metadata
license: mit
datasets:
- chaitjo/gRNAde_datasets
tags:
- rna
- biology
- rna-design
- biomolecule-design
- 3d-design
- inverse-folding
- inverse-design
gRNAde Model Checkpoints
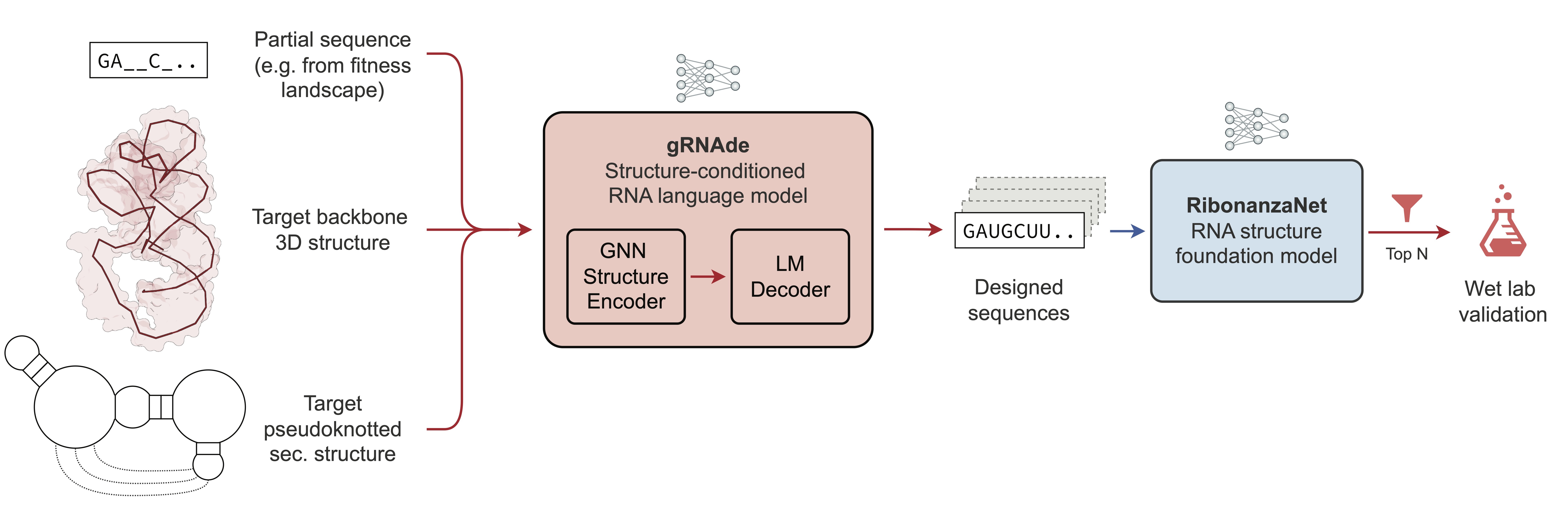
This repository contains pre-trained model checkpoints for gRNAde, a generative AI framework for RNA inverse design.
📦 Model Checkpoints
.
├── gRNAde_drop3d@0.75_maxlen@500.h5 # Main gRNAde model checkpoint
├── rhofold/ # RhoFold checkpoint (optional)
├── ribonanzanet/ # RibonanzaNet checkpoint
└── ribonanzanet_sec_struct/ # RibonanzaNet secondary structure checkpoint
gRNAde Model
- File:
gRNAde_drop3d@0.75_maxlen@500.h5 - Description: Core gRNAde model trained on RNA structures from PDB (≤4Å resolution, RNASolo database, Oct 2023 cutoff)
- Architecture: Geometric Graph Neural Network conditioned on 3D backbone coordinates and secondary structures
- Training: 75% 3D coordinate dropout, maximum sequence length 500 nucleotides
- Use case: Generating RNA sequences for target 3D structures and secondary structures
RibonanzaNet
- Directory:
ribonanzanet/ - Description: RNA structure foundation model for predicting per-nucleotide SHAPE reactivity profiles
- Use case: High-throughput computational screening of designed sequences
- Reference: Trained on Ribonanza dataset with diverse natural and synthetic RNAs
RibonanzaNet Secondary Structure
- Directory:
ribonanzanet_sec_struct/ - Description: RibonanzaNet variant for predicting pseudoknotted secondary structures
- Use case: Alternative structural screening metric and OpenKnot score calculation
RhoFold (Optional)
- Directory:
rhofold/ - Description: RNA sequence to 3D structure prediction tool
- Use case: Predicting 3D structures of designed sequences (not used by default in the pipeline)
🚀 Quick Start
After setting up the gRNAde codebase, checkpoints can be downloaded manually and placed in the appropriate directory, or using HuggingFace CLI (recommended):
# Ensure you are in the base directory
cd ~/geometric-rna-design
# Install HuggingFace CLI (https://huggingface.co/docs/huggingface_hub/main/en/guides/cli)
curl -LsSf https://hf.co/cli/install.sh | bash
# alternate: pip install -U "huggingface_hub", or brew install huggingface-cli
hf auth login
# Download all checkpoints to checkpoints/ directory
huggingface-cli download chaitjo/gRNAde --local-dir checkpoints/
Alternatively, download specific files:
# Download only the main gRNAde checkpoint
huggingface-cli download chaitjo/gRNAde gRNAde_drop3d@0.75_maxlen@500.h5 --local-dir checkpoints/
# Download only RibonanzaNet checkpoints (required for design pipeline)
huggingface-cli download chaitjo/gRNAde ribonanzanet/ --local-dir checkpoints/
huggingface-cli download chaitjo/gRNAde ribonanzanet_sec_struct/ --local-dir checkpoints/
Citations
@article{joshi2025generative,
title={Generative inverse design of RNA structure and function with g{RNA}de},
author={Joshi, Chaitanya K and Gianni, Edoardo and Kwok, Samantha LY and Mathis, Simon V and Lio, Pietro and Holliger, Philipp},
journal={bioRxiv},
year={2025},
publisher={Cold Spring Harbor Laboratory}
}
@inproceedings{joshi2025grnade,
title={g{RNA}de: Geometric Deep Learning for 3D RNA inverse design},
author={Joshi, Chaitanya K and Jamasb, Arian R and Vi{\~n}as, Ramon and Harris, Charles and Mathis, Simon V and Morehead, Alex and Anand, Rishabh and Li{\`o}, Pietro},
booktitle={International Conference on Learning Representations (ICLR)},
year={2025},
}
@incollection{joshi2024grnade,
title={g{RNA}de: A Geometric Deep Learning pipeline for 3D RNA inverse design},
author={Joshi, Chaitanya K and Li{\`o}, Pietro},
booktitle={RNA Design: Methods and Protocols},
pages={121--135},
year={2024},
publisher={Springer}
}