instance_id stringlengths 13 45 | pull_number int64 7 30.1k | repo stringclasses 83
values | version stringclasses 68
values | base_commit stringlengths 40 40 | created_at stringdate 2013-05-16 18:15:55 2025-01-08 15:12:50 | patch stringlengths 347 35.2k | test_patch stringlengths 432 113k | non_py_patch stringlengths 0 18.3k | new_components listlengths 0 40 | FAIL_TO_PASS listlengths 1 2.53k | PASS_TO_PASS listlengths 0 1.7k | problem_statement stringlengths 607 52.7k | hints_text stringlengths 0 57.4k | environment_setup_commit stringclasses 167
values |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
sympy__sympy-19681 | 19,681 | sympy/sympy | 1.7 | 9104980668755d0611af1de37d06e8401a0e2f98 | 2020-07-01T13:06:49Z | diff --git a/sympy/vector/__init__.py b/sympy/vector/__init__.py
index 718cb9575cad..29f825f9e95b 100644
--- a/sympy/vector/__init__.py
+++ b/sympy/vector/__init__.py
@@ -15,6 +15,7 @@
SpaceOrienter, QuaternionOrienter)
from sympy.vector.operators import Gradient, Divergence, Curl,... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 6bdc452c62c2..68b39b3ec04a 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -4892,6 +4892,12 @@ def test_sympy__vector__deloperator__Del():
assert _test_args(Del())
+def test_sympy__vector__implici... | [
{
"components": [
{
"doc": "Represents an implicit region in space.\n\nExample\n=======\n\n>>> from sympy import Eq\n>>> from sympy.abc import x, y, z, t\n>>> from sympy.vector import ImplicitRegion\n\n>>> ImplicitRegion((x, y), x**2 + y**2 - 4)\nImplicitRegion((x, y), x**2 + y**2 - 4)\n>>> Implic... | [
"test_sympy__vector__implicitregion__ImplicitRegion",
"test_ImplicitRegion",
"test_regular_point",
"test_singular_points_and_multiplicty"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Adding classes to represent implicitly defined regions
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to... | 1b4529a95ef641c2fc15889091b281644069d20e | ||
Project-MONAI__MONAI-640 | 640 | Project-MONAI/MONAI | null | 39f6df581d72562c43207ddf408195c10bf845e7 | 2020-06-28T23:34:41Z | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index b02db7677b..1c8b84152f 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -373,6 +373,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`LabelToContour`
+~~~~~~~~~~~~~~~~
+.. autoclass:: LabelToC... | diff --git a/tests/test_label_to_contour.py b/tests/test_label_to_contour.py
index e90dcc73c6..cd9bf88de4 100644
--- a/tests/test_label_to_contour.py
+++ b/tests/test_label_to_contour.py
@@ -171,10 +171,6 @@ def test_contour(self):
error_input = torch.rand(1, 2, 3, 4, 5, 6)
self.assertRaises(RuntimeEr... | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index b02db7677b..1c8b84152f 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -373,6 +373,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`LabelToContour`
+~~~~~~~~~~~~~~~~
+.. autoclass:: LabelToC... | [
{
"components": [
{
"doc": "dictionary-based wrapper of :py:class:monai.transforms.LabelToContour.",
"lines": [
190,
210
],
"name": "LabelToContourd",
"signature": "class LabelToContourd(MapTransform):",
"type": "class"
},
{
... | [
"tests/test_label_to_contour.py::TestContour::test_contour",
"tests/test_label_to_contourd.py::TestContourd::test_contour"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
634 dictionary level LabelToContourd
Fixes #634 .
### Description
This PR refined the `LabelToContour` transform and added dictionary version.
### Status
**Ready**
### Types of changes
<!-... | Here is the discussion in the issues of the pull request.
<issues>
add dictionary level LabelToContourd transform
**Is your feature request related to a problem? Please describe.**
As we already have array level LabelToContour transform, need to update the document and add dictionary level version.
----------
--------... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
Project-MONAI__MONAI-639 | 639 | Project-MONAI/MONAI | null | e8c26a2baa539624c36aa1262e6de9009c74cfda | 2020-06-28T16:44:30Z | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index 673d4bae91..b02db7677b 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -343,6 +343,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`Lambda`
+~~~~~~~~
+.. autoclass:: Lambda
+ :members:
+ ... | diff --git a/tests/test_lambda.py b/tests/test_lambda.py
new file mode 100644
index 0000000000..96ebd88705
--- /dev/null
+++ b/tests/test_lambda.py
@@ -0,0 +1,41 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in compliance wit... | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index 673d4bae91..b02db7677b 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -343,6 +343,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`Lambda`
+~~~~~~~~
+.. autoclass:: Lambda
+ :members:
+ ... | [
{
"components": [
{
"doc": "Apply a user-defined lambda as a transform.\n\nFor example:\n\n.. code-block:: python\n :emphasize-lines: 2\n\n image = np.ones((10, 2, 2))\n lambd = Lambda(func=lambda x: x[:4, :, :])\n print(lambd(image).shape)\n (4, 2, 2)\n\nArgs:\n func: Lambda/fun... | [
"tests/test_lambda.py::TestLambda::test_lambda_identity",
"tests/test_lambda.py::TestLambda::test_lambda_slicing",
"tests/test_lambdad.py::TestLambdad::test_lambdad_identity",
"tests/test_lambdad.py::TestLambdad::test_lambdad_slicing"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
630 Lambda Transform
Fixes #630
### Description
Adding array and dictionary lambda transform.
### Status
**Ready**
### Types of changes
<!--- Put an `x` in all the boxes that apply, and r... | Here is the discussion in the issues of the pull request.
<issues>
lambda transform
**Is your feature request related to a problem? Please describe.**
this feature request is about providing a transform that supports use-defined simple lambda functions.
a use case came from the experiments is, for example,
only re... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
Project-MONAI__MONAI-615 | 615 | Project-MONAI/MONAI | null | d25e4e56ef9c5e56fefbce0478b61e1f2a114c9f | 2020-06-24T16:05:39Z | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index 8dc3bef951..673d4bae91 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -187,6 +187,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`ScaleIntensityRangePercentiles`
+~~~~~~~~~~~~~~~~~~~~~~~~~... | diff --git a/tests/test_scale_intensity_range_percentiles.py b/tests/test_scale_intensity_range_percentiles.py
new file mode 100644
index 0000000000..ace11fcc8c
--- /dev/null
+++ b/tests/test_scale_intensity_range_percentiles.py
@@ -0,0 +1,60 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, V... | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index 8dc3bef951..673d4bae91 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -187,6 +187,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`ScaleIntensityRangePercentiles`
+~~~~~~~~~~~~~~~~~~~~~~~~~... | [
{
"components": [
{
"doc": "Apply range scaling to a numpy array based on the intensity distribution of the input.\n\nBy default this transform will scale from [lower_intensity_percentile, upper_intensity_percentile] to [b_min, b_max], where \n{lower,upper}_intensity_percentile are the intensity v... | [
"tests/test_scale_intensity_range_percentiles.py::TestScaleIntensityRangePercentiles::test_invalid_instantiation",
"tests/test_scale_intensity_range_percentiles.py::TestScaleIntensityRangePercentiles::test_relative_scaling",
"tests/test_scale_intensity_range_percentiles.py::TestScaleIntensityRangePercentiles::t... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
614 ScaleIntensityRangePercentiles transform
Fixes #614
### Description
Adds a data transform to rescale an image based on upper and lower intensity percentiles.
### Status
Ready
TODOs:
... | Here is the discussion in the issues of the pull request.
<issues>
Add percentile version of ScaleIntensityRange
**Describe the solution you'd like**
A variant of `monai.transforms.ScaleIntensityRange` that has upper and lower intensity percentile parameters instead of `a_min` and `a_max`.
So this transform... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
matplotlib__matplotlib-17709 | 17,709 | matplotlib/matplotlib | 3.2 | ae47ad26eda9c69b182837feaece416c6017d29f | 2020-06-22T02:27:58Z | diff --git a/doc/api/colors_api.rst b/doc/api/colors_api.rst
index d0c05aeaef35..e6fad2993f5d 100644
--- a/doc/api/colors_api.rst
+++ b/doc/api/colors_api.rst
@@ -20,6 +20,7 @@ Classes
BoundaryNorm
Colormap
+ CenteredNorm
LightSource
LinearSegmentedColormap
ListedColormap
diff --git a/doc/users/... | diff --git a/lib/matplotlib/tests/test_colors.py b/lib/matplotlib/tests/test_colors.py
index 97790de87617..f174d7232c33 100644
--- a/lib/matplotlib/tests/test_colors.py
+++ b/lib/matplotlib/tests/test_colors.py
@@ -359,6 +359,56 @@ def test_BoundaryNorm():
assert_array_equal(cmshould(mynorm(x)), cmref(refnorm(x)))... | diff --git a/doc/api/colors_api.rst b/doc/api/colors_api.rst
index d0c05aeaef35..e6fad2993f5d 100644
--- a/doc/api/colors_api.rst
+++ b/doc/api/colors_api.rst
@@ -20,6 +20,7 @@ Classes
BoundaryNorm
Colormap
+ CenteredNorm
LightSource
LinearSegmentedColormap
ListedColormap
diff --git a/doc/users/... | [
{
"components": [
{
"doc": "",
"lines": [
1258,
1350
],
"name": "CenteredNorm",
"signature": "class CenteredNorm(Normalize):",
"type": "class"
},
{
"doc": "Normalize symmetrical data around a center (0 by default).\n\n... | [
"lib/matplotlib/tests/test_colors.py::test_CenteredNorm"
] | [
"lib/matplotlib/tests/test_colors.py::test_create_lookup_table[5-result0]",
"lib/matplotlib/tests/test_colors.py::test_create_lookup_table[2-result1]",
"lib/matplotlib/tests/test_colors.py::test_create_lookup_table[1-result2]",
"lib/matplotlib/tests/test_colors.py::test_resample",
"lib/matplotlib/tests/test... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Enh: SymNorm for normalizing symmetrical data around a center
## PR Summary
Adds `SymNorm`, a new norm that automatically creates linear colormaps that are symmetrical around a center. Useful for s... | ff821ba3249401fe7f5fdb11cb7ac5d0564c2697 | |
Project-MONAI__MONAI-599 | 599 | Project-MONAI/MONAI | null | 9d0396ba76cb054a623a71e68af80e056040047d | 2020-06-20T14:24:02Z | diff --git a/monai/transforms/post/array.py b/monai/transforms/post/array.py
index a6988ccd86..c50833e4c2 100644
--- a/monai/transforms/post/array.py
+++ b/monai/transforms/post/array.py
@@ -292,3 +292,60 @@ def __call__(self, img):
output = img
return output
+
+
+class LabelToContour(Transform)... | diff --git a/tests/test_label_to_contour.py b/tests/test_label_to_contour.py
new file mode 100644
index 0000000000..e90dcc73c6
--- /dev/null
+++ b/tests/test_label_to_contour.py
@@ -0,0 +1,180 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use thi... | [
{
"components": [
{
"doc": "Return the contour flag of objects in mask images that only compose of 0 and 1, with Laplace kernel\n set as default for edge detection.\n\nArgs:\n kernel_type: the method applied to do edge detection.",
"lines": [
297,
351
],
... | [
"tests/test_label_to_contour.py::TestContour::test_contour"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add edge detection for mask image
### Description
Add edge detection for mask image
### Status
**Ready**
### Types of changes
<!--- Put an `x` in all the boxes that apply, and remove the not ... | e73257caa79309dcce1e93abf1632f4bfd75b11f | ||
joke2k__faker-1206 | 1,206 | joke2k/faker | null | 7ecd296e5393f33697d98d1e3b54b9cbf9cd2629 | 2020-06-19T23:15:41Z | diff --git a/faker/providers/person/__init__.py b/faker/providers/person/__init__.py
index f19f9f7144..cfb53278cc 100644
--- a/faker/providers/person/__init__.py
+++ b/faker/providers/person/__init__.py
@@ -67,6 +67,14 @@ def name_male(self):
pattern = self.random_element(formats)
return self.generato... | diff --git a/tests/providers/test_person.py b/tests/providers/test_person.py
index 3aec73e704..add22acf14 100644
--- a/tests/providers/test_person.py
+++ b/tests/providers/test_person.py
@@ -7,6 +7,8 @@
from faker import Faker
from faker.providers.person.ar_AA import Provider as ArProvider
from faker.providers.perso... | [
{
"components": [
{
"doc": "",
"lines": [
70,
76
],
"name": "Provider.name_nonbinary",
"signature": "def name_nonbinary(self):",
"type": "function"
},
{
"doc": "",
"lines": [
91,
94
... | [
"tests/providers/test_person.py::TestUs::test_first_names",
"tests/providers/test_person.py::TestUs::test_last_names",
"tests/providers/test_person.py::TestUs::test_prefix",
"tests/providers/test_person.py::TestUs::test_suffix"
] | [
"tests/providers/test_person.py::TestAr::test_first_name",
"tests/providers/test_person.py::TestAr::test_last_name",
"tests/providers/test_person.py::TestJaJP::test_person",
"tests/providers/test_person.py::TestNeNP::test_names",
"tests/providers/test_person.py::TestFiFI::test_gender_first_names",
"tests/... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Extend Person Provider to support nonbinary suffixes and prefixes
### What does this change
* Extends Person Provider to support nonbinary suffixes and prefixes
* Provides a nonbinary name formatt... | Here is the discussion in the issues of the pull request.
<issues>
Feature Request: Support nonbinary gender options in Person Provider
* Faker version: < Faker 4.1.1
* OS: All
Many applications support more than a straightforward binary Male/Female gender selection and require adequate test data. Presently, Faker ... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | |
Project-MONAI__MONAI-582 | 582 | Project-MONAI/MONAI | null | c98da0db72c5bbef53f191612a0812b880ee312c | 2020-06-17T08:00:56Z | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index 7c99f638d7..6cd9143607 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -641,6 +641,12 @@ Dictionary-based Transforms
:members:
:special-members: __call__
+`ConcatItemsd`
+~~~~~~~~~~~~~~
+.. autoclass:: Con... | diff --git a/tests/test_concat_itemsd.py b/tests/test_concat_itemsd.py
new file mode 100644
index 0000000000..5b6261b8f7
--- /dev/null
+++ b/tests/test_concat_itemsd.py
@@ -0,0 +1,43 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file exc... | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index 7c99f638d7..6cd9143607 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -641,6 +641,12 @@ Dictionary-based Transforms
:members:
:special-members: __call__
+`ConcatItemsd`
+~~~~~~~~~~~~~~
+.. autoclass:: Con... | [
{
"components": [
{
"doc": "Concatenate specified items from data dictionary together on the first dim to construct a big array.\nExpect all the items are numpy array or PyTorch Tensor.",
"lines": [
367,
405
],
"name": "ConcatItemsd",
"signature"... | [
"tests/test_concat_itemsd.py::TestConcatItemsd::test_numpy_values",
"tests/test_concat_itemsd.py::TestConcatItemsd::test_tensor_values"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
580 add ConcatItems transform
Fixes #580 .
### Description
This PR implemented `ConcatItems` transform.
### Status
**Ready**
### Types of changes
<!--- Put an `x` in all the boxes that app... | Here is the discussion in the issues of the pull request.
<issues>
Add ConcatItems transform
**Is your feature request related to a problem? Please describe.**
In order to combine several transform results together, need a `ConcatItems` transform.
For example, take 3 different intensity ranges from 1 image and then c... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
scikit-learn__scikit-learn-17612 | 17,612 | scikit-learn/scikit-learn | 0.24 | 071ce4bf8672a5fb9b3df66c2b2b5515a09a52d0 | 2020-06-16T14:20:53Z | diff --git a/doc/whats_new/v0.24.rst b/doc/whats_new/v0.24.rst
index 6b39fb01e35b0..19aa962da8269 100644
--- a/doc/whats_new/v0.24.rst
+++ b/doc/whats_new/v0.24.rst
@@ -91,6 +91,11 @@ Changelog
when ``strategy='most_frequent'`` or ``strategy='constant'``.
:pr:`17526` by :user:`Ayako YAGI <yagi-3>` and :user:`Juan... | diff --git a/sklearn/impute/tests/test_impute.py b/sklearn/impute/tests/test_impute.py
index 53580f32c34ca..8a3643a7628c5 100644
--- a/sklearn/impute/tests/test_impute.py
+++ b/sklearn/impute/tests/test_impute.py
@@ -1383,3 +1383,68 @@ def test_imputation_order(order, idx_order):
random_... | diff --git a/doc/whats_new/v0.24.rst b/doc/whats_new/v0.24.rst
index 6b39fb01e35b0..19aa962da8269 100644
--- a/doc/whats_new/v0.24.rst
+++ b/doc/whats_new/v0.24.rst
@@ -91,6 +91,11 @@ Changelog
when ``strategy='most_frequent'`` or ``strategy='constant'``.
:pr:`17526` by :user:`Ayako YAGI <yagi-3>` and :user:`Juan... | [
{
"components": [
{
"doc": "Convert the data back to the original representation.\n\nInverts the `transform` operation performed on an array.\nThis operation can only be performed after :class:`SimpleImputer` is\ninstantiated with `add_indicator=True`.\n\nNote that ``inverse_transform`` can only i... | [
"sklearn/impute/tests/test_impute.py::test_simple_imputation_inverse_transform[-1]",
"sklearn/impute/tests/test_impute.py::test_simple_imputation_inverse_transform[nan]",
"sklearn/impute/tests/test_impute.py::test_simple_imputation_inverse_transform_exceptions[-1]",
"sklearn/impute/tests/test_impute.py::test_... | [
"sklearn/impute/tests/test_impute.py::test_imputation_shape[mean]",
"sklearn/impute/tests/test_impute.py::test_imputation_shape[median]",
"sklearn/impute/tests/test_impute.py::test_imputation_shape[most_frequent]",
"sklearn/impute/tests/test_impute.py::test_imputation_shape[constant]",
"sklearn/impute/tests... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
ENH Add inverse_transform feature to SimpleImputer
<!--
Thanks for contributing a pull request! Please ensure you have taken a look at
the contribution guidelines: https://github.com/scikit-learn/sc... | Here is the discussion in the issues of the pull request.
<issues>
MissingIndicator.inverse_transform and SimpleImputer.inverse_transform needed
#### Describe the workflow you want to enable
When using MissingIndicator and SimpleImputer, I need to inverse the transform. If a missing indicator is present, the missing... | 54ce4222694819ad52d544ce5cba5da274c34ab7 |
joke2k__faker-1199 | 1,199 | joke2k/faker | null | 845d757e882db400332af8b7365181515c01df90 | 2020-06-15T23:40:39Z | diff --git a/faker/providers/currency/es_ES/__init__.py b/faker/providers/currency/es_ES/__init__.py
new file mode 100644
index 0000000000..e7f9006b9c
--- /dev/null
+++ b/faker/providers/currency/es_ES/__init__.py
@@ -0,0 +1,171 @@
+from .. import Provider as CurrencyProvider
+
+
+class Provider(CurrencyProvider):
+ ... | diff --git a/tests/providers/test_currency.py b/tests/providers/test_currency.py
index 9432b7a4db..d88231693c 100644
--- a/tests/providers/test_currency.py
+++ b/tests/providers/test_currency.py
@@ -88,3 +88,25 @@ def test_currency_name(self, faker, num_samples):
for _ in range(num_samples):
name ... | [
{
"components": [
{
"doc": "",
"lines": [
4,
170
],
"name": "Provider",
"signature": "class Provider(CurrencyProvider):",
"type": "class"
}
],
"file": "faker/providers/currency/es_ES/__init__.py"
}
] | [
"tests/providers/test_currency.py::TestEsEs::test_currency",
"tests/providers/test_currency.py::TestEsEs::test_currency_name"
] | [
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency",
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency_code",
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency_name",
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency_symbol_no_... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add es_ES currency provider.
### What does this changes
Add Spanish currency names to `currency` provider.
----------
</request>
There are several new functions or classes that need to be impleme... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
facebookresearch__hydra-683 | 683 | facebookresearch/hydra | null | c29ba76b4b024ed3705db7fde3166f28bcd2fca6 | 2020-06-15T17:33:40Z | diff --git a/news/683.plugin b/news/683.plugin
new file mode 100644
index 00000000000..cc5a21c1cca
--- /dev/null
+++ b/news/683.plugin
@@ -0,0 +1,1 @@
+Add [Redis Queue](https://python-rq.org/) Launcher plugin ([Jan-Matthis](https://github.com/jan-matthis))
\ No newline at end of file
diff --git a/plugins/hydra_rq_laun... | diff --git a/plugins/hydra_rq_launcher/tests/__init__.py b/plugins/hydra_rq_launcher/tests/__init__.py
new file mode 100644
index 00000000000..168f9979a46
--- /dev/null
+++ b/plugins/hydra_rq_launcher/tests/__init__.py
@@ -0,0 +1,1 @@
+# Copyright (c) Facebook, Inc. and its affiliates. All Rights Reserved
diff --git a/... | diff --git a/news/683.plugin b/news/683.plugin
new file mode 100644
index 00000000000..cc5a21c1cca
--- /dev/null
+++ b/news/683.plugin
@@ -0,0 +1,1 @@
+Add [Redis Queue](https://python-rq.org/) Launcher plugin ([Jan-Matthis](https://github.com/jan-matthis))
\ No newline at end of file
diff --git a/plugins/hydra_rq_laun... | [
{
"components": [
{
"doc": "",
"lines": [
12,
15
],
"name": "my_app",
"signature": "def my_app(cfg):",
"type": "function"
}
],
"file": "plugins/hydra_rq_launcher/example/my_app.py"
},
{
"components": [
{
... | [
"plugins/hydra_rq_launcher/tests/test_rq_launcher.py::test_discovery"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
RQ launcher
## Motivation
This PR adds a launcher plugin for [Redis Queue (RQ)](https://python-rq.org):
> RQ (Redis Queue) is a simple Python library for queueing jobs and processing them in the... | 9deaa6cbf46af47355bab1d28b6d4991921de1ee | |
pydata__xarray-4155 | 4,155 | pydata/xarray | 0.12 | a198218ddabe557adbb04311b3234ec8d20419e7 | 2020-06-15T14:42:32Z | diff --git a/doc/whats-new.rst b/doc/whats-new.rst
index 2ad2a426532..7e76066cb33 100644
--- a/doc/whats-new.rst
+++ b/doc/whats-new.rst
@@ -162,9 +162,10 @@ New Features
Enhancements
~~~~~~~~~~~~
- Performance improvement of :py:meth:`DataArray.interp` and :py:func:`Dataset.interp`
- For orthogonal linear- and nea... | diff --git a/xarray/tests/test_dataarray.py b/xarray/tests/test_dataarray.py
index e0da3f1527f..d7e88735fbf 100644
--- a/xarray/tests/test_dataarray.py
+++ b/xarray/tests/test_dataarray.py
@@ -3147,7 +3147,8 @@ def test_upsample_interpolate_regression_1605(self):
@requires_dask
@requires_scipy
- def test... | diff --git a/doc/whats-new.rst b/doc/whats-new.rst
index 2ad2a426532..7e76066cb33 100644
--- a/doc/whats-new.rst
+++ b/doc/whats-new.rst
@@ -162,9 +162,10 @@ New Features
Enhancements
~~~~~~~~~~~~
- Performance improvement of :py:meth:`DataArray.interp` and :py:func:`Dataset.interp`
- For orthogonal linear- and nea... | [
{
"components": [
{
"doc": "Wrapper for `_interpnd` through `blockwise`\n\nThe first half arrays in `coords` are original coordinates,\nthe other half are destination coordinates",
"lines": [
770,
799
],
"name": "_dask_aware_interpnd",
"signature... | [
"xarray/tests/test_interp.py::test_interpolate_1d[1-x-linear]",
"xarray/tests/test_interp.py::test_interpolate_1d[1-x-cubic]",
"xarray/tests/test_interp.py::test_interpolate_1d[1-y-linear]",
"xarray/tests/test_interp.py::test_interpolate_1d[1-y-cubic]",
"xarray/tests/test_interp.py::test_interpolate_vectori... | [
"xarray/tests/test_interp.py::test_keywargs",
"xarray/tests/test_interp.py::test_interpolate_1d[0-x-linear]",
"xarray/tests/test_interp.py::test_interpolate_1d[0-x-cubic]",
"xarray/tests/test_interp.py::test_interpolate_1d[0-y-linear]",
"xarray/tests/test_interp.py::test_interpolate_1d[0-y-cubic]",
"xarra... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implement interp for interpolating between chunks of data (dask)
In a project of mine I need to interpolate a dask-based xarray between chunk of data.
When using the current official `interp` funct... | Here is the discussion in the issues of the pull request.
<issues>
Feature request: Implement interp for interpolating between chunks of data (dask)
In a project of mine I need to interpolate a dask-based xarray between chunk of data.
I made it work using monkey patching. I'm pretty sure that you can write it better... | 1c198a191127c601d091213c4b3292a8bb3054e1 |
Project-MONAI__MONAI-555 | 555 | Project-MONAI/MONAI | null | f03373c0428bddbd9ff132013e9d86cc6952b79a | 2020-06-12T21:56:49Z | diff --git a/monai/transforms/utility/array.py b/monai/transforms/utility/array.py
index dfba0ff582..d533e55be5 100644
--- a/monai/transforms/utility/array.py
+++ b/monai/transforms/utility/array.py
@@ -23,6 +23,18 @@
from monai.transforms.compose import Transform
+class Identity(Transform):
+ """
+ Identity... | diff --git a/tests/test_identity.py b/tests/test_identity.py
new file mode 100644
index 0000000000..7a4b2de291
--- /dev/null
+++ b/tests/test_identity.py
@@ -0,0 +1,28 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in complian... | [
{
"components": [
{
"doc": "Identity transform output is same as the input.",
"lines": [
26,
35
],
"name": "Identity",
"signature": "class Identity(Transform):",
"type": "class"
},
{
"doc": "",
"lines": [
... | [
"tests/test_identity.py::TestIdentity::test_identity",
"tests/test_identityd.py::TestIdentityd::test_identityd"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Identity transform
Add identity transform
### Description
Add identity transform to monai.transforms.utility module. Identity transform's output is same as its input and it can be used as a testin... | e73257caa79309dcce1e93abf1632f4bfd75b11f | ||
sympy__sympy-19539 | 19,539 | sympy/sympy | 1.7 | 822906143dee2c7951835697c463cb40115bf63e | 2020-06-11T20:54:41Z | diff --git a/sympy/vector/__init__.py b/sympy/vector/__init__.py
index 19b52ecd0c06..1353d561b847 100644
--- a/sympy/vector/__init__.py
+++ b/sympy/vector/__init__.py
@@ -15,6 +15,7 @@
SpaceOrienter, QuaternionOrienter)
from sympy.vector.operators import Gradient, Divergence, Curl,... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 131a69f339da..043c1f43998e 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -4856,6 +4856,14 @@ def test_sympy__vector__deloperator__Del():
assert _test_args(Del())
+def test_sympy__vector__integra... | [
{
"components": [
{
"doc": "Represents integral of a scalar or vector field\nover a Parametric Region\n\nExamples\n========\n\n>>> from sympy import cos, sin, pi\n>>> from sympy.vector import CoordSys3D, ParametricRegion, ParametricIntegral\n>>> from sympy.abc import r, t, theta, phi\n\n>>> C = Co... | [
"test_sympy__vector__integrals__ParametricIntegral",
"test_lineintegrals",
"test_surfaceintegrals"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[WIP] Adding ParametricIntegral class
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or ... | 1b4529a95ef641c2fc15889091b281644069d20e | ||
sympy__sympy-19529 | 19,529 | sympy/sympy | 1.7 | a3da22a6f4a48d54585a95d51bc63e87d12db52a | 2020-06-11T07:49:09Z | diff --git a/sympy/stats/__init__.py b/sympy/stats/__init__.py
index b050fc4515b6..7ae96dd005e4 100644
--- a/sympy/stats/__init__.py
+++ b/sympy/stats/__init__.py
@@ -143,6 +143,8 @@
'Probability', 'Expectation', 'Variance', 'Covariance',
+ 'ExpectationMatrix', 'VarianceMatrix', 'CrossCovarianceMatrix'
+
]... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 05c27fdc6980..81b7779a415e 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -1717,6 +1717,22 @@ def test_sympy__stats__random_matrix_models__CircularSymplecticEnsemble():
from sympy.stats import Circul... | [
{
"components": [
{
"doc": "Expectation of a random matrix expression.\n\nExamples\n========\n\n>>> from sympy.stats import ExpectationMatrix, Normal\n>>> from sympy.stats.rv import RandomMatrixSymbol\n>>> from sympy import symbols, MatrixSymbol, Matrix\n>>> k = symbols(\"k\")\n>>> A, B = MatrixSy... | [
"test_sympy__stats__symbolic_multivariate_probability__ExpectationMatrix",
"test_sympy__stats__symbolic_multivariate_probability__VarianceMatrix",
"test_sympy__stats__symbolic_multivariate_probability__CrossCovarianceMatrix",
"test_multivariate_expectation",
"test_multivariate_variance"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[GSoC] Added symbolic Expectation, Variance and CrossCovariance matrix
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
Adde... | 1b4529a95ef641c2fc15889091b281644069d20e | ||
Project-MONAI__MONAI-526 | 526 | Project-MONAI/MONAI | null | edeac8f994bc4ad18b4da936395c8b3bd8ee1edc | 2020-06-10T03:43:54Z | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index b6e0190f51..f4e02b012f 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -97,6 +97,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`ToNumpy`
+~~~~~~~~~
+.. autoclass:: ToNumpy
+ :members:
+... | diff --git a/tests/test_to_numpy.py b/tests/test_to_numpy.py
new file mode 100644
index 0000000000..fa41ac2b4e
--- /dev/null
+++ b/tests/test_to_numpy.py
@@ -0,0 +1,39 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in complian... | diff --git a/docs/source/transforms.rst b/docs/source/transforms.rst
index b6e0190f51..f4e02b012f 100644
--- a/docs/source/transforms.rst
+++ b/docs/source/transforms.rst
@@ -97,6 +97,12 @@ Vanilla Transforms
:members:
:special-members: __call__
+`ToNumpy`
+~~~~~~~~~
+.. autoclass:: ToNumpy
+ :members:
+... | [
{
"components": [
{
"doc": "Converts the input Tensor data to numpy array.",
"lines": [
137,
145
],
"name": "ToNumpy",
"signature": "class ToNumpy(Transform):",
"type": "class"
},
{
"doc": "",
"lines": [
... | [
"tests/test_to_numpy.py::TestToNumpy::test_numpy_input",
"tests/test_to_numpy.py::TestToNumpy::test_tensor_input",
"tests/test_to_numpyd.py::TestToNumpyd::test_numpy_input",
"tests/test_to_numpyd.py::TestToNumpyd::test_tensor_input"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
525 add ToNumpy utility transform
Fixes #525 .
### Description
This PR implemented the ToNumpy transform.
### Status
**Ready**
### Types of changes
<!--- Put an `x` in all the boxes that a... | Here is the discussion in the issues of the pull request.
<issues>
Add ToNumpy transform to support 3rd party transforms
**Is your feature request related to a problem? Please describe.**
As we already have `ToTensor` transform to connect other 3rd party transforms that work for Tensor data only, we also need the `ToN... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
scikit-learn__scikit-learn-17478 | 17,478 | scikit-learn/scikit-learn | 0.24 | e57fab1265969686a3c218c52dce507e7bd9486c | 2020-06-06T17:17:21Z | diff --git a/doc/whats_new/v0.24.rst b/doc/whats_new/v0.24.rst
index 2aebbaed470ab..8a9b081562bd6 100644
--- a/doc/whats_new/v0.24.rst
+++ b/doc/whats_new/v0.24.rst
@@ -91,6 +91,11 @@ Changelog
samples between the train and test set on each fold.
:pr:`13204` by :user:`Kyle Kosic <kykosic>`.
+- |Feature| :class:... | diff --git a/sklearn/model_selection/tests/test_search.py b/sklearn/model_selection/tests/test_search.py
index 8a20872ac762a..6cdc19d378362 100644
--- a/sklearn/model_selection/tests/test_search.py
+++ b/sklearn/model_selection/tests/test_search.py
@@ -56,6 +56,7 @@
from sklearn.tree import DecisionTreeClassifier
fro... | diff --git a/doc/whats_new/v0.24.rst b/doc/whats_new/v0.24.rst
index 2aebbaed470ab..8a9b081562bd6 100644
--- a/doc/whats_new/v0.24.rst
+++ b/doc/whats_new/v0.24.rst
@@ -91,6 +91,11 @@ Changelog
samples between the train and test set on each fold.
:pr:`13204` by :user:`Kyle Kosic <kykosic>`.
+- |Feature| :class:... | [
{
"components": [
{
"doc": "Call score_samples on the estimator with the best found parameters.\n\nOnly available if ``refit=True`` and the underlying estimator supports\n``score_samples``.\n\nParameters\n----------\nX : iterable\n Data to predict on. Must fulfill input requirements\n of the... | [
"sklearn/model_selection/tests/test_search.py::test_search_cv_score_samples_error[search_cv0]",
"sklearn/model_selection/tests/test_search.py::test_search_cv_score_samples_error[search_cv1]",
"sklearn/model_selection/tests/test_search.py::test_search_cv_score_samples_method[search_cv0]",
"sklearn/model_select... | [
"sklearn/model_selection/tests/test_search.py::test_validate_parameter_input[0-TypeError-Parameter",
"sklearn/model_selection/tests/test_search.py::test_validate_parameter_input[input1-TypeError-Parameter",
"sklearn/model_selection/tests/test_search.py::test_validate_parameter_input[input2-TypeError-Parameter.*... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
ENH add score_samples method in base search CV and add tests
#### Reference Issues/PRs
Fixes https://github.com/scikit-learn/scikit-learn/issues/12542.
Building on and closes https://github.com/scik... | 54ce4222694819ad52d544ce5cba5da274c34ab7 | |
sympy__sympy-19500 | 19,500 | sympy/sympy | 1.7 | 25fbcce5b1a4c7e3956e6062930f4a44ce95a632 | 2020-06-06T09:48:50Z | diff --git a/sympy/stats/__init__.py b/sympy/stats/__init__.py
index b050fc4515b6..cc9160adec12 100644
--- a/sympy/stats/__init__.py
+++ b/sympy/stats/__init__.py
@@ -106,7 +106,7 @@
'skewness', 'kurtosis', 'covariance', 'dependent', 'entropy', 'independent',
'random_symbols', 'correlation', 'factorial_moment... | diff --git a/sympy/stats/tests/test_stochastic_process.py b/sympy/stats/tests/test_stochastic_process.py
index ae8800bc0aa5..9ee71deabab0 100644
--- a/sympy/stats/tests/test_stochastic_process.py
+++ b/sympy/stats/tests/test_stochastic_process.py
@@ -3,11 +3,12 @@
Piecewise)
from sympy.stats import... | [
{
"components": [
{
"doc": "This function is used to sample from stochastic process.\n\nParameters\n==========\n\nprocess: StochasticProcess\n Process used to extract the samples. It must be an instance of\n StochasticProcess\n\nExamples\n========\n\n>>> from sympy.stats import sample_stocha... | [
"test_DiscreteMarkovChain",
"test_ContinuousMarkovChain"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[GSoC] Added Sampling of Stochastic Process(DTMC)
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
Added sampling of Stochas... | Here is the discussion in the issues of the pull request.
<issues>
[GSoC Discussion] Sampling from Stochatic Processes
After completing the sampling from the external libraries, we can think of sampling from the stochastic processes. Something similar to the sample path that is followed by the stochastic process, we ca... | 1b4529a95ef641c2fc15889091b281644069d20e | |
Project-MONAI__MONAI-465 | 465 | Project-MONAI/MONAI | null | 718d11abb2310ab74321256032a264488a7883b4 | 2020-06-01T14:18:19Z | diff --git a/docs/source/data.rst b/docs/source/data.rst
index 73a6a698bc..d998605bf8 100644
--- a/docs/source/data.rst
+++ b/docs/source/data.rst
@@ -87,3 +87,7 @@ Utilities
.. automodule:: monai.data.utils
:members:
+
+Decathalon DataLoader
+~~~~~~~~~~~~~~~~~~~~~
+.. autofunction:: monai.data.load_decathalon_da... | diff --git a/tests/test_load_decathalon_datalist.py b/tests/test_load_decathalon_datalist.py
new file mode 100644
index 0000000000..4afe151482
--- /dev/null
+++ b/tests/test_load_decathalon_datalist.py
@@ -0,0 +1,104 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License")... | diff --git a/docs/source/data.rst b/docs/source/data.rst
index 73a6a698bc..d998605bf8 100644
--- a/docs/source/data.rst
+++ b/docs/source/data.rst
@@ -87,3 +87,7 @@ Utilities
.. automodule:: monai.data.utils
:members:
+
+Decathalon DataLoader
+~~~~~~~~~~~~~~~~~~~~~
+.. autofunction:: monai.data.load_decathalon_da... | [
{
"components": [
{
"doc": "",
"lines": [
16,
25
],
"name": "_compute_path",
"signature": "def _compute_path(base_dir, element):",
"type": "function"
},
{
"doc": "",
"lines": [
28,
37
... | [
"tests/test_load_decathalon_datalist.py::TestLoadDecathalonDatalist::test_cls_values",
"tests/test_load_decathalon_datalist.py::TestLoadDecathalonDatalist::test_seg_no_basedir",
"tests/test_load_decathalon_datalist.py::TestLoadDecathalonDatalist::test_seg_no_labels",
"tests/test_load_decathalon_datalist.py::T... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
461 add support to load decathalon datalist
Fixes #461 .
### Description
As Decathalon challenge dataset is very rich and very popular in medical domain, many researchers and students use Decathal... | Here is the discussion in the issues of the pull request.
<issues>
Add utility function to load dataset from JSON file
**Is your feature request related to a problem? Please describe.**
Most of public datasets has a separate file to store `datalist`, let's add a JSON parser first based on the Decathalon dataset format... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
sympy__sympy-19472 | 19,472 | sympy/sympy | 1.7 | eee759c28fa118bbf3fa27382e9c0df93d20fa88 | 2020-06-01T10:28:58Z | diff --git a/sympy/vector/__init__.py b/sympy/vector/__init__.py
index 5b1c03f17eeb..19b52ecd0c06 100644
--- a/sympy/vector/__init__.py
+++ b/sympy/vector/__init__.py
@@ -14,6 +14,7 @@
from sympy.vector.orienters import (AxisOrienter, BodyOrienter,
SpaceOrienter, QuaternionOrienter... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 05c27fdc6980..1d5827c17e63 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -4919,6 +4919,12 @@ def test_sympy__vector__orienters__QuaternionOrienter():
assert _test_args(QuaternionOrienter(a, b, c, d)... | [
{
"components": [
{
"doc": "Represents a parametric region in space.\n\nExamples\n========\n\n>>> from sympy import cos, sin, pi\n>>> from sympy.abc import r, theta, t, a, b, x, y\n>>> from sympy.vector import ParametricRegion\n\n>>> ParametricRegion(t, (t, t**2), limits={t: (-1, 2)})\nParametricR... | [
"test_sympy__vector__parametricregion__ParametricRegion"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implementation of class to represent a parametric region in space
Added an implementation of ParamRegion class.
<!-- Your title above should be a short description of what
was changed. Do not includ... | 1b4529a95ef641c2fc15889091b281644069d20e | ||
sympy__sympy-19463 | 19,463 | sympy/sympy | 1.7 | 4063fa8437fc4319bdbdf1735075a2f9853f4498 | 2020-05-30T05:28:37Z | diff --git a/sympy/core/add.py b/sympy/core/add.py
index a37140b836cc..651f6b627caa 100644
--- a/sympy/core/add.py
+++ b/sympy/core/add.py
@@ -5,7 +5,7 @@
from .parameters import global_parameters
from .logic import _fuzzy_group, fuzzy_or, fuzzy_not
from .singleton import S
-from .operations import AssocOp
+from .op... | diff --git a/sympy/core/tests/test_operations.py b/sympy/core/tests/test_operations.py
index 7555398f4395..6f07446bd17a 100644
--- a/sympy/core/tests/test_operations.py
+++ b/sympy/core/tests/test_operations.py
@@ -1,8 +1,9 @@
-from sympy import Integer, S, symbols, Mul
+from sympy import Integer, S, Symbol, symbols, E... | [
{
"components": [
{
"doc": "",
"lines": [
167,
168
],
"name": "Expr._add_handler",
"signature": "def _add_handler(self):",
"type": "function"
},
{
"doc": "",
"lines": [
171,
172
],
... | [
"test_lattice_simple",
"test_lattice_shortcircuit",
"test_lattice_print",
"test_lattice_make_args",
"test_issue_14025",
"test_AssocOp_flatten",
"test_add_dispatcher",
"test_rational",
"test_large_rational",
"test_negative_real",
"test_expand",
"test_issue_3449",
"test_issue_3866",
"test_ne... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Introduce general add, mul, and pow function
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Iss... | Here is the discussion in the issues of the pull request.
<issues>
Add Basic._exec_constructor_hook
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or PRs
<!-- If this pull request fixes an issue, write "Fixe... | 1b4529a95ef641c2fc15889091b281644069d20e | |
huggingface__datasets-214 | 214 | huggingface/datasets | null | f9ef1ee270848e322f06e546751aeffde67349c5 | 2020-05-28T16:21:40Z | diff --git a/datasets/wikipedia/wikipedia.py b/datasets/wikipedia/wikipedia.py
index 5733781ccf8..6f36e2bc534 100644
--- a/datasets/wikipedia/wikipedia.py
+++ b/datasets/wikipedia/wikipedia.py
@@ -25,9 +25,9 @@
import xml.etree.cElementTree as etree
import apache_beam as beam
-import mwparserfromhell
import six
... | diff --git a/tests/test_arrow_dataset.py b/tests/test_arrow_dataset.py
new file mode 100644
index 00000000000..88142afd586
--- /dev/null
+++ b/tests/test_arrow_dataset.py
@@ -0,0 +1,114 @@
+import os
+import tempfile
+from unittest import TestCase
+
+import numpy as np
+import pyarrow as pa
+
+from nlp.arrow_reader imp... | [
{
"components": [
{
"doc": "Apply a filter function to all the elements in the table in batches\nand update the table so that the dataset only includes examples according to the filter function.\n\nArgs:\n `function` (`callable`): with one of the following signature:\n - `function(exampl... | [
"tests/test_arrow_dataset.py::BaseDatasetTest::test_filter"
] | [
"tests/test_arrow_dataset.py::BaseDatasetTest::test_map",
"tests/test_arrow_dataset.py::BaseDatasetTest::test_map_batched",
"tests/test_arrow_dataset.py::BaseDatasetTest::test_remove_colums"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[arrow_dataset.py] add new filter function
The `.map()` function is super useful, but can IMO a bit tedious when filtering certain examples.
I think, filtering out examples is also a very common oper... | 5142a8cf61d8a4495eda3d91dc4283a6df01ea14 | ||
Project-MONAI__MONAI-401 | 401 | Project-MONAI/MONAI | null | dc7cd0ec25d4b27f321a31f13e707769922c66b3 | 2020-05-19T12:12:37Z | diff --git a/monai/handlers/__init__.py b/monai/handlers/__init__.py
index 9c6491478d..7977e4e49a 100644
--- a/monai/handlers/__init__.py
+++ b/monai/handlers/__init__.py
@@ -17,4 +17,5 @@
from .segmentation_saver import SegmentationSaver
from .stats_handler import StatsHandler
from .tensorboard_handlers import Tens... | diff --git a/tests/test_handler_lr_scheduler.py b/tests/test_handler_lr_scheduler.py
new file mode 100644
index 0000000000..4f18500587
--- /dev/null
+++ b/tests/test_handler_lr_scheduler.py
@@ -0,0 +1,63 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may ... | [
{
"components": [
{
"doc": "ignite handler to update the Learning Rate based on PyTorch LR scheduler.",
"lines": [
18,
56
],
"name": "LrScheduleHandler",
"signature": "class LrScheduleHandler:",
"type": "class"
},
{
"d... | [
"tests/test_handler_lr_scheduler.py::TestHandlerLrSchedule::test_content"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
400 add LrScheduleHandler for engines
Fixes #400 .
### Description
This PR implemented the LrScheduleHandler to adjust the learning rate during training.
### Status
**Ready**
### Types of c... | Here is the discussion in the issues of the pull request.
<issues>
Add LrScheduleHandler for engines
**Is your feature request related to a problem? Please describe.**
When developing a training program based on ignite, we need LrScheduleHandler to update the learning rate when "epoch completed".
----------
----------... | e73257caa79309dcce1e93abf1632f4bfd75b11f | |
sympy__sympy-19355 | 19,355 | sympy/sympy | 1.7 | 94fb720696f5f5d12bad8bc813699fd696afd2fb | 2020-05-18T09:55:36Z | diff --git a/sympy/matrices/eigen.py b/sympy/matrices/eigen.py
index b9e63b8ae9d6..359009f5d784 100644
--- a/sympy/matrices/eigen.py
+++ b/sympy/matrices/eigen.py
@@ -76,7 +76,9 @@ def _eigenvects_mpmath(M):
# This functions is a candidate for caching if it gets implemented for matrices.
-def _eigenvals(M, error_w... | diff --git a/sympy/matrices/tests/test_eigen.py b/sympy/matrices/tests/test_eigen.py
index 1a422c01f175..2e8b2c861916 100644
--- a/sympy/matrices/tests/test_eigen.py
+++ b/sympy/matrices/tests/test_eigen.py
@@ -2,8 +2,6 @@
Rational, Symbol, N, I, Abs, sqrt, exp, Float, sin,
cos, symbols)
from sympy.matrices ... | [
{
"components": [
{
"doc": "",
"lines": [
179,
205
],
"name": "_eigenvals_list",
"signature": "def _eigenvals_list( M, error_when_incomplete=True, simplify=False, **flags):",
"type": "function"
},
{
"doc": "",
... | [
"test_eigen"
] | [
"test_float_eigenvals",
"test_issue_8240",
"test_eigenvals",
"test_eigenvects",
"test_left_eigenvects",
"test_diagonalize",
"test_is_diagonalizable",
"test_jordan_form",
"test_singular_values",
"test___eq__"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Apply connected components decomposition for computing eigenvalues
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### R... | 1b4529a95ef641c2fc15889091b281644069d20e | ||
sympy__sympy-19339 | 19,339 | sympy/sympy | 1.7 | 94fb720696f5f5d12bad8bc813699fd696afd2fb | 2020-05-17T05:46:37Z | diff --git a/doc/src/modules/matrices/expressions.rst b/doc/src/modules/matrices/expressions.rst
index 03e212b8cc1f..63224873befd 100644
--- a/doc/src/modules/matrices/expressions.rst
+++ b/doc/src/modules/matrices/expressions.rst
@@ -54,6 +54,8 @@ Matrix Expressions Core Reference
:members:
.. autoclass:: ZeroMat... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 9758cde7cc51..05c27fdc6980 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -3099,6 +3099,15 @@ def test_sympy__matrices__expressions__permutation__MatrixPermute():
A = MatrixSymbol('A', 3, 3)
ass... | diff --git a/doc/src/modules/matrices/expressions.rst b/doc/src/modules/matrices/expressions.rst
index 03e212b8cc1f..63224873befd 100644
--- a/doc/src/modules/matrices/expressions.rst
+++ b/doc/src/modules/matrices/expressions.rst
@@ -54,6 +54,8 @@ Matrix Expressions Core Reference
:members:
.. autoclass:: ZeroMat... | [
{
"components": [
{
"doc": "Returns a companion matrix of a polynomial.\n\nExamples\n========\n\n>>> from sympy import Matrix, Poly, Symbol, symbols\n>>> x = Symbol('x')\n>>> c0, c1, c2, c3, c4 = symbols('c0:5')\n>>> p = Poly(c0 + c1*x + c2*x**2 + c3*x**3 + c4*x**4 + x**5, x)\n>>> Matrix.companion... | [
"test_sympy__matrices__expressions__companion__CompanionMatrix",
"test_creation",
"test_entry"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add companion matrix
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or PRs
<!-- If this... | Here is the discussion in the issues of the pull request.
<issues>
Add companion matrix of a polynomial
There should be a convenient way to get the companion matrix of a polynomial:
https://en.wikipedia.org/wiki/Companion_matrix
This is in some sense the reverse of getting the characteristic polynomial of a matrix.... | 1b4529a95ef641c2fc15889091b281644069d20e |
sphinx-doc__sphinx-7670 | 7,670 | sphinx-doc/sphinx | 3.1 | 3419079fb0d1f0eecd845eff0d12b367d34cd5e9 | 2020-05-16T07:58:26Z | diff --git a/CHANGES b/CHANGES
index b73d9f0e140..ae466b9e633 100644
--- a/CHANGES
+++ b/CHANGES
@@ -84,6 +84,7 @@ Features added
* #7734: napoleon: overescaped trailing underscore on attribute
* #7683: Add ``allowed_exceptions`` parameter to ``Sphinx.emit()`` to allow
handlers to raise specified exceptions
+* #72... | diff --git a/tests/test_domain_cpp.py b/tests/test_domain_cpp.py
index 116acecfb17..961646131b8 100644
--- a/tests/test_domain_cpp.py
+++ b/tests/test_domain_cpp.py
@@ -828,6 +828,15 @@ def test_templates():
check('type', 'template<C T = int&> {key}A', {2: 'I_1CE1A'}, key='using')
+def test_requires_clauses():... | diff --git a/CHANGES b/CHANGES
index b73d9f0e140..ae466b9e633 100644
--- a/CHANGES
+++ b/CHANGES
@@ -84,6 +84,7 @@ Features added
* #7734: napoleon: overescaped trailing underscore on attribute
* #7683: Add ``allowed_exceptions`` parameter to ``Sphinx.emit()`` to allow
handlers to raise specified exceptions
+* #72... | [
{
"components": [
{
"doc": "",
"lines": [
3571,
3581
],
"name": "ASTRequiresClause",
"signature": "class ASTRequiresClause(ASTBase):",
"type": "class"
},
{
"doc": "",
"lines": [
3572,
3573
... | [
"tests/test_domain_cpp.py::test_requires_clauses"
] | [
"tests/test_domain_cpp.py::test_fundamental_types",
"tests/test_domain_cpp.py::test_expressions",
"tests/test_domain_cpp.py::test_type_definitions",
"tests/test_domain_cpp.py::test_concept_definitions",
"tests/test_domain_cpp.py::test_member_definitions",
"tests/test_domain_cpp.py::test_function_definitio... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
C++, parse (trailing) requires clauses
### Feature or Bugfix
- Feature
### Purpose
C++20 allows explicit constraints on function templates with requires clauses. They can either be just after the... | Here is the discussion in the issues of the pull request.
<issues>
C++20 requires clause not supported
Could you please add the support for C++ [requires clauses](https://en.cppreference.com/w/cpp/language/constraints)?
I am the author of [mp-units](https://github.com/mpusz/units) which is a Physical Units Library t... | 3419079fb0d1f0eecd845eff0d12b367d34cd5e9 |
joke2k__faker-1180 | 1,180 | joke2k/faker | null | 81a318cdcfc5bdf36f83e3db91956e30d303971c | 2020-05-14T20:16:07Z | diff --git a/faker/providers/currency/__init__.py b/faker/providers/currency/__init__.py
index f6842e0e71..bce353ffa9 100644
--- a/faker/providers/currency/__init__.py
+++ b/faker/providers/currency/__init__.py
@@ -237,6 +237,8 @@ class Provider(BaseProvider):
'XCD': '\u0024', 'YER': '\uFDFC', 'ZWD': '\u0024',... | diff --git a/tests/providers/test_currency.py b/tests/providers/test_currency.py
index 81bd2220c7..64b14bcf54 100644
--- a/tests/providers/test_currency.py
+++ b/tests/providers/test_currency.py
@@ -67,6 +67,11 @@ def test_cryptocurrency_name(self, faker, num_samples):
name = faker.cryptocurrency_name()
... | [
{
"components": [
{
"doc": "",
"lines": [
268,
272
],
"name": "Provider.pricetag",
"signature": "def pricetag(self):",
"type": "function"
}
],
"file": "faker/providers/currency/__init__.py"
},
{
"components": [
... | [
"tests/providers/test_currency.py::TestCurrencyProvider::test_pricetag",
"tests/providers/test_currency.py::TestRuRu::test_pricetag",
"tests/providers/test_currency.py::TestCsCz::test_pricetag",
"tests/providers/test_currency.py::TestDeAt::test_pricetag",
"tests/providers/test_currency.py::TestDeDe::test_pr... | [
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency",
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency_code",
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency_name",
"tests/providers/test_currency.py::TestCurrencyProvider::test_currency_symbol_no_... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Create new currency provider pricetag()
This is a new currency provider `pricetag()` that creates a random price in the respective local currency using the format for thousands/decimals.
In the cur... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
scikit-learn__scikit-learn-17225 | 17,225 | scikit-learn/scikit-learn | 0.24 | 2f26540ee99cb4519d7471933359913c7be36ac9 | 2020-05-14T18:04:04Z | diff --git a/doc/whats_new/v0.24.rst b/doc/whats_new/v0.24.rst
index db6959fcc164f..76ec91d93e264 100644
--- a/doc/whats_new/v0.24.rst
+++ b/doc/whats_new/v0.24.rst
@@ -76,7 +76,13 @@ Changelog
attribute name/path or a `callable` for extracting feature importance from
the estimator. :pr:`15361` by :user:`Venkata... | diff --git a/sklearn/ensemble/tests/test_gradient_boosting_loss_functions.py b/sklearn/ensemble/tests/test_gradient_boosting_loss_functions.py
index 6b24f90d0239d..b5bc17eeeb14c 100644
--- a/sklearn/ensemble/tests/test_gradient_boosting_loss_functions.py
+++ b/sklearn/ensemble/tests/test_gradient_boosting_loss_function... | diff --git a/doc/whats_new/v0.24.rst b/doc/whats_new/v0.24.rst
index db6959fcc164f..76ec91d93e264 100644
--- a/doc/whats_new/v0.24.rst
+++ b/doc/whats_new/v0.24.rst
@@ -76,7 +76,13 @@ Changelog
attribute name/path or a `callable` for extracting feature importance from
the estimator. :pr:`15361` by :user:`Venkata... | [
{
"components": [
{
"doc": "Implements a simplified version of np.take_along_axis if numpy\nversion < 1.15",
"lines": [
170,
203
],
"name": "_take_along_axis",
"signature": "def _take_along_axis(arr, indices, axis):",
"type": "function"
... | [
"sklearn/utils/tests/test_stats.py::test_weighted_percentile_2d"
] | [
"sklearn/ensemble/tests/test_gradient_boosting_loss_functions.py::test_binomial_deviance",
"sklearn/ensemble/tests/test_gradient_boosting_loss_functions.py::test_sample_weight_smoke",
"sklearn/ensemble/tests/test_gradient_boosting_loss_functions.py::test_sample_weight_init_estimators",
"sklearn/ensemble/tests... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
ENH Sample weights for median_absolute_error
<!--
Thanks for contributing a pull request! Please ensure you have taken a look at
the contribution guidelines: https://github.com/scikit-learn/scikit-l... | 54ce4222694819ad52d544ce5cba5da274c34ab7 | |
slackapi__python-slack-sdk-686 | 686 | slackapi/python-slack-sdk | null | 58134fefafcc1287bc156cc74da58495db88e0d4 | 2020-05-14T05:33:45Z | diff --git a/slack/signature/__init__.py b/slack/signature/__init__.py
new file mode 100644
index 000000000..21b38c0fc
--- /dev/null
+++ b/slack/signature/__init__.py
@@ -0,0 +1,1 @@
+from .verifier import SignatureVerifier # noqa
diff --git a/slack/signature/verifier.py b/slack/signature/verifier.py
new file mode 100... | diff --git a/tests/signature/test_signature_verifier.py b/tests/signature/test_signature_verifier.py
new file mode 100644
index 000000000..8015733e6
--- /dev/null
+++ b/tests/signature/test_signature_verifier.py
@@ -0,0 +1,86 @@
+import unittest
+
+from slack.signature import SignatureVerifier
+
+
+class MockClock:
+ ... | [
{
"components": [
{
"doc": "",
"lines": [
7,
9
],
"name": "Clock",
"signature": "class Clock:",
"type": "class"
},
{
"doc": "",
"lines": [
8,
9
],
"name": "Clock.now",
... | [
"tests/signature/test_signature_verifier.py::TestSignatureVerifier::test_generate_signature",
"tests/signature/test_signature_verifier.py::TestSignatureVerifier::test_is_valid",
"tests/signature/test_signature_verifier.py::TestSignatureVerifier::test_is_valid_none",
"tests/signature/test_signature_verifier.py... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add SignatureVerifier for request verification
### Summary
This pull request adds the official implementation of Slack request verification. We have been having this logic as a static method in `B... | 2997a1786c4fd969b00ce69af888ebae8e8ebed0 | ||
scikit-learn__scikit-learn-17169 | 17,169 | scikit-learn/scikit-learn | 1.0 | 579e7de7f38f9f514ff2b2be049e67b14e723d17 | 2020-05-09T14:08:48Z | diff --git a/doc/modules/classes.rst b/doc/modules/classes.rst
index c658bc6b12452..d56914f874b42 100644
--- a/doc/modules/classes.rst
+++ b/doc/modules/classes.rst
@@ -560,6 +560,7 @@ From text
feature_selection.chi2
feature_selection.f_classif
feature_selection.f_regression
+ feature_selection.r_regress... | diff --git a/sklearn/feature_selection/tests/test_feature_select.py b/sklearn/feature_selection/tests/test_feature_select.py
index 61f709094147e..852c8228b2a76 100644
--- a/sklearn/feature_selection/tests/test_feature_select.py
+++ b/sklearn/feature_selection/tests/test_feature_select.py
@@ -4,11 +4,12 @@
import itert... | diff --git a/doc/modules/classes.rst b/doc/modules/classes.rst
index c658bc6b12452..d56914f874b42 100644
--- a/doc/modules/classes.rst
+++ b/doc/modules/classes.rst
@@ -560,6 +560,7 @@ From text
feature_selection.chi2
feature_selection.f_classif
feature_selection.f_regression
+ feature_selection.r_regress... | [
{
"components": [
{
"doc": "Compute Pearson's r for each features and the target.\n\nPearson's r is also known as the Pearson correlation coefficient.\n\n.. versionadded:: 1.0\n\nLinear model for testing the individual effect of each of many regressors.\nThis is a scoring function to be used in a ... | [
"sklearn/feature_selection/tests/test_feature_select.py::test_f_oneway_vs_scipy_stats",
"sklearn/feature_selection/tests/test_feature_select.py::test_f_oneway_ints",
"sklearn/feature_selection/tests/test_feature_select.py::test_f_classif",
"sklearn/feature_selection/tests/test_feature_select.py::test_r_regres... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[MRG] Refactor `feature_selection.f_regression` and introduce `feature_selection.r_regression`
<!--
Thanks for contributing a pull request! Please ensure you have taken a look at
the contribution gu... | Here is the discussion in the issues of the pull request.
<issues>
[refactor] f_regression into r_regression and a wrapper for performance
#### Description
I suggest to refactor `f_regression` into two separate functions: first computing Pearson R `r_regression` and the second computing f-statistic and p-value (I gues... | 3c732b9f6a77e95dfa6beb154ca2e1e7848b74f9 |
matplotlib__matplotlib-17371 | 17,371 | matplotlib/matplotlib | 3.2 | 52761deb15b9c0aca05112befcdefce5cea0a2ad | 2020-05-09T03:06:10Z | diff --git a/doc/api/next_api_changes/behavior/17371-IHI.rst b/doc/api/next_api_changes/behavior/17371-IHI.rst
new file mode 100644
index 000000000000..34fa9e03ae85
--- /dev/null
+++ b/doc/api/next_api_changes/behavior/17371-IHI.rst
@@ -0,0 +1,7 @@
+ioff and ion can be used as context managers
+~~~~~~~~~~~~~~~~~~~~~~~~... | diff --git a/lib/matplotlib/tests/test_pyplot.py b/lib/matplotlib/tests/test_pyplot.py
index 9b9873c3cb4a..b52483ea7937 100644
--- a/lib/matplotlib/tests/test_pyplot.py
+++ b/lib/matplotlib/tests/test_pyplot.py
@@ -81,3 +81,75 @@ def test_nrows_error():
plt.subplot(nrows=1)
with pytest.raises(TypeError):
... | diff --git a/doc/api/next_api_changes/behavior/17371-IHI.rst b/doc/api/next_api_changes/behavior/17371-IHI.rst
new file mode 100644
index 000000000000..34fa9e03ae85
--- /dev/null
+++ b/doc/api/next_api_changes/behavior/17371-IHI.rst
@@ -0,0 +1,7 @@
+ioff and ion can be used as context managers
+~~~~~~~~~~~~~~~~~~~~~~~~... | [
{
"components": [
{
"doc": "Context manager for `.ioff`.\n\nThe state is changed in ``__init__()`` instead of ``__enter__()``. The\nlatter is a no-op. This allows using `.ioff` both as a function and\nas a context.",
"lines": [
367,
390
],
"name": "_Ioff... | [
"lib/matplotlib/tests/test_pyplot.py::test_ioff",
"lib/matplotlib/tests/test_pyplot.py::test_ion",
"lib/matplotlib/tests/test_pyplot.py::test_nested_ion_ioff"
] | [
"lib/matplotlib/tests/test_pyplot.py::test_pyplot_up_to_date",
"lib/matplotlib/tests/test_pyplot.py::test_copy_docstring_and_deprecators",
"lib/matplotlib/tests/test_pyplot.py::test_pyplot_box",
"lib/matplotlib/tests/test_pyplot.py::test_stackplot_smoke",
"lib/matplotlib/tests/test_pyplot.py::test_nrows_err... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
add context manager functionality to ion and ioff
## PR Summary
Closes: https://github.com/matplotlib/matplotlib/issues/17013
Also closes: https://github.com/matplotlib/ipympl/issues/220
This all... | ff821ba3249401fe7f5fdb11cb7ac5d0564c2697 | |
sphinx-doc__sphinx-7597 | 7,597 | sphinx-doc/sphinx | 3.1 | c13ecd243709d1e210a030be5aa09b7714e35730 | 2020-05-02T13:44:52Z | diff --git a/CHANGES b/CHANGES
index 3dcbe2b1f75..d426fa48c87 100644
--- a/CHANGES
+++ b/CHANGES
@@ -70,6 +70,7 @@ Features added
* C++, parse trailing return types.
* #7143: py domain: Add ``:final:`` option to :rst:dir:`py:class:`,
:rst:dir:`py:exception:` and :rst:dir:`py:method:` directives
+* #7596: py domain... | diff --git a/tests/test_domain_py.py b/tests/test_domain_py.py
index 5a1d73cfe66..08b3da21e6e 100644
--- a/tests/test_domain_py.py
+++ b/tests/test_domain_py.py
@@ -420,7 +420,8 @@ def test_pydata_signature(app):
doctree = restructuredtext.parse(app, text)
assert_node(doctree, (addnodes.index,
... | diff --git a/CHANGES b/CHANGES
index 3dcbe2b1f75..d426fa48c87 100644
--- a/CHANGES
+++ b/CHANGES
@@ -70,6 +70,7 @@ Features added
* C++, parse trailing return types.
* #7143: py domain: Add ``:final:`` option to :rst:dir:`py:class:`,
:rst:dir:`py:exception:` and :rst:dir:`py:method:` directives
+* #7596: py domain... | [
{
"components": [
{
"doc": "Convert a type string to a cross reference node.",
"lines": [
80,
88
],
"name": "type_to_xref",
"signature": "def type_to_xref(text: str) -> addnodes.pending_xref:",
"type": "function"
}
],
"file"... | [
"tests/test_domain_py.py::test_pydata_signature",
"tests/test_domain_py.py::test_pyattribute"
] | [
"tests/test_domain_py.py::test_function_signatures",
"tests/test_domain_py.py::test_domain_py_xrefs",
"tests/test_domain_py.py::test_domain_py_objects",
"tests/test_domain_py.py::test_resolve_xref_for_properties",
"tests/test_domain_py.py::test_domain_py_find_obj",
"tests/test_domain_py.py::test_get_full_... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Close #7596: py domain: Change a type annotation for variables to a hyperlink
### Feature or Bugfix
- Feature
### Purpose
- refs: #7596
----------
</request>
There are several new functions or... | Here is the discussion in the issues of the pull request.
<issues>
py domain: Change a type annotation for variables to a hyperlink
**Is your feature request related to a problem? Please describe.**
py domain: Change a type annotation for variables to a hyperlink
**Describe the solution you'd like**
`type` optio... | 3419079fb0d1f0eecd845eff0d12b367d34cd5e9 |
sphinx-doc__sphinx-7590 | 7,590 | sphinx-doc/sphinx | 3.1 | 2e506c5ab457cba743bb47eb5b8c8eb9dd51d23d | 2020-05-01T18:29:11Z | diff --git a/CHANGES b/CHANGES
index 3942c4240ae..33c641fc2b2 100644
--- a/CHANGES
+++ b/CHANGES
@@ -63,6 +63,7 @@ Features added
* #7543: html theme: Add top and bottom margins to tables
* C and C++: allow semicolon in the end of declarations.
* C++, parse parameterized noexcept specifiers.
+* #7294: C++, parse exp... | diff --git a/tests/test_domain_cpp.py b/tests/test_domain_cpp.py
index 9db741ae5d9..84bd4571894 100644
--- a/tests/test_domain_cpp.py
+++ b/tests/test_domain_cpp.py
@@ -146,37 +146,48 @@ class Config:
exprCheck(expr, 'L' + expr + 'E')
expr = i + l + u
exprCheck(expr, '... | diff --git a/CHANGES b/CHANGES
index 3942c4240ae..33c641fc2b2 100644
--- a/CHANGES
+++ b/CHANGES
@@ -63,6 +63,7 @@ Features added
* #7543: html theme: Add top and bottom margins to tables
* C and C++: allow semicolon in the end of declarations.
* C++, parse parameterized noexcept specifiers.
+* #7294: C++, parse exp... | [
{
"components": [
{
"doc": "",
"lines": [
903,
918
],
"name": "ASTUserDefinedLiteral",
"signature": "class ASTUserDefinedLiteral(ASTLiteral):",
"type": "class"
},
{
"doc": "",
"lines": [
904,
... | [
"tests/test_domain_cpp.py::test_expressions"
] | [
"tests/test_domain_cpp.py::test_fundamental_types",
"tests/test_domain_cpp.py::test_type_definitions",
"tests/test_domain_cpp.py::test_concept_definitions",
"tests/test_domain_cpp.py::test_member_definitions",
"tests/test_domain_cpp.py::test_function_definitions",
"tests/test_domain_cpp.py::test_operators... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
C++, parse expressions with user-defined literals
### Feature or Bugfix
- Feature
- Bugfix
### Relates
Fixes #7294
----------
</request>
There are several new functions or classes that nee... | Here is the discussion in the issues of the pull request.
<issues>
C++ User Defined Literals not supported
The code as below
```cpp
namespace units::si {
inline constexpr auto planck_constant = 6.62607015e-34q_J * 1q_s;
}
```
causes the following error:
```
WARNING: Invalid definition: Expected end of... | 3419079fb0d1f0eecd845eff0d12b367d34cd5e9 |
Project-MONAI__MONAI-325 | 325 | Project-MONAI/MONAI | null | 57abc3b502684b5145200d1d385cf9f02521b347 | 2020-04-29T11:01:32Z | diff --git a/docs/source/losses.rst b/docs/source/losses.rst
index c5be59aa38..17ce6d8b68 100644
--- a/docs/source/losses.rst
+++ b/docs/source/losses.rst
@@ -22,6 +22,10 @@ Segmentation Losses
.. autoclass:: GeneralizedDiceLoss
:members:
+`FocalLoss`
+~~~~~~~~~~~
+.. autoclass:: monai.losses.focal_loss.FocalLo... | diff --git a/tests/test_focal_loss.py b/tests/test_focal_loss.py
new file mode 100644
index 0000000000..1a51b9ca2a
--- /dev/null
+++ b/tests/test_focal_loss.py
@@ -0,0 +1,218 @@
+# Copyright 2020 MONAI Consortium
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in c... | diff --git a/docs/source/losses.rst b/docs/source/losses.rst
index c5be59aa38..17ce6d8b68 100644
--- a/docs/source/losses.rst
+++ b/docs/source/losses.rst
@@ -22,6 +22,10 @@ Segmentation Losses
.. autoclass:: GeneralizedDiceLoss
:members:
+`FocalLoss`
+~~~~~~~~~~~
+.. autoclass:: monai.losses.focal_loss.FocalLo... | [
{
"components": [
{
"doc": "PyTorch implementation of the Focal Loss.\n[1] \"Focal Loss for Dense Object Detection\", T. Lin et al., ICCV 2017",
"lines": [
17,
98
],
"name": "FocalLoss",
"signature": "class FocalLoss(_WeightedLoss):",
"ty... | [
"tests/test_focal_loss.py::TestFocalLoss::test_bin_seg_2d",
"tests/test_focal_loss.py::TestFocalLoss::test_bin_seg_3d",
"tests/test_focal_loss.py::TestFocalLoss::test_consistency_with_cross_entropy_2d",
"tests/test_focal_loss.py::TestFocalLoss::test_consistency_with_cross_entropy_classification",
"tests/tes... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Focal loss
Fixes #115 Port focal loss .
### Description
Add an implementation of the Focal loss.
### Status
**Ready**
### Types of changes
- [x] New functionality added
- [x] Added unit t... | Here is the discussion in the issues of the pull request.
<issues>
Port focal loss
a subtask of #84, port a focal loss function into Monai
e.g.:
https://arxiv.org/abs/1708.02002
----------
Hi, I don't know what is the current state of this, but if it helps I have already an implementation of the Focal loss in PyTo... | e73257caa79309dcce1e93abf1632f4bfd75b11f |
scikit-learn__scikit-learn-17036 | 17,036 | scikit-learn/scikit-learn | 1.0 | 03245ee3afe5ee9e2ff626e2290f02748d95e497 | 2020-04-25T14:56:29Z | diff --git a/doc/modules/classes.rst b/doc/modules/classes.rst
index 3edd8adee8191..d56c7b5d8eafe 100644
--- a/doc/modules/classes.rst
+++ b/doc/modules/classes.rst
@@ -994,6 +994,7 @@ details.
metrics.mean_poisson_deviance
metrics.mean_gamma_deviance
metrics.mean_tweedie_deviance
+ metrics.d2_tweedie_sco... | diff --git a/sklearn/metrics/tests/test_common.py b/sklearn/metrics/tests/test_common.py
index 939371b01fc27..47e6bec38388f 100644
--- a/sklearn/metrics/tests/test_common.py
+++ b/sklearn/metrics/tests/test_common.py
@@ -29,6 +29,7 @@
from sklearn.metrics import cohen_kappa_score
from sklearn.metrics import confusion... | diff --git a/doc/modules/classes.rst b/doc/modules/classes.rst
index 3edd8adee8191..d56c7b5d8eafe 100644
--- a/doc/modules/classes.rst
+++ b/doc/modules/classes.rst
@@ -994,6 +994,7 @@ details.
metrics.mean_poisson_deviance
metrics.mean_gamma_deviance
metrics.mean_tweedie_deviance
+ metrics.d2_tweedie_sco... | [
{
"components": [
{
"doc": "D^2 regression score function, percentage of Tweedie deviance explained.\n\nBest possible score is 1.0 and it can be negative (because the model can be\narbitrarily worse). A model that always uses the empirical mean of `y_true` as\nconstant prediction, disregarding the... | [
"sklearn/metrics/tests/test_common.py::test_symmetry_consistency",
"sklearn/metrics/tests/test_common.py::test_symmetric_metric[accuracy_score]",
"sklearn/metrics/tests/test_common.py::test_symmetric_metric[cohen_kappa_score]",
"sklearn/metrics/tests/test_common.py::test_symmetric_metric[f1_score]",
"sklear... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
FEA add d2_tweedie_score
#### Reference Issues/PRs
Resolves #15244.
#### What does this implement/fix? Explain your changes.
Add `d2_tweedie_score` metric.
#### Open questions
- For `mean... | Here is the discussion in the issues of the pull request.
<issues>
Add d2_tweedie_score
As discussed in https://github.com/scikit-learn/scikit-learn/pull/14300#discussion_r329370799 it might be useful to add the `d2_deviance_score` that is a generalization of `r2_score` to non normal error distributions. In the same wa... | 3c732b9f6a77e95dfa6beb154ca2e1e7848b74f9 |
matplotlib__matplotlib-17233 | 17,233 | matplotlib/matplotlib | 3.2 | 4356b677a33430b5c9e34e8b18e07cfd630a0704 | 2020-04-24T05:06:24Z | diff --git a/doc/api/api_changes_3.3/deprecations.rst b/doc/api/api_changes_3.3/deprecations.rst
index 70595495a4a0..f80732851d15 100644
--- a/doc/api/api_changes_3.3/deprecations.rst
+++ b/doc/api/api_changes_3.3/deprecations.rst
@@ -593,3 +593,11 @@ APIs which support the values True, False, and "TeX" for ``ismath``.... | diff --git a/lib/matplotlib/tests/test_backend_pdf.py b/lib/matplotlib/tests/test_backend_pdf.py
index 92fc22bdc7e3..4e125992138a 100644
--- a/lib/matplotlib/tests/test_backend_pdf.py
+++ b/lib/matplotlib/tests/test_backend_pdf.py
@@ -1,3 +1,4 @@
+import datetime
import io
import os
from pathlib import Path
@@ -7,6 ... | diff --git a/doc/api/api_changes_3.3/deprecations.rst b/doc/api/api_changes_3.3/deprecations.rst
index 70595495a4a0..f80732851d15 100644
--- a/doc/api/api_changes_3.3/deprecations.rst
+++ b/doc/api/api_changes_3.3/deprecations.rst
@@ -593,3 +593,11 @@ APIs which support the values True, False, and "TeX" for ``ismath``.... | [
{
"components": [
{
"doc": "Create a PDF infoDict based on user-supplied metadata.\n\nA default ``Creator``, ``Producer``, and ``CreationDate`` are added, though\nthe user metadata may override it. The date may be the current time, or a\ntime set by the ``SOURCE_DATE_EPOCH`` environment variable.\... | [
"lib/matplotlib/tests/test_backend_pdf.py::test_savefig_metadata",
"lib/matplotlib/tests/test_backend_pdf.py::test_multipage_metadata"
] | [
"lib/matplotlib/tests/test_backend_pdf.py::test_type42",
"lib/matplotlib/tests/test_backend_pdf.py::test_multipage_pagecount",
"lib/matplotlib/tests/test_backend_pdf.py::test_multipage_properfinalize",
"lib/matplotlib/tests/test_backend_pdf.py::test_multipage_keep_empty",
"lib/matplotlib/tests/test_backend_... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Improve PDF metadata support in PGF
## PR Summary
The PGF backend only supported PDF metadata using the multi-page `PdfPages` class; this adds support for adding metadata to single-figure saving di... | ff821ba3249401fe7f5fdb11cb7ac5d0564c2697 | |
rytilahti__python-miio-675 | 675 | rytilahti/python-miio | null | 8f16c1b335599e1d02f8217819687843bb687dc8 | 2020-04-19T01:30:20Z | diff --git a/docs/vacuum.rst b/docs/vacuum.rst
index 32f17d709..cef86ad64 100644
--- a/docs/vacuum.rst
+++ b/docs/vacuum.rst
@@ -30,9 +30,7 @@ Status reporting
Fanspeed: 60
Cleaning since: 0:00:00
Cleaned area: 0.0 m²
- DND enabled: 0
- Map present: 1
- in_cleaning: 0
+ Water box attached: Fa... | diff --git a/miio/tests/test_vacuum.py b/miio/tests/test_vacuum.py
index 8c628fa41..72edfcfff 100644
--- a/miio/tests/test_vacuum.py
+++ b/miio/tests/test_vacuum.py
@@ -33,6 +33,7 @@ def __init__(self, *args, **kwargs):
"battery": 100,
"fan_power": 20,
"msg_seq": 320,
+ ... | diff --git a/docs/vacuum.rst b/docs/vacuum.rst
index 32f17d709..cef86ad64 100644
--- a/docs/vacuum.rst
+++ b/docs/vacuum.rst
@@ -30,9 +30,7 @@ Status reporting
Fanspeed: 60
Cleaning since: 0:00:00
Cleaned area: 0.0 m²
- DND enabled: 0
- Map present: 1
- in_cleaning: 0
+ Water box attached: Fa... | [
{
"components": [
{
"doc": "Return True is water box is installed.",
"lines": [
171,
173
],
"name": "VacuumStatus.is_water_box_attached",
"signature": "def is_water_box_attached(self) -> bool:",
"type": "function"
}
],
"file... | [
"miio/tests/test_vacuum.py::TestVacuum::test_status"
] | [
"miio/tests/test_vacuum.py::TestVacuum::test_goto",
"miio/tests/test_vacuum.py::TestVacuum::test_home",
"miio/tests/test_vacuum.py::TestVacuum::test_pause",
"miio/tests/test_vacuum.py::TestVacuum::test_spot",
"miio/tests/test_vacuum.py::TestVacuum::test_start_and_stop",
"miio/tests/test_vacuum.py::TestVac... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Xiaomi vacuum. Add property for water box (water tank) attach status
----------
</request>
There are several new functions or classes that need to be implemented, using the definitions below:
<<NE... | 62427d2f796e603520acca3b57b29ec3e6489bca | |
sphinx-doc__sphinx-7485 | 7,485 | sphinx-doc/sphinx | 3.1 | cec31c75d8f38d9bfbde39ead60da106bd39c641 | 2020-04-14T22:31:14Z | diff --git a/.gitignore b/.gitignore
index b72664183d2..8d33409d5bb 100644
--- a/.gitignore
+++ b/.gitignore
@@ -9,6 +9,7 @@
.mypy_cache/
.pytest_cache/
.ropeproject/
+.vscode/
TAGS
.tags
.tox/
diff --git a/sphinx/domains/cpp.py b/sphinx/domains/cpp.py
index e09a56b06f6..3365bf786df 100644
--- a/sphinx/domains/cp... | diff --git a/tests/test_domain_cpp.py b/tests/test_domain_cpp.py
index 0b757139afb..0180a11d3b4 100644
--- a/tests/test_domain_cpp.py
+++ b/tests/test_domain_cpp.py
@@ -393,7 +393,7 @@ def test_function_definitions():
x = 'std::vector<std::pair<std::string, int>> &module::test(register int ' \
'foo, bar, ... | diff --git a/.gitignore b/.gitignore
index b72664183d2..8d33409d5bb 100644
--- a/.gitignore
+++ b/.gitignore
@@ -9,6 +9,7 @@
.mypy_cache/
.pytest_cache/
.ropeproject/
+.vscode/
TAGS
.tags
.tox/
| [
{
"components": [
{
"doc": "",
"lines": [
1816,
1831
],
"name": "ASTNoexceptSpec",
"signature": "class ASTNoexceptSpec(ASTBase):",
"type": "class"
},
{
"doc": "",
"lines": [
1817,
1818
... | [
"tests/test_domain_cpp.py::test_function_definitions"
] | [
"tests/test_domain_cpp.py::test_fundamental_types",
"tests/test_domain_cpp.py::test_expressions",
"tests/test_domain_cpp.py::test_type_definitions",
"tests/test_domain_cpp.py::test_concept_definitions",
"tests/test_domain_cpp.py::test_member_definitions",
"tests/test_domain_cpp.py::test_operators",
"tes... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
C++, add support for parameterized noexcept specifier in function dec…
…larations
Subject: C++, Add support for parameterized noexcept specifier in function declarations
<!--
Before posting a p... | 3419079fb0d1f0eecd845eff0d12b367d34cd5e9 | |
sphinx-doc__sphinx-7471 | 7,471 | sphinx-doc/sphinx | 3.0 | 39cd463740d6ec955a5e216eb07d09a51204ecb4 | 2020-04-13T08:20:53Z | diff --git a/CHANGES b/CHANGES
index 84d3c266944..81674bef788 100644
--- a/CHANGES
+++ b/CHANGES
@@ -13,6 +13,9 @@ Deprecated
Features added
--------------
+* C, parse attributes and add :confval:`c_id_attributes`
+ and :confval:`c_paren_attributes` to support user-defined attributes.
+
Bugs fixed
----------
d... | diff --git a/tests/test_domain_c.py b/tests/test_domain_c.py
index 3255efc5535..009f51644d4 100644
--- a/tests/test_domain_c.py
+++ b/tests/test_domain_c.py
@@ -7,28 +7,20 @@
:copyright: Copyright 2007-2020 by the Sphinx team, see AUTHORS.
:license: BSD, see LICENSE for details.
"""
-
-import re
-
import py... | diff --git a/CHANGES b/CHANGES
index 84d3c266944..81674bef788 100644
--- a/CHANGES
+++ b/CHANGES
@@ -13,6 +13,9 @@ Deprecated
Features added
--------------
+* C, parse attributes and add :confval:`c_id_attributes`
+ and :confval:`c_paren_attributes` to support user-defined attributes.
+
Bugs fixed
----------
d... | [
{
"components": [
{
"doc": "",
"lines": [
1994,
1995
],
"name": "DefinitionParser.id_attributes",
"signature": "def id_attributes(self):",
"type": "function"
},
{
"doc": "",
"lines": [
1998,
... | [
"tests/test_domain_c.py::test_expressions",
"tests/test_domain_c.py::test_type_definitions",
"tests/test_domain_c.py::test_macro_definitions",
"tests/test_domain_c.py::test_member_definitions",
"tests/test_domain_c.py::test_function_definitions",
"tests/test_domain_c.py::test_union_definitions",
"tests/... | [
"tests/test_domain_c.py::test_anon_definitions",
"tests/test_domain_c.py::test_build_domain_c",
"tests/test_domain_c.py::test_build_domain_c_anon_dup_decl",
"tests/test_domain_c.py::test_cfunction",
"tests/test_domain_c.py::test_cmember",
"tests/test_domain_c.py::test_cvar",
"tests/test_domain_cpp.py::t... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
C, parse attributes
### Feature or Bugfix
- Feature
- Bugfix
The 3.0 version of the C domain got more strict, resulting in warnings for declarations that are not technically C, but are rather com... | 39cd463740d6ec955a5e216eb07d09a51204ecb4 | |
sphinx-doc__sphinx-7468 | 7,468 | sphinx-doc/sphinx | 3.1 | f978490dfe2f65170e92ad2ccc836cc8ab01c1fa | 2020-04-12T12:21:43Z | diff --git a/CHANGES b/CHANGES
index 863a5ea2473..ad5b6bd11ee 100644
--- a/CHANGES
+++ b/CHANGES
@@ -16,6 +16,8 @@ Features added
* LaTeX: Make the ``toplevel_sectioning`` setting optional in LaTeX theme
* #7410: Allow to suppress "circular toctree references detected" warnings using
:confval:`suppress_warnings`
+... | diff --git a/tests/roots/test-domain-c/namespace.rst b/tests/roots/test-domain-c/namespace.rst
new file mode 100644
index 00000000000..c220d38e7c7
--- /dev/null
+++ b/tests/roots/test-domain-c/namespace.rst
@@ -0,0 +1,21 @@
+.. c:namespace:: NS
+
+.. c:var:: int NSVar
+
+.. c:namespace:: NULL
+
+.. c:var:: int NULLVar
... | diff --git a/CHANGES b/CHANGES
index 863a5ea2473..ad5b6bd11ee 100644
--- a/CHANGES
+++ b/CHANGES
@@ -16,6 +16,8 @@ Features added
* LaTeX: Make the ``toplevel_sectioning`` setting optional in LaTeX theme
* #7410: Allow to suppress "circular toctree references detected" warnings using
:confval:`suppress_warnings`
+... | [
{
"components": [
{
"doc": "",
"lines": [
2932,
2933
],
"name": "DefinitionParser.parse_namespace_object",
"signature": "def parse_namespace_object(self) -> ASTNestedName:",
"type": "function"
},
{
"doc": "This directi... | [
"tests/test_domain_c.py::test_build_domain_c"
] | [
"tests/test_domain_c.py::test_expressions",
"tests/test_domain_c.py::test_type_definitions",
"tests/test_domain_c.py::test_macro_definitions",
"tests/test_domain_c.py::test_member_definitions",
"tests/test_domain_c.py::test_function_definitions",
"tests/test_domain_c.py::test_union_definitions",
"tests/... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
C, add scoping directives
Subject: add scoping directives to the C domain in the same style as the C++ domain.
### Feature or Bugfix
<!-- please choose -->
- Feature
### Relates
Will give mor... | 3419079fb0d1f0eecd845eff0d12b367d34cd5e9 | |
sympy__sympy-19094 | 19,094 | sympy/sympy | 1.6 | 64d28fe0534f6993695d11244ea740f783958dc8 | 2020-04-09T13:09:33Z | diff --git a/sympy/geometry/line.py b/sympy/geometry/line.py
index 12c162cc60bb..cd52e1ad45b9 100644
--- a/sympy/geometry/line.py
+++ b/sympy/geometry/line.py
@@ -18,28 +18,28 @@
"""
from __future__ import division, print_function
-
from sympy import Expr
from sympy.core import S, sympify
from sympy.core.compati... | diff --git a/sympy/geometry/tests/test_line.py b/sympy/geometry/tests/test_line.py
index c953ae6377c0..89af9cfc844d 100644
--- a/sympy/geometry/tests/test_line.py
+++ b/sympy/geometry/tests/test_line.py
@@ -784,6 +784,20 @@ def test_parameter_value():
raises(ValueError, lambda: l.parameter_value((0, 0), t))
+d... | [
{
"components": [
{
"doc": "Returns the perpendicular lines which pass through the intersections\nof self and other that are in the same plane.\n\nParameters\n==========\n\nline : Line3D\n\nReturns\n=======\n\nlist: two Line instances\n\nExamples\n========\n\n>>> from sympy.geometry import Point3D... | [
"test_bisectors"
] | [
"test_object_from_equation",
"test_angle_between",
"test_closing_angle",
"test_smallest_angle",
"test_svg",
"test_arbitrary_point",
"test_are_concurrent_2d",
"test_are_concurrent_3d",
"test_arguments",
"test_basic_properties_2d",
"test_basic_properties_3d",
"test_contains",
"test_contains_no... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
add Line.bisectors
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or PRs
<!-- If this p... | Here is the discussion in the issues of the pull request.
<issues>
Added bisection method for 3D lines
This PR adds a new method for generating the bisection line of two 3D lines. It currently needs a code quality revision and a better documentation.
<!-- Your title above should be a short description of what
was c... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
sympy__sympy-19077 | 19,077 | sympy/sympy | 1.6 | 116722f31a57dac8c7e9f30c6b46c034326958bd | 2020-04-05T18:05:50Z | diff --git a/sympy/combinatorics/__init__.py b/sympy/combinatorics/__init__.py
index f55034471876..c04fb9915992 100644
--- a/sympy/combinatorics/__init__.py
+++ b/sympy/combinatorics/__init__.py
@@ -6,7 +6,7 @@
RGS_rank, RGS_unrank, RGS_enum)
from sympy.combinatorics.polyhedron import (Polyhedron, tetrahedron, cu... | diff --git a/sympy/combinatorics/tests/test_perm_groups.py b/sympy/combinatorics/tests/test_perm_groups.py
index ad6eaa941f97..17a72a4689f0 100644
--- a/sympy/combinatorics/tests/test_perm_groups.py
+++ b/sympy/combinatorics/tests/test_perm_groups.py
@@ -1,5 +1,5 @@
from sympy.combinatorics.perm_groups import (Permuta... | [
{
"components": [
{
"doc": "The class defining the lazy form of SymmetricGroup.\n\ndeg : int",
"lines": [
5083,
5163
],
"name": "SymmetricPermutationGroup",
"signature": "class SymmetricPermutationGroup(Basic):",
"type": "class"
},
... | [
"test_has",
"test_generate",
"test_order",
"test_equality",
"test_stabilizer",
"test_center",
"test_centralizer",
"test_coset_rank",
"test_coset_factor",
"test_orbits",
"test_is_normal",
"test_eq",
"test_derived_subgroup",
"test_is_solvable",
"test_rubik1",
"test_direct_product",
"te... | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
perm_groups.py: Add Coset class
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or PRs
<... | Here is the discussion in the issues of the pull request.
<issues>
Permutation * PermutationGroup should give the coset
```
In [5]: a
Out[5]: Permutation(1, 2)
In [6]: b
Out[6]: Permutation(0, 1)
In [7]: G
Out[7]:
PermutationGroup([
Permutation(1, 2),
Permutation(2)(0, 1)])
In [8]: a*G
----------------... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
joke2k__faker-1147 | 1,147 | joke2k/faker | null | 5f058d9da0d1c3fcc9c237065aec5ad5473d633b | 2020-04-04T18:26:53Z | diff --git a/faker/providers/color/es_ES/__init__.py b/faker/providers/color/es_ES/__init__.py
new file mode 100644
index 0000000000..24cf88e123
--- /dev/null
+++ b/faker/providers/color/es_ES/__init__.py
@@ -0,0 +1,155 @@
+from collections import OrderedDict
+
+from .. import Provider as ColorProvider
+
+localized = T... | diff --git a/tests/providers/test_color.py b/tests/providers/test_color.py
index a126f57231..1c673864f6 100644
--- a/tests/providers/test_color.py
+++ b/tests/providers/test_color.py
@@ -7,6 +7,7 @@
from faker import Faker
from faker.providers.color import RandomColor
+from faker.providers.color.es_ES import Provid... | [
{
"components": [
{
"doc": "",
"lines": [
8,
154
],
"name": "Provider",
"signature": "class Provider(ColorProvider):",
"type": "class"
}
],
"file": "faker/providers/color/es_ES/__init__.py"
}
] | [
"tests/providers/test_color.py::TestColor::test_color",
"tests/providers/test_color.py::TestColor::test_hex_color",
"tests/providers/test_color.py::TestColor::test_rgb_color",
"tests/providers/test_color.py::TestColor::test_rgb_css_color",
"tests/providers/test_color.py::TestColor::test_safe_hex_color",
"... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added color provider for es_ES locale.
----------
</request>
There are several new functions or classes that need to be implemented, using the definitions below:
<<NEW DEFINITIONS>>
There are se... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
joke2k__faker-1146 | 1,146 | joke2k/faker | null | 5f058d9da0d1c3fcc9c237065aec5ad5473d633b | 2020-04-04T15:56:52Z | diff --git a/faker/providers/bank/es_ES/__init__.py b/faker/providers/bank/es_ES/__init__.py
new file mode 100644
index 0000000000..807d57e42e
--- /dev/null
+++ b/faker/providers/bank/es_ES/__init__.py
@@ -0,0 +1,6 @@
+from .. import Provider as BankProvider
+
+
+class Provider(BankProvider):
+ bban_format = '######... | diff --git a/tests/providers/test_bank.py b/tests/providers/test_bank.py
index a6cc524d24..2ed83dee72 100644
--- a/tests/providers/test_bank.py
+++ b/tests/providers/test_bank.py
@@ -110,3 +110,19 @@ def test_bban(self):
def test_iban(self):
iban = self.fake.iban()
assert re.match(r"PT\d{21}", ib... | [
{
"components": [
{
"doc": "",
"lines": [
4,
6
],
"name": "Provider",
"signature": "class Provider(BankProvider):",
"type": "class"
}
],
"file": "faker/providers/bank/es_ES/__init__.py"
}
] | [
"tests/providers/test_bank.py::TestEsES::test_bban",
"tests/providers/test_bank.py::TestEsES::test_iban"
] | [
"tests/providers/test_bank.py::TestNoNO::test_bban",
"tests/providers/test_bank.py::TestFiFi::test_bban",
"tests/providers/test_bank.py::TestFiFi::test_iban",
"tests/providers/test_bank.py::TestPlPL::test_bban",
"tests/providers/test_bank.py::TestPlPL::test_iban",
"tests/providers/test_bank.py::TestEnGB::... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added bank provider for es_ES locale.
----------
</request>
There are several new functions or classes that need to be implemented, using the definitions below:
<<NEW DEFINITIONS>>
There are sever... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
joke2k__faker-1145 | 1,145 | joke2k/faker | null | 0e1ddfdcb1c692227d56f8ffef6cf65d3cd50d31 | 2020-04-02T15:10:34Z | diff --git a/faker/proxy.py b/faker/proxy.py
index 7225f96f93..b91a0709c0 100644
--- a/faker/proxy.py
+++ b/faker/proxy.py
@@ -55,6 +55,14 @@ def __init__(self, locale=None, providers=None,
self._locales = locales
self._factories = list(self._factory_map.values())
+ def __dir__(self):
+ at... | diff --git a/tests/test_proxy.py b/tests/test_proxy.py
index 69d3dcb01f..7aa9338596 100644
--- a/tests/test_proxy.py
+++ b/tests/test_proxy.py
@@ -299,3 +299,18 @@ def test_multiple_locale_factory_selection_unsupported_method(self):
self.fake = Faker(['en_US', 'en_PH'])
with self.assertRaises(Attribut... | [
{
"components": [
{
"doc": "",
"lines": [
58,
64
],
"name": "Faker.__dir__",
"signature": "def __dir__(self):",
"type": "function"
}
],
"file": "faker/proxy.py"
}
] | [
"tests/test_proxy.py::TestFakerProxyClass::test_dir_include_all_providers_attribute_in_list"
] | [
"tests/test_proxy.py::TestFakerProxyClass::test_dunder_getitem",
"tests/test_proxy.py::TestFakerProxyClass::test_items",
"tests/test_proxy.py::TestFakerProxyClass::test_locale_as_list",
"tests/test_proxy.py::TestFakerProxyClass::test_locale_as_ordereddict",
"tests/test_proxy.py::TestFakerProxyClass::test_lo... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implements __dir__ method to Faker proxy
### What does this changes
Implements __dir__ magic method to Faker proxy. This allows when trying to list/show all methods of all providers in IPython, for... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
sympy__sympy-19045 | 19,045 | sympy/sympy | 1.6 | 63ab534ebc9bb8fa47ca80aea5a1b0eca743fb9e | 2020-04-01T10:30:02Z | diff --git a/sympy/matrices/common.py b/sympy/matrices/common.py
index 721be20b8734..61d0c7d56df5 100644
--- a/sympy/matrices/common.py
+++ b/sympy/matrices/common.py
@@ -175,6 +175,16 @@ def entry(i, j):
def _eval_tolist(self):
return [list(self[i,:]) for i in range(self.rows)]
+ def _eval_todok(sel... | diff --git a/sympy/matrices/expressions/tests/test_blockmatrix.py b/sympy/matrices/expressions/tests/test_blockmatrix.py
index 9bc99b0ca0be..ed79dbac26c0 100644
--- a/sympy/matrices/expressions/tests/test_blockmatrix.py
+++ b/sympy/matrices/expressions/tests/test_blockmatrix.py
@@ -172,6 +172,7 @@ def test_BlockDiagMat... | [
{
"components": [
{
"doc": "",
"lines": [
178,
186
],
"name": "MatrixShaping._eval_todok",
"signature": "def _eval_todok(self):",
"type": "function"
},
{
"doc": "Return the matrix as dictionary of keys.\n\nExamples\n==... | [
"test_BlockDiagMatrix",
"test_todok"
] | [
"test_bc_matmul",
"test_bc_matadd",
"test_bc_transpose",
"test_bc_dist_diag",
"test_block_plus_ident",
"test_BlockMatrix",
"test_block_collapse_explicit_matrices",
"test_issue_17624",
"test_issue_18618",
"test_BlockMatrix_trace",
"test_BlockMatrix_Determinant",
"test_squareBlockMatrix",
"tes... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Matrix connected components decomposition
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
joke2k__faker-1144 | 1,144 | joke2k/faker | null | 0e1ddfdcb1c692227d56f8ffef6cf65d3cd50d31 | 2020-03-31T22:38:24Z | diff --git a/faker/providers/automotive/es_ES/__init__.py b/faker/providers/automotive/es_ES/__init__.py
new file mode 100644
index 0000000000..ebbd7f10fd
--- /dev/null
+++ b/faker/providers/automotive/es_ES/__init__.py
@@ -0,0 +1,99 @@
+# -*- coding: utf-8 -*-
+
+import re
+
+from .. import Provider as AutomotiveProvi... | diff --git a/tests/providers/test_automotive.py b/tests/providers/test_automotive.py
index ddffb94500..a23b2664dc 100644
--- a/tests/providers/test_automotive.py
+++ b/tests/providers/test_automotive.py
@@ -146,3 +146,34 @@ def test_fr_FR_plate_format(self):
plate = self.fake.license_plate()
assert is... | [
{
"components": [
{
"doc": "",
"lines": [
8,
99
],
"name": "Provider",
"signature": "class Provider(AutomotiveProvider):",
"type": "class"
},
{
"doc": "",
"lines": [
80,
85
],
... | [
"tests/providers/test_automotive.py::TestEsES::test_es_ES_plate_format",
"tests/providers/test_automotive.py::TestEsES::test_es_ES_plate_new_format",
"tests/providers/test_automotive.py::TestEsES::test_es_ES_plate_old_format",
"tests/providers/test_automotive.py::TestEsES::test_es_ES_plate_old_format_explicit... | [
"tests/providers/test_automotive.py::TestPtBR::test_plate_has_been_generated",
"tests/providers/test_automotive.py::TestPtPT::test_pt_PT_plate_format",
"tests/providers/test_automotive.py::TestHuHU::test_hu_HU_plate_format",
"tests/providers/test_automotive.py::TestDeDe::test_de_DE_plate_format",
"tests/pro... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added provider for Spanish (Spain) license plates.
### What does this changes
Add the provider es_ES for license plates (automotive provider).
----------
</request>
There are several new functions ... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
sympy__sympy-18953 | 18,953 | sympy/sympy | 1.6 | 1c952df73f11d7251ab0b974840862b88540fcbd | 2020-03-25T14:47:46Z | diff --git a/sympy/matrices/common.py b/sympy/matrices/common.py
index 443dfd6dd55b..721be20b8734 100644
--- a/sympy/matrices/common.py
+++ b/sympy/matrices/common.py
@@ -2019,6 +2019,51 @@ def replace(self, F, G, map=False, simultaneous=True, exact=None):
return self.applyfunc(
lambda x: x.replac... | diff --git a/sympy/matrices/tests/test_commonmatrix.py b/sympy/matrices/tests/test_commonmatrix.py
index cfb43fa05700..f25ffe2f6c36 100644
--- a/sympy/matrices/tests/test_commonmatrix.py
+++ b/sympy/matrices/tests/test_commonmatrix.py
@@ -522,6 +522,13 @@ def test_replace_map():
assert N == K
+def test_rot90()... | [
{
"components": [
{
"doc": "Rotates Matrix by 90 degrees\n\nParameters\n==========\n\nk : int\n Specifies how many times the matrix is rotated by 90 degrees\n (clockwise when positive, counter-clockwise when negative).\n\nExamples\n========\n\n>>> from sympy import Matrix, symbols\n>>> A = M... | [
"test_rot90"
] | [
"test__MinimalMatrix",
"test_vec",
"test_tolist",
"test_row_col_del",
"test_get_diag_blocks1",
"test_get_diag_blocks2",
"test_shape",
"test_reshape",
"test_row_col",
"test_row_join",
"test_col_join",
"test_row_insert",
"test_col_insert",
"test_extract",
"test_hstack",
"test_vstack",
... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
for 90 degree rotation of matrix elements
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues... | Here is the discussion in the issues of the pull request.
<issues>
90 degree rotation of matrix elements
I think that transpose matrix elements clockwise is often given as a coding test problem.
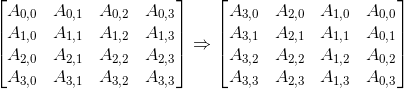
I see lo... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
sphinx-doc__sphinx-7380 | 7,380 | sphinx-doc/sphinx | 3.0 | 6deb592a2fadbf88b7a00a332837a81a3198a830 | 2020-03-25T13:08:37Z | diff --git a/CHANGES b/CHANGES
index a75d24e2e80..42bae4b8204 100644
--- a/CHANGES
+++ b/CHANGES
@@ -20,6 +20,8 @@ Bugs fixed
* #7370: autosummary: raises UnboundLocalError when unknown module given
* #7367: C++, alternate operator spellings are now supported.
* C, alternate operator spellings are now supported.
+* ... | diff --git a/tests/test_domain_cpp.py b/tests/test_domain_cpp.py
index c6d0df8aaf7..2faa3d4ea54 100644
--- a/tests/test_domain_cpp.py
+++ b/tests/test_domain_cpp.py
@@ -274,6 +274,9 @@ class Config:
exprCheck('a xor_eq 5', 'eO1aL5E')
exprCheck('a |= 5', 'oR1aL5E')
exprCheck('a or_eq 5', 'oR1aL5E')
+ e... | diff --git a/CHANGES b/CHANGES
index a75d24e2e80..42bae4b8204 100644
--- a/CHANGES
+++ b/CHANGES
@@ -20,6 +20,8 @@ Bugs fixed
* #7370: autosummary: raises UnboundLocalError when unknown module given
* #7367: C++, alternate operator spellings are now supported.
* C, alternate operator spellings are now supported.
+* ... | [
{
"components": [
{
"doc": "",
"lines": [
1573,
1595
],
"name": "ASTCommaExpr",
"signature": "class ASTCommaExpr(ASTExpression):",
"type": "class"
},
{
"doc": "",
"lines": [
1574,
1576
... | [
"tests/test_domain_cpp.py::test_expressions",
"tests/test_domain_cpp.py::test_class_definitions"
] | [
"tests/test_domain_cpp.py::test_fundamental_types",
"tests/test_domain_cpp.py::test_type_definitions",
"tests/test_domain_cpp.py::test_concept_definitions",
"tests/test_domain_cpp.py::test_member_definitions",
"tests/test_domain_cpp.py::test_function_definitions",
"tests/test_domain_cpp.py::test_operators... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
C++, comma operator, pack expansion, error messages
Subject: add missing features in the C++ domain and improve certain error messages.
### Feature or Bugfix
- Feature
- Bugfix
- Refactoring
... | Here is the discussion in the issues of the pull request.
<issues>
cpp domain parens in template parameter packs fails
**Describe the bug**
I have C++ code with parentheses in the template parameter list documented as:
```
.. cpp:class:: template <std::integer_sequence<bool, (static_cast<void>(Bs), false)>> foo
... | 39cd463740d6ec955a5e216eb07d09a51204ecb4 |
joke2k__faker-1140 | 1,140 | joke2k/faker | null | cd62db546bd0fc2b69928eda39bd232b7aaa9c5b | 2020-03-24T22:23:43Z | diff --git a/faker/providers/automotive/no_NO/__init__.py b/faker/providers/automotive/no_NO/__init__.py
new file mode 100644
index 0000000000..4fc5352baf
--- /dev/null
+++ b/faker/providers/automotive/no_NO/__init__.py
@@ -0,0 +1,9 @@
+from .. import Provider as AutomotiveProvider
+
+
+class Provider(AutomotiveProvide... | diff --git a/tests/providers/test_automotive.py b/tests/providers/test_automotive.py
index a23b2664dc..71166f97c9 100644
--- a/tests/providers/test_automotive.py
+++ b/tests/providers/test_automotive.py
@@ -148,6 +148,17 @@ def test_fr_FR_plate_format(self):
assert self.pattern.match(plate)
+class TestNoNO... | [
{
"components": [
{
"doc": "",
"lines": [
4,
8
],
"name": "Provider",
"signature": "class Provider(AutomotiveProvider):",
"type": "class"
}
],
"file": "faker/providers/automotive/no_NO/__init__.py"
}
] | [
"tests/providers/test_automotive.py::TestNoNO::test_sv_SE_plate_format"
] | [
"tests/providers/test_automotive.py::TestPtBR::test_plate_has_been_generated",
"tests/providers/test_automotive.py::TestPtPT::test_pt_PT_plate_format",
"tests/providers/test_automotive.py::TestHuHU::test_hu_HU_plate_format",
"tests/providers/test_automotive.py::TestDeDe::test_de_DE_plate_format",
"tests/pro... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added provider for Norwegian license plates.
### What does this changes
Added license plate provider for no_NO locale.
### What was wrong
Missing implementation.
### How this fixes it
... | Here is the discussion in the issues of the pull request.
<issues>
Missing implementation of no_NO license plates.
* Faker version:
* OS:
no_NO license plates are not implemeted.
### Steps to reproduce
### Expected behavior
Should be of the form
> AB 12345
### Actual behavior
#1140
----------
-----... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | |
Project-MONAI__MONAI-202 | 202 | Project-MONAI/MONAI | null | 1cbdcc30a0bde26e1f5516ea3086cdfdf33da0b0 | 2020-03-23T12:33:16Z | diff --git a/examples/segmentation_3d/unet_evaluation_array.py b/examples/segmentation_3d/unet_evaluation_array.py
index 305a513c1e..60d81cf883 100644
--- a/examples/segmentation_3d/unet_evaluation_array.py
+++ b/examples/segmentation_3d/unet_evaluation_array.py
@@ -95,7 +95,7 @@ def _sliding_window_processor(engine, b... | diff --git a/tests/integration_sliding_window.py b/tests/integration_sliding_window.py
index 91fd2994be..4cc1e6b5b1 100644
--- a/tests/integration_sliding_window.py
+++ b/tests/integration_sliding_window.py
@@ -58,7 +58,7 @@ def _sliding_window_processor(_engine, batch):
infer_engine = Engine(_sliding_window_proce... | [
{
"components": [
{
"doc": "save the data in a dictionary format cache, and write to a CSV file finally.\nTypically, the data can be classification predictions, call `save` for single data\nor call `save_batch` to save a batch of data together, and call `finalize` to write\nthe cached data into CS... | [
"tests/test_csv_saver.py::TestCSVSaver::test_saved_content",
"tests/test_handler_classification_saver.py::TestHandlerClassificationSaver::test_saved_content",
"tests/test_handler_segmentation_saver.py::TestHandlerSegmentationSaver::test_saved_content",
"tests/test_nifti_saver.py::TestNiftiSaver::test_saved_co... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
201 refactor event handlers for flexibility
Fixes #201 .
### Description
This PR refactor the event handlers to split the code structure to 2 levels, then we can call the same functions even witho... | Here is the discussion in the issues of the pull request.
<issues>
refactor event handlers for flexibility
**Is your feature request related to a problem? Please describe.**
Currently, we implemented many logics in the event handlers directly, it's not flexible to use.
**Describe the solution you'd like**
Split ha... | e73257caa79309dcce1e93abf1632f4bfd75b11f | |
sphinx-doc__sphinx-7346 | 7,346 | sphinx-doc/sphinx | 3.0 | 79989ce40e4aa168a354d85e41bfebc97a5650bb | 2020-03-21T05:42:42Z | diff --git a/CHANGES b/CHANGES
index ecba1c5e52e..fc03bdb323a 100644
--- a/CHANGES
+++ b/CHANGES
@@ -83,6 +83,7 @@ Features added
* #6417: py domain: Allow to make a style for arguments of functions and methods
* #7238, #7239: py domain: Emit a warning on describing a python object if the
entry is already added as... | diff --git a/sphinx/testing/util.py b/sphinx/testing/util.py
index 75cd0f411bb..450241f5513 100644
--- a/sphinx/testing/util.py
+++ b/sphinx/testing/util.py
@@ -63,7 +63,7 @@ def assert_node(node: Node, cls: Any = None, xpath: str = "", **kwargs: Any) ->
'The node%s has %d child nodes, not one'... | diff --git a/CHANGES b/CHANGES
index ecba1c5e52e..fc03bdb323a 100644
--- a/CHANGES
+++ b/CHANGES
@@ -83,6 +83,7 @@ Features added
* #6417: py domain: Allow to make a style for arguments of functions and methods
* #7238, #7239: py domain: Emit a warning on describing a python object if the
entry is already added as... | [
{
"components": [
{
"doc": "Parse type annotation.",
"lines": [
71,
120
],
"name": "_parse_annotation",
"signature": "def _parse_annotation(annotation: str) -> List[Node]:",
"type": "function"
},
{
"doc": "",
"... | [
"tests/test_domain_py.py::test_function_signatures",
"tests/test_domain_py.py::test_domain_py_xrefs",
"tests/test_domain_py.py::test_domain_py_objects",
"tests/test_domain_py.py::test_resolve_xref_for_properties",
"tests/test_domain_py.py::test_domain_py_find_obj",
"tests/test_domain_py.py::test_get_full_... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Close #7341: py domain: type annotations are converted to cross refs
### Feature or Bugfix
- Feature
### Purpose
- refs: #7341
- With this change, the example on #7341 is converted to this im... | Here is the discussion in the issues of the pull request.
<issues>
Autodoc type hints for other classes link to other class
**Describe the solution you'd like**
If I have a class as an annotation type, and the class is in the project or part of my intersphinx, it should link to that class' definition if it exists.
... | 39cd463740d6ec955a5e216eb07d09a51204ecb4 |
joke2k__faker-1136 | 1,136 | joke2k/faker | null | d5123ba865cbcc89d82ee3a30992119f09c4e5b9 | 2020-03-19T13:19:56Z | diff --git a/faker/providers/internet/__init__.py b/faker/providers/internet/__init__.py
index 60dab82429..e42ff0bd1f 100644
--- a/faker/providers/internet/__init__.py
+++ b/faker/providers/internet/__init__.py
@@ -222,6 +222,33 @@ def domain_word(self):
company = self._to_ascii(company_elements.pop(0))
... | diff --git a/tests/test_factory.py b/tests/test_factory.py
index a93df7cffe..9ba552042b 100644
--- a/tests/test_factory.py
+++ b/tests/test_factory.py
@@ -889,6 +889,14 @@ def test_http_method(self):
assert expected_methods == sorted(got_methods)
+ def test_dga(self):
+ faker = Faker()
+
+ ... | [
{
"components": [
{
"doc": "Generates a domain name by given date\nhttps://en.wikipedia.org/wiki/Domain_generation_algorithm\n\n:type year: int\n:type month: int\n:type day: int\n:type tld: str\n:type length: int\n:rtype: str",
"lines": [
225,
250
],
"na... | [
"tests/test_factory.py::FactoryTestCase::test_dga"
] | [
"tests/test_factory.py::FactoryTestCase::test_add_provider_gives_priority_to_newly_added_provider",
"tests/test_factory.py::FactoryTestCase::test_binary",
"tests/test_factory.py::FactoryTestCase::test_cli_seed",
"tests/test_factory.py::FactoryTestCase::test_cli_seed_with_repeat",
"tests/test_factory.py::Fac... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add dga by date
### What does this changes
Add Domain Generator Algorithm by date method and test for it
----------
</request>
There are several new functions or classes that need to be imple... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
matplotlib__matplotlib-16832 | 16,832 | matplotlib/matplotlib | 3.2 | 005c7a67bd997fa5f0aee75f9f4ffdf14709f8f9 | 2020-03-19T11:51:34Z | diff --git a/doc/users/next_whats_new/2020-03-31-path-size-methods.rst b/doc/users/next_whats_new/2020-03-31-path-size-methods.rst
new file mode 100644
index 000000000000..d2347fb3b9e5
--- /dev/null
+++ b/doc/users/next_whats_new/2020-03-31-path-size-methods.rst
@@ -0,0 +1,27 @@
+
+Functions to compute a Path's size
+-... | diff --git a/lib/matplotlib/tests/test_path.py b/lib/matplotlib/tests/test_path.py
index b61a92654dc3..2a9ccb4662b0 100644
--- a/lib/matplotlib/tests/test_path.py
+++ b/lib/matplotlib/tests/test_path.py
@@ -49,6 +49,37 @@ def test_contains_points_negative_radius():
np.testing.assert_equal(result, [True, False, Fal... | diff --git a/doc/users/next_whats_new/2020-03-31-path-size-methods.rst b/doc/users/next_whats_new/2020-03-31-path-size-methods.rst
new file mode 100644
index 000000000000..d2347fb3b9e5
--- /dev/null
+++ b/doc/users/next_whats_new/2020-03-31-path-size-methods.rst
@@ -0,0 +1,27 @@
+
+Functions to compute a Path's size
+-... | [
{
"components": [
{
"doc": "",
"lines": [
16,
21
],
"name": "_comb",
"signature": "def _comb(n, k):",
"type": "function"
},
{
"doc": "Evaluate the Bezier curve at point(s) t in [0, 1].\n\nParameters\n----------\nt : fl... | [
"lib/matplotlib/tests/test_path.py::test_exact_extents[path0-extents0]",
"lib/matplotlib/tests/test_path.py::test_exact_extents[path1-extents1]"
] | [
"lib/matplotlib/tests/test_path.py::test_empty_closed_path",
"lib/matplotlib/tests/test_path.py::test_readonly_path",
"lib/matplotlib/tests/test_path.py::test_point_in_path",
"lib/matplotlib/tests/test_path.py::test_contains_points_negative_radius",
"lib/matplotlib/tests/test_path.py::test_exact_extents[pat... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Correctly compute path extents
## PR Summary
Closes #16830 and #7184.
This code will be needed to implement `MarkerStyle` normalization correctly (building on the work now approved in #16773).
... | Here is the discussion in the issues of the pull request.
<issues>
`_path.get_extents` does not correctly handle bezier curves
<!--To help us understand and resolve your issue, please fill out the form to the best of your ability.-->
<!--You can feel free to delete the sections that do not apply.-->
### Bug report
... | ff821ba3249401fe7f5fdb11cb7ac5d0564c2697 |
sympy__sympy-18908 | 18,908 | sympy/sympy | 1.6 | 5b92c4497fcc6f1df4aac23b9c001ff323ffb421 | 2020-03-18T23:58:55Z | diff --git a/sympy/printing/pycode.py b/sympy/printing/pycode.py
index 1d04195fce7d..9ecdf83469a6 100644
--- a/sympy/printing/pycode.py
+++ b/sympy/printing/pycode.py
@@ -970,6 +970,26 @@ def _print_fresnelc(self, expr):
self._module_format("scipy.special.fresnel"),
self._print(expr.ar... | diff --git a/sympy/printing/tests/test_pycode.py b/sympy/printing/tests/test_pycode.py
index 1f061217d4fb..0ecfb2caa38f 100644
--- a/sympy/printing/tests/test_pycode.py
+++ b/sympy/printing/tests/test_pycode.py
@@ -284,3 +284,39 @@ def test_beta():
prntr = MpmathPrinter()
assert prntr.doprint(expr) == 'mpm... | [
{
"components": [
{
"doc": "",
"lines": [
973,
976
],
"name": "SciPyPrinter._print_airyai",
"signature": "def _print_airyai(self, expr):",
"type": "function"
},
{
"doc": "",
"lines": [
978,
... | [
"test_airy"
] | [
"test_PythonCodePrinter",
"test_PythonCodePrinter_standard",
"test_MpmathPrinter",
"test_NumPyPrinter",
"test_SciPyPrinter",
"test_pycode_reserved_words",
"test_sqrt",
"test_printmethod",
"test_codegen_ast_nodes",
"test_issue_14283",
"test_NumPyPrinter_print_seq",
"test_issue_16535_16536",
"... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added Airy functionality to the SciPyPrinter class
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to oth... | Here is the discussion in the issues of the pull request.
<issues>
Add more SciPy functions to code printer
Here is a list of special functions supported in SciPy: https://docs.scipy.org/doc/scipy/reference/special.html
Many of them are not supported in the SciPyPrinter and should be added.
----------
For each we al... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
sympy__sympy-18900 | 18,900 | sympy/sympy | 1.6 | 68f89c4b2abeb689d7d8425d1c6d55d4c62e5697 | 2020-03-18T12:21:27Z | diff --git a/sympy/physics/continuum_mechanics/beam.py b/sympy/physics/continuum_mechanics/beam.py
index 7132650d97ad..efdb9be531a4 100644
--- a/sympy/physics/continuum_mechanics/beam.py
+++ b/sympy/physics/continuum_mechanics/beam.py
@@ -72,7 +72,7 @@ class Beam(object):
+ Piecewise(((x - 2)**4, x - 2 > 0), ... | diff --git a/sympy/physics/continuum_mechanics/tests/test_beam.py b/sympy/physics/continuum_mechanics/tests/test_beam.py
index 509d509442b1..89edfb414978 100644
--- a/sympy/physics/continuum_mechanics/tests/test_beam.py
+++ b/sympy/physics/continuum_mechanics/tests/test_beam.py
@@ -16,6 +16,8 @@ def test_Beam():
E... | [
{
"components": [
{
"doc": "",
"lines": [
158,
159
],
"name": "Beam.area",
"signature": "def area(self, a):",
"type": "function"
},
{
"doc": "Returns an expression representing the Shear Stress\ncurve of the Beam objec... | [
"test_Beam",
"test_Beam3D"
] | [
"test_insufficient_bconditions",
"test_statically_indeterminate",
"test_beam_units",
"test_variable_moment",
"test_composite_beam",
"test_point_cflexure",
"test_remove_load",
"test_apply_support",
"test_max_shear_force",
"test_max_bmoment",
"test_max_deflection",
"test_polar_moment_Beam3D",
... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Functions and test cases added
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or PRs
<!... | Here is the discussion in the issues of the pull request.
<issues>
No function to calculate shear stress and axial stress
In `beam.py` file of `continuum_mechanics` folder there is no function to calculate shear stress and axial stress.
----------
@moorepants @czgdp1807 I would like to work on this. Is it available?
I ... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
sphinx-doc__sphinx-7327 | 7,327 | sphinx-doc/sphinx | 3.0 | 385f7ed40ef843db94ec33ad377aa87d63f27a27 | 2020-03-17T16:38:18Z | diff --git a/sphinx/util/logging.py b/sphinx/util/logging.py
index fb2ec29005f..a2dee807d33 100644
--- a/sphinx/util/logging.py
+++ b/sphinx/util/logging.py
@@ -220,16 +220,15 @@ def pending_warnings() -> Generator[logging.Handler, None, None]:
@contextmanager
-def pending_logging() -> Generator[MemoryHandler, Non... | diff --git a/tests/test_util_logging.py b/tests/test_util_logging.py
index 1581275ee23..85646112dd2 100644
--- a/tests/test_util_logging.py
+++ b/tests/test_util_logging.py
@@ -233,6 +233,20 @@ def test_warning_location(app, status, warning):
assert colorize('red', 'WARNING: message7') in warning.getvalue()
+d... | [
{
"components": [
{
"doc": "Contextmanager to suppress logging all logs temporary.\n\nFor example::\n\n >>> with suppress_logging():\n >>> logger.warning('Warning message!') # suppressed\n >>> some_long_process()\n >>>",
"lines": [
223,
248
... | [
"tests/test_util_logging.py::test_suppress_logging"
] | [
"tests/test_util_logging.py::test_info_and_warning",
"tests/test_util_logging.py::test_verbosity_filter",
"tests/test_util_logging.py::test_nonl_info_log",
"tests/test_util_logging.py::test_is_suppressed_warning",
"tests/test_util_logging.py::test_suppress_warnings",
"tests/test_util_logging.py::test_warn... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add suppress_logging()
### Feature or Bugfix
- Refactoring
### Purpose
As a helper for C/C++ domain, this adds suppress_logging(). It works as a context manager and suppresses all loggings durin... | 39cd463740d6ec955a5e216eb07d09a51204ecb4 | ||
sympy__sympy-18861 | 18,861 | sympy/sympy | 1.6 | e9f44d0b4497650df2f2f62c39fd1ebc940456fc | 2020-03-14T11:44:27Z | diff --git a/sympy/__init__.py b/sympy/__init__.py
index 71497002f990..8353b01b100a 100644
--- a/sympy/__init__.py
+++ b/sympy/__init__.py
@@ -160,7 +160,7 @@ def __sympy_debug():
ratsimp, ratsimpmodprime)
from .sets import (Set, Interval, Union, EmptySet, FiniteSet, ProductSet,
- Intersection, image... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 6fbd62089d70..735b85ce459e 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -894,6 +894,11 @@ def test_sympy__sets__sets__SymmetricDifference():
assert _test_args(SymmetricDifference(FiniteSet(1, 2, 3)... | [
{
"components": [
{
"doc": "Represents the disjoint union (also known as the external disjoint union)\nof a finite number of sets.\n\nExamples\n========\n\n>>> from sympy import DisjointUnion, FiniteSet, Interval, Union, Symbol\n>>> A = FiniteSet(1, 2, 3)\n>>> B = Interval(0, 5)\n>>> DisjointUnion... | [
"test_sympy__sets__sets__DisjointUnion",
"test_imageset",
"test_is_empty",
"test_is_finiteset",
"test_deprecated_is_EmptySet",
"test_interval_arguments",
"test_interval_symbolic_end_points",
"test_interval_is_empty",
"test_union",
"test_union_iter",
"test_union_is_empty",
"test_difference",
... | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implementing disjoint union
#### References to other Issues or PRs
Fixes #18833
#### Brief description of what is fixed or changed
Created a `DisjointUnion` object derived from `sympy.sets.sets... | Here is the discussion in the issues of the pull request.
<issues>
disjoint_union() in sets
Has evaluating the disjoint union of a collection of sets been implemented in SymPy? I could not find it therefore I'd like to implement it. Here's the definition: [https://en.wikipedia.org/wiki/Disjoint_union](url)
I was thi... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
RDFLib__rdflib-968 | 968 | RDFLib/rdflib | null | 0e5efef78702575e4abff4d8076eac4e2bd9d5f0 | 2020-03-13T11:56:00Z | diff --git a/rdflib/graph.py b/rdflib/graph.py
index 4a27e6de7..f68300cb1 100644
--- a/rdflib/graph.py
+++ b/rdflib/graph.py
@@ -1269,6 +1269,55 @@ def do_de_skolemize2(t):
return retval
+ def cbd(self, resource):
+ """Retrieves the Concise Bounded Description of a Resource from a Graph
+
+ ... | diff --git a/test/test_graph_cbd.py b/test/test_graph_cbd.py
new file mode 100644
index 000000000..aedc9dd07
--- /dev/null
+++ b/test/test_graph_cbd.py
@@ -0,0 +1,124 @@
+import unittest
+from rdflib import Graph, Namespace
+
+
+class CbdTestCase(unittest.TestCase):
+ """Tests the Graph class' cbd() function"""
+
+ ... | [
{
"components": [
{
"doc": "Retrieves the Concise Bounded Description of a Resource from a Graph\n\nConcise Bounded Description (CBD) is defined in [1] as:\n\nGiven a particular node (the starting node) in a particular RDF graph (the source graph), a subgraph of that\nparticular graph, taken to co... | [
"test/test_graph_cbd.py::CbdTestCase::testCbd",
"test/test_graph_cbd.py::CbdTestCase::testCbdReified"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Concise Bounded Description
This PR implements a Graph() function cbd() for 'Concise Bounded Description'. It extracts a subgraph from a source graph as per the rules in the W3C member submission [1].... | 0c11debb5178157baeac27b735e49a757916d2a6 | ||
sphinx-doc__sphinx-7298 | 7,298 | sphinx-doc/sphinx | 4.0 | e1a02dc4e721ede76567ced99742ec660fd2a803 | 2020-03-11T15:55:52Z | diff --git a/CHANGES b/CHANGES
index a183eba66b1..2a043fab768 100644
--- a/CHANGES
+++ b/CHANGES
@@ -66,6 +66,7 @@ Features added
the location where the object is defined
* #7199: py domain: Add :confval:`python_use_unqualified_type_names` to suppress
the module name of the python reference if it can be resolved... | diff --git a/tests/roots/test-domain-py/module.rst b/tests/roots/test-domain-py/module.rst
index dce3fa5acce..4a280681207 100644
--- a/tests/roots/test-domain-py/module.rst
+++ b/tests/roots/test-domain-py/module.rst
@@ -18,8 +18,7 @@ module
* Link to :py:meth:`module_a.submodule.ModTopLevel.mod_child_1`
-.. p... | diff --git a/CHANGES b/CHANGES
index a183eba66b1..2a043fab768 100644
--- a/CHANGES
+++ b/CHANGES
@@ -66,6 +66,7 @@ Features added
the location where the object is defined
* #7199: py domain: Add :confval:`python_use_unqualified_type_names` to suppress
the module name of the python reference if it can be resolved... | [
{
"components": [
{
"doc": "Description of an attribute.",
"lines": [
828,
865
],
"name": "PyProperty",
"signature": "class PyProperty(PyObject):",
"type": "class"
},
{
"doc": "",
"lines": [
837,
... | [
"tests/test_domain_py.py::test_domain_py_xrefs",
"tests/test_domain_py.py::test_resolve_xref_for_properties",
"tests/test_domain_py.py::test_pyproperty"
] | [
"tests/test_domain_py.py::test_function_signatures",
"tests/test_domain_py.py::test_domain_py_xrefs_abbreviations",
"tests/test_domain_py.py::test_domain_py_objects",
"tests/test_domain_py.py::test_domain_py_find_obj",
"tests/test_domain_py.py::test_get_full_qualified_name",
"tests/test_domain_py.py::test... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
py domain: Add py:property directive to describe a property (refs: #7068)
### Feature or Bugfix
- Feature
- Refactoring
### Purpose
- refs: #7068
----------
</request>
There are several new f... | cd75f8fea1cdd509615b58ba2b32756795fbaa03 | |
scikit-learn__scikit-learn-16625 | 16,625 | scikit-learn/scikit-learn | 0.24 | 6ca9eab67e1054a1f9508dfa286e0542c8bab5e3 | 2020-03-03T18:13:38Z | diff --git a/doc/modules/classes.rst b/doc/modules/classes.rst
index e05d08781181f..6dbab18d94a0c 100644
--- a/doc/modules/classes.rst
+++ b/doc/modules/classes.rst
@@ -966,6 +966,7 @@ details.
metrics.recall_score
metrics.roc_auc_score
metrics.roc_curve
+ metrics.top_k_accuracy_score
metrics.zero_one... | diff --git a/sklearn/metrics/tests/test_common.py b/sklearn/metrics/tests/test_common.py
index e503a64a47769..6688ddc2aa834 100644
--- a/sklearn/metrics/tests/test_common.py
+++ b/sklearn/metrics/tests/test_common.py
@@ -59,6 +59,7 @@
from sklearn.metrics import zero_one_loss
from sklearn.metrics import ndcg_score
f... | diff --git a/doc/modules/classes.rst b/doc/modules/classes.rst
index e05d08781181f..6dbab18d94a0c 100644
--- a/doc/modules/classes.rst
+++ b/doc/modules/classes.rst
@@ -966,6 +966,7 @@ details.
metrics.recall_score
metrics.roc_auc_score
metrics.roc_curve
+ metrics.top_k_accuracy_score
metrics.zero_one... | [
{
"components": [
{
"doc": "Top-k Accuracy classification score.\n\nThis metric computes the number of times where the correct label is among\nthe top `k` labels predicted (ranked by predicted scores). Note that the\nmultilabel case isn't covered here.\n\nRead more in the :ref:`User Guide <top_k_a... | [
"sklearn/metrics/tests/test_common.py::test_symmetry_consistency",
"sklearn/metrics/tests/test_common.py::test_symmetric_metric[accuracy_score]",
"sklearn/metrics/tests/test_common.py::test_symmetric_metric[cohen_kappa_score]",
"sklearn/metrics/tests/test_common.py::test_symmetric_metric[f1_score]",
"sklear... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
FEA Top k accuracy metric
#### Reference Issues/PRs
Closes #10488
Fixes #10144
Fixes #8234
#### What does this implement/fix? Explain your changes.
This implements a top-k accuracy classifica... | Here is the discussion in the issues of the pull request.
<issues>
Top k accuracy metric
**Reference Issues**
Fixes #10144
Fixes #8234
**What does this implement/fix? Explain your changes.**
This implements a top-k accuracy classification metric, for use with probabilistic class predictions in multiclass classi... | 54ce4222694819ad52d544ce5cba5da274c34ab7 |
matplotlib__matplotlib-16603 | 16,603 | matplotlib/matplotlib | 3.1 | c9be5a6985cc919703959902e9a8d795cfde03ad | 2020-02-28T22:07:53Z | diff --git a/doc/users/next_whats_new/2020-05-tac.rst b/doc/users/next_whats_new/2020-05-tac.rst
new file mode 100644
index 000000000000..de3427161951
--- /dev/null
+++ b/doc/users/next_whats_new/2020-05-tac.rst
@@ -0,0 +1,37 @@
+Add API for composing semantic axes layouts from text or nested lists
+-------------------... | diff --git a/lib/matplotlib/tests/test_figure.py b/lib/matplotlib/tests/test_figure.py
index 36f0c2f76ca0..9eec9cb97a7c 100644
--- a/lib/matplotlib/tests/test_figure.py
+++ b/lib/matplotlib/tests/test_figure.py
@@ -588,3 +588,186 @@ def test_animated_with_canvas_change(fig_test, fig_ref):
ax_test = fig_test.subp... | diff --git a/doc/users/next_whats_new/2020-05-tac.rst b/doc/users/next_whats_new/2020-05-tac.rst
new file mode 100644
index 000000000000..de3427161951
--- /dev/null
+++ b/doc/users/next_whats_new/2020-05-tac.rst
@@ -0,0 +1,37 @@
+Add API for composing semantic axes layouts from text or nested lists
+-------------------... | [
{
"components": [
{
"doc": "",
"lines": [
169,
184
],
"name": "SubplotBase.__repr__",
"signature": "def __repr__(self):",
"type": "function"
}
],
"file": "lib/matplotlib/axes/_subplots.py"
},
{
"components": [
... | [
"lib/matplotlib/tests/test_figure.py::TestSubplotMosaic::test_basic[x0-png]",
"lib/matplotlib/tests/test_figure.py::TestSubplotMosaic::test_basic[x1-png]",
"lib/matplotlib/tests/test_figure.py::TestSubplotMosaic::test_all_nested[png]",
"lib/matplotlib/tests/test_figure.py::TestSubplotMosaic::test_nested[png]"... | [
"lib/matplotlib/tests/test_figure.py::test_align_labels[png]",
"lib/matplotlib/tests/test_figure.py::test_figure_label",
"lib/matplotlib/tests/test_figure.py::test_fignum_exists",
"lib/matplotlib/tests/test_figure.py::test_clf_keyword",
"lib/matplotlib/tests/test_figure.py::test_figure[png]",
"lib/matplot... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
axes collage
## PR Summary
This is an attempt at implementing a nicer API for laying out complex axes schemes like patchwork for ggplot. This is following on from a conversation @anntzer and @stor... | c9be5a6985cc919703959902e9a8d795cfde03ad | |
Project-MONAI__MONAI-127 | 127 | Project-MONAI/MONAI | null | 5ae87f8e5056f23cb3e86394f4094bf5f61aa32f | 2020-02-28T14:16:03Z | diff --git a/docs/source/highlights.md b/docs/source/highlights.md
index f7895bf65d..113fc1a1c5 100644
--- a/docs/source/highlights.md
+++ b/docs/source/highlights.md
@@ -72,7 +72,16 @@ torch.backends.cudnn.benchmark = False
There are domain-specific loss functions in the medical research area which are different from... | diff --git a/tests/test_unet.py b/tests/test_unet.py
index 5b8e85f915..be661a4dbb 100644
--- a/tests/test_unet.py
+++ b/tests/test_unet.py
@@ -14,8 +14,23 @@
import torch
from parameterized import parameterized
+from monai.networks.layers.factories import Norm, Act
from monai.networks.nets.unet import UNet
+
+TE... | diff --git a/docs/source/highlights.md b/docs/source/highlights.md
index f7895bf65d..113fc1a1c5 100644
--- a/docs/source/highlights.md
+++ b/docs/source/highlights.md
@@ -72,7 +72,16 @@ torch.backends.cudnn.benchmark = False
There are domain-specific loss functions in the medical research area which are different from... | [
{
"components": [
{
"doc": "Factory object for creating layers, this uses given factory functions to actually produce the types or constructing\ncallables. These functions are referred to by name and can be added at any time.",
"lines": [
68,
141
],
"nam... | [
"tests/test_unet.py::TestUNET::test_shape_0",
"tests/test_unet.py::TestUNET::test_shape_1",
"tests/test_unet.py::TestUNET::test_shape_2",
"tests/test_unet.py::TestUNET::test_shape_3",
"tests/test_unet.py::TestUNET::test_shape_4",
"tests/test_unet.py::TestUNET::test_shape_5",
"tests/test_unet.py::TestUNE... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
81 layer factory
Fixes #81.
### Description
Adds a new layer factory mechanism which is extensible and more generic.
### Status
**Work in progress**
### Types of changes
<!--- Put an `x... | Here is the discussion in the issues of the pull request.
<issues>
Create layer factory mechanism
----------
--------------------
</issues> | e73257caa79309dcce1e93abf1632f4bfd75b11f |
joke2k__faker-1122 | 1,122 | joke2k/faker | null | 28ff1840387c3ff020a442143a10c54820d697f0 | 2020-02-27T15:02:22Z | diff --git a/faker/providers/automotive/fr_FR/__init__.py b/faker/providers/automotive/fr_FR/__init__.py
new file mode 100644
index 0000000000..3fc9c7a210
--- /dev/null
+++ b/faker/providers/automotive/fr_FR/__init__.py
@@ -0,0 +1,11 @@
+from .. import Provider as AutomotiveProvider
+
+
+class Provider(AutomotiveProvid... | diff --git a/tests/providers/test_automotive.py b/tests/providers/test_automotive.py
index 7ad2874fc7..ddffb94500 100644
--- a/tests/providers/test_automotive.py
+++ b/tests/providers/test_automotive.py
@@ -133,3 +133,16 @@ def test_ru_RU_plate_format(self):
def test_vehicle_category(self):
category = sel... | [
{
"components": [
{
"doc": "",
"lines": [
4,
10
],
"name": "Provider",
"signature": "class Provider(AutomotiveProvider):",
"type": "class"
}
],
"file": "faker/providers/automotive/fr_FR/__init__.py"
}
] | [
"tests/providers/test_automotive.py::TestFrFR::test_fr_FR_plate_format"
] | [
"tests/providers/test_automotive.py::TestPtBR::test_plate_has_been_generated",
"tests/providers/test_automotive.py::TestPtPT::test_pt_PT_plate_format",
"tests/providers/test_automotive.py::TestHuHU::test_hu_HU_plate_format",
"tests/providers/test_automotive.py::TestDeDe::test_de_DE_plate_format",
"tests/pro... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add generating random french licence plates
* Add french licence plates
* Add corresponding tests
Closes #1118
----------
</request>
There are several new functions or classes that need to be... | Here is the discussion in the issues of the pull request.
<issues>
Faker missing french licence plate
I noticed that Faker was missing the implementation of French licence plates.
I would interested in adding this new feature :)
----------
--------------------
</issues> | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | |
joke2k__faker-1119 | 1,119 | joke2k/faker | null | 28ff1840387c3ff020a442143a10c54820d697f0 | 2020-02-27T06:04:52Z | diff --git a/faker/providers/internet/__init__.py b/faker/providers/internet/__init__.py
index 79a978163b..60dab82429 100644
--- a/faker/providers/internet/__init__.py
+++ b/faker/providers/internet/__init__.py
@@ -76,6 +76,10 @@ class Provider(BaseProvider):
'.html', '.html', '.html', '.htm', '.htm', '.php', ... | diff --git a/tests/test_factory.py b/tests/test_factory.py
index 148b1790c3..a93df7cffe 100644
--- a/tests/test_factory.py
+++ b/tests/test_factory.py
@@ -876,6 +876,19 @@ def test_port_number(self):
assert 1024 <= faker.port_number(is_user=True) <= 49151
assert 49152 <= faker.port_number(is_d... | [
{
"components": [
{
"doc": "Returns random HTTP method\nhttps://developer.mozilla.org/en-US/docs/Web/HTTP/Methods\n\n:rtype: str",
"lines": [
228,
235
],
"name": "Provider.http_method",
"signature": "def http_method(self):",
"type": "func... | [
"tests/test_factory.py::FactoryTestCase::test_http_method"
] | [
"tests/test_factory.py::FactoryTestCase::test_add_provider_gives_priority_to_newly_added_provider",
"tests/test_factory.py::FactoryTestCase::test_binary",
"tests/test_factory.py::FactoryTestCase::test_cli_seed",
"tests/test_factory.py::FactoryTestCase::test_cli_seed_with_repeat",
"tests/test_factory.py::Fac... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add HTTP methods random generator
### What does this changes
Add HTTP method random generator in internet provider. Also test for it.
----------
</request>
There are several new functions or ... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
sympy__sympy-18671 | 18,671 | sympy/sympy | 1.6 | 94b9de669f41dd76416c50f8097a22fdc9e8513d | 2020-02-17T00:09:13Z | diff --git a/sympy/ntheory/__init__.py b/sympy/ntheory/__init__.py
index 819cc748ede5..e35a76b75f9d 100644
--- a/sympy/ntheory/__init__.py
+++ b/sympy/ntheory/__init__.py
@@ -6,7 +6,8 @@
randprime, Sieve, sieve, primorial, cycle_length, composite, compositepi
from .primetest import isprime
from .factor_ import d... | diff --git a/sympy/ntheory/tests/test_factor_.py b/sympy/ntheory/tests/test_factor_.py
index 8555157864e0..afea0d38eb04 100644
--- a/sympy/ntheory/tests/test_factor_.py
+++ b/sympy/ntheory/tests/test_factor_.py
@@ -4,7 +4,7 @@
from sympy.ntheory import (totient,
factorint, primefactors, divisors, nextprime,
- ... | [
{
"components": [
{
"doc": "return the largest integer ``m`` such that ``p**m`` divides ``n!``\nwithout calculating the factorial of ``n``.\n\n\nExamples\n========\n\n>>> from sympy.ntheory import multiplicity_in_factorial\n>>> from sympy import factorial\n\n>>> multiplicity_in_factorial(2, 3)\n1\... | [
"test_trailing_bitcount",
"test_multiplicity",
"test_multiplicity_in_factorial",
"test_perfect_power",
"test_factorint",
"test_divisors_and_divisor_count",
"test_proper_divisors_and_proper_divisor_count",
"test_udivisors_and_udivisor_count",
"test_issue_6981",
"test_totient",
"test_reduced_totie... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
ntheory: Added a new function multiplicity_in_factorial()
Reference: #10904
Closes: #10904
#### Brief description of what is fixed or changed
This pull request adds a function named multiplic... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
sympy__sympy-18667 | 18,667 | sympy/sympy | 1.6 | cd86e3c3335a7f43379185c239619c576522ef4a | 2020-02-16T11:52:24Z | diff --git a/sympy/combinatorics/schur_number.py b/sympy/combinatorics/schur_number.py
new file mode 100644
index 000000000000..1d8947e2809f
--- /dev/null
+++ b/sympy/combinatorics/schur_number.py
@@ -0,0 +1,152 @@
+"""
+The Schur number S(k) is the largest integer n for which the interval [1,n]
+can be partitioned int... | diff --git a/sympy/combinatorics/tests/test_schur_number.py b/sympy/combinatorics/tests/test_schur_number.py
new file mode 100644
index 000000000000..77d1d7aca0ad
--- /dev/null
+++ b/sympy/combinatorics/tests/test_schur_number.py
@@ -0,0 +1,55 @@
+from sympy.core import S, Rational
+from sympy.combinatorics.schur_numbe... | [
{
"components": [
{
"doc": "This function creates a SchurNumber object\nwhich is evaluated for k <= 4 otherwise only\nthe lower bound information can be retrieved.\n\nExamples\n========\n\n>>> from sympy.combinatorics.schur_number import SchurNumber\n\nSince S(3) = 13, hence the output is a number... | [
"test_schur_partition"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Addition of schur number in combinatorics
#### References to other Issues or PRs
Revives #14493
Closes #14493
#### Brief description of what is fixed or changed
--> Added schur number in combi... | Here is the discussion in the issues of the pull request.
<issues>
Added new feature Schur_Number
<!-- I have added a new feature in the combinatorics module the Schur_number -->
The Schur number S(k) is the largest integer n for which the interval [1,n] can be partitioned into k sum-free sets. http://mathworld.... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
sympy__sympy-18659 | 18,659 | sympy/sympy | 1.6 | b26825a9c906bb3d7e4f043600c3e25bb1afd6a8 | 2020-02-15T05:16:59Z | diff --git a/doc/src/modules/ntheory.rst b/doc/src/modules/ntheory.rst
index 51654726a16e..aa86c6df8131 100644
--- a/doc/src/modules/ntheory.rst
+++ b/doc/src/modules/ntheory.rst
@@ -177,6 +177,9 @@ Ntheory Functions Reference
.. automodule:: sympy.ntheory.continued_fraction
:members:
+.. automodule:: sympy.nthe... | diff --git a/sympy/ntheory/tests/test_digits.py b/sympy/ntheory/tests/test_digits.py
new file mode 100644
index 000000000000..23f22d7a5dd3
--- /dev/null
+++ b/sympy/ntheory/tests/test_digits.py
@@ -0,0 +1,81 @@
+import os
+import random
+import re
+import sys
+
+from collections import Counter
+
+from sympy.ntheory imp... | diff --git a/doc/src/modules/ntheory.rst b/doc/src/modules/ntheory.rst
index 51654726a16e..aa86c6df8131 100644
--- a/doc/src/modules/ntheory.rst
+++ b/doc/src/modules/ntheory.rst
@@ -177,6 +177,9 @@ Ntheory Functions Reference
.. automodule:: sympy.ntheory.continued_fraction
:members:
+.. automodule:: sympy.nthe... | [
{
"components": [
{
"doc": "Returs an ordered dict. frequency counter for the digits of a decimal\ninteger ``n`` representing an integer in a base ``base``, where the\nbase digits are mapped to the positive decimal integers, e.g. the\nbase ``11`` integer ``10321``, with digits ``10``, ``3``, ``2``... | [
"test_count_digits",
"test_is_palindromic__base_smaller_than_2__raises_value_error",
"test_is_palindromic__n_not_int_literal__raises_type_error",
"test_is_palindromic__valid_palindrome__returns_true"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add ntheory module for arithmetic properties of integers
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References ... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
prometheus__client_python-512 | 512 | prometheus/client_python | null | ce7063fc2957716aa6fa9dc4f49bd970ad1249ed | 2020-02-14T20:51:14Z | diff --git a/README.md b/README.md
index c7161fd4..586f5315 100644
--- a/README.md
+++ b/README.md
@@ -306,6 +306,19 @@ from prometheus_client import start_wsgi_server
start_wsgi_server(8000)
```
+#### ASGI
+
+To use Prometheus with [ASGI](http://asgi.readthedocs.org/en/latest/), there is
+`make_asgi_app` which cre... | diff --git a/tests/test_asgi.py b/tests/test_asgi.py
new file mode 100644
index 00000000..b2d9a70f
--- /dev/null
+++ b/tests/test_asgi.py
@@ -0,0 +1,123 @@
+from __future__ import absolute_import, unicode_literals
+
+import sys
+from unittest import TestCase
+
+from prometheus_client import CollectorRegistry, Counter, ... | diff --git a/README.md b/README.md
index c7161fd4..586f5315 100644
--- a/README.md
+++ b/README.md
@@ -306,6 +306,19 @@ from prometheus_client import start_wsgi_server
start_wsgi_server(8000)
```
+#### ASGI
+
+To use Prometheus with [ASGI](http://asgi.readthedocs.org/en/latest/), there is
+`make_asgi_app` which cre... | [
{
"components": [
{
"doc": "Create a ASGI app which serves the metrics from a registry.",
"lines": [
7,
34
],
"name": "make_asgi_app",
"signature": "def make_asgi_app(registry=REGISTRY):",
"type": "function"
}
],
"file": "pr... | [
"tests/test_wsgi.py::WSGITest::test_report_metrics_1",
"tests/test_wsgi.py::WSGITest::test_report_metrics_2",
"tests/test_wsgi.py::WSGITest::test_report_metrics_3",
"tests/test_wsgi.py::WSGITest::test_report_metrics_4"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added ASGI application
This PR introduces an ASGI counterpart to the current WSGI app.
The ASGI code itself is moved into a seperate module `asgi.py`, which is conditionally included as it is only ... | 09a5ae30602a7a81f6174dae4ba08b93ee7feed2 | |
fairlearn__fairlearn-288 | 288 | fairlearn/fairlearn | null | 2ffe87c71ebbde66f058c758da39258cc59fafc6 | 2020-02-10T18:21:59Z | diff --git a/fairlearn/_input_validation.py b/fairlearn/_input_validation.py
index 6b44e0aa4..38bfa2f6e 100644
--- a/fairlearn/_input_validation.py
+++ b/fairlearn/_input_validation.py
@@ -3,18 +3,113 @@
import numpy as np
import pandas as pd
+from sklearn.utils.validation import check_X_y, check_consistent_length,... | diff --git a/test/unit/constants.py b/test/unit/constants.py
new file mode 100644
index 000000000..bacb39ce4
--- /dev/null
+++ b/test/unit/constants.py
@@ -0,0 +1,6 @@
+# Copyright (c) Microsoft Corporation. All rights reserved.
+# Licensed under the MIT License.
+
+
+MULTIPLE_SENSITIVE_FEATURE_COMPRESSION_SKIP_REASON ... | [
{
"components": [
{
"doc": "Validate input data and return the data in an appropriate format.\n\n:param X: The feature matrix\n:type X: numpy.ndarray or pandas.DataFrame\n:param y: The label vector\n:type y: numpy.ndarray, pandas.DataFrame, pandas.Series, or list\n:param expect_y: if True y needs ... | [
"test/unit/postprocessing/test_curve_utilities.py::test_assert_interpolated_curve",
"test/unit/postprocessing/test_curve_utilities.py::test_interpolate_curve",
"test/unit/postprocessing/test_curve_utilities.py::test_convex_hull[base_points0-expected_remaining_indices0]",
"test/unit/postprocessing/test_curve_u... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
multiple sensitive features - postprocessing
This is the trimmed down version of #280 for just postprocessing. After this is merged I'll update #280 which will then only be about reductions.
--------... | Here is the discussion in the issues of the pull request.
<issues>
Enhancements for GroupMetricResult and GroupMetricSet
Both `GroupMetricResult` and `GroupMetricSet` should have comparators implemented, to ease equality checking in tests.
Also, `GroupMetricSet` should take care of ensuring that the `groups` propert... | 403da1fec74bdf2da28dc49487ccd72caa6f6976 | |
sympy__sympy-18599 | 18,599 | sympy/sympy | 1.6 | 1a66b86d23cfc606ad3ad4185aaec40ee9b822a5 | 2020-02-07T16:25:08Z | diff --git a/sympy/solvers/diophantine/diophantine.py b/sympy/solvers/diophantine/diophantine.py
index 318dc5f2fbc0..be94c6fcd30d 100644
--- a/sympy/solvers/diophantine/diophantine.py
+++ b/sympy/solvers/diophantine/diophantine.py
@@ -1,7 +1,8 @@
from __future__ import print_function, division
from sympy.core.add i... | diff --git a/sympy/solvers/diophantine/tests/test_diophantine.py b/sympy/solvers/diophantine/tests/test_diophantine.py
index f5b0d39d6c1c..d4c31b8e5b1b 100644
--- a/sympy/solvers/diophantine/tests/test_diophantine.py
+++ b/sympy/solvers/diophantine/tests/test_diophantine.py
@@ -15,7 +15,7 @@
classify_diop, base_so... | [
{
"components": [
{
"doc": "Container for a set of solutions to a particular diophantine equation.\n\nThe base representation is a set of tuples representing each of the solutions.\n\nParameters\n==========\n\nsymbols : list\n List of free symbols in the original equation.\nparameters: list (op... | [
"test_input_format",
"test_nosols",
"test_univariate",
"test_classify_diop",
"test_linear",
"test_quadratic_simple_hyperbolic_case",
"test_quadratic_elliptical_case",
"test_quadratic_parabolic_case",
"test_quadratic_perfect_square",
"test_quadratic_non_perfect_square",
"test_issue_9106",
"test... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Update internal diophantine solvers to have consistent return types
#### References to other Issues or PRs
#18493
#### Brief description of what is fixed or changed
Add a new class `DiophantineSo... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
sympy__sympy-18595 | 18,595 | sympy/sympy | 1.6 | b17ef6effe278d5b861d65896cc53442a6370d8f | 2020-02-07T05:43:51Z | diff --git a/sympy/discrete/recurrences.py b/sympy/discrete/recurrences.py
index bddaf14378da..291123f30fd4 100644
--- a/sympy/discrete/recurrences.py
+++ b/sympy/discrete/recurrences.py
@@ -109,6 +109,32 @@ def linrec(coeffs, init, n):
else:
b += [S.Zero]*(k - len(b)) # remaining initial values default t... | diff --git a/sympy/matrices/tests/test_commonmatrix.py b/sympy/matrices/tests/test_commonmatrix.py
index df8bbe858af5..2faa1610f033 100644
--- a/sympy/matrices/tests/test_commonmatrix.py
+++ b/sympy/matrices/tests/test_commonmatrix.py
@@ -757,7 +757,8 @@ def test_power():
assert A**1 == A
assert (ArithmeticOn... | [
{
"components": [
{
"doc": "Compute the coefficients of n'th term in linear recursion\nsequence defined by c.\n\n`x^k = c_0 x^{k-1} + c_1 x^{k-2} + \\cdots + c_{k-1}`.\n\nIt computes the coefficients by using binary exponentiation.\nThis function is used by `linrec` and `_eval_pow_by_cayley`.\n\nP... | [
"test_power"
] | [
"test__MinimalMatrix",
"test_vec",
"test_tolist",
"test_row_col_del",
"test_get_diag_blocks1",
"test_get_diag_blocks2",
"test_shape",
"test_reshape",
"test_row_col",
"test_row_join",
"test_col_join",
"test_row_insert",
"test_col_insert",
"test_extract",
"test_hstack",
"test_vstack",
... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Faster matrix exponentiation
<!-- BEGIN RELEASE NOTES -->
* matrices
* Faster Matrix exponentiation using Cayley Hamilton Theorem
<!-- END RELEASE NOTES -->
----------
When A = Matrix([[2, 3, 4,... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
scikit-learn__scikit-learn-16326 | 16,326 | scikit-learn/scikit-learn | 0.24 | 87eea7d409592e6b586bf2c9c1749f57c94c055b | 2020-01-30T19:37:38Z | diff --git a/doc/whats_new/v0.23.rst b/doc/whats_new/v0.23.rst
index 04e10be72cd54..fcc55e68bb985 100644
--- a/doc/whats_new/v0.23.rst
+++ b/doc/whats_new/v0.23.rst
@@ -621,6 +621,11 @@ Changelog
:mod:`sklearn.naive_bayes`
.............................
+- |Enhancement| Adds a parameter `min_categories` to
+ :class... | diff --git a/sklearn/tests/test_naive_bayes.py b/sklearn/tests/test_naive_bayes.py
index a9f558eeae703..e95eccc0c747d 100644
--- a/sklearn/tests/test_naive_bayes.py
+++ b/sklearn/tests/test_naive_bayes.py
@@ -676,6 +676,7 @@ def test_categoricalnb():
clf = CategoricalNB(alpha=1, fit_prior=False)
clf.fit(X3,... | diff --git a/doc/whats_new/v0.23.rst b/doc/whats_new/v0.23.rst
index 04e10be72cd54..fcc55e68bb985 100644
--- a/doc/whats_new/v0.23.rst
+++ b/doc/whats_new/v0.23.rst
@@ -621,6 +621,11 @@ Changelog
:mod:`sklearn.naive_bayes`
.............................
+- |Enhancement| Adds a parameter `min_categories` to
+ :class... | [
{
"components": [
{
"doc": "",
"lines": [
1225,
1246
],
"name": "CategoricalNB._validate_n_categories",
"signature": "def _validate_n_categories(X, min_categories):",
"type": "function"
}
],
"file": "sklearn/naive_bayes.py"
... | [
"sklearn/tests/test_naive_bayes.py::test_categoricalnb",
"sklearn/tests/test_naive_bayes.py::test_categoricalnb_with_min_categories[3-exp_X1_count0-exp_X2_count0-new_X0-exp_n_categories_0]",
"sklearn/tests/test_naive_bayes.py::test_categoricalnb_with_min_categories[min_categories1-exp_X1_count1-exp_X2_count1-ne... | [
"sklearn/tests/test_naive_bayes.py::test_gnb",
"sklearn/tests/test_naive_bayes.py::test_gnb_prior",
"sklearn/tests/test_naive_bayes.py::test_gnb_sample_weight",
"sklearn/tests/test_naive_bayes.py::test_gnb_neg_priors",
"sklearn/tests/test_naive_bayes.py::test_gnb_priors",
"sklearn/tests/test_naive_bayes.p... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
ENH Add min_categories parameter for each feature to CategoricalNB
<!--
Thanks for contributing a pull request! Please ensure you have taken a look at
the contribution guidelines: https://github.com... | Here is the discussion in the issues of the pull request.
<issues>
[MRG] Raise an error for categories unseen during training
#### Reference Issues/PRs
Fixes #16028
#### What does this implement/fix? Explain your changes.
Raises an error when CategoricalNB encounters new categories present during testing but never... | 54ce4222694819ad52d544ce5cba5da274c34ab7 |
sympy__sympy-18354 | 18,354 | sympy/sympy | 1.6 | 77c9b6904d8ad45ac5b3c624adc5b4abcab41be5 | 2020-01-16T10:11:47Z | diff --git a/sympy/combinatorics/perm_groups.py b/sympy/combinatorics/perm_groups.py
index 085005fcb34d..c25e15c5d718 100644
--- a/sympy/combinatorics/perm_groups.py
+++ b/sympy/combinatorics/perm_groups.py
@@ -2567,6 +2567,92 @@ def minimal_block(self, points):
new_reps = {}
return [new_reps.setdefau... | diff --git a/sympy/combinatorics/tests/test_perm_groups.py b/sympy/combinatorics/tests/test_perm_groups.py
index 8deda10723ff..ad6eaa941f97 100644
--- a/sympy/combinatorics/tests/test_perm_groups.py
+++ b/sympy/combinatorics/tests/test_perm_groups.py
@@ -1097,3 +1097,22 @@ def test_is_symmetric():
a = Permutation(... | [
{
"components": [
{
"doc": "Return the conjugacy class of an element in the group.\n\nThe conjugacy class of an element ``g`` in a group ``G`` is the set of\nelements ``x`` in ``G`` that are conjugate with ``g``, i.e. for which\n\n ``g = xax^{-1}``\n\nfor some ``a`` in ``G``.\n\nNote that conju... | [
"test_conjugacy_class"
] | [
"test_has",
"test_generate",
"test_order",
"test_equality",
"test_stabilizer",
"test_center",
"test_centralizer",
"test_coset_rank",
"test_coset_factor",
"test_orbits",
"test_is_normal",
"test_eq",
"test_derived_subgroup",
"test_is_solvable",
"test_rubik1",
"test_direct_product",
"te... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Combinatorics: Added naive algorithm for computing conjugacy classes in permutation groups
Reference: https://github.com/sympy/sympy/pull/9160
#### Brief description of what is fixed or changed
... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
sympy__sympy-18335 | 18,335 | sympy/sympy | 1.6 | 6cd0b9fade27be198ee24987be0eac709413d297 | 2020-01-14T17:32:57Z | diff --git a/sympy/geometry/polygon.py b/sympy/geometry/polygon.py
index dd6a0b74b1d8..1f55d0b608e2 100644
--- a/sympy/geometry/polygon.py
+++ b/sympy/geometry/polygon.py
@@ -1370,6 +1370,40 @@ def __contains__(self, o):
return False
+ def bisectors(p, prec=None):
+ """Returns angle bisectors of ... | diff --git a/sympy/geometry/tests/test_polygon.py b/sympy/geometry/tests/test_polygon.py
index 44fe30c5562e..42ca46b7d715 100644
--- a/sympy/geometry/tests/test_polygon.py
+++ b/sympy/geometry/tests/test_polygon.py
@@ -1,8 +1,9 @@
-from sympy import Abs, Rational, Float, S, Symbol, symbols, cos, pi, sqrt, oo
+from symp... | [
{
"components": [
{
"doc": "Returns angle bisectors of a polygon. If prec is given\nthen approximate the point defining the ray to that precision.\n\nThe distance between the points defining the bisector ray is 1.\n\nExamples\n========\n\n>>> from sympy import Polygon, Point\n>>> p = Polygon(Point... | [
"test_bisectors"
] | [
"test_convex_hull",
"test_encloses",
"test_triangle_kwargs",
"test_transform",
"test_reflect",
"test_incenter",
"test_inradius",
"test_incircle",
"test_exradii",
"test_medians",
"test_medial",
"test_nine_point_circle",
"test_eulerline",
"test_intersection",
"test_parameter_value",
"tes... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[feature] Add bisectors method for Polygon class
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
pydata__xarray-3691 | 3,691 | pydata/xarray | 0.12 | d4b7a608bab0e7c140937b0b59ca45115d205145 | 2020-01-13T17:15:23Z | diff --git a/doc/groupby.rst b/doc/groupby.rst
index d0c0b1849f9..804799aebd1 100644
--- a/doc/groupby.rst
+++ b/doc/groupby.rst
@@ -63,6 +63,12 @@ You can also iterate over groups in ``(label, group)`` pairs:
list(ds.groupby("letters"))
+You can index out a particular group:
+
+.. ipython:: python
+
+ ds.g... | diff --git a/xarray/tests/test_groupby.py b/xarray/tests/test_groupby.py
index 85f729a9f7a..5ef7677c499 100644
--- a/xarray/tests/test_groupby.py
+++ b/xarray/tests/test_groupby.py
@@ -549,4 +549,17 @@ def test_groupby_none_group_name():
assert "group" in mean.dims
+def test_groupby_getitem(dataset):
+
+ as... | diff --git a/doc/groupby.rst b/doc/groupby.rst
index d0c0b1849f9..804799aebd1 100644
--- a/doc/groupby.rst
+++ b/doc/groupby.rst
@@ -63,6 +63,12 @@ You can also iterate over groups in ``(label, group)`` pairs:
list(ds.groupby("letters"))
+You can index out a particular group:
+
+.. ipython:: python
+
+ ds.g... | [
{
"components": [
{
"doc": "Get DataArray or Dataset corresponding to a particular group label.",
"lines": [
426,
430
],
"name": "GroupBy.__getitem__",
"signature": "def __getitem__(self, key):",
"type": "function"
}
],
"fil... | [
"xarray/tests/test_groupby.py::test_groupby_getitem"
] | [
"xarray/tests/test_groupby.py::test_consolidate_slices",
"xarray/tests/test_groupby.py::test_groupby_dims_property",
"xarray/tests/test_groupby.py::test_multi_index_groupby_map",
"xarray/tests/test_groupby.py::test_multi_index_groupby_sum",
"xarray/tests/test_groupby.py::test_groupby_da_datetime",
"xarray... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implement GroupBy.__getitem__
<!-- Feel free to remove check-list items aren't relevant to your change -->
- [x] Tests added
- [x] Passes `black . && mypy . && flake8`
- [x] Fully documented, ... | 1c198a191127c601d091213c4b3292a8bb3054e1 | |
sympy__sympy-18213 | 18,213 | sympy/sympy | 1.6 | b4f1aa3540fe68d078d76e78ba59d022dd6df39f | 2020-01-03T16:00:16Z | diff --git a/sympy/geometry/point.py b/sympy/geometry/point.py
index 7d8e40e617ad..c3d3da39c9b0 100644
--- a/sympy/geometry/point.py
+++ b/sympy/geometry/point.py
@@ -1029,6 +1029,21 @@ def translate(self, x=0, y=0):
"""
return Point(self.x + x, self.y + y)
+ @property
+ def coordinates(self):... | diff --git a/sympy/geometry/tests/test_point.py b/sympy/geometry/tests/test_point.py
index 7d6ea9a1ca95..cf16e11d232d 100644
--- a/sympy/geometry/tests/test_point.py
+++ b/sympy/geometry/tests/test_point.py
@@ -172,6 +172,16 @@ def test_point3D():
raises(ValueError, lambda: Point3D(0, 0, 0) + 10)
+ # Test c... | [
{
"components": [
{
"doc": "Returns the two coordinates of the Point.\n\nExamples\n========\n\n>>> from sympy import Point2D\n>>> p = Point2D(0, 1)\n>>> p.coordinates\n(0, 1)",
"lines": [
1033,
1045
],
"name": "Point2D.coordinates",
"signature": ... | [
"test_point3D",
"test_Point2D"
] | [
"test_point",
"test_issue_9214",
"test_issue_11617",
"test_transform",
"test_concyclic_doctest_bug",
"test_arguments",
"test_unit",
"test_dot",
"test__normalize_dimension"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Adding a property for Point's coordinates
From my point of view, there should be a method for getting the coordinates of a point. Currently, the way I do this is using something like this:
`(point.x,... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
matplotlib__matplotlib-16049 | 16,049 | matplotlib/matplotlib | 3.1 | 160f711245081d2004c030946f85868135e58720 | 2019-12-30T21:47:56Z | diff --git a/lib/matplotlib/gridspec.py b/lib/matplotlib/gridspec.py
index 678f03e4a647..29c8f2d6864d 100644
--- a/lib/matplotlib/gridspec.py
+++ b/lib/matplotlib/gridspec.py
@@ -522,6 +522,11 @@ def __init__(self, gridspec, num1, num2=None):
else:
self._layoutbox = None
+ def __repr__(self):... | diff --git a/lib/matplotlib/tests/test_gridspec.py b/lib/matplotlib/tests/test_gridspec.py
index 70d1ee132851..fa931ce086e5 100644
--- a/lib/matplotlib/tests/test_gridspec.py
+++ b/lib/matplotlib/tests/test_gridspec.py
@@ -24,3 +24,8 @@ def test_height_ratios():
"""
with pytest.raises(ValueError):
gr... | [
{
"components": [
{
"doc": "",
"lines": [
525,
528
],
"name": "SubplotSpec.__repr__",
"signature": "def __repr__(self):",
"type": "function"
}
],
"file": "lib/matplotlib/gridspec.py"
}
] | [
"lib/matplotlib/tests/test_gridspec.py::test_repr"
] | [
"lib/matplotlib/tests/test_gridspec.py::test_equal",
"lib/matplotlib/tests/test_gridspec.py::test_width_ratios",
"lib/matplotlib/tests/test_gridspec.py::test_height_ratios"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add __repr__ to SubplotSpec.
This allows printing subplotspecs as e.g. `GridSpec(3, 3)[1:3, 2:3]`
which can help with debugging. (Note that `GridSpec.__repr__` was
already implemented previously.)
... | c9be5a6985cc919703959902e9a8d795cfde03ad | ||
sympy__sympy-18182 | 18,182 | sympy/sympy | 1.6 | 121ccd76879eabcb9567cb6f88d976f761f171d1 | 2019-12-30T19:57:44Z | diff --git a/sympy/physics/quantum/__init__.py b/sympy/physics/quantum/__init__.py
index e20da6c56122..bf08e1f7a383 100644
--- a/sympy/physics/quantum/__init__.py
+++ b/sympy/physics/quantum/__init__.py
@@ -22,7 +22,8 @@
'get_basis', 'enumerate_states',
'KetBase', 'BraBase', 'StateBase', 'State', 'Ket', 'Br... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 0ae4924e81c2..d10294200798 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -3665,6 +3665,21 @@ def test_sympy__physics__quantum__state__StateBase():
assert _test_args(StateBase(0))
+def test_sympy... | [
{
"components": [
{
"doc": "General abstract quantum state used as a base class for Ket and Bra.",
"lines": [
623,
625
],
"name": "OrthogonalState",
"signature": "class OrthogonalState(State, StateBase):",
"type": "class"
},
{... | [
"test_sympy__physics__quantum__state__OrthogonalBra",
"test_sympy__physics__quantum__state__OrthogonalKet",
"test_sympy__physics__quantum__state__OrthogonalState",
"test_ket",
"test_bra",
"test_ops",
"test_time_dep_ket",
"test_time_dep_bra",
"test_bra_ket_dagger",
"test_wavefunction"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add OrthogonalBra and OrthogonalKet
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other Issues or PR... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
sympy__sympy-18173 | 18,173 | sympy/sympy | 1.6 | 921379b4a781a781bf2543f81e785ca45d7370f1 | 2019-12-29T22:47:40Z | diff --git a/sympy/stats/__init__.py b/sympy/stats/__init__.py
index cfc55bdf154d..f2b5c81a806e 100644
--- a/sympy/stats/__init__.py
+++ b/sympy/stats/__init__.py
@@ -67,7 +67,7 @@
'StochasticProcess', 'DiscreteTimeStochasticProcess',
'DiscreteMarkovChain', 'TransitionMatrixOf', 'StochasticStateSpaceOf',
- ... | diff --git a/sympy/core/tests/test_args.py b/sympy/core/tests/test_args.py
index 410ab0855315..0dd4507e301f 100644
--- a/sympy/core/tests/test_args.py
+++ b/sympy/core/tests/test_args.py
@@ -1645,6 +1645,10 @@ def test_sympy__stats__stochastic_process_types__ContinuousMarkovChain():
from sympy import MatrixSymbol
... | [
{
"components": [
{
"doc": "Converts an arbitrary expression to a type that can be used inside SymPy.\nAs generally strings are unwise to use in the expressions,\nit returns the Symbol of argument if the string type argument is passed.\n\nParameters\n=========\n\narg: The parameter to be converted... | [
"test_sympy__stats__stochastic_process_types__BernoulliProcess",
"test_DiscreteMarkovChain",
"test_ContinuousMarkovChain"
] | [
"test_all_classes_are_tested",
"test_sympy__assumptions__assume__AppliedPredicate",
"test_sympy__assumptions__assume__Predicate",
"test_sympy__assumptions__sathandlers__UnevaluatedOnFree",
"test_sympy__assumptions__sathandlers__AllArgs",
"test_sympy__assumptions__sathandlers__AnyArgs",
"test_sympy__assu... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added Bernoulli Process
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
Added Bernoulli Process, discrete time and discrete... | Here is the discussion in the issues of the pull request.
<issues>
Add Bernoulli process to sympy.stats.stochastic_process_types
Since, the stochastic processes have already been added to `sympy.stats`. So, the purpose of https://github.com/sympy/sympy/pull/15058 is partially over. We now need to add Bernoulli process ... | 6daf556517f9fa61a7805fe93d8a366688781617 | |
joke2k__faker-1088 | 1,088 | joke2k/faker | null | 6c4fde452829f539e27432a07de7ca7ba4289712 | 2019-12-29T13:17:07Z | diff --git a/faker/providers/internet/__init__.py b/faker/providers/internet/__init__.py
index 1628e9bbfc..a24f367a23 100644
--- a/faker/providers/internet/__init__.py
+++ b/faker/providers/internet/__init__.py
@@ -480,6 +480,25 @@ def mac_address(self):
mac = [self.generator.random.randint(0x00, 0xff) for _ i... | diff --git a/tests/test_factory.py b/tests/test_factory.py
index 92c09cc754..aa7bb8c5da 100644
--- a/tests/test_factory.py
+++ b/tests/test_factory.py
@@ -876,6 +876,15 @@ def test_ipv6(self):
re.compile(r'^([0-9a-f]{0,4}:){2,7}[0-9a-f]{0,4}/\d{1,3}$').search(
address))
+ def ... | [
{
"components": [
{
"doc": "Returns a network port number\nhttps://tools.ietf.org/html/rfc6335\n\n:param is_system: System or well-known ports\n:param is_user: User or registered ports\n:param is_dynamic: Dynamic / private / ephemeral ports\n:rtype: int",
"lines": [
483,
... | [
"tests/test_factory.py::FactoryTestCase::test_port_number"
] | [
"tests/test_factory.py::FactoryTestCase::test_add_provider_gives_priority_to_newly_added_provider",
"tests/test_factory.py::FactoryTestCase::test_binary",
"tests/test_factory.py::FactoryTestCase::test_cli_seed",
"tests/test_factory.py::FactoryTestCase::test_cli_seed_with_repeat",
"tests/test_factory.py::Fac... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add port number gen
### What does this changes
Add network port number generator and test for it
----------
</request>
There are several new functions or classes that need to be implemented, ... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
sympy__sympy-18085 | 18,085 | sympy/sympy | 1.6 | 9da013ad0ddc3cd96fe505f2e47c63e372040916 | 2019-12-20T06:24:57Z | diff --git a/doc/src/modules/ntheory.rst b/doc/src/modules/ntheory.rst
index 68994cdbb647..51654726a16e 100644
--- a/doc/src/modules/ntheory.rst
+++ b/doc/src/modules/ntheory.rst
@@ -56,8 +56,12 @@ Ntheory Functions Reference
.. autofunction:: divisors
+.. autofunction:: proper_divisors
+
.. autofunction:: diviso... | diff --git a/sympy/ntheory/tests/test_factor_.py b/sympy/ntheory/tests/test_factor_.py
index 59d81096ff45..eb58251209bb 100644
--- a/sympy/ntheory/tests/test_factor_.py
+++ b/sympy/ntheory/tests/test_factor_.py
@@ -8,10 +8,16 @@
primerange, pollard_rho, perfect_power, multiplicity,
trailing, divisor_count, pr... | diff --git a/doc/src/modules/ntheory.rst b/doc/src/modules/ntheory.rst
index 68994cdbb647..51654726a16e 100644
--- a/doc/src/modules/ntheory.rst
+++ b/doc/src/modules/ntheory.rst
@@ -56,8 +56,12 @@ Ntheory Functions Reference
.. autofunction:: divisors
+.. autofunction:: proper_divisors
+
.. autofunction:: diviso... | [
{
"components": [
{
"doc": "Return all divisors of n except n, sorted by default.\nIf generator is ``True`` an unordered generator is returned.\n\nExamples\n========\n\n>>> from sympy import proper_divisors, proper_divisor_count\n>>> proper_divisors(24)\n[1, 2, 3, 4, 6, 8, 12]\n>>> proper_divisor_... | [
"test_trailing_bitcount",
"test_multiplicity",
"test_perfect_power",
"test_factorint",
"test_divisors_and_divisor_count",
"test_proper_divisors_and_proper_divisor_count",
"test_udivisors_and_udivisor_count",
"test_issue_6981",
"test_totient",
"test_reduced_totient",
"test_divisor_sigma",
"test... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added proper_divisor and proper_divisor_count functions
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References t... | Here is the discussion in the issues of the pull request.
<issues>
Function for proper divisors
A function returning the list of proper divisors seems to be missing
----------
--------------------
</issues> | 6daf556517f9fa61a7805fe93d8a366688781617 |
pygments__pygments-1336 | 1,336 | pygments/pygments | null | 27d7a3073073419c60df423310382bf55e7ed867 | 2019-12-11T05:38:01Z | diff --git a/doc/languages.rst b/doc/languages.rst
index c976b6af69..f83b60ffd3 100644
--- a/doc/languages.rst
+++ b/doc/languages.rst
@@ -248,6 +248,7 @@ Other markup
* `Nix language <https://nixos.org/nix/>`_
* NSIS scripts
* Notmuch
+* `PEG <https://bford.info/packrat/>`_
* POV-Ray scenes
* `Puppet <https://pup... | diff --git a/tests/test_grammar_notation.py b/tests/test_grammar_notation.py
new file mode 100644
index 0000000000..b20ca97536
--- /dev/null
+++ b/tests/test_grammar_notation.py
@@ -0,0 +1,94 @@
+# -*- coding: utf-8 -*-
+"""
+ Basic Grammar Notation Tests
+ ~~~~~~~~~~~~~~~~~~~~~~~~~~~~
+
+ :copyright: Copyrigh... | diff --git a/doc/languages.rst b/doc/languages.rst
index c976b6af69..f83b60ffd3 100644
--- a/doc/languages.rst
+++ b/doc/languages.rst
@@ -248,6 +248,7 @@ Other markup
* `Nix language <https://nixos.org/nix/>`_
* NSIS scripts
* Notmuch
+* `PEG <https://bford.info/packrat/>`_
* POV-Ray scenes
* `Puppet <https://pup... | [
{
"components": [
{
"doc": "This lexer is for `Parsing Expression Grammars\n<https://bford.info/pub/lang/peg.pdf>`_ (PEG).\n\nVarious implementations of PEG have made different decisions\nregarding the syntax, so let's try to be accommodating:\n\n* `<-`, `←`, `:`, and `=` are all accepted as rule ... | [
"tests/test_grammar_notation.py::test_peg_basic",
"tests/test_grammar_notation.py::test_peg_operators",
"tests/test_grammar_notation.py::test_peg_modified_strings"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add a PEG (Parsing Expression Grammar) lexer
[Parsing Expression Grammars](https://en.wikipedia.org/wiki/Parsing_expression_grammar) (PEGs) have been around for about 15 years and they are fairly popu... | e08bdbba2fa78270dba5ca700d053569f85d0351 | |
joke2k__faker-1075 | 1,075 | joke2k/faker | null | 6894660b8f6cfdd73f99fda1d60274ad9e42eb20 | 2019-12-09T09:31:01Z | diff --git a/faker/providers/misc/__init__.py b/faker/providers/misc/__init__.py
index 0247ecddb8..068977a123 100644
--- a/faker/providers/misc/__init__.py
+++ b/faker/providers/misc/__init__.py
@@ -1,6 +1,7 @@
# coding=utf-8
from __future__ import unicode_literals
+import csv
import hashlib
import io
import str... | diff --git a/tests/providers/test_misc.py b/tests/providers/test_misc.py
index f476901025..ef41c69039 100644
--- a/tests/providers/test_misc.py
+++ b/tests/providers/test_misc.py
@@ -1,13 +1,20 @@
# coding=utf-8
from __future__ import unicode_literals
+import csv
import io
+import itertools
import tarfile
import... | [
{
"components": [
{
"doc": "Generic method that returns delimiter-separated values\n\nThis method's signature is mostly the same as the signature of csv.writer with some\nadditional keyword arguments for controlling data generation. Dialects and formatting\nparameters are passed to the csv.writer ... | [
"tests/providers/test_misc.py::TestMisc::test_csv_helper_method",
"tests/providers/test_misc.py::TestMisc::test_dsv_csvwriter_kwargs",
"tests/providers/test_misc.py::TestMisc::test_dsv_data_columns",
"tests/providers/test_misc.py::TestMisc::test_dsv_no_header",
"tests/providers/test_misc.py::TestMisc::test_... | [
"tests/providers/test_misc.py::TestMisc::test_tar_compression_py3",
"tests/providers/test_misc.py::TestMisc::test_tar_exact_minimum_size",
"tests/providers/test_misc.py::TestMisc::test_tar_invalid_file",
"tests/providers/test_misc.py::TestMisc::test_tar_one_byte_undersized",
"tests/providers/test_misc.py::T... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add provider methods for dsv files
### What does this change
Allows generation of delimiter-separated values with a mostly same API to `csv.writer`
Related to #603
----------
</request>
There... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
joke2k__faker-1072 | 1,072 | joke2k/faker | null | 94b166f1305cf4d9fb3611a92713403b5e64d439 | 2019-12-06T10:12:34Z | diff --git a/faker/providers/person/pl_PL/__init__.py b/faker/providers/person/pl_PL/__init__.py
index 025b50f135..5f0b35c2db 100644
--- a/faker/providers/person/pl_PL/__init__.py
+++ b/faker/providers/person/pl_PL/__init__.py
@@ -741,7 +741,7 @@ def pesel(self, date_of_birth=None, sex=None):
@staticmethod
... | diff --git a/tests/providers/test_person.py b/tests/providers/test_person.py
index ae42f17c8e..58d0720a1b 100644
--- a/tests/providers/test_person.py
+++ b/tests/providers/test_person.py
@@ -21,8 +21,7 @@
from faker.providers.person.cs_CZ import Provider as CsCZProvider
from faker.providers.person.pl_PL import Provid... | [
{
"components": [
{
"doc": "Returns 10 digit of Number of tax identification.\nPolish: Numer identyfikacji podatkowej (NIP).\n\nhttps://pl.wikipedia.org/wiki/NIP\nlist of codes\nhttp://www.algorytm.org/numery-identyfikacyjne/nip.html",
"lines": [
829,
859
],
... | [
"tests/providers/test_person.py::TestPlPL::test_nip"
] | [
"tests/providers/test_person.py::TestAr::test_first_name",
"tests/providers/test_person.py::TestAr::test_last_name",
"tests/providers/test_person.py::TestJaJP::test_person",
"tests/providers/test_person.py::TestNeNP::test_names",
"tests/providers/test_person.py::TestFiFI::test_gender_first_names",
"tests/... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add NIP generator
### What does this changes
Add NIP generator
Number of tax identification.
Polish: Numer identyfikacji podatkowej (NIP)
----------
</request>
There are several new functions... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
joke2k__faker-1069 | 1,069 | joke2k/faker | null | 1ec4a6a9233431e8db3a6d9fc1e54ad76a8f95ca | 2019-12-05T13:11:47Z | diff --git a/faker/providers/misc/__init__.py b/faker/providers/misc/__init__.py
index 0e4030d39f..0247ecddb8 100644
--- a/faker/providers/misc/__init__.py
+++ b/faker/providers/misc/__init__.py
@@ -2,9 +2,14 @@
from __future__ import unicode_literals
import hashlib
+import io
import string
+import tarfile
import... | diff --git a/tests/providers/test_misc.py b/tests/providers/test_misc.py
index 0fd1741431..f476901025 100644
--- a/tests/providers/test_misc.py
+++ b/tests/providers/test_misc.py
@@ -1,5 +1,11 @@
+# coding=utf-8
+
+from __future__ import unicode_literals
+import io
+import tarfile
import unittest
import uuid
+import ... | [
{
"components": [
{
"doc": "Returns a random valid zip archive\n\n:param uncompressed_size: Total size of files before compression, 16 KiB by default\n:param num_files: Number of files archived in resulting zip file, 1 file by default\n:param min_file_size: Minimum size of each file before compres... | [
"tests/providers/test_misc.py::TestMisc::test_tar_compression_py3",
"tests/providers/test_misc.py::TestMisc::test_tar_exact_minimum_size",
"tests/providers/test_misc.py::TestMisc::test_tar_invalid_file",
"tests/providers/test_misc.py::TestMisc::test_tar_one_byte_undersized",
"tests/providers/test_misc.py::T... | [
"tests/providers/test_misc.py::TestMisc::test_uuid4",
"tests/providers/test_misc.py::TestMisc::test_uuid4_int",
"tests/providers/test_misc.py::TestMisc::test_uuid4_seedability",
"tests/providers/test_misc.py::TestMisc::test_uuid4_uuid_object"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Add provider methods for generating popular file types
Aims to fix #603
Note:
Currently, I only wrote provider methods for zip and tar files. I am also in the process of finishing provider metho... | Here is the discussion in the issues of the pull request.
<issues>
Feature Request: Fake Binary Content
Let's say I want to write unit tests for my web service that has a file upload feature.
It would be great if I can fake random binary content of popular binary file types, especially images and documents, like:
-... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | |
sympy__sympy-17974 | 17,974 | sympy/sympy | 1.6 | 029936987c7b35fcc6b347b580d4c154792344e7 | 2019-11-27T16:05:29Z | diff --git a/sympy/polys/multivariate_resultants.py b/sympy/polys/multivariate_resultants.py
index 92eac7292736..05c5e562c5b3 100644
--- a/sympy/polys/multivariate_resultants.py
+++ b/sympy/polys/multivariate_resultants.py
@@ -10,7 +10,7 @@
"""
from sympy import IndexedBase, Matrix, Mul, Poly
-from sympy import rem... | diff --git a/sympy/polys/tests/test_multivariate_resultants.py b/sympy/polys/tests/test_multivariate_resultants.py
index b6b838b10128..2b7a15edd254 100644
--- a/sympy/polys/tests/test_multivariate_resultants.py
+++ b/sympy/polys/tests/test_multivariate_resultants.py
@@ -54,6 +54,20 @@ def test_get_max_degrees():
... | [
{
"components": [
{
"doc": "Test for the validity of the Kapur-Saxena-Yang precondition.\n\nThe precondition requires that the column corresponding to the\nmonomial 1 = x_1 ^ 0 * x_2 ^ 0 * ... * x_n ^ 0 is not a linear\ncombination of the remaining ones. In sympy notation this is\nthe last column.... | [
"test_KSY_precondition",
"test_delete_zero_rows_and_columns",
"test_product_leading_entries",
"test_get_KSY_Dixon_resultant_example_one",
"test_get_KSY_Dixon_resultant_example_two"
] | [
"test_dixon_resultant_init",
"test_get_dixon_polynomial_numerical",
"test_get_max_degrees",
"test_get_dixon_matrix",
"test_get_dixon_matrix_example_two",
"test_macaulay_resultant_init",
"test_get_degree_m",
"test_get_size",
"test_macaulay_example_one"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
polys: Iplemented the Kapur-Saxena-Yang approach to Dixon's resultant
Iplemented the Kapur-Saxena-Yang approach to Dixon's resultant
<!-- Your title above should be a short description of what
was... | 6daf556517f9fa61a7805fe93d8a366688781617 | ||
rytilahti__python-miio-579 | 579 | rytilahti/python-miio | null | 8cb8fb6dc1a0a7c5a8df451932b352d767e9e9e1 | 2019-11-22T13:49:12Z | diff --git a/miio/toiletlid.py b/miio/toiletlid.py
index 1fa724347..9f6b59998 100644
--- a/miio/toiletlid.py
+++ b/miio/toiletlid.py
@@ -1,6 +1,6 @@
import enum
import logging
-from typing import Any, Dict
+from typing import Any, Dict, List
import click
@@ -27,6 +27,14 @@ class AmbientLightColor(enum.Enum):
... | diff --git a/miio/tests/test_toiletlid.py b/miio/tests/test_toiletlid.py
index a226968c3..015c3acb1 100644
--- a/miio/tests/test_toiletlid.py
+++ b/miio/tests/test_toiletlid.py
@@ -30,29 +30,65 @@ def __init__(self, *args, **kwargs):
self.state = {
"is_on": False,
"work_state": 1,
+ ... | [
{
"components": [
{
"doc": "",
"lines": [
30,
35
],
"name": "ToiletlidOperatingMode",
"signature": "class ToiletlidOperatingMode(enum.Enum):",
"type": "class"
},
{
"doc": "Device working mode",
"lines": [
... | [
"miio/tests/test_toiletlid.py::TestToiletlidV1::test_bind_xiaomi_band",
"miio/tests/test_toiletlid.py::TestToiletlidV1::test_unbind_xiaomi_band"
] | [
"miio/tests/test_toiletlid.py::TestToiletlidV1::test_get_all_user_info",
"miio/tests/test_toiletlid.py::TestToiletlidV1::test_nozzle_clean",
"miio/tests/test_toiletlid.py::TestToiletlidV1::test_set_ambient_light",
"miio/tests/test_toiletlid.py::TestToiletlidV1::test_status"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Improve toiletlid various parameters
1.add work_state analysis
2.add xiaomi_band interface
----------
</request>
There are several new functions or classes that need to be implemented, using the de... | 62427d2f796e603520acca3b57b29ec3e6489bca | ||
joke2k__faker-1059 | 1,059 | joke2k/faker | null | 07d4a2078c346e260d474df3cbceebb62bc6e289 | 2019-11-15T22:08:15Z | diff --git a/faker/providers/color/fa_IR/__init__.py b/faker/providers/color/fa_IR/__init__.py
new file mode 100644
index 0000000000..ace185e8fb
--- /dev/null
+++ b/faker/providers/color/fa_IR/__init__.py
@@ -0,0 +1,156 @@
+from collections import OrderedDict
+
+from .. import Provider as ColorProvider
+
+
+class Provi... | diff --git a/tests/providers/test_color.py b/tests/providers/test_color.py
index cfb5968c8a..a126f57231 100644
--- a/tests/providers/test_color.py
+++ b/tests/providers/test_color.py
@@ -7,6 +7,7 @@
from faker import Faker
from faker.providers.color import RandomColor
+from faker.providers.color.fa_IR import Provid... | [
{
"components": [
{
"doc": "",
"lines": [
6,
155
],
"name": "Provider",
"signature": "class Provider(ColorProvider):",
"type": "class"
}
],
"file": "faker/providers/color/fa_IR/__init__.py"
}
] | [
"tests/providers/test_color.py::TestColor::test_color",
"tests/providers/test_color.py::TestColor::test_hex_color",
"tests/providers/test_color.py::TestColor::test_rgb_color",
"tests/providers/test_color.py::TestColor::test_rgb_css_color",
"tests/providers/test_color.py::TestColor::test_safe_hex_color",
"... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
add color for fa_IR
### What does this changes
translated colors to Persian
### What was wrong
lack of Persian colors
### How this fixes it
add fa_IR directory and __init__.py with tra... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
dpkp__kafka-python-1955 | 1,955 | dpkp/kafka-python | null | e3362aca8c12a07ebe88575b073c91475585f21d | 2019-11-15T18:32:10Z | diff --git a/kafka/admin/acl_resource.py b/kafka/admin/acl_resource.py
index 7a012d2fa..fd997a10a 100644
--- a/kafka/admin/acl_resource.py
+++ b/kafka/admin/acl_resource.py
@@ -112,6 +112,24 @@ def __repr__(self):
resource=self.resource_pattern
)
+ def __eq__(self, other):
+ return all... | diff --git a/test/test_acl_comparisons.py b/test/test_acl_comparisons.py
new file mode 100644
index 000000000..291bf0e2f
--- /dev/null
+++ b/test/test_acl_comparisons.py
@@ -0,0 +1,92 @@
+from kafka.admin.acl_resource import ACL
+from kafka.admin.acl_resource import ACLOperation
+from kafka.admin.acl_resource import AC... | [
{
"components": [
{
"doc": "",
"lines": [
115,
121
],
"name": "ACLFilter.__eq__",
"signature": "def __eq__(self, other):",
"type": "function"
},
{
"doc": "",
"lines": [
124,
130
],
... | [
"test/test_acl_comparisons.py::test_same_acls_are_same"
] | [
"test/test_acl_comparisons.py::test_different_acls_are_different",
"test/test_acl_comparisons.py::test_different_acls_are_different_with_glob_topics"
] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implement __eq__ and __hash__ for ACL objects
This allows for direct comparison of ACL objects, and allows us to create `set`s of them
I've found it useful for some work I'm doing to allow us to ma... | 53dc740bce8ef19c32fad2881021d1f6bb055f7a | ||
joke2k__faker-1055 | 1,055 | joke2k/faker | null | 4b7a1898c3d86cafeca18d08ad612da475bdbe26 | 2019-11-15T05:22:35Z | diff --git a/faker/providers/color/__init__.py b/faker/providers/color/__init__.py
index 92bb669d0c..4d039dc653 100644
--- a/faker/providers/color/__init__.py
+++ b/faker/providers/color/__init__.py
@@ -2,6 +2,7 @@
from __future__ import unicode_literals
from collections import OrderedDict
+from .color import Rando... | diff --git a/tests/providers/test_color.py b/tests/providers/test_color.py
index 24f00314b9..073a911d6c 100644
--- a/tests/providers/test_color.py
+++ b/tests/providers/test_color.py
@@ -1,8 +1,11 @@
import unittest
+import re
from re import search
from faker import Faker
+from faker.providers.color import RandomC... | [
{
"components": [
{
"doc": "Creates a color in specified format\n\n:param hue: monochrome, red, orange, yellow, green, blue, purple, pink, a number\n from 0 to 360, or a tuple/list of 2 numbers from 0 to 360\n:param luminosity: bright, dark, light, or random\n:param color_format: hsv, h... | [
"tests/providers/test_color.py::TestColor::test_color",
"tests/providers/test_color.py::TestColor::test_hex_color",
"tests/providers/test_color.py::TestColor::test_rgb_color",
"tests/providers/test_color.py::TestColor::test_rgb_css_color",
"tests/providers/test_color.py::TestColor::test_safe_hex_color",
"... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Improve color provider
### What does this changes
This will add a new color provider method `color()` that allows specifying a hue, a luminosity, and a color format to generate an matching color.
... | Here is the discussion in the issues of the pull request.
<issues>
Improve color provider
see: https://github.com/davidmerfield/randomColor
----------
--------------------
</issues> | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | |
joke2k__faker-1054 | 1,054 | joke2k/faker | null | 4b7a1898c3d86cafeca18d08ad612da475bdbe26 | 2019-11-14T19:32:14Z | diff --git a/faker/providers/company/ru_RU/__init__.py b/faker/providers/company/ru_RU/__init__.py
index afd895a245..36bfc77898 100644
--- a/faker/providers/company/ru_RU/__init__.py
+++ b/faker/providers/company/ru_RU/__init__.py
@@ -1,9 +1,20 @@
# coding=utf-8
from __future__ import unicode_literals
+
+from faker... | diff --git a/tests/providers/test_company.py b/tests/providers/test_company.py
index 5e71f20d92..2b16d60313 100644
--- a/tests/providers/test_company.py
+++ b/tests/providers/test_company.py
@@ -2,19 +2,21 @@
from __future__ import unicode_literals
-import unittest
import re
+import unittest
import six
from... | [
{
"components": [
{
"doc": "",
"lines": [
9,
15
],
"name": "calculate_checksum",
"signature": "def calculate_checksum(value):",
"type": "function"
},
{
"doc": "Returns tax identification number for businesses (ru. иден... | [
"tests/providers/test_company.py::TestRuRu::test_businesses_inn",
"tests/providers/test_company.py::TestRuRu::test_businesses_ogrn",
"tests/providers/test_company.py::TestRuRu::test_calculate_checksum_nine_digits",
"tests/providers/test_company.py::TestRuRu::test_individuals_inn",
"tests/providers/test_comp... | [
"tests/providers/test_company.py::TestFiFI::test_company_business_id",
"tests/providers/test_company.py::TestHyAm::test_bs",
"tests/providers/test_company.py::TestHyAm::test_catch_phrase",
"tests/providers/test_company.py::TestHyAm::test_company",
"tests/providers/test_company.py::TestHyAm::test_company_suf... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
company details to russian provider
For Russian provider added TIN, PPC, PSRN generation
----------
</request>
There are several new functions or classes that need to be implemented, using the defin... | 6edfdbf6ae90b0153309e3bf066aa3b2d16494a7 | ||
dpkp__kafka-python-1951 | 1,951 | dpkp/kafka-python | null | 3d98741be0e9608a352221b476cf3aa2d86777be | 2019-11-13T22:05:20Z | diff --git a/kafka/protocol/api.py b/kafka/protocol/api.py
index efaf63ea2..64276fc17 100644
--- a/kafka/protocol/api.py
+++ b/kafka/protocol/api.py
@@ -3,7 +3,7 @@
import abc
from kafka.protocol.struct import Struct
-from kafka.protocol.types import Int16, Int32, String, Schema
+from kafka.protocol.types import In... | diff --git a/test/test_object_conversion.py b/test/test_object_conversion.py
new file mode 100644
index 000000000..9b1ff2131
--- /dev/null
+++ b/test/test_object_conversion.py
@@ -0,0 +1,236 @@
+from kafka.protocol.admin import Request
+from kafka.protocol.admin import Response
+from kafka.protocol.types import Schema
... | [
{
"components": [
{
"doc": "",
"lines": [
50,
51
],
"name": "Request.to_object",
"signature": "def to_object(self):",
"type": "function"
},
{
"doc": "",
"lines": [
72,
73
],
... | [
"test/test_object_conversion.py::TestObjectConversion::test_get_item[Request]",
"test/test_object_conversion.py::TestObjectConversion::test_get_item[Response]",
"test/test_object_conversion.py::TestObjectConversion::test_with_empty_schema[Request]",
"test/test_object_conversion.py::TestObjectConversion::test_... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Implement methods to convert a Struct object to a pythonic object
There are a handful of places that call out for converting response tuples to more pythonic objects. This is a stab at it - I'm happy ... | 53dc740bce8ef19c32fad2881021d1f6bb055f7a | ||
scikit-learn__scikit-learn-15557 | 15,557 | scikit-learn/scikit-learn | 0.22 | c96e0958da46ebef482a4084cdda3285d5f5ad23 | 2019-11-07T00:03:59Z | diff --git a/doc/whats_new/v0.22.rst b/doc/whats_new/v0.22.rst
index 525ae77311064..4d5ee03853e16 100644
--- a/doc/whats_new/v0.22.rst
+++ b/doc/whats_new/v0.22.rst
@@ -405,6 +405,15 @@ Changelog
estimator's constructor but not stored as attributes on the instance.
:pr:`14464` by `Joel Nothman`_.
+- |Feature| G... | diff --git a/sklearn/gaussian_process/tests/_mini_sequence_kernel.py b/sklearn/gaussian_process/tests/_mini_sequence_kernel.py
new file mode 100644
index 0000000000000..c260a361e1e71
--- /dev/null
+++ b/sklearn/gaussian_process/tests/_mini_sequence_kernel.py
@@ -0,0 +1,51 @@
+from sklearn.gaussian_process.kernels impor... | diff --git a/doc/whats_new/v0.22.rst b/doc/whats_new/v0.22.rst
index 525ae77311064..4d5ee03853e16 100644
--- a/doc/whats_new/v0.22.rst
+++ b/doc/whats_new/v0.22.rst
@@ -405,6 +405,15 @@ Changelog
estimator's constructor but not stored as attributes on the instance.
:pr:`14464` by `Joel Nothman`_.
+- |Feature| G... | [
{
"components": [
{
"doc": "A minimal (but valid) convolutional kernel for sequences of variable\nlengths.",
"lines": [
50,
101
],
"name": "SequenceKernel",
"signature": "class SequenceKernel(GenericKernelMixin, Kernel):",
"type": "class"... | [
"sklearn/gaussian_process/tests/test_gpc.py::test_predict_consistent[kernel0]",
"sklearn/gaussian_process/tests/test_gpc.py::test_predict_consistent[kernel1]",
"sklearn/gaussian_process/tests/test_gpc.py::test_predict_consistent[kernel2]",
"sklearn/gaussian_process/tests/test_gpc.py::test_predict_consistent[k... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[MRG+2] Allowing Gaussian process kernels on structured data - updated
Fixes #13936.
This PR is a from-scratch refresh of #13954 (with all previous modifications incorporated) based on the latest m... | Here is the discussion in the issues of the pull request.
<issues>
GaussianProcessRegressor unnecessarily forcing training samples to be array_like
The current interface precludes the application of the GPR module to cases where the training samples are sequences, trees, graphs etc. In fact, GPR should not care about w... | c96e0958da46ebef482a4084cdda3285d5f5ad23 |
falconry__falcon-1603 | 1,603 | falconry/falcon | null | b68c4ff5b8ddf83f84d0e9d7875fb478995a70eb | 2019-11-02T15:26:06Z | diff --git a/docs/_newsfragments/1603-smart-exception-handler-precedence.feature.rst b/docs/_newsfragments/1603-smart-exception-handler-precedence.feature.rst
new file mode 100644
index 000000000..ba23500ae
--- /dev/null
+++ b/docs/_newsfragments/1603-smart-exception-handler-precedence.feature.rst
@@ -0,0 +1,1 @@
+Exce... | diff --git a/tests/test_error_handlers.py b/tests/test_error_handlers.py
index b46d72185..35aac39fe 100644
--- a/tests/test_error_handlers.py
+++ b/tests/test_error_handlers.py
@@ -91,14 +91,14 @@ def test_subclass_error(self, client):
assert result.status_code == 723
assert result.text == 'error: Cus... | diff --git a/docs/_newsfragments/1603-smart-exception-handler-precedence.feature.rst b/docs/_newsfragments/1603-smart-exception-handler-precedence.feature.rst
new file mode 100644
index 000000000..ba23500ae
--- /dev/null
+++ b/docs/_newsfragments/1603-smart-exception-handler-precedence.feature.rst
@@ -0,0 +1,1 @@
+Exce... | [
{
"components": [
{
"doc": "",
"lines": [
807,
821
],
"name": "App._find_error_handler",
"signature": "def _find_error_handler(self, ex):",
"type": "function"
}
],
"file": "falcon/app.py"
}
] | [
"tests/test_error_handlers.py::TestErrorHandler::test_error_precedence_subclass_order_indifference"
] | [
"tests/test_error_handlers.py::TestErrorHandler::test_caught_error",
"tests/test_error_handlers.py::TestErrorHandler::test_uncaught_python_error[None-application/json-{\"]",
"tests/test_error_handlers.py::TestErrorHandler::test_uncaught_python_error[get_headers1-application/json-{\"]",
"tests/test_error_handl... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
feat(API): on exception, select most specific error handler available
# Summary of Changes
Rather than selecting error handlers in LIFO order of registration, select the error handler corresponding... | 77d5e6394a88ead151c9469494749f95f06b24bf | |
pvlib__pvlib-python-807 | 807 | pvlib/pvlib-python | 0.6 | e326fa53038f616d949e4f981dab6187d2ca9470 | 2019-11-01T14:54:52Z | diff --git a/docs/sphinx/source/api.rst b/docs/sphinx/source/api.rst
index 7241f78f4b..b9c7a82651 100644
--- a/docs/sphinx/source/api.rst
+++ b/docs/sphinx/source/api.rst
@@ -560,9 +560,19 @@ Bifacial
========
Methods for calculating back surface irradiance
------------------------------------------------
.. aut... | diff --git a/pvlib/test/test_scaling.py b/pvlib/test/test_scaling.py
new file mode 100644
index 0000000000..62f37794d2
--- /dev/null
+++ b/pvlib/test/test_scaling.py
@@ -0,0 +1,146 @@
+import numpy as np
+import pandas as pd
+
+import pytest
+from numpy.testing import assert_almost_equal
+
+from pvlib import scaling
+f... | diff --git a/docs/sphinx/source/api.rst b/docs/sphinx/source/api.rst
index 7241f78f4b..b9c7a82651 100644
--- a/docs/sphinx/source/api.rst
+++ b/docs/sphinx/source/api.rst
@@ -560,9 +560,19 @@ Bifacial
========
Methods for calculating back surface irradiance
------------------------------------------------
.. aut... | [
{
"components": [
{
"doc": "Compute spatial aggregation time series smoothing on clear sky index based\non the Wavelet Variability model of Lave et al [1-2]. Implementation is\nbasically a port of the Matlab version of the code [3].\n\nParameters\n----------\nclearsky_index : numeric or pandas.Ser... | [
"pvlib/test/test_scaling.py::test_latlon_to_xy_zero",
"pvlib/test/test_scaling.py::test_latlon_to_xy_single",
"pvlib/test/test_scaling.py::test_latlon_to_xy_array",
"pvlib/test/test_scaling.py::test_latlon_to_xy_list",
"pvlib/test/test_scaling.py::test_compute_wavelet_series",
"pvlib/test/test_scaling.py:... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Create scaling.py and implement WVM model
- [X] Closes #806
- [X] I am familiar with the [contributing guidelines](http://pvlib-python.readthedocs.io/en/latest/contributing.html).
- [x] Fully t... | Here is the discussion in the issues of the pull request.
<issues>
Add Wavelet Variability Model (WVM) for calculating spatial smoothing of irradiance
> > Should I spin this off to a separate issue, since it might be different (and more compartmented) than the broader downscaling discussion?
>
> Yes. Let's start a n... | 7fc595a13bcd42e3269c0806f5505ac907af9730 |
lark-parser__lark-466 | 466 | lark-parser/lark | null | f07359c31683805f4004fe2d6f37dec84b7c094f | 2019-10-31T12:19:33Z | diff --git a/lark/visitors.py b/lark/visitors.py
index 7d40e7479..c6e4f6bed 100644
--- a/lark/visitors.py
+++ b/lark/visitors.py
@@ -186,6 +186,11 @@ def visit(self, tree):
self._call_userfunc(subtree)
return tree
+ def visit_topdown(self,tree):
+ for subtree in tree.iter_subtrees_topd... | diff --git a/tests/test_trees.py b/tests/test_trees.py
index 4216bd6c5..edd2a8b91 100644
--- a/tests/test_trees.py
+++ b/tests/test_trees.py
@@ -7,7 +7,7 @@
import functools
from lark.tree import Tree
-from lark.visitors import Transformer, Interpreter, visit_children_decor, v_args, Discard
+from lark.visitors impo... | [
{
"components": [
{
"doc": "",
"lines": [
189,
192
],
"name": "Visitor.visit_topdown",
"signature": "def visit_topdown(self,tree):",
"type": "function"
},
{
"doc": "",
"lines": [
209,
216
... | [
"tests/test_trees.py::TestTrees::test_visitor"
] | [
"tests/test_trees.py::TestTrees::test_deepcopy",
"tests/test_trees.py::TestTrees::test_discard",
"tests/test_trees.py::TestTrees::test_interp",
"tests/test_trees.py::TestTrees::test_iter_subtrees",
"tests/test_trees.py::TestTrees::test_iter_subtrees_topdown",
"tests/test_trees.py::TestTrees::test_partial"... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
added visit_topdown methods to Visitor classes
I think it would be useful have visit_top_down methods in visitors.
----------
Looks good, but please include a test that shows that it works. You can re... | 5faea9223cc54d1dbd0985cf830d05a10a7729ec | ||
sympy__sympy-17806 | 17,806 | sympy/sympy | 1.5 | d85c30ef80cb6a4b9c53250bf4f549c8759b1141 | 2019-10-27T05:32:54Z | diff --git a/sympy/functions/special/elliptic_integrals.py b/sympy/functions/special/elliptic_integrals.py
index adee570a6cf3..d713a2feb841 100644
--- a/sympy/functions/special/elliptic_integrals.py
+++ b/sympy/functions/special/elliptic_integrals.py
@@ -94,6 +94,12 @@ def _eval_is_zero(self):
if m.is_infinite... | diff --git a/sympy/functions/special/tests/test_elliptic_integrals.py b/sympy/functions/special/tests/test_elliptic_integrals.py
index 98626d47ccd0..d9bb8261100d 100644
--- a/sympy/functions/special/tests/test_elliptic_integrals.py
+++ b/sympy/functions/special/tests/test_elliptic_integrals.py
@@ -1,5 +1,5 @@
from sym... | [
{
"components": [
{
"doc": "",
"lines": [
97,
101
],
"name": "elliptic_k._eval_rewrite_as_Integral",
"signature": "def _eval_rewrite_as_Integral(self, *args):",
"type": "function"
},
{
"doc": "",
"lines": [
... | [
"test_K",
"test_F",
"test_E"
] | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Added `rewrite(Integral)` to elliptic_integrals
<!-- Your title above should be a short description of what
was changed. Do not include the issue number in the title. -->
#### References to other ... | Here is the discussion in the issues of the pull request.
<issues>
[WIP] elliptic_integrals: add "as_integral" property
Elliptic integrals are special functions defined in terms of integrals.
Allow each of them to be rewritten as integrals. Add tests.
----------
--------------------
</issues> | c72f122f67553e1af930bac6c35732d2a0bbb776 | |
fairlearn__fairlearn-102 | 102 | fairlearn/fairlearn | null | d01b0f8cb0808667a022427faf5d26c85bc93c0b | 2019-10-23T15:50:58Z | diff --git a/fairlearn/metrics/__init__.py b/fairlearn/metrics/__init__.py
index 105b75a8d..ed5765d53 100644
--- a/fairlearn/metrics/__init__.py
+++ b/fairlearn/metrics/__init__.py
@@ -14,6 +14,7 @@
from .balanced_root_mean_squared_error import balanced_root_mean_squared_error
from .extra_metrics import specificity_s... | diff --git a/test/unit/metrics/test_mean_predictions.py b/test/unit/metrics/test_mean_predictions.py
new file mode 100644
index 000000000..d8d47c03e
--- /dev/null
+++ b/test/unit/metrics/test_mean_predictions.py
@@ -0,0 +1,61 @@
+# Copyright (c) Microsoft Corporation. All rights reserved.
+# Licensed under the MIT Lice... | [
{
"components": [
{
"doc": "Returns the (weighted) mean prediction. The true\nvalues are ignored, but required as an argument in order\nto maintain a consistent interface",
"lines": [
7,
18
],
"name": "mean_prediction",
"signature": "def mean_pre... | [
"test/unit/metrics/test_mean_predictions.py::test_mean_prediction_unweighted",
"test/unit/metrics/test_mean_predictions.py::test_mean_prediction_weighted",
"test/unit/metrics/test_mean_predictions.py::test_mean_overprediction_unweighted",
"test/unit/metrics/test_mean_predictions.py::test_mean_overprediction_w... | [] | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
Metrics related to regression predictions
Add the following metrics:
- Mean prediction
- Mean overprediction
- Mean underprediction
----------
</request>
There are several new functions or classe... | 403da1fec74bdf2da28dc49487ccd72caa6f6976 | ||
scikit-learn__scikit-learn-15304 | 15,304 | scikit-learn/scikit-learn | 0.22 | bbc23691cb9c5a7f2174d58c934f233b90f51ab2 | 2019-10-20T14:47:50Z | diff --git a/doc/whats_new/v0.22.rst b/doc/whats_new/v0.22.rst
index c19d777eec363..154af3721777d 100644
--- a/doc/whats_new/v0.22.rst
+++ b/doc/whats_new/v0.22.rst
@@ -118,6 +118,11 @@ Changelog
with a target matrix `Y` in which the first column was constant.
:issue:`13609` by :user:`Camila Williamson <camilaagw... | diff --git a/sklearn/cross_decomposition/tests/test_pls.py b/sklearn/cross_decomposition/tests/test_pls.py
index a5993dea87c14..f4973143bb058 100644
--- a/sklearn/cross_decomposition/tests/test_pls.py
+++ b/sklearn/cross_decomposition/tests/test_pls.py
@@ -80,6 +80,12 @@ def check_ortho(M, err_msg):
assert_array_a... | diff --git a/doc/whats_new/v0.22.rst b/doc/whats_new/v0.22.rst
index c19d777eec363..154af3721777d 100644
--- a/doc/whats_new/v0.22.rst
+++ b/doc/whats_new/v0.22.rst
@@ -118,6 +118,11 @@ Changelog
with a target matrix `Y` in which the first column was constant.
:issue:`13609` by :user:`Camila Williamson <camilaagw... | [
{
"components": [
{
"doc": "Transform data back to its original space.\n\nParameters\n----------\nX : array-like of shape (n_samples, n_components)\n New data, where n_samples is the number of samples\n and n_components is the number of pls components.\n\nReturns\n-------\nx_reconstructed : ... | [
"sklearn/cross_decomposition/tests/test_pls.py::test_pls"
] | [
"sklearn/cross_decomposition/tests/test_pls.py::test_convergence_fail",
"sklearn/cross_decomposition/tests/test_pls.py::test_PLSSVD",
"sklearn/cross_decomposition/tests/test_pls.py::test_univariate_pls_regression",
"sklearn/cross_decomposition/tests/test_pls.py::test_predict_transform_copy",
"sklearn/cross_... | This is a feature request which requires a new feature to add in the code repository.
<<NEW FEATURE REQUEST>>
<request>
[MRG] Added inverse_transform for pls base object and test
<!--
Thanks for contributing a pull request! Please ensure you have taken a look at
the contribution guidelines: https://github.com/scikit-... | c96e0958da46ebef482a4084cdda3285d5f5ad23 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.