ if right_shift:
+ shift += align_stride
+ else:
+ break
+ steps += 1
+
+ field_block.shift = shift
+ # print("Aligned field_block: ",field_block.name,"Corrected Shift:",
+ # field_block.shift,", dimensions:", field_block.dimensions,
+ # "origin:", field_block.origin,'\n')
+ # print("End Alignment")
+
+ final_align = None
+ if config.outputs.show_image_level >= 2:
+ initial_align = self.draw_template_layout(img, template, shifted=False)
+ final_align = self.draw_template_layout(
+ img, template, shifted=True, draw_qvals=True
+ )
+ # appendSaveImg(4,mean_vals)
+ self.append_save_img(2, initial_align)
+ self.append_save_img(2, final_align)
+
+ if auto_align:
+ final_align = np.hstack((initial_align, final_align))
+ self.append_save_img(5, img)
+
+ # Get mean bubbleValues n other stats
+ all_q_vals, all_q_strip_arrs, all_q_std_vals = [], [], []
+ total_q_strip_no = 0
+ for field_block in template.field_blocks:
+ box_w, box_h = field_block.bubble_dimensions
+ q_std_vals = []
+ for field_block_bubbles in field_block.traverse_bubbles:
+ q_strip_vals = []
+ for pt in field_block_bubbles:
+ # shifted
+ x, y = (pt.x + field_block.shift, pt.y)
+ rect = [y, y + box_h, x, x + box_w]
+ q_strip_vals.append(
+ cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0]
+ # detectCross(img, rect) ? 100 : 0
+ )
+ q_std_vals.append(round(np.std(q_strip_vals), 2))
+ all_q_strip_arrs.append(q_strip_vals)
+ # _, _, _ = get_global_threshold(q_strip_vals, "QStrip Plot",
+ # plot_show=False, sort_in_plot=True)
+ # hist = getPlotImg()
+ # InteractionUtils.show("QStrip "+field_block_bubbles[0].field_label, hist, 0, 1,config=config)
+ all_q_vals.extend(q_strip_vals)
+ # print(total_q_strip_no, field_block_bubbles[0].field_label, q_std_vals[len(q_std_vals)-1])
+ total_q_strip_no += 1
+ all_q_std_vals.extend(q_std_vals)
+
+ global_std_thresh, _, _ = self.get_global_threshold(
+ all_q_std_vals
+ ) # , "Q-wise Std-dev Plot", plot_show=True, sort_in_plot=True)
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # Note: Plotting takes Significant times here --> Change Plotting args
+ # to support show_image_level
+ # , "Mean Intensity Histogram",plot_show=True, sort_in_plot=True)
+ global_thr, _, _ = self.get_global_threshold(all_q_vals, looseness=4)
+
+ logger.info(
+ f"Thresholding: \tglobal_thr: {round(global_thr, 2)} \tglobal_std_THR: {round(global_std_thresh, 2)}\t{'(Looks like a Xeroxed OMR)' if (global_thr == 255) else ''}"
+ )
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # if(config.outputs.show_image_level>=1):
+ # hist = getPlotImg()
+ # InteractionUtils.show("Hist", hist, 0, 1,config=config)
+ # appendSaveImg(4,hist)
+ # appendSaveImg(5,hist)
+ # appendSaveImg(2,hist)
+
+ per_omr_threshold_avg, total_q_strip_no, total_q_box_no = 0, 0, 0
+ for field_block in template.field_blocks:
+ block_q_strip_no = 1
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ s, d = field_block.origin, field_block.dimensions
+ key = field_block.name[:3]
+ # cv2.rectangle(final_marked,(s[0]+shift,s[1]),(s[0]+shift+d[0],
+ # s[1]+d[1]),CLR_BLACK,3)
+ for field_block_bubbles in field_block.traverse_bubbles:
+ # All Black or All White case
+ no_outliers = all_q_std_vals[total_q_strip_no] < global_std_thresh
+ # print(total_q_strip_no, field_block_bubbles[0].field_label,
+ # all_q_std_vals[total_q_strip_no], "no_outliers:", no_outliers)
+ per_q_strip_threshold = self.get_local_threshold(
+ all_q_strip_arrs[total_q_strip_no],
+ global_thr,
+ no_outliers,
+ f"Mean Intensity Histogram for {key}.{field_block_bubbles[0].field_label}.{block_q_strip_no}",
+ config.outputs.show_image_level >= 6,
+ )
+ # print(field_block_bubbles[0].field_label,key,block_q_strip_no, "THR: ",
+ # round(per_q_strip_threshold,2))
+ per_omr_threshold_avg += per_q_strip_threshold
+
+ # Note: Little debugging visualization - view the particular Qstrip
+ # if(
+ # 0
+ # # or "q17" in (field_block_bubbles[0].field_label)
+ # # or (field_block_bubbles[0].field_label+str(block_q_strip_no))=="q15"
+ # ):
+ # st, end = qStrip
+ # InteractionUtils.show("QStrip: "+key+"-"+str(block_q_strip_no),
+ # img[st[1] : end[1], st[0]+shift : end[0]+shift],0,config=config)
+
+ # TODO: get rid of total_q_box_no
+ detected_bubbles = []
+ for bubble in field_block_bubbles:
+ bubble_is_marked = (
+ per_q_strip_threshold > all_q_vals[total_q_box_no]
+ )
+ total_q_box_no += 1
+ if bubble_is_marked:
+ detected_bubbles.append(bubble)
+ x, y, field_value = (
+ bubble.x + field_block.shift,
+ bubble.y,
+ bubble.field_value,
+ )
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 12), int(y + box_h / 12)),
+ (
+ int(x + box_w - box_w / 12),
+ int(y + box_h - box_h / 12),
+ ),
+ constants.CLR_DARK_GRAY,
+ 3,
+ )
+
+ cv2.putText(
+ final_marked,
+ str(field_value),
+ (x, y),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ (20, 20, 10),
+ int(1 + 3.5 * constants.TEXT_SIZE),
+ )
+ else:
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (
+ int(x + box_w - box_w / 10),
+ int(y + box_h - box_h / 10),
+ ),
+ constants.CLR_GRAY,
+ -1,
+ )
+
+ for bubble in detected_bubbles:
+ field_label, field_value = (
+ bubble.field_label,
+ bubble.field_value,
+ )
+ # Only send rolls multi-marked in the directory
+ multi_marked_local = field_label in omr_response
+ omr_response[field_label] = (
+ (omr_response[field_label] + field_value)
+ if multi_marked_local
+ else field_value
+ )
+ # TODO: generalize this into identifier
+ # multi_roll = multi_marked_local and "Roll" in str(q)
+ multi_marked = multi_marked or multi_marked_local
+
+ if len(detected_bubbles) == 0:
+ field_label = field_block_bubbles[0].field_label
+ omr_response[field_label] = field_block.empty_val
+
+ if config.outputs.show_image_level >= 5:
+ if key in all_c_box_vals:
+ q_nums[key].append(f"{key[:2]}_c{str(block_q_strip_no)}")
+ all_c_box_vals[key].append(
+ all_q_strip_arrs[total_q_strip_no]
+ )
+
+ block_q_strip_no += 1
+ total_q_strip_no += 1
+ # /for field_block
+
+ per_omr_threshold_avg /= total_q_strip_no
+ per_omr_threshold_avg = round(per_omr_threshold_avg, 2)
+ # Translucent
+ cv2.addWeighted(
+ final_marked, alpha, transp_layer, 1 - alpha, 0, final_marked
+ )
+ # Box types
+ if config.outputs.show_image_level >= 6:
+ # plt.draw()
+ f, axes = plt.subplots(len(all_c_box_vals), sharey=True)
+ f.canvas.manager.set_window_title(name)
+ ctr = 0
+ type_name = {
+ "int": "Integer",
+ "mcq": "MCQ",
+ "med": "MED",
+ "rol": "Roll",
+ }
+ for k, boxvals in all_c_box_vals.items():
+ axes[ctr].title.set_text(type_name[k] + " Type")
+ axes[ctr].boxplot(boxvals)
+ # thrline=axes[ctr].axhline(per_omr_threshold_avg,color='red',ls='--')
+ # thrline.set_label("Average THR")
+ axes[ctr].set_ylabel("Intensity")
+ axes[ctr].set_xticklabels(q_nums[k])
+ # axes[ctr].legend()
+ ctr += 1
+ # imshow will do the waiting
+ plt.tight_layout(pad=0.5)
+ plt.show()
+
+ if config.outputs.show_image_level >= 3 and final_align is not None:
+ final_align = ImageUtils.resize_util_h(
+ final_align, int(config.dimensions.display_height)
+ )
+ # [final_align.shape[1],0])
+ InteractionUtils.show(
+ "Template Alignment Adjustment", final_align, 0, 0, config=config
+ )
+
+ if config.outputs.save_detections and save_dir is not None:
+ if multi_roll:
+ save_dir = save_dir.joinpath("_MULTI_")
+ image_path = str(save_dir.joinpath(name))
+ ImageUtils.save_img(image_path, final_marked)
+
+ self.append_save_img(2, final_marked)
+
+ if save_dir is not None:
+ for i in range(config.outputs.save_image_level):
+ self.save_image_stacks(i + 1, name, save_dir)
+
+ return omr_response, final_marked, multi_marked, multi_roll
+
+ except Exception as e:
+ raise e
+
+ @staticmethod
+ def draw_template_layout(img, template, shifted=True, draw_qvals=False, border=-1):
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ final_align = img.copy()
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ if shifted:
+ cv2.rectangle(
+ final_align,
+ (s[0] + shift, s[1]),
+ (s[0] + shift + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ else:
+ cv2.rectangle(
+ final_align,
+ (s[0], s[1]),
+ (s[0] + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ for field_block_bubbles in field_block.traverse_bubbles:
+ for pt in field_block_bubbles:
+ x, y = (pt.x + field_block.shift, pt.y) if shifted else (pt.x, pt.y)
+ cv2.rectangle(
+ final_align,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (int(x + box_w - box_w / 10), int(y + box_h - box_h / 10)),
+ constants.CLR_GRAY,
+ border,
+ )
+ if draw_qvals:
+ rect = [y, y + box_h, x, x + box_w]
+ cv2.putText(
+ final_align,
+ f"{int(cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0])}",
+ (rect[2] + 2, rect[0] + (box_h * 2) // 3),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ 0.6,
+ constants.CLR_BLACK,
+ 2,
+ )
+ if shifted:
+ text_in_px = cv2.getTextSize(
+ field_block.name, cv2.FONT_HERSHEY_SIMPLEX, constants.TEXT_SIZE, 4
+ )
+ cv2.putText(
+ final_align,
+ field_block.name,
+ (int(s[0] + d[0] - text_in_px[0][0]), int(s[1] - text_in_px[0][1])),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ constants.CLR_BLACK,
+ 4,
+ )
+ return final_align
+
+ def get_global_threshold(
+ self,
+ q_vals_orig,
+ plot_title=None,
+ plot_show=True,
+ sort_in_plot=True,
+ looseness=1,
+ ):
+ """
+ Note: Cannot assume qStrip has only-gray or only-white bg
+ (in which case there is only one jump).
+ So there will be either 1 or 2 jumps.
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ 2 Jumps :
+ ......
+ |||||| <-- wrong THR
+ ....||||||
+ |||||||||| <-- safe THR
+ ..||||||||||
+ ||||||||||||
+
+ The abstract "First LARGE GAP" is perfect for this.
+ Current code is considering ONLY TOP 2 jumps(>= MIN_GAP) to be big,
+ gives the smaller one
+
+ """
+ config = self.tuning_config
+ PAGE_TYPE_FOR_THRESHOLD, MIN_JUMP, JUMP_DELTA = map(
+ config.threshold_params.get,
+ [
+ "PAGE_TYPE_FOR_THRESHOLD",
+ "MIN_JUMP",
+ "JUMP_DELTA",
+ ],
+ )
+
+ global_default_threshold = (
+ constants.GLOBAL_PAGE_THRESHOLD_WHITE
+ if PAGE_TYPE_FOR_THRESHOLD == "white"
+ else constants.GLOBAL_PAGE_THRESHOLD_BLACK
+ )
+
+ # Sort the Q bubbleValues
+ # TODO: Change var name of q_vals
+ q_vals = sorted(q_vals_orig)
+ # Find the FIRST LARGE GAP and set it as threshold:
+ ls = (looseness + 1) // 2
+ l = len(q_vals) - ls
+ max1, thr1 = MIN_JUMP, global_default_threshold
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - ls] + jump / 2
+
+ # NOTE: thr2 is deprecated, thus is JUMP_DELTA
+ # Make use of the fact that the JUMP_DELTA(Vertical gap ofc) between
+ # values at detected jumps would be atleast 20
+ max2, thr2 = MIN_JUMP, global_default_threshold
+ # Requires atleast 1 gray box to be present (Roll field will ensure this)
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ new_thr = q_vals[i - ls] + jump / 2
+ if jump > max2 and abs(thr1 - new_thr) > JUMP_DELTA:
+ max2 = jump
+ thr2 = new_thr
+ # global_thr = min(thr1,thr2)
+ global_thr, j_low, j_high = thr1, thr1 - max1 // 2, thr1 + max1 // 2
+
+ # # For normal images
+ # thresholdRead = 116
+ # if(thr1 > thr2 and thr2 > thresholdRead):
+ # print("Note: taking safer thr line.")
+ # global_thr, j_low, j_high = thr2, thr2 - max2//2, thr2 + max2//2
+
+ if plot_title:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals_orig)), q_vals if sort_in_plot else q_vals_orig)
+ ax.set_title(plot_title)
+ thrline = ax.axhline(global_thr, color="green", ls="--", linewidth=5)
+ thrline.set_label("Global Threshold")
+ thrline = ax.axhline(thr2, color="red", ls=":", linewidth=3)
+ thrline.set_label("THR2 Line")
+ # thrline=ax.axhline(j_low,color='red',ls='-.', linewidth=3)
+ # thrline=ax.axhline(j_high,color='red',ls='-.', linewidth=3)
+ # thrline.set_label("Boundary Line")
+ # ax.set_ylabel("Mean Intensity")
+ ax.set_ylabel("Values")
+ ax.set_xlabel("Position")
+ ax.legend()
+ if plot_show:
+ plt.title(plot_title)
+ plt.show()
+
+ return global_thr, j_low, j_high
+
+ def get_local_threshold(
+ self, q_vals, global_thr, no_outliers, plot_title=None, plot_show=True
+ ):
+ """
+ TODO: Update this documentation too-
+ //No more - Assumption : Colwise background color is uniformly gray or white,
+ but not alternating. In this case there is atmost one jump.
+
+ 0 Jump :
+ <-- safe THR?
+ .......
+ ...|||||||
+ |||||||||| <-- safe THR?
+ // How to decide given range is above or below gray?
+ -> global q_vals shall absolutely help here. Just run same function
+ on total q_vals instead of colwise _//
+ How to decide it is this case of 0 jumps
+
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ """
+ config = self.tuning_config
+ # Sort the Q bubbleValues
+ q_vals = sorted(q_vals)
+
+ # Small no of pts cases:
+ # base case: 1 or 2 pts
+ if len(q_vals) < 3:
+ thr1 = (
+ global_thr
+ if np.max(q_vals) - np.min(q_vals) < config.threshold_params.MIN_GAP
+ else np.mean(q_vals)
+ )
+ else:
+ # qmin, qmax, qmean, qstd = round(np.min(q_vals),2), round(np.max(q_vals),2),
+ # round(np.mean(q_vals),2), round(np.std(q_vals),2)
+ # GVals = [round(abs(q-qmean),2) for q in q_vals]
+ # gmean, gstd = round(np.mean(GVals),2), round(np.std(GVals),2)
+ # # DISCRETION: Pretty critical factor in reading response
+ # # Doesn't work well for small number of values.
+ # DISCRETION = 2.7 # 2.59 was closest hit, 3.0 is too far
+ # L2MaxGap = round(max([abs(g-gmean) for g in GVals]),2)
+ # if(L2MaxGap > DISCRETION*gstd):
+ # no_outliers = False
+
+ # # ^Stackoverflow method
+ # print(field_label, no_outliers,"qstd",round(np.std(q_vals),2), "gstd", gstd,
+ # "Gaps in gvals",sorted([round(abs(g-gmean),2) for g in GVals],reverse=True),
+ # '\t',round(DISCRETION*gstd,2), L2MaxGap)
+
+ # else:
+ # Find the LARGEST GAP and set it as threshold: //(FIRST LARGE GAP)
+ l = len(q_vals) - 1
+ max1, thr1 = config.threshold_params.MIN_JUMP, 255
+ for i in range(1, l):
+ jump = q_vals[i + 1] - q_vals[i - 1]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - 1] + jump / 2
+ # print(field_label,q_vals,max1)
+
+ confident_jump = (
+ config.threshold_params.MIN_JUMP
+ + config.threshold_params.CONFIDENT_SURPLUS
+ )
+ # If not confident, then only take help of global_thr
+ if max1 < confident_jump:
+ if no_outliers:
+ # All Black or All White case
+ thr1 = global_thr
+ else:
+ # TODO: Low confidence parameters here
+ pass
+
+ # if(thr1 == 255):
+ # print("Warning: threshold is unexpectedly 255! (Outlier Delta issue?)",plot_title)
+
+ # Make a common plot function to show local and global thresholds
+ if plot_show and plot_title is not None:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals)), q_vals)
+ thrline = ax.axhline(thr1, color="green", ls=("-."), linewidth=3)
+ thrline.set_label("Local Threshold")
+ thrline = ax.axhline(global_thr, color="red", ls=":", linewidth=5)
+ thrline.set_label("Global Threshold")
+ ax.set_title(plot_title)
+ ax.set_ylabel("Bubble Mean Intensity")
+ ax.set_xlabel("Bubble Number(sorted)")
+ ax.legend()
+ # TODO append QStrip to this plot-
+ # appendSaveImg(6,getPlotImg())
+ if plot_show:
+ plt.show()
+ return thr1
+
+ def append_save_img(self, key, img):
+ if self.save_image_level >= int(key):
+ self.save_img_list[key].append(img.copy())
+
+ def save_image_stacks(self, key, filename, save_dir):
+ config = self.tuning_config
+ if self.save_image_level >= int(key) and self.save_img_list[key] != []:
+ name = os.path.splitext(filename)[0]
+ result = np.hstack(
+ tuple(
+ [
+ ImageUtils.resize_util_h(img, config.dimensions.display_height)
+ for img in self.save_img_list[key]
+ ]
+ )
+ )
+ result = ImageUtils.resize_util(
+ result,
+ min(
+ len(self.save_img_list[key]) * config.dimensions.display_width // 3,
+ int(config.dimensions.display_width * 2.5),
+ ),
+ )
+ ImageUtils.save_img(f"{save_dir}stack/{name}_{str(key)}_stack.jpg", result)
+
+ def reset_all_save_img(self):
+ for i in range(self.save_image_level):
+ self.save_img_list[i + 1] = []
diff --git a/OMRChecker/src/defaults/__init__.py b/OMRChecker/src/defaults/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..db3d42e87b3e47d0cbee08032e65e87ac1bd8165
--- /dev/null
+++ b/OMRChecker/src/defaults/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+# Use all imports relative to root directory
+# (https://chrisyeh96.github.io/2017/08/08/definitive-guide-python-imports.html)
+from src.defaults.config import * # NOQA
+from src.defaults.template import * # NOQA
diff --git a/OMRChecker/src/defaults/config.py b/OMRChecker/src/defaults/config.py
new file mode 100644
index 0000000000000000000000000000000000000000..6d1315fb45b0e736ad7daedcea5f3b23e3a9078d
--- /dev/null
+++ b/OMRChecker/src/defaults/config.py
@@ -0,0 +1,35 @@
+from dotmap import DotMap
+
+CONFIG_DEFAULTS = DotMap(
+ {
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666,
+ },
+ "threshold_params": {
+ "GAMMA_LOW": 0.7,
+ "MIN_GAP": 30,
+ "MIN_JUMP": 25,
+ "CONFIDENT_SURPLUS": 5,
+ "JUMP_DELTA": 30,
+ "PAGE_TYPE_FOR_THRESHOLD": "white",
+ },
+ "alignment_params": {
+ # Note: 'auto_align' enables automatic template alignment, use if the scans show slight misalignments.
+ "auto_align": False,
+ "match_col": 5,
+ "max_steps": 20,
+ "stride": 1,
+ "thickness": 3,
+ },
+ "outputs": {
+ "show_image_level": 0,
+ "save_image_level": 0,
+ "save_detections": True,
+ "filter_out_multimarked_files": False,
+ },
+ },
+ _dynamic=False,
+)
diff --git a/OMRChecker/src/defaults/template.py b/OMRChecker/src/defaults/template.py
new file mode 100644
index 0000000000000000000000000000000000000000..efd75420f01cf49f72a2e285734c0d79cafe4837
--- /dev/null
+++ b/OMRChecker/src/defaults/template.py
@@ -0,0 +1,6 @@
+TEMPLATE_DEFAULTS = {
+ "preProcessors": [],
+ "emptyValue": "",
+ "customLabels": {},
+ "outputColumns": [],
+}
diff --git a/OMRChecker/src/entry.py b/OMRChecker/src/entry.py
new file mode 100644
index 0000000000000000000000000000000000000000..94d3137c3ac99502399cbb40d6d378cd8249d8de
--- /dev/null
+++ b/OMRChecker/src/entry.py
@@ -0,0 +1,371 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import os
+from csv import QUOTE_NONNUMERIC
+from pathlib import Path
+from time import time
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src import constants
+from src.defaults import CONFIG_DEFAULTS
+from src.evaluation import EvaluationConfig, evaluate_concatenated_response
+from src.logger import console, logger
+from src.template import Template
+from src.utils.file import Paths, setup_dirs_for_paths, setup_outputs_for_template
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils, Stats
+from src.utils.parsing import get_concatenated_response, open_config_with_defaults
+
+# Load processors
+STATS = Stats()
+
+
+def entry_point(input_dir, args):
+ if not os.path.exists(input_dir):
+ raise Exception(f"Given input directory does not exist: '{input_dir}'")
+ curr_dir = input_dir
+ return process_dir(input_dir, curr_dir, args)
+
+
+def print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+):
+ logger.info("")
+ table = Table(title="Current Configurations", show_header=False, show_lines=False)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Value", style="magenta")
+ table.add_row("Directory Path", f"{curr_dir}")
+ table.add_row("Count of Images", f"{len(omr_files)}")
+ table.add_row("Set Layout Mode ", "ON" if args["setLayout"] else "OFF")
+ pre_processor_names = [pp.__class__.__name__ for pp in template.pre_processors]
+ table.add_row(
+ "Markers Detection",
+ "ON" if "CropOnMarkers" in pre_processor_names else "OFF",
+ )
+ table.add_row("Auto Alignment", f"{tuning_config.alignment_params.auto_align}")
+ table.add_row("Detected Template Path", f"{template}")
+ if local_config_path:
+ table.add_row("Detected Local Config", f"{local_config_path}")
+ if evaluation_config:
+ table.add_row("Detected Evaluation Config", f"{evaluation_config}")
+
+ table.add_row(
+ "Detected pre-processors",
+ ", ".join(pre_processor_names),
+ )
+ console.print(table, justify="center")
+
+
+def process_dir(
+ root_dir,
+ curr_dir,
+ args,
+ template=None,
+ tuning_config=CONFIG_DEFAULTS,
+ evaluation_config=None,
+):
+ # Update local tuning_config (in current recursion stack)
+ local_config_path = curr_dir.joinpath(constants.CONFIG_FILENAME)
+ if os.path.exists(local_config_path):
+ tuning_config = open_config_with_defaults(local_config_path)
+
+ # Update local template (in current recursion stack)
+ local_template_path = curr_dir.joinpath(constants.TEMPLATE_FILENAME)
+ local_template_exists = os.path.exists(local_template_path)
+ if local_template_exists:
+ template = Template(
+ local_template_path,
+ tuning_config,
+ )
+ # Look for subdirectories for processing
+ subdirs = [d for d in curr_dir.iterdir() if d.is_dir()]
+
+ output_dir = Path(args["output_dir"], curr_dir.relative_to(root_dir))
+ paths = Paths(output_dir)
+
+ # look for images in current dir to process
+ exts = ("*.[pP][nN][gG]", "*.[jJ][pP][gG]", "*.[jJ][pP][eE][gG]")
+ omr_files = sorted([f for ext in exts for f in curr_dir.glob(ext)])
+
+ # Exclude images (take union over all pre_processors)
+ excluded_files = []
+ if template:
+ for pp in template.pre_processors:
+ excluded_files.extend(Path(p) for p in pp.exclude_files())
+
+ local_evaluation_path = curr_dir.joinpath(constants.EVALUATION_FILENAME)
+ if not args["setLayout"] and os.path.exists(local_evaluation_path):
+ if not local_template_exists:
+ logger.warning(
+ f"Found an evaluation file without a parent template file: {local_evaluation_path}"
+ )
+ evaluation_config = EvaluationConfig(
+ curr_dir,
+ local_evaluation_path,
+ template,
+ tuning_config,
+ )
+
+ excluded_files.extend(
+ Path(exclude_file) for exclude_file in evaluation_config.get_exclude_files()
+ )
+
+ omr_files = [f for f in omr_files if f not in excluded_files]
+
+ if omr_files:
+ if not template:
+ logger.error(
+ f"Found images, but no template in the directory tree \
+ of '{curr_dir}'. \nPlace {constants.TEMPLATE_FILENAME} in the \
+ appropriate directory."
+ )
+ raise Exception(
+ f"No template file found in the directory tree of {curr_dir}"
+ )
+
+ setup_dirs_for_paths(paths)
+ outputs_namespace = setup_outputs_for_template(paths, template)
+
+ print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+ )
+ if args["setLayout"]:
+ show_template_layouts(omr_files, template, tuning_config)
+ else:
+ process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+ )
+
+ elif not subdirs:
+ # Each subdirectory should have images or should be non-leaf
+ logger.info(
+ f"No valid images or sub-folders found in {curr_dir}.\
+ Empty directories not allowed."
+ )
+
+ # recursively process sub-folders
+ for d in subdirs:
+ process_dir(
+ root_dir,
+ d,
+ args,
+ template,
+ tuning_config,
+ evaluation_config,
+ )
+
+
+def show_template_layouts(omr_files, template, tuning_config):
+ for file_path in omr_files:
+ file_name = file_path.name
+ file_path = str(file_path)
+ in_omr = cv2.imread(file_path, cv2.IMREAD_GRAYSCALE)
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+ template_layout = template.image_instance_ops.draw_template_layout(
+ in_omr, template, shifted=False, border=2
+ )
+ InteractionUtils.show(
+ f"Template Layout: {file_name}", template_layout, 1, 1, config=tuning_config
+ )
+
+
+def process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+):
+ start_time = int(time())
+ files_counter = 0
+ STATS.files_not_moved = 0
+
+ for file_path in omr_files:
+ files_counter += 1
+ file_name = file_path.name
+
+ in_omr = cv2.imread(str(file_path), cv2.IMREAD_GRAYSCALE)
+
+ logger.info("")
+ logger.info(
+ f"({files_counter}) Opening image: \t'{file_path}'\tResolution: {in_omr.shape}"
+ )
+
+ template.image_instance_ops.reset_all_save_img()
+
+ template.image_instance_ops.append_save_img(1, in_omr)
+
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+
+ if in_omr is None:
+ # Error OMR case
+ new_file_path = outputs_namespace.paths.errors_dir.joinpath(file_name)
+ outputs_namespace.OUTPUT_SET.append(
+ [file_name] + outputs_namespace.empty_resp
+ )
+ if check_and_move(
+ constants.ERROR_CODES.NO_MARKER_ERR, file_path, new_file_path
+ ):
+ err_line = [
+ file_name,
+ file_path,
+ new_file_path,
+ "NA",
+ ] + outputs_namespace.empty_resp
+ pd.DataFrame(err_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Errors"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ continue
+
+ # uniquify
+ file_id = str(file_name)
+ save_dir = outputs_namespace.paths.save_marked_dir
+ (
+ response_dict,
+ final_marked,
+ multi_marked,
+ _,
+ ) = template.image_instance_ops.read_omr_response(
+ template, image=in_omr, name=file_id, save_dir=save_dir
+ )
+
+ # TODO: move inner try catch here
+ # concatenate roll nos, set unmarked responses, etc
+ omr_response = get_concatenated_response(response_dict, template)
+
+ if (
+ evaluation_config is None
+ or not evaluation_config.get_should_explain_scoring()
+ ):
+ logger.info(f"Read Response: \n{omr_response}")
+
+ score = 0
+ if evaluation_config is not None:
+ score = evaluate_concatenated_response(
+ omr_response, evaluation_config, file_path, outputs_namespace.paths.evaluation_dir
+ )
+ logger.info(
+ f"(/{files_counter}) Graded with score: {round(score, 2)}\t for file: '{file_id}'"
+ )
+ else:
+ logger.info(f"(/{files_counter}) Processed file: '{file_id}'")
+
+ if tuning_config.outputs.show_image_level >= 2:
+ InteractionUtils.show(
+ f"Final Marked Bubbles : '{file_id}'",
+ ImageUtils.resize_util_h(
+ final_marked, int(tuning_config.dimensions.display_height * 1.3)
+ ),
+ 1,
+ 1,
+ config=tuning_config,
+ )
+
+ resp_array = []
+ for k in template.output_columns:
+ resp_array.append(omr_response[k])
+
+ outputs_namespace.OUTPUT_SET.append([file_name] + resp_array)
+
+ if multi_marked == 0 or not tuning_config.outputs.filter_out_multimarked_files:
+ STATS.files_not_moved += 1
+ new_file_path = save_dir.joinpath(file_id)
+ # Enter into Results sheet-
+ results_line = [file_name, file_path, new_file_path, score] + resp_array
+ # Write/Append to results_line file(opened in append mode)
+ pd.DataFrame(results_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Results"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ else:
+ # multi_marked file
+ logger.info(f"[{files_counter}] Found multi-marked file: '{file_id}'")
+ new_file_path = outputs_namespace.paths.multi_marked_dir.joinpath(file_name)
+ if check_and_move(
+ constants.ERROR_CODES.MULTI_BUBBLE_WARN, file_path, new_file_path
+ ):
+ mm_line = [file_name, file_path, new_file_path, "NA"] + resp_array
+ pd.DataFrame(mm_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["MultiMarked"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ # else:
+ # TODO: Add appropriate record handling here
+ # pass
+
+ print_stats(start_time, files_counter, tuning_config)
+
+
+def check_and_move(error_code, file_path, filepath2):
+ # TODO: fix file movement into error/multimarked/invalid etc again
+ STATS.files_not_moved += 1
+ return True
+
+
+def print_stats(start_time, files_counter, tuning_config):
+ time_checking = max(1, round(time() - start_time, 2))
+ log = logger.info
+ log("")
+ log(f"{'Total file(s) moved': <27}: {STATS.files_moved}")
+ log(f"{'Total file(s) not moved': <27}: {STATS.files_not_moved}")
+ log("--------------------------------")
+ log(
+ f"{'Total file(s) processed': <27}: {files_counter} ({'Sum Tallied!' if files_counter == (STATS.files_moved + STATS.files_not_moved) else 'Not Tallying!'})"
+ )
+
+ if tuning_config.outputs.show_image_level <= 0:
+ log(
+ f"\nFinished Checking {files_counter} file(s) in {round(time_checking, 1)} seconds i.e. ~{round(time_checking / 60, 1)} minute(s)."
+ )
+ log(
+ f"{'OMR Processing Rate': <27}: \t ~ {round(time_checking / files_counter, 2)} seconds/OMR"
+ )
+ log(
+ f"{'OMR Processing Speed': <27}: \t ~ {round((files_counter * 60) / time_checking, 2)} OMRs/minute"
+ )
+ else:
+ log(f"\n{'Total script time': <27}: {time_checking} seconds")
+
+ if tuning_config.outputs.show_image_level <= 1:
+ log(
+ "\nTip: To see some awesome visuals, open config.json and increase 'show_image_level'"
+ )
diff --git a/OMRChecker/src/evaluation.py b/OMRChecker/src/evaluation.py
new file mode 100644
index 0000000000000000000000000000000000000000..55c2455a2967063e7604d0f0c904c09720e1064d
--- /dev/null
+++ b/OMRChecker/src/evaluation.py
@@ -0,0 +1,546 @@
+import ast
+import os
+import re
+from copy import deepcopy
+from csv import QUOTE_NONNUMERIC
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src.logger import console, logger
+from src.schemas.constants import (
+ BONUS_SECTION_PREFIX,
+ DEFAULT_SECTION_KEY,
+ MARKING_VERDICT_TYPES,
+)
+from src.utils.parsing import (
+ get_concatenated_response,
+ open_evaluation_with_validation,
+ parse_fields,
+ parse_float_or_fraction,
+)
+
+
+class AnswerMatcher:
+ def __init__(self, answer_item, section_marking_scheme):
+ self.section_marking_scheme = section_marking_scheme
+ self.answer_item = answer_item
+ self.answer_type = self.validate_and_get_answer_type(answer_item)
+ self.set_defaults_from_scheme(section_marking_scheme)
+
+ @staticmethod
+ def is_a_marking_score(answer_element):
+ # Note: strict type checking is already done at schema validation level,
+ # Here we focus on overall struct type
+ return type(answer_element) == str or type(answer_element) == int
+
+ @staticmethod
+ def is_standard_answer(answer_element):
+ return type(answer_element) == str and len(answer_element) >= 1
+
+ def validate_and_get_answer_type(self, answer_item):
+ if self.is_standard_answer(answer_item):
+ return "standard"
+ elif type(answer_item) == list:
+ if (
+ # Array of answer elements: ['A', 'B', 'AB']
+ len(answer_item) >= 2
+ and all(
+ self.is_standard_answer(answers_or_score)
+ for answers_or_score in answer_item
+ )
+ ):
+ return "multiple-correct"
+ elif (
+ # Array of two-tuples: [['A', 1], ['B', 1], ['C', 3], ['AB', 2]]
+ len(answer_item) >= 1
+ and all(
+ type(answer_and_score) == list and len(answer_and_score) == 2
+ for answer_and_score in answer_item
+ )
+ and all(
+ self.is_standard_answer(allowed_answer)
+ and self.is_a_marking_score(answer_score)
+ for allowed_answer, answer_score in answer_item
+ )
+ ):
+ return "multiple-correct-weighted"
+
+ logger.critical(
+ f"Unable to determine answer type for answer item: {answer_item}"
+ )
+ raise Exception("Unable to determine answer type")
+
+ def set_defaults_from_scheme(self, section_marking_scheme):
+ answer_type = self.answer_type
+ self.empty_val = section_marking_scheme.empty_val
+ answer_item = self.answer_item
+ self.marking = deepcopy(section_marking_scheme.marking)
+ # TODO: reuse part of parse_scheme_marking here -
+ if answer_type == "standard":
+ # no local overrides
+ pass
+ elif answer_type == "multiple-correct":
+ # override marking scheme scores for each allowed answer
+ for allowed_answer in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = self.marking["correct"]
+ elif answer_type == "multiple-correct-weighted":
+ # Note: No override using marking scheme as answer scores are provided in answer_item
+ for allowed_answer, answer_score in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = parse_float_or_fraction(
+ answer_score
+ )
+
+ def get_marking_scheme(self):
+ return self.section_marking_scheme
+
+ def get_section_explanation(self):
+ answer_type = self.answer_type
+ if answer_type in ["standard", "multiple-correct"]:
+ return self.section_marking_scheme.section_key
+ elif answer_type == "multiple-correct-weighted":
+ return f"Custom: {self.marking}"
+
+ def get_verdict_marking(self, marked_answer):
+ answer_type = self.answer_type
+ question_verdict = "incorrect"
+ if answer_type == "standard":
+ question_verdict = self.get_standard_verdict(marked_answer)
+ elif answer_type == "multiple-correct":
+ question_verdict = self.get_multiple_correct_verdict(marked_answer)
+ elif answer_type == "multiple-correct-weighted":
+ question_verdict = self.get_multiple_correct_weighted_verdict(marked_answer)
+ return question_verdict, self.marking[question_verdict]
+
+ def get_standard_verdict(self, marked_answer):
+ allowed_answer = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer == allowed_answer:
+ return "correct"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_verdict(self, marked_answer):
+ allowed_answers = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_weighted_verdict(self, marked_answer):
+ allowed_answers = [
+ allowed_answer for allowed_answer, _answer_score in self.answer_item
+ ]
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def __str__(self):
+ return f"{self.answer_item}"
+
+
+class SectionMarkingScheme:
+ def __init__(self, section_key, section_scheme, empty_val):
+ # TODO: get local empty_val from qblock
+ self.empty_val = empty_val
+ self.section_key = section_key
+ # DEFAULT marking scheme follows a shorthand
+ if section_key == DEFAULT_SECTION_KEY:
+ self.questions = None
+ self.marking = self.parse_scheme_marking(section_scheme)
+ else:
+ self.questions = parse_fields(section_key, section_scheme["questions"])
+ self.marking = self.parse_scheme_marking(section_scheme["marking"])
+
+ def __str__(self):
+ return self.section_key
+
+ def parse_scheme_marking(self, marking):
+ parsed_marking = {}
+ for verdict_type in MARKING_VERDICT_TYPES:
+ verdict_marking = parse_float_or_fraction(marking[verdict_type])
+ if (
+ verdict_marking > 0
+ and verdict_type == "incorrect"
+ and not self.section_key.startswith(BONUS_SECTION_PREFIX)
+ ):
+ logger.warning(
+ f"Found positive marks({round(verdict_marking, 2)}) for incorrect answer in the schema '{self.section_key}'. For Bonus sections, add a prefix 'BONUS_' to them."
+ )
+ parsed_marking[verdict_type] = verdict_marking
+
+ return parsed_marking
+
+ def match_answer(self, marked_answer, answer_matcher):
+ question_verdict, verdict_marking = answer_matcher.get_verdict_marking(
+ marked_answer
+ )
+
+ return verdict_marking, question_verdict
+
+
+class EvaluationConfig:
+ """Note: this instance will be reused for multiple omr sheets"""
+
+ def __init__(self, curr_dir, evaluation_path, template, tuning_config):
+ self.path = evaluation_path
+ evaluation_json = open_evaluation_with_validation(evaluation_path)
+ options, marking_schemes, source_type = map(
+ evaluation_json.get, ["options", "marking_schemes", "source_type"]
+ )
+ self.should_explain_scoring = options.get("should_explain_scoring", False)
+ self.has_non_default_section = False
+ self.exclude_files = []
+ self.enable_evaluation_table_to_csv = options.get(
+ "enable_evaluation_table_to_csv", False
+ )
+
+ if source_type == "csv":
+ csv_path = curr_dir.joinpath(options["answer_key_csv_path"])
+ if not os.path.exists(csv_path):
+ logger.warning(f"Answer key csv does not exist at: '{csv_path}'.")
+
+ answer_key_image_path = options.get("answer_key_image_path", None)
+ if os.path.exists(csv_path):
+ # TODO: CSV parsing/validation for each row with a (qNo, ) pair
+ answer_key = pd.read_csv(
+ csv_path,
+ header=None,
+ names=["question", "answer"],
+ converters={"question": str, "answer": self.parse_answer_column},
+ )
+
+ self.questions_in_order = answer_key["question"].to_list()
+ answers_in_order = answer_key["answer"].to_list()
+ elif not answer_key_image_path:
+ raise Exception(f"Answer key csv not found at '{csv_path}'")
+ else:
+ image_path = str(curr_dir.joinpath(answer_key_image_path))
+ if not os.path.exists(image_path):
+ raise Exception(f"Answer key image not found at '{image_path}'")
+
+ # self.exclude_files.append(image_path)
+
+ logger.debug(
+ f"Attempting to generate answer key from image: '{image_path}'"
+ )
+ # TODO: use a common function for below changes?
+ in_omr = cv2.imread(image_path, cv2.IMREAD_GRAYSCALE)
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ image_path, in_omr, template
+ )
+ if in_omr is None:
+ raise Exception(
+ f"Could not read answer key from image {image_path}"
+ )
+ (
+ response_dict,
+ _final_marked,
+ _multi_marked,
+ _multi_roll,
+ ) = template.image_instance_ops.read_omr_response(
+ template,
+ image=in_omr,
+ name=image_path,
+ save_dir=None,
+ )
+ omr_response = get_concatenated_response(response_dict, template)
+
+ empty_val = template.global_empty_val
+ empty_answer_regex = (
+ rf"{re.escape(empty_val)}+" if empty_val != "" else r"^$"
+ )
+
+ if "questions_in_order" in options:
+ self.questions_in_order = self.parse_questions_in_order(
+ options["questions_in_order"]
+ )
+ empty_answered_questions = [
+ question
+ for question in self.questions_in_order
+ if re.search(empty_answer_regex, omr_response[question])
+ ]
+ if len(empty_answered_questions) > 0:
+ logger.error(
+ f"Found empty answers for questions: {empty_answered_questions}, empty value used: '{empty_val}'"
+ )
+ raise Exception(
+ f"Found empty answers in file '{image_path}'. Please check your template again in the --setLayout mode."
+ )
+ else:
+ logger.warning(
+ f"questions_in_order not provided, proceeding to use non-empty values as answer key"
+ )
+ self.questions_in_order = sorted(
+ question
+ for (question, answer) in omr_response.items()
+ if not re.search(empty_answer_regex, answer)
+ )
+ answers_in_order = [
+ omr_response[question] for question in self.questions_in_order
+ ]
+ # TODO: save the CSV
+ else:
+ self.questions_in_order = self.parse_questions_in_order(
+ options["questions_in_order"]
+ )
+ answers_in_order = options["answers_in_order"]
+
+ self.validate_questions(answers_in_order)
+
+ self.section_marking_schemes, self.question_to_scheme = {}, {}
+ for section_key, section_scheme in marking_schemes.items():
+ section_marking_scheme = SectionMarkingScheme(
+ section_key, section_scheme, template.global_empty_val
+ )
+ if section_key != DEFAULT_SECTION_KEY:
+ self.section_marking_schemes[section_key] = section_marking_scheme
+ for q in section_marking_scheme.questions:
+ # TODO: check the answer key for custom scheme here?
+ self.question_to_scheme[q] = section_marking_scheme
+ self.has_non_default_section = True
+ else:
+ self.default_marking_scheme = section_marking_scheme
+
+ self.validate_marking_schemes()
+
+ self.question_to_answer_matcher = self.parse_answers_and_map_questions(
+ answers_in_order
+ )
+ self.validate_answers(answers_in_order, tuning_config)
+
+ def __str__(self):
+ return str(self.path)
+
+ # Externally called methods have higher abstraction level.
+ def prepare_and_validate_omr_response(self, omr_response):
+ self.reset_explanation_table()
+
+ omr_response_questions = set(omr_response.keys())
+ all_questions = set(self.questions_in_order)
+ missing_questions = sorted(all_questions.difference(omr_response_questions))
+ if len(missing_questions) > 0:
+ logger.critical(f"Missing OMR response for: {missing_questions}")
+ raise Exception(
+ f"Some questions are missing in the OMR response for the given answer key"
+ )
+
+ prefixed_omr_response_questions = set(
+ [k for k in omr_response.keys() if k.startswith("q")]
+ )
+ missing_prefixed_questions = sorted(
+ prefixed_omr_response_questions.difference(all_questions)
+ )
+ if len(missing_prefixed_questions) > 0:
+ logger.warning(
+ f"No answer given for potential questions in OMR response: {missing_prefixed_questions}"
+ )
+
+ def match_answer_for_question(self, current_score, question, marked_answer):
+ answer_matcher = self.question_to_answer_matcher[question]
+ question_verdict, delta = answer_matcher.get_verdict_marking(marked_answer)
+ self.conditionally_add_explanation(
+ answer_matcher,
+ delta,
+ marked_answer,
+ question_verdict,
+ question,
+ current_score,
+ )
+ return delta
+
+ def conditionally_print_explanation(self):
+ if self.should_explain_scoring:
+ console.print(self.explanation_table, justify="center")
+
+ # Explanation Table to CSV
+ def conditionally_save_explanation_csv(self, file_path, evaluation_output_dir):
+ if self.enable_evaluation_table_to_csv:
+ data = {col.header: col._cells for col in self.explanation_table.columns}
+
+ output_path = os.path.join(
+ evaluation_output_dir,
+ f"{file_path.stem}_evaluation.csv",
+ )
+
+ pd.DataFrame(data, dtype=str).to_csv(
+ output_path,
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ index=False,
+ )
+
+ def get_should_explain_scoring(self):
+ return self.should_explain_scoring
+
+ def get_exclude_files(self):
+ return self.exclude_files
+
+ @staticmethod
+ def parse_answer_column(answer_column):
+ if answer_column[0] == "[":
+ # multiple-correct-weighted or multiple-correct
+ parsed_answer = ast.literal_eval(answer_column)
+ elif "," in answer_column:
+ # multiple-correct
+ parsed_answer = answer_column.split(",")
+ else:
+ # single-correct
+ parsed_answer = answer_column
+ return parsed_answer
+
+ def parse_questions_in_order(self, questions_in_order):
+ return parse_fields("questions_in_order", questions_in_order)
+
+ def validate_answers(self, answers_in_order, tuning_config):
+ answer_matcher_map = self.question_to_answer_matcher
+ if tuning_config.outputs.filter_out_multimarked_files:
+ multi_marked_answer = False
+ for question, answer_item in zip(self.questions_in_order, answers_in_order):
+ answer_type = answer_matcher_map[question].answer_type
+ if answer_type == "standard":
+ if len(answer_item) > 1:
+ multi_marked_answer = True
+ if answer_type == "multiple-correct":
+ for single_answer in answer_item:
+ if len(single_answer) > 1:
+ multi_marked_answer = True
+ break
+ if answer_type == "multiple-correct-weighted":
+ for single_answer, _answer_score in answer_item:
+ if len(single_answer) > 1:
+ multi_marked_answer = True
+
+ if multi_marked_answer:
+ raise Exception(
+ f"Provided answer key contains multiple correct answer(s), but config.filter_out_multimarked_files is True. Scoring will get skipped."
+ )
+
+ def validate_questions(self, answers_in_order):
+ questions_in_order = self.questions_in_order
+ len_questions_in_order, len_answers_in_order = len(questions_in_order), len(
+ answers_in_order
+ )
+ if len_questions_in_order != len_answers_in_order:
+ logger.critical(
+ f"questions_in_order({len_questions_in_order}): {questions_in_order}\nanswers_in_order({len_answers_in_order}): {answers_in_order}"
+ )
+ raise Exception(
+ f"Unequal lengths for questions_in_order and answers_in_order ({len_questions_in_order} != {len_answers_in_order})"
+ )
+
+ def validate_marking_schemes(self):
+ section_marking_schemes = self.section_marking_schemes
+ section_questions = set()
+ for section_key, section_scheme in section_marking_schemes.items():
+ if section_key == DEFAULT_SECTION_KEY:
+ continue
+ current_set = set(section_scheme.questions)
+ if not section_questions.isdisjoint(current_set):
+ raise Exception(
+ f"Section '{section_key}' has overlapping question(s) with other sections"
+ )
+ section_questions = section_questions.union(current_set)
+
+ all_questions = set(self.questions_in_order)
+ missing_questions = sorted(section_questions.difference(all_questions))
+ if len(missing_questions) > 0:
+ logger.critical(f"Missing answer key for: {missing_questions}")
+ raise Exception(
+ f"Some questions are missing in the answer key for the given marking scheme"
+ )
+
+ def parse_answers_and_map_questions(self, answers_in_order):
+ question_to_answer_matcher = {}
+ for question, answer_item in zip(self.questions_in_order, answers_in_order):
+ section_marking_scheme = self.get_marking_scheme_for_question(question)
+ answer_matcher = AnswerMatcher(answer_item, section_marking_scheme)
+ question_to_answer_matcher[question] = answer_matcher
+ if (
+ answer_matcher.answer_type == "multiple-correct-weighted"
+ and section_marking_scheme.section_key != DEFAULT_SECTION_KEY
+ ):
+ logger.warning(
+ f"The custom scheme '{section_marking_scheme}' will not apply to question '{question}' as it will use the given answer weights f{answer_item}"
+ )
+ return question_to_answer_matcher
+
+ # Then unfolding lower abstraction levels
+ def reset_explanation_table(self):
+ self.explanation_table = None
+ self.prepare_explanation_table()
+
+ def prepare_explanation_table(self):
+ # TODO: provide a way to export this as csv/pdf
+ if not self.should_explain_scoring:
+ return
+ table = Table(title="Evaluation Explanation Table", show_lines=True)
+ table.add_column("Question")
+ table.add_column("Marked")
+ table.add_column("Answer(s)")
+ table.add_column("Verdict")
+ table.add_column("Delta")
+ table.add_column("Score")
+ # TODO: Add max and min score in explanation (row-wise and total)
+ if self.has_non_default_section:
+ table.add_column("Section")
+ self.explanation_table = table
+
+ def get_marking_scheme_for_question(self, question):
+ return self.question_to_scheme.get(question, self.default_marking_scheme)
+
+ def conditionally_add_explanation(
+ self,
+ answer_matcher,
+ delta,
+ marked_answer,
+ question_verdict,
+ question,
+ current_score,
+ ):
+ if self.should_explain_scoring:
+ next_score = current_score + delta
+ # Conditionally add cells
+ row = [
+ item
+ for item in [
+ question,
+ marked_answer,
+ str(answer_matcher),
+ str.title(question_verdict),
+ str(round(delta, 2)),
+ str(round(next_score, 2)),
+ (
+ answer_matcher.get_section_explanation()
+ if self.has_non_default_section
+ else None
+ ),
+ ]
+ if item is not None
+ ]
+ self.explanation_table.add_row(*row)
+
+
+def evaluate_concatenated_response(
+ concatenated_response, evaluation_config, file_path, evaluation_output_dir
+):
+ evaluation_config.prepare_and_validate_omr_response(concatenated_response)
+ current_score = 0.0
+ for question in evaluation_config.questions_in_order:
+ marked_answer = concatenated_response[question]
+ delta = evaluation_config.match_answer_for_question(
+ current_score, question, marked_answer
+ )
+ current_score += delta
+
+ evaluation_config.conditionally_print_explanation()
+ evaluation_config.conditionally_save_explanation_csv(file_path, evaluation_output_dir)
+
+ return current_score
diff --git a/OMRChecker/src/logger.py b/OMRChecker/src/logger.py
new file mode 100644
index 0000000000000000000000000000000000000000..d78988b8e8b6dc5d123c58fd0068c13bf9e67f42
--- /dev/null
+++ b/OMRChecker/src/logger.py
@@ -0,0 +1,68 @@
+import logging
+from typing import Union
+
+from rich.console import Console
+from rich.logging import RichHandler
+
+FORMAT = "%(message)s"
+
+# TODO: set logging level from config.json dynamically
+logging.basicConfig(
+ level=logging.INFO,
+ format="%(message)s",
+ datefmt="[%X]",
+ handlers=[RichHandler(rich_tracebacks=True)],
+)
+
+
+class Logger:
+ def __init__(
+ self,
+ name,
+ level: Union[int, str] = logging.NOTSET,
+ message_format="%(message)s",
+ date_format="[%X]",
+ ):
+ self.log = logging.getLogger(name)
+ self.log.setLevel(level)
+ self.log.__format__ = message_format
+ self.log.__date_format__ = date_format
+
+ def debug(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("debug", *msg, sep=sep)
+
+ def info(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("info", *msg, sep=sep)
+
+ def warning(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("warning", *msg, sep=sep)
+
+ def error(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("error", *msg, sep=sep)
+
+ def critical(self, *msg: object, sep=" ", end="\n") -> None:
+ return self.logutil("critical", *msg, sep=sep)
+
+ def stringify(func):
+ def inner(self, method_type: str, *msg: object, sep=" "):
+ nmsg = []
+ for v in msg:
+ if not isinstance(v, str):
+ v = str(v)
+ nmsg.append(v)
+ return func(self, method_type, *nmsg, sep=sep)
+
+ return inner
+
+ # set stack level to 3 so that the caller of this function is logged, not this function itself.
+ # stack-frame - self.log.debug - logutil - stringify - log method - caller
+ @stringify
+ def logutil(self, method_type: str, *msg: object, sep=" ") -> None:
+ func = getattr(self.log, method_type, None)
+ if not func:
+ raise AttributeError(f"Logger has no method {method_type}")
+ return func(sep.join(msg), stacklevel=4)
+
+
+logger = Logger(__name__)
+console = Console()
diff --git a/OMRChecker/src/processors/CropOnMarkers.py b/OMRChecker/src/processors/CropOnMarkers.py
new file mode 100644
index 0000000000000000000000000000000000000000..3801d88af313c83dd514ab33ab9bde1cf2afd16d
--- /dev/null
+++ b/OMRChecker/src/processors/CropOnMarkers.py
@@ -0,0 +1,233 @@
+import os
+
+import cv2
+import numpy as np
+
+from src.logger import logger
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class CropOnMarkers(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ config = self.tuning_config
+ marker_ops = self.options
+ self.threshold_circles = []
+ # img_utils = ImageUtils()
+
+ # options with defaults
+ self.marker_path = os.path.join(
+ self.relative_dir, marker_ops.get("relativePath", "omr_marker.jpg")
+ )
+ self.min_matching_threshold = marker_ops.get("min_matching_threshold", 0.3)
+ self.max_matching_variation = marker_ops.get("max_matching_variation", 0.41)
+ self.marker_rescale_range = tuple(
+ int(r) for r in marker_ops.get("marker_rescale_range", (35, 100))
+ )
+ self.marker_rescale_steps = int(marker_ops.get("marker_rescale_steps", 10))
+ self.apply_erode_subtract = marker_ops.get("apply_erode_subtract", True)
+ self.marker = self.load_marker(marker_ops, config)
+
+ def __str__(self):
+ return self.marker_path
+
+ def exclude_files(self):
+ return [self.marker_path]
+
+ def apply_filter(self, image, file_path):
+ config = self.tuning_config
+ image_instance_ops = self.image_instance_ops
+ image_eroded_sub = ImageUtils.normalize_util(
+ image
+ if self.apply_erode_subtract
+ else (image - cv2.erode(image, kernel=np.ones((5, 5)), iterations=5))
+ )
+ # Quads on warped image
+ quads = {}
+ h1, w1 = image_eroded_sub.shape[:2]
+ midh, midw = h1 // 3, w1 // 2
+ origins = [[0, 0], [midw, 0], [0, midh], [midw, midh]]
+ quads[0] = image_eroded_sub[0:midh, 0:midw]
+ quads[1] = image_eroded_sub[0:midh, midw:w1]
+ quads[2] = image_eroded_sub[midh:h1, 0:midw]
+ quads[3] = image_eroded_sub[midh:h1, midw:w1]
+
+ # Draw Quadlines
+ image_eroded_sub[:, midw : midw + 2] = 255
+ image_eroded_sub[midh : midh + 2, :] = 255
+
+ best_scale, all_max_t = self.getBestMatch(image_eroded_sub)
+ if best_scale is None:
+ if config.outputs.show_image_level >= 1:
+ InteractionUtils.show("Quads", image_eroded_sub, config=config)
+ return None
+
+ optimal_marker = ImageUtils.resize_util_h(
+ self.marker, u_height=int(self.marker.shape[0] * best_scale)
+ )
+ _h, w = optimal_marker.shape[:2]
+ centres = []
+ sum_t, max_t = 0, 0
+ quarter_match_log = "Matching Marker: "
+ for k in range(0, 4):
+ res = cv2.matchTemplate(quads[k], optimal_marker, cv2.TM_CCOEFF_NORMED)
+ max_t = res.max()
+ quarter_match_log += f"Quarter{str(k + 1)}: {str(round(max_t, 3))}\t"
+ if (
+ max_t < self.min_matching_threshold
+ or abs(all_max_t - max_t) >= self.max_matching_variation
+ ):
+ logger.error(
+ file_path,
+ "\nError: No circle found in Quad",
+ k + 1,
+ "\n\t min_matching_threshold",
+ self.min_matching_threshold,
+ "\t max_matching_variation",
+ self.max_matching_variation,
+ "\t max_t",
+ max_t,
+ "\t all_max_t",
+ all_max_t,
+ )
+ if config.outputs.show_image_level >= 1:
+ InteractionUtils.show(
+ f"No markers: {file_path}",
+ image_eroded_sub,
+ 0,
+ config=config,
+ )
+ InteractionUtils.show(
+ f"res_Q{str(k + 1)} ({str(max_t)})",
+ res,
+ 1,
+ config=config,
+ )
+ return None
+
+ pt = np.argwhere(res == max_t)[0]
+ pt = [pt[1], pt[0]]
+ pt[0] += origins[k][0]
+ pt[1] += origins[k][1]
+ # print(">>",pt)
+ image = cv2.rectangle(
+ image, tuple(pt), (pt[0] + w, pt[1] + _h), (150, 150, 150), 2
+ )
+ # display:
+ image_eroded_sub = cv2.rectangle(
+ image_eroded_sub,
+ tuple(pt),
+ (pt[0] + w, pt[1] + _h),
+ (50, 50, 50) if self.apply_erode_subtract else (155, 155, 155),
+ 4,
+ )
+ centres.append([pt[0] + w / 2, pt[1] + _h / 2])
+ sum_t += max_t
+
+ logger.info(quarter_match_log)
+ logger.info(f"Optimal Scale: {best_scale}")
+ # analysis data
+ self.threshold_circles.append(sum_t / 4)
+
+ image = ImageUtils.four_point_transform(image, np.array(centres))
+ # appendSaveImg(1,image_eroded_sub)
+ # appendSaveImg(1,image_norm)
+
+ image_instance_ops.append_save_img(2, image_eroded_sub)
+ # Debugging image -
+ # res = cv2.matchTemplate(image_eroded_sub,optimal_marker,cv2.TM_CCOEFF_NORMED)
+ # res[ : , midw:midw+2] = 255
+ # res[ midh:midh+2, : ] = 255
+ # show("Markers Matching",res)
+ if config.outputs.show_image_level >= 2 and config.outputs.show_image_level < 4:
+ image_eroded_sub = ImageUtils.resize_util_h(
+ image_eroded_sub, image.shape[0]
+ )
+ image_eroded_sub[:, -5:] = 0
+ h_stack = np.hstack((image_eroded_sub, image))
+ InteractionUtils.show(
+ f"Warped: {file_path}",
+ ImageUtils.resize_util(
+ h_stack, int(config.dimensions.display_width * 1.6)
+ ),
+ 0,
+ 0,
+ [0, 0],
+ config=config,
+ )
+ # iterations : Tuned to 2.
+ # image_eroded_sub = image_norm - cv2.erode(image_norm, kernel=np.ones((5,5)),iterations=2)
+ return image
+
+ def load_marker(self, marker_ops, config):
+ if not os.path.exists(self.marker_path):
+ logger.error(
+ "Marker not found at path provided in template:",
+ self.marker_path,
+ )
+ exit(31)
+
+ marker = cv2.imread(self.marker_path, cv2.IMREAD_GRAYSCALE)
+
+ if "sheetToMarkerWidthRatio" in marker_ops:
+ marker = ImageUtils.resize_util(
+ marker,
+ config.dimensions.processing_width
+ / int(marker_ops["sheetToMarkerWidthRatio"]),
+ )
+ marker = cv2.GaussianBlur(marker, (5, 5), 0)
+ marker = cv2.normalize(
+ marker, None, alpha=0, beta=255, norm_type=cv2.NORM_MINMAX
+ )
+
+ if self.apply_erode_subtract:
+ marker -= cv2.erode(marker, kernel=np.ones((5, 5)), iterations=5)
+
+ return marker
+
+ # Resizing the marker within scaleRange at rate of descent_per_step to
+ # find the best match.
+ def getBestMatch(self, image_eroded_sub):
+ config = self.tuning_config
+ descent_per_step = (
+ self.marker_rescale_range[1] - self.marker_rescale_range[0]
+ ) // self.marker_rescale_steps
+ _h, _w = self.marker.shape[:2]
+ res, best_scale = None, None
+ all_max_t = 0
+
+ for r0 in np.arange(
+ self.marker_rescale_range[1],
+ self.marker_rescale_range[0],
+ -1 * descent_per_step,
+ ): # reverse order
+ s = float(r0 * 1 / 100)
+ if s == 0.0:
+ continue
+ rescaled_marker = ImageUtils.resize_util_h(
+ self.marker, u_height=int(_h * s)
+ )
+ # res is the black image with white dots
+ res = cv2.matchTemplate(
+ image_eroded_sub, rescaled_marker, cv2.TM_CCOEFF_NORMED
+ )
+
+ max_t = res.max()
+ if all_max_t < max_t:
+ # print('Scale: '+str(s)+', Circle Match: '+str(round(max_t*100,2))+'%')
+ best_scale, all_max_t = s, max_t
+
+ if all_max_t < self.min_matching_threshold:
+ logger.warning(
+ "\tTemplate matching too low! Consider rechecking preProcessors applied before this."
+ )
+ if config.outputs.show_image_level >= 1:
+ InteractionUtils.show("res", res, 1, 0, config=config)
+
+ if best_scale is None:
+ logger.warning(
+ "No matchings for given scaleRange:", self.marker_rescale_range
+ )
+ return best_scale, all_max_t
diff --git a/OMRChecker/src/processors/CropPage.py b/OMRChecker/src/processors/CropPage.py
new file mode 100644
index 0000000000000000000000000000000000000000..49a6782597aaeae6ee00de28eedef5d40f1fc5a6
--- /dev/null
+++ b/OMRChecker/src/processors/CropPage.py
@@ -0,0 +1,112 @@
+"""
+https://www.pyimagesearch.com/2015/04/06/zero-parameter-automatic-canny-edge-detection-with-python-and-opencv/
+"""
+import cv2
+import numpy as np
+
+from src.logger import logger
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils
+
+MIN_PAGE_AREA = 80000
+
+
+def normalize(image):
+ return cv2.normalize(image, 0, 255, norm_type=cv2.NORM_MINMAX)
+
+
+def check_max_cosine(approx):
+ # assumes 4 pts present
+ max_cosine = 0
+ min_cosine = 1.5
+ for i in range(2, 5):
+ cosine = abs(angle(approx[i % 4], approx[i - 2], approx[i - 1]))
+ max_cosine = max(cosine, max_cosine)
+ min_cosine = min(cosine, min_cosine)
+
+ if max_cosine >= 0.35:
+ logger.warning("Quadrilateral is not a rectangle.")
+ return False
+ return True
+
+
+def validate_rect(approx):
+ return len(approx) == 4 and check_max_cosine(approx.reshape(4, 2))
+
+
+def angle(p_1, p_2, p_0):
+ dx1 = float(p_1[0] - p_0[0])
+ dy1 = float(p_1[1] - p_0[1])
+ dx2 = float(p_2[0] - p_0[0])
+ dy2 = float(p_2[1] - p_0[1])
+ return (dx1 * dx2 + dy1 * dy2) / np.sqrt(
+ (dx1 * dx1 + dy1 * dy1) * (dx2 * dx2 + dy2 * dy2) + 1e-10
+ )
+
+
+class CropPage(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ cropping_ops = self.options
+ self.morph_kernel = tuple(
+ int(x) for x in cropping_ops.get("morphKernel", [10, 10])
+ )
+
+ def apply_filter(self, image, file_path):
+ image = normalize(cv2.GaussianBlur(image, (3, 3), 0))
+
+ # Resize should be done with another preprocessor is needed
+ sheet = self.find_page(image, file_path)
+ if len(sheet) == 0:
+ logger.error(
+ f"\tError: Paper boundary not found for: '{file_path}'\nHave you accidentally included CropPage preprocessor?"
+ )
+ return None
+
+ logger.info(f"Found page corners: \t {sheet.tolist()}")
+
+ # Warp layer 1
+ image = ImageUtils.four_point_transform(image, sheet)
+
+ # Return preprocessed image
+ return image
+
+ def find_page(self, image, file_path):
+ config = self.tuning_config
+
+ image = normalize(image)
+
+ _ret, image = cv2.threshold(image, 200, 255, cv2.THRESH_TRUNC)
+ image = normalize(image)
+
+ kernel = cv2.getStructuringElement(cv2.MORPH_RECT, self.morph_kernel)
+
+ # Close the small holes, i.e. Complete the edges on canny image
+ closed = cv2.morphologyEx(image, cv2.MORPH_CLOSE, kernel)
+
+ edge = cv2.Canny(closed, 185, 55)
+
+ if config.outputs.show_image_level >= 5:
+ InteractionUtils.show("edge", edge, config=config)
+
+ # findContours returns outer boundaries in CW and inner ones, ACW.
+ cnts = ImageUtils.grab_contours(
+ cv2.findContours(edge, cv2.RETR_LIST, cv2.CHAIN_APPROX_SIMPLE)
+ )
+ # convexHull to resolve disordered curves due to noise
+ cnts = [cv2.convexHull(c) for c in cnts]
+ cnts = sorted(cnts, key=cv2.contourArea, reverse=True)[:5]
+ sheet = []
+ for c in cnts:
+ if cv2.contourArea(c) < MIN_PAGE_AREA:
+ continue
+ peri = cv2.arcLength(c, True)
+ approx = cv2.approxPolyDP(c, epsilon=0.025 * peri, closed=True)
+ if validate_rect(approx):
+ sheet = np.reshape(approx, (4, -1))
+ cv2.drawContours(image, [approx], -1, (0, 255, 0), 2)
+ cv2.drawContours(edge, [approx], -1, (255, 255, 255), 10)
+ break
+
+ return sheet
diff --git a/OMRChecker/src/processors/FeatureBasedAlignment.py b/OMRChecker/src/processors/FeatureBasedAlignment.py
new file mode 100644
index 0000000000000000000000000000000000000000..c7b0fb26cc0f3efd4eb27fa8b1e3461b91bced0a
--- /dev/null
+++ b/OMRChecker/src/processors/FeatureBasedAlignment.py
@@ -0,0 +1,94 @@
+"""
+Image based feature alignment
+Credits: https://www.learnopencv.com/image-alignment-feature-based-using-opencv-c-python/
+"""
+import cv2
+import numpy as np
+
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class FeatureBasedAlignment(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+ config = self.tuning_config
+
+ # process reference image
+ self.ref_path = self.relative_dir.joinpath(options["reference"])
+ ref_img = cv2.imread(str(self.ref_path), cv2.IMREAD_GRAYSCALE)
+ self.ref_img = ImageUtils.resize_util(
+ ref_img,
+ config.dimensions.processing_width,
+ config.dimensions.processing_height,
+ )
+ # get options with defaults
+ self.max_features = int(options.get("maxFeatures", 500))
+ self.good_match_percent = options.get("goodMatchPercent", 0.15)
+ self.transform_2_d = options.get("2d", False)
+ # Extract keypoints and description of source image
+ self.orb = cv2.ORB_create(self.max_features)
+ self.to_keypoints, self.to_descriptors = self.orb.detectAndCompute(
+ self.ref_img, None
+ )
+
+ def __str__(self):
+ return self.ref_path.name
+
+ def exclude_files(self):
+ return [self.ref_path]
+
+ def apply_filter(self, image, _file_path):
+ config = self.tuning_config
+ # Convert images to grayscale
+ # im1Gray = cv2.cvtColor(im1, cv2.COLOR_BGR2GRAY)
+ # im2Gray = cv2.cvtColor(im2, cv2.COLOR_BGR2GRAY)
+
+ image = cv2.normalize(image, 0, 255, norm_type=cv2.NORM_MINMAX)
+
+ # Detect ORB features and compute descriptors.
+ from_keypoints, from_descriptors = self.orb.detectAndCompute(image, None)
+
+ # Match features.
+ matcher = cv2.DescriptorMatcher_create(
+ cv2.DESCRIPTOR_MATCHER_BRUTEFORCE_HAMMING
+ )
+
+ # create BFMatcher object (alternate matcher)
+ # matcher = cv2.BFMatcher(cv2.NORM_HAMMING, crossCheck=True)
+
+ matches = np.array(matcher.match(from_descriptors, self.to_descriptors, None))
+
+ # Sort matches by score
+ matches = sorted(matches, key=lambda x: x.distance, reverse=False)
+
+ # Remove not so good matches
+ num_good_matches = int(len(matches) * self.good_match_percent)
+ matches = matches[:num_good_matches]
+
+ # Draw top matches
+ if config.outputs.show_image_level > 2:
+ im_matches = cv2.drawMatches(
+ image, from_keypoints, self.ref_img, self.to_keypoints, matches, None
+ )
+ InteractionUtils.show("Aligning", im_matches, resize=True, config=config)
+
+ # Extract location of good matches
+ points1 = np.zeros((len(matches), 2), dtype=np.float32)
+ points2 = np.zeros((len(matches), 2), dtype=np.float32)
+
+ for i, match in enumerate(matches):
+ points1[i, :] = from_keypoints[match.queryIdx].pt
+ points2[i, :] = self.to_keypoints[match.trainIdx].pt
+
+ # Find homography
+ height, width = self.ref_img.shape
+ if self.transform_2_d:
+ m, _inliers = cv2.estimateAffine2D(points1, points2)
+ return cv2.warpAffine(image, m, (width, height))
+
+ # Use homography
+ h, _mask = cv2.findHomography(points1, points2, cv2.RANSAC)
+ return cv2.warpPerspective(image, h, (width, height))
diff --git a/OMRChecker/src/processors/builtins.py b/OMRChecker/src/processors/builtins.py
new file mode 100644
index 0000000000000000000000000000000000000000..3e3f54683ee104c67bdd422c0fa3cdacfcc4fd8d
--- /dev/null
+++ b/OMRChecker/src/processors/builtins.py
@@ -0,0 +1,54 @@
+import cv2
+import numpy as np
+
+from src.processors.interfaces.ImagePreprocessor import ImagePreprocessor
+
+
+class Levels(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+
+ def output_level(value, low, high, gamma):
+ if value <= low:
+ return 0
+ if value >= high:
+ return 255
+ inv_gamma = 1.0 / gamma
+ return (((value - low) / (high - low)) ** inv_gamma) * 255
+
+ self.gamma = np.array(
+ [
+ output_level(
+ i,
+ int(255 * options.get("low", 0)),
+ int(255 * options.get("high", 1)),
+ options.get("gamma", 1.0),
+ )
+ for i in np.arange(0, 256)
+ ]
+ ).astype("uint8")
+
+ def apply_filter(self, image, _file_path):
+ return cv2.LUT(image, self.gamma)
+
+
+class MedianBlur(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+ self.kSize = int(options.get("kSize", 5))
+
+ def apply_filter(self, image, _file_path):
+ return cv2.medianBlur(image, self.kSize)
+
+
+class GaussianBlur(ImagePreprocessor):
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+ options = self.options
+ self.kSize = tuple(int(x) for x in options.get("kSize", (3, 3)))
+ self.sigmaX = int(options.get("sigmaX", 0))
+
+ def apply_filter(self, image, _file_path):
+ return cv2.GaussianBlur(image, self.kSize, self.sigmaX)
diff --git a/OMRChecker/src/processors/interfaces/ImagePreprocessor.py b/OMRChecker/src/processors/interfaces/ImagePreprocessor.py
new file mode 100644
index 0000000000000000000000000000000000000000..619a8ad711f48affc0ebaad0c6a2bf34ee2f184d
--- /dev/null
+++ b/OMRChecker/src/processors/interfaces/ImagePreprocessor.py
@@ -0,0 +1,18 @@
+# Use all imports relative to root directory
+from src.processors.manager import Processor
+
+
+class ImagePreprocessor(Processor):
+ """Base class for an extension that applies some preprocessing to the input image"""
+
+ def __init__(self, *args, **kwargs):
+ super().__init__(*args, **kwargs)
+
+ def apply_filter(self, image, filename):
+ """Apply filter to the image and returns modified image"""
+ raise NotImplementedError
+
+ @staticmethod
+ def exclude_files():
+ """Returns a list of file paths that should be excluded from processing"""
+ return []
diff --git a/OMRChecker/src/processors/manager.py b/OMRChecker/src/processors/manager.py
new file mode 100644
index 0000000000000000000000000000000000000000..18f4346c736513fea4d141d2489e37bf920003d4
--- /dev/null
+++ b/OMRChecker/src/processors/manager.py
@@ -0,0 +1,80 @@
+"""
+Processor/Extension framework
+Adapated from https://github.com/gdiepen/python_processor_example
+"""
+import inspect
+import pkgutil
+
+from src.logger import logger
+
+
+class Processor:
+ """Base class that each processor must inherit from."""
+
+ def __init__(
+ self,
+ options=None,
+ relative_dir=None,
+ image_instance_ops=None,
+ ):
+ self.options = options
+ self.relative_dir = relative_dir
+ self.image_instance_ops = image_instance_ops
+ self.tuning_config = image_instance_ops.tuning_config
+ self.description = "UNKNOWN"
+
+
+class ProcessorManager:
+ """Upon creation, this class will read the processors package for modules
+ that contain a class definition that is inheriting from the Processor class
+ """
+
+ def __init__(self, processors_dir="src.processors"):
+ """Constructor that initiates the reading of all available processors
+ when an instance of the ProcessorCollection object is created
+ """
+ self.processors_dir = processors_dir
+ self.reload_processors()
+
+ @staticmethod
+ def get_name_filter(processor_name):
+ def filter_function(member):
+ return inspect.isclass(member) and member.__module__ == processor_name
+
+ return filter_function
+
+ def reload_processors(self):
+ """Reset the list of all processors and initiate the walk over the main
+ provided processor package to load all available processors
+ """
+ self.processors = {}
+ self.seen_paths = []
+
+ logger.info(f'Loading processors from "{self.processors_dir}"...')
+ self.walk_package(self.processors_dir)
+
+ def walk_package(self, package):
+ """walk the supplied package to retrieve all processors"""
+ imported_package = __import__(package, fromlist=["blah"])
+ loaded_packages = []
+ for _, processor_name, ispkg in pkgutil.walk_packages(
+ imported_package.__path__, imported_package.__name__ + "."
+ ):
+ if not ispkg and processor_name != __name__:
+ processor_module = __import__(processor_name, fromlist=["blah"])
+ # https://stackoverflow.com/a/46206754/6242649
+ clsmembers = inspect.getmembers(
+ processor_module,
+ ProcessorManager.get_name_filter(processor_name),
+ )
+ for _, c in clsmembers:
+ # Only add classes that are a sub class of Processor, but NOT Processor itself
+ if issubclass(c, Processor) & (c is not Processor):
+ self.processors[c.__name__] = c
+ loaded_packages.append(c.__name__)
+
+ logger.info(f"Loaded processors: {loaded_packages}")
+
+
+# Singleton export
+PROCESSOR_MANAGER = ProcessorManager()
diff --git a/OMRChecker/src/schemas/__init__.py b/OMRChecker/src/schemas/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..e325daf6b22ed9b0e1c37a4d251cc58e040b431d
--- /dev/null
+++ b/OMRChecker/src/schemas/__init__.py
@@ -0,0 +1,18 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+from jsonschema import Draft202012Validator
+
+from src.schemas.config_schema import CONFIG_SCHEMA
+from src.schemas.evaluation_schema import EVALUATION_SCHEMA
+from src.schemas.template_schema import TEMPLATE_SCHEMA
+
+SCHEMA_JSONS = {
+ "config": CONFIG_SCHEMA,
+ "evaluation": EVALUATION_SCHEMA,
+ "template": TEMPLATE_SCHEMA,
+}
+
+SCHEMA_VALIDATORS = {
+ "config": Draft202012Validator(CONFIG_SCHEMA),
+ "evaluation": Draft202012Validator(EVALUATION_SCHEMA),
+ "template": Draft202012Validator(TEMPLATE_SCHEMA),
+}
diff --git a/OMRChecker/src/schemas/config_schema.py b/OMRChecker/src/schemas/config_schema.py
new file mode 100644
index 0000000000000000000000000000000000000000..ffb41f45df1373c5fd0ab3bbe8132e74ef77fc73
--- /dev/null
+++ b/OMRChecker/src/schemas/config_schema.py
@@ -0,0 +1,57 @@
+CONFIG_SCHEMA = {
+ "$schema": "https://json-schema.org/draft/2020-12/schema",
+ "$id": "https://github.com/Udayraj123/OMRChecker/tree/master/src/schemas/config-schema.json",
+ "title": "Config Schema",
+ "description": "OMRChecker config schema for custom tuning",
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "dimensions": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "display_height": {"type": "integer"},
+ "display_width": {"type": "integer"},
+ "processing_height": {"type": "integer"},
+ "processing_width": {"type": "integer"},
+ },
+ },
+ "threshold_params": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "GAMMA_LOW": {"type": "number", "minimum": 0, "maximum": 1},
+ "MIN_GAP": {"type": "integer", "minimum": 10, "maximum": 100},
+ "MIN_JUMP": {"type": "integer", "minimum": 10, "maximum": 100},
+ "CONFIDENT_SURPLUS": {"type": "integer", "minimum": 0, "maximum": 20},
+ "JUMP_DELTA": {"type": "integer", "minimum": 10, "maximum": 100},
+ "PAGE_TYPE_FOR_THRESHOLD": {
+ "enum": ["white", "black"],
+ "type": "string",
+ },
+ },
+ },
+ "alignment_params": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "auto_align": {"type": "boolean"},
+ "match_col": {"type": "integer", "minimum": 0, "maximum": 10},
+ "max_steps": {"type": "integer", "minimum": 1, "maximum": 100},
+ "stride": {"type": "integer", "minimum": 1, "maximum": 10},
+ "thickness": {"type": "integer", "minimum": 1, "maximum": 10},
+ },
+ },
+ "outputs": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "show_image_level": {"type": "integer", "minimum": 0, "maximum": 6},
+ "save_image_level": {"type": "integer", "minimum": 0, "maximum": 6},
+ "save_detections": {"type": "boolean"},
+ # This option moves multimarked files into a separate folder for manual checking, skipping evaluation
+ "filter_out_multimarked_files": {"type": "boolean"},
+ },
+ },
+ },
+}
diff --git a/OMRChecker/src/schemas/constants.py b/OMRChecker/src/schemas/constants.py
new file mode 100644
index 0000000000000000000000000000000000000000..34b9b24d3fac2f60ecbba2d20d8f039d729db11c
--- /dev/null
+++ b/OMRChecker/src/schemas/constants.py
@@ -0,0 +1,17 @@
+DEFAULT_SECTION_KEY = "DEFAULT"
+
+BONUS_SECTION_PREFIX = "BONUS"
+
+MARKING_VERDICT_TYPES = ["correct", "incorrect", "unmarked"]
+
+ARRAY_OF_STRINGS = {
+ "type": "array",
+ "items": {"type": "string"},
+}
+
+FIELD_STRING_TYPE = {
+ "type": "string",
+ "pattern": "^([^\\.]+|[^\\.\\d]+\\d+\\.{2,3}\\d+)$",
+}
+
+FIELD_STRING_REGEX_GROUPS = r"([^\.\d]+)(\d+)\.{2,3}(\d+)"
diff --git a/OMRChecker/src/schemas/evaluation_schema.py b/OMRChecker/src/schemas/evaluation_schema.py
new file mode 100644
index 0000000000000000000000000000000000000000..c51af58b849eaa3202815ca1c9358ec34839975b
--- /dev/null
+++ b/OMRChecker/src/schemas/evaluation_schema.py
@@ -0,0 +1,151 @@
+from src.schemas.constants import (
+ ARRAY_OF_STRINGS,
+ DEFAULT_SECTION_KEY,
+ FIELD_STRING_TYPE,
+)
+
+marking_score_regex = "-?(\\d+)(/(\\d+))?"
+
+marking_score = {
+ "oneOf": [
+ {"type": "string", "pattern": marking_score_regex},
+ {"type": "number"},
+ ]
+}
+
+marking_object_properties = {
+ "additionalProperties": False,
+ "required": ["correct", "incorrect", "unmarked"],
+ "type": "object",
+ "properties": {
+ # TODO: can support streak marking if we allow array of marking_scores here
+ "correct": marking_score,
+ "incorrect": marking_score,
+ "unmarked": marking_score,
+ },
+}
+
+EVALUATION_SCHEMA = {
+ "$schema": "https://json-schema.org/draft/2020-12/schema",
+ "$id": "https://github.com/Udayraj123/OMRChecker/tree/master/src/schemas/evaluation-schema.json",
+ "title": "Evaluation Schema",
+ "description": "OMRChecker evaluation schema i.e. the marking scheme",
+ "type": "object",
+ "additionalProperties": True,
+ "required": ["source_type", "options", "marking_schemes"],
+ "properties": {
+ "additionalProperties": False,
+ "source_type": {"type": "string", "enum": ["csv", "custom"]},
+ "options": {"type": "object"},
+ "marking_schemes": {
+ "type": "object",
+ "required": [DEFAULT_SECTION_KEY],
+ "patternProperties": {
+ f"^{DEFAULT_SECTION_KEY}$": marking_object_properties,
+ f"^(?!{DEFAULT_SECTION_KEY}$).*": {
+ "additionalProperties": False,
+ "required": ["marking", "questions"],
+ "type": "object",
+ "properties": {
+ "questions": {
+ "oneOf": [
+ FIELD_STRING_TYPE,
+ {
+ "type": "array",
+ "items": FIELD_STRING_TYPE,
+ },
+ ]
+ },
+ "marking": marking_object_properties,
+ },
+ },
+ },
+ },
+ },
+ "allOf": [
+ {
+ "if": {"properties": {"source_type": {"const": "csv"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "additionalProperties": False,
+ "required": ["answer_key_csv_path"],
+ "dependentRequired": {
+ "answer_key_image_path": [
+ "answer_key_csv_path",
+ "questions_in_order",
+ ]
+ },
+ "type": "object",
+ "properties": {
+ "should_explain_scoring": {"type": "boolean"},
+ "answer_key_csv_path": {"type": "string"},
+ "answer_key_image_path": {"type": "string"},
+ "questions_in_order": ARRAY_OF_STRINGS,
+ },
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"source_type": {"const": "custom"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "additionalProperties": False,
+ "required": ["answers_in_order", "questions_in_order"],
+ "type": "object",
+ "properties": {
+ "should_explain_scoring": {"type": "boolean"},
+ "answers_in_order": {
+ "oneOf": [
+ {
+ "type": "array",
+ "items": {
+ "oneOf": [
+ # "standard": single correct, multi-marked single-correct
+ # Example: "q1" --> '67'
+ {"type": "string"},
+ # "multiple-correct": multiple-correct (for ambiguous/bonus questions)
+ # Example: "q1" --> [ 'A', 'B' ]
+ {
+ "type": "array",
+ "items": {"type": "string"},
+ "minItems": 2,
+ },
+ # "multiple-correct-weighted": array of answer-wise weights (marking scheme not applicable)
+ # Example 1: "q1" --> [['A', 1], ['B', 2], ['C', 3]] or
+ # Example 2: "q2" --> [['A', 1], ['B', 1], ['AB', 2]]
+ {
+ "type": "array",
+ "items": {
+ "type": "array",
+ "items": False,
+ "minItems": 2,
+ "maxItems": 2,
+ "prefixItems": [
+ {"type": "string"},
+ marking_score,
+ ],
+ },
+ },
+ # Multiple-correct with custom marking scheme
+ # ["A", ["1", "2", "3"]],
+ # [["A", "B", "AB"], ["1", "2", "3"]]
+ ],
+ },
+ },
+ ]
+ },
+ "questions_in_order": ARRAY_OF_STRINGS,
+ "enable_evaluation_table_to_csv": {
+ "type": "boolean",
+ "default": False,
+ },
+ },
+ }
+ }
+ },
+ },
+ ],
+}
diff --git a/OMRChecker/src/schemas/template_schema.py b/OMRChecker/src/schemas/template_schema.py
new file mode 100644
index 0000000000000000000000000000000000000000..12b7ffdd8577054c03db25f12815d91813026c40
--- /dev/null
+++ b/OMRChecker/src/schemas/template_schema.py
@@ -0,0 +1,226 @@
+from src.constants import FIELD_TYPES
+from src.schemas.constants import ARRAY_OF_STRINGS, FIELD_STRING_TYPE
+
+positive_number = {"type": "number", "minimum": 0}
+positive_integer = {"type": "integer", "minimum": 0}
+two_positive_integers = {
+ "type": "array",
+ "prefixItems": [
+ positive_integer,
+ positive_integer,
+ ],
+ "maxItems": 2,
+ "minItems": 2,
+}
+two_positive_numbers = {
+ "type": "array",
+ "prefixItems": [
+ positive_number,
+ positive_number,
+ ],
+ "maxItems": 2,
+ "minItems": 2,
+}
+zero_to_one_number = {
+ "type": "number",
+ "minimum": 0,
+ "maximum": 1,
+}
+
+TEMPLATE_SCHEMA = {
+ "$schema": "https://json-schema.org/draft/2020-12/schema",
+ "$id": "https://github.com/Udayraj123/OMRChecker/tree/master/src/schemas/template-schema.json",
+ "title": "Template Validation Schema",
+ "description": "OMRChecker input template schema",
+ "type": "object",
+ "required": [
+ "bubbleDimensions",
+ "pageDimensions",
+ "preProcessors",
+ "fieldBlocks",
+ ],
+ "additionalProperties": False,
+ "properties": {
+ "bubbleDimensions": {
+ **two_positive_integers,
+ "description": "The dimensions of the overlay bubble area: [width, height]",
+ },
+ "customLabels": {
+ "description": "The customLabels contain fields that need to be joined together before generating the results sheet",
+ "type": "object",
+ "patternProperties": {
+ "^.*$": {"type": "array", "items": FIELD_STRING_TYPE}
+ },
+ },
+ "outputColumns": {
+ "type": "array",
+ "items": FIELD_STRING_TYPE,
+ "description": "The ordered list of columns to be contained in the output csv(default order: alphabetical)",
+ },
+ "pageDimensions": {
+ **two_positive_integers,
+ "description": "The dimensions(width, height) to which the page will be resized to before applying template",
+ },
+ "preProcessors": {
+ "description": "Custom configuration values to use in the template's directory",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "name": {
+ "type": "string",
+ "enum": [
+ "CropOnMarkers",
+ "CropPage",
+ "FeatureBasedAlignment",
+ "GaussianBlur",
+ "Levels",
+ "MedianBlur",
+ ],
+ },
+ },
+ "required": ["name", "options"],
+ "allOf": [
+ {
+ "if": {"properties": {"name": {"const": "CropOnMarkers"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "apply_erode_subtract": {"type": "boolean"},
+ "marker_rescale_range": two_positive_numbers,
+ "marker_rescale_steps": {"type": "number"},
+ "max_matching_variation": {"type": "number"},
+ "min_matching_threshold": {"type": "number"},
+ "relativePath": {"type": "string"},
+ "sheetToMarkerWidthRatio": {"type": "number"},
+ },
+ "required": ["relativePath"],
+ }
+ }
+ },
+ },
+ {
+ "if": {
+ "properties": {"name": {"const": "FeatureBasedAlignment"}}
+ },
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "2d": {"type": "boolean"},
+ "goodMatchPercent": {"type": "number"},
+ "maxFeatures": {"type": "integer"},
+ "reference": {"type": "string"},
+ },
+ "required": ["reference"],
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "Levels"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "gamma": zero_to_one_number,
+ "high": zero_to_one_number,
+ "low": zero_to_one_number,
+ },
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "MedianBlur"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {"kSize": {"type": "integer"}},
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "GaussianBlur"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "kSize": two_positive_integers,
+ "sigmaX": {"type": "number"},
+ },
+ }
+ }
+ },
+ },
+ {
+ "if": {"properties": {"name": {"const": "CropPage"}}},
+ "then": {
+ "properties": {
+ "options": {
+ "type": "object",
+ "additionalProperties": False,
+ "properties": {
+ "morphKernel": two_positive_integers
+ },
+ }
+ }
+ },
+ },
+ ],
+ },
+ },
+ "fieldBlocks": {
+ "description": "The fieldBlocks denote small groups of adjacent fields",
+ "type": "object",
+ "patternProperties": {
+ "^.*$": {
+ "type": "object",
+ "required": [
+ "origin",
+ "bubblesGap",
+ "labelsGap",
+ "fieldLabels",
+ ],
+ "oneOf": [
+ {"required": ["fieldType"]},
+ {"required": ["bubbleValues", "direction"]},
+ ],
+ "properties": {
+ "bubbleDimensions": two_positive_numbers,
+ "bubblesGap": positive_number,
+ "bubbleValues": ARRAY_OF_STRINGS,
+ "direction": {
+ "type": "string",
+ "enum": ["horizontal", "vertical"],
+ },
+ "emptyValue": {"type": "string"},
+ "fieldLabels": {"type": "array", "items": FIELD_STRING_TYPE},
+ "labelsGap": positive_number,
+ "origin": two_positive_integers,
+ "fieldType": {
+ "type": "string",
+ "enum": list(FIELD_TYPES.keys()),
+ },
+ },
+ }
+ },
+ },
+ "emptyValue": {
+ "description": "The value to be used in case of empty bubble detected at global level.",
+ "type": "string",
+ },
+ },
+}
diff --git a/OMRChecker/src/template.py b/OMRChecker/src/template.py
new file mode 100644
index 0000000000000000000000000000000000000000..1ab6c9173bb9bf237d8902eaef004835ed8a1446
--- /dev/null
+++ b/OMRChecker/src/template.py
@@ -0,0 +1,327 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+from src.constants import FIELD_TYPES
+from src.core import ImageInstanceOps
+from src.logger import logger
+from src.processors.manager import PROCESSOR_MANAGER
+from src.utils.parsing import (
+ custom_sort_output_columns,
+ open_template_with_defaults,
+ parse_fields,
+)
+
+
+class Template:
+ def __init__(self, template_path, tuning_config):
+ self.path = template_path
+ self.image_instance_ops = ImageInstanceOps(tuning_config)
+
+ json_object = open_template_with_defaults(template_path)
+ (
+ custom_labels_object,
+ field_blocks_object,
+ output_columns_array,
+ pre_processors_object,
+ self.bubble_dimensions,
+ self.global_empty_val,
+ self.options,
+ self.page_dimensions,
+ ) = map(
+ json_object.get,
+ [
+ "customLabels",
+ "fieldBlocks",
+ "outputColumns",
+ "preProcessors",
+ "bubbleDimensions",
+ "emptyValue",
+ "options",
+ "pageDimensions",
+ ],
+ )
+
+ self.parse_output_columns(output_columns_array)
+ self.setup_pre_processors(pre_processors_object, template_path.parent)
+ self.setup_field_blocks(field_blocks_object)
+ self.parse_custom_labels(custom_labels_object)
+
+ non_custom_columns, all_custom_columns = (
+ list(self.non_custom_labels),
+ list(custom_labels_object.keys()),
+ )
+
+ if len(self.output_columns) == 0:
+ self.fill_output_columns(non_custom_columns, all_custom_columns)
+
+ self.validate_template_columns(non_custom_columns, all_custom_columns)
+
+ def parse_output_columns(self, output_columns_array):
+ self.output_columns = parse_fields(f"Output Columns", output_columns_array)
+
+ def setup_pre_processors(self, pre_processors_object, relative_dir):
+ # load image pre_processors
+ self.pre_processors = []
+ for pre_processor in pre_processors_object:
+ ProcessorClass = PROCESSOR_MANAGER.processors[pre_processor["name"]]
+ pre_processor_instance = ProcessorClass(
+ options=pre_processor["options"],
+ relative_dir=relative_dir,
+ image_instance_ops=self.image_instance_ops,
+ )
+ self.pre_processors.append(pre_processor_instance)
+
+ def setup_field_blocks(self, field_blocks_object):
+ # Add field_blocks
+ self.field_blocks = []
+ self.all_parsed_labels = set()
+ for block_name, field_block_object in field_blocks_object.items():
+ self.parse_and_add_field_block(block_name, field_block_object)
+
+ def parse_custom_labels(self, custom_labels_object):
+ all_parsed_custom_labels = set()
+ self.custom_labels = {}
+ for custom_label, label_strings in custom_labels_object.items():
+ parsed_labels = parse_fields(f"Custom Label: {custom_label}", label_strings)
+ parsed_labels_set = set(parsed_labels)
+ self.custom_labels[custom_label] = parsed_labels
+
+ missing_custom_labels = sorted(
+ parsed_labels_set.difference(self.all_parsed_labels)
+ )
+ if len(missing_custom_labels) > 0:
+ logger.critical(
+ f"For '{custom_label}', Missing labels - {missing_custom_labels}"
+ )
+ raise Exception(
+ f"Missing field block label(s) in the given template for {missing_custom_labels} from '{custom_label}'"
+ )
+
+ if not all_parsed_custom_labels.isdisjoint(parsed_labels_set):
+ # Note: this can be made a warning, but it's a choice
+ logger.critical(
+ f"field strings overlap for labels: {label_strings} and existing custom labels: {all_parsed_custom_labels}"
+ )
+ raise Exception(
+ f"The field strings for custom label '{custom_label}' overlap with other existing custom labels"
+ )
+
+ all_parsed_custom_labels.update(parsed_labels)
+
+ self.non_custom_labels = self.all_parsed_labels.difference(
+ all_parsed_custom_labels
+ )
+
+ def fill_output_columns(self, non_custom_columns, all_custom_columns):
+ all_template_columns = non_custom_columns + all_custom_columns
+ # Typical case: sort alpha-numerical (natural sort)
+ self.output_columns = sorted(
+ all_template_columns, key=custom_sort_output_columns
+ )
+
+ def validate_template_columns(self, non_custom_columns, all_custom_columns):
+ output_columns_set = set(self.output_columns)
+ all_custom_columns_set = set(all_custom_columns)
+
+ missing_output_columns = sorted(
+ output_columns_set.difference(all_custom_columns_set).difference(
+ self.all_parsed_labels
+ )
+ )
+ if len(missing_output_columns) > 0:
+ logger.critical(f"Missing output columns: {missing_output_columns}")
+ raise Exception(
+ f"Some columns are missing in the field blocks for the given output columns"
+ )
+
+ all_template_columns_set = set(non_custom_columns + all_custom_columns)
+ missing_label_columns = sorted(
+ all_template_columns_set.difference(output_columns_set)
+ )
+ if len(missing_label_columns) > 0:
+ logger.warning(
+ f"Some label columns are not covered in the given output columns: {missing_label_columns}"
+ )
+
+ def parse_and_add_field_block(self, block_name, field_block_object):
+ field_block_object = self.pre_fill_field_block(field_block_object)
+ block_instance = FieldBlock(block_name, field_block_object)
+ self.field_blocks.append(block_instance)
+ self.validate_parsed_labels(field_block_object["fieldLabels"], block_instance)
+
+ def pre_fill_field_block(self, field_block_object):
+ if "fieldType" in field_block_object:
+ field_block_object = {
+ **field_block_object,
+ **FIELD_TYPES[field_block_object["fieldType"]],
+ }
+ else:
+ field_block_object = {**field_block_object, "fieldType": "__CUSTOM__"}
+
+ return {
+ "direction": "vertical",
+ "emptyValue": self.global_empty_val,
+ "bubbleDimensions": self.bubble_dimensions,
+ **field_block_object,
+ }
+
+ def validate_parsed_labels(self, field_labels, block_instance):
+ parsed_field_labels, block_name = (
+ block_instance.parsed_field_labels,
+ block_instance.name,
+ )
+ field_labels_set = set(parsed_field_labels)
+ if not self.all_parsed_labels.isdisjoint(field_labels_set):
+ # Note: in case of two fields pointing to same column, use a custom column instead of same field labels.
+ logger.critical(
+ f"An overlap found between field string: {field_labels} in block '{block_name}' and existing labels: {self.all_parsed_labels}"
+ )
+ raise Exception(
+ f"The field strings for field block {block_name} overlap with other existing fields"
+ )
+ self.all_parsed_labels.update(field_labels_set)
+
+ page_width, page_height = self.page_dimensions
+ block_width, block_height = block_instance.dimensions
+ [block_start_x, block_start_y] = block_instance.origin
+
+ block_end_x, block_end_y = (
+ block_start_x + block_width,
+ block_start_y + block_height,
+ )
+
+ if (
+ block_end_x >= page_width
+ or block_end_y >= page_height
+ or block_start_x < 0
+ or block_start_y < 0
+ ):
+ raise Exception(
+ f"Overflowing field block '{block_name}' with origin {block_instance.origin} and dimensions {block_instance.dimensions} in template with dimensions {self.page_dimensions}"
+ )
+
+ def __str__(self):
+ return str(self.path)
+
+
+class FieldBlock:
+ def __init__(self, block_name, field_block_object):
+ self.name = block_name
+ self.shift = 0
+ self.setup_field_block(field_block_object)
+
+ def setup_field_block(self, field_block_object):
+ # case mapping
+ (
+ bubble_dimensions,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ field_labels,
+ field_type,
+ labels_gap,
+ origin,
+ self.empty_val,
+ ) = map(
+ field_block_object.get,
+ [
+ "bubbleDimensions",
+ "bubbleValues",
+ "bubblesGap",
+ "direction",
+ "fieldLabels",
+ "fieldType",
+ "labelsGap",
+ "origin",
+ "emptyValue",
+ ],
+ )
+ self.parsed_field_labels = parse_fields(
+ f"Field Block Labels: {self.name}", field_labels
+ )
+ self.origin = origin
+ self.bubble_dimensions = bubble_dimensions
+ self.calculate_block_dimensions(
+ bubble_dimensions,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ labels_gap,
+ )
+ self.generate_bubble_grid(
+ bubble_values,
+ bubbles_gap,
+ direction,
+ field_type,
+ labels_gap,
+ )
+
+ def calculate_block_dimensions(
+ self,
+ bubble_dimensions,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ labels_gap,
+ ):
+ _h, _v = (1, 0) if (direction == "vertical") else (0, 1)
+
+ values_dimension = int(
+ bubbles_gap * (len(bubble_values) - 1) + bubble_dimensions[_h]
+ )
+ fields_dimension = int(
+ labels_gap * (len(self.parsed_field_labels) - 1) + bubble_dimensions[_v]
+ )
+ self.dimensions = (
+ [fields_dimension, values_dimension]
+ if (direction == "vertical")
+ else [values_dimension, fields_dimension]
+ )
+
+ def generate_bubble_grid(
+ self,
+ bubble_values,
+ bubbles_gap,
+ direction,
+ field_type,
+ labels_gap,
+ ):
+ _h, _v = (1, 0) if (direction == "vertical") else (0, 1)
+ self.traverse_bubbles = []
+ # Generate the bubble grid
+ lead_point = [float(self.origin[0]), float(self.origin[1])]
+ for field_label in self.parsed_field_labels:
+ bubble_point = lead_point.copy()
+ field_bubbles = []
+ for bubble_value in bubble_values:
+ field_bubbles.append(
+ Bubble(bubble_point.copy(), field_label, field_type, bubble_value)
+ )
+ bubble_point[_h] += bubbles_gap
+ self.traverse_bubbles.append(field_bubbles)
+ lead_point[_v] += labels_gap
+
+
+class Bubble:
+ """
+ Container for a Point Box on the OMR
+
+ field_label is the point's property- field to which this point belongs to
+ It can be used as a roll number column as well. (eg roll1)
+ It can also correspond to a single digit of integer type Q (eg q5d1)
+ """
+
+ def __init__(self, pt, field_label, field_type, field_value):
+ self.x = round(pt[0])
+ self.y = round(pt[1])
+ self.field_label = field_label
+ self.field_type = field_type
+ self.field_value = field_value
+
+ def __str__(self):
+ return str([self.x, self.y])
diff --git a/OMRChecker/src/tests/__init__.py b/OMRChecker/src/tests/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..28a0acd8198648687d60beb6d8dded390f95a29a
--- /dev/null
+++ b/OMRChecker/src/tests/__init__.py
@@ -0,0 +1 @@
+# https://stackoverflow.com/a/50169991/6242649
diff --git a/OMRChecker/src/tests/__snapshots__/test_all_samples.ambr b/OMRChecker/src/tests/__snapshots__/test_all_samples.ambr
new file mode 100644
index 0000000000000000000000000000000000000000..45df1372201a0679ded211fbc424716db37f7632
--- /dev/null
+++ b/OMRChecker/src/tests/__snapshots__/test_all_samples.ambr
@@ -0,0 +1,302 @@
+# serializer version: 1
+# name: test_run_answer_key_using_csv
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "adrian_omr.png","samples/answer-key/using-csv/adrian_omr.png","outputs/answer-key/using-csv/CheckedOMRs/adrian_omr.png","5.0","C","E","A","B","B"
+
+ ''',
+ })
+# ---
+# name: test_run_answer_key_weighted_answers
+ dict({
+ 'images/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'images/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'images/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "adrian_omr.png","samples/answer-key/weighted-answers/images/adrian_omr.png","outputs/answer-key/weighted-answers/images/CheckedOMRs/adrian_omr.png","5.5","B","E","A","C","B"
+ "adrian_omr_2.png","samples/answer-key/weighted-answers/images/adrian_omr_2.png","outputs/answer-key/weighted-answers/images/CheckedOMRs/adrian_omr_2.png","10.0","C","E","A","B","B"
+
+ ''',
+ })
+# ---
+# name: test_run_community_Antibodyy
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6"
+ "simple_omr_sheet.jpg","samples/community/Antibodyy/simple_omr_sheet.jpg","outputs/community/Antibodyy/CheckedOMRs/simple_omr_sheet.jpg","0","A","C","B","D","E","B"
+
+ ''',
+ })
+# ---
+# name: test_run_community_Sandeep_1507
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Booklet_No","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Booklet_No","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Booklet_No","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+ "omr-1.png","samples/community/Sandeep-1507/omr-1.png","outputs/community/Sandeep-1507/CheckedOMRs/omr-1.png","0","0190880","D","C","B","A","A","B","C","D","D","C","B","A","D","A","B","C","D","","B","D","C","A","C","C","B","A","D","A","AC","C","B","D","C","B","A","B","B","D","D","A","C","B","D","A","C","B","D","B","D","A","A","B","C","D","C","B","A","D","D","A","B","C","D","C","B","A","B","C","D","A","B","C","D","B","A","C","D","C","B","A","D","B","D","A","A","B","A","C","B","D","C","D","B","A","C","C","B","D","B","C","B","A","D","C","B","A","B","C","D","A","A","A","B","B","A","B","C","D","A","A","D","C","B","A","","A","B","C","D","D","D","B","B","C","C","D","C","C","D","D","C","C","B","B","A","A","D","D","B","A","D","C","B","A","A","D","D","B","B","A","A","B","C","D","D","C","B","A","B","D","A","C","C","C","A","A","B","B","D","D","A","A","B","C","D","B","D","A","B","C","D","AD","C","D","B","C","A","B","C","D"
+ "omr-2.png","samples/community/Sandeep-1507/omr-2.png","outputs/community/Sandeep-1507/CheckedOMRs/omr-2.png","0","0no22nonono","A","B","B","A","D","C","B","D","C","D","D","D","B","B","D","D","D","B","C","C","A","A","B","A","D","A","A","B","A","C","A","C","D","D","D","","","C","C","B","B","B","","D","","C","D","","D","B","A","D","B","A","C","A","C","A","C","B","A","D","C","B","C","B","C","D","B","B","D","C","C","D","D","A","D","A","D","C","B","D","C","A","C","","C","B","B","","A","A","D","","B","A","","C","A","D","D","C","C","A","C","A","C","D","A","A","A","D","D","B","C","B","B","B","D","A","C","D","D","A","A","A","C","D","C","C","B","D","A","A","C","B","","D","A","C","C","C","","","","A","C","","D","A","B","A","A","C","A","D","B","B","A","D","A","B","C","A","C","D","D","D","C","A","C","A","C","D","A","A","A","D","A","B","A","B","C","B","A","","B","C","D","D","","","D","C","C","C","","C","A",""
+ "omr-3.png","samples/community/Sandeep-1507/omr-3.png","outputs/community/Sandeep-1507/CheckedOMRs/omr-3.png","0","0nononono73","B","A","C","D","A","D","D","A","C","A","A","B","C","A","A","C","A","B","A","D","C","C","A","D","D","C","C","C","A","C","C","B","B","D","D","C","","","C","B","","","D","A","A","A","A","","A","C","C","C","D","C","","A","B","C","D","B","C","C","C","D","A","B","B","B","D","D","B","B","C","D","B","D","A","B","A","B","C","A","C","A","C","D","","","A","B","","B","C","D","A","D","D","","","C","D","B","B","A","A","D","D","B","A","B","B","C","C","D","D","C","A","D","C","D","C","C","B","C","D","C","D","A","B","D","C","B","D","B","B","","D","","B","D","B","B","C","A","D","","C","","C","","B","C","A","B","B","D","D","D","B","A","D","D","A","D","D","C","B","B","D","C","B","A","C","D","A","D","D","A","C","A","B","D","C","C","C","A","D","","","B","B","","C","C","B","B","C","","","B"
+
+ ''',
+ })
+# ---
+# name: test_run_community_Shamanth
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q21","q22","q23","q24","q25","q26","q27","q28"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q21","q22","q23","q24","q25","q26","q27","q28"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q21","q22","q23","q24","q25","q26","q27","q28"
+ "omr_sheet_01.png","samples/community/Shamanth/omr_sheet_01.png","outputs/community/Shamanth/CheckedOMRs/omr_sheet_01.png","0","A","B","C","D","A","C","C","D"
+
+ ''',
+ })
+# ---
+# name: test_run_community_UPSC_mock
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "answer_key.jpg","samples/community/UPSC-mock/answer_key.jpg","outputs/community/UPSC-mock/CheckedOMRs/answer_key.jpg","200.0","","","","C","D","A","C","C","C","B","A","C","C","B","D","B","D","C","C","B","D","B","D","C","C","C","B","D","D","D","B","A","D","D","C","A","B","C","A","D","A","A","A","D","D","B","A","B","C","B","A","C","D","C","D","A","B","C","A","C","C","C","D","B","C","C","C","C","A","D","A","D","A","D","C","C","D","C","D","A","A","C","B","C","D","C","A","B","C","B","D","A","A","C","A","B","D","C","D","A","C","B","A"
+
+ ''',
+ 'scan-angles/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'scan-angles/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'scan-angles/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","Subject Code","bookletNo","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "angle-1.jpg","samples/community/UPSC-mock/scan-angles/angle-1.jpg","outputs/community/UPSC-mock/scan-angles/CheckedOMRs/angle-1.jpg","70.66666666666669","","","","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+ "angle-2.jpg","samples/community/UPSC-mock/scan-angles/angle-2.jpg","outputs/community/UPSC-mock/scan-angles/CheckedOMRs/angle-2.jpg","70.66666666666669","","","","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+ "angle-3.jpg","samples/community/UPSC-mock/scan-angles/angle-3.jpg","outputs/community/UPSC-mock/scan-angles/CheckedOMRs/angle-3.jpg","70.66666666666669","","","","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+
+ ''',
+ })
+# ---
+# name: test_run_community_UmarFarootAPS
+ dict({
+ 'scans/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll_no","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'scans/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll_no","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+
+ ''',
+ 'scans/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll_no","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100","q101","q102","q103","q104","q105","q106","q107","q108","q109","q110","q111","q112","q113","q114","q115","q116","q117","q118","q119","q120","q121","q122","q123","q124","q125","q126","q127","q128","q129","q130","q131","q132","q133","q134","q135","q136","q137","q138","q139","q140","q141","q142","q143","q144","q145","q146","q147","q148","q149","q150","q151","q152","q153","q154","q155","q156","q157","q158","q159","q160","q161","q162","q163","q164","q165","q166","q167","q168","q169","q170","q171","q172","q173","q174","q175","q176","q177","q178","q179","q180","q181","q182","q183","q184","q185","q186","q187","q188","q189","q190","q191","q192","q193","q194","q195","q196","q197","q198","q199","q200"
+ "scan-type-1.jpg","samples/community/UmarFarootAPS/scans/scan-type-1.jpg","outputs/community/UmarFarootAPS/scans/CheckedOMRs/scan-type-1.jpg","49.0","2468","A","C","B","C","A","D","B","C","B","D","C","A","C","D","B","C","A","B","C","A","C","B","D","C","A","B","D","C","A","C","B","D","B","A","C","D","B","C","A","C","D","A","C","D","A","B","D","C","A","C","D","B","C","A","C","D","B","C","D","A","B","C","B","C","D","B","D","A","C","B","D","A","B","C","B","A","C","D","B","A","C","B","C","B","A","D","B","A","C","D","B","D","B","C","B","D","A","C","B","C","B","C","D","B","C","A","B","C","A","D","C","B","D","B","A","B","C","D","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","B","A","C","B","A","C","A","B","C","B","C","B","A","C","A","C","B","B","C","B","A","C","A","B","A","B","A","B","C","D","B","C","A","C","D","C","A","C","B","A","C","A","B","C","B","D","A","B","C","D","C","B","B","C","A","B","C","B"
+ "scan-type-2.jpg","samples/community/UmarFarootAPS/scans/scan-type-2.jpg","outputs/community/UmarFarootAPS/scans/CheckedOMRs/scan-type-2.jpg","20.0","0234","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","C","D","C","B","A","B","A","D","","","AD","","","","A","D","","","","","","","D","A","","D","","A","","D","","","","A","","","C","","","D","","","A","","","","D","","C","","A","","C","","D","B","B","","","A","","D","","","","D","","","","","A","D","","","B","","","D","","","A","","","D","","","","","","D","","","","A","D","","","A","","B","","D","","","","C","C","D","D","A","","D","","A","D","","","D","","B","D","","","D","","D","B","","","","D","","A","","","","D","","B","","","","","","D","","","A","","","A","","D","","","D"
+
+ ''',
+ })
+# ---
+# name: test_run_community_ibrahimkilic
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "yes_no_questionnarie.jpg","samples/community/ibrahimkilic/yes_no_questionnarie.jpg","outputs/community/ibrahimkilic/CheckedOMRs/yes_no_questionnarie.jpg","0","no","no","no","no","no"
+
+ ''',
+ })
+# ---
+# name: test_run_sample1
+ dict({
+ 'MobileCamera/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+
+ ''',
+ 'MobileCamera/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+
+ ''',
+ 'MobileCamera/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+ "sheet1.jpg","samples/sample1/MobileCamera/sheet1.jpg","outputs/sample1/MobileCamera/CheckedOMRs/sheet1.jpg","0","E503110026","B","","D","B","6","11","20","7","16","B","D","C","D","A","D","B","A","C","C","D"
+
+ ''',
+ })
+# ---
+# name: test_run_sample2
+ dict({
+ 'AdrianSample/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'AdrianSample/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+
+ ''',
+ 'AdrianSample/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
+ "adrian_omr.png","samples/sample2/AdrianSample/adrian_omr.png","outputs/sample2/AdrianSample/CheckedOMRs/adrian_omr.png","0","B","E","A","C","B"
+ "adrian_omr_2.png","samples/sample2/AdrianSample/adrian_omr_2.png","outputs/sample2/AdrianSample/CheckedOMRs/adrian_omr_2.png","0","C","E","A","B","B"
+
+ ''',
+ })
+# ---
+# name: test_run_sample3
+ dict({
+ 'colored-thick-sheet/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'colored-thick-sheet/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'colored-thick-sheet/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "rgb-100-gsm.jpg","samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg","outputs/sample3/colored-thick-sheet/CheckedOMRs/rgb-100-gsm.jpg","0","D","D","A","","C","C","B","","A","C","C","D","A","D","A","C","A","D","B","D","D","C","D","D","D","D","","B","A","D","D","C","","B","","C","D","","","A","","A","C","C","B","C","A","A","C","","C","","D","B","C","","B","C","D","","","C","C","","C","A","B","C","","","","","D","D","C","D","A","","","B","","B","D","C","C","","D","","D","C","D","A","","A","","","A","C","B","A"
+
+ ''',
+ 'xeroxed-thin-sheet/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'xeroxed-thin-sheet/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+
+ ''',
+ 'xeroxed-thin-sheet/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22","q23","q24","q25","q26","q27","q28","q29","q30","q31","q32","q33","q34","q35","q36","q37","q38","q39","q40","q41","q42","q43","q44","q45","q46","q47","q48","q49","q50","q51","q52","q53","q54","q55","q56","q57","q58","q59","q60","q61","q62","q63","q64","q65","q66","q67","q68","q69","q70","q71","q72","q73","q74","q75","q76","q77","q78","q79","q80","q81","q82","q83","q84","q85","q86","q87","q88","q89","q90","q91","q92","q93","q94","q95","q96","q97","q98","q99","q100"
+ "grayscale-80-gsm.jpg","samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg","outputs/sample3/xeroxed-thin-sheet/CheckedOMRs/grayscale-80-gsm.jpg","0","C","D","A","C","C","C","B","A","C","C","B","D","B","D","C","C","B","D","B","D","C","C","C","B","D","D","D","B","A","D","D","C","A","B","C","A","D","A","A","A","D","D","B","A","B","C","B","A","C","D","C","D","A","B","C","A","C","C","C","D","B","C","C","C","C","A","D","A","D","A","D","C","C","D","C","D","A","A","C","B","C","D","C","A","B","C","B","D","A","A","C","A","B","D","C","D","A","C","B","A"
+
+ ''',
+ })
+# ---
+# name: test_run_sample4
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11"
+ "IMG_20201116_143512.jpg","samples/sample4/IMG_20201116_143512.jpg","outputs/sample4/CheckedOMRs/IMG_20201116_143512.jpg","33.0","B","D","C","B","D","C","BC","A","C","D","C"
+ "IMG_20201116_150717658.jpg","samples/sample4/IMG_20201116_150717658.jpg","outputs/sample4/CheckedOMRs/IMG_20201116_150717658.jpg","33.0","B","D","C","B","D","C","BC","A","C","D","C"
+ "IMG_20201116_150750830.jpg","samples/sample4/IMG_20201116_150750830.jpg","outputs/sample4/CheckedOMRs/IMG_20201116_150750830.jpg","-2.0","A","","D","C","AC","A","D","B","C","D","D"
+
+ ''',
+ })
+# ---
+# name: test_run_sample5
+ dict({
+ 'ScanBatch1/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch1/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch1/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+ "camscanner-1.jpg","samples/sample5/ScanBatch1/camscanner-1.jpg","outputs/sample5/ScanBatch1/CheckedOMRs/camscanner-1.jpg","-4.0","E204420102","D","C","A","C","B","08","52","21","85","36","B","C","A","A","D","C","C","AD","A","A","D",""
+
+ ''',
+ 'ScanBatch2/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch2/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+
+ ''',
+ 'ScanBatch2/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20","q21","q22"
+ "camscanner-2.jpg","samples/sample5/ScanBatch2/camscanner-2.jpg","outputs/sample5/ScanBatch2/CheckedOMRs/camscanner-2.jpg","55.0","E204420109","C","C","B","C","C","01","19","10","10","18","D","A","D","D","D","C","C","C","C","D","B","A"
+
+ ''',
+ })
+# ---
+# name: test_run_sample6
+ dict({
+ 'Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+ "reference.png","samples/sample6/reference.png","outputs/sample6/CheckedOMRs/reference.png","0","A"
+
+ ''',
+ 'doc-scans/Manual/ErrorFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'doc-scans/Manual/MultiMarkedFiles.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+
+ ''',
+ 'doc-scans/Results/Results_05AM.csv': '''
+ "file_id","input_path","output_path","score","Roll"
+ "sample_roll_01.jpg","samples/sample6/doc-scans/sample_roll_01.jpg","outputs/sample6/doc-scans/CheckedOMRs/sample_roll_01.jpg","0","A0188877Y"
+ "sample_roll_02.jpg","samples/sample6/doc-scans/sample_roll_02.jpg","outputs/sample6/doc-scans/CheckedOMRs/sample_roll_02.jpg","0","A0203959W"
+ "sample_roll_03.jpg","samples/sample6/doc-scans/sample_roll_03.jpg","outputs/sample6/doc-scans/CheckedOMRs/sample_roll_03.jpg","0","A0204729A"
+
+ ''',
+ })
+# ---
diff --git a/OMRChecker/src/tests/test_all_samples.py b/OMRChecker/src/tests/test_all_samples.py
new file mode 100644
index 0000000000000000000000000000000000000000..79095bf5e96da15680d667ae8216bb801864943e
--- /dev/null
+++ b/OMRChecker/src/tests/test_all_samples.py
@@ -0,0 +1,112 @@
+import os
+import shutil
+from glob import glob
+
+from src.tests.utils import run_entry_point, setup_mocker_patches
+
+
+def read_file(path):
+ with open(path) as file:
+ return file.read()
+
+
+def run_sample(mocker, sample_path):
+ setup_mocker_patches(mocker)
+
+ input_path = os.path.join("samples", sample_path)
+ output_dir = os.path.join("outputs", sample_path)
+ if os.path.exists(output_dir):
+ print(
+ f"Warning: output directory already exists: {output_dir}. This may affect the test execution."
+ )
+
+ run_entry_point(input_path, output_dir)
+
+ sample_outputs = extract_sample_outputs(output_dir)
+
+ print(f"Note: removing output directory: {output_dir}")
+ shutil.rmtree(output_dir)
+
+ return sample_outputs
+
+
+EXT = "*.csv"
+
+
+def extract_sample_outputs(output_dir):
+ sample_outputs = {}
+ for _dir, _subdir, _files in os.walk(output_dir):
+ for file in glob(os.path.join(_dir, EXT)):
+ relative_path = os.path.relpath(file, output_dir)
+ sample_outputs[relative_path] = read_file(file)
+ return sample_outputs
+
+
+def test_run_answer_key_using_csv(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "answer-key/using-csv")
+ assert snapshot == sample_outputs
+
+
+def test_run_answer_key_weighted_answers(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "answer-key/weighted-answers")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample1(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample1")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample2(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample2")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample3(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample3")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample4(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample4")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample5(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample5")
+ assert snapshot == sample_outputs
+
+
+def test_run_sample6(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "sample6")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_Antibodyy(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/Antibodyy")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_ibrahimkilic(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/ibrahimkilic")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_Sandeep_1507(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/Sandeep-1507")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_Shamanth(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/Shamanth")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_UmarFarootAPS(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/UmarFarootAPS")
+ assert snapshot == sample_outputs
+
+
+def test_run_community_UPSC_mock(mocker, snapshot):
+ sample_outputs = run_sample(mocker, "community/UPSC-mock")
+ assert snapshot == sample_outputs
diff --git a/OMRChecker/src/tests/test_edge_cases.py b/OMRChecker/src/tests/test_edge_cases.py
new file mode 100644
index 0000000000000000000000000000000000000000..155d37223006680928dbbbd75054fcbfcc2fb33a
--- /dev/null
+++ b/OMRChecker/src/tests/test_edge_cases.py
@@ -0,0 +1,95 @@
+import os
+from pathlib import Path
+
+import pandas as pd
+
+from src.tests.test_samples.sample2.boilerplate import (
+ CONFIG_BOILERPLATE,
+ TEMPLATE_BOILERPLATE,
+)
+from src.tests.utils import (
+ generate_write_jsons_and_run,
+ remove_file,
+ run_entry_point,
+ setup_mocker_patches,
+)
+
+FROZEN_TIMESTAMP = "1970-01-01"
+CURRENT_DIR = Path("src/tests")
+BASE_SAMPLE_PATH = CURRENT_DIR.joinpath("test_samples", "sample2")
+BASE_RESULTS_CSV_PATH = os.path.join(
+ "outputs", BASE_SAMPLE_PATH, "Results", "Results_05AM.csv"
+)
+BASE_MULTIMARKED_CSV_PATH = os.path.join(
+ "outputs", BASE_SAMPLE_PATH, "Manual", "MultiMarkedFiles.csv"
+)
+
+
+def run_sample(mocker, input_path):
+ setup_mocker_patches(mocker)
+ output_dir = os.path.join("outputs", input_path)
+ run_entry_point(input_path, output_dir)
+
+
+def extract_output_data(path):
+ output_data = pd.read_csv(path, keep_default_na=False)
+ return output_data
+
+
+write_jsons_and_run = generate_write_jsons_and_run(
+ run_sample,
+ sample_path=BASE_SAMPLE_PATH,
+ template_boilerplate=TEMPLATE_BOILERPLATE,
+ config_boilerplate=CONFIG_BOILERPLATE,
+)
+
+
+def test_config_low_dimensions(mocker):
+ def modify_config(config):
+ config["dimensions"]["processing_height"] = 1000
+ config["dimensions"]["processing_width"] = 1000
+
+ exception = write_jsons_and_run(mocker, modify_config=modify_config)
+
+ assert str(exception) == "No Error"
+
+
+def test_different_bubble_dimensions(mocker):
+ # Prevent appending to output csv:
+ remove_file(BASE_RESULTS_CSV_PATH)
+ remove_file(BASE_MULTIMARKED_CSV_PATH)
+
+ exception = write_jsons_and_run(mocker)
+ assert str(exception) == "No Error"
+ original_output_data = extract_output_data(BASE_RESULTS_CSV_PATH)
+
+ def modify_template(template):
+ # Incorrect global bubble size
+ template["bubbleDimensions"] = [5, 5]
+ # Correct bubble size for MCQBlock1a1
+ template["fieldBlocks"]["MCQBlock1a1"]["bubbleDimensions"] = [32, 32]
+ # Incorrect bubble size for MCQBlock1a11
+ template["fieldBlocks"]["MCQBlock1a11"]["bubbleDimensions"] = [10, 10]
+
+ remove_file(BASE_RESULTS_CSV_PATH)
+ remove_file(BASE_MULTIMARKED_CSV_PATH)
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == "No Error"
+
+ results_output_data = extract_output_data(BASE_RESULTS_CSV_PATH)
+
+ assert results_output_data.empty
+
+ output_data = extract_output_data(BASE_MULTIMARKED_CSV_PATH)
+
+ equal_columns = [f"q{i}" for i in range(1, 18)]
+ assert (
+ output_data[equal_columns].iloc[0].to_list()
+ == original_output_data[equal_columns].iloc[0].to_list()
+ )
+
+ unequal_columns = [f"q{i}" for i in range(168, 185)]
+ assert not (
+ output_data[unequal_columns].iloc[0].to_list()
+ == original_output_data[unequal_columns].iloc[0].to_list()
+ )
diff --git a/OMRChecker/src/tests/test_samples/sample1/boilerplate.py b/OMRChecker/src/tests/test_samples/sample1/boilerplate.py
new file mode 100644
index 0000000000000000000000000000000000000000..f35c77c1f7081925edbbf8a207b933d45916e7f0
--- /dev/null
+++ b/OMRChecker/src/tests/test_samples/sample1/boilerplate.py
@@ -0,0 +1,14 @@
+TEMPLATE_BOILERPLATE = {
+ "pageDimensions": [300, 400],
+ "bubbleDimensions": [25, 25],
+ "preProcessors": [{"name": "CropPage", "options": {"morphKernel": [10, 10]}}],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [65, 60],
+ "fieldLabels": ["q1..5"],
+ "labelsGap": 52,
+ "bubblesGap": 41,
+ }
+ },
+}
diff --git a/OMRChecker/src/tests/test_samples/sample1/sample.png b/OMRChecker/src/tests/test_samples/sample1/sample.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/src/tests/test_samples/sample1/sample.png differ
diff --git a/OMRChecker/src/tests/test_samples/sample2/boilerplate.py b/OMRChecker/src/tests/test_samples/sample2/boilerplate.py
new file mode 100644
index 0000000000000000000000000000000000000000..22ddfb3a061a96191533c436f0f626401dc7fdfc
--- /dev/null
+++ b/OMRChecker/src/tests/test_samples/sample2/boilerplate.py
@@ -0,0 +1,39 @@
+TEMPLATE_BOILERPLATE = {
+ "pageDimensions": [2550, 3300],
+ "bubbleDimensions": [32, 32],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17,
+ },
+ }
+ ],
+ "fieldBlocks": {
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [197, 300],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": ["q1..17"],
+ },
+ "MCQBlock1a11": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [1770, 1310],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": ["q168..184"],
+ },
+ },
+}
+
+CONFIG_BOILERPLATE = {
+ "dimensions": {
+ "display_height": 960,
+ "display_width": 1280,
+ "processing_height": 1640,
+ "processing_width": 1332,
+ },
+ "outputs": {"show_image_level": 0, "filter_out_multimarked_files": True},
+}
diff --git a/OMRChecker/src/tests/test_samples/sample2/omr_marker.jpg b/OMRChecker/src/tests/test_samples/sample2/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/src/tests/test_samples/sample2/omr_marker.jpg differ
diff --git a/OMRChecker/src/tests/test_samples/sample2/sample.jpg b/OMRChecker/src/tests/test_samples/sample2/sample.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..25d75107cf4885f2974c52cf42003255be285a3e
--- /dev/null
+++ b/OMRChecker/src/tests/test_samples/sample2/sample.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18b85df487c0bc98eb9139949c3fbca2bad57e8f224fe48b495e0951660f65d
+size 237392
diff --git a/OMRChecker/src/tests/test_template_validations.py b/OMRChecker/src/tests/test_template_validations.py
new file mode 100644
index 0000000000000000000000000000000000000000..6a4a84f52469b89a3ac0e1f5f24b9944040ede5f
--- /dev/null
+++ b/OMRChecker/src/tests/test_template_validations.py
@@ -0,0 +1,159 @@
+import os
+from pathlib import Path
+
+from src.tests.test_samples.sample1.boilerplate import TEMPLATE_BOILERPLATE
+from src.tests.utils import (
+ generate_write_jsons_and_run,
+ run_entry_point,
+ setup_mocker_patches,
+)
+
+FROZEN_TIMESTAMP = "1970-01-01"
+CURRENT_DIR = Path("src/tests")
+BASE_SAMPLE_PATH = CURRENT_DIR.joinpath("test_samples", "sample1")
+BASE_SAMPLE_TEMPLATE_PATH = BASE_SAMPLE_PATH.joinpath("template.json")
+
+
+def run_sample(mocker, input_path):
+ setup_mocker_patches(mocker)
+ output_dir = os.path.join("outputs", input_path)
+ run_entry_point(input_path, output_dir)
+
+
+write_jsons_and_run = generate_write_jsons_and_run(
+ run_sample,
+ sample_path=BASE_SAMPLE_PATH,
+ template_boilerplate=TEMPLATE_BOILERPLATE,
+)
+
+
+def test_no_input_dir(mocker):
+ try:
+ run_sample(mocker, "X")
+ except Exception as e:
+ assert str(e) == "Given input directory does not exist: 'X'"
+
+
+def test_no_template(mocker):
+ if os.path.exists(BASE_SAMPLE_TEMPLATE_PATH):
+ os.remove(BASE_SAMPLE_TEMPLATE_PATH)
+ try:
+ run_sample(mocker, BASE_SAMPLE_PATH)
+ except Exception as e:
+ assert (
+ str(e)
+ == "No template file found in the directory tree of src/tests/test_samples/sample1"
+ )
+
+
+def test_empty_template(mocker):
+ def modify_template(_):
+ return {}
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == f"Provided Template JSON is Invalid: '{BASE_SAMPLE_TEMPLATE_PATH}'"
+ )
+
+
+def test_invalid_field_type(mocker):
+ def modify_template(template):
+ template["fieldBlocks"]["MCQ_Block_1"]["fieldType"] = "X"
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == f"Provided Template JSON is Invalid: '{BASE_SAMPLE_TEMPLATE_PATH}'"
+ )
+
+
+def test_overflow_labels(mocker):
+ def modify_template(template):
+ template["fieldBlocks"]["MCQ_Block_1"]["fieldLabels"] = ["q1..100"]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "Overflowing field block 'MCQ_Block_1' with origin [65, 60] and dimensions [189, 5173] in template with dimensions [300, 400]"
+ )
+
+
+def test_overflow_safe_dimensions(mocker):
+ def modify_template(template):
+ template["pageDimensions"] = [255, 400]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == "No Error"
+
+
+def test_field_strings_overlap(mocker):
+ def modify_template(template):
+ template["fieldBlocks"] = {
+ **template["fieldBlocks"],
+ "New_Block": {
+ **template["fieldBlocks"]["MCQ_Block_1"],
+ "fieldLabels": ["q5"],
+ },
+ }
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == (
+ "The field strings for field block New_Block overlap with other existing fields"
+ )
+
+
+def test_custom_label_strings_overlap_single(mocker):
+ def modify_template(template):
+ template["customLabels"] = {
+ "label1": ["q1..2", "q2..3"],
+ }
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "Given field string 'q2..3' has overlapping field(s) with other fields in 'Custom Label: label1': ['q1..2', 'q2..3']"
+ )
+
+
+def test_custom_label_strings_overlap_multiple(mocker):
+ def modify_template(template):
+ template["customLabels"] = {
+ "label1": ["q1..2"],
+ "label2": ["q2..3"],
+ }
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "The field strings for custom label 'label2' overlap with other existing custom labels"
+ )
+
+
+def test_missing_field_block_labels(mocker):
+ def modify_template(template):
+ template["customLabels"] = {"Combined": ["qX", "qY"]}
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert (
+ str(exception)
+ == "Missing field block label(s) in the given template for ['qX', 'qY'] from 'Combined'"
+ )
+
+
+def test_missing_output_columns(mocker):
+ def modify_template(template):
+ template["outputColumns"] = ["qX", "q1..5"]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == (
+ "Some columns are missing in the field blocks for the given output columns"
+ )
+
+
+def test_safe_missing_label_columns(mocker):
+ def modify_template(template):
+ template["outputColumns"] = ["q1..4"]
+
+ exception = write_jsons_and_run(mocker, modify_template=modify_template)
+ assert str(exception) == "No Error"
diff --git a/OMRChecker/src/tests/utils.py b/OMRChecker/src/tests/utils.py
new file mode 100644
index 0000000000000000000000000000000000000000..2b24215e319a6c35463662e1951982deb89cb910
--- /dev/null
+++ b/OMRChecker/src/tests/utils.py
@@ -0,0 +1,97 @@
+import json
+import os
+from copy import deepcopy
+
+from freezegun import freeze_time
+
+from main import entry_point_for_args
+
+FROZEN_TIMESTAMP = "1970-01-01"
+
+
+def setup_mocker_patches(mocker):
+ mock_imshow = mocker.patch("cv2.imshow")
+ mock_imshow.return_value = True
+
+ mock_destroy_all_windows = mocker.patch("cv2.destroyAllWindows")
+ mock_destroy_all_windows.return_value = True
+
+ mock_wait_key = mocker.patch("cv2.waitKey")
+ mock_wait_key.return_value = ord("q")
+
+
+def run_entry_point(input_path, output_dir):
+ args = {
+ "autoAlign": False,
+ "debug": False,
+ "input_paths": [input_path],
+ "output_dir": output_dir,
+ "setLayout": False,
+ "silent": True,
+ }
+ with freeze_time(FROZEN_TIMESTAMP):
+ entry_point_for_args(args)
+
+
+def write_modified(modify_content, boilerplate, sample_json_path):
+ if boilerplate is None:
+ return
+
+ content = deepcopy(boilerplate)
+
+ if modify_content is not None:
+ returned_value = modify_content(content)
+ if returned_value is not None:
+ content = returned_value
+
+ with open(sample_json_path, "w") as f:
+ json.dump(content, f)
+
+
+def remove_file(path):
+ if os.path.exists(path):
+ os.remove(path)
+
+
+def generate_write_jsons_and_run(
+ run_sample,
+ sample_path,
+ template_boilerplate=None,
+ config_boilerplate=None,
+ evaluation_boilerplate=None,
+):
+ if (template_boilerplate or config_boilerplate or evaluation_boilerplate) is None:
+ raise Exception(
+ f"No boilerplates found. Provide atleast one boilerplate to write json."
+ )
+
+ def write_jsons_and_run(
+ mocker,
+ modify_template=None,
+ modify_config=None,
+ modify_evaluation=None,
+ ):
+ sample_template_path, sample_config_path, sample_evaluation_path = (
+ sample_path.joinpath("template.json"),
+ sample_path.joinpath("config.json"),
+ sample_path.joinpath("evaluation.json"),
+ )
+ write_modified(modify_template, template_boilerplate, sample_template_path)
+ write_modified(modify_config, config_boilerplate, sample_config_path)
+ write_modified(
+ modify_evaluation, evaluation_boilerplate, sample_evaluation_path
+ )
+
+ exception = "No Error"
+ try:
+ run_sample(mocker, sample_path)
+ except Exception as e:
+ exception = e
+
+ remove_file(sample_template_path)
+ remove_file(sample_config_path)
+ remove_file(sample_evaluation_path)
+
+ return exception
+
+ return write_jsons_and_run
diff --git a/OMRChecker/src/utils/__init__.py b/OMRChecker/src/utils/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..e69de29bb2d1d6434b8b29ae775ad8c2e48c5391
diff --git a/OMRChecker/src/utils/file.py b/OMRChecker/src/utils/file.py
new file mode 100644
index 0000000000000000000000000000000000000000..89d827b49c62d78c66510547f1f312a0263e5496
--- /dev/null
+++ b/OMRChecker/src/utils/file.py
@@ -0,0 +1,95 @@
+import argparse
+import json
+import os
+from csv import QUOTE_NONNUMERIC
+from time import localtime, strftime
+
+import pandas as pd
+
+from src.logger import logger
+
+
+def load_json(path, **rest):
+ try:
+ with open(path, "r") as f:
+ loaded = json.load(f, **rest)
+ except json.decoder.JSONDecodeError as error:
+ logger.critical(f"Error when loading json file at: '{path}'\n{error}")
+ exit(1)
+ return loaded
+
+
+class Paths:
+ def __init__(self, output_dir):
+ self.output_dir = output_dir
+ self.save_marked_dir = output_dir.joinpath("CheckedOMRs")
+ self.results_dir = output_dir.joinpath("Results")
+ self.manual_dir = output_dir.joinpath("Manual")
+ self.evaluation_dir = output_dir.joinpath("Evaluation")
+ self.errors_dir = self.manual_dir.joinpath("ErrorFiles")
+ self.multi_marked_dir = self.manual_dir.joinpath("MultiMarkedFiles")
+
+
+def setup_dirs_for_paths(paths):
+ logger.info("Checking Directories...")
+ for save_output_dir in [paths.save_marked_dir]:
+ if not os.path.exists(save_output_dir):
+ logger.info(f"Created : {save_output_dir}")
+ os.makedirs(save_output_dir)
+ os.mkdir(save_output_dir.joinpath("stack"))
+ os.mkdir(save_output_dir.joinpath("_MULTI_"))
+ os.mkdir(save_output_dir.joinpath("_MULTI_", "stack"))
+
+ for save_output_dir in [paths.manual_dir, paths.results_dir, paths.evaluation_dir]:
+ if not os.path.exists(save_output_dir):
+ logger.info(f"Created : {save_output_dir}")
+ os.makedirs(save_output_dir)
+
+ for save_output_dir in [paths.multi_marked_dir, paths.errors_dir]:
+ if not os.path.exists(save_output_dir):
+ logger.info(f"Created : {save_output_dir}")
+ os.makedirs(save_output_dir)
+
+
+def setup_outputs_for_template(paths, template):
+ # TODO: consider moving this into a class instance
+ ns = argparse.Namespace()
+ logger.info("Checking Files...")
+
+ # Include current output paths
+ ns.paths = paths
+
+ ns.empty_resp = [""] * len(template.output_columns)
+ ns.sheetCols = [
+ "file_id",
+ "input_path",
+ "output_path",
+ "score",
+ ] + template.output_columns
+ ns.OUTPUT_SET = []
+ ns.files_obj = {}
+ TIME_NOW_HRS = strftime("%I%p", localtime())
+ ns.filesMap = {
+ "Results": os.path.join(paths.results_dir, f"Results_{TIME_NOW_HRS}.csv"),
+ "MultiMarked": os.path.join(paths.manual_dir, "MultiMarkedFiles.csv"),
+ "Errors": os.path.join(paths.manual_dir, "ErrorFiles.csv"),
+ }
+
+ for file_key, file_name in ns.filesMap.items():
+ if not os.path.exists(file_name):
+ logger.info(f"Created new file: '{file_name}'")
+ # moved handling of files to pandas csv writer
+ ns.files_obj[file_key] = file_name
+ # Create Header Columns
+ pd.DataFrame([ns.sheetCols], dtype=str).to_csv(
+ ns.files_obj[file_key],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ else:
+ logger.info(f"Present : appending to '{file_name}'")
+ ns.files_obj[file_key] = open(file_name, "a")
+
+ return ns
diff --git a/OMRChecker/src/utils/image.py b/OMRChecker/src/utils/image.py
new file mode 100644
index 0000000000000000000000000000000000000000..cb546b0151225306fa2804958e515b6017687255
--- /dev/null
+++ b/OMRChecker/src/utils/image.py
@@ -0,0 +1,155 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import cv2
+import matplotlib.pyplot as plt
+import numpy as np
+
+from src.logger import logger
+
+plt.rcParams["figure.figsize"] = (10.0, 8.0)
+CLAHE_HELPER = cv2.createCLAHE(clipLimit=5.0, tileGridSize=(8, 8))
+
+
+class ImageUtils:
+ """A Static-only Class to hold common image processing utilities & wrappers over OpenCV functions"""
+
+ @staticmethod
+ def save_img(path, final_marked):
+ logger.info(f"Saving Image to '{path}'")
+ cv2.imwrite(path, final_marked)
+
+ @staticmethod
+ def resize_util(img, u_width, u_height=None):
+ if u_height is None:
+ h, w = img.shape[:2]
+ u_height = int(h * u_width / w)
+ return cv2.resize(img, (int(u_width), int(u_height)))
+
+ @staticmethod
+ def resize_util_h(img, u_height, u_width=None):
+ if u_width is None:
+ h, w = img.shape[:2]
+ u_width = int(w * u_height / h)
+ return cv2.resize(img, (int(u_width), int(u_height)))
+
+ @staticmethod
+ def grab_contours(cnts):
+ # source: imutils package
+
+ # if the length the contours tuple returned by cv2.findContours
+ # is '2' then we are using either OpenCV v2.4, v4-beta, or
+ # v4-official
+ if len(cnts) == 2:
+ cnts = cnts[0]
+
+ # if the length of the contours tuple is '3' then we are using
+ # either OpenCV v3, v4-pre, or v4-alpha
+ elif len(cnts) == 3:
+ cnts = cnts[1]
+
+ # otherwise OpenCV has changed their cv2.findContours return
+ # signature yet again and I have no idea WTH is going on
+ else:
+ raise Exception(
+ (
+ "Contours tuple must have length 2 or 3, "
+ "otherwise OpenCV changed their cv2.findContours return "
+ "signature yet again. Refer to OpenCV's documentation "
+ "in that case"
+ )
+ )
+
+ # return the actual contours array
+ return cnts
+
+ @staticmethod
+ def normalize_util(img, alpha=0, beta=255):
+ return cv2.normalize(img, alpha, beta, norm_type=cv2.NORM_MINMAX)
+
+ @staticmethod
+ def auto_canny(image, sigma=0.93):
+ # compute the median of the single channel pixel intensities
+ v = np.median(image)
+
+ # apply automatic Canny edge detection using the computed median
+ lower = int(max(0, (1.0 - sigma) * v))
+ upper = int(min(255, (1.0 + sigma) * v))
+ edged = cv2.Canny(image, lower, upper)
+
+ # return the edged image
+ return edged
+
+ @staticmethod
+ def adjust_gamma(image, gamma=1.0):
+ # build a lookup table mapping the pixel values [0, 255] to
+ # their adjusted gamma values
+ inv_gamma = 1.0 / gamma
+ table = np.array(
+ [((i / 255.0) ** inv_gamma) * 255 for i in np.arange(0, 256)]
+ ).astype("uint8")
+
+ # apply gamma correction using the lookup table
+ return cv2.LUT(image, table)
+
+ @staticmethod
+ def four_point_transform(image, pts):
+ # obtain a consistent order of the points and unpack them
+ # individually
+ rect = ImageUtils.order_points(pts)
+ (tl, tr, br, bl) = rect
+
+ # compute the width of the new image, which will be the
+ width_a = np.sqrt(((br[0] - bl[0]) ** 2) + ((br[1] - bl[1]) ** 2))
+ width_b = np.sqrt(((tr[0] - tl[0]) ** 2) + ((tr[1] - tl[1]) ** 2))
+
+ max_width = max(int(width_a), int(width_b))
+ # max_width = max(int(np.linalg.norm(br-bl)), int(np.linalg.norm(tr-tl)))
+
+ # compute the height of the new image, which will be the
+ height_a = np.sqrt(((tr[0] - br[0]) ** 2) + ((tr[1] - br[1]) ** 2))
+ height_b = np.sqrt(((tl[0] - bl[0]) ** 2) + ((tl[1] - bl[1]) ** 2))
+ max_height = max(int(height_a), int(height_b))
+ # max_height = max(int(np.linalg.norm(tr-br)), int(np.linalg.norm(tl-br)))
+
+ # now that we have the dimensions of the new image, construct
+ # the set of destination points to obtain a "birds eye view",
+ # (i.e. top-down view) of the image, again specifying points
+ # in the top-left, top-right, bottom-right, and bottom-left
+ # order
+ dst = np.array(
+ [
+ [0, 0],
+ [max_width - 1, 0],
+ [max_width - 1, max_height - 1],
+ [0, max_height - 1],
+ ],
+ dtype="float32",
+ )
+
+ transform_matrix = cv2.getPerspectiveTransform(rect, dst)
+ warped = cv2.warpPerspective(image, transform_matrix, (max_width, max_height))
+
+ # return the warped image
+ return warped
+
+ @staticmethod
+ def order_points(pts):
+ rect = np.zeros((4, 2), dtype="float32")
+
+ # the top-left point will have the smallest sum, whereas
+ # the bottom-right point will have the largest sum
+ s = pts.sum(axis=1)
+ rect[0] = pts[np.argmin(s)]
+ rect[2] = pts[np.argmax(s)]
+ diff = np.diff(pts, axis=1)
+ rect[1] = pts[np.argmin(diff)]
+ rect[3] = pts[np.argmax(diff)]
+
+ # return the ordered coordinates
+ return rect
diff --git a/OMRChecker/src/utils/interaction.py b/OMRChecker/src/utils/interaction.py
new file mode 100644
index 0000000000000000000000000000000000000000..72968ec3951bd5bc21fabe545b0f87c0ef3fe896
--- /dev/null
+++ b/OMRChecker/src/utils/interaction.py
@@ -0,0 +1,110 @@
+from dataclasses import dataclass
+
+import cv2
+try:
+ from screeninfo import get_monitors
+ monitor_window = get_monitors()[0]
+ _monitor_width, _monitor_height = monitor_window.width, monitor_window.height
+except Exception:
+ _monitor_width, _monitor_height = 1920, 1080
+
+from src.logger import logger
+from src.utils.image import ImageUtils
+
+
+@dataclass
+class ImageMetrics:
+ # TODO: Move TEXT_SIZE, etc here and find a better class name
+ window_width, window_height = _monitor_width, _monitor_height
+ # for positioning image windows
+ window_x, window_y = 0, 0
+ reset_pos = [0, 0]
+
+
+class InteractionUtils:
+ """Perform primary functions such as displaying images and reading responses"""
+
+ image_metrics = ImageMetrics()
+
+ @staticmethod
+ def show(name, origin, pause=1, resize=False, reset_pos=None, config=None):
+ image_metrics = InteractionUtils.image_metrics
+ if origin is None:
+ logger.info(f"'{name}' - NoneType image to show!")
+ if pause:
+ cv2.destroyAllWindows()
+ return
+ if resize:
+ if not config:
+ raise Exception("config not provided for resizing the image to show")
+ img = ImageUtils.resize_util(origin, config.dimensions.display_width)
+ else:
+ img = origin
+
+ if not is_window_available(name):
+ cv2.namedWindow(name)
+
+ cv2.imshow(name, img)
+
+ if reset_pos:
+ image_metrics.window_x = reset_pos[0]
+ image_metrics.window_y = reset_pos[1]
+
+ cv2.moveWindow(
+ name,
+ image_metrics.window_x,
+ image_metrics.window_y,
+ )
+
+ h, w = img.shape[:2]
+
+ # Set next window position
+ margin = 25
+ w += margin
+ h += margin
+
+ w, h = w // 2, h // 2
+ if image_metrics.window_x + w > image_metrics.window_width:
+ image_metrics.window_x = 0
+ if image_metrics.window_y + h > image_metrics.window_height:
+ image_metrics.window_y = 0
+ else:
+ image_metrics.window_y += h
+ else:
+ image_metrics.window_x += w
+
+ if pause:
+ logger.info(
+ f"Showing '{name}'\n\t Press Q on image to continue. Press Ctrl + C in terminal to exit"
+ )
+
+ wait_q()
+ InteractionUtils.image_metrics.window_x = 0
+ InteractionUtils.image_metrics.window_y = 0
+
+
+@dataclass
+class Stats:
+ # TODO Fill these for stats
+ # Move qbox_vals here?
+ # badThresholds = []
+ # veryBadPoints = []
+ files_moved = 0
+ files_not_moved = 0
+
+
+def wait_q():
+ esc_key = 27
+ while cv2.waitKey(1) & 0xFF not in [ord("q"), esc_key]:
+ pass
+ cv2.destroyAllWindows()
+
+
+def is_window_available(name: str) -> bool:
+ """Checks if a window is available"""
+ try:
+ cv2.getWindowProperty(name, cv2.WND_PROP_VISIBLE)
+ return True
+ except Exception as e:
+ print(e)
+ return False
diff --git a/OMRChecker/src/utils/parsing.py b/OMRChecker/src/utils/parsing.py
new file mode 100644
index 0000000000000000000000000000000000000000..b500271ca038fc0654fba6d823ea517b70719fa1
--- /dev/null
+++ b/OMRChecker/src/utils/parsing.py
@@ -0,0 +1,113 @@
+import re
+from copy import deepcopy
+from fractions import Fraction
+
+from deepmerge import Merger
+from dotmap import DotMap
+
+from src.constants import FIELD_LABEL_NUMBER_REGEX
+from src.defaults import CONFIG_DEFAULTS, TEMPLATE_DEFAULTS
+from src.schemas.constants import FIELD_STRING_REGEX_GROUPS
+from src.utils.file import load_json
+from src.utils.validations import (
+ validate_config_json,
+ validate_evaluation_json,
+ validate_template_json,
+)
+
+OVERRIDE_MERGER = Merger(
+ # pass in a list of tuples,with the
+ # strategies you are looking to apply
+ # to each type.
+ [
+ # (list, ["prepend"]),
+ (dict, ["merge"])
+ ],
+ # next, choose the fallback strategies,
+ # applied to all other types:
+ ["override"],
+ # finally, choose the strategies in
+ # the case where the types conflict:
+ ["override"],
+)
+
+
+def get_concatenated_response(omr_response, template):
+ # Multi-column/multi-row questions which need to be concatenated
+ concatenated_response = {}
+ for field_label, concatenate_keys in template.custom_labels.items():
+ custom_label = "".join([omr_response[k] for k in concatenate_keys])
+ concatenated_response[field_label] = custom_label
+
+ for field_label in template.non_custom_labels:
+ concatenated_response[field_label] = omr_response[field_label]
+
+ return concatenated_response
+
+
+def open_config_with_defaults(config_path):
+ user_tuning_config = load_json(config_path)
+ user_tuning_config = OVERRIDE_MERGER.merge(
+ deepcopy(CONFIG_DEFAULTS), user_tuning_config
+ )
+ validate_config_json(user_tuning_config, config_path)
+ # https://github.com/drgrib/dotmap/issues/74
+ return DotMap(user_tuning_config, _dynamic=False)
+
+
+def open_template_with_defaults(template_path):
+ user_template = load_json(template_path)
+ user_template = OVERRIDE_MERGER.merge(deepcopy(TEMPLATE_DEFAULTS), user_template)
+ validate_template_json(user_template, template_path)
+ return user_template
+
+
+def open_evaluation_with_validation(evaluation_path):
+ user_evaluation_config = load_json(evaluation_path)
+ validate_evaluation_json(user_evaluation_config, evaluation_path)
+ return user_evaluation_config
+
+
+def parse_fields(key, fields):
+ parsed_fields = []
+ fields_set = set()
+ for field_string in fields:
+ fields_array = parse_field_string(field_string)
+ current_set = set(fields_array)
+ if not fields_set.isdisjoint(current_set):
+ raise Exception(
+ f"Given field string '{field_string}' has overlapping field(s) with other fields in '{key}': {fields}"
+ )
+ fields_set.update(current_set)
+ parsed_fields.extend(fields_array)
+ return parsed_fields
+
+
+def parse_field_string(field_string):
+ if "." in field_string:
+ field_prefix, start, end = re.findall(FIELD_STRING_REGEX_GROUPS, field_string)[
+ 0
+ ]
+ start, end = int(start), int(end)
+ if start >= end:
+ raise Exception(
+ f"Invalid range in fields string: '{field_string}', start: {start} is not less than end: {end}"
+ )
+ return [
+ f"{field_prefix}{field_number}" for field_number in range(start, end + 1)
+ ]
+ else:
+ return [field_string]
+
+
+def custom_sort_output_columns(field_label):
+ label_prefix, label_suffix = re.findall(FIELD_LABEL_NUMBER_REGEX, field_label)[0]
+ return [label_prefix, int(label_suffix) if len(label_suffix) > 0 else 0]
+
+
+def parse_float_or_fraction(result):
+ if type(result) == str and "/" in result:
+ result = float(Fraction(result))
+ else:
+ result = float(result)
+ return result
diff --git a/OMRChecker/src/utils/validations.py b/OMRChecker/src/utils/validations.py
new file mode 100644
index 0000000000000000000000000000000000000000..84332a0bc85cd784bfb9782450c9e510676cc656
--- /dev/null
+++ b/OMRChecker/src/utils/validations.py
@@ -0,0 +1,115 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import re
+
+import jsonschema
+from jsonschema import validate
+from rich.table import Table
+
+from src.logger import console, logger
+from src.schemas import SCHEMA_JSONS, SCHEMA_VALIDATORS
+
+
+def validate_evaluation_json(json_data, evaluation_path):
+ logger.info(f"Loading evaluation.json: {evaluation_path}")
+ try:
+ validate(instance=json_data, schema=SCHEMA_JSONS["evaluation"])
+ except jsonschema.exceptions.ValidationError as _err: # NOQA
+ table = Table(show_lines=True)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Error", style="magenta")
+
+ errors = sorted(
+ SCHEMA_VALIDATORS["evaluation"].iter_errors(json_data),
+ key=lambda e: e.path,
+ )
+ for error in errors:
+ key, validator, msg = parse_validation_error(error)
+ if validator == "required":
+ requiredProperty = re.findall(r"'(.*?)'", msg)[0]
+ table.add_row(
+ f"{key}.{requiredProperty}",
+ msg + ". Make sure the spelling of the key is correct",
+ )
+ else:
+ table.add_row(key, msg)
+ console.print(table, justify="center")
+ raise Exception(
+ f"Provided Evaluation JSON is Invalid: '{evaluation_path}'"
+ ) from None
+
+
+def validate_template_json(json_data, template_path):
+ logger.info(f"Loading template.json: {template_path}")
+ try:
+ validate(instance=json_data, schema=SCHEMA_JSONS["template"])
+ except jsonschema.exceptions.ValidationError as _err: # NOQA
+ table = Table(show_lines=True)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Error", style="magenta")
+
+ errors = sorted(
+ SCHEMA_VALIDATORS["template"].iter_errors(json_data),
+ key=lambda e: e.path,
+ )
+ for error in errors:
+ key, validator, msg = parse_validation_error(error)
+
+ # Print preProcessor name in case of options error
+ if key == "preProcessors":
+ preProcessorName = json_data["preProcessors"][error.path[1]]["name"]
+ preProcessorKey = error.path[2]
+ table.add_row(f"{key}.{preProcessorName}.{preProcessorKey}", msg)
+ elif validator == "required":
+ requiredProperty = re.findall(r"'(.*?)'", msg)[0]
+ table.add_row(
+ f"{key}.{requiredProperty}",
+ f"{msg}. Check for spelling errors and make sure it is in camelCase",
+ )
+ else:
+ table.add_row(key, msg)
+ console.print(table, justify="center")
+ raise Exception(
+ f"Provided Template JSON is Invalid: '{template_path}'"
+ ) from None
+
+
+def validate_config_json(json_data, config_path):
+ logger.info(f"Loading config.json: {config_path}")
+ try:
+ validate(instance=json_data, schema=SCHEMA_JSONS["config"])
+ except jsonschema.exceptions.ValidationError as _err: # NOQA
+ table = Table(show_lines=True)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Error", style="magenta")
+ errors = sorted(
+ SCHEMA_VALIDATORS["config"].iter_errors(json_data),
+ key=lambda e: e.path,
+ )
+ for error in errors:
+ key, validator, msg = parse_validation_error(error)
+
+ if validator == "required":
+ requiredProperty = re.findall(r"'(.*?)'", msg)[0]
+ table.add_row(
+ f"{key}.{requiredProperty}",
+ f"{msg}. Check for spelling errors and make sure it is in camelCase",
+ )
+ else:
+ table.add_row(key, msg)
+ console.print(table, justify="center")
+ raise Exception(f"Provided config JSON is Invalid: '{config_path}'") from None
+
+
+def parse_validation_error(error):
+ return (
+ (error.path[0] if len(error.path) > 0 else "$root"),
+ error.validator,
+ error.message,
+ )
diff --git a/README.md b/README.md
index a49d94a9a4c6b64ee5291b5d4cfa41e385b0c51b..1ff342f984e5b3c4275cf804a348a0460c18a4fe 100644
--- a/README.md
+++ b/README.md
@@ -1,10 +1,90 @@
----
-title: Ocr Omr Backend
-emoji: 📚
-colorFrom: indigo
-colorTo: indigo
-sdk: docker
-pinned: false
----
-
-Check out the configuration reference at https://huggingface.co/docs/hub/spaces-config-reference
+# OCR Backend
+
+Backend API for OCR on handwritten images.
+
+## Setup
+
+1. Create a virtual environment:
+ ```bash
+ python -m venv venv
+ ```
+2. Activate the environment:
+ * Windows:
+ ```bash
+ .\venv\Scripts\activate
+ ```
+3. Install dependencies:
+ ```bash
+ pip install -r requirements.txt
+ ```
+4. **For Tesseract OCR:** Install Tesseract on your system. Download from [Tesseract GitHub](https://github.com/tesseract-ocr/tesseract).
+ * If `pytesseract` can't find Tesseract, you might need to set the path in `app.py`:
+ ```python
+ pytesseract.pytesseract.tesseract_cmd = r'C:\Program Files\Tesseract-OCR\tesseract.exe'
+ ```
+5. **For PDF processing:** Install Poppler. Download from [Poppler for Windows](https://github.com/oschwartz10612/poppler-windows/releases).
+ * You will need to update the `poppler_path` in `app.py` to point to the `bin` directory of your Poppler installation (e.g., `r'C:\Program Files\poppler-0.68.0\bin'`)
+
+## Run
+
+```bash
+python app.py
+```
+
+API will be at `http://127.0.0.1:5000`.
+
+## API Endpoints
+
+### `POST /easyocr`
+
+Uses EasyOCR to extract text from images.
+
+**Request:** `multipart/form-data` with `images` (one or more image files).
+
+**Example (curl):**
+
+```bash
+curl -X POST -F "images=@/path/to/your/image1.png" http://127.0.0.1:5000/easyocr
+```
+
+### `POST /tesseract`
+
+Uses Tesseract OCR to extract text from images.
+
+**Request:** `multipart/form-data` with `images` (one or more image files).
+
+**Example (curl):**
+
+```bash
+curl -X POST -F "images=@/path/to/your/image1.png" http://127.0.0.1:5000/tesseract
+```
+
+### `POST /process_question_paper`
+
+Processes an image or PDF of a question paper to extract questions and answers.
+
+**Request:** `multipart/form-data` with `file` (a single image or PDF file).
+
+**Example (curl for image):**
+
+```bash
+curl -X POST -F "file=@/path/to/your/question_paper.png" http://127.0.0.1:5000/process_question_paper
+```
+
+**Example (curl for PDF):**
+
+```bash
+curl -X POST -F "file=@/path/to/your/question_paper.pdf" http://127.0.0.1:5000/process_question_paper
+```
+
+### `GET /evaluate_answers`
+
+Compares OCR extracted texts with the answers from the last processed question paper.
+
+**Request:** None (GET request).
+
+**Example (curl):**
+
+```bash
+curl -X GET http://127.0.0.1:5000/evaluate_answers
+```
diff --git a/app.py b/app.py
new file mode 100644
index 0000000000000000000000000000000000000000..6706a773d328a3aaf4cdcd993304a233bc9c962a
--- /dev/null
+++ b/app.py
@@ -0,0 +1,2242 @@
+import sys
+import platform
+import easyocr
+from pdf2image import convert_from_path, convert_from_bytes
+from flask import Flask, request, jsonify
+from flask_cors import CORS
+from dataclasses import dataclass
+from typing import List, Tuple, Optional, Dict, Any
+from collections import defaultdict
+import numpy as np
+import cv2
+import pytesseract
+from PIL import Image
+import os
+import tempfile
+import difflib
+import re
+from fuzzywuzzy import fuzz
+from dotenv import load_dotenv
+import google.generativeai as genai
+import asyncio
+import base64
+import io
+import json
+import pandas as pd
+import subprocess
+
+# Import the SupabaseHandler
+import uuid
+from datetime import datetime
+from supabase import create_client, Client
+
+_tesseract_cmd = os.getenv("TESSERACT_CMD")
+if _tesseract_cmd:
+ pytesseract.pytesseract.tesseract_cmd = _tesseract_cmd
+elif platform.system() == "Windows":
+ pytesseract.pytesseract.tesseract_cmd = r"C:\Program Files\Tesseract-OCR\tesseract.exe"
+
+
+def _get_poppler_path():
+ env_path = os.getenv("POPPLER_PATH")
+ if env_path:
+ return env_path
+ if platform.system() == "Windows":
+ # Check common install locations
+ candidates = [
+ r'C:\Program Files\poppler\Library\bin',
+ r'C:\Program Files\poppler\poppler-24.08.0\Library\bin',
+ ]
+ # Also scan for any versioned poppler directory
+ poppler_base = r'C:\Program Files\poppler'
+ if os.path.isdir(poppler_base):
+ for entry in os.listdir(poppler_base):
+ candidate = os.path.join(poppler_base, entry, 'Library', 'bin')
+ if candidate not in candidates:
+ candidates.append(candidate)
+ for path in candidates:
+ if os.path.isdir(path):
+ return path
+ return None
+
+
+load_dotenv()
+GEMINI_API_KEY = os.getenv("GEMINI_API_KEY")
+genai.configure(api_key=GEMINI_API_KEY)
+
+app = Flask(__name__)
+CORS(app)
+reader = easyocr.Reader(['en'])
+
+# Global variables to store processing results
+ocr_extracted_texts = []
+last_processed_question_paper_object = None
+last_processed_omr_key = None # Global variable to store OMR answer key
+last_processed_omr_results = None # Global variable to store OMR processing results
+porcessed_omr_results = []
+OMR_IMAGES = []
+
+class SupabaseHandler:
+ def __init__(self):
+ url: str = os.getenv("SUPABASE_URL")
+ key: str = os.getenv("SUPABASE_ANON_KEY")
+ if not url or not key:
+ raise ValueError("Supabase URL and ANON_KEY must be set in environment variables")
+ self.supabase: Client = create_client(url, key)
+
+ def store_evaluation_result(self, teacher_email, evaluation_data, exam_name=None):
+ """
+ Store evaluation result in Supabase with a unique key and exam name
+ Returns the unique key for retrieval
+ """
+ try:
+ # Generate unique key
+ unique_key = str(uuid.uuid4())
+
+ # Prepare data for storage
+ storage_data = {
+ "unique_key": unique_key,
+ "teacher_email": teacher_email,
+ "evaluation_data": evaluation_data,
+ "exam_name": exam_name, # Add exam name field
+ "created_at": datetime.utcnow().isoformat(),
+ "total_students": evaluation_data.get("total_students", 0)
+ }
+
+ # Insert into Supabase
+ result = self.supabase.table("evaluation_results").insert(storage_data).execute()
+
+ if result.data:
+ print(f"Successfully stored evaluation result with key: {unique_key} for exam: {exam_name}")
+ return unique_key
+ else:
+ print("Failed to store evaluation result")
+ return None
+
+ except Exception as e:
+ print(f"Error storing evaluation result: {str(e)}")
+ return None
+
+ def get_evaluation_result(self, unique_key):
+ """
+ Retrieve evaluation result by unique key
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("*").eq("unique_key", unique_key).execute()
+
+ if result.data and len(result.data) > 0:
+ return result.data[0]
+ else:
+ return None
+
+ except Exception as e:
+ print(f"Error retrieving evaluation result: {str(e)}")
+ return None
+
+ def get_teacher_evaluations(self, teacher_email):
+ """
+ Get all evaluation results for a specific teacher
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("unique_key", "created_at", "total_students", "exam_name").eq("teacher_email", teacher_email).order("created_at", desc=True).execute()
+
+ if result.data:
+ return result.data
+ else:
+ return []
+
+ except Exception as e:
+ print(f"Error retrieving teacher evaluations: {str(e)}")
+ return []
+
+class QuestionPaper:
+ def __init__(self, path=None):
+ self.questions = []
+ self.answers = []
+ self.path = path
+ def clean_answers(self):
+ # Remove unwanted patterns from answers
+ unwanted_patterns = [
+ "Time: 15 MinutesMarks: 20",
+ "Time: 15 Minutes Marks: 20",
+ "GENERAL KNOWLEDGE QUESTION PAPER WITH ANSWERS",
+ "GENERAL KNOWLEDGE QUESTION PAPER",
+ ]
+ # Filter out unwanted answers
+ cleaned_answers = []
+ for answer in self.answers:
+ if answer.strip() and answer.strip() not in unwanted_patterns:
+ # Also check if it doesn't match any unwanted pattern with regex
+ is_unwanted = False
+ for pattern in unwanted_patterns:
+ if pattern and re.search(re.escape(pattern), answer, re.IGNORECASE):
+ is_unwanted = True
+ break
+ if not is_unwanted:
+ cleaned_answers.append(answer.strip())
+
+ self.answers = cleaned_answers
+
+ def add_question(self, question_text):
+ self.questions.append(question_text)
+
+ def add_answer(self, answer_text):
+ self.answers.append(answer_text)
+
+ def to_dict(self):
+ return {
+ 'questions': self.questions,
+ 'answers': self.answers
+ }
+
+class OMRAnswerKey:
+ def __init__(self):
+ self.answers = {} # Dictionary mapping question numbers to correct options
+ self.total_marks = 0
+ self.marks_per_question = 1
+ self.negative_marking = 0
+ self.title = ""
+ self.duration = ""
+ self.total_questions = 0
+ self.path = None
+ self.questions = [] # List to store questions if needed
+ self.question_data = [] # List to store complete question data with options
+
+ def __str__(self):
+ return f"OMR Answer Key: {self.title}\nTotal Questions: {self.total_questions}\nAnswers: {self.answers}"
+
+ def set_answers(self, answers: dict):
+ """Set the answer key with question numbers as keys and correct options (A,B,C,D) as values"""
+ self.answers = {int(k): v.upper() for k, v in answers.items() if v.upper() in ['A', 'B', 'C', 'D']}
+ self.total_questions = len(self.answers)
+
+ def set_marking_scheme(self, marks_per_question: float, negative_marking: float = 0):
+ """Set the marking scheme for the answer key"""
+ self.marks_per_question = marks_per_question
+ self.negative_marking = negative_marking
+ self.total_marks = self.total_questions * marks_per_question
+
+ def set_metadata(self, title: str, duration: str):
+ """Set metadata for the answer key"""
+ self.title = title
+ self.duration = duration
+
+ def set_question_data(self, question_data):
+ """Store complete question data including options"""
+ self.question_data = question_data
+ self.questions = [f"{q['number']}. {q['question']}" for q in question_data]
+ self.answers = {q['number']: q['answer'] for q in question_data if q['answer']}
+ self.total_questions = len(question_data)
+
+ def get_question_details(self, question_number):
+ """Get complete details for a specific question"""
+ for q in self.question_data:
+ print(f"Checking question number: {q['number']} with {question_number}")
+ if str(q['number']) == str(question_number):
+ return q
+ return None
+
+ def to_dict(self):
+ return {
+ 'title': self.title,
+ 'duration': self.duration,
+ 'total_questions': self.total_questions,
+ 'answers': self.answers,
+ 'total_marks': self.total_marks,
+ 'marks_per_question': self.marks_per_question,
+ 'negative_marking': self.negative_marking,
+ 'questions': self.questions,
+ 'question_data': self.question_data # Include complete question data
+ }
+
+
+
+def parse_question_paper_text(text):
+ """
+ Improved parsing function that correctly identifies questions and answers
+ """
+ lines = [line.strip() for line in text.split('\n') if line.strip()]
+
+ questions = []
+ answers = []
+
+ # Patterns to ignore (headers, footers, etc.)
+ ignore_patterns = [
+ r'GENERAL KNOWLEDGE QUESTION PAPER.*',
+ r'Time:\s*\d+\s*Minutes.*Marks:\s*\d+',
+ r'Time:\s*\d+\s*MinutesMarks:\s*\d+',
+ r'^\s*$' # Empty lines
+ ]
+
+ # Filter out unwanted lines
+ filtered_lines = []
+ for line in lines:
+ should_ignore = False
+ for pattern in ignore_patterns:
+ if re.match(pattern, line, re.IGNORECASE):
+ should_ignore = True
+ break
+ if not should_ignore:
+ filtered_lines.append(line)
+
+ # Pattern to identify questions (starts with number followed by dot/parenthesis)
+ question_pattern = r'^\d+\s*[.)]\s*(.+)'
+
+ i = 0
+ while i < len(filtered_lines):
+ current_line = filtered_lines[i].strip()
+
+ # Check if current line is a question
+ question_match = re.match(question_pattern, current_line)
+ if question_match:
+ # This is a question
+ question_text = question_match.group(1).strip()
+ questions.append(f"{current_line}") # Keep the full question with number
+
+ # Look for the answer in the next line
+ if i + 1 < len(filtered_lines):
+ next_line = filtered_lines[i + 1].strip()
+ # If next line is not a question (doesn't start with number), it's likely an answer
+ if not re.match(question_pattern, next_line):
+ answers.append(next_line)
+ i += 2 # Skip both question and answer
+ else:
+ # Next line is also a question, so this question might not have an answer
+ # Or the answer might be embedded in the same line
+ # Try to extract answer from the question line itself if it contains common answer patterns
+ answers.append("") # Placeholder for missing answer
+ i += 1
+ else:
+ # Last line and it's a question without answer
+ answers.append("")
+ i += 1
+ else:
+ # This line doesn't match question pattern, skip it or try to pair it with previous question
+ if len(questions) > len(answers):
+ # We have more questions than answers, this might be an answer
+ answers.append(current_line)
+ i += 1
+
+ # Ensure we have equal number of questions and answers
+ while len(answers) < len(questions):
+ answers.append("")
+ while len(questions) < len(answers):
+ questions.append(f"Question {len(questions) + 1}")
+
+ return questions, answers
+
+def improved_clean_and_parse_ocr_text(ocr_text):
+ """
+ Improved parsing with better answer extraction logic
+ """
+ # Remove special characters but keep important ones
+ cleaned_text = re.sub(r'[|@~¥#$%^&*()_+=\[\]{}\\:";\'<>?,./]', ' ', ocr_text)
+
+ # Split by newlines and filter out empty strings
+ lines = [line.strip() for line in cleaned_text.split('\n') if line.strip()]
+
+ individual_answers = []
+
+ # Try to find numbered patterns first
+ numbered_pattern = re.compile(r'(\d+)\s*[.)]\s*([^0-9]+?)(?=\d+\s*[.)]|$)', re.MULTILINE | re.DOTALL)
+ matches = numbered_pattern.findall(cleaned_text)
+
+ if matches:
+ # If we found numbered patterns, use them
+ for number, answer in matches:
+ answer = answer.strip()
+ if answer and len(answer) > 1:
+ individual_answers.append(answer)
+ else:
+ # Fallback to line-by-line processing
+ for line in lines:
+ # Remove leading numbers and punctuation
+ cleaned_line = re.sub(r'^\d+\s*[.)]\s*', '', line).strip()
+ if cleaned_line and len(cleaned_line) > 1:
+ individual_answers.append(cleaned_line)
+
+ return individual_answers
+
+def find_best_match(student_answer, correct_answers, threshold=0.6):
+ """
+ Find the best matching correct answer for a student answer
+ """
+ best_score = 0
+ best_match = None
+
+ for correct_answer in correct_answers:
+ # Use multiple similarity metrics
+ ratio_score = difflib.SequenceMatcher(None, student_answer.lower(), correct_answer.lower()).ratio()
+ fuzzy_score = fuzz.ratio(student_answer.lower(), correct_answer.lower()) / 100.0
+ partial_score = fuzz.partial_ratio(student_answer.lower(), correct_answer.lower()) / 100.0
+
+ # Take the maximum of all scores
+ combined_score = max(ratio_score, fuzzy_score, partial_score)
+
+ if combined_score > best_score:
+ best_score = combined_score
+ best_match = correct_answer
+
+ # Only return match if it meets the threshold
+ if best_score >= threshold:
+ return best_match, best_score
+ else:
+ return None, best_score
+
+def extract_roll_number(student_answer_path):
+ """
+ Extract roll number from student answer sheet using OCR
+ """
+ try:
+ student_answer_image = Image.open(student_answer_path)
+ text = pytesseract.image_to_string(student_answer_image)
+
+ # Look for common roll number patterns
+ roll_patterns = [
+ r'(?i)roll\s*no\s*[:\-]?\s*(\w+)',
+ r'(?i)roll\s*number\s*[:\-]?\s*(\w+)',
+ r'(?i)roll\s*[:\-]?\s*(\w+)',
+ r'(?i)reg\s*no\s*[:\-]?\s*(\w+)',
+ r'(?i)registration\s*[:\-]?\s*(\w+)'
+ ]
+
+ for pattern in roll_patterns:
+ match = re.search(pattern, text)
+ if match:
+ return match.group(1).strip()
+
+ # If no explicit roll number found, try to find number sequences
+ number_sequences = re.findall(r'\b\d{2,}\b', text)
+ if number_sequences:
+ return number_sequences[0] # Return first significant number sequence
+
+ return "Unknown"
+ except Exception as e:
+ print(f"Error extracting roll number: {str(e)}")
+ return "Unknown"
+
+# OMR Section
+
+
+@dataclass
+class BubbleLocation:
+ """Stores information about each bubble"""
+ question_num: int
+ option: str
+ center: Tuple[int, int]
+ radius: int
+ filled: bool = False
+ fill_ratio: float = 0.0
+
+
+
+class CorrectedOMRReader:
+ def __init__(self, image_path: str = None, image_array: np.ndarray = None):
+ """Initialize the OMR Reader with an image"""
+ if image_array is not None:
+ self.image = image_array
+ self.image_path = None
+ elif image_path is not None:
+ self.image = cv2.imread(image_path)
+ self.image_path = image_path
+ else:
+ raise ValueError("Either image_array or image_path must be provided")
+
+ if self.image is None:
+ raise ValueError("Could not load image")
+
+ self.gray = cv2.cvtColor(self.image, cv2.COLOR_BGR2GRAY)
+ self.height, self.width = self.gray.shape
+ self.bubbles = []
+ self.answers = {}
+
+ # Expected grid parameters
+ self.expected_radius = 15 # Approximate bubble radius
+ self.grid_params = {
+ 'rows': 20, # Maximum rows
+ 'cols': 3, # 3 columns of questions
+ 'options': 4 # 4 options per question (A, B, C, D)
+ }
+
+ def preprocess_for_detection(self):
+ """Preprocess specifically for bubble DETECTION (not fill detection)"""
+ blurred = cv2.GaussianBlur(self.gray, (3, 3), 0)
+ _, thresh = cv2.threshold(blurred, 200, 255, cv2.THRESH_BINARY)
+ self.detection_thresh = cv2.bitwise_not(thresh)
+ return self.detection_thresh
+
+ def find_bubble_grid(self):
+ """Find bubble locations using grid detection"""
+ bubbles = []
+
+ param_sets = [
+ {'dp': 1.0, 'minDist': 20, 'param1': 50, 'param2': 28, 'minRadius': 10, 'maxRadius': 20},
+ {'dp': 1.1, 'minDist': 22, 'param1': 45, 'param2': 25, 'minRadius': 11, 'maxRadius': 19},
+ {'dp': 1.2, 'minDist': 25, 'param1': 40, 'param2': 30, 'minRadius': 9, 'maxRadius': 21},
+ ]
+
+ for params in param_sets:
+ circles = cv2.HoughCircles(
+ self.gray,
+ cv2.HOUGH_GRADIENT,
+ dp=params['dp'],
+ minDist=params['minDist'],
+ param1=params['param1'],
+ param2=params['param2'],
+ minRadius=params['minRadius'],
+ maxRadius=params['maxRadius']
+ )
+
+ if circles is not None:
+ circles = np.round(circles[0, :]).astype("int")
+ for (x, y, r) in circles:
+ is_dup = False
+ for bub in bubbles:
+ if np.sqrt((x - bub[0])**2 + (y - bub[1])**2) < 15:
+ is_dup = True
+ break
+ if not is_dup:
+ bubbles.append((x, y, r))
+
+ print(f" Found {len(bubbles)} bubbles with Hough Circles")
+
+ if len(bubbles) < 180:
+ template_bubbles = self.template_matching_detection()
+ bubbles.extend(template_bubbles)
+ print(f" Added {len(template_bubbles)} bubbles with template matching")
+
+ return bubbles
+
+ def template_matching_detection(self):
+ """Use template matching to find bubble locations"""
+ bubbles = []
+ template_size = 30
+ template = np.zeros((template_size, template_size), dtype=np.uint8)
+ cv2.circle(template, (template_size//2, template_size//2), 12, 255, 2)
+
+ result = cv2.matchTemplate(self.gray, template, cv2.TM_CCOEFF_NORMED)
+ threshold = 0.5
+ locations = np.where(result >= threshold)
+
+ for pt in zip(*locations[::-1]):
+ center_x = pt[0] + template_size // 2
+ center_y = pt[1] + template_size // 2
+ too_close = False
+ for (bx, by, _) in bubbles:
+ if np.sqrt((center_x - bx)**2 + (center_y - by)**2) < 20:
+ too_close = True
+ break
+ if not too_close:
+ bubbles.append((center_x, center_y, 12))
+
+ return bubbles
+
+ def detect_bubbles_by_contours(self):
+ """Detect bubbles using contours - focusing on circular shapes"""
+ bubbles = []
+ edge_params = [(30, 100), (50, 150), (20, 80)]
+
+ for low, high in edge_params:
+ edges = cv2.Canny(self.gray, low, high)
+ contours, _ = cv2.findContours(edges, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
+
+ for contour in contours:
+ area = cv2.contourArea(contour)
+ if 150 < area < 900:
+ (x, y), radius = cv2.minEnclosingCircle(contour)
+ perimeter = cv2.arcLength(contour, True)
+ if perimeter > 0:
+ circularity = 4 * np.pi * area / (perimeter * perimeter)
+ if circularity > 0.6 and 8 < radius < 22:
+ is_dup = False
+ for bub in bubbles:
+ if np.sqrt((x - bub[0])**2 + (y - bub[1])**2) < 15:
+ is_dup = True
+ break
+ if not is_dup:
+ bubbles.append((int(x), int(y), int(radius)))
+
+ return bubbles
+
+ def organize_and_filter_bubbles(self, all_bubbles):
+ if not all_bubbles:
+ return []
+
+ filtered_bubbles = []
+ for bubble in all_bubbles:
+ is_duplicate = False
+ for existing in filtered_bubbles:
+ dist = np.sqrt((bubble[0] - existing[0])**2 + (bubble[1] - existing[1])**2)
+ if dist < 15:
+ is_duplicate = True
+ break
+ if not is_duplicate:
+ filtered_bubbles.append(bubble)
+
+ filtered_bubbles.sort(key=lambda b: (b[1], b[0]))
+
+ rows = []
+ current_row = []
+ row_threshold = 20
+
+ for bubble in filtered_bubbles:
+ if not current_row:
+ current_row.append(bubble)
+ else:
+ avg_y = np.mean([b[1] for b in current_row])
+ if abs(bubble[1] - avg_y) < row_threshold:
+ current_row.append(bubble)
+ else:
+ if len(current_row) >= 4:
+ current_row.sort(key=lambda b: b[0])
+ rows.append(current_row)
+ current_row = [bubble]
+
+ if len(current_row) >= 4:
+ current_row.sort(key=lambda b: b[0])
+ rows.append(current_row)
+
+ return rows
+
+ def map_to_questions(self, bubble_rows):
+ mapped_bubbles = []
+ options = ['A', 'B', 'C', 'D']
+
+ if not bubble_rows:
+ return mapped_bubbles
+
+ col1_max = self.width * 0.35
+ col2_max = self.width * 0.68
+
+ for row_idx, row in enumerate(bubble_rows[:20]):
+ col1 = [b for b in row if b[0] < col1_max]
+ col2 = [b for b in row if col1_max <= b[0] < col2_max]
+ col3 = [b for b in row if b[0] >= col2_max]
+
+ if len(col1) >= 4:
+ col1_sorted = sorted(col1, key=lambda b: b[0])[:4]
+ q_num = row_idx + 1
+ for opt_idx, bubble in enumerate(col1_sorted):
+ mapped_bubbles.append(BubbleLocation(q_num, options[opt_idx], (bubble[0], bubble[1]), bubble[2]))
+
+ if len(col2) >= 4:
+ col2_sorted = sorted(col2, key=lambda b: b[0])[:4]
+ q_num = row_idx + 21
+ for opt_idx, bubble in enumerate(col2_sorted):
+ mapped_bubbles.append(BubbleLocation(q_num, options[opt_idx], (bubble[0], bubble[1]), bubble[2]))
+
+ if row_idx < 10 and len(col3) >= 4:
+ col3_sorted = sorted(col3, key=lambda b: b[0])[:4]
+ q_num = row_idx + 41
+ for opt_idx, bubble in enumerate(col3_sorted):
+ mapped_bubbles.append(BubbleLocation(q_num, options[opt_idx], (bubble[0], bubble[1]), bubble[2]))
+
+ return mapped_bubbles
+
+ def analyze_bubble_fill(self, bubble: BubbleLocation):
+ mask = np.zeros(self.gray.shape, dtype=np.uint8)
+ cv2.circle(mask, bubble.center, max(bubble.radius - 5, 5), 255, -1)
+ mean_val = cv2.mean(self.gray, mask=mask)[0]
+
+ large_ring_mask = np.zeros(self.gray.shape, dtype=np.uint8)
+ cv2.circle(large_ring_mask, bubble.center, bubble.radius + 10, 255, -1)
+ cv2.circle(large_ring_mask, bubble.center, bubble.radius + 5, 0, -1)
+ surrounding_mean = cv2.mean(self.gray, mask=large_ring_mask)[0]
+
+ bubble.darkness_score = surrounding_mean - mean_val
+ darkness_threshold = 50
+ absolute_darkness_threshold = 150 # 150
+ bubble.filled = (bubble.darkness_score > darkness_threshold) and (mean_val < absolute_darkness_threshold)
+
+ pixels = self.gray[mask > 0]
+ if len(pixels) > 0:
+ std_dev = np.std(pixels)
+ if std_dev > 25 and mean_val < 170:
+ bubble.filled = True
+ if mean_val < 120:
+ bubble.filled = True
+
+ return bubble.filled
+
+ def process(self):
+ """Main processing pipeline"""
+ print("Starting corrected OMR processing...")
+ print("Detecting bubble locations...")
+
+ all_bubbles = []
+ circles = self.find_bubble_grid()
+ all_bubbles.extend(circles)
+ contour_bubbles = self.detect_bubbles_by_contours()
+ all_bubbles.extend(contour_bubbles)
+ print(f" Contour bubbles found: {len(contour_bubbles)}")
+ print(f"Total bubbles detected: {len(all_bubbles)}")
+
+ if len(all_bubbles) < 180:
+ print("Not enough bubbles detected, using grid-based approach...")
+ grid_bubbles = self.detect_by_grid_assumption()
+ all_bubbles.extend(grid_bubbles)
+ print(f"Added {len(grid_bubbles)} bubbles from grid assumption")
+
+ print("Organizing bubbles into grid...")
+ bubble_rows = self.organize_and_filter_bubbles(all_bubbles)
+ print(f"Organized into {len(bubble_rows)} rows")
+
+ print("Mapping bubbles to questions...")
+ self.bubbles = self.map_to_questions(bubble_rows)
+ print(f"Mapped {len(self.bubbles)} bubble locations")
+
+ print("Analyzing filled bubbles...")
+ for bubble in self.bubbles:
+ self.analyze_bubble_fill(bubble)
+
+ print("Extracting final answers...")
+ self.extract_answers()
+
+ return self.answers
+
+ def detect_by_grid_assumption(self):
+ bubbles = []
+ col_starts = [60, 360, 660]
+ bubble_spacing_x = 45
+ bubble_spacing_y = 28
+ start_y = 50
+
+ for col_idx, col_x in enumerate(col_starts):
+ num_rows = 20 if col_idx < 2 else 10
+ for row in range(num_rows):
+ y = start_y + row * bubble_spacing_y
+ for opt in range(4):
+ x = col_x + opt * bubble_spacing_x
+ exists = False
+ for existing in bubbles:
+ if np.sqrt((x - existing[0])**2 + (y - existing[1])**2) < 20:
+ exists = True
+ break
+ if not exists:
+ bubbles.append((x, y, 13))
+ return bubbles
+
+ def extract_answers(self):
+ questions = defaultdict(list)
+ for bubble in self.bubbles:
+ questions[bubble.question_num].append(bubble)
+
+ self.answers = {}
+ for q_num in sorted(questions.keys()):
+ q_bubbles = questions[q_num]
+ filled = [b for b in q_bubbles if b.filled]
+
+ if not filled:
+ self.answers[q_num] = "---"
+ elif len(filled) == 1:
+ self.answers[q_num] = filled[0].option
+ else:
+ filled.sort(key=lambda b: b.darkness_score, reverse=True)
+ self.answers[q_num] = filled[0].option
+
+ return self.answers
+
+ def visualize_results(self):
+ result_img = self.image.copy()
+ for bubble in self.bubbles:
+ if bubble.filled:
+ cv2.circle(result_img, bubble.center, bubble.radius, (0, 255, 0), 2)
+ text = f"Q{bubble.question_num}:{bubble.option}"
+ cv2.putText(result_img, text,
+ (bubble.center[0] - 25, bubble.center[1] - bubble.radius - 5),
+ cv2.FONT_HERSHEY_SIMPLEX, 0.3, (0, 0, 255), 1)
+ else:
+ cv2.circle(result_img, bubble.center, bubble.radius, (100, 100, 255), 1)
+ return result_img
+
+ def display_results(self):
+ print("\n" + "="*60)
+ print("DETECTED ANSWERS")
+ print("="*60)
+
+ for i in range(1, 21):
+ row_str = ""
+ ans1 = self.answers.get(i, "---")
+ row_str += f"Q{i:2d}: {ans1:^4} | "
+
+ if i + 20 <= 40:
+ ans2 = self.answers.get(i + 20, "---")
+ row_str += f"Q{i+20:2d}: {ans2:^4} | "
+ else:
+ row_str += " " * 13 + "| "
+
+ if i + 40 <= 50:
+ ans3 = self.answers.get(i + 40, "---")
+ row_str += f"Q{i+40:2d}: {ans3:^4}"
+
+ print(row_str)
+
+ print("\n" + "="*60)
+ print("SUMMARY")
+ print("="*60)
+ answered = sum(1 for v in self.answers.values() if v != "---")
+ print(f"Questions detected: {len(self.answers)}")
+ print(f"Answered: {answered}")
+ print(f"Unanswered: {len(self.answers) - answered}")
+
+
+
+
+def process_single_image(image_data) -> Dict[str, Any]:
+ """Process a single image and return results with fixed indexing"""
+ try:
+ # Convert image data to numpy array
+ if isinstance(image_data, str):
+ # Base64 encoded image
+ image_bytes = base64.b64decode(image_data)
+ image = Image.open(io.BytesIO(image_bytes))
+ image_array = cv2.cvtColor(np.array(image), cv2.COLOR_RGB2BGR)
+ else:
+ # Direct file upload
+ image = Image.open(image_data)
+ image_array = cv2.cvtColor(np.array(image), cv2.COLOR_RGB2BGR)
+
+ # Process the image using the new CorrectedOMRReader
+ reader = CorrectedOMRReader(image_array=image_array)
+ answers = reader.process()
+
+ # No need for indexing fix in the new implementation
+ fixed_answers = answers
+
+ # Calculate CORRECTED statistics for 50 questions total
+ total_questions = 50 # Fixed to always be 50
+ answered = sum(1 for v in fixed_answers.values() if v is not None)
+ unanswered = total_questions - answered
+
+ # Format answers for JSON (convert None to "null" string)
+ formatted_answers = {}
+ for q_num in range(1, total_questions + 1):
+ answer = fixed_answers.get(q_num)
+ formatted_answers[str(q_num)] = answer if answer is not None else "null"
+
+ return {
+ "success": True,
+ "answers": formatted_answers,
+ "summary": {
+ "total_questions": total_questions,
+ "answered": answered,
+ "unanswered": unanswered
+ }
+ }
+
+ except Exception as e:
+ return {
+ "success": False,
+ "error": str(e),
+ "answers": {},
+ "summary": {
+ "total_questions": 50,
+ "answered": 0,
+ "unanswered": 50
+ }
+ }
+
+
+
+
+@app.route('/health', methods=['GET'])
+def health_check():
+ """Health check endpoint"""
+ return jsonify({
+ "status": "healthy",
+ "message": "OMR API is running"
+ })
+
+@app.route('/', methods=['GET'])
+def home():
+ """Home endpoint with API documentation"""
+ return jsonify({
+ "message": "OMR Processing API",
+ "version": "1.0",
+ "endpoints": {
+ "/process_omr": {
+ "method": "POST",
+ "description": "Process OMR answer sheets",
+ "accepts": [
+ "Multipart form data with 'images' field",
+ "JSON with base64 encoded images in 'images' array"
+ ],
+ "returns": "JSON with detected answers and summary"
+ },
+ "/health": {
+ "method": "GET",
+ "description": "Health check endpoint"
+ }
+ },
+ "example_response": {
+ "success": True,
+ "answers": {
+ "1": "A",
+ "2": "B",
+ "3": "null"
+ },
+ "summary": {
+ "total_questions": 50,
+ "answered": 45,
+ "unanswered": 5
+ }
+ }
+ })
+
+
+# <----------------->
+
+@app.route('/easyocr', methods=['POST'])
+def easyocr_image():
+ if 'images' not in request.files:
+ return jsonify({'error': 'No image files provided'}), 400
+
+ images = request.files.getlist('images')
+ extracted_texts = []
+
+ for image_file in images:
+ try:
+ # Save the image to a temporary file
+ with tempfile.NamedTemporaryFile(delete=False, suffix='.png') as temp_image_file:
+ image_file.save(temp_image_file.name)
+ temp_path = temp_image_file.name
+
+ try:
+ image_np = np.frombuffer(open(temp_path, 'rb').read(), np.uint8)
+ image = cv2.imdecode(image_np, cv2.IMREAD_COLOR)
+
+ # Perform OCR
+ result = reader.readtext(image)
+
+ # Extract text from the result
+ text = " ".join([item[1] for item in result])
+ extracted_texts.append(text)
+ ocr_extracted_texts.append(text)
+ finally:
+ # Clean up temp file
+ if os.path.exists(temp_path):
+ os.unlink(temp_path)
+ except Exception as e:
+ extracted_texts.append(f"Error processing image with EasyOCR: {str(e)}")
+
+ return jsonify({'extracted_texts': extracted_texts})
+
+@app.route('/tesseract', methods=['POST'])
+def tesseract_image():
+ if 'images' not in request.files:
+ return jsonify({'error': 'No image files provided'}), 400
+
+ images = request.files.getlist('images')
+ extracted_texts = []
+
+ for image_file in images:
+ try:
+ # Save the image to a temporary file
+ with tempfile.NamedTemporaryFile(delete=False, suffix='.png') as temp_image_file:
+ image_file.save(temp_image_file.name)
+ temp_path = temp_image_file.name
+
+ try:
+ with Image.open(temp_path) as image:
+ # Perform OCR using Tesseract
+ text = pytesseract.image_to_string(image)
+ extracted_texts.append(text.strip())
+ ocr_extracted_texts.append(text.strip())
+ finally:
+ # Clean up the temporary file
+ if os.path.exists(temp_path):
+ os.unlink(temp_path)
+ except Exception as e:
+ extracted_texts.append(f"Error processing image with Tesseract: {str(e)}")
+
+ return jsonify({'extracted_texts': extracted_texts})
+
+@app.route('/process_question_paper', methods=['POST'])
+def process_question_paper():
+ global last_processed_question_paper_object
+
+ if 'file' not in request.files:
+ return jsonify({'error': 'No file provided'}), 400
+
+ file = request.files['file']
+ if file.filename == '':
+ return jsonify({'error': 'No file selected'}), 400
+
+ question_paper = QuestionPaper()
+
+ try:
+ # Create Images directory if it doesn't exist
+ images_dir = os.path.join(app.root_path, 'Images')
+ os.makedirs(images_dir, exist_ok=True)
+
+ if file.filename.lower().endswith('.pdf'):
+ question_paper_filename = "question_paper.pdf"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ # Initialize the global object with the path
+ question_paper.path = question_paper_path
+
+ # For PDF processing
+ images_from_pdf = convert_from_path(question_paper_path, poppler_path=_get_poppler_path())
+
+ all_text = ""
+ for page_image in images_from_pdf:
+ text = pytesseract.image_to_string(page_image)
+ all_text += text + "\n"
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(all_text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ else:
+ # Process as image
+ question_paper_filename = "question_paper.png"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ question_paper.path = question_paper_path
+
+ image = Image.open(question_paper_path)
+ text = pytesseract.image_to_string(image)
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ # Clean the answers (remove any remaining unwanted patterns)
+ question_paper.clean_answers()
+
+ # Store the processed question paper globally
+ last_processed_question_paper_object = question_paper
+
+ return jsonify(question_paper.to_dict())
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+
+def gemini_evaluate_answer_sheet_with_roll(question_paper_path, student_answer_path, questions, correct_answers, paddle_results):
+ """
+ Evaluate entire answer sheet using Gemini and extract roll number
+ """
+ try:
+ model = genai.GenerativeModel('gemini-2.5-flash')
+
+ # Create the expected answers list for the prompt
+ expected_answers_text = "\n".join([f"{i+1}. {answer}" for i, answer in enumerate(correct_answers)])
+
+ prompt_text = f"""You are an OCR Assitant for an evaluvation script.
+ You will be given an image of a question paper and an image of a student's handwritten answers along with traditional OCR evaluvations.
+ Your task is assist the traditional OCR in overcoming its limitation with handwritten text the image may have bad quality handwritten text which the OCR may fail to extract and evaluvate properly, this is where you come in.
+ Your task is to Just do a double check of the OCR results and correct any mistakes or missing answers. and provide the result in a structured way.
+
+ Expected correct answers:
+ {expected_answers_text}
+ Traditional OCR Evaluation Results:
+ {paddle_results}
+
+ Instructions:
+ - First, identify and extract the student's roll number from the answer sheet
+ - Compare the student's handwritten answers with the expected answers above
+ - Small spelling mistakes should be ignored and considered correct
+ - If an answer has been crossed out or strikethrough, consider it incorrect
+ - Be lenient with handwriting recognition issues
+ - Look for answers by question numbers (1, 2, 3, etc.)
+
+ Please evaluate ALL questions and respond in this EXACT JSON format:
+ {{
+ "roll_number": "extracted_roll_number_here",
+ "evaluations": [
+ {{"question_number": 1, "status": "Correct"}},
+ {{"question_number": 2, "status": "Wrong"}},
+ {{"question_number": 3, "status": "Missing"}},
+ ...
+ ]
+ }}
+
+ For roll_number: Look for patterns like "Roll No:", "Roll Number:", "Reg No:", or any number sequence that appears to be a student identifier.
+
+ For each question, use ONLY one of these three status values:
+ - "Correct" - if the student's answer matches the expected answer (allowing for minor spelling)
+ - "Wrong" - if the student's answer is clearly different from the expected answer
+ - "Missing" - if no answer is visible for this question number
+
+ Respond with ONLY the JSON format above, no other text.
+
+ ! Note
+ Ignore texts like `GENERAL KNOWLEDGE QUESTION PAPER WITH ANSWERS` and the final output should only have actual questions.
+
+ """
+
+ # Handle PDF vs Image for question paper
+ if question_paper_path.lower().endswith('.pdf'):
+ # Convert PDF to images
+ pdf_images = convert_from_path(question_paper_path, poppler_path=_get_poppler_path())
+ question_paper_img = pdf_images[0] # Use first page
+ else:
+ question_paper_img = Image.open(question_paper_path)
+
+ # Load student answer image
+ student_answer_img = Image.open(student_answer_path)
+
+ # Create content for the model
+ content = [prompt_text, question_paper_img, student_answer_img]
+
+ response = model.generate_content(content)
+ result_text = response.text.strip()
+
+ print(f"Gemini response: {result_text}")
+
+ # Try to parse JSON response
+ import json
+ try:
+ # Clean the response - sometimes Gemini adds markdown formatting
+ if "```json" in result_text:
+ result_text = result_text.split("```json")[1].split("```")[0].strip()
+ elif "```" in result_text:
+ result_text = result_text.split("```")[1].strip()
+
+ parsed_result = json.loads(result_text)
+ return parsed_result["roll_number"], parsed_result["evaluations"]
+ except (json.JSONDecodeError, KeyError) as e:
+ print(f"Failed to parse JSON response: {e}")
+ print(f"Raw response: {result_text}")
+ # Fallback - extract roll number using OCR and create default "Error" results
+ roll_number = extract_roll_number(student_answer_path)
+ return roll_number, [{"question_number": i+1, "status": "Error"} for i in range(len(correct_answers))]
+
+ except Exception as e:
+ print(f"Error in Gemini evaluation: {str(e)}")
+ # Return error status for all questions with OCR extracted roll number
+ roll_number = extract_roll_number(student_answer_path)
+ return roll_number, [{"question_number": i+1, "status": "Error"} for i in range(len(correct_answers))]
+
+def quick_match(correct_list, messy_student_list, min_score=80):
+ """Quick function to match messy student answers"""
+ from fuzzywuzzy import process
+ import re
+
+ results = []
+ used = set()
+
+ for item in messy_student_list:
+ # Extract content
+ content = re.sub(r'^\d+\.?\s*', '', str(item)).strip()
+ if content and content != '-':
+ # Find best match
+ match = process.extractOne(content, correct_list)
+ if match and match[1] >= min_score:
+ q_num = correct_list.index(match[0]) + 1
+ if q_num not in used:
+ used.add(q_num)
+ results.append((item, q_num, match[0], match[1]))
+
+ return results
+
+def process_with_paddle_ocr(image_path, correct_answers):
+ """
+ Process an image with PaddleOCR and perform similarity matching with correct answers
+ Returns:
+ tuple: (extracted_text, similarity_scores, average_similarity)
+ """
+ try:
+ # Initialize PaddleOCR
+ from paddleocr import PaddleOCR
+ print("Initializing PaddleOCR...")
+ ocr = PaddleOCR(
+ use_doc_orientation_classify=True,
+ use_doc_unwarping=False,
+ use_textline_orientation=False
+ )
+ print("PaddleOCR initialized.")
+ # Read and process the image
+ # result = ocr.ocr(image_path, cls=True)
+ print("Preditcing")
+ result = ocr.predict(image_path)
+ print("PaddleOCR processing completed.")
+ # print(f"PaddleOCR result: {result}")
+ print("Correct Answers are:")
+ print(correct_answers)
+ for res in result:
+ words = res["rec_texts"]
+ print(f"PaddleOCR extracted words: {words}")
+ # words = result["rec_texts"]
+ result = quick_match(correct_answers, words, min_score=85)
+ print(f"PaddleOCR matched results: {result}")
+
+ return result
+
+ except Exception as e:
+ print(f"Error in PaddleOCR processing: {str(e)}")
+ return None, [], 0
+
+# OCR Evaluvation Endpoint
+@app.route('/evaluate_answers', methods=['POST'])
+def evaluate_answers():
+ global ocr_extracted_texts
+ if 'student_answers' not in request.files:
+ return jsonify({"error": "Missing student answers"}), 400
+
+ student_answer_files = request.files.getlist('student_answers')
+
+ # Get teacher email and exam name from the request
+ teacher_email = request.form.get('teacher_email', 'unknown@example.com')
+ exam_name = request.form.get('exam_name', 'Untitled Exam') # Get exam name from form data
+
+ # Retrieve the question paper object
+ question_paper = last_processed_question_paper_object
+
+ if last_processed_question_paper_object is None:
+ return jsonify({'error': 'Question paper not found or processed yet'}), 404
+
+ student_answer_paths = []
+ try:
+ # Save student answer files temporarily
+ for student_answer_file in student_answer_files:
+ with tempfile.NamedTemporaryFile(delete=False, suffix='.png') as temp_ans_file:
+ student_answer_file.save(temp_ans_file.name)
+ student_answer_paths.append(temp_ans_file.name)
+
+ # Process each student's answer sheet
+ all_students_results = []
+
+ if question_paper.path and os.path.exists(question_paper.path):
+ print(f"Starting Gemini evaluation for exam: {exam_name} with {len(student_answer_paths)} students...")
+
+ for idx, student_answer_path in enumerate(student_answer_paths):
+ print(f"Processing answer sheet {idx + 1} with PaddleOCR...")
+
+ # First process with PaddleOCR
+ results = process_with_paddle_ocr(
+ student_answer_path,
+ question_paper.answers
+ )
+
+ roll_number, sheet_evaluations = gemini_evaluate_answer_sheet_with_roll(
+ question_paper.path,
+ student_answer_path,
+ question_paper.questions,
+ question_paper.answers,
+ results
+ )
+
+
+
+
+ # Process the results for this student
+ student_results = []
+ for eval_result in sheet_evaluations:
+ question_num = eval_result["question_number"]
+ if 1 <= question_num <= len(question_paper.questions):
+ student_results.append({
+ 'question_number': question_num,
+ 'question_text': question_paper.questions[question_num - 1],
+ 'correct_answer': question_paper.answers[question_num - 1],
+ 'status': eval_result["status"]
+ })
+
+ # Calculate summary for this student
+ correct_count = sum(1 for result in student_results if result['status'] == 'Correct')
+ total_questions = len(student_results)
+ score_percentage = (correct_count / total_questions) * 100 if total_questions > 0 else 0
+
+ student_summary = {
+ 'roll_number': roll_number,
+ 'total_questions': len(question_paper.answers),
+ 'correct_answers': correct_count,
+ 'wrong_answers': sum(1 for result in student_results if result['status'] == 'Wrong'),
+ 'missing_answers': sum(1 for result in student_results if result['status'] == 'Missing'),
+ 'error_answers': sum(1 for result in student_results if result['status'] == 'Error'),
+ 'score_percentage': round(score_percentage, 2),
+ 'evaluation_results': student_results,
+ 'ocr_results': {
+ 'extracted_text': results,
+ }
+ }
+
+ all_students_results.append(student_summary)
+
+ final_results = {
+ 'exam_name': exam_name, # Include exam name in results
+ 'total_students': len(student_answer_paths),
+ 'students_evaluated': all_students_results
+ }
+
+ # STORE THE RESULTS IN SUPABASE WITH EXAM NAME
+ try:
+ supabase_handler = SupabaseHandler()
+ unique_key = supabase_handler.store_evaluation_result(teacher_email, final_results, exam_name)
+
+ if unique_key:
+ # Add the unique key to the response
+ final_results['unique_key'] = unique_key
+ final_results['storage_success'] = True
+ print(f"Results stored successfully with key: {unique_key} for exam: {exam_name}")
+ else:
+ final_results['storage_success'] = False
+ final_results['storage_error'] = "Failed to store results in database"
+ print("Failed to store results in Supabase")
+
+ except Exception as storage_error:
+ print(f"Error storing results: {str(storage_error)}")
+ final_results['storage_success'] = False
+ final_results['storage_error'] = str(storage_error)
+
+ return jsonify(final_results)
+ else:
+ return jsonify({
+ 'error': 'Question paper file not found for Gemini evaluation.'
+ })
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+ finally:
+ # Clean up temporary student answer files
+ for path in student_answer_paths:
+ try:
+ if os.path.exists(path):
+ os.unlink(path)
+ except PermissionError:
+ pass # File still locked on Windows; OS will clean up temp dir
+
+# Get Evaluation
+@app.route('/get_evaluation_result/', methods=['GET'])
+def get_evaluation_result(unique_key):
+ """
+ Get evaluation result by unique key
+ """
+ try:
+ supabase_handler = SupabaseHandler()
+ result = supabase_handler.get_evaluation_result(unique_key)
+
+ if result:
+ return jsonify({
+ 'success': True,
+ 'data': result
+ })
+ else:
+ return jsonify({
+ 'error': 'Evaluation result not found'
+ }), 404
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+
+# Get Teacher Evaluation
+@app.route('/get_teacher_evaluations/', methods=['GET'])
+def get_teacher_evaluations(teacher_email):
+ """
+ Get all evaluation results for a specific teacher
+ """
+ try:
+ supabase_handler = SupabaseHandler()
+ results = supabase_handler.get_teacher_evaluations(teacher_email)
+
+ return jsonify({
+ 'success': True,
+ 'data': results,
+ 'total_evaluations': len(results)
+ })
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
+
+# Get OMR Answer Key
+@app.route('/get_omr_answer_key', methods=['GET'])
+def get_omr_answer_key():
+ """Get the currently stored OMR answer key"""
+ global last_processed_omr_key
+
+ if last_processed_omr_key is None:
+ return jsonify({
+ 'error': 'No answer key has been processed yet'
+ }), 404
+
+ return jsonify({
+ 'success': True,
+ 'answer_key': last_processed_omr_key.to_dict()
+ })
+
+def omr_gemini_process(error_questions, correct_answers, image_file):
+ """
+ Use Gemini to assist in evaluating OMR sheets, especially for error questions
+ """
+ try:
+
+ model = genai.GenerativeModel('gemini-2.5-flash')
+
+ prompt_text = f"""
+ You are an OMR Assistant for an evaluvation script.
+ Your main purpose is to assist in the process.
+
+ Correct Answers to questions sorted by question number: {correct_answers}
+ Error Question numbers: {error_questions}
+ Your task:
+ - From the given image identify the student name and roll number
+ - if for some reason the traditional OMR Processing failed to detect some answers, those question numbers will be provided to you, you should look into those questions form the given image and correct answers.
+ - Only provide answer for the questions that are in the error list.
+ - You can ignore the rest of the question
+ - if Error question is empty, just extract the roll number and name
+
+
+ Please evaluate ALL questions and respond in this EXACT JSON format:
+ {{
+ "roll_number": "extracted_roll_number_here",
+ "evaluations": [
+ {{"question_number": 1, "status": "Correct"}},
+ {{"question_number": 2, "status": "Wrong"}},
+ {{"question_number": 3, "status": "Missing"}},
+ ...
+ ]
+ }}
+
+ """
+ student_answer_img = image_file
+
+ content = [prompt_text, student_answer_img]
+
+ response = model.generate_content(content)
+ result_text = response.text.strip()
+
+ print(f"Gemini response: {result_text}")
+
+ import json
+ try:
+ # Clean the response - sometimes Gemini adds markdown formatting
+ if "```json" in result_text:
+ result_text = result_text.split("```json")[1].split("```")[0].strip()
+ elif "```" in result_text:
+ result_text = result_text.split("```")[1].strip()
+
+ parsed_result = json.loads(result_text)
+ return parsed_result["roll_number"], parsed_result["evaluations"]
+ except (json.JSONDecodeError, KeyError) as e:
+ print(f"Failed to parse JSON response: {e}")
+ print(f"Raw response: {result_text}")
+ # Fallback - extract roll number using OCR and create default "Error" results
+ roll_number = extract_roll_number(os.path.join("OMRChecker", "inputs", "OMRImage.jpg"))
+ return roll_number, [{"question_number": i+1, "status": "Error"} for i in range(len(correct_answers))]
+
+ except Exception as e:
+ print(f"Error in OMR Gemini processing: {str(e)}")
+ return "Unknown", [{"question_number": q, "status": "Error"} for q in error_questions]
+
+@app.route('/evaluate_omr', methods=['POST'])
+def evaluate_omr():
+ """
+ Evaluate OMR answers against stored answer key
+ """
+ global last_processed_omr_key, last_processed_omr_results, porcessed_omr_results, OMR_IMAGES
+
+ # Get teacher email and exam name from the request
+ teacher_email = request.form.get('teacher_email', 'unknown@example.com')
+ exam_name = request.form.get('exam_name', 'Untitled Exam') # Get exam name from form data
+ if not last_processed_omr_key:
+ return jsonify({
+ 'error': 'No answer key has been processed. Please process an answer key first.'
+ }), 400
+
+ if not last_processed_omr_results:
+ return jsonify({
+ 'error': 'No OMR sheet has been processed. Please process an OMR sheet first.'
+ }), 400
+
+ try:
+ # Get the marked answers from the processed OMR
+ if isinstance(last_processed_omr_results, list):
+ omr_data = last_processed_omr_results[0] # Take first sheet if multiple
+ else:
+ omr_data = last_processed_omr_results
+ student_datas = []
+ for idx, omr_data in enumerate(porcessed_omr_results):
+ marked_answers = omr_data
+ image_file = OMR_IMAGES[idx]
+
+ # Get correct answers from answer key (only for questions that exist)
+ correct_answers = last_processed_omr_key.answers
+ total_questions_in_key = len(correct_answers)
+
+ # Evaluate answers only for questions that exist in the answer key
+ evaluation_details = []
+ correct_count = 0
+ wrong_count = 0
+ missing_count = 0
+ error_questions = []
+
+ for q_num in sorted(correct_answers.keys()):
+ print(f"Evaluating Question {q_num}")
+ print(f"Correct Answer: {correct_answers[q_num]} | Marked Answer: {marked_answers.get(str(q_num))}")
+ correct_ans = correct_answers[q_num]
+ marked_ans = marked_answers.get(str(q_num))
+
+ if marked_ans is None or marked_ans == '' or len(str(marked_ans)) > 1 or marked_ans == 'nan':
+ status = 'Missing'
+ error_questions.append(q_num)
+ missing_count += 1
+ elif marked_ans.upper() == correct_ans.upper():
+ status = 'Correct'
+ correct_count += 1
+ else:
+ status = 'Wrong'
+ wrong_count += 1
+
+ evaluation_details.append({
+ 'question_number': q_num,
+ 'question_text': last_processed_omr_key.questions[q_num - 1] if q_num <= len(last_processed_omr_key.questions) else f"Question {q_num}",
+ 'correct_answer': correct_ans,
+ 'marked_answer': marked_ans if marked_ans != 'null' else None,
+ 'status': status
+ })
+
+ roll_no, gemini_result = omr_gemini_process(
+ error_questions,
+ last_processed_omr_key.answers,
+ image_file
+ )
+
+ for err_idx in error_questions:
+ for gemini_eval in gemini_result:
+ if gemini_eval["question_number"] == err_idx:
+ correct_ans = last_processed_omr_key.answers[err_idx]
+ marked_ans = None # Since it was an error question
+ status = gemini_eval["status"]
+
+ if status == "Correct":
+ correct_count += 1
+ # wrong_count -= 1 # Adjust wrong count
+ missing_count -= 1 # Adjust missing count
+ elif status == "Wrong":
+ wrong_count += 1
+ missing_count -= 1 # Adjust missing count
+ elif status == "Missing":
+ missing_count += 1
+
+ # Update the evaluation details
+ for eval_detail in evaluation_details:
+ if eval_detail['question_number'] == err_idx:
+ eval_detail.update({
+ 'marked_answer': marked_ans,
+ 'status': status
+ })
+ break
+ break
+
+ # Calculate score
+ total_score = correct_count * last_processed_omr_key.marks_per_question
+
+ if last_processed_omr_key.negative_marking > 0:
+ total_score -= wrong_count * last_processed_omr_key.negative_marking
+
+ max_score = total_questions_in_key * last_processed_omr_key.marks_per_question
+ student_summary = {
+ 'roll_number': roll_no,
+ 'total_questions': len(last_processed_omr_key.answers),
+ 'correct_answers': correct_count,
+ 'wrong_answers': wrong_count,
+ 'missing_answers': missing_count,
+ 'error_answers': len(error_questions),
+ 'score_percentage': correct_count / len(last_processed_omr_key.answers) * 100 if len(last_processed_omr_key.answers) > 0 else 0,
+ 'evaluation_results': evaluation_details,
+ 'ocr_results': {
+ 'extracted_text': gemini_result,
+ }
+ }
+ student_datas.append(student_summary)
+
+ # Format the data in the required structure for Supabase
+ formatted_evaluation_data = {
+ 'exam_name': exam_name, # Include exam name in results
+ 'total_students': len(student_datas),
+ 'students_evaluated': student_datas
+ }
+
+ # Store results in Supabase (optional — skip if credentials not configured)
+ unique_key = None
+ try:
+ supabase_handler = SupabaseHandler()
+ unique_key = supabase_handler.store_evaluation_result(teacher_email, formatted_evaluation_data, exam_name)
+ except Exception as supa_err:
+ print(f"Supabase storage skipped: {supa_err}")
+
+ # Prepare answer key info
+ answer_key_info = {
+ "title": getattr(last_processed_omr_key, 'title', 'Untitled'),
+ "marks_per_question": last_processed_omr_key.marks_per_question,
+ "negative_marking": last_processed_omr_key.negative_marking
+ }
+
+ # Return response in the same format as stored in Supabase
+ final_result = {
+ "success": True,
+ "unique_key": unique_key,
+ #**formatted_evaluation_data, # Include all the formatted data
+ "additional_info": {
+ "answer_key_info": answer_key_info
+ }
+ }
+
+ return jsonify(final_result)
+
+ except Exception as e:
+ return jsonify({
+ "success": False,
+ "error": f"Evaluation failed: {str(e)}"
+ }), 500
+
+def process_with_gemini(evaluation_details, evaluation_summary, omr_data):
+ """
+ Use Gemini to independently evaluate the OMR sheet and extract student details
+ """
+ global last_processed_omr_key
+
+ try:
+ model = genai.GenerativeModel('gemini-2.5-flash')
+
+ # Prepare the questions and correct answers for Gemini
+ questions_and_answers = ""
+ for i, (q_num, correct_answer) in enumerate(sorted(last_processed_omr_key.answers.items())):
+ question_text = last_processed_omr_key.questions[i] if i < len(last_processed_omr_key.questions) else f"Question {q_num}"
+ questions_and_answers += f"Question {q_num}: {question_text}\nCorrect Answer: {correct_answer}\n\n"
+
+ prompt = f"""
+ You are a teacher grading an OMR answer sheet.
+
+STUDENT INFO: Extract the student's name and roll number from the image.
+
+GRADING TASK: For each question, identify which bubble (A, B, C, or D) is filled/darkened, then compare with the correct answer.
+
+QUESTIONS AND CORRECT ANSWERS:
+{questions_and_answers}
+
+IMPORTANT: Look carefully at each row of bubbles. A filled bubble will be darkened/shaded, while empty bubbles will be white/clear.
+
+Respond in this EXACT JSON format:
+{{
+ "student_info": {{
+ "name": "extracted student name",
+ "roll_no": "extracted roll number"
+ }},
+ "gemini_evaluation": [
+ {{"question": 1, "marked_answer": "C", "correct_answer": "C", "status": "Correct"}},
+ {{"question": 2, "marked_answer": "D", "correct_answer": "D", "status": "Correct"}},
+ // ... continue for all questions
+ ]
+}}
+
+For status: use "Correct", "Wrong", or "Missing" only.
+For marked_answer: use "A", "B", "C", "D", or null if no bubble is clearly filled.
+
+ """
+ # Get the image - we need to retrieve it from the last processed OMR
+ # Since we don't store the image directly, we'll need to work with what we have
+ # For now, let's assume we have access to the image file
+
+ # Check if we have image data stored
+ if 'image_data' in omr_data:
+ # If we have base64 image data
+ image_data = omr_data['image_data']
+ image_bytes = base64.b64decode(image_data)
+ image = Image.open(io.BytesIO(image_bytes))
+ elif 'filename' in omr_data:
+ # Try to find the image file
+ try:
+ # Look for the image in common locations
+ possible_paths = [
+ f"Images/{omr_data['filename']}",
+ f"temp/{omr_data['filename']}",
+ omr_data['filename']
+ ]
+
+ image = None
+ for path in possible_paths:
+ if os.path.exists(path):
+ image = Image.open(path)
+ break
+
+ if image is None:
+ # If we can't find the image, return a fallback result
+ return {
+ "student_info": {
+ "name": "Image not available",
+ "roll_number": "Image not available"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": ["Original image not available for verification"],
+ "notes": "Could not verify due to missing image file"
+ },
+ "gemini_evaluation": []
+ }
+ except Exception as e:
+ print(f"Error loading image: {str(e)}")
+ return {
+ "student_info": {
+ "name": "Error loading image",
+ "roll_number": "Error loading image"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": [f"Error loading image: {str(e)}"],
+ "notes": "Image processing failed"
+ },
+ "gemini_evaluation": []
+ }
+ else:
+ # No image reference available
+ return {
+ "student_info": {
+ "name": "No image data",
+ "roll_number": "No image data"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": ["No image data available"],
+ "notes": "Cannot verify without image"
+ },
+ "gemini_evaluation": []
+ }
+
+ # Generate content with Gemini
+ response = model.generate_content([prompt, image])
+ result_text = response.text.strip()
+
+ print(f"Gemini raw response: {result_text}")
+
+ # Parse the JSON response
+ try:
+ # Clean the response - remove markdown formatting if present
+ if "```json" in result_text:
+ result_text = result_text.split("```json")[1].split("```")[0].strip()
+ elif "```" in result_text:
+ result_text = result_text.split("```")[1].strip()
+
+ parsed_result = json.loads(result_text)
+
+ # Update summary counts and score based on the evaluation
+ if 'gemini_evaluation' in parsed_result:
+ correct_count = sum(1 for item in parsed_result['gemini_evaluation'] if item.get('status') == 'Correct')
+ wrong_count = sum(1 for item in parsed_result['gemini_evaluation'] if item.get('status') == 'Wrong')
+ missing_count = sum(1 for item in parsed_result['gemini_evaluation'] if item.get('status') == 'Missing')
+
+ score = (correct_count * last_processed_omr_key.marks_per_question) - (wrong_count * last_processed_omr_key.negative_marking)
+ max_score = len(last_processed_omr_key.answers) * last_processed_omr_key.marks_per_question
+
+ parsed_result['summary'] = {
+ "total_questions": len(last_processed_omr_key.answers),
+ "correct_count": correct_count,
+ "wrong_count": wrong_count,
+ "missing_count": missing_count,
+ "score": score,
+ "max_score": max_score,
+ "percentage": round((score / max_score) * 100, 2) if max_score > 0 else 0
+ }
+
+ return parsed_result
+
+ except json.JSONDecodeError as e:
+ print(f"Failed to parse Gemini JSON response: {e}")
+ print(f"Raw response: {result_text}")
+
+ # Fallback response with extracted text attempt
+ return {
+ "student_info": {
+ "name": "Parse error",
+ "roll_number": "Parse error"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": ["Failed to parse Gemini response"],
+ "notes": f"JSON parse error: {str(e)}"
+ },
+ "gemini_evaluation": [],
+ "raw_response": result_text # Include raw response for debugging
+ }
+ # print(f"Error in Gemini processing: {str(e)}")
+ except Exception as e:
+ return {
+ "student_info": {
+ "name": "Processing error",
+ "roll_number": "Processing error"
+ },
+ "verification": {
+ "evaluation_correct": "unknown",
+ "confidence": "low",
+ "discrepancies": [f"Gemini processing error: {str(e)}"],
+ "notes": "Failed to process with Gemini"
+ },
+ "gemini_evaluation": []
+ }
+def compare_evaluations(our_evaluation, gemini_evaluation):
+ """
+ Compare our automated evaluation with Gemini's independent evaluation
+ """
+ if not gemini_evaluation:
+ return {
+ "comparison_available": False,
+ "reason": "Gemini evaluation not available"
+ }
+
+ matches = 0
+ differences = []
+ total_compared = 0
+
+ # Create a lookup for our evaluation
+ our_eval_lookup = {detail['question_number']: detail for detail in our_evaluation}
+
+ for gemini_item in gemini_evaluation:
+ q_num = gemini_item.get('question')
+ if q_num in our_eval_lookup:
+ total_compared += 1
+ our_status = our_eval_lookup[q_num]['status']
+ gemini_status = gemini_item.get('status')
+
+ if our_status == gemini_status:
+ matches += 1
+ else:
+ differences.append({
+ "question": q_num,
+ "our_evaluation": {
+ "marked_answer": our_eval_lookup[q_num]['marked_answer'],
+ "status": our_status
+ },
+ "gemini_evaluation": {
+ "marked_answer": gemini_item.get('marked_answer'),
+ "status": gemini_status
+ }
+ })
+
+ agreement_rate = (matches / total_compared) * 100 if total_compared > 0 else 0
+
+ return {
+ "comparison_available": True,
+ "total_questions_compared": total_compared,
+ "agreements": matches,
+ "differences_count": len(differences),
+ "agreement_rate": round(agreement_rate, 2),
+ "differences": differences
+ }
+
+
+
+
+# Also need to modify the process_omr endpoint to store image data for later use
+@app.route('/process_omr', methods=['POST'])
+def process_omr_enhanced():
+ """
+ Enhanced OMR processing that stores image data for later Gemini processing
+ """
+ global last_processed_omr_results
+ global OMR_IMAGES
+ global porcessed_omr_results
+
+ OMR_IMAGES = []
+ porcessed_omr_results = []
+ try:
+ results = []
+ print("Starting OMR processing...")
+ # Check if files were uploaded
+ if 'images' in request.files:
+ files = request.files.getlist('images')
+ results = []
+ for idx, file in enumerate(files):
+ if file.filename == '':
+ continue
+
+ print(f"===================================== Processing file {file.filename} =====================================")
+ name, extension = os.path.splitext(file.filename)
+ filename = os.path.join("OMRChecker", "inputs", "OMRImage" + extension)
+ file.save(filename)
+ OMR_IMAGES.append(Image.open(filename))
+
+ result = subprocess.run([sys.executable, os.path.join('OMRChecker', 'main.py'), '--inputDir=' + os.path.join('OMRChecker', 'inputs')])
+
+ print("OMR Finished Processing Successfully")
+ folder = os.path.join("outputs", "Results")
+ csv_files = [f for f in os.listdir(folder) if f.endswith(".csv")]
+ print("CSV FILES:", csv_files)
+ result_file = os.path.join(folder, csv_files[0])
+ print("Found Result File", result_file)
+ df = pd.read_csv(result_file)
+
+ # Convert to JSON
+ data_json = df.to_json(orient="records")
+ parsed_json = json.loads(data_json)
+
+ columns_dict = df.to_dict(orient="list")
+
+ print(columns_dict)
+ questions_only = {k.replace("q", ""): v[0] for k, v in columns_dict.items() if k.startswith("q")}
+
+ last_processed_omr_results = questions_only
+ porcessed_omr_results.append(questions_only)
+
+ if os.path.exists(result_file):
+ os.remove(result_file)
+ print(f"{result_file} deleted")
+
+ return jsonify(parsed_json)
+
+ else:
+ return jsonify({
+ "success": False,
+ "error": "No images provided. Use 'images' field for file uploads.",
+ "results": []
+ }), 400
+
+ except Exception as e:
+ return jsonify({
+ "success": False,
+ "error": f"Server error: {str(e)}",
+ "results": []
+ }), 500
+
+@app.route('/get_question_details/', methods=['GET'])
+def get_question_details(question_number):
+ """Get detailed information about a specific question"""
+ global last_processed_omr_key
+
+ if last_processed_omr_key is None:
+ return jsonify({
+ 'error': 'No answer key has been processed yet'
+ }), 404
+
+ question_data = last_processed_omr_key.get_question_details(question_number)
+ if question_data is None:
+ return jsonify({
+ 'error': f'Question number {question_number} not found'
+ }), 404
+
+ return jsonify({
+ 'success': True,
+ 'question_data': question_data
+ })
+
+@app.route('/debug_parsing', methods=['GET'])
+def debug_parsing():
+ """
+ Debug endpoint to see how OCR text is being parsed
+ """
+ if not ocr_extracted_texts:
+ return jsonify({'error': 'No OCR extracted texts available.'}), 400
+
+ debug_results = []
+
+ for ocr_text in ocr_extracted_texts:
+ parsed_answers = improved_clean_and_parse_ocr_text(ocr_text)
+ debug_results.append({
+ 'original_ocr_text': ocr_text,
+ 'parsed_answers': parsed_answers
+ })
+
+ return jsonify({'debug_results': debug_results})
+
+def extract_omr_metadata(text: str) -> tuple:
+ """Extract title and duration from the question paper text"""
+ title = ""
+ duration = ""
+
+ # Look for title (usually in first few lines, often in caps)
+ lines = text.split('\n')
+ for line in lines[:5]: # Check first 5 lines
+ if line.strip().upper() == line.strip() and len(line.strip()) > 10:
+ title = line.strip()
+ break
+
+ # Look for duration/time
+ time_pattern = r'Time:\s*(\d+)\s*(minutes|mins|min)'
+ duration_match = re.search(time_pattern, text, re.IGNORECASE)
+ if duration_match:
+ duration = f"{duration_match.group(1)} minutes"
+
+ return title, duration
+
+def extract_omr_answers(text: str) -> dict:
+ """Extract answers from the question paper text"""
+ answers = {}
+ questions = []
+ question_data = []
+ current_question = None
+
+ print("\nStarting answer extraction...")
+
+ # Split text into lines and process line by line
+ lines = [line.strip() for line in text.split('\n') if line.strip()]
+
+ # Skip header lines until we find the first question
+ started = False
+ current_dict = None
+
+ for line in lines:
+ print(f"Processing line: {line}")
+
+ # Skip header or empty lines
+ if not started:
+ if line.startswith('1.'):
+ started = True
+ else:
+ continue
+
+ # Check for new question
+ question_match = re.match(r'^(\d+)[.)](.*?)$', line)
+ if question_match:
+ # Save previous question if exists
+ if current_dict:
+ question_data.append(current_dict)
+
+ # Start new question
+ q_num = int(question_match.group(1))
+ q_text = question_match.group(2).strip()
+ current_dict = {
+ 'number': q_num,
+ 'question': q_text,
+ 'options': {},
+ 'answer': None
+ }
+ continue
+
+ # Check for options
+ option_match = re.match(r'^([A-D])[).](.*?)$', line)
+ if option_match and current_dict is not None:
+ opt_letter = option_match.group(1)
+ opt_text = option_match.group(2).strip()
+ current_dict['options'][opt_letter] = opt_text
+ continue
+
+ # Check for answer
+ answer_match = re.match(r'^\s*Answer[:\s]*([A-D]|.+)$', line, re.IGNORECASE)
+ if answer_match and current_dict is not None:
+ answer = answer_match.group(1).strip()
+ # print(f"For Question: {current_dict['number']}, Options are:")
+ # print(current_dict['options'])
+ for opt_letter, opt_text in current_dict['options'].items():
+ if answer.lower() == opt_text.lower():
+ answer = opt_letter
+ break
+ current_dict['answer'] = answer
+ continue
+
+ # Add last question
+ if current_dict:
+ question_data.append(current_dict)
+
+ print("\nExtracted Question Data:")
+ for q in question_data:
+ print(f"\nQuestion {q['number']}:")
+ print(f"Text: {q['question']}")
+ print(f"Options: {q['options']}")
+ print(f"Answer: {q['answer']}")
+
+ # Add to return format
+ if q['answer']:
+ answers[q['number']] = q['answer']
+ questions.append(f"{q['number']}. {q['question']}")
+
+ print(f"\nExtracted {len(questions)} questions and {len(answers)} answers")
+ print("Questions:", questions)
+ print("Answers:", answers)
+
+ return answers, questions
+
+def debug_text_extraction(text: str):
+ """Helper function to debug text extraction issues"""
+ print("=== Extracted Text ===")
+ print(text)
+ print("\n=== Line by Line Analysis ===")
+ for line in text.split('\n'):
+ if line.strip():
+ print(f"Line: {line.strip()}")
+
+@app.route('/process_omr_answer_key', methods=['POST'])
+def process_omr_answer_key():
+ """
+ Process OMR answer key from either:
+ 1. JSON format with direct answers
+ 2. PDF/Image of question paper with answers marked
+
+ For JSON format:
+ {
+ "answers": {
+ "1": "A",
+ "2": "B",
+ ...
+ },
+ "marks_per_question": 1.0, # optional, defaults to 1
+ "negative_marking": 0.0 # optional, defaults to 0
+ }
+
+ For PDF/Image:
+ multipart/form-data with 'file' field containing the question paper
+ """
+ global last_processed_omr_key
+
+ try:
+ omr_key = OMRAnswerKey()
+
+ # Check if file upload or JSON
+ if 'file' in request.files:
+ file = request.files['file']
+ if file.filename == '':
+ return jsonify({'error': 'No file selected'}), 400
+
+ # Create Images directory if it doesn't exist
+ images_dir = os.path.join(app.root_path, 'Images')
+ os.makedirs(images_dir, exist_ok=True)
+
+ if file.filename.lower().endswith('.pdf'):
+ # Save and process PDF
+ answer_key_path = os.path.join(images_dir, "omr_answer_key.pdf")
+ file.save(answer_key_path)
+ omr_key.path = answer_key_path
+
+ # Convert PDF to images and extract text
+ all_text = ""
+ try:
+ print(f"\nProcessing PDF file: {answer_key_path}")
+ images_from_pdf = convert_from_path(
+ answer_key_path,
+ poppler_path=_get_poppler_path(),
+ dpi=300 # Increase DPI for better quality
+ )
+ print(f"Converted PDF to {len(images_from_pdf)} images")
+
+ for idx, page_image in enumerate(images_from_pdf):
+ print(f"\nProcessing page {idx + 1}")
+
+ # Preprocess the image for better OCR
+ # Convert to numpy array
+ img_np = np.array(page_image)
+
+ # Convert to grayscale
+ gray = cv2.cvtColor(img_np, cv2.COLOR_RGB2GRAY)
+
+ # Apply thresholding to get black and white image
+ _, threshold = cv2.threshold(gray, 0, 255, cv2.THRESH_BINARY + cv2.THRESH_OTSU)
+
+ # Save processed image for debugging
+ debug_image_path = os.path.join(images_dir, f"debug_page_{idx + 1}.png")
+ cv2.imwrite(debug_image_path, threshold)
+ print(f"Saved processed image to {debug_image_path}")
+
+ # Configure Tesseract parameters for better accuracy
+ custom_config = r'--oem 3 --psm 6'
+ text = pytesseract.image_to_string(threshold, config=custom_config)
+ print(f"Extracted text length: {len(text)}")
+ all_text += text + "\n"
+
+ print("\nTotal extracted text length:", len(all_text))
+
+ except Exception as e:
+ print(f"Error during PDF processing: {str(e)}")
+ raise
+
+ # Debug the extracted text
+ print("\nDebugging PDF extraction:")
+ debug_text_extraction(all_text)
+
+ # Extract metadata and answers
+ title, duration = extract_omr_metadata(all_text)
+ answers, questions = extract_omr_answers(all_text)
+
+ print("\nExtracted answers:", answers)
+
+ omr_key.set_metadata(title, duration)
+ omr_key.set_answers(answers)
+ omr_key.questions = questions
+
+ else:
+ # Process as image
+ answer_key_path = os.path.join(images_dir, "omr_answer_key.png")
+ file.save(answer_key_path)
+ omr_key.path = answer_key_path
+
+ image = Image.open(answer_key_path)
+ text = pytesseract.image_to_string(image)
+
+ # Debug the extracted text
+ print("\nDebugging Image extraction:")
+ debug_text_extraction(text)
+
+ # Extract metadata and answers
+ title, duration = extract_omr_metadata(text)
+ answers, questions = extract_omr_answers(text)
+
+ print("\nStructured Extraction Results:")
+ print("Title:", title)
+ print("Duration:", duration)
+ print("\nQuestions found:", len(questions))
+ print("Answers found:", len(answers))
+ print("\nAnswers:", answers)
+
+ omr_key.set_metadata(title, duration)
+ omr_key.set_answers(answers)
+ omr_key.questions = questions
+
+ # Set default marking scheme
+ marks_per_question = float(request.form.get('marks_per_question', 1.0))
+ negative_marking = float(request.form.get('negative_marking', 0.0))
+
+ else:
+ # Process JSON input
+ if not request.is_json:
+ return jsonify({'error': 'Request must be JSON or file upload'}), 400
+
+ data = request.get_json()
+
+ if 'answers' not in data:
+ return jsonify({'error': 'Answer key must be provided'}), 400
+
+ # Validate answer format
+ answer_key = data['answers']
+ for q_num, answer in answer_key.items():
+ try:
+ q_num = int(q_num)
+ if not isinstance(answer, str) or answer.upper() not in ['A', 'B', 'C', 'D']:
+ return jsonify({
+ 'error': f'Invalid answer format for question {q_num}. Must be A, B, C, or D'
+ }), 400
+ except ValueError:
+ return jsonify({
+ 'error': f'Question numbers must be integers, got {q_num}'
+ }), 400
+
+ # Set the answers
+ omr_key.set_answers(answer_key)
+
+ # Set metadata if provided
+ title = data.get('title', '')
+ duration = data.get('duration', '')
+ omr_key.set_metadata(title, duration)
+
+ # Set marking scheme
+ marks_per_question = float(data.get('marks_per_question', 1.0))
+ negative_marking = float(data.get('negative_marking', 0.0))
+
+ # Set marking scheme
+ omr_key.set_marking_scheme(marks_per_question, negative_marking)
+
+ # Store globally
+ last_processed_omr_key = omr_key
+
+ return jsonify({
+ 'success': True,
+ 'message': 'OMR answer key processed successfully',
+ 'answer_key': omr_key.to_dict()
+ })
+
+ except Exception as e:
+ return jsonify({
+ 'error': f'Failed to process answer key: {str(e)}'
+ }), 500
+
+if __name__ == '__main__':
+ app.run(
+ host="0.0.0.0",
+ port=int(os.environ.get("PORT", 5000)),
+ debug=os.environ.get("FLASK_DEBUG", "false").lower() == "true"
+ )
\ No newline at end of file
diff --git a/output/test_ocr_res_img.jpg b/output/test_ocr_res_img.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3a857a483398ec504cad6afd36a4b0d2f6e317b8
--- /dev/null
+++ b/output/test_ocr_res_img.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:31eac088a5ed8769b2837fc5064db5461b8dbb972264a44aa3c5693d81b70675
+size 562726
diff --git a/output/test_preprocessed_img.jpg b/output/test_preprocessed_img.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0d44308ec253f162d92fb2ad3d3141d64fcc2bfc
--- /dev/null
+++ b/output/test_preprocessed_img.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:99af06ba58b1a2ac6ae490f63ba0ce86b4df86dd20625abbdfa70a73c0de590d
+size 1318246
diff --git a/output/test_res.json b/output/test_res.json
new file mode 100644
index 0000000000000000000000000000000000000000..da0d897b478568b9d30fca643fb466695efe18da
--- /dev/null
+++ b/output/test_res.json
@@ -0,0 +1,805 @@
+{
+ "input_path": "Images\\test.jpg",
+ "page_index": null,
+ "model_settings": {
+ "use_doc_preprocessor": true,
+ "use_textline_orientation": false
+ },
+ "doc_preprocessor_res": {
+ "input_path": null,
+ "page_index": null,
+ "model_settings": {
+ "use_doc_orientation_classify": true,
+ "use_doc_unwarping": false
+ },
+ "angle": 0
+ },
+ "dt_polys": [
+ [
+ [
+ 209,
+ 389
+ ],
+ [
+ 426,
+ 386
+ ],
+ [
+ 427,
+ 446
+ ],
+ [
+ 210,
+ 449
+ ]
+ ],
+ [
+ [
+ 68,
+ 421
+ ],
+ [
+ 135,
+ 421
+ ],
+ [
+ 135,
+ 485
+ ],
+ [
+ 68,
+ 485
+ ]
+ ],
+ [
+ [
+ 220,
+ 498
+ ],
+ [
+ 381,
+ 492
+ ],
+ [
+ 383,
+ 567
+ ],
+ [
+ 222,
+ 572
+ ]
+ ],
+ [
+ [
+ 80,
+ 524
+ ],
+ [
+ 151,
+ 524
+ ],
+ [
+ 151,
+ 584
+ ],
+ [
+ 80,
+ 584
+ ]
+ ],
+ [
+ [
+ 88,
+ 630
+ ],
+ [
+ 146,
+ 636
+ ],
+ [
+ 138,
+ 712
+ ],
+ [
+ 80,
+ 706
+ ]
+ ],
+ [
+ [
+ 222,
+ 630
+ ],
+ [
+ 559,
+ 626
+ ],
+ [
+ 560,
+ 702
+ ],
+ [
+ 223,
+ 706
+ ]
+ ],
+ [
+ [
+ 175,
+ 744
+ ],
+ [
+ 520,
+ 738
+ ],
+ [
+ 522,
+ 814
+ ],
+ [
+ 177,
+ 820
+ ]
+ ],
+ [
+ [
+ 83,
+ 772
+ ],
+ [
+ 131,
+ 772
+ ],
+ [
+ 131,
+ 819
+ ],
+ [
+ 83,
+ 819
+ ]
+ ],
+ [
+ [
+ 209,
+ 859
+ ],
+ [
+ 628,
+ 838
+ ],
+ [
+ 632,
+ 914
+ ],
+ [
+ 213,
+ 935
+ ]
+ ],
+ [
+ [
+ 85,
+ 878
+ ],
+ [
+ 140,
+ 878
+ ],
+ [
+ 140,
+ 936
+ ],
+ [
+ 85,
+ 936
+ ]
+ ],
+ [
+ [
+ 202,
+ 997
+ ],
+ [
+ 343,
+ 989
+ ],
+ [
+ 348,
+ 1065
+ ],
+ [
+ 207,
+ 1073
+ ]
+ ],
+ [
+ [
+ 81,
+ 1009
+ ],
+ [
+ 149,
+ 1009
+ ],
+ [
+ 149,
+ 1081
+ ],
+ [
+ 81,
+ 1081
+ ]
+ ],
+ [
+ [
+ 209,
+ 1121
+ ],
+ [
+ 508,
+ 1110
+ ],
+ [
+ 512,
+ 1196
+ ],
+ [
+ 212,
+ 1207
+ ]
+ ],
+ [
+ [
+ 0,
+ 1132
+ ],
+ [
+ 2160,
+ 1052
+ ],
+ [
+ 2160,
+ 2938
+ ],
+ [
+ 0,
+ 2938
+ ]
+ ],
+ [
+ [
+ 77,
+ 1136
+ ],
+ [
+ 157,
+ 1136
+ ],
+ [
+ 157,
+ 1205
+ ],
+ [
+ 77,
+ 1205
+ ]
+ ],
+ [
+ [
+ 86,
+ 1277
+ ],
+ [
+ 501,
+ 1252
+ ],
+ [
+ 504,
+ 1314
+ ],
+ [
+ 90,
+ 1339
+ ]
+ ],
+ [
+ [
+ 80,
+ 1396
+ ],
+ [
+ 465,
+ 1366
+ ],
+ [
+ 470,
+ 1431
+ ],
+ [
+ 85,
+ 1460
+ ]
+ ]
+ ],
+ "text_det_params": {
+ "limit_side_len": 64,
+ "limit_type": "min",
+ "thresh": 0.3,
+ "max_side_limit": 4000,
+ "box_thresh": 0.6,
+ "unclip_ratio": 1.5
+ },
+ "text_type": "general",
+ "textline_orientation_angles": [
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1,
+ -1
+ ],
+ "text_rec_score_thresh": 0.0,
+ "return_word_box": false,
+ "rec_texts": [
+ "Venus",
+ "4",
+ "Beu",
+ "5.",
+ "6",
+ "Graphite",
+ "cheetah",
+ "7.",
+ "Bluewhale",
+ "8.",
+ "Sun",
+ "9.",
+ "Babbage",
+ "-",
+ "10.",
+ "1. sahara",
+ "12.Mount"
+ ],
+ "rec_scores": [
+ 0.9222331047058105,
+ 0.9949488639831543,
+ 0.8761672377586365,
+ 0.769047737121582,
+ 0.9965930581092834,
+ 0.9706618785858154,
+ 0.9598208665847778,
+ 0.9705947637557983,
+ 0.9192090630531311,
+ 0.7382487058639526,
+ 0.9669068455696106,
+ 0.981717586517334,
+ 0.9927512407302856,
+ 0.11772612482309341,
+ 0.9063168168067932,
+ 0.926537811756134,
+ 0.9096950888633728
+ ],
+ "rec_polys": [
+ [
+ [
+ 209,
+ 389
+ ],
+ [
+ 426,
+ 386
+ ],
+ [
+ 427,
+ 446
+ ],
+ [
+ 210,
+ 449
+ ]
+ ],
+ [
+ [
+ 68,
+ 421
+ ],
+ [
+ 135,
+ 421
+ ],
+ [
+ 135,
+ 485
+ ],
+ [
+ 68,
+ 485
+ ]
+ ],
+ [
+ [
+ 220,
+ 498
+ ],
+ [
+ 381,
+ 492
+ ],
+ [
+ 383,
+ 567
+ ],
+ [
+ 222,
+ 572
+ ]
+ ],
+ [
+ [
+ 80,
+ 524
+ ],
+ [
+ 151,
+ 524
+ ],
+ [
+ 151,
+ 584
+ ],
+ [
+ 80,
+ 584
+ ]
+ ],
+ [
+ [
+ 88,
+ 630
+ ],
+ [
+ 146,
+ 636
+ ],
+ [
+ 138,
+ 712
+ ],
+ [
+ 80,
+ 706
+ ]
+ ],
+ [
+ [
+ 222,
+ 630
+ ],
+ [
+ 559,
+ 626
+ ],
+ [
+ 560,
+ 702
+ ],
+ [
+ 223,
+ 706
+ ]
+ ],
+ [
+ [
+ 175,
+ 744
+ ],
+ [
+ 520,
+ 738
+ ],
+ [
+ 522,
+ 814
+ ],
+ [
+ 177,
+ 820
+ ]
+ ],
+ [
+ [
+ 83,
+ 772
+ ],
+ [
+ 131,
+ 772
+ ],
+ [
+ 131,
+ 819
+ ],
+ [
+ 83,
+ 819
+ ]
+ ],
+ [
+ [
+ 209,
+ 859
+ ],
+ [
+ 628,
+ 838
+ ],
+ [
+ 632,
+ 914
+ ],
+ [
+ 213,
+ 935
+ ]
+ ],
+ [
+ [
+ 85,
+ 878
+ ],
+ [
+ 140,
+ 878
+ ],
+ [
+ 140,
+ 936
+ ],
+ [
+ 85,
+ 936
+ ]
+ ],
+ [
+ [
+ 202,
+ 997
+ ],
+ [
+ 343,
+ 989
+ ],
+ [
+ 348,
+ 1065
+ ],
+ [
+ 207,
+ 1073
+ ]
+ ],
+ [
+ [
+ 81,
+ 1009
+ ],
+ [
+ 149,
+ 1009
+ ],
+ [
+ 149,
+ 1081
+ ],
+ [
+ 81,
+ 1081
+ ]
+ ],
+ [
+ [
+ 209,
+ 1121
+ ],
+ [
+ 508,
+ 1110
+ ],
+ [
+ 512,
+ 1196
+ ],
+ [
+ 212,
+ 1207
+ ]
+ ],
+ [
+ [
+ 0,
+ 1132
+ ],
+ [
+ 2160,
+ 1052
+ ],
+ [
+ 2160,
+ 2938
+ ],
+ [
+ 0,
+ 2938
+ ]
+ ],
+ [
+ [
+ 77,
+ 1136
+ ],
+ [
+ 157,
+ 1136
+ ],
+ [
+ 157,
+ 1205
+ ],
+ [
+ 77,
+ 1205
+ ]
+ ],
+ [
+ [
+ 86,
+ 1277
+ ],
+ [
+ 501,
+ 1252
+ ],
+ [
+ 504,
+ 1314
+ ],
+ [
+ 90,
+ 1339
+ ]
+ ],
+ [
+ [
+ 80,
+ 1396
+ ],
+ [
+ 465,
+ 1366
+ ],
+ [
+ 470,
+ 1431
+ ],
+ [
+ 85,
+ 1460
+ ]
+ ]
+ ],
+ "rec_boxes": [
+ [
+ 209,
+ 386,
+ 427,
+ 449
+ ],
+ [
+ 68,
+ 421,
+ 135,
+ 485
+ ],
+ [
+ 220,
+ 492,
+ 383,
+ 572
+ ],
+ [
+ 80,
+ 524,
+ 151,
+ 584
+ ],
+ [
+ 80,
+ 630,
+ 146,
+ 712
+ ],
+ [
+ 222,
+ 626,
+ 560,
+ 706
+ ],
+ [
+ 175,
+ 738,
+ 522,
+ 820
+ ],
+ [
+ 83,
+ 772,
+ 131,
+ 819
+ ],
+ [
+ 209,
+ 838,
+ 632,
+ 935
+ ],
+ [
+ 85,
+ 878,
+ 140,
+ 936
+ ],
+ [
+ 202,
+ 989,
+ 348,
+ 1073
+ ],
+ [
+ 81,
+ 1009,
+ 149,
+ 1081
+ ],
+ [
+ 209,
+ 1110,
+ 512,
+ 1207
+ ],
+ [
+ 0,
+ 1052,
+ 2160,
+ 2938
+ ],
+ [
+ 77,
+ 1136,
+ 157,
+ 1205
+ ],
+ [
+ 86,
+ 1252,
+ 504,
+ 1339
+ ],
+ [
+ 80,
+ 1366,
+ 470,
+ 1460
+ ]
+ ]
+}
\ No newline at end of file
diff --git a/requirements.txt b/requirements.txt
new file mode 100644
index 0000000000000000000000000000000000000000..7a4abd3119741fdb3c41345dd7933e84c4c69aa0
--- /dev/null
+++ b/requirements.txt
@@ -0,0 +1,24 @@
+easyocr
+opencv-python
+flask
+flask-cors
+pytesseract
+pillow
+pdf2image
+fuzzywuzzy
+python-levenshtein
+google-generativeai
+python-dotenv
+supabase
+paddlepaddle
+paddleocr
+openpyxl
+deepmerge>=1.1.0
+dotmap>=1.3.30
+jsonschema>=4.17.3
+matplotlib>=3.7.1
+numpy>=1.25.0
+pandas>=2.0.2
+rich>=13.4.2
+screeninfo>=0.8.1
+requests
diff --git a/server.py b/server.py
new file mode 100644
index 0000000000000000000000000000000000000000..b63634859c02cf1cefd7ce57eb98acc513aa7a34
--- /dev/null
+++ b/server.py
@@ -0,0 +1,182 @@
+class QuestionPaper:
+ def __init__(self, path=None):
+ self.questions = []
+ self.answers = []
+ self.path = path
+
+ def clean_answers(self):
+ # Remove unwanted patterns from answers
+ unwanted_patterns = [
+ "Time: 15 MinutesMarks: 20",
+ "Time: 15 Minutes Marks: 20",
+ "GENERAL KNOWLEDGE QUESTION PAPER WITH ANSWERS",
+ "GENERAL KNOWLEDGE QUESTION PAPER",
+ "" # Empty strings
+ ]
+
+ # Filter out unwanted answers
+ cleaned_answers = []
+ for answer in self.answers:
+ if answer.strip() and answer.strip() not in unwanted_patterns:
+ # Also check if it doesn't match any unwanted pattern with regex
+ is_unwanted = False
+ for pattern in unwanted_patterns:
+ if pattern and re.search(re.escape(pattern), answer, re.IGNORECASE):
+ is_unwanted = True
+ break
+ if not is_unwanted:
+ cleaned_answers.append(answer.strip())
+
+ self.answers = cleaned_answers
+
+ def add_question(self, question_text):
+ self.questions.append(question_text)
+
+ def add_answer(self, answer_text):
+ self.answers.append(answer_text)
+
+ def to_dict(self):
+ return {
+ 'questions': self.questions,
+ 'answers': self.answers
+ }
+
+def parse_question_paper_text(text):
+ """
+ Improved parsing function that correctly identifies questions and answers
+ """
+ lines = [line.strip() for line in text.split('\n') if line.strip()]
+
+ questions = []
+ answers = []
+
+ # Patterns to ignore (headers, footers, etc.)
+ ignore_patterns = [
+ r'GENERAL KNOWLEDGE QUESTION PAPER.*',
+ r'Time:\s*\d+\s*Minutes.*Marks:\s*\d+',
+ r'Time:\s*\d+\s*MinutesMarks:\s*\d+',
+ r'^\s*$' # Empty lines
+ ]
+
+ # Filter out unwanted lines
+ filtered_lines = []
+ for line in lines:
+ should_ignore = False
+ for pattern in ignore_patterns:
+ if re.match(pattern, line, re.IGNORECASE):
+ should_ignore = True
+ break
+ if not should_ignore:
+ filtered_lines.append(line)
+
+ # Pattern to identify questions (starts with number followed by dot/parenthesis)
+ question_pattern = r'^\d+\s*[.)]\s*(.+)'
+
+ i = 0
+ while i < len(filtered_lines):
+ current_line = filtered_lines[i].strip()
+
+ # Check if current line is a question
+ question_match = re.match(question_pattern, current_line)
+ if question_match:
+ # This is a question
+ question_text = question_match.group(1).strip()
+ questions.append(f"{current_line}") # Keep the full question with number
+
+ # Look for the answer in the next line
+ if i + 1 < len(filtered_lines):
+ next_line = filtered_lines[i + 1].strip()
+ # If next line is not a question (doesn't start with number), it's likely an answer
+ if not re.match(question_pattern, next_line):
+ answers.append(next_line)
+ i += 2 # Skip both question and answer
+ else:
+ # Next line is also a question, so this question might not have an answer
+ # Or the answer might be embedded in the same line
+ # Try to extract answer from the question line itself if it contains common answer patterns
+ answers.append("") # Placeholder for missing answer
+ i += 1
+ else:
+ # Last line and it's a question without answer
+ answers.append("")
+ i += 1
+ else:
+ # This line doesn't match question pattern, skip it or try to pair it with previous question
+ if len(questions) > len(answers):
+ # We have more questions than answers, this might be an answer
+ answers.append(current_line)
+ i += 1
+
+ # Ensure we have equal number of questions and answers
+ while len(answers) < len(questions):
+ answers.append("")
+ while len(questions) < len(answers):
+ questions.append(f"Question {len(questions) + 1}")
+
+ return questions, answers
+
+@app.route('/process_question_paper', methods=['POST'])
+def process_question_paper():
+ global last_processed_question_paper_object
+
+ if 'file' not in request.files:
+ return jsonify({'error': 'No file provided'}), 400
+
+ file = request.files['file']
+ if file.filename == '':
+ return jsonify({'error': 'No file selected'}), 400
+
+ question_paper = QuestionPaper()
+
+ try:
+ # Create Images directory if it doesn't exist
+ images_dir = os.path.join(app.root_path, 'Images')
+ os.makedirs(images_dir, exist_ok=True)
+
+ if file.filename.lower().endswith('.pdf'):
+ question_paper_filename = "question_paper.pdf"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ # Initialize the global object with the path
+ question_paper.path = question_paper_path
+
+ # For PDF processing
+ images_from_pdf = convert_from_path(question_paper_path, poppler_path=r'C:\Program Files\poppler\Library\bin')
+
+ all_text = ""
+ for page_image in images_from_pdf:
+ text = pytesseract.image_to_string(page_image)
+ all_text += text + "\n"
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(all_text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ else:
+ # Process as image
+ question_paper_filename = "question_paper.png"
+ question_paper_path = os.path.join(images_dir, question_paper_filename)
+ file.save(question_paper_path)
+
+ question_paper.path = question_paper_path
+
+ image = Image.open(question_paper_path)
+ text = pytesseract.image_to_string(image)
+
+ # Use improved parsing
+ questions, answers = parse_question_paper_text(text)
+ question_paper.questions = questions
+ question_paper.answers = answers
+
+ # Clean the answers (remove any remaining unwanted patterns)
+ question_paper.clean_answers()
+
+ # Store the processed question paper globally
+ last_processed_question_paper_object = question_paper
+
+ return jsonify(question_paper.to_dict())
+
+ except Exception as e:
+ return jsonify({'error': str(e)}), 500
\ No newline at end of file
diff --git a/supabase_handler.py b/supabase_handler.py
new file mode 100644
index 0000000000000000000000000000000000000000..252c666b5bc0798342655da4beefe6f63c30e90a
--- /dev/null
+++ b/supabase_handler.py
@@ -0,0 +1,80 @@
+import os
+import uuid
+from datetime import datetime
+from supabase import create_client, Client
+from dotenv import load_dotenv
+
+load_dotenv()
+
+class SupabaseHandler:
+ def __init__(self):
+ url: str = os.getenv("SUPABASE_URL")
+ key: str = os.getenv("SUPABASE_ANON_KEY")
+ if not url or not key:
+ raise ValueError("SUPABASE_URL and SUPABASE_ANON_KEY must be set in environment variables")
+ self.supabase: Client = create_client(url, key)
+
+ def store_evaluation_result(self, teacher_email, evaluation_data):
+ """
+ Store evaluation result in Supabase with a unique key
+ Returns the unique key for retrieval
+ """
+ try:
+ # Generate unique key
+ unique_key = str(uuid.uuid4())
+
+ # Prepare data for storage
+ storage_data = {
+ "unique_key": unique_key,
+ "teacher_email": teacher_email,
+ "evaluation_data": evaluation_data,
+ "created_at": datetime.utcnow().isoformat(),
+ "total_students": evaluation_data.get("total_students", 0)
+ }
+
+ # Insert into Supabase
+ result = self.supabase.table("evaluation_results").insert(storage_data).execute()
+
+ if result.data:
+ print(f"Successfully stored evaluation result with key: {unique_key}")
+ return unique_key
+ else:
+ print("Failed to store evaluation result")
+ return None
+
+ except Exception as e:
+ print(f"Error storing evaluation result: {str(e)}")
+ return None
+
+ def get_evaluation_result(self, unique_key):
+ """
+ Retrieve evaluation result by unique key
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("*").eq("unique_key", unique_key).execute()
+
+ if result.data and len(result.data) > 0:
+ return result.data[0]
+ else:
+ return None
+
+ except Exception as e:
+ print(f"Error retrieving evaluation result: {str(e)}")
+ return None
+
+ def get_teacher_evaluations(self, teacher_email):
+ """
+ Get all evaluation results for a specific teacher
+ """
+ try:
+ result = self.supabase.table("evaluation_results").select("unique_key", "created_at", "total_students").eq("teacher_email", teacher_email).order("created_at", desc=True).execute()
+
+ if result.data:
+ return result.data
+ else:
+ return []
+
+ except Exception as e:
+ print(f"Error retrieving teacher evaluations: {str(e)}")
+ return []
+
diff --git a/test_endpoints.py b/test_endpoints.py
new file mode 100644
index 0000000000000000000000000000000000000000..03ae6de089292ac54acd9c12873773ad35fa8b7f
--- /dev/null
+++ b/test_endpoints.py
@@ -0,0 +1,236 @@
+"""
+Test script for OCR/OMR backend endpoints.
+Usage: python test_endpoints.py [base_url]
+Default base_url: http://localhost:5000
+"""
+
+import sys
+import os
+import requests
+
+BASE_URL = sys.argv[1] if len(sys.argv) > 1 else "http://localhost:5000"
+IMAGES_DIR = os.path.join(os.path.dirname(__file__), "Images")
+
+results = []
+# Track prerequisite passes for dependent tests
+prereqs = {
+ "question_paper": False,
+ "omr_answer_key": False,
+ "process_omr": False,
+}
+
+
+def report(name, passed, detail=""):
+ status = "PASS" if passed else "FAIL"
+ results.append((name, status))
+ msg = f"[{status}] {name}"
+ if detail:
+ msg += f" -- {detail}"
+ print(msg)
+ return passed
+
+
+def skip(name, reason):
+ results.append((name, "SKIP"))
+ print(f"[SKIP] {name} -- {reason}")
+
+
+# ---------------------------------------------------------------------------
+# 1. Health check
+# ---------------------------------------------------------------------------
+def test_health():
+ try:
+ r = requests.get(f"{BASE_URL}/health", timeout=10)
+ return report("/health", r.status_code == 200, f"status={r.status_code}")
+ except Exception as e:
+ return report("/health", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 2. Root endpoint
+# ---------------------------------------------------------------------------
+def test_root():
+ try:
+ r = requests.get(f"{BASE_URL}/", timeout=10)
+ return report("/ (root)", r.status_code == 200, f"status={r.status_code}")
+ except Exception as e:
+ return report("/ (root)", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 3. EasyOCR
+# ---------------------------------------------------------------------------
+def test_easyocr():
+ img = os.path.join(IMAGES_DIR, "test.jpg")
+ if not os.path.exists(img):
+ return report("/easyocr", False, "test.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/easyocr", files={"images": f}, timeout=60)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", keys={list(data.keys()) if isinstance(data, dict) else 'list'}"
+ return report("/easyocr", ok, detail)
+ except Exception as e:
+ return report("/easyocr", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 4. Tesseract
+# ---------------------------------------------------------------------------
+def test_tesseract():
+ img = os.path.join(IMAGES_DIR, "OCRSheet.jpg")
+ if not os.path.exists(img):
+ return report("/tesseract", False, "OCRSheet.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/tesseract", files={"images": f}, timeout=60)
+ ok = r.status_code == 200
+ return report("/tesseract", ok, f"status={r.status_code}")
+ except Exception as e:
+ return report("/tesseract", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 5. Process question paper (PDF)
+# ---------------------------------------------------------------------------
+def test_question_paper_pdf():
+ img = os.path.join(IMAGES_DIR, "question_paper.pdf")
+ if not os.path.exists(img):
+ return report("/process_question_paper (pdf)", False, "question_paper.pdf not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_question_paper", files={"file": f}, timeout=120)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", questions={len(data.get('questions', []))}"
+ prereqs["question_paper"] = True
+ return report("/process_question_paper (pdf)", ok, detail)
+ except Exception as e:
+ return report("/process_question_paper (pdf)", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 6. Process question paper (image)
+# ---------------------------------------------------------------------------
+def test_question_paper_png():
+ img = os.path.join(IMAGES_DIR, "question_paper.png")
+ if not os.path.exists(img):
+ return report("/process_question_paper (png)", False, "question_paper.png not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_question_paper", files={"file": f}, timeout=120)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", questions={len(data.get('questions', []))}"
+ prereqs["question_paper"] = True
+ return report("/process_question_paper (png)", ok, detail)
+ except Exception as e:
+ return report("/process_question_paper (png)", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 7. Process OMR answer key
+# ---------------------------------------------------------------------------
+def test_omr_answer_key():
+ img = os.path.join(IMAGES_DIR, "omr_answer_key.pdf")
+ if not os.path.exists(img):
+ return report("/process_omr_answer_key", False, "omr_answer_key.pdf not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_omr_answer_key", files={"file": f}, timeout=120)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ data = r.json()
+ detail += f", success={data.get('success')}"
+ prereqs["omr_answer_key"] = True
+ return report("/process_omr_answer_key", ok, detail)
+ except Exception as e:
+ return report("/process_omr_answer_key", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 8. Process OMR sheet
+# ---------------------------------------------------------------------------
+def test_process_omr():
+ img = os.path.join(IMAGES_DIR, "OMRSheet.jpg")
+ if not os.path.exists(img):
+ return report("/process_omr", False, "OMRSheet.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/process_omr", files={"images": f}, timeout=180)
+ ok = r.status_code == 200
+ detail = f"status={r.status_code}"
+ if ok:
+ prereqs["process_omr"] = True
+ return report("/process_omr", ok, detail)
+ except Exception as e:
+ return report("/process_omr", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 9. Evaluate OMR (depends on 7 + 8)
+# ---------------------------------------------------------------------------
+def test_evaluate_omr():
+ if not prereqs["omr_answer_key"] or not prereqs["process_omr"]:
+ skip("/evaluate_omr", "requires /process_omr_answer_key and /process_omr to pass first")
+ return
+ try:
+ r = requests.post(f"{BASE_URL}/evaluate_omr", timeout=120)
+ ok = r.status_code == 200
+ return report("/evaluate_omr", ok, f"status={r.status_code}")
+ except Exception as e:
+ return report("/evaluate_omr", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# 10. Evaluate answers (depends on 5/6)
+# ---------------------------------------------------------------------------
+def test_evaluate_answers():
+ if not prereqs["question_paper"]:
+ skip("/evaluate_answers", "requires /process_question_paper to pass first")
+ return
+ img = os.path.join(IMAGES_DIR, "test.jpg")
+ if not os.path.exists(img):
+ return report("/evaluate_answers", False, "test.jpg not found")
+ try:
+ with open(img, "rb") as f:
+ r = requests.post(f"{BASE_URL}/evaluate_answers", files={"student_answers": f}, timeout=120)
+ ok = r.status_code == 200
+ return report("/evaluate_answers", ok, f"status={r.status_code}")
+ except Exception as e:
+ return report("/evaluate_answers", False, str(e))
+
+
+# ---------------------------------------------------------------------------
+# Run all tests in order
+# ---------------------------------------------------------------------------
+if __name__ == "__main__":
+ print(f"\nTesting backend at: {BASE_URL}\n" + "=" * 60)
+
+ test_health()
+ test_root()
+ test_easyocr()
+ test_tesseract()
+ test_question_paper_pdf()
+ test_question_paper_png()
+ test_omr_answer_key()
+ test_process_omr()
+ test_evaluate_omr()
+ test_evaluate_answers()
+
+ # Summary
+ print("\n" + "=" * 60)
+ passed = sum(1 for _, s in results if s == "PASS")
+ failed = sum(1 for _, s in results if s == "FAIL")
+ skipped = sum(1 for _, s in results if s == "SKIP")
+ total = len(results)
+ print(f"Results: {passed}/{total} passed, {failed} failed, {skipped} skipped")
+ sys.exit(0 if failed == 0 else 1)
 +
+
 +
+
+
+ +
+

 +
+
+
+
 [](https://www.paypal.me/Udayraj123/500)
+
+_Find OMRChecker on_ [**_Product Hunt_**](https://www.producthunt.com/posts/omr-checker/) **|** [**_Reddit_**](https://www.reddit.com/r/computervision/comments/ccbj6f/omrchecker_grade_exams_using_python_and_opencv/) **|** [**Discord**](https://discord.gg/qFv2Vqf) **|** [**Linkedin**](https://www.linkedin.com/pulse/open-source-talks-udayraj-udayraj-deshmukh/) **|** [**goodfirstissue.dev**](https://goodfirstissue.dev/language/python) **|** [**codepeak.tech**](https://www.codepeak.tech/) **|** [**fossoverflow.dev**](https://fossoverflow.dev/projects) **|** [**Interview on Console by CodeSee**](https://console.substack.com/p/console-140) **|** [**Open Source Hub**](https://opensourcehub.io/udayraj123/omrchecker)
+
+
+
diff --git a/OMRChecker/docs/assets/colored_output.jpg b/OMRChecker/docs/assets/colored_output.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3cafa473b9a1134f845858be4cf0e98f3bd3bb7c
Binary files /dev/null and b/OMRChecker/docs/assets/colored_output.jpg differ
diff --git a/OMRChecker/inputs/OMRImage.jpg b/OMRChecker/inputs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c9e3dcccfc8af7c60becb3a4cc25f650861db98e
--- /dev/null
+++ b/OMRChecker/inputs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4439b6a479cb9fea92de9656b900a528052616ae8e8962a6c33b8e81ed7f327a
+size 267398
diff --git a/OMRChecker/inputs/template.json b/OMRChecker/inputs/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..823fe2e58a4182ef04d1fff99ecf146b5ec53aae
--- /dev/null
+++ b/OMRChecker/inputs/template.json
@@ -0,0 +1,37 @@
+{
+ "pageDimensions": [
+ 1122,
+ 1600
+ ],
+ "bubbleDimensions": [
+ 48,
+ 50
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 100,
+ 175
+ ],
+ "bubblesGap": 55,
+ "labelsGap": 67,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q02block": {
+ "origin": [
+ 100,
+ 845
+ ],
+ "bubblesGap": 58,
+ "labelsGap": 70,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ }
+
+}
\ No newline at end of file
diff --git a/OMRChecker/main.py b/OMRChecker/main.py
new file mode 100644
index 0000000000000000000000000000000000000000..19e3ebd43d4be2a096cf709b04817247f9952506
--- /dev/null
+++ b/OMRChecker/main.py
@@ -0,0 +1,99 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+
+import argparse
+import sys
+from pathlib import Path
+
+from src.entry import entry_point
+from src.logger import logger
+
+
+def parse_args():
+ # construct the argument parse and parse the arguments
+ argparser = argparse.ArgumentParser()
+
+ argparser.add_argument(
+ "-i",
+ "--inputDir",
+ default=["inputs"],
+ # https://docs.python.org/3/library/argparse.html#nargs
+ nargs="*",
+ required=False,
+ type=str,
+ dest="input_paths",
+ help="Specify an input directory.",
+ )
+
+ argparser.add_argument(
+ "-d",
+ "--debug",
+ required=False,
+ dest="debug",
+ action="store_false",
+ help="Enables debugging mode for showing detailed errors",
+ )
+
+ argparser.add_argument(
+ "-o",
+ "--outputDir",
+ default="outputs",
+ required=False,
+ dest="output_dir",
+ help="Specify an output directory.",
+ )
+
+ argparser.add_argument(
+ "-a",
+ "--autoAlign",
+ required=False,
+ dest="autoAlign",
+ action="store_true",
+ help="(experimental) Enables automatic template alignment - \
+ use if the scans show slight misalignments.",
+ )
+
+ argparser.add_argument(
+ "-l",
+ "--setLayout",
+ required=False,
+ dest="setLayout",
+ action="store_true",
+ help="Set up OMR template layout - modify your json file and \
+ run again until the template is set.",
+ )
+
+ (
+ args,
+ unknown,
+ ) = argparser.parse_known_args()
+
+ args = vars(args)
+
+ if len(unknown) > 0:
+ logger.warning(f"\nError: Unknown arguments: {unknown}", unknown)
+ argparser.print_help()
+ exit(11)
+ return args
+
+
+def entry_point_for_args(args):
+ if args["debug"] is True:
+ # Disable tracebacks
+ sys.tracebacklimit = 0
+ for root in args["input_paths"]:
+ entry_point(
+ Path(root),
+ args,
+ )
+
+
+if __name__ == "__main__":
+ args = parse_args()
+ entry_point_for_args(args)
diff --git a/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..d8d1a2325170014745edf3503b40e7c7bda160dc
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:f16c45dda1d02bf06e33a6d219c8282c363bf3943b94960eede1efe2bc6204d9
+size 408744
diff --git a/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3a968368fa191241dda21ea0c41f5d2d840f2e52
--- /dev/null
+++ b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:7e9678555042535c83ab920d0b616d4f99f9f08017629d46a15eb64d7c3abd3c
+size 407394
diff --git a/OMRChecker/outputs/Images/Manual/ErrorFiles.csv b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Results/Results_10PM.csv b/OMRChecker/outputs/Images/Results/Results_10PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..f3b83ec4b12f9e002ca0824ee43fc41f1f9ae30a
--- /dev/null
+++ b/OMRChecker/outputs/Images/Results/Results_10PM.csv
@@ -0,0 +1,2 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\Images\OMRSheet.jpg","outputs\Images\CheckedOMRs\OMRSheet.jpg","0","C","D","A","C","B","C","C","D","D","B","B","B","D","C","C","ABCD","B","ABCD","C","ABCD"
diff --git a/OMRChecker/outputs/Manual/ErrorFiles.csv b/OMRChecker/outputs/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..a9bb85c601677eefe49c6a75b39d5bd404506322
--- /dev/null
+++ b/OMRChecker/outputs/Manual/ErrorFiles.csv
@@ -0,0 +1,3 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
diff --git a/OMRChecker/outputs/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/pyproject.toml b/OMRChecker/pyproject.toml
new file mode 100644
index 0000000000000000000000000000000000000000..efa55e8fc8bf1623d8569abd406fef4d81270291
--- /dev/null
+++ b/OMRChecker/pyproject.toml
@@ -0,0 +1,18 @@
+[tool.black]
+exclude = '''
+(
+ /(
+ \.eggs # exclude a few common directories in the
+ | \.git # root of the project
+ | \.venv
+ | _build
+ | build
+ | dist
+ )/
+ | foo.py # also separately exclude a file named foo.py in
+ # the root of the project
+)
+'''
+include = '\.pyi?$'
+line-length = 88
+target-version = ['py37']
diff --git a/OMRChecker/pytest.ini b/OMRChecker/pytest.ini
new file mode 100644
index 0000000000000000000000000000000000000000..417c4a2dd810294aa9afaa9b4f73464fc9808c96
--- /dev/null
+++ b/OMRChecker/pytest.ini
@@ -0,0 +1,6 @@
+# pytest.ini
+[pytest]
+minversion = 7.0
+addopts = -qq --capture=no
+testpaths =
+ src/tests
diff --git a/OMRChecker/requirements.dev.txt b/OMRChecker/requirements.dev.txt
new file mode 100644
index 0000000000000000000000000000000000000000..fb31b5f80558481a3f2a5edbb741b363a1bd1089
--- /dev/null
+++ b/OMRChecker/requirements.dev.txt
@@ -0,0 +1,7 @@
+-r requirements.txt
+flake8>=6.0.0
+freezegun>=1.2.2
+pre-commit>=3.3.3
+pytest-mock>=3.11.1
+pytest>=7.4.0
+syrupy>=4.0.4
diff --git a/OMRChecker/requirements.txt b/OMRChecker/requirements.txt
new file mode 100644
index 0000000000000000000000000000000000000000..d0c3af875835b492a48ef689c3dc9335eb7cdca7
--- /dev/null
+++ b/OMRChecker/requirements.txt
@@ -0,0 +1,8 @@
+deepmerge>=1.1.0
+dotmap>=1.3.30
+jsonschema>=4.17.3
+matplotlib>=3.7.1
+numpy>=1.25.0
+pandas>=2.0.2
+rich>=13.4.2
+screeninfo>=0.8.1
diff --git a/OMRChecker/samples/answer-key/using-csv/adrian_omr.png b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png differ
diff --git a/OMRChecker/samples/answer-key/using-csv/answer_key.csv b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..acf09be611a66dfaeb7127e7249eadfbba6b0881
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
@@ -0,0 +1,5 @@
+q1,C
+q2,E
+q3,A
+q4,B
+q5,B
\ No newline at end of file
diff --git a/OMRChecker/samples/answer-key/using-csv/evaluation.json b/OMRChecker/samples/answer-key/using-csv/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/using-csv/template.json b/OMRChecker/samples/answer-key/using-csv/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/evaluation.json b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..09508b18e3490efcd68d6cb0b57a03d6e203f2ca
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
@@ -0,0 +1,35 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..5"
+ ],
+ "answers_in_order": [
+ "C",
+ "E",
+ [
+ "A",
+ "C"
+ ],
+ [
+ [
+ "B",
+ 2
+ ],
+ [
+ "C",
+ "3/2"
+ ]
+ ],
+ "C"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/answer-key/weighted-answers/template.json b/OMRChecker/samples/answer-key/weighted-answers/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..661d5f4fa73d7eea57e73837cf3862a25b9efcc3
Binary files /dev/null and b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg differ
diff --git a/OMRChecker/samples/community/Antibodyy/template.json b/OMRChecker/samples/community/Antibodyy/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..55e016e8eb402156d62f6553067b68abb37377ee
--- /dev/null
+++ b/OMRChecker/samples/community/Antibodyy/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 299,
+ 398
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 79
+ ],
+ "bubblesGap": 43,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "q1..6"
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-1.png b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
new file mode 100644
index 0000000000000000000000000000000000000000..5ed00d194a806d58c3db1dab7831796df7d4aaef
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:ed3024e5b709fbbdb1bcbef35dcfa50203ddb811f74b26e657d3f1b86c7e8c94
+size 381247
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-2.png b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d250db2c94980defb7ab78f9114ca56c1f223024
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:571109a06a09b792694a0ecac329283da199ac4495b1f10ffb7e5925c76a75c9
+size 130973
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-3.png b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
new file mode 100644
index 0000000000000000000000000000000000000000..c414be1c6fa93808d3419715e2e610884e233926
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:3976c705b45d889b06acc4bf82b27f129ad55c4875ee2aa6eb01aecf5a0250f6
+size 144032
diff --git a/OMRChecker/samples/community/Sandeep-1507/template.json b/OMRChecker/samples/community/Sandeep-1507/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bdb929ccd1a61c3ae54df4cdd6a8cf4281369733
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/template.json
@@ -0,0 +1,234 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 15,
+ 15
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Booklet_No": [
+ "b1..7"
+ ]
+ },
+ "fieldBlocks": {
+ "Booklet_No": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 112,
+ 530
+ ],
+ "fieldLabels": [
+ "b1..7"
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 28,
+ "labelsGap": 26.5
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 100
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 370
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q21..35"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 638
+ ]
+ },
+ "MCQBlock2a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 100
+ ]
+ },
+ "MCQBlock2a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 370
+ ]
+ },
+ "MCQBlock2a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q71..85"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 638
+ ]
+ },
+ "MCQBlock3a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q101..110"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 100
+ ]
+ },
+ "MCQBlock3a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q111..120"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 370
+ ]
+ },
+ "MCQBlock3a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q121..135"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 638
+ ]
+ },
+ "MCQBlock4a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q151..160"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 100
+ ]
+ },
+ "MCQBlock4a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q161..170"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 370
+ ]
+ },
+ "MCQBlock4a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q171..185"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 638
+ ]
+ },
+ "MCQBlock1a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q36..50"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 480,
+ 1061
+ ]
+ },
+ "MCQBlock2a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q86..100"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 648,
+ 1061
+ ]
+ },
+ "MCQBlock3a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q136..150"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 1061
+ ]
+ },
+ "MCQBlock4a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q186..200"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.6,
+ "origin": [
+ 986,
+ 1061
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Shamanth/omr_sheet_01.png b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
new file mode 100644
index 0000000000000000000000000000000000000000..7c317db90f80fdd8d3dcfbbadcb1f9b764e619e3
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:9dd72b86ce9c6544f4a628187b4021df5746e9579222e7598c4d88846f2fbeb3
+size 107647
diff --git a/OMRChecker/samples/community/Shamanth/template.json b/OMRChecker/samples/community/Shamanth/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..7d8c34d5e4b7af5ca94415b5096c82300156bcbc
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/template.json
@@ -0,0 +1,25 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 78,
+ 41
+ ],
+ "fieldLabels": [
+ "q21..28"
+ ],
+ "bubblesGap": 56,
+ "labelsGap": 46
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/answer_key.jpg b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..9d39b263e8fe5c23e391ad39bd0905efbf976038
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:13bb31818680ca8497ee4c640a15d4acab2048cf11c973d758c8c75fd71c73be
+size 291737
diff --git a/OMRChecker/samples/community/UPSC-mock/config.json b/OMRChecker/samples/community/UPSC-mock/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..388060a33b04db4b550d0efecafd32aa277355d5
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 1800,
+ "display_width": 2400,
+ "processing_height": 2400,
+ "processing_width": 1800
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/evaluation.json b/OMRChecker/samples/community/UPSC-mock/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..3271cbd5e39161d428d765145f34d811701bdc64
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/evaluation.json
@@ -0,0 +1,18 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "answer_key_image_path": "answer_key.jpg",
+ "questions_in_order": [
+ "q1..100"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "2",
+ "incorrect": "-2/3",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..5dd9ae6adbd327c7995bfcd5f53989002e240cf2
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:020705f4618fd55a72dbc35719565307b844dbd28838c790fe858c3ebe5fb36e
+size 160882
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c696b2e2d39118dec97d4f4c7479bfbc2dc6fc66
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1bb28daa10d276768d4fe3fd3f162ff8d3556cb805c4cf3f362e84d00d808095
+size 182385
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..edb7bbb600db8175e5beaa707c58d5a95972ec1e
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:695e6cc5a497bcf114a101533ee0464a223ecb8f914bc28a8d356533bcf6e341
+size 179112
diff --git a/OMRChecker/samples/community/UPSC-mock/template.json b/OMRChecker/samples/community/UPSC-mock/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..2fadab79ca7f7f9adb03a85cde30f1ba2ccca588
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/template.json
@@ -0,0 +1,195 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 30,
+ 25
+ ],
+ "customLabels": {
+ "Subject Code": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "Roll": [
+ "roll1..10"
+ ]
+ },
+ "fieldBlocks": {
+ "bookletNo": {
+ "origin": [
+ 595,
+ 545
+ ],
+ "bubblesGap": 68,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "bookletNo"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "C",
+ "D"
+ ],
+ "direction": "vertical"
+ },
+ "subjectCode": {
+ "origin": [
+ 912,
+ 512
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.5,
+ "fieldLabels": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "roll": {
+ "origin": [
+ 1200,
+ 510
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.8,
+ "fieldLabels": [
+ "roll1..10"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "q01block": {
+ "origin": [
+ 500,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 500,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 495,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1125,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1125,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..ffd67e704a23baa49bec476e04ba6b1796ff9b52
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
@@ -0,0 +1,200 @@
+q1,C
+q2,C
+q3,"D,E"
+q4,"A,AB"
+q5,"[['A', '1'], ['B', '2']]"
+q6,"['A', 'B']"
+q7,C
+q8,D
+q9,B
+q10,B
+q11,A
+q12,A
+q13,C
+q14,B
+q15,D
+q16,B
+q17,C
+q18,A
+q19,B
+q20,D
+q21,D
+q22,C
+q23,A
+q24,C
+q25,D
+q26,C
+q27,C
+q28,B
+q29,A
+q30,D
+q31,C
+q32,B
+q33,B
+q34,C
+q35,A
+q36,D
+q37,C
+q38,B
+q39,C
+q40,A
+q41,A
+q42,C
+q43,D
+q44,D
+q45,B
+q46,C
+q47,C
+q48,A
+q49,C
+q50,B
+q51,B
+q52,C
+q53,D
+q54,C
+q55,B
+q56,B
+q57,A
+q58,A
+q59,D
+q60,C
+q61,C
+q62,B
+q63,A
+q64,C
+q65,D
+q66,C
+q67,B
+q68,A
+q69,B
+q70,B
+q71,C
+q72,B
+q73,C
+q74,A
+q75,A
+q76,C
+q77,D
+q78,D
+q79,B
+q80,A
+q81,B
+q82,C
+q83,D
+q84,C
+q85,A
+q86,C
+q87,D
+q88,B
+q89,C
+q90,B
+q91,B
+q92,A
+q93,C
+q94,D
+q95,C
+q96,B
+q97,B
+q98,A
+q99,A
+q100,A
+q101,A
+q102,B
+q103,C
+q104,C
+q105,A
+q106,D
+q107,B
+q108,A
+q109,C
+q110,B
+q111,B
+q112,C
+q113,C
+q114,B
+q115,D
+q116,B
+q117,A
+q118,C
+q119,D
+q120,C
+q121,C
+q122,A
+q123,B
+q124,C
+q125,D
+q126,C
+q127,C
+q128,D
+q129,D
+q130,A
+q131,A
+q132,C
+q133,B
+q134,C
+q135,D
+q136,B
+q137,C
+q138,A
+q139,B
+q140,D
+q141,D
+q142,C
+q143,D
+q144,A
+q145,A
+q146,C
+q147,A
+q148,D
+q149,D
+q150,B
+q151,A
+q152,B
+q153,B
+q154,D
+q155,D
+q156,B
+q157,A
+q158,B
+q159,A
+q160,C
+q161,D
+q162,C
+q163,A
+q164,B
+q165,D
+q166,D
+q167,C
+q168,C
+q169,C
+q170,D
+q171,A
+q172,A
+q173,C
+q174,C
+q175,B
+q176,D
+q177,A
+q178,B
+q179,B
+q180,C
+q181,D
+q182,C
+q183,B
+q184,B
+q185,C
+q186,D
+q187,D
+q188,A
+q189,A
+q190,B
+q191,C
+q192,B
+q193,D
+q194,C
+q195,B
+q196,B
+q197,A
+q198,B
+q199,B
+q200,A
diff --git a/OMRChecker/samples/community/UmarFarootAPS/config.json b/OMRChecker/samples/community/UmarFarootAPS/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..5a9d75731b7f5ae8c6fd803a7426b29ac39c6bfb
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 960,
+ "display_width": 1280,
+ "processing_height": 1640,
+ "processing_width": 1332
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/evaluation.json b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg differ
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..73d60b9caa9860aa6a9bb75956fd3430b1aee44e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:49e1a14e0c1be809282b6e43c913e17598380a33f607a5c46723764c32e17b59
+size 304722
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..25d75107cf4885f2974c52cf42003255be285a3e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18b85df487c0bc98eb9139949c3fbca2bad57e8f224fe48b495e0951660f65d
+size 237392
diff --git a/OMRChecker/samples/community/UmarFarootAPS/template.json b/OMRChecker/samples/community/UmarFarootAPS/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..f303edabdf3bc64c41c50152ae72d09f5a0de913
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 2550,
+ 3300
+ ],
+ "bubbleDimensions": [
+ 32,
+ 32
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll_no": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ]
+ },
+ "fieldBlocks": {
+ "Roll_no": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 2169,
+ 180
+ ],
+ "fieldLabels": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ],
+ "bubblesGap": 61,
+ "labelsGap": 93
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q1..17"
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q18..34"
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q35..50"
+ ]
+ },
+ "MCQBlock1a4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q51..67"
+ ]
+ },
+ "MCQBlock1a5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q68..84"
+ ]
+ },
+ "MCQBlock1a6": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q85..100"
+ ]
+ },
+ "MCQBlock1a7": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q101..117"
+ ]
+ },
+ "MCQBlock1a8": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q118..134"
+ ]
+ },
+ "MCQBlock1a9": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q135..150"
+ ]
+ },
+ "MCQBlock1a10": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q151..167"
+ ]
+ },
+ "MCQBlock1a11": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q168..184"
+ ]
+ },
+ "MCQBlock1a12": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q185..200"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..b9d70c6be1aa35398e2ac125f3e7bea0ff924fd0
Binary files /dev/null and b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg differ
diff --git a/OMRChecker/samples/community/dxuian/template.json b/OMRChecker/samples/community/dxuian/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e055f557537fa531a0546c2155146763c8d937a6
--- /dev/null
+++ b/OMRChecker/samples/community/dxuian/template.json
@@ -0,0 +1,48 @@
+{
+ "pageDimensions": [707, 484],
+ "bubbleDimensions": [15, 10],
+ "fieldBlocks": {
+ "Column1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [82, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q1", "Q2", "Q3", "Q4", "Q5", "Q6", "Q7", "Q8", "Q9", "Q10", "Q11", "Q12", "Q13", "Q14", "Q15", "Q16", "Q17", "Q18", "Q19", "Q20"]
+ },
+ "Column2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [205, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q21", "Q22", "Q23", "Q24", "Q25", "Q26", "Q27", "Q28", "Q29", "Q30", "Q31", "Q32", "Q33", "Q34", "Q35", "Q36", "Q37", "Q38", "Q39", "Q40"]
+ },
+ "Column3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [327, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q41", "Q42", "Q43", "Q44", "Q45", "Q46", "Q47", "Q48", "Q49", "Q50", "Q51", "Q52", "Q53", "Q54", "Q55", "Q56", "Q57", "Q58", "Q59", "Q60"]
+ },
+ "Column4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [450, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q61", "Q62", "Q63", "Q64", "Q65", "Q66", "Q67", "Q68", "Q69", "Q70", "Q71", "Q72", "Q73", "Q74", "Q75", "Q76", "Q77", "Q78", "Q79", "Q80"]
+ },
+ "Column5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [573, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q81", "Q82", "Q83", "Q84", "Q85", "Q86", "Q87", "Q88", "Q89", "Q90", "Q91", "Q92", "Q93", "Q94", "Q95", "Q96", "Q97", "Q98", "Q99", "Q100"]
+ }
+ },
+
+ "emptyValue": "-"
+}
\ No newline at end of file
diff --git a/OMRChecker/samples/community/ibrahimkilic/template.json b/OMRChecker/samples/community/ibrahimkilic/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..3064ee19192fc7474370c0f518a51bea9a30b639
--- /dev/null
+++ b/OMRChecker/samples/community/ibrahimkilic/template.json
@@ -0,0 +1,30 @@
+{
+ "pageDimensions": [
+ 299,
+ 328
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "emptyValue": "no",
+ "fieldBlocks": {
+ "YesNoBlock1": {
+ "direction": "horizontal",
+ "bubbleValues": [
+ "yes"
+ ],
+ "origin": [
+ 15,
+ 55
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 48,
+ "labelsGap": 48,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..e9436f41ee8db1269e2255ee902b73de2525d502
Binary files /dev/null and b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg differ
diff --git a/OMRChecker/samples/community/samuelIkoli/template.json b/OMRChecker/samples/community/samuelIkoli/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..19c46f7c96bb07cc0a5a1731bbb8febeba9686e9
--- /dev/null
+++ b/OMRChecker/samples/community/samuelIkoli/template.json
@@ -0,0 +1,28 @@
+{
+ "pageDimensions": [630, 404],
+ "bubbleDimensions": [20, 15],
+ "customLabels": {},
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [33, 6],
+ "fieldLabels": ["q1", "q2", "q3", "q4", "q5", "q6", "q7", "q8", "q9", "q10", "q11", "q12", "q13", "q14", "q15", "q16", "q17", "q18", "q19", "q20"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock2": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [248, 6],
+ "fieldLabels": ["q21", "q22", "q23", "q24", "q25", "q26", "q27", "q28", "q29", "q30", "q31", "q32", "q33", "q34", "q35", "q36", "q37", "q38", "q39", "q40"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock3": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [465, 6],
+ "fieldLabels": ["q41", "q42", "q43", "q44", "q45", "q46", "q47", "q48", "q49", "q50", "q51", "q52", "q53", "q54", "q55", "q56", "q57", "q58", "q59", "q60"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg
new file mode 100644
index 0000000000000000000000000000000000000000..fcd61e808e4a2f33095ea0dbef0ec27741450465
Binary files /dev/null and b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg differ
diff --git a/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6626eb476d38cb16156dc15a48d30bcd41ebaac1
--- /dev/null
+++ b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1be7b0fc432e811fd07b07fe325d9ca8d0d97782eea9f95ed62ff0e47076e1cd
+size 162012
diff --git a/OMRChecker/samples/sample1/config.json b/OMRChecker/samples/sample1/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample1/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample1/omr_marker.jpg b/OMRChecker/samples/sample1/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample1/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample1/template.json b/OMRChecker/samples/sample1/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e4795ad56c919618354f18d4de7a14f14642df07
--- /dev/null
+++ b/OMRChecker/samples/sample1/template.json
@@ -0,0 +1,197 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q5": [
+ "q5_1",
+ "q5_2"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubblesGap": 41,
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "fieldLabels": [
+ "Medium"
+ ],
+ "labelsGap": 0,
+ "origin": [
+ 170,
+ 282
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "roll1..9"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 58,
+ "origin": [
+ 225,
+ 282
+ ]
+ },
+ "Int_Block_Q5": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q5_1",
+ "q5_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 903,
+ 282
+ ]
+ },
+ "Int_Block_Q6": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1077,
+ 282
+ ]
+ },
+ "Int_Block_Q7": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1240,
+ 282
+ ]
+ },
+ "Int_Block_Q8": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1410,
+ 282
+ ]
+ },
+ "Int_Block_Q9": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1580,
+ 282
+ ]
+ },
+ "MCQ_Block_Q1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..4"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 860
+ ]
+ },
+ "MCQ_Block_Q10": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q10..13"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 1195
+ ]
+ },
+ "MCQ_Block_Q14": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q14..16"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 860
+ ]
+ },
+ "MCQ_Block_Q17": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q17..20"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 1195
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ },
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/sample2/config.json b/OMRChecker/samples/sample2/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample2/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample2/template.json b/OMRChecker/samples/sample2/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/sample2/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample3/README.md b/OMRChecker/samples/sample3/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..4b1e325dfa284cf676a1f3d9e37d426f5d20ebb6
--- /dev/null
+++ b/OMRChecker/samples/sample3/README.md
@@ -0,0 +1,31 @@
+## Observation
+The OMR layout is slightly different on colored thick papers vs on xeroxed thin papers. The shift becomes noticible in case of OMR with large number of questions.
+
+We overlapped a colored OMR sheet with a xerox copy of the same OMR sheet(both printed on A4 papers) and observed that there is a great amount of layout sheet as we reach the bottom of the OMR.
+
+Link to an explainer with a real life example: [Google Drive](https://drive.google.com/drive/folders/1GpZTmpEhEjSALJEMjHwDafzWKgEoCTOI?usp=sharing)
+
+## Reasons for shifts in Template layout:
+Listing out a few reasons for the above observation:
+### Printer margin setting
+The margin settings for different printers may be different for the same OMR layout. Thus causing the print to become elongated either horizontally, vertically or both ways.
+
+### The Fan-out effect
+The fan-out effect is usually observed in a sheet fed offset press. Depending on how the papers are made, their dimensions have a tendency to change when they are exposed to moisture or water.
+
+The standard office papers(80 gsm) can easily capture moisture and change shape e.g. in case they get stored for multiple days in a place where the weather is highly humid.
+
+Below are some examples of the GSM ranges:
+
+- 74gsm to 90gsm – This is the basic standard office paper, used in your laser printers.
+- 100gsm to 120gsm – This is stationary paper used for standard letterheads, complimentary slips.
+- 130 to 170gsm – Mostly used for leaflets, posters, single-sided flyers, and brochures.
+
+## Solution
+
+It is recommended to scan each types of prints into different folders and use a separate template.json layout for each of the folders. The same is presented in this sample folder.
+
+## References
+
+- [Paper dimensional stability in sheet-fed offset printing](https://www.diva-portal.org/smash/get/diva2:517895/FULLTEXT01.pdf)
+- An analysis of a few ["Interesting" bubble sheets](https://imgur.com/a/10qwL)
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..4877db1feb1c758bc35b41f543dc9bb14d6ca62f
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18eba01e5d0c466b4b750aa20e710e7354489ed9bef91966c2349007fee349d
+size 137681
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/template.json b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ab35b30f1e3ec35d1c4698cc68b76c4a47d6eb37
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 504,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 504,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 500,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1120,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1120,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample3/config.json b/OMRChecker/samples/sample3/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample3/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..2be38bbfb8e8075f0da00d5fb56d4689d2e72e26
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:eab4df0efac2614b2deddcadc0b754aa72c668170e5d9af49ad77c877198556c
+size 187367
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd499ade7e54d5061817985e40effb13fce8f012
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 492,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 492,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 492,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 487,
+ 1920
+ ],
+ "bubblesGap": 61.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 807,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 803,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 803,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 803,
+ 1920
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1115,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1115,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample4/IMG_20201116_143512.jpg b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6c817d4f620b6bc728eeb410bd9c1da8d27a5cfe
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0b1f0e0afd8a2c40deb7f7c992c9fb6046eb1b28a5f9a674e80ce3418c7fc307
+size 421479
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..023f66bc9eb9100a8500d84b82f7725141962082
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8c3f847e6c37c96562ee17b3d69cb919bb52743f8b95d02679ed62e20dbd5096
+size 386960
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..21869f5c27011e067f1fd8cb3a4e6ecb0e4c7e11
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0bdc5eca48c13d19f228ef7252b5e855b5c62926ba1e9e3297fc6a46e074b1be
+size 307198
diff --git a/OMRChecker/samples/sample4/config.json b/OMRChecker/samples/sample4/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..6c82782a3f6486145d6367eabe6910cca5268214
--- /dev/null
+++ b/OMRChecker/samples/sample4/config.json
@@ -0,0 +1,13 @@
+{
+ "dimensions": {
+ "display_width": 1189,
+ "display_height": 1682
+ },
+ "threshold_params": {
+ "MIN_JUMP": 30
+ },
+ "outputs": {
+ "filter_out_multimarked_files": false,
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample4/evaluation.json b/OMRChecker/samples/sample4/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..8ea935e70ded2a9c53c13bf22b5904f1ef3fa923
--- /dev/null
+++ b/OMRChecker/samples/sample4/evaluation.json
@@ -0,0 +1,34 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..11"
+ ],
+ "answers_in_order": [
+ "B",
+ "D",
+ "C",
+ "B",
+ "D",
+ "C",
+ [
+ "B",
+ "C",
+ "BC"
+ ],
+ "A",
+ "C",
+ "D",
+ "C"
+ ],
+ "should_explain_scoring": true,
+ "enable_evaluation_table_to_csv": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample4/template.json b/OMRChecker/samples/sample4/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..d1bef2985939f8ea9c01bfdec28e0d5b6bab11a3
--- /dev/null
+++ b/OMRChecker/samples/sample4/template.json
@@ -0,0 +1,45 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 30,
+ 30
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ },
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 134,
+ 684
+ ],
+ "fieldLabels": [
+ "q1..11"
+ ],
+ "bubblesGap": 79,
+ "labelsGap": 62
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/README.md b/OMRChecker/samples/sample5/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..b70cedd6cbf22de260649c08e855d929d5e8ae87
--- /dev/null
+++ b/OMRChecker/samples/sample5/README.md
@@ -0,0 +1,6 @@
+### OMRChecker Sample
+
+This sample demonstrates multiple things, namely -
+- Running OMRChecker on images scanned using popular document scanning apps
+- Using a common template.json file for sub-folders (e.g. multiple scan batches)
+- Using evaluation.json file with custom marking (without streak-based marking)
diff --git a/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..72b450e07dd7d4c4a18287183128ad7edbf5607a
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:08bb2e9fecf23be6850485cb7776a604aa2efe9800855ea763dee7ade5362323
+size 127651
diff --git a/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..46fb2ac36c0d5c2366fdf9a7ed56eb933578a60f
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0da09483868ad499925fefa284153e9544822894335e16cea859057740c3c063
+size 127863
diff --git a/OMRChecker/samples/sample5/config.json b/OMRChecker/samples/sample5/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample5/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample5/evaluation.json b/OMRChecker/samples/sample5/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..706e49ee9d9e7d445ca98ea87a1a140ce53da54d
--- /dev/null
+++ b/OMRChecker/samples/sample5/evaluation.json
@@ -0,0 +1,93 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..22"
+ ],
+ "answers_in_order": [
+ "C",
+ "C",
+ "B",
+ "C",
+ "C",
+ [
+ "1",
+ "01"
+ ],
+ "19",
+ "10",
+ "10",
+ "18",
+ "D",
+ "A",
+ "D",
+ "D",
+ "D",
+ "C",
+ "C",
+ "C",
+ "C",
+ "D",
+ "B",
+ "A"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ },
+ "BOOMERANG_1": {
+ "questions": [
+ "q1..5"
+ ],
+ "marking": {
+ "correct": 4,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "PROXIMITY_1": {
+ "questions": [
+ "q6..10"
+ ],
+ "marking": {
+ "correct": 3,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_1": {
+ "questions": [
+ "q11..14"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "POWER_SECTION_1": {
+ "questions": [
+ "q15..18"
+ ],
+ "marking": {
+ "correct": 1,
+ "incorrect": 0,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_2": {
+ "questions": [
+ "q19..22"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/omr_marker.jpg b/OMRChecker/samples/sample5/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample5/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample5/template.json b/OMRChecker/samples/sample5/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ac2436c02a8a19024e8050ff7c02dca9aa7ea016
--- /dev/null
+++ b/OMRChecker/samples/sample5/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ],
+ "q10": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "origin": [
+ 200,
+ 215
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "Medium"
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 261,
+ 210
+ ],
+ "bubblesGap": 46.5,
+ "labelsGap": 58,
+ "fieldLabels": [
+ "roll1..9"
+ ]
+ },
+ "Int1": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 935,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ]
+ },
+ "Int2": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1100,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ]
+ },
+ "Int3": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1275,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ]
+ },
+ "Int4": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1449,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "Int5": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1620,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "Mcq1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 198,
+ 826
+ ],
+ "bubblesGap": 93,
+ "labelsGap": 62,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ },
+ "Mcq2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 830
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q11..14"
+ ]
+ },
+ "Mcq3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 1270
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q15..18"
+ ]
+ },
+ "Mcq4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1481,
+ 830
+ ],
+ "bubblesGap": 73,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q19..22"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/config.json b/OMRChecker/samples/sample6/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..2ac30e718a44f7336aafda1b0f57aec1320fc72c
--- /dev/null
+++ b/OMRChecker/samples/sample6/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_width": 2480,
+ "display_height": 3508,
+ "processing_width": 1653,
+ "processing_height": 2339
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..fdaa0f4112e15037a31bc90da204b7c19114820b
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:de8a5ed5b7360a71db83b49a3d0d6e1af1482339d1116286a064d285be42e37e
+size 246590
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..bf56d121c1061302fc4ce87fca61bb8584ea9e4a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:aca51de61ba3ee76e00e22a666d8cd2b72190af674fa051c9cf70346e9ee4d0a
+size 244049
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..22bfa55de7a687729b733a615ad78dad64ac467a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:950dbf86fc152f464be30ebaeb518c61067b0c494ab5fd70f05de10b5dfabb12
+size 244321
diff --git a/OMRChecker/samples/sample6/readme.md b/OMRChecker/samples/sample6/readme.md
new file mode 100644
index 0000000000000000000000000000000000000000..d8491b2a049eb4e721e007e8e8bc4fe5ca536802
--- /dev/null
+++ b/OMRChecker/samples/sample6/readme.md
@@ -0,0 +1,21 @@
+# Demo for feature-based alignment
+
+## Background
+OMR is used to match student roll on final exam scripts. Scripts are scanned using a document scanner and the cover pages are extracted for OMR. Even though a document scanner does not produce any warpped perspective, the alignment is not perfect, causing some rotation and translation in the scans.
+
+The scripts in this sample were specifically selected incorrectly marked scripts to demonstrate how feature-based alignment can correct transformation errors using a reference image. In the actual batch. 156 out of 532 scripts were incorrectly marked. With feature-based alignment, all scripts were correctly marked.
+
+## Usage
+Two template files are given in the sample folder, one with feature-based alignment (template_fb_align), the other without (template_no_fb_align).
+
+## Additional Notes
+
+### Reference Image
+When using a reference image for feature-based alignment, it is better not to have many repeated patterns as it is causes ambiguity when trying to match similar feature points. The bubbles in an OMR form are identical and should not be used for feature-extraction.
+
+Thus, the reference image should be cleared of any bubbles. Forms with lots of text as in this example would be effective.
+
+Note the reference image in this sample was generated from a vector pdf, and not from a scanned blank, producing in a perfectly aligned reference.
+
+### Level adjustment
+The bubbles on the scripts were not shaded dark enough. Thus, a level adjustment was done to bring the black point to 70% to darken the light shading. White point was brought down to 80% to remove the light-grey background in the columns.
diff --git a/OMRChecker/samples/sample6/reference.png b/OMRChecker/samples/sample6/reference.png
new file mode 100644
index 0000000000000000000000000000000000000000..e2e692f4638e9c7fa2d7d159e044c778b18acdc9
--- /dev/null
+++ b/OMRChecker/samples/sample6/reference.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8379ba055209c065def041fc1fb1cea444b913d835c1cfeffa40091446e5e3e5
+size 178069
diff --git a/OMRChecker/samples/sample6/template.json b/OMRChecker/samples/sample6/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_fb_align.json b/OMRChecker/samples/sample6/template_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd45b9c8fd3ba024b951ff4e7965c98bb56211ee
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_fb_align.json
@@ -0,0 +1,118 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "FeatureBasedAlignment",
+ "options": {
+ "reference": "reference.png",
+ "maxFeatures": 1000,
+ "2d": true
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_no_fb_align.json b/OMRChecker/samples/sample6/template_no_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_no_fb_align.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/src/__init__.py b/OMRChecker/src/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..5c1baacaa53bb1b4219d33eee566c6a613ab88d6
--- /dev/null
+++ b/OMRChecker/src/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+from src.logger import logger
+
+# It takes a few seconds for the imports
+logger.info(f"Loading OMRChecker modules...")
diff --git a/OMRChecker/src/constants.py b/OMRChecker/src/constants.py
new file mode 100644
index 0000000000000000000000000000000000000000..628eb75ac892ddc184cc99429036aa9196889c83
--- /dev/null
+++ b/OMRChecker/src/constants.py
@@ -0,0 +1,54 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+from dotmap import DotMap
+
+# Filenames
+TEMPLATE_FILENAME = "template.json"
+EVALUATION_FILENAME = "evaluation.json"
+CONFIG_FILENAME = "config.json"
+
+FIELD_LABEL_NUMBER_REGEX = r"([^\d]+)(\d*)"
+#
+ERROR_CODES = DotMap(
+ {
+ "MULTI_BUBBLE_WARN": 1,
+ "NO_MARKER_ERR": 2,
+ },
+ _dynamic=False,
+)
+
+FIELD_TYPES = {
+ "QTYPE_INT": {
+ "bubbleValues": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"],
+ "direction": "vertical",
+ },
+ "QTYPE_INT_FROM_1": {
+ "bubbleValues": ["1", "2", "3", "4", "5", "6", "7", "8", "9", "0"],
+ "direction": "vertical",
+ },
+ "QTYPE_MCQ4": {"bubbleValues": ["A", "B", "C", "D"], "direction": "horizontal"},
+ "QTYPE_MCQ5": {
+ "bubbleValues": ["A", "B", "C", "D", "E"],
+ "direction": "horizontal",
+ },
+ #
+ # You can create and append custom field types here-
+ #
+}
+
+# TODO: move to interaction.py
+TEXT_SIZE = 0.95
+CLR_BLACK = (50, 150, 150)
+CLR_WHITE = (250, 250, 250)
+CLR_GRAY = (130, 130, 130)
+CLR_DARK_GRAY = (100, 100, 100)
+
+# TODO: move to config.json
+GLOBAL_PAGE_THRESHOLD_WHITE = 200
+GLOBAL_PAGE_THRESHOLD_BLACK = 100
diff --git a/OMRChecker/src/core.py b/OMRChecker/src/core.py
new file mode 100644
index 0000000000000000000000000000000000000000..15279cf49fb9fb12c88611bcc69611ea524344dc
--- /dev/null
+++ b/OMRChecker/src/core.py
@@ -0,0 +1,721 @@
+import os
+from collections import defaultdict
+from typing import Any
+
+import cv2
+import matplotlib.pyplot as plt
+import numpy as np
+
+import src.constants as constants
+from src.logger import logger
+from src.utils.image import CLAHE_HELPER, ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class ImageInstanceOps:
+ """Class to hold fine-tuned utilities for a group of images. One instance for each processing directory."""
+
+ save_img_list: Any = defaultdict(list)
+
+ def __init__(self, tuning_config):
+ super().__init__()
+ self.tuning_config = tuning_config
+ self.save_image_level = tuning_config.outputs.save_image_level
+
+ def apply_preprocessors(self, file_path, in_omr, template):
+ tuning_config = self.tuning_config
+ # resize to conform to template
+ in_omr = ImageUtils.resize_util(
+ in_omr,
+ tuning_config.dimensions.processing_width,
+ tuning_config.dimensions.processing_height,
+ )
+
+ # run pre_processors in sequence
+ for pre_processor in template.pre_processors:
+ in_omr = pre_processor.apply_filter(in_omr, file_path)
+ return in_omr
+
+ def read_omr_response(self, template, image, name, save_dir=None):
+ config = self.tuning_config
+ auto_align = config.alignment_params.auto_align
+ try:
+ img = image.copy()
+ # origDim = img.shape[:2]
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ if img.max() > img.min():
+ img = ImageUtils.normalize_util(img)
+ # Processing copies
+ transp_layer = img.copy()
+ final_marked = img.copy()
+
+ morph = img.copy()
+ self.append_save_img(3, morph)
+
+ if auto_align:
+ # Note: clahe is good for morphology, bad for thresholding
+ morph = CLAHE_HELPER.apply(morph)
+ self.append_save_img(3, morph)
+ # Remove shadows further, make columns/boxes darker (less gamma)
+ morph = ImageUtils.adjust_gamma(
+ morph, config.threshold_params.GAMMA_LOW
+ )
+ # TODO: all numbers should come from either constants or config
+ _, morph = cv2.threshold(morph, 220, 220, cv2.THRESH_TRUNC)
+ morph = ImageUtils.normalize_util(morph)
+ self.append_save_img(3, morph)
+ if config.outputs.show_image_level >= 4:
+ InteractionUtils.show("morph1", morph, 0, 1, config)
+
+ # Move them to data class if needed
+ # Overlay Transparencies
+ alpha = 0.65
+ omr_response = {}
+ multi_marked, multi_roll = 0, 0
+
+ # TODO Make this part useful for visualizing status checks
+ # blackVals=[0]
+ # whiteVals=[255]
+
+ if config.outputs.show_image_level >= 5:
+ all_c_box_vals = {"int": [], "mcq": []}
+ # TODO: simplify this logic
+ q_nums = {"int": [], "mcq": []}
+
+ # Find Shifts for the field_blocks --> Before calculating threshold!
+ if auto_align:
+ # print("Begin Alignment")
+ # Open : erode then dilate
+ v_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (2, 10))
+ morph_v = cv2.morphologyEx(
+ morph, cv2.MORPH_OPEN, v_kernel, iterations=3
+ )
+ _, morph_v = cv2.threshold(morph_v, 200, 200, cv2.THRESH_TRUNC)
+ morph_v = 255 - ImageUtils.normalize_util(morph_v)
+
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morphed_vertical", morph_v, 0, 1, config=config
+ )
+
+ # InteractionUtils.show("morph1",morph,0,1,config=config)
+ # InteractionUtils.show("morphed_vertical",morph_v,0,1,config=config)
+
+ self.append_save_img(3, morph_v)
+
+ morph_thr = 60 # for Mobile images, 40 for scanned Images
+ _, morph_v = cv2.threshold(morph_v, morph_thr, 255, cv2.THRESH_BINARY)
+ # kernel best tuned to 5x5 now
+ morph_v = cv2.erode(morph_v, np.ones((5, 5), np.uint8), iterations=2)
+
+ self.append_save_img(3, morph_v)
+ # h_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (10, 2))
+ # morph_h = cv2.morphologyEx(morph, cv2.MORPH_OPEN, h_kernel, iterations=3)
+ # ret, morph_h = cv2.threshold(morph_h,200,200,cv2.THRESH_TRUNC)
+ # morph_h = 255 - normalize_util(morph_h)
+ # InteractionUtils.show("morph_h",morph_h,0,1,config=config)
+ # _, morph_h = cv2.threshold(morph_h,morph_thr,255,cv2.THRESH_BINARY)
+ # morph_h = cv2.erode(morph_h, np.ones((5,5),np.uint8), iterations = 2)
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morph_thr_eroded", morph_v, 0, 1, config=config
+ )
+
+ self.append_save_img(6, morph_v)
+
+ # template relative alignment code
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+
+ match_col, max_steps, align_stride, thk = map(
+ config.alignment_params.get,
+ [
+ "match_col",
+ "max_steps",
+ "stride",
+ "thickness",
+ ],
+ )
+ shift, steps = 0, 0
+ while steps < max_steps:
+ left_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0] + shift - thk : -thk + s[0] + shift + match_col,
+ ]
+ )
+ right_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0]
+ + shift
+ - match_col
+ + d[0]
+ + thk : thk
+ + s[0]
+ + shift
+ + d[0],
+ ]
+ )
+
+ # For demonstration purposes-
+ # if(field_block.name == "int1"):
+ # ret = morph_v.copy()
+ # cv2.rectangle(ret,
+ # (s[0]+shift-thk,s[1]),
+ # (s[0]+shift+thk+d[0],s[1]+d[1]),
+ # constants.CLR_WHITE,
+ # 3)
+ # appendSaveImg(6,ret)
+ # print(shift, left_mean, right_mean)
+ left_shift, right_shift = left_mean > 100, right_mean > 100
+ if left_shift:
+ if right_shift:
+ break
+ else:
+ shift -= align_stride
+ else:
+ if right_shift:
+ shift += align_stride
+ else:
+ break
+ steps += 1
+
+ field_block.shift = shift
+ # print("Aligned field_block: ",field_block.name,"Corrected Shift:",
+ # field_block.shift,", dimensions:", field_block.dimensions,
+ # "origin:", field_block.origin,'\n')
+ # print("End Alignment")
+
+ final_align = None
+ if config.outputs.show_image_level >= 2:
+ initial_align = self.draw_template_layout(img, template, shifted=False)
+ final_align = self.draw_template_layout(
+ img, template, shifted=True, draw_qvals=True
+ )
+ # appendSaveImg(4,mean_vals)
+ self.append_save_img(2, initial_align)
+ self.append_save_img(2, final_align)
+
+ if auto_align:
+ final_align = np.hstack((initial_align, final_align))
+ self.append_save_img(5, img)
+
+ # Get mean bubbleValues n other stats
+ all_q_vals, all_q_strip_arrs, all_q_std_vals = [], [], []
+ total_q_strip_no = 0
+ for field_block in template.field_blocks:
+ box_w, box_h = field_block.bubble_dimensions
+ q_std_vals = []
+ for field_block_bubbles in field_block.traverse_bubbles:
+ q_strip_vals = []
+ for pt in field_block_bubbles:
+ # shifted
+ x, y = (pt.x + field_block.shift, pt.y)
+ rect = [y, y + box_h, x, x + box_w]
+ q_strip_vals.append(
+ cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0]
+ # detectCross(img, rect) ? 100 : 0
+ )
+ q_std_vals.append(round(np.std(q_strip_vals), 2))
+ all_q_strip_arrs.append(q_strip_vals)
+ # _, _, _ = get_global_threshold(q_strip_vals, "QStrip Plot",
+ # plot_show=False, sort_in_plot=True)
+ # hist = getPlotImg()
+ # InteractionUtils.show("QStrip "+field_block_bubbles[0].field_label, hist, 0, 1,config=config)
+ all_q_vals.extend(q_strip_vals)
+ # print(total_q_strip_no, field_block_bubbles[0].field_label, q_std_vals[len(q_std_vals)-1])
+ total_q_strip_no += 1
+ all_q_std_vals.extend(q_std_vals)
+
+ global_std_thresh, _, _ = self.get_global_threshold(
+ all_q_std_vals
+ ) # , "Q-wise Std-dev Plot", plot_show=True, sort_in_plot=True)
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # Note: Plotting takes Significant times here --> Change Plotting args
+ # to support show_image_level
+ # , "Mean Intensity Histogram",plot_show=True, sort_in_plot=True)
+ global_thr, _, _ = self.get_global_threshold(all_q_vals, looseness=4)
+
+ logger.info(
+ f"Thresholding: \tglobal_thr: {round(global_thr, 2)} \tglobal_std_THR: {round(global_std_thresh, 2)}\t{'(Looks like a Xeroxed OMR)' if (global_thr == 255) else ''}"
+ )
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # if(config.outputs.show_image_level>=1):
+ # hist = getPlotImg()
+ # InteractionUtils.show("Hist", hist, 0, 1,config=config)
+ # appendSaveImg(4,hist)
+ # appendSaveImg(5,hist)
+ # appendSaveImg(2,hist)
+
+ per_omr_threshold_avg, total_q_strip_no, total_q_box_no = 0, 0, 0
+ for field_block in template.field_blocks:
+ block_q_strip_no = 1
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ s, d = field_block.origin, field_block.dimensions
+ key = field_block.name[:3]
+ # cv2.rectangle(final_marked,(s[0]+shift,s[1]),(s[0]+shift+d[0],
+ # s[1]+d[1]),CLR_BLACK,3)
+ for field_block_bubbles in field_block.traverse_bubbles:
+ # All Black or All White case
+ no_outliers = all_q_std_vals[total_q_strip_no] < global_std_thresh
+ # print(total_q_strip_no, field_block_bubbles[0].field_label,
+ # all_q_std_vals[total_q_strip_no], "no_outliers:", no_outliers)
+ per_q_strip_threshold = self.get_local_threshold(
+ all_q_strip_arrs[total_q_strip_no],
+ global_thr,
+ no_outliers,
+ f"Mean Intensity Histogram for {key}.{field_block_bubbles[0].field_label}.{block_q_strip_no}",
+ config.outputs.show_image_level >= 6,
+ )
+ # print(field_block_bubbles[0].field_label,key,block_q_strip_no, "THR: ",
+ # round(per_q_strip_threshold,2))
+ per_omr_threshold_avg += per_q_strip_threshold
+
+ # Note: Little debugging visualization - view the particular Qstrip
+ # if(
+ # 0
+ # # or "q17" in (field_block_bubbles[0].field_label)
+ # # or (field_block_bubbles[0].field_label+str(block_q_strip_no))=="q15"
+ # ):
+ # st, end = qStrip
+ # InteractionUtils.show("QStrip: "+key+"-"+str(block_q_strip_no),
+ # img[st[1] : end[1], st[0]+shift : end[0]+shift],0,config=config)
+
+ # TODO: get rid of total_q_box_no
+ detected_bubbles = []
+ for bubble in field_block_bubbles:
+ bubble_is_marked = (
+ per_q_strip_threshold > all_q_vals[total_q_box_no]
+ )
+ total_q_box_no += 1
+ if bubble_is_marked:
+ detected_bubbles.append(bubble)
+ x, y, field_value = (
+ bubble.x + field_block.shift,
+ bubble.y,
+ bubble.field_value,
+ )
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 12), int(y + box_h / 12)),
+ (
+ int(x + box_w - box_w / 12),
+ int(y + box_h - box_h / 12),
+ ),
+ constants.CLR_DARK_GRAY,
+ 3,
+ )
+
+ cv2.putText(
+ final_marked,
+ str(field_value),
+ (x, y),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ (20, 20, 10),
+ int(1 + 3.5 * constants.TEXT_SIZE),
+ )
+ else:
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (
+ int(x + box_w - box_w / 10),
+ int(y + box_h - box_h / 10),
+ ),
+ constants.CLR_GRAY,
+ -1,
+ )
+
+ for bubble in detected_bubbles:
+ field_label, field_value = (
+ bubble.field_label,
+ bubble.field_value,
+ )
+ # Only send rolls multi-marked in the directory
+ multi_marked_local = field_label in omr_response
+ omr_response[field_label] = (
+ (omr_response[field_label] + field_value)
+ if multi_marked_local
+ else field_value
+ )
+ # TODO: generalize this into identifier
+ # multi_roll = multi_marked_local and "Roll" in str(q)
+ multi_marked = multi_marked or multi_marked_local
+
+ if len(detected_bubbles) == 0:
+ field_label = field_block_bubbles[0].field_label
+ omr_response[field_label] = field_block.empty_val
+
+ if config.outputs.show_image_level >= 5:
+ if key in all_c_box_vals:
+ q_nums[key].append(f"{key[:2]}_c{str(block_q_strip_no)}")
+ all_c_box_vals[key].append(
+ all_q_strip_arrs[total_q_strip_no]
+ )
+
+ block_q_strip_no += 1
+ total_q_strip_no += 1
+ # /for field_block
+
+ per_omr_threshold_avg /= total_q_strip_no
+ per_omr_threshold_avg = round(per_omr_threshold_avg, 2)
+ # Translucent
+ cv2.addWeighted(
+ final_marked, alpha, transp_layer, 1 - alpha, 0, final_marked
+ )
+ # Box types
+ if config.outputs.show_image_level >= 6:
+ # plt.draw()
+ f, axes = plt.subplots(len(all_c_box_vals), sharey=True)
+ f.canvas.manager.set_window_title(name)
+ ctr = 0
+ type_name = {
+ "int": "Integer",
+ "mcq": "MCQ",
+ "med": "MED",
+ "rol": "Roll",
+ }
+ for k, boxvals in all_c_box_vals.items():
+ axes[ctr].title.set_text(type_name[k] + " Type")
+ axes[ctr].boxplot(boxvals)
+ # thrline=axes[ctr].axhline(per_omr_threshold_avg,color='red',ls='--')
+ # thrline.set_label("Average THR")
+ axes[ctr].set_ylabel("Intensity")
+ axes[ctr].set_xticklabels(q_nums[k])
+ # axes[ctr].legend()
+ ctr += 1
+ # imshow will do the waiting
+ plt.tight_layout(pad=0.5)
+ plt.show()
+
+ if config.outputs.show_image_level >= 3 and final_align is not None:
+ final_align = ImageUtils.resize_util_h(
+ final_align, int(config.dimensions.display_height)
+ )
+ # [final_align.shape[1],0])
+ InteractionUtils.show(
+ "Template Alignment Adjustment", final_align, 0, 0, config=config
+ )
+
+ if config.outputs.save_detections and save_dir is not None:
+ if multi_roll:
+ save_dir = save_dir.joinpath("_MULTI_")
+ image_path = str(save_dir.joinpath(name))
+ ImageUtils.save_img(image_path, final_marked)
+
+ self.append_save_img(2, final_marked)
+
+ if save_dir is not None:
+ for i in range(config.outputs.save_image_level):
+ self.save_image_stacks(i + 1, name, save_dir)
+
+ return omr_response, final_marked, multi_marked, multi_roll
+
+ except Exception as e:
+ raise e
+
+ @staticmethod
+ def draw_template_layout(img, template, shifted=True, draw_qvals=False, border=-1):
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ final_align = img.copy()
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ if shifted:
+ cv2.rectangle(
+ final_align,
+ (s[0] + shift, s[1]),
+ (s[0] + shift + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ else:
+ cv2.rectangle(
+ final_align,
+ (s[0], s[1]),
+ (s[0] + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ for field_block_bubbles in field_block.traverse_bubbles:
+ for pt in field_block_bubbles:
+ x, y = (pt.x + field_block.shift, pt.y) if shifted else (pt.x, pt.y)
+ cv2.rectangle(
+ final_align,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (int(x + box_w - box_w / 10), int(y + box_h - box_h / 10)),
+ constants.CLR_GRAY,
+ border,
+ )
+ if draw_qvals:
+ rect = [y, y + box_h, x, x + box_w]
+ cv2.putText(
+ final_align,
+ f"{int(cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0])}",
+ (rect[2] + 2, rect[0] + (box_h * 2) // 3),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ 0.6,
+ constants.CLR_BLACK,
+ 2,
+ )
+ if shifted:
+ text_in_px = cv2.getTextSize(
+ field_block.name, cv2.FONT_HERSHEY_SIMPLEX, constants.TEXT_SIZE, 4
+ )
+ cv2.putText(
+ final_align,
+ field_block.name,
+ (int(s[0] + d[0] - text_in_px[0][0]), int(s[1] - text_in_px[0][1])),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ constants.CLR_BLACK,
+ 4,
+ )
+ return final_align
+
+ def get_global_threshold(
+ self,
+ q_vals_orig,
+ plot_title=None,
+ plot_show=True,
+ sort_in_plot=True,
+ looseness=1,
+ ):
+ """
+ Note: Cannot assume qStrip has only-gray or only-white bg
+ (in which case there is only one jump).
+ So there will be either 1 or 2 jumps.
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ 2 Jumps :
+ ......
+ |||||| <-- wrong THR
+ ....||||||
+ |||||||||| <-- safe THR
+ ..||||||||||
+ ||||||||||||
+
+ The abstract "First LARGE GAP" is perfect for this.
+ Current code is considering ONLY TOP 2 jumps(>= MIN_GAP) to be big,
+ gives the smaller one
+
+ """
+ config = self.tuning_config
+ PAGE_TYPE_FOR_THRESHOLD, MIN_JUMP, JUMP_DELTA = map(
+ config.threshold_params.get,
+ [
+ "PAGE_TYPE_FOR_THRESHOLD",
+ "MIN_JUMP",
+ "JUMP_DELTA",
+ ],
+ )
+
+ global_default_threshold = (
+ constants.GLOBAL_PAGE_THRESHOLD_WHITE
+ if PAGE_TYPE_FOR_THRESHOLD == "white"
+ else constants.GLOBAL_PAGE_THRESHOLD_BLACK
+ )
+
+ # Sort the Q bubbleValues
+ # TODO: Change var name of q_vals
+ q_vals = sorted(q_vals_orig)
+ # Find the FIRST LARGE GAP and set it as threshold:
+ ls = (looseness + 1) // 2
+ l = len(q_vals) - ls
+ max1, thr1 = MIN_JUMP, global_default_threshold
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - ls] + jump / 2
+
+ # NOTE: thr2 is deprecated, thus is JUMP_DELTA
+ # Make use of the fact that the JUMP_DELTA(Vertical gap ofc) between
+ # values at detected jumps would be atleast 20
+ max2, thr2 = MIN_JUMP, global_default_threshold
+ # Requires atleast 1 gray box to be present (Roll field will ensure this)
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ new_thr = q_vals[i - ls] + jump / 2
+ if jump > max2 and abs(thr1 - new_thr) > JUMP_DELTA:
+ max2 = jump
+ thr2 = new_thr
+ # global_thr = min(thr1,thr2)
+ global_thr, j_low, j_high = thr1, thr1 - max1 // 2, thr1 + max1 // 2
+
+ # # For normal images
+ # thresholdRead = 116
+ # if(thr1 > thr2 and thr2 > thresholdRead):
+ # print("Note: taking safer thr line.")
+ # global_thr, j_low, j_high = thr2, thr2 - max2//2, thr2 + max2//2
+
+ if plot_title:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals_orig)), q_vals if sort_in_plot else q_vals_orig)
+ ax.set_title(plot_title)
+ thrline = ax.axhline(global_thr, color="green", ls="--", linewidth=5)
+ thrline.set_label("Global Threshold")
+ thrline = ax.axhline(thr2, color="red", ls=":", linewidth=3)
+ thrline.set_label("THR2 Line")
+ # thrline=ax.axhline(j_low,color='red',ls='-.', linewidth=3)
+ # thrline=ax.axhline(j_high,color='red',ls='-.', linewidth=3)
+ # thrline.set_label("Boundary Line")
+ # ax.set_ylabel("Mean Intensity")
+ ax.set_ylabel("Values")
+ ax.set_xlabel("Position")
+ ax.legend()
+ if plot_show:
+ plt.title(plot_title)
+ plt.show()
+
+ return global_thr, j_low, j_high
+
+ def get_local_threshold(
+ self, q_vals, global_thr, no_outliers, plot_title=None, plot_show=True
+ ):
+ """
+ TODO: Update this documentation too-
+ //No more - Assumption : Colwise background color is uniformly gray or white,
+ but not alternating. In this case there is atmost one jump.
+
+ 0 Jump :
+ <-- safe THR?
+ .......
+ ...|||||||
+ |||||||||| <-- safe THR?
+ // How to decide given range is above or below gray?
+ -> global q_vals shall absolutely help here. Just run same function
+ on total q_vals instead of colwise _//
+ How to decide it is this case of 0 jumps
+
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ """
+ config = self.tuning_config
+ # Sort the Q bubbleValues
+ q_vals = sorted(q_vals)
+
+ # Small no of pts cases:
+ # base case: 1 or 2 pts
+ if len(q_vals) < 3:
+ thr1 = (
+ global_thr
+ if np.max(q_vals) - np.min(q_vals) < config.threshold_params.MIN_GAP
+ else np.mean(q_vals)
+ )
+ else:
+ # qmin, qmax, qmean, qstd = round(np.min(q_vals),2), round(np.max(q_vals),2),
+ # round(np.mean(q_vals),2), round(np.std(q_vals),2)
+ # GVals = [round(abs(q-qmean),2) for q in q_vals]
+ # gmean, gstd = round(np.mean(GVals),2), round(np.std(GVals),2)
+ # # DISCRETION: Pretty critical factor in reading response
+ # # Doesn't work well for small number of values.
+ # DISCRETION = 2.7 # 2.59 was closest hit, 3.0 is too far
+ # L2MaxGap = round(max([abs(g-gmean) for g in GVals]),2)
+ # if(L2MaxGap > DISCRETION*gstd):
+ # no_outliers = False
+
+ # # ^Stackoverflow method
+ # print(field_label, no_outliers,"qstd",round(np.std(q_vals),2), "gstd", gstd,
+ # "Gaps in gvals",sorted([round(abs(g-gmean),2) for g in GVals],reverse=True),
+ # '\t',round(DISCRETION*gstd,2), L2MaxGap)
+
+ # else:
+ # Find the LARGEST GAP and set it as threshold: //(FIRST LARGE GAP)
+ l = len(q_vals) - 1
+ max1, thr1 = config.threshold_params.MIN_JUMP, 255
+ for i in range(1, l):
+ jump = q_vals[i + 1] - q_vals[i - 1]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - 1] + jump / 2
+ # print(field_label,q_vals,max1)
+
+ confident_jump = (
+ config.threshold_params.MIN_JUMP
+ + config.threshold_params.CONFIDENT_SURPLUS
+ )
+ # If not confident, then only take help of global_thr
+ if max1 < confident_jump:
+ if no_outliers:
+ # All Black or All White case
+ thr1 = global_thr
+ else:
+ # TODO: Low confidence parameters here
+ pass
+
+ # if(thr1 == 255):
+ # print("Warning: threshold is unexpectedly 255! (Outlier Delta issue?)",plot_title)
+
+ # Make a common plot function to show local and global thresholds
+ if plot_show and plot_title is not None:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals)), q_vals)
+ thrline = ax.axhline(thr1, color="green", ls=("-."), linewidth=3)
+ thrline.set_label("Local Threshold")
+ thrline = ax.axhline(global_thr, color="red", ls=":", linewidth=5)
+ thrline.set_label("Global Threshold")
+ ax.set_title(plot_title)
+ ax.set_ylabel("Bubble Mean Intensity")
+ ax.set_xlabel("Bubble Number(sorted)")
+ ax.legend()
+ # TODO append QStrip to this plot-
+ # appendSaveImg(6,getPlotImg())
+ if plot_show:
+ plt.show()
+ return thr1
+
+ def append_save_img(self, key, img):
+ if self.save_image_level >= int(key):
+ self.save_img_list[key].append(img.copy())
+
+ def save_image_stacks(self, key, filename, save_dir):
+ config = self.tuning_config
+ if self.save_image_level >= int(key) and self.save_img_list[key] != []:
+ name = os.path.splitext(filename)[0]
+ result = np.hstack(
+ tuple(
+ [
+ ImageUtils.resize_util_h(img, config.dimensions.display_height)
+ for img in self.save_img_list[key]
+ ]
+ )
+ )
+ result = ImageUtils.resize_util(
+ result,
+ min(
+ len(self.save_img_list[key]) * config.dimensions.display_width // 3,
+ int(config.dimensions.display_width * 2.5),
+ ),
+ )
+ ImageUtils.save_img(f"{save_dir}stack/{name}_{str(key)}_stack.jpg", result)
+
+ def reset_all_save_img(self):
+ for i in range(self.save_image_level):
+ self.save_img_list[i + 1] = []
diff --git a/OMRChecker/src/defaults/__init__.py b/OMRChecker/src/defaults/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..db3d42e87b3e47d0cbee08032e65e87ac1bd8165
--- /dev/null
+++ b/OMRChecker/src/defaults/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+# Use all imports relative to root directory
+# (https://chrisyeh96.github.io/2017/08/08/definitive-guide-python-imports.html)
+from src.defaults.config import * # NOQA
+from src.defaults.template import * # NOQA
diff --git a/OMRChecker/src/defaults/config.py b/OMRChecker/src/defaults/config.py
new file mode 100644
index 0000000000000000000000000000000000000000..6d1315fb45b0e736ad7daedcea5f3b23e3a9078d
--- /dev/null
+++ b/OMRChecker/src/defaults/config.py
@@ -0,0 +1,35 @@
+from dotmap import DotMap
+
+CONFIG_DEFAULTS = DotMap(
+ {
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666,
+ },
+ "threshold_params": {
+ "GAMMA_LOW": 0.7,
+ "MIN_GAP": 30,
+ "MIN_JUMP": 25,
+ "CONFIDENT_SURPLUS": 5,
+ "JUMP_DELTA": 30,
+ "PAGE_TYPE_FOR_THRESHOLD": "white",
+ },
+ "alignment_params": {
+ # Note: 'auto_align' enables automatic template alignment, use if the scans show slight misalignments.
+ "auto_align": False,
+ "match_col": 5,
+ "max_steps": 20,
+ "stride": 1,
+ "thickness": 3,
+ },
+ "outputs": {
+ "show_image_level": 0,
+ "save_image_level": 0,
+ "save_detections": True,
+ "filter_out_multimarked_files": False,
+ },
+ },
+ _dynamic=False,
+)
diff --git a/OMRChecker/src/defaults/template.py b/OMRChecker/src/defaults/template.py
new file mode 100644
index 0000000000000000000000000000000000000000..efd75420f01cf49f72a2e285734c0d79cafe4837
--- /dev/null
+++ b/OMRChecker/src/defaults/template.py
@@ -0,0 +1,6 @@
+TEMPLATE_DEFAULTS = {
+ "preProcessors": [],
+ "emptyValue": "",
+ "customLabels": {},
+ "outputColumns": [],
+}
diff --git a/OMRChecker/src/entry.py b/OMRChecker/src/entry.py
new file mode 100644
index 0000000000000000000000000000000000000000..94d3137c3ac99502399cbb40d6d378cd8249d8de
--- /dev/null
+++ b/OMRChecker/src/entry.py
@@ -0,0 +1,371 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import os
+from csv import QUOTE_NONNUMERIC
+from pathlib import Path
+from time import time
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src import constants
+from src.defaults import CONFIG_DEFAULTS
+from src.evaluation import EvaluationConfig, evaluate_concatenated_response
+from src.logger import console, logger
+from src.template import Template
+from src.utils.file import Paths, setup_dirs_for_paths, setup_outputs_for_template
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils, Stats
+from src.utils.parsing import get_concatenated_response, open_config_with_defaults
+
+# Load processors
+STATS = Stats()
+
+
+def entry_point(input_dir, args):
+ if not os.path.exists(input_dir):
+ raise Exception(f"Given input directory does not exist: '{input_dir}'")
+ curr_dir = input_dir
+ return process_dir(input_dir, curr_dir, args)
+
+
+def print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+):
+ logger.info("")
+ table = Table(title="Current Configurations", show_header=False, show_lines=False)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Value", style="magenta")
+ table.add_row("Directory Path", f"{curr_dir}")
+ table.add_row("Count of Images", f"{len(omr_files)}")
+ table.add_row("Set Layout Mode ", "ON" if args["setLayout"] else "OFF")
+ pre_processor_names = [pp.__class__.__name__ for pp in template.pre_processors]
+ table.add_row(
+ "Markers Detection",
+ "ON" if "CropOnMarkers" in pre_processor_names else "OFF",
+ )
+ table.add_row("Auto Alignment", f"{tuning_config.alignment_params.auto_align}")
+ table.add_row("Detected Template Path", f"{template}")
+ if local_config_path:
+ table.add_row("Detected Local Config", f"{local_config_path}")
+ if evaluation_config:
+ table.add_row("Detected Evaluation Config", f"{evaluation_config}")
+
+ table.add_row(
+ "Detected pre-processors",
+ ", ".join(pre_processor_names),
+ )
+ console.print(table, justify="center")
+
+
+def process_dir(
+ root_dir,
+ curr_dir,
+ args,
+ template=None,
+ tuning_config=CONFIG_DEFAULTS,
+ evaluation_config=None,
+):
+ # Update local tuning_config (in current recursion stack)
+ local_config_path = curr_dir.joinpath(constants.CONFIG_FILENAME)
+ if os.path.exists(local_config_path):
+ tuning_config = open_config_with_defaults(local_config_path)
+
+ # Update local template (in current recursion stack)
+ local_template_path = curr_dir.joinpath(constants.TEMPLATE_FILENAME)
+ local_template_exists = os.path.exists(local_template_path)
+ if local_template_exists:
+ template = Template(
+ local_template_path,
+ tuning_config,
+ )
+ # Look for subdirectories for processing
+ subdirs = [d for d in curr_dir.iterdir() if d.is_dir()]
+
+ output_dir = Path(args["output_dir"], curr_dir.relative_to(root_dir))
+ paths = Paths(output_dir)
+
+ # look for images in current dir to process
+ exts = ("*.[pP][nN][gG]", "*.[jJ][pP][gG]", "*.[jJ][pP][eE][gG]")
+ omr_files = sorted([f for ext in exts for f in curr_dir.glob(ext)])
+
+ # Exclude images (take union over all pre_processors)
+ excluded_files = []
+ if template:
+ for pp in template.pre_processors:
+ excluded_files.extend(Path(p) for p in pp.exclude_files())
+
+ local_evaluation_path = curr_dir.joinpath(constants.EVALUATION_FILENAME)
+ if not args["setLayout"] and os.path.exists(local_evaluation_path):
+ if not local_template_exists:
+ logger.warning(
+ f"Found an evaluation file without a parent template file: {local_evaluation_path}"
+ )
+ evaluation_config = EvaluationConfig(
+ curr_dir,
+ local_evaluation_path,
+ template,
+ tuning_config,
+ )
+
+ excluded_files.extend(
+ Path(exclude_file) for exclude_file in evaluation_config.get_exclude_files()
+ )
+
+ omr_files = [f for f in omr_files if f not in excluded_files]
+
+ if omr_files:
+ if not template:
+ logger.error(
+ f"Found images, but no template in the directory tree \
+ of '{curr_dir}'. \nPlace {constants.TEMPLATE_FILENAME} in the \
+ appropriate directory."
+ )
+ raise Exception(
+ f"No template file found in the directory tree of {curr_dir}"
+ )
+
+ setup_dirs_for_paths(paths)
+ outputs_namespace = setup_outputs_for_template(paths, template)
+
+ print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+ )
+ if args["setLayout"]:
+ show_template_layouts(omr_files, template, tuning_config)
+ else:
+ process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+ )
+
+ elif not subdirs:
+ # Each subdirectory should have images or should be non-leaf
+ logger.info(
+ f"No valid images or sub-folders found in {curr_dir}.\
+ Empty directories not allowed."
+ )
+
+ # recursively process sub-folders
+ for d in subdirs:
+ process_dir(
+ root_dir,
+ d,
+ args,
+ template,
+ tuning_config,
+ evaluation_config,
+ )
+
+
+def show_template_layouts(omr_files, template, tuning_config):
+ for file_path in omr_files:
+ file_name = file_path.name
+ file_path = str(file_path)
+ in_omr = cv2.imread(file_path, cv2.IMREAD_GRAYSCALE)
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+ template_layout = template.image_instance_ops.draw_template_layout(
+ in_omr, template, shifted=False, border=2
+ )
+ InteractionUtils.show(
+ f"Template Layout: {file_name}", template_layout, 1, 1, config=tuning_config
+ )
+
+
+def process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+):
+ start_time = int(time())
+ files_counter = 0
+ STATS.files_not_moved = 0
+
+ for file_path in omr_files:
+ files_counter += 1
+ file_name = file_path.name
+
+ in_omr = cv2.imread(str(file_path), cv2.IMREAD_GRAYSCALE)
+
+ logger.info("")
+ logger.info(
+ f"({files_counter}) Opening image: \t'{file_path}'\tResolution: {in_omr.shape}"
+ )
+
+ template.image_instance_ops.reset_all_save_img()
+
+ template.image_instance_ops.append_save_img(1, in_omr)
+
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+
+ if in_omr is None:
+ # Error OMR case
+ new_file_path = outputs_namespace.paths.errors_dir.joinpath(file_name)
+ outputs_namespace.OUTPUT_SET.append(
+ [file_name] + outputs_namespace.empty_resp
+ )
+ if check_and_move(
+ constants.ERROR_CODES.NO_MARKER_ERR, file_path, new_file_path
+ ):
+ err_line = [
+ file_name,
+ file_path,
+ new_file_path,
+ "NA",
+ ] + outputs_namespace.empty_resp
+ pd.DataFrame(err_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Errors"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ continue
+
+ # uniquify
+ file_id = str(file_name)
+ save_dir = outputs_namespace.paths.save_marked_dir
+ (
+ response_dict,
+ final_marked,
+ multi_marked,
+ _,
+ ) = template.image_instance_ops.read_omr_response(
+ template, image=in_omr, name=file_id, save_dir=save_dir
+ )
+
+ # TODO: move inner try catch here
+ # concatenate roll nos, set unmarked responses, etc
+ omr_response = get_concatenated_response(response_dict, template)
+
+ if (
+ evaluation_config is None
+ or not evaluation_config.get_should_explain_scoring()
+ ):
+ logger.info(f"Read Response: \n{omr_response}")
+
+ score = 0
+ if evaluation_config is not None:
+ score = evaluate_concatenated_response(
+ omr_response, evaluation_config, file_path, outputs_namespace.paths.evaluation_dir
+ )
+ logger.info(
+ f"(/{files_counter}) Graded with score: {round(score, 2)}\t for file: '{file_id}'"
+ )
+ else:
+ logger.info(f"(/{files_counter}) Processed file: '{file_id}'")
+
+ if tuning_config.outputs.show_image_level >= 2:
+ InteractionUtils.show(
+ f"Final Marked Bubbles : '{file_id}'",
+ ImageUtils.resize_util_h(
+ final_marked, int(tuning_config.dimensions.display_height * 1.3)
+ ),
+ 1,
+ 1,
+ config=tuning_config,
+ )
+
+ resp_array = []
+ for k in template.output_columns:
+ resp_array.append(omr_response[k])
+
+ outputs_namespace.OUTPUT_SET.append([file_name] + resp_array)
+
+ if multi_marked == 0 or not tuning_config.outputs.filter_out_multimarked_files:
+ STATS.files_not_moved += 1
+ new_file_path = save_dir.joinpath(file_id)
+ # Enter into Results sheet-
+ results_line = [file_name, file_path, new_file_path, score] + resp_array
+ # Write/Append to results_line file(opened in append mode)
+ pd.DataFrame(results_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Results"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ else:
+ # multi_marked file
+ logger.info(f"[{files_counter}] Found multi-marked file: '{file_id}'")
+ new_file_path = outputs_namespace.paths.multi_marked_dir.joinpath(file_name)
+ if check_and_move(
+ constants.ERROR_CODES.MULTI_BUBBLE_WARN, file_path, new_file_path
+ ):
+ mm_line = [file_name, file_path, new_file_path, "NA"] + resp_array
+ pd.DataFrame(mm_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["MultiMarked"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ # else:
+ # TODO: Add appropriate record handling here
+ # pass
+
+ print_stats(start_time, files_counter, tuning_config)
+
+
+def check_and_move(error_code, file_path, filepath2):
+ # TODO: fix file movement into error/multimarked/invalid etc again
+ STATS.files_not_moved += 1
+ return True
+
+
+def print_stats(start_time, files_counter, tuning_config):
+ time_checking = max(1, round(time() - start_time, 2))
+ log = logger.info
+ log("")
+ log(f"{'Total file(s) moved': <27}: {STATS.files_moved}")
+ log(f"{'Total file(s) not moved': <27}: {STATS.files_not_moved}")
+ log("--------------------------------")
+ log(
+ f"{'Total file(s) processed': <27}: {files_counter} ({'Sum Tallied!' if files_counter == (STATS.files_moved + STATS.files_not_moved) else 'Not Tallying!'})"
+ )
+
+ if tuning_config.outputs.show_image_level <= 0:
+ log(
+ f"\nFinished Checking {files_counter} file(s) in {round(time_checking, 1)} seconds i.e. ~{round(time_checking / 60, 1)} minute(s)."
+ )
+ log(
+ f"{'OMR Processing Rate': <27}: \t ~ {round(time_checking / files_counter, 2)} seconds/OMR"
+ )
+ log(
+ f"{'OMR Processing Speed': <27}: \t ~ {round((files_counter * 60) / time_checking, 2)} OMRs/minute"
+ )
+ else:
+ log(f"\n{'Total script time': <27}: {time_checking} seconds")
+
+ if tuning_config.outputs.show_image_level <= 1:
+ log(
+ "\nTip: To see some awesome visuals, open config.json and increase 'show_image_level'"
+ )
diff --git a/OMRChecker/src/evaluation.py b/OMRChecker/src/evaluation.py
new file mode 100644
index 0000000000000000000000000000000000000000..55c2455a2967063e7604d0f0c904c09720e1064d
--- /dev/null
+++ b/OMRChecker/src/evaluation.py
@@ -0,0 +1,546 @@
+import ast
+import os
+import re
+from copy import deepcopy
+from csv import QUOTE_NONNUMERIC
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src.logger import console, logger
+from src.schemas.constants import (
+ BONUS_SECTION_PREFIX,
+ DEFAULT_SECTION_KEY,
+ MARKING_VERDICT_TYPES,
+)
+from src.utils.parsing import (
+ get_concatenated_response,
+ open_evaluation_with_validation,
+ parse_fields,
+ parse_float_or_fraction,
+)
+
+
+class AnswerMatcher:
+ def __init__(self, answer_item, section_marking_scheme):
+ self.section_marking_scheme = section_marking_scheme
+ self.answer_item = answer_item
+ self.answer_type = self.validate_and_get_answer_type(answer_item)
+ self.set_defaults_from_scheme(section_marking_scheme)
+
+ @staticmethod
+ def is_a_marking_score(answer_element):
+ # Note: strict type checking is already done at schema validation level,
+ # Here we focus on overall struct type
+ return type(answer_element) == str or type(answer_element) == int
+
+ @staticmethod
+ def is_standard_answer(answer_element):
+ return type(answer_element) == str and len(answer_element) >= 1
+
+ def validate_and_get_answer_type(self, answer_item):
+ if self.is_standard_answer(answer_item):
+ return "standard"
+ elif type(answer_item) == list:
+ if (
+ # Array of answer elements: ['A', 'B', 'AB']
+ len(answer_item) >= 2
+ and all(
+ self.is_standard_answer(answers_or_score)
+ for answers_or_score in answer_item
+ )
+ ):
+ return "multiple-correct"
+ elif (
+ # Array of two-tuples: [['A', 1], ['B', 1], ['C', 3], ['AB', 2]]
+ len(answer_item) >= 1
+ and all(
+ type(answer_and_score) == list and len(answer_and_score) == 2
+ for answer_and_score in answer_item
+ )
+ and all(
+ self.is_standard_answer(allowed_answer)
+ and self.is_a_marking_score(answer_score)
+ for allowed_answer, answer_score in answer_item
+ )
+ ):
+ return "multiple-correct-weighted"
+
+ logger.critical(
+ f"Unable to determine answer type for answer item: {answer_item}"
+ )
+ raise Exception("Unable to determine answer type")
+
+ def set_defaults_from_scheme(self, section_marking_scheme):
+ answer_type = self.answer_type
+ self.empty_val = section_marking_scheme.empty_val
+ answer_item = self.answer_item
+ self.marking = deepcopy(section_marking_scheme.marking)
+ # TODO: reuse part of parse_scheme_marking here -
+ if answer_type == "standard":
+ # no local overrides
+ pass
+ elif answer_type == "multiple-correct":
+ # override marking scheme scores for each allowed answer
+ for allowed_answer in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = self.marking["correct"]
+ elif answer_type == "multiple-correct-weighted":
+ # Note: No override using marking scheme as answer scores are provided in answer_item
+ for allowed_answer, answer_score in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = parse_float_or_fraction(
+ answer_score
+ )
+
+ def get_marking_scheme(self):
+ return self.section_marking_scheme
+
+ def get_section_explanation(self):
+ answer_type = self.answer_type
+ if answer_type in ["standard", "multiple-correct"]:
+ return self.section_marking_scheme.section_key
+ elif answer_type == "multiple-correct-weighted":
+ return f"Custom: {self.marking}"
+
+ def get_verdict_marking(self, marked_answer):
+ answer_type = self.answer_type
+ question_verdict = "incorrect"
+ if answer_type == "standard":
+ question_verdict = self.get_standard_verdict(marked_answer)
+ elif answer_type == "multiple-correct":
+ question_verdict = self.get_multiple_correct_verdict(marked_answer)
+ elif answer_type == "multiple-correct-weighted":
+ question_verdict = self.get_multiple_correct_weighted_verdict(marked_answer)
+ return question_verdict, self.marking[question_verdict]
+
+ def get_standard_verdict(self, marked_answer):
+ allowed_answer = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer == allowed_answer:
+ return "correct"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_verdict(self, marked_answer):
+ allowed_answers = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_weighted_verdict(self, marked_answer):
+ allowed_answers = [
+ allowed_answer for allowed_answer, _answer_score in self.answer_item
+ ]
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def __str__(self):
+ return f"{self.answer_item}"
+
+
+class SectionMarkingScheme:
+ def __init__(self, section_key, section_scheme, empty_val):
+ # TODO: get local empty_val from qblock
+ self.empty_val = empty_val
+ self.section_key = section_key
+ # DEFAULT marking scheme follows a shorthand
+ if section_key == DEFAULT_SECTION_KEY:
+ self.questions = None
+ self.marking = self.parse_scheme_marking(section_scheme)
+ else:
+ self.questions = parse_fields(section_key, section_scheme["questions"])
+ self.marking = self.parse_scheme_marking(section_scheme["marking"])
+
+ def __str__(self):
+ return self.section_key
+
+ def parse_scheme_marking(self, marking):
+ parsed_marking = {}
+ for verdict_type in MARKING_VERDICT_TYPES:
+ verdict_marking = parse_float_or_fraction(marking[verdict_type])
+ if (
+ verdict_marking > 0
+ and verdict_type == "incorrect"
+ and not self.section_key.startswith(BONUS_SECTION_PREFIX)
+ ):
+ logger.warning(
+ f"Found positive marks({round(verdict_marking, 2)}) for incorrect answer in the schema '{self.section_key}'. For Bonus sections, add a prefix 'BONUS_' to them."
+ )
+ parsed_marking[verdict_type] = verdict_marking
+
+ return parsed_marking
+
+ def match_answer(self, marked_answer, answer_matcher):
+ question_verdict, verdict_marking = answer_matcher.get_verdict_marking(
+ marked_answer
+ )
+
+ return verdict_marking, question_verdict
+
+
+class EvaluationConfig:
+ """Note: this instance will be reused for multiple omr sheets"""
+
+ def __init__(self, curr_dir, evaluation_path, template, tuning_config):
+ self.path = evaluation_path
+ evaluation_json = open_evaluation_with_validation(evaluation_path)
+ options, marking_schemes, source_type = map(
+ evaluation_json.get, ["options", "marking_schemes", "source_type"]
+ )
+ self.should_explain_scoring = options.get("should_explain_scoring", False)
+ self.has_non_default_section = False
+ self.exclude_files = []
+ self.enable_evaluation_table_to_csv = options.get(
+ "enable_evaluation_table_to_csv", False
+ )
+
+ if source_type == "csv":
+ csv_path = curr_dir.joinpath(options["answer_key_csv_path"])
+ if not os.path.exists(csv_path):
+ logger.warning(f"Answer key csv does not exist at: '{csv_path}'.")
+
+ answer_key_image_path = options.get("answer_key_image_path", None)
+ if os.path.exists(csv_path):
+ # TODO: CSV parsing/validation for each row with a (qNo,
[](https://www.paypal.me/Udayraj123/500)
+
+_Find OMRChecker on_ [**_Product Hunt_**](https://www.producthunt.com/posts/omr-checker/) **|** [**_Reddit_**](https://www.reddit.com/r/computervision/comments/ccbj6f/omrchecker_grade_exams_using_python_and_opencv/) **|** [**Discord**](https://discord.gg/qFv2Vqf) **|** [**Linkedin**](https://www.linkedin.com/pulse/open-source-talks-udayraj-udayraj-deshmukh/) **|** [**goodfirstissue.dev**](https://goodfirstissue.dev/language/python) **|** [**codepeak.tech**](https://www.codepeak.tech/) **|** [**fossoverflow.dev**](https://fossoverflow.dev/projects) **|** [**Interview on Console by CodeSee**](https://console.substack.com/p/console-140) **|** [**Open Source Hub**](https://opensourcehub.io/udayraj123/omrchecker)
+
+
+
diff --git a/OMRChecker/docs/assets/colored_output.jpg b/OMRChecker/docs/assets/colored_output.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3cafa473b9a1134f845858be4cf0e98f3bd3bb7c
Binary files /dev/null and b/OMRChecker/docs/assets/colored_output.jpg differ
diff --git a/OMRChecker/inputs/OMRImage.jpg b/OMRChecker/inputs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c9e3dcccfc8af7c60becb3a4cc25f650861db98e
--- /dev/null
+++ b/OMRChecker/inputs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:4439b6a479cb9fea92de9656b900a528052616ae8e8962a6c33b8e81ed7f327a
+size 267398
diff --git a/OMRChecker/inputs/template.json b/OMRChecker/inputs/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..823fe2e58a4182ef04d1fff99ecf146b5ec53aae
--- /dev/null
+++ b/OMRChecker/inputs/template.json
@@ -0,0 +1,37 @@
+{
+ "pageDimensions": [
+ 1122,
+ 1600
+ ],
+ "bubbleDimensions": [
+ 48,
+ 50
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 100,
+ 175
+ ],
+ "bubblesGap": 55,
+ "labelsGap": 67,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q02block": {
+ "origin": [
+ 100,
+ 845
+ ],
+ "bubblesGap": 58,
+ "labelsGap": 70,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ }
+
+}
\ No newline at end of file
diff --git a/OMRChecker/main.py b/OMRChecker/main.py
new file mode 100644
index 0000000000000000000000000000000000000000..19e3ebd43d4be2a096cf709b04817247f9952506
--- /dev/null
+++ b/OMRChecker/main.py
@@ -0,0 +1,99 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+
+import argparse
+import sys
+from pathlib import Path
+
+from src.entry import entry_point
+from src.logger import logger
+
+
+def parse_args():
+ # construct the argument parse and parse the arguments
+ argparser = argparse.ArgumentParser()
+
+ argparser.add_argument(
+ "-i",
+ "--inputDir",
+ default=["inputs"],
+ # https://docs.python.org/3/library/argparse.html#nargs
+ nargs="*",
+ required=False,
+ type=str,
+ dest="input_paths",
+ help="Specify an input directory.",
+ )
+
+ argparser.add_argument(
+ "-d",
+ "--debug",
+ required=False,
+ dest="debug",
+ action="store_false",
+ help="Enables debugging mode for showing detailed errors",
+ )
+
+ argparser.add_argument(
+ "-o",
+ "--outputDir",
+ default="outputs",
+ required=False,
+ dest="output_dir",
+ help="Specify an output directory.",
+ )
+
+ argparser.add_argument(
+ "-a",
+ "--autoAlign",
+ required=False,
+ dest="autoAlign",
+ action="store_true",
+ help="(experimental) Enables automatic template alignment - \
+ use if the scans show slight misalignments.",
+ )
+
+ argparser.add_argument(
+ "-l",
+ "--setLayout",
+ required=False,
+ dest="setLayout",
+ action="store_true",
+ help="Set up OMR template layout - modify your json file and \
+ run again until the template is set.",
+ )
+
+ (
+ args,
+ unknown,
+ ) = argparser.parse_known_args()
+
+ args = vars(args)
+
+ if len(unknown) > 0:
+ logger.warning(f"\nError: Unknown arguments: {unknown}", unknown)
+ argparser.print_help()
+ exit(11)
+ return args
+
+
+def entry_point_for_args(args):
+ if args["debug"] is True:
+ # Disable tracebacks
+ sys.tracebacklimit = 0
+ for root in args["input_paths"]:
+ entry_point(
+ Path(root),
+ args,
+ )
+
+
+if __name__ == "__main__":
+ args = parse_args()
+ entry_point_for_args(args)
diff --git a/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..23529519c2c8778ff89dd2a8f5c7ce602a34a826
--- /dev/null
+++ b/OMRChecker/outputs/AdrianSample/Results/Results_07PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5"
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRImage.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..d8d1a2325170014745edf3503b40e7c7bda160dc
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:f16c45dda1d02bf06e33a6d219c8282c363bf3943b94960eede1efe2bc6204d9
+size 408744
diff --git a/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..aa91e5cd858dfa684d288a51acb4bc1b59b838cd
--- /dev/null
+++ b/OMRChecker/outputs/CheckedOMRs/OcrSheetMarked.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:b9104889d79a25e0550e8e1bd258f1c8520ae1162e41b567d5cdfebad64c2ab4
+size 401565
diff --git a/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..3a968368fa191241dda21ea0c41f5d2d840f2e52
--- /dev/null
+++ b/OMRChecker/outputs/Images/CheckedOMRs/OMRSheet.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:7e9678555042535c83ab920d0b616d4f99f9f08017629d46a15eb64d7c3abd3c
+size 407394
diff --git a/OMRChecker/outputs/Images/Manual/ErrorFiles.csv b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Images/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/Images/Results/Results_10PM.csv b/OMRChecker/outputs/Images/Results/Results_10PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..f3b83ec4b12f9e002ca0824ee43fc41f1f9ae30a
--- /dev/null
+++ b/OMRChecker/outputs/Images/Results/Results_10PM.csv
@@ -0,0 +1,2 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\Images\OMRSheet.jpg","outputs\Images\CheckedOMRs\OMRSheet.jpg","0","C","D","A","C","B","C","C","D","D","B","B","B","D","C","C","ABCD","B","ABCD","C","ABCD"
diff --git a/OMRChecker/outputs/Manual/ErrorFiles.csv b/OMRChecker/outputs/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..a9bb85c601677eefe49c6a75b39d5bd404506322
--- /dev/null
+++ b/OMRChecker/outputs/Manual/ErrorFiles.csv
@@ -0,0 +1,3 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
+"OMRSheet.jpg","inputs\OMRSheet.jpg","outputs\Manual\ErrorFiles\OMRSheet.jpg","NA","","","","","","","","","","","","","","","","","","","",""
diff --git a/OMRChecker/outputs/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..84597b76dd63112496537204330caf1a7d436621
--- /dev/null
+++ b/OMRChecker/outputs/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/ErrorFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Manual/MultiMarkedFiles.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
new file mode 100644
index 0000000000000000000000000000000000000000..16614444af8b0e2c2eb65aeabd26d58f85ddce32
--- /dev/null
+++ b/OMRChecker/outputs/MobileCamera/Results/Results_09PM.csv
@@ -0,0 +1 @@
+"file_id","input_path","output_path","score","Roll","q1","q2","q3","q4","q5","q6","q7","q8","q9","q10","q11","q12","q13","q14","q15","q16","q17","q18","q19","q20"
diff --git a/OMRChecker/pyproject.toml b/OMRChecker/pyproject.toml
new file mode 100644
index 0000000000000000000000000000000000000000..efa55e8fc8bf1623d8569abd406fef4d81270291
--- /dev/null
+++ b/OMRChecker/pyproject.toml
@@ -0,0 +1,18 @@
+[tool.black]
+exclude = '''
+(
+ /(
+ \.eggs # exclude a few common directories in the
+ | \.git # root of the project
+ | \.venv
+ | _build
+ | build
+ | dist
+ )/
+ | foo.py # also separately exclude a file named foo.py in
+ # the root of the project
+)
+'''
+include = '\.pyi?$'
+line-length = 88
+target-version = ['py37']
diff --git a/OMRChecker/pytest.ini b/OMRChecker/pytest.ini
new file mode 100644
index 0000000000000000000000000000000000000000..417c4a2dd810294aa9afaa9b4f73464fc9808c96
--- /dev/null
+++ b/OMRChecker/pytest.ini
@@ -0,0 +1,6 @@
+# pytest.ini
+[pytest]
+minversion = 7.0
+addopts = -qq --capture=no
+testpaths =
+ src/tests
diff --git a/OMRChecker/requirements.dev.txt b/OMRChecker/requirements.dev.txt
new file mode 100644
index 0000000000000000000000000000000000000000..fb31b5f80558481a3f2a5edbb741b363a1bd1089
--- /dev/null
+++ b/OMRChecker/requirements.dev.txt
@@ -0,0 +1,7 @@
+-r requirements.txt
+flake8>=6.0.0
+freezegun>=1.2.2
+pre-commit>=3.3.3
+pytest-mock>=3.11.1
+pytest>=7.4.0
+syrupy>=4.0.4
diff --git a/OMRChecker/requirements.txt b/OMRChecker/requirements.txt
new file mode 100644
index 0000000000000000000000000000000000000000..d0c3af875835b492a48ef689c3dc9335eb7cdca7
--- /dev/null
+++ b/OMRChecker/requirements.txt
@@ -0,0 +1,8 @@
+deepmerge>=1.1.0
+dotmap>=1.3.30
+jsonschema>=4.17.3
+matplotlib>=3.7.1
+numpy>=1.25.0
+pandas>=2.0.2
+rich>=13.4.2
+screeninfo>=0.8.1
diff --git a/OMRChecker/samples/answer-key/using-csv/adrian_omr.png b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/using-csv/adrian_omr.png differ
diff --git a/OMRChecker/samples/answer-key/using-csv/answer_key.csv b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..acf09be611a66dfaeb7127e7249eadfbba6b0881
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/answer_key.csv
@@ -0,0 +1,5 @@
+q1,C
+q2,E
+q3,A
+q4,B
+q5,B
\ No newline at end of file
diff --git a/OMRChecker/samples/answer-key/using-csv/evaluation.json b/OMRChecker/samples/answer-key/using-csv/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/using-csv/template.json b/OMRChecker/samples/answer-key/using-csv/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/using-csv/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/evaluation.json b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..09508b18e3490efcd68d6cb0b57a03d6e203f2ca
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/evaluation.json
@@ -0,0 +1,35 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..5"
+ ],
+ "answers_in_order": [
+ "C",
+ "E",
+ [
+ "A",
+ "C"
+ ],
+ [
+ [
+ "B",
+ 2
+ ],
+ [
+ "C",
+ "3/2"
+ ]
+ ],
+ "C"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/answer-key/weighted-answers/images/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/answer-key/weighted-answers/template.json b/OMRChecker/samples/answer-key/weighted-answers/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/answer-key/weighted-answers/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..661d5f4fa73d7eea57e73837cf3862a25b9efcc3
Binary files /dev/null and b/OMRChecker/samples/community/Antibodyy/simple_omr_sheet.jpg differ
diff --git a/OMRChecker/samples/community/Antibodyy/template.json b/OMRChecker/samples/community/Antibodyy/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..55e016e8eb402156d62f6553067b68abb37377ee
--- /dev/null
+++ b/OMRChecker/samples/community/Antibodyy/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 299,
+ 398
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 79
+ ],
+ "bubblesGap": 43,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "q1..6"
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-1.png b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
new file mode 100644
index 0000000000000000000000000000000000000000..5ed00d194a806d58c3db1dab7831796df7d4aaef
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-1.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:ed3024e5b709fbbdb1bcbef35dcfa50203ddb811f74b26e657d3f1b86c7e8c94
+size 381247
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-2.png b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d250db2c94980defb7ab78f9114ca56c1f223024
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-2.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:571109a06a09b792694a0ecac329283da199ac4495b1f10ffb7e5925c76a75c9
+size 130973
diff --git a/OMRChecker/samples/community/Sandeep-1507/omr-3.png b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
new file mode 100644
index 0000000000000000000000000000000000000000..c414be1c6fa93808d3419715e2e610884e233926
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/omr-3.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:3976c705b45d889b06acc4bf82b27f129ad55c4875ee2aa6eb01aecf5a0250f6
+size 144032
diff --git a/OMRChecker/samples/community/Sandeep-1507/template.json b/OMRChecker/samples/community/Sandeep-1507/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bdb929ccd1a61c3ae54df4cdd6a8cf4281369733
--- /dev/null
+++ b/OMRChecker/samples/community/Sandeep-1507/template.json
@@ -0,0 +1,234 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 15,
+ 15
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Booklet_No": [
+ "b1..7"
+ ]
+ },
+ "fieldBlocks": {
+ "Booklet_No": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 112,
+ 530
+ ],
+ "fieldLabels": [
+ "b1..7"
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 28,
+ "labelsGap": 26.5
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 100
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 370
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q21..35"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 476,
+ 638
+ ]
+ },
+ "MCQBlock2a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 100
+ ]
+ },
+ "MCQBlock2a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 370
+ ]
+ },
+ "MCQBlock2a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q71..85"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 645,
+ 638
+ ]
+ },
+ "MCQBlock3a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q101..110"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 100
+ ]
+ },
+ "MCQBlock3a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q111..120"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 370
+ ]
+ },
+ "MCQBlock3a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q121..135"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 638
+ ]
+ },
+ "MCQBlock4a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q151..160"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 100
+ ]
+ },
+ "MCQBlock4a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q161..170"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 370
+ ]
+ },
+ "MCQBlock4a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q171..185"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 983,
+ 638
+ ]
+ },
+ "MCQBlock1a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q36..50"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 480,
+ 1061
+ ]
+ },
+ "MCQBlock2a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q86..100"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 648,
+ 1061
+ ]
+ },
+ "MCQBlock3a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q136..150"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.7,
+ "origin": [
+ 815,
+ 1061
+ ]
+ },
+ "MCQBlock4a": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q186..200"
+ ],
+ "bubblesGap": 28.7,
+ "labelsGap": 26.6,
+ "origin": [
+ 986,
+ 1061
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/Shamanth/omr_sheet_01.png b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
new file mode 100644
index 0000000000000000000000000000000000000000..7c317db90f80fdd8d3dcfbbadcb1f9b764e619e3
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/omr_sheet_01.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:9dd72b86ce9c6544f4a628187b4021df5746e9579222e7598c4d88846f2fbeb3
+size 107647
diff --git a/OMRChecker/samples/community/Shamanth/template.json b/OMRChecker/samples/community/Shamanth/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..7d8c34d5e4b7af5ca94415b5096c82300156bcbc
--- /dev/null
+++ b/OMRChecker/samples/community/Shamanth/template.json
@@ -0,0 +1,25 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 78,
+ 41
+ ],
+ "fieldLabels": [
+ "q21..28"
+ ],
+ "bubblesGap": 56,
+ "labelsGap": 46
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/answer_key.jpg b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..9d39b263e8fe5c23e391ad39bd0905efbf976038
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/answer_key.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:13bb31818680ca8497ee4c640a15d4acab2048cf11c973d758c8c75fd71c73be
+size 291737
diff --git a/OMRChecker/samples/community/UPSC-mock/config.json b/OMRChecker/samples/community/UPSC-mock/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..388060a33b04db4b550d0efecafd32aa277355d5
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 1800,
+ "display_width": 2400,
+ "processing_height": 2400,
+ "processing_width": 1800
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/evaluation.json b/OMRChecker/samples/community/UPSC-mock/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..3271cbd5e39161d428d765145f34d811701bdc64
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/evaluation.json
@@ -0,0 +1,18 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "answer_key_image_path": "answer_key.jpg",
+ "questions_in_order": [
+ "q1..100"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "2",
+ "incorrect": "-2/3",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..5dd9ae6adbd327c7995bfcd5f53989002e240cf2
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:020705f4618fd55a72dbc35719565307b844dbd28838c790fe858c3ebe5fb36e
+size 160882
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..c696b2e2d39118dec97d4f4c7479bfbc2dc6fc66
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1bb28daa10d276768d4fe3fd3f162ff8d3556cb805c4cf3f362e84d00d808095
+size 182385
diff --git a/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..edb7bbb600db8175e5beaa707c58d5a95972ec1e
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/scan-angles/angle-3.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:695e6cc5a497bcf114a101533ee0464a223ecb8f914bc28a8d356533bcf6e341
+size 179112
diff --git a/OMRChecker/samples/community/UPSC-mock/template.json b/OMRChecker/samples/community/UPSC-mock/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..2fadab79ca7f7f9adb03a85cde30f1ba2ccca588
--- /dev/null
+++ b/OMRChecker/samples/community/UPSC-mock/template.json
@@ -0,0 +1,195 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 30,
+ 25
+ ],
+ "customLabels": {
+ "Subject Code": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "Roll": [
+ "roll1..10"
+ ]
+ },
+ "fieldBlocks": {
+ "bookletNo": {
+ "origin": [
+ 595,
+ 545
+ ],
+ "bubblesGap": 68,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "bookletNo"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "C",
+ "D"
+ ],
+ "direction": "vertical"
+ },
+ "subjectCode": {
+ "origin": [
+ 912,
+ 512
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.5,
+ "fieldLabels": [
+ "subjectCode1",
+ "subjectCode2"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "roll": {
+ "origin": [
+ 1200,
+ 510
+ ],
+ "bubblesGap": 33,
+ "labelsGap": 42.8,
+ "fieldLabels": [
+ "roll1..10"
+ ],
+ "fieldType": "QTYPE_INT"
+ },
+ "q01block": {
+ "origin": [
+ 500,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 500,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 495,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1589
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1925
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1125,
+ 927
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1125,
+ 1258
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
new file mode 100644
index 0000000000000000000000000000000000000000..ffd67e704a23baa49bec476e04ba6b1796ff9b52
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/answer_key.csv
@@ -0,0 +1,200 @@
+q1,C
+q2,C
+q3,"D,E"
+q4,"A,AB"
+q5,"[['A', '1'], ['B', '2']]"
+q6,"['A', 'B']"
+q7,C
+q8,D
+q9,B
+q10,B
+q11,A
+q12,A
+q13,C
+q14,B
+q15,D
+q16,B
+q17,C
+q18,A
+q19,B
+q20,D
+q21,D
+q22,C
+q23,A
+q24,C
+q25,D
+q26,C
+q27,C
+q28,B
+q29,A
+q30,D
+q31,C
+q32,B
+q33,B
+q34,C
+q35,A
+q36,D
+q37,C
+q38,B
+q39,C
+q40,A
+q41,A
+q42,C
+q43,D
+q44,D
+q45,B
+q46,C
+q47,C
+q48,A
+q49,C
+q50,B
+q51,B
+q52,C
+q53,D
+q54,C
+q55,B
+q56,B
+q57,A
+q58,A
+q59,D
+q60,C
+q61,C
+q62,B
+q63,A
+q64,C
+q65,D
+q66,C
+q67,B
+q68,A
+q69,B
+q70,B
+q71,C
+q72,B
+q73,C
+q74,A
+q75,A
+q76,C
+q77,D
+q78,D
+q79,B
+q80,A
+q81,B
+q82,C
+q83,D
+q84,C
+q85,A
+q86,C
+q87,D
+q88,B
+q89,C
+q90,B
+q91,B
+q92,A
+q93,C
+q94,D
+q95,C
+q96,B
+q97,B
+q98,A
+q99,A
+q100,A
+q101,A
+q102,B
+q103,C
+q104,C
+q105,A
+q106,D
+q107,B
+q108,A
+q109,C
+q110,B
+q111,B
+q112,C
+q113,C
+q114,B
+q115,D
+q116,B
+q117,A
+q118,C
+q119,D
+q120,C
+q121,C
+q122,A
+q123,B
+q124,C
+q125,D
+q126,C
+q127,C
+q128,D
+q129,D
+q130,A
+q131,A
+q132,C
+q133,B
+q134,C
+q135,D
+q136,B
+q137,C
+q138,A
+q139,B
+q140,D
+q141,D
+q142,C
+q143,D
+q144,A
+q145,A
+q146,C
+q147,A
+q148,D
+q149,D
+q150,B
+q151,A
+q152,B
+q153,B
+q154,D
+q155,D
+q156,B
+q157,A
+q158,B
+q159,A
+q160,C
+q161,D
+q162,C
+q163,A
+q164,B
+q165,D
+q166,D
+q167,C
+q168,C
+q169,C
+q170,D
+q171,A
+q172,A
+q173,C
+q174,C
+q175,B
+q176,D
+q177,A
+q178,B
+q179,B
+q180,C
+q181,D
+q182,C
+q183,B
+q184,B
+q185,C
+q186,D
+q187,D
+q188,A
+q189,A
+q190,B
+q191,C
+q192,B
+q193,D
+q194,C
+q195,B
+q196,B
+q197,A
+q198,B
+q199,B
+q200,A
diff --git a/OMRChecker/samples/community/UmarFarootAPS/config.json b/OMRChecker/samples/community/UmarFarootAPS/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..5a9d75731b7f5ae8c6fd803a7426b29ac39c6bfb
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 960,
+ "display_width": 1280,
+ "processing_height": 1640,
+ "processing_width": 1332
+ },
+ "outputs": {
+ "show_image_level": 0
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/evaluation.json b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..69b0ef8c94df580b6190a41bb1eeacba73323762
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/evaluation.json
@@ -0,0 +1,14 @@
+{
+ "source_type": "csv",
+ "options": {
+ "answer_key_csv_path": "answer_key.csv",
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/community/UmarFarootAPS/omr_marker.jpg differ
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..73d60b9caa9860aa6a9bb75956fd3430b1aee44e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:49e1a14e0c1be809282b6e43c913e17598380a33f607a5c46723764c32e17b59
+size 304722
diff --git a/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..25d75107cf4885f2974c52cf42003255be285a3e
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/scans/scan-type-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18b85df487c0bc98eb9139949c3fbca2bad57e8f224fe48b495e0951660f65d
+size 237392
diff --git a/OMRChecker/samples/community/UmarFarootAPS/template.json b/OMRChecker/samples/community/UmarFarootAPS/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..f303edabdf3bc64c41c50152ae72d09f5a0de913
--- /dev/null
+++ b/OMRChecker/samples/community/UmarFarootAPS/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 2550,
+ 3300
+ ],
+ "bubbleDimensions": [
+ 32,
+ 32
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll_no": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ]
+ },
+ "fieldBlocks": {
+ "Roll_no": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 2169,
+ 180
+ ],
+ "fieldLabels": [
+ "r1",
+ "r2",
+ "r3",
+ "r4"
+ ],
+ "bubblesGap": 61,
+ "labelsGap": 93
+ },
+ "MCQBlock1a1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q1..17"
+ ]
+ },
+ "MCQBlock1a2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q18..34"
+ ]
+ },
+ "MCQBlock1a3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 197,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q35..50"
+ ]
+ },
+ "MCQBlock1a4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q51..67"
+ ]
+ },
+ "MCQBlock1a5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q68..84"
+ ]
+ },
+ "MCQBlock1a6": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 725,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q85..100"
+ ]
+ },
+ "MCQBlock1a7": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q101..117"
+ ]
+ },
+ "MCQBlock1a8": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q118..134"
+ ]
+ },
+ "MCQBlock1a9": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1250,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q135..150"
+ ]
+ },
+ "MCQBlock1a10": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 300
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q151..167"
+ ]
+ },
+ "MCQBlock1a11": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 1310
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q168..184"
+ ]
+ },
+ "MCQBlock1a12": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1770,
+ 2316
+ ],
+ "bubblesGap": 92,
+ "labelsGap": 59.6,
+ "fieldLabels": [
+ "q185..200"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..b9d70c6be1aa35398e2ac125f3e7bea0ff924fd0
Binary files /dev/null and b/OMRChecker/samples/community/dxuian/omrcollegesheet.jpg differ
diff --git a/OMRChecker/samples/community/dxuian/template.json b/OMRChecker/samples/community/dxuian/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e055f557537fa531a0546c2155146763c8d937a6
--- /dev/null
+++ b/OMRChecker/samples/community/dxuian/template.json
@@ -0,0 +1,48 @@
+{
+ "pageDimensions": [707, 484],
+ "bubbleDimensions": [15, 10],
+ "fieldBlocks": {
+ "Column1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [82, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q1", "Q2", "Q3", "Q4", "Q5", "Q6", "Q7", "Q8", "Q9", "Q10", "Q11", "Q12", "Q13", "Q14", "Q15", "Q16", "Q17", "Q18", "Q19", "Q20"]
+ },
+ "Column2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [205, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q21", "Q22", "Q23", "Q24", "Q25", "Q26", "Q27", "Q28", "Q29", "Q30", "Q31", "Q32", "Q33", "Q34", "Q35", "Q36", "Q37", "Q38", "Q39", "Q40"]
+ },
+ "Column3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [327, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q41", "Q42", "Q43", "Q44", "Q45", "Q46", "Q47", "Q48", "Q49", "Q50", "Q51", "Q52", "Q53", "Q54", "Q55", "Q56", "Q57", "Q58", "Q59", "Q60"]
+ },
+ "Column4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [450, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q61", "Q62", "Q63", "Q64", "Q65", "Q66", "Q67", "Q68", "Q69", "Q70", "Q71", "Q72", "Q73", "Q74", "Q75", "Q76", "Q77", "Q78", "Q79", "Q80"]
+ },
+ "Column5": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [573, 35],
+ "bubblesGap": 21,
+ "labelsGap": 22.7,
+ "bubbleCount": 20,
+ "fieldLabels": ["Q81", "Q82", "Q83", "Q84", "Q85", "Q86", "Q87", "Q88", "Q89", "Q90", "Q91", "Q92", "Q93", "Q94", "Q95", "Q96", "Q97", "Q98", "Q99", "Q100"]
+ }
+ },
+
+ "emptyValue": "-"
+}
\ No newline at end of file
diff --git a/OMRChecker/samples/community/ibrahimkilic/template.json b/OMRChecker/samples/community/ibrahimkilic/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..3064ee19192fc7474370c0f518a51bea9a30b639
--- /dev/null
+++ b/OMRChecker/samples/community/ibrahimkilic/template.json
@@ -0,0 +1,30 @@
+{
+ "pageDimensions": [
+ 299,
+ 328
+ ],
+ "bubbleDimensions": [
+ 20,
+ 20
+ ],
+ "emptyValue": "no",
+ "fieldBlocks": {
+ "YesNoBlock1": {
+ "direction": "horizontal",
+ "bubbleValues": [
+ "yes"
+ ],
+ "origin": [
+ 15,
+ 55
+ ],
+ "emptyValue": "no",
+ "bubblesGap": 48,
+ "labelsGap": 48,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ }
+ },
+ "preProcessors": []
+}
diff --git a/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..e9436f41ee8db1269e2255ee902b73de2525d502
Binary files /dev/null and b/OMRChecker/samples/community/ibrahimkilic/yes_no_questionnarie.jpg differ
diff --git a/OMRChecker/samples/community/samuelIkoli/template.json b/OMRChecker/samples/community/samuelIkoli/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..19c46f7c96bb07cc0a5a1731bbb8febeba9686e9
--- /dev/null
+++ b/OMRChecker/samples/community/samuelIkoli/template.json
@@ -0,0 +1,28 @@
+{
+ "pageDimensions": [630, 404],
+ "bubbleDimensions": [20, 15],
+ "customLabels": {},
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [33, 6],
+ "fieldLabels": ["q1", "q2", "q3", "q4", "q5", "q6", "q7", "q8", "q9", "q10", "q11", "q12", "q13", "q14", "q15", "q16", "q17", "q18", "q19", "q20"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock2": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [248, 6],
+ "fieldLabels": ["q21", "q22", "q23", "q24", "q25", "q26", "q27", "q28", "q29", "q30", "q31", "q32", "q33", "q34", "q35", "q36", "q37", "q38", "q39", "q40"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ },
+ "MCQBlock3": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [465, 6],
+ "fieldLabels": ["q41", "q42", "q43", "q44", "q45", "q46", "q47", "q48", "q49", "q50", "q51", "q52", "q53", "q54", "q55", "q56", "q57", "q58", "q59", "q60"],
+ "bubblesGap": 33,
+ "labelsGap": 20
+ }
+ }
+}
diff --git a/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg
new file mode 100644
index 0000000000000000000000000000000000000000..fcd61e808e4a2f33095ea0dbef0ec27741450465
Binary files /dev/null and b/OMRChecker/samples/community/samuelIkoli/waec_sample.jpeg differ
diff --git a/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6626eb476d38cb16156dc15a48d30bcd41ebaac1
--- /dev/null
+++ b/OMRChecker/samples/sample1/MobileCamera/sheet1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:1be7b0fc432e811fd07b07fe325d9ca8d0d97782eea9f95ed62ff0e47076e1cd
+size 162012
diff --git a/OMRChecker/samples/sample1/config.json b/OMRChecker/samples/sample1/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample1/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample1/omr_marker.jpg b/OMRChecker/samples/sample1/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample1/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample1/template.json b/OMRChecker/samples/sample1/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..e4795ad56c919618354f18d4de7a14f14642df07
--- /dev/null
+++ b/OMRChecker/samples/sample1/template.json
@@ -0,0 +1,197 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q5": [
+ "q5_1",
+ "q5_2"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubblesGap": 41,
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "fieldLabels": [
+ "Medium"
+ ],
+ "labelsGap": 0,
+ "origin": [
+ 170,
+ 282
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "roll1..9"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 58,
+ "origin": [
+ 225,
+ 282
+ ]
+ },
+ "Int_Block_Q5": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q5_1",
+ "q5_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 903,
+ 282
+ ]
+ },
+ "Int_Block_Q6": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1077,
+ 282
+ ]
+ },
+ "Int_Block_Q7": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 60,
+ "origin": [
+ 1240,
+ 282
+ ]
+ },
+ "Int_Block_Q8": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1410,
+ 282
+ ]
+ },
+ "Int_Block_Q9": {
+ "fieldType": "QTYPE_INT",
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "origin": [
+ 1580,
+ 282
+ ]
+ },
+ "MCQ_Block_Q1": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q1..4"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 860
+ ]
+ },
+ "MCQ_Block_Q10": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q10..13"
+ ],
+ "bubblesGap": 59,
+ "labelsGap": 50,
+ "origin": [
+ 121,
+ 1195
+ ]
+ },
+ "MCQ_Block_Q14": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q14..16"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 860
+ ]
+ },
+ "MCQ_Block_Q17": {
+ "fieldType": "QTYPE_MCQ4",
+ "fieldLabels": [
+ "q17..20"
+ ],
+ "bubblesGap": 57,
+ "labelsGap": 50,
+ "origin": [
+ 905,
+ 1195
+ ]
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ },
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
new file mode 100644
index 0000000000000000000000000000000000000000..215ef154d463851576758b4eeababd196359e9ac
--- /dev/null
+++ b/OMRChecker/samples/sample2/AdrianSample/adrian_omr.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:85b11881d4b3a62002dfb411a2a4fa1b5ee3816d4d26e07924681e29c3fe68e4
+size 105167
diff --git a/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png
new file mode 100644
index 0000000000000000000000000000000000000000..d8db0994df2dcfaeb67ff667abd2edb55b47f927
Binary files /dev/null and b/OMRChecker/samples/sample2/AdrianSample/adrian_omr_2.png differ
diff --git a/OMRChecker/samples/sample2/config.json b/OMRChecker/samples/sample2/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample2/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample2/template.json b/OMRChecker/samples/sample2/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..25db408b27c65a7ec5b570507403b9e439a26452
--- /dev/null
+++ b/OMRChecker/samples/sample2/template.json
@@ -0,0 +1,35 @@
+{
+ "pageDimensions": [
+ 300,
+ 400
+ ],
+ "bubbleDimensions": [
+ 25,
+ 25
+ ],
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQ_Block_1": {
+ "fieldType": "QTYPE_MCQ5",
+ "origin": [
+ 65,
+ 60
+ ],
+ "fieldLabels": [
+ "q1..5"
+ ],
+ "labelsGap": 52,
+ "bubblesGap": 41
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample3/README.md b/OMRChecker/samples/sample3/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..4b1e325dfa284cf676a1f3d9e37d426f5d20ebb6
--- /dev/null
+++ b/OMRChecker/samples/sample3/README.md
@@ -0,0 +1,31 @@
+## Observation
+The OMR layout is slightly different on colored thick papers vs on xeroxed thin papers. The shift becomes noticible in case of OMR with large number of questions.
+
+We overlapped a colored OMR sheet with a xerox copy of the same OMR sheet(both printed on A4 papers) and observed that there is a great amount of layout sheet as we reach the bottom of the OMR.
+
+Link to an explainer with a real life example: [Google Drive](https://drive.google.com/drive/folders/1GpZTmpEhEjSALJEMjHwDafzWKgEoCTOI?usp=sharing)
+
+## Reasons for shifts in Template layout:
+Listing out a few reasons for the above observation:
+### Printer margin setting
+The margin settings for different printers may be different for the same OMR layout. Thus causing the print to become elongated either horizontally, vertically or both ways.
+
+### The Fan-out effect
+The fan-out effect is usually observed in a sheet fed offset press. Depending on how the papers are made, their dimensions have a tendency to change when they are exposed to moisture or water.
+
+The standard office papers(80 gsm) can easily capture moisture and change shape e.g. in case they get stored for multiple days in a place where the weather is highly humid.
+
+Below are some examples of the GSM ranges:
+
+- 74gsm to 90gsm – This is the basic standard office paper, used in your laser printers.
+- 100gsm to 120gsm – This is stationary paper used for standard letterheads, complimentary slips.
+- 130 to 170gsm – Mostly used for leaflets, posters, single-sided flyers, and brochures.
+
+## Solution
+
+It is recommended to scan each types of prints into different folders and use a separate template.json layout for each of the folders. The same is presented in this sample folder.
+
+## References
+
+- [Paper dimensional stability in sheet-fed offset printing](https://www.diva-portal.org/smash/get/diva2:517895/FULLTEXT01.pdf)
+- An analysis of a few ["Interesting" bubble sheets](https://imgur.com/a/10qwL)
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..4877db1feb1c758bc35b41f543dc9bb14d6ca62f
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/rgb-100-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:e18eba01e5d0c466b4b750aa20e710e7354489ed9bef91966c2349007fee349d
+size 137681
diff --git a/OMRChecker/samples/sample3/colored-thick-sheet/template.json b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ab35b30f1e3ec35d1c4698cc68b76c4a47d6eb37
--- /dev/null
+++ b/OMRChecker/samples/sample3/colored-thick-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 504,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 504,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 500,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 500,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 811,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 811,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 811,
+ 1562
+ ],
+ "bubblesGap": 61.25,
+ "labelsGap": 32.5,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 811,
+ 1885
+ ],
+ "bubblesGap": 62.25,
+ "labelsGap": 33.5,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1120,
+ 927
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1120,
+ 1242
+ ],
+ "bubblesGap": 60.35,
+ "labelsGap": 31.75,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample3/config.json b/OMRChecker/samples/sample3/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample3/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..2be38bbfb8e8075f0da00d5fb56d4689d2e72e26
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/grayscale-80-gsm.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:eab4df0efac2614b2deddcadc0b754aa72c668170e5d9af49ad77c877198556c
+size 187367
diff --git a/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd499ade7e54d5061817985e40effb13fce8f012
--- /dev/null
+++ b/OMRChecker/samples/sample3/xeroxed-thin-sheet/template.json
@@ -0,0 +1,143 @@
+{
+ "pageDimensions": [
+ 1800,
+ 2400
+ ],
+ "bubbleDimensions": [
+ 23,
+ 20
+ ],
+ "fieldBlocks": {
+ "q01block": {
+ "origin": [
+ 492,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q1..10"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q11block": {
+ "origin": [
+ 492,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q11..20"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q21block": {
+ "origin": [
+ 492,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q21..30"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q31block": {
+ "origin": [
+ 487,
+ 1920
+ ],
+ "bubblesGap": 61.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q31..40"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q41block": {
+ "origin": [
+ 807,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q41..50"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q51block": {
+ "origin": [
+ 803,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q51..60"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q61block": {
+ "origin": [
+ 803,
+ 1589
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q61..70"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q71block": {
+ "origin": [
+ 803,
+ 1920
+ ],
+ "bubblesGap": 60.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q71..80"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q81block": {
+ "origin": [
+ 1115,
+ 924
+ ],
+ "bubblesGap": 58.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q81..90"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ },
+ "q91block": {
+ "origin": [
+ 1115,
+ 1258
+ ],
+ "bubblesGap": 59.75,
+ "labelsGap": 32.65,
+ "fieldLabels": [
+ "q91..100"
+ ],
+ "fieldType": "QTYPE_MCQ4"
+ }
+ },
+ "preProcessors": [
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ]
+}
diff --git a/OMRChecker/samples/sample4/IMG_20201116_143512.jpg b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..6c817d4f620b6bc728eeb410bd9c1da8d27a5cfe
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_143512.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0b1f0e0afd8a2c40deb7f7c992c9fb6046eb1b28a5f9a674e80ce3418c7fc307
+size 421479
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..023f66bc9eb9100a8500d84b82f7725141962082
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150717658.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8c3f847e6c37c96562ee17b3d69cb919bb52743f8b95d02679ed62e20dbd5096
+size 386960
diff --git a/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..21869f5c27011e067f1fd8cb3a4e6ecb0e4c7e11
--- /dev/null
+++ b/OMRChecker/samples/sample4/IMG_20201116_150750830.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0bdc5eca48c13d19f228ef7252b5e855b5c62926ba1e9e3297fc6a46e074b1be
+size 307198
diff --git a/OMRChecker/samples/sample4/config.json b/OMRChecker/samples/sample4/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..6c82782a3f6486145d6367eabe6910cca5268214
--- /dev/null
+++ b/OMRChecker/samples/sample4/config.json
@@ -0,0 +1,13 @@
+{
+ "dimensions": {
+ "display_width": 1189,
+ "display_height": 1682
+ },
+ "threshold_params": {
+ "MIN_JUMP": 30
+ },
+ "outputs": {
+ "filter_out_multimarked_files": false,
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample4/evaluation.json b/OMRChecker/samples/sample4/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..8ea935e70ded2a9c53c13bf22b5904f1ef3fa923
--- /dev/null
+++ b/OMRChecker/samples/sample4/evaluation.json
@@ -0,0 +1,34 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..11"
+ ],
+ "answers_in_order": [
+ "B",
+ "D",
+ "C",
+ "B",
+ "D",
+ "C",
+ [
+ "B",
+ "C",
+ "BC"
+ ],
+ "A",
+ "C",
+ "D",
+ "C"
+ ],
+ "should_explain_scoring": true,
+ "enable_evaluation_table_to_csv": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "3",
+ "incorrect": "-1",
+ "unmarked": "0"
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample4/template.json b/OMRChecker/samples/sample4/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..d1bef2985939f8ea9c01bfdec28e0d5b6bab11a3
--- /dev/null
+++ b/OMRChecker/samples/sample4/template.json
@@ -0,0 +1,45 @@
+{
+ "pageDimensions": [
+ 1189,
+ 1682
+ ],
+ "bubbleDimensions": [
+ 30,
+ 30
+ ],
+ "preProcessors": [
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ },
+ {
+ "name": "CropPage",
+ "options": {
+ "morphKernel": [
+ 10,
+ 10
+ ]
+ }
+ }
+ ],
+ "fieldBlocks": {
+ "MCQBlock1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 134,
+ 684
+ ],
+ "fieldLabels": [
+ "q1..11"
+ ],
+ "bubblesGap": 79,
+ "labelsGap": 62
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/README.md b/OMRChecker/samples/sample5/README.md
new file mode 100644
index 0000000000000000000000000000000000000000..b70cedd6cbf22de260649c08e855d929d5e8ae87
--- /dev/null
+++ b/OMRChecker/samples/sample5/README.md
@@ -0,0 +1,6 @@
+### OMRChecker Sample
+
+This sample demonstrates multiple things, namely -
+- Running OMRChecker on images scanned using popular document scanning apps
+- Using a common template.json file for sub-folders (e.g. multiple scan batches)
+- Using evaluation.json file with custom marking (without streak-based marking)
diff --git a/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..72b450e07dd7d4c4a18287183128ad7edbf5607a
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch1/camscanner-1.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:08bb2e9fecf23be6850485cb7776a604aa2efe9800855ea763dee7ade5362323
+size 127651
diff --git a/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..46fb2ac36c0d5c2366fdf9a7ed56eb933578a60f
--- /dev/null
+++ b/OMRChecker/samples/sample5/ScanBatch2/camscanner-2.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:0da09483868ad499925fefa284153e9544822894335e16cea859057740c3c063
+size 127863
diff --git a/OMRChecker/samples/sample5/config.json b/OMRChecker/samples/sample5/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..086b4c5961137266d3ad134250638930dd855296
--- /dev/null
+++ b/OMRChecker/samples/sample5/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample5/evaluation.json b/OMRChecker/samples/sample5/evaluation.json
new file mode 100644
index 0000000000000000000000000000000000000000..706e49ee9d9e7d445ca98ea87a1a140ce53da54d
--- /dev/null
+++ b/OMRChecker/samples/sample5/evaluation.json
@@ -0,0 +1,93 @@
+{
+ "source_type": "custom",
+ "options": {
+ "questions_in_order": [
+ "q1..22"
+ ],
+ "answers_in_order": [
+ "C",
+ "C",
+ "B",
+ "C",
+ "C",
+ [
+ "1",
+ "01"
+ ],
+ "19",
+ "10",
+ "10",
+ "18",
+ "D",
+ "A",
+ "D",
+ "D",
+ "D",
+ "C",
+ "C",
+ "C",
+ "C",
+ "D",
+ "B",
+ "A"
+ ],
+ "should_explain_scoring": true
+ },
+ "marking_schemes": {
+ "DEFAULT": {
+ "correct": "1",
+ "incorrect": "0",
+ "unmarked": "0"
+ },
+ "BOOMERANG_1": {
+ "questions": [
+ "q1..5"
+ ],
+ "marking": {
+ "correct": 4,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "PROXIMITY_1": {
+ "questions": [
+ "q6..10"
+ ],
+ "marking": {
+ "correct": 3,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_1": {
+ "questions": [
+ "q11..14"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ },
+ "POWER_SECTION_1": {
+ "questions": [
+ "q15..18"
+ ],
+ "marking": {
+ "correct": 1,
+ "incorrect": 0,
+ "unmarked": 0
+ }
+ },
+ "FIBONACCI_SECTION_2": {
+ "questions": [
+ "q19..22"
+ ],
+ "marking": {
+ "correct": 2,
+ "incorrect": -1,
+ "unmarked": 0
+ }
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample5/omr_marker.jpg b/OMRChecker/samples/sample5/omr_marker.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..0929feec8c97fc00e6f3e55a74eb826f593379e3
Binary files /dev/null and b/OMRChecker/samples/sample5/omr_marker.jpg differ
diff --git a/OMRChecker/samples/sample5/template.json b/OMRChecker/samples/sample5/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..ac2436c02a8a19024e8050ff7c02dca9aa7ea016
--- /dev/null
+++ b/OMRChecker/samples/sample5/template.json
@@ -0,0 +1,188 @@
+{
+ "pageDimensions": [
+ 1846,
+ 1500
+ ],
+ "bubbleDimensions": [
+ 40,
+ 40
+ ],
+ "preProcessors": [
+ {
+ "name": "CropOnMarkers",
+ "options": {
+ "relativePath": "omr_marker.jpg",
+ "sheetToMarkerWidthRatio": 17
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "Medium",
+ "roll1..9"
+ ],
+ "q6": [
+ "q6_1",
+ "q6_2"
+ ],
+ "q7": [
+ "q7_1",
+ "q7_2"
+ ],
+ "q8": [
+ "q8_1",
+ "q8_2"
+ ],
+ "q9": [
+ "q9_1",
+ "q9_2"
+ ],
+ "q10": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Medium": {
+ "bubbleValues": [
+ "E",
+ "H"
+ ],
+ "direction": "vertical",
+ "origin": [
+ 200,
+ 215
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 0,
+ "fieldLabels": [
+ "Medium"
+ ]
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 261,
+ 210
+ ],
+ "bubblesGap": 46.5,
+ "labelsGap": 58,
+ "fieldLabels": [
+ "roll1..9"
+ ]
+ },
+ "Int1": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 935,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q6_1",
+ "q6_2"
+ ]
+ },
+ "Int2": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1100,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q7_1",
+ "q7_2"
+ ]
+ },
+ "Int3": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1275,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q8_1",
+ "q8_2"
+ ]
+ },
+ "Int4": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1449,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q9_1",
+ "q9_2"
+ ]
+ },
+ "Int5": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1620,
+ 211
+ ],
+ "bubblesGap": 46,
+ "labelsGap": 57,
+ "fieldLabels": [
+ "q10_1",
+ "q10_2"
+ ]
+ },
+ "Mcq1": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 198,
+ 826
+ ],
+ "bubblesGap": 93,
+ "labelsGap": 62,
+ "fieldLabels": [
+ "q1..5"
+ ]
+ },
+ "Mcq2": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 830
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q11..14"
+ ]
+ },
+ "Mcq3": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 833,
+ 1270
+ ],
+ "bubblesGap": 71,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q15..18"
+ ]
+ },
+ "Mcq4": {
+ "fieldType": "QTYPE_MCQ4",
+ "origin": [
+ 1481,
+ 830
+ ],
+ "bubblesGap": 73,
+ "labelsGap": 61,
+ "fieldLabels": [
+ "q19..22"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/config.json b/OMRChecker/samples/sample6/config.json
new file mode 100644
index 0000000000000000000000000000000000000000..2ac30e718a44f7336aafda1b0f57aec1320fc72c
--- /dev/null
+++ b/OMRChecker/samples/sample6/config.json
@@ -0,0 +1,11 @@
+{
+ "dimensions": {
+ "display_width": 2480,
+ "display_height": 3508,
+ "processing_width": 1653,
+ "processing_height": 2339
+ },
+ "outputs": {
+ "show_image_level": 5
+ }
+}
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..fdaa0f4112e15037a31bc90da204b7c19114820b
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_01.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:de8a5ed5b7360a71db83b49a3d0d6e1af1482339d1116286a064d285be42e37e
+size 246590
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..bf56d121c1061302fc4ce87fca61bb8584ea9e4a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_02.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:aca51de61ba3ee76e00e22a666d8cd2b72190af674fa051c9cf70346e9ee4d0a
+size 244049
diff --git a/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
new file mode 100644
index 0000000000000000000000000000000000000000..22bfa55de7a687729b733a615ad78dad64ac467a
--- /dev/null
+++ b/OMRChecker/samples/sample6/doc-scans/sample_roll_03.jpg
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:950dbf86fc152f464be30ebaeb518c61067b0c494ab5fd70f05de10b5dfabb12
+size 244321
diff --git a/OMRChecker/samples/sample6/readme.md b/OMRChecker/samples/sample6/readme.md
new file mode 100644
index 0000000000000000000000000000000000000000..d8491b2a049eb4e721e007e8e8bc4fe5ca536802
--- /dev/null
+++ b/OMRChecker/samples/sample6/readme.md
@@ -0,0 +1,21 @@
+# Demo for feature-based alignment
+
+## Background
+OMR is used to match student roll on final exam scripts. Scripts are scanned using a document scanner and the cover pages are extracted for OMR. Even though a document scanner does not produce any warpped perspective, the alignment is not perfect, causing some rotation and translation in the scans.
+
+The scripts in this sample were specifically selected incorrectly marked scripts to demonstrate how feature-based alignment can correct transformation errors using a reference image. In the actual batch. 156 out of 532 scripts were incorrectly marked. With feature-based alignment, all scripts were correctly marked.
+
+## Usage
+Two template files are given in the sample folder, one with feature-based alignment (template_fb_align), the other without (template_no_fb_align).
+
+## Additional Notes
+
+### Reference Image
+When using a reference image for feature-based alignment, it is better not to have many repeated patterns as it is causes ambiguity when trying to match similar feature points. The bubbles in an OMR form are identical and should not be used for feature-extraction.
+
+Thus, the reference image should be cleared of any bubbles. Forms with lots of text as in this example would be effective.
+
+Note the reference image in this sample was generated from a vector pdf, and not from a scanned blank, producing in a perfectly aligned reference.
+
+### Level adjustment
+The bubbles on the scripts were not shaded dark enough. Thus, a level adjustment was done to bring the black point to 70% to darken the light shading. White point was brought down to 80% to remove the light-grey background in the columns.
diff --git a/OMRChecker/samples/sample6/reference.png b/OMRChecker/samples/sample6/reference.png
new file mode 100644
index 0000000000000000000000000000000000000000..e2e692f4638e9c7fa2d7d159e044c778b18acdc9
--- /dev/null
+++ b/OMRChecker/samples/sample6/reference.png
@@ -0,0 +1,3 @@
+version https://git-lfs.github.com/spec/v1
+oid sha256:8379ba055209c065def041fc1fb1cea444b913d835c1cfeffa40091446e5e3e5
+size 178069
diff --git a/OMRChecker/samples/sample6/template.json b/OMRChecker/samples/sample6/template.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_fb_align.json b/OMRChecker/samples/sample6/template_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..bd45b9c8fd3ba024b951ff4e7965c98bb56211ee
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_fb_align.json
@@ -0,0 +1,118 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "FeatureBasedAlignment",
+ "options": {
+ "reference": "reference.png",
+ "maxFeatures": 1000,
+ "2d": true
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/samples/sample6/template_no_fb_align.json b/OMRChecker/samples/sample6/template_no_fb_align.json
new file mode 100644
index 0000000000000000000000000000000000000000..80e8914dd4fd04814fd96ecededa6c8fd06a1c4c
--- /dev/null
+++ b/OMRChecker/samples/sample6/template_no_fb_align.json
@@ -0,0 +1,110 @@
+{
+ "pageDimensions": [
+ 2480,
+ 3508
+ ],
+ "bubbleDimensions": [
+ 42,
+ 42
+ ],
+ "preProcessors": [
+ {
+ "name": "Levels",
+ "options": {
+ "low": 0.7,
+ "high": 0.8
+ }
+ },
+ {
+ "name": "GaussianBlur",
+ "options": {
+ "kSize": [
+ 3,
+ 3
+ ],
+ "sigmaX": 0
+ }
+ }
+ ],
+ "customLabels": {
+ "Roll": [
+ "stu",
+ "roll1..7",
+ "check_1",
+ "check_2"
+ ]
+ },
+ "fieldBlocks": {
+ "Check1": {
+ "origin": [
+ 2033,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_1"
+ ],
+ "bubbleValues": [
+ "A",
+ "B",
+ "E",
+ "H",
+ "J",
+ "L",
+ "M"
+ ],
+ "direction": "vertical"
+ },
+ "Check2": {
+ "origin": [
+ 2083,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "check_2"
+ ],
+ "bubbleValues": [
+ "N",
+ "R",
+ "U",
+ "W",
+ "X",
+ "Y"
+ ],
+ "direction": "vertical"
+ },
+ "Stu": {
+ "origin": [
+ 1636,
+ 1290
+ ],
+ "bubblesGap": 50,
+ "labelsGap": 50,
+ "fieldLabels": [
+ "stu"
+ ],
+ "bubbleValues": [
+ "U",
+ "A",
+ "HT",
+ "GT"
+ ],
+ "direction": "vertical"
+ },
+ "Roll": {
+ "fieldType": "QTYPE_INT",
+ "origin": [
+ 1685,
+ 1290
+ ],
+ "bubblesGap": 50.5,
+ "labelsGap": 50.5,
+ "fieldLabels": [
+ "roll1..7"
+ ]
+ }
+ }
+}
diff --git a/OMRChecker/src/__init__.py b/OMRChecker/src/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..5c1baacaa53bb1b4219d33eee566c6a613ab88d6
--- /dev/null
+++ b/OMRChecker/src/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+from src.logger import logger
+
+# It takes a few seconds for the imports
+logger.info(f"Loading OMRChecker modules...")
diff --git a/OMRChecker/src/constants.py b/OMRChecker/src/constants.py
new file mode 100644
index 0000000000000000000000000000000000000000..628eb75ac892ddc184cc99429036aa9196889c83
--- /dev/null
+++ b/OMRChecker/src/constants.py
@@ -0,0 +1,54 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+from dotmap import DotMap
+
+# Filenames
+TEMPLATE_FILENAME = "template.json"
+EVALUATION_FILENAME = "evaluation.json"
+CONFIG_FILENAME = "config.json"
+
+FIELD_LABEL_NUMBER_REGEX = r"([^\d]+)(\d*)"
+#
+ERROR_CODES = DotMap(
+ {
+ "MULTI_BUBBLE_WARN": 1,
+ "NO_MARKER_ERR": 2,
+ },
+ _dynamic=False,
+)
+
+FIELD_TYPES = {
+ "QTYPE_INT": {
+ "bubbleValues": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"],
+ "direction": "vertical",
+ },
+ "QTYPE_INT_FROM_1": {
+ "bubbleValues": ["1", "2", "3", "4", "5", "6", "7", "8", "9", "0"],
+ "direction": "vertical",
+ },
+ "QTYPE_MCQ4": {"bubbleValues": ["A", "B", "C", "D"], "direction": "horizontal"},
+ "QTYPE_MCQ5": {
+ "bubbleValues": ["A", "B", "C", "D", "E"],
+ "direction": "horizontal",
+ },
+ #
+ # You can create and append custom field types here-
+ #
+}
+
+# TODO: move to interaction.py
+TEXT_SIZE = 0.95
+CLR_BLACK = (50, 150, 150)
+CLR_WHITE = (250, 250, 250)
+CLR_GRAY = (130, 130, 130)
+CLR_DARK_GRAY = (100, 100, 100)
+
+# TODO: move to config.json
+GLOBAL_PAGE_THRESHOLD_WHITE = 200
+GLOBAL_PAGE_THRESHOLD_BLACK = 100
diff --git a/OMRChecker/src/core.py b/OMRChecker/src/core.py
new file mode 100644
index 0000000000000000000000000000000000000000..15279cf49fb9fb12c88611bcc69611ea524344dc
--- /dev/null
+++ b/OMRChecker/src/core.py
@@ -0,0 +1,721 @@
+import os
+from collections import defaultdict
+from typing import Any
+
+import cv2
+import matplotlib.pyplot as plt
+import numpy as np
+
+import src.constants as constants
+from src.logger import logger
+from src.utils.image import CLAHE_HELPER, ImageUtils
+from src.utils.interaction import InteractionUtils
+
+
+class ImageInstanceOps:
+ """Class to hold fine-tuned utilities for a group of images. One instance for each processing directory."""
+
+ save_img_list: Any = defaultdict(list)
+
+ def __init__(self, tuning_config):
+ super().__init__()
+ self.tuning_config = tuning_config
+ self.save_image_level = tuning_config.outputs.save_image_level
+
+ def apply_preprocessors(self, file_path, in_omr, template):
+ tuning_config = self.tuning_config
+ # resize to conform to template
+ in_omr = ImageUtils.resize_util(
+ in_omr,
+ tuning_config.dimensions.processing_width,
+ tuning_config.dimensions.processing_height,
+ )
+
+ # run pre_processors in sequence
+ for pre_processor in template.pre_processors:
+ in_omr = pre_processor.apply_filter(in_omr, file_path)
+ return in_omr
+
+ def read_omr_response(self, template, image, name, save_dir=None):
+ config = self.tuning_config
+ auto_align = config.alignment_params.auto_align
+ try:
+ img = image.copy()
+ # origDim = img.shape[:2]
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ if img.max() > img.min():
+ img = ImageUtils.normalize_util(img)
+ # Processing copies
+ transp_layer = img.copy()
+ final_marked = img.copy()
+
+ morph = img.copy()
+ self.append_save_img(3, morph)
+
+ if auto_align:
+ # Note: clahe is good for morphology, bad for thresholding
+ morph = CLAHE_HELPER.apply(morph)
+ self.append_save_img(3, morph)
+ # Remove shadows further, make columns/boxes darker (less gamma)
+ morph = ImageUtils.adjust_gamma(
+ morph, config.threshold_params.GAMMA_LOW
+ )
+ # TODO: all numbers should come from either constants or config
+ _, morph = cv2.threshold(morph, 220, 220, cv2.THRESH_TRUNC)
+ morph = ImageUtils.normalize_util(morph)
+ self.append_save_img(3, morph)
+ if config.outputs.show_image_level >= 4:
+ InteractionUtils.show("morph1", morph, 0, 1, config)
+
+ # Move them to data class if needed
+ # Overlay Transparencies
+ alpha = 0.65
+ omr_response = {}
+ multi_marked, multi_roll = 0, 0
+
+ # TODO Make this part useful for visualizing status checks
+ # blackVals=[0]
+ # whiteVals=[255]
+
+ if config.outputs.show_image_level >= 5:
+ all_c_box_vals = {"int": [], "mcq": []}
+ # TODO: simplify this logic
+ q_nums = {"int": [], "mcq": []}
+
+ # Find Shifts for the field_blocks --> Before calculating threshold!
+ if auto_align:
+ # print("Begin Alignment")
+ # Open : erode then dilate
+ v_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (2, 10))
+ morph_v = cv2.morphologyEx(
+ morph, cv2.MORPH_OPEN, v_kernel, iterations=3
+ )
+ _, morph_v = cv2.threshold(morph_v, 200, 200, cv2.THRESH_TRUNC)
+ morph_v = 255 - ImageUtils.normalize_util(morph_v)
+
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morphed_vertical", morph_v, 0, 1, config=config
+ )
+
+ # InteractionUtils.show("morph1",morph,0,1,config=config)
+ # InteractionUtils.show("morphed_vertical",morph_v,0,1,config=config)
+
+ self.append_save_img(3, morph_v)
+
+ morph_thr = 60 # for Mobile images, 40 for scanned Images
+ _, morph_v = cv2.threshold(morph_v, morph_thr, 255, cv2.THRESH_BINARY)
+ # kernel best tuned to 5x5 now
+ morph_v = cv2.erode(morph_v, np.ones((5, 5), np.uint8), iterations=2)
+
+ self.append_save_img(3, morph_v)
+ # h_kernel = cv2.getStructuringElement(cv2.MORPH_RECT, (10, 2))
+ # morph_h = cv2.morphologyEx(morph, cv2.MORPH_OPEN, h_kernel, iterations=3)
+ # ret, morph_h = cv2.threshold(morph_h,200,200,cv2.THRESH_TRUNC)
+ # morph_h = 255 - normalize_util(morph_h)
+ # InteractionUtils.show("morph_h",morph_h,0,1,config=config)
+ # _, morph_h = cv2.threshold(morph_h,morph_thr,255,cv2.THRESH_BINARY)
+ # morph_h = cv2.erode(morph_h, np.ones((5,5),np.uint8), iterations = 2)
+ if config.outputs.show_image_level >= 3:
+ InteractionUtils.show(
+ "morph_thr_eroded", morph_v, 0, 1, config=config
+ )
+
+ self.append_save_img(6, morph_v)
+
+ # template relative alignment code
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+
+ match_col, max_steps, align_stride, thk = map(
+ config.alignment_params.get,
+ [
+ "match_col",
+ "max_steps",
+ "stride",
+ "thickness",
+ ],
+ )
+ shift, steps = 0, 0
+ while steps < max_steps:
+ left_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0] + shift - thk : -thk + s[0] + shift + match_col,
+ ]
+ )
+ right_mean = np.mean(
+ morph_v[
+ s[1] : s[1] + d[1],
+ s[0]
+ + shift
+ - match_col
+ + d[0]
+ + thk : thk
+ + s[0]
+ + shift
+ + d[0],
+ ]
+ )
+
+ # For demonstration purposes-
+ # if(field_block.name == "int1"):
+ # ret = morph_v.copy()
+ # cv2.rectangle(ret,
+ # (s[0]+shift-thk,s[1]),
+ # (s[0]+shift+thk+d[0],s[1]+d[1]),
+ # constants.CLR_WHITE,
+ # 3)
+ # appendSaveImg(6,ret)
+ # print(shift, left_mean, right_mean)
+ left_shift, right_shift = left_mean > 100, right_mean > 100
+ if left_shift:
+ if right_shift:
+ break
+ else:
+ shift -= align_stride
+ else:
+ if right_shift:
+ shift += align_stride
+ else:
+ break
+ steps += 1
+
+ field_block.shift = shift
+ # print("Aligned field_block: ",field_block.name,"Corrected Shift:",
+ # field_block.shift,", dimensions:", field_block.dimensions,
+ # "origin:", field_block.origin,'\n')
+ # print("End Alignment")
+
+ final_align = None
+ if config.outputs.show_image_level >= 2:
+ initial_align = self.draw_template_layout(img, template, shifted=False)
+ final_align = self.draw_template_layout(
+ img, template, shifted=True, draw_qvals=True
+ )
+ # appendSaveImg(4,mean_vals)
+ self.append_save_img(2, initial_align)
+ self.append_save_img(2, final_align)
+
+ if auto_align:
+ final_align = np.hstack((initial_align, final_align))
+ self.append_save_img(5, img)
+
+ # Get mean bubbleValues n other stats
+ all_q_vals, all_q_strip_arrs, all_q_std_vals = [], [], []
+ total_q_strip_no = 0
+ for field_block in template.field_blocks:
+ box_w, box_h = field_block.bubble_dimensions
+ q_std_vals = []
+ for field_block_bubbles in field_block.traverse_bubbles:
+ q_strip_vals = []
+ for pt in field_block_bubbles:
+ # shifted
+ x, y = (pt.x + field_block.shift, pt.y)
+ rect = [y, y + box_h, x, x + box_w]
+ q_strip_vals.append(
+ cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0]
+ # detectCross(img, rect) ? 100 : 0
+ )
+ q_std_vals.append(round(np.std(q_strip_vals), 2))
+ all_q_strip_arrs.append(q_strip_vals)
+ # _, _, _ = get_global_threshold(q_strip_vals, "QStrip Plot",
+ # plot_show=False, sort_in_plot=True)
+ # hist = getPlotImg()
+ # InteractionUtils.show("QStrip "+field_block_bubbles[0].field_label, hist, 0, 1,config=config)
+ all_q_vals.extend(q_strip_vals)
+ # print(total_q_strip_no, field_block_bubbles[0].field_label, q_std_vals[len(q_std_vals)-1])
+ total_q_strip_no += 1
+ all_q_std_vals.extend(q_std_vals)
+
+ global_std_thresh, _, _ = self.get_global_threshold(
+ all_q_std_vals
+ ) # , "Q-wise Std-dev Plot", plot_show=True, sort_in_plot=True)
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # Note: Plotting takes Significant times here --> Change Plotting args
+ # to support show_image_level
+ # , "Mean Intensity Histogram",plot_show=True, sort_in_plot=True)
+ global_thr, _, _ = self.get_global_threshold(all_q_vals, looseness=4)
+
+ logger.info(
+ f"Thresholding: \tglobal_thr: {round(global_thr, 2)} \tglobal_std_THR: {round(global_std_thresh, 2)}\t{'(Looks like a Xeroxed OMR)' if (global_thr == 255) else ''}"
+ )
+ # plt.show()
+ # hist = getPlotImg()
+ # InteractionUtils.show("StdHist", hist, 0, 1,config=config)
+
+ # if(config.outputs.show_image_level>=1):
+ # hist = getPlotImg()
+ # InteractionUtils.show("Hist", hist, 0, 1,config=config)
+ # appendSaveImg(4,hist)
+ # appendSaveImg(5,hist)
+ # appendSaveImg(2,hist)
+
+ per_omr_threshold_avg, total_q_strip_no, total_q_box_no = 0, 0, 0
+ for field_block in template.field_blocks:
+ block_q_strip_no = 1
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ s, d = field_block.origin, field_block.dimensions
+ key = field_block.name[:3]
+ # cv2.rectangle(final_marked,(s[0]+shift,s[1]),(s[0]+shift+d[0],
+ # s[1]+d[1]),CLR_BLACK,3)
+ for field_block_bubbles in field_block.traverse_bubbles:
+ # All Black or All White case
+ no_outliers = all_q_std_vals[total_q_strip_no] < global_std_thresh
+ # print(total_q_strip_no, field_block_bubbles[0].field_label,
+ # all_q_std_vals[total_q_strip_no], "no_outliers:", no_outliers)
+ per_q_strip_threshold = self.get_local_threshold(
+ all_q_strip_arrs[total_q_strip_no],
+ global_thr,
+ no_outliers,
+ f"Mean Intensity Histogram for {key}.{field_block_bubbles[0].field_label}.{block_q_strip_no}",
+ config.outputs.show_image_level >= 6,
+ )
+ # print(field_block_bubbles[0].field_label,key,block_q_strip_no, "THR: ",
+ # round(per_q_strip_threshold,2))
+ per_omr_threshold_avg += per_q_strip_threshold
+
+ # Note: Little debugging visualization - view the particular Qstrip
+ # if(
+ # 0
+ # # or "q17" in (field_block_bubbles[0].field_label)
+ # # or (field_block_bubbles[0].field_label+str(block_q_strip_no))=="q15"
+ # ):
+ # st, end = qStrip
+ # InteractionUtils.show("QStrip: "+key+"-"+str(block_q_strip_no),
+ # img[st[1] : end[1], st[0]+shift : end[0]+shift],0,config=config)
+
+ # TODO: get rid of total_q_box_no
+ detected_bubbles = []
+ for bubble in field_block_bubbles:
+ bubble_is_marked = (
+ per_q_strip_threshold > all_q_vals[total_q_box_no]
+ )
+ total_q_box_no += 1
+ if bubble_is_marked:
+ detected_bubbles.append(bubble)
+ x, y, field_value = (
+ bubble.x + field_block.shift,
+ bubble.y,
+ bubble.field_value,
+ )
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 12), int(y + box_h / 12)),
+ (
+ int(x + box_w - box_w / 12),
+ int(y + box_h - box_h / 12),
+ ),
+ constants.CLR_DARK_GRAY,
+ 3,
+ )
+
+ cv2.putText(
+ final_marked,
+ str(field_value),
+ (x, y),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ (20, 20, 10),
+ int(1 + 3.5 * constants.TEXT_SIZE),
+ )
+ else:
+ cv2.rectangle(
+ final_marked,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (
+ int(x + box_w - box_w / 10),
+ int(y + box_h - box_h / 10),
+ ),
+ constants.CLR_GRAY,
+ -1,
+ )
+
+ for bubble in detected_bubbles:
+ field_label, field_value = (
+ bubble.field_label,
+ bubble.field_value,
+ )
+ # Only send rolls multi-marked in the directory
+ multi_marked_local = field_label in omr_response
+ omr_response[field_label] = (
+ (omr_response[field_label] + field_value)
+ if multi_marked_local
+ else field_value
+ )
+ # TODO: generalize this into identifier
+ # multi_roll = multi_marked_local and "Roll" in str(q)
+ multi_marked = multi_marked or multi_marked_local
+
+ if len(detected_bubbles) == 0:
+ field_label = field_block_bubbles[0].field_label
+ omr_response[field_label] = field_block.empty_val
+
+ if config.outputs.show_image_level >= 5:
+ if key in all_c_box_vals:
+ q_nums[key].append(f"{key[:2]}_c{str(block_q_strip_no)}")
+ all_c_box_vals[key].append(
+ all_q_strip_arrs[total_q_strip_no]
+ )
+
+ block_q_strip_no += 1
+ total_q_strip_no += 1
+ # /for field_block
+
+ per_omr_threshold_avg /= total_q_strip_no
+ per_omr_threshold_avg = round(per_omr_threshold_avg, 2)
+ # Translucent
+ cv2.addWeighted(
+ final_marked, alpha, transp_layer, 1 - alpha, 0, final_marked
+ )
+ # Box types
+ if config.outputs.show_image_level >= 6:
+ # plt.draw()
+ f, axes = plt.subplots(len(all_c_box_vals), sharey=True)
+ f.canvas.manager.set_window_title(name)
+ ctr = 0
+ type_name = {
+ "int": "Integer",
+ "mcq": "MCQ",
+ "med": "MED",
+ "rol": "Roll",
+ }
+ for k, boxvals in all_c_box_vals.items():
+ axes[ctr].title.set_text(type_name[k] + " Type")
+ axes[ctr].boxplot(boxvals)
+ # thrline=axes[ctr].axhline(per_omr_threshold_avg,color='red',ls='--')
+ # thrline.set_label("Average THR")
+ axes[ctr].set_ylabel("Intensity")
+ axes[ctr].set_xticklabels(q_nums[k])
+ # axes[ctr].legend()
+ ctr += 1
+ # imshow will do the waiting
+ plt.tight_layout(pad=0.5)
+ plt.show()
+
+ if config.outputs.show_image_level >= 3 and final_align is not None:
+ final_align = ImageUtils.resize_util_h(
+ final_align, int(config.dimensions.display_height)
+ )
+ # [final_align.shape[1],0])
+ InteractionUtils.show(
+ "Template Alignment Adjustment", final_align, 0, 0, config=config
+ )
+
+ if config.outputs.save_detections and save_dir is not None:
+ if multi_roll:
+ save_dir = save_dir.joinpath("_MULTI_")
+ image_path = str(save_dir.joinpath(name))
+ ImageUtils.save_img(image_path, final_marked)
+
+ self.append_save_img(2, final_marked)
+
+ if save_dir is not None:
+ for i in range(config.outputs.save_image_level):
+ self.save_image_stacks(i + 1, name, save_dir)
+
+ return omr_response, final_marked, multi_marked, multi_roll
+
+ except Exception as e:
+ raise e
+
+ @staticmethod
+ def draw_template_layout(img, template, shifted=True, draw_qvals=False, border=-1):
+ img = ImageUtils.resize_util(
+ img, template.page_dimensions[0], template.page_dimensions[1]
+ )
+ final_align = img.copy()
+ for field_block in template.field_blocks:
+ s, d = field_block.origin, field_block.dimensions
+ box_w, box_h = field_block.bubble_dimensions
+ shift = field_block.shift
+ if shifted:
+ cv2.rectangle(
+ final_align,
+ (s[0] + shift, s[1]),
+ (s[0] + shift + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ else:
+ cv2.rectangle(
+ final_align,
+ (s[0], s[1]),
+ (s[0] + d[0], s[1] + d[1]),
+ constants.CLR_BLACK,
+ 3,
+ )
+ for field_block_bubbles in field_block.traverse_bubbles:
+ for pt in field_block_bubbles:
+ x, y = (pt.x + field_block.shift, pt.y) if shifted else (pt.x, pt.y)
+ cv2.rectangle(
+ final_align,
+ (int(x + box_w / 10), int(y + box_h / 10)),
+ (int(x + box_w - box_w / 10), int(y + box_h - box_h / 10)),
+ constants.CLR_GRAY,
+ border,
+ )
+ if draw_qvals:
+ rect = [y, y + box_h, x, x + box_w]
+ cv2.putText(
+ final_align,
+ f"{int(cv2.mean(img[rect[0] : rect[1], rect[2] : rect[3]])[0])}",
+ (rect[2] + 2, rect[0] + (box_h * 2) // 3),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ 0.6,
+ constants.CLR_BLACK,
+ 2,
+ )
+ if shifted:
+ text_in_px = cv2.getTextSize(
+ field_block.name, cv2.FONT_HERSHEY_SIMPLEX, constants.TEXT_SIZE, 4
+ )
+ cv2.putText(
+ final_align,
+ field_block.name,
+ (int(s[0] + d[0] - text_in_px[0][0]), int(s[1] - text_in_px[0][1])),
+ cv2.FONT_HERSHEY_SIMPLEX,
+ constants.TEXT_SIZE,
+ constants.CLR_BLACK,
+ 4,
+ )
+ return final_align
+
+ def get_global_threshold(
+ self,
+ q_vals_orig,
+ plot_title=None,
+ plot_show=True,
+ sort_in_plot=True,
+ looseness=1,
+ ):
+ """
+ Note: Cannot assume qStrip has only-gray or only-white bg
+ (in which case there is only one jump).
+ So there will be either 1 or 2 jumps.
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ 2 Jumps :
+ ......
+ |||||| <-- wrong THR
+ ....||||||
+ |||||||||| <-- safe THR
+ ..||||||||||
+ ||||||||||||
+
+ The abstract "First LARGE GAP" is perfect for this.
+ Current code is considering ONLY TOP 2 jumps(>= MIN_GAP) to be big,
+ gives the smaller one
+
+ """
+ config = self.tuning_config
+ PAGE_TYPE_FOR_THRESHOLD, MIN_JUMP, JUMP_DELTA = map(
+ config.threshold_params.get,
+ [
+ "PAGE_TYPE_FOR_THRESHOLD",
+ "MIN_JUMP",
+ "JUMP_DELTA",
+ ],
+ )
+
+ global_default_threshold = (
+ constants.GLOBAL_PAGE_THRESHOLD_WHITE
+ if PAGE_TYPE_FOR_THRESHOLD == "white"
+ else constants.GLOBAL_PAGE_THRESHOLD_BLACK
+ )
+
+ # Sort the Q bubbleValues
+ # TODO: Change var name of q_vals
+ q_vals = sorted(q_vals_orig)
+ # Find the FIRST LARGE GAP and set it as threshold:
+ ls = (looseness + 1) // 2
+ l = len(q_vals) - ls
+ max1, thr1 = MIN_JUMP, global_default_threshold
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - ls] + jump / 2
+
+ # NOTE: thr2 is deprecated, thus is JUMP_DELTA
+ # Make use of the fact that the JUMP_DELTA(Vertical gap ofc) between
+ # values at detected jumps would be atleast 20
+ max2, thr2 = MIN_JUMP, global_default_threshold
+ # Requires atleast 1 gray box to be present (Roll field will ensure this)
+ for i in range(ls, l):
+ jump = q_vals[i + ls] - q_vals[i - ls]
+ new_thr = q_vals[i - ls] + jump / 2
+ if jump > max2 and abs(thr1 - new_thr) > JUMP_DELTA:
+ max2 = jump
+ thr2 = new_thr
+ # global_thr = min(thr1,thr2)
+ global_thr, j_low, j_high = thr1, thr1 - max1 // 2, thr1 + max1 // 2
+
+ # # For normal images
+ # thresholdRead = 116
+ # if(thr1 > thr2 and thr2 > thresholdRead):
+ # print("Note: taking safer thr line.")
+ # global_thr, j_low, j_high = thr2, thr2 - max2//2, thr2 + max2//2
+
+ if plot_title:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals_orig)), q_vals if sort_in_plot else q_vals_orig)
+ ax.set_title(plot_title)
+ thrline = ax.axhline(global_thr, color="green", ls="--", linewidth=5)
+ thrline.set_label("Global Threshold")
+ thrline = ax.axhline(thr2, color="red", ls=":", linewidth=3)
+ thrline.set_label("THR2 Line")
+ # thrline=ax.axhline(j_low,color='red',ls='-.', linewidth=3)
+ # thrline=ax.axhline(j_high,color='red',ls='-.', linewidth=3)
+ # thrline.set_label("Boundary Line")
+ # ax.set_ylabel("Mean Intensity")
+ ax.set_ylabel("Values")
+ ax.set_xlabel("Position")
+ ax.legend()
+ if plot_show:
+ plt.title(plot_title)
+ plt.show()
+
+ return global_thr, j_low, j_high
+
+ def get_local_threshold(
+ self, q_vals, global_thr, no_outliers, plot_title=None, plot_show=True
+ ):
+ """
+ TODO: Update this documentation too-
+ //No more - Assumption : Colwise background color is uniformly gray or white,
+ but not alternating. In this case there is atmost one jump.
+
+ 0 Jump :
+ <-- safe THR?
+ .......
+ ...|||||||
+ |||||||||| <-- safe THR?
+ // How to decide given range is above or below gray?
+ -> global q_vals shall absolutely help here. Just run same function
+ on total q_vals instead of colwise _//
+ How to decide it is this case of 0 jumps
+
+ 1 Jump :
+ ......
+ ||||||
+ |||||| <-- risky THR
+ |||||| <-- safe THR
+ ....||||||
+ ||||||||||
+
+ """
+ config = self.tuning_config
+ # Sort the Q bubbleValues
+ q_vals = sorted(q_vals)
+
+ # Small no of pts cases:
+ # base case: 1 or 2 pts
+ if len(q_vals) < 3:
+ thr1 = (
+ global_thr
+ if np.max(q_vals) - np.min(q_vals) < config.threshold_params.MIN_GAP
+ else np.mean(q_vals)
+ )
+ else:
+ # qmin, qmax, qmean, qstd = round(np.min(q_vals),2), round(np.max(q_vals),2),
+ # round(np.mean(q_vals),2), round(np.std(q_vals),2)
+ # GVals = [round(abs(q-qmean),2) for q in q_vals]
+ # gmean, gstd = round(np.mean(GVals),2), round(np.std(GVals),2)
+ # # DISCRETION: Pretty critical factor in reading response
+ # # Doesn't work well for small number of values.
+ # DISCRETION = 2.7 # 2.59 was closest hit, 3.0 is too far
+ # L2MaxGap = round(max([abs(g-gmean) for g in GVals]),2)
+ # if(L2MaxGap > DISCRETION*gstd):
+ # no_outliers = False
+
+ # # ^Stackoverflow method
+ # print(field_label, no_outliers,"qstd",round(np.std(q_vals),2), "gstd", gstd,
+ # "Gaps in gvals",sorted([round(abs(g-gmean),2) for g in GVals],reverse=True),
+ # '\t',round(DISCRETION*gstd,2), L2MaxGap)
+
+ # else:
+ # Find the LARGEST GAP and set it as threshold: //(FIRST LARGE GAP)
+ l = len(q_vals) - 1
+ max1, thr1 = config.threshold_params.MIN_JUMP, 255
+ for i in range(1, l):
+ jump = q_vals[i + 1] - q_vals[i - 1]
+ if jump > max1:
+ max1 = jump
+ thr1 = q_vals[i - 1] + jump / 2
+ # print(field_label,q_vals,max1)
+
+ confident_jump = (
+ config.threshold_params.MIN_JUMP
+ + config.threshold_params.CONFIDENT_SURPLUS
+ )
+ # If not confident, then only take help of global_thr
+ if max1 < confident_jump:
+ if no_outliers:
+ # All Black or All White case
+ thr1 = global_thr
+ else:
+ # TODO: Low confidence parameters here
+ pass
+
+ # if(thr1 == 255):
+ # print("Warning: threshold is unexpectedly 255! (Outlier Delta issue?)",plot_title)
+
+ # Make a common plot function to show local and global thresholds
+ if plot_show and plot_title is not None:
+ _, ax = plt.subplots()
+ ax.bar(range(len(q_vals)), q_vals)
+ thrline = ax.axhline(thr1, color="green", ls=("-."), linewidth=3)
+ thrline.set_label("Local Threshold")
+ thrline = ax.axhline(global_thr, color="red", ls=":", linewidth=5)
+ thrline.set_label("Global Threshold")
+ ax.set_title(plot_title)
+ ax.set_ylabel("Bubble Mean Intensity")
+ ax.set_xlabel("Bubble Number(sorted)")
+ ax.legend()
+ # TODO append QStrip to this plot-
+ # appendSaveImg(6,getPlotImg())
+ if plot_show:
+ plt.show()
+ return thr1
+
+ def append_save_img(self, key, img):
+ if self.save_image_level >= int(key):
+ self.save_img_list[key].append(img.copy())
+
+ def save_image_stacks(self, key, filename, save_dir):
+ config = self.tuning_config
+ if self.save_image_level >= int(key) and self.save_img_list[key] != []:
+ name = os.path.splitext(filename)[0]
+ result = np.hstack(
+ tuple(
+ [
+ ImageUtils.resize_util_h(img, config.dimensions.display_height)
+ for img in self.save_img_list[key]
+ ]
+ )
+ )
+ result = ImageUtils.resize_util(
+ result,
+ min(
+ len(self.save_img_list[key]) * config.dimensions.display_width // 3,
+ int(config.dimensions.display_width * 2.5),
+ ),
+ )
+ ImageUtils.save_img(f"{save_dir}stack/{name}_{str(key)}_stack.jpg", result)
+
+ def reset_all_save_img(self):
+ for i in range(self.save_image_level):
+ self.save_img_list[i + 1] = []
diff --git a/OMRChecker/src/defaults/__init__.py b/OMRChecker/src/defaults/__init__.py
new file mode 100644
index 0000000000000000000000000000000000000000..db3d42e87b3e47d0cbee08032e65e87ac1bd8165
--- /dev/null
+++ b/OMRChecker/src/defaults/__init__.py
@@ -0,0 +1,5 @@
+# https://docs.python.org/3/tutorial/modules.html#:~:text=The%20__init__.py,on%20the%20module%20search%20path.
+# Use all imports relative to root directory
+# (https://chrisyeh96.github.io/2017/08/08/definitive-guide-python-imports.html)
+from src.defaults.config import * # NOQA
+from src.defaults.template import * # NOQA
diff --git a/OMRChecker/src/defaults/config.py b/OMRChecker/src/defaults/config.py
new file mode 100644
index 0000000000000000000000000000000000000000..6d1315fb45b0e736ad7daedcea5f3b23e3a9078d
--- /dev/null
+++ b/OMRChecker/src/defaults/config.py
@@ -0,0 +1,35 @@
+from dotmap import DotMap
+
+CONFIG_DEFAULTS = DotMap(
+ {
+ "dimensions": {
+ "display_height": 2480,
+ "display_width": 1640,
+ "processing_height": 820,
+ "processing_width": 666,
+ },
+ "threshold_params": {
+ "GAMMA_LOW": 0.7,
+ "MIN_GAP": 30,
+ "MIN_JUMP": 25,
+ "CONFIDENT_SURPLUS": 5,
+ "JUMP_DELTA": 30,
+ "PAGE_TYPE_FOR_THRESHOLD": "white",
+ },
+ "alignment_params": {
+ # Note: 'auto_align' enables automatic template alignment, use if the scans show slight misalignments.
+ "auto_align": False,
+ "match_col": 5,
+ "max_steps": 20,
+ "stride": 1,
+ "thickness": 3,
+ },
+ "outputs": {
+ "show_image_level": 0,
+ "save_image_level": 0,
+ "save_detections": True,
+ "filter_out_multimarked_files": False,
+ },
+ },
+ _dynamic=False,
+)
diff --git a/OMRChecker/src/defaults/template.py b/OMRChecker/src/defaults/template.py
new file mode 100644
index 0000000000000000000000000000000000000000..efd75420f01cf49f72a2e285734c0d79cafe4837
--- /dev/null
+++ b/OMRChecker/src/defaults/template.py
@@ -0,0 +1,6 @@
+TEMPLATE_DEFAULTS = {
+ "preProcessors": [],
+ "emptyValue": "",
+ "customLabels": {},
+ "outputColumns": [],
+}
diff --git a/OMRChecker/src/entry.py b/OMRChecker/src/entry.py
new file mode 100644
index 0000000000000000000000000000000000000000..94d3137c3ac99502399cbb40d6d378cd8249d8de
--- /dev/null
+++ b/OMRChecker/src/entry.py
@@ -0,0 +1,371 @@
+"""
+
+ OMRChecker
+
+ Author: Udayraj Deshmukh
+ Github: https://github.com/Udayraj123
+
+"""
+import os
+from csv import QUOTE_NONNUMERIC
+from pathlib import Path
+from time import time
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src import constants
+from src.defaults import CONFIG_DEFAULTS
+from src.evaluation import EvaluationConfig, evaluate_concatenated_response
+from src.logger import console, logger
+from src.template import Template
+from src.utils.file import Paths, setup_dirs_for_paths, setup_outputs_for_template
+from src.utils.image import ImageUtils
+from src.utils.interaction import InteractionUtils, Stats
+from src.utils.parsing import get_concatenated_response, open_config_with_defaults
+
+# Load processors
+STATS = Stats()
+
+
+def entry_point(input_dir, args):
+ if not os.path.exists(input_dir):
+ raise Exception(f"Given input directory does not exist: '{input_dir}'")
+ curr_dir = input_dir
+ return process_dir(input_dir, curr_dir, args)
+
+
+def print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+):
+ logger.info("")
+ table = Table(title="Current Configurations", show_header=False, show_lines=False)
+ table.add_column("Key", style="cyan", no_wrap=True)
+ table.add_column("Value", style="magenta")
+ table.add_row("Directory Path", f"{curr_dir}")
+ table.add_row("Count of Images", f"{len(omr_files)}")
+ table.add_row("Set Layout Mode ", "ON" if args["setLayout"] else "OFF")
+ pre_processor_names = [pp.__class__.__name__ for pp in template.pre_processors]
+ table.add_row(
+ "Markers Detection",
+ "ON" if "CropOnMarkers" in pre_processor_names else "OFF",
+ )
+ table.add_row("Auto Alignment", f"{tuning_config.alignment_params.auto_align}")
+ table.add_row("Detected Template Path", f"{template}")
+ if local_config_path:
+ table.add_row("Detected Local Config", f"{local_config_path}")
+ if evaluation_config:
+ table.add_row("Detected Evaluation Config", f"{evaluation_config}")
+
+ table.add_row(
+ "Detected pre-processors",
+ ", ".join(pre_processor_names),
+ )
+ console.print(table, justify="center")
+
+
+def process_dir(
+ root_dir,
+ curr_dir,
+ args,
+ template=None,
+ tuning_config=CONFIG_DEFAULTS,
+ evaluation_config=None,
+):
+ # Update local tuning_config (in current recursion stack)
+ local_config_path = curr_dir.joinpath(constants.CONFIG_FILENAME)
+ if os.path.exists(local_config_path):
+ tuning_config = open_config_with_defaults(local_config_path)
+
+ # Update local template (in current recursion stack)
+ local_template_path = curr_dir.joinpath(constants.TEMPLATE_FILENAME)
+ local_template_exists = os.path.exists(local_template_path)
+ if local_template_exists:
+ template = Template(
+ local_template_path,
+ tuning_config,
+ )
+ # Look for subdirectories for processing
+ subdirs = [d for d in curr_dir.iterdir() if d.is_dir()]
+
+ output_dir = Path(args["output_dir"], curr_dir.relative_to(root_dir))
+ paths = Paths(output_dir)
+
+ # look for images in current dir to process
+ exts = ("*.[pP][nN][gG]", "*.[jJ][pP][gG]", "*.[jJ][pP][eE][gG]")
+ omr_files = sorted([f for ext in exts for f in curr_dir.glob(ext)])
+
+ # Exclude images (take union over all pre_processors)
+ excluded_files = []
+ if template:
+ for pp in template.pre_processors:
+ excluded_files.extend(Path(p) for p in pp.exclude_files())
+
+ local_evaluation_path = curr_dir.joinpath(constants.EVALUATION_FILENAME)
+ if not args["setLayout"] and os.path.exists(local_evaluation_path):
+ if not local_template_exists:
+ logger.warning(
+ f"Found an evaluation file without a parent template file: {local_evaluation_path}"
+ )
+ evaluation_config = EvaluationConfig(
+ curr_dir,
+ local_evaluation_path,
+ template,
+ tuning_config,
+ )
+
+ excluded_files.extend(
+ Path(exclude_file) for exclude_file in evaluation_config.get_exclude_files()
+ )
+
+ omr_files = [f for f in omr_files if f not in excluded_files]
+
+ if omr_files:
+ if not template:
+ logger.error(
+ f"Found images, but no template in the directory tree \
+ of '{curr_dir}'. \nPlace {constants.TEMPLATE_FILENAME} in the \
+ appropriate directory."
+ )
+ raise Exception(
+ f"No template file found in the directory tree of {curr_dir}"
+ )
+
+ setup_dirs_for_paths(paths)
+ outputs_namespace = setup_outputs_for_template(paths, template)
+
+ print_config_summary(
+ curr_dir,
+ omr_files,
+ template,
+ tuning_config,
+ local_config_path,
+ evaluation_config,
+ args,
+ )
+ if args["setLayout"]:
+ show_template_layouts(omr_files, template, tuning_config)
+ else:
+ process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+ )
+
+ elif not subdirs:
+ # Each subdirectory should have images or should be non-leaf
+ logger.info(
+ f"No valid images or sub-folders found in {curr_dir}.\
+ Empty directories not allowed."
+ )
+
+ # recursively process sub-folders
+ for d in subdirs:
+ process_dir(
+ root_dir,
+ d,
+ args,
+ template,
+ tuning_config,
+ evaluation_config,
+ )
+
+
+def show_template_layouts(omr_files, template, tuning_config):
+ for file_path in omr_files:
+ file_name = file_path.name
+ file_path = str(file_path)
+ in_omr = cv2.imread(file_path, cv2.IMREAD_GRAYSCALE)
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+ template_layout = template.image_instance_ops.draw_template_layout(
+ in_omr, template, shifted=False, border=2
+ )
+ InteractionUtils.show(
+ f"Template Layout: {file_name}", template_layout, 1, 1, config=tuning_config
+ )
+
+
+def process_files(
+ omr_files,
+ template,
+ tuning_config,
+ evaluation_config,
+ outputs_namespace,
+):
+ start_time = int(time())
+ files_counter = 0
+ STATS.files_not_moved = 0
+
+ for file_path in omr_files:
+ files_counter += 1
+ file_name = file_path.name
+
+ in_omr = cv2.imread(str(file_path), cv2.IMREAD_GRAYSCALE)
+
+ logger.info("")
+ logger.info(
+ f"({files_counter}) Opening image: \t'{file_path}'\tResolution: {in_omr.shape}"
+ )
+
+ template.image_instance_ops.reset_all_save_img()
+
+ template.image_instance_ops.append_save_img(1, in_omr)
+
+ in_omr = template.image_instance_ops.apply_preprocessors(
+ file_path, in_omr, template
+ )
+
+ if in_omr is None:
+ # Error OMR case
+ new_file_path = outputs_namespace.paths.errors_dir.joinpath(file_name)
+ outputs_namespace.OUTPUT_SET.append(
+ [file_name] + outputs_namespace.empty_resp
+ )
+ if check_and_move(
+ constants.ERROR_CODES.NO_MARKER_ERR, file_path, new_file_path
+ ):
+ err_line = [
+ file_name,
+ file_path,
+ new_file_path,
+ "NA",
+ ] + outputs_namespace.empty_resp
+ pd.DataFrame(err_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Errors"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ continue
+
+ # uniquify
+ file_id = str(file_name)
+ save_dir = outputs_namespace.paths.save_marked_dir
+ (
+ response_dict,
+ final_marked,
+ multi_marked,
+ _,
+ ) = template.image_instance_ops.read_omr_response(
+ template, image=in_omr, name=file_id, save_dir=save_dir
+ )
+
+ # TODO: move inner try catch here
+ # concatenate roll nos, set unmarked responses, etc
+ omr_response = get_concatenated_response(response_dict, template)
+
+ if (
+ evaluation_config is None
+ or not evaluation_config.get_should_explain_scoring()
+ ):
+ logger.info(f"Read Response: \n{omr_response}")
+
+ score = 0
+ if evaluation_config is not None:
+ score = evaluate_concatenated_response(
+ omr_response, evaluation_config, file_path, outputs_namespace.paths.evaluation_dir
+ )
+ logger.info(
+ f"(/{files_counter}) Graded with score: {round(score, 2)}\t for file: '{file_id}'"
+ )
+ else:
+ logger.info(f"(/{files_counter}) Processed file: '{file_id}'")
+
+ if tuning_config.outputs.show_image_level >= 2:
+ InteractionUtils.show(
+ f"Final Marked Bubbles : '{file_id}'",
+ ImageUtils.resize_util_h(
+ final_marked, int(tuning_config.dimensions.display_height * 1.3)
+ ),
+ 1,
+ 1,
+ config=tuning_config,
+ )
+
+ resp_array = []
+ for k in template.output_columns:
+ resp_array.append(omr_response[k])
+
+ outputs_namespace.OUTPUT_SET.append([file_name] + resp_array)
+
+ if multi_marked == 0 or not tuning_config.outputs.filter_out_multimarked_files:
+ STATS.files_not_moved += 1
+ new_file_path = save_dir.joinpath(file_id)
+ # Enter into Results sheet-
+ results_line = [file_name, file_path, new_file_path, score] + resp_array
+ # Write/Append to results_line file(opened in append mode)
+ pd.DataFrame(results_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["Results"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ else:
+ # multi_marked file
+ logger.info(f"[{files_counter}] Found multi-marked file: '{file_id}'")
+ new_file_path = outputs_namespace.paths.multi_marked_dir.joinpath(file_name)
+ if check_and_move(
+ constants.ERROR_CODES.MULTI_BUBBLE_WARN, file_path, new_file_path
+ ):
+ mm_line = [file_name, file_path, new_file_path, "NA"] + resp_array
+ pd.DataFrame(mm_line, dtype=str).T.to_csv(
+ outputs_namespace.files_obj["MultiMarked"],
+ mode="a",
+ quoting=QUOTE_NONNUMERIC,
+ header=False,
+ index=False,
+ )
+ # else:
+ # TODO: Add appropriate record handling here
+ # pass
+
+ print_stats(start_time, files_counter, tuning_config)
+
+
+def check_and_move(error_code, file_path, filepath2):
+ # TODO: fix file movement into error/multimarked/invalid etc again
+ STATS.files_not_moved += 1
+ return True
+
+
+def print_stats(start_time, files_counter, tuning_config):
+ time_checking = max(1, round(time() - start_time, 2))
+ log = logger.info
+ log("")
+ log(f"{'Total file(s) moved': <27}: {STATS.files_moved}")
+ log(f"{'Total file(s) not moved': <27}: {STATS.files_not_moved}")
+ log("--------------------------------")
+ log(
+ f"{'Total file(s) processed': <27}: {files_counter} ({'Sum Tallied!' if files_counter == (STATS.files_moved + STATS.files_not_moved) else 'Not Tallying!'})"
+ )
+
+ if tuning_config.outputs.show_image_level <= 0:
+ log(
+ f"\nFinished Checking {files_counter} file(s) in {round(time_checking, 1)} seconds i.e. ~{round(time_checking / 60, 1)} minute(s)."
+ )
+ log(
+ f"{'OMR Processing Rate': <27}: \t ~ {round(time_checking / files_counter, 2)} seconds/OMR"
+ )
+ log(
+ f"{'OMR Processing Speed': <27}: \t ~ {round((files_counter * 60) / time_checking, 2)} OMRs/minute"
+ )
+ else:
+ log(f"\n{'Total script time': <27}: {time_checking} seconds")
+
+ if tuning_config.outputs.show_image_level <= 1:
+ log(
+ "\nTip: To see some awesome visuals, open config.json and increase 'show_image_level'"
+ )
diff --git a/OMRChecker/src/evaluation.py b/OMRChecker/src/evaluation.py
new file mode 100644
index 0000000000000000000000000000000000000000..55c2455a2967063e7604d0f0c904c09720e1064d
--- /dev/null
+++ b/OMRChecker/src/evaluation.py
@@ -0,0 +1,546 @@
+import ast
+import os
+import re
+from copy import deepcopy
+from csv import QUOTE_NONNUMERIC
+
+import cv2
+import pandas as pd
+from rich.table import Table
+
+from src.logger import console, logger
+from src.schemas.constants import (
+ BONUS_SECTION_PREFIX,
+ DEFAULT_SECTION_KEY,
+ MARKING_VERDICT_TYPES,
+)
+from src.utils.parsing import (
+ get_concatenated_response,
+ open_evaluation_with_validation,
+ parse_fields,
+ parse_float_or_fraction,
+)
+
+
+class AnswerMatcher:
+ def __init__(self, answer_item, section_marking_scheme):
+ self.section_marking_scheme = section_marking_scheme
+ self.answer_item = answer_item
+ self.answer_type = self.validate_and_get_answer_type(answer_item)
+ self.set_defaults_from_scheme(section_marking_scheme)
+
+ @staticmethod
+ def is_a_marking_score(answer_element):
+ # Note: strict type checking is already done at schema validation level,
+ # Here we focus on overall struct type
+ return type(answer_element) == str or type(answer_element) == int
+
+ @staticmethod
+ def is_standard_answer(answer_element):
+ return type(answer_element) == str and len(answer_element) >= 1
+
+ def validate_and_get_answer_type(self, answer_item):
+ if self.is_standard_answer(answer_item):
+ return "standard"
+ elif type(answer_item) == list:
+ if (
+ # Array of answer elements: ['A', 'B', 'AB']
+ len(answer_item) >= 2
+ and all(
+ self.is_standard_answer(answers_or_score)
+ for answers_or_score in answer_item
+ )
+ ):
+ return "multiple-correct"
+ elif (
+ # Array of two-tuples: [['A', 1], ['B', 1], ['C', 3], ['AB', 2]]
+ len(answer_item) >= 1
+ and all(
+ type(answer_and_score) == list and len(answer_and_score) == 2
+ for answer_and_score in answer_item
+ )
+ and all(
+ self.is_standard_answer(allowed_answer)
+ and self.is_a_marking_score(answer_score)
+ for allowed_answer, answer_score in answer_item
+ )
+ ):
+ return "multiple-correct-weighted"
+
+ logger.critical(
+ f"Unable to determine answer type for answer item: {answer_item}"
+ )
+ raise Exception("Unable to determine answer type")
+
+ def set_defaults_from_scheme(self, section_marking_scheme):
+ answer_type = self.answer_type
+ self.empty_val = section_marking_scheme.empty_val
+ answer_item = self.answer_item
+ self.marking = deepcopy(section_marking_scheme.marking)
+ # TODO: reuse part of parse_scheme_marking here -
+ if answer_type == "standard":
+ # no local overrides
+ pass
+ elif answer_type == "multiple-correct":
+ # override marking scheme scores for each allowed answer
+ for allowed_answer in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = self.marking["correct"]
+ elif answer_type == "multiple-correct-weighted":
+ # Note: No override using marking scheme as answer scores are provided in answer_item
+ for allowed_answer, answer_score in answer_item:
+ self.marking[f"correct-{allowed_answer}"] = parse_float_or_fraction(
+ answer_score
+ )
+
+ def get_marking_scheme(self):
+ return self.section_marking_scheme
+
+ def get_section_explanation(self):
+ answer_type = self.answer_type
+ if answer_type in ["standard", "multiple-correct"]:
+ return self.section_marking_scheme.section_key
+ elif answer_type == "multiple-correct-weighted":
+ return f"Custom: {self.marking}"
+
+ def get_verdict_marking(self, marked_answer):
+ answer_type = self.answer_type
+ question_verdict = "incorrect"
+ if answer_type == "standard":
+ question_verdict = self.get_standard_verdict(marked_answer)
+ elif answer_type == "multiple-correct":
+ question_verdict = self.get_multiple_correct_verdict(marked_answer)
+ elif answer_type == "multiple-correct-weighted":
+ question_verdict = self.get_multiple_correct_weighted_verdict(marked_answer)
+ return question_verdict, self.marking[question_verdict]
+
+ def get_standard_verdict(self, marked_answer):
+ allowed_answer = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer == allowed_answer:
+ return "correct"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_verdict(self, marked_answer):
+ allowed_answers = self.answer_item
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def get_multiple_correct_weighted_verdict(self, marked_answer):
+ allowed_answers = [
+ allowed_answer for allowed_answer, _answer_score in self.answer_item
+ ]
+ if marked_answer == self.empty_val:
+ return "unmarked"
+ elif marked_answer in allowed_answers:
+ return f"correct-{marked_answer}"
+ else:
+ return "incorrect"
+
+ def __str__(self):
+ return f"{self.answer_item}"
+
+
+class SectionMarkingScheme:
+ def __init__(self, section_key, section_scheme, empty_val):
+ # TODO: get local empty_val from qblock
+ self.empty_val = empty_val
+ self.section_key = section_key
+ # DEFAULT marking scheme follows a shorthand
+ if section_key == DEFAULT_SECTION_KEY:
+ self.questions = None
+ self.marking = self.parse_scheme_marking(section_scheme)
+ else:
+ self.questions = parse_fields(section_key, section_scheme["questions"])
+ self.marking = self.parse_scheme_marking(section_scheme["marking"])
+
+ def __str__(self):
+ return self.section_key
+
+ def parse_scheme_marking(self, marking):
+ parsed_marking = {}
+ for verdict_type in MARKING_VERDICT_TYPES:
+ verdict_marking = parse_float_or_fraction(marking[verdict_type])
+ if (
+ verdict_marking > 0
+ and verdict_type == "incorrect"
+ and not self.section_key.startswith(BONUS_SECTION_PREFIX)
+ ):
+ logger.warning(
+ f"Found positive marks({round(verdict_marking, 2)}) for incorrect answer in the schema '{self.section_key}'. For Bonus sections, add a prefix 'BONUS_' to them."
+ )
+ parsed_marking[verdict_type] = verdict_marking
+
+ return parsed_marking
+
+ def match_answer(self, marked_answer, answer_matcher):
+ question_verdict, verdict_marking = answer_matcher.get_verdict_marking(
+ marked_answer
+ )
+
+ return verdict_marking, question_verdict
+
+
+class EvaluationConfig:
+ """Note: this instance will be reused for multiple omr sheets"""
+
+ def __init__(self, curr_dir, evaluation_path, template, tuning_config):
+ self.path = evaluation_path
+ evaluation_json = open_evaluation_with_validation(evaluation_path)
+ options, marking_schemes, source_type = map(
+ evaluation_json.get, ["options", "marking_schemes", "source_type"]
+ )
+ self.should_explain_scoring = options.get("should_explain_scoring", False)
+ self.has_non_default_section = False
+ self.exclude_files = []
+ self.enable_evaluation_table_to_csv = options.get(
+ "enable_evaluation_table_to_csv", False
+ )
+
+ if source_type == "csv":
+ csv_path = curr_dir.joinpath(options["answer_key_csv_path"])
+ if not os.path.exists(csv_path):
+ logger.warning(f"Answer key csv does not exist at: '{csv_path}'.")
+
+ answer_key_image_path = options.get("answer_key_image_path", None)
+ if os.path.exists(csv_path):
+ # TODO: CSV parsing/validation for each row with a (qNo,