title stringlengths 1 185 | diff stringlengths 0 32.2M | body stringlengths 0 123k ⌀ | url stringlengths 57 58 | created_at stringlengths 20 20 | closed_at stringlengths 20 20 | merged_at stringlengths 20 20 ⌀ | updated_at stringlengths 20 20 |
|---|---|---|---|---|---|---|---|
CLN: remove cmath, closes #23209 | diff --git a/pandas/_libs/src/headers/cmath b/pandas/_libs/src/headers/cmath
deleted file mode 100644
index 632e1fc2390d0..0000000000000
--- a/pandas/_libs/src/headers/cmath
+++ /dev/null
@@ -1,36 +0,0 @@
-#ifndef _PANDAS_MATH_H_
-#define _PANDAS_MATH_H_
-
-// MSVC 2017 has a bug where `x == x` can be true for NaNs.
-// MSC_VER from https://stackoverflow.com/a/70630/1889400
-// Place upper bound on this check once a fixed MSVC is released.
-#if defined(_MSC_VER) && (_MSC_VER < 1800)
-#include <cmath>
-// In older versions of Visual Studio there wasn't a std::signbit defined
-// This defines it using _copysign
-namespace std {
- __inline int isnan(double x) { return _isnan(x); }
- __inline int signbit(double num) { return _copysign(1.0, num) < 0; }
- __inline int notnan(double x) { return !isnan(x); }
-}
-#elif defined(_MSC_VER) && (_MSC_VER >= 1900)
-#include <cmath>
-namespace std {
- __inline int isnan(double x) { return _isnan(x); }
- __inline int notnan(double x) { return !isnan(x); }
-}
-#elif defined(_MSC_VER)
-#include <cmath>
-namespace std {
- __inline int isnan(double x) { return _isnan(x); }
- __inline int notnan(double x) { return x == x; }
-}
-#else
-#include <cmath>
-
-namespace std {
- __inline int notnan(double x) { return x == x; }
-}
-
-#endif
-#endif
diff --git a/pandas/_libs/window.pyx b/pandas/_libs/window.pyx
index 62066c5f66ea3..b75dfbd6a9f36 100644
--- a/pandas/_libs/window.pyx
+++ b/pandas/_libs/window.pyx
@@ -9,15 +9,11 @@ from libc.stdlib cimport malloc, free
import numpy as np
cimport numpy as cnp
from numpy cimport ndarray, int64_t, float64_t, float32_t
+from numpy.math cimport signbit, isnan, sqrtl as sqrt
+# sqrtl is square root specialized to "long double"
cnp.import_array()
-cdef extern from "src/headers/cmath" namespace "std":
- bint isnan(float64_t) nogil
- bint notnan(float64_t) nogil
- int signbit(float64_t) nogil
- float64_t sqrt(float64_t x) nogil
-
cimport pandas._libs.util as util
from pandas._libs.util cimport numeric
@@ -384,7 +380,7 @@ def roll_count(ndarray[float64_t] values, int64_t win, int64_t minp,
count_x = 0.0
for j in range(s, e):
val = values[j]
- if notnan(val):
+ if not isnan(val):
count_x += 1.0
else:
@@ -392,13 +388,13 @@ def roll_count(ndarray[float64_t] values, int64_t win, int64_t minp,
# calculate deletes
for j in range(start[i - 1], s):
val = values[j]
- if notnan(val):
+ if not isnan(val):
count_x -= 1.0
# calculate adds
for j in range(end[i - 1], e):
val = values[j]
- if notnan(val):
+ if not isnan(val):
count_x += 1.0
if count_x >= minp:
@@ -429,7 +425,7 @@ cdef inline void add_sum(float64_t val, int64_t *nobs, float64_t *sum_x) nogil:
""" add a value from the sum calc """
# Not NaN
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] + 1
sum_x[0] = sum_x[0] + val
@@ -437,7 +433,7 @@ cdef inline void add_sum(float64_t val, int64_t *nobs, float64_t *sum_x) nogil:
cdef inline void remove_sum(float64_t val, int64_t *nobs, float64_t *sum_x) nogil:
""" remove a value from the sum calc """
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] - 1
sum_x[0] = sum_x[0] - val
@@ -545,7 +541,7 @@ cdef inline void add_mean(float64_t val, Py_ssize_t *nobs, float64_t *sum_x,
""" add a value from the mean calc """
# Not NaN
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] + 1
sum_x[0] = sum_x[0] + val
if signbit(val):
@@ -556,7 +552,7 @@ cdef inline void remove_mean(float64_t val, Py_ssize_t *nobs, float64_t *sum_x,
Py_ssize_t *neg_ct) nogil:
""" remove a value from the mean calc """
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] - 1
sum_x[0] = sum_x[0] - val
if signbit(val):
@@ -683,7 +679,7 @@ cdef inline void remove_var(float64_t val, float64_t *nobs, float64_t *mean_x,
cdef:
float64_t delta
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] - 1
if nobs[0]:
# a part of Welford's method for the online variance-calculation
@@ -772,7 +768,7 @@ def roll_var(ndarray[float64_t] values, int64_t win, int64_t minp,
val = values[i]
prev = values[i - win]
- if notnan(val):
+ if not isnan(val):
if prev == prev:
# Adding one observation and removing another one
@@ -837,7 +833,7 @@ cdef inline void add_skew(float64_t val, int64_t *nobs,
""" add a value from the skew calc """
# Not NaN
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] + 1
# seriously don't ask me why this is faster
@@ -852,7 +848,7 @@ cdef inline void remove_skew(float64_t val, int64_t *nobs,
""" remove a value from the skew calc """
# Not NaN
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] - 1
# seriously don't ask me why this is faster
@@ -979,7 +975,7 @@ cdef inline void add_kurt(float64_t val, int64_t *nobs,
""" add a value from the kurotic calc """
# Not NaN
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] + 1
# seriously don't ask me why this is faster
@@ -995,7 +991,7 @@ cdef inline void remove_kurt(float64_t val, int64_t *nobs,
""" remove a value from the kurotic calc """
# Not NaN
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] - 1
# seriously don't ask me why this is faster
@@ -1116,7 +1112,7 @@ def roll_median_c(ndarray[float64_t] values, int64_t win, int64_t minp,
# setup
for j in range(s, e):
val = values[j]
- if notnan(val):
+ if not isnan(val):
nobs += 1
err = skiplist_insert(sl, val) != 1
if err:
@@ -1127,7 +1123,7 @@ def roll_median_c(ndarray[float64_t] values, int64_t win, int64_t minp,
# calculate adds
for j in range(end[i - 1], e):
val = values[j]
- if notnan(val):
+ if not isnan(val):
nobs += 1
err = skiplist_insert(sl, val) != 1
if err:
@@ -1136,7 +1132,7 @@ def roll_median_c(ndarray[float64_t] values, int64_t win, int64_t minp,
# calculate deletes
for j in range(start[i - 1], s):
val = values[j]
- if notnan(val):
+ if not isnan(val):
skiplist_remove(sl, val)
nobs -= 1
@@ -1510,7 +1506,7 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
# setup
for j in range(s, e):
val = values[j]
- if notnan(val):
+ if not isnan(val):
nobs += 1
skiplist_insert(skiplist, val)
@@ -1519,14 +1515,14 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
# calculate adds
for j in range(end[i - 1], e):
val = values[j]
- if notnan(val):
+ if not isnan(val):
nobs += 1
skiplist_insert(skiplist, val)
# calculate deletes
for j in range(start[i - 1], s):
val = values[j]
- if notnan(val):
+ if not isnan(val):
skiplist_remove(skiplist, val)
nobs -= 1
@@ -1885,7 +1881,7 @@ cdef inline void remove_weighted_var(float64_t val,
cdef:
float64_t temp, q, r
- if notnan(val):
+ if not isnan(val):
nobs[0] = nobs[0] - 1
if nobs[0]:
@@ -1955,7 +1951,7 @@ def roll_weighted_var(float64_t[:] values, float64_t[:] weights,
w = weights[i % win_n]
pre_w = weights[(i - win_n) % win_n]
- if notnan(val):
+ if not isnan(val):
if pre_val == pre_val:
remove_weighted_var(pre_val, pre_w, &t,
&sum_w, &mean, &nobs)
| cc @chris-b1 | https://api.github.com/repos/pandas-dev/pandas/pulls/29491 | 2019-11-08T19:19:29Z | 2019-11-08T23:51:20Z | null | 2019-11-21T19:59:35Z |
CLN: annotation in reshape.merge | diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index bc23d50c634d5..2674b7ee95088 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -6,6 +6,7 @@
import datetime
from functools import partial
import string
+from typing import TYPE_CHECKING, Optional, Tuple, Union
import warnings
import numpy as np
@@ -39,6 +40,7 @@
from pandas.core.dtypes.missing import isna, na_value_for_dtype
from pandas import Categorical, Index, MultiIndex
+from pandas._typing import FrameOrSeries
import pandas.core.algorithms as algos
from pandas.core.arrays.categorical import _recode_for_categories
import pandas.core.common as com
@@ -46,22 +48,25 @@
from pandas.core.internals import _transform_index, concatenate_block_managers
from pandas.core.sorting import is_int64_overflow_possible
+if TYPE_CHECKING:
+ from pandas import DataFrame, Series # noqa:F401
+
@Substitution("\nleft : DataFrame")
@Appender(_merge_doc, indents=0)
def merge(
left,
right,
- how="inner",
+ how: str = "inner",
on=None,
left_on=None,
right_on=None,
- left_index=False,
- right_index=False,
- sort=False,
+ left_index: bool = False,
+ right_index: bool = False,
+ sort: bool = False,
suffixes=("_x", "_y"),
- copy=True,
- indicator=False,
+ copy: bool = True,
+ indicator: bool = False,
validate=None,
):
op = _MergeOperation(
@@ -86,7 +91,9 @@ def merge(
merge.__doc__ = _merge_doc % "\nleft : DataFrame"
-def _groupby_and_merge(by, on, left, right, _merge_pieces, check_duplicates=True):
+def _groupby_and_merge(
+ by, on, left, right, _merge_pieces, check_duplicates: bool = True
+):
"""
groupby & merge; we are always performing a left-by type operation
@@ -172,7 +179,7 @@ def merge_ordered(
right_by=None,
fill_method=None,
suffixes=("_x", "_y"),
- how="outer",
+ how: str = "outer",
):
"""
Perform merge with optional filling/interpolation.
@@ -298,14 +305,14 @@ def merge_asof(
on=None,
left_on=None,
right_on=None,
- left_index=False,
- right_index=False,
+ left_index: bool = False,
+ right_index: bool = False,
by=None,
left_by=None,
right_by=None,
suffixes=("_x", "_y"),
tolerance=None,
- allow_exact_matches=True,
+ allow_exact_matches: bool = True,
direction="backward",
):
"""
@@ -533,33 +540,33 @@ def merge_asof(
# TODO: only copy DataFrames when modification necessary
class _MergeOperation:
"""
- Perform a database (SQL) merge operation between two DataFrame objects
- using either columns as keys or their row indexes
+ Perform a database (SQL) merge operation between two DataFrame or Series
+ objects using either columns as keys or their row indexes
"""
_merge_type = "merge"
def __init__(
self,
- left,
- right,
- how="inner",
+ left: Union["Series", "DataFrame"],
+ right: Union["Series", "DataFrame"],
+ how: str = "inner",
on=None,
left_on=None,
right_on=None,
axis=1,
- left_index=False,
- right_index=False,
- sort=True,
+ left_index: bool = False,
+ right_index: bool = False,
+ sort: bool = True,
suffixes=("_x", "_y"),
- copy=True,
- indicator=False,
+ copy: bool = True,
+ indicator: bool = False,
validate=None,
):
- left = validate_operand(left)
- right = validate_operand(right)
- self.left = self.orig_left = left
- self.right = self.orig_right = right
+ _left = _validate_operand(left)
+ _right = _validate_operand(right)
+ self.left = self.orig_left = _validate_operand(_left) # type: "DataFrame"
+ self.right = self.orig_right = _validate_operand(_right) # type: "DataFrame"
self.how = how
self.axis = axis
@@ -577,7 +584,7 @@ def __init__(
self.indicator = indicator
if isinstance(self.indicator, str):
- self.indicator_name = self.indicator
+ self.indicator_name = self.indicator # type: Optional[str]
elif isinstance(self.indicator, bool):
self.indicator_name = "_merge" if self.indicator else None
else:
@@ -597,11 +604,11 @@ def __init__(
)
# warn user when merging between different levels
- if left.columns.nlevels != right.columns.nlevels:
+ if _left.columns.nlevels != _right.columns.nlevels:
msg = (
"merging between different levels can give an unintended "
"result ({left} levels on the left, {right} on the right)"
- ).format(left=left.columns.nlevels, right=right.columns.nlevels)
+ ).format(left=_left.columns.nlevels, right=_right.columns.nlevels)
warnings.warn(msg, UserWarning)
self._validate_specification()
@@ -658,7 +665,9 @@ def get_result(self):
return result
- def _indicator_pre_merge(self, left, right):
+ def _indicator_pre_merge(
+ self, left: "DataFrame", right: "DataFrame"
+ ) -> Tuple["DataFrame", "DataFrame"]:
columns = left.columns.union(right.columns)
@@ -878,7 +887,12 @@ def _get_join_info(self):
return join_index, left_indexer, right_indexer
def _create_join_index(
- self, index, other_index, indexer, other_indexer, how="left"
+ self,
+ index: Index,
+ other_index: Index,
+ indexer,
+ other_indexer,
+ how: str = "left",
):
"""
Create a join index by rearranging one index to match another
@@ -1263,7 +1277,9 @@ def _validate(self, validate: str):
raise ValueError("Not a valid argument for validate")
-def _get_join_indexers(left_keys, right_keys, sort=False, how="inner", **kwargs):
+def _get_join_indexers(
+ left_keys, right_keys, sort: bool = False, how: str = "inner", **kwargs
+):
"""
Parameters
@@ -1410,13 +1426,13 @@ def __init__(
on=None,
left_on=None,
right_on=None,
- left_index=False,
- right_index=False,
+ left_index: bool = False,
+ right_index: bool = False,
axis=1,
suffixes=("_x", "_y"),
- copy=True,
+ copy: bool = True,
fill_method=None,
- how="outer",
+ how: str = "outer",
):
self.fill_method = fill_method
@@ -1508,18 +1524,18 @@ def __init__(
on=None,
left_on=None,
right_on=None,
- left_index=False,
- right_index=False,
+ left_index: bool = False,
+ right_index: bool = False,
by=None,
left_by=None,
right_by=None,
axis=1,
suffixes=("_x", "_y"),
- copy=True,
+ copy: bool = True,
fill_method=None,
- how="asof",
+ how: str = "asof",
tolerance=None,
- allow_exact_matches=True,
+ allow_exact_matches: bool = True,
direction="backward",
):
@@ -1757,13 +1773,15 @@ def flip(xs):
return func(left_values, right_values, self.allow_exact_matches, tolerance)
-def _get_multiindex_indexer(join_keys, index, sort):
+def _get_multiindex_indexer(join_keys, index: MultiIndex, sort: bool):
# bind `sort` argument
fkeys = partial(_factorize_keys, sort=sort)
# left & right join labels and num. of levels at each location
- rcodes, lcodes, shape = map(list, zip(*map(fkeys, index.levels, join_keys)))
+ mapped = (fkeys(index.levels[n], join_keys[n]) for n in range(len(index.levels)))
+ zipped = zip(*mapped)
+ rcodes, lcodes, shape = [list(x) for x in zipped]
if sort:
rcodes = list(map(np.take, rcodes, index.codes))
else:
@@ -1791,7 +1809,7 @@ def _get_multiindex_indexer(join_keys, index, sort):
return libjoin.left_outer_join(lkey, rkey, count, sort=sort)
-def _get_single_indexer(join_key, index, sort=False):
+def _get_single_indexer(join_key, index, sort: bool = False):
left_key, right_key, count = _factorize_keys(join_key, index, sort=sort)
left_indexer, right_indexer = libjoin.left_outer_join(
@@ -1801,7 +1819,7 @@ def _get_single_indexer(join_key, index, sort=False):
return left_indexer, right_indexer
-def _left_join_on_index(left_ax, right_ax, join_keys, sort=False):
+def _left_join_on_index(left_ax: Index, right_ax: Index, join_keys, sort: bool = False):
if len(join_keys) > 1:
if not (
(isinstance(right_ax, MultiIndex) and len(join_keys) == right_ax.nlevels)
@@ -1915,7 +1933,7 @@ def _factorize_keys(lk, rk, sort=True):
return llab, rlab, count
-def _sort_labels(uniques, left, right):
+def _sort_labels(uniques: np.ndarray, left, right):
if not isinstance(uniques, np.ndarray):
# tuplesafe
uniques = Index(uniques).values
@@ -1930,7 +1948,7 @@ def _sort_labels(uniques, left, right):
return new_left, new_right
-def _get_join_keys(llab, rlab, shape, sort):
+def _get_join_keys(llab, rlab, shape, sort: bool):
# how many levels can be done without overflow
pred = lambda i: not is_int64_overflow_possible(shape[:i])
@@ -1970,7 +1988,7 @@ def _any(x) -> bool:
return x is not None and com.any_not_none(*x)
-def validate_operand(obj):
+def _validate_operand(obj: FrameOrSeries) -> "DataFrame":
if isinstance(obj, ABCDataFrame):
return obj
elif isinstance(obj, ABCSeries):
@@ -1985,7 +2003,7 @@ def validate_operand(obj):
)
-def _items_overlap_with_suffix(left, lsuffix, right, rsuffix):
+def _items_overlap_with_suffix(left: Index, lsuffix, right: Index, rsuffix):
"""
If two indices overlap, add suffixes to overlapping entries.
| This will need multiple passes, trying to keep a moderately-sized diff. | https://api.github.com/repos/pandas-dev/pandas/pulls/29490 | 2019-11-08T17:06:10Z | 2019-11-12T23:43:05Z | 2019-11-12T23:43:05Z | 2019-11-13T14:29:10Z |
ENH: Support arrow/parquet roundtrip for nullable integer / string extension dtypes | diff --git a/doc/source/development/extending.rst b/doc/source/development/extending.rst
index e341dcb8318bc..89d43e8a43825 100644
--- a/doc/source/development/extending.rst
+++ b/doc/source/development/extending.rst
@@ -251,6 +251,48 @@ To use a test, subclass it:
See https://github.com/pandas-dev/pandas/blob/master/pandas/tests/extension/base/__init__.py
for a list of all the tests available.
+.. _extending.extension.arrow:
+
+Compatibility with Apache Arrow
+^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
+
+An ``ExtensionArray`` can support conversion to / from ``pyarrow`` arrays
+(and thus support for example serialization to the Parquet file format)
+by implementing two methods: ``ExtensionArray.__arrow_array__`` and
+``ExtensionDtype.__from_arrow__``.
+
+The ``ExtensionArray.__arrow_array__`` ensures that ``pyarrow`` knowns how
+to convert the specific extension array into a ``pyarrow.Array`` (also when
+included as a column in a pandas DataFrame):
+
+.. code-block:: python
+
+ class MyExtensionArray(ExtensionArray):
+ ...
+
+ def __arrow_array__(self, type=None):
+ # convert the underlying array values to a pyarrow Array
+ import pyarrow
+ return pyarrow.array(..., type=type)
+
+The ``ExtensionDtype.__from_arrow__`` method then controls the conversion
+back from pyarrow to a pandas ExtensionArray. This method receives a pyarrow
+``Array`` or ``ChunkedArray`` as only argument and is expected to return the
+appropriate pandas ``ExtensionArray`` for this dtype and the passed values:
+
+.. code-block:: none
+
+ class ExtensionDtype:
+ ...
+
+ def __from_arrow__(self, array: pyarrow.Array/ChunkedArray) -> ExtensionArray:
+ ...
+
+See more in the `Arrow documentation <https://arrow.apache.org/docs/python/extending_types.html>`__.
+
+Those methods have been implemented for the nullable integer and string extension
+dtypes included in pandas, and ensure roundtrip to pyarrow and the Parquet file format.
+
.. _extension dtype dtypes: https://github.com/pandas-dev/pandas/blob/master/pandas/core/dtypes/dtypes.py
.. _extension dtype source: https://github.com/pandas-dev/pandas/blob/master/pandas/core/dtypes/base.py
.. _extension array source: https://github.com/pandas-dev/pandas/blob/master/pandas/core/arrays/base.py
diff --git a/doc/source/user_guide/io.rst b/doc/source/user_guide/io.rst
index 6e45d6748c2a5..fa47a5944f7bf 100644
--- a/doc/source/user_guide/io.rst
+++ b/doc/source/user_guide/io.rst
@@ -4716,6 +4716,9 @@ Several caveats.
* The ``pyarrow`` engine preserves the ``ordered`` flag of categorical dtypes with string types. ``fastparquet`` does not preserve the ``ordered`` flag.
* Non supported types include ``Period`` and actual Python object types. These will raise a helpful error message
on an attempt at serialization.
+* The ``pyarrow`` engine preserves extension data types such as the nullable integer and string data
+ type (requiring pyarrow >= 1.0.0, and requiring the extension type to implement the needed protocols,
+ see the :ref:`extension types documentation <extending.extension.arrow>`).
You can specify an ``engine`` to direct the serialization. This can be one of ``pyarrow``, or ``fastparquet``, or ``auto``.
If the engine is NOT specified, then the ``pd.options.io.parquet.engine`` option is checked; if this is also ``auto``,
diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 0027343a13b60..de8273f890436 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -114,6 +114,9 @@ Other enhancements
- Added ``encoding`` argument to :meth:`DataFrame.to_string` for non-ascii text (:issue:`28766`)
- Added ``encoding`` argument to :func:`DataFrame.to_html` for non-ascii text (:issue:`28663`)
- :meth:`Styler.background_gradient` now accepts ``vmin`` and ``vmax`` arguments (:issue:`12145`)
+- Roundtripping DataFrames with nullable integer or string data types to parquet
+ (:meth:`~DataFrame.to_parquet` / :func:`read_parquet`) using the `'pyarrow'` engine
+ now preserve those data types with pyarrow >= 1.0.0 (:issue:`20612`).
Build Changes
^^^^^^^^^^^^^
diff --git a/pandas/core/arrays/integer.py b/pandas/core/arrays/integer.py
index af7755fb1373d..63296b4a26354 100644
--- a/pandas/core/arrays/integer.py
+++ b/pandas/core/arrays/integer.py
@@ -85,6 +85,35 @@ def construct_array_type(cls):
"""
return IntegerArray
+ def __from_arrow__(self, array):
+ """Construct IntegerArray from passed pyarrow Array/ChunkedArray"""
+ import pyarrow
+
+ if isinstance(array, pyarrow.Array):
+ chunks = [array]
+ else:
+ # pyarrow.ChunkedArray
+ chunks = array.chunks
+
+ results = []
+ for arr in chunks:
+ buflist = arr.buffers()
+ data = np.frombuffer(buflist[1], dtype=self.type)[
+ arr.offset : arr.offset + len(arr)
+ ]
+ bitmask = buflist[0]
+ if bitmask is not None:
+ mask = pyarrow.BooleanArray.from_buffers(
+ pyarrow.bool_(), len(arr), [None, bitmask]
+ )
+ mask = np.asarray(mask)
+ else:
+ mask = np.ones(len(arr), dtype=bool)
+ int_arr = IntegerArray(data.copy(), ~mask, copy=False)
+ results.append(int_arr)
+
+ return IntegerArray._concat_same_type(results)
+
def integer_array(values, dtype=None, copy=False):
"""
diff --git a/pandas/core/arrays/string_.py b/pandas/core/arrays/string_.py
index 7c487b227de20..8599b5e39f34a 100644
--- a/pandas/core/arrays/string_.py
+++ b/pandas/core/arrays/string_.py
@@ -85,6 +85,24 @@ def construct_array_type(cls) -> "Type[StringArray]":
def __repr__(self) -> str:
return "StringDtype"
+ def __from_arrow__(self, array):
+ """Construct StringArray from passed pyarrow Array/ChunkedArray"""
+ import pyarrow
+
+ if isinstance(array, pyarrow.Array):
+ chunks = [array]
+ else:
+ # pyarrow.ChunkedArray
+ chunks = array.chunks
+
+ results = []
+ for arr in chunks:
+ # using _from_sequence to ensure None is convered to np.nan
+ str_arr = StringArray._from_sequence(np.array(arr))
+ results.append(str_arr)
+
+ return StringArray._concat_same_type(results)
+
class StringArray(PandasArray):
"""
diff --git a/pandas/tests/arrays/string_/test_string.py b/pandas/tests/arrays/string_/test_string.py
index efe2b4e0b2deb..1ce62d8f8b3d9 100644
--- a/pandas/tests/arrays/string_/test_string.py
+++ b/pandas/tests/arrays/string_/test_string.py
@@ -171,3 +171,19 @@ def test_arrow_array():
arr = pa.array(data)
expected = pa.array(list(data), type=pa.string(), from_pandas=True)
assert arr.equals(expected)
+
+
+@td.skip_if_no("pyarrow", min_version="0.15.1.dev")
+def test_arrow_roundtrip():
+ # roundtrip possible from arrow 1.0.0
+ import pyarrow as pa
+
+ data = pd.array(["a", "b", None], dtype="string")
+ df = pd.DataFrame({"a": data})
+ table = pa.table(df)
+ assert table.field("a").type == "string"
+ result = table.to_pandas()
+ assert isinstance(result["a"].dtype, pd.StringDtype)
+ tm.assert_frame_equal(result, df)
+ # ensure the missing value is represented by NaN and not None
+ assert np.isnan(result.loc[2, "a"])
diff --git a/pandas/tests/arrays/test_integer.py b/pandas/tests/arrays/test_integer.py
index 025366e5b210b..443a0c7e71616 100644
--- a/pandas/tests/arrays/test_integer.py
+++ b/pandas/tests/arrays/test_integer.py
@@ -829,6 +829,18 @@ def test_arrow_array(data):
assert arr.equals(expected)
+@td.skip_if_no("pyarrow", min_version="0.15.1.dev")
+def test_arrow_roundtrip(data):
+ # roundtrip possible from arrow 1.0.0
+ import pyarrow as pa
+
+ df = pd.DataFrame({"a": data})
+ table = pa.table(df)
+ assert table.field("a").type == str(data.dtype.numpy_dtype)
+ result = table.to_pandas()
+ tm.assert_frame_equal(result, df)
+
+
@pytest.mark.parametrize(
"pandasmethname, kwargs",
[
diff --git a/pandas/tests/io/test_parquet.py b/pandas/tests/io/test_parquet.py
index 5dd671c659263..bcbbee3b86769 100644
--- a/pandas/tests/io/test_parquet.py
+++ b/pandas/tests/io/test_parquet.py
@@ -514,13 +514,19 @@ def test_additional_extension_arrays(self, pa):
"b": pd.Series(["a", None, "c"], dtype="string"),
}
)
- # currently de-serialized as plain int / object
- expected = df.assign(a=df.a.astype("int64"), b=df.b.astype("object"))
+ if LooseVersion(pyarrow.__version__) >= LooseVersion("0.15.1.dev"):
+ expected = df
+ else:
+ # de-serialized as plain int / object
+ expected = df.assign(a=df.a.astype("int64"), b=df.b.astype("object"))
check_round_trip(df, pa, expected=expected)
df = pd.DataFrame({"a": pd.Series([1, 2, 3, None], dtype="Int64")})
- # if missing values in integer, currently de-serialized as float
- expected = df.assign(a=df.a.astype("float64"))
+ if LooseVersion(pyarrow.__version__) >= LooseVersion("0.15.1.dev"):
+ expected = df
+ else:
+ # if missing values in integer, currently de-serialized as float
+ expected = df.assign(a=df.a.astype("float64"))
check_round_trip(df, pa, expected=expected)
| xref https://github.com/pandas-dev/pandas/issues/20612
This implements the `__from_arrow__` method for integer/string extension types, so roundtripping to arrow/parquet fully works. | https://api.github.com/repos/pandas-dev/pandas/pulls/29483 | 2019-11-08T13:42:47Z | 2019-11-19T15:17:45Z | 2019-11-19T15:17:45Z | 2019-11-19T15:19:03Z |
TYPING: change to FrameOrSeries Alias in pandas._typing | diff --git a/pandas/_typing.py b/pandas/_typing.py
index 445eff9e19e47..df2d327af92a3 100644
--- a/pandas/_typing.py
+++ b/pandas/_typing.py
@@ -21,8 +21,8 @@
from pandas.core.arrays.base import ExtensionArray # noqa: F401
from pandas.core.dtypes.dtypes import ExtensionDtype # noqa: F401
from pandas.core.indexes.base import Index # noqa: F401
+ from pandas.core.frame import DataFrame # noqa: F401
from pandas.core.series import Series # noqa: F401
- from pandas.core.generic import NDFrame # noqa: F401
AnyArrayLike = TypeVar("AnyArrayLike", "ExtensionArray", "Index", "Series", np.ndarray)
@@ -31,7 +31,7 @@
Dtype = Union[str, np.dtype, "ExtensionDtype"]
FilePathOrBuffer = Union[str, Path, IO[AnyStr]]
-FrameOrSeries = TypeVar("FrameOrSeries", bound="NDFrame")
+FrameOrSeries = TypeVar("FrameOrSeries", "DataFrame", "Series")
Scalar = Union[str, int, float, bool]
Axis = Union[str, int]
Ordered = Optional[bool]
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index e73be29d5b104..8b20371de603b 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -14,7 +14,17 @@ class providing the base-class of operations.
import inspect
import re
import types
-from typing import FrozenSet, Hashable, Iterable, List, Optional, Tuple, Type, Union
+from typing import (
+ FrozenSet,
+ Generic,
+ Hashable,
+ Iterable,
+ List,
+ Optional,
+ Tuple,
+ Type,
+ Union,
+)
import numpy as np
@@ -41,6 +51,7 @@ class providing the base-class of operations.
)
from pandas.core.dtypes.missing import isna, notna
+from pandas._typing import FrameOrSeries
from pandas.core import nanops
import pandas.core.algorithms as algorithms
from pandas.core.arrays import Categorical, try_cast_to_ea
@@ -336,13 +347,13 @@ def _group_selection_context(groupby):
groupby._reset_group_selection()
-class _GroupBy(PandasObject, SelectionMixin):
+class _GroupBy(PandasObject, SelectionMixin, Generic[FrameOrSeries]):
_group_selection = None
_apply_whitelist = frozenset() # type: FrozenSet[str]
def __init__(
self,
- obj: NDFrame,
+ obj: FrameOrSeries,
keys=None,
axis: int = 0,
level=None,
@@ -391,7 +402,7 @@ def __init__(
mutated=self.mutated,
)
- self.obj = obj
+ self.obj = obj # type: FrameOrSeries
self.axis = obj._get_axis_number(axis)
self.grouper = grouper
self.exclusions = set(exclusions) if exclusions else set()
@@ -1662,7 +1673,9 @@ def backfill(self, limit=None):
@Substitution(name="groupby")
@Substitution(see_also=_common_see_also)
- def nth(self, n: Union[int, List[int]], dropna: Optional[str] = None) -> DataFrame:
+ def nth(
+ self, n: Union[int, List[int]], dropna: Optional[str] = None
+ ) -> FrameOrSeries:
"""
Take the nth row from each group if n is an int, or a subset of rows
if n is a list of ints.
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index 370abe75e1327..c950ccd6ed1e4 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -21,7 +21,7 @@
)
from pandas.core.dtypes.generic import ABCSeries
-from pandas._typing import FrameOrSeries
+from pandas._typing import FrameOrSeries, Union
import pandas.core.algorithms as algorithms
from pandas.core.arrays import Categorical, ExtensionArray
import pandas.core.common as com
@@ -249,7 +249,7 @@ def __init__(
self,
index: Index,
grouper=None,
- obj: Optional[FrameOrSeries] = None,
+ obj: Optional[Union[DataFrame, Series]] = None,
name=None,
level=None,
sort: bool = True,
@@ -570,8 +570,7 @@ def get_grouper(
all_in_columns_index = all(
g in obj.columns or g in obj.index.names for g in keys
)
- else:
- assert isinstance(obj, Series)
+ elif isinstance(obj, Series):
all_in_columns_index = all(g in obj.index.names for g in keys)
if not all_in_columns_index:
| @WillAyd
I think we should discuss reverting #28173 and only use `TypeVar("FrameOrSeries", bound="NDFrame")` in `core.generic`. perhaps call it `_NDFrameT` to avoid confusion.
This PR is to show the changes required to keep mypy green if we wanted to revert.
It also reverts the change in #29458, see https://github.com/pandas-dev/pandas/pull/29458#discussion_r343733263
`_GroupBy` is defined as a generic class, but not sure about `Grouping`
I've created it as a draft PR, since I think we should wait for more issues to surface going forward to help decide. The number of changes needed at this stage is not significant enough to warrant a hasty decision. | https://api.github.com/repos/pandas-dev/pandas/pulls/29480 | 2019-11-08T10:24:34Z | 2019-12-02T14:20:22Z | null | 2019-12-02T14:20:23Z |
CLN: annotations in core.apply | diff --git a/pandas/core/apply.py b/pandas/core/apply.py
index d9f6bdae288ed..9c5806a3fe945 100644
--- a/pandas/core/apply.py
+++ b/pandas/core/apply.py
@@ -1,4 +1,6 @@
+import abc
import inspect
+from typing import TYPE_CHECKING, Iterator, Type
import numpy as np
@@ -13,14 +15,17 @@
)
from pandas.core.dtypes.generic import ABCSeries
+if TYPE_CHECKING:
+ from pandas import DataFrame, Series, Index
+
def frame_apply(
- obj,
+ obj: "DataFrame",
func,
axis=0,
- raw=False,
+ raw: bool = False,
result_type=None,
- ignore_failures=False,
+ ignore_failures: bool = False,
args=None,
kwds=None,
):
@@ -28,7 +33,7 @@ def frame_apply(
axis = obj._get_axis_number(axis)
if axis == 0:
- klass = FrameRowApply
+ klass = FrameRowApply # type: Type[FrameApply]
elif axis == 1:
klass = FrameColumnApply
@@ -43,8 +48,38 @@ def frame_apply(
)
-class FrameApply:
- def __init__(self, obj, func, raw, result_type, ignore_failures, args, kwds):
+class FrameApply(metaclass=abc.ABCMeta):
+
+ # ---------------------------------------------------------------
+ # Abstract Methods
+ axis: int
+
+ @property
+ @abc.abstractmethod
+ def result_index(self) -> "Index":
+ pass
+
+ @property
+ @abc.abstractmethod
+ def result_columns(self) -> "Index":
+ pass
+
+ @abc.abstractmethod
+ def series_generator(self) -> Iterator["Series"]:
+ pass
+
+ # ---------------------------------------------------------------
+
+ def __init__(
+ self,
+ obj: "DataFrame",
+ func,
+ raw: bool,
+ result_type,
+ ignore_failures: bool,
+ args,
+ kwds,
+ ):
self.obj = obj
self.raw = raw
self.ignore_failures = ignore_failures
@@ -76,11 +111,11 @@ def f(x):
self.res_columns = None
@property
- def columns(self):
+ def columns(self) -> "Index":
return self.obj.columns
@property
- def index(self):
+ def index(self) -> "Index":
return self.obj.index
@cache_readonly
@@ -88,11 +123,11 @@ def values(self):
return self.obj.values
@cache_readonly
- def dtypes(self):
+ def dtypes(self) -> "Series":
return self.obj.dtypes
@property
- def agg_axis(self):
+ def agg_axis(self) -> "Index":
return self.obj._get_agg_axis(self.axis)
def get_result(self):
@@ -127,7 +162,7 @@ def get_result(self):
# broadcasting
if self.result_type == "broadcast":
- return self.apply_broadcast()
+ return self.apply_broadcast(self.obj)
# one axis empty
elif not all(self.obj.shape):
@@ -191,7 +226,7 @@ def apply_raw(self):
else:
return self.obj._constructor_sliced(result, index=self.agg_axis)
- def apply_broadcast(self, target):
+ def apply_broadcast(self, target: "DataFrame") -> "DataFrame":
result_values = np.empty_like(target.values)
# axis which we want to compare compliance
@@ -317,19 +352,19 @@ def wrap_results(self):
class FrameRowApply(FrameApply):
axis = 0
- def apply_broadcast(self):
- return super().apply_broadcast(self.obj)
+ def apply_broadcast(self, target: "DataFrame") -> "DataFrame":
+ return super().apply_broadcast(target)
@property
def series_generator(self):
return (self.obj._ixs(i, axis=1) for i in range(len(self.columns)))
@property
- def result_index(self):
+ def result_index(self) -> "Index":
return self.columns
@property
- def result_columns(self):
+ def result_columns(self) -> "Index":
return self.index
def wrap_results_for_axis(self):
@@ -351,8 +386,8 @@ def wrap_results_for_axis(self):
class FrameColumnApply(FrameApply):
axis = 1
- def apply_broadcast(self):
- result = super().apply_broadcast(self.obj.T)
+ def apply_broadcast(self, target: "DataFrame") -> "DataFrame":
+ result = super().apply_broadcast(target.T)
return result.T
@property
@@ -364,11 +399,11 @@ def series_generator(self):
)
@property
- def result_index(self):
+ def result_index(self) -> "Index":
return self.index
@property
- def result_columns(self):
+ def result_columns(self) -> "Index":
return self.columns
def wrap_results_for_axis(self):
@@ -392,7 +427,7 @@ def wrap_results_for_axis(self):
return result
- def infer_to_same_shape(self):
+ def infer_to_same_shape(self) -> "DataFrame":
""" infer the results to the same shape as the input object """
results = self.results
| make `apply_broadcast` have consistent signature.
After this I plan to do a pass to make these classes much less stateful | https://api.github.com/repos/pandas-dev/pandas/pulls/29477 | 2019-11-08T04:21:41Z | 2019-11-12T23:44:19Z | 2019-11-12T23:44:19Z | 2019-11-13T00:02:06Z |
TYPES: __len__, is_all_dates, inferred_type | diff --git a/pandas/_libs/hashtable_class_helper.pxi.in b/pandas/_libs/hashtable_class_helper.pxi.in
index c39d6d60d4ea5..b207fcb66948d 100644
--- a/pandas/_libs/hashtable_class_helper.pxi.in
+++ b/pandas/_libs/hashtable_class_helper.pxi.in
@@ -100,7 +100,7 @@ cdef class {{name}}Vector:
PyMem_Free(self.data)
self.data = NULL
- def __len__(self):
+ def __len__(self) -> int:
return self.data.n
cpdef to_array(self):
@@ -168,7 +168,7 @@ cdef class StringVector:
PyMem_Free(self.data)
self.data = NULL
- def __len__(self):
+ def __len__(self) -> int:
return self.data.n
def to_array(self):
@@ -212,7 +212,7 @@ cdef class ObjectVector:
self.ao = np.empty(_INIT_VEC_CAP, dtype=object)
self.data = <PyObject**>self.ao.data
- def __len__(self):
+ def __len__(self) -> int:
return self.n
cdef inline append(self, object obj):
@@ -270,7 +270,7 @@ cdef class {{name}}HashTable(HashTable):
size_hint = min(size_hint, _SIZE_HINT_LIMIT)
kh_resize_{{dtype}}(self.table, size_hint)
- def __len__(self):
+ def __len__(self) -> int:
return self.table.size
def __dealloc__(self):
@@ -897,7 +897,7 @@ cdef class PyObjectHashTable(HashTable):
kh_destroy_pymap(self.table)
self.table = NULL
- def __len__(self):
+ def __len__(self) -> int:
return self.table.size
def __contains__(self, object key):
diff --git a/pandas/_libs/internals.pyx b/pandas/_libs/internals.pyx
index db9f16d46e48c..00d647711b53a 100644
--- a/pandas/_libs/internals.pyx
+++ b/pandas/_libs/internals.pyx
@@ -66,7 +66,7 @@ cdef class BlockPlacement:
def __repr__(self) -> str:
return str(self)
- def __len__(self):
+ def __len__(self) -> int:
cdef:
slice s = self._ensure_has_slice()
if s is not None:
diff --git a/pandas/core/arrays/categorical.py b/pandas/core/arrays/categorical.py
index 39470c7420086..73d1db9bda8ed 100644
--- a/pandas/core/arrays/categorical.py
+++ b/pandas/core/arrays/categorical.py
@@ -1940,7 +1940,7 @@ def take_nd(self, indexer, allow_fill=None, fill_value=None):
take = take_nd
- def __len__(self):
+ def __len__(self) -> int:
"""
The length of this Categorical.
"""
diff --git a/pandas/core/arrays/datetimelike.py b/pandas/core/arrays/datetimelike.py
index 4b83dd0cfff09..f93db4695d38f 100644
--- a/pandas/core/arrays/datetimelike.py
+++ b/pandas/core/arrays/datetimelike.py
@@ -396,7 +396,7 @@ def size(self) -> int:
"""The number of elements in this array."""
return np.prod(self.shape)
- def __len__(self):
+ def __len__(self) -> int:
return len(self._data)
def __getitem__(self, key):
diff --git a/pandas/core/arrays/integer.py b/pandas/core/arrays/integer.py
index 08b53e54b91ef..41d8bffd8c131 100644
--- a/pandas/core/arrays/integer.py
+++ b/pandas/core/arrays/integer.py
@@ -469,7 +469,7 @@ def __setitem__(self, key, value):
self._data[key] = value
self._mask[key] = mask
- def __len__(self):
+ def __len__(self) -> int:
return len(self._data)
@property
diff --git a/pandas/core/arrays/interval.py b/pandas/core/arrays/interval.py
index cc41797e7872b..cb482665b3534 100644
--- a/pandas/core/arrays/interval.py
+++ b/pandas/core/arrays/interval.py
@@ -489,7 +489,7 @@ def _validate(self):
def __iter__(self):
return iter(np.asarray(self))
- def __len__(self):
+ def __len__(self) -> int:
return len(self.left)
def __getitem__(self, value):
diff --git a/pandas/core/computation/expr.py b/pandas/core/computation/expr.py
index 39653c3d695b2..929c9e69d56ac 100644
--- a/pandas/core/computation/expr.py
+++ b/pandas/core/computation/expr.py
@@ -837,7 +837,7 @@ def __call__(self):
def __repr__(self) -> str:
return printing.pprint_thing(self.terms)
- def __len__(self):
+ def __len__(self) -> int:
return len(self.expr)
def parse(self):
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 7e3c2200dbabc..ebee8b10896be 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -1023,7 +1023,7 @@ def itertuples(self, index=True, name="Pandas"):
# fallback to regular tuples
return zip(*arrays)
- def __len__(self):
+ def __len__(self) -> int:
"""
Returns length of info axis, but here we use the index.
"""
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index c3788baec030a..bcdffe695e96a 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -1951,7 +1951,7 @@ def items(self):
def iteritems(self):
return self.items()
- def __len__(self):
+ def __len__(self) -> int:
"""Returns length of info axis"""
return len(self._info_axis)
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index e73be29d5b104..fd45d60b02277 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -399,7 +399,7 @@ def __init__(
# we accept no other args
validate_kwargs("group", kwargs, {})
- def __len__(self):
+ def __len__(self) -> int:
return len(self.groups)
def __repr__(self) -> str:
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index c9697c530628a..ee124ba3851b1 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -649,10 +649,12 @@ def _engine(self):
# Array-Like Methods
# ndarray compat
- def __len__(self):
+ def __len__(self) -> int:
"""
Return the length of the Index.
"""
+ # Assertion needed for mypy, see GH#29475
+ assert self._data is not None
return len(self._data)
def __array__(self, dtype=None):
@@ -1807,7 +1809,7 @@ def inferred_type(self):
return lib.infer_dtype(self, skipna=False)
@cache_readonly
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
return is_datetime_array(ensure_object(self.values))
# --------------------------------------------------------------------
diff --git a/pandas/core/indexes/category.py b/pandas/core/indexes/category.py
index e5a8edb56e413..0187b47ab50a1 100644
--- a/pandas/core/indexes/category.py
+++ b/pandas/core/indexes/category.py
@@ -366,7 +366,7 @@ def _format_attrs(self):
# --------------------------------------------------------------------
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
return "categorical"
@property
diff --git a/pandas/core/indexes/datetimes.py b/pandas/core/indexes/datetimes.py
index 2d0ecf1b936da..4a3ee57084a8a 100644
--- a/pandas/core/indexes/datetimes.py
+++ b/pandas/core/indexes/datetimes.py
@@ -1235,13 +1235,13 @@ def is_type_compatible(self, typ):
return typ == self.inferred_type or typ == "datetime"
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
# b/c datetime is represented as microseconds since the epoch, make
# sure we can't have ambiguous indexing
return "datetime64"
@property
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
return True
def insert(self, loc, item):
diff --git a/pandas/core/indexes/interval.py b/pandas/core/indexes/interval.py
index 0c077702b4cb4..d4305f4c40288 100644
--- a/pandas/core/indexes/interval.py
+++ b/pandas/core/indexes/interval.py
@@ -468,7 +468,7 @@ def itemsize(self):
warnings.simplefilter("ignore")
return self.left.itemsize + self.right.itemsize
- def __len__(self):
+ def __len__(self) -> int:
return len(self.left)
@cache_readonly
@@ -524,7 +524,7 @@ def dtype(self):
return self._data.dtype
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
"""Return a string of the type inferred from the values"""
return "interval"
@@ -1357,7 +1357,7 @@ def func(self, other, sort=sort):
return func
@property
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
"""
This is False even when left/right contain datetime-like objects,
as the check is done on the Interval itself
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index 19769d5b029a1..a6a6de6c13c04 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -1217,7 +1217,7 @@ def format(
# --------------------------------------------------------------------
- def __len__(self):
+ def __len__(self) -> int:
return len(self.codes[0])
def _get_names(self):
@@ -1322,7 +1322,7 @@ def _constructor(self):
return MultiIndex.from_tuples
@cache_readonly
- def inferred_type(self):
+ def inferred_type(self) -> str:
return "mixed"
def _get_level_number(self, level):
@@ -1791,7 +1791,7 @@ def to_flat_index(self):
return Index(self.values, tupleize_cols=False)
@property
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
return False
def is_lexsorted(self):
diff --git a/pandas/core/indexes/numeric.py b/pandas/core/indexes/numeric.py
index 46bb8eafee3b9..01924dc4b79f1 100644
--- a/pandas/core/indexes/numeric.py
+++ b/pandas/core/indexes/numeric.py
@@ -133,7 +133,7 @@ def _concat_same_dtype(self, indexes, name):
return result.rename(name)
@property
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
"""
Checks that all the labels are datetime objects
"""
@@ -227,7 +227,7 @@ class Int64Index(IntegerIndex):
_default_dtype = np.int64
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
"""Always 'integer' for ``Int64Index``"""
return "integer"
@@ -282,7 +282,7 @@ class UInt64Index(IntegerIndex):
_default_dtype = np.uint64
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
"""Always 'integer' for ``UInt64Index``"""
return "integer"
@@ -355,7 +355,7 @@ class Float64Index(NumericIndex):
_default_dtype = np.float64
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
"""Always 'floating' for ``Float64Index``"""
return "floating"
diff --git a/pandas/core/indexes/period.py b/pandas/core/indexes/period.py
index ca7be9ba512da..3bcb9ba345713 100644
--- a/pandas/core/indexes/period.py
+++ b/pandas/core/indexes/period.py
@@ -574,7 +574,7 @@ def searchsorted(self, value, side="left", sorter=None):
return self._ndarray_values.searchsorted(value, side=side, sorter=sorter)
@property
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
return True
@property
@@ -591,7 +591,7 @@ def is_full(self):
return ((values[1:] - values[:-1]) < 2).all()
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
# b/c data is represented as ints make sure we can't have ambiguous
# indexing
return "period"
diff --git a/pandas/core/indexes/range.py b/pandas/core/indexes/range.py
index 5fa3431fc97c0..67791417f1bb5 100644
--- a/pandas/core/indexes/range.py
+++ b/pandas/core/indexes/range.py
@@ -698,7 +698,7 @@ def _concat_same_dtype(self, indexes, name):
# In this case return an empty range index.
return RangeIndex(0, 0).rename(name)
- def __len__(self):
+ def __len__(self) -> int:
"""
return the length of the RangeIndex
"""
diff --git a/pandas/core/indexes/timedeltas.py b/pandas/core/indexes/timedeltas.py
index 2324b8cf74c46..8114b4a772f28 100644
--- a/pandas/core/indexes/timedeltas.py
+++ b/pandas/core/indexes/timedeltas.py
@@ -602,11 +602,11 @@ def is_type_compatible(self, typ):
return typ == self.inferred_type or typ == "timedelta"
@property
- def inferred_type(self):
+ def inferred_type(self) -> str:
return "timedelta64"
@property
- def is_all_dates(self):
+ def is_all_dates(self) -> bool:
return True
def insert(self, loc, item):
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 9402a3ef9a763..5508cf3ca522e 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -288,7 +288,7 @@ def __repr__(self) -> str:
return result
- def __len__(self):
+ def __len__(self) -> int:
return len(self.values)
def __getstate__(self):
diff --git a/pandas/core/internals/managers.py b/pandas/core/internals/managers.py
index f86a294475ee2..617844bee2115 100644
--- a/pandas/core/internals/managers.py
+++ b/pandas/core/internals/managers.py
@@ -321,7 +321,7 @@ def _post_setstate(self):
self._known_consolidated = False
self._rebuild_blknos_and_blklocs()
- def __len__(self):
+ def __len__(self) -> int:
return len(self.items)
def __repr__(self) -> str:
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 15f405e244d0f..7327c2d543836 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -712,7 +712,7 @@ def put(self, *args, **kwargs):
)
self._values.put(*args, **kwargs)
- def __len__(self):
+ def __len__(self) -> int:
"""
Return the length of the Series.
"""
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index e98802888e582..ee08e2abb2289 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -540,7 +540,7 @@ def __contains__(self, key):
return True
return False
- def __len__(self):
+ def __len__(self) -> int:
return len(self.groups())
def __repr__(self) -> str:
diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index 5c930e01c735d..b537200dd7664 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -1879,7 +1879,7 @@ def test_concat_iterables(self):
tm.assert_frame_equal(concat(deque((df1, df2)), ignore_index=True), expected)
class CustomIterator1:
- def __len__(self):
+ def __len__(self) -> int:
return 2
def __getitem__(self, index):
diff --git a/scripts/validate_docstrings.py b/scripts/validate_docstrings.py
index 1d0f4b583bd0c..7c6f2fea97933 100755
--- a/scripts/validate_docstrings.py
+++ b/scripts/validate_docstrings.py
@@ -250,7 +250,7 @@ def __init__(self, name):
self.clean_doc = pydoc.getdoc(obj)
self.doc = NumpyDocString(self.clean_doc)
- def __len__(self):
+ def __len__(self) -> int:
return len(self.raw_doc)
@staticmethod
| Operation Get This Over With, part 2 | https://api.github.com/repos/pandas-dev/pandas/pulls/29475 | 2019-11-08T03:05:20Z | 2019-11-11T23:08:30Z | 2019-11-11T23:08:30Z | 2019-11-11T23:11:29Z |
CLN: simplify _shallow_copy | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 8c8037091559d..f236fea93278c 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -634,20 +634,19 @@ def _is_builtin_func(self, arg):
class ShallowMixin:
_attributes = [] # type: List[str]
- def _shallow_copy(self, obj=None, obj_type=None, **kwargs):
+ def _shallow_copy(self, obj=None, **kwargs):
"""
return a new object with the replacement attributes
"""
if obj is None:
obj = self._selected_obj.copy()
- if obj_type is None:
- obj_type = self._constructor
- if isinstance(obj, obj_type):
+
+ if isinstance(obj, self._constructor):
obj = obj.obj
for attr in self._attributes:
if attr not in kwargs:
kwargs[attr] = getattr(self, attr)
- return obj_type(obj, **kwargs)
+ return self._constructor(obj, **kwargs)
class IndexOpsMixin:
diff --git a/pandas/core/groupby/base.py b/pandas/core/groupby/base.py
index fc3bb69afd0cb..fed387cbeade4 100644
--- a/pandas/core/groupby/base.py
+++ b/pandas/core/groupby/base.py
@@ -11,22 +11,6 @@ class GroupByMixin:
Provide the groupby facilities to the mixed object.
"""
- @staticmethod
- def _dispatch(name, *args, **kwargs):
- """
- Dispatch to apply.
- """
-
- def outer(self, *args, **kwargs):
- def f(x):
- x = self._shallow_copy(x, groupby=self._groupby)
- return getattr(x, name)(*args, **kwargs)
-
- return self._groupby.apply(f)
-
- outer.__name__ = name
- return outer
-
def _gotitem(self, key, ndim, subset=None):
"""
Sub-classes to define. Return a sliced object.
diff --git a/pandas/core/window/common.py b/pandas/core/window/common.py
index 2ad5a1eb6faed..3fd567f97edae 100644
--- a/pandas/core/window/common.py
+++ b/pandas/core/window/common.py
@@ -26,7 +26,23 @@
"""
-class _GroupByMixin(GroupByMixin):
+def _dispatch(name: str, *args, **kwargs):
+ """
+ Dispatch to apply.
+ """
+
+ def outer(self, *args, **kwargs):
+ def f(x):
+ x = self._shallow_copy(x, groupby=self._groupby)
+ return getattr(x, name)(*args, **kwargs)
+
+ return self._groupby.apply(f)
+
+ outer.__name__ = name
+ return outer
+
+
+class WindowGroupByMixin(GroupByMixin):
"""
Provide the groupby facilities.
"""
@@ -41,9 +57,9 @@ def __init__(self, obj, *args, **kwargs):
self._groupby.grouper.mutated = True
super().__init__(obj, *args, **kwargs)
- count = GroupByMixin._dispatch("count")
- corr = GroupByMixin._dispatch("corr", other=None, pairwise=None)
- cov = GroupByMixin._dispatch("cov", other=None, pairwise=None)
+ count = _dispatch("count")
+ corr = _dispatch("corr", other=None, pairwise=None)
+ cov = _dispatch("cov", other=None, pairwise=None)
def _apply(
self, func, name=None, window=None, center=None, check_minp=None, **kwargs
@@ -53,6 +69,7 @@ def _apply(
performing the original function call on the grouped object.
"""
+ # TODO: can we de-duplicate with _dispatch?
def f(x, name=name, *args):
x = self._shallow_copy(x)
diff --git a/pandas/core/window/expanding.py b/pandas/core/window/expanding.py
index 47bd8f2ec593b..55389d2fc7d9f 100644
--- a/pandas/core/window/expanding.py
+++ b/pandas/core/window/expanding.py
@@ -3,7 +3,7 @@
from pandas.compat.numpy import function as nv
from pandas.util._decorators import Appender, Substitution
-from pandas.core.window.common import _doc_template, _GroupByMixin, _shared_docs
+from pandas.core.window.common import WindowGroupByMixin, _doc_template, _shared_docs
from pandas.core.window.rolling import _Rolling_and_Expanding
@@ -250,7 +250,7 @@ def corr(self, other=None, pairwise=None, **kwargs):
return super().corr(other=other, pairwise=pairwise, **kwargs)
-class ExpandingGroupby(_GroupByMixin, Expanding):
+class ExpandingGroupby(WindowGroupByMixin, Expanding):
"""
Provide a expanding groupby implementation.
"""
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index bf5ea9c457e8a..75f9a1c628d72 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -39,9 +39,9 @@
import pandas.core.common as com
from pandas.core.index import Index, ensure_index
from pandas.core.window.common import (
+ WindowGroupByMixin,
_doc_template,
_flex_binary_moment,
- _GroupByMixin,
_offset,
_require_min_periods,
_shared_docs,
@@ -1917,7 +1917,7 @@ def corr(self, other=None, pairwise=None, **kwargs):
Rolling.__doc__ = Window.__doc__
-class RollingGroupby(_GroupByMixin, Rolling):
+class RollingGroupby(WindowGroupByMixin, Rolling):
"""
Provide a rolling groupby implementation.
"""
| The class hierarchy of in/around groupby is really tough, this simplifies it a little bit | https://api.github.com/repos/pandas-dev/pandas/pulls/29474 | 2019-11-07T23:50:22Z | 2019-11-08T14:39:32Z | 2019-11-08T14:39:32Z | 2019-11-08T15:18:41Z |
AreaPlot: add support for step drawstyle | diff --git a/pandas/plotting/_matplotlib/core.py b/pandas/plotting/_matplotlib/core.py
index 541dca715e814..9fa531bef859b 100644
--- a/pandas/plotting/_matplotlib/core.py
+++ b/pandas/plotting/_matplotlib/core.py
@@ -1257,6 +1257,11 @@ def _plot(
# need to remove label, because subplots uses mpl legend as it is
line_kwds = kwds.copy()
line_kwds.pop("label")
+
+ # need to replace "step" argument with equivalent "drawstyle"
+ if "step" in line_kwds:
+ line_kwds["drawstyle"] = "steps-" + line_kwds["step"]
+ line_kwds.pop("step")
lines = MPLPlot._plot(ax, x, y_values, style=style, **line_kwds)
# get data from the line to get coordinates for fill_between
| - [ ] closes #29451
- [ ] tests added / passed
- [ ] passes `black pandas`
- [ ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29473 | 2019-11-07T23:48:26Z | 2020-01-01T20:54:16Z | null | 2020-03-04T11:06:41Z |
API: Rename BaseGrouper.recons_codes to BaseGrouper.reconstructed_codes | diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 3b8c3148f5177..3ef726e569970 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -655,7 +655,7 @@ def value_counts(
rep = partial(np.repeat, repeats=np.add.reduceat(inc, idx))
# multi-index components
- codes = self.grouper.recons_codes
+ codes = self.grouper.reconstructed_codes
codes = [rep(level_codes) for level_codes in codes] + [llab(lab, inc)]
levels = [ping.group_index for ping in self.grouper.groupings] + [lev]
names = self.grouper.names + [self._selection_name]
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 9599ce0bf39a9..e438db6c620ec 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,7 +7,7 @@
"""
import collections
-from typing import List, Optional, Sequence, Type
+from typing import List, Optional, Sequence, Tuple, Type
import numpy as np
@@ -216,11 +216,11 @@ def indices(self):
return get_indexer_dict(codes_list, keys)
@property
- def codes(self):
+ def codes(self) -> List[np.ndarray]:
return [ping.codes for ping in self.groupings]
@property
- def levels(self):
+ def levels(self) -> List[Index]:
return [ping.group_index for ping in self.groupings]
@property
@@ -264,7 +264,7 @@ def group_info(self):
return comp_ids, obs_group_ids, ngroups
@cache_readonly
- def codes_info(self):
+ def codes_info(self) -> np.ndarray:
# return the codes of items in original grouped axis
codes, _, _ = self.group_info
if self.indexer is not None:
@@ -272,8 +272,8 @@ def codes_info(self):
codes = codes[sorter]
return codes
- def _get_compressed_codes(self):
- all_codes = [ping.codes for ping in self.groupings]
+ def _get_compressed_codes(self) -> Tuple[np.ndarray, np.ndarray]:
+ all_codes = self.codes
if len(all_codes) > 1:
group_index = get_group_index(all_codes, self.shape, sort=True, xnull=True)
return compress_group_index(group_index, sort=self.sort)
@@ -286,9 +286,9 @@ def ngroups(self) -> int:
return len(self.result_index)
@property
- def recons_codes(self):
+ def reconstructed_codes(self) -> List[np.ndarray]:
+ codes = self.codes
comp_ids, obs_ids, _ = self.group_info
- codes = (ping.codes for ping in self.groupings)
return decons_obs_group_ids(comp_ids, obs_ids, self.shape, codes, xnull=True)
@cache_readonly
@@ -296,7 +296,7 @@ def result_index(self):
if not self.compressed and len(self.groupings) == 1:
return self.groupings[0].result_index.rename(self.names[0])
- codes = self.recons_codes
+ codes = self.reconstructed_codes
levels = [ping.result_index for ping in self.groupings]
result = MultiIndex(
levels=levels, codes=codes, verify_integrity=False, names=self.names
@@ -308,7 +308,7 @@ def get_group_levels(self):
return [self.groupings[0].result_index]
name_list = []
- for ping, codes in zip(self.groupings, self.recons_codes):
+ for ping, codes in zip(self.groupings, self.reconstructed_codes):
codes = ensure_platform_int(codes)
levels = ping.result_index.take(codes)
@@ -768,7 +768,7 @@ def group_info(self):
)
@cache_readonly
- def recons_codes(self):
+ def reconstructed_codes(self) -> List[np.ndarray]:
# get unique result indices, and prepend 0 as groupby starts from the first
return [np.r_[0, np.flatnonzero(self.bins[1:] != self.bins[:-1]) + 1]]
| Rename ``BaseGrouper.recons_codes`` to ``BaseGrouper.reconstructed_codes``. This is an internal attribute, so ok to rename.
Also adds some typing. | https://api.github.com/repos/pandas-dev/pandas/pulls/29471 | 2019-11-07T19:26:23Z | 2019-11-08T04:41:07Z | 2019-11-08T04:41:06Z | 2019-11-08T04:41:11Z |
CLN: remove dead code, closes #28898 | diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 40efc4c65476a..c9111812e42b0 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -2731,38 +2731,6 @@ def transpose(self, *args, **kwargs):
T = property(transpose)
- # ----------------------------------------------------------------------
- # Picklability
-
- # legacy pickle formats
- def _unpickle_frame_compat(self, state): # pragma: no cover
- if len(state) == 2: # pragma: no cover
- series, idx = state
- columns = sorted(series)
- else:
- series, cols, idx = state
- columns = com._unpickle_array(cols)
-
- index = com._unpickle_array(idx)
- self._data = self._init_dict(series, index, columns, None)
-
- def _unpickle_matrix_compat(self, state): # pragma: no cover
- # old unpickling
- (vals, idx, cols), object_state = state
-
- index = com._unpickle_array(idx)
- dm = DataFrame(vals, index=index, columns=com._unpickle_array(cols), copy=False)
-
- if object_state is not None:
- ovals, _, ocols = object_state
- objects = DataFrame(
- ovals, index=index, columns=com._unpickle_array(ocols), copy=False
- )
-
- dm = dm.join(objects)
-
- self._data = dm._data
-
# ----------------------------------------------------------------------
# Indexing Methods
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index bafc37d478fdb..d488a2e57aa52 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -2089,18 +2089,8 @@ def __setstate__(self, state):
else:
self._unpickle_series_compat(state)
- elif isinstance(state[0], dict):
- if len(state) == 5:
- self._unpickle_sparse_frame_compat(state)
- else:
- self._unpickle_frame_compat(state)
- elif len(state) == 4:
- self._unpickle_panel_compat(state)
elif len(state) == 2:
self._unpickle_series_compat(state)
- else: # pragma: no cover
- # old pickling format, for compatibility
- self._unpickle_matrix_compat(state)
self._item_cache = {}
diff --git a/pandas/io/pickle.py b/pandas/io/pickle.py
index 8f9bae0f7a28f..5e066c4f9ecbd 100644
--- a/pandas/io/pickle.py
+++ b/pandas/io/pickle.py
@@ -1,10 +1,7 @@
""" pickle compat """
-from io import BytesIO
import pickle
import warnings
-from numpy.lib.format import read_array
-
from pandas.compat import PY36, pickle_compat as pc
from pandas.io.common import _get_handle, _stringify_path
@@ -164,12 +161,3 @@ def read_pickle(path, compression="infer"):
f.close()
for _f in fh:
_f.close()
-
-
-# compat with sparse pickle / unpickle
-
-
-def _unpickle_array(bytes):
- arr = read_array(BytesIO(bytes))
-
- return arr
| - [x] closes #28898
| https://api.github.com/repos/pandas-dev/pandas/pulls/29470 | 2019-11-07T18:59:27Z | 2019-11-07T21:21:44Z | 2019-11-07T21:21:43Z | 2019-11-07T21:57:36Z |
PERF: MultiIndex.get_loc | diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index fe91a588c7dde..19769d5b029a1 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -1981,11 +1981,11 @@ def remove_unused_levels(self):
return result
@property
- def nlevels(self):
+ def nlevels(self) -> int:
"""
Integer number of levels in this MultiIndex.
"""
- return len(self.levels)
+ return len(self._levels)
@property
def levshape(self):
| - [x] closes #29311
```
In [4]: mi_med = pd.MultiIndex.from_product(
...: [np.arange(1000), np.arange(10), list("A")], names=["one", "two", "three"]
...: )
In [5]: %timeit mi_med.get_loc((999, 9, "A"))
master --> 42.6 µs ± 411 ns per loop (mean ± std. dev. of 7 runs, 10000 loops each)
PR --> 11.4 µs ± 120 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
```
| https://api.github.com/repos/pandas-dev/pandas/pulls/29469 | 2019-11-07T18:40:15Z | 2019-11-07T21:20:57Z | 2019-11-07T21:20:57Z | 2019-11-07T21:56:53Z |
CLN: annotate __str__ and __repr__ methods | diff --git a/pandas/_libs/internals.pyx b/pandas/_libs/internals.pyx
index ff143fea892ae..db9f16d46e48c 100644
--- a/pandas/_libs/internals.pyx
+++ b/pandas/_libs/internals.pyx
@@ -53,7 +53,7 @@ cdef class BlockPlacement:
self._as_array = arr
self._has_array = True
- def __str__(self):
+ def __str__(self) -> str:
cdef:
slice s = self._ensure_has_slice()
if s is not None:
@@ -63,7 +63,7 @@ cdef class BlockPlacement:
return '%s(%r)' % (self.__class__.__name__, v)
- def __repr__(self):
+ def __repr__(self) -> str:
return str(self)
def __len__(self):
diff --git a/pandas/_libs/interval.pyx b/pandas/_libs/interval.pyx
index b13ce7c294f37..2bd38524852ec 100644
--- a/pandas/_libs/interval.pyx
+++ b/pandas/_libs/interval.pyx
@@ -377,7 +377,7 @@ cdef class Interval(IntervalMixin):
return left, right
- def __repr__(self):
+ def __repr__(self) -> str:
left, right = self._repr_base()
name = type(self).__name__
@@ -385,7 +385,7 @@ cdef class Interval(IntervalMixin):
name=name, left=left, right=right, closed=self.closed)
return repr_str
- def __str__(self):
+ def __str__(self) -> str:
left, right = self._repr_base()
start_symbol = '[' if self.closed_left else '('
diff --git a/pandas/_libs/intervaltree.pxi.in b/pandas/_libs/intervaltree.pxi.in
index 6e3be19f2b73e..8cb51be36645e 100644
--- a/pandas/_libs/intervaltree.pxi.in
+++ b/pandas/_libs/intervaltree.pxi.in
@@ -195,7 +195,7 @@ cdef class IntervalTree(IntervalMixin):
return (result.to_array().astype('intp'),
missing.to_array().astype('intp'))
- def __repr__(self):
+ def __repr__(self) -> str:
return ('<IntervalTree[{dtype},{closed}]: '
'{n_elements} elements>'.format(
dtype=self.dtype, closed=self.closed,
@@ -394,7 +394,7 @@ cdef class {{dtype_title}}Closed{{closed_title}}IntervalNode:
else:
result.extend(self.center_left_indices)
- def __repr__(self):
+ def __repr__(self) -> str:
if self.is_leaf_node:
return ('<{{dtype_title}}Closed{{closed_title}}IntervalNode: '
'%s elements (terminal)>' % self.n_elements)
diff --git a/pandas/_libs/sparse.pyx b/pandas/_libs/sparse.pyx
index 6abaaca010b00..1944f9592829c 100644
--- a/pandas/_libs/sparse.pyx
+++ b/pandas/_libs/sparse.pyx
@@ -51,7 +51,7 @@ cdef class IntIndex(SparseIndex):
args = (self.length, self.indices)
return IntIndex, args
- def __repr__(self):
+ def __repr__(self) -> str:
output = 'IntIndex\n'
output += 'Indices: %s\n' % repr(self.indices)
return output
@@ -341,7 +341,7 @@ cdef class BlockIndex(SparseIndex):
args = (self.length, self.blocs, self.blengths)
return BlockIndex, args
- def __repr__(self):
+ def __repr__(self) -> str:
output = 'BlockIndex\n'
output += 'Block locations: %s\n' % repr(self.blocs)
output += 'Block lengths: %s' % repr(self.blengths)
diff --git a/pandas/_libs/tslibs/c_timestamp.pyx b/pandas/_libs/tslibs/c_timestamp.pyx
index 032363d867196..8e4143a053ba3 100644
--- a/pandas/_libs/tslibs/c_timestamp.pyx
+++ b/pandas/_libs/tslibs/c_timestamp.pyx
@@ -124,7 +124,7 @@ cdef class _Timestamp(datetime):
# now __reduce_ex__ is defined and higher priority than __reduce__
return self.__reduce__()
- def __repr__(self):
+ def __repr__(self) -> str:
stamp = self._repr_base
zone = None
diff --git a/pandas/_libs/tslibs/nattype.pyx b/pandas/_libs/tslibs/nattype.pyx
index 241aff0e19112..e491d6111a919 100644
--- a/pandas/_libs/tslibs/nattype.pyx
+++ b/pandas/_libs/tslibs/nattype.pyx
@@ -259,10 +259,10 @@ cdef class _NaT(datetime):
"""
return self.to_datetime64()
- def __repr__(self):
+ def __repr__(self) -> str:
return 'NaT'
- def __str__(self):
+ def __str__(self) -> str:
return 'NaT'
def isoformat(self, sep='T'):
diff --git a/pandas/_libs/tslibs/offsets.pyx b/pandas/_libs/tslibs/offsets.pyx
index aaefab6ee7ff6..434252677f1a1 100644
--- a/pandas/_libs/tslibs/offsets.pyx
+++ b/pandas/_libs/tslibs/offsets.pyx
@@ -422,7 +422,7 @@ class _BaseOffset:
# that allows us to use methods that can go in a `cdef class`
return self * 1
- def __repr__(self):
+ def __repr__(self) -> str:
className = getattr(self, '_outputName', type(self).__name__)
if abs(self.n) != 1:
diff --git a/pandas/_libs/tslibs/period.pyx b/pandas/_libs/tslibs/period.pyx
index e297d11c5144d..2512fdb891e3e 100644
--- a/pandas/_libs/tslibs/period.pyx
+++ b/pandas/_libs/tslibs/period.pyx
@@ -2215,12 +2215,12 @@ cdef class _Period:
def freqstr(self):
return self.freq.freqstr
- def __repr__(self):
+ def __repr__(self) -> str:
base, mult = get_freq_code(self.freq)
formatted = period_format(self.ordinal, base)
return "Period('%s', '%s')" % (formatted, self.freqstr)
- def __str__(self):
+ def __str__(self) -> str:
"""
Return a string representation for a particular DataFrame
"""
diff --git a/pandas/_libs/tslibs/timedeltas.pyx b/pandas/_libs/tslibs/timedeltas.pyx
index 8435f1cd7d732..9d8ed62388655 100644
--- a/pandas/_libs/tslibs/timedeltas.pyx
+++ b/pandas/_libs/tslibs/timedeltas.pyx
@@ -1142,10 +1142,10 @@ cdef class _Timedelta(timedelta):
return fmt.format(**comp_dict)
- def __repr__(self):
+ def __repr__(self) -> str:
return "Timedelta('{val}')".format(val=self._repr_base(format='long'))
- def __str__(self):
+ def __str__(self) -> str:
return self._repr_base(format='long')
def __bool__(self):
diff --git a/pandas/core/arrays/base.py b/pandas/core/arrays/base.py
index 7333254831838..2980f0d4cb906 100644
--- a/pandas/core/arrays/base.py
+++ b/pandas/core/arrays/base.py
@@ -913,7 +913,7 @@ def view(self, dtype=None) -> Union[ABCExtensionArray, np.ndarray]:
# Printing
# ------------------------------------------------------------------------
- def __repr__(self):
+ def __repr__(self) -> str:
from pandas.io.formats.printing import format_object_summary
template = "{class_name}{data}\nLength: {length}, dtype: {dtype}"
diff --git a/pandas/core/arrays/categorical.py b/pandas/core/arrays/categorical.py
index ce174baa66a97..d3e9c6a415879 100644
--- a/pandas/core/arrays/categorical.py
+++ b/pandas/core/arrays/categorical.py
@@ -2048,7 +2048,7 @@ def _get_repr(self, length=True, na_rep="NaN", footer=True):
result = formatter.to_string()
return str(result)
- def __repr__(self):
+ def __repr__(self) -> str:
"""
String representation.
"""
diff --git a/pandas/core/arrays/integer.py b/pandas/core/arrays/integer.py
index 630c3e50f2c09..08b53e54b91ef 100644
--- a/pandas/core/arrays/integer.py
+++ b/pandas/core/arrays/integer.py
@@ -45,7 +45,7 @@ class _IntegerDtype(ExtensionDtype):
type = None # type: Type
na_value = np.nan
- def __repr__(self):
+ def __repr__(self) -> str:
sign = "U" if self.is_unsigned_integer else ""
return "{sign}Int{size}Dtype()".format(sign=sign, size=8 * self.itemsize)
diff --git a/pandas/core/arrays/interval.py b/pandas/core/arrays/interval.py
index 869019cd3d222..cc41797e7872b 100644
--- a/pandas/core/arrays/interval.py
+++ b/pandas/core/arrays/interval.py
@@ -860,7 +860,7 @@ def _format_data(self):
return summary
- def __repr__(self):
+ def __repr__(self) -> str:
template = (
"{class_name}"
"{data}\n"
diff --git a/pandas/core/arrays/numpy_.py b/pandas/core/arrays/numpy_.py
index bf7404e8997c6..6f2bb095a014d 100644
--- a/pandas/core/arrays/numpy_.py
+++ b/pandas/core/arrays/numpy_.py
@@ -44,7 +44,7 @@ def __init__(self, dtype):
self._name = dtype.name
self._type = dtype.type
- def __repr__(self):
+ def __repr__(self) -> str:
return "PandasDtype({!r})".format(self.name)
@property
diff --git a/pandas/core/arrays/sparse/array.py b/pandas/core/arrays/sparse/array.py
index e1691de234335..075cdf09d531f 100644
--- a/pandas/core/arrays/sparse/array.py
+++ b/pandas/core/arrays/sparse/array.py
@@ -1515,7 +1515,7 @@ def _add_comparison_ops(cls):
# ----------
# Formatting
# -----------
- def __repr__(self):
+ def __repr__(self) -> str:
return "{self}\nFill: {fill}\n{index}".format(
self=printing.pprint_thing(self),
fill=printing.pprint_thing(self.fill_value),
diff --git a/pandas/core/arrays/sparse/dtype.py b/pandas/core/arrays/sparse/dtype.py
index 6fd73ae14fff1..4de958cc386b9 100644
--- a/pandas/core/arrays/sparse/dtype.py
+++ b/pandas/core/arrays/sparse/dtype.py
@@ -165,7 +165,7 @@ def subtype(self):
def name(self):
return "Sparse[{}, {}]".format(self.subtype.name, self.fill_value)
- def __repr__(self):
+ def __repr__(self) -> str:
return self.name
@classmethod
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 61dc5f35cadf7..bab358aa5dc8d 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -55,7 +55,7 @@ def _constructor(self):
"""class constructor (for this class it's just `__class__`"""
return self.__class__
- def __repr__(self):
+ def __repr__(self) -> str:
"""
Return a string representation for a particular object.
"""
diff --git a/pandas/core/computation/expr.py b/pandas/core/computation/expr.py
index 72367c8fb7a4f..39653c3d695b2 100644
--- a/pandas/core/computation/expr.py
+++ b/pandas/core/computation/expr.py
@@ -834,7 +834,7 @@ def assigner(self):
def __call__(self):
return self.terms(self.env)
- def __repr__(self):
+ def __repr__(self) -> str:
return printing.pprint_thing(self.terms)
def __len__(self):
diff --git a/pandas/core/computation/ops.py b/pandas/core/computation/ops.py
index dc0f381414970..fe74b6994be7c 100644
--- a/pandas/core/computation/ops.py
+++ b/pandas/core/computation/ops.py
@@ -82,7 +82,7 @@ def __init__(self, name, env, side=None, encoding=None):
def local_name(self):
return self.name.replace(_LOCAL_TAG, "")
- def __repr__(self):
+ def __repr__(self) -> str:
return pprint_thing(self.name)
def __call__(self, *args, **kwargs):
@@ -182,7 +182,7 @@ def _resolve_name(self):
def name(self):
return self.value
- def __repr__(self):
+ def __repr__(self) -> str:
# in python 2 str() of float
# can truncate shorter than repr()
return repr(self.name)
@@ -204,7 +204,7 @@ def __init__(self, op, operands, *args, **kwargs):
def __iter__(self):
return iter(self.operands)
- def __repr__(self):
+ def __repr__(self) -> str:
"""
Print a generic n-ary operator and its operands using infix notation.
"""
@@ -557,7 +557,7 @@ def __call__(self, env):
operand = self.operand(env)
return self.func(operand)
- def __repr__(self):
+ def __repr__(self) -> str:
return pprint_thing("{0}({1})".format(self.op, self.operand))
@property
@@ -582,7 +582,7 @@ def __call__(self, env):
with np.errstate(all="ignore"):
return self.func.func(*operands)
- def __repr__(self):
+ def __repr__(self) -> str:
operands = map(str, self.operands)
return pprint_thing("{0}({1})".format(self.op, ",".join(operands)))
diff --git a/pandas/core/computation/pytables.py b/pandas/core/computation/pytables.py
index 81658ab23ba46..3a2ea30cbc8b9 100644
--- a/pandas/core/computation/pytables.py
+++ b/pandas/core/computation/pytables.py
@@ -229,7 +229,7 @@ def convert_values(self):
class FilterBinOp(BinOp):
- def __repr__(self):
+ def __repr__(self) -> str:
return pprint_thing(
"[Filter : [{lhs}] -> [{op}]".format(lhs=self.filter[0], op=self.filter[1])
)
@@ -295,7 +295,7 @@ def evaluate(self):
class ConditionBinOp(BinOp):
- def __repr__(self):
+ def __repr__(self) -> str:
return pprint_thing("[Condition : [{cond}]]".format(cond=self.condition))
def invert(self):
@@ -545,7 +545,7 @@ def __init__(self, where, queryables=None, encoding=None, scope_level=0):
)
self.terms = self.parse()
- def __repr__(self):
+ def __repr__(self) -> str:
if self.terms is not None:
return pprint_thing(self.terms)
return pprint_thing(self.expr)
diff --git a/pandas/core/computation/scope.py b/pandas/core/computation/scope.py
index b11411eb2dc66..81c7b04bf3284 100644
--- a/pandas/core/computation/scope.py
+++ b/pandas/core/computation/scope.py
@@ -139,7 +139,7 @@ def __init__(
self.resolvers = DeepChainMap(*resolvers)
self.temps = {}
- def __repr__(self):
+ def __repr__(self) -> str:
scope_keys = _get_pretty_string(list(self.scope.keys()))
res_keys = _get_pretty_string(list(self.resolvers.keys()))
unicode_str = "{name}(scope={scope_keys}, resolvers={res_keys})"
diff --git a/pandas/core/dtypes/base.py b/pandas/core/dtypes/base.py
index 59ef17e3d121f..7d98a42e06257 100644
--- a/pandas/core/dtypes/base.py
+++ b/pandas/core/dtypes/base.py
@@ -71,7 +71,7 @@ class property**.
_metadata = () # type: Tuple[str, ...]
- def __str__(self):
+ def __str__(self) -> str:
return self.name
def __eq__(self, other):
diff --git a/pandas/core/dtypes/dtypes.py b/pandas/core/dtypes/dtypes.py
index 7dca588e33839..4a4ad076f14ca 100644
--- a/pandas/core/dtypes/dtypes.py
+++ b/pandas/core/dtypes/dtypes.py
@@ -415,7 +415,7 @@ def __eq__(self, other: Any) -> bool:
return True
return hash(self) == hash(other)
- def __repr__(self):
+ def __repr__(self) -> str_type:
tpl = "CategoricalDtype(categories={}ordered={})"
if self.categories is None:
data = "None, "
@@ -752,7 +752,7 @@ def construct_from_string(cls, string):
raise TypeError("Could not construct DatetimeTZDtype")
- def __str__(self):
+ def __str__(self) -> str_type:
return "datetime64[{unit}, {tz}]".format(unit=self.unit, tz=self.tz)
@property
@@ -889,7 +889,7 @@ def construct_from_string(cls, string):
pass
raise TypeError("could not construct PeriodDtype")
- def __str__(self):
+ def __str__(self) -> str_type:
return self.name
@property
@@ -1068,7 +1068,7 @@ def construct_from_string(cls, string):
def type(self):
return Interval
- def __str__(self):
+ def __str__(self) -> str_type:
if self.subtype is None:
return "interval"
return "interval[{subtype}]".format(subtype=self.subtype)
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index bafc37d478fdb..2c778b7011f76 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -2107,7 +2107,7 @@ def __setstate__(self, state):
# ----------------------------------------------------------------------
# Rendering Methods
- def __repr__(self):
+ def __repr__(self) -> str:
# string representation based upon iterating over self
# (since, by definition, `PandasContainers` are iterable)
prepr = "[%s]" % ",".join(map(pprint_thing, self))
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 81ba594c97391..31d6e2206f569 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -402,7 +402,7 @@ def __init__(
def __len__(self):
return len(self.groups)
- def __repr__(self):
+ def __repr__(self) -> str:
# TODO: Better repr for GroupBy object
return object.__repr__(self)
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index dc6336b17ac1e..ff3b4b1096ecb 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -210,7 +210,7 @@ def _set_grouper(self, obj, sort=False):
def groups(self):
return self.grouper.groups
- def __repr__(self):
+ def __repr__(self) -> str:
attrs_list = (
"{}={!r}".format(attr_name, getattr(self, attr_name))
for attr_name in self._attributes
@@ -372,7 +372,7 @@ def __init__(
self.grouper = self.grouper.astype("timedelta64[ns]")
- def __repr__(self):
+ def __repr__(self) -> str:
return "Grouping({0})".format(self.name)
def __iter__(self):
diff --git a/pandas/core/indexes/frozen.py b/pandas/core/indexes/frozen.py
index 4791ea2b70691..08c86b81b59c0 100644
--- a/pandas/core/indexes/frozen.py
+++ b/pandas/core/indexes/frozen.py
@@ -105,10 +105,10 @@ def _disabled(self, *args, **kwargs):
)
)
- def __str__(self):
+ def __str__(self) -> str:
return pprint_thing(self, quote_strings=True, escape_chars=("\t", "\r", "\n"))
- def __repr__(self):
+ def __repr__(self) -> str:
return "%s(%s)" % (self.__class__.__name__, str(self))
__setitem__ = __setslice__ = __delitem__ = __delslice__ = _disabled
@@ -148,7 +148,7 @@ def values(self):
arr = self.view(np.ndarray).copy()
return arr
- def __repr__(self):
+ def __repr__(self) -> str:
"""
Return a string representation for this object.
"""
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 448d2faf8b85f..fd5d3f2247a90 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -268,7 +268,7 @@ def make_block_same_class(self, values, placement=None, ndim=None, dtype=None):
values, placement=placement, ndim=ndim, klass=self.__class__, dtype=dtype
)
- def __repr__(self):
+ def __repr__(self) -> str:
# don't want to print out all of the items here
name = pprint_thing(self.__class__.__name__)
if self._is_single_block:
diff --git a/pandas/core/internals/concat.py b/pandas/core/internals/concat.py
index 4ba485c85d8ba..f981c00fdad36 100644
--- a/pandas/core/internals/concat.py
+++ b/pandas/core/internals/concat.py
@@ -120,7 +120,7 @@ def __init__(self, block, shape, indexers=None):
self.indexers = indexers
self.shape = shape
- def __repr__(self):
+ def __repr__(self) -> str:
return "{name}({block!r}, {indexers})".format(
name=self.__class__.__name__, block=self.block, indexers=self.indexers
)
diff --git a/pandas/core/internals/managers.py b/pandas/core/internals/managers.py
index 21ae820cfcee6..74cebd8b59fba 100644
--- a/pandas/core/internals/managers.py
+++ b/pandas/core/internals/managers.py
@@ -325,7 +325,7 @@ def _post_setstate(self):
def __len__(self):
return len(self.items)
- def __repr__(self):
+ def __repr__(self) -> str:
output = pprint_thing(self.__class__.__name__)
for i, ax in enumerate(self.axes):
if i == 0:
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index e418461883e6c..6d877bf666881 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -86,7 +86,7 @@ def __init__(self, obj, groupby=None, axis=0, kind=None, **kwargs):
if self.groupby is not None:
self.groupby._set_grouper(self._convert_obj(obj), sort=True)
- def __str__(self):
+ def __str__(self) -> str:
"""
Provide a nice str repr of our rolling object.
"""
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 73a05b4cdfa66..bdc319c6d7bab 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -1555,7 +1555,7 @@ def reset_index(self, level=None, drop=False, name=None, inplace=False):
# ----------------------------------------------------------------------
# Rendering Methods
- def __repr__(self):
+ def __repr__(self) -> str:
"""
Return a string representation for a particular Series.
"""
diff --git a/pandas/errors/__init__.py b/pandas/errors/__init__.py
index 883af5c2e62f0..73cc40ae0e0d3 100644
--- a/pandas/errors/__init__.py
+++ b/pandas/errors/__init__.py
@@ -174,7 +174,7 @@ def __init__(self, class_instance, methodtype="method"):
self.methodtype = methodtype
self.class_instance = class_instance
- def __str__(self):
+ def __str__(self) -> str:
if self.methodtype == "classmethod":
name = self.class_instance.__name__
else:
diff --git a/pandas/io/msgpack/exceptions.py b/pandas/io/msgpack/exceptions.py
index 40f5a8af8f583..2966f69920930 100644
--- a/pandas/io/msgpack/exceptions.py
+++ b/pandas/io/msgpack/exceptions.py
@@ -19,7 +19,7 @@ def __init__(self, unpacked, extra):
self.unpacked = unpacked
self.extra = extra
- def __str__(self):
+ def __str__(self) -> str:
return "unpack(b) received extra data."
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index 35e6d53127e59..8580e0069ccdf 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -543,7 +543,7 @@ def __contains__(self, key):
def __len__(self):
return len(self.groups())
- def __repr__(self):
+ def __repr__(self) -> str:
return "{type}\nFile path: {path}\n".format(
type=type(self), path=pprint_thing(self._path)
)
@@ -1725,7 +1725,7 @@ def set_table(self, table):
self.table = table
return self
- def __repr__(self):
+ def __repr__(self) -> str:
temp = tuple(
map(pprint_thing, (self.name, self.cname, self.axis, self.pos, self.kind))
)
@@ -2052,7 +2052,7 @@ def __init__(
self.set_data(data)
self.set_metadata(metadata)
- def __repr__(self):
+ def __repr__(self) -> str:
temp = tuple(
map(
pprint_thing, (self.name, self.cname, self.dtype, self.kind, self.shape)
@@ -2518,7 +2518,7 @@ def pandas_type(self):
def format_type(self):
return "fixed"
- def __repr__(self):
+ def __repr__(self) -> str:
""" return a pretty representation of myself """
self.infer_axes()
s = self.shape
@@ -3213,7 +3213,7 @@ def table_type_short(self):
def format_type(self):
return "table"
- def __repr__(self):
+ def __repr__(self) -> str:
""" return a pretty representation of myself """
self.infer_axes()
dc = ",dc->[{columns}]".format(
diff --git a/pandas/io/stata.py b/pandas/io/stata.py
index 07475f224bd5f..d62c3f7d2e3b8 100644
--- a/pandas/io/stata.py
+++ b/pandas/io/stata.py
@@ -862,10 +862,10 @@ def __init__(self, value):
lambda self: self._value, doc="The binary representation of the missing value."
)
- def __str__(self):
+ def __str__(self) -> str:
return self.string
- def __repr__(self):
+ def __repr__(self) -> str:
# not perfect :-/
return "{cls}({obj})".format(cls=self.__class__, obj=self)
diff --git a/pandas/tests/extension/decimal/array.py b/pandas/tests/extension/decimal/array.py
index a1988744d76a1..93816e3a8a613 100644
--- a/pandas/tests/extension/decimal/array.py
+++ b/pandas/tests/extension/decimal/array.py
@@ -22,7 +22,7 @@ class DecimalDtype(ExtensionDtype):
def __init__(self, context=None):
self.context = context or decimal.getcontext()
- def __repr__(self):
+ def __repr__(self) -> str:
return "DecimalDtype(context={})".format(self.context)
@classmethod
diff --git a/pandas/tests/frame/test_alter_axes.py b/pandas/tests/frame/test_alter_axes.py
index 11d73fc37105e..9b76be18b0e88 100644
--- a/pandas/tests/frame/test_alter_axes.py
+++ b/pandas/tests/frame/test_alter_axes.py
@@ -341,7 +341,7 @@ def __init__(self, name, color):
self.name = name
self.color = color
- def __str__(self):
+ def __str__(self) -> str:
return "<Thing {self.name!r}>".format(self=self)
# necessary for pretty KeyError
@@ -380,7 +380,7 @@ def test_set_index_custom_label_hashable_iterable(self):
class Thing(frozenset):
# need to stabilize repr for KeyError (due to random order in sets)
- def __repr__(self):
+ def __repr__(self) -> str:
tmp = sorted(list(self))
# double curly brace prints one brace in format string
return "frozenset({{{}}})".format(", ".join(map(repr, tmp)))
@@ -418,7 +418,7 @@ def __init__(self, name, color):
self.name = name
self.color = color
- def __str__(self):
+ def __str__(self) -> str:
return "<Thing {self.name!r}>".format(self=self)
thing1 = Thing("One", "red")
diff --git a/pandas/tests/indexing/test_indexing.py b/pandas/tests/indexing/test_indexing.py
index d611dc5497cca..d6d3763981131 100644
--- a/pandas/tests/indexing/test_indexing.py
+++ b/pandas/tests/indexing/test_indexing.py
@@ -591,7 +591,7 @@ class TO:
def __init__(self, value):
self.value = value
- def __str__(self):
+ def __str__(self) -> str:
return "[{0}]".format(self.value)
__repr__ = __str__
diff --git a/pandas/tests/internals/test_internals.py b/pandas/tests/internals/test_internals.py
index 16f14f35fdbae..ee7fca6ec7672 100644
--- a/pandas/tests/internals/test_internals.py
+++ b/pandas/tests/internals/test_internals.py
@@ -1185,10 +1185,10 @@ def __init__(self, value, dtype):
def __array__(self):
return np.array(self.value, dtype=self.dtype)
- def __str__(self):
+ def __str__(self) -> str:
return "DummyElement({}, {})".format(self.value, self.dtype)
- def __repr__(self):
+ def __repr__(self) -> str:
return str(self)
def astype(self, dtype, copy=False):
diff --git a/pandas/tests/io/json/test_pandas.py b/pandas/tests/io/json/test_pandas.py
index eaa46c4e9dc9b..aa065b6e13079 100644
--- a/pandas/tests/io/json/test_pandas.py
+++ b/pandas/tests/io/json/test_pandas.py
@@ -594,7 +594,7 @@ def __init__(self, hexed):
self.hexed = hexed
self.binary = bytes.fromhex(hexed)
- def __str__(self):
+ def __str__(self) -> str:
return self.hexed
hexed = "574b4454ba8c5eb4f98a8f45"
diff --git a/pandas/tests/io/json/test_ujson.py b/pandas/tests/io/json/test_ujson.py
index 20e2690084e2a..8dcc77fc2fbc1 100644
--- a/pandas/tests/io/json/test_ujson.py
+++ b/pandas/tests/io/json/test_ujson.py
@@ -621,7 +621,7 @@ def __init__(self, val):
def recursive_attr(self):
return _TestObject("recursive_attr")
- def __str__(self):
+ def __str__(self) -> str:
return str(self.val)
msg = "Maximum recursion level reached"
diff --git a/pandas/tests/series/test_repr.py b/pandas/tests/series/test_repr.py
index 9f881f5a5aa29..f1661ad034e4c 100644
--- a/pandas/tests/series/test_repr.py
+++ b/pandas/tests/series/test_repr.py
@@ -227,7 +227,7 @@ class County:
name = "San Sebastián"
state = "PR"
- def __repr__(self):
+ def __repr__(self) -> str:
return self.name + ", " + self.state
cat = pd.Categorical([County() for _ in range(61)])
diff --git a/pandas/tests/series/test_ufunc.py b/pandas/tests/series/test_ufunc.py
index 8144a3931b9b8..c8a127f89bf91 100644
--- a/pandas/tests/series/test_ufunc.py
+++ b/pandas/tests/series/test_ufunc.py
@@ -285,7 +285,7 @@ def __add__(self, other):
def __eq__(self, other):
return type(other) is Thing and self.value == other.value
- def __repr__(self):
+ def __repr__(self) -> str:
return "Thing({})".format(self.value)
s = pd.Series([Thing(1), Thing(2)])
diff --git a/pandas/tseries/holiday.py b/pandas/tseries/holiday.py
index eb8600031439f..d4f02286ff8d6 100644
--- a/pandas/tseries/holiday.py
+++ b/pandas/tseries/holiday.py
@@ -183,7 +183,7 @@ class from pandas.tseries.offsets
assert days_of_week is None or type(days_of_week) == tuple
self.days_of_week = days_of_week
- def __repr__(self):
+ def __repr__(self) -> str:
info = ""
if self.year is not None:
info += "year={year}, ".format(year=self.year)
diff --git a/pandas/util/_depr_module.py b/pandas/util/_depr_module.py
index 54f090ede3fc4..45e7db9281837 100644
--- a/pandas/util/_depr_module.py
+++ b/pandas/util/_depr_module.py
@@ -38,7 +38,7 @@ def __dir__(self):
deprmodule = self._import_deprmod()
return dir(deprmodule)
- def __repr__(self):
+ def __repr__(self) -> str:
deprmodule = self._import_deprmod()
return repr(deprmodule)
| Operation: Get This Over With | https://api.github.com/repos/pandas-dev/pandas/pulls/29467 | 2019-11-07T18:16:09Z | 2019-11-07T21:01:52Z | 2019-11-07T21:01:52Z | 2019-11-07T21:07:21Z |
PR09 batch 4 | diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 40efc4c65476a..301cfa53e3e0b 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -5881,13 +5881,13 @@ def pivot(self, index=None, columns=None, values=None):
hierarchical columns whose top level are the function names
(inferred from the function objects themselves)
If dict is passed, the key is column to aggregate and value
- is function or list of functions
+ is function or list of functions.
fill_value : scalar, default None
- Value to replace missing values with
+ Value to replace missing values with.
margins : bool, default False
- Add all row / columns (e.g. for subtotal / grand totals)
+ Add all row / columns (e.g. for subtotal / grand totals).
dropna : bool, default True
- Do not include columns whose entries are all NaN
+ Do not include columns whose entries are all NaN.
margins_name : str, default 'All'
Name of the row / column that will contain the totals
when margins is True.
@@ -5901,6 +5901,7 @@ def pivot(self, index=None, columns=None, values=None):
Returns
-------
DataFrame
+ An Excel style pivot table.
See Also
--------
diff --git a/pandas/core/indexes/interval.py b/pandas/core/indexes/interval.py
index c9554016630cd..0c077702b4cb4 100644
--- a/pandas/core/indexes/interval.py
+++ b/pandas/core/indexes/interval.py
@@ -1407,24 +1407,24 @@ def interval_range(
Parameters
----------
start : numeric or datetime-like, default None
- Left bound for generating intervals
+ Left bound for generating intervals.
end : numeric or datetime-like, default None
- Right bound for generating intervals
+ Right bound for generating intervals.
periods : int, default None
- Number of periods to generate
+ Number of periods to generate.
freq : numeric, str, or DateOffset, default None
The length of each interval. Must be consistent with the type of start
and end, e.g. 2 for numeric, or '5H' for datetime-like. Default is 1
for numeric and 'D' for datetime-like.
name : str, default None
- Name of the resulting IntervalIndex
+ Name of the resulting IntervalIndex.
closed : {'left', 'right', 'both', 'neither'}, default 'right'
Whether the intervals are closed on the left-side, right-side, both
or neither.
Returns
-------
- rng : IntervalIndex
+ IntervalIndex
See Also
--------
diff --git a/pandas/core/indexes/period.py b/pandas/core/indexes/period.py
index a0f16789621c7..ca7be9ba512da 100644
--- a/pandas/core/indexes/period.py
+++ b/pandas/core/indexes/period.py
@@ -997,28 +997,28 @@ def memory_usage(self, deep=False):
def period_range(start=None, end=None, periods=None, freq=None, name=None):
"""
- Return a fixed frequency PeriodIndex, with day (calendar) as the default
- frequency.
+ Return a fixed frequency PeriodIndex.
+
+ The day (calendar) is the default frequency.
Parameters
----------
start : str or period-like, default None
- Left bound for generating periods
+ Left bound for generating periods.
end : str or period-like, default None
- Right bound for generating periods
+ Right bound for generating periods.
periods : int, default None
- Number of periods to generate
+ Number of periods to generate.
freq : str or DateOffset, optional
Frequency alias. By default the freq is taken from `start` or `end`
if those are Period objects. Otherwise, the default is ``"D"`` for
daily frequency.
-
name : str, default None
- Name of the resulting PeriodIndex
+ Name of the resulting PeriodIndex.
Returns
-------
- prng : PeriodIndex
+ PeriodIndex
Notes
-----
diff --git a/pandas/core/reshape/melt.py b/pandas/core/reshape/melt.py
index 98fee491e0a73..f7d9462d2ec32 100644
--- a/pandas/core/reshape/melt.py
+++ b/pandas/core/reshape/melt.py
@@ -206,12 +206,12 @@ def wide_to_long(df, stubnames, i, j, sep: str = "", suffix: str = r"\d+"):
Parameters
----------
df : DataFrame
- The wide-format DataFrame
+ The wide-format DataFrame.
stubnames : str or list-like
The stub name(s). The wide format variables are assumed to
start with the stub names.
i : str or list-like
- Column(s) to use as id variable(s)
+ Column(s) to use as id variable(s).
j : str
The name of the sub-observation variable. What you wish to name your
suffix in the long format.
@@ -219,14 +219,14 @@ def wide_to_long(df, stubnames, i, j, sep: str = "", suffix: str = r"\d+"):
A character indicating the separation of the variable names
in the wide format, to be stripped from the names in the long format.
For example, if your column names are A-suffix1, A-suffix2, you
- can strip the hyphen by specifying `sep='-'`
+ can strip the hyphen by specifying `sep='-'`.
suffix : str, default '\\d+'
A regular expression capturing the wanted suffixes. '\\d+' captures
numeric suffixes. Suffixes with no numbers could be specified with the
negated character class '\\D+'. You can also further disambiguate
suffixes, for example, if your wide variables are of the form
A-one, B-two,.., and you have an unrelated column A-rating, you can
- ignore the last one by specifying `suffix='(!?one|two)'`
+ ignore the last one by specifying `suffix='(!?one|two)'`.
.. versionchanged:: 0.23.0
When all suffixes are numeric, they are cast to int64/float64.
@@ -360,7 +360,7 @@ def wide_to_long(df, stubnames, i, j, sep: str = "", suffix: str = r"\d+"):
>>> stubnames = sorted(
... set([match[0] for match in df.columns.str.findall(
- ... r'[A-B]\(.*\)').values if match != [] ])
+ ... r'[A-B]\(.*\)').values if match != []])
... )
>>> list(stubnames)
['A(weekly)', 'B(weekly)']
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index a189b2cd1ab84..956642b51ce97 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -176,9 +176,10 @@ def merge_ordered(
how="outer",
):
"""
- Perform merge with optional filling/interpolation designed for ordered
- data like time series data. Optionally perform group-wise merge (see
- examples).
+ Perform merge with optional filling/interpolation.
+
+ Designed for ordered data like time series data. Optionally
+ perform group-wise merge (see examples).
Parameters
----------
@@ -189,18 +190,18 @@ def merge_ordered(
left_on : label or list, or array-like
Field names to join on in left DataFrame. Can be a vector or list of
vectors of the length of the DataFrame to use a particular vector as
- the join key instead of columns
+ the join key instead of columns.
right_on : label or list, or array-like
Field names to join on in right DataFrame or vector/list of vectors per
- left_on docs
+ left_on docs.
left_by : column name or list of column names
Group left DataFrame by group columns and merge piece by piece with
- right DataFrame
+ right DataFrame.
right_by : column name or list of column names
Group right DataFrame by group columns and merge piece by piece with
- left DataFrame
+ left DataFrame.
fill_method : {'ffill', None}, default None
- Interpolation method for data
+ Interpolation method for data.
suffixes : Sequence, default is ("_x", "_y")
A length-2 sequence where each element is optionally a string
indicating the suffix to add to overlapping column names in
@@ -214,13 +215,13 @@ def merge_ordered(
* left: use only keys from left frame (SQL: left outer join)
* right: use only keys from right frame (SQL: right outer join)
* outer: use union of keys from both frames (SQL: full outer join)
- * inner: use intersection of keys from both frames (SQL: inner join)
+ * inner: use intersection of keys from both frames (SQL: inner join).
Returns
-------
- merged : DataFrame
- The output type will the be same as 'left', if it is a subclass
- of DataFrame.
+ DataFrame
+ The merged DataFrame output type will the be same as
+ 'left', if it is a subclass of DataFrame.
See Also
--------
@@ -229,15 +230,21 @@ def merge_ordered(
Examples
--------
- >>> A >>> B
- key lvalue group key rvalue
- 0 a 1 a 0 b 1
- 1 c 2 a 1 c 2
- 2 e 3 a 2 d 3
+ >>> A
+ key lvalue group
+ 0 a 1 a
+ 1 c 2 a
+ 2 e 3 a
3 a 1 b
4 c 2 b
5 e 3 b
+ >>> B
+ Key rvalue
+ 0 b 1
+ 1 c 2
+ 2 d 3
+
>>> merge_ordered(A, B, fill_method='ffill', left_by='group')
group key lvalue rvalue
0 a a 1 NaN
diff --git a/pandas/core/tools/datetimes.py b/pandas/core/tools/datetimes.py
index 70143e4603a4b..bb8d15896b727 100644
--- a/pandas/core/tools/datetimes.py
+++ b/pandas/core/tools/datetimes.py
@@ -577,14 +577,12 @@ def to_datetime(
Parameters
----------
- arg : int, float, str, datetime, list, tuple, 1-d array, Series
- or DataFrame/dict-like
-
+ arg : int, float, str, datetime, list, tuple, 1-d array, Series DataFrame/dict-like
+ The object to convert to a datetime.
errors : {'ignore', 'raise', 'coerce'}, default 'raise'
-
- - If 'raise', then invalid parsing will raise an exception
- - If 'coerce', then invalid parsing will be set as NaT
- - If 'ignore', then invalid parsing will return the input
+ - If 'raise', then invalid parsing will raise an exception.
+ - If 'coerce', then invalid parsing will be set as NaT.
+ - If 'ignore', then invalid parsing will return the input.
dayfirst : bool, default False
Specify a date parse order if `arg` is str or its list-likes.
If True, parses dates with the day first, eg 10/11/12 is parsed as
@@ -605,7 +603,6 @@ def to_datetime(
Return UTC DatetimeIndex if True (converting any tz-aware
datetime.datetime objects as well).
box : bool, default True
-
- If True returns a DatetimeIndex or Index-like object
- If False returns ndarray of values.
@@ -615,17 +612,17 @@ def to_datetime(
respectively.
format : str, default None
- strftime to parse time, eg "%d/%m/%Y", note that "%f" will parse
+ The strftime to parse time, eg "%d/%m/%Y", note that "%f" will parse
all the way up to nanoseconds.
See strftime documentation for more information on choices:
- https://docs.python.org/3/library/datetime.html#strftime-and-strptime-behavior
+ https://docs.python.org/3/library/datetime.html#strftime-and-strptime-behavior.
exact : bool, True by default
-
+ Behaves as:
- If True, require an exact format match.
- If False, allow the format to match anywhere in the target string.
unit : str, default 'ns'
- unit of the arg (D,s,ms,us,ns) denote the unit, which is an
+ The unit of the arg (D,s,ms,us,ns) denote the unit, which is an
integer or float number. This will be based off the origin.
Example, with unit='ms' and origin='unix' (the default), this
would calculate the number of milliseconds to the unix epoch start.
@@ -652,11 +649,12 @@ def to_datetime(
.. versionadded:: 0.23.0
.. versionchanged:: 0.25.0
- - changed default value from False to True
+ - changed default value from False to True.
Returns
-------
- ret : datetime if parsing succeeded.
+ datetime
+ If parsing succeeded.
Return type depends on input:
- list-like: DatetimeIndex
@@ -712,10 +710,10 @@ def to_datetime(
4 3/12/2000
dtype: object
- >>> %timeit pd.to_datetime(s,infer_datetime_format=True) # doctest: +SKIP
+ >>> %timeit pd.to_datetime(s, infer_datetime_format=True) # doctest: +SKIP
100 loops, best of 3: 10.4 ms per loop
- >>> %timeit pd.to_datetime(s,infer_datetime_format=False) # doctest: +SKIP
+ >>> %timeit pd.to_datetime(s, infer_datetime_format=False) # doctest: +SKIP
1 loop, best of 3: 471 ms per loop
Using a unix epoch time
diff --git a/pandas/core/util/hashing.py b/pandas/core/util/hashing.py
index 011ea1b8e42f2..23c370638b572 100644
--- a/pandas/core/util/hashing.py
+++ b/pandas/core/util/hashing.py
@@ -67,11 +67,11 @@ def hash_pandas_object(
Parameters
----------
index : bool, default True
- include the index in the hash (if Series/DataFrame)
+ Include the index in the hash (if Series/DataFrame).
encoding : str, default 'utf8'
- encoding for data & key when strings
+ Encoding for data & key when strings.
hash_key : str, default _default_hash_key
- hash_key for string key to encode
+ Hash_key for string key to encode.
categorize : bool, default True
Whether to first categorize object arrays before hashing. This is more
efficient when the array contains duplicate values.
@@ -253,9 +253,9 @@ def hash_array(
----------
vals : ndarray, Categorical
encoding : str, default 'utf8'
- encoding for data & key when strings
+ Encoding for data & key when strings.
hash_key : str, default _default_hash_key
- hash_key for string key to encode
+ Hash_key for string key to encode.
categorize : bool, default True
Whether to first categorize object arrays before hashing. This is more
efficient when the array contains duplicate values.
diff --git a/pandas/io/excel/_base.py b/pandas/io/excel/_base.py
index 1f1ad55969d6f..d0ab6dd37596c 100644
--- a/pandas/io/excel/_base.py
+++ b/pandas/io/excel/_base.py
@@ -79,8 +79,6 @@
subset of data is selected with ``usecols``, index_col
is based on the subset.
usecols : int, str, list-like, or callable default None
- Return a subset of the columns.
-
* If None, then parse all columns.
* If int, then indicates last column to be parsed.
@@ -98,6 +96,8 @@
* If callable, then evaluate each column name against it and parse the
column if the callable returns ``True``.
+ Returns a subset of the columns according to behavior above.
+
.. versionadded:: 0.24.0
squeeze : bool, default False
diff --git a/pandas/io/packers.py b/pandas/io/packers.py
index c0ace7996e1b9..253441ab25813 100644
--- a/pandas/io/packers.py
+++ b/pandas/io/packers.py
@@ -191,7 +191,7 @@ def read_msgpack(path_or_buf, encoding="utf-8", iterator=False, **kwargs):
``StringIO``.
encoding : Encoding for decoding msgpack str type
iterator : boolean, if True, return an iterator to the unpacker
- (default is False)
+ (default is False).
Returns
-------
| chunk of #28602 | https://api.github.com/repos/pandas-dev/pandas/pulls/29466 | 2019-11-07T15:58:25Z | 2019-11-08T15:49:14Z | 2019-11-08T15:49:14Z | 2020-01-06T16:47:00Z |
Pin dateutil to 2.8.0 in requirements | diff --git a/environment.yml b/environment.yml
index e9ac76f5bc52c..6d95fbc1a017e 100644
--- a/environment.yml
+++ b/environment.yml
@@ -5,7 +5,7 @@ dependencies:
# required
- numpy>=1.15
- python=3.7
- - python-dateutil>=2.6.1
+ - python-dateutil>=2.6.1,<=2.8.0
- pytz
# benchmarks
diff --git a/requirements-dev.txt b/requirements-dev.txt
index 13e2c95126f0c..439aa9853dda1 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -1,5 +1,5 @@
numpy>=1.15
-python-dateutil>=2.6.1
+python-dateutil>=2.6.1,<=2.8.0
pytz
asv
cython>=0.29.13
@@ -64,4 +64,4 @@ xlsxwriter
xlwt
odfpy
pyreadstat
-git+https://github.com/pandas-dev/pandas-sphinx-theme.git@master
\ No newline at end of file
+git+https://github.com/pandas-dev/pandas-sphinx-theme.git@master
| botocore has pinned dateutil to <=2.8.0, which conflicts with our dateutil here and leads to
```
ERROR: botocore 1.13.12 has requirement python-dateutil<2.8.1,>=2.1; python_version >= "2.7", but you'll have python-dateutil 2.8.1 which is incompatible.
```
See https://github.com/boto/botocore/commit/e87e7a745fd972815b235a9ee685232745aa94f9
Closes #29465 | https://api.github.com/repos/pandas-dev/pandas/pulls/29464 | 2019-11-07T15:29:45Z | 2019-12-01T01:02:29Z | null | 2019-12-01T01:02:30Z |
CLN: type up core.groupby.grouper.get_grouper | diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 31d6e2206f569..e73be29d5b104 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -379,9 +379,9 @@ def __init__(
self.mutated = kwargs.pop("mutated", False)
if grouper is None:
- from pandas.core.groupby.grouper import _get_grouper
+ from pandas.core.groupby.grouper import get_grouper
- grouper, exclusions, obj = _get_grouper(
+ grouper, exclusions, obj = get_grouper(
obj,
keys,
axis=axis,
@@ -1802,9 +1802,9 @@ def nth(self, n: Union[int, List[int]], dropna: Optional[str] = None) -> DataFra
# create a grouper with the original parameters, but on dropped
# object
- from pandas.core.groupby.grouper import _get_grouper
+ from pandas.core.groupby.grouper import get_grouper
- grouper, _, _ = _get_grouper(
+ grouper, _, _ = get_grouper(
dropped,
key=self.keys,
axis=self.axis,
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index ff3b4b1096ecb..370abe75e1327 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -3,7 +3,7 @@
split-apply-combine paradigm.
"""
-from typing import Optional, Tuple
+from typing import Hashable, List, Optional, Tuple
import warnings
import numpy as np
@@ -26,7 +26,6 @@
from pandas.core.arrays import Categorical, ExtensionArray
import pandas.core.common as com
from pandas.core.frame import DataFrame
-from pandas.core.generic import NDFrame
from pandas.core.groupby.categorical import recode_for_groupby, recode_from_groupby
from pandas.core.groupby.ops import BaseGrouper
from pandas.core.index import CategoricalIndex, Index, MultiIndex
@@ -134,7 +133,7 @@ def _get_grouper(self, obj, validate=True):
"""
self._set_grouper(obj)
- self.grouper, exclusions, self.obj = _get_grouper(
+ self.grouper, exclusions, self.obj = get_grouper(
self.obj,
[self.key],
axis=self.axis,
@@ -429,8 +428,8 @@ def groups(self) -> dict:
return self.index.groupby(Categorical.from_codes(self.codes, self.group_index))
-def _get_grouper(
- obj: NDFrame,
+def get_grouper(
+ obj: FrameOrSeries,
key=None,
axis: int = 0,
level=None,
@@ -438,9 +437,9 @@ def _get_grouper(
observed=False,
mutated=False,
validate=True,
-):
+) -> Tuple[BaseGrouper, List[Hashable], FrameOrSeries]:
"""
- create and return a BaseGrouper, which is an internal
+ Create and return a BaseGrouper, which is an internal
mapping of how to create the grouper indexers.
This may be composed of multiple Grouping objects, indicating
multiple groupers
@@ -456,9 +455,9 @@ def _get_grouper(
a BaseGrouper.
If observed & we have a categorical grouper, only show the observed
- values
+ values.
- If validate, then check for key/level overlaps
+ If validate, then check for key/level overlaps.
"""
group_axis = obj._get_axis(axis)
@@ -517,7 +516,7 @@ def _get_grouper(
if key.key is None:
return grouper, [], obj
else:
- return grouper, {key.key}, obj
+ return grouper, [key.key], obj
# already have a BaseGrouper, just return it
elif isinstance(key, BaseGrouper):
@@ -530,10 +529,8 @@ def _get_grouper(
# unhashable elements of `key`. Any unhashable elements implies that
# they wanted a list of keys.
# https://github.com/pandas-dev/pandas/issues/18314
- is_tuple = isinstance(key, tuple)
- all_hashable = is_tuple and is_hashable(key)
-
- if is_tuple:
+ if isinstance(key, tuple):
+ all_hashable = is_hashable(key)
if (
all_hashable and key not in obj and set(key).issubset(obj)
) or not all_hashable:
@@ -573,7 +570,8 @@ def _get_grouper(
all_in_columns_index = all(
g in obj.columns or g in obj.index.names for g in keys
)
- elif isinstance(obj, Series):
+ else:
+ assert isinstance(obj, Series)
all_in_columns_index = all(g in obj.index.names for g in keys)
if not all_in_columns_index:
@@ -586,8 +584,8 @@ def _get_grouper(
else:
levels = [level] * len(keys)
- groupings = []
- exclusions = []
+ groupings = [] # type: List[Grouping]
+ exclusions = [] # type: List[Hashable]
# if the actual grouper should be obj[key]
def is_in_axis(key) -> bool:
| Add types + make some minor cleanups. | https://api.github.com/repos/pandas-dev/pandas/pulls/29458 | 2019-11-07T07:33:28Z | 2019-11-08T08:54:52Z | 2019-11-08T08:54:52Z | 2019-11-08T14:58:58Z |
DEPR: is_extension_type | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 370e1c09d33aa..8c2b140cc2311 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -213,7 +213,8 @@ Deprecations
- ``Index.set_value`` has been deprecated. For a given index ``idx``, array ``arr``,
value in ``idx`` of ``idx_val`` and a new value of ``val``, ``idx.set_value(arr, idx_val, val)``
is equivalent to ``arr[idx.get_loc(idx_val)] = val``, which should be used instead (:issue:`28621`).
--
+- :func:`is_extension_type` is deprecated, :func:`is_extension_array_dtype` should be used instead (:issue:`29457`)
+
.. _whatsnew_1000.prior_deprecations:
diff --git a/pandas/core/apply.py b/pandas/core/apply.py
index f402154dc91ca..e7b088658ac5d 100644
--- a/pandas/core/apply.py
+++ b/pandas/core/apply.py
@@ -7,7 +7,7 @@
from pandas.core.dtypes.common import (
is_dict_like,
- is_extension_type,
+ is_extension_array_dtype,
is_list_like,
is_sequence,
)
@@ -230,7 +230,7 @@ def apply_standard(self):
# as demonstrated in gh-12244
if (
self.result_type in ["reduce", None]
- and not self.dtypes.apply(is_extension_type).any()
+ and not self.dtypes.apply(is_extension_array_dtype).any()
# Disallow complex_internals since libreduction shortcut
# cannot handle MultiIndex
and not self.agg_axis._has_complex_internals
diff --git a/pandas/core/arrays/datetimes.py b/pandas/core/arrays/datetimes.py
index 788cd2a3ce5b7..7cd103d12fa8a 100644
--- a/pandas/core/arrays/datetimes.py
+++ b/pandas/core/arrays/datetimes.py
@@ -31,7 +31,7 @@
is_datetime64_ns_dtype,
is_datetime64tz_dtype,
is_dtype_equal,
- is_extension_type,
+ is_extension_array_dtype,
is_float_dtype,
is_object_dtype,
is_period_dtype,
@@ -2131,7 +2131,7 @@ def maybe_convert_dtype(data, copy):
data = data.categories.take(data.codes, fill_value=NaT)._values
copy = False
- elif is_extension_type(data) and not is_datetime64tz_dtype(data):
+ elif is_extension_array_dtype(data) and not is_datetime64tz_dtype(data):
# Includes categorical
# TODO: We have no tests for these
data = np.array(data, dtype=np.object_)
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 61dc5f35cadf7..a1985f4afc754 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -23,7 +23,6 @@
is_datetime64tz_dtype,
is_datetimelike,
is_extension_array_dtype,
- is_extension_type,
is_list_like,
is_object_dtype,
is_scalar,
@@ -1268,7 +1267,7 @@ def _map_values(self, mapper, na_action=None):
# use the built in categorical series mapper which saves
# time by mapping the categories instead of all values
return self._values.map(mapper)
- if is_extension_type(self.dtype):
+ if is_extension_array_dtype(self.dtype):
values = self._values
else:
values = self.values
@@ -1279,7 +1278,8 @@ def _map_values(self, mapper, na_action=None):
return new_values
# we must convert to python types
- if is_extension_type(self.dtype):
+ if is_extension_array_dtype(self.dtype) and hasattr(self._values, "map"):
+ # GH#23179 some EAs do not have `map`
values = self._values
if na_action is not None:
raise NotImplementedError
diff --git a/pandas/core/construction.py b/pandas/core/construction.py
index 5e8b28267f24f..c0b08beead0ca 100644
--- a/pandas/core/construction.py
+++ b/pandas/core/construction.py
@@ -27,7 +27,6 @@
is_categorical_dtype,
is_datetime64_ns_dtype,
is_extension_array_dtype,
- is_extension_type,
is_float_dtype,
is_integer_dtype,
is_iterator,
@@ -527,7 +526,7 @@ def _try_cast(
and not (is_iterator(subarr) or isinstance(subarr, np.ndarray))
):
subarr = construct_1d_object_array_from_listlike(subarr)
- elif not is_extension_type(subarr):
+ elif not is_extension_array_dtype(subarr):
subarr = construct_1d_ndarray_preserving_na(subarr, dtype, copy=copy)
except OutOfBoundsDatetime:
# in case of out of bound datetime64 -> always raise
diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index fad80d6bf5745..98874fce288bc 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -30,7 +30,6 @@
is_datetimelike,
is_dtype_equal,
is_extension_array_dtype,
- is_extension_type,
is_float,
is_float_dtype,
is_integer,
@@ -633,7 +632,7 @@ def infer_dtype_from_array(arr, pandas_dtype: bool = False):
if not is_list_like(arr):
arr = [arr]
- if pandas_dtype and is_extension_type(arr):
+ if pandas_dtype and is_extension_array_dtype(arr):
return arr.dtype, arr
elif isinstance(arr, ABCSeries):
@@ -695,7 +694,7 @@ def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
# We allow arbitrary fill values for object dtype
raise ValueError("fill_value must be a scalar")
- if is_extension_type(values):
+ if is_extension_array_dtype(values):
if copy:
values = values.copy()
else:
diff --git a/pandas/core/dtypes/common.py b/pandas/core/dtypes/common.py
index 2a46d335ff512..41cbc731e18c4 100644
--- a/pandas/core/dtypes/common.py
+++ b/pandas/core/dtypes/common.py
@@ -1674,6 +1674,8 @@ def is_extension_type(arr):
"""
Check whether an array-like is of a pandas extension class instance.
+ .. deprecated:: 1.0.0
+
Extension classes include categoricals, pandas sparse objects (i.e.
classes represented within the pandas library and not ones external
to it like scipy sparse matrices), and datetime-like arrays.
@@ -1716,6 +1718,12 @@ def is_extension_type(arr):
>>> is_extension_type(s)
True
"""
+ warnings.warn(
+ "'is_extension_type' is deprecated and will be removed in a future "
+ "version. Use 'is_extension_array_dtype' instead.",
+ FutureWarning,
+ stacklevel=2,
+ )
if is_categorical(arr):
return True
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 40efc4c65476a..b005b70eedc7e 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -71,7 +71,6 @@
is_dict_like,
is_dtype_equal,
is_extension_array_dtype,
- is_extension_type,
is_float_dtype,
is_hashable,
is_integer,
@@ -3690,7 +3689,7 @@ def reindexer(value):
value = maybe_cast_to_datetime(value, infer_dtype)
# return internal types directly
- if is_extension_type(value) or is_extension_array_dtype(value):
+ if is_extension_array_dtype(value):
return value
# broadcast across multiple columns if necessary
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 448d2faf8b85f..ce889ea95f782 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -37,7 +37,6 @@
is_datetime64tz_dtype,
is_dtype_equal,
is_extension_array_dtype,
- is_extension_type,
is_float_dtype,
is_integer,
is_integer_dtype,
@@ -2605,10 +2604,6 @@ def should_store(self, value):
value.dtype.type,
(np.integer, np.floating, np.complexfloating, np.datetime64, np.bool_),
)
- or
- # TODO(ExtensionArray): remove is_extension_type
- # when all extension arrays have been ported.
- is_extension_type(value)
or is_extension_array_dtype(value)
)
@@ -3168,7 +3163,7 @@ def _putmask_preserve(nv, n):
# change the dtype if needed
dtype, _ = maybe_promote(n.dtype)
- if is_extension_type(v.dtype) and is_object_dtype(dtype):
+ if is_extension_array_dtype(v.dtype) and is_object_dtype(dtype):
v = v._internal_get_values(dtype)
else:
v = v.astype(dtype)
diff --git a/pandas/core/internals/managers.py b/pandas/core/internals/managers.py
index 21ae820cfcee6..d32e026351e22 100644
--- a/pandas/core/internals/managers.py
+++ b/pandas/core/internals/managers.py
@@ -20,7 +20,6 @@
_NS_DTYPE,
is_datetimelike_v_numeric,
is_extension_array_dtype,
- is_extension_type,
is_list_like,
is_numeric_v_string_like,
is_scalar,
@@ -1034,11 +1033,7 @@ def set(self, item, value):
# FIXME: refactor, clearly separate broadcasting & zip-like assignment
# can prob also fix the various if tests for sparse/categorical
- # TODO(EA): Remove an is_extension_ when all extension types satisfy
- # the interface
- value_is_extension_type = is_extension_type(value) or is_extension_array_dtype(
- value
- )
+ value_is_extension_type = is_extension_array_dtype(value)
# categorical/sparse/datetimetz
if value_is_extension_type:
diff --git a/pandas/core/reshape/melt.py b/pandas/core/reshape/melt.py
index 98fee491e0a73..9ccd36871050f 100644
--- a/pandas/core/reshape/melt.py
+++ b/pandas/core/reshape/melt.py
@@ -4,7 +4,7 @@
from pandas.util._decorators import Appender
-from pandas.core.dtypes.common import is_extension_type, is_list_like
+from pandas.core.dtypes.common import is_extension_array_dtype, is_list_like
from pandas.core.dtypes.concat import concat_compat
from pandas.core.dtypes.generic import ABCMultiIndex
from pandas.core.dtypes.missing import notna
@@ -103,7 +103,7 @@ def melt(
mdata = {}
for col in id_vars:
id_data = frame.pop(col)
- if is_extension_type(id_data):
+ if is_extension_array_dtype(id_data):
id_data = concat([id_data] * K, ignore_index=True)
else:
id_data = np.tile(id_data.values, K)
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 73a05b4cdfa66..ffaecfde6e10f 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -28,7 +28,6 @@
is_datetimelike,
is_dict_like,
is_extension_array_dtype,
- is_extension_type,
is_integer,
is_iterator,
is_list_like,
@@ -3958,7 +3957,8 @@ def f(x):
return f(self)
# row-wise access
- if is_extension_type(self.dtype):
+ if is_extension_array_dtype(self.dtype) and hasattr(self._values, "map"):
+ # GH#23179 some EAs do not have `map`
mapped = self._values.map(f)
else:
values = self.astype(object).values
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index 35e6d53127e59..77b2db20ac2a9 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -26,7 +26,7 @@
is_categorical_dtype,
is_datetime64_dtype,
is_datetime64tz_dtype,
- is_extension_type,
+ is_extension_array_dtype,
is_list_like,
is_timedelta64_dtype,
)
@@ -2827,7 +2827,7 @@ def write_multi_index(self, key, index):
zip(index.levels, index.codes, index.names)
):
# write the level
- if is_extension_type(lev):
+ if is_extension_array_dtype(lev):
raise NotImplementedError(
"Saving a MultiIndex with an extension dtype is not supported."
)
diff --git a/pandas/tests/api/test_types.py b/pandas/tests/api/test_types.py
index 24f325643479c..e9f68692a9863 100644
--- a/pandas/tests/api/test_types.py
+++ b/pandas/tests/api/test_types.py
@@ -18,7 +18,6 @@ class TestTypes(Base):
"is_datetime64_ns_dtype",
"is_datetime64tz_dtype",
"is_dtype_equal",
- "is_extension_type",
"is_float",
"is_float_dtype",
"is_int64_dtype",
@@ -51,7 +50,7 @@ class TestTypes(Base):
"infer_dtype",
"is_extension_array_dtype",
]
- deprecated = ["is_period", "is_datetimetz"]
+ deprecated = ["is_period", "is_datetimetz", "is_extension_type"]
dtypes = ["CategoricalDtype", "DatetimeTZDtype", "PeriodDtype", "IntervalDtype"]
def test_types(self):
| It is mostly redundant with `is_extension_array_dtype`, and having both is confusing.
xref #23179. | https://api.github.com/repos/pandas-dev/pandas/pulls/29457 | 2019-11-07T05:24:40Z | 2019-11-08T14:37:45Z | 2019-11-08T14:37:45Z | 2023-12-11T23:50:52Z |
CLN: type annotations in groupby.grouper, groupby.ops | diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index 370abe75e1327..e6e3ee62459ca 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -119,7 +119,7 @@ def __init__(self, key=None, level=None, freq=None, axis=0, sort=False):
def ax(self):
return self.grouper
- def _get_grouper(self, obj, validate=True):
+ def _get_grouper(self, obj, validate: bool = True):
"""
Parameters
----------
@@ -143,17 +143,18 @@ def _get_grouper(self, obj, validate=True):
)
return self.binner, self.grouper, self.obj
- def _set_grouper(self, obj, sort=False):
+ def _set_grouper(self, obj: FrameOrSeries, sort: bool = False):
"""
given an object and the specifications, setup the internal grouper
for this particular specification
Parameters
----------
- obj : the subject object
+ obj : Series or DataFrame
sort : bool, default False
whether the resulting grouper should be sorted
"""
+ assert obj is not None
if self.key is not None and self.level is not None:
raise ValueError("The Grouper cannot specify both a key and a level!")
@@ -211,13 +212,13 @@ def groups(self):
def __repr__(self) -> str:
attrs_list = (
- "{}={!r}".format(attr_name, getattr(self, attr_name))
+ "{name}={val!r}".format(name=attr_name, val=getattr(self, attr_name))
for attr_name in self._attributes
if getattr(self, attr_name) is not None
)
attrs = ", ".join(attrs_list)
cls_name = self.__class__.__name__
- return "{}({})".format(cls_name, attrs)
+ return "{cls}({attrs})".format(cls=cls_name, attrs=attrs)
class Grouping:
@@ -372,7 +373,7 @@ def __init__(
self.grouper = self.grouper.astype("timedelta64[ns]")
def __repr__(self) -> str:
- return "Grouping({0})".format(self.name)
+ return "Grouping({name})".format(name=self.name)
def __iter__(self):
return iter(self.indices)
@@ -433,10 +434,10 @@ def get_grouper(
key=None,
axis: int = 0,
level=None,
- sort=True,
- observed=False,
- mutated=False,
- validate=True,
+ sort: bool = True,
+ observed: bool = False,
+ mutated: bool = False,
+ validate: bool = True,
) -> Tuple[BaseGrouper, List[Hashable], FrameOrSeries]:
"""
Create and return a BaseGrouper, which is an internal
@@ -670,7 +671,7 @@ def is_in_obj(gpr) -> bool:
return grouper, exclusions, obj
-def _is_label_like(val):
+def _is_label_like(val) -> bool:
return isinstance(val, (str, tuple)) or (val is not None and is_scalar(val))
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 6796239cf3fd9..e6cf46de5c350 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -36,6 +36,7 @@
)
from pandas.core.dtypes.missing import _maybe_fill, isna
+from pandas._typing import FrameOrSeries
import pandas.core.algorithms as algorithms
from pandas.core.base import SelectionMixin
import pandas.core.common as com
@@ -89,12 +90,16 @@ def __init__(
self._filter_empty_groups = self.compressed = len(groupings) != 1
self.axis = axis
- self.groupings = groupings # type: Sequence[grouper.Grouping]
+ self._groupings = list(groupings) # type: List[grouper.Grouping]
self.sort = sort
self.group_keys = group_keys
self.mutated = mutated
self.indexer = indexer
+ @property
+ def groupings(self) -> List["grouper.Grouping"]:
+ return self._groupings
+
@property
def shape(self):
return tuple(ping.ngroups for ping in self.groupings)
@@ -106,7 +111,7 @@ def __iter__(self):
def nkeys(self) -> int:
return len(self.groupings)
- def get_iterator(self, data, axis=0):
+ def get_iterator(self, data: FrameOrSeries, axis: int = 0):
"""
Groupby iterator
@@ -120,7 +125,7 @@ def get_iterator(self, data, axis=0):
for key, (i, group) in zip(keys, splitter):
yield key, group
- def _get_splitter(self, data, axis=0):
+ def _get_splitter(self, data: FrameOrSeries, axis: int = 0) -> "DataSplitter":
comp_ids, _, ngroups = self.group_info
return get_splitter(data, comp_ids, ngroups, axis=axis)
@@ -142,13 +147,13 @@ def _get_group_keys(self):
# provide "flattened" iterator for multi-group setting
return get_flattened_iterator(comp_ids, ngroups, self.levels, self.codes)
- def apply(self, f, data, axis: int = 0):
+ def apply(self, f, data: FrameOrSeries, axis: int = 0):
mutated = self.mutated
splitter = self._get_splitter(data, axis=axis)
group_keys = self._get_group_keys()
result_values = None
- sdata = splitter._get_sorted_data()
+ sdata = splitter._get_sorted_data() # type: FrameOrSeries
if sdata.ndim == 2 and np.any(sdata.dtypes.apply(is_extension_array_dtype)):
# calling splitter.fast_apply will raise TypeError via apply_frame_axis0
# if we pass EA instead of ndarray
@@ -157,7 +162,7 @@ def apply(self, f, data, axis: int = 0):
elif (
com.get_callable_name(f) not in base.plotting_methods
- and hasattr(splitter, "fast_apply")
+ and isinstance(splitter, FrameSplitter)
and axis == 0
# with MultiIndex, apply_frame_axis0 would raise InvalidApply
# TODO: can we make this check prettier?
@@ -229,8 +234,7 @@ def names(self):
def size(self) -> Series:
"""
- Compute group sizes
-
+ Compute group sizes.
"""
ids, _, ngroup = self.group_info
ids = ensure_platform_int(ids)
@@ -292,7 +296,7 @@ def reconstructed_codes(self) -> List[np.ndarray]:
return decons_obs_group_ids(comp_ids, obs_ids, self.shape, codes, xnull=True)
@cache_readonly
- def result_index(self):
+ def result_index(self) -> Index:
if not self.compressed and len(self.groupings) == 1:
return self.groupings[0].result_index.rename(self.names[0])
@@ -628,7 +632,7 @@ def agg_series(self, obj: Series, func):
raise
return self._aggregate_series_pure_python(obj, func)
- def _aggregate_series_fast(self, obj, func):
+ def _aggregate_series_fast(self, obj: Series, func):
# At this point we have already checked that
# - obj.index is not a MultiIndex
# - obj is backed by an ndarray, not ExtensionArray
@@ -646,7 +650,7 @@ def _aggregate_series_fast(self, obj, func):
result, counts = grouper.get_result()
return result, counts
- def _aggregate_series_pure_python(self, obj, func):
+ def _aggregate_series_pure_python(self, obj: Series, func):
group_index, _, ngroups = self.group_info
@@ -703,7 +707,12 @@ class BinGrouper(BaseGrouper):
"""
def __init__(
- self, bins, binlabels, filter_empty=False, mutated=False, indexer=None
+ self,
+ bins,
+ binlabels,
+ filter_empty: bool = False,
+ mutated: bool = False,
+ indexer=None,
):
self.bins = ensure_int64(bins)
self.binlabels = ensure_index(binlabels)
@@ -737,7 +746,7 @@ def _get_grouper(self):
"""
return self
- def get_iterator(self, data: NDFrame, axis: int = 0):
+ def get_iterator(self, data: FrameOrSeries, axis: int = 0):
"""
Groupby iterator
@@ -809,11 +818,9 @@ def names(self):
return [self.binlabels.name]
@property
- def groupings(self):
- from pandas.core.groupby.grouper import Grouping
-
+ def groupings(self) -> "List[grouper.Grouping]":
return [
- Grouping(lvl, lvl, in_axis=False, level=None, name=name)
+ grouper.Grouping(lvl, lvl, in_axis=False, level=None, name=name)
for lvl, name in zip(self.levels, self.names)
]
@@ -854,7 +861,7 @@ def _is_indexed_like(obj, axes) -> bool:
class DataSplitter:
- def __init__(self, data, labels, ngroups, axis: int = 0):
+ def __init__(self, data: FrameOrSeries, labels, ngroups: int, axis: int = 0):
self.data = data
self.labels = ensure_int64(labels)
self.ngroups = ngroups
@@ -885,15 +892,15 @@ def __iter__(self):
for i, (start, end) in enumerate(zip(starts, ends)):
yield i, self._chop(sdata, slice(start, end))
- def _get_sorted_data(self):
+ def _get_sorted_data(self) -> FrameOrSeries:
return self.data.take(self.sort_idx, axis=self.axis)
- def _chop(self, sdata, slice_obj: slice):
+ def _chop(self, sdata, slice_obj: slice) -> NDFrame:
raise AbstractMethodError(self)
class SeriesSplitter(DataSplitter):
- def _chop(self, sdata, slice_obj: slice):
+ def _chop(self, sdata: Series, slice_obj: slice) -> Series:
return sdata._get_values(slice_obj)
@@ -905,14 +912,14 @@ def fast_apply(self, f, names):
sdata = self._get_sorted_data()
return libreduction.apply_frame_axis0(sdata, f, names, starts, ends)
- def _chop(self, sdata, slice_obj: slice):
+ def _chop(self, sdata: DataFrame, slice_obj: slice) -> DataFrame:
if self.axis == 0:
return sdata.iloc[slice_obj]
else:
return sdata._slice(slice_obj, axis=1)
-def get_splitter(data: NDFrame, *args, **kwargs):
+def get_splitter(data: FrameOrSeries, *args, **kwargs) -> DataSplitter:
if isinstance(data, Series):
klass = SeriesSplitter # type: Type[DataSplitter]
else:
| @simonjayhawkins mypy is still giving a couple of complaints I could use your help sorting out:
```
pandas/core/groupby/ops.py:791: error: Signature of "groupings" incompatible with supertype "BaseGrouper"
pandas/core/groupby/ops.py:872: error: Argument 1 of "_chop" is incompatible with supertype "DataSplitter"; supertype defines the argument type as "NDFrame"
pandas/core/groupby/ops.py:884: error: Argument 1 of "_chop" is incompatible with supertype "DataSplitter"; supertype defines the argument type as "NDFrame"
```
For the groupings complaint, AFAICT the attribute has the same annotation, but in the subclass its a property instead of defined in `__init__`. For the other two, I annotated an argument with `NDFrame` in the base class and overrode with `Series` and `DataFrame` in the subclasses. What is the preferred idiom for this pattern? | https://api.github.com/repos/pandas-dev/pandas/pulls/29456 | 2019-11-07T05:18:19Z | 2019-11-13T00:39:19Z | 2019-11-13T00:39:19Z | 2019-11-13T12:12:10Z |
TST: add test for empty frame groupby dtypes consistency | diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 79c9fe2b60bd9..a535fcc511daa 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -1490,6 +1490,20 @@ def test_frame_dict_constructor_empty_series(self):
DataFrame({"foo": s1, "bar": s2, "baz": s3})
DataFrame.from_dict({"foo": s1, "baz": s3, "bar": s2})
+ @pytest.mark.parametrize("d", [4, "d"])
+ def test_empty_frame_groupby_dtypes_consistency(self, d):
+ # GH 20888
+ group_keys = ["a", "b", "c"]
+ df = DataFrame({"a": [1], "b": [2], "c": [3], "d": [d]})
+
+ g = df[df.a == 2].groupby(group_keys)
+ result = g.first().index
+ expected = MultiIndex(
+ levels=[[1], [2], [3]], codes=[[], [], []], names=["a", "b", "c"]
+ )
+
+ tm.assert_index_equal(result, expected)
+
def test_multiindex_na_repr(self):
# only an issue with long columns
df3 = DataFrame(
| - [x] closes #20888
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29455 | 2019-11-07T02:12:32Z | 2019-11-08T04:17:29Z | 2019-11-08T04:17:28Z | 2019-11-08T04:17:35Z |
TST: add test for df.where() with category dtype | diff --git a/pandas/tests/frame/test_dtypes.py b/pandas/tests/frame/test_dtypes.py
index 68844aeeb081e..c29f5e78b033f 100644
--- a/pandas/tests/frame/test_dtypes.py
+++ b/pandas/tests/frame/test_dtypes.py
@@ -815,6 +815,22 @@ def test_astype_extension_dtypes_duplicate_col(self, dtype):
expected = concat([a1.astype(dtype), a2.astype(dtype)], axis=1)
tm.assert_frame_equal(result, expected)
+ @pytest.mark.parametrize("kwargs", [dict(), dict(other=None)])
+ def test_df_where_with_category(self, kwargs):
+ # GH 16979
+ df = DataFrame(np.arange(2 * 3).reshape(2, 3), columns=list("ABC"))
+ mask = np.array([[True, False, True], [False, True, True]])
+
+ # change type to category
+ df.A = df.A.astype("category")
+ df.B = df.B.astype("category")
+ df.C = df.C.astype("category")
+
+ result = df.A.where(mask[:, 0], **kwargs)
+ expected = Series(pd.Categorical([0, np.nan], categories=[0, 3]), name="A")
+
+ tm.assert_series_equal(result, expected)
+
@pytest.mark.parametrize(
"dtype", [{100: "float64", 200: "uint64"}, "category", "float64"]
)
| - [x] xref #16979
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29454 | 2019-11-07T01:45:46Z | 2019-11-08T19:57:37Z | 2019-11-08T19:57:37Z | 2019-11-08T19:57:37Z |
CLN: remove is_datetimelike | diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 23ba0ac1c737e..26099a94834e8 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -29,7 +29,6 @@
is_complex_dtype,
is_datetime64_any_dtype,
is_datetime64_ns_dtype,
- is_datetimelike,
is_extension_array_dtype,
is_float_dtype,
is_integer,
@@ -834,7 +833,7 @@ def mode(values, dropna: bool = True) -> ABCSeries:
return Series(values.values.mode(dropna=dropna), name=values.name)
return values.mode(dropna=dropna)
- if dropna and is_datetimelike(values):
+ if dropna and needs_i8_conversion(values.dtype):
mask = values.isnull()
values = values[~mask]
diff --git a/pandas/core/arrays/categorical.py b/pandas/core/arrays/categorical.py
index ce174baa66a97..73716fdeb42bb 100644
--- a/pandas/core/arrays/categorical.py
+++ b/pandas/core/arrays/categorical.py
@@ -25,7 +25,6 @@
ensure_platform_int,
is_categorical_dtype,
is_datetime64_dtype,
- is_datetimelike,
is_dict_like,
is_dtype_equal,
is_extension_array_dtype,
@@ -37,6 +36,7 @@
is_scalar,
is_sequence,
is_timedelta64_dtype,
+ needs_i8_conversion,
)
from pandas.core.dtypes.dtypes import CategoricalDtype
from pandas.core.dtypes.generic import ABCDataFrame, ABCIndexClass, ABCSeries
@@ -1533,7 +1533,7 @@ def get_values(self):
def _internal_get_values(self):
# if we are a datetime and period index, return Index to keep metadata
- if is_datetimelike(self.categories):
+ if needs_i8_conversion(self.categories):
return self.categories.take(self._codes, fill_value=np.nan)
elif is_integer_dtype(self.categories) and -1 in self._codes:
return self.categories.astype("object").take(self._codes, fill_value=np.nan)
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 61dc5f35cadf7..eeb0b72e301dd 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -21,7 +21,6 @@
is_categorical_dtype,
is_datetime64_ns_dtype,
is_datetime64tz_dtype,
- is_datetimelike,
is_extension_array_dtype,
is_extension_type,
is_list_like,
@@ -1172,7 +1171,7 @@ def tolist(self):
--------
numpy.ndarray.tolist
"""
- if is_datetimelike(self._values):
+ if self.dtype.kind in ["m", "M"]:
return [com.maybe_box_datetimelike(x) for x in self._values]
elif is_extension_array_dtype(self._values):
return list(self._values)
@@ -1194,7 +1193,7 @@ def __iter__(self):
iterator
"""
# We are explicitly making element iterators.
- if is_datetimelike(self._values):
+ if self.dtype.kind in ["m", "M"]:
return map(com.maybe_box_datetimelike, self._values)
elif is_extension_array_dtype(self._values):
return iter(self._values)
diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index fad80d6bf5745..bbed3a545e478 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -27,7 +27,6 @@
is_datetime64_ns_dtype,
is_datetime64tz_dtype,
is_datetime_or_timedelta_dtype,
- is_datetimelike,
is_dtype_equal,
is_extension_array_dtype,
is_extension_type,
@@ -274,7 +273,7 @@ def maybe_upcast_putmask(result: np.ndarray, mask: np.ndarray, other):
# in np.place:
# NaN -> NaT
# integer or integer array -> date-like array
- if is_datetimelike(result.dtype):
+ if result.dtype.kind in ["m", "M"]:
if is_scalar(other):
if isna(other):
other = result.dtype.type("nat")
diff --git a/pandas/core/dtypes/common.py b/pandas/core/dtypes/common.py
index 2a46d335ff512..c3e98d4009135 100644
--- a/pandas/core/dtypes/common.py
+++ b/pandas/core/dtypes/common.py
@@ -799,54 +799,6 @@ def is_datetime_arraylike(arr):
return getattr(arr, "inferred_type", None) == "datetime"
-def is_datetimelike(arr):
- """
- Check whether an array-like is a datetime-like array-like.
-
- Acceptable datetime-like objects are (but not limited to) datetime
- indices, periodic indices, and timedelta indices.
-
- Parameters
- ----------
- arr : array-like
- The array-like to check.
-
- Returns
- -------
- boolean
- Whether or not the array-like is a datetime-like array-like.
-
- Examples
- --------
- >>> is_datetimelike([1, 2, 3])
- False
- >>> is_datetimelike(pd.Index([1, 2, 3]))
- False
- >>> is_datetimelike(pd.DatetimeIndex([1, 2, 3]))
- True
- >>> is_datetimelike(pd.DatetimeIndex([1, 2, 3], tz="US/Eastern"))
- True
- >>> is_datetimelike(pd.PeriodIndex([], freq="A"))
- True
- >>> is_datetimelike(np.array([], dtype=np.datetime64))
- True
- >>> is_datetimelike(pd.Series([], dtype="timedelta64[ns]"))
- True
- >>>
- >>> dtype = DatetimeTZDtype("ns", tz="US/Eastern")
- >>> s = pd.Series([], dtype=dtype)
- >>> is_datetimelike(s)
- True
- """
-
- return (
- is_datetime64_dtype(arr)
- or is_datetime64tz_dtype(arr)
- or is_timedelta64_dtype(arr)
- or isinstance(arr, ABCPeriodIndex)
- )
-
-
def is_dtype_equal(source, target):
"""
Check if two dtypes are equal.
@@ -1446,9 +1398,8 @@ def is_numeric(x):
"""
return is_integer_dtype(x) or is_float_dtype(x)
- is_datetimelike = needs_i8_conversion
- return (is_datetimelike(a) and is_numeric(b)) or (
- is_datetimelike(b) and is_numeric(a)
+ return (needs_i8_conversion(a) and is_numeric(b)) or (
+ needs_i8_conversion(b) and is_numeric(a)
)
diff --git a/pandas/core/dtypes/missing.py b/pandas/core/dtypes/missing.py
index 322011eb8e263..22e38a805f996 100644
--- a/pandas/core/dtypes/missing.py
+++ b/pandas/core/dtypes/missing.py
@@ -17,7 +17,6 @@
is_complex_dtype,
is_datetime64_dtype,
is_datetime64tz_dtype,
- is_datetimelike,
is_datetimelike_v_numeric,
is_dtype_equal,
is_extension_array_dtype,
@@ -494,7 +493,7 @@ def _infer_fill_value(val):
if not is_list_like(val):
val = [val]
val = np.array(val, copy=False)
- if is_datetimelike(val):
+ if needs_i8_conversion(val):
return np.array("NaT", dtype=val.dtype)
elif is_object_dtype(val.dtype):
dtype = lib.infer_dtype(ensure_object(val), skipna=False)
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 511b87dab087e..3b8c3148f5177 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -40,7 +40,6 @@
ensure_int64,
ensure_platform_int,
is_bool,
- is_datetimelike,
is_dict_like,
is_integer_dtype,
is_interval_dtype,
@@ -48,6 +47,7 @@
is_numeric_dtype,
is_object_dtype,
is_scalar,
+ needs_i8_conversion,
)
from pandas.core.dtypes.missing import _isna_ndarraylike, isna, notna
@@ -1287,7 +1287,7 @@ def first_not_none(values):
# if we have date/time like in the original, then coerce dates
# as we are stacking can easily have object dtypes here
so = self._selected_obj
- if so.ndim == 2 and so.dtypes.apply(is_datetimelike).any():
+ if so.ndim == 2 and so.dtypes.apply(needs_i8_conversion).any():
result = _recast_datetimelike_result(result)
else:
result = result._convert(datetime=True)
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index a189b2cd1ab84..30857a51debd1 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -24,7 +24,6 @@
is_bool_dtype,
is_categorical_dtype,
is_datetime64tz_dtype,
- is_datetimelike,
is_dtype_equal,
is_extension_array_dtype,
is_float_dtype,
@@ -1120,9 +1119,9 @@ def _maybe_coerce_merge_keys(self):
raise ValueError(msg)
# datetimelikes must match exactly
- elif is_datetimelike(lk) and not is_datetimelike(rk):
+ elif needs_i8_conversion(lk) and not needs_i8_conversion(rk):
raise ValueError(msg)
- elif not is_datetimelike(lk) and is_datetimelike(rk):
+ elif not needs_i8_conversion(lk) and needs_i8_conversion(rk):
raise ValueError(msg)
elif is_datetime64tz_dtype(lk) and not is_datetime64tz_dtype(rk):
raise ValueError(msg)
@@ -1637,7 +1636,7 @@ def _get_merge_keys(self):
)
)
- if is_datetimelike(lt):
+ if needs_i8_conversion(lt):
if not isinstance(self.tolerance, datetime.timedelta):
raise MergeError(msg)
if self.tolerance < Timedelta(0):
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 73a05b4cdfa66..6440d2f03cf1a 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -25,7 +25,6 @@
is_categorical,
is_categorical_dtype,
is_datetime64_dtype,
- is_datetimelike,
is_dict_like,
is_extension_array_dtype,
is_extension_type,
@@ -2886,7 +2885,7 @@ def combine_first(self, other):
new_index = self.index.union(other.index)
this = self.reindex(new_index, copy=False)
other = other.reindex(new_index, copy=False)
- if is_datetimelike(this) and not is_datetimelike(other):
+ if this.dtype.kind == "M" and other.dtype.kind != "M":
other = to_datetime(other)
return this.where(notna(this), other)
diff --git a/pandas/tests/dtypes/test_common.py b/pandas/tests/dtypes/test_common.py
index 894d6a40280b7..5e409b85049ae 100644
--- a/pandas/tests/dtypes/test_common.py
+++ b/pandas/tests/dtypes/test_common.py
@@ -309,21 +309,6 @@ def test_is_datetime_arraylike():
assert com.is_datetime_arraylike(pd.DatetimeIndex([1, 2, 3]))
-def test_is_datetimelike():
- assert not com.is_datetimelike([1, 2, 3])
- assert not com.is_datetimelike(pd.Index([1, 2, 3]))
-
- assert com.is_datetimelike(pd.DatetimeIndex([1, 2, 3]))
- assert com.is_datetimelike(pd.PeriodIndex([], freq="A"))
- assert com.is_datetimelike(np.array([], dtype=np.datetime64))
- assert com.is_datetimelike(pd.Series([], dtype="timedelta64[ns]"))
- assert com.is_datetimelike(pd.DatetimeIndex(["2000"], tz="US/Eastern"))
-
- dtype = DatetimeTZDtype("ns", tz="US/Eastern")
- s = pd.Series([], dtype=dtype)
- assert com.is_datetimelike(s)
-
-
integer_dtypes = [] # type: List
| - [x] closes #23914
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
AFAICT this is strictly dominated by needs_i8_conversion | https://api.github.com/repos/pandas-dev/pandas/pulls/29452 | 2019-11-07T00:28:50Z | 2019-11-07T21:23:32Z | 2019-11-07T21:23:31Z | 2019-11-07T21:56:05Z |
CLN: remove is_stringlike | diff --git a/pandas/core/dtypes/common.py b/pandas/core/dtypes/common.py
index 3f4ebc88c1c8a..2a46d335ff512 100644
--- a/pandas/core/dtypes/common.py
+++ b/pandas/core/dtypes/common.py
@@ -45,7 +45,6 @@
is_re_compilable,
is_scalar,
is_sequence,
- is_string_like,
)
from pandas._typing import ArrayLike
@@ -1383,8 +1382,8 @@ def is_numeric_v_string_like(a, b):
is_a_string_array = is_a_array and is_string_like_dtype(a)
is_b_string_array = is_b_array and is_string_like_dtype(b)
- is_a_scalar_string_like = not is_a_array and is_string_like(a)
- is_b_scalar_string_like = not is_b_array and is_string_like(b)
+ is_a_scalar_string_like = not is_a_array and isinstance(a, str)
+ is_b_scalar_string_like = not is_b_array and isinstance(b, str)
return (
(is_a_numeric_array and is_b_scalar_string_like)
diff --git a/pandas/core/dtypes/inference.py b/pandas/core/dtypes/inference.py
index e69e703f3a96c..61fa7940c1bce 100644
--- a/pandas/core/dtypes/inference.py
+++ b/pandas/core/dtypes/inference.py
@@ -67,30 +67,6 @@ def is_number(obj):
return isinstance(obj, (Number, np.number))
-def is_string_like(obj):
- """
- Check if the object is a string.
-
- Parameters
- ----------
- obj : The object to check
-
- Examples
- --------
- >>> is_string_like("foo")
- True
- >>> is_string_like(1)
- False
-
- Returns
- -------
- is_str_like : bool
- Whether `obj` is a string or not.
- """
-
- return isinstance(obj, str)
-
-
def _iterable_not_string(obj):
"""
Check if the object is an iterable but not a string.
diff --git a/pandas/core/indexes/datetimes.py b/pandas/core/indexes/datetimes.py
index e5cecd090e061..2d0ecf1b936da 100644
--- a/pandas/core/indexes/datetimes.py
+++ b/pandas/core/indexes/datetimes.py
@@ -16,7 +16,6 @@
is_integer,
is_list_like,
is_scalar,
- is_string_like,
)
from pandas.core.dtypes.concat import concat_compat
from pandas.core.dtypes.dtypes import DatetimeTZDtype
@@ -1659,7 +1658,7 @@ def bdate_range(
msg = "freq must be specified for bdate_range; use date_range instead"
raise TypeError(msg)
- if is_string_like(freq) and freq.startswith("C"):
+ if isinstance(freq, str) and freq.startswith("C"):
try:
weekmask = weekmask or "Mon Tue Wed Thu Fri"
freq = prefix_mapping[freq](holidays=holidays, weekmask=weekmask)
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 7b65816dc06b9..73a05b4cdfa66 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -34,7 +34,6 @@
is_list_like,
is_object_dtype,
is_scalar,
- is_string_like,
is_timedelta64_dtype,
)
from pandas.core.dtypes.generic import (
@@ -4539,7 +4538,7 @@ def to_csv(self, *args, **kwargs):
# passed as second argument (while the first is the same)
maybe_sep = args[1]
- if not (is_string_like(maybe_sep) and len(maybe_sep) == 1):
+ if not (isinstance(maybe_sep, str) and len(maybe_sep) == 1):
# old signature
warnings.warn(
"The signature of `Series.to_csv` was aligned "
diff --git a/pandas/core/strings.py b/pandas/core/strings.py
index f1a67d0892cad..7194d1cf08e4a 100644
--- a/pandas/core/strings.py
+++ b/pandas/core/strings.py
@@ -19,7 +19,6 @@
is_list_like,
is_re,
is_scalar,
- is_string_like,
)
from pandas.core.dtypes.generic import (
ABCDataFrame,
@@ -601,7 +600,7 @@ def str_replace(arr, pat, repl, n=-1, case=None, flags=0, regex=True):
"""
# Check whether repl is valid (GH 13438, GH 15055)
- if not (is_string_like(repl) or callable(repl)):
+ if not (isinstance(repl, str) or callable(repl)):
raise TypeError("repl must be a string or callable")
is_compiled_re = is_re(pat)
diff --git a/pandas/io/formats/style.py b/pandas/io/formats/style.py
index 9865087a26ae3..dce0afd8670b2 100644
--- a/pandas/io/formats/style.py
+++ b/pandas/io/formats/style.py
@@ -18,7 +18,7 @@
from pandas.compat._optional import import_optional_dependency
from pandas.util._decorators import Appender
-from pandas.core.dtypes.common import is_float, is_string_like
+from pandas.core.dtypes.common import is_float
import pandas as pd
from pandas.api.types import is_dict_like, is_list_like
@@ -1488,7 +1488,7 @@ def _get_level_lengths(index, hidden_elements=None):
def _maybe_wrap_formatter(formatter):
- if is_string_like(formatter):
+ if isinstance(formatter, str):
return lambda x: formatter.format(x)
elif callable(formatter):
return formatter
| its now just an alias for `isinstance(obj, str)` | https://api.github.com/repos/pandas-dev/pandas/pulls/29450 | 2019-11-06T22:23:54Z | 2019-11-07T07:34:28Z | 2019-11-07T07:34:28Z | 2019-11-07T17:18:30Z |
DOC: Improving (hopefully) the documintation | diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index 2d6c8e1008ce1..c2d6a3bc4906d 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -84,8 +84,8 @@ cpdef ndarray[int64_t, ndim=1] unique_deltas(const int64_t[:] arr):
Returns
-------
- result : ndarray[int64_t]
- result is sorted
+ ndarray[int64_t]
+ An ordered ndarray[int64_t]
"""
cdef:
Py_ssize_t i, n = len(arr)
@@ -150,9 +150,10 @@ def is_lexsorted(list_of_arrays: list) -> bint:
@cython.wraparound(False)
def groupsort_indexer(const int64_t[:] index, Py_ssize_t ngroups):
"""
- compute a 1-d indexer that is an ordering of the passed index,
- ordered by the groups. This is a reverse of the label
- factorization process.
+ Compute a 1-d indexer.
+
+ The indexer is an ordering of the passed index,
+ ordered by the groups.
Parameters
----------
@@ -161,7 +162,14 @@ def groupsort_indexer(const int64_t[:] index, Py_ssize_t ngroups):
ngroups: int64
number of groups
- return a tuple of (1-d indexer ordered by groups, group counts)
+ Returns
+ -------
+ tuple
+ 1-d indexer ordered by groups, group counts
+
+ Notes
+ -----
+ This is a reverse of the label factorization process.
"""
cdef:
@@ -391,6 +399,7 @@ def _validate_limit(nobs: int, limit=None) -> int:
Returns
-------
int
+ The limit.
"""
if limit is None:
lim = nobs
@@ -669,7 +678,8 @@ def is_monotonic(ndarray[algos_t, ndim=1] arr, bint timelike):
"""
Returns
-------
- is_monotonic_inc, is_monotonic_dec, is_unique
+ tuple
+ is_monotonic_inc, is_monotonic_dec, is_unique
"""
cdef:
Py_ssize_t i, n
| - [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` | https://api.github.com/repos/pandas-dev/pandas/pulls/29449 | 2019-11-06T22:13:18Z | 2019-11-07T16:10:39Z | 2019-11-07T16:10:39Z | 2019-11-07T20:52:15Z |
TST: add test for indexing with single/double tuples | diff --git a/pandas/tests/frame/indexing/test_indexing.py b/pandas/tests/frame/indexing/test_indexing.py
index 24a431fe42cf8..9a7cd4ace686f 100644
--- a/pandas/tests/frame/indexing/test_indexing.py
+++ b/pandas/tests/frame/indexing/test_indexing.py
@@ -2624,6 +2624,17 @@ def test_index_namedtuple(self):
result = df.loc[IndexType("foo", "bar")]["A"]
assert result == 1
+ @pytest.mark.parametrize("tpl", [tuple([1]), tuple([1, 2])])
+ def test_index_single_double_tuples(self, tpl):
+ # GH 20991
+ idx = pd.Index([tuple([1]), tuple([1, 2])], name="A", tupleize_cols=False)
+ df = DataFrame(index=idx)
+
+ result = df.loc[[tpl]]
+ idx = pd.Index([tpl], name="A", tupleize_cols=False)
+ expected = DataFrame(index=idx)
+ tm.assert_frame_equal(result, expected)
+
def test_boolean_indexing(self):
idx = list(range(3))
cols = ["A", "B", "C"]
| - [x] closes #20991
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` | https://api.github.com/repos/pandas-dev/pandas/pulls/29448 | 2019-11-06T21:56:29Z | 2019-11-20T17:14:29Z | 2019-11-20T17:14:29Z | 2019-11-20T17:15:58Z |
ENH: Add ORC reader | diff --git a/doc/source/getting_started/install.rst b/doc/source/getting_started/install.rst
index 9f3ab22496ae7..14530a9010a1c 100644
--- a/doc/source/getting_started/install.rst
+++ b/doc/source/getting_started/install.rst
@@ -258,7 +258,7 @@ matplotlib 2.2.2 Visualization
openpyxl 2.4.8 Reading / writing for xlsx files
pandas-gbq 0.8.0 Google Big Query access
psycopg2 PostgreSQL engine for sqlalchemy
-pyarrow 0.12.0 Parquet and feather reading / writing
+pyarrow 0.12.0 Parquet, ORC (requires 0.13.0), and feather reading / writing
pymysql 0.7.11 MySQL engine for sqlalchemy
pyreadstat SPSS files (.sav) reading
pytables 3.4.2 HDF5 reading / writing
diff --git a/doc/source/reference/io.rst b/doc/source/reference/io.rst
index 91f4942d03b0d..6d2d405a15850 100644
--- a/doc/source/reference/io.rst
+++ b/doc/source/reference/io.rst
@@ -98,6 +98,13 @@ Parquet
read_parquet
+ORC
+~~~
+.. autosummary::
+ :toctree: api/
+
+ read_orc
+
SAS
~~~
.. autosummary::
diff --git a/doc/source/user_guide/io.rst b/doc/source/user_guide/io.rst
index fa47a5944f7bf..972f36aecad24 100644
--- a/doc/source/user_guide/io.rst
+++ b/doc/source/user_guide/io.rst
@@ -28,6 +28,7 @@ The pandas I/O API is a set of top level ``reader`` functions accessed like
binary;`HDF5 Format <https://support.hdfgroup.org/HDF5/whatishdf5.html>`__;:ref:`read_hdf<io.hdf5>`;:ref:`to_hdf<io.hdf5>`
binary;`Feather Format <https://github.com/wesm/feather>`__;:ref:`read_feather<io.feather>`;:ref:`to_feather<io.feather>`
binary;`Parquet Format <https://parquet.apache.org/>`__;:ref:`read_parquet<io.parquet>`;:ref:`to_parquet<io.parquet>`
+ binary;`ORC Format <//https://orc.apache.org/>`__;:ref:`read_orc<io.orc>`;
binary;`Msgpack <https://msgpack.org/index.html>`__;:ref:`read_msgpack<io.msgpack>`;:ref:`to_msgpack<io.msgpack>`
binary;`Stata <https://en.wikipedia.org/wiki/Stata>`__;:ref:`read_stata<io.stata_reader>`;:ref:`to_stata<io.stata_writer>`
binary;`SAS <https://en.wikipedia.org/wiki/SAS_(software)>`__;:ref:`read_sas<io.sas_reader>`;
@@ -4858,6 +4859,17 @@ The above example creates a partitioned dataset that may look like:
except OSError:
pass
+.. _io.orc:
+
+ORC
+---
+
+.. versionadded:: 1.0.0
+
+Similar to the :ref:`parquet <io.parquet>` format, the `ORC Format <//https://orc.apache.org/>`__ is a binary columnar serialization
+for data frames. It is designed to make reading data frames efficient. Pandas provides *only* a reader for the
+ORC format, :func:`~pandas.read_orc`. This requires the `pyarrow <https://arrow.apache.org/docs/python/>`__ library.
+
.. _io.sql:
SQL queries
@@ -5761,6 +5773,3 @@ Space on disk (in bytes)
24009288 Oct 10 06:43 test_fixed_compress.hdf
24458940 Oct 10 06:44 test_table.hdf
24458940 Oct 10 06:44 test_table_compress.hdf
-
-
-
diff --git a/pandas/__init__.py b/pandas/__init__.py
index a60aa08b89f84..f72a12b58edcb 100644
--- a/pandas/__init__.py
+++ b/pandas/__init__.py
@@ -168,6 +168,7 @@
# misc
read_clipboard,
read_parquet,
+ read_orc,
read_feather,
read_gbq,
read_html,
diff --git a/pandas/io/api.py b/pandas/io/api.py
index 725e82604ca7f..e20aa18324a34 100644
--- a/pandas/io/api.py
+++ b/pandas/io/api.py
@@ -10,6 +10,7 @@
from pandas.io.gbq import read_gbq
from pandas.io.html import read_html
from pandas.io.json import read_json
+from pandas.io.orc import read_orc
from pandas.io.packers import read_msgpack, to_msgpack
from pandas.io.parquet import read_parquet
from pandas.io.parsers import read_csv, read_fwf, read_table
diff --git a/pandas/io/orc.py b/pandas/io/orc.py
new file mode 100644
index 0000000000000..bbefe447cb7fe
--- /dev/null
+++ b/pandas/io/orc.py
@@ -0,0 +1,57 @@
+""" orc compat """
+
+import distutils
+from typing import TYPE_CHECKING, List, Optional
+
+from pandas._typing import FilePathOrBuffer
+
+from pandas.io.common import get_filepath_or_buffer
+
+if TYPE_CHECKING:
+ from pandas import DataFrame
+
+
+def read_orc(
+ path: FilePathOrBuffer, columns: Optional[List[str]] = None, **kwargs,
+) -> "DataFrame":
+ """
+ Load an ORC object from the file path, returning a DataFrame.
+
+ .. versionadded:: 1.0.0
+
+ Parameters
+ ----------
+ path : str, path object or file-like object
+ Any valid string path is acceptable. The string could be a URL. Valid
+ URL schemes include http, ftp, s3, and file. For file URLs, a host is
+ expected. A local file could be:
+ ``file://localhost/path/to/table.orc``.
+
+ If you want to pass in a path object, pandas accepts any
+ ``os.PathLike``.
+
+ By file-like object, we refer to objects with a ``read()`` method,
+ such as a file handler (e.g. via builtin ``open`` function)
+ or ``StringIO``.
+ columns : list, default None
+ If not None, only these columns will be read from the file.
+ **kwargs
+ Any additional kwargs are passed to pyarrow.
+
+ Returns
+ -------
+ DataFrame
+ """
+
+ # we require a newer version of pyarrow than we support for parquet
+ import pyarrow
+
+ if distutils.version.LooseVersion(pyarrow.__version__) < "0.13.0":
+ raise ImportError("pyarrow must be >= 0.13.0 for read_orc")
+
+ import pyarrow.orc
+
+ path, _, _, _ = get_filepath_or_buffer(path)
+ orc_file = pyarrow.orc.ORCFile(path)
+ result = orc_file.read(columns=columns, **kwargs).to_pandas()
+ return result
diff --git a/pandas/tests/api/test_api.py b/pandas/tests/api/test_api.py
index 76141dceae930..870d7fd6e44c1 100644
--- a/pandas/tests/api/test_api.py
+++ b/pandas/tests/api/test_api.py
@@ -167,6 +167,7 @@ class TestPDApi(Base):
"read_table",
"read_feather",
"read_parquet",
+ "read_orc",
"read_spss",
]
diff --git a/pandas/tests/io/data/orc/TestOrcFile.decimal.orc b/pandas/tests/io/data/orc/TestOrcFile.decimal.orc
new file mode 100644
index 0000000000000..cb0f7b9d767a3
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.decimal.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.emptyFile.orc b/pandas/tests/io/data/orc/TestOrcFile.emptyFile.orc
new file mode 100644
index 0000000000000..ecdadcbff1346
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.emptyFile.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.test1.orc b/pandas/tests/io/data/orc/TestOrcFile.test1.orc
new file mode 100644
index 0000000000000..4fb0beff86897
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.test1.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.testDate1900.orc b/pandas/tests/io/data/orc/TestOrcFile.testDate1900.orc
new file mode 100644
index 0000000000000..f51ffdbd03a43
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.testDate1900.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.testDate2038.orc b/pandas/tests/io/data/orc/TestOrcFile.testDate2038.orc
new file mode 100644
index 0000000000000..cd11fa8a4e91d
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.testDate2038.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.testSnappy.orc b/pandas/tests/io/data/orc/TestOrcFile.testSnappy.orc
new file mode 100644
index 0000000000000..aa6cc9c9ba1a7
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.testSnappy.orc differ
diff --git a/pandas/tests/io/test_orc.py b/pandas/tests/io/test_orc.py
new file mode 100644
index 0000000000000..9f3ec274007d0
--- /dev/null
+++ b/pandas/tests/io/test_orc.py
@@ -0,0 +1,227 @@
+""" test orc compat """
+import datetime
+import os
+
+import numpy as np
+import pytest
+
+import pandas as pd
+from pandas import read_orc
+import pandas.util.testing as tm
+
+pytest.importorskip("pyarrow", minversion="0.13.0")
+pytest.importorskip("pyarrow.orc")
+
+pytestmark = pytest.mark.filterwarnings(
+ "ignore:RangeIndex.* is deprecated:DeprecationWarning"
+)
+
+
+@pytest.fixture
+def dirpath(datapath):
+ return datapath("io", "data", "orc")
+
+
+def test_orc_reader_empty(dirpath):
+ columns = [
+ "boolean1",
+ "byte1",
+ "short1",
+ "int1",
+ "long1",
+ "float1",
+ "double1",
+ "bytes1",
+ "string1",
+ ]
+ dtypes = [
+ "bool",
+ "int8",
+ "int16",
+ "int32",
+ "int64",
+ "float32",
+ "float64",
+ "object",
+ "object",
+ ]
+ expected = pd.DataFrame(index=pd.RangeIndex(0))
+ for colname, dtype in zip(columns, dtypes):
+ expected[colname] = pd.Series(dtype=dtype)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.emptyFile.orc")
+ got = read_orc(inputfile, columns=columns)
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_basic(dirpath):
+ data = {
+ "boolean1": np.array([False, True], dtype="bool"),
+ "byte1": np.array([1, 100], dtype="int8"),
+ "short1": np.array([1024, 2048], dtype="int16"),
+ "int1": np.array([65536, 65536], dtype="int32"),
+ "long1": np.array([9223372036854775807, 9223372036854775807], dtype="int64"),
+ "float1": np.array([1.0, 2.0], dtype="float32"),
+ "double1": np.array([-15.0, -5.0], dtype="float64"),
+ "bytes1": np.array([b"\x00\x01\x02\x03\x04", b""], dtype="object"),
+ "string1": np.array(["hi", "bye"], dtype="object"),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.test1.orc")
+ got = read_orc(inputfile, columns=data.keys())
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_decimal(dirpath):
+ from decimal import Decimal
+
+ # Only testing the first 10 rows of data
+ data = {
+ "_col0": np.array(
+ [
+ Decimal("-1000.50000"),
+ Decimal("-999.60000"),
+ Decimal("-998.70000"),
+ Decimal("-997.80000"),
+ Decimal("-996.90000"),
+ Decimal("-995.10000"),
+ Decimal("-994.11000"),
+ Decimal("-993.12000"),
+ Decimal("-992.13000"),
+ Decimal("-991.14000"),
+ ],
+ dtype="object",
+ )
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.decimal.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_date_low(dirpath):
+ data = {
+ "time": np.array(
+ [
+ "1900-05-05 12:34:56.100000",
+ "1900-05-05 12:34:56.100100",
+ "1900-05-05 12:34:56.100200",
+ "1900-05-05 12:34:56.100300",
+ "1900-05-05 12:34:56.100400",
+ "1900-05-05 12:34:56.100500",
+ "1900-05-05 12:34:56.100600",
+ "1900-05-05 12:34:56.100700",
+ "1900-05-05 12:34:56.100800",
+ "1900-05-05 12:34:56.100900",
+ ],
+ dtype="datetime64[ns]",
+ ),
+ "date": np.array(
+ [
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ ],
+ dtype="object",
+ ),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.testDate1900.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_date_high(dirpath):
+ data = {
+ "time": np.array(
+ [
+ "2038-05-05 12:34:56.100000",
+ "2038-05-05 12:34:56.100100",
+ "2038-05-05 12:34:56.100200",
+ "2038-05-05 12:34:56.100300",
+ "2038-05-05 12:34:56.100400",
+ "2038-05-05 12:34:56.100500",
+ "2038-05-05 12:34:56.100600",
+ "2038-05-05 12:34:56.100700",
+ "2038-05-05 12:34:56.100800",
+ "2038-05-05 12:34:56.100900",
+ ],
+ dtype="datetime64[ns]",
+ ),
+ "date": np.array(
+ [
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ ],
+ dtype="object",
+ ),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.testDate2038.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_snappy_compressed(dirpath):
+ data = {
+ "int1": np.array(
+ [
+ -1160101563,
+ 1181413113,
+ 2065821249,
+ -267157795,
+ 172111193,
+ 1752363137,
+ 1406072123,
+ 1911809390,
+ -1308542224,
+ -467100286,
+ ],
+ dtype="int32",
+ ),
+ "string1": np.array(
+ [
+ "f50dcb8",
+ "382fdaaa",
+ "90758c6",
+ "9e8caf3f",
+ "ee97332b",
+ "d634da1",
+ "2bea4396",
+ "d67d89e8",
+ "ad71007e",
+ "e8c82066",
+ ],
+ dtype="object",
+ ),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.testSnappy.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
| - [x] closes #25229
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
Added an ORC reader following the `read_parquet` API. Still need to give some additional love to the docstrings but this is at least ready for some discussion and eyes on it. | https://api.github.com/repos/pandas-dev/pandas/pulls/29447 | 2019-11-06T21:54:14Z | 2019-12-11T08:11:52Z | 2019-12-11T08:11:52Z | 2019-12-23T20:57:03Z |
DOC: Remove errant backslashes from the Ecosystem tab on new website. | diff --git a/web/pandas/community/ecosystem.md b/web/pandas/community/ecosystem.md
index cf242e86f879f..af6fd1ac77605 100644
--- a/web/pandas/community/ecosystem.md
+++ b/web/pandas/community/ecosystem.md
@@ -86,12 +86,12 @@ models to emphasize patterns in a dataset.
### [yhat/ggpy](https://github.com/yhat/ggpy)
-Hadley Wickham\'s [ggplot2](https://ggplot2.tidyverse.org/) is a
+Hadley Wickham's [ggplot2](https://ggplot2.tidyverse.org/) is a
foundational exploratory visualization package for the R language. Based
-on [\"The Grammar of
-Graphics\"](https://www.cs.uic.edu/~wilkinson/TheGrammarOfGraphics/GOG.html)
+on ["The Grammar of
+Graphics"](https://www.cs.uic.edu/~wilkinson/TheGrammarOfGraphics/GOG.html)
it provides a powerful, declarative and extremely general way to
-generate bespoke plots of any kind of data. It\'s really quite
+generate bespoke plots of any kind of data. It's really quite
incredible. Various implementations to other languages are available,
but a faithful implementation for Python users has long been missing.
Although still young (as of Jan-2014), the
@@ -100,9 +100,7 @@ quickly in that direction.
### [IPython Vega](https://github.com/vega/ipyvega)
-[IPython Vega](https://github.com/vega/ipyvega) leverages [Vega
-\<https://github.com/trifacta/vega\>]\_\_ to create plots
-within Jupyter Notebook.
+[IPython Vega](https://github.com/vega/ipyvega) leverages [Vega](https://github.com/vega/vega) to create plots within Jupyter Notebook.
### [Plotly](https://plot.ly/python)
@@ -158,8 +156,8 @@ for pandas `display.` settings.
### [quantopian/qgrid](https://github.com/quantopian/qgrid)
-qgrid is \"an interactive grid for sorting and filtering DataFrames in
-IPython Notebook\" built with SlickGrid.
+qgrid is "an interactive grid for sorting and filtering DataFrames in
+IPython Notebook" built with SlickGrid.
### [Spyder](https://www.spyder-ide.org/)
@@ -172,8 +170,8 @@ environment like MATLAB or Rstudio.
Its [Variable
Explorer](https://docs.spyder-ide.org/variableexplorer.html) allows
users to view, manipulate and edit pandas `Index`, `Series`, and
-`DataFrame` objects like a \"spreadsheet\", including copying and
-modifying values, sorting, displaying a \"heatmap\", converting data
+`DataFrame` objects like a "spreadsheet", including copying and
+modifying values, sorting, displaying a "heatmap", converting data
types and more. Pandas objects can also be renamed, duplicated, new
columns added, copyed/pasted to/from the clipboard (as TSV), and
saved/loaded to/from a file. Spyder can also import data from a variety
@@ -181,8 +179,8 @@ of plain text and binary files or the clipboard into a new pandas
DataFrame via a sophisticated import wizard.
Most pandas classes, methods and data attributes can be autocompleted in
-Spyder\'s [Editor](https://docs.spyder-ide.org/editor.html) and [IPython
-Console](https://docs.spyder-ide.org/ipythonconsole.html), and Spyder\'s
+Spyder's [Editor](https://docs.spyder-ide.org/editor.html) and [IPython
+Console](https://docs.spyder-ide.org/ipythonconsole.html), and Spyder's
[Help pane](https://docs.spyder-ide.org/help.html) can retrieve and
render Numpydoc documentation on pandas objects in rich text with Sphinx
both automatically and on-demand.
@@ -355,7 +353,7 @@ which work well with pandas' data containers.
### [cyberpandas](https://cyberpandas.readthedocs.io/en/latest)
Cyberpandas provides an extension type for storing arrays of IP
-Addresses. These arrays can be stored inside pandas\' Series and
+Addresses. These arrays can be stored inside pandas' Series and
DataFrame.
## Accessors
@@ -364,7 +362,7 @@ A directory of projects providing
`extension accessors <extending.register-accessors>`. This is for users to discover new accessors and for library
authors to coordinate on the namespace.
- Library Accessor Classes
- ------------------------------------------------------------- ---------- -----------------------
- [cyberpandas](https://cyberpandas.readthedocs.io/en/latest) `ip` `Series`
- [pdvega](https://altair-viz.github.io/pdvega/) `vgplot` `Series`, `DataFrame`
+ | Library | Accessor | Classes |
+ | ------------------------------------------------------------|----------|-----------------------|
+ | [cyberpandas](https://cyberpandas.readthedocs.io/en/latest) | `ip` | `Series` |
+ | [pdvega](https://altair-viz.github.io/pdvega/) | `vgplot` | `Series`, `DataFrame` |
| Pandas Sprint at PyData NYC :-)
Fixed at the direction of @datapythonista
The only file touched is a markdown file.
- [ ] closes #xxxx
- [ ] tests added / passed
- [ ] passes `black pandas`
- [ ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29446 | 2019-11-06T21:11:17Z | 2019-11-07T16:31:50Z | 2019-11-07T16:31:50Z | 2019-11-07T16:31:52Z |
fixup pip env | diff --git a/requirements-dev.txt b/requirements-dev.txt
index e7df704925485..13e2c95126f0c 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -1,5 +1,4 @@
numpy>=1.15
-python==3.7
python-dateutil>=2.6.1
pytz
asv
diff --git a/scripts/generate_pip_deps_from_conda.py b/scripts/generate_pip_deps_from_conda.py
index f1c7c3298fb26..6f809669d917f 100755
--- a/scripts/generate_pip_deps_from_conda.py
+++ b/scripts/generate_pip_deps_from_conda.py
@@ -19,7 +19,7 @@
import yaml
-EXCLUDE = {"python=3"}
+EXCLUDE = {"python"}
RENAME = {"pytables": "tables", "pyqt": "pyqt5", "dask-core": "dask"}
@@ -33,15 +33,15 @@ def conda_package_to_pip(package):
- A package requiring a specific version, in conda is defined with a single
equal (e.g. ``pandas=1.0``) and in pip with two (e.g. ``pandas==1.0``)
"""
- if package in EXCLUDE:
- return
-
package = re.sub("(?<=[^<>])=", "==", package).strip()
+
for compare in ("<=", ">=", "=="):
if compare not in package:
continue
pkg, version = package.split(compare)
+ if pkg in EXCLUDE:
+ return
if pkg in RENAME:
return "".join((RENAME[pkg], compare, version))
| Closes https://github.com/pandas-dev/pandas/issues/29443
cc @MomIsBestFriend | https://api.github.com/repos/pandas-dev/pandas/pulls/29445 | 2019-11-06T21:07:27Z | 2019-11-07T14:28:18Z | 2019-11-07T14:28:17Z | 2019-11-07T14:28:22Z |
Adding more documentation for upsampling with replacement and error m… | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index b40a64420a0be..1cb1f745fb61b 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -333,6 +333,7 @@ Numeric
- :class:`DataFrame` flex inequality comparisons methods (:meth:`DataFrame.lt`, :meth:`DataFrame.le`, :meth:`DataFrame.gt`, :meth: `DataFrame.ge`) with object-dtype and ``complex`` entries failing to raise ``TypeError`` like their :class:`Series` counterparts (:issue:`28079`)
- Bug in :class:`DataFrame` logical operations (`&`, `|`, `^`) not matching :class:`Series` behavior by filling NA values (:issue:`28741`)
- Bug in :meth:`DataFrame.interpolate` where specifying axis by name references variable before it is assigned (:issue:`29142`)
+- Improved error message when using `frac` > 1 and `replace` = False (:issue:`27451`)
-
Conversion
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index bafc37d478fdb..ffe8e794a03ea 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -4934,6 +4934,10 @@ def sample(
numpy.random.choice: Generates a random sample from a given 1-D numpy
array.
+ Notes
+ -----
+ If `frac` > 1, `replacement` should be set to `True`.
+
Examples
--------
>>> df = pd.DataFrame({'num_legs': [2, 4, 8, 0],
@@ -4964,6 +4968,20 @@ def sample(
dog 4 0 2
fish 0 0 8
+ An upsample sample of the ``DataFrame`` with replacement:
+ Note that `replace` parameter has to be `True` for `frac` parameter > 1.
+
+ >>> df.sample(frac=2, replace=True, random_state=1)
+ num_legs num_wings num_specimen_seen
+ dog 4 0 2
+ fish 0 0 8
+ falcon 2 2 10
+ falcon 2 2 10
+ fish 0 0 8
+ dog 4 0 2
+ fish 0 0 8
+ dog 4 0 2
+
Using a DataFrame column as weights. Rows with larger value in the
`num_specimen_seen` column are more likely to be sampled.
@@ -5039,6 +5057,11 @@ def sample(
# If no frac or n, default to n=1.
if n is None and frac is None:
n = 1
+ elif frac is not None and frac > 1 and not replace:
+ raise ValueError(
+ "Replace has to be set to `True` when "
+ "upsampling the population `frac` > 1."
+ )
elif n is not None and frac is None and n % 1 != 0:
raise ValueError("Only integers accepted as `n` values")
elif n is None and frac is not None:
diff --git a/pandas/tests/generic/test_generic.py b/pandas/tests/generic/test_generic.py
index a7506f3d60b3c..c180511e31619 100644
--- a/pandas/tests/generic/test_generic.py
+++ b/pandas/tests/generic/test_generic.py
@@ -322,6 +322,7 @@ def test_sample(self):
self._compare(
o.sample(n=4, random_state=seed), o.sample(n=4, random_state=seed)
)
+
self._compare(
o.sample(frac=0.7, random_state=seed),
o.sample(frac=0.7, random_state=seed),
@@ -337,6 +338,15 @@ def test_sample(self):
o.sample(frac=0.7, random_state=np.random.RandomState(test)),
)
+ self._compare(
+ o.sample(
+ frac=2, replace=True, random_state=np.random.RandomState(test)
+ ),
+ o.sample(
+ frac=2, replace=True, random_state=np.random.RandomState(test)
+ ),
+ )
+
os1, os2 = [], []
for _ in range(2):
np.random.seed(test)
@@ -424,6 +434,17 @@ def test_sample(self):
weights_with_None[5] = 0.5
self._compare(o.sample(n=1, axis=0, weights=weights_with_None), o.iloc[5:6])
+ def test_sample_upsampling_without_replacement(self):
+ # GH27451
+
+ df = pd.DataFrame({"A": list("abc")})
+ msg = (
+ "Replace has to be set to `True` when "
+ "upsampling the population `frac` > 1."
+ )
+ with pytest.raises(ValueError, match=msg):
+ df.sample(frac=2, replace=False)
+
def test_size_compat(self):
# GH8846
# size property should be defined
| …essage in case replacement is set to False
- [X] closes #27451
- [x] tests added / passed
- [X] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29444 | 2019-11-06T20:41:21Z | 2019-11-08T01:03:24Z | 2019-11-08T01:03:24Z | 2019-11-08T01:03:38Z |
TST: Add docstrings to arithmetic fixtures | diff --git a/pandas/tests/arithmetic/conftest.py b/pandas/tests/arithmetic/conftest.py
index 774ff14398bdb..1f8fdfd671856 100644
--- a/pandas/tests/arithmetic/conftest.py
+++ b/pandas/tests/arithmetic/conftest.py
@@ -21,7 +21,24 @@ def id_func(x):
@pytest.fixture(params=[1, np.array(1, dtype=np.int64)])
def one(request):
- # zero-dim integer array behaves like an integer
+ """
+ Several variants of integer value 1. The zero-dim integer array
+ behaves like an integer.
+
+ This fixture can be used to check that datetimelike indexes handle
+ addition and subtraction of integers and zero-dimensional arrays
+ of integers.
+
+ Examples
+ --------
+ >>> dti = pd.date_range('2016-01-01', periods=2, freq='H')
+ >>> dti
+ DatetimeIndex(['2016-01-01 00:00:00', '2016-01-01 01:00:00'],
+ dtype='datetime64[ns]', freq='H')
+ >>> dti + one
+ DatetimeIndex(['2016-01-01 01:00:00', '2016-01-01 02:00:00'],
+ dtype='datetime64[ns]', freq='H')
+ """
return request.param
@@ -40,8 +57,21 @@ def one(request):
@pytest.fixture(params=zeros)
def zero(request):
- # For testing division by (or of) zero for Index with length 5, this
- # gives several scalar-zeros and length-5 vector-zeros
+ """
+ Several types of scalar zeros and length 5 vectors of zeros.
+
+ This fixture can be used to check that numeric-dtype indexes handle
+ division by any zero numeric-dtype.
+
+ Uses vector of length 5 for broadcasting with `numeric_idx` fixture,
+ which creates numeric-dtype vectors also of length 5.
+
+ Examples
+ --------
+ >>> arr = pd.RangeIndex(5)
+ >>> arr / zeros
+ Float64Index([nan, inf, inf, inf, inf], dtype='float64')
+ """
return request.param
| Relates to #19159
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29441 | 2019-11-06T20:27:23Z | 2019-11-18T01:40:48Z | 2019-11-18T01:40:48Z | 2019-11-18T01:40:59Z |
add unit tests for issue #19351 | diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 79c9fe2b60bd9..a1f58922ea0ca 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -358,6 +358,49 @@ def test_unstack(self):
# test that int32 work
self.ymd.astype(np.int32).unstack()
+ @pytest.mark.parametrize(
+ "result_rows,result_columns,index_product,expected_row",
+ [
+ (
+ [[1, 1, None, None, 30.0, None], [2, 2, None, None, 30.0, None]],
+ [u"ix1", u"ix2", u"col1", u"col2", u"col3", u"col4"],
+ 2,
+ [None, None, 30.0, None],
+ ),
+ (
+ [[1, 1, None, None, 30.0], [2, 2, None, None, 30.0]],
+ [u"ix1", u"ix2", u"col1", u"col2", u"col3"],
+ 2,
+ [None, None, 30.0],
+ ),
+ (
+ [[1, 1, None, None, 30.0], [2, None, None, None, 30.0]],
+ [u"ix1", u"ix2", u"col1", u"col2", u"col3"],
+ None,
+ [None, None, 30.0],
+ ),
+ ],
+ )
+ def test_unstack_partial(
+ self, result_rows, result_columns, index_product, expected_row
+ ):
+ # check for regressions on this issue:
+ # https://github.com/pandas-dev/pandas/issues/19351
+ # make sure DataFrame.unstack() works when its run on a subset of the DataFrame
+ # and the Index levels contain values that are not present in the subset
+ result = pd.DataFrame(result_rows, columns=result_columns).set_index(
+ [u"ix1", "ix2"]
+ )
+ result = result.iloc[1:2].unstack("ix2")
+ expected = pd.DataFrame(
+ [expected_row],
+ columns=pd.MultiIndex.from_product(
+ [result_columns[2:], [index_product]], names=[None, "ix2"]
+ ),
+ index=pd.Index([2], name="ix1"),
+ )
+ tm.assert_frame_equal(result, expected)
+
def test_unstack_multiple_no_empty_columns(self):
index = MultiIndex.from_tuples(
[(0, "foo", 0), (0, "bar", 0), (1, "baz", 1), (1, "qux", 1)]
| - [x] closes #19351
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
This is a unit test for an already fixed problem.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29438 | 2019-11-06T19:33:12Z | 2019-11-06T21:23:27Z | 2019-11-06T21:23:27Z | 2019-11-06T21:23:33Z |
CLN: remove unnecessary check in MultiIndex | diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index 2e3f440573a0f..fe91a588c7dde 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -2164,6 +2164,7 @@ def drop(self, codes, level=None, errors="raise"):
codes : array-like
Must be a list of tuples
level : int or level name, default None
+ errors : str, default 'raise'
Returns
-------
@@ -2172,18 +2173,11 @@ def drop(self, codes, level=None, errors="raise"):
if level is not None:
return self._drop_from_level(codes, level)
- try:
- if not isinstance(codes, (np.ndarray, Index)):
+ if not isinstance(codes, (np.ndarray, Index)):
+ try:
codes = com.index_labels_to_array(codes)
- indexer = self.get_indexer(codes)
- mask = indexer == -1
- if mask.any():
- if errors != "ignore":
- raise ValueError(
- "codes {codes} not contained in axis".format(codes=codes[mask])
- )
- except Exception:
- pass
+ except ValueError:
+ pass
inds = []
for level_codes in codes:
| We raise a ValueError and immediately ignore it. | https://api.github.com/repos/pandas-dev/pandas/pulls/29437 | 2019-11-06T19:06:28Z | 2019-11-06T21:22:46Z | 2019-11-06T21:22:46Z | 2019-11-06T21:30:26Z |
CLN: core.computation, mostly typing | diff --git a/pandas/core/computation/align.py b/pandas/core/computation/align.py
index 3e1e5ed89d877..92374caa29b10 100644
--- a/pandas/core/computation/align.py
+++ b/pandas/core/computation/align.py
@@ -33,7 +33,7 @@ def _zip_axes_from_type(typ, new_axes):
return axes
-def _any_pandas_objects(terms):
+def _any_pandas_objects(terms) -> bool:
"""Check a sequence of terms for instances of PandasObject."""
return any(isinstance(term.value, PandasObject) for term in terms)
@@ -144,7 +144,8 @@ def _reconstruct_object(typ, obj, axes, dtype):
obj : object
The value to use in the type constructor
axes : dict
- The axes to use to construct the resulting pandas object
+ The axes to use to construct the resulting pandas object.
+ dtype : numpy dtype
Returns
-------
diff --git a/pandas/core/computation/engines.py b/pandas/core/computation/engines.py
index dc6378e83d229..d6e6bd62a8985 100644
--- a/pandas/core/computation/engines.py
+++ b/pandas/core/computation/engines.py
@@ -22,7 +22,7 @@ def _check_ne_builtin_clash(expr):
Parameters
----------
- terms : Term
+ expr : Term
Terms can contain
"""
names = expr.names
@@ -46,8 +46,9 @@ def __init__(self, expr):
self.aligned_axes = None
self.result_type = None
- def convert(self):
- """Convert an expression for evaluation.
+ def convert(self) -> str:
+ """
+ Convert an expression for evaluation.
Defaults to return the expression as a string.
"""
@@ -75,10 +76,9 @@ def evaluate(self):
)
@property
- def _is_aligned(self):
+ def _is_aligned(self) -> bool:
return self.aligned_axes is not None and self.result_type is not None
- @abc.abstractmethod
def _evaluate(self):
"""
Return an evaluated expression.
@@ -93,7 +93,11 @@ def _evaluate(self):
-----
Must be implemented by subclasses.
"""
- pass
+ # mypy complains if we use @abc.abstractmethod, so we do use
+ # AbstractMethodError instead
+ from pandas.errors import AbstractMethodError
+
+ raise AbstractMethodError(self)
class NumExprEngine(AbstractEngine):
@@ -101,10 +105,7 @@ class NumExprEngine(AbstractEngine):
has_neg_frac = True
- def __init__(self, expr):
- super().__init__(expr)
-
- def convert(self):
+ def convert(self) -> str:
return str(super().convert())
def _evaluate(self):
@@ -137,9 +138,6 @@ class PythonEngine(AbstractEngine):
has_neg_frac = False
- def __init__(self, expr):
- super().__init__(expr)
-
def evaluate(self):
return self.expr()
diff --git a/pandas/core/computation/eval.py b/pandas/core/computation/eval.py
index 461561a80a7e5..335ce16257faa 100644
--- a/pandas/core/computation/eval.py
+++ b/pandas/core/computation/eval.py
@@ -15,7 +15,7 @@
from pandas.io.formats.printing import pprint_thing
-def _check_engine(engine):
+def _check_engine(engine) -> str:
"""
Make sure a valid engine is passed.
@@ -64,7 +64,7 @@ def _check_engine(engine):
return engine
-def _check_parser(parser):
+def _check_parser(parser: str):
"""
Make sure a valid parser is passed.
@@ -97,14 +97,13 @@ def _check_resolvers(resolvers):
)
-def _check_expression(expr):
+def _check_expression(expr: str):
"""
Make sure an expression is not an empty string
Parameters
----------
- expr : object
- An object that can be converted to a string
+ expr : str
Raises
------
@@ -115,7 +114,7 @@ def _check_expression(expr):
raise ValueError("expr cannot be an empty string")
-def _convert_expression(expr):
+def _convert_expression(expr) -> str:
"""
Convert an object to an expression.
@@ -144,7 +143,7 @@ def _convert_expression(expr):
return s
-def _check_for_locals(expr, stack_level, parser):
+def _check_for_locals(expr, stack_level: int, parser):
from pandas.core.computation.expr import tokenize_string
at_top_of_stack = stack_level == 0
@@ -168,15 +167,15 @@ def _check_for_locals(expr, stack_level, parser):
def eval(
expr,
- parser="pandas",
+ parser: str = "pandas",
engine=None,
truediv=True,
local_dict=None,
global_dict=None,
resolvers=(),
- level=0,
+ level: int = 0,
target=None,
- inplace=False,
+ inplace: bool = False,
):
"""
Evaluate a Python expression as a string using various backends.
@@ -192,7 +191,7 @@ def eval(
Parameters
----------
- expr : str or unicode
+ expr : str
The expression to evaluate. This string cannot contain any Python
`statements
<https://docs.python.org/3/reference/simple_stmts.html#simple-statements>`__,
@@ -232,7 +231,7 @@ def eval(
``DataFrame.index`` and ``DataFrame.columns``
variables that refer to their respective :class:`~pandas.DataFrame`
instance attributes.
- level : int, optional
+ level : int, default 0
The number of prior stack frames to traverse and add to the current
scope. Most users will **not** need to change this parameter.
target : object, optional, default None
diff --git a/pandas/core/computation/expr.py b/pandas/core/computation/expr.py
index 39653c3d695b2..cf9ed96dfbed7 100644
--- a/pandas/core/computation/expr.py
+++ b/pandas/core/computation/expr.py
@@ -7,7 +7,7 @@
import itertools as it
import operator
import tokenize
-from typing import Type
+from typing import Optional, Type
import numpy as np
@@ -40,7 +40,7 @@
import pandas.io.formats.printing as printing
-def tokenize_string(source):
+def tokenize_string(source: str):
"""
Tokenize a Python source code string.
@@ -68,7 +68,8 @@ def tokenize_string(source):
def _rewrite_assign(tok):
- """Rewrite the assignment operator for PyTables expressions that use ``=``
+ """
+ Rewrite the assignment operator for PyTables expressions that use ``=``
as a substitute for ``==``.
Parameters
@@ -86,7 +87,8 @@ def _rewrite_assign(tok):
def _replace_booleans(tok):
- """Replace ``&`` with ``and`` and ``|`` with ``or`` so that bitwise
+ """
+ Replace ``&`` with ``and`` and ``|`` with ``or`` so that bitwise
precedence is changed to boolean precedence.
Parameters
@@ -110,7 +112,8 @@ def _replace_booleans(tok):
def _replace_locals(tok):
- """Replace local variables with a syntactically valid name.
+ """
+ Replace local variables with a syntactically valid name.
Parameters
----------
@@ -135,7 +138,8 @@ def _replace_locals(tok):
def _clean_spaces_backtick_quoted_names(tok):
- """Clean up a column name if surrounded by backticks.
+ """
+ Clean up a column name if surrounded by backticks.
Backtick quoted string are indicated by a certain tokval value. If a string
is a backtick quoted token it will processed by
@@ -303,7 +307,8 @@ def f(self, *args, **kwargs):
def disallow(nodes):
- """Decorator to disallow certain nodes from parsing. Raises a
+ """
+ Decorator to disallow certain nodes from parsing. Raises a
NotImplementedError instead.
Returns
@@ -324,7 +329,8 @@ def disallowed(cls):
def _op_maker(op_class, op_symbol):
- """Return a function to create an op class with its symbol already passed.
+ """
+ Return a function to create an op class with its symbol already passed.
Returns
-------
@@ -332,8 +338,8 @@ def _op_maker(op_class, op_symbol):
"""
def f(self, node, *args, **kwargs):
- """Return a partial function with an Op subclass with an operator
- already passed.
+ """
+ Return a partial function with an Op subclass with an operator already passed.
Returns
-------
@@ -813,18 +819,27 @@ class Expr:
parser : str, optional, default 'pandas'
env : Scope, optional, default None
truediv : bool, optional, default True
- level : int, optional, default 2
+ level : int, optional, default 0
"""
def __init__(
- self, expr, engine="numexpr", parser="pandas", env=None, truediv=True, level=0
+ self,
+ expr,
+ engine: str = "numexpr",
+ parser: str = "pandas",
+ env=None,
+ truediv: bool = True,
+ level: int = 0,
):
self.expr = expr
self.env = env or Scope(level=level + 1)
self.engine = engine
self.parser = parser
self.env.scope["truediv"] = truediv
- self._visitor = _parsers[parser](self.env, self.engine, self.parser)
+ self._visitor = _parsers[parser](
+ self.env, self.engine, self.parser
+ ) # type: Optional[BaseExprVisitor]
+ assert isinstance(self._visitor, BaseExprVisitor), type(self._visitor)
self.terms = self.parse()
@property
@@ -837,7 +852,7 @@ def __call__(self):
def __repr__(self) -> str:
return printing.pprint_thing(self.terms)
- def __len__(self):
+ def __len__(self) -> int:
return len(self.expr)
def parse(self):
diff --git a/pandas/core/computation/expressions.py b/pandas/core/computation/expressions.py
index 46bc762e1a0b3..d7c38af5539cb 100644
--- a/pandas/core/computation/expressions.py
+++ b/pandas/core/computation/expressions.py
@@ -157,7 +157,7 @@ def _where_numexpr(cond, a, b):
set_use_numexpr(get_option("compute.use_numexpr"))
-def _has_bool_dtype(x):
+def _has_bool_dtype(x) -> bool:
if isinstance(x, ABCDataFrame):
return "bool" in x.dtypes
try:
diff --git a/pandas/core/computation/ops.py b/pandas/core/computation/ops.py
index fe74b6994be7c..9f24b895b9f10 100644
--- a/pandas/core/computation/ops.py
+++ b/pandas/core/computation/ops.py
@@ -57,10 +57,10 @@ class UndefinedVariableError(NameError):
def __init__(self, name, is_local):
if is_local:
- msg = "local variable {0!r} is not defined"
+ msg = "local variable {name!r} is not defined"
else:
- msg = "name {0!r} is not defined"
- super().__init__(msg.format(name))
+ msg = "name {name!r} is not defined"
+ super().__init__(msg.format(name=name))
class Term:
@@ -79,7 +79,7 @@ def __init__(self, name, env, side=None, encoding=None):
self.encoding = encoding
@property
- def local_name(self):
+ def local_name(self) -> str:
return self.name.replace(_LOCAL_TAG, "")
def __repr__(self) -> str:
@@ -120,7 +120,7 @@ def update(self, value):
self.value = value
@property
- def is_scalar(self):
+ def is_scalar(self) -> bool:
return is_scalar(self._value)
@property
@@ -139,14 +139,14 @@ def type(self):
return_type = type
@property
- def raw(self):
+ def raw(self) -> str:
return pprint_thing(
- "{0}(name={1!r}, type={2})"
- "".format(self.__class__.__name__, self.name, self.type)
+ "{cls}(name={name!r}, type={typ})"
+ "".format(cls=self.__class__.__name__, name=self.name, typ=self.type)
)
@property
- def is_datetime(self):
+ def is_datetime(self) -> bool:
try:
t = self.type.type
except AttributeError:
@@ -167,7 +167,7 @@ def name(self):
return self._name
@property
- def ndim(self):
+ def ndim(self) -> int:
return self._value.ndim
@@ -209,8 +209,8 @@ def __repr__(self) -> str:
Print a generic n-ary operator and its operands using infix notation.
"""
# recurse over the operands
- parened = ("({0})".format(pprint_thing(opr)) for opr in self.operands)
- return pprint_thing(" {0} ".format(self.op).join(parened))
+ parened = ("({opr})".format(opr=pprint_thing(opr)) for opr in self.operands)
+ return pprint_thing(" {op} ".format(op=self.op).join(parened))
@property
def return_type(self):
@@ -220,7 +220,7 @@ def return_type(self):
return _result_type_many(*(term.type for term in com.flatten(self)))
@property
- def has_invalid_return_type(self):
+ def has_invalid_return_type(self) -> bool:
types = self.operand_types
obj_dtype_set = frozenset([np.dtype("object")])
return self.return_type == object and types - obj_dtype_set
@@ -230,11 +230,11 @@ def operand_types(self):
return frozenset(term.type for term in com.flatten(self))
@property
- def is_scalar(self):
+ def is_scalar(self) -> bool:
return all(operand.is_scalar for operand in self.operands)
@property
- def is_datetime(self):
+ def is_datetime(self) -> bool:
try:
t = self.return_type.type
except AttributeError:
@@ -339,7 +339,7 @@ def _cast_inplace(terms, acceptable_dtypes, dtype):
term.update(new_value)
-def is_term(obj):
+def is_term(obj) -> bool:
return isinstance(obj, Term)
@@ -354,7 +354,7 @@ class BinOp(Op):
right : Term or Op
"""
- def __init__(self, op, lhs, rhs, **kwargs):
+ def __init__(self, op: str, lhs, rhs, **kwargs):
super().__init__(op, (lhs, rhs))
self.lhs = lhs
self.rhs = rhs
@@ -369,9 +369,10 @@ def __init__(self, op, lhs, rhs, **kwargs):
# has to be made a list for python3
keys = list(_binary_ops_dict.keys())
raise ValueError(
- "Invalid binary operator {0!r}, valid"
- " operators are {1}".format(op, keys)
+ "Invalid binary operator {op!r}, valid"
+ " operators are {keys}".format(op=op, keys=keys)
)
+ assert not kwargs, kwargs
def __call__(self, env):
"""
@@ -396,7 +397,7 @@ def __call__(self, env):
return self.func(left, right)
- def evaluate(self, env, engine, parser, term_type, eval_in_python):
+ def evaluate(self, env, engine: str, parser: str, term_type, eval_in_python):
"""
Evaluate a binary operation *before* being passed to the engine.
@@ -446,6 +447,7 @@ def evaluate(self, env, engine, parser, term_type, eval_in_python):
def convert_values(self):
"""Convert datetimes to a comparable value in an expression.
"""
+ assert self.encoding is None, self.encoding
def stringify(value):
if self.encoding is not None:
@@ -488,7 +490,7 @@ def _disallow_scalar_only_bool_ops(self):
raise NotImplementedError("cannot evaluate scalar only bool ops")
-def isnumeric(dtype):
+def isnumeric(dtype) -> bool:
return issubclass(np.dtype(dtype).type, np.number)
@@ -549,8 +551,8 @@ def __init__(self, op, operand):
self.func = _unary_ops_dict[op]
except KeyError:
raise ValueError(
- "Invalid unary operator {0!r}, valid operators "
- "are {1}".format(op, _unary_ops_syms)
+ "Invalid unary operator {op!r}, valid operators "
+ "are {syms}".format(op=op, syms=_unary_ops_syms)
)
def __call__(self, env):
@@ -558,7 +560,7 @@ def __call__(self, env):
return self.func(operand)
def __repr__(self) -> str:
- return pprint_thing("{0}({1})".format(self.op, self.operand))
+ return pprint_thing("{op}({operand})".format(op=self.op, operand=self.operand))
@property
def return_type(self):
@@ -583,12 +585,12 @@ def __call__(self, env):
return self.func.func(*operands)
def __repr__(self) -> str:
- operands = map(str, self.operands)
- return pprint_thing("{0}({1})".format(self.op, ",".join(operands)))
+ operands = ",".join(str(x) for x in self.operands)
+ return pprint_thing("{op}({operands})".format(op=self.op, operands=operands))
class FuncNode:
- def __init__(self, name):
+ def __init__(self, name: str):
from pandas.core.computation.check import _NUMEXPR_INSTALLED, _NUMEXPR_VERSION
if name not in _mathops or (
@@ -596,7 +598,7 @@ def __init__(self, name):
and _NUMEXPR_VERSION < LooseVersion("2.6.9")
and name in ("floor", "ceil")
):
- raise ValueError('"{0}" is not a supported function'.format(name))
+ raise ValueError('"{name}" is not a supported function'.format(name=name))
self.name = name
self.func = getattr(np, name)
diff --git a/pandas/core/computation/pytables.py b/pandas/core/computation/pytables.py
index 3a2ea30cbc8b9..f1be87477a2c8 100644
--- a/pandas/core/computation/pytables.py
+++ b/pandas/core/computation/pytables.py
@@ -2,6 +2,7 @@
import ast
from functools import partial
+from typing import Optional
import numpy as np
@@ -33,9 +34,6 @@ def __new__(cls, name, env, side=None, encoding=None):
klass = Constant if not isinstance(name, str) else cls
return object.__new__(klass)
- def __init__(self, name, env, side=None, encoding=None):
- super().__init__(name, env, side=side, encoding=encoding)
-
def _resolve_name(self):
# must be a queryables
if self.side == "left":
@@ -56,9 +54,6 @@ def value(self):
class Constant(Term):
- def __init__(self, value, env, side=None, encoding=None):
- super().__init__(value, env, side=side, encoding=encoding)
-
def _resolve_name(self):
return self._name
@@ -129,12 +124,12 @@ def conform(self, rhs):
return rhs
@property
- def is_valid(self):
+ def is_valid(self) -> bool:
""" return True if this is a valid field """
return self.lhs in self.queryables
@property
- def is_in_table(self):
+ def is_in_table(self) -> bool:
""" return True if this is a valid column name for generation (e.g. an
actual column in the table) """
return self.queryables.get(self.lhs) is not None
@@ -200,7 +195,7 @@ def stringify(value):
return TermValue(v, v, kind)
elif kind == "bool":
if isinstance(v, str):
- v = not v.strip().lower() in [
+ v = v.strip().lower() not in [
"false",
"f",
"no",
@@ -253,6 +248,7 @@ def evaluate(self):
rhs = self.conform(self.rhs)
values = [TermValue(v, v, self.kind).value for v in rhs]
+ # TODO: Isnt TermValue(v, v, self.kind).value just `v`?
if self.is_in_table:
@@ -478,7 +474,8 @@ def _validate_where(w):
class Expr(expr.Expr):
- """ hold a pytables like expression, comprised of possibly multiple 'terms'
+ """
+ Hold a pytables like expression, comprised of possibly multiple 'terms'.
Parameters
----------
@@ -486,6 +483,7 @@ class Expr(expr.Expr):
queryables : a "kinds" map (dict of column name -> kind), or None if column
is non-indexable
encoding : an encoding that will encode the query terms
+ scope_level : int, default 0
Returns
-------
@@ -505,7 +503,7 @@ class Expr(expr.Expr):
"major_axis>=20130101"
"""
- def __init__(self, where, queryables=None, encoding=None, scope_level=0):
+ def __init__(self, where, queryables=None, encoding=None, scope_level: int = 0):
where = _validate_where(where)
@@ -513,7 +511,7 @@ def __init__(self, where, queryables=None, encoding=None, scope_level=0):
self.condition = None
self.filter = None
self.terms = None
- self._visitor = None
+ self._visitor = None # type: Optional[ExprVisitor]
# capture the environment if needed
local_dict = DeepChainMap()
@@ -523,13 +521,16 @@ def __init__(self, where, queryables=None, encoding=None, scope_level=0):
where = where.expr
elif isinstance(where, (list, tuple)):
+ # TODO: could disallow tuple arg?
+ where = list(where)
for idx, w in enumerate(where):
if isinstance(w, Expr):
local_dict = w.env.scope
else:
w = _validate_where(w)
where[idx] = w
- where = " & ".join(map("({})".format, com.flatten(where))) # noqa
+ wheres = ["({x})".format(x=x) for x in com.flatten(where)]
+ where = " & ".join(wheres)
self.expr = where
self.env = Scope(scope_level + 1, local_dict=local_dict)
@@ -574,7 +575,7 @@ def evaluate(self):
class TermValue:
""" hold a term value the we use to construct a condition/filter """
- def __init__(self, value, converted, kind):
+ def __init__(self, value, converted, kind: str):
self.value = value
self.converted = converted
self.kind = kind
@@ -593,7 +594,7 @@ def tostring(self, encoding):
return self.converted
-def maybe_expression(s):
+def maybe_expression(s) -> bool:
""" loose checking if s is a pytables-acceptable expression """
if not isinstance(s, str):
return False
diff --git a/pandas/core/computation/scope.py b/pandas/core/computation/scope.py
index 81c7b04bf3284..36710b774cc5a 100644
--- a/pandas/core/computation/scope.py
+++ b/pandas/core/computation/scope.py
@@ -9,6 +9,7 @@
import pprint
import struct
import sys
+from typing import Mapping, Tuple
import numpy as np
@@ -17,7 +18,7 @@
def _ensure_scope(
- level, global_dict=None, local_dict=None, resolvers=(), target=None, **kwargs
+ level: int, global_dict=None, local_dict=None, resolvers=(), target=None, **kwargs
):
"""Ensure that we are grabbing the correct scope."""
return Scope(
@@ -29,9 +30,11 @@ def _ensure_scope(
)
-def _replacer(x):
- """Replace a number with its hexadecimal representation. Used to tag
- temporary variables with their calling scope's id.
+def _replacer(x) -> str:
+ """
+ Replace a number with its hexadecimal representation.
+
+ Used to tag temporary variables with their calling scope's id.
"""
# get the hex repr of the binary char and remove 0x and pad by pad_size
# zeros
@@ -44,7 +47,7 @@ def _replacer(x):
return hex(hexin)
-def _raw_hex_id(obj):
+def _raw_hex_id(obj) -> str:
"""Return the padded hexadecimal id of ``obj``."""
# interpret as a pointer since that's what really what id returns
packed = struct.pack("@P", id(obj))
@@ -63,7 +66,7 @@ def _raw_hex_id(obj):
}
-def _get_pretty_string(obj):
+def _get_pretty_string(obj) -> str:
"""
Return a prettier version of obj.
@@ -106,7 +109,12 @@ class Scope:
__slots__ = ["level", "scope", "target", "resolvers", "temps"]
def __init__(
- self, level, global_dict=None, local_dict=None, resolvers=(), target=None
+ self,
+ level: int,
+ global_dict=None,
+ local_dict=None,
+ resolvers: Tuple = (),
+ target=None,
):
self.level = level + 1
@@ -127,17 +135,21 @@ def __init__(
# shallow copy here because we don't want to replace what's in
# scope when we align terms (alignment accesses the underlying
# numpy array of pandas objects)
- self.scope = self.scope.new_child((global_dict or frame.f_globals).copy())
+ self.scope = DeepChainMap(
+ self.scope.new_child((global_dict or frame.f_globals).copy())
+ )
if not isinstance(local_dict, Scope):
- self.scope = self.scope.new_child((local_dict or frame.f_locals).copy())
+ self.scope = DeepChainMap(
+ self.scope.new_child((local_dict or frame.f_locals).copy())
+ )
finally:
del frame
# assumes that resolvers are going from outermost scope to inner
if isinstance(local_dict, Scope):
resolvers += tuple(local_dict.resolvers.maps)
- self.resolvers = DeepChainMap(*resolvers)
- self.temps = {}
+ self.resolvers = DeepChainMap(*resolvers) # type: DeepChainMap
+ self.temps = {} # type: Mapping
def __repr__(self) -> str:
scope_keys = _get_pretty_string(list(self.scope.keys()))
@@ -148,19 +160,20 @@ def __repr__(self) -> str:
)
@property
- def has_resolvers(self):
- """Return whether we have any extra scope.
+ def has_resolvers(self) -> bool:
+ """
+ Return whether we have any extra scope.
For example, DataFrames pass Their columns as resolvers during calls to
``DataFrame.eval()`` and ``DataFrame.query()``.
Returns
-------
- hr : bool
+ bool
"""
return bool(len(self.resolvers))
- def resolve(self, key, is_local):
+ def resolve(self, key: str, is_local: bool):
"""
Resolve a variable name in a possibly local context.
@@ -202,7 +215,7 @@ def resolve(self, key, is_local):
raise UndefinedVariableError(key, is_local)
- def swapkey(self, old_key, new_key, new_value=None):
+ def swapkey(self, old_key: str, new_key: str, new_value=None):
"""
Replace a variable name, with a potentially new value.
@@ -223,6 +236,7 @@ def swapkey(self, old_key, new_key, new_value=None):
maps.append(self.temps)
for mapping in maps:
+ assert isinstance(mapping, (DeepChainMap, dict)), type(mapping)
if old_key in mapping:
mapping[new_key] = new_value
return
@@ -250,7 +264,7 @@ def _get_vars(self, stack, scopes):
# scope after the loop
del frame
- def update(self, level):
+ def update(self, level: int):
"""
Update the current scope by going back `level` levels.
@@ -270,7 +284,7 @@ def update(self, level):
finally:
del stack[:], stack
- def add_tmp(self, value):
+ def add_tmp(self, value) -> str:
"""
Add a temporary variable to the scope.
@@ -281,7 +295,7 @@ def add_tmp(self, value):
Returns
-------
- name : basestring
+ name : str
The name of the temporary variable created.
"""
name = "{name}_{num}_{hex_id}".format(
@@ -290,6 +304,7 @@ def add_tmp(self, value):
# add to inner most scope
assert name not in self.temps
+ assert isinstance(self.temps, dict)
self.temps[name] = value
assert name in self.temps
@@ -297,12 +312,12 @@ def add_tmp(self, value):
return name
@property
- def ntemps(self):
+ def ntemps(self) -> int:
"""The number of temporary variables in this scope"""
return len(self.temps)
@property
- def full_scope(self):
+ def full_scope(self) -> DeepChainMap:
"""
Return the full scope for use with passing to engines transparently
as a mapping.
| cc @simonjayhawkins
Largely using annotations as an excuse to put eyeballs on parts of the code that would otherwise be left alone. | https://api.github.com/repos/pandas-dev/pandas/pulls/29436 | 2019-11-06T18:03:15Z | 2019-11-11T15:38:34Z | null | 2019-11-21T19:59:49Z |
Pr09 batch 3 | diff --git a/pandas/_libs/lib.pyx b/pandas/_libs/lib.pyx
index 328b67b6722f1..a14efd3313eaf 100644
--- a/pandas/_libs/lib.pyx
+++ b/pandas/_libs/lib.pyx
@@ -125,7 +125,7 @@ def is_scalar(val: object) -> bool:
- Interval
- DateOffset
- Fraction
- - Number
+ - Number.
Returns
-------
@@ -867,9 +867,10 @@ def is_list_like(obj: object, allow_sets: bool = True):
Parameters
----------
- obj : The object to check
- allow_sets : boolean, default True
- If this parameter is False, sets will not be considered list-like
+ obj : object
+ The object to check.
+ allow_sets : bool, default True
+ If this parameter is False, sets will not be considered list-like.
.. versionadded:: 0.24.0
diff --git a/pandas/core/arrays/interval.py b/pandas/core/arrays/interval.py
index 4039cc91fb554..869019cd3d222 100644
--- a/pandas/core/arrays/interval.py
+++ b/pandas/core/arrays/interval.py
@@ -260,9 +260,9 @@ def _from_factorized(cls, values, original):
Whether the intervals are closed on the left-side, right-side, both
or neither.
copy : bool, default False
- copy the data
+ Copy the data.
dtype : dtype or None, default None
- If None, dtype will be inferred
+ If None, dtype will be inferred.
.. versionadded:: 0.23.0
@@ -383,16 +383,16 @@ def from_arrays(cls, left, right, closed="right", copy=False, dtype=None):
Parameters
----------
data : array-like (1-dimensional)
- Array of tuples
+ Array of tuples.
closed : {'left', 'right', 'both', 'neither'}, default 'right'
Whether the intervals are closed on the left-side, right-side, both
or neither.
copy : bool, default False
- by-default copy the data, this is compat only and ignored
+ By-default copy the data, this is compat only and ignored.
dtype : dtype or None, default None
- If None, dtype will be inferred
+ If None, dtype will be inferred.
- ..versionadded:: 0.23.0
+ .. versionadded:: 0.23.0
Returns
-------
diff --git a/pandas/core/dtypes/concat.py b/pandas/core/dtypes/concat.py
index f2176f573207c..a62d3d0f4e65b 100644
--- a/pandas/core/dtypes/concat.py
+++ b/pandas/core/dtypes/concat.py
@@ -185,13 +185,14 @@ def concat_categorical(to_concat, axis=0):
def union_categoricals(to_union, sort_categories=False, ignore_order=False):
"""
- Combine list-like of Categorical-like, unioning categories. All
- categories must have the same dtype.
+ Combine list-like of Categorical-like, unioning categories.
+
+ All categories must have the same dtype.
Parameters
----------
- to_union : list-like of Categorical, CategoricalIndex,
- or Series with dtype='category'
+ to_union : list-like
+ Categorical, CategoricalIndex, or Series with dtype='category'.
sort_categories : bool, default False
If true, resulting categories will be lexsorted, otherwise
they will be ordered as they appear in the data.
@@ -201,7 +202,7 @@ def union_categoricals(to_union, sort_categories=False, ignore_order=False):
Returns
-------
- result : Categorical
+ Categorical
Raises
------
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index 9d7ddcf3c7727..e418461883e6c 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -441,7 +441,7 @@ def pad(self, limit=None):
Parameters
----------
limit : int, optional
- limit of how many values to fill
+ Limit of how many values to fill.
Returns
-------
@@ -856,7 +856,7 @@ def var(self, ddof=1, *args, **kwargs):
Parameters
----------
ddof : int, default 1
- degrees of freedom
+ Degrees of freedom.
Returns
-------
@@ -1237,11 +1237,11 @@ def _upsample(self, method, limit=None, fill_value=None):
Parameters
----------
method : string {'backfill', 'bfill', 'pad', 'ffill'}
- method for upsampling
+ Method for upsampling.
limit : int, default None
- Maximum size gap to fill when reindexing
+ Maximum size gap to fill when reindexing.
fill_value : scalar, default None
- Value to use for missing values
+ Value to use for missing values.
See Also
--------
diff --git a/pandas/io/html.py b/pandas/io/html.py
index 7da7a819f81e8..9a368907b65aa 100644
--- a/pandas/io/html.py
+++ b/pandas/io/html.py
@@ -960,7 +960,7 @@ def read_html(
This value is converted to a regular expression so that there is
consistent behavior between Beautiful Soup and lxml.
- flavor : str or None, container of strings
+ flavor : str or None
The parsing engine to use. 'bs4' and 'html5lib' are synonymous with
each other, they are both there for backwards compatibility. The
default of ``None`` tries to use ``lxml`` to parse and if that fails it
@@ -974,7 +974,7 @@ def read_html(
The column (or list of columns) to use to create the index.
skiprows : int or list-like or slice or None, optional
- 0-based. Number of rows to skip after parsing the column integer. If a
+ Number of rows to skip after parsing the column integer. 0-based. If a
sequence of integers or a slice is given, will skip the rows indexed by
that sequence. Note that a single element sequence means 'skip the nth
row' whereas an integer means 'skip n rows'.
@@ -1024,18 +1024,19 @@ def read_html(
transformed content.
na_values : iterable, default None
- Custom NA values
+ Custom NA values.
keep_default_na : bool, default True
If na_values are specified and keep_default_na is False the default NaN
- values are overridden, otherwise they're appended to
+ values are overridden, otherwise they're appended to.
displayed_only : bool, default True
- Whether elements with "display: none" should be parsed
+ Whether elements with "display: none" should be parsed.
Returns
-------
- dfs : list of DataFrames
+ dfs
+ A list of DataFrames.
See Also
--------
diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index f3b0226547c78..5a2f189ad8d10 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -593,14 +593,14 @@ def assert_index_equal(
check_less_precise : bool or int, default False
Specify comparison precision. Only used when check_exact is False.
5 digits (False) or 3 digits (True) after decimal points are compared.
- If int, then specify the digits to compare
+ If int, then specify the digits to compare.
check_exact : bool, default True
Whether to compare number exactly.
check_categorical : bool, default True
Whether to compare internal Categorical exactly.
obj : str, default 'Index'
Specify object name being compared, internally used to show appropriate
- assertion message
+ assertion message.
"""
__tracebackhide__ = True
@@ -1273,10 +1273,7 @@ def assert_frame_equal(
check whether it is equivalent to 1 within the specified precision.
check_names : bool, default True
Whether to check that the `names` attribute for both the `index`
- and `column` attributes of the DataFrame is identical, i.e.
-
- * left.index.names == right.index.names
- * left.columns.names == right.columns.names
+ and `column` attributes of the DataFrame is identical.
by_blocks : bool, default False
Specify how to compare internal data. If False, compare by columns.
If True, compare by blocks.
| part of #28602 | https://api.github.com/repos/pandas-dev/pandas/pulls/29434 | 2019-11-06T17:03:42Z | 2019-11-06T20:36:34Z | 2019-11-06T20:36:34Z | 2020-01-06T16:47:00Z |
CI: workaround numpydev bug | diff --git a/ci/azure/posix.yml b/ci/azure/posix.yml
index 281107559a38c..62b15bae6d2ce 100644
--- a/ci/azure/posix.yml
+++ b/ci/azure/posix.yml
@@ -45,13 +45,16 @@ jobs:
PATTERN: "not slow and not network"
LOCALE_OVERRIDE: "zh_CN.UTF-8"
- py37_np_dev:
- ENV_FILE: ci/deps/azure-37-numpydev.yaml
- CONDA_PY: "37"
- PATTERN: "not slow and not network"
- TEST_ARGS: "-W error"
- PANDAS_TESTING_MODE: "deprecate"
- EXTRA_APT: "xsel"
+ # https://github.com/pandas-dev/pandas/issues/29432
+ # py37_np_dev:
+ # ENV_FILE: ci/deps/azure-37-numpydev.yaml
+ # CONDA_PY: "37"
+ # PATTERN: "not slow and not network"
+ # TEST_ARGS: "-W error"
+ # PANDAS_TESTING_MODE: "deprecate"
+ # EXTRA_APT: "xsel"
+ # # TODO:
+ # continueOnError: true
steps:
- script: |
| We don't want this long-term. But there's no easy way
to skip this for numpydev, since it errors in setup.
xref #29432 (keep open till numpydev is fixed) | https://api.github.com/repos/pandas-dev/pandas/pulls/29433 | 2019-11-06T16:31:35Z | 2019-11-06T19:10:04Z | 2019-11-06T19:10:03Z | 2019-11-06T21:21:01Z |
Cleanup env | diff --git a/environment.yml b/environment.yml
index 443dc483aedf8..e9ac76f5bc52c 100644
--- a/environment.yml
+++ b/environment.yml
@@ -1,11 +1,10 @@
name: pandas-dev
channels:
- - defaults
- conda-forge
dependencies:
# required
- numpy>=1.15
- - python=3
+ - python=3.7
- python-dateutil>=2.6.1
- pytz
@@ -22,7 +21,7 @@ dependencies:
- flake8-comprehensions # used by flake8, linting of unnecessary comprehensions
- flake8-rst>=0.6.0,<=0.7.0 # linting of code blocks in rst files
- isort # check that imports are in the right order
- - mypy
+ - mypy=0.720
- pycodestyle # used by flake8
# documentation
@@ -54,7 +53,6 @@ dependencies:
- moto # mock S3
- pytest>=4.0.2
- pytest-cov
- - pytest-mock
- pytest-xdist
- seaborn
- statsmodels
diff --git a/requirements-dev.txt b/requirements-dev.txt
index 7a378cd2f2697..e7df704925485 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -1,4 +1,5 @@
numpy>=1.15
+python==3.7
python-dateutil>=2.6.1
pytz
asv
@@ -9,7 +10,7 @@ flake8
flake8-comprehensions
flake8-rst>=0.6.0,<=0.7.0
isort
-mypy
+mypy==0.720
pycodestyle
gitpython
sphinx
@@ -32,7 +33,6 @@ hypothesis>=3.82
moto
pytest>=4.0.2
pytest-cov
-pytest-mock
pytest-xdist
seaborn
statsmodels
| Closes #29330
This was most likely due to inconsistent constraints between conda-forge & defaults.
Also, pinning to 3.7 for now until the 3.8 buildout is done to make the solver's life a bit easier. | https://api.github.com/repos/pandas-dev/pandas/pulls/29431 | 2019-11-06T16:09:15Z | 2019-11-06T18:12:52Z | 2019-11-06T18:12:51Z | 2019-11-06T18:13:05Z |
CLN: Replaced '%' string formating to '.format' formatting | diff --git a/pandas/_libs/parsers.pyx b/pandas/_libs/parsers.pyx
index 8b9842ba087a5..aed1ee1c6f6e5 100644
--- a/pandas/_libs/parsers.pyx
+++ b/pandas/_libs/parsers.pyx
@@ -638,18 +638,19 @@ cdef class TextReader:
elif len(zip_names) == 0:
raise ValueError('Zero files found in compressed '
- 'zip file %s', source)
+ 'zip file {source}'.format(source=source))
else:
raise ValueError('Multiple files found in compressed '
- 'zip file %s', str(zip_names))
+ 'zip file {zip_names}'
+ .format(zip_names=str(zip_names)))
elif self.compression == 'xz':
if isinstance(source, str):
source = _get_lzma_file(lzma)(source, 'rb')
else:
source = _get_lzma_file(lzma)(filename=source)
else:
- raise ValueError('Unrecognized compression type: %s' %
- self.compression)
+ raise ValueError('Unrecognized compression type: {compression_type}'
+ .format(compression_type=self.compression))
if b'utf-16' in (self.encoding or b''):
# we need to read utf-16 through UTF8Recoder.
@@ -703,8 +704,10 @@ cdef class TextReader:
self.parser.cb_io = &buffer_rd_bytes
self.parser.cb_cleanup = &del_rd_source
else:
- raise IOError('Expected file path name or file-like object,'
- ' got %s type' % type(source))
+ raise IOError('Expected file path name or file-like object, '
+ 'got {source_type} type'
+ .format(source_type=type(source))
+ )
cdef _get_header(self):
# header is now a list of lists, so field_count should use header[0]
@@ -741,11 +744,12 @@ cdef class TextReader:
self.parser.lines < hr):
msg = self.orig_header
if isinstance(msg, list):
- msg = "[%s], len of %d," % (
- ','.join(str(m) for m in msg), len(msg))
+ msg = "[{msg_lst}], len of {orig_header_len},".format(
+ msg_lst=(','.join(str(m) for m in msg)),
+ orig_msg_len=len(msg))
raise ParserError(
- 'Passed header=%s but only %d lines in file'
- % (msg, self.parser.lines))
+ 'Passed header={msg} but only {line_count} lines in file'
+ .format(msg=msg, line_count=self.parser.lines))
else:
field_count = self.parser.line_fields[hr]
@@ -779,7 +783,10 @@ cdef class TextReader:
if not self.has_mi_columns and self.mangle_dupe_cols:
while count > 0:
counts[name] = count + 1
- name = '%s.%d' % (name, count)
+ name = '{name}.{count}'.format(
+ name=name,
+ count=count
+ )
count = counts.get(name, 0)
if old_name == '':
@@ -990,7 +997,9 @@ cdef class TextReader:
cdef _end_clock(self, what):
if self.verbose:
elapsed = time.time() - self.clocks.pop(-1)
- print('%s took: %.2f ms' % (what, elapsed * 1000))
+ print('{what} took: {elapsed} ms'
+ .format(what=what, elapsed=round(elapsed * 1000, 2))
+ )
def set_noconvert(self, i):
self.noconvert.add(i)
@@ -1662,7 +1671,8 @@ cdef _to_fw_string(parser_t *parser, int64_t col, int64_t line_start,
char *data
ndarray result
- result = np.empty(line_end - line_start, dtype='|S%d' % width)
+ result = np.empty(line_end - line_start,
+ dtype='|S{width}'.format(width=width))
data = <char*>result.data
with nogil:
@@ -2176,8 +2186,9 @@ def _concatenate_chunks(list chunks):
if warning_columns:
warning_names = ','.join(warning_columns)
warning_message = " ".join([
- "Columns (%s) have mixed types." % warning_names,
+ "Columns {col_name} have mixed types."
"Specify dtype option on import or set low_memory=False."
+ .format(col_name=warning_names)
])
warnings.warn(warning_message, DtypeWarning, stacklevel=8)
return result
| REF issue #16130
- [ ] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
P.S
**Need code review** | https://api.github.com/repos/pandas-dev/pandas/pulls/29430 | 2019-11-06T10:47:11Z | 2019-11-06T15:20:35Z | null | 2019-11-11T08:49:32Z |
BUG: Styling user guide points to a wrong nbviewer link | diff --git a/doc/source/user_guide/style.ipynb b/doc/source/user_guide/style.ipynb
index 006f928c037bd..e0dc2e734e660 100644
--- a/doc/source/user_guide/style.ipynb
+++ b/doc/source/user_guide/style.ipynb
@@ -6,7 +6,7 @@
"source": [
"# Styling\n",
"\n",
- "This document is written as a Jupyter Notebook, and can be viewed or downloaded [here](http://nbviewer.ipython.org/github/pandas-dev/pandas/blob/master/doc/source/style.ipynb).\n",
+ "This document is written as a Jupyter Notebook, and can be viewed or downloaded [here](http://nbviewer.ipython.org/github/pandas-dev/pandas/blob/master/doc/source/user_guide/style.ipynb).\n",
"\n",
"You can apply **conditional formatting**, the visual styling of a DataFrame\n",
"depending on the data within, by using the ``DataFrame.style`` property.\n",
| Just missing the 'user_guide' part => one line change.
- [ ] closes #xxxx (NOT THAT I KNOW)
- [ ] tests added / passed (NO CODE CHANGE)
- [ ] passes `black pandas` (NO CODE CHANGE)
- [ ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` (NO CODE CHANGE)
- [ ] whatsnew entry (NO CHANGE)
| https://api.github.com/repos/pandas-dev/pandas/pulls/29429 | 2019-11-06T09:13:30Z | 2019-11-06T16:45:13Z | 2019-11-06T16:45:13Z | 2019-11-06T16:45:18Z |
REF: Separate window bounds calculation from aggregation functions | diff --git a/pandas/_libs/window.pyx b/pandas/_libs/window.pyx
index d6bad0f20d760..303b4f6f24eac 100644
--- a/pandas/_libs/window.pyx
+++ b/pandas/_libs/window.pyx
@@ -96,280 +96,20 @@ def _check_minp(win, minp, N, floor=None) -> int:
# Physical description: 366 p.
# Series: Prentice-Hall Series in Automatic Computation
-# ----------------------------------------------------------------------
-# The indexer objects for rolling
-# These define start/end indexers to compute offsets
-
-
-cdef class WindowIndexer:
-
- cdef:
- ndarray start, end
- int64_t N, minp, win
- bint is_variable
-
- def get_data(self):
- return (self.start, self.end, <int64_t>self.N,
- <int64_t>self.win, <int64_t>self.minp,
- self.is_variable)
-
-
-cdef class MockFixedWindowIndexer(WindowIndexer):
- """
-
- We are just checking parameters of the indexer,
- and returning a consistent API with fixed/variable
- indexers.
-
- Parameters
- ----------
- values: ndarray
- values data array
- win: int64_t
- window size
- minp: int64_t
- min number of obs in a window to consider non-NaN
- index: object
- index of the values
- floor: optional
- unit for flooring
- left_closed: bint
- left endpoint closedness
- right_closed: bint
- right endpoint closedness
-
- """
- def __init__(self, ndarray values, int64_t win, int64_t minp,
- bint left_closed, bint right_closed,
- object index=None, object floor=None):
-
- assert index is None
- self.is_variable = 0
- self.N = len(values)
- self.minp = _check_minp(win, minp, self.N, floor=floor)
- self.start = np.empty(0, dtype='int64')
- self.end = np.empty(0, dtype='int64')
- self.win = win
-
-
-cdef class FixedWindowIndexer(WindowIndexer):
- """
- create a fixed length window indexer object
- that has start & end, that point to offsets in
- the index object; these are defined based on the win
- arguments
-
- Parameters
- ----------
- values: ndarray
- values data array
- win: int64_t
- window size
- minp: int64_t
- min number of obs in a window to consider non-NaN
- index: object
- index of the values
- floor: optional
- unit for flooring the unit
- left_closed: bint
- left endpoint closedness
- right_closed: bint
- right endpoint closedness
-
- """
- def __init__(self, ndarray values, int64_t win, int64_t minp,
- bint left_closed, bint right_closed,
- object index=None, object floor=None):
- cdef:
- ndarray[int64_t] start_s, start_e, end_s, end_e
-
- assert index is None
- self.is_variable = 0
- self.N = len(values)
- self.minp = _check_minp(win, minp, self.N, floor=floor)
-
- start_s = np.zeros(win, dtype='int64')
- start_e = np.arange(win, self.N, dtype='int64') - win + 1
- self.start = np.concatenate([start_s, start_e])
-
- end_s = np.arange(win, dtype='int64') + 1
- end_e = start_e + win
- self.end = np.concatenate([end_s, end_e])
- self.win = win
-
-
-cdef class VariableWindowIndexer(WindowIndexer):
- """
- create a variable length window indexer object
- that has start & end, that point to offsets in
- the index object; these are defined based on the win
- arguments
-
- Parameters
- ----------
- values: ndarray
- values data array
- win: int64_t
- window size
- minp: int64_t
- min number of obs in a window to consider non-NaN
- index: ndarray
- index of the values
- left_closed: bint
- left endpoint closedness
- True if the left endpoint is closed, False if open
- right_closed: bint
- right endpoint closedness
- True if the right endpoint is closed, False if open
- floor: optional
- unit for flooring the unit
- """
- def __init__(self, ndarray values, int64_t win, int64_t minp,
- bint left_closed, bint right_closed, ndarray index,
- object floor=None):
-
- self.is_variable = 1
- self.N = len(index)
- self.minp = _check_minp(win, minp, self.N, floor=floor)
-
- self.start = np.empty(self.N, dtype='int64')
- self.start.fill(-1)
-
- self.end = np.empty(self.N, dtype='int64')
- self.end.fill(-1)
-
- self.build(index, win, left_closed, right_closed)
-
- # max window size
- self.win = (self.end - self.start).max()
-
- def build(self, const int64_t[:] index, int64_t win, bint left_closed,
- bint right_closed):
-
- cdef:
- ndarray[int64_t] start, end
- int64_t start_bound, end_bound, N
- Py_ssize_t i, j
-
- start = self.start
- end = self.end
- N = self.N
-
- start[0] = 0
-
- # right endpoint is closed
- if right_closed:
- end[0] = 1
- # right endpoint is open
- else:
- end[0] = 0
-
- with nogil:
-
- # start is start of slice interval (including)
- # end is end of slice interval (not including)
- for i in range(1, N):
- end_bound = index[i]
- start_bound = index[i] - win
-
- # left endpoint is closed
- if left_closed:
- start_bound -= 1
-
- # advance the start bound until we are
- # within the constraint
- start[i] = i
- for j in range(start[i - 1], i):
- if index[j] > start_bound:
- start[i] = j
- break
-
- # end bound is previous end
- # or current index
- if index[end[i - 1]] <= end_bound:
- end[i] = i + 1
- else:
- end[i] = end[i - 1]
-
- # right endpoint is open
- if not right_closed:
- end[i] -= 1
-
-
-def get_window_indexer(values, win, minp, index, closed,
- floor=None, use_mock=True):
- """
- Return the correct window indexer for the computation.
-
- Parameters
- ----------
- values: 1d ndarray
- win: integer, window size
- minp: integer, minimum periods
- index: 1d ndarray, optional
- index to the values array
- closed: string, default None
- {'right', 'left', 'both', 'neither'}
- window endpoint closedness. Defaults to 'right' in
- VariableWindowIndexer and to 'both' in FixedWindowIndexer
- floor: optional
- unit for flooring the unit
- use_mock: boolean, default True
- if we are a fixed indexer, return a mock indexer
- instead of the FixedWindow Indexer. This is a type
- compat Indexer that allows us to use a standard
- code path with all of the indexers.
-
- Returns
- -------
- tuple of 1d int64 ndarrays of the offsets & data about the window
-
- """
-
- cdef:
- bint left_closed = False
- bint right_closed = False
-
- assert closed is None or closed in ['right', 'left', 'both', 'neither']
-
- # if windows is variable, default is 'right', otherwise default is 'both'
- if closed is None:
- closed = 'right' if index is not None else 'both'
-
- if closed in ['right', 'both']:
- right_closed = True
-
- if closed in ['left', 'both']:
- left_closed = True
-
- if index is not None:
- indexer = VariableWindowIndexer(values, win, minp, left_closed,
- right_closed, index, floor)
- elif use_mock:
- indexer = MockFixedWindowIndexer(values, win, minp, left_closed,
- right_closed, index, floor)
- else:
- indexer = FixedWindowIndexer(values, win, minp, left_closed,
- right_closed, index, floor)
- return indexer.get_data()
-
# ----------------------------------------------------------------------
# Rolling count
# this is only an impl for index not None, IOW, freq aware
-def roll_count(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_count(ndarray[float64_t] values, ndarray[int64_t] start, ndarray[int64_t] end,
+ int64_t minp):
cdef:
float64_t val, count_x = 0.0
- int64_t s, e, nobs, N
+ int64_t s, e, nobs, N = len(values)
Py_ssize_t i, j
- int64_t[:] start, end
ndarray[float64_t] output
- start, end, N, win, minp, _ = get_window_indexer(values, win,
- minp, index, closed)
output = np.empty(N, dtype=float)
with nogil:
@@ -442,80 +182,75 @@ cdef inline void remove_sum(float64_t val, int64_t *nobs, float64_t *sum_x) nogi
sum_x[0] = sum_x[0] - val
-def roll_sum(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_sum_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
cdef:
- float64_t val, prev_x, sum_x = 0
- int64_t s, e, range_endpoint
- int64_t nobs = 0, i, j, N
- bint is_variable
- int64_t[:] start, end
+ float64_t sum_x = 0
+ int64_t s, e
+ int64_t nobs = 0, i, j, N = len(values)
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed,
- floor=0)
output = np.empty(N, dtype=float)
- # for performance we are going to iterate
- # fixed windows separately, makes the code more complex as we have 2 paths
- # but is faster
+ with nogil:
- if is_variable:
+ for i in range(0, N):
+ s = start[i]
+ e = end[i]
- # variable window
- with nogil:
+ if i == 0:
- for i in range(0, N):
- s = start[i]
- e = end[i]
+ # setup
+ sum_x = 0.0
+ nobs = 0
+ for j in range(s, e):
+ add_sum(values[j], &nobs, &sum_x)
- if i == 0:
+ else:
- # setup
- sum_x = 0.0
- nobs = 0
- for j in range(s, e):
- add_sum(values[j], &nobs, &sum_x)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ remove_sum(values[j], &nobs, &sum_x)
- else:
+ # calculate adds
+ for j in range(end[i - 1], e):
+ add_sum(values[j], &nobs, &sum_x)
- # calculate deletes
- for j in range(start[i - 1], s):
- remove_sum(values[j], &nobs, &sum_x)
+ output[i] = calc_sum(minp, nobs, sum_x)
- # calculate adds
- for j in range(end[i - 1], e):
- add_sum(values[j], &nobs, &sum_x)
+ return output
- output[i] = calc_sum(minp, nobs, sum_x)
- else:
+def roll_sum_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
+ cdef:
+ float64_t val, prev_x, sum_x = 0
+ int64_t range_endpoint
+ int64_t nobs = 0, i, N = len(values)
+ ndarray[float64_t] output
- # fixed window
+ output = np.empty(N, dtype=float)
- range_endpoint = int_max(minp, 1) - 1
+ range_endpoint = int_max(minp, 1) - 1
- with nogil:
+ with nogil:
- for i in range(0, range_endpoint):
- add_sum(values[i], &nobs, &sum_x)
- output[i] = NaN
+ for i in range(0, range_endpoint):
+ add_sum(values[i], &nobs, &sum_x)
+ output[i] = NaN
- for i in range(range_endpoint, N):
- val = values[i]
- add_sum(val, &nobs, &sum_x)
+ for i in range(range_endpoint, N):
+ val = values[i]
+ add_sum(val, &nobs, &sum_x)
- if i > win - 1:
- prev_x = values[i - win]
- remove_sum(prev_x, &nobs, &sum_x)
+ if i > win - 1:
+ prev_x = values[i - win]
+ remove_sum(prev_x, &nobs, &sum_x)
- output[i] = calc_sum(minp, nobs, sum_x)
+ output[i] = calc_sum(minp, nobs, sum_x)
return output
-
# ----------------------------------------------------------------------
# Rolling mean
@@ -563,77 +298,75 @@ cdef inline void remove_mean(float64_t val, Py_ssize_t *nobs, float64_t *sum_x,
neg_ct[0] = neg_ct[0] - 1
-def roll_mean(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_mean_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
- float64_t val, prev_x, result, sum_x = 0
- int64_t s, e
- bint is_variable
- Py_ssize_t nobs = 0, i, j, neg_ct = 0, N
- int64_t[:] start, end
+ float64_t val, prev_x, sum_x = 0
+ Py_ssize_t nobs = 0, i, neg_ct = 0, N = len(values)
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
- # for performance we are going to iterate
- # fixed windows separately, makes the code more complex as we have 2 paths
- # but is faster
+ with nogil:
+ for i in range(minp - 1):
+ val = values[i]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
+ output[i] = NaN
+
+ for i in range(minp - 1, N):
+ val = values[i]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
- if is_variable:
+ if i > win - 1:
+ prev_x = values[i - win]
+ remove_mean(prev_x, &nobs, &sum_x, &neg_ct)
- with nogil:
+ output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
- for i in range(0, N):
- s = start[i]
- e = end[i]
+ return output
- if i == 0:
- # setup
- sum_x = 0.0
- nobs = 0
- for j in range(s, e):
- val = values[j]
- add_mean(val, &nobs, &sum_x, &neg_ct)
+def roll_mean_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ cdef:
+ float64_t val, sum_x = 0
+ int64_t s, e
+ Py_ssize_t nobs = 0, i, j, neg_ct = 0, N = len(values)
+ ndarray[float64_t] output
- else:
+ output = np.empty(N, dtype=float)
- # calculate deletes
- for j in range(start[i - 1], s):
- val = values[j]
- remove_mean(val, &nobs, &sum_x, &neg_ct)
+ with nogil:
- # calculate adds
- for j in range(end[i - 1], e):
- val = values[j]
- add_mean(val, &nobs, &sum_x, &neg_ct)
+ for i in range(0, N):
+ s = start[i]
+ e = end[i]
- output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
+ if i == 0:
- else:
+ # setup
+ sum_x = 0.0
+ nobs = 0
+ for j in range(s, e):
+ val = values[j]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
- with nogil:
- for i in range(minp - 1):
- val = values[i]
- add_mean(val, &nobs, &sum_x, &neg_ct)
- output[i] = NaN
+ else:
- for i in range(minp - 1, N):
- val = values[i]
- add_mean(val, &nobs, &sum_x, &neg_ct)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ val = values[j]
+ remove_mean(val, &nobs, &sum_x, &neg_ct)
- if i > win - 1:
- prev_x = values[i - win]
- remove_mean(prev_x, &nobs, &sum_x, &neg_ct)
+ # calculate adds
+ for j in range(end[i - 1], e):
+ val = values[j]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
- output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
+ output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
return output
-
# ----------------------------------------------------------------------
# Rolling variance
@@ -696,8 +429,8 @@ cdef inline void remove_var(float64_t val, float64_t *nobs, float64_t *mean_x,
ssqdm_x[0] = 0
-def roll_var(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed, int ddof=1):
+def roll_var_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win, int ddof=1):
"""
Numerically stable implementation using Welford's method.
"""
@@ -705,98 +438,102 @@ def roll_var(ndarray[float64_t] values, int64_t win, int64_t minp,
float64_t mean_x = 0, ssqdm_x = 0, nobs = 0,
float64_t val, prev, delta, mean_x_old
int64_t s, e
- bint is_variable
- Py_ssize_t i, j, N
- int64_t[:] start, end
+ Py_ssize_t i, j, N = len(values)
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
# Check for windows larger than array, addresses #7297
win = min(win, N)
- # for performance we are going to iterate
- # fixed windows separately, makes the code more complex as we
- # have 2 paths but is faster
+ with nogil:
- if is_variable:
+ # Over the first window, observations can only be added, never
+ # removed
+ for i in range(win):
+ add_var(values[i], &nobs, &mean_x, &ssqdm_x)
+ output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
- with nogil:
+ # a part of Welford's method for the online variance-calculation
+ # https://en.wikipedia.org/wiki/Algorithms_for_calculating_variance
- for i in range(0, N):
+ # After the first window, observations can both be added and
+ # removed
+ for i in range(win, N):
+ val = values[i]
+ prev = values[i - win]
- s = start[i]
- e = end[i]
+ if notnan(val):
+ if prev == prev:
- # Over the first window, observations can only be added
- # never removed
- if i == 0:
+ # Adding one observation and removing another one
+ delta = val - prev
+ mean_x_old = mean_x
- for j in range(s, e):
- add_var(values[j], &nobs, &mean_x, &ssqdm_x)
+ mean_x += delta / nobs
+ ssqdm_x += ((nobs - 1) * val
+ + (nobs + 1) * prev
+ - 2 * nobs * mean_x_old) * delta / nobs
else:
+ add_var(val, &nobs, &mean_x, &ssqdm_x)
+ elif prev == prev:
+ remove_var(prev, &nobs, &mean_x, &ssqdm_x)
- # After the first window, observations can both be added
- # and removed
+ output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+
+ return output
- # calculate adds
- for j in range(end[i - 1], e):
- add_var(values[j], &nobs, &mean_x, &ssqdm_x)
- # calculate deletes
- for j in range(start[i - 1], s):
- remove_var(values[j], &nobs, &mean_x, &ssqdm_x)
+def roll_var_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int ddof=1):
+ """
+ Numerically stable implementation using Welford's method.
+ """
+ cdef:
+ float64_t mean_x = 0, ssqdm_x = 0, nobs = 0,
+ float64_t val, prev, delta, mean_x_old
+ int64_t s, e
+ Py_ssize_t i, j, N = len(values)
+ ndarray[float64_t] output
- output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+ output = np.empty(N, dtype=float)
- else:
+ with nogil:
- with nogil:
+ for i in range(0, N):
- # Over the first window, observations can only be added, never
- # removed
- for i in range(win):
- add_var(values[i], &nobs, &mean_x, &ssqdm_x)
- output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+ s = start[i]
+ e = end[i]
- # a part of Welford's method for the online variance-calculation
- # https://en.wikipedia.org/wiki/Algorithms_for_calculating_variance
+ # Over the first window, observations can only be added
+ # never removed
+ if i == 0:
- # After the first window, observations can both be added and
- # removed
- for i in range(win, N):
- val = values[i]
- prev = values[i - win]
+ for j in range(s, e):
+ add_var(values[j], &nobs, &mean_x, &ssqdm_x)
- if notnan(val):
- if prev == prev:
+ else:
- # Adding one observation and removing another one
- delta = val - prev
- mean_x_old = mean_x
+ # After the first window, observations can both be added
+ # and removed
- mean_x += delta / nobs
- ssqdm_x += ((nobs - 1) * val
- + (nobs + 1) * prev
- - 2 * nobs * mean_x_old) * delta / nobs
+ # calculate adds
+ for j in range(end[i - 1], e):
+ add_var(values[j], &nobs, &mean_x, &ssqdm_x)
- else:
- add_var(val, &nobs, &mean_x, &ssqdm_x)
- elif prev == prev:
- remove_var(prev, &nobs, &mean_x, &ssqdm_x)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ remove_var(values[j], &nobs, &mean_x, &ssqdm_x)
- output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+ output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
return output
-
# ----------------------------------------------------------------------
# Rolling skewness
+
cdef inline float64_t calc_skew(int64_t minp, int64_t nobs,
float64_t x, float64_t xx,
float64_t xxx) nogil:
@@ -861,76 +598,80 @@ cdef inline void remove_skew(float64_t val, int64_t *nobs,
xxx[0] = xxx[0] - val * val * val
-def roll_skew(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_skew_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
float64_t val, prev
float64_t x = 0, xx = 0, xxx = 0
- int64_t nobs = 0, i, j, N
+ int64_t nobs = 0, i, j, N = len(values)
int64_t s, e
- bint is_variable
- int64_t[:] start, end
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
- if is_variable:
+ with nogil:
+ for i in range(minp - 1):
+ val = values[i]
+ add_skew(val, &nobs, &x, &xx, &xxx)
+ output[i] = NaN
- with nogil:
+ for i in range(minp - 1, N):
+ val = values[i]
+ add_skew(val, &nobs, &x, &xx, &xxx)
- for i in range(0, N):
+ if i > win - 1:
+ prev = values[i - win]
+ remove_skew(prev, &nobs, &x, &xx, &xxx)
- s = start[i]
- e = end[i]
+ output[i] = calc_skew(minp, nobs, x, xx, xxx)
- # Over the first window, observations can only be added
- # never removed
- if i == 0:
+ return output
- for j in range(s, e):
- val = values[j]
- add_skew(val, &nobs, &x, &xx, &xxx)
- else:
+def roll_skew_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ cdef:
+ float64_t val, prev
+ float64_t x = 0, xx = 0, xxx = 0
+ int64_t nobs = 0, i, j, N = len(values)
+ int64_t s, e
+ ndarray[float64_t] output
+
+ output = np.empty(N, dtype=float)
- # After the first window, observations can both be added
- # and removed
+ with nogil:
- # calculate adds
- for j in range(end[i - 1], e):
- val = values[j]
- add_skew(val, &nobs, &x, &xx, &xxx)
+ for i in range(0, N):
- # calculate deletes
- for j in range(start[i - 1], s):
- val = values[j]
- remove_skew(val, &nobs, &x, &xx, &xxx)
+ s = start[i]
+ e = end[i]
- output[i] = calc_skew(minp, nobs, x, xx, xxx)
+ # Over the first window, observations can only be added
+ # never removed
+ if i == 0:
- else:
+ for j in range(s, e):
+ val = values[j]
+ add_skew(val, &nobs, &x, &xx, &xxx)
- with nogil:
- for i in range(minp - 1):
- val = values[i]
- add_skew(val, &nobs, &x, &xx, &xxx)
- output[i] = NaN
+ else:
- for i in range(minp - 1, N):
- val = values[i]
- add_skew(val, &nobs, &x, &xx, &xxx)
+ # After the first window, observations can both be added
+ # and removed
- if i > win - 1:
- prev = values[i - win]
- remove_skew(prev, &nobs, &x, &xx, &xxx)
+ # calculate adds
+ for j in range(end[i - 1], e):
+ val = values[j]
+ add_skew(val, &nobs, &x, &xx, &xxx)
- output[i] = calc_skew(minp, nobs, x, xx, xxx)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ val = values[j]
+ remove_skew(val, &nobs, &x, &xx, &xxx)
- return output
+ output[i] = calc_skew(minp, nobs, x, xx, xxx)
+ return output
# ----------------------------------------------------------------------
# Rolling kurtosis
@@ -1005,69 +746,73 @@ cdef inline void remove_kurt(float64_t val, int64_t *nobs,
xxxx[0] = xxxx[0] - val * val * val * val
-def roll_kurt(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_kurt_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
float64_t val, prev
float64_t x = 0, xx = 0, xxx = 0, xxxx = 0
- int64_t nobs = 0, i, j, N
+ int64_t nobs = 0, i, j, N = len(values)
int64_t s, e
- bint is_variable
- int64_t[:] start, end
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
- if is_variable:
+ with nogil:
- with nogil:
+ for i in range(minp - 1):
+ add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
+ output[i] = NaN
- for i in range(0, N):
+ for i in range(minp - 1, N):
+ add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
- s = start[i]
- e = end[i]
+ if i > win - 1:
+ prev = values[i - win]
+ remove_kurt(prev, &nobs, &x, &xx, &xxx, &xxxx)
- # Over the first window, observations can only be added
- # never removed
- if i == 0:
+ output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
- for j in range(s, e):
- add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+ return output
- else:
- # After the first window, observations can both be added
- # and removed
+def roll_kurt_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ cdef:
+ float64_t val, prev
+ float64_t x = 0, xx = 0, xxx = 0, xxxx = 0
+ int64_t nobs = 0, i, j, s, e, N = len(values)
+ ndarray[float64_t] output
- # calculate adds
- for j in range(end[i - 1], e):
- add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+ output = np.empty(N, dtype=float)
- # calculate deletes
- for j in range(start[i - 1], s):
- remove_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+ with nogil:
- output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
+ for i in range(0, N):
- else:
+ s = start[i]
+ e = end[i]
- with nogil:
+ # Over the first window, observations can only be added
+ # never removed
+ if i == 0:
- for i in range(minp - 1):
- add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
- output[i] = NaN
+ for j in range(s, e):
+ add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
- for i in range(minp - 1, N):
- add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
+ else:
- if i > win - 1:
- prev = values[i - win]
- remove_kurt(prev, &nobs, &x, &xx, &xxx, &xxxx)
+ # After the first window, observations can both be added
+ # and removed
- output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
+ # calculate adds
+ for j in range(end[i - 1], e):
+ add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ remove_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+
+ output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
return output
@@ -1076,31 +821,26 @@ def roll_kurt(ndarray[float64_t] values, int64_t win, int64_t minp,
# Rolling median, min, max
-def roll_median_c(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_median_c(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
float64_t val, res, prev
- bint err = 0, is_variable
+ bint err = 0
int ret = 0
skiplist_t *sl
Py_ssize_t i, j
- int64_t nobs = 0, N, s, e
+ int64_t nobs = 0, N = len(values), s, e
int midpoint
- int64_t[:] start, end
ndarray[float64_t] output
# we use the Fixed/Variable Indexer here as the
# actual skiplist ops outweigh any window computation costs
- start, end, N, win, minp, is_variable = get_window_indexer(
- values, win,
- minp, index, closed,
- use_mock=False)
output = np.empty(N, dtype=float)
- if win == 0:
+ if win == 0 or (end - start).max() == 0:
output[:] = NaN
return output
-
+ win = (end - start).max()
sl = skiplist_init(<int>win)
if sl == NULL:
raise MemoryError("skiplist_init failed")
@@ -1209,76 +949,89 @@ cdef inline numeric calc_mm(int64_t minp, Py_ssize_t nobs,
return result
-def roll_max(ndarray[numeric] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_max_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
"""
Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
Parameters
----------
- values: numpy array
- window: int, size of rolling window
- minp: if number of observations in window
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
is below this, output a NaN
- index: ndarray, optional
+ index : ndarray, optional
index for window computation
- closed: 'right', 'left', 'both', 'neither'
+ closed : 'right', 'left', 'both', 'neither'
make the interval closed on the right, left,
both or neither endpoints
"""
- return _roll_min_max(values, win, minp, index, closed=closed, is_max=1)
+ return _roll_min_max_fixed(values, start, end, minp, win, is_max=1)
-def roll_min(ndarray[numeric] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_max_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
"""
Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
Parameters
----------
- values: numpy array
- window: int, size of rolling window
- minp: if number of observations in window
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
is below this, output a NaN
- index: ndarray, optional
+ index : ndarray, optional
index for window computation
+ closed : 'right', 'left', 'both', 'neither'
+ make the interval closed on the right, left,
+ both or neither endpoints
"""
- return _roll_min_max(values, win, minp, index, is_max=0, closed=closed)
+ return _roll_min_max_variable(values, start, end, minp, is_max=1)
-cdef _roll_min_max(ndarray[numeric] values, int64_t win, int64_t minp,
- object index, object closed, bint is_max):
+def roll_min_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
"""
- Moving min/max of 1d array of any numeric type along axis=0
- ignoring NaNs.
+ Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
+
+ Parameters
+ ----------
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
+ is below this, output a NaN
+ index : ndarray, optional
+ index for window computation
"""
- cdef:
- ndarray[int64_t] starti, endi
- int64_t N
- bint is_variable
+ return _roll_min_max_fixed(values, start, end, minp, win, is_max=0)
- starti, endi, N, win, minp, is_variable = get_window_indexer(
- values, win,
- minp, index, closed)
- if is_variable:
- return _roll_min_max_variable(values, starti, endi, N, win, minp,
- is_max)
- else:
- return _roll_min_max_fixed(values, N, win, minp, is_max)
+def roll_min_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ """
+ Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
+
+ Parameters
+ ----------
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
+ is below this, output a NaN
+ index : ndarray, optional
+ index for window computation
+ """
+ return _roll_min_max_variable(values, start, end, minp, is_max=0)
cdef _roll_min_max_variable(ndarray[numeric] values,
ndarray[int64_t] starti,
ndarray[int64_t] endi,
- int64_t N,
- int64_t win,
int64_t minp,
bint is_max):
cdef:
numeric ai
int64_t i, close_offset, curr_win_size
- Py_ssize_t nobs = 0
+ Py_ssize_t nobs = 0, N = len(values)
deque Q[int64_t] # min/max always the front
deque W[int64_t] # track the whole window for nobs compute
ndarray[float64_t, ndim=1] output
@@ -1353,15 +1106,16 @@ cdef _roll_min_max_variable(ndarray[numeric] values,
cdef _roll_min_max_fixed(ndarray[numeric] values,
- int64_t N,
- int64_t win,
+ ndarray[int64_t] starti,
+ ndarray[int64_t] endi,
int64_t minp,
+ int64_t win,
bint is_max):
cdef:
numeric ai
bint should_replace
int64_t i, removed, window_i,
- Py_ssize_t nobs = 0
+ Py_ssize_t nobs = 0, N = len(values)
int64_t* death
numeric* ring
numeric* minvalue
@@ -1457,8 +1211,8 @@ interpolation_types = {
}
-def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
- int64_t minp, object index, object closed,
+def roll_quantile(ndarray[float64_t, cast=True] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win,
float64_t quantile, str interpolation):
"""
O(N log(window)) implementation using skip list
@@ -1466,10 +1220,8 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
cdef:
float64_t val, prev, midpoint, idx_with_fraction
skiplist_t *skiplist
- int64_t nobs = 0, i, j, s, e, N
+ int64_t nobs = 0, i, j, s, e, N = len(values)
Py_ssize_t idx
- bint is_variable
- int64_t[:] start, end
ndarray[float64_t] output
float64_t vlow, vhigh
InterpolationType interpolation_type
@@ -1485,16 +1237,12 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
# we use the Fixed/Variable Indexer here as the
# actual skiplist ops outweigh any window computation costs
- start, end, N, win, minp, is_variable = get_window_indexer(
- values, win,
- minp, index, closed,
- use_mock=False)
output = np.empty(N, dtype=float)
- if win == 0:
+ if win == 0 or (end - start).max() == 0:
output[:] = NaN
return output
-
+ win = (end - start).max()
skiplist = skiplist_init(<int>win)
if skiplist == NULL:
raise MemoryError("skiplist_init failed")
@@ -1575,18 +1323,17 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
return output
-def roll_generic(object obj,
- int64_t win, int64_t minp, object index, object closed,
- int offset, object func, bint raw,
- object args, object kwargs):
+def roll_generic_fixed(object obj,
+ ndarray[int64_t] start, ndarray[int64_t] end,
+ int64_t minp, int64_t win,
+ int offset, object func, bint raw,
+ object args, object kwargs):
cdef:
ndarray[float64_t] output, counts, bufarr
ndarray[float64_t, cast=True] arr
float64_t *buf
float64_t *oldbuf
- int64_t nobs = 0, i, j, s, e, N
- bint is_variable
- int64_t[:] start, end
+ int64_t nobs = 0, i, j, s, e, N = len(start)
n = len(obj)
if n == 0:
@@ -1599,36 +1346,13 @@ def roll_generic(object obj,
if not arr.flags.c_contiguous:
arr = arr.copy('C')
- counts = roll_sum(np.concatenate([np.isfinite(arr).astype(float),
- np.array([0.] * offset)]),
- win, minp, index, closed)[offset:]
-
- start, end, N, win, minp, is_variable = get_window_indexer(arr, win,
- minp, index,
- closed,
- floor=0)
+ counts = roll_sum_fixed(np.concatenate([np.isfinite(arr).astype(float),
+ np.array([0.] * offset)]),
+ start, end, minp, win)[offset:]
output = np.empty(N, dtype=float)
- if is_variable:
- # variable window arr or series
-
- if offset != 0:
- raise ValueError("unable to roll_generic with a non-zero offset")
-
- for i in range(0, N):
- s = start[i]
- e = end[i]
-
- if counts[i] >= minp:
- if raw:
- output[i] = func(arr[s:e], *args, **kwargs)
- else:
- output[i] = func(obj.iloc[s:e], *args, **kwargs)
- else:
- output[i] = NaN
-
- elif not raw:
+ if not raw:
# series
for i in range(N):
if counts[i] >= minp:
@@ -1672,6 +1396,53 @@ def roll_generic(object obj,
return output
+def roll_generic_variable(object obj,
+ ndarray[int64_t] start, ndarray[int64_t] end,
+ int64_t minp,
+ int offset, object func, bint raw,
+ object args, object kwargs):
+ cdef:
+ ndarray[float64_t] output, counts, bufarr
+ ndarray[float64_t, cast=True] arr
+ float64_t *buf
+ float64_t *oldbuf
+ int64_t nobs = 0, i, j, s, e, N = len(start)
+
+ n = len(obj)
+ if n == 0:
+ return obj
+
+ arr = np.asarray(obj)
+
+ # ndarray input
+ if raw:
+ if not arr.flags.c_contiguous:
+ arr = arr.copy('C')
+
+ counts = roll_sum_variable(np.concatenate([np.isfinite(arr).astype(float),
+ np.array([0.] * offset)]),
+ start, end, minp)[offset:]
+
+ output = np.empty(N, dtype=float)
+
+ if offset != 0:
+ raise ValueError("unable to roll_generic with a non-zero offset")
+
+ for i in range(0, N):
+ s = start[i]
+ e = end[i]
+
+ if counts[i] >= minp:
+ if raw:
+ output[i] = func(arr[s:e], *args, **kwargs)
+ else:
+ output[i] = func(obj.iloc[s:e], *args, **kwargs)
+ else:
+ output[i] = NaN
+
+ return output
+
+
# ----------------------------------------------------------------------
# Rolling sum and mean for weighted window
diff --git a/pandas/_libs/window_indexer.pyx b/pandas/_libs/window_indexer.pyx
new file mode 100644
index 0000000000000..8f49a8b9462d3
--- /dev/null
+++ b/pandas/_libs/window_indexer.pyx
@@ -0,0 +1,165 @@
+# cython: boundscheck=False, wraparound=False, cdivision=True
+
+import numpy as np
+from numpy cimport ndarray, int64_t
+
+# ----------------------------------------------------------------------
+# The indexer objects for rolling
+# These define start/end indexers to compute offsets
+
+
+class MockFixedWindowIndexer:
+ """
+
+ We are just checking parameters of the indexer,
+ and returning a consistent API with fixed/variable
+ indexers.
+
+ Parameters
+ ----------
+ values: ndarray
+ values data array
+ win: int64_t
+ window size
+ index: object
+ index of the values
+ closed: string
+ closed behavior
+ """
+ def __init__(self, ndarray values, int64_t win, object closed, object index=None):
+
+ self.start = np.empty(0, dtype='int64')
+ self.end = np.empty(0, dtype='int64')
+
+ def get_window_bounds(self):
+ return self.start, self.end
+
+
+class FixedWindowIndexer:
+ """
+ create a fixed length window indexer object
+ that has start & end, that point to offsets in
+ the index object; these are defined based on the win
+ arguments
+
+ Parameters
+ ----------
+ values: ndarray
+ values data array
+ win: int64_t
+ window size
+ index: object
+ index of the values
+ closed: string
+ closed behavior
+ """
+ def __init__(self, ndarray values, int64_t win, object closed, object index=None):
+ cdef:
+ ndarray[int64_t, ndim=1] start_s, start_e, end_s, end_e
+ int64_t N = len(values)
+
+ start_s = np.zeros(win, dtype='int64')
+ start_e = np.arange(win, N, dtype='int64') - win + 1
+ self.start = np.concatenate([start_s, start_e])[:N]
+
+ end_s = np.arange(win, dtype='int64') + 1
+ end_e = start_e + win
+ self.end = np.concatenate([end_s, end_e])[:N]
+
+ def get_window_bounds(self):
+ return self.start, self.end
+
+
+class VariableWindowIndexer:
+ """
+ create a variable length window indexer object
+ that has start & end, that point to offsets in
+ the index object; these are defined based on the win
+ arguments
+
+ Parameters
+ ----------
+ values: ndarray
+ values data array
+ win: int64_t
+ window size
+ index: ndarray
+ index of the values
+ closed: string
+ closed behavior
+ """
+ def __init__(self, ndarray values, int64_t win, object closed, ndarray index):
+ cdef:
+ bint left_closed = False
+ bint right_closed = False
+ int64_t N = len(index)
+
+ # if windows is variable, default is 'right', otherwise default is 'both'
+ if closed is None:
+ closed = 'right' if index is not None else 'both'
+
+ if closed in ['right', 'both']:
+ right_closed = True
+
+ if closed in ['left', 'both']:
+ left_closed = True
+
+ self.start, self.end = self.build(index, win, left_closed, right_closed, N)
+
+ @staticmethod
+ def build(const int64_t[:] index, int64_t win, bint left_closed,
+ bint right_closed, int64_t N):
+
+ cdef:
+ ndarray[int64_t] start, end
+ int64_t start_bound, end_bound
+ Py_ssize_t i, j
+
+ start = np.empty(N, dtype='int64')
+ start.fill(-1)
+ end = np.empty(N, dtype='int64')
+ end.fill(-1)
+
+ start[0] = 0
+
+ # right endpoint is closed
+ if right_closed:
+ end[0] = 1
+ # right endpoint is open
+ else:
+ end[0] = 0
+
+ with nogil:
+
+ # start is start of slice interval (including)
+ # end is end of slice interval (not including)
+ for i in range(1, N):
+ end_bound = index[i]
+ start_bound = index[i] - win
+
+ # left endpoint is closed
+ if left_closed:
+ start_bound -= 1
+
+ # advance the start bound until we are
+ # within the constraint
+ start[i] = i
+ for j in range(start[i - 1], i):
+ if index[j] > start_bound:
+ start[i] = j
+ break
+
+ # end bound is previous end
+ # or current index
+ if index[end[i - 1]] <= end_bound:
+ end[i] = i + 1
+ else:
+ end[i] = end[i - 1]
+
+ # right endpoint is open
+ if not right_closed:
+ end[i] -= 1
+ return start, end
+
+ def get_window_bounds(self):
+ return self.start, self.end
diff --git a/pandas/core/window/common.py b/pandas/core/window/common.py
index 3fd567f97edae..453fd12495543 100644
--- a/pandas/core/window/common.py
+++ b/pandas/core/window/common.py
@@ -1,5 +1,6 @@
"""Common utility functions for rolling operations"""
from collections import defaultdict
+from typing import Callable, Optional
import warnings
import numpy as np
@@ -62,12 +63,20 @@ def __init__(self, obj, *args, **kwargs):
cov = _dispatch("cov", other=None, pairwise=None)
def _apply(
- self, func, name=None, window=None, center=None, check_minp=None, **kwargs
+ self,
+ func: Callable,
+ center: bool,
+ require_min_periods: int = 0,
+ floor: int = 1,
+ is_weighted: bool = False,
+ name: Optional[str] = None,
+ **kwargs,
):
"""
Dispatch to apply; we are stripping all of the _apply kwargs and
performing the original function call on the grouped object.
"""
+ kwargs.pop("floor", None)
# TODO: can we de-duplicate with _dispatch?
def f(x, name=name, *args):
@@ -267,6 +276,44 @@ def _use_window(minp, window):
return minp
+def calculate_min_periods(
+ window: int,
+ min_periods: Optional[int],
+ num_values: int,
+ required_min_periods: int,
+ floor: int,
+) -> int:
+ """
+ Calculates final minimum periods value for rolling aggregations.
+
+ Parameters
+ ----------
+ window : passed window value
+ min_periods : passed min periods value
+ num_values : total number of values
+ required_min_periods : required min periods per aggregation function
+ floor : required min periods per aggregation function
+
+ Returns
+ -------
+ min_periods : int
+ """
+ if min_periods is None:
+ min_periods = window
+ else:
+ min_periods = max(required_min_periods, min_periods)
+ if min_periods > window:
+ raise ValueError(
+ "min_periods {min_periods} must be <= "
+ "window {window}".format(min_periods=min_periods, window=window)
+ )
+ elif min_periods > num_values:
+ min_periods = num_values + 1
+ elif min_periods < 0:
+ raise ValueError("min_periods must be >= 0")
+ return max(min_periods, floor)
+
+
def _zsqrt(x):
with np.errstate(all="ignore"):
result = np.sqrt(x)
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index bec350f6b7d8b..fd2e8aa2ad02f 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -3,6 +3,7 @@
similar to how we have a Groupby object.
"""
from datetime import timedelta
+from functools import partial
from textwrap import dedent
from typing import Callable, Dict, List, Optional, Set, Tuple, Union
import warnings
@@ -10,6 +11,7 @@
import numpy as np
import pandas._libs.window as libwindow
+import pandas._libs.window_indexer as libwindow_indexer
from pandas.compat._optional import import_optional_dependency
from pandas.compat.numpy import function as nv
from pandas.util._decorators import Appender, Substitution, cache_readonly
@@ -43,10 +45,10 @@
_doc_template,
_flex_binary_moment,
_offset,
- _require_min_periods,
_shared_docs,
_use_window,
_zsqrt,
+ calculate_min_periods,
)
@@ -366,39 +368,55 @@ def _center_window(self, result, window) -> np.ndarray:
result = np.copy(result[tuple(lead_indexer)])
return result
- def _get_roll_func(
- self, cfunc: Callable, check_minp: Callable, index: np.ndarray, **kwargs
- ) -> Callable:
+ def _get_roll_func(self, func_name: str) -> Callable:
"""
Wrap rolling function to check values passed.
Parameters
----------
- cfunc : callable
+ func_name : str
Cython function used to calculate rolling statistics
- check_minp : callable
- function to check minimum period parameter
- index : ndarray
- used for variable window
Returns
-------
func : callable
"""
+ window_func = getattr(libwindow, func_name, None)
+ if window_func is None:
+ raise ValueError(
+ "we do not support this function "
+ "in libwindow.{func_name}".format(func_name=func_name)
+ )
+ return window_func
- def func(arg, window, min_periods=None, closed=None):
- minp = check_minp(min_periods, window)
- return cfunc(arg, window, minp, index, closed, **kwargs)
+ def _get_cython_func_type(self, func):
+ """
+ Return a variable or fixed cython function type.
- return func
+ Variable algorithms do not use window while fixed do.
+ """
+ if self.is_freq_type:
+ return self._get_roll_func("{}_variable".format(func))
+ return partial(
+ self._get_roll_func("{}_fixed".format(func)), win=self._get_window()
+ )
+
+ def _get_window_indexer(self):
+ """
+ Return an indexer class that will compute the window start and end bounds
+ """
+ if self.is_freq_type:
+ return libwindow_indexer.VariableWindowIndexer
+ return libwindow_indexer.FixedWindowIndexer
def _apply(
self,
- func: Union[str, Callable],
+ func: Callable,
+ center: bool,
+ require_min_periods: int = 0,
+ floor: int = 1,
+ is_weighted: bool = False,
name: Optional[str] = None,
- window: Optional[Union[int, str]] = None,
- center: Optional[bool] = None,
- check_minp: Optional[Callable] = None,
**kwargs,
):
"""
@@ -408,13 +426,13 @@ def _apply(
Parameters
----------
- func : str/callable to apply
- name : str, optional
- name of this function
- window : int/str, default to _get_window()
- window length or offset
- center : bool, default to self.center
- check_minp : function, default to _use_window
+ func : callable function to apply
+ center : bool
+ require_min_periods : int
+ floor: int
+ is_weighted
+ name: str,
+ compatibility with groupby.rolling
**kwargs
additional arguments for rolling function and window function
@@ -422,20 +440,13 @@ def _apply(
-------
y : type of input
"""
-
- if center is None:
- center = self.center
-
- if check_minp is None:
- check_minp = _use_window
-
- if window is None:
- win_type = self._get_win_type(kwargs)
- window = self._get_window(win_type=win_type)
+ win_type = self._get_win_type(kwargs)
+ window = self._get_window(win_type=win_type)
blocks, obj = self._create_blocks()
block_list = list(blocks)
index_as_array = self._get_index()
+ window_indexer = self._get_window_indexer()
results = []
exclude = [] # type: List[Scalar]
@@ -455,36 +466,27 @@ def _apply(
results.append(values.copy())
continue
- # if we have a string function name, wrap it
- if isinstance(func, str):
- cfunc = getattr(libwindow, func, None)
- if cfunc is None:
- raise ValueError(
- "we do not support this function "
- "in libwindow.{func}".format(func=func)
- )
-
- func = self._get_roll_func(cfunc, check_minp, index_as_array, **kwargs)
-
# calculation function
- if center:
- offset = _offset(window, center)
- additional_nans = np.array([np.NaN] * offset)
+ offset = _offset(window, center) if center else 0
+ additional_nans = np.array([np.nan] * offset)
+
+ if not is_weighted:
def calc(x):
- return func(
- np.concatenate((x, additional_nans)),
- window,
- min_periods=self.min_periods,
- closed=self.closed,
+ x = np.concatenate((x, additional_nans))
+ min_periods = calculate_min_periods(
+ window, self.min_periods, len(x), require_min_periods, floor
)
+ start, end = window_indexer(
+ x, window, self.closed, index_as_array
+ ).get_window_bounds()
+ return func(x, start, end, min_periods)
else:
def calc(x):
- return func(
- x, window, min_periods=self.min_periods, closed=self.closed
- )
+ x = np.concatenate((x, additional_nans))
+ return func(x, window, self.min_periods)
with np.errstate(all="ignore"):
if values.ndim > 1:
@@ -995,8 +997,8 @@ def _get_window(
# GH #15662. `False` makes symmetric window, rather than periodic.
return sig.get_window(win_type, window, False).astype(float)
- def _get_roll_func(
- self, cfunc: Callable, check_minp: Callable, index: np.ndarray, **kwargs
+ def _get_weighted_roll_func(
+ self, cfunc: Callable, check_minp: Callable, **kwargs
) -> Callable:
def func(arg, window, min_periods=None, closed=None):
minp = check_minp(min_periods, len(window))
@@ -1070,25 +1072,38 @@ def aggregate(self, func, *args, **kwargs):
@Appender(_shared_docs["sum"])
def sum(self, *args, **kwargs):
nv.validate_window_func("sum", args, kwargs)
- return self._apply("roll_weighted_sum", **kwargs)
+ window_func = self._get_roll_func("roll_weighted_sum")
+ window_func = self._get_weighted_roll_func(window_func, _use_window)
+ return self._apply(
+ window_func, center=self.center, is_weighted=True, name="sum", **kwargs
+ )
@Substitution(name="window")
@Appender(_shared_docs["mean"])
def mean(self, *args, **kwargs):
nv.validate_window_func("mean", args, kwargs)
- return self._apply("roll_weighted_mean", **kwargs)
+ window_func = self._get_roll_func("roll_weighted_mean")
+ window_func = self._get_weighted_roll_func(window_func, _use_window)
+ return self._apply(
+ window_func, center=self.center, is_weighted=True, name="mean", **kwargs
+ )
@Substitution(name="window", versionadded="\n.. versionadded:: 1.0.0\n")
@Appender(_shared_docs["var"])
def var(self, ddof=1, *args, **kwargs):
nv.validate_window_func("var", args, kwargs)
- return self._apply("roll_weighted_var", ddof=ddof, **kwargs)
+ window_func = partial(self._get_roll_func("roll_weighted_var"), ddof=ddof)
+ window_func = self._get_weighted_roll_func(window_func, _use_window)
+ kwargs.pop("name", None)
+ return self._apply(
+ window_func, center=self.center, is_weighted=True, name="var", **kwargs
+ )
@Substitution(name="window", versionadded="\n.. versionadded:: 1.0.0\n")
@Appender(_shared_docs["std"])
def std(self, ddof=1, *args, **kwargs):
nv.validate_window_func("std", args, kwargs)
- return _zsqrt(self.var(ddof=ddof, **kwargs))
+ return _zsqrt(self.var(ddof=ddof, name="std", **kwargs))
class _Rolling(_Window):
@@ -1203,9 +1218,9 @@ def apply(self, func, raw=None, args=(), kwargs={}):
from pandas import Series
kwargs.pop("_level", None)
+ kwargs.pop("floor", None)
window = self._get_window()
offset = _offset(window, self.center)
- index_as_array = self._get_index()
# TODO: default is for backward compat
# change to False in the future
@@ -1221,28 +1236,31 @@ def apply(self, func, raw=None, args=(), kwargs={}):
)
raw = True
- def f(arg, window, min_periods, closed):
- minp = _use_window(min_periods, window)
+ window_func = partial(
+ self._get_cython_func_type("roll_generic"),
+ args=args,
+ kwargs=kwargs,
+ raw=raw,
+ offset=offset,
+ func=func,
+ )
+
+ def apply_func(values, begin, end, min_periods, raw=raw):
if not raw:
- arg = Series(arg, index=self.obj.index)
- return libwindow.roll_generic(
- arg,
- window,
- minp,
- index_as_array,
- closed,
- offset,
- func,
- raw,
- args,
- kwargs,
- )
+ values = Series(values, index=self.obj.index)
+ return window_func(values, begin, end, min_periods)
- return self._apply(f, func, args=args, kwargs=kwargs, center=False, raw=raw)
+ # TODO: Why do we always pass center=False?
+ # name=func for WindowGroupByMixin._apply
+ return self._apply(apply_func, center=False, floor=0, name=func)
def sum(self, *args, **kwargs):
nv.validate_window_func("sum", args, kwargs)
- return self._apply("roll_sum", "sum", **kwargs)
+ window_func = self._get_cython_func_type("roll_sum")
+ kwargs.pop("floor", None)
+ return self._apply(
+ window_func, center=self.center, floor=0, name="sum", **kwargs
+ )
_shared_docs["max"] = dedent(
"""
@@ -1257,7 +1275,8 @@ def sum(self, *args, **kwargs):
def max(self, *args, **kwargs):
nv.validate_window_func("max", args, kwargs)
- return self._apply("roll_max", "max", **kwargs)
+ window_func = self._get_cython_func_type("roll_max")
+ return self._apply(window_func, center=self.center, name="max", **kwargs)
_shared_docs["min"] = dedent(
"""
@@ -1298,11 +1317,13 @@ def max(self, *args, **kwargs):
def min(self, *args, **kwargs):
nv.validate_window_func("min", args, kwargs)
- return self._apply("roll_min", "min", **kwargs)
+ window_func = self._get_cython_func_type("roll_min")
+ return self._apply(window_func, center=self.center, name="min", **kwargs)
def mean(self, *args, **kwargs):
nv.validate_window_func("mean", args, kwargs)
- return self._apply("roll_mean", "mean", **kwargs)
+ window_func = self._get_cython_func_type("roll_mean")
+ return self._apply(window_func, center=self.center, name="mean", **kwargs)
_shared_docs["median"] = dedent(
"""
@@ -1342,27 +1363,40 @@ def mean(self, *args, **kwargs):
)
def median(self, **kwargs):
- return self._apply("roll_median_c", "median", **kwargs)
+ window_func = self._get_roll_func("roll_median_c")
+ window_func = partial(window_func, win=self._get_window())
+ return self._apply(window_func, center=self.center, name="median", **kwargs)
def std(self, ddof=1, *args, **kwargs):
nv.validate_window_func("std", args, kwargs)
- window = self._get_window()
- index_as_array = self._get_index()
+ kwargs.pop("require_min_periods", None)
+ window_func = self._get_cython_func_type("roll_var")
- def f(arg, *args, **kwargs):
- minp = _require_min_periods(1)(self.min_periods, window)
- return _zsqrt(
- libwindow.roll_var(arg, window, minp, index_as_array, self.closed, ddof)
- )
+ def zsqrt_func(values, begin, end, min_periods):
+ return _zsqrt(window_func(values, begin, end, min_periods, ddof=ddof))
+ # ddof passed again for compat with groupby.rolling
return self._apply(
- f, "std", check_minp=_require_min_periods(1), ddof=ddof, **kwargs
+ zsqrt_func,
+ center=self.center,
+ require_min_periods=1,
+ name="std",
+ ddof=ddof,
+ **kwargs,
)
def var(self, ddof=1, *args, **kwargs):
nv.validate_window_func("var", args, kwargs)
+ kwargs.pop("require_min_periods", None)
+ window_func = partial(self._get_cython_func_type("roll_var"), ddof=ddof)
+ # ddof passed again for compat with groupby.rolling
return self._apply(
- "roll_var", "var", check_minp=_require_min_periods(1), ddof=ddof, **kwargs
+ window_func,
+ center=self.center,
+ require_min_periods=1,
+ name="var",
+ ddof=ddof,
+ **kwargs,
)
_shared_docs[
@@ -1377,8 +1411,14 @@ def var(self, ddof=1, *args, **kwargs):
"""
def skew(self, **kwargs):
+ window_func = self._get_cython_func_type("roll_skew")
+ kwargs.pop("require_min_periods", None)
return self._apply(
- "roll_skew", "skew", check_minp=_require_min_periods(3), **kwargs
+ window_func,
+ center=self.center,
+ require_min_periods=3,
+ name="skew",
+ **kwargs,
)
_shared_docs["kurt"] = dedent(
@@ -1414,8 +1454,14 @@ def skew(self, **kwargs):
)
def kurt(self, **kwargs):
+ window_func = self._get_cython_func_type("roll_kurt")
+ kwargs.pop("require_min_periods", None)
return self._apply(
- "roll_kurt", "kurt", check_minp=_require_min_periods(4), **kwargs
+ window_func,
+ center=self.center,
+ require_min_periods=4,
+ name="kurt",
+ **kwargs,
)
_shared_docs["quantile"] = dedent(
@@ -1475,33 +1521,22 @@ def kurt(self, **kwargs):
)
def quantile(self, quantile, interpolation="linear", **kwargs):
- window = self._get_window()
- index_as_array = self._get_index()
-
- def f(arg, *args, **kwargs):
- minp = _use_window(self.min_periods, window)
- if quantile == 1.0:
- return libwindow.roll_max(
- arg, window, minp, index_as_array, self.closed
- )
- elif quantile == 0.0:
- return libwindow.roll_min(
- arg, window, minp, index_as_array, self.closed
- )
- else:
- return libwindow.roll_quantile(
- arg,
- window,
- minp,
- index_as_array,
- self.closed,
- quantile,
- interpolation,
- )
+ if quantile == 1.0:
+ window_func = self._get_cython_func_type("roll_max")
+ elif quantile == 0.0:
+ window_func = self._get_cython_func_type("roll_min")
+ else:
+ window_func = partial(
+ self._get_roll_func("roll_quantile"),
+ win=self._get_window(),
+ quantile=quantile,
+ interpolation=interpolation,
+ )
- return self._apply(
- f, "quantile", quantile=quantile, interpolation=interpolation, **kwargs
- )
+ # Pass through for groupby.rolling
+ kwargs["quantile"] = quantile
+ kwargs["interpolation"] = interpolation
+ return self._apply(window_func, center=self.center, name="quantile", **kwargs)
_shared_docs[
"cov"
@@ -1856,7 +1891,8 @@ def count(self):
# different impl for freq counting
if self.is_freq_type:
- return self._apply("roll_count", "count")
+ window_func = self._get_roll_func("roll_count")
+ return self._apply(window_func, center=self.center, name="count")
return super().count()
diff --git a/setup.py b/setup.py
index 545765ecb114d..0915b6aba113a 100755
--- a/setup.py
+++ b/setup.py
@@ -344,6 +344,7 @@ class CheckSDist(sdist_class):
"pandas/_libs/tslibs/resolution.pyx",
"pandas/_libs/tslibs/parsing.pyx",
"pandas/_libs/tslibs/tzconversion.pyx",
+ "pandas/_libs/window_indexer.pyx",
"pandas/_libs/writers.pyx",
"pandas/io/sas/sas.pyx",
]
@@ -683,6 +684,7 @@ def srcpath(name=None, suffix=".pyx", subdir="src"):
},
"_libs.testing": {"pyxfile": "_libs/testing"},
"_libs.window": {"pyxfile": "_libs/window", "language": "c++", "suffix": ".cpp"},
+ "_libs.window_indexer": {"pyxfile": "_libs/window_indexer"},
"_libs.writers": {"pyxfile": "_libs/writers"},
"io.sas._sas": {"pyxfile": "io/sas/sas"},
"io.msgpack._packer": {
| - [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Pre-req for https://github.com/pandas-dev/pandas/issues/28987
Currently many of the aggregation functions in `window.pyx` follow the form:
```
def roll_func(values, window, minp, N, closed):
# calculate window bounds _and_ validate arguments
start, end, ... = get_window_bounds(values, window, minp, N, ...)
for i in range(values):
s = start[i]
....
```
This PR refactors out the window bound calculation into `window_indexer.pyx` and validation so the aggregation functions can be of the form:
```
def roll_func(values, start, end, minp):
for i in range(values):
s = start[i]
....
```
The methods therefore in `rolling.py` now have the following pattern:
1. Fetch the correct cython aggregation function (whether the window is fixed or variable), and prep it with kwargs if needed
2. Compute the `start` and `end` window bounds from functionality in `window_indexer.pyx`
3. Pass in the `values`, `start`, `end`, `min periods` into the aggregation function. | https://api.github.com/repos/pandas-dev/pandas/pulls/29428 | 2019-11-06T07:30:05Z | 2019-11-21T12:59:31Z | 2019-11-21T12:59:31Z | 2019-11-26T14:22:18Z |
REF: separate out ShallowMixin | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 1a2f906f97152..65e531f96614a 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -4,7 +4,7 @@
import builtins
from collections import OrderedDict
import textwrap
-from typing import Dict, FrozenSet, Optional
+from typing import Dict, FrozenSet, List, Optional
import warnings
import numpy as np
@@ -569,7 +569,7 @@ def _aggregate_multiple_funcs(self, arg, _level, _axis):
try:
new_res = colg.aggregate(a)
- except (TypeError, DataError):
+ except TypeError:
pass
else:
results.append(new_res)
@@ -618,6 +618,23 @@ def _aggregate_multiple_funcs(self, arg, _level, _axis):
raise ValueError("cannot combine transform and aggregation operations")
return result
+ def _get_cython_func(self, arg: str) -> Optional[str]:
+ """
+ if we define an internal function for this argument, return it
+ """
+ return self._cython_table.get(arg)
+
+ def _is_builtin_func(self, arg):
+ """
+ if we define an builtin function for this argument, return it,
+ otherwise return the arg
+ """
+ return self._builtin_table.get(arg, arg)
+
+
+class ShallowMixin:
+ _attributes = [] # type: List[str]
+
def _shallow_copy(self, obj=None, obj_type=None, **kwargs):
"""
return a new object with the replacement attributes
@@ -633,19 +650,6 @@ def _shallow_copy(self, obj=None, obj_type=None, **kwargs):
kwargs[attr] = getattr(self, attr)
return obj_type(obj, **kwargs)
- def _get_cython_func(self, arg: str) -> Optional[str]:
- """
- if we define an internal function for this argument, return it
- """
- return self._cython_table.get(arg)
-
- def _is_builtin_func(self, arg):
- """
- if we define an builtin function for this argument, return it,
- otherwise return the arg
- """
- return self._builtin_table.get(arg, arg)
-
class IndexOpsMixin:
"""
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index e68a2efc3f4e6..9d7ddcf3c7727 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -17,7 +17,7 @@
from pandas.core.dtypes.generic import ABCDataFrame, ABCSeries
import pandas.core.algorithms as algos
-from pandas.core.base import DataError
+from pandas.core.base import DataError, ShallowMixin
from pandas.core.generic import _shared_docs
from pandas.core.groupby.base import GroupByMixin
from pandas.core.groupby.generic import SeriesGroupBy
@@ -34,7 +34,7 @@
_shared_docs_kwargs = dict() # type: Dict[str, str]
-class Resampler(_GroupBy):
+class Resampler(_GroupBy, ShallowMixin):
"""
Class for resampling datetimelike data, a groupby-like operation.
See aggregate, transform, and apply functions on this object.
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index 68eb1f630bfc3..0718acd6360bf 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -35,7 +35,7 @@
)
from pandas._typing import Axis, FrameOrSeries, Scalar
-from pandas.core.base import DataError, PandasObject, SelectionMixin
+from pandas.core.base import DataError, PandasObject, SelectionMixin, ShallowMixin
import pandas.core.common as com
from pandas.core.index import Index, ensure_index
from pandas.core.window.common import (
@@ -50,7 +50,7 @@
)
-class _Window(PandasObject, SelectionMixin):
+class _Window(PandasObject, ShallowMixin, SelectionMixin):
_attributes = [
"window",
"min_periods",
| - [x] closes #28938
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Stops catching DataError in the 1D case for _aggregate_multiple_funcs. This change is mostly unrelated, but shares the process of reasoning about what cases need _shallow_copy/DataError. | https://api.github.com/repos/pandas-dev/pandas/pulls/29427 | 2019-11-06T03:11:19Z | 2019-11-06T19:11:31Z | 2019-11-06T19:11:30Z | 2019-11-06T19:24:46Z |
BUG: fix TypeErrors raised within _python_agg_general | diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 873a31e658625..fa4a184e8f7a4 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -899,10 +899,21 @@ def _python_agg_general(self, func, *args, **kwargs):
output = {}
for name, obj in self._iterate_slices():
try:
- result, counts = self.grouper.agg_series(obj, f)
+ # if this function is invalid for this dtype, we will ignore it.
+ func(obj[:0])
except TypeError:
continue
- else:
+ except AssertionError:
+ raise
+ except Exception:
+ # Our function depends on having a non-empty argument
+ # See test_groupby_agg_err_catching
+ pass
+
+ result, counts = self.grouper.agg_series(obj, f)
+ if result is not None:
+ # TODO: only 3 test cases get None here, do something
+ # in those cases
output[name] = self._try_cast(result, obj, numeric_only=True)
if len(output) == 0:
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 2cc0e5fde2290..5bad73bf40ff5 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -61,8 +61,7 @@ class BaseGrouper:
Parameters
----------
- axis : int
- the axis to group
+ axis : Index
groupings : array of grouping
all the grouping instances to handle in this grouper
for example for grouper list to groupby, need to pass the list
@@ -78,8 +77,15 @@ class BaseGrouper:
"""
def __init__(
- self, axis, groupings, sort=True, group_keys=True, mutated=False, indexer=None
+ self,
+ axis: Index,
+ groupings,
+ sort=True,
+ group_keys=True,
+ mutated=False,
+ indexer=None,
):
+ assert isinstance(axis, Index), axis
self._filter_empty_groups = self.compressed = len(groupings) != 1
self.axis = axis
self.groupings = groupings
@@ -623,7 +629,7 @@ def _aggregate_series_pure_python(self, obj, func):
counts = np.zeros(ngroups, dtype=int)
result = None
- splitter = get_splitter(obj, group_index, ngroups, axis=self.axis)
+ splitter = get_splitter(obj, group_index, ngroups, axis=0)
for label, group in splitter:
res = func(group)
@@ -635,8 +641,12 @@ def _aggregate_series_pure_python(self, obj, func):
counts[label] = group.shape[0]
result[label] = res
- result = lib.maybe_convert_objects(result, try_float=0)
- # TODO: try_cast back to EA?
+ if result is not None:
+ # if splitter is empty, result can be None, in which case
+ # maybe_convert_objects would raise TypeError
+ result = lib.maybe_convert_objects(result, try_float=0)
+ # TODO: try_cast back to EA?
+
return result, counts
@@ -781,6 +791,11 @@ def groupings(self):
]
def agg_series(self, obj: Series, func):
+ if is_extension_array_dtype(obj.dtype):
+ # pre-empty SeriesBinGrouper from raising TypeError
+ # TODO: watch out, this can return None
+ return self._aggregate_series_pure_python(obj, func)
+
dummy = obj[:0]
grouper = libreduction.SeriesBinGrouper(obj, func, self.bins, dummy)
return grouper.get_result()
@@ -809,12 +824,13 @@ def _is_indexed_like(obj, axes) -> bool:
class DataSplitter:
- def __init__(self, data, labels, ngroups, axis=0):
+ def __init__(self, data, labels, ngroups, axis: int = 0):
self.data = data
self.labels = ensure_int64(labels)
self.ngroups = ngroups
self.axis = axis
+ assert isinstance(axis, int), axis
@cache_readonly
def slabels(self):
@@ -837,12 +853,6 @@ def __iter__(self):
starts, ends = lib.generate_slices(self.slabels, self.ngroups)
for i, (start, end) in enumerate(zip(starts, ends)):
- # Since I'm now compressing the group ids, it's now not "possible"
- # to produce empty slices because such groups would not be observed
- # in the data
- # if start >= end:
- # raise AssertionError('Start %s must be less than end %s'
- # % (str(start), str(end)))
yield i, self._chop(sdata, slice(start, end))
def _get_sorted_data(self):
diff --git a/pandas/tests/groupby/aggregate/test_other.py b/pandas/tests/groupby/aggregate/test_other.py
index 5dad868c8c3aa..1c297f3e2ada3 100644
--- a/pandas/tests/groupby/aggregate/test_other.py
+++ b/pandas/tests/groupby/aggregate/test_other.py
@@ -602,3 +602,41 @@ def test_agg_lambda_with_timezone():
columns=["date"],
)
tm.assert_frame_equal(result, expected)
+
+
+@pytest.mark.parametrize(
+ "err_cls",
+ [
+ NotImplementedError,
+ RuntimeError,
+ KeyError,
+ IndexError,
+ OSError,
+ ValueError,
+ ArithmeticError,
+ AttributeError,
+ ],
+)
+def test_groupby_agg_err_catching(err_cls):
+ # make sure we suppress anything other than TypeError or AssertionError
+ # in _python_agg_general
+
+ # Use a non-standard EA to make sure we don't go down ndarray paths
+ from pandas.tests.extension.decimal.array import DecimalArray, make_data, to_decimal
+
+ data = make_data()[:5]
+ df = pd.DataFrame(
+ {"id1": [0, 0, 0, 1, 1], "id2": [0, 1, 0, 1, 1], "decimals": DecimalArray(data)}
+ )
+
+ expected = pd.Series(to_decimal([data[0], data[3]]))
+
+ def weird_func(x):
+ # weird function that raise something other than TypeError or IndexError
+ # in _python_agg_general
+ if len(x) == 0:
+ raise err_cls
+ return x.iloc[0]
+
+ result = df["decimals"].groupby(df["id1"]).agg(weird_func)
+ tm.assert_series_equal(result, expected, check_names=False)
| cc @jreback @WillAyd
There are a few ways in which we incorrectly raise TypeError within _python_agg_general that this fixes.
A lot of the complexity in this code comes from the fact that we drop columns on which a function is invalid instead of requiring the user to subset columns. | https://api.github.com/repos/pandas-dev/pandas/pulls/29425 | 2019-11-06T01:24:04Z | 2019-11-06T21:25:07Z | 2019-11-06T21:25:07Z | 2019-11-06T21:33:09Z |
TST: consistent result in dropping NA from CSV | diff --git a/pandas/tests/io/parser/test_na_values.py b/pandas/tests/io/parser/test_na_values.py
index f154d09358dc1..f52c6b8858fd3 100644
--- a/pandas/tests/io/parser/test_na_values.py
+++ b/pandas/tests/io/parser/test_na_values.py
@@ -536,3 +536,31 @@ def test_cast_NA_to_bool_raises_error(all_parsers, data, na_values):
dtype={"a": "bool"},
na_values=na_values,
)
+
+
+def test_str_nan_dropped(all_parsers):
+ # see gh-21131
+ parser = all_parsers
+
+ data = """File: small.csv,,
+10010010233,0123,654
+foo,,bar
+01001000155,4530,898"""
+
+ result = parser.read_csv(
+ StringIO(data),
+ header=None,
+ names=["col1", "col2", "col3"],
+ dtype={"col1": str, "col2": str, "col3": str},
+ ).dropna()
+
+ expected = DataFrame(
+ {
+ "col1": ["10010010233", "01001000155"],
+ "col2": ["0123", "4530"],
+ "col3": ["654", "898"],
+ },
+ index=[1, 3],
+ )
+
+ tm.assert_frame_equal(result, expected)
| Closes https://github.com/pandas-dev/pandas/issues/21131 | https://api.github.com/repos/pandas-dev/pandas/pulls/29424 | 2019-11-05T23:10:16Z | 2019-11-06T21:11:44Z | 2019-11-06T21:11:44Z | 2019-11-06T21:43:39Z |
TST: Test nLargest with MI grouper | diff --git a/pandas/tests/groupby/test_function.py b/pandas/tests/groupby/test_function.py
index 2d7dfe49dc038..18c4d7ceddc65 100644
--- a/pandas/tests/groupby/test_function.py
+++ b/pandas/tests/groupby/test_function.py
@@ -607,6 +607,51 @@ def test_nlargest():
tm.assert_series_equal(gb.nlargest(3, keep="last"), e)
+def test_nlargest_mi_grouper():
+ # see gh-21411
+ npr = np.random.RandomState(123456789)
+
+ dts = date_range("20180101", periods=10)
+ iterables = [dts, ["one", "two"]]
+
+ idx = MultiIndex.from_product(iterables, names=["first", "second"])
+ s = Series(npr.randn(20), index=idx)
+
+ result = s.groupby("first").nlargest(1)
+
+ exp_idx = MultiIndex.from_tuples(
+ [
+ (dts[0], dts[0], "one"),
+ (dts[1], dts[1], "one"),
+ (dts[2], dts[2], "one"),
+ (dts[3], dts[3], "two"),
+ (dts[4], dts[4], "one"),
+ (dts[5], dts[5], "one"),
+ (dts[6], dts[6], "one"),
+ (dts[7], dts[7], "one"),
+ (dts[8], dts[8], "two"),
+ (dts[9], dts[9], "one"),
+ ],
+ names=["first", "first", "second"],
+ )
+
+ exp_values = [
+ 2.2129019979039612,
+ 1.8417114045748335,
+ 0.858963679564603,
+ 1.3759151378258088,
+ 0.9430284594687134,
+ 0.5296914208183142,
+ 0.8318045593815487,
+ -0.8476703342910327,
+ 0.3804446884133735,
+ -0.8028845810770998,
+ ]
+
+ expected = Series(exp_values, index=exp_idx)
+ tm.assert_series_equal(result, expected, check_exact=False, check_less_precise=True)
+
+
def test_nsmallest():
a = Series([1, 3, 5, 7, 2, 9, 0, 4, 6, 10])
b = Series(list("a" * 5 + "b" * 5))
| Closes https://github.com/pandas-dev/pandas/issues/21411 | https://api.github.com/repos/pandas-dev/pandas/pulls/29423 | 2019-11-05T22:36:02Z | 2019-11-06T19:10:35Z | 2019-11-06T19:10:35Z | 2019-11-06T19:28:04Z |
TST: ignore _version.py | diff --git a/setup.cfg b/setup.cfg
index d4657100c1291..c7a71222ac91f 100644
--- a/setup.cfg
+++ b/setup.cfg
@@ -80,6 +80,7 @@ branch = False
omit =
*/tests/*
pandas/_typing.py
+ pandas/_version.py
plugins = Cython.Coverage
[coverage:report]
| - [x] closes #26877
The file is auto-generated, not something for us to worry about | https://api.github.com/repos/pandas-dev/pandas/pulls/29421 | 2019-11-05T22:14:11Z | 2019-11-06T19:27:15Z | 2019-11-06T19:27:15Z | 2019-11-06T20:03:02Z |
Correct type inference for UInt64Index during access | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 30a828064f812..950b8db373eef 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -345,7 +345,8 @@ Numeric
- Improved error message when using `frac` > 1 and `replace` = False (:issue:`27451`)
- Bug in numeric indexes resulted in it being possible to instantiate an :class:`Int64Index`, :class:`UInt64Index`, or :class:`Float64Index` with an invalid dtype (e.g. datetime-like) (:issue:`29539`)
- Bug in :class:`UInt64Index` precision loss while constructing from a list with values in the ``np.uint64`` range (:issue:`29526`)
--
+- Bug in :class:`NumericIndex` construction that caused indexing to fail when integers in the ``np.uint64`` range were used (:issue:`28023`)
+- Bug in :class:`NumericIndex` construction that caused :class:`UInt64Index` to be casted to :class:`Float64Index` when integers in the ``np.uint64`` range were used to index a :class:`DataFrame` (:issue:`28279`)
Conversion
^^^^^^^^^^
diff --git a/pandas/core/indexes/numeric.py b/pandas/core/indexes/numeric.py
index 29f56259dac79..747a9f75a3e00 100644
--- a/pandas/core/indexes/numeric.py
+++ b/pandas/core/indexes/numeric.py
@@ -2,7 +2,7 @@
import numpy as np
-from pandas._libs import index as libindex
+from pandas._libs import index as libindex, lib
from pandas.util._decorators import Appender, cache_readonly
from pandas.core.dtypes.cast import astype_nansafe
@@ -331,13 +331,15 @@ def _convert_scalar_indexer(self, key, kind=None):
@Appender(_index_shared_docs["_convert_arr_indexer"])
def _convert_arr_indexer(self, keyarr):
- # Cast the indexer to uint64 if possible so
- # that the values returned from indexing are
- # also uint64.
- keyarr = com.asarray_tuplesafe(keyarr)
- if is_integer_dtype(keyarr):
- return com.asarray_tuplesafe(keyarr, dtype=np.uint64)
- return keyarr
+ # Cast the indexer to uint64 if possible so that the values returned
+ # from indexing are also uint64.
+ dtype = None
+ if is_integer_dtype(keyarr) or (
+ lib.infer_dtype(keyarr, skipna=False) == "integer"
+ ):
+ dtype = np.uint64
+
+ return com.asarray_tuplesafe(keyarr, dtype=dtype)
@Appender(_index_shared_docs["_convert_index_indexer"])
def _convert_index_indexer(self, keyarr):
diff --git a/pandas/tests/indexes/test_numeric.py b/pandas/tests/indexes/test_numeric.py
index 6ee1ce5c4f2ad..37976d89ecba4 100644
--- a/pandas/tests/indexes/test_numeric.py
+++ b/pandas/tests/indexes/test_numeric.py
@@ -1209,3 +1209,29 @@ def test_range_float_union_dtype():
result = other.union(index)
tm.assert_index_equal(result, expected)
+
+
+def test_uint_index_does_not_convert_to_float64():
+ # https://github.com/pandas-dev/pandas/issues/28279
+ # https://github.com/pandas-dev/pandas/issues/28023
+ series = pd.Series(
+ [0, 1, 2, 3, 4, 5],
+ index=[
+ 7606741985629028552,
+ 17876870360202815256,
+ 17876870360202815256,
+ 13106359306506049338,
+ 8991270399732411471,
+ 8991270399732411472,
+ ],
+ )
+
+ result = series.loc[[7606741985629028552, 17876870360202815256]]
+
+ expected = UInt64Index(
+ [7606741985629028552, 17876870360202815256, 17876870360202815256],
+ dtype="uint64",
+ )
+ tm.assert_index_equal(result.index, expected)
+
+ tm.assert_equal(result, series[:3])
| - [x] closes #28023 and closes #28279
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29420 | 2019-11-05T21:28:23Z | 2019-11-27T20:47:44Z | 2019-11-27T20:47:43Z | 2019-11-28T12:21:18Z |
CLN: assorted, mostly typing | diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index a08ae66865e20..2d6c8e1008ce1 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -1150,6 +1150,77 @@ def rank_2d(rank_t[:, :] in_arr, axis=0, ties_method='average',
return ranks
+ctypedef fused diff_t:
+ float64_t
+ float32_t
+ int8_t
+ int16_t
+ int32_t
+ int64_t
+
+ctypedef fused out_t:
+ float32_t
+ float64_t
+
+
+@cython.boundscheck(False)
+@cython.wraparound(False)
+def diff_2d(ndarray[diff_t, ndim=2] arr,
+ ndarray[out_t, ndim=2] out,
+ Py_ssize_t periods, int axis):
+ cdef:
+ Py_ssize_t i, j, sx, sy, start, stop
+ bint f_contig = arr.flags.f_contiguous
+
+ # Disable for unsupported dtype combinations,
+ # see https://github.com/cython/cython/issues/2646
+ if (out_t is float32_t
+ and not (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
+ raise NotImplementedError
+ elif (out_t is float64_t
+ and (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
+ raise NotImplementedError
+ else:
+ # We put this inside an indented else block to avoid cython build
+ # warnings about unreachable code
+ sx, sy = (<object>arr).shape
+ with nogil:
+ if f_contig:
+ if axis == 0:
+ if periods >= 0:
+ start, stop = periods, sx
+ else:
+ start, stop = 0, sx + periods
+ for j in range(sy):
+ for i in range(start, stop):
+ out[i, j] = arr[i, j] - arr[i - periods, j]
+ else:
+ if periods >= 0:
+ start, stop = periods, sy
+ else:
+ start, stop = 0, sy + periods
+ for j in range(start, stop):
+ for i in range(sx):
+ out[i, j] = arr[i, j] - arr[i, j - periods]
+ else:
+ if axis == 0:
+ if periods >= 0:
+ start, stop = periods, sx
+ else:
+ start, stop = 0, sx + periods
+ for i in range(start, stop):
+ for j in range(sy):
+ out[i, j] = arr[i, j] - arr[i - periods, j]
+ else:
+ if periods >= 0:
+ start, stop = periods, sy
+ else:
+ start, stop = 0, sy + periods
+ for i in range(sx):
+ for j in range(start, stop):
+ out[i, j] = arr[i, j] - arr[i, j - periods]
+
+
# generated from template
include "algos_common_helper.pxi"
include "algos_take_helper.pxi"
diff --git a/pandas/_libs/algos_common_helper.pxi.in b/pandas/_libs/algos_common_helper.pxi.in
index ea05c4afc8fce..5bfc594602dd8 100644
--- a/pandas/_libs/algos_common_helper.pxi.in
+++ b/pandas/_libs/algos_common_helper.pxi.in
@@ -4,77 +4,6 @@ Template for each `dtype` helper function using 1-d template
WARNING: DO NOT edit .pxi FILE directly, .pxi is generated from .pxi.in
"""
-ctypedef fused diff_t:
- float64_t
- float32_t
- int8_t
- int16_t
- int32_t
- int64_t
-
-ctypedef fused out_t:
- float32_t
- float64_t
-
-
-@cython.boundscheck(False)
-@cython.wraparound(False)
-def diff_2d(ndarray[diff_t, ndim=2] arr,
- ndarray[out_t, ndim=2] out,
- Py_ssize_t periods, int axis):
- cdef:
- Py_ssize_t i, j, sx, sy, start, stop
- bint f_contig = arr.flags.f_contiguous
-
- # Disable for unsupported dtype combinations,
- # see https://github.com/cython/cython/issues/2646
- if (out_t is float32_t
- and not (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
- raise NotImplementedError
- elif (out_t is float64_t
- and (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
- raise NotImplementedError
- else:
- # We put this inside an indented else block to avoid cython build
- # warnings about unreachable code
- sx, sy = (<object>arr).shape
- with nogil:
- if f_contig:
- if axis == 0:
- if periods >= 0:
- start, stop = periods, sx
- else:
- start, stop = 0, sx + periods
- for j in range(sy):
- for i in range(start, stop):
- out[i, j] = arr[i, j] - arr[i - periods, j]
- else:
- if periods >= 0:
- start, stop = periods, sy
- else:
- start, stop = 0, sy + periods
- for j in range(start, stop):
- for i in range(sx):
- out[i, j] = arr[i, j] - arr[i, j - periods]
- else:
- if axis == 0:
- if periods >= 0:
- start, stop = periods, sx
- else:
- start, stop = 0, sx + periods
- for i in range(start, stop):
- for j in range(sy):
- out[i, j] = arr[i, j] - arr[i - periods, j]
- else:
- if periods >= 0:
- start, stop = periods, sy
- else:
- start, stop = 0, sy + periods
- for i in range(sx):
- for j in range(start, stop):
- out[i, j] = arr[i, j] - arr[i, j - periods]
-
-
# ----------------------------------------------------------------------
# ensure_dtype
# ----------------------------------------------------------------------
diff --git a/pandas/_libs/missing.pyx b/pandas/_libs/missing.pyx
index 052b081988c9e..9568ddb7fe53f 100644
--- a/pandas/_libs/missing.pyx
+++ b/pandas/_libs/missing.pyx
@@ -121,7 +121,7 @@ cpdef ndarray[uint8_t] isnaobj(ndarray arr):
@cython.wraparound(False)
@cython.boundscheck(False)
-def isnaobj_old(ndarray arr):
+def isnaobj_old(arr: ndarray) -> ndarray:
"""
Return boolean mask denoting which elements of a 1-D array are na-like,
defined as being any of:
@@ -156,7 +156,7 @@ def isnaobj_old(ndarray arr):
@cython.wraparound(False)
@cython.boundscheck(False)
-def isnaobj2d(ndarray arr):
+def isnaobj2d(arr: ndarray) -> ndarray:
"""
Return boolean mask denoting which elements of a 2-D array are na-like,
according to the criteria defined in `checknull`:
@@ -198,7 +198,7 @@ def isnaobj2d(ndarray arr):
@cython.wraparound(False)
@cython.boundscheck(False)
-def isnaobj2d_old(ndarray arr):
+def isnaobj2d_old(arr: ndarray) -> ndarray:
"""
Return boolean mask denoting which elements of a 2-D array are na-like,
according to the criteria defined in `checknull_old`:
diff --git a/pandas/_libs/window.pyx b/pandas/_libs/window.pyx
index d1adc7789a7a3..b51d61d05ce98 100644
--- a/pandas/_libs/window.pyx
+++ b/pandas/_libs/window.pyx
@@ -69,8 +69,8 @@ def _check_minp(win, minp, N, floor=None) -> int:
if not util.is_integer_object(minp):
raise ValueError("min_periods must be an integer")
if minp > win:
- raise ValueError("min_periods (%d) must be <= "
- "window (%d)" % (minp, win))
+ raise ValueError("min_periods (minp) must be <= "
+ "window (win)".format(minp=minp, win=win))
elif minp > N:
minp = N + 1
elif minp < 0:
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 1a2f906f97152..0e088a381e964 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -207,7 +207,7 @@ def _selected_obj(self):
return self.obj[self._selection]
@cache_readonly
- def ndim(self):
+ def ndim(self) -> int:
return self._selected_obj.ndim
@cache_readonly
@@ -339,7 +339,7 @@ def _aggregate(self, arg, *args, **kwargs):
obj = self._selected_obj
- def nested_renaming_depr(level=4):
+ def nested_renaming_depr(level: int = 4):
# deprecation of nested renaming
# GH 15931
msg = textwrap.dedent(
@@ -488,11 +488,11 @@ def _agg(arg, func):
# combine results
- def is_any_series():
+ def is_any_series() -> bool:
# return a boolean if we have *any* nested series
return any(isinstance(r, ABCSeries) for r in result.values())
- def is_any_frame():
+ def is_any_frame() -> bool:
# return a boolean if we have *any* nested series
return any(isinstance(r, ABCDataFrame) for r in result.values())
diff --git a/pandas/core/groupby/categorical.py b/pandas/core/groupby/categorical.py
index fcf52ecfcbbcd..399ed9ddc9ba1 100644
--- a/pandas/core/groupby/categorical.py
+++ b/pandas/core/groupby/categorical.py
@@ -8,7 +8,7 @@
)
-def recode_for_groupby(c, sort, observed):
+def recode_for_groupby(c: Categorical, sort: bool, observed: bool):
"""
Code the categories to ensure we can groupby for categoricals.
@@ -74,7 +74,7 @@ def recode_for_groupby(c, sort, observed):
return c.reorder_categories(cat.categories), None
-def recode_from_groupby(c, sort, ci):
+def recode_from_groupby(c: Categorical, sort: bool, ci):
"""
Reverse the codes_to_groupby to account for sort / observed.
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 1e38dde2096ba..8512b6c3ae530 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -21,6 +21,7 @@
Tuple,
Type,
Union,
+ cast,
)
import warnings
@@ -369,7 +370,7 @@ def _wrap_applied_output(self, keys, values, not_indexed_same=False):
# GH #6265
return Series([], name=self._selection_name, index=keys)
- def _get_index():
+ def _get_index() -> Index:
if self.grouper.nkeys > 1:
index = MultiIndex.from_tuples(keys, names=self.grouper.names)
else:
@@ -462,7 +463,7 @@ def transform(self, func, *args, **kwargs):
result.index = self._selected_obj.index
return result
- def _transform_fast(self, func, func_nm):
+ def _transform_fast(self, func, func_nm) -> Series:
"""
fast version of transform, only applicable to
builtin/cythonizable functions
@@ -512,7 +513,7 @@ def filter(self, func, dropna=True, *args, **kwargs):
wrapper = lambda x: func(x, *args, **kwargs)
# Interpret np.nan as False.
- def true_and_notna(x, *args, **kwargs):
+ def true_and_notna(x, *args, **kwargs) -> bool:
b = wrapper(x, *args, **kwargs)
return b and notna(b)
@@ -526,7 +527,7 @@ def true_and_notna(x, *args, **kwargs):
filtered = self._apply_filter(indices, dropna)
return filtered
- def nunique(self, dropna=True):
+ def nunique(self, dropna: bool = True) -> Series:
"""
Return number of unique elements in the group.
@@ -719,7 +720,7 @@ def value_counts(
out = ensure_int64(out)
return Series(out, index=mi, name=self._selection_name)
- def count(self):
+ def count(self) -> Series:
"""
Compute count of group, excluding missing values.
@@ -768,8 +769,6 @@ class DataFrameGroupBy(GroupBy):
_apply_whitelist = base.dataframe_apply_whitelist
- _block_agg_axis = 1
-
_agg_see_also_doc = dedent(
"""
See Also
@@ -944,19 +943,21 @@ def _iterate_slices(self) -> Iterable[Tuple[Optional[Hashable], Series]]:
yield label, values
- def _cython_agg_general(self, how, alt=None, numeric_only=True, min_count=-1):
+ def _cython_agg_general(
+ self, how: str, alt=None, numeric_only: bool = True, min_count: int = -1
+ ):
new_items, new_blocks = self._cython_agg_blocks(
how, alt=alt, numeric_only=numeric_only, min_count=min_count
)
return self._wrap_agged_blocks(new_items, new_blocks)
- _block_agg_axis = 0
-
- def _cython_agg_blocks(self, how, alt=None, numeric_only=True, min_count=-1):
+ def _cython_agg_blocks(
+ self, how: str, alt=None, numeric_only: bool = True, min_count: int = -1
+ ):
# TODO: the actual managing of mgr_locs is a PITA
# here, it should happen via BlockManager.combine
- data, agg_axis = self._get_data_to_aggregate()
+ data = self._get_data_to_aggregate()
if numeric_only:
data = data.get_numeric_data(copy=False)
@@ -971,7 +972,7 @@ def _cython_agg_blocks(self, how, alt=None, numeric_only=True, min_count=-1):
locs = block.mgr_locs.as_array
try:
result, _ = self.grouper.aggregate(
- block.values, how, axis=agg_axis, min_count=min_count
+ block.values, how, axis=1, min_count=min_count
)
except NotImplementedError:
# generally if we have numeric_only=False
@@ -1000,12 +1001,13 @@ def _cython_agg_blocks(self, how, alt=None, numeric_only=True, min_count=-1):
# continue and exclude the block
deleted_items.append(locs)
continue
-
- # unwrap DataFrame to get array
- assert len(result._data.blocks) == 1
- result = result._data.blocks[0].values
- if result.ndim == 1 and isinstance(result, np.ndarray):
- result = result.reshape(1, -1)
+ else:
+ result = cast(DataFrame, result)
+ # unwrap DataFrame to get array
+ assert len(result._data.blocks) == 1
+ result = result._data.blocks[0].values
+ if isinstance(result, np.ndarray) and result.ndim == 1:
+ result = result.reshape(1, -1)
finally:
assert not isinstance(result, DataFrame)
@@ -1081,11 +1083,11 @@ def _aggregate_frame(self, func, *args, **kwargs):
return self._wrap_frame_output(result, obj)
- def _aggregate_item_by_item(self, func, *args, **kwargs):
+ def _aggregate_item_by_item(self, func, *args, **kwargs) -> DataFrame:
# only for axis==0
obj = self._obj_with_exclusions
- result = OrderedDict()
+ result = OrderedDict() # type: dict
cannot_agg = []
errors = None
for item in obj:
@@ -1291,12 +1293,12 @@ def first_not_none(values):
# values are not series or array-like but scalars
else:
# only coerce dates if we find at least 1 datetime
- coerce = any(isinstance(x, Timestamp) for x in values)
+ should_coerce = any(isinstance(x, Timestamp) for x in values)
# self._selection_name not passed through to Series as the
# result should not take the name of original selection
# of columns
return Series(values, index=key_index)._convert(
- datetime=True, coerce=coerce
+ datetime=True, coerce=should_coerce
)
else:
@@ -1391,7 +1393,7 @@ def transform(self, func, *args, **kwargs):
return self._transform_fast(result, obj, func)
- def _transform_fast(self, result, obj, func_nm):
+ def _transform_fast(self, result: DataFrame, obj: DataFrame, func_nm) -> DataFrame:
"""
Fast transform path for aggregations
"""
@@ -1451,7 +1453,7 @@ def _choose_path(self, fast_path, slow_path, group):
return path, res
- def _transform_item_by_item(self, obj, wrapper):
+ def _transform_item_by_item(self, obj: DataFrame, wrapper) -> DataFrame:
# iterate through columns
output = {}
inds = []
@@ -1536,7 +1538,7 @@ def filter(self, func, dropna=True, *args, **kwargs):
return self._apply_filter(indices, dropna)
- def _gotitem(self, key, ndim, subset=None):
+ def _gotitem(self, key, ndim: int, subset=None):
"""
sub-classes to define
return a sliced object
@@ -1571,7 +1573,7 @@ def _gotitem(self, key, ndim, subset=None):
raise AssertionError("invalid ndim for _gotitem")
- def _wrap_frame_output(self, result, obj):
+ def _wrap_frame_output(self, result, obj) -> DataFrame:
result_index = self.grouper.levels[0]
if self.axis == 0:
@@ -1582,9 +1584,9 @@ def _wrap_frame_output(self, result, obj):
def _get_data_to_aggregate(self):
obj = self._obj_with_exclusions
if self.axis == 1:
- return obj.T._data, 1
+ return obj.T._data
else:
- return obj._data, 1
+ return obj._data
def _insert_inaxis_grouper_inplace(self, result):
# zip in reverse so we can always insert at loc 0
@@ -1622,7 +1624,7 @@ def _wrap_aggregated_output(self, output, names=None):
return self._reindex_output(result)._convert(datetime=True)
- def _wrap_transformed_output(self, output, names=None):
+ def _wrap_transformed_output(self, output, names=None) -> DataFrame:
return DataFrame(output, index=self.obj.index)
def _wrap_agged_blocks(self, items, blocks):
@@ -1670,7 +1672,7 @@ def count(self):
DataFrame
Count of values within each group.
"""
- data, _ = self._get_data_to_aggregate()
+ data = self._get_data_to_aggregate()
ids, _, ngroups = self.grouper.group_info
mask = ids != -1
@@ -1687,7 +1689,7 @@ def count(self):
return self._wrap_agged_blocks(data.items, list(blk))
- def nunique(self, dropna=True):
+ def nunique(self, dropna: bool = True):
"""
Return DataFrame with number of distinct observations per group for
each column.
@@ -1756,7 +1758,7 @@ def groupby_series(obj, col=None):
boxplot = boxplot_frame_groupby
-def _is_multi_agg_with_relabel(**kwargs):
+def _is_multi_agg_with_relabel(**kwargs) -> bool:
"""
Check whether kwargs passed to .agg look like multi-agg with relabeling.
@@ -1778,7 +1780,9 @@ def _is_multi_agg_with_relabel(**kwargs):
>>> _is_multi_agg_with_relabel()
False
"""
- return all(isinstance(v, tuple) and len(v) == 2 for v in kwargs.values()) and kwargs
+ return all(isinstance(v, tuple) and len(v) == 2 for v in kwargs.values()) and (
+ len(kwargs) > 0
+ )
def _normalize_keyword_aggregation(kwargs):
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 59b118431cfc9..873a31e658625 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -756,7 +756,7 @@ def _iterate_slices(self) -> Iterable[Tuple[Optional[Hashable], Series]]:
def transform(self, func, *args, **kwargs):
raise AbstractMethodError(self)
- def _cumcount_array(self, ascending=True):
+ def _cumcount_array(self, ascending: bool = True):
"""
Parameters
----------
@@ -788,7 +788,7 @@ def _cumcount_array(self, ascending=True):
rev[sorter] = np.arange(count, dtype=np.intp)
return out[rev].astype(np.int64, copy=False)
- def _try_cast(self, result, obj, numeric_only=False):
+ def _try_cast(self, result, obj, numeric_only: bool = False):
"""
Try to cast the result to our obj original type,
we may have roundtripped through object in the mean-time.
@@ -828,7 +828,7 @@ def _try_cast(self, result, obj, numeric_only=False):
return result
- def _transform_should_cast(self, func_nm):
+ def _transform_should_cast(self, func_nm: str) -> bool:
"""
Parameters
----------
@@ -844,8 +844,8 @@ def _transform_should_cast(self, func_nm):
func_nm not in base.cython_cast_blacklist
)
- def _cython_transform(self, how, numeric_only=True, **kwargs):
- output = collections.OrderedDict()
+ def _cython_transform(self, how: str, numeric_only: bool = True, **kwargs):
+ output = collections.OrderedDict() # type: dict
for name, obj in self._iterate_slices():
is_numeric = is_numeric_dtype(obj.dtype)
if numeric_only and not is_numeric:
@@ -871,10 +871,12 @@ def _wrap_aggregated_output(self, output, names=None):
def _wrap_transformed_output(self, output, names=None):
raise AbstractMethodError(self)
- def _wrap_applied_output(self, keys, values, not_indexed_same=False):
+ def _wrap_applied_output(self, keys, values, not_indexed_same: bool = False):
raise AbstractMethodError(self)
- def _cython_agg_general(self, how, alt=None, numeric_only=True, min_count=-1):
+ def _cython_agg_general(
+ self, how: str, alt=None, numeric_only: bool = True, min_count: int = -1
+ ):
output = {}
for name, obj in self._iterate_slices():
is_numeric = is_numeric_dtype(obj.dtype)
@@ -920,7 +922,7 @@ def _python_agg_general(self, func, *args, **kwargs):
return self._wrap_aggregated_output(output)
- def _concat_objects(self, keys, values, not_indexed_same=False):
+ def _concat_objects(self, keys, values, not_indexed_same: bool = False):
from pandas.core.reshape.concat import concat
def reset_identity(values):
@@ -980,10 +982,7 @@ def reset_identity(values):
values = reset_identity(values)
result = concat(values, axis=self.axis)
- if (
- isinstance(result, Series)
- and getattr(self, "_selection_name", None) is not None
- ):
+ if isinstance(result, Series) and self._selection_name is not None:
result.name = self._selection_name
@@ -1104,7 +1103,7 @@ def result_to_bool(result: np.ndarray, inference: Type) -> np.ndarray:
@Substitution(name="groupby")
@Appender(_common_see_also)
- def any(self, skipna=True):
+ def any(self, skipna: bool = True):
"""
Return True if any value in the group is truthful, else False.
@@ -1121,7 +1120,7 @@ def any(self, skipna=True):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def all(self, skipna=True):
+ def all(self, skipna: bool = True):
"""
Return True if all values in the group are truthful, else False.
@@ -1221,7 +1220,7 @@ def median(self, **kwargs):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def std(self, ddof=1, *args, **kwargs):
+ def std(self, ddof: int = 1, *args, **kwargs):
"""
Compute standard deviation of groups, excluding missing values.
@@ -1244,7 +1243,7 @@ def std(self, ddof=1, *args, **kwargs):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def var(self, ddof=1, *args, **kwargs):
+ def var(self, ddof: int = 1, *args, **kwargs):
"""
Compute variance of groups, excluding missing values.
@@ -1272,7 +1271,7 @@ def var(self, ddof=1, *args, **kwargs):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def sem(self, ddof=1):
+ def sem(self, ddof: int = 1):
"""
Compute standard error of the mean of groups, excluding missing values.
@@ -1313,7 +1312,13 @@ def _add_numeric_operations(cls):
Add numeric operations to the GroupBy generically.
"""
- def groupby_function(name, alias, npfunc, numeric_only=True, min_count=-1):
+ def groupby_function(
+ name: str,
+ alias: str,
+ npfunc,
+ numeric_only: bool = True,
+ min_count: int = -1,
+ ):
_local_template = """
Compute %(f)s of group values.
@@ -1403,7 +1408,7 @@ def last(x):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def ohlc(self):
+ def ohlc(self) -> DataFrame:
"""
Compute sum of values, excluding missing values.
@@ -1815,7 +1820,7 @@ def nth(self, n: Union[int, List[int]], dropna: Optional[str] = None) -> DataFra
return result
- def quantile(self, q=0.5, interpolation="linear"):
+ def quantile(self, q=0.5, interpolation: str = "linear"):
"""
Return group values at the given quantile, a la numpy.percentile.
@@ -1928,7 +1933,7 @@ def post_processor(vals: np.ndarray, inference: Optional[Type]) -> np.ndarray:
return result.take(indices)
@Substitution(name="groupby")
- def ngroup(self, ascending=True):
+ def ngroup(self, ascending: bool = True):
"""
Number each group from 0 to the number of groups - 1.
@@ -1997,7 +2002,7 @@ def ngroup(self, ascending=True):
return result
@Substitution(name="groupby")
- def cumcount(self, ascending=True):
+ def cumcount(self, ascending: bool = True):
"""
Number each item in each group from 0 to the length of that group - 1.
@@ -2058,7 +2063,12 @@ def cumcount(self, ascending=True):
@Substitution(name="groupby")
@Appender(_common_see_also)
def rank(
- self, method="average", ascending=True, na_option="keep", pct=False, axis=0
+ self,
+ method: str = "average",
+ ascending: bool = True,
+ na_option: str = "keep",
+ pct: bool = False,
+ axis: int = 0,
):
"""
Provide the rank of values within each group.
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 9bbe73c1851b5..2cc0e5fde2290 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,7 +7,7 @@
"""
import collections
-from typing import List, Optional
+from typing import List, Optional, Type
import numpy as np
@@ -96,7 +96,7 @@ def __iter__(self):
return iter(self.indices)
@property
- def nkeys(self):
+ def nkeys(self) -> int:
return len(self.groupings)
def get_iterator(self, data, axis=0):
@@ -135,7 +135,7 @@ def _get_group_keys(self):
# provide "flattened" iterator for multi-group setting
return get_flattened_iterator(comp_ids, ngroups, self.levels, self.labels)
- def apply(self, f, data, axis=0):
+ def apply(self, f, data, axis: int = 0):
mutated = self.mutated
splitter = self._get_splitter(data, axis=axis)
group_keys = self._get_group_keys()
@@ -220,7 +220,7 @@ def levels(self):
def names(self):
return [ping.name for ping in self.groupings]
- def size(self):
+ def size(self) -> Series:
"""
Compute group sizes
@@ -244,7 +244,7 @@ def groups(self):
return self.axis.groupby(to_groupby)
@cache_readonly
- def is_monotonic(self):
+ def is_monotonic(self) -> bool:
# return if my group orderings are monotonic
return Index(self.group_info[0]).is_monotonic
@@ -275,7 +275,7 @@ def _get_compressed_labels(self):
return ping.labels, np.arange(len(ping.group_index))
@cache_readonly
- def ngroups(self):
+ def ngroups(self) -> int:
return len(self.result_index)
@property
@@ -345,7 +345,7 @@ def _is_builtin_func(self, arg):
"""
return SelectionMixin._builtin_table.get(arg, arg)
- def _get_cython_function(self, kind, how, values, is_numeric):
+ def _get_cython_function(self, kind: str, how: str, values, is_numeric: bool):
dtype_str = values.dtype.name
@@ -386,7 +386,9 @@ def get_func(fname):
return func
- def _cython_operation(self, kind: str, values, how, axis, min_count=-1, **kwargs):
+ def _cython_operation(
+ self, kind: str, values, how: str, axis: int, min_count: int = -1, **kwargs
+ ):
assert kind in ["transform", "aggregate"]
orig_values = values
@@ -530,16 +532,23 @@ def _cython_operation(self, kind: str, values, how, axis, min_count=-1, **kwargs
return result, names
- def aggregate(self, values, how, axis=0, min_count=-1):
+ def aggregate(self, values, how: str, axis: int = 0, min_count: int = -1):
return self._cython_operation(
"aggregate", values, how, axis, min_count=min_count
)
- def transform(self, values, how, axis=0, **kwargs):
+ def transform(self, values, how: str, axis: int = 0, **kwargs):
return self._cython_operation("transform", values, how, axis, **kwargs)
def _aggregate(
- self, result, counts, values, comp_ids, agg_func, is_datetimelike, min_count=-1
+ self,
+ result,
+ counts,
+ values,
+ comp_ids,
+ agg_func,
+ is_datetimelike: bool,
+ min_count: int = -1,
):
if values.ndim > 2:
# punting for now
@@ -554,7 +563,7 @@ def _aggregate(
return result
def _transform(
- self, result, values, comp_ids, transform_func, is_datetimelike, **kwargs
+ self, result, values, comp_ids, transform_func, is_datetimelike: bool, **kwargs
):
comp_ids, _, ngroups = self.group_info
@@ -566,7 +575,7 @@ def _transform(
return result
- def agg_series(self, obj, func):
+ def agg_series(self, obj: Series, func):
if is_extension_array_dtype(obj.dtype) and obj.dtype.kind != "M":
# _aggregate_series_fast would raise TypeError when
# calling libreduction.Slider
@@ -684,7 +693,7 @@ def groups(self):
return result
@property
- def nkeys(self):
+ def nkeys(self) -> int:
return 1
def _get_grouper(self):
@@ -771,7 +780,7 @@ def groupings(self):
for lvl, name in zip(self.levels, self.names)
]
- def agg_series(self, obj, func):
+ def agg_series(self, obj: Series, func):
dummy = obj[:0]
grouper = libreduction.SeriesBinGrouper(obj, func, self.bins, dummy)
return grouper.get_result()
@@ -863,10 +872,11 @@ def _chop(self, sdata, slice_obj: slice):
return sdata._slice(slice_obj, axis=1)
-def get_splitter(data, *args, **kwargs):
+def get_splitter(data: NDFrame, *args, **kwargs):
if isinstance(data, Series):
- klass = SeriesSplitter
- elif isinstance(data, DataFrame):
+ klass = SeriesSplitter # type: Type[DataSplitter]
+ else:
+ # i.e. DataFrame
klass = FrameSplitter
return klass(data, *args, **kwargs)
diff --git a/pandas/core/internals/concat.py b/pandas/core/internals/concat.py
index 36e1b06230d7e..4ba485c85d8ba 100644
--- a/pandas/core/internals/concat.py
+++ b/pandas/core/internals/concat.py
@@ -244,7 +244,7 @@ def concatenate_join_units(join_units, concat_axis, copy):
# Concatenating join units along ax0 is handled in _merge_blocks.
raise AssertionError("Concatenating join units along axis0")
- empty_dtype, upcasted_na = get_empty_dtype_and_na(join_units)
+ empty_dtype, upcasted_na = _get_empty_dtype_and_na(join_units)
to_concat = [
ju.get_reindexed_values(empty_dtype=empty_dtype, upcasted_na=upcasted_na)
@@ -268,7 +268,7 @@ def concatenate_join_units(join_units, concat_axis, copy):
return concat_values
-def get_empty_dtype_and_na(join_units):
+def _get_empty_dtype_and_na(join_units):
"""
Return dtype and N/A values to use when concatenating specified units.
@@ -284,7 +284,7 @@ def get_empty_dtype_and_na(join_units):
if blk is None:
return np.float64, np.nan
- if is_uniform_reindex(join_units):
+ if _is_uniform_reindex(join_units):
# FIXME: integrate property
empty_dtype = join_units[0].block.dtype
upcasted_na = join_units[0].block.fill_value
@@ -398,7 +398,7 @@ def is_uniform_join_units(join_units):
)
-def is_uniform_reindex(join_units):
+def _is_uniform_reindex(join_units) -> bool:
return (
# TODO: should this be ju.block._can_hold_na?
all(ju.block and ju.block.is_extension for ju in join_units)
@@ -406,7 +406,7 @@ def is_uniform_reindex(join_units):
)
-def trim_join_unit(join_unit, length):
+def _trim_join_unit(join_unit, length):
"""
Reduce join_unit's shape along item axis to length.
@@ -486,9 +486,9 @@ def _next_or_none(seq):
for i, (plc, unit) in enumerate(next_items):
yielded_units[i] = unit
if len(plc) > min_len:
- # trim_join_unit updates unit in place, so only
+ # _trim_join_unit updates unit in place, so only
# placement needs to be sliced to skip min_len.
- next_items[i] = (plc[min_len:], trim_join_unit(unit, min_len))
+ next_items[i] = (plc[min_len:], _trim_join_unit(unit, min_len))
else:
yielded_placement = plc
next_items[i] = _next_or_none(plans[i])
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index 39e00047ea968..772ac1cd93059 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -29,15 +29,15 @@
def concat(
objs,
axis=0,
- join="outer",
+ join: str = "outer",
join_axes=None,
- ignore_index=False,
+ ignore_index: bool = False,
keys=None,
levels=None,
names=None,
- verify_integrity=False,
+ verify_integrity: bool = False,
sort=None,
- copy=True,
+ copy: bool = True,
):
"""
Concatenate pandas objects along a particular axis with optional set logic
@@ -265,14 +265,14 @@ def __init__(
self,
objs,
axis=0,
- join="outer",
+ join: str = "outer",
join_axes=None,
keys=None,
levels=None,
names=None,
- ignore_index=False,
- verify_integrity=False,
- copy=True,
+ ignore_index: bool = False,
+ verify_integrity: bool = False,
+ copy: bool = True,
sort=False,
):
if isinstance(objs, (NDFrame, str)):
@@ -324,8 +324,8 @@ def __init__(
for obj in objs:
if not isinstance(obj, (Series, DataFrame)):
msg = (
- "cannot concatenate object of type '{}';"
- " only Series and DataFrame objs are valid".format(type(obj))
+ "cannot concatenate object of type '{typ}';"
+ " only Series and DataFrame objs are valid".format(typ=type(obj))
)
raise TypeError(msg)
@@ -580,7 +580,7 @@ def _get_concat_axis(self):
return concat_axis
- def _maybe_check_integrity(self, concat_index):
+ def _maybe_check_integrity(self, concat_index: Index):
if self.verify_integrity:
if not concat_index.is_unique:
overlap = concat_index[concat_index.duplicated()].unique()
@@ -590,11 +590,11 @@ def _maybe_check_integrity(self, concat_index):
)
-def _concat_indexes(indexes):
+def _concat_indexes(indexes) -> Index:
return indexes[0].append(indexes[1:])
-def _make_concat_multiindex(indexes, keys, levels=None, names=None):
+def _make_concat_multiindex(indexes, keys, levels=None, names=None) -> MultiIndex:
if (levels is None and isinstance(keys[0], tuple)) or (
levels is not None and len(levels) > 1
@@ -715,7 +715,6 @@ def _get_series_result_type(result, objs=None):
"""
# TODO: See if we can just inline with _constructor_expanddim
# now that sparse is removed.
- from pandas import DataFrame
# concat Series with axis 1
if isinstance(result, dict):
diff --git a/pandas/core/reshape/melt.py b/pandas/core/reshape/melt.py
index c85050bc4232b..98fee491e0a73 100644
--- a/pandas/core/reshape/melt.py
+++ b/pandas/core/reshape/melt.py
@@ -188,7 +188,7 @@ def lreshape(data, groups, dropna=True, label=None):
return data._constructor(mdata, columns=id_cols + pivot_cols)
-def wide_to_long(df, stubnames, i, j, sep="", suffix=r"\d+"):
+def wide_to_long(df, stubnames, i, j, sep: str = "", suffix: str = r"\d+"):
r"""
Wide panel to long format. Less flexible but more user-friendly than melt.
@@ -419,7 +419,7 @@ def get_var_names(df, stub, sep, suffix):
pattern = re.compile(regex)
return [col for col in df.columns if pattern.match(col)]
- def melt_stub(df, stub, i, j, value_vars, sep):
+ def melt_stub(df, stub, i, j, value_vars, sep: str):
newdf = melt(
df,
id_vars=i,
@@ -456,8 +456,8 @@ def melt_stub(df, stub, i, j, value_vars, sep):
value_vars_flattened = [e for sublist in value_vars for e in sublist]
id_vars = list(set(df.columns.tolist()).difference(value_vars_flattened))
- melted = [melt_stub(df, s, i, j, v, sep) for s, v in zip(stubnames, value_vars)]
- melted = melted[0].join(melted[1:], how="outer")
+ _melted = [melt_stub(df, s, i, j, v, sep) for s, v in zip(stubnames, value_vars)]
+ melted = _melted[0].join(_melted[1:], how="outer")
if len(i) == 1:
new = df[id_vars].set_index(i).join(melted)
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index 6ef13a62ee366..a189b2cd1ab84 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -10,7 +10,7 @@
import numpy as np
-from pandas._libs import hashtable as libhashtable, lib
+from pandas._libs import Timedelta, hashtable as libhashtable, lib
import pandas._libs.join as libjoin
from pandas.errors import MergeError
from pandas.util._decorators import Appender, Substitution
@@ -36,9 +36,10 @@
is_object_dtype,
needs_i8_conversion,
)
-from pandas.core.dtypes.missing import isnull, na_value_for_dtype
+from pandas.core.dtypes.generic import ABCDataFrame, ABCSeries
+from pandas.core.dtypes.missing import isna, na_value_for_dtype
-from pandas import Categorical, DataFrame, Index, MultiIndex, Series, Timedelta
+from pandas import Categorical, Index, MultiIndex
import pandas.core.algorithms as algos
from pandas.core.arrays.categorical import _recode_for_categories
import pandas.core.common as com
@@ -1204,7 +1205,7 @@ def _validate_specification(self):
if len(self.right_on) != len(self.left_on):
raise ValueError("len(right_on) must equal len(left_on)")
- def _validate(self, validate):
+ def _validate(self, validate: str):
# Check uniqueness of each
if self.left_index:
@@ -1300,7 +1301,12 @@ def _get_join_indexers(left_keys, right_keys, sort=False, how="inner", **kwargs)
def _restore_dropped_levels_multijoin(
- left, right, dropped_level_names, join_index, lindexer, rindexer
+ left: MultiIndex,
+ right: MultiIndex,
+ dropped_level_names,
+ join_index,
+ lindexer,
+ rindexer,
):
"""
*this is an internal non-public method*
@@ -1338,7 +1344,7 @@ def _restore_dropped_levels_multijoin(
"""
- def _convert_to_mulitindex(index):
+ def _convert_to_mulitindex(index) -> MultiIndex:
if isinstance(index, MultiIndex):
return index
else:
@@ -1686,13 +1692,13 @@ def flip(xs):
msg_missings = "Merge keys contain null values on {side} side"
if not Index(left_values).is_monotonic:
- if isnull(left_values).any():
+ if isna(left_values).any():
raise ValueError(msg_missings.format(side="left"))
else:
raise ValueError(msg_sorted.format(side="left"))
if not Index(right_values).is_monotonic:
- if isnull(right_values).any():
+ if isna(right_values).any():
raise ValueError(msg_missings.format(side="right"))
else:
raise ValueError(msg_sorted.format(side="right"))
@@ -1959,9 +1965,9 @@ def _any(x) -> bool:
def validate_operand(obj):
- if isinstance(obj, DataFrame):
+ if isinstance(obj, ABCDataFrame):
return obj
- elif isinstance(obj, Series):
+ elif isinstance(obj, ABCSeries):
if obj.name is None:
raise ValueError("Cannot merge a Series without a name")
else:
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index 7537dd0ac2065..a8dcc995e48da 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -958,7 +958,7 @@ def _get_dummies_1d(
if is_object_dtype(dtype):
raise ValueError("dtype=object is not a valid dtype for get_dummies")
- def get_empty_frame(data):
+ def get_empty_frame(data) -> DataFrame:
if isinstance(data, Series):
index = data.index
else:
diff --git a/pandas/core/window/common.py b/pandas/core/window/common.py
index 0f2920b3558c9..2ad5a1eb6faed 100644
--- a/pandas/core/window/common.py
+++ b/pandas/core/window/common.py
@@ -32,7 +32,7 @@ class _GroupByMixin(GroupByMixin):
"""
def __init__(self, obj, *args, **kwargs):
- parent = kwargs.pop("parent", None) # noqa
+ kwargs.pop("parent", None)
groupby = kwargs.pop("groupby", None)
if groupby is None:
groupby, obj = obj, obj.obj
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index 68eb1f630bfc3..f6d27de132ad9 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -1642,17 +1642,18 @@ def _get_corr(a, b):
class Rolling(_Rolling_and_Expanding):
@cache_readonly
- def is_datetimelike(self):
+ def is_datetimelike(self) -> bool:
return isinstance(
self._on, (ABCDatetimeIndex, ABCTimedeltaIndex, ABCPeriodIndex)
)
@cache_readonly
- def _on(self):
+ def _on(self) -> Index:
if self.on is None:
if self.axis == 0:
return self.obj.index
- elif self.axis == 1:
+ else:
+ # i.e. self.axis == 1
return self.obj.columns
elif isinstance(self.on, Index):
return self.on
@@ -1660,9 +1661,9 @@ def _on(self):
return Index(self.obj[self.on])
else:
raise ValueError(
- "invalid on specified as {0}, "
+ "invalid on specified as {on}, "
"must be a column (of DataFrame), an Index "
- "or None".format(self.on)
+ "or None".format(on=self.on)
)
def validate(self):
@@ -1711,7 +1712,9 @@ def _validate_monotonic(self):
formatted = self.on
if self.on is None:
formatted = "index"
- raise ValueError("{0} must be monotonic".format(formatted))
+ raise ValueError(
+ "{formatted} must be monotonic".format(formatted=formatted)
+ )
def _validate_freq(self):
"""
@@ -1723,9 +1726,9 @@ def _validate_freq(self):
return to_offset(self.window)
except (TypeError, ValueError):
raise ValueError(
- "passed window {0} is not "
+ "passed window {window} is not "
"compatible with a datetimelike "
- "index".format(self.window)
+ "index".format(window=self.window)
)
_agg_see_also_doc = dedent(
diff --git a/pandas/tseries/offsets.py b/pandas/tseries/offsets.py
index 1e3f5c1ed870e..f5e40e712642e 100644
--- a/pandas/tseries/offsets.py
+++ b/pandas/tseries/offsets.py
@@ -36,8 +36,6 @@
from pandas.core.dtypes.inference import is_list_like
-from pandas.core.tools.datetimes import to_datetime
-
__all__ = [
"Day",
"BusinessDay",
@@ -2752,8 +2750,10 @@ def generate_range(start=None, end=None, periods=None, offset=BDay()):
offset = to_offset(offset)
- start = to_datetime(start)
- end = to_datetime(end)
+ start = Timestamp(start)
+ start = start if start is not NaT else None
+ end = Timestamp(end)
+ end = end if end is not NaT else None
if start and not offset.onOffset(start):
start = offset.rollforward(start)
| diff_2d no longer needs to be in the pxi.in file, so moved it to the pyx
A couple of recently-identified bugs in the groupby code are caused by passing incorrect types, so im getting more motivated to add annotations in/around the affected code. | https://api.github.com/repos/pandas-dev/pandas/pulls/29419 | 2019-11-05T19:46:33Z | 2019-11-06T18:11:04Z | 2019-11-06T18:11:04Z | 2019-11-06T18:19:32Z |
maybe_promote: Restrict fill_value to scalar for non-object dtype | diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index 542618e332f7b..fad80d6bf5745 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -339,6 +339,11 @@ def changeit():
def maybe_promote(dtype, fill_value=np.nan):
+ if not is_scalar(fill_value) and not is_object_dtype(dtype):
+ # with object dtype there is nothing to promote, and the user can
+ # pass pretty much any weird fill_value they like
+ raise ValueError("fill_value must be a scalar")
+
# if we passed an array here, determine the fill value by dtype
if isinstance(fill_value, np.ndarray):
if issubclass(fill_value.dtype.type, (np.datetime64, np.timedelta64)):
@@ -686,7 +691,8 @@ def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
dtype : if None, then use the dtype of the values, else coerce to this type
copy : if True always make a copy even if no upcast is required
"""
- if not is_scalar(fill_value):
+ if not is_scalar(fill_value) and not is_object_dtype(values.dtype):
+ # We allow arbitrary fill values for object dtype
raise ValueError("fill_value must be a scalar")
if is_extension_type(values):
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index c792460add429..448d2faf8b85f 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -1283,10 +1283,6 @@ def diff(self, n: int, axis: int = 1) -> List["Block"]:
def shift(self, periods, axis=0, fill_value=None):
""" shift the block by periods, possibly upcast """
- if not lib.is_scalar(fill_value):
- # We could go further and require e.g. self._can_hold_element(fv)
- raise ValueError("fill_value must be a scalar")
-
# convert integer to float if necessary. need to do a lot more than
# that, handle boolean etc also
new_values, fill_value = maybe_upcast(self.values, fill_value)
diff --git a/pandas/tests/dtypes/cast/test_promote.py b/pandas/tests/dtypes/cast/test_promote.py
index 5c61574eddb50..0939e35bd64fa 100644
--- a/pandas/tests/dtypes/cast/test_promote.py
+++ b/pandas/tests/dtypes/cast/test_promote.py
@@ -19,7 +19,6 @@
is_integer_dtype,
is_object_dtype,
is_scalar,
- is_string_dtype,
is_timedelta64_dtype,
)
from pandas.core.dtypes.dtypes import DatetimeTZDtype
@@ -65,42 +64,7 @@ def any_numpy_dtype_reduced(request):
return request.param
-@pytest.fixture(
- params=[(True, None), (True, object), (False, None)],
- ids=["True-None", "True-object", "False-None"],
-)
-def box(request):
- """
- Parametrized fixture determining whether/how to transform fill_value.
-
- Since fill_value is defined on a per-test basis, the actual transformation
- (based on this fixture) is executed in _check_promote.
-
- Returns
- -------
- boxed : Boolean
- Whether fill_value should be wrapped in an np.array.
- box_dtype : dtype
- The dtype to pass to np.array([fill_value], dtype=box_dtype). If None,
- then this is passed on unmodified, and corresponds to the numpy default
- dtype for the given fill_value.
-
- * (True, None) # fill_value wrapped in array with default dtype
- * (True, object) # fill_value wrapped in array with object dtype
- * (False, None) # fill_value passed on as scalar
- """
- return request.param
-
-
-def _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar=None,
- exp_val_for_array=None,
-):
+def _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar=None):
"""
Auxiliary function to unify testing of scalar/array promotion.
@@ -109,13 +73,8 @@ def _check_promote(
dtype : dtype
The value to pass on as the first argument to maybe_promote.
fill_value : scalar
- The value to pass on as the second argument to maybe_promote, either as
- a scalar, or boxed into an array (depending on the parameter `boxed`).
- boxed : Boolean
- Parameter whether fill_value should be passed to maybe_promote
- directly, or wrapped in an array (of dtype box_dtype).
- box_dtype : dtype
- The dtype to enforce when wrapping fill_value into an np.array.
+ The value to pass on as the second argument to maybe_promote as
+ a scalar.
expected_dtype : dtype
The expected dtype returned by maybe_promote (by design this is the
same regardless of whether fill_value was passed as a scalar or in an
@@ -123,25 +82,14 @@ def _check_promote(
exp_val_for_scalar : scalar
The expected value for the (potentially upcast) fill_value returned by
maybe_promote.
- exp_val_for_array : scalar
- The expected missing value marker for the expected_dtype (which is
- returned by maybe_promote when it receives an array).
"""
assert is_scalar(fill_value)
- if boxed:
- # in this case, we pass on fill_value wrapped in an array of specified
- # box_dtype; the expected value returned from maybe_promote is the
- # missing value marker for the returned dtype.
- fill_array = np.array([fill_value], dtype=box_dtype)
- result_dtype, result_fill_value = maybe_promote(dtype, fill_array)
- expected_fill_value = exp_val_for_array
- else:
- # here, we pass on fill_value as a scalar directly; the expected value
- # returned from maybe_promote is fill_value, potentially upcast to the
- # returned dtype.
- result_dtype, result_fill_value = maybe_promote(dtype, fill_value)
- expected_fill_value = exp_val_for_scalar
+ # here, we pass on fill_value as a scalar directly; the expected value
+ # returned from maybe_promote is fill_value, potentially upcast to the
+ # returned dtype.
+ result_dtype, result_fill_value = maybe_promote(dtype, fill_value)
+ expected_fill_value = exp_val_for_scalar
assert result_dtype == expected_dtype
_assert_match(result_fill_value, expected_fill_value)
@@ -280,41 +228,19 @@ def _assert_match(result_fill_value, expected_fill_value):
("uint64", np.iinfo("int64").min - 1, "object"),
],
)
-def test_maybe_promote_int_with_int(dtype, fill_value, expected_dtype, box):
+def test_maybe_promote_int_with_int(dtype, fill_value, expected_dtype):
dtype = np.dtype(dtype)
expected_dtype = np.dtype(expected_dtype)
- boxed, box_dtype = box # read from parametrized fixture
-
- if boxed:
- if expected_dtype != object:
- pytest.xfail("falsely casts to object")
- if box_dtype is None and (
- fill_value > np.iinfo("int64").max or np.iinfo("int64").min < fill_value < 0
- ):
- pytest.xfail("falsely casts to float instead of object")
# output is not a generic int, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- # no missing value marker for integers
- exp_val_for_array = None if expected_dtype != "object" else np.nan
-
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
-
-
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, None), (False, None)])
-def test_maybe_promote_int_with_float(any_int_dtype, float_dtype, box):
+
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
+
+
+def test_maybe_promote_int_with_float(any_int_dtype, float_dtype):
dtype = np.dtype(any_int_dtype)
fill_dtype = np.dtype(float_dtype)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -323,26 +249,14 @@ def test_maybe_promote_int_with_float(any_int_dtype, float_dtype, box):
expected_dtype = np.float64
# fill_value can be different float type
exp_val_for_scalar = np.float64(fill_value)
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, None), (False, None)])
-def test_maybe_promote_float_with_int(float_dtype, any_int_dtype, box):
+def test_maybe_promote_float_with_int(float_dtype, any_int_dtype):
dtype = np.dtype(float_dtype)
fill_dtype = np.dtype(any_int_dtype)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -352,17 +266,8 @@ def test_maybe_promote_float_with_int(float_dtype, any_int_dtype, box):
expected_dtype = dtype
# output is not a generic float, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize(
@@ -390,49 +295,20 @@ def test_maybe_promote_float_with_int(float_dtype, any_int_dtype, box):
("complex128", np.finfo("float32").max * (1.1 + 1j), "complex128"),
],
)
-def test_maybe_promote_float_with_float(dtype, fill_value, expected_dtype, box):
+def test_maybe_promote_float_with_float(dtype, fill_value, expected_dtype):
dtype = np.dtype(dtype)
expected_dtype = np.dtype(expected_dtype)
- boxed, box_dtype = box # read from parametrized fixture
-
- if box_dtype == object:
- pytest.xfail("falsely upcasts to object")
- elif boxed and is_float_dtype(dtype) and is_complex_dtype(expected_dtype):
- pytest.xfail("does not upcast to complex")
- elif boxed and (dtype, expected_dtype) in [
- ("float32", "float64"),
- ("float32", "complex64"),
- ("complex64", "complex128"),
- ]:
- pytest.xfail("does not upcast correctly depending on value")
# output is not a generic float, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_bool_with_any(any_numpy_dtype_reduced, box):
+def test_maybe_promote_bool_with_any(any_numpy_dtype_reduced):
dtype = np.dtype(bool)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if boxed and fill_dtype == bool:
- pytest.xfail("falsely upcasts to object")
- if boxed and box_dtype is None and fill_dtype.kind == "M":
- pytest.xfail("wrongly casts fill_value")
- if boxed and box_dtype is None and fill_dtype.kind == "m":
- pytest.xfail("wrongly casts fill_value")
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -440,50 +316,25 @@ def test_maybe_promote_bool_with_any(any_numpy_dtype_reduced, box):
# filling bool with anything but bool casts to object
expected_dtype = np.dtype(object) if fill_dtype != bool else fill_dtype
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan if fill_dtype != bool else None
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_any_with_bool(any_numpy_dtype_reduced, box):
+def test_maybe_promote_any_with_bool(any_numpy_dtype_reduced):
dtype = np.dtype(any_numpy_dtype_reduced)
fill_value = True
- boxed, box_dtype = box # read from parametrized fixture
-
- if boxed and dtype == bool:
- pytest.xfail("falsely upcasts to object")
- if boxed and dtype not in (str, object) and box_dtype is None:
- pytest.xfail("falsely upcasts to object")
# filling anything but bool with bool casts to object
expected_dtype = np.dtype(object) if dtype != bool else dtype
# output is not a generic bool, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan if dtype != bool else None
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_bytes_with_any(bytes_dtype, any_numpy_dtype_reduced, box):
+def test_maybe_promote_bytes_with_any(bytes_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(bytes_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -491,78 +342,27 @@ def test_maybe_promote_bytes_with_any(bytes_dtype, any_numpy_dtype_reduced, box)
# we never use bytes dtype internally, always promote to object
expected_dtype = np.dtype(np.object_)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization of box to add special case for bytes
-@pytest.mark.parametrize(
- "box",
- [
- (True, None), # fill_value wrapped in array with auto-dtype (fixed len)
- (True, "bytes"), # fill_value wrapped in array with generic bytes-dtype
- (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value directly
- ],
-)
-def test_maybe_promote_any_with_bytes(any_numpy_dtype_reduced, bytes_dtype, box):
+def test_maybe_promote_any_with_bytes(any_numpy_dtype_reduced, bytes_dtype):
dtype = np.dtype(any_numpy_dtype_reduced)
- fill_dtype = np.dtype(bytes_dtype)
- boxed, box_dtype = box # read from parametrized fixture
-
- if not issubclass(dtype.type, np.bytes_):
- if (
- boxed
- and (box_dtype == "bytes" or box_dtype is None)
- and not (is_string_dtype(dtype) or dtype == bool)
- ):
- pytest.xfail("does not upcast to object")
# create array of given dtype
fill_value = b"abc"
- # special case for box_dtype (cannot use fixture in parametrization)
- box_dtype = fill_dtype if box_dtype == "bytes" else box_dtype
-
# we never use bytes dtype internally, always promote to object
expected_dtype = np.dtype(np.object_)
# output is not a generic bytes, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_datetime64_with_any(
- datetime64_dtype, any_numpy_dtype_reduced, box
-):
+def test_maybe_promote_datetime64_with_any(datetime64_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(datetime64_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_datetime64_dtype(fill_dtype):
- if box_dtype == object:
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and box_dtype is None:
- pytest.xfail("does not upcast to object")
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -572,34 +372,13 @@ def test_maybe_promote_datetime64_with_any(
expected_dtype = dtype
# for datetime dtypes, scalar values get cast to to_datetime64
exp_val_for_scalar = pd.Timestamp(fill_value).to_datetime64()
- exp_val_for_array = np.datetime64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization of box to add special case for dt_dtype
-@pytest.mark.parametrize(
- "box",
- [
- (True, None), # fill_value wrapped in array with default dtype
- # disabled due to too many xfails; see GH 23982 / 25425
- # (True, 'dt_dtype'), # fill_value in array with explicit datetime dtype
- # (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value passed on as scalar
- ],
-)
@pytest.mark.parametrize(
"fill_value",
[
@@ -611,57 +390,28 @@ def test_maybe_promote_datetime64_with_any(
ids=["pd.Timestamp", "np.datetime64", "datetime.datetime", "datetime.date"],
)
def test_maybe_promote_any_with_datetime64(
- any_numpy_dtype_reduced, datetime64_dtype, fill_value, box
+ any_numpy_dtype_reduced, datetime64_dtype, fill_value
):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_datetime64_dtype(dtype):
- if boxed and (
- box_dtype == object
- or (box_dtype is None and not is_datetime64_dtype(type(fill_value)))
- ):
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and (
- box_dtype == "dt_dtype"
- or (box_dtype is None and is_datetime64_dtype(type(fill_value)))
- ):
- pytest.xfail("mix of lack of upcasting, resp. wrong missing value")
-
- # special case for box_dtype
- box_dtype = np.dtype(datetime64_dtype) if box_dtype == "dt_dtype" else box_dtype
# filling datetime with anything but datetime casts to object
if is_datetime64_dtype(dtype):
expected_dtype = dtype
# for datetime dtypes, scalar values get cast to pd.Timestamp.value
exp_val_for_scalar = pd.Timestamp(fill_value).to_datetime64()
- exp_val_for_array = np.datetime64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, object)])
+@pytest.mark.xfail(reason="Fails to upcast to object")
def test_maybe_promote_datetimetz_with_any_numpy_dtype(
- tz_aware_fixture, any_numpy_dtype_reduced, box
+ tz_aware_fixture, any_numpy_dtype_reduced
):
dtype = DatetimeTZDtype(tz=tz_aware_fixture)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -669,34 +419,18 @@ def test_maybe_promote_datetimetz_with_any_numpy_dtype(
# filling datetimetz with any numpy dtype casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
-
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
-
-
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, None), (True, object)])
-def test_maybe_promote_datetimetz_with_datetimetz(
- tz_aware_fixture, tz_aware_fixture2, box
-):
+
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
+
+
+def test_maybe_promote_datetimetz_with_datetimetz(tz_aware_fixture, tz_aware_fixture2):
dtype = DatetimeTZDtype(tz=tz_aware_fixture)
fill_dtype = DatetimeTZDtype(tz=tz_aware_fixture2)
- boxed, box_dtype = box # read from parametrized fixture
from dateutil.tz import tzlocal
if is_platform_windows() and tz_aware_fixture2 == tzlocal():
pytest.xfail("Cannot process fill_value with this dtype, see GH 24310")
- if dtype.tz == fill_dtype.tz and boxed:
- pytest.xfail("falsely upcasts")
# create array of given dtype; casts "1" to correct dtype
fill_value = pd.Series([10 ** 9], dtype=fill_dtype)[0]
@@ -705,43 +439,22 @@ def test_maybe_promote_datetimetz_with_datetimetz(
exp_val_for_scalar = fill_value
if dtype.tz == fill_dtype.tz:
expected_dtype = dtype
- exp_val_for_array = NaT
else:
expected_dtype = np.dtype(object)
- exp_val_for_array = np.nan
+ pytest.xfail("fails to cast to object")
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize("fill_value", [None, np.nan, NaT])
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(False, None)])
-def test_maybe_promote_datetimetz_with_na(tz_aware_fixture, fill_value, box):
+def test_maybe_promote_datetimetz_with_na(tz_aware_fixture, fill_value):
dtype = DatetimeTZDtype(tz=tz_aware_fixture)
- boxed, box_dtype = box # read from parametrized fixture
expected_dtype = dtype
exp_val_for_scalar = NaT
- exp_val_for_array = NaT
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize(
@@ -755,43 +468,23 @@ def test_maybe_promote_datetimetz_with_na(tz_aware_fixture, fill_value, box):
ids=["pd.Timestamp", "np.datetime64", "datetime.datetime", "datetime.date"],
)
def test_maybe_promote_any_numpy_dtype_with_datetimetz(
- any_numpy_dtype_reduced, tz_aware_fixture, fill_value, box
+ any_numpy_dtype_reduced, tz_aware_fixture, fill_value
):
dtype = np.dtype(any_numpy_dtype_reduced)
fill_dtype = DatetimeTZDtype(tz=tz_aware_fixture)
- boxed, box_dtype = box # read from parametrized fixture
fill_value = pd.Series([fill_value], dtype=fill_dtype)[0]
# filling any numpy dtype with datetimetz casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_timedelta64_with_any(
- timedelta64_dtype, any_numpy_dtype_reduced, box
-):
+def test_maybe_promote_timedelta64_with_any(timedelta64_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(timedelta64_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_timedelta64_dtype(fill_dtype):
- if box_dtype == object:
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and box_dtype is None:
- pytest.xfail("does not upcast to object")
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -801,21 +494,11 @@ def test_maybe_promote_timedelta64_with_any(
expected_dtype = dtype
# for timedelta dtypes, scalar values get cast to pd.Timedelta.value
exp_val_for_scalar = pd.Timedelta(fill_value).to_timedelta64()
- exp_val_for_array = np.timedelta64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize(
@@ -823,62 +506,26 @@ def test_maybe_promote_timedelta64_with_any(
[pd.Timedelta(days=1), np.timedelta64(24, "h"), datetime.timedelta(1)],
ids=["pd.Timedelta", "np.timedelta64", "datetime.timedelta"],
)
-# override parametrization of box to add special case for td_dtype
-@pytest.mark.parametrize(
- "box",
- [
- (True, None), # fill_value wrapped in array with default dtype
- # disabled due to too many xfails; see GH 23982 / 25425
- # (True, 'td_dtype'), # fill_value in array with explicit timedelta dtype
- (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value passed on as scalar
- ],
-)
def test_maybe_promote_any_with_timedelta64(
- any_numpy_dtype_reduced, timedelta64_dtype, fill_value, box
+ any_numpy_dtype_reduced, timedelta64_dtype, fill_value
):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_timedelta64_dtype(dtype):
- if boxed and (
- box_dtype == object
- or (box_dtype is None and not is_timedelta64_dtype(type(fill_value)))
- ):
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and box_dtype is None and is_timedelta64_dtype(type(fill_value)):
- pytest.xfail("does not upcast correctly")
-
- # special case for box_dtype
- box_dtype = np.dtype(timedelta64_dtype) if box_dtype == "td_dtype" else box_dtype
# filling anything but timedelta with timedelta casts to object
if is_timedelta64_dtype(dtype):
expected_dtype = dtype
# for timedelta dtypes, scalar values get cast to pd.Timedelta.value
exp_val_for_scalar = pd.Timedelta(fill_value).to_timedelta64()
- exp_val_for_array = np.timedelta64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_string_with_any(string_dtype, any_numpy_dtype_reduced, box):
+def test_maybe_promote_string_with_any(string_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(string_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -886,61 +533,26 @@ def test_maybe_promote_string_with_any(string_dtype, any_numpy_dtype_reduced, bo
# filling string with anything casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization of box to add special case for str
-@pytest.mark.parametrize(
- "box",
- [
- # disabled due to too many xfails; see GH 23982 / 25425
- # (True, None), # fill_value wrapped in array with default dtype
- # (True, 'str'), # fill_value wrapped in array with generic string-dtype
- (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value passed on as scalar
- ],
-)
-def test_maybe_promote_any_with_string(any_numpy_dtype_reduced, string_dtype, box):
+def test_maybe_promote_any_with_string(any_numpy_dtype_reduced, string_dtype):
dtype = np.dtype(any_numpy_dtype_reduced)
- fill_dtype = np.dtype(string_dtype)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype
fill_value = "abc"
- # special case for box_dtype (cannot use fixture in parametrization)
- box_dtype = fill_dtype if box_dtype == "str" else box_dtype
-
# filling anything with a string casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_object_with_any(object_dtype, any_numpy_dtype_reduced, box):
+def test_maybe_promote_object_with_any(object_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(object_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -948,22 +560,12 @@ def test_maybe_promote_object_with_any(object_dtype, any_numpy_dtype_reduced, bo
# filling object with anything stays object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_any_with_object(any_numpy_dtype_reduced, object_dtype, box):
+def test_maybe_promote_any_with_object(any_numpy_dtype_reduced, object_dtype):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of object dtype from a scalar value (i.e. passing
# dtypes.common.is_scalar), which can however not be cast to int/float etc.
@@ -972,27 +574,13 @@ def test_maybe_promote_any_with_object(any_numpy_dtype_reduced, object_dtype, bo
# filling object with anything stays object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize("fill_value", [None, np.nan, NaT])
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(False, None)])
-def test_maybe_promote_any_numpy_dtype_with_na(
- any_numpy_dtype_reduced, fill_value, box
-):
+def test_maybe_promote_any_numpy_dtype_with_na(any_numpy_dtype_reduced, fill_value):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
if is_integer_dtype(dtype) and fill_value is not NaT:
# integer + other missing value (np.nan / None) casts to float
@@ -1020,24 +608,7 @@ def test_maybe_promote_any_numpy_dtype_with_na(
expected_dtype = np.dtype(object)
exp_val_for_scalar = np.nan
- # array case has same expected_dtype; but returns corresponding na-marker
- if is_integer_dtype(expected_dtype):
- # integers cannot hold NaNs; maybe_promote_with_array returns None
- exp_val_for_array = None
- elif is_datetime_or_timedelta_dtype(expected_dtype):
- exp_val_for_array = expected_dtype.type("NaT", "ns")
- else: # expected_dtype = float / complex / object
- exp_val_for_array = np.nan
-
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize("dim", [0, 2, 3])
@@ -1051,12 +622,18 @@ def test_maybe_promote_dimensions(any_numpy_dtype_reduced, dim):
for _ in range(dim):
fill_array = np.expand_dims(fill_array, 0)
- # test against 1-dimensional case
- expected_dtype, expected_missing_value = maybe_promote(
- dtype, np.array([1], dtype=dtype)
- )
+ if dtype != object:
+ # test against 1-dimensional case
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ maybe_promote(dtype, np.array([1], dtype=dtype))
- result_dtype, result_missing_value = maybe_promote(dtype, fill_array)
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ maybe_promote(dtype, fill_array)
- assert result_dtype == expected_dtype
- _assert_match(result_missing_value, expected_missing_value)
+ else:
+ expected_dtype, expected_missing_value = maybe_promote(
+ dtype, np.array([1], dtype=dtype)
+ )
+ result_dtype, result_missing_value = maybe_promote(dtype, fill_array)
+ assert result_dtype == expected_dtype
+ _assert_match(result_missing_value, expected_missing_value)
diff --git a/pandas/tests/series/test_analytics.py b/pandas/tests/series/test_analytics.py
index 457c976137c11..79eaeaf051d2e 100644
--- a/pandas/tests/series/test_analytics.py
+++ b/pandas/tests/series/test_analytics.py
@@ -1028,6 +1028,24 @@ def test_shift_int(self, datetime_series):
expected = ts.astype(float).shift(1)
tm.assert_series_equal(shifted, expected)
+ def test_shift_object_non_scalar_fill(self):
+ # shift requires scalar fill_value except for object dtype
+ ser = Series(range(3))
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ ser.shift(1, fill_value=[])
+
+ df = ser.to_frame()
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ df.shift(1, fill_value=np.arange(3))
+
+ obj_ser = ser.astype(object)
+ result = obj_ser.shift(1, fill_value={})
+ assert result[0] == {}
+
+ obj_df = obj_ser.to_frame()
+ result = obj_df.shift(1, fill_value={})
+ assert result.iloc[0, 0] == {}
+
def test_shift_categorical(self):
# GH 9416
s = pd.Series(["a", "b", "c", "d"], dtype="category")
| Partially reverts #29362 by allowing non-scalar fill_value for _object_ dtypes. i.e. in 0.25.3 `pd.Series(range(3), dtype=object).shift(1, fill_value={})` would work, #29362 broke that, and this restores it. Added `test_shift_object_non_scalar_fill` for this.
With the new restriction on `maybe_promote` in place, we can get rid of all the `box` tests and simplify test_promote a _ton_. This removes about 2500 tests. This also uncovers the fact that we were failing to run some of the non-box cases, which are now xfailed. | https://api.github.com/repos/pandas-dev/pandas/pulls/29416 | 2019-11-05T17:45:37Z | 2019-11-06T19:29:03Z | 2019-11-06T19:29:03Z | 2020-04-05T17:44:46Z |
Fixed SS03 errors | diff --git a/pandas/_libs/interval.pyx b/pandas/_libs/interval.pyx
index 1a712d0c4efa8..b13ce7c294f37 100644
--- a/pandas/_libs/interval.pyx
+++ b/pandas/_libs/interval.pyx
@@ -94,7 +94,7 @@ cdef class IntervalMixin:
@property
def mid(self):
"""
- Return the midpoint of the Interval
+ Return the midpoint of the Interval.
"""
try:
return 0.5 * (self.left + self.right)
@@ -104,7 +104,9 @@ cdef class IntervalMixin:
@property
def length(self):
- """Return the length of the Interval"""
+ """
+ Return the length of the Interval.
+ """
return self.right - self.left
@property
@@ -283,15 +285,19 @@ cdef class Interval(IntervalMixin):
_typ = "interval"
cdef readonly object left
- """Left bound for the interval"""
+ """
+ Left bound for the interval.
+ """
cdef readonly object right
- """Right bound for the interval"""
+ """
+ Right bound for the interval.
+ """
cdef readonly str closed
"""
Whether the interval is closed on the left-side, right-side, both or
- neither
+ neither.
"""
def __init__(self, left, right, str closed='right'):
diff --git a/pandas/_libs/tslibs/nattype.pyx b/pandas/_libs/tslibs/nattype.pyx
index 0bd4b78d51e4e..241aff0e19112 100644
--- a/pandas/_libs/tslibs/nattype.pyx
+++ b/pandas/_libs/tslibs/nattype.pyx
@@ -464,7 +464,7 @@ class NaTType(_NaT):
"""
Timestamp.combine(date, time)
- date, time -> datetime with same date and time fields
+ date, time -> datetime with same date and time fields.
"""
)
utcnow = _make_error_func('utcnow', # noqa:E128
@@ -503,8 +503,8 @@ class NaTType(_NaT):
"""
Timestamp.fromordinal(ordinal, freq=None, tz=None)
- passed an ordinal, translate and convert to a ts
- note: by definition there cannot be any tz info on the ordinal itself
+ Passed an ordinal, translate and convert to a ts.
+ Note: by definition there cannot be any tz info on the ordinal itself.
Parameters
----------
diff --git a/pandas/_libs/tslibs/period.pyx b/pandas/_libs/tslibs/period.pyx
index aed64aff14e0a..e297d11c5144d 100644
--- a/pandas/_libs/tslibs/period.pyx
+++ b/pandas/_libs/tslibs/period.pyx
@@ -2244,7 +2244,7 @@ cdef class _Period:
containing one or several directives. The method recognizes the same
directives as the :func:`time.strftime` function of the standard Python
distribution, as well as the specific additional directives ``%f``,
- ``%F``, ``%q``. (formatting & docs originally from scikits.timeries)
+ ``%F``, ``%q``. (formatting & docs originally from scikits.timeries).
+-----------+--------------------------------+-------+
| Directive | Meaning | Notes |
diff --git a/pandas/_libs/tslibs/timestamps.pyx b/pandas/_libs/tslibs/timestamps.pyx
index 50a71d062c63f..317dc769636fb 100644
--- a/pandas/_libs/tslibs/timestamps.pyx
+++ b/pandas/_libs/tslibs/timestamps.pyx
@@ -242,8 +242,8 @@ class Timestamp(_Timestamp):
"""
Timestamp.fromordinal(ordinal, freq=None, tz=None)
- passed an ordinal, translate and convert to a ts
- note: by definition there cannot be any tz info on the ordinal itself
+ Passed an ordinal, translate and convert to a ts.
+ Note: by definition there cannot be any tz info on the ordinal itself.
Parameters
----------
@@ -333,7 +333,7 @@ class Timestamp(_Timestamp):
"""
Timestamp.combine(date, time)
- date, time -> datetime with same date and time fields
+ date, time -> datetime with same date and time fields.
"""
return cls(datetime.combine(date, time))
@@ -601,7 +601,7 @@ timedelta}, default 'raise'
@property
def dayofweek(self):
"""
- Return day of whe week.
+ Return day of the week.
"""
return self.weekday()
| Fixed SS03 errors for:
`pandas.Timestamp.combine`; `pandas.Timestamp.fromordinal`; `pandas.Period.strftime`; `pandas.Interval.closed`; `pandas.Interval.left`; `pandas.Interval.length`; `pandas.Interval.mid`; `pandas.Interval.right`.
- [x] xref to #29315
- [ ] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
Similar to #28053 I could not find methods for `pandas.Timestamp.isoweekday` or `pandas.Timestamp.weekday`. This was not resolved in #28053. I think this may be due to docstrings in the original python datetime file.
Timestamp import _Timestamp which imports datetime. Neither Timestamp nor _Timestamp have `isoweekday` or `weekday` methods.
@datapythonista | https://api.github.com/repos/pandas-dev/pandas/pulls/29410 | 2019-11-05T08:24:51Z | 2019-11-05T15:19:21Z | 2019-11-05T15:19:21Z | 2019-11-06T06:37:57Z |
CLN: assorted cleanups | diff --git a/pandas/_libs/sparse.pyx b/pandas/_libs/sparse.pyx
index 4906e45c884e9..6abaaca010b00 100644
--- a/pandas/_libs/sparse.pyx
+++ b/pandas/_libs/sparse.pyx
@@ -597,7 +597,7 @@ cdef class BlockIndex(SparseIndex):
result = np.empty(other.npoints, dtype=np.float64)
- for 0 <= i < other.nblocks:
+ for i in range(other.nblocks):
ocur = olocs[i]
ocurlen = olens[i]
@@ -746,9 +746,6 @@ cdef class BlockUnion(BlockMerge):
nend = xend[xi]
- # print 'here xi=%d, yi=%d, mode=%d, nend=%d' % (self.xi, self.yi,
- # mode, nend)
-
# done with y?
if yi == ynblocks:
self._set_current_indices(xi + 1, yi, mode)
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 009e83b861523..1e38dde2096ba 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -1124,10 +1124,6 @@ def _decide_output_index(self, output, labels):
output_keys = labels
else:
output_keys = sorted(output)
- try:
- output_keys.sort()
- except TypeError:
- pass
if isinstance(labels, MultiIndex):
output_keys = MultiIndex.from_tuples(output_keys, names=labels.names)
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 642b1e93a057a..59b118431cfc9 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -1092,9 +1092,8 @@ def result_to_bool(result: np.ndarray, inference: Type) -> np.ndarray:
return self._get_cythonized_result(
"group_any_all",
- self.grouper,
aggregate=True,
- cython_dtype=np.uint8,
+ cython_dtype=np.dtype(np.uint8),
needs_values=True,
needs_mask=True,
pre_processing=objs_to_bool,
@@ -1305,7 +1304,7 @@ def size(self):
result = self.grouper.size()
if isinstance(self.obj, Series):
- result.name = getattr(self.obj, "name", None)
+ result.name = self.obj.name
return result
@classmethod
@@ -1586,9 +1585,8 @@ def _fill(self, direction, limit=None):
return self._get_cythonized_result(
"group_fillna_indexer",
- self.grouper,
needs_mask=True,
- cython_dtype=np.int64,
+ cython_dtype=np.dtype(np.int64),
result_is_index=True,
direction=direction,
limit=limit,
@@ -1882,11 +1880,10 @@ def post_processor(vals: np.ndarray, inference: Optional[Type]) -> np.ndarray:
if is_scalar(q):
return self._get_cythonized_result(
"group_quantile",
- self.grouper,
aggregate=True,
needs_values=True,
needs_mask=True,
- cython_dtype=np.float64,
+ cython_dtype=np.dtype(np.float64),
pre_processing=pre_processor,
post_processing=post_processor,
q=q,
@@ -1896,11 +1893,10 @@ def post_processor(vals: np.ndarray, inference: Optional[Type]) -> np.ndarray:
results = [
self._get_cythonized_result(
"group_quantile",
- self.grouper,
aggregate=True,
needs_values=True,
needs_mask=True,
- cython_dtype=np.float64,
+ cython_dtype=np.dtype(np.float64),
pre_processing=pre_processor,
post_processing=post_processor,
q=qi,
@@ -2167,14 +2163,13 @@ def cummax(self, axis=0, **kwargs):
def _get_cythonized_result(
self,
- how,
- grouper,
- aggregate=False,
- cython_dtype=None,
- needs_values=False,
- needs_mask=False,
- needs_ngroups=False,
- result_is_index=False,
+ how: str,
+ cython_dtype: np.dtype,
+ aggregate: bool = False,
+ needs_values: bool = False,
+ needs_mask: bool = False,
+ needs_ngroups: bool = False,
+ result_is_index: bool = False,
pre_processing=None,
post_processing=None,
**kwargs
@@ -2185,13 +2180,11 @@ def _get_cythonized_result(
Parameters
----------
how : str, Cythonized function name to be called
- grouper : Grouper object containing pertinent group info
+ cython_dtype : np.dtype
+ Type of the array that will be modified by the Cython call.
aggregate : bool, default False
Whether the result should be aggregated to match the number of
groups
- cython_dtype : default None
- Type of the array that will be modified by the Cython call. If
- `None`, the type will be inferred from the values of each slice
needs_values : bool, default False
Whether the values should be a part of the Cython call
signature
@@ -2234,8 +2227,10 @@ def _get_cythonized_result(
"Cannot use 'pre_processing' without specifying 'needs_values'!"
)
+ grouper = self.grouper
+
labels, _, ngroups = grouper.group_info
- output = collections.OrderedDict()
+ output = collections.OrderedDict() # type: dict
base_func = getattr(libgroupby, how)
for name, obj in self._iterate_slices():
@@ -2246,9 +2241,6 @@ def _get_cythonized_result(
else:
result_sz = len(values)
- if not cython_dtype:
- cython_dtype = values.dtype
-
result = np.zeros(result_sz, dtype=cython_dtype)
func = partial(base_func, result, labels)
inferences = None
@@ -2308,8 +2300,7 @@ def shift(self, periods=1, freq=None, axis=0, fill_value=None):
return self._get_cythonized_result(
"group_shift_indexer",
- self.grouper,
- cython_dtype=np.int64,
+ cython_dtype=np.dtype(np.int64),
needs_ngroups=True,
result_is_index=True,
periods=periods,
@@ -2478,11 +2469,13 @@ def _reindex_output(self, output):
@Appender(GroupBy.__doc__)
-def groupby(obj, by, **kwds):
+def groupby(obj: NDFrame, by, **kwds):
if isinstance(obj, Series):
from pandas.core.groupby.generic import SeriesGroupBy
- klass = SeriesGroupBy
+ klass = (
+ SeriesGroupBy
+ ) # type: Union[Type["SeriesGroupBy"], Type["DataFrameGroupBy"]]
elif isinstance(obj, DataFrame):
from pandas.core.groupby.generic import DataFrameGroupBy
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 7918e463c73ac..9bbe73c1851b5 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -592,13 +592,10 @@ def agg_series(self, obj, func):
return self._aggregate_series_pure_python(obj, func)
def _aggregate_series_fast(self, obj, func):
+ # At this point we have already checked that obj.index is not a MultiIndex
+ # and that obj is backed by an ndarray, not ExtensionArray
func = self._is_builtin_func(func)
- # TODO: pre-empt this, also pre-empt get_result raising TypError if we pass a EA
- # for EAs backed by ndarray we may have a performant workaround
- if obj.index._has_complex_internals:
- raise TypeError("Incompatible index for Cython grouper")
-
group_index, _, ngroups = self.group_info
# avoids object / Series creation overhead
@@ -842,15 +839,12 @@ def __iter__(self):
def _get_sorted_data(self):
return self.data.take(self.sort_idx, axis=self.axis)
- def _chop(self, sdata, slice_obj):
- raise AbstractMethodError(self)
-
- def apply(self, f):
+ def _chop(self, sdata, slice_obj: slice):
raise AbstractMethodError(self)
class SeriesSplitter(DataSplitter):
- def _chop(self, sdata, slice_obj):
+ def _chop(self, sdata, slice_obj: slice):
return sdata._get_values(slice_obj)
@@ -862,7 +856,7 @@ def fast_apply(self, f, names):
sdata = self._get_sorted_data()
return libreduction.apply_frame_axis0(sdata, f, names, starts, ends)
- def _chop(self, sdata, slice_obj):
+ def _chop(self, sdata, slice_obj: slice):
if self.axis == 0:
return sdata.iloc[slice_obj]
else:
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 5751ce6ea730e..c9697c530628a 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -4747,10 +4747,9 @@ def get_indexer_for(self, target, **kwargs):
def _maybe_promote(self, other):
# A hack, but it works
- from pandas import DatetimeIndex
- if self.inferred_type == "date" and isinstance(other, DatetimeIndex):
- return DatetimeIndex(self), other
+ if self.inferred_type == "date" and isinstance(other, ABCDatetimeIndex):
+ return type(other)(self), other
elif self.inferred_type == "boolean":
if not is_object_dtype(self.dtype):
return self.astype("object"), other.astype("object")
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index caaf55546189c..2e3f440573a0f 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -2179,7 +2179,9 @@ def drop(self, codes, level=None, errors="raise"):
mask = indexer == -1
if mask.any():
if errors != "ignore":
- raise ValueError("codes %s not contained in axis" % codes[mask])
+ raise ValueError(
+ "codes {codes} not contained in axis".format(codes=codes[mask])
+ )
except Exception:
pass
diff --git a/pandas/core/internals/construction.py b/pandas/core/internals/construction.py
index 4a8216cc73264..05a2803b3fc2f 100644
--- a/pandas/core/internals/construction.py
+++ b/pandas/core/internals/construction.py
@@ -167,6 +167,7 @@ def init_ndarray(values, index, columns, dtype=None, copy=False):
try:
values = values.astype(dtype)
except Exception as orig:
+ # e.g. ValueError when trying to cast object dtype to float64
raise ValueError(
"failed to cast to '{dtype}' (Exception "
"was: {orig})".format(dtype=dtype, orig=orig)
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index c11915c00c59d..39e00047ea968 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -478,7 +478,7 @@ def get_result(self):
self, method="concat"
)
- def _get_result_dim(self):
+ def _get_result_dim(self) -> int:
if self._is_series and self.axis == 1:
return 2
else:
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index 9845c570ca704..6ef13a62ee366 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -1948,13 +1948,13 @@ def _get_join_keys(llab, rlab, shape, sort):
return _get_join_keys(llab, rlab, shape, sort)
-def _should_fill(lname, rname):
+def _should_fill(lname, rname) -> bool:
if not isinstance(lname, str) or not isinstance(rname, str):
return True
return lname == rname
-def _any(x):
+def _any(x) -> bool:
return x is not None and com.any_not_none(*x)
diff --git a/pandas/core/reshape/pivot.py b/pandas/core/reshape/pivot.py
index d653dd87308cf..404292fe4d539 100644
--- a/pandas/core/reshape/pivot.py
+++ b/pandas/core/reshape/pivot.py
@@ -620,7 +620,9 @@ def _normalize(table, normalize, margins, margins_name="All"):
if (margins_name not in table.iloc[-1, :].name) | (
margins_name != table.iloc[:, -1].name
):
- raise ValueError("{} not in pivoted DataFrame".format(margins_name))
+ raise ValueError(
+ "{mname} not in pivoted DataFrame".format(mname=margins_name)
+ )
column_margin = table.iloc[:-1, -1]
index_margin = table.iloc[-1, :-1]
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index d7eae1c543804..7537dd0ac2065 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -88,7 +88,7 @@ class _Unstacker:
def __init__(
self,
- values,
+ values: np.ndarray,
index,
level=-1,
value_columns=None,
@@ -985,7 +985,7 @@ def get_empty_frame(data):
else:
# PY2 embedded unicode, gh-22084
- def _make_col_name(prefix, prefix_sep, level):
+ def _make_col_name(prefix, prefix_sep, level) -> str:
fstr = "{prefix}{prefix_sep}{level}"
return fstr.format(prefix=prefix, prefix_sep=prefix_sep, level=level)
| https://api.github.com/repos/pandas-dev/pandas/pulls/29406 | 2019-11-04T23:26:44Z | 2019-11-05T15:14:55Z | 2019-11-05T15:14:54Z | 2019-11-07T18:36:53Z | |
API: Use object dtype for empty Series | diff --git a/doc/source/user_guide/missing_data.rst b/doc/source/user_guide/missing_data.rst
index 11957cfa265f5..1cc485a229123 100644
--- a/doc/source/user_guide/missing_data.rst
+++ b/doc/source/user_guide/missing_data.rst
@@ -190,7 +190,7 @@ The sum of an empty or all-NA Series or column of a DataFrame is 0.
pd.Series([np.nan]).sum()
- pd.Series([]).sum()
+ pd.Series([], dtype="float64").sum()
The product of an empty or all-NA Series or column of a DataFrame is 1.
@@ -198,7 +198,7 @@ The product of an empty or all-NA Series or column of a DataFrame is 1.
pd.Series([np.nan]).prod()
- pd.Series([]).prod()
+ pd.Series([], dtype="float64").prod()
NA values in GroupBy
diff --git a/doc/source/user_guide/scale.rst b/doc/source/user_guide/scale.rst
index ba213864ec469..0611c6334937f 100644
--- a/doc/source/user_guide/scale.rst
+++ b/doc/source/user_guide/scale.rst
@@ -358,6 +358,7 @@ results will fit in memory, so we can safely call ``compute`` without running
out of memory. At that point it's just a regular pandas object.
.. ipython:: python
+ :okwarning:
@savefig dask_resample.png
ddf[['x', 'y']].resample("1D").mean().cumsum().compute().plot()
diff --git a/doc/source/whatsnew/v0.19.0.rst b/doc/source/whatsnew/v0.19.0.rst
index 61a65415f6b57..6f6446c3f74e1 100644
--- a/doc/source/whatsnew/v0.19.0.rst
+++ b/doc/source/whatsnew/v0.19.0.rst
@@ -707,6 +707,7 @@ A ``Series`` will now correctly promote its dtype for assignment with incompat v
.. ipython:: python
+ :okwarning:
s = pd.Series()
diff --git a/doc/source/whatsnew/v0.21.0.rst b/doc/source/whatsnew/v0.21.0.rst
index a9c7937308204..f33943e423b25 100644
--- a/doc/source/whatsnew/v0.21.0.rst
+++ b/doc/source/whatsnew/v0.21.0.rst
@@ -428,6 +428,7 @@ Note that this also changes the sum of an empty ``Series``. Previously this alwa
but for consistency with the all-NaN case, this was changed to return NaN as well:
.. ipython:: python
+ :okwarning:
pd.Series([]).sum()
diff --git a/doc/source/whatsnew/v0.22.0.rst b/doc/source/whatsnew/v0.22.0.rst
index ea36b35d61740..75949a90d09a6 100644
--- a/doc/source/whatsnew/v0.22.0.rst
+++ b/doc/source/whatsnew/v0.22.0.rst
@@ -55,6 +55,7 @@ The default sum for empty or all-*NA* ``Series`` is now ``0``.
*pandas 0.22.0*
.. ipython:: python
+ :okwarning:
pd.Series([]).sum()
pd.Series([np.nan]).sum()
@@ -67,6 +68,7 @@ pandas 0.20.3 without bottleneck, or pandas 0.21.x), use the ``min_count``
keyword.
.. ipython:: python
+ :okwarning:
pd.Series([]).sum(min_count=1)
@@ -85,6 +87,7 @@ required for a non-NA sum or product.
returning ``1`` instead.
.. ipython:: python
+ :okwarning:
pd.Series([]).prod()
pd.Series([np.nan]).prod()
diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 4ce4c12483b36..771b3e484f67c 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -366,6 +366,23 @@ When :class:`Categorical` contains ``np.nan``,
pd.Categorical([1, 2, np.nan], ordered=True).min()
+
+Default dtype of empty :class:`pandas.Series`
+^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
+
+Initialising an empty :class:`pandas.Series` without specifying a dtype will raise a `DeprecationWarning` now
+(:issue:`17261`). The default dtype will change from ``float64`` to ``object`` in future releases so that it is
+consistent with the behaviour of :class:`DataFrame` and :class:`Index`.
+
+*pandas 1.0.0*
+
+.. code-block:: ipython
+
+ In [1]: pd.Series()
+ Out[2]:
+ DeprecationWarning: The default dtype for empty Series will be 'object' instead of 'float64' in a future version. Specify a dtype explicitly to silence this warning.
+ Series([], dtype: float64)
+
.. _whatsnew_1000.api_breaking.deps:
Increased minimum versions for dependencies
@@ -494,7 +511,7 @@ Removal of prior version deprecations/changes
Previously, pandas would register converters with matplotlib as a side effect of importing pandas (:issue:`18720`).
This changed the output of plots made via matplotlib plots after pandas was imported, even if you were using
-matplotlib directly rather than rather than :meth:`~DataFrame.plot`.
+matplotlib directly rather than :meth:`~DataFrame.plot`.
To use pandas formatters with a matplotlib plot, specify
diff --git a/pandas/compat/pickle_compat.py b/pandas/compat/pickle_compat.py
index aeec5e8a0400a..7dfed94482a05 100644
--- a/pandas/compat/pickle_compat.py
+++ b/pandas/compat/pickle_compat.py
@@ -64,7 +64,7 @@ def __new__(cls) -> "Series": # type: ignore
stacklevel=6,
)
- return Series()
+ return Series(dtype=object)
class _LoadSparseFrame:
diff --git a/pandas/core/apply.py b/pandas/core/apply.py
index 8c49b2b803241..ef3d8cd53596b 100644
--- a/pandas/core/apply.py
+++ b/pandas/core/apply.py
@@ -15,6 +15,8 @@
)
from pandas.core.dtypes.generic import ABCMultiIndex, ABCSeries
+from pandas.core.construction import create_series_with_explicit_dtype
+
if TYPE_CHECKING:
from pandas import DataFrame, Series, Index
@@ -203,7 +205,7 @@ def apply_empty_result(self):
if not should_reduce:
try:
- r = self.f(Series([]))
+ r = self.f(Series([], dtype=np.float64))
except Exception:
pass
else:
@@ -211,7 +213,7 @@ def apply_empty_result(self):
if should_reduce:
if len(self.agg_axis):
- r = self.f(Series([]))
+ r = self.f(Series([], dtype=np.float64))
else:
r = np.nan
@@ -346,6 +348,7 @@ def apply_series_generator(self) -> Tuple[ResType, "Index"]:
def wrap_results(
self, results: ResType, res_index: "Index"
) -> Union["Series", "DataFrame"]:
+ from pandas import Series
# see if we can infer the results
if len(results) > 0 and 0 in results and is_sequence(results[0]):
@@ -353,7 +356,17 @@ def wrap_results(
return self.wrap_results_for_axis(results, res_index)
# dict of scalars
- result = self.obj._constructor_sliced(results)
+
+ # the default dtype of an empty Series will be `object`, but this
+ # code can be hit by df.mean() where the result should have dtype
+ # float64 even if it's an empty Series.
+ constructor_sliced = self.obj._constructor_sliced
+ if constructor_sliced is Series:
+ result = create_series_with_explicit_dtype(
+ results, dtype_if_empty=np.float64
+ )
+ else:
+ result = constructor_sliced(results)
result.index = res_index
return result
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 5e613849ba8d5..b7216d2a70ee6 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -34,6 +34,7 @@
from pandas.core.accessor import DirNamesMixin
from pandas.core.algorithms import duplicated, unique1d, value_counts
from pandas.core.arrays import ExtensionArray
+from pandas.core.construction import create_series_with_explicit_dtype
import pandas.core.nanops as nanops
_shared_docs: Dict[str, str] = dict()
@@ -1132,9 +1133,14 @@ def _map_values(self, mapper, na_action=None):
# convert to an Series for efficiency.
# we specify the keys here to handle the
# possibility that they are tuples
- from pandas import Series
- mapper = Series(mapper)
+ # The return value of mapping with an empty mapper is
+ # expected to be pd.Series(np.nan, ...). As np.nan is
+ # of dtype float64 the return value of this method should
+ # be float64 as well
+ mapper = create_series_with_explicit_dtype(
+ mapper, dtype_if_empty=np.float64
+ )
if isinstance(mapper, ABCSeries):
# Since values were input this means we came from either
diff --git a/pandas/core/construction.py b/pandas/core/construction.py
index dc537d50b3419..b03c69d865301 100644
--- a/pandas/core/construction.py
+++ b/pandas/core/construction.py
@@ -4,7 +4,7 @@
These should not depend on core.internals.
"""
-from typing import Optional, Sequence, Union, cast
+from typing import TYPE_CHECKING, Any, Optional, Sequence, Union, cast
import numpy as np
import numpy.ma as ma
@@ -44,8 +44,13 @@
)
from pandas.core.dtypes.missing import isna
+from pandas._typing import ArrayLike, Dtype
import pandas.core.common as com
+if TYPE_CHECKING:
+ from pandas.core.series import Series # noqa: F401
+ from pandas.core.index import Index # noqa: F401
+
def array(
data: Sequence[object],
@@ -565,3 +570,62 @@ def _try_cast(
else:
subarr = np.array(arr, dtype=object, copy=copy)
return subarr
+
+
+def is_empty_data(data: Any) -> bool:
+ """
+ Utility to check if a Series is instantiated with empty data,
+ which does not contain dtype information.
+
+ Parameters
+ ----------
+ data : array-like, Iterable, dict, or scalar value
+ Contains data stored in Series.
+
+ Returns
+ -------
+ bool
+ """
+ is_none = data is None
+ is_list_like_without_dtype = is_list_like(data) and not hasattr(data, "dtype")
+ is_simple_empty = is_list_like_without_dtype and not data
+ return is_none or is_simple_empty
+
+
+def create_series_with_explicit_dtype(
+ data: Any = None,
+ index: Optional[Union[ArrayLike, "Index"]] = None,
+ dtype: Optional[Dtype] = None,
+ name: Optional[str] = None,
+ copy: bool = False,
+ fastpath: bool = False,
+ dtype_if_empty: Dtype = object,
+) -> "Series":
+ """
+ Helper to pass an explicit dtype when instantiating an empty Series.
+
+ This silences a DeprecationWarning described in GitHub-17261.
+
+ Parameters
+ ----------
+ data : Mirrored from Series.__init__
+ index : Mirrored from Series.__init__
+ dtype : Mirrored from Series.__init__
+ name : Mirrored from Series.__init__
+ copy : Mirrored from Series.__init__
+ fastpath : Mirrored from Series.__init__
+ dtype_if_empty : str, numpy.dtype, or ExtensionDtype
+ This dtype will be passed explicitly if an empty Series will
+ be instantiated.
+
+ Returns
+ -------
+ Series
+ """
+ from pandas.core.series import Series
+
+ if is_empty_data(data) and dtype is None:
+ dtype = dtype_if_empty
+ return Series(
+ data=data, index=index, dtype=dtype, name=name, copy=copy, fastpath=fastpath
+ )
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 601dac3a1208b..c1616efabcdba 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -7956,7 +7956,7 @@ def quantile(self, q=0.5, axis=0, numeric_only=True, interpolation="linear"):
cols = Index([], name=self.columns.name)
if is_list_like(q):
return self._constructor([], index=q, columns=cols)
- return self._constructor_sliced([], index=cols, name=q)
+ return self._constructor_sliced([], index=cols, name=q, dtype=np.float64)
result = data._data.quantile(
qs=q, axis=1, interpolation=interpolation, transposed=is_transposed
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index 9aecd97194aad..efdcfa7edbba3 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -72,6 +72,7 @@
import pandas.core.algorithms as algos
from pandas.core.base import PandasObject, SelectionMixin
import pandas.core.common as com
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.core.index import (
Index,
InvalidIndexError,
@@ -6042,9 +6043,9 @@ def fillna(
if self.ndim == 1:
if isinstance(value, (dict, ABCSeries)):
- from pandas import Series
-
- value = Series(value)
+ value = create_series_with_explicit_dtype(
+ value, dtype_if_empty=object
+ )
elif not is_list_like(value):
pass
else:
@@ -6996,7 +6997,7 @@ def asof(self, where, subset=None):
if not is_series:
from pandas import Series
- return Series(index=self.columns, name=where)
+ return Series(index=self.columns, name=where, dtype=np.float64)
return np.nan
# It's always much faster to use a *while* loop here for
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 4726cdfb05a70..9bb0b8de9ba71 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -51,6 +51,7 @@
import pandas.core.algorithms as algorithms
from pandas.core.base import DataError, SpecificationError
import pandas.core.common as com
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.core.frame import DataFrame
from pandas.core.generic import ABCDataFrame, ABCSeries, NDFrame, _shared_docs
from pandas.core.groupby import base
@@ -259,7 +260,9 @@ def aggregate(self, func=None, *args, **kwargs):
result = self._aggregate_named(func, *args, **kwargs)
index = Index(sorted(result), name=self.grouper.names[0])
- ret = Series(result, index=index)
+ ret = create_series_with_explicit_dtype(
+ result, index=index, dtype_if_empty=object
+ )
if not self.as_index: # pragma: no cover
print("Warning, ignoring as_index=True")
@@ -407,7 +410,7 @@ def _wrap_transformed_output(
def _wrap_applied_output(self, keys, values, not_indexed_same=False):
if len(keys) == 0:
# GH #6265
- return Series([], name=self._selection_name, index=keys)
+ return Series([], name=self._selection_name, index=keys, dtype=np.float64)
def _get_index() -> Index:
if self.grouper.nkeys > 1:
@@ -493,7 +496,7 @@ def _transform_general(self, func, *args, **kwargs):
result = concat(results).sort_index()
else:
- result = Series()
+ result = Series(dtype=np.float64)
# we will only try to coerce the result type if
# we have a numeric dtype, as these are *always* user-defined funcs
@@ -1205,10 +1208,18 @@ def first_not_none(values):
if v is None:
return DataFrame()
elif isinstance(v, NDFrame):
- values = [
- x if x is not None else v._constructor(**v._construct_axes_dict())
- for x in values
- ]
+
+ # this is to silence a DeprecationWarning
+ # TODO: Remove when default dtype of empty Series is object
+ kwargs = v._construct_axes_dict()
+ if v._constructor is Series:
+ backup = create_series_with_explicit_dtype(
+ **kwargs, dtype_if_empty=object
+ )
+ else:
+ backup = v._constructor(**kwargs)
+
+ values = [x if (x is not None) else backup for x in values]
v = values[0]
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 537a960f7d463..efa3d33a2a79a 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -54,7 +54,12 @@
from pandas.core.arrays.categorical import Categorical, CategoricalAccessor
from pandas.core.arrays.sparse import SparseAccessor
import pandas.core.common as com
-from pandas.core.construction import extract_array, sanitize_array
+from pandas.core.construction import (
+ create_series_with_explicit_dtype,
+ extract_array,
+ is_empty_data,
+ sanitize_array,
+)
from pandas.core.index import (
Float64Index,
Index,
@@ -177,7 +182,6 @@ class Series(base.IndexOpsMixin, generic.NDFrame):
def __init__(
self, data=None, index=None, dtype=None, name=None, copy=False, fastpath=False
):
-
# we are called internally, so short-circuit
if fastpath:
@@ -191,6 +195,18 @@ def __init__(
else:
+ if is_empty_data(data) and dtype is None:
+ # gh-17261
+ warnings.warn(
+ "The default dtype for empty Series will be 'object' instead"
+ " of 'float64' in a future version. Specify a dtype explicitly"
+ " to silence this warning.",
+ DeprecationWarning,
+ stacklevel=2,
+ )
+ # uncomment the line below when removing the DeprecationWarning
+ # dtype = np.dtype(object)
+
if index is not None:
index = ensure_index(index)
@@ -330,7 +346,11 @@ def _init_dict(self, data, index=None, dtype=None):
keys, values = [], []
# Input is now list-like, so rely on "standard" construction:
- s = Series(values, index=keys, dtype=dtype)
+
+ # TODO: passing np.float64 to not break anything yet. See GH-17261
+ s = create_series_with_explicit_dtype(
+ values, index=keys, dtype=dtype, dtype_if_empty=np.float64
+ )
# Now we just make sure the order is respected, if any
if data and index is not None:
diff --git a/pandas/core/tools/datetimes.py b/pandas/core/tools/datetimes.py
index 453d1cca2e085..3dfafd04dff0a 100644
--- a/pandas/core/tools/datetimes.py
+++ b/pandas/core/tools/datetimes.py
@@ -145,7 +145,8 @@ def _maybe_cache(arg, format, cache, convert_listlike):
"""
from pandas import Series
- cache_array = Series()
+ cache_array = Series(dtype=object)
+
if cache:
# Perform a quicker unique check
if not should_cache(arg):
diff --git a/pandas/io/html.py b/pandas/io/html.py
index b8cb6679a9562..c629c0bab7779 100644
--- a/pandas/io/html.py
+++ b/pandas/io/html.py
@@ -14,7 +14,7 @@
from pandas.core.dtypes.common import is_list_like
-from pandas import Series
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.io.common import _is_url, _validate_header_arg, urlopen
from pandas.io.formats.printing import pprint_thing
@@ -762,7 +762,8 @@ def _parse_tfoot_tr(self, table):
def _expand_elements(body):
- lens = Series([len(elem) for elem in body])
+ data = [len(elem) for elem in body]
+ lens = create_series_with_explicit_dtype(data, dtype_if_empty=object)
lens_max = lens.max()
not_max = lens[lens != lens_max]
diff --git a/pandas/io/json/_json.py b/pandas/io/json/_json.py
index 89d5b52ffbf1e..30c1c2d59e983 100644
--- a/pandas/io/json/_json.py
+++ b/pandas/io/json/_json.py
@@ -1,4 +1,5 @@
from collections import OrderedDict
+import functools
from io import StringIO
from itertools import islice
import os
@@ -14,6 +15,7 @@
from pandas import DataFrame, MultiIndex, Series, isna, to_datetime
from pandas._typing import JSONSerializable
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.core.reshape.concat import concat
from pandas.io.common import (
@@ -1006,44 +1008,34 @@ class SeriesParser(Parser):
_split_keys = ("name", "index", "data")
def _parse_no_numpy(self):
+ data = loads(self.json, precise_float=self.precise_float)
- json = self.json
- orient = self.orient
- if orient == "split":
- decoded = {
- str(k): v
- for k, v in loads(json, precise_float=self.precise_float).items()
- }
+ if self.orient == "split":
+ decoded = {str(k): v for k, v in data.items()}
self.check_keys_split(decoded)
- self.obj = Series(dtype=None, **decoded)
+ self.obj = create_series_with_explicit_dtype(**decoded)
else:
- self.obj = Series(loads(json, precise_float=self.precise_float), dtype=None)
+ self.obj = create_series_with_explicit_dtype(data, dtype_if_empty=object)
def _parse_numpy(self):
+ load_kwargs = {
+ "dtype": None,
+ "numpy": True,
+ "precise_float": self.precise_float,
+ }
+ if self.orient in ["columns", "index"]:
+ load_kwargs["labelled"] = True
+ loads_ = functools.partial(loads, **load_kwargs)
+ data = loads_(self.json)
- json = self.json
- orient = self.orient
- if orient == "split":
- decoded = loads(
- json, dtype=None, numpy=True, precise_float=self.precise_float
- )
- decoded = {str(k): v for k, v in decoded.items()}
+ if self.orient == "split":
+ decoded = {str(k): v for k, v in data.items()}
self.check_keys_split(decoded)
- self.obj = Series(**decoded)
- elif orient == "columns" or orient == "index":
- self.obj = Series(
- *loads(
- json,
- dtype=None,
- numpy=True,
- labelled=True,
- precise_float=self.precise_float,
- )
- )
+ self.obj = create_series_with_explicit_dtype(**decoded)
+ elif self.orient in ["columns", "index"]:
+ self.obj = create_series_with_explicit_dtype(*data, dtype_if_empty=object)
else:
- self.obj = Series(
- loads(json, dtype=None, numpy=True, precise_float=self.precise_float)
- )
+ self.obj = create_series_with_explicit_dtype(data, dtype_if_empty=object)
def _try_convert_types(self):
if self.obj is None:
diff --git a/pandas/plotting/_matplotlib/boxplot.py b/pandas/plotting/_matplotlib/boxplot.py
index 7bcca659ee3f6..deeeb0016142c 100644
--- a/pandas/plotting/_matplotlib/boxplot.py
+++ b/pandas/plotting/_matplotlib/boxplot.py
@@ -114,7 +114,7 @@ def maybe_color_bp(self, bp):
def _make_plot(self):
if self.subplots:
- self._return_obj = pd.Series()
+ self._return_obj = pd.Series(dtype=object)
for i, (label, y) in enumerate(self._iter_data()):
ax = self._get_ax(i)
@@ -405,7 +405,8 @@ def boxplot_frame_groupby(
)
axes = _flatten(axes)
- ret = pd.Series()
+ ret = pd.Series(dtype=object)
+
for (key, group), ax in zip(grouped, axes):
d = group.boxplot(
ax=ax, column=column, fontsize=fontsize, rot=rot, grid=grid, **kwds
diff --git a/pandas/tests/arrays/categorical/test_algos.py b/pandas/tests/arrays/categorical/test_algos.py
index dce3c4e4d5e98..da142fa0bd63c 100644
--- a/pandas/tests/arrays/categorical/test_algos.py
+++ b/pandas/tests/arrays/categorical/test_algos.py
@@ -77,7 +77,7 @@ def test_replace(to_replace, value, result):
tm.assert_categorical_equal(cat, expected)
-@pytest.mark.parametrize("empty", [[], pd.Series(), np.array([])])
+@pytest.mark.parametrize("empty", [[], pd.Series(dtype=object), np.array([])])
def test_isin_empty(empty):
s = pd.Categorical(["a", "b"])
expected = np.array([False, False], dtype=bool)
diff --git a/pandas/tests/dtypes/test_inference.py b/pandas/tests/dtypes/test_inference.py
index 75e86a2ee7ecc..3fb4e291d7d91 100644
--- a/pandas/tests/dtypes/test_inference.py
+++ b/pandas/tests/dtypes/test_inference.py
@@ -78,7 +78,7 @@ def coerce(request):
((x for x in [1, 2]), True, "generator"),
((_ for _ in []), True, "generator-empty"),
(Series([1]), True, "Series"),
- (Series([]), True, "Series-empty"),
+ (Series([], dtype=object), True, "Series-empty"),
(Series(["a"]).str, True, "StringMethods"),
(Series([], dtype="O").str, True, "StringMethods-empty"),
(Index([1]), True, "Index"),
@@ -139,7 +139,7 @@ def __getitem__(self):
def test_is_array_like():
- assert inference.is_array_like(Series([]))
+ assert inference.is_array_like(Series([], dtype=object))
assert inference.is_array_like(Series([1, 2]))
assert inference.is_array_like(np.array(["a", "b"]))
assert inference.is_array_like(Index(["2016-01-01"]))
@@ -165,7 +165,7 @@ class DtypeList(list):
{"a": 1},
{1, "a"},
Series([1]),
- Series([]),
+ Series([], dtype=object),
Series(["a"]).str,
(x for x in range(5)),
],
@@ -1404,7 +1404,7 @@ def test_is_scalar_pandas_scalars(self):
assert is_scalar(DateOffset(days=1))
def test_is_scalar_pandas_containers(self):
- assert not is_scalar(Series())
+ assert not is_scalar(Series(dtype=object))
assert not is_scalar(Series([1]))
assert not is_scalar(DataFrame())
assert not is_scalar(DataFrame([[1]]))
diff --git a/pandas/tests/dtypes/test_missing.py b/pandas/tests/dtypes/test_missing.py
index 89474cf8fa953..5e7c6e4b48682 100644
--- a/pandas/tests/dtypes/test_missing.py
+++ b/pandas/tests/dtypes/test_missing.py
@@ -90,7 +90,8 @@ def test_isna_isnull(self, isna_f):
assert not isna_f(-np.inf)
# type
- assert not isna_f(type(pd.Series()))
+ assert not isna_f(type(pd.Series(dtype=object)))
+ assert not isna_f(type(pd.Series(dtype=np.float64)))
assert not isna_f(type(pd.DataFrame()))
# series
diff --git a/pandas/tests/frame/indexing/test_indexing.py b/pandas/tests/frame/indexing/test_indexing.py
index 9a7cd4ace686f..716be92ebca3f 100644
--- a/pandas/tests/frame/indexing/test_indexing.py
+++ b/pandas/tests/frame/indexing/test_indexing.py
@@ -2572,7 +2572,7 @@ def test_xs_corner(self):
# no columns but Index(dtype=object)
df = DataFrame(index=["a", "b", "c"])
result = df.xs("a")
- expected = Series([], name="a", index=pd.Index([], dtype=object))
+ expected = Series([], name="a", index=pd.Index([]), dtype=np.float64)
tm.assert_series_equal(result, expected)
def test_xs_duplicates(self):
diff --git a/pandas/tests/frame/test_analytics.py b/pandas/tests/frame/test_analytics.py
index 005ca8d95182e..5c14c3cd2a2b5 100644
--- a/pandas/tests/frame/test_analytics.py
+++ b/pandas/tests/frame/test_analytics.py
@@ -1067,13 +1067,13 @@ def test_mean_mixed_datetime_numeric(self, tz):
tm.assert_series_equal(result, expected)
@pytest.mark.parametrize("tz", [None, "UTC"])
- def test_mean_excludeds_datetimes(self, tz):
+ def test_mean_excludes_datetimes(self, tz):
# https://github.com/pandas-dev/pandas/issues/24752
# Our long-term desired behavior is unclear, but the behavior in
# 0.24.0rc1 was buggy.
df = pd.DataFrame({"A": [pd.Timestamp("2000", tz=tz)] * 2})
result = df.mean()
- expected = pd.Series()
+ expected = pd.Series(dtype=np.float64)
tm.assert_series_equal(result, expected)
def test_mean_mixed_string_decimal(self):
@@ -1907,7 +1907,7 @@ def test_isin(self):
expected = DataFrame([df.loc[s].isin(other) for s in df.index])
tm.assert_frame_equal(result, expected)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_isin_empty(self, empty):
# GH 16991
df = DataFrame({"A": ["a", "b", "c"], "B": ["a", "e", "f"]})
diff --git a/pandas/tests/frame/test_apply.py b/pandas/tests/frame/test_apply.py
index 26a3c738750ca..eb98bdc49f976 100644
--- a/pandas/tests/frame/test_apply.py
+++ b/pandas/tests/frame/test_apply.py
@@ -105,13 +105,15 @@ def test_apply_with_reduce_empty(self):
result = empty_frame.apply(x.append, axis=1, result_type="expand")
tm.assert_frame_equal(result, empty_frame)
result = empty_frame.apply(x.append, axis=1, result_type="reduce")
- tm.assert_series_equal(result, Series([], index=pd.Index([], dtype=object)))
+ expected = Series([], index=pd.Index([], dtype=object), dtype=np.float64)
+ tm.assert_series_equal(result, expected)
empty_with_cols = DataFrame(columns=["a", "b", "c"])
result = empty_with_cols.apply(x.append, axis=1, result_type="expand")
tm.assert_frame_equal(result, empty_with_cols)
result = empty_with_cols.apply(x.append, axis=1, result_type="reduce")
- tm.assert_series_equal(result, Series([], index=pd.Index([], dtype=object)))
+ expected = Series([], index=pd.Index([], dtype=object), dtype=np.float64)
+ tm.assert_series_equal(result, expected)
# Ensure that x.append hasn't been called
assert x == []
@@ -134,7 +136,7 @@ def test_nunique_empty(self):
tm.assert_series_equal(result, expected)
result = df.T.nunique()
- expected = Series([], index=pd.Index([]))
+ expected = Series([], index=pd.Index([]), dtype=np.float64)
tm.assert_series_equal(result, expected)
def test_apply_standard_nonunique(self):
@@ -1284,16 +1286,16 @@ def func(group_col):
_get_cython_table_params(
DataFrame(),
[
- ("sum", Series()),
- ("max", Series()),
- ("min", Series()),
+ ("sum", Series(dtype="float64")),
+ ("max", Series(dtype="float64")),
+ ("min", Series(dtype="float64")),
("all", Series(dtype=bool)),
("any", Series(dtype=bool)),
- ("mean", Series()),
- ("prod", Series()),
- ("std", Series()),
- ("var", Series()),
- ("median", Series()),
+ ("mean", Series(dtype="float64")),
+ ("prod", Series(dtype="float64")),
+ ("std", Series(dtype="float64")),
+ ("var", Series(dtype="float64")),
+ ("median", Series(dtype="float64")),
],
),
_get_cython_table_params(
diff --git a/pandas/tests/frame/test_arithmetic.py b/pandas/tests/frame/test_arithmetic.py
index 88bd5a4fedfae..f6e203afb0898 100644
--- a/pandas/tests/frame/test_arithmetic.py
+++ b/pandas/tests/frame/test_arithmetic.py
@@ -470,7 +470,7 @@ def test_arith_flex_series(self, simple_frame):
def test_arith_flex_zero_len_raises(self):
# GH 19522 passing fill_value to frame flex arith methods should
# raise even in the zero-length special cases
- ser_len0 = pd.Series([])
+ ser_len0 = pd.Series([], dtype=object)
df_len0 = pd.DataFrame(columns=["A", "B"])
df = pd.DataFrame([[1, 2], [3, 4]], columns=["A", "B"])
diff --git a/pandas/tests/frame/test_asof.py b/pandas/tests/frame/test_asof.py
index 9a7d806c79dc3..89be3779e5748 100644
--- a/pandas/tests/frame/test_asof.py
+++ b/pandas/tests/frame/test_asof.py
@@ -67,7 +67,9 @@ def test_missing(self, date_range_frame):
df = date_range_frame.iloc[:N].copy()
result = df.asof("1989-12-31")
- expected = Series(index=["A", "B"], name=Timestamp("1989-12-31"))
+ expected = Series(
+ index=["A", "B"], name=Timestamp("1989-12-31"), dtype=np.float64
+ )
tm.assert_series_equal(result, expected)
result = df.asof(to_datetime(["1989-12-31"]))
diff --git a/pandas/tests/frame/test_constructors.py b/pandas/tests/frame/test_constructors.py
index ce0ebdbe56354..08dbeb9e585f1 100644
--- a/pandas/tests/frame/test_constructors.py
+++ b/pandas/tests/frame/test_constructors.py
@@ -25,6 +25,7 @@
date_range,
isna,
)
+from pandas.core.construction import create_series_with_explicit_dtype
import pandas.util.testing as tm
MIXED_FLOAT_DTYPES = ["float16", "float32", "float64"]
@@ -1216,7 +1217,9 @@ def test_constructor_list_of_series(self):
OrderedDict([["a", 1.5], ["b", 3], ["c", 4]]),
OrderedDict([["b", 3], ["c", 4], ["d", 6]]),
]
- data = [Series(d) for d in data]
+ data = [
+ create_series_with_explicit_dtype(d, dtype_if_empty=object) for d in data
+ ]
result = DataFrame(data)
sdict = OrderedDict(zip(range(len(data)), data))
@@ -1226,7 +1229,7 @@ def test_constructor_list_of_series(self):
result2 = DataFrame(data, index=np.arange(6))
tm.assert_frame_equal(result, result2)
- result = DataFrame([Series()])
+ result = DataFrame([Series(dtype=object)])
expected = DataFrame(index=[0])
tm.assert_frame_equal(result, expected)
@@ -1450,7 +1453,7 @@ def test_constructor_Series_named(self):
DataFrame(s, columns=[1, 2])
# #2234
- a = Series([], name="x")
+ a = Series([], name="x", dtype=object)
df = DataFrame(a)
assert df.columns[0] == "x"
@@ -2356,11 +2359,11 @@ def test_from_records_series_list_dict(self):
def test_to_frame_with_falsey_names(self):
# GH 16114
- result = Series(name=0).to_frame().dtypes
- expected = Series({0: np.float64})
+ result = Series(name=0, dtype=object).to_frame().dtypes
+ expected = Series({0: object})
tm.assert_series_equal(result, expected)
- result = DataFrame(Series(name=0)).dtypes
+ result = DataFrame(Series(name=0, dtype=object)).dtypes
tm.assert_series_equal(result, expected)
@pytest.mark.parametrize("dtype", [None, "uint8", "category"])
diff --git a/pandas/tests/frame/test_dtypes.py b/pandas/tests/frame/test_dtypes.py
index 6709cdcb1eebf..d8d56e90a2f31 100644
--- a/pandas/tests/frame/test_dtypes.py
+++ b/pandas/tests/frame/test_dtypes.py
@@ -656,8 +656,8 @@ def test_astype_dict_like(self, dtype_class):
# GH 16717
# if dtypes provided is empty, the resulting DataFrame
# should be the same as the original DataFrame
- dt7 = dtype_class({})
- result = df.astype(dt7)
+ dt7 = dtype_class({}) if dtype_class is dict else dtype_class({}, dtype=object)
+ equiv = df.astype(dt7)
tm.assert_frame_equal(df, equiv)
tm.assert_frame_equal(df, original)
diff --git a/pandas/tests/frame/test_quantile.py b/pandas/tests/frame/test_quantile.py
index 78953d43677fc..5ca7dd32200ee 100644
--- a/pandas/tests/frame/test_quantile.py
+++ b/pandas/tests/frame/test_quantile.py
@@ -472,7 +472,7 @@ def test_quantile_empty_no_columns(self):
df = pd.DataFrame(pd.date_range("1/1/18", periods=5))
df.columns.name = "captain tightpants"
result = df.quantile(0.5)
- expected = pd.Series([], index=[], name=0.5)
+ expected = pd.Series([], index=[], name=0.5, dtype=np.float64)
expected.index.name = "captain tightpants"
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/frame/test_replace.py b/pandas/tests/frame/test_replace.py
index 434ea6ea7b4f0..3b01ae0c3c2e8 100644
--- a/pandas/tests/frame/test_replace.py
+++ b/pandas/tests/frame/test_replace.py
@@ -1251,7 +1251,7 @@ def test_replace_with_empty_dictlike(self, mix_abc):
# GH 15289
df = DataFrame(mix_abc)
tm.assert_frame_equal(df, df.replace({}))
- tm.assert_frame_equal(df, df.replace(Series([])))
+ tm.assert_frame_equal(df, df.replace(Series([], dtype=object)))
tm.assert_frame_equal(df, df.replace({"b": {}}))
tm.assert_frame_equal(df, df.replace(Series({"b": {}})))
diff --git a/pandas/tests/generic/test_generic.py b/pandas/tests/generic/test_generic.py
index 0912a8901dc6a..0ff9d7fcdb209 100644
--- a/pandas/tests/generic/test_generic.py
+++ b/pandas/tests/generic/test_generic.py
@@ -33,6 +33,7 @@ def _construct(self, shape, value=None, dtype=None, **kwargs):
if is_scalar(value):
if value == "empty":
arr = None
+ dtype = np.float64
# remove the info axis
kwargs.pop(self._typ._info_axis_name, None)
@@ -732,13 +733,10 @@ def test_squeeze(self):
tm.assert_series_equal(df.squeeze(), df["A"])
# don't fail with 0 length dimensions GH11229 & GH8999
- empty_series = Series([], name="five")
+ empty_series = Series([], name="five", dtype=np.float64)
empty_frame = DataFrame([empty_series])
-
- [
- tm.assert_series_equal(empty_series, higher_dim.squeeze())
- for higher_dim in [empty_series, empty_frame]
- ]
+ tm.assert_series_equal(empty_series, empty_series.squeeze())
+ tm.assert_series_equal(empty_series, empty_frame.squeeze())
# axis argument
df = tm.makeTimeDataFrame(nper=1).iloc[:, :1]
@@ -898,10 +896,10 @@ def test_equals(self):
# GH 8437
a = pd.Series([False, np.nan])
b = pd.Series([False, np.nan])
- c = pd.Series(index=range(2))
- d = pd.Series(index=range(2))
- e = pd.Series(index=range(2))
- f = pd.Series(index=range(2))
+ c = pd.Series(index=range(2), dtype=object)
+ d = c.copy()
+ e = c.copy()
+ f = c.copy()
c[:-1] = d[:-1] = e[0] = f[0] = False
assert a.equals(a)
assert a.equals(b)
@@ -940,7 +938,7 @@ def test_pipe_tuple_error(self):
@pytest.mark.parametrize("box", [pd.Series, pd.DataFrame])
def test_axis_classmethods(self, box):
- obj = box()
+ obj = box(dtype=object)
values = (
list(box._AXIS_NAMES.keys())
+ list(box._AXIS_NUMBERS.keys())
diff --git a/pandas/tests/generic/test_series.py b/pandas/tests/generic/test_series.py
index 096a5aa99bd80..aaf523956aaed 100644
--- a/pandas/tests/generic/test_series.py
+++ b/pandas/tests/generic/test_series.py
@@ -224,7 +224,7 @@ def test_to_xarray_index_types(self, index):
def test_to_xarray(self):
from xarray import DataArray
- s = Series([])
+ s = Series([], dtype=object)
s.index.name = "foo"
result = s.to_xarray()
assert len(result) == 0
diff --git a/pandas/tests/groupby/test_counting.py b/pandas/tests/groupby/test_counting.py
index 9882f12714d2d..8e9554085b9ee 100644
--- a/pandas/tests/groupby/test_counting.py
+++ b/pandas/tests/groupby/test_counting.py
@@ -20,7 +20,7 @@ def test_cumcount(self):
def test_cumcount_empty(self):
ge = DataFrame().groupby(level=0)
- se = Series().groupby(level=0)
+ se = Series(dtype=object).groupby(level=0)
# edge case, as this is usually considered float
e = Series(dtype="int64")
@@ -95,7 +95,7 @@ def test_ngroup_one_group(self):
def test_ngroup_empty(self):
ge = DataFrame().groupby(level=0)
- se = Series().groupby(level=0)
+ se = Series(dtype=object).groupby(level=0)
# edge case, as this is usually considered float
e = Series(dtype="int64")
diff --git a/pandas/tests/groupby/test_filters.py b/pandas/tests/groupby/test_filters.py
index 2ce04fc774083..b3ee12b6691d7 100644
--- a/pandas/tests/groupby/test_filters.py
+++ b/pandas/tests/groupby/test_filters.py
@@ -593,5 +593,5 @@ def test_filter_dropna_with_empty_groups():
tm.assert_series_equal(result_false, expected_false)
result_true = groupped.filter(lambda x: x.mean() > 1, dropna=True)
- expected_true = pd.Series(index=pd.Index([], dtype=int))
+ expected_true = pd.Series(index=pd.Index([], dtype=int), dtype=np.float64)
tm.assert_series_equal(result_true, expected_true)
diff --git a/pandas/tests/groupby/test_function.py b/pandas/tests/groupby/test_function.py
index c41f762e9128d..4ca23c61ba920 100644
--- a/pandas/tests/groupby/test_function.py
+++ b/pandas/tests/groupby/test_function.py
@@ -1047,7 +1047,7 @@ def test_nunique_with_object():
def test_nunique_with_empty_series():
# GH 12553
- data = pd.Series(name="name")
+ data = pd.Series(name="name", dtype=object)
result = data.groupby(level=0).nunique()
expected = pd.Series(name="name", dtype="int64")
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/groupby/test_grouping.py b/pandas/tests/groupby/test_grouping.py
index e4edc64016567..2c84c2f034fc6 100644
--- a/pandas/tests/groupby/test_grouping.py
+++ b/pandas/tests/groupby/test_grouping.py
@@ -585,9 +585,18 @@ def test_list_grouper_with_nat(self):
@pytest.mark.parametrize(
"func,expected",
[
- ("transform", pd.Series(name=2, index=pd.RangeIndex(0, 0, 1))),
- ("agg", pd.Series(name=2, index=pd.Float64Index([], name=1))),
- ("apply", pd.Series(name=2, index=pd.Float64Index([], name=1))),
+ (
+ "transform",
+ pd.Series(name=2, dtype=np.float64, index=pd.RangeIndex(0, 0, 1)),
+ ),
+ (
+ "agg",
+ pd.Series(name=2, dtype=np.float64, index=pd.Float64Index([], name=1)),
+ ),
+ (
+ "apply",
+ pd.Series(name=2, dtype=np.float64, index=pd.Float64Index([], name=1)),
+ ),
],
)
def test_evaluate_with_empty_groups(self, func, expected):
@@ -602,7 +611,7 @@ def test_evaluate_with_empty_groups(self, func, expected):
def test_groupby_empty(self):
# https://github.com/pandas-dev/pandas/issues/27190
- s = pd.Series([], name="name")
+ s = pd.Series([], name="name", dtype="float64")
gr = s.groupby([])
result = gr.mean()
@@ -731,7 +740,7 @@ def test_get_group_grouped_by_tuple(self):
def test_groupby_with_empty(self):
index = pd.DatetimeIndex(())
data = ()
- series = pd.Series(data, index)
+ series = pd.Series(data, index, dtype=object)
grouper = pd.Grouper(freq="D")
grouped = series.groupby(grouper)
assert next(iter(grouped), None) is None
diff --git a/pandas/tests/indexes/datetimelike.py b/pandas/tests/indexes/datetimelike.py
index 42244626749b9..6eedfca129856 100644
--- a/pandas/tests/indexes/datetimelike.py
+++ b/pandas/tests/indexes/datetimelike.py
@@ -72,7 +72,7 @@ def test_map_callable(self):
"mapper",
[
lambda values, index: {i: e for e, i in zip(values, index)},
- lambda values, index: pd.Series(values, index),
+ lambda values, index: pd.Series(values, index, dtype=object),
],
)
def test_map_dictlike(self, mapper):
diff --git a/pandas/tests/indexes/datetimes/test_arithmetic.py b/pandas/tests/indexes/datetimes/test_arithmetic.py
index 4851dd5a55c1e..2bcaa973acd6b 100644
--- a/pandas/tests/indexes/datetimes/test_arithmetic.py
+++ b/pandas/tests/indexes/datetimes/test_arithmetic.py
@@ -100,9 +100,9 @@ def test_dti_shift_localized(self, tzstr):
def test_dti_shift_across_dst(self):
# GH 8616
idx = date_range("2013-11-03", tz="America/Chicago", periods=7, freq="H")
- s = Series(index=idx[:-1])
+ s = Series(index=idx[:-1], dtype=object)
result = s.shift(freq="H")
- expected = Series(index=idx[1:])
+ expected = Series(index=idx[1:], dtype=object)
tm.assert_series_equal(result, expected)
@pytest.mark.parametrize(
diff --git a/pandas/tests/indexes/test_base.py b/pandas/tests/indexes/test_base.py
index c0c677b076e2c..e62d50f64d8ff 100644
--- a/pandas/tests/indexes/test_base.py
+++ b/pandas/tests/indexes/test_base.py
@@ -2001,7 +2001,7 @@ def test_isin_level_kwarg_bad_label_raises(self, label, indices):
with pytest.raises(KeyError, match=msg):
index.isin([], level=label)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_isin_empty(self, empty):
# see gh-16991
index = Index(["a", "b"])
diff --git a/pandas/tests/indexing/common.py b/pandas/tests/indexing/common.py
index db6dddfdca11b..e5b2c83f29030 100644
--- a/pandas/tests/indexing/common.py
+++ b/pandas/tests/indexing/common.py
@@ -93,7 +93,7 @@ def setup_method(self, method):
self.frame_ts_rev = DataFrame(np.random.randn(4, 4), index=dates_rev)
self.frame_empty = DataFrame()
- self.series_empty = Series()
+ self.series_empty = Series(dtype=object)
# form agglomerates
for kind in self._kinds:
diff --git a/pandas/tests/indexing/multiindex/test_loc.py b/pandas/tests/indexing/multiindex/test_loc.py
index 76425c72ce4f9..b6b9f7f205394 100644
--- a/pandas/tests/indexing/multiindex/test_loc.py
+++ b/pandas/tests/indexing/multiindex/test_loc.py
@@ -48,7 +48,9 @@ def test_loc_getitem_series(self):
empty = Series(data=[], dtype=np.float64)
expected = Series(
- [], index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64)
+ [],
+ index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64),
+ dtype=np.float64,
)
result = x.loc[empty]
tm.assert_series_equal(result, expected)
@@ -70,7 +72,9 @@ def test_loc_getitem_array(self):
# empty array:
empty = np.array([])
expected = Series(
- [], index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64)
+ [],
+ index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64),
+ dtype="float64",
)
result = x.loc[empty]
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/indexing/test_iloc.py b/pandas/tests/indexing/test_iloc.py
index e4d387fd3ac38..f9bded5b266f1 100644
--- a/pandas/tests/indexing/test_iloc.py
+++ b/pandas/tests/indexing/test_iloc.py
@@ -286,7 +286,7 @@ def test_iloc_getitem_dups(self):
def test_iloc_getitem_array(self):
# array like
- s = Series(index=range(1, 4))
+ s = Series(index=range(1, 4), dtype=object)
self.check_result(
"iloc",
s.index,
@@ -499,7 +499,7 @@ def test_iloc_getitem_frame(self):
tm.assert_frame_equal(result, expected)
# with index-like
- s = Series(index=range(1, 5))
+ s = Series(index=range(1, 5), dtype=object)
result = df.iloc[s.index]
with catch_warnings(record=True):
filterwarnings("ignore", "\\n.ix", FutureWarning)
diff --git a/pandas/tests/indexing/test_indexing.py b/pandas/tests/indexing/test_indexing.py
index 25b8713eb0307..d75afd1540f22 100644
--- a/pandas/tests/indexing/test_indexing.py
+++ b/pandas/tests/indexing/test_indexing.py
@@ -895,7 +895,7 @@ def test_range_in_series_indexing(self):
# range can cause an indexing error
# GH 11652
for x in [5, 999999, 1000000]:
- s = Series(index=range(x))
+ s = Series(index=range(x), dtype=np.float64)
s.loc[range(1)] = 42
tm.assert_series_equal(s.loc[range(1)], Series(42.0, index=[0]))
diff --git a/pandas/tests/indexing/test_loc.py b/pandas/tests/indexing/test_loc.py
index cb523efb78cf4..e5e899bfb7f0d 100644
--- a/pandas/tests/indexing/test_loc.py
+++ b/pandas/tests/indexing/test_loc.py
@@ -217,7 +217,7 @@ def test_loc_getitem_label_array_like(self):
# array like
self.check_result(
"loc",
- Series(index=[0, 2, 4]).index,
+ Series(index=[0, 2, 4], dtype=object).index,
"ix",
[0, 2, 4],
typs=["ints", "uints"],
@@ -225,7 +225,7 @@ def test_loc_getitem_label_array_like(self):
)
self.check_result(
"loc",
- Series(index=[3, 6, 9]).index,
+ Series(index=[3, 6, 9], dtype=object).index,
"ix",
[3, 6, 9],
typs=["ints", "uints"],
@@ -282,7 +282,7 @@ def test_loc_to_fail(self):
# GH 7496
# loc should not fallback
- s = Series()
+ s = Series(dtype=object)
s.loc[1] = 1
s.loc["a"] = 2
@@ -794,13 +794,13 @@ def test_setitem_new_key_tz(self):
]
expected = pd.Series(vals, index=["foo", "bar"])
- ser = pd.Series()
+ ser = pd.Series(dtype=object)
ser["foo"] = vals[0]
ser["bar"] = vals[1]
tm.assert_series_equal(ser, expected)
- ser = pd.Series()
+ ser = pd.Series(dtype=object)
ser.loc["foo"] = vals[0]
ser.loc["bar"] = vals[1]
@@ -1016,7 +1016,7 @@ def test_loc_reverse_assignment(self):
data = [1, 2, 3, 4, 5, 6] + [None] * 4
expected = Series(data, index=range(2010, 2020))
- result = pd.Series(index=range(2010, 2020))
+ result = pd.Series(index=range(2010, 2020), dtype=np.float64)
result.loc[2015:2010:-1] = [6, 5, 4, 3, 2, 1]
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/indexing/test_partial.py b/pandas/tests/indexing/test_partial.py
index aa49edd51aa39..3adc206335e6f 100644
--- a/pandas/tests/indexing/test_partial.py
+++ b/pandas/tests/indexing/test_partial.py
@@ -368,19 +368,19 @@ def test_partial_set_empty_series(self):
# GH5226
# partially set with an empty object series
- s = Series()
+ s = Series(dtype=object)
s.loc[1] = 1
tm.assert_series_equal(s, Series([1], index=[1]))
s.loc[3] = 3
tm.assert_series_equal(s, Series([1, 3], index=[1, 3]))
- s = Series()
+ s = Series(dtype=object)
s.loc[1] = 1.0
tm.assert_series_equal(s, Series([1.0], index=[1]))
s.loc[3] = 3.0
tm.assert_series_equal(s, Series([1.0, 3.0], index=[1, 3]))
- s = Series()
+ s = Series(dtype=object)
s.loc["foo"] = 1
tm.assert_series_equal(s, Series([1], index=["foo"]))
s.loc["bar"] = 3
@@ -512,11 +512,11 @@ def test_partial_set_empty_frame_row(self):
def test_partial_set_empty_frame_set_series(self):
# GH 5756
# setting with empty Series
- df = DataFrame(Series())
- tm.assert_frame_equal(df, DataFrame({0: Series()}))
+ df = DataFrame(Series(dtype=object))
+ tm.assert_frame_equal(df, DataFrame({0: Series(dtype=object)}))
- df = DataFrame(Series(name="foo"))
- tm.assert_frame_equal(df, DataFrame({"foo": Series()}))
+ df = DataFrame(Series(name="foo", dtype=object))
+ tm.assert_frame_equal(df, DataFrame({"foo": Series(dtype=object)}))
def test_partial_set_empty_frame_empty_copy_assignment(self):
# GH 5932
diff --git a/pandas/tests/io/formats/test_format.py b/pandas/tests/io/formats/test_format.py
index 004a1d184537d..e875a6f137d80 100644
--- a/pandas/tests/io/formats/test_format.py
+++ b/pandas/tests/io/formats/test_format.py
@@ -1017,7 +1017,7 @@ def test_east_asian_unicode_true(self):
def test_to_string_buffer_all_unicode(self):
buf = StringIO()
- empty = DataFrame({"c/\u03c3": Series()})
+ empty = DataFrame({"c/\u03c3": Series(dtype=object)})
nonempty = DataFrame({"c/\u03c3": Series([1, 2, 3])})
print(empty, file=buf)
@@ -2765,7 +2765,7 @@ def test_to_string_length(self):
assert res == exp
def test_to_string_na_rep(self):
- s = pd.Series(index=range(100))
+ s = pd.Series(index=range(100), dtype=np.float64)
res = s.to_string(na_rep="foo", max_rows=2)
exp = "0 foo\n ..\n99 foo"
assert res == exp
diff --git a/pandas/tests/io/json/test_pandas.py b/pandas/tests/io/json/test_pandas.py
index d31aa04b223e8..bce3d1de849aa 100644
--- a/pandas/tests/io/json/test_pandas.py
+++ b/pandas/tests/io/json/test_pandas.py
@@ -53,7 +53,7 @@ def setup(self, datapath):
self.objSeries = tm.makeObjectSeries()
self.objSeries.name = "objects"
- self.empty_series = Series([], index=[])
+ self.empty_series = Series([], index=[], dtype=np.float64)
self.empty_frame = DataFrame()
self.frame = _frame.copy()
diff --git a/pandas/tests/io/pytables/test_store.py b/pandas/tests/io/pytables/test_store.py
index d79280f9ea494..d9a76fe97f813 100644
--- a/pandas/tests/io/pytables/test_store.py
+++ b/pandas/tests/io/pytables/test_store.py
@@ -2376,8 +2376,8 @@ def test_frame(self, compression, setup_path):
@td.xfail_non_writeable
def test_empty_series_frame(self, setup_path):
- s0 = Series()
- s1 = Series(name="myseries")
+ s0 = Series(dtype=object)
+ s1 = Series(name="myseries", dtype=object)
df0 = DataFrame()
df1 = DataFrame(index=["a", "b", "c"])
df2 = DataFrame(columns=["d", "e", "f"])
diff --git a/pandas/tests/io/test_html.py b/pandas/tests/io/test_html.py
index 353946a311c1a..c34f2ebace683 100644
--- a/pandas/tests/io/test_html.py
+++ b/pandas/tests/io/test_html.py
@@ -395,8 +395,7 @@ def test_empty_tables(self):
"""
Make sure that read_html ignores empty tables.
"""
- result = self.read_html(
- """
+ html = """
<table>
<thead>
<tr>
@@ -416,8 +415,7 @@ def test_empty_tables(self):
</tbody>
</table>
"""
- )
-
+ result = self.read_html(html)
assert len(result) == 1
def test_multiple_tbody(self):
diff --git a/pandas/tests/plotting/test_misc.py b/pandas/tests/plotting/test_misc.py
index 1e59fbf928876..9e947d4ba878a 100644
--- a/pandas/tests/plotting/test_misc.py
+++ b/pandas/tests/plotting/test_misc.py
@@ -34,7 +34,7 @@ def test_get_accessor_args():
msg = "should not be called with positional arguments"
with pytest.raises(TypeError, match=msg):
- func(backend_name="", data=Series(), args=["line", None], kwargs={})
+ func(backend_name="", data=Series(dtype=object), args=["line", None], kwargs={})
x, y, kind, kwargs = func(
backend_name="",
@@ -48,7 +48,10 @@ def test_get_accessor_args():
assert kwargs == {"grid": False}
x, y, kind, kwargs = func(
- backend_name="pandas.plotting._matplotlib", data=Series(), args=[], kwargs={}
+ backend_name="pandas.plotting._matplotlib",
+ data=Series(dtype=object),
+ args=[],
+ kwargs={},
)
assert x is None
assert y is None
diff --git a/pandas/tests/reductions/test_reductions.py b/pandas/tests/reductions/test_reductions.py
index 80d148c919ab2..3f78a6ac4a778 100644
--- a/pandas/tests/reductions/test_reductions.py
+++ b/pandas/tests/reductions/test_reductions.py
@@ -79,7 +79,7 @@ def test_nanops(self):
assert pd.isna(getattr(obj, opname)())
assert pd.isna(getattr(obj, opname)(skipna=False))
- obj = klass([])
+ obj = klass([], dtype=object)
assert pd.isna(getattr(obj, opname)())
assert pd.isna(getattr(obj, opname)(skipna=False))
@@ -528,7 +528,7 @@ def test_empty(self, method, unit, use_bottleneck):
with pd.option_context("use_bottleneck", use_bottleneck):
# GH#9422 / GH#18921
# Entirely empty
- s = Series([])
+ s = Series([], dtype=object)
# NA by default
result = getattr(s, method)()
assert result == unit
@@ -900,7 +900,7 @@ def test_timedelta64_analytics(self):
@pytest.mark.parametrize(
"test_input,error_type",
[
- (pd.Series([]), ValueError),
+ (pd.Series([], dtype="float64"), ValueError),
# For strings, or any Series with dtype 'O'
(pd.Series(["foo", "bar", "baz"]), TypeError),
(pd.Series([(1,), (2,)]), TypeError),
diff --git a/pandas/tests/resample/test_base.py b/pandas/tests/resample/test_base.py
index 161581e16b6fe..622b85f2a398c 100644
--- a/pandas/tests/resample/test_base.py
+++ b/pandas/tests/resample/test_base.py
@@ -139,7 +139,7 @@ def test_resample_empty_dataframe(empty_frame, freq, resample_method):
expected = df.copy()
else:
# GH14962
- expected = Series([])
+ expected = Series([], dtype=object)
if isinstance(df.index, PeriodIndex):
expected.index = df.index.asfreq(freq=freq)
diff --git a/pandas/tests/resample/test_datetime_index.py b/pandas/tests/resample/test_datetime_index.py
index f9229e8066be4..5837d526e3978 100644
--- a/pandas/tests/resample/test_datetime_index.py
+++ b/pandas/tests/resample/test_datetime_index.py
@@ -1429,10 +1429,11 @@ def test_downsample_across_dst_weekly():
tm.assert_frame_equal(result, expected)
idx = pd.date_range("2013-04-01", "2013-05-01", tz="Europe/London", freq="H")
- s = Series(index=idx)
+ s = Series(index=idx, dtype=np.float64)
result = s.resample("W").mean()
expected = Series(
- index=pd.date_range("2013-04-07", freq="W", periods=5, tz="Europe/London")
+ index=pd.date_range("2013-04-07", freq="W", periods=5, tz="Europe/London"),
+ dtype=np.float64,
)
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/resample/test_period_index.py b/pandas/tests/resample/test_period_index.py
index 93ce7a9480b35..219491367d292 100644
--- a/pandas/tests/resample/test_period_index.py
+++ b/pandas/tests/resample/test_period_index.py
@@ -594,7 +594,7 @@ def test_resample_with_dst_time_change(self):
def test_resample_bms_2752(self):
# GH2753
- foo = Series(index=pd.bdate_range("20000101", "20000201"))
+ foo = Series(index=pd.bdate_range("20000101", "20000201"), dtype=np.float64)
res1 = foo.resample("BMS").mean()
res2 = foo.resample("BMS").mean().resample("B").mean()
assert res1.index[0] == Timestamp("20000103")
diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index 63f1ef7595f31..8ef35882dcc12 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -27,6 +27,7 @@
isna,
read_csv,
)
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.tests.extension.decimal import to_decimal
import pandas.util.testing as tm
@@ -2177,7 +2178,7 @@ def test_concat_period_other_series(self):
def test_concat_empty_series(self):
# GH 11082
s1 = pd.Series([1, 2, 3], name="x")
- s2 = pd.Series(name="y")
+ s2 = pd.Series(name="y", dtype="float64")
res = pd.concat([s1, s2], axis=1)
exp = pd.DataFrame(
{"x": [1, 2, 3], "y": [np.nan, np.nan, np.nan]},
@@ -2186,7 +2187,7 @@ def test_concat_empty_series(self):
tm.assert_frame_equal(res, exp)
s1 = pd.Series([1, 2, 3], name="x")
- s2 = pd.Series(name="y")
+ s2 = pd.Series(name="y", dtype="float64")
res = pd.concat([s1, s2], axis=0)
# name will be reset
exp = pd.Series([1, 2, 3])
@@ -2194,7 +2195,7 @@ def test_concat_empty_series(self):
# empty Series with no name
s1 = pd.Series([1, 2, 3], name="x")
- s2 = pd.Series(name=None)
+ s2 = pd.Series(name=None, dtype="float64")
res = pd.concat([s1, s2], axis=1)
exp = pd.DataFrame(
{"x": [1, 2, 3], 0: [np.nan, np.nan, np.nan]},
@@ -2209,7 +2210,9 @@ def test_concat_empty_series_timelike(self, tz, values):
# GH 18447
first = Series([], dtype="M8[ns]").dt.tz_localize(tz)
- second = Series(values)
+ dtype = None if values else np.float64
+ second = Series(values, dtype=dtype)
+
expected = DataFrame(
{
0: pd.Series([pd.NaT] * len(values), dtype="M8[ns]").dt.tz_localize(tz),
@@ -2569,7 +2572,8 @@ def test_concat_odered_dict(self):
@pytest.mark.parametrize("dt", np.sctypes["float"])
def test_concat_no_unnecessary_upcast(dt, pdt):
# GH 13247
- dims = pdt().ndim
+ dims = pdt(dtype=object).ndim
+
dfs = [
pdt(np.array([1], dtype=dt, ndmin=dims)),
pdt(np.array([np.nan], dtype=dt, ndmin=dims)),
@@ -2579,7 +2583,7 @@ def test_concat_no_unnecessary_upcast(dt, pdt):
assert x.values.dtype == dt
-@pytest.mark.parametrize("pdt", [pd.Series, pd.DataFrame])
+@pytest.mark.parametrize("pdt", [create_series_with_explicit_dtype, pd.DataFrame])
@pytest.mark.parametrize("dt", np.sctypes["int"])
def test_concat_will_upcast(dt, pdt):
with catch_warnings(record=True):
@@ -2605,7 +2609,8 @@ def test_concat_empty_and_non_empty_frame_regression():
def test_concat_empty_and_non_empty_series_regression():
# GH 18187 regression test
s1 = pd.Series([1])
- s2 = pd.Series([])
+ s2 = pd.Series([], dtype=object)
+
expected = s1
result = pd.concat([s1, s2])
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/series/indexing/test_alter_index.py b/pandas/tests/series/indexing/test_alter_index.py
index 7509d21b8832f..c47b99fa38989 100644
--- a/pandas/tests/series/indexing/test_alter_index.py
+++ b/pandas/tests/series/indexing/test_alter_index.py
@@ -230,7 +230,7 @@ def test_reindex_with_datetimes():
def test_reindex_corner(datetime_series):
# (don't forget to fix this) I think it's fixed
- empty = Series()
+ empty = Series(dtype=object)
empty.reindex(datetime_series.index, method="pad") # it works
# corner case: pad empty series
@@ -539,8 +539,9 @@ def test_drop_with_ignore_errors():
def test_drop_empty_list(index, drop_labels):
# GH 21494
expected_index = [i for i in index if i not in drop_labels]
- series = pd.Series(index=index).drop(drop_labels)
- tm.assert_series_equal(series, pd.Series(index=expected_index))
+ series = pd.Series(index=index, dtype=object).drop(drop_labels)
+ expected = pd.Series(index=expected_index, dtype=object)
+ tm.assert_series_equal(series, expected)
@pytest.mark.parametrize(
@@ -554,4 +555,5 @@ def test_drop_empty_list(index, drop_labels):
def test_drop_non_empty_list(data, index, drop_labels):
# GH 21494 and GH 16877
with pytest.raises(KeyError, match="not found in axis"):
- pd.Series(data=data, index=index).drop(drop_labels)
+ dtype = object if data is None else None
+ pd.Series(data=data, index=index, dtype=dtype).drop(drop_labels)
diff --git a/pandas/tests/series/indexing/test_datetime.py b/pandas/tests/series/indexing/test_datetime.py
index fab3310fa3dfe..83c1c0ff16f4c 100644
--- a/pandas/tests/series/indexing/test_datetime.py
+++ b/pandas/tests/series/indexing/test_datetime.py
@@ -105,7 +105,7 @@ def test_series_set_value():
dates = [datetime(2001, 1, 1), datetime(2001, 1, 2)]
index = DatetimeIndex(dates)
- s = Series()._set_value(dates[0], 1.0)
+ s = Series(dtype=object)._set_value(dates[0], 1.0)
s2 = s._set_value(dates[1], np.nan)
expected = Series([1.0, np.nan], index=index)
diff --git a/pandas/tests/series/indexing/test_indexing.py b/pandas/tests/series/indexing/test_indexing.py
index 173bc9d9d6409..5bebd480ce8d4 100644
--- a/pandas/tests/series/indexing/test_indexing.py
+++ b/pandas/tests/series/indexing/test_indexing.py
@@ -105,7 +105,9 @@ def test_getitem_get(datetime_series, string_series, object_series):
# None
# GH 5652
- for s in [Series(), Series(index=list("abc"))]:
+ s1 = Series(dtype=object)
+ s2 = Series(dtype=object, index=list("abc"))
+ for s in [s1, s2]:
result = s.get(None)
assert result is None
@@ -130,7 +132,7 @@ def test_getitem_generator(string_series):
def test_type_promotion():
# GH12599
- s = pd.Series()
+ s = pd.Series(dtype=object)
s["a"] = pd.Timestamp("2016-01-01")
s["b"] = 3.0
s["c"] = "foo"
@@ -168,7 +170,7 @@ def test_getitem_out_of_bounds(datetime_series):
datetime_series[len(datetime_series)]
# GH #917
- s = Series([])
+ s = Series([], dtype=object)
with pytest.raises(IndexError, match=msg):
s[-1]
@@ -324,12 +326,12 @@ def test_setitem(datetime_series, string_series):
# Test for issue #10193
key = pd.Timestamp("2012-01-01")
- series = pd.Series()
+ series = pd.Series(dtype=object)
series[key] = 47
expected = pd.Series(47, [key])
tm.assert_series_equal(series, expected)
- series = pd.Series([], pd.DatetimeIndex([], freq="D"))
+ series = pd.Series([], pd.DatetimeIndex([], freq="D"), dtype=object)
series[key] = 47
expected = pd.Series(47, pd.DatetimeIndex([key], freq="D"))
tm.assert_series_equal(series, expected)
@@ -637,7 +639,7 @@ def test_setitem_na():
def test_timedelta_assignment():
# GH 8209
- s = Series([])
+ s = Series([], dtype=object)
s.loc["B"] = timedelta(1)
tm.assert_series_equal(s, Series(Timedelta("1 days"), index=["B"]))
diff --git a/pandas/tests/series/indexing/test_numeric.py b/pandas/tests/series/indexing/test_numeric.py
index 426a98b00827e..a641b47f2e690 100644
--- a/pandas/tests/series/indexing/test_numeric.py
+++ b/pandas/tests/series/indexing/test_numeric.py
@@ -150,7 +150,7 @@ def test_delitem():
tm.assert_series_equal(s, expected)
# empty
- s = Series()
+ s = Series(dtype=object)
with pytest.raises(KeyError, match=r"^0$"):
del s[0]
diff --git a/pandas/tests/series/test_analytics.py b/pandas/tests/series/test_analytics.py
index fe9306a06efc7..71b4819bb4da8 100644
--- a/pandas/tests/series/test_analytics.py
+++ b/pandas/tests/series/test_analytics.py
@@ -843,7 +843,7 @@ def test_isin_with_i8(self):
result = s.isin(s[0:2])
tm.assert_series_equal(result, expected)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_isin_empty(self, empty):
# see gh-16991
s = Series(["a", "b"])
diff --git a/pandas/tests/series/test_api.py b/pandas/tests/series/test_api.py
index 8acab3fa2541d..5da0ee9b5b1c0 100644
--- a/pandas/tests/series/test_api.py
+++ b/pandas/tests/series/test_api.py
@@ -266,7 +266,7 @@ def get_dir(s):
)
def test_index_tab_completion(self, index):
# dir contains string-like values of the Index.
- s = pd.Series(index=index)
+ s = pd.Series(index=index, dtype=object)
dir_s = dir(s)
for i, x in enumerate(s.index.unique(level=0)):
if i < 100:
@@ -275,7 +275,7 @@ def test_index_tab_completion(self, index):
assert x not in dir_s
def test_not_hashable(self):
- s_empty = Series()
+ s_empty = Series(dtype=object)
s = Series([1])
msg = "'Series' objects are mutable, thus they cannot be hashed"
with pytest.raises(TypeError, match=msg):
@@ -474,10 +474,11 @@ def test_str_attribute(self):
s.str.repeat(2)
def test_empty_method(self):
- s_empty = pd.Series()
+ s_empty = pd.Series(dtype=object)
assert s_empty.empty
- for full_series in [pd.Series([1]), pd.Series(index=[1])]:
+ s2 = pd.Series(index=[1], dtype=object)
+ for full_series in [pd.Series([1]), s2]:
assert not full_series.empty
def test_tab_complete_warning(self, ip):
diff --git a/pandas/tests/series/test_apply.py b/pandas/tests/series/test_apply.py
index eb4f3273f8713..8956b8b0b2d20 100644
--- a/pandas/tests/series/test_apply.py
+++ b/pandas/tests/series/test_apply.py
@@ -37,7 +37,7 @@ def test_apply(self, datetime_series):
assert s.name == rs.name
# index but no data
- s = Series(index=[1, 2, 3])
+ s = Series(index=[1, 2, 3], dtype=np.float64)
rs = s.apply(lambda x: x)
tm.assert_series_equal(s, rs)
@@ -340,7 +340,7 @@ def test_non_callable_aggregates(self):
"series, func, expected",
chain(
_get_cython_table_params(
- Series(),
+ Series(dtype=np.float64),
[
("sum", 0),
("max", np.nan),
@@ -395,8 +395,11 @@ def test_agg_cython_table(self, series, func, expected):
"series, func, expected",
chain(
_get_cython_table_params(
- Series(),
- [("cumprod", Series([], Index([]))), ("cumsum", Series([], Index([])))],
+ Series(dtype=np.float64),
+ [
+ ("cumprod", Series([], Index([]), dtype=np.float64)),
+ ("cumsum", Series([], Index([]), dtype=np.float64)),
+ ],
),
_get_cython_table_params(
Series([np.nan, 1, 2, 3]),
diff --git a/pandas/tests/series/test_combine_concat.py b/pandas/tests/series/test_combine_concat.py
index 9d02c1bdc2d9c..c6f4ce364f328 100644
--- a/pandas/tests/series/test_combine_concat.py
+++ b/pandas/tests/series/test_combine_concat.py
@@ -107,7 +107,8 @@ def test_combine_first(self):
# corner case
s = Series([1.0, 2, 3], index=[0, 1, 2])
- result = s.combine_first(Series([], index=[]))
+ empty = Series([], index=[], dtype=object)
+ result = s.combine_first(empty)
s.index = s.index.astype("O")
tm.assert_series_equal(s, result)
diff --git a/pandas/tests/series/test_constructors.py b/pandas/tests/series/test_constructors.py
index 34b11a0d008aa..293ec9580436e 100644
--- a/pandas/tests/series/test_constructors.py
+++ b/pandas/tests/series/test_constructors.py
@@ -52,8 +52,10 @@ class TestSeriesConstructors:
],
)
def test_empty_constructor(self, constructor, check_index_type):
- expected = Series()
- result = constructor()
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ expected = Series()
+ result = constructor()
+
assert len(result.index) == 0
tm.assert_series_equal(result, expected, check_index_type=check_index_type)
@@ -76,8 +78,8 @@ def test_scalar_conversion(self):
assert int(Series([1.0])) == 1
def test_constructor(self, datetime_series):
- empty_series = Series()
-
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty_series = Series()
assert datetime_series.index.is_all_dates
# Pass in Series
@@ -94,7 +96,8 @@ def test_constructor(self, datetime_series):
assert mixed[1] is np.NaN
assert not empty_series.index.is_all_dates
- assert not Series().index.is_all_dates
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ assert not Series().index.is_all_dates
# exception raised is of type Exception
with pytest.raises(Exception, match="Data must be 1-dimensional"):
@@ -113,8 +116,9 @@ def test_constructor(self, datetime_series):
@pytest.mark.parametrize("input_class", [list, dict, OrderedDict])
def test_constructor_empty(self, input_class):
- empty = Series()
- empty2 = Series(input_class())
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty = Series()
+ empty2 = Series(input_class())
# these are Index() and RangeIndex() which don't compare type equal
# but are just .equals
@@ -132,8 +136,9 @@ def test_constructor_empty(self, input_class):
if input_class is not list:
# With index:
- empty = Series(index=range(10))
- empty2 = Series(input_class(), index=range(10))
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty = Series(index=range(10))
+ empty2 = Series(input_class(), index=range(10))
tm.assert_series_equal(empty, empty2)
# With index and dtype float64:
@@ -165,7 +170,8 @@ def test_constructor_dtype_only(self, dtype, index):
assert len(result) == 0
def test_constructor_no_data_index_order(self):
- result = pd.Series(index=["b", "a", "c"])
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ result = pd.Series(index=["b", "a", "c"])
assert result.index.tolist() == ["b", "a", "c"]
def test_constructor_no_data_string_type(self):
@@ -631,7 +637,8 @@ def test_constructor_limit_copies(self, index):
assert s._data.blocks[0].values is not index
def test_constructor_pass_none(self):
- s = Series(None, index=range(5))
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ s = Series(None, index=range(5))
assert s.dtype == np.float64
s = Series(None, index=range(5), dtype=object)
@@ -639,8 +646,9 @@ def test_constructor_pass_none(self):
# GH 7431
# inference on the index
- s = Series(index=np.array([None]))
- expected = Series(index=Index([None]))
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ s = Series(index=np.array([None]))
+ expected = Series(index=Index([None]))
tm.assert_series_equal(s, expected)
def test_constructor_pass_nan_nat(self):
@@ -1029,7 +1037,7 @@ def test_constructor_dict(self):
pidx = tm.makePeriodIndex(100)
d = {pidx[0]: 0, pidx[1]: 1}
result = Series(d, index=pidx)
- expected = Series(np.nan, pidx)
+ expected = Series(np.nan, pidx, dtype=np.float64)
expected.iloc[0] = 0
expected.iloc[1] = 1
tm.assert_series_equal(result, expected)
@@ -1135,7 +1143,7 @@ def test_fromDict(self):
def test_fromValue(self, datetime_series):
- nans = Series(np.NaN, index=datetime_series.index)
+ nans = Series(np.NaN, index=datetime_series.index, dtype=np.float64)
assert nans.dtype == np.float_
assert len(nans) == len(datetime_series)
diff --git a/pandas/tests/series/test_dtypes.py b/pandas/tests/series/test_dtypes.py
index 065be966efa49..22b00425abb6b 100644
--- a/pandas/tests/series/test_dtypes.py
+++ b/pandas/tests/series/test_dtypes.py
@@ -205,7 +205,11 @@ def test_astype_dict_like(self, dtype_class):
# GH16717
# if dtypes provided is empty, it should error
- dt5 = dtype_class({})
+ if dtype_class is Series:
+ dt5 = dtype_class({}, dtype=object)
+ else:
+ dt5 = dtype_class({})
+
with pytest.raises(KeyError, match=msg):
s.astype(dt5)
@@ -408,7 +412,8 @@ def test_astype_empty_constructor_equality(self, dtype):
"m", # Generic timestamps raise a ValueError. Already tested.
):
init_empty = Series([], dtype=dtype)
- as_type_empty = Series([]).astype(dtype)
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ as_type_empty = Series([]).astype(dtype)
tm.assert_series_equal(init_empty, as_type_empty)
def test_arg_for_errors_in_astype(self):
@@ -472,7 +477,9 @@ def test_infer_objects_series(self):
tm.assert_series_equal(actual, expected)
def test_is_homogeneous_type(self):
- assert Series()._is_homogeneous_type
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty = Series()
+ assert empty._is_homogeneous_type
assert Series([1, 2])._is_homogeneous_type
assert Series(pd.Categorical([1, 2]))._is_homogeneous_type
diff --git a/pandas/tests/series/test_duplicates.py b/pandas/tests/series/test_duplicates.py
index 0f7e3e307ed19..666354e70bdd4 100644
--- a/pandas/tests/series/test_duplicates.py
+++ b/pandas/tests/series/test_duplicates.py
@@ -2,6 +2,7 @@
import pytest
from pandas import Categorical, Series
+from pandas.core.construction import create_series_with_explicit_dtype
import pandas.util.testing as tm
@@ -70,7 +71,7 @@ def test_unique_data_ownership():
)
def test_is_unique(data, expected):
# GH11946 / GH25180
- s = Series(data)
+ s = create_series_with_explicit_dtype(data, dtype_if_empty=object)
assert s.is_unique is expected
diff --git a/pandas/tests/series/test_explode.py b/pandas/tests/series/test_explode.py
index 6262da6bdfabf..e79d3c0556cf1 100644
--- a/pandas/tests/series/test_explode.py
+++ b/pandas/tests/series/test_explode.py
@@ -29,7 +29,7 @@ def test_mixed_type():
def test_empty():
- s = pd.Series()
+ s = pd.Series(dtype=object)
result = s.explode()
expected = s.copy()
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/series/test_missing.py b/pandas/tests/series/test_missing.py
index c5ce125d10ac2..72f08876e71ae 100644
--- a/pandas/tests/series/test_missing.py
+++ b/pandas/tests/series/test_missing.py
@@ -710,7 +710,7 @@ def test_fillna(self, datetime_series):
tm.assert_series_equal(result, expected)
result = s1.fillna({})
tm.assert_series_equal(result, s1)
- result = s1.fillna(Series(()))
+ result = s1.fillna(Series((), dtype=object))
tm.assert_series_equal(result, s1)
result = s2.fillna(s1)
tm.assert_series_equal(result, s2)
@@ -834,7 +834,8 @@ def test_timedelta64_nan(self):
# tm.assert_series_equal(selector, expected)
def test_dropna_empty(self):
- s = Series([])
+ s = Series([], dtype=object)
+
assert len(s.dropna()) == 0
s.dropna(inplace=True)
assert len(s) == 0
@@ -1163,7 +1164,7 @@ def test_interpolate_corners(self, kwargs):
s = Series([np.nan, np.nan])
tm.assert_series_equal(s.interpolate(**kwargs), s)
- s = Series([]).interpolate()
+ s = Series([], dtype=object).interpolate()
tm.assert_series_equal(s.interpolate(**kwargs), s)
def test_interpolate_index_values(self):
diff --git a/pandas/tests/series/test_operators.py b/pandas/tests/series/test_operators.py
index 983560d68c28c..06fe64d69fb6b 100644
--- a/pandas/tests/series/test_operators.py
+++ b/pandas/tests/series/test_operators.py
@@ -33,7 +33,7 @@ def test_logical_operators_bool_dtype_with_empty(self):
s_tft = Series([True, False, True], index=index)
s_fff = Series([False, False, False], index=index)
- s_empty = Series([])
+ s_empty = Series([], dtype=object)
res = s_tft & s_empty
expected = s_fff
@@ -408,11 +408,13 @@ def test_logical_ops_label_based(self):
# filling
# vs empty
- result = a & Series([])
+ empty = Series([], dtype=object)
+
+ result = a & empty.copy()
expected = Series([False, False, False], list("bca"))
tm.assert_series_equal(result, expected)
- result = a | Series([])
+ result = a | empty.copy()
expected = Series([True, False, True], list("bca"))
tm.assert_series_equal(result, expected)
@@ -428,7 +430,7 @@ def test_logical_ops_label_based(self):
# identity
# we would like s[s|e] == s to hold for any e, whether empty or not
for e in [
- Series([]),
+ empty.copy(),
Series([1], ["z"]),
Series(np.nan, b.index),
Series(np.nan, a.index),
@@ -797,12 +799,12 @@ def test_ops_datetimelike_align(self):
tm.assert_series_equal(result, expected)
def test_operators_corner(self, datetime_series):
- empty = Series([], index=Index([]))
+ empty = Series([], index=Index([]), dtype=np.float64)
result = datetime_series + empty
assert np.isnan(result).all()
- result = empty + Series([], index=Index([]))
+ result = empty + empty.copy()
assert len(result) == 0
# TODO: this returned NotImplemented earlier, what to do?
diff --git a/pandas/tests/series/test_quantile.py b/pandas/tests/series/test_quantile.py
index 1a4a3f523cbbe..4eb275d63e878 100644
--- a/pandas/tests/series/test_quantile.py
+++ b/pandas/tests/series/test_quantile.py
@@ -67,7 +67,7 @@ def test_quantile_multi(self, datetime_series):
result = datetime_series.quantile([])
expected = pd.Series(
- [], name=datetime_series.name, index=Index([], dtype=float)
+ [], name=datetime_series.name, index=Index([], dtype=float), dtype="float64"
)
tm.assert_series_equal(result, expected)
@@ -104,7 +104,8 @@ def test_quantile_nan(self):
assert result == expected
# all nan/empty
- cases = [Series([]), Series([np.nan, np.nan])]
+ s1 = Series([], dtype=object)
+ cases = [s1, Series([np.nan, np.nan])]
for s in cases:
res = s.quantile(0.5)
diff --git a/pandas/tests/series/test_replace.py b/pandas/tests/series/test_replace.py
index 8018ecf03960c..4125b5816422a 100644
--- a/pandas/tests/series/test_replace.py
+++ b/pandas/tests/series/test_replace.py
@@ -245,7 +245,10 @@ def test_replace_with_empty_dictlike(self):
# GH 15289
s = pd.Series(list("abcd"))
tm.assert_series_equal(s, s.replace(dict()))
- tm.assert_series_equal(s, s.replace(pd.Series([])))
+
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty_series = pd.Series([])
+ tm.assert_series_equal(s, s.replace(empty_series))
def test_replace_string_with_number(self):
# GH 15743
diff --git a/pandas/tests/series/test_repr.py b/pandas/tests/series/test_repr.py
index f1661ad034e4c..b687179f176c3 100644
--- a/pandas/tests/series/test_repr.py
+++ b/pandas/tests/series/test_repr.py
@@ -62,7 +62,7 @@ def test_name_printing(self):
s.name = None
assert "Name:" not in repr(s)
- s = Series(index=date_range("20010101", "20020101"), name="test")
+ s = Series(index=date_range("20010101", "20020101"), name="test", dtype=object)
assert "Name: test" in repr(s)
def test_repr(self, datetime_series, string_series, object_series):
@@ -75,7 +75,7 @@ def test_repr(self, datetime_series, string_series, object_series):
str(Series(tm.randn(1000), index=np.arange(1000, 0, step=-1)))
# empty
- str(Series())
+ str(Series(dtype=object))
# with NaNs
string_series[5:7] = np.NaN
diff --git a/pandas/tests/series/test_sorting.py b/pandas/tests/series/test_sorting.py
index 8039b133cae10..fd3445e271699 100644
--- a/pandas/tests/series/test_sorting.py
+++ b/pandas/tests/series/test_sorting.py
@@ -157,8 +157,8 @@ def test_sort_index_multiindex(self, level):
def test_sort_index_kind(self):
# GH #14444 & #13589: Add support for sort algo choosing
- series = Series(index=[3, 2, 1, 4, 3])
- expected_series = Series(index=[1, 2, 3, 3, 4])
+ series = Series(index=[3, 2, 1, 4, 3], dtype=object)
+ expected_series = Series(index=[1, 2, 3, 3, 4], dtype=object)
index_sorted_series = series.sort_index(kind="mergesort")
tm.assert_series_equal(expected_series, index_sorted_series)
@@ -170,13 +170,14 @@ def test_sort_index_kind(self):
tm.assert_series_equal(expected_series, index_sorted_series)
def test_sort_index_na_position(self):
- series = Series(index=[3, 2, 1, 4, 3, np.nan])
+ series = Series(index=[3, 2, 1, 4, 3, np.nan], dtype=object)
+ expected_series_first = Series(index=[np.nan, 1, 2, 3, 3, 4], dtype=object)
- expected_series_first = Series(index=[np.nan, 1, 2, 3, 3, 4])
index_sorted_series = series.sort_index(na_position="first")
tm.assert_series_equal(expected_series_first, index_sorted_series)
- expected_series_last = Series(index=[1, 2, 3, 3, 4, np.nan])
+ expected_series_last = Series(index=[1, 2, 3, 3, 4, np.nan], dtype=object)
+
index_sorted_series = series.sort_index(na_position="last")
tm.assert_series_equal(expected_series_last, index_sorted_series)
diff --git a/pandas/tests/series/test_subclass.py b/pandas/tests/series/test_subclass.py
index 6b82f890e974b..5e2d23a70e5be 100644
--- a/pandas/tests/series/test_subclass.py
+++ b/pandas/tests/series/test_subclass.py
@@ -32,4 +32,6 @@ def test_subclass_unstack(self):
tm.assert_frame_equal(res, exp)
def test_subclass_empty_repr(self):
- assert "SubclassedSeries" in repr(tm.SubclassedSeries())
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ sub_series = tm.SubclassedSeries()
+ assert "SubclassedSeries" in repr(sub_series)
diff --git a/pandas/tests/series/test_timeseries.py b/pandas/tests/series/test_timeseries.py
index 1587ae5eb7d07..6d00b9f2b09df 100644
--- a/pandas/tests/series/test_timeseries.py
+++ b/pandas/tests/series/test_timeseries.py
@@ -346,10 +346,9 @@ def test_asfreq(self):
def test_asfreq_datetimeindex_empty_series(self):
# GH 14320
- expected = Series(index=pd.DatetimeIndex(["2016-09-29 11:00"])).asfreq("H")
- result = Series(index=pd.DatetimeIndex(["2016-09-29 11:00"]), data=[3]).asfreq(
- "H"
- )
+ index = pd.DatetimeIndex(["2016-09-29 11:00"])
+ expected = Series(index=index, dtype=object).asfreq("H")
+ result = Series([3], index=index.copy()).asfreq("H")
tm.assert_index_equal(expected.index, result.index)
def test_pct_change(self, datetime_series):
@@ -410,7 +409,7 @@ def test_pct_change_periods_freq(
)
tm.assert_series_equal(rs_freq, rs_periods)
- empty_ts = Series(index=datetime_series.index)
+ empty_ts = Series(index=datetime_series.index, dtype=object)
rs_freq = empty_ts.pct_change(freq=freq, fill_method=fill_method, limit=limit)
rs_periods = empty_ts.pct_change(periods, fill_method=fill_method, limit=limit)
tm.assert_series_equal(rs_freq, rs_periods)
@@ -457,12 +456,12 @@ def test_first_last_valid(self, datetime_series):
assert ts.last_valid_index() is None
assert ts.first_valid_index() is None
- ser = Series([], index=[])
+ ser = Series([], index=[], dtype=object)
assert ser.last_valid_index() is None
assert ser.first_valid_index() is None
# GH12800
- empty = Series()
+ empty = Series(dtype=object)
assert empty.last_valid_index() is None
assert empty.first_valid_index() is None
diff --git a/pandas/tests/series/test_timezones.py b/pandas/tests/series/test_timezones.py
index c03101265f7e7..5e255e7cd5dcd 100644
--- a/pandas/tests/series/test_timezones.py
+++ b/pandas/tests/series/test_timezones.py
@@ -89,7 +89,7 @@ def test_series_tz_localize_nonexistent(self, tz, method, exp):
@pytest.mark.parametrize("tzstr", ["US/Eastern", "dateutil/US/Eastern"])
def test_series_tz_localize_empty(self, tzstr):
# GH#2248
- ser = Series()
+ ser = Series(dtype=object)
ser2 = ser.tz_localize("utc")
assert ser2.index.tz == pytz.utc
diff --git a/pandas/tests/test_algos.py b/pandas/tests/test_algos.py
index 02b50d84c6eca..e0e4beffe113a 100644
--- a/pandas/tests/test_algos.py
+++ b/pandas/tests/test_algos.py
@@ -812,7 +812,7 @@ def test_no_cast(self):
result = algos.isin(comps, values)
tm.assert_numpy_array_equal(expected, result)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_empty(self, empty):
# see gh-16991
vals = Index(["a", "b"])
diff --git a/pandas/tests/test_base.py b/pandas/tests/test_base.py
index d515a015cdbec..5c9a119400319 100644
--- a/pandas/tests/test_base.py
+++ b/pandas/tests/test_base.py
@@ -589,7 +589,7 @@ def test_value_counts_bins(self, index_or_series):
tm.assert_numpy_array_equal(s.unique(), exp)
assert s.nunique() == 3
- s = klass({})
+ s = klass({}) if klass is dict else klass({}, dtype=object)
expected = Series([], dtype=np.int64)
tm.assert_series_equal(s.value_counts(), expected, check_index_type=False)
# returned dtype differs depending on original
diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 44829423be1bb..204cdee2d9e1f 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -1538,7 +1538,7 @@ def test_frame_dict_constructor_empty_series(self):
s2 = Series(
[1, 2, 3, 4], index=MultiIndex.from_tuples([(1, 2), (1, 3), (3, 2), (3, 4)])
)
- s3 = Series()
+ s3 = Series(dtype=object)
# it works!
DataFrame({"foo": s1, "bar": s2, "baz": s3})
diff --git a/pandas/tests/test_register_accessor.py b/pandas/tests/test_register_accessor.py
index 97086f8ab1e85..6b40ff8b3fa1e 100644
--- a/pandas/tests/test_register_accessor.py
+++ b/pandas/tests/test_register_accessor.py
@@ -45,7 +45,8 @@ def test_register(obj, registrar):
with ensure_removed(obj, "mine"):
before = set(dir(obj))
registrar("mine")(MyAccessor)
- assert obj([]).mine.prop == "item"
+ o = obj([]) if obj is not pd.Series else obj([], dtype=object)
+ assert o.mine.prop == "item"
after = set(dir(obj))
assert (before ^ after) == {"mine"}
assert "mine" in obj._accessors
@@ -88,4 +89,4 @@ def __init__(self, data):
raise AttributeError("whoops")
with pytest.raises(AttributeError, match="whoops"):
- pd.Series([]).bad
+ pd.Series([], dtype=object).bad
diff --git a/pandas/tests/util/test_hashing.py b/pandas/tests/util/test_hashing.py
index df3c7fe9c9936..ebbdbd6c29842 100644
--- a/pandas/tests/util/test_hashing.py
+++ b/pandas/tests/util/test_hashing.py
@@ -207,7 +207,7 @@ def test_multiindex_objects():
Series(["a", np.nan, "c"]),
Series(["a", None, "c"]),
Series([True, False, True]),
- Series(),
+ Series(dtype=object),
Index([1, 2, 3]),
Index([True, False, True]),
DataFrame({"x": ["a", "b", "c"], "y": [1, 2, 3]}),
diff --git a/pandas/tests/window/test_moments.py b/pandas/tests/window/test_moments.py
index f1c89d3c6c1b4..2c65c9e2ac82c 100644
--- a/pandas/tests/window/test_moments.py
+++ b/pandas/tests/window/test_moments.py
@@ -108,7 +108,7 @@ def test_cmov_window_corner(self):
assert np.isnan(result).all()
# empty
- vals = pd.Series([])
+ vals = pd.Series([], dtype=object)
result = vals.rolling(5, center=True, win_type="boxcar").mean()
assert len(result) == 0
@@ -674,7 +674,7 @@ def f(x):
self._check_moment_func(np.mean, name="apply", func=f, raw=raw)
- expected = Series([])
+ expected = Series([], dtype="float64")
result = expected.rolling(10).apply(lambda x: x.mean(), raw=raw)
tm.assert_series_equal(result, expected)
@@ -1193,8 +1193,10 @@ def _check_ew(self, name=None, preserve_nan=False):
assert not result[11:].isna().any()
# check series of length 0
- result = getattr(Series().ewm(com=50, min_periods=min_periods), name)()
- tm.assert_series_equal(result, Series())
+ result = getattr(
+ Series(dtype=object).ewm(com=50, min_periods=min_periods), name
+ )()
+ tm.assert_series_equal(result, Series(dtype="float64"))
# check series of length 1
result = getattr(Series([1.0]).ewm(50, min_periods=min_periods), name)()
@@ -1214,7 +1216,7 @@ def _check_ew(self, name=None, preserve_nan=False):
def _create_consistency_data():
def create_series():
return [
- Series(),
+ Series(dtype=object),
Series([np.nan]),
Series([np.nan, np.nan]),
Series([3.0]),
@@ -1989,8 +1991,9 @@ def func(A, B, com, **kwargs):
assert not np.isnan(result.values[11:]).any()
# check series of length 0
- result = func(Series([]), Series([]), 50, min_periods=min_periods)
- tm.assert_series_equal(result, Series([]))
+ empty = Series([], dtype=np.float64)
+ result = func(empty, empty, 50, min_periods=min_periods)
+ tm.assert_series_equal(result, empty)
# check series of length 1
result = func(Series([1.0]), Series([1.0]), 50, min_periods=min_periods)
@@ -2190,7 +2193,7 @@ def test_rolling_functions_window_non_shrinkage_binary(self):
def test_moment_functions_zero_length(self):
# GH 8056
- s = Series()
+ s = Series(dtype=np.float64)
s_expected = s
df1 = DataFrame()
df1_expected = df1
@@ -2409,7 +2412,7 @@ def expanding_mean(x, min_periods=1):
# here to make this pass
self._check_expanding(expanding_mean, np.mean, preserve_nan=False)
- ser = Series([])
+ ser = Series([], dtype=np.float64)
tm.assert_series_equal(ser, ser.expanding().apply(lambda x: x.mean(), raw=raw))
# GH 8080
| - [x] closes #17261
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29405 | 2019-11-04T23:20:57Z | 2019-12-05T19:00:16Z | 2019-12-05T19:00:15Z | 2019-12-05T22:29:50Z |
BUG: GH29310 HDF file compression not working | diff --git a/pandas/_testing.py b/pandas/_testing.py
index 631d550c60534..55b0c8eda8e11 100644
--- a/pandas/_testing.py
+++ b/pandas/_testing.py
@@ -842,7 +842,7 @@ def assert_categorical_equal(
if check_category_order:
assert_index_equal(left.categories, right.categories, obj=f"{obj}.categories")
assert_numpy_array_equal(
- left.codes, right.codes, check_dtype=check_dtype, obj=f"{obj}.codes",
+ left.codes, right.codes, check_dtype=check_dtype, obj=f"{obj}.codes"
)
else:
assert_index_equal(
@@ -982,7 +982,7 @@ def _raise(left, right, err_msg):
if err_msg is None:
if left.shape != right.shape:
raise_assert_detail(
- obj, f"{obj} shapes are different", left.shape, right.shape,
+ obj, f"{obj} shapes are different", left.shape, right.shape
)
diff = 0
@@ -1326,7 +1326,7 @@ def assert_frame_equal(
# shape comparison
if left.shape != right.shape:
raise_assert_detail(
- obj, f"{obj} shape mismatch", f"{repr(left.shape)}", f"{repr(right.shape)}",
+ obj, f"{obj} shape mismatch", f"{repr(left.shape)}", f"{repr(right.shape)}"
)
if check_like:
diff --git a/pandas/core/arrays/masked.py b/pandas/core/arrays/masked.py
index 47605413ff1a6..f70292e98806d 100644
--- a/pandas/core/arrays/masked.py
+++ b/pandas/core/arrays/masked.py
@@ -50,9 +50,7 @@ def __iter__(self):
def __len__(self) -> int:
return len(self._data)
- def to_numpy(
- self, dtype=None, copy=False, na_value: "Scalar" = lib.no_default,
- ):
+ def to_numpy(self, dtype=None, copy=False, na_value: "Scalar" = lib.no_default):
"""
Convert to a NumPy Array.
diff --git a/pandas/core/arrays/period.py b/pandas/core/arrays/period.py
index d9b53aa4a867c..56e6c3aee0d2d 100644
--- a/pandas/core/arrays/period.py
+++ b/pandas/core/arrays/period.py
@@ -608,7 +608,7 @@ def _sub_period(self, other):
return new_data
def _addsub_int_array(
- self, other: np.ndarray, op: Callable[[Any, Any], Any],
+ self, other: np.ndarray, op: Callable[[Any, Any], Any]
) -> "PeriodArray":
"""
Add or subtract array of integers; equivalent to applying
diff --git a/pandas/core/groupby/__init__.py b/pandas/core/groupby/__init__.py
index 0c5d2658978b4..15d8a996b6e7b 100644
--- a/pandas/core/groupby/__init__.py
+++ b/pandas/core/groupby/__init__.py
@@ -2,10 +2,4 @@
from pandas.core.groupby.groupby import GroupBy
from pandas.core.groupby.grouper import Grouper
-__all__ = [
- "DataFrameGroupBy",
- "NamedAgg",
- "SeriesGroupBy",
- "GroupBy",
- "Grouper",
-]
+__all__ = ["DataFrameGroupBy", "NamedAgg", "SeriesGroupBy", "GroupBy", "Grouper"]
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index c158bdfbac441..c55f77cd5fc28 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -274,7 +274,7 @@ def _outer_indexer(self, left, right):
# Constructors
def __new__(
- cls, data=None, dtype=None, copy=False, name=None, tupleize_cols=True, **kwargs,
+ cls, data=None, dtype=None, copy=False, name=None, tupleize_cols=True, **kwargs
) -> "Index":
from pandas.core.indexes.range import RangeIndex
diff --git a/pandas/core/indexes/range.py b/pandas/core/indexes/range.py
index 22940f851ddb0..43abae8267d0e 100644
--- a/pandas/core/indexes/range.py
+++ b/pandas/core/indexes/range.py
@@ -84,7 +84,7 @@ class RangeIndex(Int64Index):
# Constructors
def __new__(
- cls, start=None, stop=None, step=None, dtype=None, copy=False, name=None,
+ cls, start=None, stop=None, step=None, dtype=None, copy=False, name=None
):
cls._validate_dtype(dtype)
diff --git a/pandas/core/indexes/timedeltas.py b/pandas/core/indexes/timedeltas.py
index 1dd5c065ec216..166fe0ddb5ac2 100644
--- a/pandas/core/indexes/timedeltas.py
+++ b/pandas/core/indexes/timedeltas.py
@@ -69,9 +69,7 @@ class TimedeltaDelegateMixin(DatetimelikeDelegateMixin):
typ="method",
overwrite=True,
)
-class TimedeltaIndex(
- DatetimeTimedeltaMixin, dtl.TimelikeOps, TimedeltaDelegateMixin,
-):
+class TimedeltaIndex(DatetimeTimedeltaMixin, dtl.TimelikeOps, TimedeltaDelegateMixin):
"""
Immutable ndarray of timedelta64 data, represented internally as int64, and
which can be boxed to timedelta objects.
diff --git a/pandas/core/nanops.py b/pandas/core/nanops.py
index 2bf2be082f639..8f19303fd90e6 100644
--- a/pandas/core/nanops.py
+++ b/pandas/core/nanops.py
@@ -1282,7 +1282,7 @@ def _zero_out_fperr(arg):
@disallow("M8", "m8")
def nancorr(
- a: np.ndarray, b: np.ndarray, method="pearson", min_periods: Optional[int] = None,
+ a: np.ndarray, b: np.ndarray, method="pearson", min_periods: Optional[int] = None
):
"""
a, b: ndarrays
diff --git a/pandas/core/ops/__init__.py b/pandas/core/ops/__init__.py
index 1355060efd097..3d83a44a1f1da 100644
--- a/pandas/core/ops/__init__.py
+++ b/pandas/core/ops/__init__.py
@@ -82,14 +82,7 @@
}
-COMPARISON_BINOPS: Set[str] = {
- "eq",
- "ne",
- "lt",
- "gt",
- "le",
- "ge",
-}
+COMPARISON_BINOPS: Set[str] = {"eq", "ne", "lt", "gt", "le", "ge"}
# -----------------------------------------------------------------------------
# Ops Wrapping Utilities
diff --git a/pandas/core/ops/dispatch.py b/pandas/core/ops/dispatch.py
index 61a3032c7a02c..22a99f5d2eea3 100644
--- a/pandas/core/ops/dispatch.py
+++ b/pandas/core/ops/dispatch.py
@@ -94,7 +94,7 @@ def should_series_dispatch(left, right, op):
def dispatch_to_extension_op(
- op, left: Union[ABCExtensionArray, np.ndarray], right: Any,
+ op, left: Union[ABCExtensionArray, np.ndarray], right: Any
):
"""
Assume that left or right is a Series backed by an ExtensionArray,
diff --git a/pandas/core/reshape/pivot.py b/pandas/core/reshape/pivot.py
index e250a072766e3..38babc179ec62 100644
--- a/pandas/core/reshape/pivot.py
+++ b/pandas/core/reshape/pivot.py
@@ -226,7 +226,7 @@ def _add_margins(
elif values:
marginal_result_set = _generate_marginal_results(
- table, data, values, rows, cols, aggfunc, observed, margins_name,
+ table, data, values, rows, cols, aggfunc, observed, margins_name
)
if not isinstance(marginal_result_set, tuple):
return marginal_result_set
@@ -295,7 +295,7 @@ def _compute_grand_margin(data, values, aggfunc, margins_name: str = "All"):
def _generate_marginal_results(
- table, data, values, rows, cols, aggfunc, observed, margins_name: str = "All",
+ table, data, values, rows, cols, aggfunc, observed, margins_name: str = "All"
):
if len(cols) > 0:
# need to "interleave" the margins
diff --git a/pandas/core/window/indexers.py b/pandas/core/window/indexers.py
index 921cdb3c2523f..70298d5df3606 100644
--- a/pandas/core/window/indexers.py
+++ b/pandas/core/window/indexers.py
@@ -35,7 +35,7 @@ class BaseIndexer:
"""Base class for window bounds calculations."""
def __init__(
- self, index_array: Optional[np.ndarray] = None, window_size: int = 0, **kwargs,
+ self, index_array: Optional[np.ndarray] = None, window_size: int = 0, **kwargs
):
"""
Parameters
@@ -100,7 +100,7 @@ def get_window_bounds(
) -> Tuple[np.ndarray, np.ndarray]:
return calculate_variable_window_bounds(
- num_values, self.window_size, min_periods, center, closed, self.index_array,
+ num_values, self.window_size, min_periods, center, closed, self.index_array
)
diff --git a/pandas/core/window/numba_.py b/pandas/core/window/numba_.py
index 127957943d2ff..20dd1679550f2 100644
--- a/pandas/core/window/numba_.py
+++ b/pandas/core/window/numba_.py
@@ -63,7 +63,7 @@ def impl(window, *_args):
@numba.jit(nopython=nopython, nogil=nogil, parallel=parallel)
def roll_apply(
- values: np.ndarray, begin: np.ndarray, end: np.ndarray, minimum_periods: int,
+ values: np.ndarray, begin: np.ndarray, end: np.ndarray, minimum_periods: int
) -> np.ndarray:
result = np.empty(len(begin))
for i in loop_range(len(result)):
diff --git a/pandas/io/formats/css.py b/pandas/io/formats/css.py
index b40d2a57b8106..4d6f03489725f 100644
--- a/pandas/io/formats/css.py
+++ b/pandas/io/formats/css.py
@@ -20,9 +20,7 @@ def expand(self, prop, value: str):
try:
mapping = self.SIDE_SHORTHANDS[len(tokens)]
except KeyError:
- warnings.warn(
- f'Could not expand "{prop}: {value}"', CSSWarning,
- )
+ warnings.warn(f'Could not expand "{prop}: {value}"', CSSWarning)
return
for key, idx in zip(self.SIDES, mapping):
yield prop_fmt.format(key), tokens[idx]
@@ -117,10 +115,7 @@ def __call__(self, declarations_str, inherited=None):
props[prop] = self.size_to_pt(
props[prop], em_pt=font_size, conversions=self.BORDER_WIDTH_RATIOS
)
- for prop in [
- f"margin-{side}",
- f"padding-{side}",
- ]:
+ for prop in [f"margin-{side}", f"padding-{side}"]:
if prop in props:
# TODO: support %
props[prop] = self.size_to_pt(
diff --git a/pandas/io/formats/excel.py b/pandas/io/formats/excel.py
index b0e8e4033edf2..957a98abe68dd 100644
--- a/pandas/io/formats/excel.py
+++ b/pandas/io/formats/excel.py
@@ -140,8 +140,7 @@ def build_border(self, props: Dict) -> Dict[str, Dict[str, str]]:
return {
side: {
"style": self._border_style(
- props.get(f"border-{side}-style"),
- props.get(f"border-{side}-width"),
+ props.get(f"border-{side}-style"), props.get(f"border-{side}-width")
),
"color": self.color_to_excel(props.get(f"border-{side}-color")),
}
diff --git a/pandas/io/formats/style.py b/pandas/io/formats/style.py
index 565752e269d79..89219b0994288 100644
--- a/pandas/io/formats/style.py
+++ b/pandas/io/formats/style.py
@@ -303,10 +303,7 @@ def format_attr(pair):
# ... except maybe the last for columns.names
name = self.data.columns.names[r]
- cs = [
- BLANK_CLASS if name is None else INDEX_NAME_CLASS,
- f"level{r}",
- ]
+ cs = [BLANK_CLASS if name is None else INDEX_NAME_CLASS, f"level{r}"]
name = BLANK_VALUE if name is None else name
row_es.append(
{
@@ -320,11 +317,7 @@ def format_attr(pair):
if clabels:
for c, value in enumerate(clabels[r]):
- cs = [
- COL_HEADING_CLASS,
- f"level{r}",
- f"col{c}",
- ]
+ cs = [COL_HEADING_CLASS, f"level{r}", f"col{c}"]
cs.extend(
cell_context.get("col_headings", {}).get(r, {}).get(c, [])
)
@@ -368,11 +361,7 @@ def format_attr(pair):
for r, idx in enumerate(self.data.index):
row_es = []
for c, value in enumerate(rlabels[r]):
- rid = [
- ROW_HEADING_CLASS,
- f"level{c}",
- f"row{r}",
- ]
+ rid = [ROW_HEADING_CLASS, f"level{c}", f"row{r}"]
es = {
"type": "th",
"is_visible": (_is_visible(r, c, idx_lengths) and not hidden_index),
diff --git a/pandas/io/orc.py b/pandas/io/orc.py
index bbefe447cb7fe..a590c517d970c 100644
--- a/pandas/io/orc.py
+++ b/pandas/io/orc.py
@@ -12,7 +12,7 @@
def read_orc(
- path: FilePathOrBuffer, columns: Optional[List[str]] = None, **kwargs,
+ path: FilePathOrBuffer, columns: Optional[List[str]] = None, **kwargs
) -> "DataFrame":
"""
Load an ORC object from the file path, returning a DataFrame.
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index 3e4673c890bef..9c02f1979ceb6 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -270,6 +270,7 @@ def to_hdf(
min_itemsize=min_itemsize,
nan_rep=nan_rep,
data_columns=data_columns,
+ dropna=dropna,
errors=errors,
encoding=encoding,
)
@@ -995,6 +996,7 @@ def put(
min_itemsize: Optional[Union[int, Dict[str, int]]] = None,
nan_rep=None,
data_columns: Optional[List[str]] = None,
+ dropna: Optional[bool] = False,
encoding=None,
errors: str = "strict",
):
@@ -1015,14 +1017,25 @@ def put(
append : bool, default False
This will force Table format, append the input data to the
existing.
+ complib : {'zlib', 'lzo', 'bzip2', 'blosc'}, default 'zlib'
+ Specifies the compression library to be used.
+ As of v0.20.2 these additional compressors for Blosc are supported
+ (default if no compressor specified: 'blosc:blosclz'):
+ {'blosc:blosclz', 'blosc:lz4', 'blosc:lz4hc', 'blosc:snappy',
+ 'blosc:zlib', 'blosc:zstd'}.
+ Specifying a compression library which is not available issues
+ a ValueError.
+ complevel : int, 0-9, default None
+ Specifies a compression level for data.
+ A value of 0 or None disables compression.
+ dropna : bool, default False, do not write an ALL nan row to
+ The store settable by the option 'io.hdf.dropna_table'.
data_columns : list, default None
List of columns to create as data columns, or True to
use all columns. See `here
<https://pandas.pydata.org/pandas-docs/stable/user_guide/io.html#query-via-data-columns>`__.
encoding : str, default None
Provide an encoding for strings.
- dropna : bool, default False, do not write an ALL nan row to
- The store settable by the option 'io.hdf.dropna_table'.
"""
if format is None:
format = get_option("io.hdf.default_format") or "fixed"
@@ -1037,6 +1050,7 @@ def put(
complevel=complevel,
min_itemsize=min_itemsize,
nan_rep=nan_rep,
+ dropna=dropna,
data_columns=data_columns,
encoding=encoding,
errors=errors,
@@ -1147,6 +1161,17 @@ def append(
dropna : bool, default False
Do not write an ALL nan row to the store settable
by the option 'io.hdf.dropna_table'.
+ complevel : int, 0-9, default None
+ Specifies a compression level for data.
+ A value of 0 or None disables compression.
+ complib : {'zlib', 'lzo', 'bzip2', 'blosc'}, default 'zlib'
+ Specifies the compression library to be used.
+ As of v0.20.2 these additional compressors for Blosc are supported
+ (default if no compressor specified: 'blosc:blosclz'):
+ {'blosc:blosclz', 'blosc:lz4', 'blosc:lz4hc', 'blosc:snappy',
+ 'blosc:zlib', 'blosc:zstd'}.
+ Specifying a compression library which is not available issues
+ a ValueError.
Notes
-----
@@ -2836,7 +2861,7 @@ def read_index_node(
# If the index was an empty array write_array_empty() will
# have written a sentinel. Here we relace it with the original.
if "shape" in node._v_attrs and np.prod(node._v_attrs.shape) == 0:
- data = np.empty(node._v_attrs.shape, dtype=node._v_attrs.value_type,)
+ data = np.empty(node._v_attrs.shape, dtype=node._v_attrs.value_type)
kind = _ensure_decoded(node._v_attrs.kind)
name = None
@@ -3578,10 +3603,7 @@ def _read_axes(
for a in self.axes:
a.set_info(self.info)
res = a.convert(
- values,
- nan_rep=self.nan_rep,
- encoding=self.encoding,
- errors=self.errors,
+ values, nan_rep=self.nan_rep, encoding=self.encoding, errors=self.errors
)
results.append(res)
@@ -4007,7 +4029,7 @@ def create_description(
return d
def read_coordinates(
- self, where=None, start: Optional[int] = None, stop: Optional[int] = None,
+ self, where=None, start: Optional[int] = None, stop: Optional[int] = None
):
"""select coordinates (row numbers) from a table; return the
coordinates object
@@ -4274,7 +4296,7 @@ def write_data_chunk(
self.table.flush()
def delete(
- self, where=None, start: Optional[int] = None, stop: Optional[int] = None,
+ self, where=None, start: Optional[int] = None, stop: Optional[int] = None
):
# delete all rows (and return the nrows)
@@ -4452,7 +4474,7 @@ def is_transposed(self) -> bool:
def get_object(cls, obj, transposed: bool):
return obj
- def write(self, obj, data_columns=None, **kwargs):
+ def write(self, obj, data_columns=None, dropna=None, **kwargs):
""" we are going to write this as a frame table """
if not isinstance(obj, DataFrame):
name = obj.name or "values"
@@ -4705,7 +4727,7 @@ def _convert_index(name: str, index: Index, encoding: str, errors: str) -> Index
if inferred_type == "date":
converted = np.asarray([v.toordinal() for v in values], dtype=np.int32)
return IndexCol(
- name, converted, "date", _tables().Time32Col(), index_name=index_name,
+ name, converted, "date", _tables().Time32Col(), index_name=index_name
)
elif inferred_type == "string":
@@ -4721,13 +4743,13 @@ def _convert_index(name: str, index: Index, encoding: str, errors: str) -> Index
elif inferred_type in ["integer", "floating"]:
return IndexCol(
- name, values=converted, kind=kind, typ=atom, index_name=index_name,
+ name, values=converted, kind=kind, typ=atom, index_name=index_name
)
else:
assert isinstance(converted, np.ndarray) and converted.dtype == object
assert kind == "object", kind
atom = _tables().ObjectAtom()
- return IndexCol(name, converted, kind, atom, index_name=index_name,)
+ return IndexCol(name, converted, kind, atom, index_name=index_name)
def _unconvert_index(
diff --git a/pandas/tests/arithmetic/test_interval.py b/pandas/tests/arithmetic/test_interval.py
index f9e1a515277d5..b6ebeedb6a49d 100644
--- a/pandas/tests/arithmetic/test_interval.py
+++ b/pandas/tests/arithmetic/test_interval.py
@@ -149,9 +149,7 @@ def test_compare_scalar_other(self, op, array, other):
expected = self.elementwise_comparison(op, array, other)
tm.assert_numpy_array_equal(result, expected)
- def test_compare_list_like_interval(
- self, op, array, interval_constructor,
- ):
+ def test_compare_list_like_interval(self, op, array, interval_constructor):
# same endpoints
other = interval_constructor(array.left, array.right)
result = op(array, other)
diff --git a/pandas/tests/arithmetic/test_numeric.py b/pandas/tests/arithmetic/test_numeric.py
index f55e2b98ee912..b8794ed601bf0 100644
--- a/pandas/tests/arithmetic/test_numeric.py
+++ b/pandas/tests/arithmetic/test_numeric.py
@@ -98,7 +98,7 @@ class TestNumericArraylikeArithmeticWithDatetimeLike:
# TODO: also check name retentention
@pytest.mark.parametrize("box_cls", [np.array, pd.Index, pd.Series])
@pytest.mark.parametrize(
- "left", lefts, ids=lambda x: type(x).__name__ + str(x.dtype),
+ "left", lefts, ids=lambda x: type(x).__name__ + str(x.dtype)
)
def test_mul_td64arr(self, left, box_cls):
# GH#22390
@@ -118,7 +118,7 @@ def test_mul_td64arr(self, left, box_cls):
# TODO: also check name retentention
@pytest.mark.parametrize("box_cls", [np.array, pd.Index, pd.Series])
@pytest.mark.parametrize(
- "left", lefts, ids=lambda x: type(x).__name__ + str(x.dtype),
+ "left", lefts, ids=lambda x: type(x).__name__ + str(x.dtype)
)
def test_div_td64arr(self, left, box_cls):
# GH#22390
diff --git a/pandas/tests/arrays/sparse/test_array.py b/pandas/tests/arrays/sparse/test_array.py
index baca18239b929..c506944af7d60 100644
--- a/pandas/tests/arrays/sparse/test_array.py
+++ b/pandas/tests/arrays/sparse/test_array.py
@@ -660,16 +660,12 @@ def test_getslice_tuple(self):
dense = np.array([np.nan, 0, 3, 4, 0, 5, np.nan, np.nan, 0])
sparse = SparseArray(dense)
- res = sparse[
- 4:,
- ] # noqa: E231
+ res = sparse[4:,] # noqa: E231
exp = SparseArray(dense[4:,]) # noqa: E231
tm.assert_sp_array_equal(res, exp)
sparse = SparseArray(dense, fill_value=0)
- res = sparse[
- 4:,
- ] # noqa: E231
+ res = sparse[4:,] # noqa: E231
exp = SparseArray(dense[4:,], fill_value=0) # noqa: E231
tm.assert_sp_array_equal(res, exp)
diff --git a/pandas/tests/arrays/test_array.py b/pandas/tests/arrays/test_array.py
index b1b5a9482e34f..d5f5a2bb27975 100644
--- a/pandas/tests/arrays/test_array.py
+++ b/pandas/tests/arrays/test_array.py
@@ -35,7 +35,7 @@
np.dtype("float32"),
PandasArray(np.array([1.0, 2.0], dtype=np.dtype("float32"))),
),
- (np.array([1, 2], dtype="int64"), None, IntegerArray._from_sequence([1, 2]),),
+ (np.array([1, 2], dtype="int64"), None, IntegerArray._from_sequence([1, 2])),
# String alias passes through to NumPy
([1, 2], "float32", PandasArray(np.array([1, 2], dtype="float32"))),
# Period alias
@@ -120,10 +120,10 @@
(pd.Series([1, 2]), None, PandasArray(np.array([1, 2], dtype=np.int64))),
# String
(["a", None], "string", StringArray._from_sequence(["a", None])),
- (["a", None], pd.StringDtype(), StringArray._from_sequence(["a", None]),),
+ (["a", None], pd.StringDtype(), StringArray._from_sequence(["a", None])),
# Boolean
([True, None], "boolean", BooleanArray._from_sequence([True, None])),
- ([True, None], pd.BooleanDtype(), BooleanArray._from_sequence([True, None]),),
+ ([True, None], pd.BooleanDtype(), BooleanArray._from_sequence([True, None])),
# Index
(pd.Index([1, 2]), None, PandasArray(np.array([1, 2], dtype=np.int64))),
# Series[EA] returns the EA
@@ -174,7 +174,7 @@ def test_array_copy():
period_array(["2000", "2001"], freq="D"),
),
# interval
- ([pd.Interval(0, 1), pd.Interval(1, 2)], IntervalArray.from_breaks([0, 1, 2]),),
+ ([pd.Interval(0, 1), pd.Interval(1, 2)], IntervalArray.from_breaks([0, 1, 2])),
# datetime
(
[pd.Timestamp("2000"), pd.Timestamp("2001")],
diff --git a/pandas/tests/arrays/test_boolean.py b/pandas/tests/arrays/test_boolean.py
index cc8d0cdcb518d..d472e6b26f9f5 100644
--- a/pandas/tests/arrays/test_boolean.py
+++ b/pandas/tests/arrays/test_boolean.py
@@ -650,9 +650,7 @@ def test_kleene_xor_scalar(self, other, expected):
a, pd.array([True, False, None], dtype="boolean")
)
- @pytest.mark.parametrize(
- "other", [True, False, pd.NA, [True, False, None] * 3],
- )
+ @pytest.mark.parametrize("other", [True, False, pd.NA, [True, False, None] * 3])
def test_no_masked_assumptions(self, other, all_logical_operators):
# The logical operations should not assume that masked values are False!
a = pd.arrays.BooleanArray(
diff --git a/pandas/tests/arrays/test_timedeltas.py b/pandas/tests/arrays/test_timedeltas.py
index c86b4f71ee592..a32529cb58ba3 100644
--- a/pandas/tests/arrays/test_timedeltas.py
+++ b/pandas/tests/arrays/test_timedeltas.py
@@ -46,7 +46,7 @@ def test_incorrect_dtype_raises(self):
TimedeltaArray(np.array([1, 2, 3], dtype="i8"), dtype="category")
with pytest.raises(
- ValueError, match=r"dtype int64 cannot be converted to timedelta64\[ns\]",
+ ValueError, match=r"dtype int64 cannot be converted to timedelta64\[ns\]"
):
TimedeltaArray(np.array([1, 2, 3], dtype="i8"), dtype=np.dtype("int64"))
diff --git a/pandas/tests/base/test_conversion.py b/pandas/tests/base/test_conversion.py
index 07a15d0619bb6..10427b4ae14c0 100644
--- a/pandas/tests/base/test_conversion.py
+++ b/pandas/tests/base/test_conversion.py
@@ -187,7 +187,7 @@ def test_iter_box(self):
PeriodArray,
pd.core.dtypes.dtypes.PeriodDtype("A-DEC"),
),
- (pd.IntervalIndex.from_breaks([0, 1, 2]), IntervalArray, "interval",),
+ (pd.IntervalIndex.from_breaks([0, 1, 2]), IntervalArray, "interval"),
# This test is currently failing for datetime64[ns] and timedelta64[ns].
# The NumPy type system is sufficient for representing these types, so
# we just use NumPy for Series / DataFrame columns of these types (so
@@ -316,10 +316,7 @@ def test_array_multiindex_raises():
pd.core.arrays.period_array(["2000", "2001"], freq="D"),
np.array([pd.Period("2000", freq="D"), pd.Period("2001", freq="D")]),
),
- (
- pd.core.arrays.integer_array([0, np.nan]),
- np.array([0, pd.NA], dtype=object),
- ),
+ (pd.core.arrays.integer_array([0, np.nan]), np.array([0, pd.NA], dtype=object)),
(
IntervalArray.from_breaks([0, 1, 2]),
np.array([pd.Interval(0, 1), pd.Interval(1, 2)], dtype=object),
diff --git a/pandas/tests/frame/test_arithmetic.py b/pandas/tests/frame/test_arithmetic.py
index 659b55756c4b6..959470459a426 100644
--- a/pandas/tests/frame/test_arithmetic.py
+++ b/pandas/tests/frame/test_arithmetic.py
@@ -730,9 +730,7 @@ def test_zero_len_frame_with_series_corner_cases():
def test_frame_single_columns_object_sum_axis_1():
# GH 13758
- data = {
- "One": pd.Series(["A", 1.2, np.nan]),
- }
+ data = {"One": pd.Series(["A", 1.2, np.nan])}
df = pd.DataFrame(data)
result = df.sum(axis=1)
expected = pd.Series(["A", 1.2, 0])
diff --git a/pandas/tests/frame/test_reshape.py b/pandas/tests/frame/test_reshape.py
index b3af5a7b7317e..872e89c895d3b 100644
--- a/pandas/tests/frame/test_reshape.py
+++ b/pandas/tests/frame/test_reshape.py
@@ -406,7 +406,7 @@ def test_unstack_mixed_type_name_in_multiindex(
result = df.unstack(unstack_idx)
expected = pd.DataFrame(
- expected_values, columns=expected_columns, index=expected_index,
+ expected_values, columns=expected_columns, index=expected_index
)
tm.assert_frame_equal(result, expected)
diff --git a/pandas/tests/indexes/multi/test_contains.py b/pandas/tests/indexes/multi/test_contains.py
index 49aa63210cd5e..8fb9a11d6d0c5 100644
--- a/pandas/tests/indexes/multi/test_contains.py
+++ b/pandas/tests/indexes/multi/test_contains.py
@@ -113,7 +113,7 @@ def test_contains_with_missing_value():
@pytest.mark.parametrize(
"labels,expected,level",
[
- ([("b", np.nan)], np.array([False, False, True]), None,),
+ ([("b", np.nan)], np.array([False, False, True]), None),
([np.nan, "a"], np.array([True, True, False]), 0),
(["d", np.nan], np.array([False, True, True]), 1),
],
diff --git a/pandas/tests/indexing/common.py b/pandas/tests/indexing/common.py
index 3c027b035c2b8..d078e49e0a10c 100644
--- a/pandas/tests/indexing/common.py
+++ b/pandas/tests/indexing/common.py
@@ -170,7 +170,7 @@ def check_values(self, f, func, values=False):
tm.assert_almost_equal(result, expected)
def check_result(
- self, method1, key1, method2, key2, typs=None, axes=None, fails=None,
+ self, method1, key1, method2, key2, typs=None, axes=None, fails=None
):
def _eq(axis, obj, key1, key2):
""" compare equal for these 2 keys """
diff --git a/pandas/tests/indexing/test_callable.py b/pandas/tests/indexing/test_callable.py
index 621417eb38d94..be1bd4908fc79 100644
--- a/pandas/tests/indexing/test_callable.py
+++ b/pandas/tests/indexing/test_callable.py
@@ -17,14 +17,10 @@ def test_frame_loc_callable(self):
res = df.loc[lambda x: x.A > 2]
tm.assert_frame_equal(res, df.loc[df.A > 2])
- res = df.loc[
- lambda x: x.A > 2,
- ] # noqa: E231
+ res = df.loc[lambda x: x.A > 2,] # noqa: E231
tm.assert_frame_equal(res, df.loc[df.A > 2,]) # noqa: E231
- res = df.loc[
- lambda x: x.A > 2,
- ] # noqa: E231
+ res = df.loc[lambda x: x.A > 2,] # noqa: E231
tm.assert_frame_equal(res, df.loc[df.A > 2,]) # noqa: E231
res = df.loc[lambda x: x.B == "b", :]
@@ -94,9 +90,7 @@ def test_frame_loc_callable_labels(self):
res = df.loc[lambda x: ["A", "C"]]
tm.assert_frame_equal(res, df.loc[["A", "C"]])
- res = df.loc[
- lambda x: ["A", "C"],
- ] # noqa: E231
+ res = df.loc[lambda x: ["A", "C"],] # noqa: E231
tm.assert_frame_equal(res, df.loc[["A", "C"],]) # noqa: E231
res = df.loc[lambda x: ["A", "C"], :]
diff --git a/pandas/tests/indexing/test_loc.py b/pandas/tests/indexing/test_loc.py
index 4c1436b800fc3..beb6fac522d32 100644
--- a/pandas/tests/indexing/test_loc.py
+++ b/pandas/tests/indexing/test_loc.py
@@ -116,7 +116,7 @@ def test_loc_getitem_label_out_of_range(self):
self.check_result("loc", "f", "ix", "f", typs=["floats"], fails=KeyError)
self.check_result("loc", "f", "loc", "f", typs=["floats"], fails=KeyError)
self.check_result(
- "loc", 20, "loc", 20, typs=["ints", "uints", "mixed"], fails=KeyError,
+ "loc", 20, "loc", 20, typs=["ints", "uints", "mixed"], fails=KeyError
)
self.check_result("loc", 20, "loc", 20, typs=["labels"], fails=TypeError)
self.check_result("loc", 20, "loc", 20, typs=["ts"], axes=0, fails=TypeError)
@@ -129,7 +129,7 @@ def test_loc_getitem_label_list(self):
def test_loc_getitem_label_list_with_missing(self):
self.check_result(
- "loc", [0, 1, 2], "loc", [0, 1, 2], typs=["empty"], fails=KeyError,
+ "loc", [0, 1, 2], "loc", [0, 1, 2], typs=["empty"], fails=KeyError
)
self.check_result(
"loc",
diff --git a/pandas/tests/io/formats/test_css.py b/pandas/tests/io/formats/test_css.py
index 7008cef7b28fa..f6871e7a272b3 100644
--- a/pandas/tests/io/formats/test_css.py
+++ b/pandas/tests/io/formats/test_css.py
@@ -101,11 +101,11 @@ def test_css_side_shorthands(shorthand, expansions):
top, right, bottom, left = expansions
assert_resolves(
- f"{shorthand}: 1pt", {top: "1pt", right: "1pt", bottom: "1pt", left: "1pt"},
+ f"{shorthand}: 1pt", {top: "1pt", right: "1pt", bottom: "1pt", left: "1pt"}
)
assert_resolves(
- f"{shorthand}: 1pt 4pt", {top: "1pt", right: "4pt", bottom: "1pt", left: "4pt"},
+ f"{shorthand}: 1pt 4pt", {top: "1pt", right: "4pt", bottom: "1pt", left: "4pt"}
)
assert_resolves(
@@ -191,9 +191,7 @@ def test_css_absolute_font_size(size, relative_to, resolved):
inherited = None
else:
inherited = {"font-size": relative_to}
- assert_resolves(
- f"font-size: {size}", {"font-size": resolved}, inherited=inherited,
- )
+ assert_resolves(f"font-size: {size}", {"font-size": resolved}, inherited=inherited)
@pytest.mark.parametrize(
@@ -227,6 +225,4 @@ def test_css_relative_font_size(size, relative_to, resolved):
inherited = None
else:
inherited = {"font-size": relative_to}
- assert_resolves(
- f"font-size: {size}", {"font-size": resolved}, inherited=inherited,
- )
+ assert_resolves(f"font-size: {size}", {"font-size": resolved}, inherited=inherited)
diff --git a/pandas/tests/io/formats/test_format.py b/pandas/tests/io/formats/test_format.py
index 97956489e7da6..faa55e335f2b8 100644
--- a/pandas/tests/io/formats/test_format.py
+++ b/pandas/tests/io/formats/test_format.py
@@ -2379,8 +2379,7 @@ def test_east_asian_unicode_series(self):
# object dtype, longer than unicode repr
s = Series(
- [1, 22, 3333, 44444],
- index=[1, "AB", pd.Timestamp("2011-01-01"), "あああ"],
+ [1, 22, 3333, 44444], index=[1, "AB", pd.Timestamp("2011-01-01"), "あああ"]
)
expected = (
"1 1\n"
diff --git a/pandas/tests/io/parser/test_usecols.py b/pandas/tests/io/parser/test_usecols.py
index 979eb4702cc84..e05575cd79ccc 100644
--- a/pandas/tests/io/parser/test_usecols.py
+++ b/pandas/tests/io/parser/test_usecols.py
@@ -199,7 +199,7 @@ def test_usecols_with_whitespace(all_parsers):
# Column selection by index.
([0, 1], DataFrame(data=[[1000, 2000], [4000, 5000]], columns=["2", "0"])),
# Column selection by name.
- (["0", "1"], DataFrame(data=[[2000, 3000], [5000, 6000]], columns=["0", "1"]),),
+ (["0", "1"], DataFrame(data=[[2000, 3000], [5000, 6000]], columns=["0", "1"])),
],
)
def test_usecols_with_integer_like_header(all_parsers, usecols, expected):
diff --git a/pandas/tests/io/pytables/test_store.py b/pandas/tests/io/pytables/test_store.py
index 64c4ad800f49d..74de886572fd6 100644
--- a/pandas/tests/io/pytables/test_store.py
+++ b/pandas/tests/io/pytables/test_store.py
@@ -276,6 +276,29 @@ def test_api_default_format(self, setup_path):
pd.set_option("io.hdf.default_format", None)
+ def test_api_dropna(self, setup_path):
+
+ # GH2930
+
+ df = DataFrame({"A1": np.random.randn(20)}, index=np.arange(20))
+ df.loc[0:15] = np.nan
+
+ with ensure_clean_store(setup_path) as path:
+
+ df.to_hdf(path, "df", dropna=False, format="table")
+
+ with HDFStore(path) as store:
+ result = read_hdf(store, "df")
+ tm.assert_frame_equal(result, df)
+
+ with ensure_clean_store(setup_path) as path:
+
+ df.to_hdf(path, "df2", dropna=True, format="table")
+
+ with HDFStore(path) as store:
+ result = read_hdf(store, "df2")
+ tm.assert_frame_equal(result, df[-4:])
+
def test_keys(self, setup_path):
with ensure_clean_store(setup_path) as store:
@@ -804,7 +827,7 @@ def test_complibs(self, setup_path):
gname = "foo"
# Write and read file to see if data is consistent
- df.to_hdf(tmpfile, gname, complib=lib, complevel=lvl)
+ df.to_hdf(tmpfile, gname, complib=lib, complevel=lvl, format="table")
result = pd.read_hdf(tmpfile, gname)
tm.assert_frame_equal(result, df)
diff --git a/pandas/tests/io/test_parquet.py b/pandas/tests/io/test_parquet.py
index d51c712ed5abd..cb6cfa9c98afb 100644
--- a/pandas/tests/io/test_parquet.py
+++ b/pandas/tests/io/test_parquet.py
@@ -559,7 +559,7 @@ def test_additional_extension_types(self, pa):
{
# Arrow does not yet support struct in writing to Parquet (ARROW-1644)
# "c": pd.arrays.IntervalArray.from_tuples([(0, 1), (1, 2), (3, 4)]),
- "d": pd.period_range("2012-01-01", periods=3, freq="D"),
+ "d": pd.period_range("2012-01-01", periods=3, freq="D")
}
)
check_round_trip(df, pa)
diff --git a/pandas/tests/scalar/test_na_scalar.py b/pandas/tests/scalar/test_na_scalar.py
index dcb9d66708724..6662464bca2a4 100644
--- a/pandas/tests/scalar/test_na_scalar.py
+++ b/pandas/tests/scalar/test_na_scalar.py
@@ -96,7 +96,7 @@ def test_pow_special(value, asarray):
@pytest.mark.parametrize(
- "value", [1, 1.0, True, np.bool_(True), np.int_(1), np.float_(1)],
+ "value", [1, 1.0, True, np.bool_(True), np.int_(1), np.float_(1)]
)
@pytest.mark.parametrize("asarray", [True, False])
def test_rpow_special(value, asarray):
@@ -113,9 +113,7 @@ def test_rpow_special(value, asarray):
assert result == value
-@pytest.mark.parametrize(
- "value", [-1, -1.0, np.int_(-1), np.float_(-1)],
-)
+@pytest.mark.parametrize("value", [-1, -1.0, np.int_(-1), np.float_(-1)])
@pytest.mark.parametrize("asarray", [True, False])
def test_rpow_minus_one(value, asarray):
if asarray:
@@ -175,9 +173,7 @@ def test_logical_not():
assert ~NA is NA
-@pytest.mark.parametrize(
- "shape", [(3,), (3, 3), (1, 2, 3)],
-)
+@pytest.mark.parametrize("shape", [(3,), (3, 3), (1, 2, 3)])
def test_arithmetic_ndarray(shape, all_arithmetic_functions):
op = all_arithmetic_functions
a = np.zeros(shape)
diff --git a/pandas/tests/series/methods/test_argsort.py b/pandas/tests/series/methods/test_argsort.py
index 62273e2d363fb..67576fcc764c3 100644
--- a/pandas/tests/series/methods/test_argsort.py
+++ b/pandas/tests/series/methods/test_argsort.py
@@ -9,7 +9,7 @@ class TestSeriesArgsort:
def _check_accum_op(self, name, ser, check_dtype=True):
func = getattr(np, name)
tm.assert_numpy_array_equal(
- func(ser).values, func(np.array(ser)), check_dtype=check_dtype,
+ func(ser).values, func(np.array(ser)), check_dtype=check_dtype
)
# with missing values
diff --git a/pandas/tests/series/test_cumulative.py b/pandas/tests/series/test_cumulative.py
index 885b5bf0476f2..86f09807ac657 100644
--- a/pandas/tests/series/test_cumulative.py
+++ b/pandas/tests/series/test_cumulative.py
@@ -17,7 +17,7 @@
def _check_accum_op(name, series, check_dtype=True):
func = getattr(np, name)
tm.assert_numpy_array_equal(
- func(series).values, func(np.array(series)), check_dtype=check_dtype,
+ func(series).values, func(np.array(series)), check_dtype=check_dtype
)
# with missing values
diff --git a/pandas/tests/series/test_reshaping.py b/pandas/tests/series/test_reshaping.py
index 7645fb8759a54..1fb44a2620860 100644
--- a/pandas/tests/series/test_reshaping.py
+++ b/pandas/tests/series/test_reshaping.py
@@ -75,9 +75,7 @@ def test_unstack_tuplename_in_multiindex():
expected = pd.DataFrame(
[[1, 1, 1], [1, 1, 1], [1, 1, 1]],
- columns=pd.MultiIndex.from_tuples(
- [("a",), ("b",), ("c",)], names=[("A", "a")],
- ),
+ columns=pd.MultiIndex.from_tuples([("a",), ("b",), ("c",)], names=[("A", "a")]),
index=pd.Index([1, 2, 3], name=("B", "b")),
)
tm.assert_frame_equal(result, expected)
@@ -115,6 +113,6 @@ def test_unstack_mixed_type_name_in_multiindex(
result = ser.unstack(unstack_idx)
expected = pd.DataFrame(
- expected_values, columns=expected_columns, index=expected_index,
+ expected_values, columns=expected_columns, index=expected_index
)
tm.assert_frame_equal(result, expected)
diff --git a/pandas/tests/test_strings.py b/pandas/tests/test_strings.py
index 62d26dacde67b..bbc7552bda4b0 100644
--- a/pandas/tests/test_strings.py
+++ b/pandas/tests/test_strings.py
@@ -3520,9 +3520,7 @@ def test_string_array(any_string_method):
result = getattr(b.str, method_name)(*args, **kwargs)
if isinstance(expected, Series):
- if expected.dtype == "object" and lib.is_string_array(
- expected.dropna().values,
- ):
+ if expected.dtype == "object" and lib.is_string_array(expected.dropna().values):
assert result.dtype == "string"
result = result.astype(object)
diff --git a/pandas/util/_test_decorators.py b/pandas/util/_test_decorators.py
index d8804994af426..4526df2f7b951 100644
--- a/pandas/util/_test_decorators.py
+++ b/pandas/util/_test_decorators.py
@@ -182,10 +182,10 @@ def skip_if_no(package: str, min_version: Optional[str] = None) -> Callable:
is_platform_windows(), reason="not used on win32"
)
skip_if_has_locale = pytest.mark.skipif(
- _skip_if_has_locale(), reason=f"Specific locale is set {locale.getlocale()[0]}",
+ _skip_if_has_locale(), reason=f"Specific locale is set {locale.getlocale()[0]}"
)
skip_if_not_us_locale = pytest.mark.skipif(
- _skip_if_not_us_locale(), reason=f"Specific locale is set {locale.getlocale()[0]}",
+ _skip_if_not_us_locale(), reason=f"Specific locale is set {locale.getlocale()[0]}"
)
skip_if_no_scipy = pytest.mark.skipif(
_skip_if_no_scipy(), reason="Missing SciPy requirement"
| - [x] closes #29310
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Re #29310, the complib and complevel parameters were not being passed down all the way previously, hence HDF compression not working.
I noticed that the implementation of to_hdf() specifies that compression is not allowed for fixed formats:
` if not s.is_table and complib:
raise ValueError("Compression not supported on Fixed format stores")`
I'm guessing that means the performance comparison section for https://github.com/pandas-dev/pandas/pull/28890/files will also need to be updated to remove the test_fixed_compress test @WuraolaOyewusi?
Also, after the update, the following test is currently failing:

due to a ValueError for using compression with a fixed format, and I'm not sure as to why the expected behaviour for this test is what it is? Why should setting complib disable compression? I would appreciate any further info on that.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29404 | 2019-11-04T23:06:06Z | 2020-02-07T07:55:30Z | null | 2020-02-07T07:55:31Z |
API: rename labels to codes in core/groupby | diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 8512b6c3ae530..511b87dab087e 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -655,16 +655,17 @@ def value_counts(
rep = partial(np.repeat, repeats=np.add.reduceat(inc, idx))
# multi-index components
- labels = list(map(rep, self.grouper.recons_labels)) + [llab(lab, inc)]
+ codes = self.grouper.recons_codes
+ codes = [rep(level_codes) for level_codes in codes] + [llab(lab, inc)]
levels = [ping.group_index for ping in self.grouper.groupings] + [lev]
names = self.grouper.names + [self._selection_name]
if dropna:
- mask = labels[-1] != -1
+ mask = codes[-1] != -1
if mask.all():
dropna = False
else:
- out, labels = out[mask], [label[mask] for label in labels]
+ out, codes = out[mask], [level_codes[mask] for level_codes in codes]
if normalize:
out = out.astype("float")
@@ -680,11 +681,11 @@ def value_counts(
if sort and bins is None:
cat = ids[inc][mask] if dropna else ids[inc]
sorter = np.lexsort((out if ascending else -out, cat))
- out, labels[-1] = out[sorter], labels[-1][sorter]
+ out, codes[-1] = out[sorter], codes[-1][sorter]
if bins is None:
mi = MultiIndex(
- levels=levels, codes=labels, names=names, verify_integrity=False
+ levels=levels, codes=codes, names=names, verify_integrity=False
)
if is_integer_dtype(out):
@@ -694,14 +695,14 @@ def value_counts(
# for compat. with libgroupby.value_counts need to ensure every
# bin is present at every index level, null filled with zeros
diff = np.zeros(len(out), dtype="bool")
- for lab in labels[:-1]:
- diff |= np.r_[True, lab[1:] != lab[:-1]]
+ for level_codes in codes[:-1]:
+ diff |= np.r_[True, level_codes[1:] != level_codes[:-1]]
ncat, nbin = diff.sum(), len(levels[-1])
left = [np.repeat(np.arange(ncat), nbin), np.tile(np.arange(nbin), ncat)]
- right = [diff.cumsum() - 1, labels[-1]]
+ right = [diff.cumsum() - 1, codes[-1]]
_, idx = _get_join_indexers(left, right, sort=False, how="left")
out = np.where(idx != -1, out[idx], 0)
@@ -711,7 +712,10 @@ def value_counts(
out, left[-1] = out[sorter], left[-1][sorter]
# build the multi-index w/ full levels
- codes = list(map(lambda lab: np.repeat(lab[diff], nbin), labels[:-1]))
+ def build_codes(lev_codes: np.ndarray) -> np.ndarray:
+ return np.repeat(lev_codes[diff], nbin)
+
+ codes = [build_codes(lev_codes) for lev_codes in codes[:-1]]
codes.append(left[-1])
mi = MultiIndex(levels=levels, codes=codes, names=names, verify_integrity=False)
@@ -758,7 +762,7 @@ def pct_change(self, periods=1, fill_method="pad", limit=None, freq=None):
)
)
filled = getattr(self, fill_method)(limit=limit)
- fill_grp = filled.groupby(self.grouper.labels)
+ fill_grp = filled.groupby(self.grouper.codes)
shifted = fill_grp.shift(periods=periods, freq=freq)
return (filled / shifted) - 1
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index fa4a184e8f7a4..81ba594c97391 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -2349,7 +2349,7 @@ def pct_change(self, periods=1, fill_method="pad", limit=None, freq=None, axis=0
)
)
filled = getattr(self, fill_method)(limit=limit)
- fill_grp = filled.groupby(self.grouper.labels)
+ fill_grp = filled.groupby(self.grouper.codes)
shifted = fill_grp.shift(periods=periods, freq=freq)
return (filled / shifted) - 1
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index 45d2a819ae5ad..dc6336b17ac1e 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -3,7 +3,7 @@
split-apply-combine paradigm.
"""
-from typing import Tuple
+from typing import Optional, Tuple
import warnings
import numpy as np
@@ -21,6 +21,7 @@
)
from pandas.core.dtypes.generic import ABCSeries
+from pandas._typing import FrameOrSeries
import pandas.core.algorithms as algorithms
from pandas.core.arrays import Categorical, ExtensionArray
import pandas.core.common as com
@@ -228,10 +229,10 @@ class Grouping:
----------
index : Index
grouper :
- obj :
+ obj Union[DataFrame, Series]:
name :
level :
- observed : boolean, default False
+ observed : bool, default False
If we are a Categorical, use the observed values
in_axis : if the Grouping is a column in self.obj and hence among
Groupby.exclusions list
@@ -240,25 +241,22 @@ class Grouping:
-------
**Attributes**:
* indices : dict of {group -> index_list}
- * labels : ndarray, group labels
- * ids : mapping of label -> group
- * counts : array of group counts
+ * codes : ndarray, group codes
* group_index : unique groups
* groups : dict of {group -> label_list}
"""
def __init__(
self,
- index,
+ index: Index,
grouper=None,
- obj=None,
+ obj: Optional[FrameOrSeries] = None,
name=None,
level=None,
- sort=True,
- observed=False,
- in_axis=False,
+ sort: bool = True,
+ observed: bool = False,
+ in_axis: bool = False,
):
-
self.name = name
self.level = level
self.grouper = _convert_grouper(index, grouper)
@@ -290,12 +288,12 @@ def __init__(
if self.name is None:
self.name = index.names[level]
- self.grouper, self._labels, self._group_index = index._get_grouper_for_level( # noqa: E501
+ self.grouper, self._codes, self._group_index = index._get_grouper_for_level( # noqa: E501
self.grouper, level
)
# a passed Grouper like, directly get the grouper in the same way
- # as single grouper groupby, use the group_info to get labels
+ # as single grouper groupby, use the group_info to get codes
elif isinstance(self.grouper, Grouper):
# get the new grouper; we already have disambiguated
# what key/level refer to exactly, don't need to
@@ -308,7 +306,7 @@ def __init__(
self.grouper = grouper._get_grouper()
else:
- if self.grouper is None and self.name is not None:
+ if self.grouper is None and self.name is not None and self.obj is not None:
self.grouper = self.obj[self.name]
elif isinstance(self.grouper, (list, tuple)):
@@ -324,7 +322,7 @@ def __init__(
# we make a CategoricalIndex out of the cat grouper
# preserving the categories / ordered attributes
- self._labels = self.grouper.codes
+ self._codes = self.grouper.codes
if observed:
codes = algorithms.unique1d(self.grouper.codes)
codes = codes[codes != -1]
@@ -380,11 +378,11 @@ def __repr__(self):
def __iter__(self):
return iter(self.indices)
- _labels = None
- _group_index = None
+ _codes = None # type: np.ndarray
+ _group_index = None # type: Index
@property
- def ngroups(self):
+ def ngroups(self) -> int:
return len(self.group_index)
@cache_readonly
@@ -397,38 +395,38 @@ def indices(self):
return values._reverse_indexer()
@property
- def labels(self):
- if self._labels is None:
- self._make_labels()
- return self._labels
+ def codes(self) -> np.ndarray:
+ if self._codes is None:
+ self._make_codes()
+ return self._codes
@cache_readonly
- def result_index(self):
+ def result_index(self) -> Index:
if self.all_grouper is not None:
return recode_from_groupby(self.all_grouper, self.sort, self.group_index)
return self.group_index
@property
- def group_index(self):
+ def group_index(self) -> Index:
if self._group_index is None:
- self._make_labels()
+ self._make_codes()
return self._group_index
- def _make_labels(self):
- if self._labels is None or self._group_index is None:
+ def _make_codes(self) -> None:
+ if self._codes is None or self._group_index is None:
# we have a list of groupers
if isinstance(self.grouper, BaseGrouper):
- labels = self.grouper.label_info
+ codes = self.grouper.codes_info
uniques = self.grouper.result_index
else:
- labels, uniques = algorithms.factorize(self.grouper, sort=self.sort)
+ codes, uniques = algorithms.factorize(self.grouper, sort=self.sort)
uniques = Index(uniques, name=self.name)
- self._labels = labels
+ self._codes = codes
self._group_index = uniques
@cache_readonly
- def groups(self):
- return self.index.groupby(Categorical.from_codes(self.labels, self.group_index))
+ def groups(self) -> dict:
+ return self.index.groupby(Categorical.from_codes(self.codes, self.group_index))
def _get_grouper(
@@ -678,7 +676,7 @@ def _is_label_like(val):
return isinstance(val, (str, tuple)) or (val is not None and is_scalar(val))
-def _convert_grouper(axis, grouper):
+def _convert_grouper(axis: Index, grouper):
if isinstance(grouper, dict):
return grouper.get
elif isinstance(grouper, Series):
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 5bad73bf40ff5..2c8aa1294451d 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,7 +7,7 @@
"""
import collections
-from typing import List, Optional, Type
+from typing import List, Optional, Sequence, Type
import numpy as np
@@ -41,7 +41,7 @@
import pandas.core.common as com
from pandas.core.frame import DataFrame
from pandas.core.generic import NDFrame
-from pandas.core.groupby import base
+from pandas.core.groupby import base, grouper
from pandas.core.index import Index, MultiIndex, ensure_index
from pandas.core.series import Series
from pandas.core.sorting import (
@@ -62,13 +62,13 @@ class BaseGrouper:
Parameters
----------
axis : Index
- groupings : array of grouping
+ groupings : Sequence[Grouping]
all the grouping instances to handle in this grouper
for example for grouper list to groupby, need to pass the list
- sort : boolean, default True
+ sort : bool, default True
whether this grouper will give sorted result or not
- group_keys : boolean, default True
- mutated : boolean, default False
+ group_keys : bool, default True
+ mutated : bool, default False
indexer : intp array, optional
the indexer created by Grouper
some groupers (TimeGrouper) will sort its axis and its
@@ -79,16 +79,17 @@ class BaseGrouper:
def __init__(
self,
axis: Index,
- groupings,
- sort=True,
- group_keys=True,
- mutated=False,
- indexer=None,
+ groupings: "Sequence[grouper.Grouping]",
+ sort: bool = True,
+ group_keys: bool = True,
+ mutated: bool = False,
+ indexer: Optional[np.ndarray] = None,
):
assert isinstance(axis, Index), axis
+
self._filter_empty_groups = self.compressed = len(groupings) != 1
self.axis = axis
- self.groupings = groupings
+ self.groupings = groupings # type: Sequence[grouper.Grouping]
self.sort = sort
self.group_keys = group_keys
self.mutated = mutated
@@ -139,7 +140,7 @@ def _get_group_keys(self):
comp_ids, _, ngroups = self.group_info
# provide "flattened" iterator for multi-group setting
- return get_flattened_iterator(comp_ids, ngroups, self.levels, self.labels)
+ return get_flattened_iterator(comp_ids, ngroups, self.levels, self.codes)
def apply(self, f, data, axis: int = 0):
mutated = self.mutated
@@ -210,13 +211,13 @@ def indices(self):
if len(self.groupings) == 1:
return self.groupings[0].indices
else:
- label_list = [ping.labels for ping in self.groupings]
+ codes_list = [ping.codes for ping in self.groupings]
keys = [com.values_from_object(ping.group_index) for ping in self.groupings]
- return get_indexer_dict(label_list, keys)
+ return get_indexer_dict(codes_list, keys)
@property
- def labels(self):
- return [ping.labels for ping in self.groupings]
+ def codes(self):
+ return [ping.codes for ping in self.groupings]
@property
def levels(self):
@@ -256,46 +257,46 @@ def is_monotonic(self) -> bool:
@cache_readonly
def group_info(self):
- comp_ids, obs_group_ids = self._get_compressed_labels()
+ comp_ids, obs_group_ids = self._get_compressed_codes()
ngroups = len(obs_group_ids)
comp_ids = ensure_int64(comp_ids)
return comp_ids, obs_group_ids, ngroups
@cache_readonly
- def label_info(self):
- # return the labels of items in original grouped axis
- labels, _, _ = self.group_info
+ def codes_info(self):
+ # return the codes of items in original grouped axis
+ codes, _, _ = self.group_info
if self.indexer is not None:
- sorter = np.lexsort((labels, self.indexer))
- labels = labels[sorter]
- return labels
-
- def _get_compressed_labels(self):
- all_labels = [ping.labels for ping in self.groupings]
- if len(all_labels) > 1:
- group_index = get_group_index(all_labels, self.shape, sort=True, xnull=True)
+ sorter = np.lexsort((codes, self.indexer))
+ codes = codes[sorter]
+ return codes
+
+ def _get_compressed_codes(self):
+ all_codes = [ping.codes for ping in self.groupings]
+ if len(all_codes) > 1:
+ group_index = get_group_index(all_codes, self.shape, sort=True, xnull=True)
return compress_group_index(group_index, sort=self.sort)
ping = self.groupings[0]
- return ping.labels, np.arange(len(ping.group_index))
+ return ping.codes, np.arange(len(ping.group_index))
@cache_readonly
def ngroups(self) -> int:
return len(self.result_index)
@property
- def recons_labels(self):
+ def recons_codes(self):
comp_ids, obs_ids, _ = self.group_info
- labels = (ping.labels for ping in self.groupings)
- return decons_obs_group_ids(comp_ids, obs_ids, self.shape, labels, xnull=True)
+ codes = (ping.codes for ping in self.groupings)
+ return decons_obs_group_ids(comp_ids, obs_ids, self.shape, codes, xnull=True)
@cache_readonly
def result_index(self):
if not self.compressed and len(self.groupings) == 1:
return self.groupings[0].result_index.rename(self.names[0])
- codes = self.recons_labels
+ codes = self.recons_codes
levels = [ping.result_index for ping in self.groupings]
result = MultiIndex(
levels=levels, codes=codes, verify_integrity=False, names=self.names
@@ -307,9 +308,9 @@ def get_group_levels(self):
return [self.groupings[0].result_index]
name_list = []
- for ping, labels in zip(self.groupings, self.recons_labels):
- labels = ensure_platform_int(labels)
- levels = ping.result_index.take(labels)
+ for ping, codes in zip(self.groupings, self.recons_codes):
+ codes = ensure_platform_int(codes)
+ levels = ping.result_index.take(codes)
name_list.append(levels)
@@ -490,7 +491,7 @@ def _cython_operation(
else:
out_dtype = "object"
- labels, _, _ = self.group_info
+ codes, _, _ = self.group_info
if kind == "aggregate":
result = _maybe_fill(
@@ -498,7 +499,7 @@ def _cython_operation(
)
counts = np.zeros(self.ngroups, dtype=np.int64)
result = self._aggregate(
- result, counts, values, labels, func, is_datetimelike, min_count
+ result, counts, values, codes, func, is_datetimelike, min_count
)
elif kind == "transform":
result = _maybe_fill(
@@ -507,7 +508,7 @@ def _cython_operation(
# TODO: min_count
result = self._transform(
- result, values, labels, func, is_datetimelike, **kwargs
+ result, values, codes, func, is_datetimelike, **kwargs
)
if is_integer_dtype(result) and not is_datetimelike:
diff --git a/pandas/tests/groupby/test_grouping.py b/pandas/tests/groupby/test_grouping.py
index e1fd8d7da6833..e4edc64016567 100644
--- a/pandas/tests/groupby/test_grouping.py
+++ b/pandas/tests/groupby/test_grouping.py
@@ -559,12 +559,12 @@ def test_level_preserve_order(self, sort, labels, mframe):
# GH 17537
grouped = mframe.groupby(level=0, sort=sort)
exp_labels = np.array(labels, np.intp)
- tm.assert_almost_equal(grouped.grouper.labels[0], exp_labels)
+ tm.assert_almost_equal(grouped.grouper.codes[0], exp_labels)
def test_grouping_labels(self, mframe):
grouped = mframe.groupby(mframe.index.get_level_values(0))
exp_labels = np.array([2, 2, 2, 0, 0, 1, 1, 3, 3, 3], dtype=np.intp)
- tm.assert_almost_equal(grouped.grouper.labels[0], exp_labels)
+ tm.assert_almost_equal(grouped.grouper.codes[0], exp_labels)
def test_list_grouper_with_nat(self):
# GH 14715
diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index 5a2f189ad8d10..4ba32c377a345 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -621,8 +621,8 @@ def _check_types(l, r, obj="Index"):
def _get_ilevel_values(index, level):
# accept level number only
unique = index.levels[level]
- labels = index.codes[level]
- filled = take_1d(unique.values, labels, fill_value=unique._na_value)
+ level_codes = index.codes[level]
+ filled = take_1d(unique.values, level_codes, fill_value=unique._na_value)
values = unique._shallow_copy(filled, name=index.names[level])
return values
| This PR renames the various ``*label*`` names in core/groupby to like-named ``*codes*``.
I think the name ``label`` can be confused by the single values in a index, and ``codes`` sound smore like an array of ints, so by renaming we get a cleaner nomenclature, IMO.
All these attributes/methods are internal, so no deprecations needed. | https://api.github.com/repos/pandas-dev/pandas/pulls/29402 | 2019-11-04T22:24:21Z | 2019-11-07T01:34:47Z | 2019-11-07T01:34:47Z | 2019-11-07T01:34:51Z |
removing kendall tests | diff --git a/asv_bench/benchmarks/stat_ops.py b/asv_bench/benchmarks/stat_ops.py
index ed5ebfa61594e..ec67394e55a1e 100644
--- a/asv_bench/benchmarks/stat_ops.py
+++ b/asv_bench/benchmarks/stat_ops.py
@@ -7,20 +7,14 @@
class FrameOps:
- params = [ops, ["float", "int"], [0, 1], [True, False]]
- param_names = ["op", "dtype", "axis", "use_bottleneck"]
+ params = [ops, ["float", "int"], [0, 1]]
+ param_names = ["op", "dtype", "axis"]
- def setup(self, op, dtype, axis, use_bottleneck):
+ def setup(self, op, dtype, axis):
df = pd.DataFrame(np.random.randn(100000, 4)).astype(dtype)
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
-
- nanops._USE_BOTTLENECK = use_bottleneck
self.df_func = getattr(df, op)
- def time_op(self, op, dtype, axis, use_bottleneck):
+ def time_op(self, op, dtype, axis):
self.df_func(axis=axis)
@@ -46,20 +40,14 @@ def time_op(self, level, op):
class SeriesOps:
- params = [ops, ["float", "int"], [True, False]]
- param_names = ["op", "dtype", "use_bottleneck"]
+ params = [ops, ["float", "int"]]
+ param_names = ["op", "dtype"]
- def setup(self, op, dtype, use_bottleneck):
+ def setup(self, op, dtype):
s = pd.Series(np.random.randn(100000)).astype(dtype)
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
-
- nanops._USE_BOTTLENECK = use_bottleneck
self.s_func = getattr(s, op)
- def time_op(self, op, dtype, use_bottleneck):
+ def time_op(self, op, dtype):
self.s_func()
@@ -101,61 +89,49 @@ def time_average_old(self, constructor, pct):
class Correlation:
- params = [["spearman", "kendall", "pearson"], [True, False]]
- param_names = ["method", "use_bottleneck"]
+ params = [["spearman", "kendall", "pearson"]]
+ param_names = ["method"]
- def setup(self, method, use_bottleneck):
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
+ def setup(self, method):
+ self.df = pd.DataFrame(np.random.randn(500, 15))
+ self.df2 = pd.DataFrame(np.random.randn(500, 15))
+ self.df_wide = pd.DataFrame(np.random.randn(500, 100))
+ self.df_wide_nans = self.df_wide.where(np.random.random((500, 100)) < 0.9)
+ self.s = pd.Series(np.random.randn(500))
+ self.s2 = pd.Series(np.random.randn(500))
- nanops._USE_BOTTLENECK = use_bottleneck
- self.df = pd.DataFrame(np.random.randn(1000, 30))
- self.df2 = pd.DataFrame(np.random.randn(1000, 30))
- self.df_wide = pd.DataFrame(np.random.randn(1000, 200))
- self.df_wide_nans = self.df_wide.where(np.random.random((1000, 200)) < 0.9)
- self.s = pd.Series(np.random.randn(1000))
- self.s2 = pd.Series(np.random.randn(1000))
-
- def time_corr(self, method, use_bottleneck):
+ def time_corr(self, method):
self.df.corr(method=method)
- def time_corr_wide(self, method, use_bottleneck):
+ def time_corr_wide(self, method):
self.df_wide.corr(method=method)
- def time_corr_wide_nans(self, method, use_bottleneck):
+ def time_corr_wide_nans(self, method):
self.df_wide_nans.corr(method=method)
- def peakmem_corr_wide(self, method, use_bottleneck):
+ def peakmem_corr_wide(self, method):
self.df_wide.corr(method=method)
- def time_corr_series(self, method, use_bottleneck):
+ def time_corr_series(self, method):
self.s.corr(self.s2, method=method)
- def time_corrwith_cols(self, method, use_bottleneck):
+ def time_corrwith_cols(self, method):
self.df.corrwith(self.df2, method=method)
- def time_corrwith_rows(self, method, use_bottleneck):
+ def time_corrwith_rows(self, method):
self.df.corrwith(self.df2, axis=1, method=method)
class Covariance:
- params = [[True, False]]
- param_names = ["use_bottleneck"]
-
- def setup(self, use_bottleneck):
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
+ params = []
+ param_names = []
- nanops._USE_BOTTLENECK = use_bottleneck
+ def setup(self):
self.s = pd.Series(np.random.randn(100000))
self.s2 = pd.Series(np.random.randn(100000))
- def time_cov_series(self, use_bottleneck):
+ def time_cov_series(self):
self.s.cov(self.s2)
| closes #29270
Following is the output after removing "kendall"
```
· Creating environments
· Discovering benchmarks
·· Uninstalling from conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
·· Building cd59acf5 <fix-kendall-issues> for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt................................................
·· Installing cd59acf5 <fix-kendall-issues> into conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt..
· Running 14 total benchmarks (2 commits * 1 environments * 7 benchmarks)
[ 0.00%] · For pandas commit 165d5ee4 <master> (round 1/2):
[ 0.00%] ·· Building for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt...
[ 0.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 7.14%] ··· Running (stat_ops.Correlation.time_corr--)....
[ 21.43%] ··· Running (stat_ops.Correlation.time_corrwith_cols--)..
[ 25.00%] · For pandas commit cd59acf5 <fix-kendall-issues> (round 1/2):
[ 25.00%] ·· Building for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt...
[ 25.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 32.14%] ··· Running (stat_ops.Correlation.time_corr--)....
[ 46.43%] ··· Running (stat_ops.Correlation.time_corrwith_cols--)..
[ 50.00%] · For pandas commit cd59acf5 <fix-kendall-issues> (round 2/2):
[ 50.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 53.57%] ··· stat_ops.Correlation.peakmem_corr_wide ok
[ 53.57%] ··· ========== ====== =======
-- use_bottleneck
---------- --------------
method True False
========== ====== =======
spearman 108M 108M
pearson 105M 105M
========== ====== =======
[ 57.14%] ··· stat_ops.Correlation.time_corr ok
[ 57.14%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 8.74±1ms 8.71±2ms
pearson 2.88±0.6ms 2.84±0.4ms
========== ============ ============
[ 60.71%] ··· stat_ops.Correlation.time_corr_series ok
[ 60.71%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 1.34±0.2ms 1.35±0.3ms
pearson 317±60μs 319±70μs
========== ============ ============
[ 64.29%] ··· stat_ops.Correlation.time_corr_wide ok
[ 64.29%] ··· ========== ========== ==========
-- use_bottleneck
---------- ---------------------
method True False
========== ========== ==========
spearman 286±40ms 273±10ms
pearson 176±10ms 178±10ms
========== ========== ==========
[ 67.86%] ··· stat_ops.Correlation.time_corr_wide_nans ok
[ 67.86%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 3.23±0.02s 3.20±0.01s
pearson 196±7ms 192±10ms
========== ============ ============
[ 71.43%] ··· stat_ops.Correlation.time_corrwith_cols ok
[ 71.43%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 24.2±1ms 25.5±1ms
pearson 36.1±0.3ms 36.2±0.4ms
========== ============ ============
[ 75.00%] ··· stat_ops.Correlation.time_corrwith_rows ok
[ 75.00%] ··· ========== ========= ==========
-- use_bottleneck
---------- --------------------
method True False
========== ========= ==========
spearman 549±4ms 551±3ms
pearson 884±7ms 890±20ms
========== ========= ==========
[ 75.00%] · For pandas commit 165d5ee4 <master> (round 2/2):
[ 75.00%] ·· Building for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt...
[ 75.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 78.57%] ··· stat_ops.Correlation.peakmem_corr_wide ok
[ 78.57%] ··· ========== ====== =======
-- use_bottleneck
---------- --------------
method True False
========== ====== =======
spearman 108M 108M
pearson 106M 106M
========== ====== =======
[ 82.14%] ··· stat_ops.Correlation.time_corr ok
[ 82.14%] ··· ========== ============ =============
-- use_bottleneck
---------- --------------------------
method True False
========== ============ =============
spearman 10.2±0.2ms 10.1±0.2ms
pearson 3.51±0.2ms 3.54±0.09ms
========== ============ =============
[ 85.71%] ··· stat_ops.Correlation.time_corr_series ok
[ 85.71%] ··· ========== ============= =============
-- use_bottleneck
---------- ---------------------------
method True False
========== ============= =============
spearman 1.52±0.03ms 1.26±0.01ms
pearson 316±10μs 389±0.6μs
========== ============= =============
[ 89.29%] ··· stat_ops.Correlation.time_corr_wide ok
[ 89.29%] ··· ========== ========== ==========
-- use_bottleneck
---------- ---------------------
method True False
========== ========== ==========
spearman 359±20ms 288±10ms
pearson 186±6ms 184±7ms
========== ========== ==========
[ 92.86%] ··· stat_ops.Correlation.time_corr_wide_nans ok
[ 92.86%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 3.29±0.01s 3.26±0.03s
pearson 195±6ms 203±5ms
========== ============ ============
[ 96.43%] ··· stat_ops.Correlation.time_corrwith_cols ok
[ 96.43%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 24.4±2ms 24.8±0.9ms
pearson 35.8±0.5ms 36.1±0.5ms
========== ============ ============
[100.00%] ··· stat_ops.Correlation.time_corrwith_rows ok
[100.00%] ··· ========== ========== ==========
-- use_bottleneck
---------- ---------------------
method True False
========== ========== ==========
spearman 551±6ms 555±7ms
pearson 894±10ms 899±10ms
========== ========== ==========
before after ratio
[165d5ee4] [cd59acf5]
<master> <fix-kendall-issues>
- 3.54±0.09ms 2.84±0.4ms 0.80 stat_ops.Correlation.time_corr('pearson', False)
SOME BENCHMARKS HAVE CHANGED SIGNIFICANTLY.
PERFORMANCE INCREASED.
``` | https://api.github.com/repos/pandas-dev/pandas/pulls/29401 | 2019-11-04T22:02:52Z | 2019-12-05T16:35:46Z | 2019-12-05T16:35:45Z | 2019-12-05T16:35:52Z |
REF: implement first_valid_index in core.missing | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index cbeee88d75b51..bafc37d478fdb 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -83,6 +83,7 @@
from pandas.core.indexes.period import Period, PeriodIndex
import pandas.core.indexing as indexing
from pandas.core.internals import BlockManager
+from pandas.core.missing import find_valid_index
from pandas.core.ops import _align_method_FRAME
from pandas.io.formats import format as fmt
@@ -10870,27 +10871,11 @@ def _find_valid_index(self, how: str):
-------
idx_first_valid : type of index
"""
- assert how in ["first", "last"]
- if len(self) == 0: # early stop
+ idxpos = find_valid_index(self._values, how)
+ if idxpos is None:
return None
- is_valid = ~self.isna()
-
- if self.ndim == 2:
- is_valid = is_valid.any(1) # reduce axis 1
-
- if how == "first":
- idxpos = is_valid.values[::].argmax()
-
- if how == "last":
- idxpos = len(self) - 1 - is_valid.values[::-1].argmax()
-
- chk_notna = is_valid.iat[idxpos]
- idx = self.index[idxpos]
-
- if not chk_notna:
- return None
- return idx
+ return self.index[idxpos]
@Appender(
_shared_docs["valid_index"] % {"position": "first", "klass": "Series/DataFrame"}
diff --git a/pandas/core/missing.py b/pandas/core/missing.py
index 5a1bf6d37b081..c1e63a49a0f0a 100644
--- a/pandas/core/missing.py
+++ b/pandas/core/missing.py
@@ -128,6 +128,43 @@ def clean_interp_method(method, **kwargs):
return method
+def find_valid_index(values, how: str):
+ """
+ Retrieves the index of the first valid value.
+
+ Parameters
+ ----------
+ values : ndarray or ExtensionArray
+ how : {'first', 'last'}
+ Use this parameter to change between the first or last valid index.
+
+ Returns
+ -------
+ int or None
+ """
+ assert how in ["first", "last"]
+
+ if len(values) == 0: # early stop
+ return None
+
+ is_valid = ~isna(values)
+
+ if values.ndim == 2:
+ is_valid = is_valid.any(1) # reduce axis 1
+
+ if how == "first":
+ idxpos = is_valid[::].argmax()
+
+ if how == "last":
+ idxpos = len(values) - 1 - is_valid[::-1].argmax()
+
+ chk_notna = is_valid[idxpos]
+
+ if not chk_notna:
+ return None
+ return idxpos
+
+
def interpolate_1d(
xvalues,
yvalues,
@@ -192,14 +229,10 @@ def interpolate_1d(
# default limit is unlimited GH #16282
limit = algos._validate_limit(nobs=None, limit=limit)
- from pandas import Series
-
- ys = Series(yvalues)
-
# These are sets of index pointers to invalid values... i.e. {0, 1, etc...
all_nans = set(np.flatnonzero(invalid))
- start_nans = set(range(ys.first_valid_index()))
- end_nans = set(range(1 + ys.last_valid_index(), len(valid)))
+ start_nans = set(range(find_valid_index(yvalues, "first")))
+ end_nans = set(range(1 + find_valid_index(yvalues, "last"), len(valid)))
mid_nans = all_nans - start_nans - end_nans
# Like the sets above, preserve_nans contains indices of invalid values,
| The implementation here operates on the values (ndarray or EA) instead of on the Series/DataFrame.
This lets us avoid a runtime import of Series, so core.missing joins the Simple Dependencies Club. | https://api.github.com/repos/pandas-dev/pandas/pulls/29400 | 2019-11-04T21:43:44Z | 2019-11-05T17:13:17Z | 2019-11-05T17:13:16Z | 2019-11-05T17:47:01Z |
PR09 Batch 2 | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 2fb552af717fc..1a2f906f97152 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -1073,7 +1073,7 @@ def argmax(self, axis=None, skipna=True, *args, **kwargs):
Parameters
----------
axis : {None}
- Dummy argument for consistency with Series
+ Dummy argument for consistency with Series.
skipna : bool, default True
Returns
@@ -1096,7 +1096,7 @@ def min(self, axis=None, skipna=True, *args, **kwargs):
Parameters
----------
axis : {None}
- Dummy argument for consistency with Series
+ Dummy argument for consistency with Series.
skipna : bool, default True
Returns
@@ -1137,7 +1137,7 @@ def argmin(self, axis=None, skipna=True, *args, **kwargs):
Parameters
----------
axis : {None}
- Dummy argument for consistency with Series
+ Dummy argument for consistency with Series.
skipna : bool, default True
Returns
@@ -1486,7 +1486,7 @@ def memory_usage(self, deep=False):
----------
deep : bool
Introspect the data deeply, interrogate
- `object` dtypes for system-level memory consumption
+ `object` dtypes for system-level memory consumption.
Returns
-------
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 301426d237d19..feee6dca23ac8 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -176,11 +176,11 @@ class Index(IndexOpsMixin, PandasObject):
If an actual dtype is provided, we coerce to that dtype if it's safe.
Otherwise, an error will be raised.
copy : bool
- Make a copy of input ndarray
+ Make a copy of input ndarray.
name : object
- Name to be stored in the index
+ Name to be stored in the index.
tupleize_cols : bool (default: True)
- When True, attempt to create a MultiIndex if possible
+ When True, attempt to create a MultiIndex if possible.
See Also
--------
@@ -791,13 +791,13 @@ def astype(self, dtype, copy=True):
Parameters
----------
indices : list
- Indices to be taken
+ Indices to be taken.
axis : int, optional
The axis over which to select values, always 0.
allow_fill : bool, default True
fill_value : bool, default None
If allow_fill=True and fill_value is not None, indices specified by
- -1 is regarded as NA. If Index doesn't hold NA, raise ValueError
+ -1 is regarded as NA. If Index doesn't hold NA, raise ValueError.
Returns
-------
@@ -1077,7 +1077,7 @@ def to_native_types(self, slicer=None, **kwargs):
2) quoting : bool or None
Whether or not there are quoted values in `self`
3) date_format : str
- The format used to represent date-like values
+ The format used to represent date-like values.
Returns
-------
@@ -2001,7 +2001,7 @@ def notna(self):
downcast : dict, default is None
a dict of item->dtype of what to downcast if possible,
or the string 'infer' which will try to downcast to an appropriate
- equal type (e.g. float64 to int64 if possible)
+ equal type (e.g. float64 to int64 if possible).
Returns
-------
@@ -2056,7 +2056,7 @@ def dropna(self, how="any"):
Parameters
----------
level : int or str, optional, default None
- Only return values from specified level (for MultiIndex)
+ Only return values from specified level (for MultiIndex).
.. versionadded:: 0.23.0
@@ -3413,7 +3413,7 @@ def _reindex_non_unique(self, target):
return_indexers : bool, default False
sort : bool, default False
Sort the join keys lexicographically in the result Index. If False,
- the order of the join keys depends on the join type (how keyword)
+ the order of the join keys depends on the join type (how keyword).
Returns
-------
@@ -4923,9 +4923,9 @@ def slice_indexer(self, start=None, end=None, step=None, kind=None):
Parameters
----------
start : label, default None
- If None, defaults to the beginning
+ If None, defaults to the beginning.
end : label, default None
- If None, defaults to the end
+ If None, defaults to the end.
step : int, default None
kind : str, default None
@@ -5122,11 +5122,11 @@ def slice_locs(self, start=None, end=None, step=None, kind=None):
Parameters
----------
start : label, default None
- If None, defaults to the beginning
+ If None, defaults to the beginning.
end : label, default None
- If None, defaults to the end
+ If None, defaults to the end.
step : int, defaults None
- If None, defaults to 1
+ If None, defaults to 1.
kind : {'ix', 'loc', 'getitem'} or None
Returns
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index 66deacac37789..caaf55546189c 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -737,19 +737,18 @@ def _set_levels(
def set_levels(self, levels, level=None, inplace=False, verify_integrity=True):
"""
- Set new levels on MultiIndex. Defaults to returning
- new index.
+ Set new levels on MultiIndex. Defaults to returning new index.
Parameters
----------
levels : sequence or list of sequence
- new level(s) to apply
+ New level(s) to apply.
level : int, level name, or sequence of int/level names (default None)
- level(s) to set (None for all levels)
+ Level(s) to set (None for all levels).
inplace : bool
- if True, mutates in place
+ If True, mutates in place.
verify_integrity : bool (default True)
- if True, checks that levels and codes are compatible
+ If True, checks that levels and codes are compatible.
Returns
-------
diff --git a/pandas/core/indexes/numeric.py b/pandas/core/indexes/numeric.py
index e83360dc701f3..46bb8eafee3b9 100644
--- a/pandas/core/indexes/numeric.py
+++ b/pandas/core/indexes/numeric.py
@@ -176,9 +176,9 @@ def _union(self, other, sort):
data : array-like (1-dimensional)
dtype : NumPy dtype (default: %(dtype)s)
copy : bool
- Make a copy of input ndarray
+ Make a copy of input ndarray.
name : object
- Name to be stored in the index
+ Name to be stored in the index.
Attributes
----------
diff --git a/pandas/core/indexes/range.py b/pandas/core/indexes/range.py
index 6e2d500f4c5ab..5fa3431fc97c0 100644
--- a/pandas/core/indexes/range.py
+++ b/pandas/core/indexes/range.py
@@ -51,7 +51,7 @@ class RangeIndex(Int64Index):
stop : int (default: 0)
step : int (default: 1)
name : object, optional
- Name to be stored in the index
+ Name to be stored in the index.
copy : bool, default False
Unused, accepted for homogeneity with other index types.
diff --git a/pandas/core/reshape/tile.py b/pandas/core/reshape/tile.py
index 95534755b8beb..073bb4707f890 100644
--- a/pandas/core/reshape/tile.py
+++ b/pandas/core/reshape/tile.py
@@ -275,17 +275,18 @@ def qcut(
duplicates: str = "raise",
):
"""
- Quantile-based discretization function. Discretize variable into
- equal-sized buckets based on rank or based on sample quantiles. For example
- 1000 values for 10 quantiles would produce a Categorical object indicating
- quantile membership for each data point.
+ Quantile-based discretization function.
+
+ Discretize variable into equal-sized buckets based on rank or based
+ on sample quantiles. For example 1000 values for 10 quantiles would
+ produce a Categorical object indicating quantile membership for each data point.
Parameters
----------
x : 1d ndarray or Series
q : int or list-like of int
Number of quantiles. 10 for deciles, 4 for quartiles, etc. Alternately
- array of quantiles, e.g. [0, .25, .5, .75, 1.] for quartiles
+ array of quantiles, e.g. [0, .25, .5, .75, 1.] for quartiles.
labels : array or bool, default None
Used as labels for the resulting bins. Must be of the same length as
the resulting bins. If False, return only integer indicators of the
| Another batch of commits for #28602. Also fixes a few summary formatting errors and PR08 capitalization errors. | https://api.github.com/repos/pandas-dev/pandas/pulls/29396 | 2019-11-04T14:58:19Z | 2019-11-04T16:11:23Z | 2019-11-04T16:11:23Z | 2020-01-06T16:47:02Z |
DOC: remove okwarning once pyarrow 0.12 is released | diff --git a/doc/source/user_guide/io.rst b/doc/source/user_guide/io.rst
index 173bcf7537154..f9fbc33cba966 100644
--- a/doc/source/user_guide/io.rst
+++ b/doc/source/user_guide/io.rst
@@ -4685,7 +4685,6 @@ Write to a feather file.
Read from a feather file.
.. ipython:: python
- :okwarning:
result = pd.read_feather('example.feather')
result
@@ -4764,7 +4763,6 @@ Write to a parquet file.
Read from a parquet file.
.. ipython:: python
- :okwarning:
result = pd.read_parquet('example_fp.parquet', engine='fastparquet')
result = pd.read_parquet('example_pa.parquet', engine='pyarrow')
@@ -4839,7 +4837,6 @@ Partitioning Parquet files
Parquet supports partitioning of data based on the values of one or more columns.
.. ipython:: python
- :okwarning:
df = pd.DataFrame({'a': [0, 0, 1, 1], 'b': [0, 1, 0, 1]})
df.to_parquet(fname='test', engine='pyarrow',
diff --git a/environment.yml b/environment.yml
index 4c96ab815dc90..443dc483aedf8 100644
--- a/environment.yml
+++ b/environment.yml
@@ -81,7 +81,7 @@ dependencies:
- html5lib # pandas.read_html
- lxml # pandas.read_html
- openpyxl # pandas.read_excel, DataFrame.to_excel, pandas.ExcelWriter, pandas.ExcelFile
- - pyarrow>=0.13.1 # pandas.read_paquet, DataFrame.to_parquet, pandas.read_feather, DataFrame.to_feather
+ - pyarrow>=0.13.1 # pandas.read_parquet, DataFrame.to_parquet, pandas.read_feather, DataFrame.to_feather
- pyqt>=5.9.2 # pandas.read_clipboard
- pytables>=3.4.2 # pandas.read_hdf, DataFrame.to_hdf
- python-snappy # required by pyarrow
| - [x] closes #24617
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29395 | 2019-11-04T14:05:01Z | 2019-11-04T14:53:10Z | 2019-11-04T14:53:09Z | 2019-11-04T14:53:10Z |
BUG: GH25495 incorrect dtype when using .loc to set Categorical value for column in 1-row DataFrame | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index f0ba1250b7f8d..c9e2e7e133133 100755
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -1086,6 +1086,7 @@ Indexing
- Bug when indexing with ``.loc`` where the index was a :class:`CategoricalIndex` with non-string categories didn't work (:issue:`17569`, :issue:`30225`)
- :meth:`Index.get_indexer_non_unique` could fail with ``TypeError`` in some cases, such as when searching for ints in a string index (:issue:`28257`)
- Bug in :meth:`Float64Index.get_loc` incorrectly raising ``TypeError`` instead of ``KeyError`` (:issue:`29189`)
+- Bug in :meth:`DataFrame.loc` with incorrect dtype when setting Categorical value in 1-row DataFrame (:issue:`25495`)
- :meth:`MultiIndex.get_loc` can't find missing values when input includes missing values (:issue:`19132`)
- Bug in :meth:`Series.__setitem__` incorrectly assigning values with boolean indexer when the length of new data matches the number of ``True`` values and new data is not a ``Series`` or an ``np.array`` (:issue:`30567`)
- Bug in indexing with a :class:`PeriodIndex` incorrectly accepting integers representing years, use e.g. ``ser.loc["2007"]`` instead of ``ser.loc[2007]`` (:issue:`30763`)
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index a93211edf162b..43edc246da6dd 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -876,7 +876,11 @@ def setitem(self, indexer, value):
# length checking
check_setitem_lengths(indexer, value, values)
-
+ exact_match = (
+ len(arr_value.shape)
+ and arr_value.shape[0] == values.shape[0]
+ and arr_value.size == values.size
+ )
if is_empty_indexer(indexer, arr_value):
# GH#8669 empty indexers
pass
@@ -886,14 +890,21 @@ def setitem(self, indexer, value):
# be e.g. a list; see GH#6043
values[indexer] = value
- # if we are an exact match (ex-broadcasting),
- # then use the resultant dtype
elif (
- len(arr_value.shape)
- and arr_value.shape[0] == values.shape[0]
- and arr_value.size == values.size
+ exact_match
+ and is_categorical_dtype(arr_value.dtype)
+ and not is_categorical_dtype(values)
):
+ # GH25495 - If the current dtype is not categorical,
+ # we need to create a new categorical block
values[indexer] = value
+ return self.make_block(Categorical(self.values, dtype=arr_value.dtype))
+
+ # if we are an exact match (ex-broadcasting),
+ # then use the resultant dtype
+ elif exact_match:
+ values[indexer] = value
+
try:
values = values.astype(arr_value.dtype)
except ValueError:
diff --git a/pandas/tests/frame/indexing/test_categorical.py b/pandas/tests/frame/indexing/test_categorical.py
index 5de38915f04c1..a29c193676db2 100644
--- a/pandas/tests/frame/indexing/test_categorical.py
+++ b/pandas/tests/frame/indexing/test_categorical.py
@@ -354,6 +354,16 @@ def test_functions_no_warnings(self):
df.value, range(0, 105, 10), right=False, labels=labels
)
+ def test_setitem_single_row_categorical(self):
+ # GH 25495
+ df = DataFrame({"Alpha": ["a"], "Numeric": [0]})
+ categories = pd.Categorical(df["Alpha"], categories=["a", "b", "c"])
+ df.loc[:, "Alpha"] = categories
+
+ result = df["Alpha"]
+ expected = Series(categories, index=df.index, name="Alpha")
+ tm.assert_series_equal(result, expected)
+
def test_loc_indexing_preserves_index_category_dtype(self):
# GH 15166
df = DataFrame(
| - [x] closes https://github.com/pandas-dev/pandas/issues/25495
- [x] 1 test added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29393 | 2019-11-04T09:25:45Z | 2020-01-27T12:34:43Z | 2020-01-27T12:34:43Z | 2020-01-28T15:29:45Z |
TST: Test DataFrame.rolling with window as string | diff --git a/pandas/tests/window/test_rolling.py b/pandas/tests/window/test_rolling.py
index 70ba85120af3c..72b72b31d8faa 100644
--- a/pandas/tests/window/test_rolling.py
+++ b/pandas/tests/window/test_rolling.py
@@ -1,4 +1,4 @@
-from datetime import timedelta
+from datetime import datetime, timedelta
import numpy as np
import pytest
@@ -7,7 +7,7 @@
import pandas.util._test_decorators as td
import pandas as pd
-from pandas import DataFrame, Series
+from pandas import DataFrame, Index, Series
from pandas.core.window import Rolling
from pandas.tests.window.common import Base
import pandas.util.testing as tm
@@ -361,3 +361,60 @@ def test_rolling_datetime(self, axis_frame, tz_naive_fixture):
}
)
tm.assert_frame_equal(result, expected)
+
+
+def test_rolling_window_as_string():
+ # see gh-22590
+ date_today = datetime.now()
+ days = pd.date_range(date_today, date_today + timedelta(365), freq="D")
+
+ npr = np.random.RandomState(seed=421)
+
+ data = npr.randint(1, high=100, size=len(days))
+ df = DataFrame({"DateCol": days, "metric": data})
+
+ df.set_index("DateCol", inplace=True)
+ result = df.rolling(window="21D", min_periods=2, closed="left")["metric"].agg("max")
+
+ expData = (
+ [np.nan] * 2
+ + [88.0] * 16
+ + [97.0] * 9
+ + [98.0]
+ + [99.0] * 21
+ + [95.0] * 16
+ + [93.0] * 5
+ + [89.0] * 5
+ + [96.0] * 21
+ + [94.0] * 14
+ + [90.0] * 13
+ + [88.0] * 2
+ + [90.0] * 9
+ + [96.0] * 21
+ + [95.0] * 6
+ + [91.0]
+ + [87.0] * 6
+ + [92.0] * 21
+ + [83.0] * 2
+ + [86.0] * 10
+ + [87.0] * 5
+ + [98.0] * 21
+ + [97.0] * 14
+ + [93.0] * 7
+ + [87.0] * 4
+ + [86.0] * 4
+ + [95.0] * 21
+ + [85.0] * 14
+ + [83.0] * 2
+ + [76.0] * 5
+ + [81.0] * 2
+ + [98.0] * 21
+ + [95.0] * 14
+ + [91.0] * 7
+ + [86.0]
+ + [93.0] * 3
+ + [95.0] * 20
+ )
+
+ expected = Series(expData, index=Index(days, name="DateCol"), name="metric")
+ tm.assert_series_equal(result, expected)
| Closes https://github.com/pandas-dev/pandas/issues/22590 | https://api.github.com/repos/pandas-dev/pandas/pulls/29392 | 2019-11-04T02:51:56Z | 2019-11-04T22:10:32Z | 2019-11-04T22:10:32Z | 2019-11-04T23:56:07Z |
Fix: DataFrame.append with empty list raises IndexError | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 101c5ec9137fc..19bba71edf7f2 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -233,7 +233,7 @@ Removal of prior version deprecations/changes
Previously, pandas would register converters with matplotlib as a side effect of importing pandas (:issue:`18720`).
This changed the output of plots made via matplotlib plots after pandas was imported, even if you were using
-matplotlib directly rather than rather than :meth:`~DataFrame.plot`.
+matplotlib directly rather than :meth:`~DataFrame.plot`.
To use pandas formatters with a matplotlib plot, specify
@@ -430,6 +430,7 @@ Reshaping
- :func:`qcut` and :func:`cut` now handle boolean input (:issue:`20303`)
- Fix to ensure all int dtypes can be used in :func:`merge_asof` when using a tolerance value. Previously every non-int64 type would raise an erroneous ``MergeError`` (:issue:`28870`).
- Better error message in :func:`get_dummies` when `columns` isn't a list-like value (:issue:`28383`)
+- :meth:`DataFrame.append` raised an ``IndexError`` when passed an empty ``list`` (:issue:`28769`)
Sparse
^^^^^^
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 40efc4c65476a..824afc8bcc9b4 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -6968,6 +6968,8 @@ def append(self, other, ignore_index=False, verify_integrity=False, sort=None):
3 3
4 4
"""
+ from pandas.core.reshape.concat import concat
+
if isinstance(other, (Series, dict)):
if isinstance(other, dict):
other = Series(other)
@@ -6998,17 +7000,16 @@ def append(self, other, ignore_index=False, verify_integrity=False, sort=None):
other = other._convert(datetime=True, timedelta=True)
if not self.columns.equals(combined_columns):
self = self.reindex(columns=combined_columns)
- elif isinstance(other, list) and not isinstance(other[0], DataFrame):
+ elif isinstance(other, list) and other and not isinstance(other[0], DataFrame):
other = DataFrame(other)
if (self.columns.get_indexer(other.columns) >= 0).all():
other = other.reindex(columns=self.columns)
- from pandas.core.reshape.concat import concat
-
- if isinstance(other, (list, tuple)):
+ if isinstance(other, list):
to_concat = [self] + other
else:
to_concat = [self, other]
+
return concat(
to_concat,
ignore_index=ignore_index,
diff --git a/pandas/tests/frame/test_combine_concat.py b/pandas/tests/frame/test_combine_concat.py
index e3f37e1ef3186..0ffed94fd67c9 100644
--- a/pandas/tests/frame/test_combine_concat.py
+++ b/pandas/tests/frame/test_combine_concat.py
@@ -182,6 +182,22 @@ def test_append_list_of_series_dicts(self):
expected = df.append(DataFrame(dicts), ignore_index=True, sort=True)
tm.assert_frame_equal(result, expected)
+ def test_append_empty_list(self, float_frame):
+ result = float_frame.append([])
+ expected = float_frame
+ tm.assert_frame_equal(result, expected)
+
+ def test_append_list_of_frames(self, float_frame):
+ more_dfs = [float_frame] * 2
+ result = float_frame.append(more_dfs)
+ expected = pd.concat([float_frame] + more_dfs)
+ tm.assert_frame_equal(result, expected)
+
+ def test_append_tuple_of_frames(self, float_frame):
+ more_dfs = (float_frame, float_frame)
+ with pytest.raises(TypeError):
+ float_frame.append(more_dfs)
+
def test_append_missing_cols(self):
# GH22252
# exercise the conditional branch in append method where the data
| - [x] closes #28769
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29391 | 2019-11-04T00:17:03Z | 2019-11-04T00:22:31Z | null | 2019-11-04T00:22:31Z |
changing docstring for to_csv compression to 1.0.0 | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index f88c26c7bc782..cbeee88d75b51 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -3221,7 +3221,7 @@ def to_csv(
and mode is 'zip' or inferred as 'zip', other entries passed as
additional compression options.
- .. versionchanged:: 0.25.0
+ .. versionchanged:: 1.0.0
May now be a dict with key 'method' as compression mode
and other entries as additional compression options if
| - closes #29328
- Just updating doc-string for to_csv() compression.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29390 | 2019-11-04T00:06:07Z | 2019-11-04T13:50:14Z | 2019-11-04T13:50:14Z | 2019-11-04T13:50:14Z |
CLN core.groupby | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 9586d49c555ff..2fb552af717fc 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -586,9 +586,16 @@ def _aggregate_multiple_funcs(self, arg, _level, _axis):
new_res = colg.aggregate(arg)
except (TypeError, DataError):
pass
- except ValueError:
+ except ValueError as err:
# cannot aggregate
- continue
+ if "Must produce aggregated value" in str(err):
+ # raised directly in _aggregate_named
+ pass
+ elif "no results" in str(err):
+ # raised direcly in _aggregate_multiple_funcs
+ pass
+ else:
+ raise
else:
results.append(new_res)
keys.append(col)
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 996c178bd7feb..009e83b861523 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -244,7 +244,7 @@ def aggregate(self, func=None, *args, **kwargs):
if isinstance(func, str):
return getattr(self, func)(*args, **kwargs)
- if isinstance(func, abc.Iterable):
+ elif isinstance(func, abc.Iterable):
# Catch instances of lists / tuples
# but not the class list / tuple itself.
func = _maybe_mangle_lambdas(func)
@@ -261,8 +261,6 @@ def aggregate(self, func=None, *args, **kwargs):
try:
return self._python_agg_general(func, *args, **kwargs)
- except (AssertionError, TypeError):
- raise
except (ValueError, KeyError, AttributeError, IndexError):
# TODO: IndexError can be removed here following GH#29106
# TODO: AttributeError is caused by _index_data hijinx in
@@ -325,7 +323,7 @@ def _aggregate_multiple_funcs(self, arg, _level):
if name in results:
raise SpecificationError(
"Function names must be unique, found multiple named "
- "{}".format(name)
+ "{name}".format(name=name)
)
# reset the cache so that we
@@ -1464,8 +1462,6 @@ def _transform_item_by_item(self, obj, wrapper):
for i, col in enumerate(obj):
try:
output[col] = self[col].transform(wrapper)
- except AssertionError:
- raise
except TypeError:
# e.g. trying to call nanmean with string values
pass
@@ -1538,8 +1534,8 @@ def filter(self, func, dropna=True, *args, **kwargs):
else:
# non scalars aren't allowed
raise TypeError(
- "filter function returned a %s, "
- "but expected a scalar bool" % type(res).__name__
+ "filter function returned a {typ}, "
+ "but expected a scalar bool".format(typ=type(res).__name__)
)
return self._apply_filter(indices, dropna)
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 404da096d8535..642b1e93a057a 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -344,7 +344,7 @@ def __init__(
self,
obj: NDFrame,
keys=None,
- axis=0,
+ axis: int = 0,
level=None,
grouper=None,
exclusions=None,
@@ -561,7 +561,9 @@ def __getattr__(self, attr):
return self[attr]
raise AttributeError(
- "%r object has no attribute %r" % (type(self).__name__, attr)
+ "'{typ}' object has no attribute '{attr}'".format(
+ typ=type(self).__name__, attr=attr
+ )
)
@Substitution(
@@ -2486,6 +2488,6 @@ def groupby(obj, by, **kwds):
klass = DataFrameGroupBy
else:
- raise TypeError("invalid type: {}".format(obj))
+ raise TypeError("invalid type: {obj}".format(obj=obj))
return klass(obj, by, **kwds)
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index d7eaaca5ac83a..45d2a819ae5ad 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -172,7 +172,9 @@ def _set_grouper(self, obj, sort=False):
ax = self._grouper.take(obj.index)
else:
if key not in obj._info_axis:
- raise KeyError("The grouper name {0} is not found".format(key))
+ raise KeyError(
+ "The grouper name {key} is not found".format(key=key)
+ )
ax = Index(obj[key], name=key)
else:
@@ -188,7 +190,9 @@ def _set_grouper(self, obj, sort=False):
else:
if level not in (0, ax.name):
- raise ValueError("The level {0} is not valid".format(level))
+ raise ValueError(
+ "The level {level} is not valid".format(level=level)
+ )
# possibly sort
if (self.sort or sort) and not ax.is_monotonic:
@@ -278,7 +282,9 @@ def __init__(
if level is not None:
if not isinstance(level, int):
if level not in index.names:
- raise AssertionError("Level {} not in index".format(level))
+ raise AssertionError(
+ "Level {level} not in index".format(level=level)
+ )
level = index.names.index(level)
if self.name is None:
@@ -344,7 +350,7 @@ def __init__(
):
if getattr(self.grouper, "ndim", 1) != 1:
t = self.name or str(type(self.grouper))
- raise ValueError("Grouper for '{}' not 1-dimensional".format(t))
+ raise ValueError("Grouper for '{t}' not 1-dimensional".format(t=t))
self.grouper = self.index.map(self.grouper)
if not (
hasattr(self.grouper, "__len__")
@@ -352,7 +358,9 @@ def __init__(
):
errmsg = (
"Grouper result violates len(labels) == "
- "len(data)\nresult: %s" % pprint_thing(self.grouper)
+ "len(data)\nresult: {grper}".format(
+ grper=pprint_thing(self.grouper)
+ )
)
self.grouper = None # Try for sanity
raise AssertionError(errmsg)
@@ -426,7 +434,7 @@ def groups(self):
def _get_grouper(
obj: NDFrame,
key=None,
- axis=0,
+ axis: int = 0,
level=None,
sort=True,
observed=False,
@@ -493,7 +501,9 @@ def _get_grouper(
if isinstance(level, str):
if obj.index.name != level:
raise ValueError(
- "level name {} is not the name of the index".format(level)
+ "level name {level} is not the name of the index".format(
+ level=level
+ )
)
elif level > 0 or level < -1:
raise ValueError("level > 0 or level < -1 only valid with MultiIndex")
@@ -582,7 +592,7 @@ def _get_grouper(
exclusions = []
# if the actual grouper should be obj[key]
- def is_in_axis(key):
+ def is_in_axis(key) -> bool:
if not _is_label_like(key):
items = obj._data.items
try:
@@ -594,7 +604,7 @@ def is_in_axis(key):
return True
# if the grouper is obj[name]
- def is_in_obj(gpr):
+ def is_in_obj(gpr) -> bool:
if not hasattr(gpr, "name"):
return False
try:
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 8d13c37270d7a..7918e463c73ac 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,6 +7,7 @@
"""
import collections
+from typing import List, Optional
import numpy as np
@@ -385,7 +386,7 @@ def get_func(fname):
return func
- def _cython_operation(self, kind, values, how, axis, min_count=-1, **kwargs):
+ def _cython_operation(self, kind: str, values, how, axis, min_count=-1, **kwargs):
assert kind in ["transform", "aggregate"]
orig_values = values
@@ -398,16 +399,18 @@ def _cython_operation(self, kind, values, how, axis, min_count=-1, **kwargs):
# categoricals are only 1d, so we
# are not setup for dim transforming
if is_categorical_dtype(values) or is_sparse(values):
- raise NotImplementedError("{} dtype not supported".format(values.dtype))
+ raise NotImplementedError(
+ "{dtype} dtype not supported".format(dtype=values.dtype)
+ )
elif is_datetime64_any_dtype(values):
if how in ["add", "prod", "cumsum", "cumprod"]:
raise NotImplementedError(
- "datetime64 type does not support {} operations".format(how)
+ "datetime64 type does not support {how} operations".format(how=how)
)
elif is_timedelta64_dtype(values):
if how in ["prod", "cumprod"]:
raise NotImplementedError(
- "timedelta64 type does not support {} operations".format(how)
+ "timedelta64 type does not support {how} operations".format(how=how)
)
if is_datetime64tz_dtype(values.dtype):
@@ -513,7 +516,7 @@ def _cython_operation(self, kind, values, how, axis, min_count=-1, **kwargs):
result = result[:, 0]
if how in self._name_functions:
- names = self._name_functions[how]()
+ names = self._name_functions[how]() # type: Optional[List[str]]
else:
names = None
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index 13cb0f9aed303..e68a2efc3f4e6 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -361,8 +361,6 @@ def _groupby_and_aggregate(self, how, grouper=None, *args, **kwargs):
result = grouped._aggregate_item_by_item(how, *args, **kwargs)
else:
result = grouped.aggregate(how, *args, **kwargs)
- except AssertionError:
- raise
except DataError:
# we have a non-reducing function; try to evaluate
result = grouped.apply(how, *args, **kwargs)
@@ -1450,7 +1448,7 @@ def _get_resampler(self, obj, kind=None):
raise TypeError(
"Only valid with DatetimeIndex, "
"TimedeltaIndex or PeriodIndex, "
- "but got an instance of %r" % type(ax).__name__
+ "but got an instance of '{typ}'".format(typ=type(ax).__name__)
)
def _get_grouper(self, obj, validate=True):
@@ -1463,7 +1461,7 @@ def _get_time_bins(self, ax):
if not isinstance(ax, DatetimeIndex):
raise TypeError(
"axis must be a DatetimeIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
if len(ax) == 0:
@@ -1539,7 +1537,7 @@ def _get_time_delta_bins(self, ax):
if not isinstance(ax, TimedeltaIndex):
raise TypeError(
"axis must be a TimedeltaIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
if not len(ax):
@@ -1564,7 +1562,7 @@ def _get_time_period_bins(self, ax):
if not isinstance(ax, DatetimeIndex):
raise TypeError(
"axis must be a DatetimeIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
freq = self.freq
@@ -1586,7 +1584,7 @@ def _get_period_bins(self, ax):
if not isinstance(ax, PeriodIndex):
raise TypeError(
"axis must be a PeriodIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
memb = ax.asfreq(self.freq, how=self.convention)
| Broken off from local branches doing non-CLN work.
Foreshadowing: some of the TypeErrors we are catching are being caused by `self.axis` not being an Index instead of an int in some cases. | https://api.github.com/repos/pandas-dev/pandas/pulls/29389 | 2019-11-03T22:18:33Z | 2019-11-04T13:38:04Z | 2019-11-04T13:38:04Z | 2019-11-04T14:55:19Z |
API: drop kwargs from Series.dropna, add explicit `how` parameter | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 101c5ec9137fc..8a481f194d408 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -192,6 +192,8 @@ Other API changes
Now, pandas custom formatters will only be applied to plots created by pandas, through :meth:`~DataFrame.plot`.
Previously, pandas' formatters would be applied to all plots created *after* a :meth:`~DataFrame.plot`.
See :ref:`units registration <whatsnew_1000.matplotlib_units>` for more.
+- :meth:`Series.dropna` has dropped its ``**kwargs`` argument in favor of a single ``how`` parameter.
+ Supplying anything else than ``how`` to ``**kwargs`` raised a ``TypeError`` previously (:issue:`29388`)
-
diff --git a/pandas/core/series.py b/pandas/core/series.py
index e57de0e69b366..7b65816dc06b9 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -4595,7 +4595,7 @@ def notna(self):
def notnull(self):
return super().notnull()
- def dropna(self, axis=0, inplace=False, **kwargs):
+ def dropna(self, axis=0, inplace=False, how=None):
"""
Return a new Series with missing values removed.
@@ -4608,8 +4608,8 @@ def dropna(self, axis=0, inplace=False, **kwargs):
There is only one axis to drop values from.
inplace : bool, default False
If True, do operation inplace and return None.
- **kwargs
- Not in use.
+ how : str, optional
+ Not in use. Kept for compatibility.
Returns
-------
@@ -4667,12 +4667,6 @@ def dropna(self, axis=0, inplace=False, **kwargs):
dtype: object
"""
inplace = validate_bool_kwarg(inplace, "inplace")
- kwargs.pop("how", None)
- if kwargs:
- raise TypeError(
- "dropna() got an unexpected keyword "
- 'argument "{0}"'.format(list(kwargs.keys())[0])
- )
# Validate the axis parameter
self._get_axis_number(axis or 0)
| Using ``**kwargs`` gave false type hints on what the dropna method could take for arguments, and supplying anything but ``how`` raised a TypeError already.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29388 | 2019-11-03T21:54:23Z | 2019-11-04T19:58:48Z | 2019-11-04T19:58:48Z | 2019-11-04T19:58:52Z |
DOC: Added the flag "--no-use-pep517" to contrib guide. | diff --git a/README.md b/README.md
index c299241722b7e..158d48898a7bd 100644
--- a/README.md
+++ b/README.md
@@ -190,7 +190,7 @@ or for installing in [development mode](https://pip.pypa.io/en/latest/reference/
```sh
-python -m pip install --no-build-isolation -e .
+python -m pip install -e . --no-build-isolation --no-use-pep517
```
If you have `make`, you can also use `make develop` to run the same command.
diff --git a/doc/source/development/contributing.rst b/doc/source/development/contributing.rst
index 56fac1cb6852a..eed4a7862cc5f 100644
--- a/doc/source/development/contributing.rst
+++ b/doc/source/development/contributing.rst
@@ -208,7 +208,7 @@ We'll now kick off a three-step process:
# Build and install pandas
python setup.py build_ext --inplace -j 4
- python -m pip install -e . --no-build-isolation
+ python -m pip install -e . --no-build-isolation --no-use-pep517
At this point you should be able to import pandas from your locally built version::
@@ -255,7 +255,7 @@ You'll need to have at least python3.5 installed on your system.
# Build and install pandas
python setup.py build_ext --inplace -j 0
- python -m pip install -e . --no-build-isolation
+ python -m pip install -e . --no-build-isolation --no-use-pep517
**Windows**
| - [x] closes #28633
| https://api.github.com/repos/pandas-dev/pandas/pulls/29387 | 2019-11-03T21:19:14Z | 2019-11-04T13:42:42Z | 2019-11-04T13:42:42Z | 2019-11-04T13:42:42Z |
DOC: Added --no-use-pep517 flag to the developer guide. | diff --git a/doc/source/development/contributing.rst b/doc/source/development/contributing.rst
index 56fac1cb6852a..f66cc65774f6b 100644
--- a/doc/source/development/contributing.rst
+++ b/doc/source/development/contributing.rst
@@ -255,7 +255,7 @@ You'll need to have at least python3.5 installed on your system.
# Build and install pandas
python setup.py build_ext --inplace -j 0
- python -m pip install -e . --no-build-isolation
+ python -m pip install -e . --no-build-isolation --no-use-pep517
**Windows**
|
- [x] closes #28633
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` | https://api.github.com/repos/pandas-dev/pandas/pulls/29386 | 2019-11-03T20:28:49Z | 2019-11-03T21:08:15Z | null | 2019-11-03T21:08:19Z |
REF: simplify core.algorithms, reshape.cut | diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 06ba2a7e0ccfb..ceec0652b7ce2 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -8,7 +8,7 @@
import numpy as np
-from pandas._libs import algos, hashtable as htable, lib
+from pandas._libs import Timestamp, algos, hashtable as htable, lib
from pandas._libs.tslib import iNaT
from pandas.util._decorators import Appender, Substitution, deprecate_kwarg
@@ -1440,7 +1440,9 @@ def _take_nd_object(arr, indexer, out, axis: int, fill_value, mask_info):
}
-def _get_take_nd_function(ndim, arr_dtype, out_dtype, axis: int = 0, mask_info=None):
+def _get_take_nd_function(
+ ndim: int, arr_dtype, out_dtype, axis: int = 0, mask_info=None
+):
if ndim <= 2:
tup = (arr_dtype.name, out_dtype.name)
if ndim == 1:
@@ -1474,7 +1476,7 @@ def func2(arr, indexer, out, fill_value=np.nan):
return func2
-def take(arr, indices, axis=0, allow_fill: bool = False, fill_value=None):
+def take(arr, indices, axis: int = 0, allow_fill: bool = False, fill_value=None):
"""
Take elements from an array.
@@ -1568,13 +1570,7 @@ def take(arr, indices, axis=0, allow_fill: bool = False, fill_value=None):
def take_nd(
- arr,
- indexer,
- axis=0,
- out=None,
- fill_value=np.nan,
- mask_info=None,
- allow_fill: bool = True,
+ arr, indexer, axis: int = 0, out=None, fill_value=np.nan, allow_fill: bool = True
):
"""
Specialized Cython take which sets NaN values in one pass
@@ -1597,10 +1593,6 @@ def take_nd(
maybe_promote to determine this type for any fill_value
fill_value : any, default np.nan
Fill value to replace -1 values with
- mask_info : tuple of (ndarray, boolean)
- If provided, value should correspond to:
- (indexer != -1, (indexer != -1).any())
- If not provided, it will be computed internally if necessary
allow_fill : boolean, default True
If False, indexer is assumed to contain no -1 values so no filling
will be done. This short-circuits computation of a mask. Result is
@@ -1611,6 +1603,7 @@ def take_nd(
subarray : array-like
May be the same type as the input, or cast to an ndarray.
"""
+ mask_info = None
if is_extension_array_dtype(arr):
return arr.take(indexer, fill_value=fill_value, allow_fill=allow_fill)
@@ -1632,12 +1625,9 @@ def take_nd(
dtype, fill_value = maybe_promote(arr.dtype, fill_value)
if dtype != arr.dtype and (out is None or out.dtype != dtype):
# check if promotion is actually required based on indexer
- if mask_info is not None:
- mask, needs_masking = mask_info
- else:
- mask = indexer == -1
- needs_masking = mask.any()
- mask_info = mask, needs_masking
+ mask = indexer == -1
+ needs_masking = mask.any()
+ mask_info = mask, needs_masking
if needs_masking:
if out is not None and out.dtype != dtype:
raise TypeError("Incompatible type for fill_value")
@@ -1818,12 +1808,12 @@ def searchsorted(arr, value, side="left", sorter=None):
elif not (
is_object_dtype(arr) or is_numeric_dtype(arr) or is_categorical_dtype(arr)
):
- from pandas.core.series import Series
-
# E.g. if `arr` is an array with dtype='datetime64[ns]'
# and `value` is a pd.Timestamp, we may need to convert value
- value_ser = Series(value)._values
+ value_ser = array([value]) if is_scalar(value) else array(value)
value = value_ser[0] if is_scalar(value) else value_ser
+ if isinstance(value, Timestamp) and value.tzinfo is None:
+ value = value.to_datetime64()
result = arr.searchsorted(value, side=side, sorter=sorter)
return result
diff --git a/pandas/core/reshape/tile.py b/pandas/core/reshape/tile.py
index 09db840ca4db0..95534755b8beb 100644
--- a/pandas/core/reshape/tile.py
+++ b/pandas/core/reshape/tile.py
@@ -4,6 +4,7 @@
import numpy as np
from pandas._libs import Timedelta, Timestamp
+from pandas._libs.interval import Interval
from pandas._libs.lib import infer_dtype
from pandas.core.dtypes.common import (
@@ -18,17 +19,10 @@
is_scalar,
is_timedelta64_dtype,
)
+from pandas.core.dtypes.generic import ABCSeries
from pandas.core.dtypes.missing import isna
-from pandas import (
- Categorical,
- Index,
- Interval,
- IntervalIndex,
- Series,
- to_datetime,
- to_timedelta,
-)
+from pandas import Categorical, Index, IntervalIndex, to_datetime, to_timedelta
import pandas.core.algorithms as algos
import pandas.core.nanops as nanops
@@ -206,7 +200,8 @@ def cut(
# NOTE: this binning code is changed a bit from histogram for var(x) == 0
# for handling the cut for datetime and timedelta objects
- x_is_series, series_index, name, x = _preprocess_for_cut(x)
+ original = x
+ x = _preprocess_for_cut(x)
x, dtype = _coerce_to_type(x)
if not np.iterable(bins):
@@ -268,9 +263,7 @@ def cut(
duplicates=duplicates,
)
- return _postprocess_for_cut(
- fac, bins, retbins, x_is_series, series_index, name, dtype
- )
+ return _postprocess_for_cut(fac, bins, retbins, dtype, original)
def qcut(
@@ -333,8 +326,8 @@ def qcut(
>>> pd.qcut(range(5), 4, labels=False)
array([0, 0, 1, 2, 3])
"""
- x_is_series, series_index, name, x = _preprocess_for_cut(x)
-
+ original = x
+ x = _preprocess_for_cut(x)
x, dtype = _coerce_to_type(x)
if is_integer(q):
@@ -352,9 +345,7 @@ def qcut(
duplicates=duplicates,
)
- return _postprocess_for_cut(
- fac, bins, retbins, x_is_series, series_index, name, dtype
- )
+ return _postprocess_for_cut(fac, bins, retbins, dtype, original)
def _bins_to_cuts(
@@ -544,13 +535,6 @@ def _preprocess_for_cut(x):
input to array, strip the index information and store it
separately
"""
- x_is_series = isinstance(x, Series)
- series_index = None
- name = None
-
- if x_is_series:
- series_index = x.index
- name = x.name
# Check that the passed array is a Pandas or Numpy object
# We don't want to strip away a Pandas data-type here (e.g. datetimetz)
@@ -560,19 +544,17 @@ def _preprocess_for_cut(x):
if x.ndim != 1:
raise ValueError("Input array must be 1 dimensional")
- return x_is_series, series_index, name, x
+ return x
-def _postprocess_for_cut(
- fac, bins, retbins: bool, x_is_series, series_index, name, dtype
-):
+def _postprocess_for_cut(fac, bins, retbins: bool, dtype, original):
"""
handles post processing for the cut method where
we combine the index information if the originally passed
datatype was a series
"""
- if x_is_series:
- fac = Series(fac, index=series_index, name=name)
+ if isinstance(original, ABCSeries):
+ fac = original._constructor(fac, index=original.index, name=original.name)
if not retbins:
return fac
| https://api.github.com/repos/pandas-dev/pandas/pulls/29385 | 2019-11-03T19:34:03Z | 2019-11-04T13:40:47Z | 2019-11-04T13:40:47Z | 2019-11-04T14:59:54Z | |
REF: move safe_sort to algos to avoid private/circular dependencies | diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 06ba2a7e0ccfb..fc55bfbae0900 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -14,6 +14,7 @@
from pandas.core.dtypes.cast import (
construct_1d_object_array_from_listlike,
+ infer_dtype_from_array,
maybe_promote,
)
from pandas.core.dtypes.common import (
@@ -639,8 +640,6 @@ def factorize(values, sort: bool = False, order=None, na_sentinel=-1, size_hint=
)
if sort and len(uniques) > 0:
- from pandas.core.sorting import safe_sort
-
uniques, labels = safe_sort(
uniques, labels, na_sentinel=na_sentinel, assume_unique=True, verify=False
)
@@ -1920,3 +1919,138 @@ def diff(arr, n: int, axis: int = 0):
out_arr = out_arr.astype("int64").view("timedelta64[ns]")
return out_arr
+
+
+# --------------------------------------------------------------------
+# Helper functions
+
+# Note: safe_sort is in algorithms.py instead of sorting.py because it is
+# low-dependency, is used in this module, and used private methods from
+# this module.
+def safe_sort(
+ values,
+ labels=None,
+ na_sentinel: int = -1,
+ assume_unique: bool = False,
+ verify: bool = True,
+):
+ """
+ Sort ``values`` and reorder corresponding ``labels``.
+ ``values`` should be unique if ``labels`` is not None.
+ Safe for use with mixed types (int, str), orders ints before strs.
+
+ Parameters
+ ----------
+ values : list-like
+ Sequence; must be unique if ``labels`` is not None.
+ labels : list_like
+ Indices to ``values``. All out of bound indices are treated as
+ "not found" and will be masked with ``na_sentinel``.
+ na_sentinel : int, default -1
+ Value in ``labels`` to mark "not found".
+ Ignored when ``labels`` is None.
+ assume_unique : bool, default False
+ When True, ``values`` are assumed to be unique, which can speed up
+ the calculation. Ignored when ``labels`` is None.
+ verify : bool, default True
+ Check if labels are out of bound for the values and put out of bound
+ labels equal to na_sentinel. If ``verify=False``, it is assumed there
+ are no out of bound labels. Ignored when ``labels`` is None.
+
+ .. versionadded:: 0.25.0
+
+ Returns
+ -------
+ ordered : ndarray
+ Sorted ``values``
+ new_labels : ndarray
+ Reordered ``labels``; returned when ``labels`` is not None.
+
+ Raises
+ ------
+ TypeError
+ * If ``values`` is not list-like or if ``labels`` is neither None
+ nor list-like
+ * If ``values`` cannot be sorted
+ ValueError
+ * If ``labels`` is not None and ``values`` contain duplicates.
+ """
+ if not is_list_like(values):
+ raise TypeError(
+ "Only list-like objects are allowed to be passed to safe_sort as values"
+ )
+
+ if not isinstance(values, np.ndarray) and not is_extension_array_dtype(values):
+ # don't convert to string types
+ dtype, _ = infer_dtype_from_array(values)
+ values = np.asarray(values, dtype=dtype)
+
+ def sort_mixed(values):
+ # order ints before strings, safe in py3
+ str_pos = np.array([isinstance(x, str) for x in values], dtype=bool)
+ nums = np.sort(values[~str_pos])
+ strs = np.sort(values[str_pos])
+ return np.concatenate([nums, np.asarray(strs, dtype=object)])
+
+ sorter = None
+ if (
+ not is_extension_array_dtype(values)
+ and lib.infer_dtype(values, skipna=False) == "mixed-integer"
+ ):
+ # unorderable in py3 if mixed str/int
+ ordered = sort_mixed(values)
+ else:
+ try:
+ sorter = values.argsort()
+ ordered = values.take(sorter)
+ except TypeError:
+ # try this anyway
+ ordered = sort_mixed(values)
+
+ # labels:
+
+ if labels is None:
+ return ordered
+
+ if not is_list_like(labels):
+ raise TypeError(
+ "Only list-like objects or None are allowed to be"
+ "passed to safe_sort as labels"
+ )
+ labels = ensure_platform_int(np.asarray(labels))
+
+ from pandas import Index
+
+ if not assume_unique and not Index(values).is_unique:
+ raise ValueError("values should be unique if labels is not None")
+
+ if sorter is None:
+ # mixed types
+ hash_klass, values = _get_data_algo(values)
+ t = hash_klass(len(values))
+ t.map_locations(values)
+ sorter = ensure_platform_int(t.lookup(ordered))
+
+ if na_sentinel == -1:
+ # take_1d is faster, but only works for na_sentinels of -1
+ order2 = sorter.argsort()
+ new_labels = take_1d(order2, labels, fill_value=-1)
+ if verify:
+ mask = (labels < -len(values)) | (labels >= len(values))
+ else:
+ mask = None
+ else:
+ reverse_indexer = np.empty(len(sorter), dtype=np.int_)
+ reverse_indexer.put(sorter, np.arange(len(sorter)))
+ # Out of bound indices will be masked with `na_sentinel` next, so we
+ # may deal with them here without performance loss using `mode='wrap'`
+ new_labels = reverse_indexer.take(labels, mode="wrap")
+
+ mask = labels == na_sentinel
+ if verify:
+ mask = mask | (labels < -len(values)) | (labels >= len(values))
+
+ if mask is not None:
+ np.putmask(new_labels, mask, na_sentinel)
+
+ return ordered, ensure_platform_int(new_labels)
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 187c7e2f3a7f7..4c5b7442337fb 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -73,7 +73,6 @@
import pandas.core.missing as missing
from pandas.core.ops import get_op_result_name
from pandas.core.ops.invalid import make_invalid_op
-import pandas.core.sorting as sorting
from pandas.core.strings import StringMethods
from pandas.io.formats.printing import (
@@ -2507,7 +2506,7 @@ def _union(self, other, sort):
if sort is None:
try:
- result = sorting.safe_sort(result)
+ result = algos.safe_sort(result)
except TypeError as e:
warnings.warn(
"{}, sort order is undefined for "
@@ -2603,7 +2602,7 @@ def intersection(self, other, sort=False):
taken = other.take(indexer)
if sort is None:
- taken = sorting.safe_sort(taken.values)
+ taken = algos.safe_sort(taken.values)
if self.name != other.name:
name = None
else:
@@ -2673,7 +2672,7 @@ def difference(self, other, sort=None):
the_diff = this.values.take(label_diff)
if sort is None:
try:
- the_diff = sorting.safe_sort(the_diff)
+ the_diff = algos.safe_sort(the_diff)
except TypeError:
pass
@@ -2750,7 +2749,7 @@ def symmetric_difference(self, other, result_name=None, sort=None):
the_diff = concat_compat([left_diff, right_diff])
if sort is None:
try:
- the_diff = sorting.safe_sort(the_diff)
+ the_diff = algos.safe_sort(the_diff)
except TypeError:
pass
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index ea334503a4302..9845c570ca704 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -44,7 +44,6 @@
import pandas.core.common as com
from pandas.core.frame import _merge_doc
from pandas.core.internals import _transform_index, concatenate_block_managers
-import pandas.core.sorting as sorting
from pandas.core.sorting import is_int64_overflow_possible
@@ -1912,7 +1911,7 @@ def _sort_labels(uniques, left, right):
llength = len(left)
labels = np.concatenate([left, right])
- _, new_labels = sorting.safe_sort(uniques, labels, na_sentinel=-1)
+ _, new_labels = algos.safe_sort(uniques, labels, na_sentinel=-1)
new_labels = ensure_int64(new_labels)
new_left, new_right = new_labels[:llength], new_labels[llength:]
diff --git a/pandas/core/sorting.py b/pandas/core/sorting.py
index 9b8a1a76e419c..82eb93dd4c879 100644
--- a/pandas/core/sorting.py
+++ b/pandas/core/sorting.py
@@ -4,13 +4,11 @@
from pandas._libs import algos, hashtable, lib
from pandas._libs.hashtable import unique_label_indices
-from pandas.core.dtypes.cast import infer_dtype_from_array
from pandas.core.dtypes.common import (
ensure_int64,
ensure_platform_int,
is_categorical_dtype,
is_extension_array_dtype,
- is_list_like,
)
from pandas.core.dtypes.missing import isna
@@ -389,132 +387,3 @@ def _reorder_by_uniques(uniques, labels):
uniques = algorithms.take_nd(uniques, sorter, allow_fill=False)
return uniques, labels
-
-
-def safe_sort(
- values,
- labels=None,
- na_sentinel: int = -1,
- assume_unique: bool = False,
- verify: bool = True,
-):
- """
- Sort ``values`` and reorder corresponding ``labels``.
- ``values`` should be unique if ``labels`` is not None.
- Safe for use with mixed types (int, str), orders ints before strs.
-
- Parameters
- ----------
- values : list-like
- Sequence; must be unique if ``labels`` is not None.
- labels : list_like
- Indices to ``values``. All out of bound indices are treated as
- "not found" and will be masked with ``na_sentinel``.
- na_sentinel : int, default -1
- Value in ``labels`` to mark "not found".
- Ignored when ``labels`` is None.
- assume_unique : bool, default False
- When True, ``values`` are assumed to be unique, which can speed up
- the calculation. Ignored when ``labels`` is None.
- verify : bool, default True
- Check if labels are out of bound for the values and put out of bound
- labels equal to na_sentinel. If ``verify=False``, it is assumed there
- are no out of bound labels. Ignored when ``labels`` is None.
-
- .. versionadded:: 0.25.0
-
- Returns
- -------
- ordered : ndarray
- Sorted ``values``
- new_labels : ndarray
- Reordered ``labels``; returned when ``labels`` is not None.
-
- Raises
- ------
- TypeError
- * If ``values`` is not list-like or if ``labels`` is neither None
- nor list-like
- * If ``values`` cannot be sorted
- ValueError
- * If ``labels`` is not None and ``values`` contain duplicates.
- """
- if not is_list_like(values):
- raise TypeError(
- "Only list-like objects are allowed to be passed to safe_sort as values"
- )
-
- if not isinstance(values, np.ndarray) and not is_extension_array_dtype(values):
- # don't convert to string types
- dtype, _ = infer_dtype_from_array(values)
- values = np.asarray(values, dtype=dtype)
-
- def sort_mixed(values):
- # order ints before strings, safe in py3
- str_pos = np.array([isinstance(x, str) for x in values], dtype=bool)
- nums = np.sort(values[~str_pos])
- strs = np.sort(values[str_pos])
- return np.concatenate([nums, np.asarray(strs, dtype=object)])
-
- sorter = None
- if (
- not is_extension_array_dtype(values)
- and lib.infer_dtype(values, skipna=False) == "mixed-integer"
- ):
- # unorderable in py3 if mixed str/int
- ordered = sort_mixed(values)
- else:
- try:
- sorter = values.argsort()
- ordered = values.take(sorter)
- except TypeError:
- # try this anyway
- ordered = sort_mixed(values)
-
- # labels:
-
- if labels is None:
- return ordered
-
- if not is_list_like(labels):
- raise TypeError(
- "Only list-like objects or None are allowed to be"
- "passed to safe_sort as labels"
- )
- labels = ensure_platform_int(np.asarray(labels))
-
- from pandas import Index
-
- if not assume_unique and not Index(values).is_unique:
- raise ValueError("values should be unique if labels is not None")
-
- if sorter is None:
- # mixed types
- hash_klass, values = algorithms._get_data_algo(values)
- t = hash_klass(len(values))
- t.map_locations(values)
- sorter = ensure_platform_int(t.lookup(ordered))
-
- if na_sentinel == -1:
- # take_1d is faster, but only works for na_sentinels of -1
- order2 = sorter.argsort()
- new_labels = algorithms.take_1d(order2, labels, fill_value=-1)
- if verify:
- mask = (labels < -len(values)) | (labels >= len(values))
- else:
- mask = None
- else:
- reverse_indexer = np.empty(len(sorter), dtype=np.int_)
- reverse_indexer.put(sorter, np.arange(len(sorter)))
- # Out of bound indices will be masked with `na_sentinel` next, so we
- # may deal with them here without performance loss using `mode='wrap'`
- new_labels = reverse_indexer.take(labels, mode="wrap")
-
- mask = labels == na_sentinel
- if verify:
- mask = mask | (labels < -len(values)) | (labels >= len(values))
-
- if mask is not None:
- np.putmask(new_labels, mask, na_sentinel)
-
- return ordered, ensure_platform_int(new_labels)
diff --git a/pandas/tests/indexes/test_base.py b/pandas/tests/indexes/test_base.py
index 8d0cb0edf51df..e43d340a46d9f 100644
--- a/pandas/tests/indexes/test_base.py
+++ b/pandas/tests/indexes/test_base.py
@@ -33,13 +33,13 @@
isna,
period_range,
)
+from pandas.core.algorithms import safe_sort
from pandas.core.index import (
_get_combined_index,
ensure_index,
ensure_index_from_sequences,
)
from pandas.core.indexes.api import Index, MultiIndex
-from pandas.core.sorting import safe_sort
from pandas.tests.indexes.common import Base
from pandas.tests.indexes.conftest import indices_dict
import pandas.util.testing as tm
diff --git a/pandas/tests/test_algos.py b/pandas/tests/test_algos.py
index 9dd88fd5dd25b..a64501040442d 100644
--- a/pandas/tests/test_algos.py
+++ b/pandas/tests/test_algos.py
@@ -26,7 +26,6 @@
import pandas.core.algorithms as algos
from pandas.core.arrays import DatetimeArray
import pandas.core.common as com
-from pandas.core.sorting import safe_sort
import pandas.util.testing as tm
@@ -309,7 +308,7 @@ def test_factorize_na_sentinel(self, sort, na_sentinel, data, uniques):
labels, uniques = algos.factorize(data, sort=sort, na_sentinel=na_sentinel)
if sort:
expected_labels = np.array([1, 0, na_sentinel, 1], dtype=np.intp)
- expected_uniques = safe_sort(uniques)
+ expected_uniques = algos.safe_sort(uniques)
else:
expected_labels = np.array([0, 1, na_sentinel, 0], dtype=np.intp)
expected_uniques = uniques
diff --git a/pandas/tests/test_sorting.py b/pandas/tests/test_sorting.py
index b86aaa0ed7e1f..5d7eb70817a11 100644
--- a/pandas/tests/test_sorting.py
+++ b/pandas/tests/test_sorting.py
@@ -6,6 +6,7 @@
import pytest
from pandas import DataFrame, MultiIndex, Series, array, concat, merge
+from pandas.core.algorithms import safe_sort
import pandas.core.common as com
from pandas.core.sorting import (
decons_group_index,
@@ -13,7 +14,6 @@
is_int64_overflow_possible,
lexsort_indexer,
nargsort,
- safe_sort,
)
import pandas.util.testing as tm
diff --git a/pandas/tests/window/test_pairwise.py b/pandas/tests/window/test_pairwise.py
index 56d89e15c418c..6f6d4c09526ff 100644
--- a/pandas/tests/window/test_pairwise.py
+++ b/pandas/tests/window/test_pairwise.py
@@ -3,7 +3,7 @@
import pytest
from pandas import DataFrame, Series
-from pandas.core.sorting import safe_sort
+from pandas.core.algorithms import safe_sort
import pandas.util.testing as tm
| safe_sort uses private functions from core.algorithms and is runtime-imported into core.algorithms. It also doesn't use _anything_ else defined in core.sorting. This move cleans up the dependency structure, in particular is a step towards getting #29133 working. | https://api.github.com/repos/pandas-dev/pandas/pulls/29384 | 2019-11-03T17:22:08Z | 2019-11-04T16:21:26Z | 2019-11-04T16:21:25Z | 2019-11-04T16:46:38Z |
Series repr html only | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index cb68bd0e762c4..4ef773fa3fe24 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -113,6 +113,7 @@ Other enhancements
- Implemented :meth:`pandas.core.window.Window.var` and :meth:`pandas.core.window.Window.std` functions (:issue:`26597`)
- Added ``encoding`` argument to :meth:`DataFrame.to_string` for non-ascii text (:issue:`28766`)
- Added ``encoding`` argument to :func:`DataFrame.to_html` for non-ascii text (:issue:`28663`)
+- Added :meth:`Series._repr_html_` to :class:`Series` to provide basic HTML rendering suitable for notebooks (:issue:`5563`).
- :meth:`Styler.background_gradient` now accepts ``vmin`` and ``vmax`` arguments (:issue:`12145`)
Build Changes
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 3f69dd53491c1..7d00ff054eb4e 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -1576,6 +1576,150 @@ def __repr__(self) -> str:
return result
+ def _repr_html_(self):
+ """
+ Return a html representation for a particular DataFrame.
+
+ Mainly for IPython notebook.
+ """
+ # TODO: Full independent HTML generation in SeriesFormatter, rather
+ # than depending on a limited subset of functionality via to_frame().
+ if get_option("display.notebook_repr_html"):
+ max_rows = get_option("display.max_rows")
+ min_rows = get_option("display.min_rows")
+ show_dimensions = get_option("display.show_dimensions")
+
+ formatter = fmt.DataFrameFormatter(
+ self.to_frame(),
+ columns=None,
+ col_space=None,
+ na_rep="NaN",
+ formatters=None,
+ float_format=None,
+ sparsify=None,
+ justify=None,
+ index_names=True,
+ header=False,
+ index=True,
+ bold_rows=True,
+ escape=True,
+ max_rows=max_rows,
+ min_rows=min_rows,
+ show_dimensions=False, # We do this later for a series.
+ decimal=".",
+ table_id=None,
+ render_links=False,
+ )
+ html = formatter.to_html(notebook=True).split("\n")
+
+ # Find out where the column ends - we will insert footer information here.
+ tbl_end = [
+ rownum for (rownum, row) in enumerate(html) if "</table>" in row
+ ][-1]
+
+ footer = []
+ if self.name is not None:
+ footer.append("Name: <b>{name}</b>".format(name=self.name))
+ if show_dimensions:
+ footer.append("Length: {rows}".format(rows=len(self)))
+ footer.append("dtype: <tt>{dtype}</tt>".format(dtype=self.dtype))
+
+ html.insert(tbl_end + 1, "<p>{footer}</p>".format(footer=", ".join(footer)))
+
+ return "\n".join(html)
+ else:
+ return None
+
+ @Substitution(
+ header_type="bool",
+ header="Whether to print column labels, default True",
+ col_space_type="str or int",
+ col_space="The minimum width of each column in CSS length "
+ "units. An int is assumed to be px units.\n\n"
+ " .. versionadded:: 0.25.0\n"
+ " Ability to use str",
+ )
+ @Substitution(shared_params=fmt.common_docstring, returns=fmt.return_docstring)
+ def to_html(
+ self,
+ buf=None,
+ col_space=None,
+ header=True,
+ index=True,
+ na_rep="NaN",
+ formatters=None,
+ float_format=None,
+ sparsify=None,
+ index_names=True,
+ justify=None,
+ max_rows=None,
+ show_dimensions=False,
+ decimal=".",
+ bold_rows=True,
+ classes=None,
+ escape=True,
+ notebook=False,
+ border=None,
+ series_id=None,
+ render_links=False,
+ encoding=None,
+ ):
+ """
+ Render a Series as a single-column HTML table.
+ %(shared_params)s
+ bold_rows : bool, default True
+ Make the row labels bold in the output.
+ classes : str or list or tuple, default None
+ CSS class(es) to apply to the resulting html table.
+ escape : bool, default True
+ Convert the characters <, >, and & to HTML-safe sequences.
+ notebook : {True, False}, default False
+ Whether the generated HTML is for IPython Notebook.
+ border : int
+ A ``border=border`` attribute is included in the opening
+ `<table>` tag. Default ``pd.options.display.html.border``.
+ encoding : str, default "utf-8"
+ Set character encoding
+ series_id : str, optional
+ A css id is included in the opening `<table>` tag if specified.
+ render_links : bool, default False
+ Convert URLs to HTML links.
+ %(returns)s
+ See Also
+ --------
+ to_string : Convert Series to a string.
+ """
+
+ if justify is not None and justify not in fmt._VALID_JUSTIFY_PARAMETERS:
+ raise ValueError("Invalid value for justify parameter")
+
+ formatter = fmt.SeriesFormatter(
+ self,
+ col_space=col_space,
+ na_rep=na_rep,
+ formatters=formatters,
+ float_format=float_format,
+ sparsify=sparsify,
+ justify=justify,
+ index_names=index_names,
+ header=header,
+ index=index,
+ bold_rows=bold_rows,
+ escape=escape,
+ max_rows=max_rows,
+ show_dimensions=show_dimensions,
+ decimal=decimal,
+ series_id=series_id,
+ render_links=render_links,
+ )
+ return formatter.to_html(
+ buf=buf,
+ classes=classes,
+ notebook=notebook,
+ border=border,
+ encoding=encoding,
+ )
+
def to_string(
self,
buf=None,
diff --git a/pandas/io/formats/format.py b/pandas/io/formats/format.py
index 41bddc7683764..181561080356e 100644
--- a/pandas/io/formats/format.py
+++ b/pandas/io/formats/format.py
@@ -226,7 +226,79 @@ def to_string(self) -> str:
return str("\n".join(result))
-class SeriesFormatter:
+class TableFormatter:
+
+ show_dimensions = None # type: bool
+ is_truncated = None # type: bool
+ formatters = None # type: formatters_type
+ columns = None # type: Index
+
+ @property
+ def should_show_dimensions(self) -> Optional[bool]:
+ return self.show_dimensions is True or (
+ self.show_dimensions == "truncate" and self.is_truncated
+ )
+
+ def _get_formatter(self, i: Union[str, int]) -> Optional[Callable]:
+ if isinstance(self.formatters, (list, tuple)):
+ if is_integer(i):
+ i = cast(int, i)
+ return self.formatters[i]
+ else:
+ return None
+ else:
+ if is_integer(i) and i not in self.columns:
+ i = self.columns[i]
+ return self.formatters.get(i, None)
+
+ @contextmanager
+ def get_buffer(
+ self, buf: Optional[FilePathOrBuffer[str]], encoding: Optional[str] = None
+ ):
+ """
+ Context manager to open, yield and close buffer for filenames or Path-like
+ objects, otherwise yield buf unchanged.
+ """
+ if buf is not None:
+ buf = _stringify_path(buf)
+ else:
+ buf = StringIO()
+
+ if encoding is None:
+ encoding = "utf-8"
+ elif not isinstance(buf, str):
+ raise ValueError("buf is not a file name and encoding is specified.")
+
+ if hasattr(buf, "write"):
+ yield buf
+ elif isinstance(buf, str):
+ with codecs.open(buf, "w", encoding=encoding) as f:
+ yield f
+ else:
+ raise TypeError("buf is not a file name and it has no write method")
+
+ def write_result(self, buf: IO[str]) -> None:
+ """
+ Write the result of serialization to buf.
+ """
+ raise AbstractMethodError(self)
+
+ def get_result(
+ self,
+ buf: Optional[FilePathOrBuffer[str]] = None,
+ encoding: Optional[str] = None,
+ ) -> Optional[str]:
+ """
+ Perform serialization. Write to buf or return as string if buf is None.
+ """
+ with self.get_buffer(buf, encoding=encoding) as f:
+ self.write_result(buf=f)
+ if buf is None:
+ return f.getvalue()
+ return None
+
+
+class SeriesFormatter(TableFormatter):
def __init__(
self,
series: "Series",
@@ -240,10 +312,25 @@ def __init__(
dtype: bool = True,
max_rows: Optional[int] = None,
min_rows: Optional[int] = None,
+ justify: Optional[str] = None,
+ sparsify: Optional[bool] = None,
+ formatters: Optional[formatters_type] = None,
+ show_dimensions: bool = False,
+ col_space: Optional[Union[str, int]] = None,
+ decimal: str = ".",
+ index_names: bool = True,
+ series_id: Optional[str] = None,
+ render_links: bool = False,
+ bold_rows: bool = False,
+ escape: bool = True,
):
self.series = series
self.buf = buf if buf is not None else StringIO()
self.name = name
+ if formatters is None:
+ self.formatters = {}
+ else:
+ self.formatters = formatters
self.na_rep = na_rep
self.header = header
self.length = length
@@ -251,6 +338,25 @@ def __init__(
self.max_rows = max_rows
self.min_rows = min_rows
+ self.show_dimensions = show_dimensions
+ self.col_space = col_space
+ self.decimal = decimal
+ self.show_index_names = index_names
+ self.series_id = series_id
+ self.render_links = render_links
+ self.bold_rows = bold_rows
+ self.escape = escape
+
+ if sparsify is None:
+ sparsify = get_option("display.multi_sparse")
+
+ self.sparsify = sparsify
+
+ if justify is None:
+ justify = get_option("display.colheader_justify")
+
+ self.justify = justify
+
if float_format is None:
float_format = get_option("display.float_format")
self.float_format = float_format
@@ -285,6 +391,7 @@ def _chk_truncate(self) -> None:
self.tr_row_num = None
self.tr_series = series
self.truncate_v = truncate_v
+ self.is_truncated = self.truncate_v
def _get_footer(self) -> str:
name = self.series.name
@@ -324,6 +431,14 @@ def _get_footer(self) -> str:
return str(footer)
+ @property
+ def has_index_names(self) -> bool:
+ return _has_names(self.series.index)
+
+ @property
+ def show_row_idx_names(self) -> bool:
+ return all((self.has_index_names, self.index, self.show_index_names))
+
def _get_formatted_index(self) -> Tuple[List[str], bool]:
index = self.tr_series.index
is_multi = isinstance(index, ABCMultiIndex)
@@ -384,6 +499,46 @@ def to_string(self) -> str:
return str("".join(result))
+ def _format_col(self) -> List[str]:
+ series = self.tr_series
+ formatter = None
+ return format_array(
+ series._values,
+ formatter,
+ float_format=self.float_format,
+ na_rep=self.na_rep,
+ decimal=self.decimal,
+ )
+
+ def to_html(
+ self,
+ buf: Optional[FilePathOrBuffer[str]] = None,
+ encoding: Optional[str] = None,
+ classes: Optional[Union[str, List, Tuple]] = None,
+ notebook: bool = False,
+ border: Optional[int] = None,
+ ) -> Optional[str]:
+ """
+ Render a Series to a html table.
+
+ Parameters
+ ----------
+ classes : str or list-like
+ classes to include in the `class` attribute of the opening
+ ``<table>`` tag, in addition to the default "dataframe".
+ notebook : {True, False}, optional, default False
+ Whether the generated HTML is for IPython Notebook.
+ border : int
+ A ``border=border`` attribute is included in the opening
+ ``<table>`` tag. Default ``pd.options.display.html.border``.
+ """
+ from pandas.io.formats.html import HTMLColumnFormatter, NotebookColumnFormatter
+
+ Klass = NotebookColumnFormatter if notebook else HTMLColumnFormatter
+ return Klass(self, classes=classes, border=border).get_result(
+ buf=buf, encoding=encoding
+ )
+
class TextAdjustment:
def __init__(self):
@@ -446,78 +601,6 @@ def _get_adjustment() -> TextAdjustment:
return TextAdjustment()
-class TableFormatter:
-
- show_dimensions = None # type: bool
- is_truncated = None # type: bool
- formatters = None # type: formatters_type
- columns = None # type: Index
-
- @property
- def should_show_dimensions(self) -> Optional[bool]:
- return self.show_dimensions is True or (
- self.show_dimensions == "truncate" and self.is_truncated
- )
-
- def _get_formatter(self, i: Union[str, int]) -> Optional[Callable]:
- if isinstance(self.formatters, (list, tuple)):
- if is_integer(i):
- i = cast(int, i)
- return self.formatters[i]
- else:
- return None
- else:
- if is_integer(i) and i not in self.columns:
- i = self.columns[i]
- return self.formatters.get(i, None)
-
- @contextmanager
- def get_buffer(
- self, buf: Optional[FilePathOrBuffer[str]], encoding: Optional[str] = None
- ):
- """
- Context manager to open, yield and close buffer for filenames or Path-like
- objects, otherwise yield buf unchanged.
- """
- if buf is not None:
- buf = _stringify_path(buf)
- else:
- buf = StringIO()
-
- if encoding is None:
- encoding = "utf-8"
- elif not isinstance(buf, str):
- raise ValueError("buf is not a file name and encoding is specified.")
-
- if hasattr(buf, "write"):
- yield buf
- elif isinstance(buf, str):
- with codecs.open(buf, "w", encoding=encoding) as f:
- yield f
- else:
- raise TypeError("buf is not a file name and it has no write method")
-
- def write_result(self, buf: IO[str]) -> None:
- """
- Write the result of serialization to buf.
- """
- raise AbstractMethodError(self)
-
- def get_result(
- self,
- buf: Optional[FilePathOrBuffer[str]] = None,
- encoding: Optional[str] = None,
- ) -> Optional[str]:
- """
- Perform serialization. Write to buf or return as string if buf is None.
- """
- with self.get_buffer(buf, encoding=encoding) as f:
- self.write_result(buf=f)
- if buf is None:
- return f.getvalue()
- return None
-
-
class DataFrameFormatter(TableFormatter):
"""
Render a DataFrame
@@ -968,9 +1051,9 @@ def to_html(
A ``border=border`` attribute is included in the opening
``<table>`` tag. Default ``pd.options.display.html.border``.
"""
- from pandas.io.formats.html import HTMLFormatter, NotebookFormatter
+ from pandas.io.formats.html import HTMLTableFormatter, NotebookTableFormatter
- Klass = NotebookFormatter if notebook else HTMLFormatter
+ Klass = NotebookTableFormatter if notebook else HTMLTableFormatter
return Klass(self, classes=classes, border=border).get_result(
buf=buf, encoding=encoding
)
diff --git a/pandas/io/formats/html.py b/pandas/io/formats/html.py
index 38f2e332017f0..9fd2bee674048 100644
--- a/pandas/io/formats/html.py
+++ b/pandas/io/formats/html.py
@@ -15,6 +15,7 @@
from pandas.io.common import _is_url
from pandas.io.formats.format import (
DataFrameFormatter,
+ SeriesFormatter,
TableFormatter,
buffer_put_lines,
get_level_lengths,
@@ -22,7 +23,7 @@
from pandas.io.formats.printing import pprint_thing
-class HTMLFormatter(TableFormatter):
+class HTMLTableFormatter(TableFormatter):
"""
Internal class for formatting output data in html.
This class is intended for shared functionality between
@@ -566,7 +567,7 @@ def _write_hierarchical_rows(
)
-class NotebookFormatter(HTMLFormatter):
+class NotebookTableFormatter(HTMLTableFormatter):
"""
Internal class for formatting output data in html for display in Jupyter
Notebooks. This class is intended for functionality specific to
@@ -613,3 +614,312 @@ def render(self) -> List[str]:
super().render()
self.write("</div>")
return self.elements
+
+
+class HTMLColumnFormatter(HTMLTableFormatter):
+ """
+ Internal class for formatting output data in html.
+ This class is intended for shared functionality between
+ Series.to_html() and Series._repr_html_().
+ Any logic in common with other output formatting methods
+ should ideally be inherited from classes in format.py
+ and this class responsible for only producing html markup.
+ """
+
+ indent_delta = 2
+
+ def __init__(
+ self,
+ formatter: SeriesFormatter,
+ classes: Optional[Union[str, List[str], Tuple[str, ...]]] = None,
+ border: Optional[int] = None,
+ ) -> None:
+ self.fmt = formatter
+ self.classes = classes
+
+ self.series = self.fmt.series
+ self.series_id = self.fmt.series_id
+ self.elements = [] # type: List[str]
+ self.bold_rows = self.fmt.bold_rows
+ self.escape = self.fmt.escape
+ self.show_dimensions = self.fmt.show_dimensions
+ if border is None:
+ border = cast(int, get_option("display.html.border"))
+ self.border = border
+ self.render_links = self.fmt.render_links
+
+ def render(self) -> List[str]:
+ self._write_column()
+
+ info = []
+ if self.series.name is not None:
+ info.append('Name: {name}'.format(name=self.series.name))
+ if self.should_show_dimensions:
+ info.append('Length: {rows}'.format(rows=len(self.series)))
+ info.append('dtype: <tt>{dtype}</tt>'.format(dtype=self.series.dtype))
+
+ self.write("<p>{info}</p>".format(info=', '.join(info)))
+
+ return self.elements
+
+ def _get_formatted_values(self) -> Dict[int, List[str]]:
+ return {0: self.fmt._format_col()}
+
+ def _write_column(self, indent: int = 0) -> None:
+ _classes = ["series"] # Default class.
+ use_mathjax = get_option("display.html.use_mathjax")
+ if not use_mathjax:
+ _classes.append("tex2jax_ignore")
+ if self.classes is not None:
+ if isinstance(self.classes, str):
+ self.classes = self.classes.split()
+ if not isinstance(self.classes, (list, tuple)):
+ raise TypeError(
+ "classes must be a string, list, or tuple, "
+ "not {typ}".format(typ=type(self.classes))
+ )
+ _classes.extend(self.classes)
+
+ if self.series_id is None:
+ id_section = ""
+ else:
+ id_section = ' id="{series_id}"'.format(series_id=self.series_id)
+
+ self.write(
+ '<table border="{border}" class="{cls}"{id_section}>'.format(
+ border=self.border, cls=" ".join(_classes), id_section=id_section
+ ),
+ indent,
+ )
+
+ if self.show_row_idx_names:
+ self._write_header(indent + self.indent_delta)
+
+ self._write_body(indent + self.indent_delta)
+
+ self.write("</table>", indent)
+
+ def _write_body(self, indent: int) -> None:
+ self.write("<tbody>", indent)
+ fmt_values = self._get_formatted_values()
+
+ # write values
+ if self.fmt.index and isinstance(self.series.index, ABCMultiIndex):
+ self._write_hierarchical_rows(fmt_values, indent + self.indent_delta)
+ else:
+ self._write_regular_rows(fmt_values, indent + self.indent_delta)
+
+ self.write("</tbody>", indent)
+
+ def _write_row_header(self, indent: int) -> None:
+ row = [x if x is not None else "" for x in self.series.index.names] + [""]
+ self.write_tr(row, indent, self.indent_delta, header=True)
+
+ def _write_header(self, indent: int) -> None:
+ self.write("<thead>", indent)
+
+ if self.show_row_idx_names:
+ self._write_row_header(indent + self.indent_delta)
+
+ self.write("</thead>", indent)
+
+ def _write_regular_rows(
+ self, fmt_values: Mapping[int, List[str]], indent: int
+ ) -> None:
+ truncate_v = self.fmt.truncate_v
+
+ nrows = len(self.fmt.tr_series)
+
+ if self.fmt.index:
+ fmt = self.fmt._get_formatter("__index__")
+ if fmt is not None:
+ index_values = self.fmt.tr_series.index.map(fmt)
+ else:
+ index_values = self.fmt.tr_series.index.format()
+
+ row = [] # type: List[str]
+ for i in range(nrows):
+
+ if truncate_v and i == (self.fmt.tr_row_num):
+ str_sep_row = ["..."] * len(row)
+ self.write_tr(
+ str_sep_row,
+ indent,
+ self.indent_delta,
+ tags=None,
+ nindex_levels=self.row_levels,
+ )
+
+ row = []
+ if self.fmt.index:
+ row.append(index_values[i])
+ row.extend(fmt_values[j][i] for j in range(self.ncols))
+
+ self.write_tr(
+ row, indent, self.indent_delta, tags=None, nindex_levels=self.row_levels
+ )
+
+ def _write_hierarchical_rows(
+ self, fmt_values: Mapping[int, List[str]], indent: int
+ ) -> None:
+ template = 'rowspan="{span}" valign="top"'
+
+ truncate_v = self.fmt.truncate_v
+ series = self.fmt.tr_series
+ nrows = len(series)
+
+ idx_values = series.index.format(sparsify=False, adjoin=False, names=False)
+ idx_values = list(zip(*idx_values))
+
+ if self.fmt.sparsify:
+ sentinel = object()
+ levels = series.index.format(sparsify=sentinel, adjoin=False, names=False)
+
+ level_lengths = get_level_lengths(levels, sentinel)
+ inner_lvl = len(level_lengths) - 1
+ if truncate_v:
+ # Insert ... row and adjust idx_values and
+ # level_lengths to take this into account.
+ ins_row = self.fmt.tr_row_num
+ # cast here since if truncate_v is True, self.fmt.tr_row_num is not None
+ ins_row = cast(int, ins_row)
+ inserted = False
+ for lnum, records in enumerate(level_lengths):
+ rec_new = {}
+ for tag, span in list(records.items()):
+ if tag >= ins_row:
+ rec_new[tag + 1] = span
+ elif tag + span > ins_row:
+ rec_new[tag] = span + 1
+
+ # GH 14882 - Make sure insertion done once
+ if not inserted:
+ dot_row = list(idx_values[ins_row - 1])
+ dot_row[-1] = "..."
+ idx_values.insert(ins_row, tuple(dot_row))
+ inserted = True
+ else:
+ dot_row = list(idx_values[ins_row])
+ dot_row[inner_lvl - lnum] = "..."
+ idx_values[ins_row] = tuple(dot_row)
+ else:
+ rec_new[tag] = span
+ # If ins_row lies between tags, all cols idx cols
+ # receive ...
+ if tag + span == ins_row:
+ rec_new[ins_row] = 1
+ if lnum == 0:
+ idx_values.insert(
+ ins_row, tuple(["..."] * len(level_lengths))
+ )
+
+ # GH 14882 - Place ... in correct level
+ elif inserted:
+ dot_row = list(idx_values[ins_row])
+ dot_row[inner_lvl - lnum] = "..."
+ idx_values[ins_row] = tuple(dot_row)
+ level_lengths[lnum] = rec_new
+
+ level_lengths[inner_lvl][ins_row] = 1
+ for ix_col in range(len(fmt_values)):
+ fmt_values[ix_col].insert(ins_row, "...")
+ nrows += 1
+
+ for i in range(nrows):
+ row = []
+ tags = {}
+
+ sparse_offset = 0
+ j = 0
+ for records, v in zip(level_lengths, idx_values[i]):
+ if i in records:
+ if records[i] > 1:
+ tags[j] = template.format(span=records[i])
+ else:
+ sparse_offset += 1
+ continue
+
+ j += 1
+ row.append(v)
+
+ row.append(fmt_values[0][i])
+ self.write_tr(
+ row,
+ indent,
+ self.indent_delta,
+ tags=tags,
+ nindex_levels=len(levels) - sparse_offset,
+ )
+ else:
+ row = []
+ for i in range(len(series)):
+ if truncate_v and i == (self.fmt.tr_row_num):
+ str_sep_row = ["..."] * len(row)
+ self.write_tr(
+ str_sep_row,
+ indent,
+ self.indent_delta,
+ tags=None,
+ nindex_levels=self.row_levels,
+ )
+
+ idx_values = list(
+ zip(*series.index.format(sparsify=False, adjoin=False, names=False))
+ )
+ row = []
+ row.extend(idx_values[i])
+ row.append(fmt_values[0][i])
+ self.write_tr(
+ row,
+ indent,
+ self.indent_delta,
+ tags=None,
+ nindex_levels=series.index.nlevels,
+ )
+
+ @property
+ def row_levels(self) -> int:
+ if self.fmt.index:
+ # showing (row) index
+ return self.series.index.nlevels
+ else:
+ return 0
+
+ @property
+ def ncols(self) -> int:
+ return 1
+
+
+class NotebookColumnFormatter(HTMLColumnFormatter):
+ """
+ Internal class for formatting output data in html for display in Jupyter
+ Notebooks. This class is intended for functionality specific to
+ DataFrame._repr_html_() and DataFrame.to_html(notebook=True)
+ """
+
+ def write_style(self) -> None:
+ # We use the "scoped" attribute here so that the desired
+ # style properties for the data frame are not then applied
+ # throughout the entire notebook.
+ template_first = """\
+ <style scoped>"""
+ template_last = """\
+ </style>"""
+ template_select = """\
+ .dataframe %s {
+ %s: %s;
+ }"""
+ element_props = [
+ ("tbody tr th:only-of-type", "vertical-align", "middle"),
+ ("tbody tr th", "vertical-align", "top"),
+ ]
+ template_mid = "\n\n".join(map(lambda t: template_select % t, element_props))
+ template = dedent("\n".join((template_first, template_mid, template_last)))
+ self.write(template)
+
+ def render(self) -> List[str]:
+ self.write("<div>")
+ self.write_style()
+ super().render()
+ self.write("</div>")
+ return self.elements
diff --git a/pandas/io/formats/latex.py b/pandas/io/formats/latex.py
index 6f903e770c86c..87c342b4b8659 100644
--- a/pandas/io/formats/latex.py
+++ b/pandas/io/formats/latex.py
@@ -25,7 +25,7 @@ class LatexFormatter(TableFormatter):
See Also
--------
- HTMLFormatter
+ HTMLTableFormatter
"""
def __init__(
diff --git a/pandas/tests/io/formats/test_format.py b/pandas/tests/io/formats/test_format.py
index 0f4a7a33dd115..510ec32667e20 100644
--- a/pandas/tests/io/formats/test_format.py
+++ b/pandas/tests/io/formats/test_format.py
@@ -1865,6 +1865,54 @@ def test_repr_html(self, float_frame):
tm.reset_display_options()
+ def test_repr_html_series(self):
+ data = [1, "two", 3.1, -4.2, True, np.nan]
+
+ fmt.set_option("display.max_rows", len(data))
+
+ small = pd.Series(data, name="test series")
+ sm_html = small._repr_html_()
+
+ true_sm_html = (
+ "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n"
+ " vertical-align: middle;\n }\n\n .dataframe tbody tr th"
+ " {\n vertical-align: top;\n }\n\n .dataframe thead th {\n"
+ ' text-align: right;\n }\n</style>\n<table border="1" '
+ 'class="dataframe">\n <tbody>\n <tr>\n <th>0</th>\n '
+ "<td>1</td>\n </tr>\n <tr>\n <th>1</th>\n <td>two</td>\n"
+ " </tr>\n <tr>\n <th>2</th>\n <td>3.1</td>\n </tr>\n"
+ " <tr>\n <th>3</th>\n <td>-4.2</td>\n </tr>\n <tr>\n"
+ " <th>4</th>\n <td>True</td>\n </tr>\n <tr>\n "
+ "<th>5</th>\n <td>NaN</td>\n </tr>\n </tbody>\n</table>\n"
+ "<p>Name: <b>test series</b>, Length: 6, dtype: <tt>object</tt></p>\n"
+ "</div>"
+ )
+
+ assert sm_html == true_sm_html
+
+ large = small.repeat(1000)
+ large.name = None
+ lg_html = large._repr_html_()
+
+ true_lg_html = (
+ "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n"
+ " vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n"
+ " vertical-align: top;\n }\n\n .dataframe thead th {\n"
+ ' text-align: right;\n }\n</style>\n<table border="1" '
+ 'class="dataframe">\n <tbody>\n <tr>\n <th>0</th>\n '
+ "<td>1</td>\n </tr>\n <tr>\n <th>0</th>\n <td>1</td>\n "
+ "</tr>\n <tr>\n <th>0</th>\n <td>1</td>\n </tr>\n <tr>"
+ "\n <th>...</th>\n <td>...</td>\n </tr>\n <tr>\n "
+ "<th>5</th>\n <td>NaN</td>\n </tr>\n <tr>\n <th>5</th>\n"
+ " <td>NaN</td>\n </tr>\n <tr>\n <th>5</th>\n "
+ "<td>NaN</td>\n </tr>\n </tbody>\n</table>\n<p>Length: 6000, dtype: "
+ "<tt>object</tt></p>\n</div>"
+ )
+
+ assert lg_html == true_lg_html
+
+ tm.reset_display_options()
+
def test_repr_html_mathjax(self):
df = DataFrame([[1, 2], [3, 4]])
assert "tex2jax_ignore" not in df._repr_html_()
diff --git a/pandas/tests/io/formats/test_printing.py b/pandas/tests/io/formats/test_printing.py
index f0d5ef19c4468..2d26dca281a5f 100644
--- a/pandas/tests/io/formats/test_printing.py
+++ b/pandas/tests/io/formats/test_printing.py
@@ -132,16 +132,13 @@ def test_publishes(self):
df = pd.DataFrame({"A": [1, 2]})
objects = [df["A"], df, df] # dataframe / series
- expected_keys = [
- {"text/plain", "application/vnd.dataresource+json"},
- {"text/plain", "text/html", "application/vnd.dataresource+json"},
- ]
+ expected_keys = {"text/plain", "text/html", "application/vnd.dataresource+json"}
opt = pd.option_context("display.html.table_schema", True)
- for obj, expected in zip(objects, expected_keys):
+ for obj in objects:
with opt:
formatted = self.display_formatter.format(obj)
- assert set(formatted[0].keys()) == expected
+ assert set(formatted[0].keys()) == expected_keys
with_latex = pd.option_context("display.latex.repr", True)
| - [x] closes #5563
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29383 | 2019-11-03T15:31:41Z | 2020-09-10T18:56:28Z | null | 2020-09-11T17:22:03Z |
CLN: Remove unused _isfinite, make_axis_dummies | diff --git a/pandas/core/nanops.py b/pandas/core/nanops.py
index 070891c4acb5e..7e50348962fc5 100644
--- a/pandas/core/nanops.py
+++ b/pandas/core/nanops.py
@@ -16,7 +16,6 @@
is_any_int_dtype,
is_bool_dtype,
is_complex,
- is_complex_dtype,
is_datetime64_dtype,
is_datetime64tz_dtype,
is_datetime_or_timedelta_dtype,
@@ -325,19 +324,6 @@ def _get_values(
return values, mask, dtype, dtype_max, fill_value
-def _isfinite(values):
- if is_datetime_or_timedelta_dtype(values):
- return isna(values)
- if (
- is_complex_dtype(values)
- or is_float_dtype(values)
- or is_integer_dtype(values)
- or is_bool_dtype(values)
- ):
- return ~np.isfinite(values)
- return ~np.isfinite(values.astype("float64"))
-
-
def _na_ok_dtype(dtype):
# TODO: what about datetime64tz? PeriodDtype?
return not issubclass(dtype.type, (np.integer, np.timedelta64, np.datetime64))
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index ad7081fb17703..949d8f1bfb09c 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -1046,43 +1046,7 @@ def _make_col_name(prefix, prefix_sep, level):
return DataFrame(dummy_mat, index=index, columns=dummy_cols)
-def make_axis_dummies(frame, axis="minor", transform=None):
- """
- Construct 1-0 dummy variables corresponding to designated axis
- labels
-
- Parameters
- ----------
- frame : DataFrame
- axis : {'major', 'minor'}, default 'minor'
- transform : function, default None
- Function to apply to axis labels first. For example, to
- get "day of week" dummies in a time series regression
- you might call::
-
- make_axis_dummies(panel, axis='major',
- transform=lambda d: d.weekday())
- Returns
- -------
- dummies : DataFrame
- Column names taken from chosen axis
- """
- numbers = {"major": 0, "minor": 1}
- num = numbers.get(axis, axis)
-
- items = frame.index.levels[num]
- codes = frame.index.codes[num]
- if transform is not None:
- mapped_items = items.map(transform)
- codes, items = _factorize_from_iterable(mapped_items.take(codes))
-
- values = np.eye(len(items), dtype=float)
- values = values.take(codes, axis=0)
-
- return DataFrame(values, columns=items, index=frame.index)
-
-
-def _reorder_for_extension_array_stack(arr, n_rows, n_columns):
+def _reorder_for_extension_array_stack(arr, n_rows: int, n_columns: int):
"""
Re-orders the values when stacking multiple extension-arrays.
diff --git a/pandas/tests/reshape/test_reshape.py b/pandas/tests/reshape/test_reshape.py
index 2e94eeba1d05b..b695b05c7c7db 100644
--- a/pandas/tests/reshape/test_reshape.py
+++ b/pandas/tests/reshape/test_reshape.py
@@ -645,24 +645,3 @@ def test_reshaping_multi_index_categorical(self):
index=dti.rename("major"),
)
tm.assert_frame_equal(result, expected)
-
-
-class TestMakeAxisDummies:
- def test_preserve_categorical_dtype(self):
- # GH13854
- for ordered in [False, True]:
- cidx = pd.CategoricalIndex(list("xyz"), ordered=ordered)
- midx = pd.MultiIndex(levels=[["a"], cidx], codes=[[0, 0], [0, 1]])
- df = DataFrame([[10, 11]], index=midx)
-
- expected = DataFrame(
- [[1.0, 0.0, 0.0], [0.0, 1.0, 0.0]], index=midx, columns=cidx
- )
-
- from pandas.core.reshape.reshape import make_axis_dummies
-
- result = make_axis_dummies(df)
- tm.assert_frame_equal(result, expected)
-
- result = make_axis_dummies(df, transform=lambda x: x)
- tm.assert_frame_equal(result, expected)
diff --git a/pandas/tests/test_nanops.py b/pandas/tests/test_nanops.py
index 49d1777df0751..52ad56967220f 100644
--- a/pandas/tests/test_nanops.py
+++ b/pandas/tests/test_nanops.py
@@ -704,46 +704,6 @@ def test__has_infs(self):
self.check_bool(nanops._has_infs, val.astype("f4"), correct)
self.check_bool(nanops._has_infs, val.astype("f2"), correct)
- def test__isfinite(self):
- pairs = [
- ("arr_complex", False),
- ("arr_int", False),
- ("arr_bool", False),
- ("arr_str", False),
- ("arr_utf", False),
- ("arr_complex", False),
- ("arr_complex_nan", True),
- ("arr_nan_nanj", True),
- ("arr_nan_infj", True),
- ("arr_complex_nan_infj", True),
- ]
- pairs_float = [
- ("arr_float", False),
- ("arr_nan", True),
- ("arr_float_nan", True),
- ("arr_nan_nan", True),
- ("arr_float_inf", True),
- ("arr_inf", True),
- ("arr_nan_inf", True),
- ("arr_float_nan_inf", True),
- ("arr_nan_nan_inf", True),
- ]
-
- func1 = lambda x: np.any(nanops._isfinite(x).ravel())
-
- # TODO: unused?
- # func2 = lambda x: np.any(nanops._isfinite(x).values.ravel())
-
- for arr, correct in pairs:
- val = getattr(self, arr)
- self.check_bool(func1, val, correct)
-
- for arr, correct in pairs_float:
- val = getattr(self, arr)
- self.check_bool(func1, val, correct)
- self.check_bool(func1, val.astype("f4"), correct)
- self.check_bool(func1, val.astype("f2"), correct)
-
def test__bn_ok_dtype(self):
assert nanops._bn_ok_dtype(self.arr_float.dtype, "test")
assert nanops._bn_ok_dtype(self.arr_complex.dtype, "test")
| https://api.github.com/repos/pandas-dev/pandas/pulls/29380 | 2019-11-03T04:35:44Z | 2019-11-03T14:54:47Z | 2019-11-03T14:54:47Z | 2019-11-03T15:34:09Z | |
Fixes typo | diff --git a/web/pandas/getting_started.md b/web/pandas/getting_started.md
index 9682cf90cad6f..4195cc00b2419 100644
--- a/web/pandas/getting_started.md
+++ b/web/pandas/getting_started.md
@@ -9,7 +9,7 @@ the [advanced installation page]({{ base_url}}/docs/getting_started/install.html
1. Download [Anaconda](https://www.anaconda.com/distribution/) for your operating system and
the latest Python version, run the installer, and follow the steps. Detailed instructions
on how to install Anaconda can be found in the
- [Anaconda documentation](https://docs.anaconda.com/anaconda/install/)).
+ [Anaconda documentation](https://docs.anaconda.com/anaconda/install/).
2. In the Anaconda prompt (or terminal in Linux or MacOS), start JupyterLab:
| https://api.github.com/repos/pandas-dev/pandas/pulls/29379 | 2019-11-03T03:33:15Z | 2019-11-03T14:53:47Z | 2019-11-03T14:53:47Z | 2019-11-03T14:53:51Z | |
TST: new test for incorrect series assignment | diff --git a/pandas/tests/series/indexing/test_indexing.py b/pandas/tests/series/indexing/test_indexing.py
index 4673dabca811b..5aba2920999d5 100644
--- a/pandas/tests/series/indexing/test_indexing.py
+++ b/pandas/tests/series/indexing/test_indexing.py
@@ -391,6 +391,22 @@ def test_setslice(datetime_series):
assert sl.index.is_unique is True
+def test_2d_to_1d_assignment_raises():
+ x = np.random.randn(2, 2)
+ y = pd.Series(range(2))
+
+ msg = (
+ r"shape mismatch: value array of shape \(2,2\) could not be"
+ r" broadcast to indexing result of shape \(2,\)"
+ )
+ with pytest.raises(ValueError, match=msg):
+ y.loc[range(2)] = x
+
+ msg = r"could not broadcast input array from shape \(2,2\) into shape \(2\)"
+ with pytest.raises(ValueError, match=msg):
+ y.loc[:] = x
+
+
# FutureWarning from NumPy about [slice(None, 5).
@pytest.mark.filterwarnings("ignore:Using a non-tuple:FutureWarning")
def test_basic_getitem_setitem_corner(datetime_series):
| - [x] closes #14525
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29378 | 2019-11-03T00:41:52Z | 2019-11-07T22:13:28Z | 2019-11-07T22:13:28Z | 2019-11-07T22:13:43Z |
CLN: deprivatize factorize_from_iterable | diff --git a/pandas/core/arrays/categorical.py b/pandas/core/arrays/categorical.py
index 612e54ba426f3..ce174baa66a97 100644
--- a/pandas/core/arrays/categorical.py
+++ b/pandas/core/arrays/categorical.py
@@ -2678,7 +2678,7 @@ def _convert_to_list_like(list_like):
return [list_like]
-def _factorize_from_iterable(values):
+def factorize_from_iterable(values):
"""
Factorize an input `values` into `categories` and `codes`. Preserves
categorical dtype in `categories`.
@@ -2716,9 +2716,9 @@ def _factorize_from_iterable(values):
return codes, categories
-def _factorize_from_iterables(iterables):
+def factorize_from_iterables(iterables):
"""
- A higher-level wrapper over `_factorize_from_iterable`.
+ A higher-level wrapper over `factorize_from_iterable`.
*This is an internal function*
@@ -2733,9 +2733,9 @@ def _factorize_from_iterables(iterables):
Notes
-----
- See `_factorize_from_iterable` for more info.
+ See `factorize_from_iterable` for more info.
"""
if len(iterables) == 0:
# For consistency, it should return a list of 2 lists.
return [[], []]
- return map(list, zip(*(_factorize_from_iterable(it) for it in iterables)))
+ return map(list, zip(*(factorize_from_iterable(it) for it in iterables)))
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index f968a9eb4103c..66deacac37789 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -31,7 +31,7 @@
import pandas.core.algorithms as algos
from pandas.core.arrays import Categorical
-from pandas.core.arrays.categorical import _factorize_from_iterables
+from pandas.core.arrays.categorical import factorize_from_iterables
import pandas.core.common as com
import pandas.core.indexes.base as ibase
from pandas.core.indexes.base import (
@@ -440,7 +440,7 @@ def from_arrays(cls, arrays, sortorder=None, names=_no_default_names):
if len(arrays[i]) != len(arrays[i - 1]):
raise ValueError("all arrays must be same length")
- codes, levels = _factorize_from_iterables(arrays)
+ codes, levels = factorize_from_iterables(arrays)
if names is _no_default_names:
names = [getattr(arr, "name", None) for arr in arrays]
@@ -562,7 +562,7 @@ def from_product(cls, iterables, sortorder=None, names=_no_default_names):
elif is_iterator(iterables):
iterables = list(iterables)
- codes, levels = _factorize_from_iterables(iterables)
+ codes, levels = factorize_from_iterables(iterables)
if names is _no_default_names:
names = [getattr(it, "name", None) for it in iterables]
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index bbf41fc28e9d2..c11915c00c59d 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -8,8 +8,8 @@
from pandas import DataFrame, Index, MultiIndex, Series
from pandas.core.arrays.categorical import (
- _factorize_from_iterable,
- _factorize_from_iterables,
+ factorize_from_iterable,
+ factorize_from_iterables,
)
import pandas.core.common as com
from pandas.core.generic import NDFrame
@@ -604,7 +604,7 @@ def _make_concat_multiindex(indexes, keys, levels=None, names=None):
names = [None] * len(zipped)
if levels is None:
- _, levels = _factorize_from_iterables(zipped)
+ _, levels = factorize_from_iterables(zipped)
else:
levels = [ensure_index(x) for x in levels]
else:
@@ -645,7 +645,7 @@ def _make_concat_multiindex(indexes, keys, levels=None, names=None):
levels.extend(concat_index.levels)
codes_list.extend(concat_index.codes)
else:
- codes, categories = _factorize_from_iterable(concat_index)
+ codes, categories = factorize_from_iterable(concat_index)
levels.append(categories)
codes_list.append(codes)
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index 949d8f1bfb09c..d7eae1c543804 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -22,7 +22,7 @@
import pandas.core.algorithms as algos
from pandas.core.arrays import SparseArray
-from pandas.core.arrays.categorical import _factorize_from_iterable
+from pandas.core.arrays.categorical import factorize_from_iterable
from pandas.core.construction import extract_array
from pandas.core.frame import DataFrame
from pandas.core.index import Index, MultiIndex
@@ -504,7 +504,7 @@ def stack(frame, level=-1, dropna=True):
def factorize(index):
if index.is_unique:
return index, np.arange(len(index))
- codes, categories = _factorize_from_iterable(index)
+ codes, categories = factorize_from_iterable(index)
return categories, codes
N, K = frame.shape
@@ -725,7 +725,7 @@ def _convert_level_number(level_num, columns):
new_names = list(this.index.names)
new_codes = [lab.repeat(levsize) for lab in this.index.codes]
else:
- old_codes, old_levels = _factorize_from_iterable(this.index)
+ old_codes, old_levels = factorize_from_iterable(this.index)
new_levels = [old_levels]
new_codes = [old_codes.repeat(levsize)]
new_names = [this.index.name] # something better?
@@ -949,7 +949,7 @@ def _get_dummies_1d(
from pandas.core.reshape.concat import concat
# Series avoids inconsistent NaN handling
- codes, levels = _factorize_from_iterable(Series(data))
+ codes, levels = factorize_from_iterable(Series(data))
if dtype is None:
dtype = np.uint8
| Deprivatize ``_factorize_from_iterable`` and ``_factorize_from_iterables`` | https://api.github.com/repos/pandas-dev/pandas/pulls/29377 | 2019-11-02T21:26:57Z | 2019-11-03T22:11:43Z | 2019-11-03T22:11:43Z | 2019-11-03T22:11:47Z |
TST: new test for sort index when Nan in other axis. | diff --git a/pandas/tests/frame/test_sorting.py b/pandas/tests/frame/test_sorting.py
index 9ea78b974fcbb..422d1b0239a4e 100644
--- a/pandas/tests/frame/test_sorting.py
+++ b/pandas/tests/frame/test_sorting.py
@@ -735,3 +735,12 @@ def test_sort_index_na_position_with_categories_raises(self):
with pytest.raises(ValueError):
df.sort_values(by="c", ascending=False, na_position="bad_position")
+
+ def test_sort_index_nan(self):
+
+ df = pd.DataFrame(0, columns=[], index=pd.MultiIndex.from_product([[], []]))
+ df.loc["b", "2"] = 1
+ df.loc["a", "3"] = 1
+ result = df.sort_index()
+
+ assert result.index.is_monotonic == True
| closes #12261
test added / passed
passes black pandas | https://api.github.com/repos/pandas-dev/pandas/pulls/29376 | 2019-11-02T20:45:37Z | 2020-01-01T18:33:42Z | null | 2020-01-01T18:33:43Z |
Deprecate using `xlrd` engine | diff --git a/doc/source/whatsnew/v1.1.0.rst b/doc/source/whatsnew/v1.1.0.rst
index 920919755dc23..e1b99bfcbaaf6 100644
--- a/doc/source/whatsnew/v1.1.0.rst
+++ b/doc/source/whatsnew/v1.1.0.rst
@@ -68,7 +68,8 @@ Backwards incompatible API changes
Deprecations
~~~~~~~~~~~~
-
+- :func:`read_excel` engine argument "xlrd" will no longer be the default engine and
+ will be replaced by "openpyxl" in a future version (:issue:`28547`).
-
-
diff --git a/pandas/io/excel/_base.py b/pandas/io/excel/_base.py
index 2a91381b7fbeb..ee0fb1ae19eb3 100644
--- a/pandas/io/excel/_base.py
+++ b/pandas/io/excel/_base.py
@@ -3,6 +3,7 @@
from io import BytesIO
import os
from textwrap import fill
+import warnings
from pandas._config import config
@@ -781,7 +782,7 @@ def close(self):
class ExcelFile:
"""
Class for parsing tabular excel sheets into DataFrame objects.
- Uses xlrd. See read_excel for more documentation
+ Uses xlrd, openpyxl or odf. See read_excel for more documentation
Parameters
----------
@@ -809,6 +810,11 @@ class ExcelFile:
def __init__(self, io, engine=None):
if engine is None:
engine = "xlrd"
+ warnings.warn(
+ 'The Excel reader engine will default to "openpyxl" in the future. \
+ Specify engine="openpyxl" to suppress this warning.',
+ FutureWarning,
+ )
if engine not in self._engines:
raise ValueError(f"Unknown engine: {engine}")
diff --git a/pandas/tests/io/excel/test_xlrd.py b/pandas/tests/io/excel/test_xlrd.py
index cc7e2311f362a..3e8d300f65c35 100644
--- a/pandas/tests/io/excel/test_xlrd.py
+++ b/pandas/tests/io/excel/test_xlrd.py
@@ -1,3 +1,5 @@
+import warnings
+
import pytest
import pandas as pd
@@ -26,7 +28,6 @@ def test_read_xlrd_book(read_ext, frame):
with tm.ensure_clean(read_ext) as pth:
df.to_excel(pth, sheet_name)
book = xlrd.open_workbook(pth)
-
with ExcelFile(book, engine=engine) as xl:
result = pd.read_excel(xl, sheet_name, index_col=0)
tm.assert_frame_equal(df, result)
@@ -38,6 +39,24 @@ def test_read_xlrd_book(read_ext, frame):
# TODO: test for openpyxl as well
def test_excel_table_sheet_by_index(datapath, read_ext):
path = datapath("io", "data", "excel", "test1{}".format(read_ext))
- with pd.ExcelFile(path) as excel:
- with pytest.raises(xlrd.XLRDError):
- pd.read_excel(excel, "asdf")
+ with warnings.catch_warnings():
+ warnings.simplefilter("ignore")
+ with pd.ExcelFile(path) as excel:
+ with pytest.raises(xlrd.XLRDError):
+ pd.read_excel(excel, "asdf")
+
+
+# See issue #29375
+def test_excel_file_warning_with_default_engine(datapath):
+ path = datapath("io", "data", "excel", "test1.xls")
+ with warnings.catch_warnings(record=True) as w:
+ pd.ExcelFile(path)
+ assert "default to \"openpyxl\" in the future." in str(w[-1].message)
+
+
+# See issue #29375
+def test_read_excel_warning_with_default_engine(tmpdir, datapath):
+ path = datapath("io", "data", "excel", "test1.xls")
+ with warnings.catch_warnings(record=True) as w:
+ pd.read_excel(path, "Sheet1")
+ assert "default to \"openpyxl\" in the future." in str(w[-1].message)
| - [x] closes #28547
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29375 | 2019-11-02T20:20:11Z | 2020-04-22T03:56:00Z | null | 2020-04-22T03:56:01Z |
Updated index links for 0.25.3 | diff --git a/doc/source/index.rst.template b/doc/source/index.rst.template
index 09d18d6f96197..cbdbbda7e530d 100644
--- a/doc/source/index.rst.template
+++ b/doc/source/index.rst.template
@@ -39,7 +39,7 @@ See the :ref:`overview` for more detail about what's in the library.
:hidden:
{% endif %}
{% if not single_doc %}
- What's New in 0.25.2 <whatsnew/v0.25.2>
+ What's New in 0.25.3 <whatsnew/v0.25.3>
install
getting_started/index
user_guide/index
@@ -53,7 +53,7 @@ See the :ref:`overview` for more detail about what's in the library.
whatsnew/index
{% endif %}
-* :doc:`whatsnew/v0.25.2`
+* :doc:`whatsnew/v0.25.3`
* :doc:`install`
* :doc:`getting_started/index`
| I think this was supposed to be updated for the release. Might need to retag after repush docs after this
@TomAugspurger | https://api.github.com/repos/pandas-dev/pandas/pulls/29374 | 2019-11-02T19:44:11Z | 2019-11-08T16:27:25Z | 2019-11-08T16:27:25Z | 2019-11-14T16:20:20Z |
Add documentation linking to sqlalchemy | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index f88c26c7bc782..cfbd125b7445e 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -2645,7 +2645,11 @@ def to_sql(
Name of SQL table.
con : sqlalchemy.engine.Engine or sqlite3.Connection
Using SQLAlchemy makes it possible to use any DB supported by that
- library. Legacy support is provided for sqlite3.Connection objects.
+ library. Legacy support is provided for sqlite3.Connection objects. The user
+ is responsible for engine disposal and connection closure for the SQLAlchemy
+ connectable See `here \
+ <https://docs.sqlalchemy.org/en/13/core/connections.html>`_
+
schema : str, optional
Specify the schema (if database flavor supports this). If None, use
default schema.
diff --git a/pandas/io/sql.py b/pandas/io/sql.py
index e90e19649f645..684e602f06d12 100644
--- a/pandas/io/sql.py
+++ b/pandas/io/sql.py
@@ -361,7 +361,9 @@ def read_sql(
or DBAPI2 connection (fallback mode)
Using SQLAlchemy makes it possible to use any DB supported by that
- library. If a DBAPI2 object, only sqlite3 is supported.
+ library. If a DBAPI2 object, only sqlite3 is supported. The user is responsible
+ for engine disposal and connection closure for the SQLAlchemy connectable. See
+ `here <https://docs.sqlalchemy.org/en/13/core/connections.html>`_
index_col : string or list of strings, optional, default: None
Column(s) to set as index(MultiIndex).
coerce_float : boolean, default True
| - [x] closes #23086
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29373 | 2019-11-02T19:16:28Z | 2019-11-22T16:19:28Z | 2019-11-22T16:19:27Z | 2019-11-22T16:19:34Z |
TST: new test for str to small float conversion dtype | diff --git a/pandas/tests/frame/test_dtypes.py b/pandas/tests/frame/test_dtypes.py
index 68844aeeb081e..07accdb47d252 100644
--- a/pandas/tests/frame/test_dtypes.py
+++ b/pandas/tests/frame/test_dtypes.py
@@ -1063,6 +1063,18 @@ def test_asarray_homogenous(self):
expected = np.array([[1, 1], [2, 2]], dtype="object")
tm.assert_numpy_array_equal(result, expected)
+ def test_str_to_small_float_conversion_type(self):
+ # GH 20388
+ np.random.seed(13)
+ col_data = [str(np.random.random() * 1e-12) for _ in range(5)]
+ result = pd.DataFrame(col_data, columns=["A"])
+ expected = pd.DataFrame(col_data, columns=["A"], dtype=object)
+ tm.assert_frame_equal(result, expected)
+ # change the dtype of the elements from object to float one by one
+ result.loc[result.index, "A"] = [float(x) for x in col_data]
+ expected = pd.DataFrame(col_data, columns=["A"], dtype=float)
+ tm.assert_frame_equal(result, expected)
+
class TestDataFrameDatetimeWithTZ:
def test_interleave(self, timezone_frame):
| - [x] closes #20388
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29372 | 2019-11-02T18:40:16Z | 2019-11-03T21:22:19Z | 2019-11-03T21:22:19Z | 2019-11-03T21:22:30Z |
Update setup.py to https | diff --git a/setup.py b/setup.py
index c75ad5896a439..3dd38bdb6adbb 100755
--- a/setup.py
+++ b/setup.py
@@ -165,7 +165,7 @@ def build_extensions(self):
(2-dimensional), handle the vast majority of typical use cases in finance,
statistics, social science, and many areas of engineering. For R users,
DataFrame provides everything that R's ``data.frame`` provides and much
-more. pandas is built on top of `NumPy <http://www.numpy.org>`__ and is
+more. pandas is built on top of `NumPy <https://www.numpy.org>`__ and is
intended to integrate well within a scientific computing environment with many
other 3rd party libraries.
@@ -209,11 +209,11 @@ def build_extensions(self):
LICENSE = "BSD"
AUTHOR = "The PyData Development Team"
EMAIL = "pydata@googlegroups.com"
-URL = "http://pandas.pydata.org"
+URL = "https://pandas.pydata.org"
DOWNLOAD_URL = ""
PROJECT_URLS = {
"Bug Tracker": "https://github.com/pandas-dev/pandas/issues",
- "Documentation": "http://pandas.pydata.org/pandas-docs/stable/",
+ "Documentation": "https://pandas.pydata.org/pandas-docs/stable/",
"Source Code": "https://github.com/pandas-dev/pandas",
}
CLASSIFIERS = [
| Avoids a redirect in browser when clicking these links.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29371 | 2019-11-02T18:08:06Z | 2019-11-02T19:49:01Z | 2019-11-02T19:49:01Z | 2019-11-02T19:49:06Z |
TST: Adding merge test for non-string columns [Ref 17962] | diff --git a/pandas/tests/reshape/merge/test_merge.py b/pandas/tests/reshape/merge/test_merge.py
index 37c0b57bc7581..dd51a1a6c8359 100644
--- a/pandas/tests/reshape/merge/test_merge.py
+++ b/pandas/tests/reshape/merge/test_merge.py
@@ -134,6 +134,18 @@ def test_merge_common(self):
exp = merge(self.df, self.df2, on=["key1", "key2"])
tm.assert_frame_equal(joined, exp)
+ def test_merge_non_string_columns(self):
+ # https://github.com/pandas-dev/pandas/issues/17962
+ # Checks that method runs for non string column names
+ left = pd.DataFrame(
+ {0: [1, 0, 1, 0], 1: [0, 1, 0, 0], 2: [0, 0, 2, 0], 3: [1, 0, 0, 3]}
+ )
+
+ right = left.astype(float)
+ expected = left
+ result = pd.merge(left, right)
+ tm.assert_frame_equal(expected, result)
+
def test_merge_index_as_on_arg(self):
# GH14355
| - [x] closes #17962
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29370 | 2019-11-02T17:55:39Z | 2019-11-03T21:23:01Z | 2019-11-03T21:23:01Z | 2019-11-03T21:23:12Z |
GH:11670: possible bug when calculating mean of DataFrame? | diff --git a/pandas/tests/frame/test_analytics.py b/pandas/tests/frame/test_analytics.py
index e99208ac78e15..f694689fa9dfb 100644
--- a/pandas/tests/frame/test_analytics.py
+++ b/pandas/tests/frame/test_analytics.py
@@ -1,4 +1,5 @@
from datetime import timedelta
+from decimal import Decimal
import operator
from string import ascii_lowercase
import warnings
@@ -1075,6 +1076,29 @@ def test_mean_excludeds_datetimes(self, tz):
expected = pd.Series()
tm.assert_series_equal(result, expected)
+ def test_mean_mixed_string_decimal(self):
+ # GH 11670
+ # possible bug when calculating mean of DataFrame?
+
+ d = [
+ {"A": 2, "B": None, "C": Decimal("628.00")},
+ {"A": 1, "B": None, "C": Decimal("383.00")},
+ {"A": 3, "B": None, "C": Decimal("651.00")},
+ {"A": 2, "B": None, "C": Decimal("575.00")},
+ {"A": 4, "B": None, "C": Decimal("1114.00")},
+ {"A": 1, "B": "TEST", "C": Decimal("241.00")},
+ {"A": 2, "B": None, "C": Decimal("572.00")},
+ {"A": 4, "B": None, "C": Decimal("609.00")},
+ {"A": 3, "B": None, "C": Decimal("820.00")},
+ {"A": 5, "B": None, "C": Decimal("1223.00")},
+ ]
+
+ df = pd.DataFrame(d)
+
+ result = df.mean()
+ expected = pd.Series([2.7, 681.6], index=["A", "C"])
+ tm.assert_series_equal(result, expected)
+
def test_var_std(self, datetime_frame):
result = datetime_frame.std(ddof=4)
expected = datetime_frame.apply(lambda x: x.std(ddof=4))
| - [x] closes #11670
- [x] tests added / passed
- [x] passes `black pandas`
- [x] added test test_mean_mixed_string_float | https://api.github.com/repos/pandas-dev/pandas/pulls/29369 | 2019-11-02T17:39:28Z | 2019-11-03T14:59:06Z | 2019-11-03T14:59:06Z | 2019-11-03T14:59:10Z |
Fix pipe docs | diff --git a/doc/source/getting_started/basics.rst b/doc/source/getting_started/basics.rst
index 9b97aa25a9240..125990f7cadcd 100644
--- a/doc/source/getting_started/basics.rst
+++ b/doc/source/getting_started/basics.rst
@@ -753,28 +753,51 @@ on an entire ``DataFrame`` or ``Series``, row- or column-wise, or elementwise.
Tablewise function application
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-``DataFrames`` and ``Series`` can of course just be passed into functions.
+``DataFrames`` and ``Series`` can be passed into functions.
However, if the function needs to be called in a chain, consider using the :meth:`~DataFrame.pipe` method.
-Compare the following
-.. code-block:: python
+First some setup:
+
+.. ipython:: python
- # f, g, and h are functions taking and returning ``DataFrames``
- >>> f(g(h(df), arg1=1), arg2=2, arg3=3)
+ def extract_city_name(df):
+ """
+ Chicago, IL -> Chicago for city_name column
+ """
+ df['city_name'] = df['city_and_code'].str.split(",").str.get(0)
+ return df
-with the equivalent
+ def add_country_name(df, country_name=None):
+ """
+ Chicago -> Chicago-US for city_name column
+ """
+ col = 'city_name'
+ df['city_and_country'] = df[col] + country_name
+ return df
-.. code-block:: python
+ df_p = pd.DataFrame({'city_and_code': ['Chicago, IL']})
+
+
+``extract_city_name`` and ``add_country_name`` are functions taking and returning ``DataFrames``.
+
+Now compare the following:
+
+.. ipython:: python
+
+ add_country_name(extract_city_name(df_p), country_name='US')
+
+Is equivalent to:
+
+.. ipython:: python
- >>> (df.pipe(h)
- ... .pipe(g, arg1=1)
- ... .pipe(f, arg2=2, arg3=3))
+ (df_p.pipe(extract_city_name)
+ .pipe(add_country_name, country_name="US"))
Pandas encourages the second style, which is known as method chaining.
``pipe`` makes it easy to use your own or another library's functions
in method chains, alongside pandas' methods.
-In the example above, the functions ``f``, ``g``, and ``h`` each expected the ``DataFrame`` as the first positional argument.
+In the example above, the functions ``extract_city_name`` and ``add_country_name`` each expected a ``DataFrame`` as the first positional argument.
What if the function you wish to apply takes its data as, say, the second argument?
In this case, provide ``pipe`` with a tuple of ``(callable, data_keyword)``.
``.pipe`` will route the ``DataFrame`` to the argument specified in the tuple.
diff --git a/setup.cfg b/setup.cfg
index d4657100c1291..2dcb46584f19e 100644
--- a/setup.cfg
+++ b/setup.cfg
@@ -47,7 +47,6 @@ ignore = E402, # module level import not at top of file
E711, # comparison to none should be 'if cond is none:'
exclude =
- doc/source/getting_started/basics.rst
doc/source/development/contributing_docstring.rst
| - [x] closes #27054
See the image below which highlights what i've done here
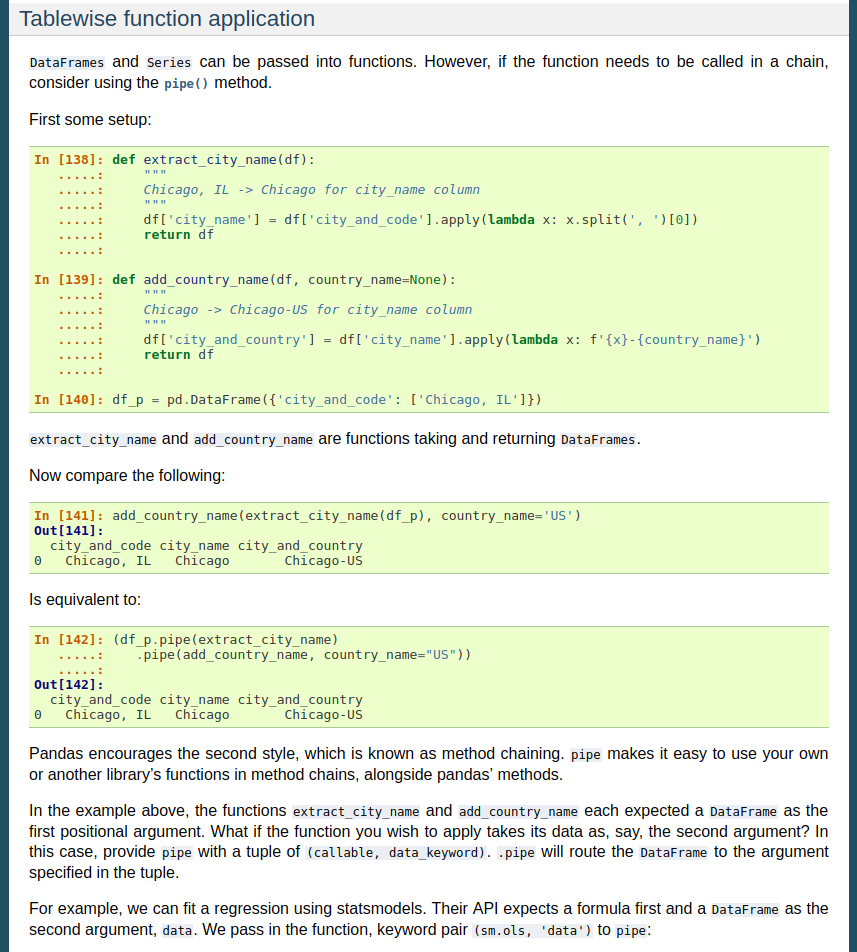
`flake8-rst doc/source --filename=/getting_started/basics.rst` runs clean
cc. @datapythonista to review | https://api.github.com/repos/pandas-dev/pandas/pulls/29368 | 2019-11-02T17:30:12Z | 2019-11-06T17:51:05Z | 2019-11-06T17:51:05Z | 2019-11-07T01:48:17Z |
TST: Adding map test for dict with np.nan key [Ref 17648] | diff --git a/pandas/tests/series/test_apply.py b/pandas/tests/series/test_apply.py
index e56294669a546..971ce5b18c323 100644
--- a/pandas/tests/series/test_apply.py
+++ b/pandas/tests/series/test_apply.py
@@ -581,6 +581,14 @@ def test_map_defaultdict(self):
expected = Series(["stuff", "blank", "blank"], index=["a", "b", "c"])
tm.assert_series_equal(result, expected)
+ def test_map_dict_na_key(self):
+ # https://github.com/pandas-dev/pandas/issues/17648
+ # Checks that np.nan key is appropriately mapped
+ s = Series([1, 2, np.nan])
+ expected = Series(["a", "b", "c"])
+ result = s.map({1: "a", 2: "b", np.nan: "c"})
+ tm.assert_series_equal(result, expected)
+
def test_map_dict_subclass_with_missing(self):
"""
Test Series.map with a dictionary subclass that defines __missing__,
| - [x] closes #17648
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29367 | 2019-11-02T17:18:17Z | 2019-11-03T01:57:00Z | 2019-11-03T01:57:00Z | 2019-11-03T01:57:09Z |
29213: Dataframe Constructor from List of List and non-iterables | diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 40efc4c65476a..d753ebc2c78ea 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -460,21 +460,29 @@ def __init__(
data = list(data)
if len(data) > 0:
if is_list_like(data[0]) and getattr(data[0], "ndim", 1) == 1:
- if is_named_tuple(data[0]) and columns is None:
- columns = data[0]._fields
- arrays, columns = to_arrays(data, columns, dtype=dtype)
- columns = ensure_index(columns)
-
- # set the index
- if index is None:
- if isinstance(data[0], Series):
- index = get_names_from_index(data)
- elif isinstance(data[0], Categorical):
- index = ibase.default_index(len(data[0]))
- else:
- index = ibase.default_index(len(data))
-
- mgr = arrays_to_mgr(arrays, columns, index, columns, dtype=dtype)
+ # try to infer that all elements are list-like as well
+ try:
+ if is_named_tuple(data[0]) and columns is None:
+ columns = data[0]._fields
+ arrays, columns = to_arrays(data, columns, dtype=dtype)
+ columns = ensure_index(columns)
+
+ # set the index
+ if index is None:
+ if isinstance(data[0], Series):
+ index = get_names_from_index(data)
+ elif isinstance(data[0], Categorical):
+ index = ibase.default_index(len(data[0]))
+ else:
+ index = ibase.default_index(len(data))
+
+ mgr = arrays_to_mgr(
+ arrays, columns, index, columns, dtype=dtype
+ )
+
+ except TypeError:
+ mgr = init_ndarray(data, index, columns, dtype=dtype, copy=copy)
+
else:
mgr = init_ndarray(data, index, columns, dtype=dtype, copy=copy)
else:
diff --git a/pandas/tests/frame/test_constructors.py b/pandas/tests/frame/test_constructors.py
index aa00cf234d9ee..1938544515ac1 100644
--- a/pandas/tests/frame/test_constructors.py
+++ b/pandas/tests/frame/test_constructors.py
@@ -1055,6 +1055,32 @@ def test_constructor_list_of_lists(self):
result = DataFrame(data)
tm.assert_frame_equal(result, expected)
+ def test_constructor_list_containing_lists_and_non_iterables(self):
+ # GH-29213
+ # First element iterable
+ result = DataFrame([[1, 2, 3], 4])
+ expected = DataFrame(Series([[1, 2, 3], 4]))
+ tm.assert_frame_equal(result, expected)
+
+ # First element non-iterable
+ result = DataFrame([4, [1, 2, 3]])
+ expected = DataFrame(Series([4, [1, 2, 3]]))
+ tm.assert_frame_equal(result, expected)
+
+ def test_constructor_from_dict_lists_and_non_iterables(self):
+ # GH-29213
+ # First dic.values() element iterable
+ dic = OrderedDict([["a", [1, 2, 3]], ["b", 4]])
+ result = DataFrame.from_dict(dic, orient="index")
+ expected = DataFrame(Series([[1, 2, 3], 4], ["a", "b"]))
+ tm.assert_frame_equal(result, expected)
+
+ # First dict.values() element non-iterable
+ dic = OrderedDict([["b", 4], ["a", [1, 2, 3]]])
+ result = DataFrame.from_dict(dic, orient="index")
+ expected = DataFrame(Series([4, [1, 2, 3]], ["b", "a"]))
+ tm.assert_frame_equal(result, expected)
+
def test_constructor_sequence_like(self):
# GH 3783
# collections.Squence like
| - [x] closes https://github.com/pandas-dev/pandas/issues/29213
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
---
## What
Whilst the original issue is from the factory method `DataFrame.from_dict(d, orient='index')`, the main issue is the *order* of elements within the list in the main constructor:
For example:
```
pd.DataFrame([[1, 2, 3], 4]) # doesn't work
# TypeError: object of type 'int' has no len()
In [2]: pd.DataFrame([4, [1, 2, 3]]) # works, creates a 1D DataFrame
Out[2]:
0
0 4
1 [1, 2, 3]
```
## Current Constructor Logic on List Argument
- Current logic looks at the [first element](https://github.com/pandas-dev/pandas/blob/master/pandas/core/frame.py#L462-L465), and *infers* that all other elements are iterables as well.
- if all elements in the list are iterables, it generates a 2D DataFrame.
- if any element from index-1 onwards is a non-iterable, it doesn't have a len() method and fails.
## Proposed Solution
- Based on the first element, *try* to infer that all elements are iterables as well.
- If not all subsequent elements are iterables, then return 1D DataFrame. This would be the same behaviour as it would have been if the first element in the list is non-iterable.
- This should not have performance degradation as there is noneed to check if all elements in the list are iterables.
## Note
- This does not solve the issue whereby we have iterables of different types (such as lists and strings...)
```
In [2]: pd.DataFrame([[1, 2, 3], 'foobar'])
Out[2]:
0 1 2 3 4 5
0 1 2 3 None None None
1 f o o b a r
``` | https://api.github.com/repos/pandas-dev/pandas/pulls/29366 | 2019-11-02T17:13:35Z | 2020-02-02T01:24:19Z | null | 2020-02-03T10:12:57Z |
Test added: uint64 multicolumn sort | diff --git a/pandas/tests/frame/test_sorting.py b/pandas/tests/frame/test_sorting.py
index 9ea78b974fcbb..6ed245b6ebb98 100644
--- a/pandas/tests/frame/test_sorting.py
+++ b/pandas/tests/frame/test_sorting.py
@@ -735,3 +735,26 @@ def test_sort_index_na_position_with_categories_raises(self):
with pytest.raises(ValueError):
df.sort_values(by="c", ascending=False, na_position="bad_position")
+
+ def test_sort_multicolumn_uint64(self):
+ # GH9918
+ # uint64 multicolumn sort
+
+ df = pd.DataFrame(
+ {
+ "a": pd.Series([18446637057563306014, 1162265347240853609]),
+ "b": pd.Series([1, 2]),
+ }
+ )
+ df["a"] = df["a"].astype(np.uint64)
+ result = df.sort_values(["a", "b"])
+
+ expected = pd.DataFrame(
+ {
+ "a": pd.Series([18446637057563306014, 1162265347240853609]),
+ "b": pd.Series([1, 2]),
+ },
+ index=pd.Index([1, 0]),
+ )
+
+ tm.assert_frame_equal(result, expected)
| - [x] closes #9918
- [x] tests added / passed
- [x] passes `black pandas`
- [x] added test test_sort_multicolumn_uint64 | https://api.github.com/repos/pandas-dev/pandas/pulls/29365 | 2019-11-02T17:02:53Z | 2019-11-03T15:05:47Z | 2019-11-03T15:05:46Z | 2019-11-03T15:05:51Z |
26302 add typing to assert star equal funcs | diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index af9fe4846b27d..af7330bda4d4e 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -10,3 +10,853 @@
FutureWarning,
stacklevel=2,
)
+
+
+# def assert_almost_equal(
+# left, right, check_dtype="equiv", check_less_precise=False, **kwargs
+# ):
+# """
+# Check that the left and right objects are approximately equal.
+
+# By approximately equal, we refer to objects that are numbers or that
+# contain numbers which may be equivalent to specific levels of precision.
+
+# Parameters
+# ----------
+# left : object
+# right : object
+# check_dtype : bool or {'equiv'}, default 'equiv'
+# Check dtype if both a and b are the same type. If 'equiv' is passed in,
+# then `RangeIndex` and `Int64Index` are also considered equivalent
+# when doing type checking.
+# check_less_precise : bool or int, default False
+# Specify comparison precision. 5 digits (False) or 3 digits (True)
+# after decimal points are compared. If int, then specify the number
+# of digits to compare.
+
+# When comparing two numbers, if the first number has magnitude less
+# than 1e-5, we compare the two numbers directly and check whether
+# they are equivalent within the specified precision. Otherwise, we
+# compare the **ratio** of the second number to the first number and
+# check whether it is equivalent to 1 within the specified precision.
+# """
+
+# if isinstance(left, pd.Index):
+# assert_index_equal(
+# left,
+# right,
+# check_exact=False,
+# exact=check_dtype,
+# check_less_precise=check_less_precise,
+# **kwargs,
+# )
+
+# elif isinstance(left, pd.Series):
+# assert_series_equal(
+# left,
+# right,
+# check_exact=False,
+# check_dtype=check_dtype,
+# check_less_precise=check_less_precise,
+# **kwargs,
+# )
+
+# elif isinstance(left, pd.DataFrame):
+# assert_frame_equal(
+# left,
+# right,
+# check_exact=False,
+# check_dtype=check_dtype,
+# check_less_precise=check_less_precise,
+# **kwargs,
+# )
+
+# else:
+# # Other sequences.
+# if check_dtype:
+# if is_number(left) and is_number(right):
+# # Do not compare numeric classes, like np.float64 and float.
+# pass
+# elif is_bool(left) and is_bool(right):
+# # Do not compare bool classes, like np.bool_ and bool.
+# pass
+# else:
+# if isinstance(left, np.ndarray) or isinstance(right, np.ndarray):
+# obj = "numpy array"
+# else:
+# obj = "Input"
+# assert_class_equal(left, right, obj=obj)
+# _testing.assert_almost_equal(
+# left,
+# right,
+# check_dtype=check_dtype,
+# check_less_precise=check_less_precise,
+# **kwargs,
+# )
+
+
+# def assert_class_equal(left, right, exact=True, obj="Input"):
+# """checks classes are equal."""
+# __tracebackhide__ = True
+
+# def repr_class(x):
+# if isinstance(x, Index):
+# # return Index as it is to include values in the error message
+# return x
+
+# try:
+# return x.__class__.__name__
+# except AttributeError:
+# return repr(type(x))
+
+# if exact == "equiv":
+# if type(left) != type(right):
+# # allow equivalence of Int64Index/RangeIndex
+# types = {type(left).__name__, type(right).__name__}
+# if len(types - {"Int64Index", "RangeIndex"}):
+# msg = "{obj} classes are not equivalent".format(obj=obj)
+# raise_assert_detail(obj, msg, repr_class(left), repr_class(right))
+# elif exact:
+# if type(left) != type(right):
+# msg = "{obj} classes are different".format(obj=obj)
+# raise_assert_detail(obj, msg, repr_class(left), repr_class(right))
+
+
+# def assert_attr_equal(attr, left, right, obj="Attributes"):
+# """checks attributes are equal. Both objects must have attribute.
+
+# Parameters
+# ----------
+# attr : str
+# Attribute name being compared.
+# left : object
+# right : object
+# obj : str, default 'Attributes'
+# Specify object name being compared, internally used to show appropriate
+# assertion message
+# """
+# __tracebackhide__ = True
+
+# left_attr = getattr(left, attr)
+# right_attr = getattr(right, attr)
+
+# if left_attr is right_attr:
+# return True
+# elif (
+# is_number(left_attr)
+# and np.isnan(left_attr)
+# and is_number(right_attr)
+# and np.isnan(right_attr)
+# ):
+# # np.nan
+# return True
+
+# try:
+# result = left_attr == right_attr
+# except TypeError:
+# # datetimetz on rhs may raise TypeError
+# result = False
+# if not isinstance(result, bool):
+# result = result.all()
+
+# if result:
+# return True
+# else:
+# msg = 'Attribute "{attr}" are different'.format(attr=attr)
+# raise_assert_detail(obj, msg, left_attr, right_attr)
+
+
+# def assert_categorical_equal(
+# left: Categorical,
+# right: Categorical,
+# check_dtype: bool = True,
+# check_category_order: bool = True,
+# obj: str = "Categorical",
+# ) -> None:
+# """Test that Categoricals are equivalent.
+
+# Parameters
+# ----------
+# left : Categorical
+# right : Categorical
+# check_dtype : bool, default True
+# Check that integer dtype of the codes are the same
+# check_category_order : bool, default True
+# Whether the order of the categories should be compared, which
+# implies identical integer codes. If False, only the resulting
+# values are compared. The ordered attribute is
+# checked regardless.
+# obj : str, default 'Categorical'
+# Specify object name being compared, internally used to show appropriate
+# assertion message
+# """
+# _check_isinstance(left, right, Categorical)
+
+# if check_category_order:
+# assert_index_equal(
+# left.categories, right.categories, obj="{obj}.categories".format(obj=obj)
+# )
+# assert_numpy_array_equal(
+# left.codes,
+# right.codes,
+# check_dtype=check_dtype,
+# obj="{obj}.codes".format(obj=obj),
+# )
+# else:
+# assert_index_equal(
+# left.categories.sort_values(),
+# right.categories.sort_values(),
+# obj="{obj}.categories".format(obj=obj),
+# )
+# assert_index_equal(
+# left.categories.take(left.codes),
+# right.categories.take(right.codes),
+# obj="{obj}.values".format(obj=obj),
+# )
+
+# assert_attr_equal("ordered", left, right, obj=obj)
+
+
+# def assert_interval_array_equal(
+# left: IntervalArray,
+# right: IntervalArray,
+# exact: str = "equiv",
+# obj: str = "IntervalArray",
+# ) -> None:
+# """Test that two IntervalArrays are equivalent.
+
+# Parameters
+# ----------
+# left, right : IntervalArray
+# The IntervalArrays to compare.
+# exact : bool or {'equiv'}, default 'equiv'
+# Whether to check the Index class, dtype and inferred_type
+# are identical. If 'equiv', then RangeIndex can be substituted for
+# Int64Index as well.
+# obj : str, default 'IntervalArray'
+# Specify object name being compared, internally used to show appropriate
+# assertion message
+# """
+# assert_index_equal(
+# left.left, right.left, exact=exact, obj="{obj}.left".format(obj=obj)
+# )
+# assert_index_equal(
+# left.right, right.right, exact=exact, obj="{obj}.left".format(obj=obj)
+# )
+# assert_attr_equal("closed", left, right, obj=obj)
+
+
+# def assert_period_array_equal(
+# left: PeriodArray, right: PeriodArray, obj: str = "PeriodArray"
+# ) -> None:
+# _check_isinstance(left, right, PeriodArray)
+
+# assert_numpy_array_equal(
+# left._data, right._data, obj="{obj}.values".format(obj=obj)
+# )
+# assert_attr_equal("freq", left, right, obj=obj)
+
+
+# def assert_datetime_array_equal(
+# left: DatetimeArray, right: DatetimeArray, obj: str = "DatetimeArray"
+# ) -> None:
+# __tracebackhide__ = True
+# _check_isinstance(left, right, DatetimeArray)
+
+# assert_numpy_array_equal(left._data, right._data, obj="{obj}._data".format(obj=obj))
+# assert_attr_equal("freq", left, right, obj=obj)
+# assert_attr_equal("tz", left, right, obj=obj)
+
+
+# def assert_timedelta_array_equal(
+# left: TimedeltaArray, right: TimedeltaArray, obj: str = "TimedeltaArray"
+# ) -> None:
+# __tracebackhide__ = True
+# _check_isinstance(left, right, TimedeltaArray)
+# assert_numpy_array_equal(left._data, right._data, obj="{obj}._data".format(obj=obj))
+# assert_attr_equal("freq", left, right, obj=obj)
+
+
+# def assert_numpy_array_equal(
+# left: np.ndarray,
+# right: np.ndarray,
+# strict_nan: bool = False,
+# check_dtype: bool = True,
+# err_msg: Optional[str] = None,
+# check_same: Optional[str] = None,
+# obj: str = "numpy array",
+# ) -> None:
+# """ Checks that 'np.ndarray' is equivalent
+
+# Parameters
+# ----------
+# left : np.ndarray or iterable
+# right : np.ndarray or iterable
+# strict_nan : bool, default False
+# If True, consider NaN and None to be different.
+# check_dtype: bool, default True
+# check dtype if both a and b are np.ndarray
+# err_msg : str, default None
+# If provided, used as assertion message
+# check_same : None|'copy'|'same', default None
+# Ensure left and right refer/do not refer to the same memory area
+# obj : str, default 'numpy array'
+# Specify object name being compared, internally used to show appropriate
+# assertion message
+# """
+# __tracebackhide__ = True
+
+# # instance validation
+# # Show a detailed error message when classes are different
+# assert_class_equal(left, right, obj=obj)
+# # both classes must be an np.ndarray
+# _check_isinstance(left, right, np.ndarray)
+
+# def _get_base(obj):
+# return obj.base if getattr(obj, "base", None) is not None else obj
+
+# left_base = _get_base(left)
+# right_base = _get_base(right)
+
+# if check_same == "same":
+# if left_base is not right_base:
+# msg = "{left!r} is not {right!r}".format(left=left_base, right=right_base)
+# raise AssertionError(msg)
+# elif check_same == "copy":
+# if left_base is right_base:
+# msg = "{left!r} is {right!r}".format(left=left_base, right=right_base)
+# raise AssertionError(msg)
+
+# def _raise(left, right, err_msg):
+# if err_msg is None:
+# if left.shape != right.shape:
+# raise_assert_detail(
+# obj,
+# "{obj} shapes are different".format(obj=obj),
+# left.shape,
+# right.shape,
+# )
+
+# diff = 0
+# for l, r in zip(left, right):
+# # count up differences
+# if not array_equivalent(l, r, strict_nan=strict_nan):
+# diff += 1
+
+# diff = diff * 100.0 / left.size
+# msg = "{obj} values are different ({pct} %)".format(
+# obj=obj, pct=np.round(diff, 5)
+# )
+# raise_assert_detail(obj, msg, left, right)
+
+# raise AssertionError(err_msg)
+
+# # compare shape and values
+# if not array_equivalent(left, right, strict_nan=strict_nan):
+# _raise(left, right, err_msg)
+
+# if check_dtype:
+# if isinstance(left, np.ndarray) and isinstance(right, np.ndarray):
+# assert_attr_equal("dtype", left, right, obj=obj)
+
+
+# def assert_extension_array_equal(
+# left, right, check_dtype=True, check_less_precise=False, check_exact=False
+# ):
+# """Check that left and right ExtensionArrays are equal.
+
+# Parameters
+# ----------
+# left, right : ExtensionArray
+# The two arrays to compare
+# check_dtype : bool, default True
+# Whether to check if the ExtensionArray dtypes are identical.
+# check_less_precise : bool or int, default False
+# Specify comparison precision. Only used when check_exact is False.
+# 5 digits (False) or 3 digits (True) after decimal points are compared.
+# If int, then specify the digits to compare.
+# check_exact : bool, default False
+# Whether to compare number exactly.
+
+# Notes
+# -----
+# Missing values are checked separately from valid values.
+# A mask of missing values is computed for each and checked to match.
+# The remaining all-valid values are cast to object dtype and checked.
+# """
+# assert isinstance(left, ExtensionArray), "left is not an ExtensionArray"
+# assert isinstance(right, ExtensionArray), "right is not an ExtensionArray"
+# if check_dtype:
+# assert_attr_equal("dtype", left, right, obj="ExtensionArray")
+
+# if hasattr(left, "asi8") and type(right) == type(left):
+# # Avoid slow object-dtype comparisons
+# assert_numpy_array_equal(left.asi8, right.asi8)
+# return
+
+# left_na = np.asarray(left.isna())
+# right_na = np.asarray(right.isna())
+# assert_numpy_array_equal(left_na, right_na, obj="ExtensionArray NA mask")
+
+# left_valid = np.asarray(left[~left_na].astype(object))
+# right_valid = np.asarray(right[~right_na].astype(object))
+# if check_exact:
+# assert_numpy_array_equal(left_valid, right_valid, obj="ExtensionArray")
+# else:
+# _testing.assert_almost_equal(
+# left_valid,
+# right_valid,
+# check_dtype=check_dtype,
+# check_less_precise=check_less_precise,
+# obj="ExtensionArray",
+# )
+
+
+# # This could be refactored to use the NDFrame.equals method
+# def assert_series_equal(
+# left: Series,
+# right: Series,
+# check_dtype: bool = True,
+# check_index_type: str = "equiv",
+# check_series_type: bool = True,
+# check_less_precise: bool = False,
+# check_names: bool = True,
+# check_exact: bool = False,
+# check_datetimelike_compat: bool = False,
+# check_categorical: bool = True,
+# obj: str = "Series",
+# ) -> None:
+# """
+# Check that left and right Series are equal.
+
+# Parameters
+# ----------
+# left : Series
+# right : Series
+# check_dtype : bool, default True
+# Whether to check the Series dtype is identical.
+# check_index_type : bool or {'equiv'}, default 'equiv'
+# Whether to check the Index class, dtype and inferred_type
+# are identical.
+# check_series_type : bool, default True
+# Whether to check the Series class is identical.
+# check_less_precise : bool or int, default False
+# Specify comparison precision. Only used when check_exact is False.
+# 5 digits (False) or 3 digits (True) after decimal points are compared.
+# If int, then specify the digits to compare.
+
+# When comparing two numbers, if the first number has magnitude less
+# than 1e-5, we compare the two numbers directly and check whether
+# they are equivalent within the specified precision. Otherwise, we
+# compare the **ratio** of the second number to the first number and
+# check whether it is equivalent to 1 within the specified precision.
+# check_names : bool, default True
+# Whether to check the Series and Index names attribute.
+# check_exact : bool, default False
+# Whether to compare number exactly.
+# check_datetimelike_compat : bool, default False
+# Compare datetime-like which is comparable ignoring dtype.
+# check_categorical : bool, default True
+# Whether to compare internal Categorical exactly.
+# obj : str, default 'Series'
+# Specify object name being compared, internally used to show appropriate
+# assertion message.
+# """
+# __tracebackhide__ = True
+
+# # instance validation
+# _check_isinstance(left, right, Series)
+
+# if check_series_type:
+# # ToDo: There are some tests using rhs is sparse
+# # lhs is dense. Should use assert_class_equal in future
+# assert isinstance(left, type(right))
+# # assert_class_equal(left, right, obj=obj)
+
+# # length comparison
+# if len(left) != len(right):
+# msg1 = "{len}, {left}".format(len=len(left), left=left.index)
+# msg2 = "{len}, {right}".format(len=len(right), right=right.index)
+# raise_assert_detail(obj, "Series length are different", msg1, msg2)
+
+# # index comparison
+# assert_index_equal(
+# left.index,
+# right.index,
+# exact=check_index_type,
+# check_names=check_names,
+# check_less_precise=check_less_precise,
+# check_exact=check_exact,
+# check_categorical=check_categorical,
+# obj="{obj}.index".format(obj=obj),
+# )
+
+# if check_dtype:
+# # We want to skip exact dtype checking when `check_categorical`
+# # is False. We'll still raise if only one is a `Categorical`,
+# # regardless of `check_categorical`
+# if (
+# is_categorical_dtype(left)
+# and is_categorical_dtype(right)
+# and not check_categorical
+# ):
+# pass
+# else:
+# assert_attr_equal(
+# "dtype", left, right, obj="Attributes of {obj}".format(obj=obj)
+# )
+
+# if check_exact:
+# assert_numpy_array_equal(
+# left._internal_get_values(),
+# right._internal_get_values(),
+# check_dtype=check_dtype,
+# obj="{obj}".format(obj=obj),
+# )
+# elif check_datetimelike_compat:
+# # we want to check only if we have compat dtypes
+# # e.g. integer and M|m are NOT compat, but we can simply check
+# # the values in that case
+# if needs_i8_conversion(left) or needs_i8_conversion(right):
+
+# # datetimelike may have different objects (e.g. datetime.datetime
+# # vs Timestamp) but will compare equal
+# if not Index(left.values).equals(Index(right.values)):
+# msg = (
+# "[datetimelike_compat=True] {left} is not equal to {right}."
+# ).format(left=left.values, right=right.values)
+# raise AssertionError(msg)
+# else:
+# assert_numpy_array_equal(
+# left._internal_get_values(),
+# right._internal_get_values(),
+# check_dtype=check_dtype,
+# )
+# elif is_interval_dtype(left) or is_interval_dtype(right):
+# left_array = cast(IntervalArray, left.array)
+# right_array = cast(IntervalArray, right.array)
+# assert_interval_array_equal(left_array, right_array)
+# elif is_extension_array_dtype(left.dtype) and is_datetime64tz_dtype(left.dtype):
+# # .values is an ndarray, but ._values is the ExtensionArray.
+# # TODO: Use .array
+# assert is_extension_array_dtype(right.dtype)
+# assert_extension_array_equal(left._values, right._values)
+# elif (
+# is_extension_array_dtype(left)
+# and not is_categorical_dtype(left)
+# and is_extension_array_dtype(right)
+# and not is_categorical_dtype(right)
+# ):
+# assert_extension_array_equal(left.array, right.array)
+# else:
+# _testing.assert_almost_equal(
+# left._internal_get_values(),
+# right._internal_get_values(),
+# check_less_precise=check_less_precise,
+# check_dtype=check_dtype,
+# obj="{obj}".format(obj=obj),
+# )
+
+# # metadata comparison
+# if check_names:
+# assert_attr_equal("name", left, right, obj=obj)
+
+# if check_categorical:
+# if is_categorical_dtype(left) or is_categorical_dtype(right):
+# assert_categorical_equal(
+# left.values, right.values, obj="{obj} category".format(obj=obj)
+# )
+
+
+# # This could be refactored to use the NDFrame.equals method
+# def assert_frame_equal(
+# left: Any,
+# right: Any,
+# check_dtype: bool = True,
+# check_index_type: str = "equiv",
+# check_column_type: str = "equiv",
+# check_frame_type: bool = True,
+# check_less_precise: bool = False,
+# check_names: bool = True,
+# by_blocks: bool = False,
+# check_exact: bool = False,
+# check_datetimelike_compat: bool = False,
+# check_categorical: bool = True,
+# check_like: bool = False,
+# obj: str = "DataFrame",
+# ) -> None:
+# """
+# Check that left and right DataFrame are equal.
+
+# This function is intended to compare two DataFrames and output any
+# differences. Is is mostly intended for use in unit tests.
+# Additional parameters allow varying the strictness of the
+# equality checks performed.
+
+# Parameters
+# ----------
+# left : Any
+# First DataFrame to compare.
+# right : Any
+# Second DataFrame to compare.
+# check_dtype : bool, default True
+# Whether to check the DataFrame dtype is identical.
+# check_index_type : bool or {'equiv'}, default 'equiv'
+# Whether to check the Index class, dtype and inferred_type
+# are identical.
+# check_column_type : bool or {'equiv'}, default 'equiv'
+# Whether to check the columns class, dtype and inferred_type
+# are identical. Is passed as the ``exact`` argument of
+# :func:`assert_index_equal`.
+# check_frame_type : bool, default True
+# Whether to check the DataFrame class is identical.
+# check_less_precise : bool or int, default False
+# Specify comparison precision. Only used when check_exact is False.
+# 5 digits (False) or 3 digits (True) after decimal points are compared.
+# If int, then specify the digits to compare.
+
+# When comparing two numbers, if the first number has magnitude less
+# than 1e-5, we compare the two numbers directly and check whether
+# they are equivalent within the specified precision. Otherwise, we
+# compare the **ratio** of the second number to the first number and
+# check whether it is equivalent to 1 within the specified precision.
+# check_names : bool, default True
+# Whether to check that the `names` attribute for both the `index`
+# and `column` attributes of the DataFrame is identical.
+# by_blocks : bool, default False
+# Specify how to compare internal data. If False, compare by columns.
+# If True, compare by blocks.
+# check_exact : bool, default False
+# Whether to compare number exactly.
+# check_datetimelike_compat : bool, default False
+# Compare datetime-like which is comparable ignoring dtype.
+# check_categorical : bool, default True
+# Whether to compare internal Categorical exactly.
+# check_like : bool, default False
+# If True, ignore the order of index & columns.
+# Note: index labels must match their respective rows
+# (same as in columns) - same labels must be with the same data.
+# obj : str, default 'DataFrame'
+# Specify object name being compared, internally used to show appropriate
+# assertion message.
+
+# See Also
+# --------
+# assert_series_equal : Equivalent method for asserting Series equality.
+# DataFrame.equals : Check DataFrame equality.
+
+# Examples
+# --------
+# This example shows comparing two DataFrames that are equal
+# but with columns of differing dtypes.
+
+# >>> from pandas.util.testing import assert_frame_equal
+# >>> df1 = pd.DataFrame({'a': [1, 2], 'b': [3, 4]})
+# >>> df2 = pd.DataFrame({'a': [1, 2], 'b': [3.0, 4.0]})
+
+# df1 equals itself.
+
+# >>> assert_frame_equal(df1, df1)
+
+# df1 differs from df2 as column 'b' is of a different type.
+
+# >>> assert_frame_equal(df1, df2)
+# Traceback (most recent call last):
+# ...
+# AssertionError: Attributes of DataFrame.iloc[:, 1] are different
+
+# Attribute "dtype" are different
+# [left]: int64
+# [right]: float64
+
+# Ignore differing dtypes in columns with check_dtype.
+
+# >>> assert_frame_equal(df1, df2, check_dtype=False)
+# """
+# __tracebackhide__ = True
+
+# # instance validation
+# _check_isinstance(left, right, DataFrame)
+
+# if check_frame_type:
+# assert isinstance(left, type(right))
+# # assert_class_equal(left, right, obj=obj)
+
+# # shape comparison
+# if left.shape != right.shape:
+# raise_assert_detail(
+# obj,
+# "{obj} shape mismatch".format(obj=obj),
+# "{shape!r}".format(shape=left.shape),
+# "{shape!r}".format(shape=right.shape),
+# )
+
+# if check_like:
+# left, right = left.reindex_like(right), right
+
+# # index comparison
+# assert_index_equal(
+# left.index,
+# right.index,
+# exact=check_index_type,
+# check_names=check_names,
+# check_less_precise=check_less_precise,
+# check_exact=check_exact,
+# check_categorical=check_categorical,
+# obj="{obj}.index".format(obj=obj),
+# )
+
+# # column comparison
+# assert_index_equal(
+# left.columns,
+# right.columns,
+# exact=check_column_type,
+# check_names=check_names,
+# check_less_precise=check_less_precise,
+# check_exact=check_exact,
+# check_categorical=check_categorical,
+# obj="{obj}.columns".format(obj=obj),
+# )
+
+# # compare by blocks
+# if by_blocks:
+# rblocks = right._to_dict_of_blocks()
+# lblocks = left._to_dict_of_blocks()
+# for dtype in list(set(list(lblocks.keys()) + list(rblocks.keys()))):
+# assert dtype in lblocks
+# assert dtype in rblocks
+# assert_frame_equal(
+# lblocks[dtype], rblocks[dtype], check_dtype=check_dtype, obj=obj
+# )
+
+# # compare by columns
+# else:
+# for i, col in enumerate(left.columns):
+# assert col in right
+# lcol = left.iloc[:, i]
+# rcol = right.iloc[:, i]
+# assert_series_equal(
+# lcol,
+# rcol,
+# check_dtype=check_dtype,
+# check_index_type=check_index_type,
+# check_less_precise=check_less_precise,
+# check_exact=check_exact,
+# check_names=check_names,
+# check_datetimelike_compat=check_datetimelike_compat,
+# check_categorical=check_categorical,
+# obj="{obj}.iloc[:, {idx}]".format(obj=obj, idx=i),
+# )
+
+
+# def assert_equal(
+# left: Union[DataFrame, AnyArrayLike],
+# right: Union[DataFrame, AnyArrayLike],
+# **kwargs,
+# ) -> None:
+# """
+# Wrapper for tm.assert_*_equal to dispatch to the appropriate test function.
+
+# Parameters
+# ----------
+# left : Index, Series, DataFrame, ExtensionArray, or np.ndarray
+# right : Index, Series, DataFrame, ExtensionArray, or np.ndarray
+# **kwargs
+# """
+# __tracebackhide__ = True
+
+# if isinstance(left, Index):
+# right = cast(Index, right)
+# assert_index_equal(left, right, **kwargs)
+# elif isinstance(left, Series):
+# right = cast(Series, right)
+# assert_series_equal(left, right, **kwargs)
+# elif isinstance(left, DataFrame):
+# right = cast(DataFrame, right)
+# assert_frame_equal(left, right, **kwargs)
+# elif isinstance(left, IntervalArray):
+# right = cast(IntervalArray, right)
+# assert_interval_array_equal(left, right, **kwargs)
+# elif isinstance(left, PeriodArray):
+# right = cast(PeriodArray, right)
+# assert_period_array_equal(left, right, **kwargs)
+# elif isinstance(left, DatetimeArray):
+# right = cast(DatetimeArray, right)
+# assert_datetime_array_equal(left, right, **kwargs)
+# elif isinstance(left, TimedeltaArray):
+# right = cast(TimedeltaArray, right)
+# assert_timedelta_array_equal(left, right, **kwargs)
+# elif isinstance(left, ExtensionArray):
+# right = cast(ExtensionArray, right)
+# assert_extension_array_equal(left, right, **kwargs)
+# elif isinstance(left, np.ndarray):
+# right = cast(np.ndarray, right)
+# assert_numpy_array_equal(left, right, **kwargs)
+# elif isinstance(left, str):
+# assert kwargs == {}
+# assert left == right
+# else:
+# raise NotImplementedError(type(left))
+
+
+# def assert_sp_array_equal(
+# left: pd.SparseArray,
+# right: pd.SparseArray,
+# check_dtype: bool = True,
+# check_kind: bool = True,
+# check_fill_value: bool = True,
+# consolidate_block_indices: bool = False,
+# ):
+# """Check that the left and right SparseArray are equal.
+
+# Parameters
+# ----------
+# left : SparseArray
+# right : SparseArray
+# check_dtype : bool, default True
+# Whether to check the data dtype is identical.
+# check_kind : bool, default True
+# Whether to just the kind of the sparse index for each column.
+# check_fill_value : bool, default True
+# Whether to check that left.fill_value matches right.fill_value
+# consolidate_block_indices : bool, default False
+# Whether to consolidate contiguous blocks for sparse arrays with
+# a BlockIndex. Some operations, e.g. concat, will end up with
+# block indices that could be consolidated. Setting this to true will
+# create a new BlockIndex for that array, with consolidated
+# block indices.
+# """
+
+# _check_isinstance(left, right, pd.SparseArray)
+
+# assert_numpy_array_equal(left.sp_values, right.sp_values, check_dtype=check_dtype)
+
+# # SparseIndex comparison
+# assert isinstance(left.sp_index, pd._libs.sparse.SparseIndex)
+# assert isinstance(right.sp_index, pd._libs.sparse.SparseIndex)
+
+# if not check_kind:
+# left_index = left.sp_index.to_block_index()
+# right_index = right.sp_index.to_block_index()
+# else:
+# left_index = left.sp_index
+# right_index = right.sp_index
+
+# if consolidate_block_indices and left.kind == "block":
+# # we'll probably remove this hack...
+# left_index = left_index.to_int_index().to_block_index()
+# right_index = right_index.to_int_index().to_block_index()
+
+# if not left_index.equals(right_index):
+# raise_assert_detail(
+# "SparseArray.index", "index are not equal", left_index, right_index
+# )
+# else:
+# # Just ensure a
+# pass
+
+# if check_fill_value:
+# assert_attr_equal("fill_value", left, right)
+# if check_dtype:
+# assert_attr_equal("dtype", left, right)
+# assert_numpy_array_equal(left.to_dense(), right.to_dense(), check_dtype=check_dtype)
| - [X] closes #26302
- [X] passes `black pandas`
- [X] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29364 | 2019-11-02T16:40:07Z | 2020-03-11T03:14:08Z | null | 2020-03-11T03:14:08Z |
Add documentation linking to sqlalchemy | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index f88c26c7bc782..3ba805fa71e8a 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -2646,6 +2646,10 @@ def to_sql(
con : sqlalchemy.engine.Engine or sqlite3.Connection
Using SQLAlchemy makes it possible to use any DB supported by that
library. Legacy support is provided for sqlite3.Connection objects.
+
+ Closing the connection is handled by the SQLAlchemy Engine. See `here \
+ <https://docs.sqlalchemy.org/en/13/core/connections.html>`_
+
schema : str, optional
Specify the schema (if database flavor supports this). If None, use
default schema.
diff --git a/pandas/io/sql.py b/pandas/io/sql.py
index e90e19649f645..57e9ae3de6b67 100644
--- a/pandas/io/sql.py
+++ b/pandas/io/sql.py
@@ -362,6 +362,10 @@ def read_sql(
Using SQLAlchemy makes it possible to use any DB supported by that
library. If a DBAPI2 object, only sqlite3 is supported.
+
+ Closing the connection is handled by the SQLAlchemy Engine. See `here \
+ <https://docs.sqlalchemy.org/en/13/core/connections.html>`_
+
index_col : string or list of strings, optional, default: None
Column(s) to set as index(MultiIndex).
coerce_float : boolean, default True
| - [x] closes #23086
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ n/a] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29363 | 2019-11-02T16:35:15Z | 2019-11-02T19:15:52Z | null | 2019-11-02T19:17:05Z |
Disallow non-scalar fill_value in maybe_upcast | diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index 304eeac87f64d..69c2e7fef365f 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -686,6 +686,8 @@ def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
dtype : if None, then use the dtype of the values, else coerce to this type
copy : if True always make a copy even if no upcast is required
"""
+ if not is_scalar(fill_value):
+ raise ValueError("fill_value must be a scalar")
if is_extension_type(values):
if copy:
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 51108d9a5a573..1f5a14a41e6a3 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -1286,6 +1286,10 @@ def diff(self, n: int, axis: int = 1) -> List["Block"]:
def shift(self, periods, axis=0, fill_value=None):
""" shift the block by periods, possibly upcast """
+ if not lib.is_scalar(fill_value):
+ # We could go further and require e.g. self._can_hold_element(fv)
+ raise ValueError("fill_value must be a scalar")
+
# convert integer to float if necessary. need to do a lot more than
# that, handle boolean etc also
new_values, fill_value = maybe_upcast(self.values, fill_value)
diff --git a/pandas/core/internals/construction.py b/pandas/core/internals/construction.py
index 176f4acd113fe..4a8216cc73264 100644
--- a/pandas/core/internals/construction.py
+++ b/pandas/core/internals/construction.py
@@ -97,6 +97,9 @@ def masked_rec_array_to_mgr(data, index, columns, dtype, copy):
# fill if needed
new_arrays = []
for fv, arr, col in zip(fill_value, arrays, arr_columns):
+ # TODO: numpy docs suggest fv must be scalar, but could it be
+ # non-scalar for object dtype?
+ assert lib.is_scalar(fv), fv
mask = ma.getmaskarray(data[col])
if mask.any():
arr, fv = maybe_upcast(arr, fill_value=fv, copy=True)
| Along with #29331 and #29332 this allows us to rule out non-scalar fill_value being passed to maybe_promote for _almost_ all cases, the last few of which we'll have to address individually.
The restriction in `Block.shift` could be an API change if you squint and tilt your head.
The assertion in masked_rec_array_to_mgr is based on my read of the numpy docs, but it'd be worth double-checking. | https://api.github.com/repos/pandas-dev/pandas/pulls/29362 | 2019-11-02T16:27:26Z | 2019-11-04T14:41:06Z | 2019-11-04T14:41:06Z | 2019-11-04T14:59:25Z |
CLN: requested follow-ups | diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index e3c7fef6f048f..a08ae66865e20 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -380,6 +380,18 @@ ctypedef fused algos_t:
def _validate_limit(nobs: int, limit=None) -> int:
+ """
+ Check that the `limit` argument is a positive integer.
+
+ Parameters
+ ----------
+ nobs : int
+ limit : object
+
+ Returns
+ -------
+ int
+ """
if limit is None:
lim = nobs
else:
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 51108d9a5a573..448d2faf8b85f 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -7,7 +7,7 @@
import numpy as np
-from pandas._libs import NaT, lib, tslib, writers
+from pandas._libs import NaT, algos as libalgos, lib, tslib, writers
from pandas._libs.index import convert_scalar
import pandas._libs.internals as libinternals
from pandas._libs.tslibs import Timedelta, conversion
@@ -393,10 +393,7 @@ def fillna(self, value, limit=None, inplace=False, downcast=None):
mask = isna(self.values)
if limit is not None:
- if not is_integer(limit):
- raise ValueError("Limit must be an integer")
- if limit < 1:
- raise ValueError("Limit must be greater than 0")
+ limit = libalgos._validate_limit(None, limit=limit)
mask[mask.cumsum(self.ndim - 1) > limit] = False
if not self._can_hold_na:
diff --git a/pandas/core/missing.py b/pandas/core/missing.py
index f2655c126b9e5..5a1bf6d37b081 100644
--- a/pandas/core/missing.py
+++ b/pandas/core/missing.py
@@ -11,7 +11,6 @@
ensure_float64,
is_datetime64_dtype,
is_datetime64tz_dtype,
- is_integer,
is_integer_dtype,
is_numeric_v_string_like,
is_scalar,
@@ -191,13 +190,7 @@ def interpolate_1d(
)
# default limit is unlimited GH #16282
- if limit is None:
- # limit = len(xvalues)
- pass
- elif not is_integer(limit):
- raise ValueError("Limit must be an integer")
- elif limit < 1:
- raise ValueError("Limit must be greater than 0")
+ limit = algos._validate_limit(nobs=None, limit=limit)
from pandas import Series
diff --git a/pandas/tests/reductions/test_reductions.py b/pandas/tests/reductions/test_reductions.py
index a04f8f0df1151..4dfe561831ced 100644
--- a/pandas/tests/reductions/test_reductions.py
+++ b/pandas/tests/reductions/test_reductions.py
@@ -299,15 +299,6 @@ def test_timedelta_ops(self):
result = td.to_frame().std()
assert result[0] == expected
- # invalid ops
- for op in ["skew", "kurt", "sem", "prod", "var"]:
- msg = "reduction operation '{}' not allowed for this dtype"
- with pytest.raises(TypeError, match=msg.format(op)):
- getattr(td, op)()
-
- with pytest.raises(TypeError, match=msg.format(op)):
- getattr(td.to_frame(), op)(numeric_only=False)
-
# GH#10040
# make sure NaT is properly handled by median()
s = Series([Timestamp("2015-02-03"), Timestamp("2015-02-07")])
@@ -318,6 +309,22 @@ def test_timedelta_ops(self):
)
assert s.diff().median() == timedelta(days=6)
+ @pytest.mark.parametrize("opname", ["skew", "kurt", "sem", "prod", "var"])
+ def test_invalid_td64_reductions(self, opname):
+ s = Series(
+ [Timestamp("20130101") + timedelta(seconds=i * i) for i in range(10)]
+ )
+ td = s.diff()
+
+ msg = "reduction operation '{op}' not allowed for this dtype"
+ msg = msg.format(op=opname)
+
+ with pytest.raises(TypeError, match=msg):
+ getattr(td, opname)()
+
+ with pytest.raises(TypeError, match=msg):
+ getattr(td.to_frame(), opname)(numeric_only=False)
+
def test_minmax_tz(self, tz_naive_fixture):
tz = tz_naive_fixture
# monotonic
| Also use the new _validate_limit in two places in the non-cython code | https://api.github.com/repos/pandas-dev/pandas/pulls/29360 | 2019-11-02T16:10:56Z | 2019-11-02T19:48:23Z | 2019-11-02T19:48:23Z | 2019-11-02T20:58:25Z |
GH 16051: DataFrame.replace() overwrites when values are non-numeric | diff --git a/pandas/tests/frame/test_replace.py b/pandas/tests/frame/test_replace.py
index 5eb2416d0dcd7..c30efa121262f 100644
--- a/pandas/tests/frame/test_replace.py
+++ b/pandas/tests/frame/test_replace.py
@@ -1295,3 +1295,30 @@ def test_replace_method(self, to_replace, method, expected):
result = df.replace(to_replace=to_replace, value=None, method=method)
expected = DataFrame(expected)
tm.assert_frame_equal(result, expected)
+
+ @pytest.mark.parametrize(
+ "df, to_replace, exp",
+ [
+ (
+ {"col1": [1, 2, 3], "col2": [4, 5, 6]},
+ {4: 5, 5: 6, 6: 7},
+ {"col1": [1, 2, 3], "col2": [5, 6, 7]},
+ ),
+ (
+ {"col1": [1, 2, 3], "col2": ["4", "5", "6"]},
+ {"4": "5", "5": "6", "6": "7"},
+ {"col1": [1, 2, 3], "col2": ["5", "6", "7"]},
+ ),
+ ],
+ )
+ def test_replace_commutative(self, df, to_replace, exp):
+ # GH 16051
+ # DataFrame.replace() overwrites when values are non-numeric
+ # also added to data frame whilst issue was for series
+
+ df = pd.DataFrame(df)
+
+ expected = pd.DataFrame(exp)
+ result = df.replace(to_replace)
+
+ tm.assert_frame_equal(result, expected)
diff --git a/pandas/tests/series/test_replace.py b/pandas/tests/series/test_replace.py
index 86a54922fcf86..ebfd468e034f9 100644
--- a/pandas/tests/series/test_replace.py
+++ b/pandas/tests/series/test_replace.py
@@ -306,6 +306,24 @@ def test_replace_with_no_overflowerror(self):
expected = pd.Series([0, 1, "100000000000000000001"])
tm.assert_series_equal(result, expected)
+ @pytest.mark.parametrize(
+ "ser, to_replace, exp",
+ [
+ ([1, 2, 3], {1: 2, 2: 3, 3: 4}, [2, 3, 4]),
+ (["1", "2", "3"], {"1": "2", "2": "3", "3": "4"}, ["2", "3", "4"]),
+ ],
+ )
+ def test_replace_commutative(self, ser, to_replace, exp):
+ # GH 16051
+ # DataFrame.replace() overwrites when values are non-numeric
+
+ series = pd.Series(ser)
+
+ expected = pd.Series(exp)
+ result = series.replace(to_replace)
+
+ tm.assert_series_equal(result, expected)
+
@pytest.mark.parametrize(
"ser, exp", [([1, 2, 3], [1, True, 3]), (["x", 2, 3], ["x", True, 3])]
)
@@ -316,4 +334,5 @@ def test_replace_no_cast(self, ser, exp):
series = pd.Series(ser)
result = series.replace(2, True)
expected = pd.Series(exp)
+
tm.assert_series_equal(result, expected)
| - [x] closes #16051
- [x] tests added and passed
- [x] passes `black pandas`
- [x] added new test test_replace_commutative | https://api.github.com/repos/pandas-dev/pandas/pulls/29359 | 2019-11-02T15:36:33Z | 2019-11-04T16:54:55Z | 2019-11-04T16:54:55Z | 2019-11-04T16:55:03Z |
TST: Apply method broken for empty integer series with datetime index | diff --git a/pandas/tests/series/test_apply.py b/pandas/tests/series/test_apply.py
index e56294669a546..8eac79ae826c3 100644
--- a/pandas/tests/series/test_apply.py
+++ b/pandas/tests/series/test_apply.py
@@ -170,6 +170,12 @@ def test_apply_categorical_with_nan_values(self, series):
expected = expected.astype(object)
tm.assert_series_equal(result, expected)
+ def test_apply_empty_integer_series_with_datetime_index(self):
+ # GH 21245
+ s = pd.Series([], index=pd.date_range(start="2018-01-01", periods=0), dtype=int)
+ result = s.apply(lambda x: x)
+ tm.assert_series_equal(result, s)
+
class TestSeriesAggregate:
def test_transform(self, string_series):
| I added a unit test for an edge case that was failing. Using the apply method on an empty integer series with a datetime index would throw an error.
- [x] closes #21245
- [x] 1 test added
- [x] passes pandas and flake8
| https://api.github.com/repos/pandas-dev/pandas/pulls/29358 | 2019-11-02T15:23:05Z | 2019-11-02T19:51:08Z | 2019-11-02T19:51:08Z | 2019-11-02T19:51:13Z |
ensure consistent structure for groupby on index and column | diff --git a/pandas/tests/groupby/test_grouping.py b/pandas/tests/groupby/test_grouping.py
index e1fd8d7da6833..979e7b2dd7ffc 100644
--- a/pandas/tests/groupby/test_grouping.py
+++ b/pandas/tests/groupby/test_grouping.py
@@ -14,6 +14,7 @@
date_range,
)
from pandas.core.groupby.grouper import Grouping
+from pandas.core.indexes.frozen import FrozenList
import pandas.util.testing as tm
# selection
@@ -641,6 +642,27 @@ def test_groupby_level_index_value_all_na(self):
)
tm.assert_frame_equal(result, expected)
+ @pytest.mark.parametrize(
+ "df",
+ [
+ pd.DataFrame([[1, 2, 3]], columns=["a", "b", "c"]).set_index("a"),
+ pd.DataFrame([[1, 2, 3], [4, 5, 6]], columns=["a", "b", "c"]).set_index(
+ "a"
+ ),
+ pd.DataFrame(
+ [[1, 2, 3], [4, 5, 6], [7, 8, 9]], columns=["a", "b", "c"]
+ ).set_index("a"),
+ ],
+ )
+ @pytest.mark.parametrize(
+ "method", ["mean", "median", "prod", "min", "max", "sum", "std", "var"]
+ )
+ def test_groupby_on_index_and_column_consistent_structure(self, df, method):
+ # https://github.com/pandas-dev/pandas/issues/17681
+ df_gb = df.groupby(["a", "c"])
+ result = getattr(df_gb, method)()
+ assert result.index.names == FrozenList(["a", "c"])
+
# get_group
# --------------------------------
| - [x] closes #17681
- [ ] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29357 | 2019-11-02T15:18:30Z | 2019-12-17T17:26:40Z | null | 2019-12-17T17:26:41Z |
TST: new test for subset of a MultiIndex dtype | diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 79c9fe2b60bd9..4430628ce3d92 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -1932,6 +1932,15 @@ def test_repeat(self):
m_df = Series(data, index=m_idx)
assert m_df.repeat(3).shape == (3 * len(data),)
+ def test_subsets_multiindex_dtype(self):
+ # GH 20757
+ data = [["x", 1]]
+ columns = [("a", "b", np.nan), ("a", "c", 0.0)]
+ df = DataFrame(data, columns=pd.MultiIndex.from_tuples(columns))
+ expected = df.dtypes.a.b
+ result = df.a.b.dtypes
+ tm.assert_series_equal(result, expected)
+
class TestSorted(Base):
""" everything you wanted to test about sorting """
| Tried unsuccessfully reproducing Issue #20757 on pandas: 0.23.0 and 0.26.0.dev0+734.g0de99558b.dirty. That's why I decided to keep the input data from Issue #20757 unchanged.
- [x] closes #20757
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29356 | 2019-11-02T15:11:47Z | 2019-11-16T22:06:10Z | 2019-11-16T22:06:09Z | 2019-11-16T22:06:14Z |
TST: Test type issue fix in empty groupby from DataFrame with categorical | diff --git a/pandas/tests/groupby/test_categorical.py b/pandas/tests/groupby/test_categorical.py
index 22a23407b2521..a187781ea214c 100644
--- a/pandas/tests/groupby/test_categorical.py
+++ b/pandas/tests/groupby/test_categorical.py
@@ -781,6 +781,22 @@ def test_categorical_no_compress():
tm.assert_numpy_array_equal(result, exp)
+def test_groupby_empty_with_category():
+ # GH-9614
+ # test fix for when group by on None resulted in
+ # coercion of dtype categorical -> float
+ df = pd.DataFrame(
+ {"A": [None] * 3, "B": pd.Categorical(["train", "train", "test"])}
+ )
+ result = df.groupby("A").first()["B"]
+ expected = pd.Series(
+ pd.Categorical([], categories=["test", "train"]),
+ index=pd.Series([], dtype="object", name="A"),
+ name="B",
+ )
+ tm.assert_series_equal(result, expected)
+
+
def test_sort():
# http://stackoverflow.com/questions/23814368/sorting-pandas-
| TST: Test type issue fix in empty groupby from DataFrame with categorical
closes #9614 | https://api.github.com/repos/pandas-dev/pandas/pulls/29355 | 2019-11-02T14:58:13Z | 2019-11-13T01:57:20Z | 2019-11-13T01:57:19Z | 2019-11-13T01:57:24Z |
Pv feature2 | diff --git a/pandas/tests/series/test_replace.py b/pandas/tests/series/test_replace.py
index e9d5a4b105a35..86a54922fcf86 100644
--- a/pandas/tests/series/test_replace.py
+++ b/pandas/tests/series/test_replace.py
@@ -305,3 +305,15 @@ def test_replace_with_no_overflowerror(self):
result = s.replace(["100000000000000000000"], [1])
expected = pd.Series([0, 1, "100000000000000000001"])
tm.assert_series_equal(result, expected)
+
+ @pytest.mark.parametrize(
+ "ser, exp", [([1, 2, 3], [1, True, 3]), (["x", 2, 3], ["x", True, 3])]
+ )
+ def test_replace_no_cast(self, ser, exp):
+ # GH 9113
+ # BUG: replace int64 dtype with bool coerces to int64
+
+ series = pd.Series(ser)
+ result = series.replace(2, True)
+ expected = pd.Series(exp)
+ tm.assert_series_equal(result, expected)
| - [x] closes #9113
- [x] tests added / passed
- [x] passes `black pandas`
- [x] new test added: test_replace_no_cast() | https://api.github.com/repos/pandas-dev/pandas/pulls/29354 | 2019-11-02T14:50:34Z | 2019-11-03T01:42:18Z | 2019-11-03T01:42:18Z | 2019-11-03T01:42:36Z |
BUG: Issue #29128 Series.var not returning the correct result | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 664fcc91bacc4..2ae3379a6a23c 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -335,6 +335,7 @@ Numeric
- :class:`DataFrame` flex inequality comparisons methods (:meth:`DataFrame.lt`, :meth:`DataFrame.le`, :meth:`DataFrame.gt`, :meth: `DataFrame.ge`) with object-dtype and ``complex`` entries failing to raise ``TypeError`` like their :class:`Series` counterparts (:issue:`28079`)
- Bug in :class:`DataFrame` logical operations (`&`, `|`, `^`) not matching :class:`Series` behavior by filling NA values (:issue:`28741`)
- Bug in :meth:`DataFrame.interpolate` where specifying axis by name references variable before it is assigned (:issue:`29142`)
+- Bug in :meth:`Series.var` not computing the right value with a nullable integer dtype series not passing through ddof argument (:issue:`29128`)
- Improved error message when using `frac` > 1 and `replace` = False (:issue:`27451`)
-
diff --git a/pandas/core/arrays/integer.py b/pandas/core/arrays/integer.py
index 08b53e54b91ef..86e19508f2adc 100644
--- a/pandas/core/arrays/integer.py
+++ b/pandas/core/arrays/integer.py
@@ -652,7 +652,7 @@ def _reduce(self, name, skipna=True, **kwargs):
data[mask] = self._na_value
op = getattr(nanops, "nan" + name)
- result = op(data, axis=0, skipna=skipna, mask=mask)
+ result = op(data, axis=0, skipna=skipna, mask=mask, **kwargs)
# if we have a boolean op, don't coerce
if name in ["any", "all"]:
diff --git a/pandas/tests/arrays/test_integer.py b/pandas/tests/arrays/test_integer.py
index 793de66767cc3..025366e5b210b 100644
--- a/pandas/tests/arrays/test_integer.py
+++ b/pandas/tests/arrays/test_integer.py
@@ -829,6 +829,26 @@ def test_arrow_array(data):
assert arr.equals(expected)
+@pytest.mark.parametrize(
+ "pandasmethname, kwargs",
+ [
+ ("var", {"ddof": 0}),
+ ("var", {"ddof": 1}),
+ ("kurtosis", {}),
+ ("skew", {}),
+ ("sem", {}),
+ ],
+)
+def test_stat_method(pandasmethname, kwargs):
+ s = pd.Series(data=[1, 2, 3, 4, 5, 6, np.nan, np.nan], dtype="Int64")
+ pandasmeth = getattr(s, pandasmethname)
+ result = pandasmeth(**kwargs)
+ s2 = pd.Series(data=[1, 2, 3, 4, 5, 6], dtype="Int64")
+ pandasmeth = getattr(s2, pandasmethname)
+ expected = pandasmeth(**kwargs)
+ assert expected == result
+
+
# TODO(jreback) - these need testing / are broken
# shift
| - [ x ] closes #29128
- [ x ] tests added / passed
- [ x ] passes `black pandas`
- [ x ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ x ] whatsnew entry
This PR closes https://github.com/pandas-dev/pandas/issues/29128 | https://api.github.com/repos/pandas-dev/pandas/pulls/29353 | 2019-11-02T14:48:42Z | 2019-11-19T04:23:55Z | 2019-11-19T04:23:55Z | 2019-11-19T04:24:04Z |
TST: Add test to check category dtype remains unchanged after concat. | diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index b537200dd7664..46dafbc4e1ec8 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -2747,6 +2747,22 @@ def test_concat_categorical_tz():
tm.assert_series_equal(result, expected)
+def test_concat_categorical_unchanged():
+ # GH-12007
+ # test fix for when concat on categorical and float
+ # coerces dtype categorical -> float
+ df = pd.DataFrame(pd.Series(["a", "b", "c"], dtype="category", name="A"))
+ ser = pd.Series([0, 1, 2], index=[0, 1, 3], name="B")
+ result = pd.concat([df, ser], axis=1)
+ expected = pd.DataFrame(
+ {
+ "A": pd.Series(["a", "b", "c", np.nan], dtype="category"),
+ "B": pd.Series([0, 1, np.nan, 2], dtype="float"),
+ }
+ )
+ tm.assert_equal(result, expected)
+
+
def test_concat_datetimeindex_freq():
# GH 3232
# Monotonic index result
| - closes #12007
- 1 tests added / passed | https://api.github.com/repos/pandas-dev/pandas/pulls/29352 | 2019-11-02T14:22:46Z | 2019-11-12T23:07:55Z | 2019-11-12T23:07:55Z | 2019-11-12T23:07:59Z |
pin black, xref gh-29341 | diff --git a/environment.yml b/environment.yml
index 163bd08b93c9e..4c96ab815dc90 100644
--- a/environment.yml
+++ b/environment.yml
@@ -16,7 +16,7 @@ dependencies:
- cython>=0.29.13
# code checks
- - black
+ - black<=19.3b0
- cpplint
- flake8
- flake8-comprehensions # used by flake8, linting of unnecessary comprehensions
diff --git a/requirements-dev.txt b/requirements-dev.txt
index 8a9974d393297..7a378cd2f2697 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -3,7 +3,7 @@ python-dateutil>=2.6.1
pytz
asv
cython>=0.29.13
-black
+black<=19.3b0
cpplint
flake8
flake8-comprehensions
| xref #29341
pinning until we can update the codebase | https://api.github.com/repos/pandas-dev/pandas/pulls/29351 | 2019-11-02T13:57:20Z | 2019-11-02T15:02:06Z | 2019-11-02T15:02:06Z | 2019-11-04T16:28:05Z |
Stable python 3.8.0 | diff --git a/.travis.yml b/.travis.yml
index b9fa06304d387..398dd07089ef9 100644
--- a/.travis.yml
+++ b/.travis.yml
@@ -32,7 +32,7 @@ matrix:
include:
- dist: bionic
# 18.04
- python: 3.8-dev
+ python: 3.8.0
env:
- JOB="3.8-dev" PATTERN="(not slow and not network)"
| Python 3.8.0 stable release is now available on travis https://travis-ci.community/t/add-python-3-8-support/5463 we can use it?
The 3.8-dev snapshot seemed to cause some issues here:
https://travis-ci.org/pandas-dev/pandas/jobs/606411398?utm_medium=notification&utm_source=github_status
related - https://github.com/pandas-dev/pandas/issues/26626 | https://api.github.com/repos/pandas-dev/pandas/pulls/29350 | 2019-11-02T13:40:57Z | 2019-11-02T15:51:27Z | 2019-11-02T15:51:27Z | 2019-11-02T16:09:16Z |
GH14422: BUG: to_numeric doesn't work uint64 numbers | diff --git a/pandas/tests/tools/test_numeric.py b/pandas/tests/tools/test_numeric.py
index 55f83e492e2cc..082277796e602 100644
--- a/pandas/tests/tools/test_numeric.py
+++ b/pandas/tests/tools/test_numeric.py
@@ -567,6 +567,24 @@ def test_downcast_limits(dtype, downcast, min_max):
assert series.dtype == dtype
+@pytest.mark.parametrize(
+ "ser,expected",
+ [
+ (
+ pd.Series([0, 9223372036854775808]),
+ pd.Series([0, 9223372036854775808], dtype=np.uint64),
+ )
+ ],
+)
+def test_downcast_uint64(ser, expected):
+ # see gh-14422:
+ # BUG: to_numeric doesn't work uint64 numbers
+
+ result = pd.to_numeric(ser, downcast="unsigned")
+
+ tm.assert_series_equal(result, expected)
+
+
@pytest.mark.parametrize(
"data,exp_data",
[
| -closes #14422
-passes `black pandas`
-added test test_downcast_uint64_exception() | https://api.github.com/repos/pandas-dev/pandas/pulls/29348 | 2019-11-02T13:07:14Z | 2019-11-05T17:19:53Z | 2019-11-05T17:19:53Z | 2019-11-05T17:19:56Z |
Update contributing.rst | diff --git a/doc/source/development/contributing.rst b/doc/source/development/contributing.rst
index 1f77c19f02301..56fac1cb6852a 100644
--- a/doc/source/development/contributing.rst
+++ b/doc/source/development/contributing.rst
@@ -482,7 +482,7 @@ reducing the turn-around time for checking your changes.
python make.py --no-api
# compile the docs with only a single section, relative to the "source" folder.
- # For example, compiling only this guide (docs/source/development/contributing.rst)
+ # For example, compiling only this guide (doc/source/development/contributing.rst)
python make.py clean
python make.py --single development/contributing.rst
| Correct Docs path
| https://api.github.com/repos/pandas-dev/pandas/pulls/29347 | 2019-11-02T13:06:48Z | 2019-11-02T14:15:51Z | 2019-11-02T14:15:51Z | 2019-11-02T14:15:55Z |
TST: Adding styler applymap multindex & code test [Ref: #25858] | diff --git a/pandas/tests/io/formats/test_style.py b/pandas/tests/io/formats/test_style.py
index 0f1402d7da389..0e88f5433c33b 100644
--- a/pandas/tests/io/formats/test_style.py
+++ b/pandas/tests/io/formats/test_style.py
@@ -376,6 +376,25 @@ def color_negative_red(val):
(df.style.applymap(color_negative_red, subset=idx[:, idx["b", "d"]]).render())
+ def test_applymap_subset_multiindex_code(self):
+ # https://github.com/pandas-dev/pandas/issues/25858
+ # Checks styler.applymap works with multindex when codes are provided
+ codes = np.array([[0, 0, 1, 1], [0, 1, 0, 1]])
+ columns = pd.MultiIndex(
+ levels=[["a", "b"], ["%", "#"]], codes=codes, names=["", ""]
+ )
+ df = DataFrame(
+ [[1, -1, 1, 1], [-1, 1, 1, 1]], index=["hello", "world"], columns=columns
+ )
+ pct_subset = pd.IndexSlice[:, pd.IndexSlice[:, "%":"%"]]
+
+ def color_negative_red(val):
+ color = "red" if val < 0 else "black"
+ return "color: %s" % color
+
+ df.loc[pct_subset]
+ df.style.applymap(color_negative_red, subset=pct_subset)
+
def test_where_with_one_style(self):
# GH 17474
def f(x):
| - [x] closes #25858
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29346 | 2019-11-02T12:44:13Z | 2019-11-05T20:57:40Z | 2019-11-05T20:57:39Z | 2019-11-05T20:57:49Z |
ensure consistency between columns aggregates with missing values present | diff --git a/pandas/tests/frame/test_apply.py b/pandas/tests/frame/test_apply.py
index 16d17b04423b7..a1172610b847e 100644
--- a/pandas/tests/frame/test_apply.py
+++ b/pandas/tests/frame/test_apply.py
@@ -1359,3 +1359,14 @@ def test_apply_datetime_tz_issue(self):
expected = pd.Series(index=timestamps, data=timestamps)
tm.assert_series_equal(result, expected)
+
+ @pytest.mark.parametrize("df", [pd.DataFrame({"A": ["a", None], "B": ["c", "d"]})])
+ @pytest.mark.parametrize("method", ["min", "max", "sum"])
+ def test_consistency_of_aggregates_of_columns_with_missing_values(self, df, method):
+ # GH 16832
+ none_in_first_column_result = getattr(df[["A", "B"]], method)()
+ none_in_second_column_result = getattr(df[["B", "A"]], method)()
+
+ tm.assert_series_equal(
+ none_in_first_column_result, none_in_second_column_result
+ )
| - [x] closes #16832
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29345 | 2019-11-02T12:35:00Z | 2019-11-02T14:09:40Z | 2019-11-02T14:09:40Z | 2019-11-02T14:09:44Z |
TST: Test for Boolean Series with missing to Categorical dtype | diff --git a/pandas/tests/series/test_dtypes.py b/pandas/tests/series/test_dtypes.py
index 6ee120f3bec64..8f628d045a7f4 100644
--- a/pandas/tests/series/test_dtypes.py
+++ b/pandas/tests/series/test_dtypes.py
@@ -377,6 +377,15 @@ def test_astype_categorical_to_categorical(
result = s.astype("category")
tm.assert_series_equal(result, expected)
+ def test_astype_bool_missing_to_categorical(self):
+ # GH-19182
+ s = Series([True, False, np.nan])
+ assert s.dtypes == np.object_
+
+ result = s.astype(CategoricalDtype(categories=[True, False]))
+ expected = Series(Categorical([True, False, np.nan], categories=[True, False]))
+ tm.assert_series_equal(result, expected)
+
def test_astype_categoricaldtype(self):
s = Series(["a", "b", "a"])
result = s.astype(CategoricalDtype(["a", "b"], ordered=True))
| - [x] closes #19182
- [x] 1 test added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29344 | 2019-11-02T12:30:06Z | 2019-11-02T20:12:28Z | 2019-11-02T20:12:28Z | 2019-11-04T01:00:44Z |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.