 -
-
-
- - Watch the Demo Video -
-
-  -
-
-
-
- Watch the Demo Video
-
-  -
-
-  -
-
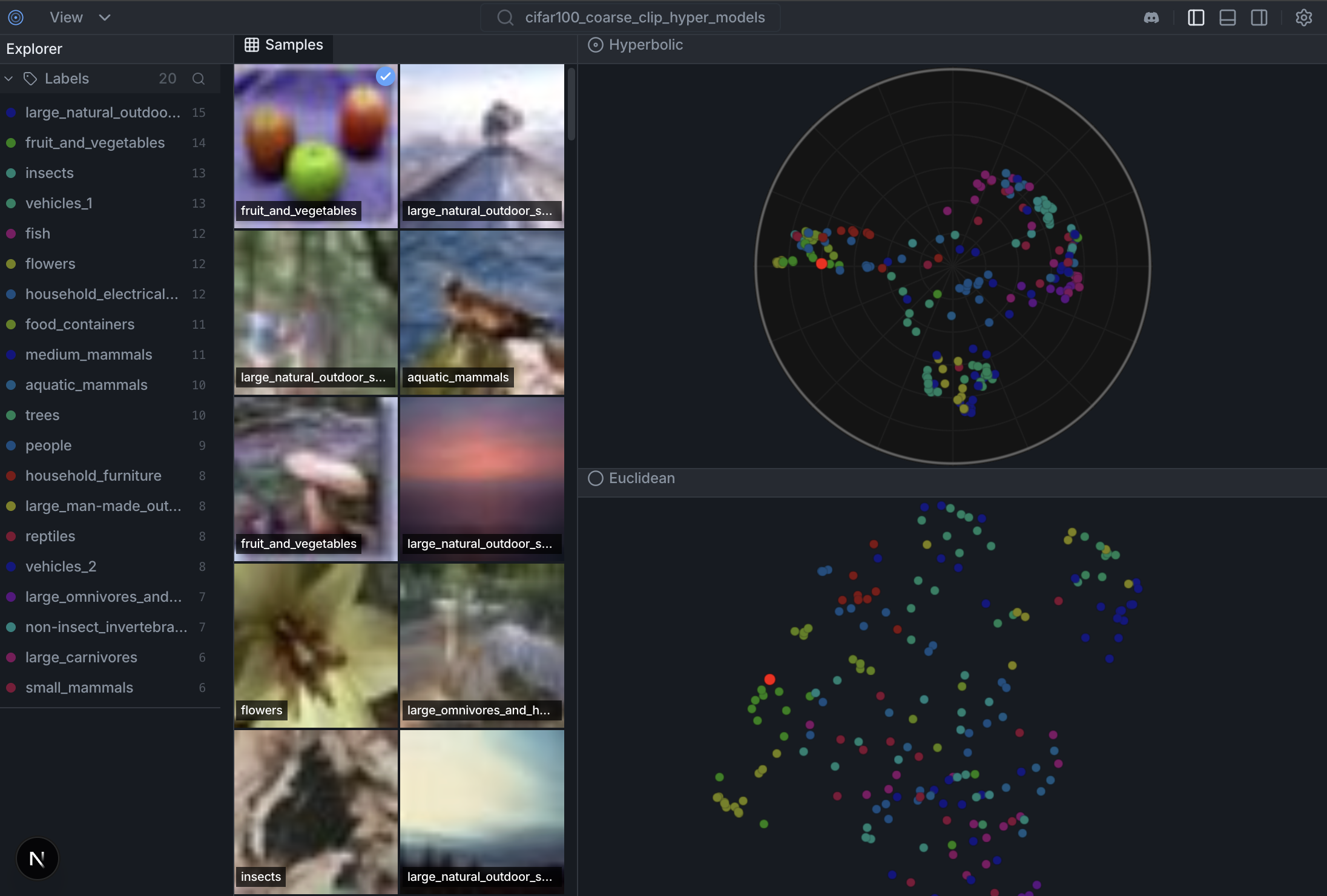
- Drag to Pan. Experience the "infinite" space. - Notice how the red "Rare" points expand and separate as you bring them towards the center. -
-- Make sure the HyperView backend is running on port 6262. -
-HyperView is running in Colab. " - f"" - "Open HyperView in a new tab.
" - ) - ) - display(HTML(f"{app_url}
")) - return - except Exception: - # Fall through to the generic notebook behavior. - pass - - # Default: open in a new browser tab (works well for Jupyter). - try: - from IPython.display import HTML, Javascript, display - - display( - HTML( - "HyperView is running. " - f"Open in a new tab." - "
" - ) - ) - - # Best-effort auto-open. Some browsers may block popups. - display(Javascript(f'window.open("{self.url}", "_blank");')) - except ImportError: - print(f"IPython not installed. Please visit {self.url} in your browser.") - - def open_browser(self): - """Open the visualizer in a browser window.""" - webbrowser.open(self.url) - - -def launch( - dataset: Dataset, - port: int = 6262, - host: str = "127.0.0.1", - open_browser: bool = True, - notebook: bool | None = None, - height: int = 800, - reuse_server: bool = False, -) -> Session: - """Launch the HyperView visualization server. - - Note: - HyperView's UI requires 2D layouts (Euclidean + Poincare). If they are - missing but high-dimensional embeddings exist, this function will compute - them automatically. - - Args: - dataset: The dataset to visualize. - port: Port to run the server on. - host: Host to bind to. - open_browser: Whether to open a browser window. - notebook: Whether to display in a notebook. If None, auto-detects. - height: Height of the iframe in the notebook. - reuse_server: If True, and the requested port is already serving HyperView, - attach to the existing server instead of starting a new one. For safety, - this will only attach when the existing server reports the same dataset - name (via `/__hyperview__/health`). - - Returns: - A Session object. - - Example: - >>> import hyperview as hv - >>> dataset = hv.Dataset("my_dataset") - >>> dataset.add_images_dir("/path/to/images", label_from_folder=True) - >>> dataset.compute_embeddings() - >>> dataset.compute_visualization() - >>> hv.launch(dataset) - """ - if notebook is None: - # Colab is always a notebook environment, even if _is_notebook() fails to detect it - notebook = _is_notebook() or _is_colab() - - if _is_colab() and host == "127.0.0.1": - # Colab port forwarding/proxying is most reliable when the server binds - # to all interfaces. - host = "0.0.0.0" - - # Preflight: avoid doing expensive work if the port is already in use. - # If it's already serving HyperView and reuse_server=True, we can safely attach. - connect_host = "127.0.0.1" if host == "0.0.0.0" else host - health_url = f"http://{connect_host}:{port}/__hyperview__/health" - - if _can_connect(connect_host, port, timeout_s=0.2): - health = _try_read_health(health_url, timeout_s=0.2) - if health is not None and health.name == "hyperview": - if not reuse_server: - raise RuntimeError( - "HyperView failed to start because the port is already serving " - f"HyperView (port={port}, dataset={health.dataset}, " - f"session_id={health.session_id}, pid={health.pid}). " - "Choose a different port, stop the existing server, or pass " - "reuse_server=True to attach." - ) - - if health.dataset is not None and health.dataset != dataset.name: - raise RuntimeError( - "HyperView refused to attach to the existing server because it is " - f"serving a different dataset (port={port}, dataset={health.dataset}). " - f"Requested dataset={dataset.name}. Stop the existing server or " - "choose a different port." - ) - - session = Session(dataset, host, port) - if health.session_id is not None: - session.session_id = health.session_id - - if notebook: - if _is_colab(): - print( - f"\nHyperView is already running (Colab, port={session.port}). " - "Use the link below to open it." - ) - else: - print( - f"\nHyperView is already running at {session.url} (port={session.port}). " - "Opening a new tab..." - ) - session.show(height=height) - else: - print(f"\nHyperView is already running at {session.url} (port={session.port}).") - if open_browser: - session.open_browser() - - return session - - raise RuntimeError( - "HyperView failed to start because the port is already in use " - f"by a non-HyperView service (port={port}). Choose a different " - "port or stop the process listening on that port." - ) - - # The frontend requires 2D coords from /api/embeddings. - # Ensure at least one layout exists; do not auto-generate optional geometries. - layouts = dataset.list_layouts() - spaces = dataset.list_spaces() - - if not spaces: - raise ValueError( - "HyperView launch requires 2D projections for the UI. " - "No projections or embedding spaces were found. " - "Call `dataset.compute_embeddings()` and `dataset.compute_visualization()` " - "before `hv.launch()`." - ) - - if not layouts: - default_space_key = spaces[0].space_key - print("No layouts found. Computing euclidean visualization...") - dataset.compute_visualization(space_key=default_space_key, geometry="euclidean") - - session = Session(dataset, host, port) - - if notebook: - session.start(background=True) - if _is_colab(): - print( - f"\nHyperView is running (Colab, port={session.port}). " - "Use the link below to open it." - ) - else: - print(f"\nHyperView is running at {session.url}. Opening a new tab...") - session.show(height=height) - else: - session.start(background=True) - print(" Press Ctrl+C to stop.\n") - print(f"\nHyperView is running at {session.url}") - - if open_browser: - session.open_browser() - - try: - while True: - # Keep the main thread alive so the daemon server thread can run. - time.sleep(0.25) - if session._server_thread is not None and not session._server_thread.is_alive(): - raise RuntimeError("HyperView server stopped unexpectedly.") - except KeyboardInterrupt: - pass - finally: - session.stop() - if session._server_thread is not None: - session._server_thread.join(timeout=2.0) - - return session - - -def _is_notebook() -> bool: - """Check if running in a notebook environment.""" - try: - from IPython import get_ipython - except ImportError: - return False - - shell = get_ipython() - return shell is not None and shell.__class__.__name__ == "ZMQInteractiveShell" - - -def _is_colab() -> bool: - """Check if running inside a Google Colab notebook runtime.""" - if os.environ.get("COLAB_RELEASE_TAG"): - return True - try: - import google.colab # type: ignore[import-not-found] - - return True - except ImportError: - return False diff --git a/src/hyperview/cli.py b/src/hyperview/cli.py deleted file mode 100644 index 9e5486ca24bc7b22ad4850d45158f662f3e9715e..0000000000000000000000000000000000000000 --- a/src/hyperview/cli.py +++ /dev/null @@ -1,159 +0,0 @@ -"""Command-line interface for HyperView.""" - -import argparse -import sys - -from hyperview import Dataset, launch - - -def main(): - """Main CLI entry point.""" - parser = argparse.ArgumentParser( - prog="hyperview", - description="HyperView - Dataset visualization with hyperbolic embeddings", - ) - subparsers = parser.add_subparsers(dest="command", help="Available commands") - - # Demo command - demo_parser = subparsers.add_parser("demo", help="Run a demo with sample data") - demo_parser.add_argument( - "--samples", - type=int, - default=500, - help="Number of samples to load (default: 500)", - ) - demo_parser.add_argument( - "--port", - type=int, - default=6262, - help="Port to run the server on (default: 6262)", - ) - demo_parser.add_argument( - "--host", - type=str, - default="127.0.0.1", - help="Host to bind the server to (default: 127.0.0.1)", - ) - demo_parser.add_argument( - "--no-browser", - action="store_true", - help="Do not open a browser window automatically", - ) - demo_parser.add_argument( - "--reuse-server", - action="store_true", - help=( - "If the port is already serving HyperView, attach instead of failing. " - "For safety, this only attaches when the existing server reports the same dataset name." - ), - ) - - # Serve command - serve_parser = subparsers.add_parser("serve", help="Serve a saved dataset") - serve_parser.add_argument("dataset", help="Path to saved dataset JSON file") - serve_parser.add_argument( - "--port", - type=int, - default=6262, - help="Port to run the server on (default: 6262)", - ) - serve_parser.add_argument( - "--host", - type=str, - default="127.0.0.1", - help="Host to bind the server to (default: 127.0.0.1)", - ) - serve_parser.add_argument( - "--no-browser", - action="store_true", - help="Do not open a browser window automatically", - ) - serve_parser.add_argument( - "--reuse-server", - action="store_true", - help=( - "If the port is already serving HyperView, attach instead of failing. " - "For safety, this only attaches when the existing server reports the same dataset name." - ), - ) - - args = parser.parse_args() - - if args.command == "demo": - run_demo( - args.samples, - args.port, - host=args.host, - open_browser=not args.no_browser, - reuse_server=args.reuse_server, - ) - elif args.command == "serve": - serve_dataset( - args.dataset, - args.port, - host=args.host, - open_browser=not args.no_browser, - reuse_server=args.reuse_server, - ) - else: - parser.print_help() - sys.exit(1) - - -def run_demo( - num_samples: int = 500, - port: int = 6262, - *, - host: str = "127.0.0.1", - open_browser: bool = True, - reuse_server: bool = False, -): - """Run a demo with CIFAR-10 data.""" - print("Loading CIFAR-10 dataset...") - dataset = Dataset("cifar10_demo") - - added, skipped = dataset.add_from_huggingface( - "uoft-cs/cifar10", - split="train", - image_key="img", - label_key="label", - max_samples=num_samples, - ) - if skipped > 0: - print(f"Loaded {added} samples ({skipped} already present)") - else: - print(f"Loaded {added} samples") - - print("Computing embeddings...") - dataset.compute_embeddings(show_progress=True) - print("Embeddings computed") - - print("Computing visualizations...") - # Compute both euclidean and poincare layouts - dataset.compute_visualization(geometry="euclidean") - dataset.compute_visualization(geometry="poincare") - print("Visualizations ready") - - launch(dataset, port=port, host=host, open_browser=open_browser, reuse_server=reuse_server) - - -def serve_dataset( - filepath: str, - port: int = 6262, - *, - host: str = "127.0.0.1", - open_browser: bool = True, - reuse_server: bool = False, -): - """Serve a saved dataset.""" - from hyperview import Dataset, launch - - print(f"Loading dataset from {filepath}...") - dataset = Dataset.load(filepath) - print(f"Loaded {len(dataset)} samples") - - launch(dataset, port=port, host=host, open_browser=open_browser, reuse_server=reuse_server) - - -if __name__ == "__main__": - main() diff --git a/src/hyperview/core/__init__.py b/src/hyperview/core/__init__.py deleted file mode 100644 index 2f09c78cc3b71fb293208678553e3774c9bf0032..0000000000000000000000000000000000000000 --- a/src/hyperview/core/__init__.py +++ /dev/null @@ -1,6 +0,0 @@ -"""Core data structures for HyperView.""" - -from hyperview.core.dataset import Dataset -from hyperview.core.sample import Sample - -__all__ = ["Dataset", "Sample"] diff --git a/src/hyperview/core/dataset.py b/src/hyperview/core/dataset.py deleted file mode 100644 index a943e8a419d7c3a5ebe755ebf6b2d8931f939b1a..0000000000000000000000000000000000000000 --- a/src/hyperview/core/dataset.py +++ /dev/null @@ -1,670 +0,0 @@ -"""Dataset class for managing collections of samples.""" - -from __future__ import annotations - -import hashlib -import json -import uuid -from collections.abc import Callable, Iterator -from pathlib import Path -from typing import Any, cast - -import numpy as np -from datasets import DownloadConfig, load_dataset -from PIL import Image - -from hyperview.core.sample import Sample -from hyperview.storage.backend import StorageBackend -from hyperview.storage.schema import make_layout_key - - -class Dataset: - """A collection of samples with support for embeddings and visualization. - - Datasets are automatically persisted to LanceDB by default, providing: - - Automatic persistence (no need to call save()) - - Vector similarity search - - Efficient storage and retrieval - - Embeddings are stored separately from samples, keyed by model_id. - Layouts (2D projections) are stored per layout_key (space + method). - - Examples: - # Create a new dataset (auto-persisted) - dataset = hv.Dataset("my_dataset") - dataset.add_images_dir("/path/to/images") - - # Create an in-memory dataset (for testing) - dataset = hv.Dataset("temp", persist=False) - """ - - def __init__( - self, - name: str | None = None, - persist: bool = True, - storage: StorageBackend | None = None, - ): - """Initialize a new dataset. - - Args: - name: Optional name for the dataset. - persist: If True (default), use LanceDB for persistence. - If False, use in-memory storage. - storage: Optional custom storage backend. If provided, persist is ignored. - """ - self.name = name or f"dataset_{uuid.uuid4().hex[:8]}" - - # Initialize storage backend - if storage is not None: - self._storage = storage - elif persist: - from hyperview.storage import LanceDBBackend, StorageConfig - - config = StorageConfig.default() - self._storage = LanceDBBackend(self.name, config) - else: - from hyperview.storage import MemoryBackend - self._storage = MemoryBackend(self.name) - - # Color palette for deterministic label color assignment - _COLOR_PALETTE = [ - "#e6194b", "#3cb44b", "#ffe119", "#4363d8", "#f58231", - "#911eb4", "#46f0f0", "#f032e6", "#bcf60c", "#fabebe", - "#008080", "#e6beff", "#9a6324", "#fffac8", "#800000", - "#aaffc3", "#808000", "#ffd8b1", "#000075", "#808080", - ] - - def __len__(self) -> int: - return len(self._storage) - - def __iter__(self) -> Iterator[Sample]: - return iter(self._storage) - - def __getitem__(self, sample_id: str) -> Sample: - sample = self._storage.get_sample(sample_id) - if sample is None: - raise KeyError(sample_id) - return sample - - def add_sample(self, sample: Sample) -> None: - """Add a sample to the dataset (idempotent).""" - self._storage.add_sample(sample) - - def _ingest_samples( - self, - samples: list[Sample], - *, - skip_existing: bool = True, - ) -> tuple[int, int]: - """Shared ingestion helper for batch sample insertion. - - Handles deduplication uniformly. - - Args: - samples: List of samples to ingest. - skip_existing: If True, skip samples that already exist in storage. - - Returns: - Tuple of (num_added, num_skipped). - """ - if not samples: - return 0, 0 - - skipped = 0 - if skip_existing: - all_ids = [s.id for s in samples] - existing_ids = self._storage.get_existing_ids(all_ids) - if existing_ids: - samples = [s for s in samples if s.id not in existing_ids] - skipped = len(all_ids) - len(samples) - - if not samples: - return 0, skipped - - self._storage.add_samples_batch(samples) - - return len(samples), skipped - - def add_image( - self, - filepath: str, - label: str | None = None, - metadata: dict[str, Any] | None = None, - sample_id: str | None = None, - ) -> Sample: - """Add a single image to the dataset. - - Args: - filepath: Path to the image file. - label: Optional label for the image. - metadata: Optional metadata dictionary. - sample_id: Optional custom ID. If not provided, one will be generated. - - Returns: - The created Sample. - """ - if sample_id is None: - sample_id = hashlib.md5(filepath.encode()).hexdigest()[:12] - - sample = Sample( - id=sample_id, - filepath=filepath, - label=label, - metadata=metadata or {}, - ) - self.add_sample(sample) - return sample - - def add_images_dir( - self, - directory: str, - extensions: tuple[str, ...] = (".jpg", ".jpeg", ".png", ".webp"), - label_from_folder: bool = False, - recursive: bool = True, - skip_existing: bool = True, - ) -> tuple[int, int]: - """Add all images from a directory. - - Args: - directory: Path to the directory containing images. - extensions: Tuple of valid file extensions. - label_from_folder: If True, use parent folder name as label. - recursive: If True, search subdirectories. - skip_existing: If True (default), skip samples that already exist. - - Returns: - Tuple of (num_added, num_skipped). - """ - directory_path = Path(directory) - if not directory_path.exists(): - raise ValueError(f"Directory does not exist: {directory_path}") - - samples = [] - pattern = "**/*" if recursive else "*" - - for path in directory_path.glob(pattern): - if path.is_file() and path.suffix.lower() in extensions: - label = path.parent.name if label_from_folder else None - sample_id = hashlib.md5(str(path).encode()).hexdigest()[:12] - sample = Sample( - id=sample_id, - filepath=str(path), - label=label, - metadata={}, - ) - samples.append(sample) - - # Use shared ingestion helper - return self._ingest_samples(samples, skip_existing=skip_existing) - - def add_from_huggingface( - self, - dataset_name: str, - split: str = "train", - image_key: str = "img", - label_key: str | None = "fine_label", - label_names_key: str | None = None, - max_samples: int | None = None, - show_progress: bool = True, - skip_existing: bool = True, - image_format: str = "auto", - ) -> tuple[int, int]: - """Load samples from a HuggingFace dataset. - - Images are downloaded to disk at ~/.hyperview/media/huggingface/{dataset}/{split}/ - This ensures images persist across sessions and embeddings can be computed - at any time, similar to FiftyOne's approach. - - Args: - dataset_name: Name of the HuggingFace dataset. - split: Dataset split to use. - image_key: Key for the image column. - label_key: Key for the label column (can be None). - label_names_key: Key for label names in dataset info. - max_samples: Maximum number of samples to load. - show_progress: Whether to print progress. - skip_existing: If True (default), skip samples that already exist in storage. - image_format: Image format to save: "auto" (detect from source, fallback PNG), - "png" (lossless), or "jpeg" (smaller files). - - Returns: - Tuple of (num_added, num_skipped). - """ - from hyperview.storage import StorageConfig - - # HuggingFace `load_dataset()` can be surprisingly slow even when the dataset - # is already cached, due to Hub reachability checks in some environments. - # For a fast path, first try loading in "offline" mode (cache-only), and - # fall back to an online load if the dataset isn't cached yet. - try: - ds = cast( - Any, - load_dataset( - dataset_name, - split=split, - download_config=DownloadConfig(local_files_only=True), - ), - ) - except Exception: - ds = cast(Any, load_dataset(dataset_name, split=split)) - - # Get label names if available - label_names = None - if label_key and label_names_key: - if label_names_key in ds.features: - label_names = ds.features[label_names_key].names - elif label_key: - if hasattr(ds.features[label_key], "names"): - label_names = ds.features[label_key].names - - # Extract dataset metadata for robust sample IDs - config_name = getattr(ds.info, "config_name", None) or "default" - fingerprint = ds._fingerprint[:8] if hasattr(ds, "_fingerprint") and ds._fingerprint else "unknown" - version = str(ds.info.version) if ds.info.version else None - - # Get media directory for this dataset - config = StorageConfig.default() - media_dir = config.get_huggingface_media_dir(dataset_name, split) - - samples = [] - total = len(ds) if max_samples is None else min(len(ds), max_samples) - - if show_progress: - print(f"Loading {total} samples from {dataset_name}...") - - iterator = range(total) - - for i in iterator: - item = ds[i] - image = item[image_key] - - # Handle PIL Image or numpy array - if isinstance(image, Image.Image): - pil_image = image - else: - pil_image = Image.fromarray(np.asarray(image)) - - # Get label - label = None - if label_key and label_key in item: - label_idx = item[label_key] - if label_names and isinstance(label_idx, int): - label = label_names[label_idx] - else: - label = str(label_idx) - - # Generate robust sample ID with config and fingerprint - safe_name = dataset_name.replace("/", "_") - sample_id = f"{safe_name}_{config_name}_{fingerprint}_{split}_{i}" - - # Determine image format and extension - if image_format == "auto": - # Try to preserve original format, fallback to PNG - original_format = getattr(pil_image, "format", None) - if original_format in ("JPEG", "JPG"): - save_format = "JPEG" - ext = ".jpg" - else: - save_format = "PNG" - ext = ".png" - elif image_format == "jpeg": - save_format = "JPEG" - ext = ".jpg" - else: - save_format = "PNG" - ext = ".png" - - # Enhanced metadata with dataset info - metadata = { - "source": dataset_name, - "config": config_name, - "split": split, - "index": i, - "fingerprint": ds._fingerprint if hasattr(ds, "_fingerprint") else None, - "version": version, - } - - image_path = media_dir / f"{sample_id}{ext}" - if not image_path.exists(): - if save_format == "JPEG" or pil_image.mode in ("RGBA", "P", "L"): - pil_image = pil_image.convert("RGB") - pil_image.save(image_path, format=save_format) - - sample = Sample( - id=sample_id, - filepath=str(image_path), - label=label, - metadata=metadata, - ) - - samples.append(sample) - - # Use shared ingestion helper - num_added, skipped = self._ingest_samples(samples, skip_existing=skip_existing) - - if show_progress: - print(f"Images saved to: {media_dir}") - if skipped > 0: - print(f"Skipped {skipped} existing samples") - - return num_added, skipped - - def compute_embeddings( - self, - model: str | ModelSpec = "openai/clip-vit-base-patch32", - batch_size: int = 32, - show_progress: bool = True, - ) -> str: - """Compute embeddings for samples that don't have them yet. - - Embeddings are stored in a dedicated space keyed by model_id. - - Args: - model: EmbedAnything HuggingFace `model_id` to use. - batch_size: Batch size for processing. - show_progress: Whether to show progress bar. - - Returns: - space_key for the embedding space. - """ - from hyperview.embeddings.pipelines import compute_embeddings - from hyperview.embeddings.providers import ModelSpec - - if isinstance(model, ModelSpec): - model_spec = model - else: - model_spec = ModelSpec(provider="embed_anything", model_id=model) - space_key, _num_computed, _num_skipped = compute_embeddings( - storage=self._storage, - model_spec=model_spec, - batch_size=batch_size, - show_progress=show_progress, - ) - return space_key - - def compute_visualization( - self, - space_key: str | None = None, - method: str = "umap", - geometry: str = "euclidean", - n_neighbors: int = 15, - min_dist: float = 0.1, - metric: str = "cosine", - force: bool = False, - ) -> str: - """Compute 2D projections for visualization. - - Args: - space_key: Embedding space to project. If None, uses the first available. - method: Projection method ('umap' supported). - geometry: Output geometry type ('euclidean' or 'poincare'). - n_neighbors: Number of neighbors for UMAP. - min_dist: Minimum distance for UMAP. - metric: Distance metric for UMAP. - force: Force recomputation even if layout exists. - - Returns: - layout_key for the computed layout. - """ - from hyperview.embeddings.pipelines import compute_layout - - return compute_layout( - storage=self._storage, - space_key=space_key, - method=method, - geometry=geometry, - n_neighbors=n_neighbors, - min_dist=min_dist, - metric=metric, - force=force, - show_progress=True, - ) - - def list_spaces(self) -> list[Any]: - """List all embedding spaces in this dataset.""" - return self._storage.list_spaces() - - def list_layouts(self) -> list[Any]: - """List all layouts in this dataset (returns LayoutInfo objects).""" - return self._storage.list_layouts() - - def find_similar( - self, - sample_id: str, - k: int = 10, - space_key: str | None = None, - ) -> list[tuple[Sample, float]]: - """Find k most similar samples to a given sample. - - Args: - sample_id: ID of the query sample. - k: Number of neighbors to return. - space_key: Embedding space to search in. If None, uses first available. - - Returns: - List of (sample, distance) tuples, sorted by distance ascending. - """ - return self._storage.find_similar(sample_id, k, space_key) - - def find_similar_by_vector( - self, - vector: list[float], - k: int = 10, - space_key: str | None = None, - ) -> list[tuple[Sample, float]]: - """Find k most similar samples to a given vector. - - Args: - vector: Query vector. - k: Number of neighbors to return. - space_key: Embedding space to search in. If None, uses first available. - - Returns: - List of (sample, distance) tuples, sorted by distance ascending. - """ - return self._storage.find_similar_by_vector(vector, k, space_key) - - @staticmethod - def _compute_label_color(label: str, palette: list[str]) -> str: - """Compute a deterministic color for a label.""" - digest = hashlib.md5(label.encode("utf-8")).digest() - idx = int.from_bytes(digest[:4], "big") % len(palette) - return palette[idx] - - def get_label_colors(self) -> dict[str, str]: - """Get the color mapping for labels (computed deterministically).""" - labels = self._storage.get_unique_labels() - return {label: self._compute_label_color(label, self._COLOR_PALETTE) for label in labels} - - def set_coords( - self, - geometry: str, - ids: list[str], - coords: np.ndarray | list[list[float]], - ) -> str: - """Set precomputed 2D coordinates for visualization. - - Use this when you have precomputed 2D projections and want to skip - embedding computation. Useful for smoke tests or external projections. - - Args: - geometry: "euclidean" or "poincare". - ids: List of sample IDs. - coords: (N, 2) array of coordinates. - - Returns: - The layout_key for the stored coordinates. - - Example: - >>> dataset.set_coords("euclidean", ["s0", "s1"], [[0.1, 0.2], [0.3, 0.4]]) - >>> dataset.set_coords("poincare", ["s0", "s1"], [[0.1, 0.2], [0.3, 0.4]]) - >>> hv.launch(dataset) - """ - if geometry not in ("euclidean", "poincare"): - raise ValueError(f"geometry must be 'euclidean' or 'poincare', got '{geometry}'") - - coords_arr = np.asarray(coords, dtype=np.float32) - if coords_arr.ndim != 2 or coords_arr.shape[1] != 2: - raise ValueError(f"coords must be (N, 2), got shape {coords_arr.shape}") - - # Ensure a synthetic space exists (required by launch()) - space_key = "precomputed" - if not any(s.space_key == space_key for s in self._storage.list_spaces()): - precomputed_config = { - "provider": "precomputed", - "geometry": "unknown", # Precomputed coords don't have a source embedding geometry - } - self._storage.ensure_space(space_key, dim=2, config=precomputed_config) - - layout_key = make_layout_key(space_key, method="precomputed", geometry=geometry) - - # Ensure layout registry entry exists - self._storage.ensure_layout( - layout_key=layout_key, - space_key=space_key, - method="precomputed", - geometry=geometry, - params=None, - ) - - self._storage.add_layout_coords(layout_key, list(ids), coords_arr) - return layout_key - - @property - def samples(self) -> list[Sample]: - """Get all samples as a list.""" - return self._storage.get_all_samples() - - @property - def labels(self) -> list[str]: - """Get unique labels in the dataset.""" - return self._storage.get_unique_labels() - - def filter(self, predicate: Callable[[Sample], bool]) -> list[Sample]: - """Filter samples based on a predicate function.""" - return self._storage.filter(predicate) - - def get_samples_paginated( - self, - offset: int = 0, - limit: int = 100, - label: str | None = None, - ) -> tuple[list[Sample], int]: - """Get paginated samples. - - This avoids loading all samples into memory and is used by the server - API for efficient pagination. - """ - return self._storage.get_samples_paginated(offset=offset, limit=limit, label=label) - - def get_samples_by_ids(self, sample_ids: list[str]) -> list[Sample]: - """Retrieve multiple samples by ID. - - The returned list is aligned to the input order and skips missing IDs. - """ - return self._storage.get_samples_by_ids(sample_ids) - - def get_visualization_data( - self, - layout_key: str, - ) -> tuple[list[str], list[str | None], np.ndarray]: - """Get visualization data (ids, labels, coords) for a layout.""" - layout_ids, layout_coords = self._storage.get_layout_coords(layout_key) - if not layout_ids: - return [], [], np.empty((0, 2), dtype=np.float32) - - labels_by_id = self._storage.get_labels_by_ids(layout_ids) - - ids: list[str] = [] - labels: list[str | None] = [] - coords: list[np.ndarray] = [] - - for i, sample_id in enumerate(layout_ids): - if sample_id in labels_by_id: - ids.append(sample_id) - labels.append(labels_by_id[sample_id]) - coords.append(layout_coords[i]) - - if not coords: - return [], [], np.empty((0, 2), dtype=np.float32) - - return ids, labels, np.asarray(coords, dtype=np.float32) - - - def get_lasso_candidates_aabb( - self, - *, - layout_key: str, - x_min: float, - x_max: float, - y_min: float, - y_max: float, - ) -> tuple[list[str], np.ndarray]: - """Return candidate (id, xy) rows within an AABB for a layout.""" - return self._storage.get_lasso_candidates_aabb( - layout_key=layout_key, - x_min=x_min, - x_max=x_max, - y_min=y_min, - y_max=y_max, - ) - - def save(self, filepath: str, include_thumbnails: bool = True) -> None: - """Export dataset to a JSON file. - - Args: - filepath: Path to save the JSON file. - include_thumbnails: Whether to include cached thumbnails. - """ - samples = self._storage.get_all_samples() - if include_thumbnails: - for s in samples: - s.cache_thumbnail() - - data = { - "name": self.name, - "samples": [ - { - "id": s.id, - "filepath": s.filepath, - "label": s.label, - "metadata": s.metadata, - "thumbnail_base64": s.thumbnail_base64 if include_thumbnails else None, - } - for s in samples - ], - } - with open(filepath, "w") as f: - json.dump(data, f) - - @classmethod - def load(cls, filepath: str, persist: bool = False) -> "Dataset": - """Load dataset from a JSON file. - - Args: - filepath: Path to the JSON file. - persist: If True, persist the loaded data to LanceDB. - If False (default), keep in memory only. - - Returns: - Dataset instance. - """ - with open(filepath) as f: - data = json.load(f) - - dataset = cls(name=data["name"], persist=persist) - - # Add samples - samples = [] - for s_data in data["samples"]: - sample = Sample( - id=s_data["id"], - filepath=s_data["filepath"], - label=s_data.get("label"), - metadata=s_data.get("metadata", {}), - thumbnail_base64=s_data.get("thumbnail_base64"), - ) - samples.append(sample) - - dataset._storage.add_samples_batch(samples) - return dataset diff --git a/src/hyperview/core/sample.py b/src/hyperview/core/sample.py deleted file mode 100644 index d0e3f38afb2541915f830ad7278e5e225b1a141d..0000000000000000000000000000000000000000 --- a/src/hyperview/core/sample.py +++ /dev/null @@ -1,95 +0,0 @@ -"""Sample class representing a single data point in a dataset.""" - -import base64 -import io -from pathlib import Path -from typing import Any - -from PIL import Image -from pydantic import BaseModel, Field - - -class Sample(BaseModel): - """A single sample in a HyperView dataset. - - Samples are pure metadata containers. Embeddings and layouts are stored - separately in dedicated tables (per embedding space / per layout). - """ - - id: str = Field(..., description="Unique identifier for the sample") - filepath: str = Field(..., description="Path to the image file") - label: str | None = Field(default=None, description="Label for the sample") - metadata: dict[str, Any] = Field(default_factory=dict, description="Additional metadata") - thumbnail_base64: str | None = Field(default=None, description="Cached thumbnail as base64") - width: int | None = Field(default=None, description="Image width in pixels") - height: int | None = Field(default=None, description="Image height in pixels") - - model_config = {"arbitrary_types_allowed": True} - - @property - def filename(self) -> str: - """Get the filename from the filepath.""" - return Path(self.filepath).name - - def load_image(self) -> Image.Image: - """Load the image from disk.""" - return Image.open(self.filepath) - - def get_thumbnail(self, size: tuple[int, int] = (128, 128)) -> Image.Image: - """Get a thumbnail of the image. Also captures original dimensions.""" - img = self.load_image() - # Capture original dimensions while we have the image loaded - if self.width is None or self.height is None: - self.width, self.height = img.size - img.thumbnail(size, Image.Resampling.LANCZOS) - return img - - def _encode_thumbnail(self, size: tuple[int, int] = (128, 128)) -> str: - """Encode thumbnail as base64 JPEG.""" - thumb = self.get_thumbnail(size) - if thumb.mode in ("RGBA", "P"): - thumb = thumb.convert("RGB") - buffer = io.BytesIO() - thumb.save(buffer, format="JPEG", quality=85) - return base64.b64encode(buffer.getvalue()).decode("utf-8") - - def get_thumbnail_base64(self, size: tuple[int, int] = (128, 128)) -> str: - """Get thumbnail as base64 encoded string.""" - return self.thumbnail_base64 or self._encode_thumbnail(size) - - def cache_thumbnail(self, size: tuple[int, int] = (128, 128)) -> None: - """Cache the thumbnail as base64 for persistence.""" - if self.thumbnail_base64 is None: - self.thumbnail_base64 = self._encode_thumbnail(size) - - def to_api_dict(self, include_thumbnail: bool = True) -> dict[str, Any]: - """Convert to dictionary for API response.""" - # Ensure dimensions are populated (loads image if needed but not cached) - if self.width is None or self.height is None: - self.ensure_dimensions() - - data = { - "id": self.id, - "filepath": self.filepath, - "filename": self.filename, - "label": self.label, - "metadata": self.metadata, - "width": self.width, - "height": self.height, - } - if include_thumbnail: - data["thumbnail"] = self.get_thumbnail_base64() - return data - - def ensure_dimensions(self) -> None: - """Load image dimensions if not already set.""" - if self.width is None or self.height is None: - try: - img = self.load_image() - self.width, self.height = img.size - except Exception: - # If image can't be loaded, leave as None - pass - - - diff --git a/src/hyperview/core/selection.py b/src/hyperview/core/selection.py deleted file mode 100644 index 1dee065d6b8e177425439152def48cd132406aa5..0000000000000000000000000000000000000000 --- a/src/hyperview/core/selection.py +++ /dev/null @@ -1,53 +0,0 @@ -"""Selection / geometry helpers. - -This module contains small, backend-agnostic utilities used by selection endpoints -(e.g. lasso selection over 2D embeddings). -""" - -from __future__ import annotations - -import numpy as np - - -def points_in_polygon(points_xy: np.ndarray, polygon_xy: np.ndarray) -> np.ndarray: - """Vectorized point-in-polygon (even-odd rule / ray casting). - - Args: - points_xy: Array of shape (m, 2) with point coordinates. - polygon_xy: Array of shape (n, 2) with polygon vertices. - - Returns: - Boolean mask of length m, True where point lies inside polygon. - - Notes: - Boundary points may be classified as outside depending on floating point - ties (common for lasso selection tools). - """ - if polygon_xy.shape[0] < 3: - return np.zeros((points_xy.shape[0],), dtype=bool) - - x = points_xy[:, 0] - y = points_xy[:, 1] - poly_x = polygon_xy[:, 0] - poly_y = polygon_xy[:, 1] - - inside = np.zeros((points_xy.shape[0],), dtype=bool) - j = polygon_xy.shape[0] - 1 - - for i in range(polygon_xy.shape[0]): - xi = poly_x[i] - yi = poly_y[i] - xj = poly_x[j] - yj = poly_y[j] - - # Half-open y-interval to avoid double-counting vertices. - intersects = (yi > y) != (yj > y) - - denom = yj - yi - # denom == 0 => intersects is always False; add tiny epsilon to avoid warnings. - x_intersect = (xj - xi) * (y - yi) / (denom + 1e-30) + xi - - inside ^= intersects & (x < x_intersect) - j = i - - return inside diff --git a/src/hyperview/embeddings/__init__.py b/src/hyperview/embeddings/__init__.py deleted file mode 100644 index 14ff04dcdfcc580a95b4a6f9eb80014de80c1a96..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/__init__.py +++ /dev/null @@ -1,37 +0,0 @@ -"""Embedding computation, projection, and provider modules.""" - -from hyperview.embeddings.compute import EmbeddingComputer -from hyperview.embeddings.providers import ( - BaseEmbeddingProvider, - ModelSpec, - get_provider, - list_providers, - make_provider_aware_space_key, - register_provider, -) - - -def __getattr__(name: str): - """Lazy import for heavy dependencies (UMAP/numba).""" - if name == "ProjectionEngine": - from hyperview.embeddings.projection import ProjectionEngine - return ProjectionEngine - if name == "EmbedAnythingProvider": - from hyperview.embeddings.providers.embed_anything import EmbedAnythingProvider - return EmbedAnythingProvider - raise AttributeError(f"module {__name__!r} has no attribute {name!r}") - - -__all__ = [ - "EmbeddingComputer", - "ProjectionEngine", - # Provider types - "BaseEmbeddingProvider", - "EmbedAnythingProvider", - "ModelSpec", - # Provider utilities - "get_provider", - "list_providers", - "register_provider", - "make_provider_aware_space_key", -] diff --git a/src/hyperview/embeddings/compute.py b/src/hyperview/embeddings/compute.py deleted file mode 100644 index d2026b528bc3b37b74aafd84c35cee71fb8d8fc4..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/compute.py +++ /dev/null @@ -1,89 +0,0 @@ -"""Image embedding computation via EmbedAnything.""" - -import os -import tempfile -from pathlib import Path - -import numpy as np -from embed_anything import EmbeddingModel -from PIL import Image - -from hyperview.core.sample import Sample - - -class EmbeddingComputer: - """Compute embeddings for image samples using EmbedAnything.""" - - def __init__(self, model: str): - """Initialize the embedding computer. - - Args: - model: HuggingFace model ID to load via EmbedAnything. - """ - if not model or not model.strip(): - raise ValueError("model must be a non-empty HuggingFace model_id") - - self.model_id = model - self._model: EmbeddingModel | None = None - - def _get_model(self) -> EmbeddingModel: - """Lazily initialize the EmbedAnything model.""" - if self._model is None: - self._model = EmbeddingModel.from_pretrained_hf(model_id=self.model_id) - return self._model - - def _load_rgb_image(self, sample: Sample) -> Image.Image: - """Load an image and normalize it to RGB. - - For file-backed samples, returns an in-memory copy and closes the file - handle immediately to avoid leaking descriptors during batch processing. - """ - with sample.load_image() as img: - img.load() - if img.mode != "RGB": - return img.convert("RGB") - return img.copy() - - def _embed_file(self, file_path: str) -> np.ndarray: - model = self._get_model() - result = model.embed_file(file_path) - - if not result: - raise RuntimeError(f"EmbedAnything returned no embeddings for: {file_path}") - if len(result) != 1: - raise RuntimeError( - f"Expected 1 embedding for an image file, got {len(result)}: {file_path}" - ) - - return np.asarray(result[0].embedding, dtype=np.float32) - - def _embed_pil_image(self, image: Image.Image) -> np.ndarray: - temp_fd, temp_path = tempfile.mkstemp(suffix=".png") - os.close(temp_fd) - try: - image.save(temp_path, format="PNG") - return self._embed_file(temp_path) - finally: - Path(temp_path).unlink(missing_ok=True) - - def compute_single(self, sample: Sample) -> np.ndarray: - """Compute embedding for a single sample.""" - image = self._load_rgb_image(sample) - return self._embed_pil_image(image) - - def compute_batch( - self, - samples: list[Sample], - batch_size: int = 32, - show_progress: bool = True, - ) -> list[np.ndarray]: - """Compute embeddings for a list of samples.""" - if batch_size <= 0: - raise ValueError("batch_size must be > 0") - self._get_model() - - if show_progress: - print(f"Computing embeddings for {len(samples)} samples...") - - return [self.compute_single(sample) for sample in samples] - diff --git a/src/hyperview/embeddings/pipelines.py b/src/hyperview/embeddings/pipelines.py deleted file mode 100644 index 578a9f363205a9ccd1efb186b33392c4b9337891..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/pipelines.py +++ /dev/null @@ -1,191 +0,0 @@ -"""Compute orchestration pipelines for HyperView. - -These functions coordinate embedding computation and 2D layout/projection -computation, persisting results into the configured storage backend. -""" - -from __future__ import annotations - -import numpy as np - -from hyperview.embeddings.providers import ModelSpec, get_provider, make_provider_aware_space_key -from hyperview.storage.backend import StorageBackend -from hyperview.storage.schema import make_layout_key - - -def compute_embeddings( - storage: StorageBackend, - model_spec: ModelSpec, - batch_size: int = 32, - show_progress: bool = True, -) -> tuple[str, int, int]: - """Compute embeddings for samples that don't have them yet. - - Args: - storage: Storage backend to read samples from and write embeddings to. - model_spec: Model specification (provider, model_id, geometry, etc.). - batch_size: Batch size for processing. - show_progress: Whether to show progress bar. - - Returns: - Tuple of (space_key, num_computed, num_skipped). - - Raises: - ValueError: If no samples in storage or provider not found. - """ - provider = get_provider(model_spec.provider) - - all_samples = storage.get_all_samples() - if not all_samples: - raise ValueError("No samples in storage") - - # Generate space key before computing (deterministic from spec) - space_key = make_provider_aware_space_key(model_spec) - - # Check which samples need embeddings - missing_ids = storage.get_missing_embedding_ids(space_key) - - # If space doesn't exist yet, all samples are missing - if not storage.get_space(space_key): - missing_ids = [s.id for s in all_samples] - - num_skipped = len(all_samples) - len(missing_ids) - - if not missing_ids: - if show_progress: - print(f"All {len(all_samples)} samples already have embeddings in space '{space_key}'") - return space_key, 0, num_skipped - - samples_to_embed = storage.get_samples_by_ids(missing_ids) - - if show_progress and num_skipped > 0: - print(f"Skipped {num_skipped} samples with existing embeddings") - - # Compute all embeddings in one pass (no separate probe) - embeddings = provider.compute_embeddings( - samples=samples_to_embed, - model_spec=model_spec, - batch_size=batch_size, - show_progress=show_progress, - ) - - dim = embeddings.shape[1] - - # Ensure space exists (create if needed) - config = provider.get_space_config(model_spec, dim) - storage.ensure_space( - model_id=model_spec.model_id, - dim=dim, - config=config, - space_key=space_key, - ) - - # Store embeddings - ids = [s.id for s in samples_to_embed] - storage.add_embeddings(space_key, ids, embeddings) - - return space_key, len(ids), num_skipped - - -def compute_layout( - storage: StorageBackend, - space_key: str | None = None, - method: str = "umap", - geometry: str = "euclidean", - n_neighbors: int = 15, - min_dist: float = 0.1, - metric: str = "cosine", - force: bool = False, - show_progress: bool = True, -) -> str: - """Compute 2D layout/projection for visualization. - - Args: - storage: Storage backend with embeddings. - space_key: Embedding space to project. If None, uses the first available. - method: Projection method ('umap' supported). - geometry: Output geometry type ('euclidean' or 'poincare'). - n_neighbors: Number of neighbors for UMAP. - min_dist: Minimum distance for UMAP. - metric: Distance metric for UMAP. - force: Force recomputation even if layout exists. - show_progress: Whether to print progress messages. - - Returns: - layout_key for the computed layout. - - Raises: - ValueError: If no embedding spaces, space not found, or insufficient samples. - """ - from hyperview.embeddings.projection import ProjectionEngine - - if method != "umap": - raise ValueError(f"Invalid method: {method}. Only 'umap' is supported.") - - if geometry not in ("euclidean", "poincare"): - raise ValueError(f"Invalid geometry: {geometry}. Must be 'euclidean' or 'poincare'.") - - if space_key is None: - spaces = storage.list_spaces() - if not spaces: - raise ValueError("No embedding spaces. Call compute_embeddings() first.") - space_key = spaces[0].space_key - - space = storage.get_space(space_key) - if space is None: - raise ValueError(f"Space not found: {space_key}") - - input_geometry = space.geometry - curvature = (space.config or {}).get("curvature") - - ids, vectors = storage.get_embeddings(space_key) - if len(ids) == 0: - raise ValueError(f"No embeddings in space '{space_key}'. Call compute_embeddings() first.") - - if len(ids) < 3: - raise ValueError(f"Need at least 3 samples for visualization, have {len(ids)}") - - layout_params = { - "n_neighbors": n_neighbors, - "min_dist": min_dist, - "metric": metric, - } - layout_key = make_layout_key(space_key, method, geometry, layout_params) - - if not force: - existing_layout = storage.get_layout(layout_key) - if existing_layout is not None: - existing_ids, _ = storage.get_layout_coords(layout_key) - if set(existing_ids) == set(ids): - if show_progress: - print(f"Layout '{layout_key}' already exists with {len(ids)} points") - return layout_key - if show_progress: - print("Layout exists but has different samples, recomputing...") - - if show_progress: - print(f"Computing {geometry} {method} layout for {len(ids)} samples...") - - storage.ensure_layout( - layout_key=layout_key, - space_key=space_key, - method=method, - geometry=geometry, - params=layout_params, - ) - - engine = ProjectionEngine() - coords = engine.project( - vectors, - input_geometry=input_geometry, - output_geometry=geometry, - curvature=curvature, - method=method, - n_neighbors=n_neighbors, - min_dist=min_dist, - metric=metric, - ) - - storage.add_layout_coords(layout_key, ids, coords) - - return layout_key diff --git a/src/hyperview/embeddings/projection.py b/src/hyperview/embeddings/projection.py deleted file mode 100644 index 83dff50655cf11f5dd5984e0a1710afd852d3e43..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/projection.py +++ /dev/null @@ -1,267 +0,0 @@ -"""Projection methods for dimensionality reduction.""" - -import logging -import warnings - -import numpy as np -import umap - -logger = logging.getLogger(__name__) - - -class ProjectionEngine: - """Engine for projecting high-dimensional embeddings to 2D.""" - - def to_poincare_ball( - self, - hyperboloid_embeddings: np.ndarray, - curvature: float | None = None, - clamp_radius: float = 0.999999, - ) -> np.ndarray: - """Convert hyperboloid (Lorentz) coordinates to Poincaré ball coordinates. - - Input is expected to be shape (N, D+1) with first coordinate being time-like. - Points are assumed to satisfy: t^2 - ||x||^2 = 1/c (c > 0). - - Returns Poincaré ball coordinates of shape (N, D) in the unit ball. - - Notes: - - Many hyperbolic libraries parameterize curvature as a positive number c - where the manifold has sectional curvature -c. - - We map to the unit ball for downstream distance metrics (UMAP 'poincare'). - """ - if hyperboloid_embeddings.ndim != 2 or hyperboloid_embeddings.shape[1] < 2: - raise ValueError( - "hyperboloid_embeddings must have shape (N, D+1) with D>=1" - ) - - c = float(curvature) if curvature is not None else 1.0 - if c <= 0: - raise ValueError(f"curvature must be > 0, got {c}") - - # Radius R = 1/sqrt(c) for curvature -c - R = 1.0 / np.sqrt(c) - - t = hyperboloid_embeddings[:, :1] - x = hyperboloid_embeddings[:, 1:] - - # Map to ball radius R: u_R = x / (t + R) - denom = t + R - u_R = x / denom - - # Rescale to unit ball: u = u_R / R = sqrt(c) * u_R - u = u_R / R - - # Numerical guard: ensure inside the unit ball - radii = np.linalg.norm(u, axis=1) - mask = radii >= clamp_radius - if np.any(mask): - u[mask] = u[mask] / radii[mask][:, np.newaxis] * clamp_radius - - return u.astype(np.float32) - - def project( - self, - embeddings: np.ndarray, - *, - input_geometry: str = "euclidean", - output_geometry: str = "euclidean", - curvature: float | None = None, - method: str = "umap", - n_neighbors: int = 15, - min_dist: float = 0.1, - metric: str = "cosine", - random_state: int = 42, - ) -> np.ndarray: - """Project embeddings to 2D with geometry-aware preprocessing. - - This separates two concerns: - 1) Geometry/model transforms for the *input* embeddings (e.g. hyperboloid -> Poincaré) - 2) Dimensionality reduction / layout (currently UMAP) - - Args: - embeddings: Input embeddings (N x D) or hyperboloid (N x D+1). - input_geometry: Geometry/model of the input embeddings (euclidean, hyperboloid). - output_geometry: Geometry of the output coordinates (euclidean, poincare). - curvature: Curvature parameter for hyperbolic embeddings (positive c). - method: Layout method (currently only 'umap'). - n_neighbors: UMAP neighbors. - min_dist: UMAP min_dist. - metric: Input metric (used for euclidean inputs). - random_state: Random seed. - - Returns: - 2D coordinates (N x 2). - """ - if method != "umap": - raise ValueError(f"Invalid method: {method}. Only 'umap' is supported.") - - prepared = embeddings - prepared_metric: str = metric - - if input_geometry == "hyperboloid": - # Convert to unit Poincaré ball and use UMAP's built-in hyperbolic distance. - prepared = self.to_poincare_ball(embeddings, curvature=curvature) - prepared_metric = "poincare" - - if output_geometry == "poincare": - return self.project_to_poincare( - prepared, - n_neighbors=n_neighbors, - min_dist=min_dist, - metric=prepared_metric, - random_state=random_state, - ) - - if output_geometry == "euclidean": - return self.project_umap( - prepared, - n_neighbors=n_neighbors, - min_dist=min_dist, - metric=prepared_metric, - n_components=2, - random_state=random_state, - ) - - raise ValueError( - f"Invalid output_geometry: {output_geometry}. Must be 'euclidean' or 'poincare'." - ) - - def project_umap( - self, - embeddings: np.ndarray, - n_neighbors: int = 15, - min_dist: float = 0.1, - metric: str = "cosine", - n_components: int = 2, - random_state: int = 42, - ) -> np.ndarray: - """Project embeddings to Euclidean 2D using UMAP.""" - n_neighbors = min(n_neighbors, len(embeddings) - 1) - if n_neighbors < 2: - n_neighbors = 2 - - n_jobs = 1 if random_state is not None else -1 - - reducer = umap.UMAP( - n_neighbors=n_neighbors, - min_dist=min_dist, - n_components=n_components, - metric=metric, - random_state=random_state, - n_jobs=n_jobs, - ) - - coords = reducer.fit_transform(embeddings) - coords = self._normalize_coords(coords) - - return coords - - def project_to_poincare( - self, - embeddings: np.ndarray, - n_neighbors: int = 15, - min_dist: float = 0.1, - metric: str = "cosine", - random_state: int = 42, - ) -> np.ndarray: - """Project embeddings to the Poincaré disk using UMAP with hyperboloid output.""" - n_neighbors = min(n_neighbors, len(embeddings) - 1) - if n_neighbors < 2: - n_neighbors = 2 - - n_jobs = 1 if random_state is not None else -1 - - # Suppress warning about missing gradient for poincare metric (only affects inverse_transform) - with warnings.catch_warnings(): - warnings.filterwarnings("ignore", message="gradient function is not yet implemented") - reducer = umap.UMAP( - n_neighbors=n_neighbors, - min_dist=min_dist, - n_components=2, - metric=metric, - output_metric="hyperboloid", - random_state=random_state, - n_jobs=n_jobs, - ) - spatial_coords = reducer.fit_transform(embeddings) - - squared_norm = np.sum(spatial_coords**2, axis=1) - t = np.sqrt(1 + squared_norm) - - # Project to Poincaré disk: u = x / (1 + t) - denom = 1 + t - poincare_coords = spatial_coords / denom[:, np.newaxis] - - # Clamp to unit disk for numerical stability - radii = np.linalg.norm(poincare_coords, axis=1) - max_radius = 0.999 - mask = radii > max_radius - if np.any(mask): - logger.warning(f"Clamping {np.sum(mask)} points to unit disk.") - poincare_coords[mask] = ( - poincare_coords[mask] / radii[mask][:, np.newaxis] * max_radius - ) - - poincare_coords = self._center_poincare(poincare_coords) - poincare_coords = self._scale_poincare(poincare_coords, factor=0.65) - - return poincare_coords - - def _scale_poincare(self, coords: np.ndarray, factor: float) -> np.ndarray: - """Scale points towards the origin in hyperbolic space. - - Scales hyperbolic distance from origin by `factor`. If factor < 1, points move closer to center. - """ - radii = np.linalg.norm(coords, axis=1) - mask = radii > 1e-6 - - r = radii[mask] - r = np.minimum(r, 0.9999999) - r_new = np.tanh(factor * np.arctanh(r)) - - scale_ratios = np.ones_like(radii) - scale_ratios[mask] = r_new / r - - return coords * scale_ratios[:, np.newaxis] - - def _center_poincare(self, coords: np.ndarray) -> np.ndarray: - """Center points in the Poincaré disk using a Möbius transformation.""" - if len(coords) == 0: - return coords - - z = coords[:, 0] + 1j * coords[:, 1] - centroid = np.mean(z) - - if np.abs(centroid) > 0.99 or np.abs(centroid) < 1e-6: - return coords - - # Möbius transformation: w = (z - a) / (1 - conj(a) * z) - a = centroid - w = (z - a) / (1 - np.conj(a) * z) - - return np.stack([w.real, w.imag], axis=1) - - def _normalize_coords(self, coords: np.ndarray) -> np.ndarray: - """Normalize coordinates to [-1, 1] range.""" - if len(coords) == 0: - return coords - - coords = coords - coords.mean(axis=0) - max_abs = np.abs(coords).max() - if max_abs > 0: - coords = coords / max_abs * 0.95 - - return coords - - def poincare_distance(self, u: np.ndarray, v: np.ndarray) -> float: - """Compute the Poincaré distance between two points.""" - u_norm_sq = np.sum(u**2) - v_norm_sq = np.sum(v**2) - diff_norm_sq = np.sum((u - v) ** 2) - - u_norm_sq = min(u_norm_sq, 0.99999) - v_norm_sq = min(v_norm_sq, 0.99999) - - delta = 2 * diff_norm_sq / ((1 - u_norm_sq) * (1 - v_norm_sq)) - return np.arccosh(1 + delta) diff --git a/src/hyperview/embeddings/providers/__init__.py b/src/hyperview/embeddings/providers/__init__.py deleted file mode 100644 index 55fca86a161d201117fad86dc9af27056c2b11f4..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/providers/__init__.py +++ /dev/null @@ -1,200 +0,0 @@ -"""Embedding provider abstraction for HyperView.""" - -from __future__ import annotations - -import hashlib -from importlib import import_module -import json -from abc import ABC, abstractmethod -from dataclasses import dataclass -from typing import Any - -import numpy as np - -from hyperview.core.sample import Sample - - -@dataclass -class ModelSpec: - """Structured specification for an embedding model. - - Attributes: - provider: Provider identifier (e.g., "embed_anything", "hycoclip") - model_id: Model identifier (HuggingFace model_id, checkpoint path, etc.) - checkpoint: Optional checkpoint path or URL for weight-only models - config_path: Optional config path for models that need it - output_geometry: Geometry of the embedding space ("euclidean", "hyperboloid") - curvature: Hyperbolic curvature (only relevant for hyperbolic geometries) - """ - - provider: str - model_id: str - checkpoint: str | None = None - config_path: str | None = None - output_geometry: str = "euclidean" - curvature: float | None = None - - def to_dict(self) -> dict[str, Any]: - """Convert to JSON-serializable dict.""" - d: dict[str, Any] = { - "provider": self.provider, - "model_id": self.model_id, - "geometry": self.output_geometry, - } - if self.checkpoint: - d["checkpoint"] = self.checkpoint - if self.config_path: - d["config_path"] = self.config_path - if self.curvature is not None: - d["curvature"] = self.curvature - return d - - @classmethod - def from_dict(cls, d: dict[str, Any]) -> ModelSpec: - """Create from dict (e.g., loaded from JSON).""" - return cls( - provider=d["provider"], - model_id=d["model_id"], - checkpoint=d.get("checkpoint"), - config_path=d.get("config_path"), - output_geometry=d.get("geometry", "euclidean"), - curvature=d.get("curvature"), - ) - - def content_hash(self) -> str: - """Generate a short hash of the spec for collision-resistant keys.""" - content = json.dumps(self.to_dict(), sort_keys=True) - return hashlib.sha256(content.encode()).hexdigest()[:12] - - -class BaseEmbeddingProvider(ABC): - """Base class for embedding providers.""" - - @property - @abstractmethod - def provider_id(self) -> str: - """Unique identifier for this provider.""" - ... - - @abstractmethod - def compute_embeddings( - self, - samples: list[Sample], - model_spec: ModelSpec, - batch_size: int = 32, - show_progress: bool = True, - ) -> np.ndarray: - """Compute embeddings for samples. - - Returns: - Array of shape (N, D) where N is len(samples) and D is embedding dim. - """ - ... - - def get_space_config(self, model_spec: ModelSpec, dim: int) -> dict[str, Any]: - """Get config dict for SpaceInfo.config_json. - - Args: - model_spec: Model specification. - dim: Embedding dimension. - - Returns: - Config dict with provider, geometry, model_id, dim, and any extras. - """ - return { - **model_spec.to_dict(), - "dim": dim, - } - - -_PROVIDER_CLASSES: dict[str, type[BaseEmbeddingProvider]] = {} -_PROVIDER_INSTANCES: dict[str, BaseEmbeddingProvider] = {} - - -_KNOWN_PROVIDER_MODULES: dict[str, str] = { - "embed_anything": "hyperview.embeddings.providers.embed_anything", - "hycoclip": "hyperview.embeddings.providers.hycoclip", - "hycoclip_onnx": "hyperview.embeddings.providers.hycoclip_onnx", -} - - -def register_provider(provider_id: str, provider_class: type[BaseEmbeddingProvider]) -> None: - """Register a new embedding provider class.""" - _PROVIDER_CLASSES[provider_id] = provider_class - # Clear cached instance if re-registering - _PROVIDER_INSTANCES.pop(provider_id, None) - - -def _try_auto_register(provider_id: str, *, silent: bool = True) -> None: - """Attempt to auto-register a provider by importing its module. - - Args: - provider_id: Provider identifier. - silent: If True, swallow ImportError (used when listing providers). - If False, let ImportError propagate (used when explicitly requesting - a provider via get_provider()). - """ - - module_name = _KNOWN_PROVIDER_MODULES.get(provider_id) - if not module_name: - return - - if silent: - try: - import_module(module_name) - except ImportError: - return - else: - import_module(module_name) - - -def get_provider(provider_id: str) -> BaseEmbeddingProvider: - """Get a provider singleton instance by ID. - - Providers are cached to preserve model state across calls. - """ - if provider_id not in _PROVIDER_CLASSES: - _try_auto_register(provider_id, silent=False) - - if provider_id not in _PROVIDER_CLASSES: - available = ", ".join(sorted(_PROVIDER_CLASSES.keys())) or "(none registered)" - raise ValueError( - f"Unknown embedding provider: '{provider_id}'. " - f"Available: {available}" - ) - - if provider_id not in _PROVIDER_INSTANCES: - _PROVIDER_INSTANCES[provider_id] = _PROVIDER_CLASSES[provider_id]() - - return _PROVIDER_INSTANCES[provider_id] - - -def list_providers() -> list[str]: - """List available provider IDs.""" - # Trigger auto-registration for known providers - for pid in _KNOWN_PROVIDER_MODULES: - _try_auto_register(pid, silent=True) - return list(_PROVIDER_CLASSES.keys()) - - -def make_provider_aware_space_key(model_spec: ModelSpec) -> str: - """Generate a collision-resistant space_key from a ModelSpec. - - Format: {provider}__{slugified_model_id}__{content_hash} - """ - from hyperview.storage.schema import slugify_model_id - - slug = slugify_model_id(model_spec.model_id) - content_hash = model_spec.content_hash() - - return f"{model_spec.provider}__{slug}__{content_hash}" - - -__all__ = [ - "BaseEmbeddingProvider", - "ModelSpec", - "get_provider", - "list_providers", - "make_provider_aware_space_key", - "register_provider", -] diff --git a/src/hyperview/embeddings/providers/embed_anything.py b/src/hyperview/embeddings/providers/embed_anything.py deleted file mode 100644 index 60370df7649a916e610ef179b963c9b5c9f88503..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/providers/embed_anything.py +++ /dev/null @@ -1,57 +0,0 @@ -"""EmbedAnything embedding provider - default lightweight provider.""" - -from __future__ import annotations - -from typing import Any - -import numpy as np - -from hyperview.core.sample import Sample -from hyperview.embeddings.providers import ( - BaseEmbeddingProvider, - ModelSpec, - register_provider, -) - -__all__ = ["EmbedAnythingProvider"] - - -class EmbedAnythingProvider(BaseEmbeddingProvider): - """Default embedding provider using EmbedAnything. - - Supports HuggingFace vision models via EmbedAnything's inference engine. - Model is cached per model_id to avoid repeated initialization. - """ - - def __init__(self) -> None: - self._computers: dict[str, Any] = {} # model_id -> EmbeddingComputer - - @property - def provider_id(self) -> str: - return "embed_anything" - - def _get_computer(self, model_id: str) -> Any: - """Get or create an EmbeddingComputer for the given model_id.""" - if model_id not in self._computers: - from hyperview.embeddings.compute import EmbeddingComputer - - self._computers[model_id] = EmbeddingComputer(model=model_id) - return self._computers[model_id] - - def compute_embeddings( - self, - samples: list[Sample], - model_spec: ModelSpec, - batch_size: int = 32, - show_progress: bool = True, - ) -> np.ndarray: - """Compute embeddings using EmbedAnything.""" - computer = self._get_computer(model_spec.model_id) - embeddings = computer.compute_batch( - samples, batch_size=batch_size, show_progress=show_progress - ) - return np.array(embeddings, dtype=np.float32) - - -# Auto-register on import -register_provider("embed_anything", EmbedAnythingProvider) diff --git a/src/hyperview/embeddings/providers/hycoclip.py b/src/hyperview/embeddings/providers/hycoclip.py deleted file mode 100644 index 3f8a81c9b276916a976ffa9f601bf8614f3320d3..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/providers/hycoclip.py +++ /dev/null @@ -1,279 +0,0 @@ -"""Clean HyCoCLIP embedding provider (PyTorch) - no external hycoclip package. - -This is a minimal reimplementation that loads HyCoCLIP weights directly. -Only depends on torch, timm, and numpy. - -Architecture: -- ViT backbone (timm) -- Linear projection to embedding space -- Exponential map to hyperboloid (Lorentz model) - -Checkpoints: https://huggingface.co/avik-pal/hycoclip - -Requirements: - uv sync --extra ml -""" - -from __future__ import annotations - -import math -from pathlib import Path -from typing import Any - -import numpy as np - -from hyperview.core.sample import Sample -from hyperview.embeddings.providers import ( - BaseEmbeddingProvider, - ModelSpec, - register_provider, -) - -__all__ = ["HyCoCLIPProvider"] - - -HYCOCLIP_CHECKPOINTS: dict[str, str] = { - "hycoclip_vit_s": "hf://avik-pal/hycoclip#hycoclip_vit_s.pth", - "hycoclip_vit_b": "hf://avik-pal/hycoclip#hycoclip_vit_b.pth", - "meru_vit_s": "hf://avik-pal/hycoclip#meru_vit_s.pth", - "meru_vit_b": "hf://avik-pal/hycoclip#meru_vit_b.pth", -} - - -def _exp_map_lorentz(x: "torch.Tensor", c: float) -> "torch.Tensor": - """Exponential map from tangent space at the hyperboloid vertex. - - Maps Euclidean tangent vectors at the origin onto the Lorentz (hyperboloid) - model of hyperbolic space with curvature -c. - - Output is ordered as (t, x1, ..., xD) and satisfies: - t^2 - ||x||^2 = 1/c - - This matches HyCoCLIP/MERU exp_map0 numerics by clamping the sinh input for - stability and inferring the time component from the hyperboloid constraint. - - Args: - x: Euclidean tangent vectors at the origin, shape (..., D). - c: Positive curvature parameter (hyperbolic curvature is -c). - - Returns: - Hyperboloid coordinates, shape (..., D + 1). - """ - import torch - - if c <= 0: - raise ValueError(f"curvature c must be > 0, got {c}") - - # Compute in float32 under AMP to avoid float16/bfloat16 overflow. - if x.dtype in (torch.float16, torch.bfloat16): - x = x.float() - - sqrt_c = math.sqrt(c) - rc_xnorm = sqrt_c * torch.norm(x, dim=-1, keepdim=True) - - eps = 1e-8 - sinh_input = torch.clamp(rc_xnorm, min=eps, max=math.asinh(2**15)) - spatial = torch.sinh(sinh_input) * x / torch.clamp(rc_xnorm, min=eps) - - t = torch.sqrt((1.0 / c) + torch.sum(spatial * spatial, dim=-1, keepdim=True)) - return torch.cat([t, spatial], dim=-1) - - -def _create_encoder( - embed_dim: int = 512, - curvature: float = 0.1, - vit_model: str = "vit_small_patch16_224", -) -> "nn.Module": - """Create HyCoCLIP image encoder using timm ViT backbone.""" - import timm - import torch.nn as nn - - class HyCoCLIPImageEncoder(nn.Module): - def __init__(self) -> None: - super().__init__() - self.backbone = timm.create_model(vit_model, pretrained=False, num_classes=0) - backbone_dim = int(getattr(self.backbone, "embed_dim")) - self.proj = nn.Linear(backbone_dim, embed_dim, bias=False) - self.curvature = curvature - self.embed_dim = embed_dim - - def forward(self, x: "torch.Tensor") -> "torch.Tensor": - features = self.backbone(x) - spatial = self.proj(features) - return _exp_map_lorentz(spatial, self.curvature) - - return HyCoCLIPImageEncoder() - - -def _load_encoder(checkpoint_path: str, device: str = "cpu") -> Any: - """Load HyCoCLIP image encoder from checkpoint.""" - import torch - - ckpt = torch.load(checkpoint_path, map_location="cpu", weights_only=False) - state = ckpt["model"] - - # Extract curvature (stored as log) - curvature = torch.exp(state["curv"]).item() - - # Determine model variant from checkpoint - proj_shape = state["visual_proj.weight"].shape - embed_dim = proj_shape[0] - backbone_dim = proj_shape[1] - - vit_models = { - 384: "vit_small_patch16_224", - 768: "vit_base_patch16_224", - 1024: "vit_large_patch16_224", - } - vit_model = vit_models.get(backbone_dim, "vit_small_patch16_224") - - model = _create_encoder(embed_dim=embed_dim, curvature=curvature, vit_model=vit_model) - - # Remap checkpoint keys - new_state = {} - for key, value in state.items(): - if key.startswith("visual."): - new_state["backbone." + key[7:]] = value - elif key == "visual_proj.weight": - new_state["proj.weight"] = value - - model.load_state_dict(new_state, strict=False) - return model.to(device).eval() - - -class HyCoCLIPProvider(BaseEmbeddingProvider): - """Clean HyCoCLIP provider (PyTorch) - no hycoclip package dependency. - - Requires: torch, torchvision, timm (install via `uv sync --extra ml`) - """ - - def __init__(self) -> None: - self._model: Any = None - self._model_spec: ModelSpec | None = None - self._device: Any = None - self._transform: Any = None - - @property - def provider_id(self) -> str: - return "hycoclip" - - def _get_device(self) -> Any: - import torch - - if self._device is None: - self._device = torch.device("cuda" if torch.cuda.is_available() else "cpu") - return self._device - - def _get_transform(self) -> Any: - if self._transform is None: - from torchvision import transforms - - self._transform = transforms.Compose([ - transforms.Resize(224, transforms.InterpolationMode.BICUBIC), - transforms.CenterCrop(224), - transforms.ToTensor(), - transforms.Normalize( - mean=[0.48145466, 0.4578275, 0.40821073], - std=[0.26862954, 0.26130258, 0.27577711], - ), - ]) - return self._transform - - def _resolve_checkpoint(self, checkpoint: str) -> Path: - """Resolve checkpoint path, downloading from HuggingFace if needed.""" - # Handle HuggingFace Hub URLs: hf://repo_id#filename - if checkpoint.startswith("hf://"): - from huggingface_hub import hf_hub_download - - path = checkpoint[5:] - if "#" not in path: - raise ValueError(f"HF checkpoint must include filename: {checkpoint}") - repo_id, filename = path.split("#", 1) - return Path(hf_hub_download(repo_id=repo_id, filename=filename)).resolve() - - # Local path - path = Path(checkpoint).expanduser().resolve() - if not path.exists(): - raise FileNotFoundError(f"Checkpoint not found: {path}") - return path - - def _load_model(self, model_spec: ModelSpec) -> None: - if self._model is not None and self._model_spec == model_spec: - return - - # Auto-resolve checkpoint from model_id if not provided - checkpoint = model_spec.checkpoint - if not checkpoint: - checkpoint = HYCOCLIP_CHECKPOINTS.get(model_spec.model_id) - if not checkpoint: - available = ", ".join(sorted(HYCOCLIP_CHECKPOINTS.keys())) - raise ValueError( - f"Unknown HyCoCLIP model_id: '{model_spec.model_id}'. " - f"Known models: {available}. " - f"Or provide 'checkpoint' path explicitly." - ) - - checkpoint_path = self._resolve_checkpoint(checkpoint) - self._model = _load_encoder(str(checkpoint_path), str(self._get_device())) - self._model_spec = model_spec - - def compute_embeddings( - self, - samples: list["Sample"], - model_spec: ModelSpec, - batch_size: int = 32, - show_progress: bool = True, - ) -> np.ndarray: - """Compute hyperboloid embeddings for samples.""" - import torch - - self._load_model(model_spec) - assert self._model is not None - - device = self._get_device() - transform = self._get_transform() - - if show_progress: - print(f"Computing HyCoCLIP embeddings for {len(samples)} samples...") - - all_embeddings = [] - - for i in range(0, len(samples), batch_size): - batch_samples = samples[i : i + batch_size] - - images = [] - for sample in batch_samples: - img = sample.load_image() - if img.mode != "RGB": - img = img.convert("RGB") - images.append(transform(img)) - - batch_tensor = torch.stack(images).to(device) - - with torch.no_grad(), torch.amp.autocast( - device_type=device.type, enabled=device.type == "cuda" - ): - embeddings = self._model(batch_tensor) - - all_embeddings.append(embeddings.cpu().numpy()) - - return np.concatenate(all_embeddings, axis=0) - - def get_space_config(self, model_spec: ModelSpec, dim: int) -> dict[str, Any]: - """Return embedding space configuration with curvature.""" - self._load_model(model_spec) - assert self._model is not None - - return { - "provider": self.provider_id, - "model_id": model_spec.model_id, - "geometry": "hyperboloid", - "checkpoint": model_spec.checkpoint, - "dim": dim, - "curvature": self._model.curvature, - "spatial_dim": self._model.embed_dim, - } - - -# Auto-register on import -register_provider("hycoclip", HyCoCLIPProvider) diff --git a/src/hyperview/embeddings/providers/hycoclip_onnx.py b/src/hyperview/embeddings/providers/hycoclip_onnx.py deleted file mode 100644 index 6d01dbef54c29ecfb03319342f08f6f4bfeae58f..0000000000000000000000000000000000000000 --- a/src/hyperview/embeddings/providers/hycoclip_onnx.py +++ /dev/null @@ -1,210 +0,0 @@ -"""HyCoCLIP ONNX embedding provider - torch-free runtime. - -This provider runs an ONNX-exported HyCoCLIP/MERU image encoder with -`onnxruntime` to compute *hyperboloid (Lorentz)* embeddings. - -Outputs: -- Embeddings are returned in hyperboloid format with shape (N, D+1), where the - first coordinate is the time component. - -Requirements: -- onnxruntime (already included via embed-anything) -- An exported ONNX model (and its external weights .data file) produced by - `hyperbolic_model_zoo/hycoclip_onnx/export_onnx.py`. - -Why this exists: -- Torch is required to *export* HyCoCLIP to ONNX. -- Torch is NOT required at runtime once you have the ONNX artifacts. -""" - -from __future__ import annotations - -from pathlib import Path -from typing import Any - -import numpy as np - -from hyperview.core.sample import Sample -from hyperview.embeddings.providers import ( - BaseEmbeddingProvider, - ModelSpec, - register_provider, -) - -__all__ = ["HyCoCLIPOnnxProvider"] - - -def _preprocess_rgb_image_to_chw_float01(img: Any, size: int = 224) -> np.ndarray: - """Resize shortest side to `size`, center crop, return (3,H,W) float32 in [0,1].""" - from PIL import Image - - if not isinstance(img, Image.Image): - raise TypeError(f"Expected PIL.Image.Image, got {type(img)}") - - w, h = img.size - if w <= 0 or h <= 0: - raise ValueError(f"Invalid image size: {w}x{h}") - - # Resize shortest side to `size`. - scale = float(size) / float(min(w, h)) - new_w = int(round(w * scale)) - new_h = int(round(h * scale)) - img = img.resize((new_w, new_h), resample=Image.Resampling.BICUBIC) - - # Center crop. - left = int(round((new_w - size) / 2.0)) - top = int(round((new_h - size) / 2.0)) - img = img.crop((left, top, left + size, top + size)) - - arr = np.asarray(img, dtype=np.float32) / 255.0 - arr = np.transpose(arr, (2, 0, 1)) - return arr - - -class HyCoCLIPOnnxProvider(BaseEmbeddingProvider): - """ONNX HyCoCLIP provider - torch-free runtime. - - Uses onnxruntime for inference. No PyTorch required at runtime. - """ - - def __init__(self) -> None: - self._session: Any = None - self._model_spec: ModelSpec | None = None - self._input_name: str | None = None - self._output_names: list[str] | None = None - self._curvature: float | None = None - - @property - def provider_id(self) -> str: - return "hycoclip_onnx" - - def _resolve_onnx_path(self, model_spec: ModelSpec) -> Path: - if not model_spec.checkpoint: - raise ValueError( - "HyCoCLIP ONNX provider requires 'checkpoint' to be a path/URL to a .onnx file." - ) - - checkpoint = model_spec.checkpoint - - # Handle HuggingFace Hub URLs (hf://repo_id/path/to/model.onnx) - if checkpoint.startswith("hf://"): - from huggingface_hub import hf_hub_download - - # Parse: hf://mnm-matin/hyperbolic-clip/hycoclip-vit-s/model.onnx - path_part = checkpoint[5:] # Remove "hf://" - parts = path_part.split("/", 2) - if len(parts) < 3: - raise ValueError( - f"Invalid hf:// URL format: {checkpoint}. " - "Expected: hf://owner/repo/path/to/file.onnx" - ) - repo_id = f"{parts[0]}/{parts[1]}" - filename = parts[2] - - downloaded_path = hf_hub_download(repo_id=repo_id, filename=filename) - return Path(downloaded_path) - - path = Path(checkpoint).expanduser() - if not path.exists(): - raise FileNotFoundError(f"ONNX model not found: {path}") - checkpoint_suffix = Path(checkpoint).suffix.lower() - if path.suffix.lower() != ".onnx" and checkpoint_suffix != ".onnx": - raise ValueError(f"Expected a .onnx file, got: {path}") - return path - - def _ensure_session(self, model_spec: ModelSpec) -> None: - if self._session is not None and self._model_spec == model_spec: - return - - import onnxruntime as ort - - onnx_path = self._resolve_onnx_path(model_spec) - - # Default to CPU for maximum compatibility. - available = ort.get_available_providers() - providers = ( - ["CPUExecutionProvider"] if "CPUExecutionProvider" in available else list(available) - ) - - self._session = ort.InferenceSession(str(onnx_path), providers=providers) - self._input_name = self._session.get_inputs()[0].name - self._output_names = [o.name for o in self._session.get_outputs()] - self._curvature = None - self._model_spec = model_spec - - def compute_embeddings( - self, - samples: list[Sample], - model_spec: ModelSpec, - batch_size: int = 32, - show_progress: bool = True, - ) -> np.ndarray: - """Compute hyperboloid embeddings (t, x) with shape (N, D+1).""" - self._ensure_session(model_spec) - - assert self._session is not None - assert self._input_name is not None - - output_names = self._output_names or [] - if not output_names: - raise RuntimeError("ONNX session has no outputs") - - # Prefer named outputs if present. - emb_name = ( - "embedding_hyperboloid" if "embedding_hyperboloid" in output_names else output_names[0] - ) - curv_name = "curvature" if "curvature" in output_names else None - - if batch_size != 1 and show_progress: - print("HyCoCLIP-ONNX export currently runs with batch_size=1; overriding") - batch_size = 1 - - all_embeddings: list[np.ndarray] = [] - - if show_progress: - print(f"Computing HyCoCLIP-ONNX embeddings for {len(samples)} samples...") - - for i in range(0, len(samples), batch_size): - batch_samples = samples[i : i + batch_size] - - images = [] - for sample in batch_samples: - with sample.load_image() as img: - img.load() - if img.mode != "RGB": - img = img.convert("RGB") - chw = _preprocess_rgb_image_to_chw_float01(img.copy(), size=224) - images.append(chw) - - batch_np = np.stack(images, axis=0).astype(np.float32) - - outputs = self._session.run( - [name for name in (emb_name, curv_name) if name is not None], - {self._input_name: batch_np}, - ) - - hyper = np.asarray(outputs[0], dtype=np.float32) - if hyper.ndim != 2: - raise RuntimeError(f"Expected (B,D) embeddings, got shape={hyper.shape}") - - # Capture curvature once. - if curv_name is not None and self._curvature is None and len(outputs) > 1: - curv_arr = np.asarray(outputs[1]).reshape(-1) - if curv_arr.size > 0: - self._curvature = float(curv_arr[0]) - - all_embeddings.append(hyper) - - return np.vstack(all_embeddings) - - def get_space_config(self, model_spec: ModelSpec, dim: int) -> dict[str, Any]: - config = super().get_space_config(model_spec, dim) - config["geometry"] = "hyperboloid" - if self._curvature is not None: - config["curvature"] = self._curvature - config["spatial_dim"] = dim - 1 - return config - - -# Auto-register on import -register_provider("hycoclip_onnx", HyCoCLIPOnnxProvider) diff --git a/src/hyperview/server/__init__.py b/src/hyperview/server/__init__.py deleted file mode 100644 index a2db27419e7f6b9e2bfc32b76bb0aab94b1380e0..0000000000000000000000000000000000000000 --- a/src/hyperview/server/__init__.py +++ /dev/null @@ -1,5 +0,0 @@ -"""FastAPI server for HyperView.""" - -from hyperview.server.app import create_app - -__all__ = ["create_app"] diff --git a/src/hyperview/server/app.py b/src/hyperview/server/app.py deleted file mode 100644 index 26e897a62082082a8ba4e78b173f67eb15d0ab7c..0000000000000000000000000000000000000000 --- a/src/hyperview/server/app.py +++ /dev/null @@ -1,399 +0,0 @@ -"""FastAPI application for HyperView.""" - -import os -from pathlib import Path -from typing import Any - -from fastapi import Depends, FastAPI, HTTPException, Query -from fastapi.middleware.cors import CORSMiddleware -from fastapi.responses import JSONResponse -from fastapi.staticfiles import StaticFiles -from pydantic import BaseModel - -import numpy as np - -from hyperview.core.dataset import Dataset -from hyperview.core.selection import points_in_polygon - -# Global dataset reference (set by launch()) -_current_dataset: Dataset | None = None -_current_session_id: str | None = None - - -class SelectionRequest(BaseModel): - """Request model for selection sync.""" - - sample_ids: list[str] - - -class LassoSelectionRequest(BaseModel): - """Request model for lasso selection queries.""" - - layout_key: str # e.g., "openai_clip-vit-base-patch32__umap" - # Polygon vertices in data space, interleaved: [x0, y0, x1, y1, ...] - polygon: list[float] - offset: int = 0 - limit: int = 100 - include_thumbnails: bool = True - - -class SampleResponse(BaseModel): - """Response model for a sample.""" - - id: str - filepath: str - filename: str - label: str | None - thumbnail: str | None - metadata: dict - width: int | None = None - height: int | None = None - - -class LayoutInfoResponse(BaseModel): - """Response model for layout info.""" - - layout_key: str - space_key: str - method: str - geometry: str - count: int - params: dict[str, Any] | None - - -class SpaceInfoResponse(BaseModel): - """Response model for embedding space info.""" - - space_key: str - model_id: str - dim: int - count: int - provider: str - geometry: str - config: dict[str, Any] | None - - -class DatasetResponse(BaseModel): - """Response model for dataset info.""" - - name: str - num_samples: int - labels: list[str] - label_colors: dict[str, str] - spaces: list[SpaceInfoResponse] - layouts: list[LayoutInfoResponse] - - -class EmbeddingsResponse(BaseModel): - """Response model for embeddings data (for scatter plot).""" - - layout_key: str - geometry: str - ids: list[str] - labels: list[str | None] - coords: list[list[float]] - label_colors: dict[str, str] - - -class SimilarSampleResponse(BaseModel): - """Response model for a similar sample with distance.""" - - id: str - filepath: str - filename: str - label: str | None - thumbnail: str | None - distance: float - metadata: dict - - -class SimilaritySearchResponse(BaseModel): - """Response model for similarity search results.""" - - query_id: str - k: int - results: list[SimilarSampleResponse] - - -def create_app(dataset: Dataset | None = None, session_id: str | None = None) -> FastAPI: - """Create the FastAPI application. - - Args: - dataset: Optional dataset to serve. If None, uses global dataset. - - Returns: - FastAPI application instance. - """ - global _current_dataset, _current_session_id - if dataset is not None: - _current_dataset = dataset - if session_id is not None: - _current_session_id = session_id - - app = FastAPI( - title="HyperView", - description="Dataset visualization with hyperbolic embeddings", - version="0.1.0", - ) - - def get_dataset() -> Dataset: - """Dependency that returns the current dataset or raises 404.""" - if _current_dataset is None: - raise HTTPException(status_code=404, detail="No dataset loaded") - return _current_dataset - - # CORS middleware for development - app.add_middleware( - CORSMiddleware, - allow_origins=["*"], - allow_credentials=True, - allow_methods=["*"], - allow_headers=["*"], - ) - - @app.get("/__hyperview__/health") - async def hyperview_health(): - return { - "name": "hyperview", - "version": app.version, - "session_id": _current_session_id, - "dataset": _current_dataset.name if _current_dataset is not None else None, - "pid": os.getpid(), - } - - @app.get("/api/dataset", response_model=DatasetResponse) - async def get_dataset_info(ds: Dataset = Depends(get_dataset)): - """Get dataset metadata.""" - spaces = ds.list_spaces() - space_dicts = [s.to_api_dict() for s in spaces] - - layouts = ds.list_layouts() - layout_dicts = [l.to_api_dict() for l in layouts] - - return DatasetResponse( - name=ds.name, - num_samples=len(ds), - labels=ds.labels, - label_colors=ds.get_label_colors(), - spaces=space_dicts, - layouts=layout_dicts, - ) - - @app.get("/api/samples") - async def get_samples( - ds: Dataset = Depends(get_dataset), - offset: int = Query(0, ge=0), - limit: int = Query(100, ge=1, le=1000), - label: str | None = None, - ): - """Get paginated samples with thumbnails.""" - samples, total = ds.get_samples_paginated( - offset=offset, limit=limit, label=label - ) - - return { - "total": total, - "offset": offset, - "limit": limit, - "samples": [s.to_api_dict(include_thumbnail=True) for s in samples], - } - - @app.get("/api/samples/{sample_id}", response_model=SampleResponse) - async def get_sample(sample_id: str, ds: Dataset = Depends(get_dataset)): - """Get a single sample by ID.""" - try: - sample = ds[sample_id] - return SampleResponse(**sample.to_api_dict()) - except KeyError: - raise HTTPException(status_code=404, detail=f"Sample not found: {sample_id}") - - @app.post("/api/samples/batch") - async def get_samples_batch(request: SelectionRequest, ds: Dataset = Depends(get_dataset)): - """Get multiple samples by their IDs.""" - samples = ds.get_samples_by_ids(request.sample_ids) - return {"samples": [s.to_api_dict(include_thumbnail=True) for s in samples]} - - @app.get("/api/embeddings", response_model=EmbeddingsResponse) - async def get_embeddings(ds: Dataset = Depends(get_dataset), layout_key: str | None = None): - """Get embedding coordinates for visualization.""" - layouts = ds.list_layouts() - if not layouts: - raise HTTPException( - status_code=400, detail="No layouts computed. Call compute_visualization() first." - ) - - # Find the requested layout - layout_info = None - if layout_key is None: - layout_info = layouts[0] - layout_key = layout_info.layout_key - else: - layout_info = next((l for l in layouts if l.layout_key == layout_key), None) - if layout_info is None: - raise HTTPException(status_code=404, detail=f"Layout not found: {layout_key}") - - ids, labels, coords = ds.get_visualization_data(layout_key) - - if not ids: - raise HTTPException(status_code=400, detail=f"No data in layout '{layout_key}'.") - - return EmbeddingsResponse( - layout_key=layout_key, - geometry=layout_info.geometry, - ids=ids, - labels=labels, - coords=coords.tolist(), - label_colors=ds.get_label_colors(), - ) - - @app.get("/api/spaces") - async def get_spaces(ds: Dataset = Depends(get_dataset)): - """Get all embedding spaces.""" - spaces = ds.list_spaces() - return {"spaces": [s.to_api_dict() for s in spaces]} - - @app.get("/api/layouts") - async def get_layouts(ds: Dataset = Depends(get_dataset)): - """Get all available layouts.""" - layouts = ds.list_layouts() - return {"layouts": [l.to_api_dict() for l in layouts]} - - @app.post("/api/selection") - async def sync_selection(request: SelectionRequest): - """Sync selection state (for future use).""" - return {"status": "ok", "selected": request.sample_ids} - - @app.post("/api/selection/lasso") - async def lasso_selection(request: LassoSelectionRequest, ds: Dataset = Depends(get_dataset)): - """Compute a lasso selection over the current embeddings. - - Returns a total selected count and a paginated page of selected samples. - - Notes: - - Selection is performed in *data space* (the same coordinates returned - by /api/embeddings). - - For now we use an in-memory scan with a tight AABB prefilter. - """ - if request.offset < 0: - raise HTTPException(status_code=400, detail="offset must be >= 0") - if request.limit < 1 or request.limit > 2000: - raise HTTPException(status_code=400, detail="limit must be between 1 and 2000") - - if len(request.polygon) < 6 or len(request.polygon) % 2 != 0: - raise HTTPException( - status_code=400, - detail="polygon must be an even-length list with at least 3 vertices", - ) - - poly = np.asarray(request.polygon, dtype=np.float32).reshape((-1, 2)) - if not np.all(np.isfinite(poly)): - raise HTTPException(status_code=400, detail="polygon must contain only finite numbers") - - # Tight AABB prefilter. - x_min = float(np.min(poly[:, 0])) - x_max = float(np.max(poly[:, 0])) - y_min = float(np.min(poly[:, 1])) - y_max = float(np.max(poly[:, 1])) - - candidate_ids, candidate_coords = ds.get_lasso_candidates_aabb( - layout_key=request.layout_key, - x_min=x_min, - x_max=x_max, - y_min=y_min, - y_max=y_max, - ) - - if candidate_coords.size == 0: - return {"total": 0, "offset": request.offset, "limit": request.limit, "sample_ids": [], "samples": []} - - inside_mask = points_in_polygon(candidate_coords, poly) - if not np.any(inside_mask): - return {"total": 0, "offset": request.offset, "limit": request.limit, "sample_ids": [], "samples": []} - - selected_ids = [candidate_ids[i] for i in np.flatnonzero(inside_mask)] - total = len(selected_ids) - - start = int(request.offset) - end = int(request.offset + request.limit) - sample_ids = selected_ids[start:end] - - samples = ds.get_samples_by_ids(sample_ids) - sample_dicts = [s.to_api_dict(include_thumbnail=request.include_thumbnails) for s in samples] - - return { - "total": total, - "offset": request.offset, - "limit": request.limit, - "sample_ids": sample_ids, - "samples": sample_dicts, - } - - @app.get("/api/search/similar/{sample_id}", response_model=SimilaritySearchResponse) - async def search_similar( - sample_id: str, - ds: Dataset = Depends(get_dataset), - k: int = Query(10, ge=1, le=100), - space_key: str | None = None, - ): - """Return k nearest neighbors for a given sample.""" - try: - similar = ds.find_similar( - sample_id, k=k, space_key=space_key - ) - except ValueError as e: - raise HTTPException(status_code=400, detail=str(e)) - except KeyError: - raise HTTPException(status_code=404, detail=f"Sample not found: {sample_id}") - - results = [] - for sample, distance in similar: - try: - thumbnail = sample.get_thumbnail_base64() - except Exception: - thumbnail = None - - results.append( - SimilarSampleResponse( - id=sample.id, - filepath=sample.filepath, - filename=sample.filename, - label=sample.label, - thumbnail=thumbnail, - distance=distance, - metadata=sample.metadata, - ) - ) - - return SimilaritySearchResponse( - query_id=sample_id, - k=k, - results=results, - ) - - @app.get("/api/thumbnail/{sample_id}") - async def get_thumbnail(sample_id: str, ds: Dataset = Depends(get_dataset)): - """Get thumbnail image for a sample.""" - try: - sample = ds[sample_id] - thumbnail_b64 = sample.get_thumbnail_base64() - return JSONResponse({"thumbnail": thumbnail_b64}) - except KeyError: - raise HTTPException(status_code=404, detail=f"Sample not found: {sample_id}") - - # Serve static frontend files - static_dir = Path(__file__).parent / "static" - if static_dir.exists(): - app.mount("/", StaticFiles(directory=str(static_dir), html=True), name="static") - else: - # Fallback: serve a simple HTML page - @app.get("/") - async def root(): - return {"message": "HyperView API", "docs": "/docs"} - - return app - - -def set_dataset(dataset: Dataset) -> None: - """Set the global dataset for the server.""" - global _current_dataset - _current_dataset = dataset diff --git a/src/hyperview/server/static/404.html b/src/hyperview/server/static/404.html deleted file mode 100644 index 7d84c4644e74ddb20f4cbd03dfb1c8ba576a6fa8..0000000000000000000000000000000000000000 --- a/src/hyperview/server/static/404.html +++ /dev/null @@ -1 +0,0 @@ -