File size: 7,933 Bytes
a548e03 b7bb405 a548e03 |
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 |
---
dataset_name: "CROssBARv2-KG"
tags:
- biomedical
- knowledge-graph
- bioinformatics
- biology
- chemistry
- systems-biology
task_categories:
- graph-construction
- link-prediction
- node-classification
- graph-ml
pretty_name: "CROssBARv2 Knowledge Graph Dataset"
language:
- en
size_categories:
- 10M<n<100M
---
# CROssBARv2-KG
This repository provides the dataset for the CROssBARv2 Knowledge Graph (KG), a heterogeneous and general-purpose biomedical KG-based system.
The CROssBARv2 KG comprises approximately **2.7 million** nodes spanning **14 distinct node types** and around **12.6 million edges** representing **51 different edge types**, all integrated from **34 biological data sources**. We also incorporated several ontologies (e.g., Gene Ontology, Mondo Disease Ontology) along with rich metadata captured as node and edge properties.
Building upon this foundation, we further enhanced the semantic depth of CROssBARv2. This was achieved by generating and storing embeddings for key biological entities, such as proteins, drugs, and Gene Ontology terms. These embeddings are managed using the native [vector index](https://neo4j.com/developer/genai-ecosystem/vector-search/) feature in Neo4j, enabling powerful semantic similarity searches.
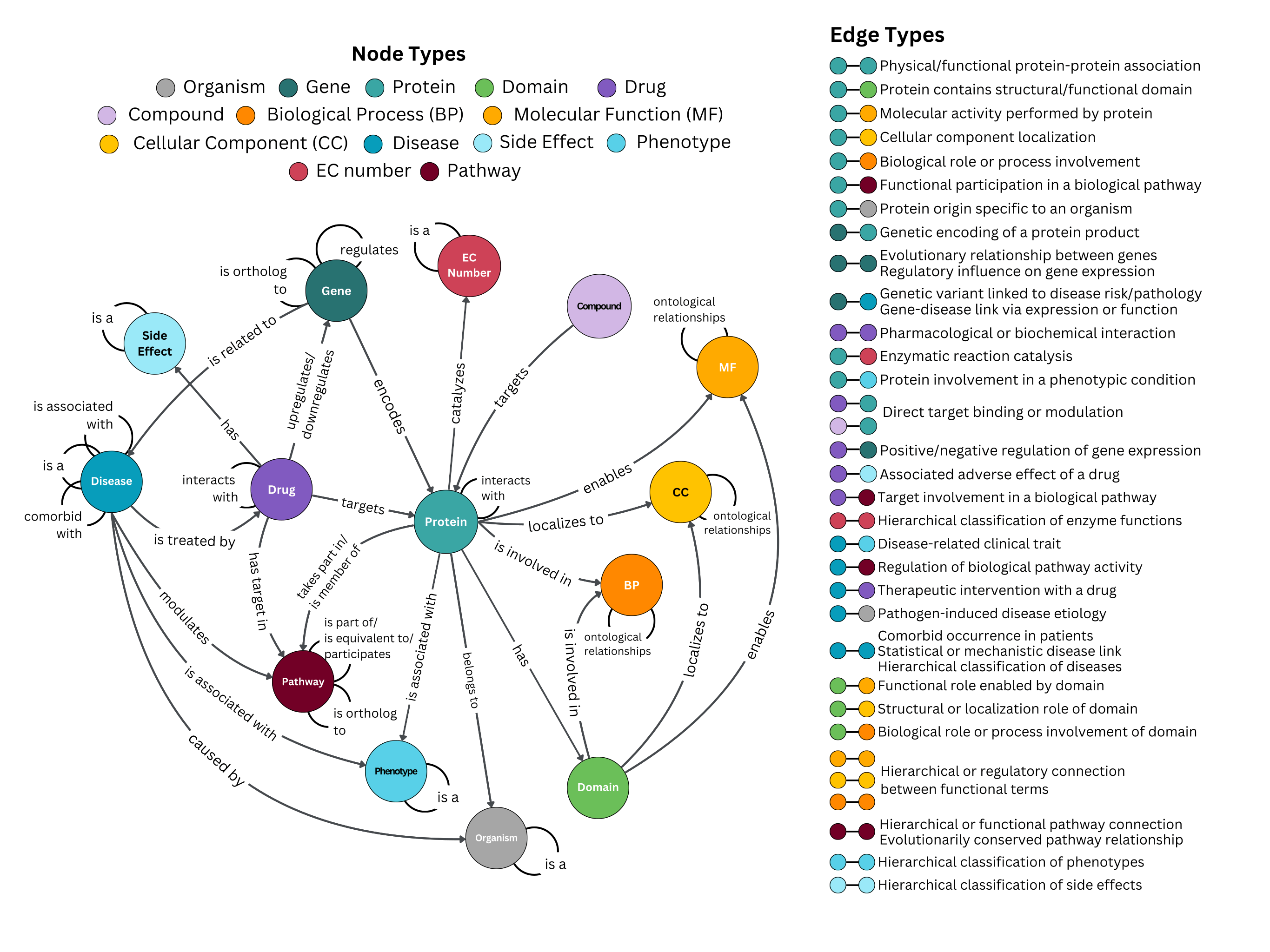
# Data
The dataset is organized into two primary directories:
- [**`nodes/`**](https://huggingface.co/datasets/HUBioDataLab/CROssBARv2-KG/tree/main/nodes): Contains CSV files defining the biological entities.
- [**`edges/`**](https://huggingface.co/datasets/HUBioDataLab/CROssBARv2-KG/tree/main/edges): Contains CSV files defining the relationships between these entities.
## nodes
Each file corresponds to a specific biological entity type and contains a primary identifier column that provides a unique ID along with additional metadata fields. Identifiers follow the **Compact URI (CURIE)** standard (e.g., `uniprot:Q9H161`) as defined in the [Bioregistry](https://bioregistry.io/).
The table below lists the primary identifier column name for each node file:
| **File** | **ID Column Name** |
| :--- | :--- |
| Cellular_component.csv | id |
| Biological_process.csv | id |
| Molecular_function.csv | id |
| Protein.csv | id |
| Phenotype.csv | hpo_id |
| Pathway.csv | pathway_id |
| Ec.csv | ec_number |
| Compound.csv | compound_id |
| Gene.csv | id |
| Side_effect.csv | meddra_id |
| Drug.csv | drugbank_id |
| Drug.csv | drugbank_id |
| Organism.csv | id |
| Domain.csv | id |
| Disease.csv | disease_id |
The following table demonstrates the CURIE format used for each node type within the KG:
| **Node Type** | **CURIE** |
|--------------------------------------------------|------------------------|
| Protein | uniprot:Q9H161 |
| Gene | ncbigene:60529 |
| OrganismTaxon | ncbitaxon:9606 |
| ProteinDomain | interpro:IPR000001 |
| Drug | drugbank:DB00821 |
| Compound | chembl:CHEMBL6228 |
| GOTerm (BiologicalProcess, MolecularFunction, CellularComponent) | go:0016072 |
| Disease | mondo:0054666 |
| Phenotype | hp:0000012 |
| SideEffect | meddra:10073487 |
| EcNumber | eccode:1.1.1.- |
## edges
Each file represents a relationship between two biological entity types in the KG.
Every edge file contains **source** and **target** columns, which specify the identifiers of the nodes being linked.
Other columns serve as metadata for the relationship.
The table below lists the filename along with the source and target identifier column names and a description of the edge type:
| **File** | **Source Column** | **Target Column** | **Description** |
| :--- | :--- | :--- | :--- |
| Pathway_orthology.csv | pathway1_id | pathway2_id | Evolutionarily conserved pathway relationship |
| Go_to_go.csv | source | target | Hierarchical or regulatory connection between functional terms |
| Gene_to_disease_edge.csv | gene_id | disease_id | <p>Gene-disease link via expression or function<br> Genetic variant linked to disease risk/pathology<p> |
| Ec_hierarchy.csv | child_id | parent_id | Hierarchical classification of enzyme functions |
| Disease_to_drug_edge.csv | disease_id | drug_id | Therapeutic intervention with a drug |
| DTI.csv | drugbank_id | uniprot_id | Direct target binding or modulation |
| Phenotype_hierarchical_edges.csv | child_id | parent_id | Hierarchical classification of phenotypes |
| Protein_to_phenotype.csv | protein_id | hpo_id | Protein involvement in a phenotypic condition |
| Drug_to_side_effect.csv | drugbank_id | meddra_id | Associated adverse effect of a drug |
| Disease_to_disease_comorbidity_edge.csv | disease1 | disease2 | Comorbid occurrence in patients |
| DGI.csv | entrez_id | drugbank_id | Positive/negative regulation of gene expression |
| Drug_to_pathway.csv | drug_id | pathway_id | Target involvement in a biological pathway |
| PPI.csv | uniprot_a | uniprot_b | Physical/functional protein-protein association |
| Disease_to_disease_association_edge.csv | disease_id1 | disease_id2 | Statistical or mechanistic disease link |
| Phenotype_to_disease.csv | hpo_id | disease_id | Disease-related clinical trait |
| Orthology.csv | entrez_a | entrez_b | Evolutionary relationship between genes |
| Side_effect_hierarchy.csv | child_id | parent_id | Hierarchical classification of side effects |
| Organism_to_disease_edge.csv | organism_id | disease_id | Pathogen-induced disease etiology |
| DDI.csv | drug1 | drug2 | Pharmacological or biochemical interaction |
| Protein_to_ec.csv | protein_id | ec_id | Enzymatic reaction catalysis |
| Reactome_hierarchical_edges.csv | child_id | parent_id | Hierarchical or functional pathway connection |
| Tf_gene_edges.csv | tf | target | Regulatory influence on gene expression |
| Disease_hiererchical_edges.csv | child_id | parent_id | Hierarchical classification of diseases |
| Protein_to_pathway.csv | uniprot_id | pathway_id | Functional participation in a biological pathway |
| Protein_has_domain.csv | source_id | target_id | Protein contains structural/functional domain |
| Protein_belongs_to_organism.csv | source_id | target_id | Protein origin specific to an organism |
| Protein_to_go.csv | source | target | Molecular activity performed by protein / Cellular component localization / Biological role or process involvement |
| Pathway_to_pathway.csv | pathway_id1 | pathway_id2 | Hierarchical or functional pathway connection |
| Gene_encodes_protein.csv | source_id | target_id | Genetic encoding of a protein product |
| Disease_to_pathway.csv | disease_id | pathway_id | Regulation of biological pathway activity |
| Domain_to_go.csv | source | target | Functional role enabled by domain / Structural or localization role of domain / Biological role or process involvement of domain |
| CTI.csv | chembl | uniprot_id | Direct target binding or modulation |
# How to Use
You can easily load this dataset using the Hugging Face `datasets` library:
```python
from datasets import load_dataset
# Example 1: Load the Protein nodes
proteins = load_dataset("HUBioDataLab/CROssBARv2-KG", data_files="nodes/Protein.csv")
# Example 2: Load Drug-Target Interactions (Edges)
dti_edges = load_dataset("HUBioDataLab/CROssBARv2-KG", data_files="edges/DTI.csv")
```
|