| # Model Overview | |
| Body CT segmentation models are evolving. Starting from abdominal multi-organ segmentation model [1]. Now the community is developing hundreds of target anatomies. In this bundle, we provide re-trained models for (3D) segmentation of 104 whole-body segments. | |
| This model is trained using the SegResNet [3] network. The model is trained using TotalSegmentator datasets [2]. | |
|  | |
| Figure source from the TotalSegmentator [2]. | |
| ### MONAI Label Showcase | |
| - We highlight the use of this bundle to use and visualize in MONAI Label + 3D Slicer integration. | |
| 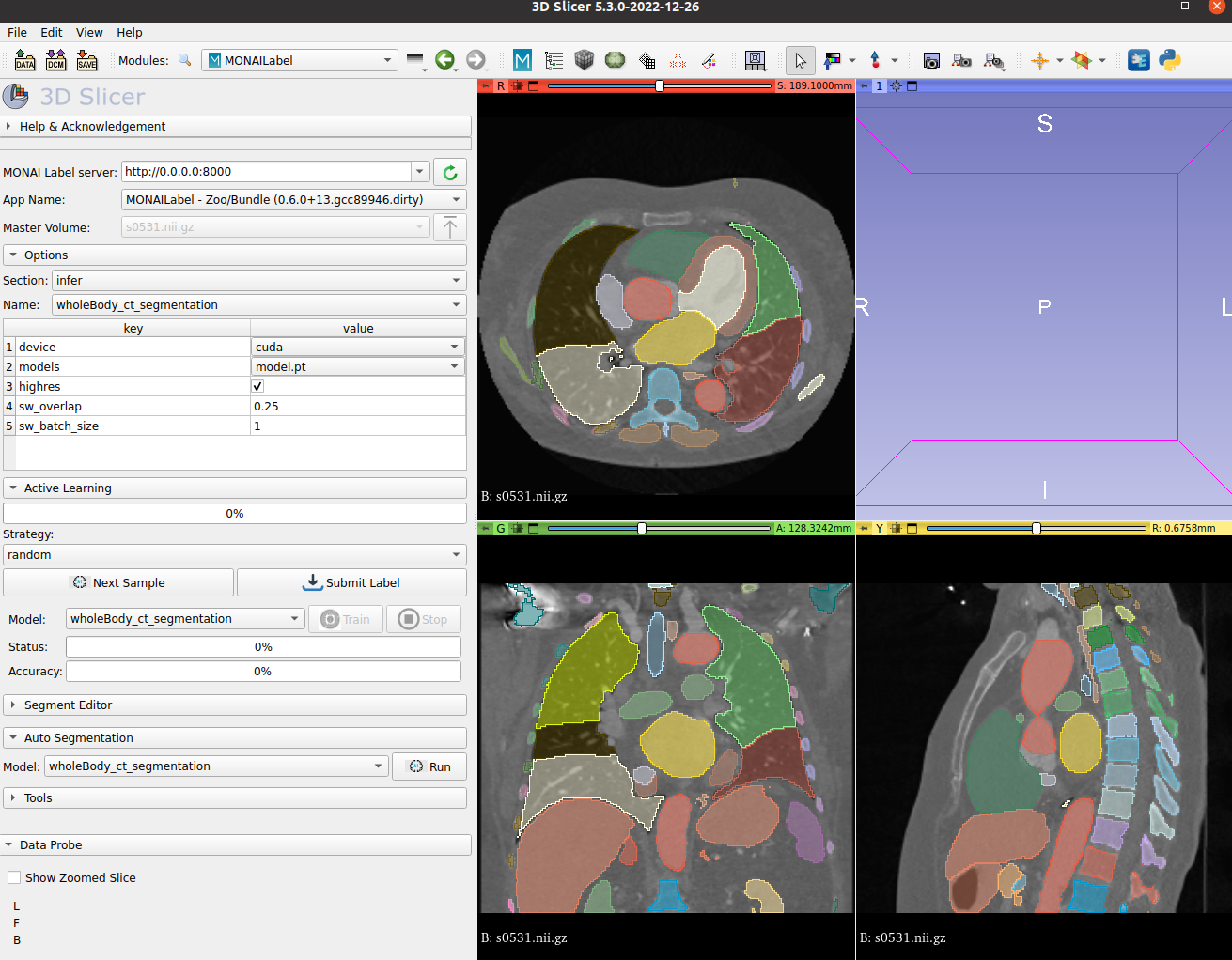 <br> | |
| ## Data | |
| The training set is the 104 whole-body structures from the TotalSegmentator released datasets. Users can find more details on the datasets at https://github.com/wasserth/TotalSegmentator. All rights and licenses are reserved to the original authors. | |
| - Target: 104 structures | |
| - Modality: CT | |
| - Source: TotalSegmentator | |
| - Challenge: Large volumes of structures in CT images | |
| ### Preprocessing | |
| To use the bundle, users need to download the data and merge all annotated labels into one NIFTI file. Each file contains 0-104 values, each value represents one anatomy class. We provide sample datasets and step-by-step instructions on how to get prepared: | |
| Instruction on how to start with the prepared sample dataset: | |
| 1. Download the sample set with this [link](https://drive.google.com/file/d/1DtDmERVMjks1HooUhggOKAuDm0YIEunG/view?usp=share_link). | |
| 2. Unzip the dataset into a workspace folder. | |
| 3. There will be three sub-folders, each with several preprocessed CT volumes: | |
| - imagesTr: 20 samples of training scans and validation scans. | |
| - labelsTr: 20 samples of pre-processed label files. | |
| - imagesTs: 5 samples of sample testing scans. | |
| 4. Usage: users can add `--dataset_dir <totalSegmentator_mergedLabel_samples>` to the bundle run command to specify the data path. | |
| Instruction on how to merge labels with the raw dataset: | |
| - There are 104 binary masks associated with each CT scan, each mask corresponds to anatomy. These pixel-level labels are class-exclusive, users can assign each anatomy a class number then merge to a single NIFTI file as the ground truth label file. The order of anatomies can be found [here](https://github.com/Project-MONAI/model-zoo/blob/dev/models/wholeBody_ct_segmentation/configs/metadata.json). | |
| ## Training Configuration | |
| The segmentation of 104 tissues is formulated as voxel-wise multi-label segmentation. The model is optimized with the gradient descent method minimizing Dice + cross-entropy loss between the predicted mask and ground truth segmentation. | |
| The training was performed with the following: | |
| - GPU: 48 GB of GPU memory | |
| - Actual Model Input: 96 x 96 x 96 | |
| - AMP: True | |
| - Optimizer: AdamW | |
| - Learning Rate: 1e-4 | |
| - Loss: DiceCELoss | |
| ## Evaluation Configuration | |
| The model predicts 105 channels output at the same time using softmax and argmax. It requires higher GPU memory when calculating | |
| metrics between predicted masked and ground truth. The consumption of hardware requirements, such as GPU memory is dependent on the input CT volume size. | |
| The recommended evaluation configuration and the metrics were acquired with the following hardware: | |
| - GPU: equal to or larger than 48 GB of GPU memory | |
| - Model: high resolution model pre-trained at a slice thickness of 1.5 mm. | |
| Note: there are two pre-trained models provided. The default is the high resolution model, evaluation pipeline at slice thickness of **1.5mm**, | |
| users can use the lower resolution model if out of memory (OOM) occurs, which the model is pre-trained with CT scans at a slice thickness of **3.0mm**. | |
| Users can also use the inference pipeline for predicted masks, we provide detailed GPU memory consumption in the following sections. | |
| ### Memory Consumption | |
| - Dataset Manager: CacheDataset | |
| - Data Size: 1000 3D Volumes | |
| - Cache Rate: 0.4 | |
| - Single GPU - System RAM Usage: 83G | |
| - Multi GPU (8 GPUs) - System RAM Usage: 666G | |
| ### Memory Consumption Warning | |
| If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate `cache_rate` in the configurations within range [0, 1] to minimize the System RAM requirements. | |
| ### Input | |
| One channel | |
| - CT image | |
| ### Output | |
| 105 channels | |
| - Label 0: Background (everything else) | |
| - label 1-105: Foreground classes (104) | |
| ## Resource Requirements and Latency Benchmarks | |
| ### GPU Consumption Warning | |
| The model is trained with 104 classes in single instance, for predicting 104 structures, the GPU consumption can be large. | |
| For inference pipeline, please refer to the following section for benchmarking results. Normally, a CT scans with 300 slices will take about 27G memory, if your CT is larger, please prepare larger GPU memory or use CPU for inference. | |
| ### High-Resolution and Low-Resolution Models | |
| We retrained two versions of the totalSegmentator models, following the original paper and implementation. | |
| To meet multiple demands according to computation resources and performance, we provide a 1.5 mm model and a 3.0 mm model, both models are trained with 104 foreground output channels. | |
| In this bundle, we configured a parameter called `highres`, users can set it to `true` when using 1.5 mm model, and set it to `false` to use the 3.0 mm model. The high-resolution model is named `model.pt` by default, the low-resolution model is named `model_lowres.pt`. | |
| In MONAI Label use case, users can set the parameter in 3D Slicer plugin to control which model to infer and train. | |
| - Pretrained Checkpoints | |
| - 1.5 mm model: [Download link](https://drive.google.com/file/d/1PHpFWboimEXmMSe2vBra6T8SaCMC2SHT/view?usp=share_link) | |
| - 3.0 mm model: [Download link](https://drive.google.com/file/d/1c3osYscnr6710ObqZZS8GkZJQlWlc7rt/view?usp=share_link) | |
| Latencies and memory performance of using the bundle with MONAI Label: | |
| Tested Image Dimension: **(512, 512, 397)**, the slice thickness is **1.5mm** in this case. After resample to **1.5** isotropic resolution, the dimension is **(287, 287, 397)** | |
| ### 1.5 mm (highres) model (Single Model with 104 foreground classes) | |
| Benchmarking on GPU: Memory: **28.73G** | |
| - `++ Latencies => Total: 6.0277; Pre: 1.6228; Inferer: 4.1153; Invert: 0.0000; Post: 0.0897; Write: 0.1995` | |
| Benchmarking on CPU: Memory: **26G** | |
| - `++ Latencies => Total: 38.3108; Pre: 1.6643; Inferer: 30.3018; Invert: 0.0000; Post: 6.1656; Write: 0.1786` | |
| ### 3.0 mm (lowres) model (single model with 104 foreground classes) | |
| GPU: Memory: **5.89G** | |
| - `++ Latencies => Total: 1.9993; Pre: 1.2363; Inferer: 0.5207; Invert: 0.0000; Post: 0.0358; Write: 0.2060` | |
| CPU: Memory: **2.3G** | |
| - `++ Latencies => Total: 6.6138; Pre: 1.3192; Inferer: 3.6746; Invert: 0.0000; Post: 1.4431; Write: 0.1760` | |
| ## Performance | |
| ### 1.5 mm Model Training | |
| #### Training Accuracy | |
| 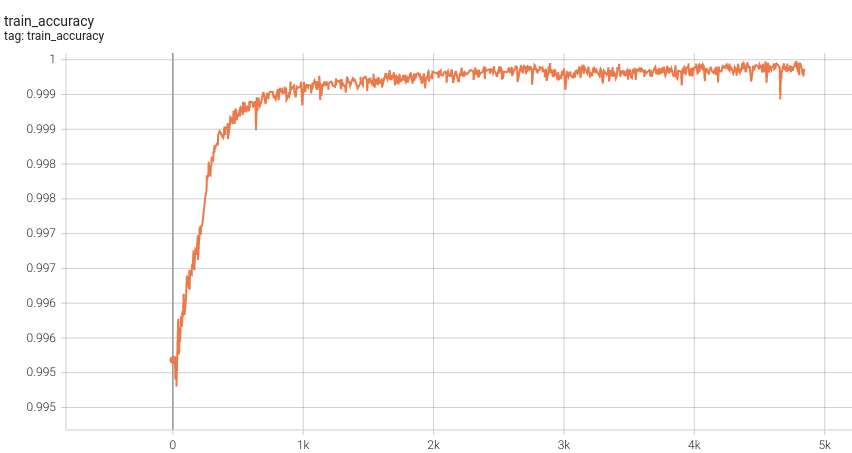 <br> | |
| #### Validation Dice | |
| 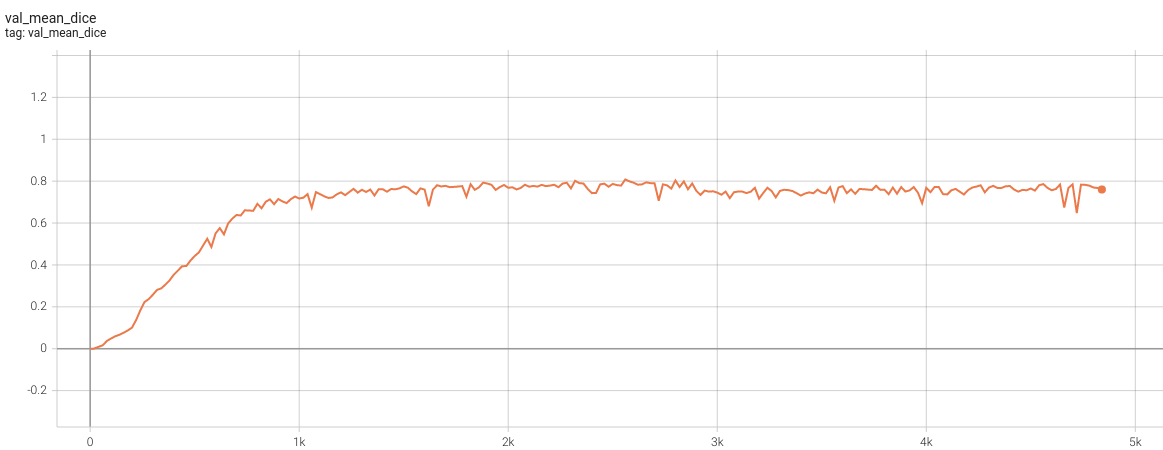 <br> | |
| Please note that this bundle is non-deterministic because of the trilinear interpolation used in the network. Therefore, reproducing the training process may not get exactly the same performance. | |
| Please refer to https://pytorch.org/docs/stable/notes/randomness.html#reproducibility for more details about reproducibility. | |
| #### TensorRT speedup | |
| This bundle supports acceleration with TensorRT. The table below displays the speedup ratios observed on an A100 80G GPU. | |
| | method | torch_fp32(ms) | torch_amp(ms) | trt_fp32(ms) | trt_fp16(ms) | speedup amp | speedup fp32 | speedup fp16 | amp vs fp16| | |
| | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | | |
| | model computation | 88.20 | 37.1 | 39.2 | 36.9 | 2.38 | 2.25 | 2.39 | 1.01 | | |
| | end2end | 3717.14 | 2596.77 | 2517.29 | 2501.37 | 1.43 | 1.48 | 1.49 | 1.04 | | |
| Where: | |
| - `model computation` means the speedup ratio of model's inference with a random input without preprocessing and postprocessing | |
| - `end2end` means run the bundle end-to-end with the TensorRT based model. | |
| - `torch_fp32` and `torch_amp` are for the PyTorch models with or without `amp` mode. | |
| - `trt_fp32` and `trt_fp16` are for the TensorRT based models converted in corresponding precision. | |
| - `speedup amp`, `speedup fp32` and `speedup fp16` are the speedup ratios of corresponding models versus the PyTorch float32 model | |
| - `amp vs fp16` is the speedup ratio between the PyTorch amp model and the TensorRT float16 based model. | |
| This result is benchmarked under: | |
| - TensorRT: 8.6.1+cuda12.0 | |
| - Torch-TensorRT Version: 1.4.0 | |
| - CPU Architecture: x86-64 | |
| - OS: ubuntu 20.04 | |
| - Python version:3.8.10 | |
| - CUDA version: 12.1 | |
| - GPU models and configuration: A100 80G | |
| ## MONAI Bundle Commands | |
| In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file. | |
| For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html). | |
| #### Execute training: | |
| ``` | |
| python -m monai.bundle run --config_file configs/train.json | |
| ``` | |
| Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`: | |
| ``` | |
| python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path> | |
| ``` | |
| #### Override the `train` config to execute multi-GPU training: | |
| ``` | |
| torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']" | |
| ``` | |
| Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html). | |
| #### Override the `train` config to execute evaluation with the trained model: | |
| ``` | |
| python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']" | |
| ``` | |
| #### Override the `train` config and `evaluate` config to execute multi-GPU evaluation: | |
| ``` | |
| torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json','configs/multi_gpu_evaluate.json']" | |
| ``` | |
| #### Execute inference: | |
| ``` | |
| python -m monai.bundle run --config_file configs/inference.json | |
| ``` | |
| #### Execute inference with Data Samples: | |
| ``` | |
| python -m monai.bundle run --config_file configs/inference.json --datalist "['sampledata/imagesTr/s0037.nii.gz','sampledata/imagesTr/s0038.nii.gz']" | |
| ``` | |
| #### Export checkpoint to TensorRT based models with fp32 or fp16 precision: | |
| ``` | |
| python -m monai.bundle trt_export --net_id network_def --filepath models/model_trt.ts --ckpt_file models/model.pt --meta_file configs/metadata.json --config_file configs/inference.json --precision <fp32/fp16> --use_trace "True" | |
| ``` | |
| #### Execute inference with the TensorRT model: | |
| ``` | |
| python -m monai.bundle run --config_file "['configs/inference.json', 'configs/inference_trt.json']" | |
| ``` | |
| # References | |
| [1] Tang, Y., Gao, R., Lee, H.H., Han, S., Chen, Y., Gao, D., Nath, V., Bermudez, C., Savona, M.R., Abramson, R.G. and Bao, S., 2021. High-resolution 3D abdominal segmentation with random patch network fusion. Medical image analysis, 69, p.101894. | |
| [2] Wasserthal, J., Meyer, M., Breit, H.C., Cyriac, J., Yang, S. and Segeroth, M., 2022. TotalSegmentator: robust segmentation of 104 anatomical structures in CT images. arXiv preprint arXiv:2208.05868. | |
| [3] Myronenko, A., Siddiquee, M.M.R., Yang, D., He, Y. and Xu, D., 2022. Automated head and neck tumor segmentation from 3D PET/CT. arXiv preprint arXiv:2209.10809. | |
| # License | |
| Copyright (c) MONAI Consortium | |
| Licensed under the Apache License, Version 2.0 (the "License"); | |
| you may not use this file except in compliance with the License. | |
| You may obtain a copy of the License at | |
| http://www.apache.org/licenses/LICENSE-2.0 | |
| Unless required by applicable law or agreed to in writing, software | |
| distributed under the License is distributed on an "AS IS" BASIS, | |
| WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. | |
| See the License for the specific language governing permissions and | |
| limitations under the License. | |