File size: 2,485 Bytes
7183b27 582038e 7183b27 d5505d6 7183b27 19dd68e 7183b27 d5505d6 7183b27 582038e 7183b27 d5505d6 7183b27 582038e 7183b27 22e905d 20b8a25 7183b27 582038e 7183b27 18ae1b8 7183b27 72f160c 7183b27 147b396 58ed971 9c7467e e831d36 5ac51ca 7183b27 |
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 |
---
license: gpl-3.0
datasets:
- Kaynaaf/Brain-Tumour-MRI
metrics:
- accuracy 0.90
- precision 0.90
library_name: keras
tags:
- medical
- healthcare
---
# Model Card
An Image Classifier that predicts the presence of certain Brain tumours from their MRI scans
## Model Details
A 134M Parameter ConvNet designed for classification of Brain tumours in MRI scans.
## Paper
Interpretable Deep Learning for Brain Tumor Diagnosis: Occlusion Sensitivity-Driven Explainability in MRI Classification
DOI: [10.21015/vtse.v13i2.2082](10.21015/vtse.v13i2.2082)
## Uses
### Direct Use
Load the model, finetune the model if needed or just go straight towards generating inferences using the model.
### Downstream Use
Finetune the model on other diagnostic scans, though the model only accepts grayscale images of size 256x256.
## How to Get Started with the Model
[](https://colab.research.google.com/drive/1SfK9d2In3JHDvyXH4jpwznVGEG_wXRuQ?usp=sharing)
## Training
The colab notebook used to train the model can be found below
[](https://colab.research.google.com/drive/1SfK9d2In3JHDvyXH4jpwznVGEG_wXRuQ?usp=sharing)
## Evaluation
### Metrics
| Class | Precision | Recall | F1-Score | Support |
|-------------|-----------|--------|----------|---------|
| Glioma | 0.96 | 0.87 | 0.91 | 300 |
| Meningioma | 0.84 | 0.71 | 0.77 | 306 |
| No Tumor | 0.88 | 1.00 | 0.93 | 405 |
| Pituitary | 0.93 | 0.99 | 0.96 | 300 |
| **Accuracy**| | | **0.90** | 1311 |
| **Macro Avg** | 0.90 | 0.89 | 0.89 | 1311 |
| **Weighted Avg** | 0.90 | 0.90 | 0.90 | 1311 |
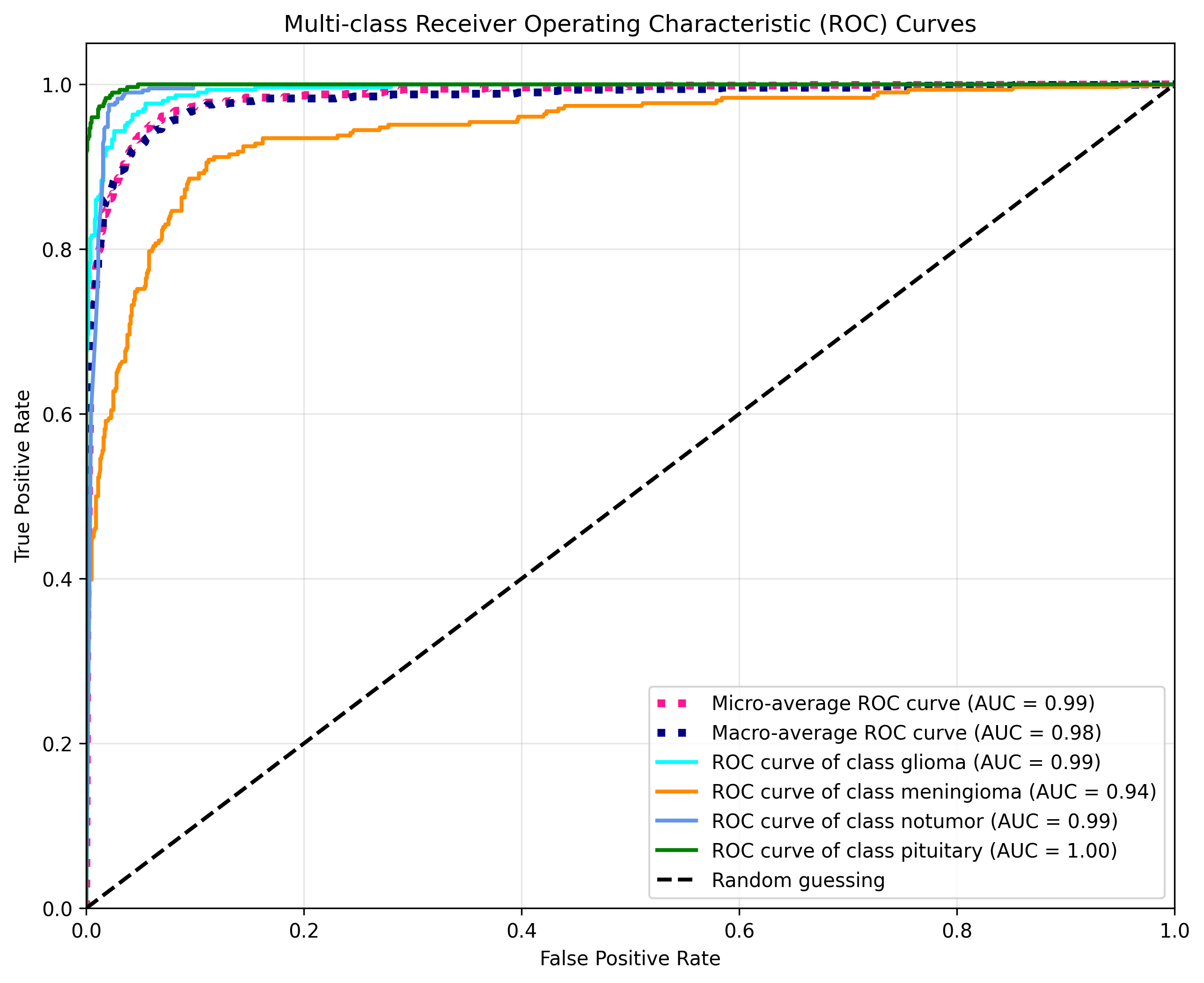
### Results
This model was developed for my project that can be found on github [here](https://github.com/Kaynaaf/BrainMRI-Classifier)
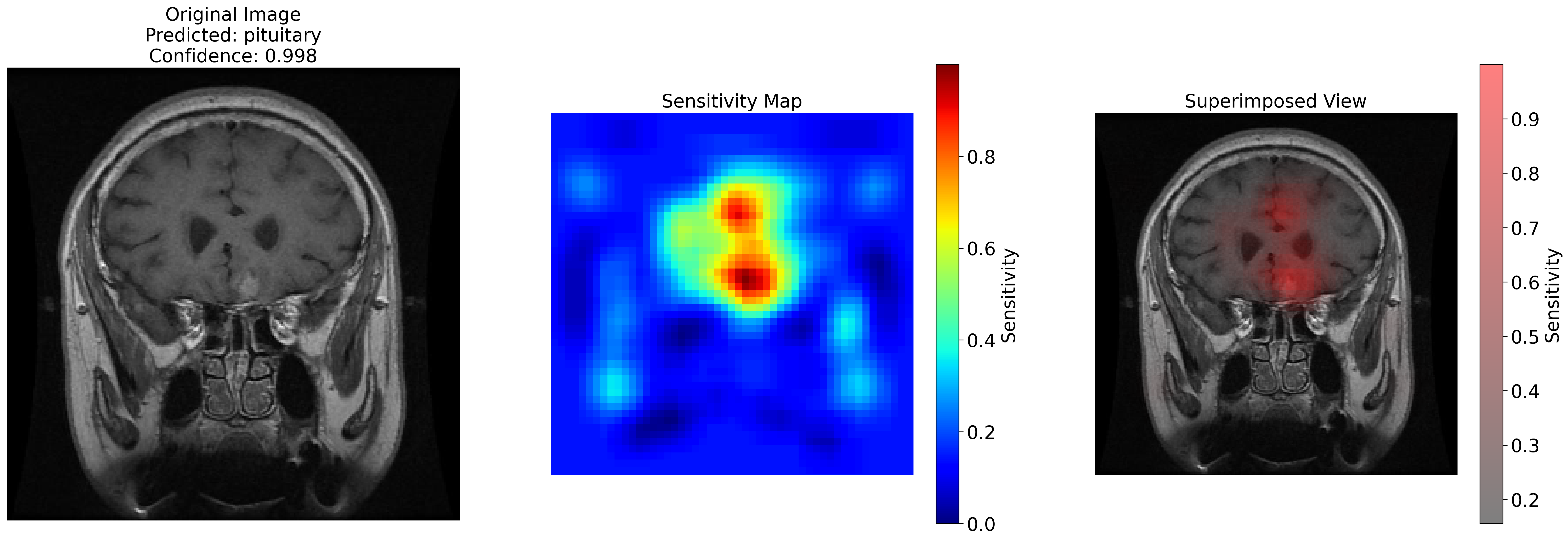
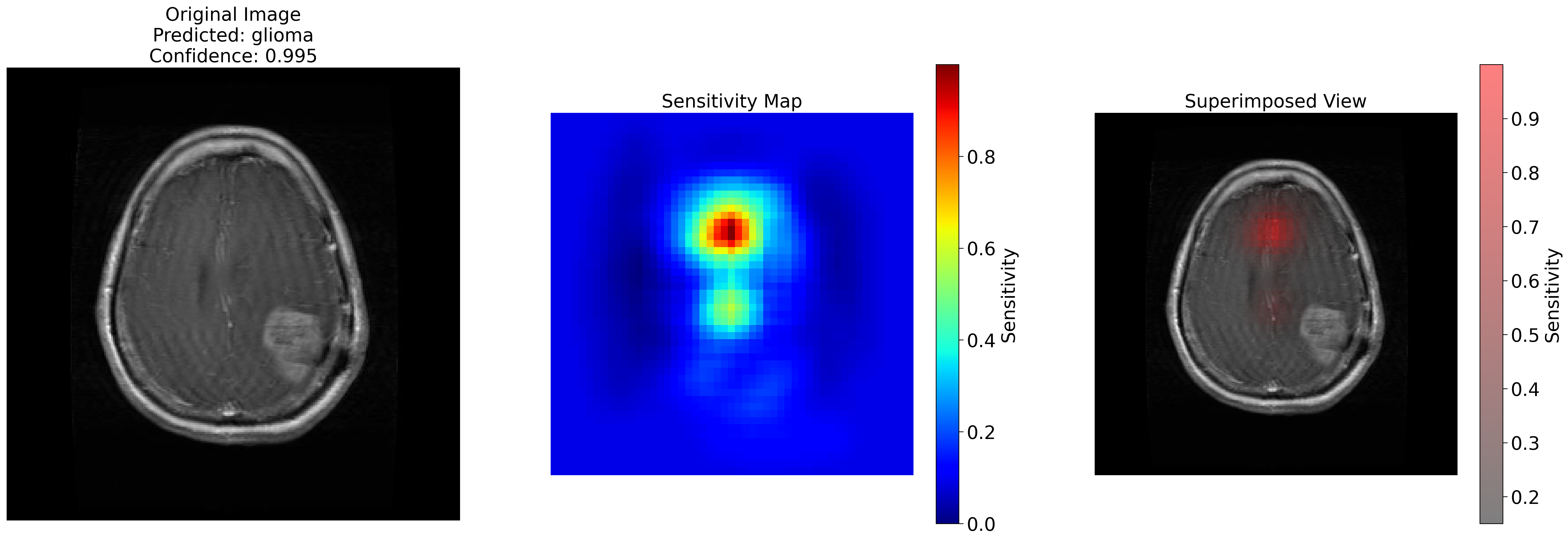
. This project involved generating sensitivity maps to explain the predictions of the model.
These maps assign values to areas of the image that act as feature importance markers.
|