You need to agree to share your contact information to access this model
This repository is publicly accessible, but you have to accept the conditions to access its files and content.
These weights are released under CC BY-NC 4.0 — strictly non-commercial,
research and educational use only. By requesting access you agree to:
- Use the weights only for non-commercial research.
- Cite the SPARC paper in any derived publication.
- Not redistribute the weights to third parties.
Log in or Sign Up to review the conditions and access this model content.
SPARC
Gene-program-aware survival modelling from H&E whole-slide images.
This repository hosts the trained model weights for the SPARC paper (Ayed, Cohn, et al.). Code, configs, training scripts, and figure-regeneration notebooks live at github.com/aziz-ayed/SPARC.
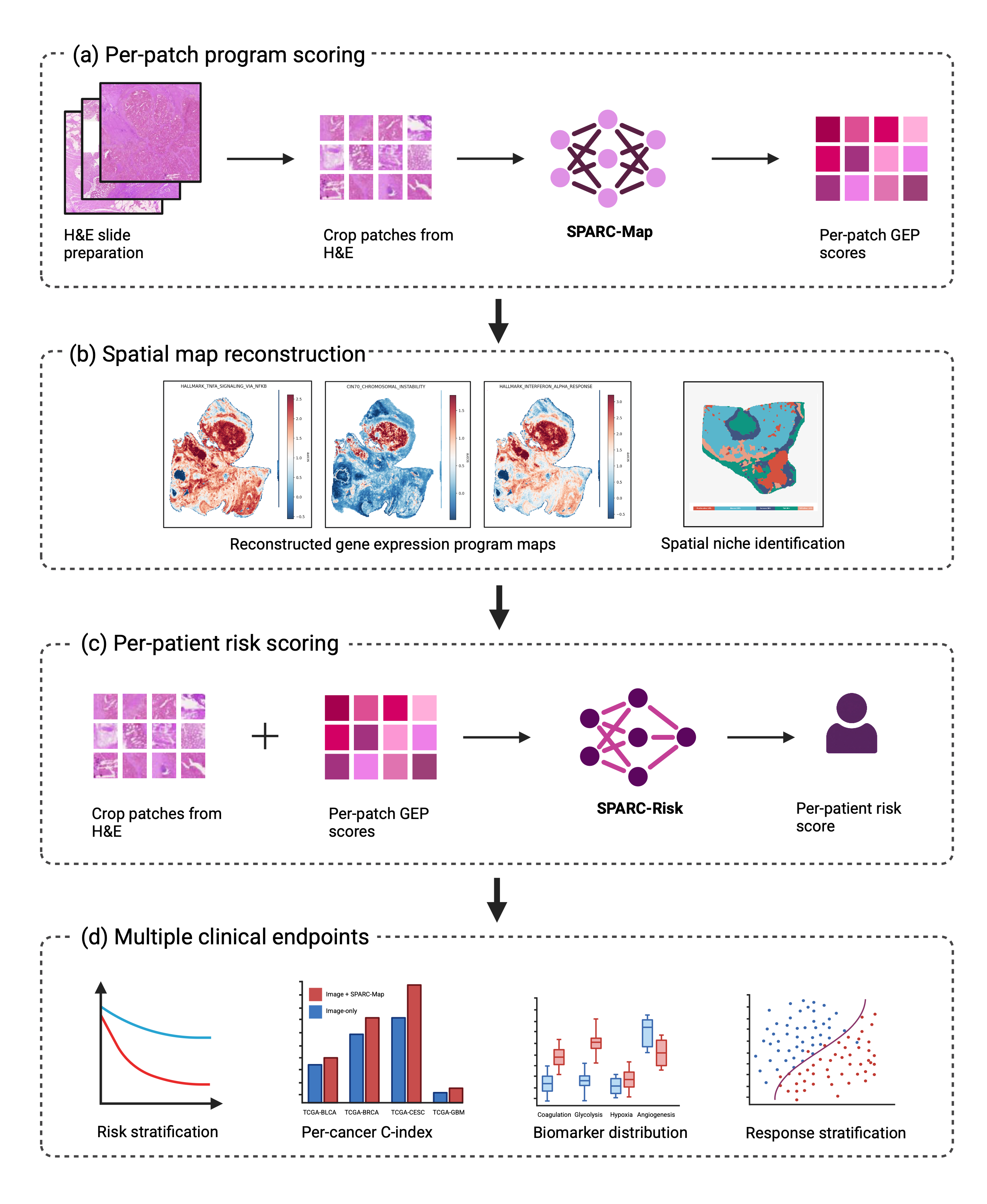
SPARC is a two-stage pipeline:
- SPARC-Map predicts 40 hallmark gene-expression-program (GEP) scores per H&E patch, recovering a spatial molecular map of each slide.
- SPARC-Risk fuses those per-patch GEP scores with the same H&E features through a signature-query attention head and a cancer-aware gate, producing a single per-patient risk score.
These weights cover the SPARC-Risk model and the image-only baseline used for ablations.
What you get
| Folder | Model | Description |
|---|---|---|
sparc_risk/ |
SPARC-Risk (canonical) | Signature-query fusion + H&E. The model reported throughout the paper. |
image_only/ |
Image-only baseline | Same backbone, GEP pathway disabled. Use for direct ablation against SPARC-Risk. |
Each folder contains 5 checkpoints — fold_0_best.pt through
fold_4_best.pt — corresponding to the 5-fold cross-validation splits
described in the paper and in
data/mmp_hybrid_splits_v2_20cancer.csv.
Every .pt carries both model_state_dict and the original training
config, so the model can be rebuilt with one line:
import torch
from sparc.models.factory import build_model
ckpt = torch.load("sparc_risk/fold_0_best.pt", map_location="cpu", weights_only=False)
model = build_model(ckpt["config"])
model.load_state_dict(ckpt["model_state_dict"])
model.eval()
Quick start
# 1. Install the SPARC package
git clone https://github.com/aziz-ayed/SPARC.git && cd SPARC
conda env create -f environment.yml
conda activate sparc
# 2. Accept the license on https://huggingface.co/azizayed/SPARC, then:
pip install -U "huggingface_hub[cli]"
hf auth login
hf download azizayed/SPARC --local-dir checkpoints
# 3. Inference on an external cohort (e.g. NLST lung)
python -m inference.run \
--cohort nlst \
--checkpoint_dir checkpoints/sparc_risk \
--gpus 0,1,2,3
The download produces:
checkpoints/
├── sparc_risk/ fold_{0..4}_best.pt
└── image_only/ fold_{0..4}_best.pt
Architecture (SPARC-Risk)
| Component | Setting |
|---|---|
| Image backbone | H-optimus-1 (1536-dim) |
| Patch size / magnification | 224 px @ 20× |
| Max patches per slide | 4096 |
| Fusion | Signature-query cross-attention (64-NN, 4 heads) |
| Aggregator | Gated attention MIL |
| Head | Discrete-time NLL survival, 4 bins |
| Cancer conditioning | Per-cancer learned gate |
| Hidden dim | 256 |
| Trainable params | ≈ 2.6 M |
| Optimiser / schedule | Adam, lr 1 × 10⁻⁴, cosine T_max 20 |
| Random seed | 1337 |
Full config + reproduction recipe: configs/sparc_risk.yaml.
Training data
5-fold patient-level cross-validation over 20 TCGA cancer types
(BLCA, BRCA, CESC, COAD, ESCA, GBM, HNSC, KIRC, KIRP, LGG, LIHC, LUAD,
LUSC, PAAD, READ, SARC, SKCM, STAD, UCEC, plus a held-out evaluation
split). Splits derive from the MMP hybrid scheme of Mahmood et al. and
are released alongside the code at
data/mmp_hybrid_splits_v2_20cancer.csv.
External validation cohorts (not used for training) — NLST lung, SurGen CRC, Yale breast, ovarian — are described in the paper.
Intended use
These weights are intended for non-commercial biomedical research and education only. Acceptable uses include:
- Reproducing the SPARC paper's results.
- Benchmarking against SPARC-Risk in computational-pathology research.
- Methodological extensions (new fusion designs, additional cohorts, ablation studies).
Citation
The SPARC paper is currently under review. Once a preprint or accepted version is available, a BibTeX entry will be added here. In the meantime, if you use these weights, please link back to github.com/aziz-ayed/SPARC and contact the corresponding author at azizayed@mit.edu.
License
These weights are released under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC 4.0). For commercial licensing, please contact the authors via the corresponding GitHub issues page.