|
|
--- |
|
|
license: apache-2.0 |
|
|
pipeline_tag: image-segmentation |
|
|
tags: |
|
|
- medical |
|
|
--- |
|
|
# U-Net Transplant: Model Merging for 3D Medical Segmentation |
|
|
 |
|
|
|
|
|
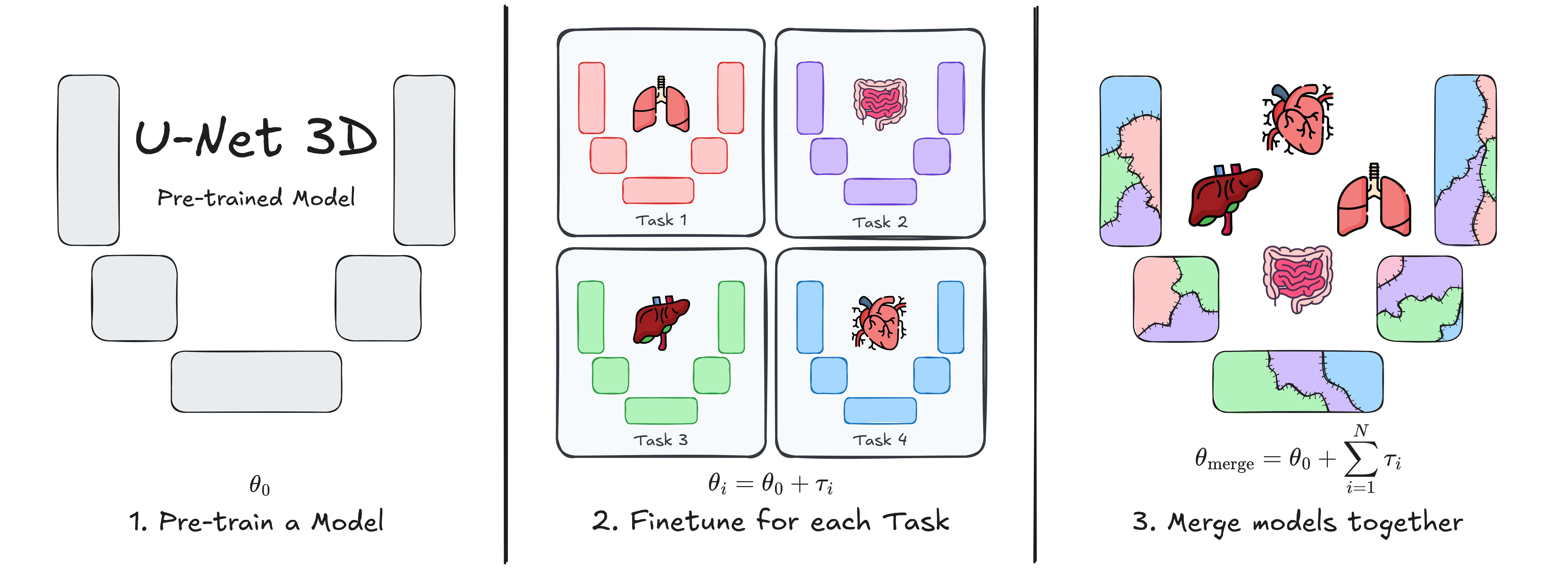
This repository contains the implementation of **U-Net Transplant**, a framework for efficient model merging in 3D medical image segmentation. Model merging enables the combination of specialized segmentation models without requiring full retraining, offering a flexible and privacy-conscious solution for updating AI models in clinical applications. |
|
|
|
|
|
Our approach leverages **task vectors** and encourages **wide minima** during pre-training to enhance the effectiveness of model merging. We evaluate this method using the **ToothFairy2** and **BTCV Abdomen** datasets with a standard **3D U-Net** architecture, demonstrating its ability to integrate multiple specialized segmentation tasks into a single model. |
|
|
|
|
|
|
|
|
# Pretrain and Task Vector Checkpoints |
|
|
The related checkpoints and task vectors used in the paper will be available from the 23rd June 2025. |
|
|
|
|
|
|
|
|
# How to Run |
|
|
|
|
|
### 1. Clone the Repository |
|
|
```bash |
|
|
git clone git@github.com:LucaLumetti/UNetTransplant.git |
|
|
cd UNetTransplant |
|
|
``` |
|
|
|
|
|
### 2. Setup Environment |
|
|
```bash |
|
|
python -m venv env |
|
|
source env/bin/activate |
|
|
pip install -r requirements.txt |
|
|
``` |
|
|
|
|
|
### 3. Downloads |
|
|
Ensure the datasets are downloaded and organized following the nnUNet dataset format. |
|
|
|
|
|
- **BTCV Abdomen**: [Download Here](https://www.synapse.org/Synapse:syn3193805/wiki/217753) |
|
|
- **ToothFairy2**: [Download Here](https://ditto.ing.unimore.it/toothfairy2/) |
|
|
- **AMOS**: [Download Here](https://zenodo.org/records/7262581) |
|
|
- **ZhimingCui**: Available upon request from the authors ([Paper](https://www.nature.com/articles/s41467-022-29637-2)) |
|
|
|
|
|
You can also download pretrained checkpoints and task vectors: |
|
|
```bash |
|
|
#!/bin/bash |
|
|
|
|
|
BASE_ABDOMEN="https://huggingface.co/Lumett/UNetTransplant/resolve/main/Abdomen" |
|
|
BASE_TOOTHFAIRY="https://huggingface.co/Lumett/UNetTransplant/resolve/main/ToothFairy" |
|
|
|
|
|
abdomen_files=( |
|
|
Pretrain_AMOS.pth |
|
|
TaskVector_Kidney_Abdomen.pth |
|
|
TaskVector_Liver_Abdomen.pth |
|
|
TaskVector_Spleen_Abdomen.pth |
|
|
TaskVector_Stomach_Abdomen.pth |
|
|
) |
|
|
|
|
|
toothfairy_files=( |
|
|
Pretrain_Cui.pth |
|
|
TaskVector_Canals_ToothFairy2.pth |
|
|
TaskVector_Mandible_ToothFairy2.pth |
|
|
TaskVector_Teeth_ToothFairy2.pth |
|
|
TaskVector_Pharynx_ToothFairy2.pth |
|
|
) |
|
|
|
|
|
echo "🩻 Downloading Abdomen files..." |
|
|
for file in "${abdomen_files[@]}"; do |
|
|
wget -c "${BASE_ABDOMEN}/${file}" |
|
|
done |
|
|
|
|
|
echo "🦷 Downloading ToothFairy files..." |
|
|
for file in "${toothfairy_files[@]}"; do |
|
|
wget -c "${BASE_TOOTHFAIRY}/${file}" |
|
|
done |
|
|
``` |
|
|
|
|
|
### 4. Running the U-Net Transplant Framework |
|
|
|
|
|
The main script for running experiments is `main.py`. It requires specifying the type of experiment and a configuration file that defines dataset, model, optimizer, and training parameters. |
|
|
|
|
|
#### Command Structure |
|
|
```bash |
|
|
python main.py --experiment <EXPERIMENT_TYPE> --config <CONFIG_PATH> [--expname <NAME>] [--override <PARAMS>] |
|
|
``` |
|
|
|
|
|
#### Arguments |
|
|
- **`--experiment`**: Specifies the type of experiment to run. |
|
|
- `"PretrainExperiment"` → Pretrains the model from scratch. |
|
|
- `"TaskVectorTrainExperiment"` → Trains a task vector using a pretrained checkpoint. |
|
|
|
|
|
- **`--config`**: Path to the configuration file, which defines dataset, model, and training settings. |
|
|
|
|
|
- **`--expname`** (optional): Custom experiment name. If not provided, the config filename is used. |
|
|
|
|
|
- **`--override`** (optional): Allows overriding config values at runtime. Example: |
|
|
```bash |
|
|
python main.py --experiment PretrainExperiment --config configs/default.yaml --override DataConfig.BATCH_SIZE=4 OptimizerConfig.LR=0.01 |
|
|
``` |
|
|
|
|
|
#### Configuration File |
|
|
The configuration file defines: |
|
|
- **Dataset** (`DataConfig`): Path, batch size, patch size, and datasets used. |
|
|
- **Model** (`BackboneConfig` & `HeadsConfig`): Architecture, checkpoints, and initialization. |
|
|
- **Optimizer** (`OptimizerConfig`): Learning rates, weight decay, and momentum. |
|
|
- **Loss Function** (`LossConfig`): Defines the loss function used. |
|
|
- **Training** (`TrainConfig`): Number of epochs, checkpoint saving, and resume options. |
|
|
|
|
|
Check [the provided configs](https://github.com/LucaLumetti/UNetTransplant/tree/main/configs/miccai2025) for examples. |
|
|
|
|
|
#### Example Commands |
|
|
1. **Pretraining a model**: |
|
|
```bash |
|
|
python main.py --experiment PretrainExperiment --config configs/miccai2025/pretrain_stable.yaml |
|
|
``` |
|
|
2. **Training a task vector from a checkpoint**: |
|
|
```bash |
|
|
python main.py --experiment TaskVectorTrainExperiment --config configs/miccai2025/finetune.yaml --override BackboneConfig.PRETRAIN_CHECKPOINTS="/path/to/checkpoint.pth" |
|
|
``` |
|
|
|
|
|
For further details, refer to the config files used in our experiments under the `configs` folder. |
|
|
|
|
|
### 5. Cite |
|
|
If you used our work, please cite it: |
|
|
``` |
|
|
@incollection{lumetti2025u, |
|
|
title={U-Net Transplant: The Role of Pre-training for Model Merging in 3D Medical Segmentation}, |
|
|
author={Lumetti, Luca and Capitani, Giacomo and Ficarra, Elisa and Grana, Costantino and Calderara, Simone and Porrello, Angelo and Bolelli, Federico and others}, |
|
|
booktitle={Medical Image Computing and Computer Assisted Intervention--MICCAI 2025}, |
|
|
year={2025} |
|
|
} |
|
|
``` |