title stringlengths 1 185 | diff stringlengths 0 32.2M | body stringlengths 0 123k ⌀ | url stringlengths 57 58 | created_at stringlengths 20 20 | closed_at stringlengths 20 20 | merged_at stringlengths 20 20 ⌀ | updated_at stringlengths 20 20 |
|---|---|---|---|---|---|---|---|
CLN: type up core.groupby.grouper.get_grouper | diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 31d6e2206f569..e73be29d5b104 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -379,9 +379,9 @@ def __init__(
self.mutated = kwargs.pop("mutated", False)
if grouper is None:
- from pandas.core.groupby.grouper import _get_grouper
+ from pandas.core.groupby.grouper import get_grouper
- grouper, exclusions, obj = _get_grouper(
+ grouper, exclusions, obj = get_grouper(
obj,
keys,
axis=axis,
@@ -1802,9 +1802,9 @@ def nth(self, n: Union[int, List[int]], dropna: Optional[str] = None) -> DataFra
# create a grouper with the original parameters, but on dropped
# object
- from pandas.core.groupby.grouper import _get_grouper
+ from pandas.core.groupby.grouper import get_grouper
- grouper, _, _ = _get_grouper(
+ grouper, _, _ = get_grouper(
dropped,
key=self.keys,
axis=self.axis,
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index ff3b4b1096ecb..370abe75e1327 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -3,7 +3,7 @@
split-apply-combine paradigm.
"""
-from typing import Optional, Tuple
+from typing import Hashable, List, Optional, Tuple
import warnings
import numpy as np
@@ -26,7 +26,6 @@
from pandas.core.arrays import Categorical, ExtensionArray
import pandas.core.common as com
from pandas.core.frame import DataFrame
-from pandas.core.generic import NDFrame
from pandas.core.groupby.categorical import recode_for_groupby, recode_from_groupby
from pandas.core.groupby.ops import BaseGrouper
from pandas.core.index import CategoricalIndex, Index, MultiIndex
@@ -134,7 +133,7 @@ def _get_grouper(self, obj, validate=True):
"""
self._set_grouper(obj)
- self.grouper, exclusions, self.obj = _get_grouper(
+ self.grouper, exclusions, self.obj = get_grouper(
self.obj,
[self.key],
axis=self.axis,
@@ -429,8 +428,8 @@ def groups(self) -> dict:
return self.index.groupby(Categorical.from_codes(self.codes, self.group_index))
-def _get_grouper(
- obj: NDFrame,
+def get_grouper(
+ obj: FrameOrSeries,
key=None,
axis: int = 0,
level=None,
@@ -438,9 +437,9 @@ def _get_grouper(
observed=False,
mutated=False,
validate=True,
-):
+) -> Tuple[BaseGrouper, List[Hashable], FrameOrSeries]:
"""
- create and return a BaseGrouper, which is an internal
+ Create and return a BaseGrouper, which is an internal
mapping of how to create the grouper indexers.
This may be composed of multiple Grouping objects, indicating
multiple groupers
@@ -456,9 +455,9 @@ def _get_grouper(
a BaseGrouper.
If observed & we have a categorical grouper, only show the observed
- values
+ values.
- If validate, then check for key/level overlaps
+ If validate, then check for key/level overlaps.
"""
group_axis = obj._get_axis(axis)
@@ -517,7 +516,7 @@ def _get_grouper(
if key.key is None:
return grouper, [], obj
else:
- return grouper, {key.key}, obj
+ return grouper, [key.key], obj
# already have a BaseGrouper, just return it
elif isinstance(key, BaseGrouper):
@@ -530,10 +529,8 @@ def _get_grouper(
# unhashable elements of `key`. Any unhashable elements implies that
# they wanted a list of keys.
# https://github.com/pandas-dev/pandas/issues/18314
- is_tuple = isinstance(key, tuple)
- all_hashable = is_tuple and is_hashable(key)
-
- if is_tuple:
+ if isinstance(key, tuple):
+ all_hashable = is_hashable(key)
if (
all_hashable and key not in obj and set(key).issubset(obj)
) or not all_hashable:
@@ -573,7 +570,8 @@ def _get_grouper(
all_in_columns_index = all(
g in obj.columns or g in obj.index.names for g in keys
)
- elif isinstance(obj, Series):
+ else:
+ assert isinstance(obj, Series)
all_in_columns_index = all(g in obj.index.names for g in keys)
if not all_in_columns_index:
@@ -586,8 +584,8 @@ def _get_grouper(
else:
levels = [level] * len(keys)
- groupings = []
- exclusions = []
+ groupings = [] # type: List[Grouping]
+ exclusions = [] # type: List[Hashable]
# if the actual grouper should be obj[key]
def is_in_axis(key) -> bool:
| Add types + make some minor cleanups. | https://api.github.com/repos/pandas-dev/pandas/pulls/29458 | 2019-11-07T07:33:28Z | 2019-11-08T08:54:52Z | 2019-11-08T08:54:52Z | 2019-11-08T14:58:58Z |
DEPR: is_extension_type | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 370e1c09d33aa..8c2b140cc2311 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -213,7 +213,8 @@ Deprecations
- ``Index.set_value`` has been deprecated. For a given index ``idx``, array ``arr``,
value in ``idx`` of ``idx_val`` and a new value of ``val``, ``idx.set_value(arr, idx_val, val)``
is equivalent to ``arr[idx.get_loc(idx_val)] = val``, which should be used instead (:issue:`28621`).
--
+- :func:`is_extension_type` is deprecated, :func:`is_extension_array_dtype` should be used instead (:issue:`29457`)
+
.. _whatsnew_1000.prior_deprecations:
diff --git a/pandas/core/apply.py b/pandas/core/apply.py
index f402154dc91ca..e7b088658ac5d 100644
--- a/pandas/core/apply.py
+++ b/pandas/core/apply.py
@@ -7,7 +7,7 @@
from pandas.core.dtypes.common import (
is_dict_like,
- is_extension_type,
+ is_extension_array_dtype,
is_list_like,
is_sequence,
)
@@ -230,7 +230,7 @@ def apply_standard(self):
# as demonstrated in gh-12244
if (
self.result_type in ["reduce", None]
- and not self.dtypes.apply(is_extension_type).any()
+ and not self.dtypes.apply(is_extension_array_dtype).any()
# Disallow complex_internals since libreduction shortcut
# cannot handle MultiIndex
and not self.agg_axis._has_complex_internals
diff --git a/pandas/core/arrays/datetimes.py b/pandas/core/arrays/datetimes.py
index 788cd2a3ce5b7..7cd103d12fa8a 100644
--- a/pandas/core/arrays/datetimes.py
+++ b/pandas/core/arrays/datetimes.py
@@ -31,7 +31,7 @@
is_datetime64_ns_dtype,
is_datetime64tz_dtype,
is_dtype_equal,
- is_extension_type,
+ is_extension_array_dtype,
is_float_dtype,
is_object_dtype,
is_period_dtype,
@@ -2131,7 +2131,7 @@ def maybe_convert_dtype(data, copy):
data = data.categories.take(data.codes, fill_value=NaT)._values
copy = False
- elif is_extension_type(data) and not is_datetime64tz_dtype(data):
+ elif is_extension_array_dtype(data) and not is_datetime64tz_dtype(data):
# Includes categorical
# TODO: We have no tests for these
data = np.array(data, dtype=np.object_)
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 61dc5f35cadf7..a1985f4afc754 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -23,7 +23,6 @@
is_datetime64tz_dtype,
is_datetimelike,
is_extension_array_dtype,
- is_extension_type,
is_list_like,
is_object_dtype,
is_scalar,
@@ -1268,7 +1267,7 @@ def _map_values(self, mapper, na_action=None):
# use the built in categorical series mapper which saves
# time by mapping the categories instead of all values
return self._values.map(mapper)
- if is_extension_type(self.dtype):
+ if is_extension_array_dtype(self.dtype):
values = self._values
else:
values = self.values
@@ -1279,7 +1278,8 @@ def _map_values(self, mapper, na_action=None):
return new_values
# we must convert to python types
- if is_extension_type(self.dtype):
+ if is_extension_array_dtype(self.dtype) and hasattr(self._values, "map"):
+ # GH#23179 some EAs do not have `map`
values = self._values
if na_action is not None:
raise NotImplementedError
diff --git a/pandas/core/construction.py b/pandas/core/construction.py
index 5e8b28267f24f..c0b08beead0ca 100644
--- a/pandas/core/construction.py
+++ b/pandas/core/construction.py
@@ -27,7 +27,6 @@
is_categorical_dtype,
is_datetime64_ns_dtype,
is_extension_array_dtype,
- is_extension_type,
is_float_dtype,
is_integer_dtype,
is_iterator,
@@ -527,7 +526,7 @@ def _try_cast(
and not (is_iterator(subarr) or isinstance(subarr, np.ndarray))
):
subarr = construct_1d_object_array_from_listlike(subarr)
- elif not is_extension_type(subarr):
+ elif not is_extension_array_dtype(subarr):
subarr = construct_1d_ndarray_preserving_na(subarr, dtype, copy=copy)
except OutOfBoundsDatetime:
# in case of out of bound datetime64 -> always raise
diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index fad80d6bf5745..98874fce288bc 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -30,7 +30,6 @@
is_datetimelike,
is_dtype_equal,
is_extension_array_dtype,
- is_extension_type,
is_float,
is_float_dtype,
is_integer,
@@ -633,7 +632,7 @@ def infer_dtype_from_array(arr, pandas_dtype: bool = False):
if not is_list_like(arr):
arr = [arr]
- if pandas_dtype and is_extension_type(arr):
+ if pandas_dtype and is_extension_array_dtype(arr):
return arr.dtype, arr
elif isinstance(arr, ABCSeries):
@@ -695,7 +694,7 @@ def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
# We allow arbitrary fill values for object dtype
raise ValueError("fill_value must be a scalar")
- if is_extension_type(values):
+ if is_extension_array_dtype(values):
if copy:
values = values.copy()
else:
diff --git a/pandas/core/dtypes/common.py b/pandas/core/dtypes/common.py
index 2a46d335ff512..41cbc731e18c4 100644
--- a/pandas/core/dtypes/common.py
+++ b/pandas/core/dtypes/common.py
@@ -1674,6 +1674,8 @@ def is_extension_type(arr):
"""
Check whether an array-like is of a pandas extension class instance.
+ .. deprecated:: 1.0.0
+
Extension classes include categoricals, pandas sparse objects (i.e.
classes represented within the pandas library and not ones external
to it like scipy sparse matrices), and datetime-like arrays.
@@ -1716,6 +1718,12 @@ def is_extension_type(arr):
>>> is_extension_type(s)
True
"""
+ warnings.warn(
+ "'is_extension_type' is deprecated and will be removed in a future "
+ "version. Use 'is_extension_array_dtype' instead.",
+ FutureWarning,
+ stacklevel=2,
+ )
if is_categorical(arr):
return True
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 40efc4c65476a..b005b70eedc7e 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -71,7 +71,6 @@
is_dict_like,
is_dtype_equal,
is_extension_array_dtype,
- is_extension_type,
is_float_dtype,
is_hashable,
is_integer,
@@ -3690,7 +3689,7 @@ def reindexer(value):
value = maybe_cast_to_datetime(value, infer_dtype)
# return internal types directly
- if is_extension_type(value) or is_extension_array_dtype(value):
+ if is_extension_array_dtype(value):
return value
# broadcast across multiple columns if necessary
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 448d2faf8b85f..ce889ea95f782 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -37,7 +37,6 @@
is_datetime64tz_dtype,
is_dtype_equal,
is_extension_array_dtype,
- is_extension_type,
is_float_dtype,
is_integer,
is_integer_dtype,
@@ -2605,10 +2604,6 @@ def should_store(self, value):
value.dtype.type,
(np.integer, np.floating, np.complexfloating, np.datetime64, np.bool_),
)
- or
- # TODO(ExtensionArray): remove is_extension_type
- # when all extension arrays have been ported.
- is_extension_type(value)
or is_extension_array_dtype(value)
)
@@ -3168,7 +3163,7 @@ def _putmask_preserve(nv, n):
# change the dtype if needed
dtype, _ = maybe_promote(n.dtype)
- if is_extension_type(v.dtype) and is_object_dtype(dtype):
+ if is_extension_array_dtype(v.dtype) and is_object_dtype(dtype):
v = v._internal_get_values(dtype)
else:
v = v.astype(dtype)
diff --git a/pandas/core/internals/managers.py b/pandas/core/internals/managers.py
index 21ae820cfcee6..d32e026351e22 100644
--- a/pandas/core/internals/managers.py
+++ b/pandas/core/internals/managers.py
@@ -20,7 +20,6 @@
_NS_DTYPE,
is_datetimelike_v_numeric,
is_extension_array_dtype,
- is_extension_type,
is_list_like,
is_numeric_v_string_like,
is_scalar,
@@ -1034,11 +1033,7 @@ def set(self, item, value):
# FIXME: refactor, clearly separate broadcasting & zip-like assignment
# can prob also fix the various if tests for sparse/categorical
- # TODO(EA): Remove an is_extension_ when all extension types satisfy
- # the interface
- value_is_extension_type = is_extension_type(value) or is_extension_array_dtype(
- value
- )
+ value_is_extension_type = is_extension_array_dtype(value)
# categorical/sparse/datetimetz
if value_is_extension_type:
diff --git a/pandas/core/reshape/melt.py b/pandas/core/reshape/melt.py
index 98fee491e0a73..9ccd36871050f 100644
--- a/pandas/core/reshape/melt.py
+++ b/pandas/core/reshape/melt.py
@@ -4,7 +4,7 @@
from pandas.util._decorators import Appender
-from pandas.core.dtypes.common import is_extension_type, is_list_like
+from pandas.core.dtypes.common import is_extension_array_dtype, is_list_like
from pandas.core.dtypes.concat import concat_compat
from pandas.core.dtypes.generic import ABCMultiIndex
from pandas.core.dtypes.missing import notna
@@ -103,7 +103,7 @@ def melt(
mdata = {}
for col in id_vars:
id_data = frame.pop(col)
- if is_extension_type(id_data):
+ if is_extension_array_dtype(id_data):
id_data = concat([id_data] * K, ignore_index=True)
else:
id_data = np.tile(id_data.values, K)
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 73a05b4cdfa66..ffaecfde6e10f 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -28,7 +28,6 @@
is_datetimelike,
is_dict_like,
is_extension_array_dtype,
- is_extension_type,
is_integer,
is_iterator,
is_list_like,
@@ -3958,7 +3957,8 @@ def f(x):
return f(self)
# row-wise access
- if is_extension_type(self.dtype):
+ if is_extension_array_dtype(self.dtype) and hasattr(self._values, "map"):
+ # GH#23179 some EAs do not have `map`
mapped = self._values.map(f)
else:
values = self.astype(object).values
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index 35e6d53127e59..77b2db20ac2a9 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -26,7 +26,7 @@
is_categorical_dtype,
is_datetime64_dtype,
is_datetime64tz_dtype,
- is_extension_type,
+ is_extension_array_dtype,
is_list_like,
is_timedelta64_dtype,
)
@@ -2827,7 +2827,7 @@ def write_multi_index(self, key, index):
zip(index.levels, index.codes, index.names)
):
# write the level
- if is_extension_type(lev):
+ if is_extension_array_dtype(lev):
raise NotImplementedError(
"Saving a MultiIndex with an extension dtype is not supported."
)
diff --git a/pandas/tests/api/test_types.py b/pandas/tests/api/test_types.py
index 24f325643479c..e9f68692a9863 100644
--- a/pandas/tests/api/test_types.py
+++ b/pandas/tests/api/test_types.py
@@ -18,7 +18,6 @@ class TestTypes(Base):
"is_datetime64_ns_dtype",
"is_datetime64tz_dtype",
"is_dtype_equal",
- "is_extension_type",
"is_float",
"is_float_dtype",
"is_int64_dtype",
@@ -51,7 +50,7 @@ class TestTypes(Base):
"infer_dtype",
"is_extension_array_dtype",
]
- deprecated = ["is_period", "is_datetimetz"]
+ deprecated = ["is_period", "is_datetimetz", "is_extension_type"]
dtypes = ["CategoricalDtype", "DatetimeTZDtype", "PeriodDtype", "IntervalDtype"]
def test_types(self):
| It is mostly redundant with `is_extension_array_dtype`, and having both is confusing.
xref #23179. | https://api.github.com/repos/pandas-dev/pandas/pulls/29457 | 2019-11-07T05:24:40Z | 2019-11-08T14:37:45Z | 2019-11-08T14:37:45Z | 2023-12-11T23:50:52Z |
CLN: type annotations in groupby.grouper, groupby.ops | diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index 370abe75e1327..e6e3ee62459ca 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -119,7 +119,7 @@ def __init__(self, key=None, level=None, freq=None, axis=0, sort=False):
def ax(self):
return self.grouper
- def _get_grouper(self, obj, validate=True):
+ def _get_grouper(self, obj, validate: bool = True):
"""
Parameters
----------
@@ -143,17 +143,18 @@ def _get_grouper(self, obj, validate=True):
)
return self.binner, self.grouper, self.obj
- def _set_grouper(self, obj, sort=False):
+ def _set_grouper(self, obj: FrameOrSeries, sort: bool = False):
"""
given an object and the specifications, setup the internal grouper
for this particular specification
Parameters
----------
- obj : the subject object
+ obj : Series or DataFrame
sort : bool, default False
whether the resulting grouper should be sorted
"""
+ assert obj is not None
if self.key is not None and self.level is not None:
raise ValueError("The Grouper cannot specify both a key and a level!")
@@ -211,13 +212,13 @@ def groups(self):
def __repr__(self) -> str:
attrs_list = (
- "{}={!r}".format(attr_name, getattr(self, attr_name))
+ "{name}={val!r}".format(name=attr_name, val=getattr(self, attr_name))
for attr_name in self._attributes
if getattr(self, attr_name) is not None
)
attrs = ", ".join(attrs_list)
cls_name = self.__class__.__name__
- return "{}({})".format(cls_name, attrs)
+ return "{cls}({attrs})".format(cls=cls_name, attrs=attrs)
class Grouping:
@@ -372,7 +373,7 @@ def __init__(
self.grouper = self.grouper.astype("timedelta64[ns]")
def __repr__(self) -> str:
- return "Grouping({0})".format(self.name)
+ return "Grouping({name})".format(name=self.name)
def __iter__(self):
return iter(self.indices)
@@ -433,10 +434,10 @@ def get_grouper(
key=None,
axis: int = 0,
level=None,
- sort=True,
- observed=False,
- mutated=False,
- validate=True,
+ sort: bool = True,
+ observed: bool = False,
+ mutated: bool = False,
+ validate: bool = True,
) -> Tuple[BaseGrouper, List[Hashable], FrameOrSeries]:
"""
Create and return a BaseGrouper, which is an internal
@@ -670,7 +671,7 @@ def is_in_obj(gpr) -> bool:
return grouper, exclusions, obj
-def _is_label_like(val):
+def _is_label_like(val) -> bool:
return isinstance(val, (str, tuple)) or (val is not None and is_scalar(val))
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 6796239cf3fd9..e6cf46de5c350 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -36,6 +36,7 @@
)
from pandas.core.dtypes.missing import _maybe_fill, isna
+from pandas._typing import FrameOrSeries
import pandas.core.algorithms as algorithms
from pandas.core.base import SelectionMixin
import pandas.core.common as com
@@ -89,12 +90,16 @@ def __init__(
self._filter_empty_groups = self.compressed = len(groupings) != 1
self.axis = axis
- self.groupings = groupings # type: Sequence[grouper.Grouping]
+ self._groupings = list(groupings) # type: List[grouper.Grouping]
self.sort = sort
self.group_keys = group_keys
self.mutated = mutated
self.indexer = indexer
+ @property
+ def groupings(self) -> List["grouper.Grouping"]:
+ return self._groupings
+
@property
def shape(self):
return tuple(ping.ngroups for ping in self.groupings)
@@ -106,7 +111,7 @@ def __iter__(self):
def nkeys(self) -> int:
return len(self.groupings)
- def get_iterator(self, data, axis=0):
+ def get_iterator(self, data: FrameOrSeries, axis: int = 0):
"""
Groupby iterator
@@ -120,7 +125,7 @@ def get_iterator(self, data, axis=0):
for key, (i, group) in zip(keys, splitter):
yield key, group
- def _get_splitter(self, data, axis=0):
+ def _get_splitter(self, data: FrameOrSeries, axis: int = 0) -> "DataSplitter":
comp_ids, _, ngroups = self.group_info
return get_splitter(data, comp_ids, ngroups, axis=axis)
@@ -142,13 +147,13 @@ def _get_group_keys(self):
# provide "flattened" iterator for multi-group setting
return get_flattened_iterator(comp_ids, ngroups, self.levels, self.codes)
- def apply(self, f, data, axis: int = 0):
+ def apply(self, f, data: FrameOrSeries, axis: int = 0):
mutated = self.mutated
splitter = self._get_splitter(data, axis=axis)
group_keys = self._get_group_keys()
result_values = None
- sdata = splitter._get_sorted_data()
+ sdata = splitter._get_sorted_data() # type: FrameOrSeries
if sdata.ndim == 2 and np.any(sdata.dtypes.apply(is_extension_array_dtype)):
# calling splitter.fast_apply will raise TypeError via apply_frame_axis0
# if we pass EA instead of ndarray
@@ -157,7 +162,7 @@ def apply(self, f, data, axis: int = 0):
elif (
com.get_callable_name(f) not in base.plotting_methods
- and hasattr(splitter, "fast_apply")
+ and isinstance(splitter, FrameSplitter)
and axis == 0
# with MultiIndex, apply_frame_axis0 would raise InvalidApply
# TODO: can we make this check prettier?
@@ -229,8 +234,7 @@ def names(self):
def size(self) -> Series:
"""
- Compute group sizes
-
+ Compute group sizes.
"""
ids, _, ngroup = self.group_info
ids = ensure_platform_int(ids)
@@ -292,7 +296,7 @@ def reconstructed_codes(self) -> List[np.ndarray]:
return decons_obs_group_ids(comp_ids, obs_ids, self.shape, codes, xnull=True)
@cache_readonly
- def result_index(self):
+ def result_index(self) -> Index:
if not self.compressed and len(self.groupings) == 1:
return self.groupings[0].result_index.rename(self.names[0])
@@ -628,7 +632,7 @@ def agg_series(self, obj: Series, func):
raise
return self._aggregate_series_pure_python(obj, func)
- def _aggregate_series_fast(self, obj, func):
+ def _aggregate_series_fast(self, obj: Series, func):
# At this point we have already checked that
# - obj.index is not a MultiIndex
# - obj is backed by an ndarray, not ExtensionArray
@@ -646,7 +650,7 @@ def _aggregate_series_fast(self, obj, func):
result, counts = grouper.get_result()
return result, counts
- def _aggregate_series_pure_python(self, obj, func):
+ def _aggregate_series_pure_python(self, obj: Series, func):
group_index, _, ngroups = self.group_info
@@ -703,7 +707,12 @@ class BinGrouper(BaseGrouper):
"""
def __init__(
- self, bins, binlabels, filter_empty=False, mutated=False, indexer=None
+ self,
+ bins,
+ binlabels,
+ filter_empty: bool = False,
+ mutated: bool = False,
+ indexer=None,
):
self.bins = ensure_int64(bins)
self.binlabels = ensure_index(binlabels)
@@ -737,7 +746,7 @@ def _get_grouper(self):
"""
return self
- def get_iterator(self, data: NDFrame, axis: int = 0):
+ def get_iterator(self, data: FrameOrSeries, axis: int = 0):
"""
Groupby iterator
@@ -809,11 +818,9 @@ def names(self):
return [self.binlabels.name]
@property
- def groupings(self):
- from pandas.core.groupby.grouper import Grouping
-
+ def groupings(self) -> "List[grouper.Grouping]":
return [
- Grouping(lvl, lvl, in_axis=False, level=None, name=name)
+ grouper.Grouping(lvl, lvl, in_axis=False, level=None, name=name)
for lvl, name in zip(self.levels, self.names)
]
@@ -854,7 +861,7 @@ def _is_indexed_like(obj, axes) -> bool:
class DataSplitter:
- def __init__(self, data, labels, ngroups, axis: int = 0):
+ def __init__(self, data: FrameOrSeries, labels, ngroups: int, axis: int = 0):
self.data = data
self.labels = ensure_int64(labels)
self.ngroups = ngroups
@@ -885,15 +892,15 @@ def __iter__(self):
for i, (start, end) in enumerate(zip(starts, ends)):
yield i, self._chop(sdata, slice(start, end))
- def _get_sorted_data(self):
+ def _get_sorted_data(self) -> FrameOrSeries:
return self.data.take(self.sort_idx, axis=self.axis)
- def _chop(self, sdata, slice_obj: slice):
+ def _chop(self, sdata, slice_obj: slice) -> NDFrame:
raise AbstractMethodError(self)
class SeriesSplitter(DataSplitter):
- def _chop(self, sdata, slice_obj: slice):
+ def _chop(self, sdata: Series, slice_obj: slice) -> Series:
return sdata._get_values(slice_obj)
@@ -905,14 +912,14 @@ def fast_apply(self, f, names):
sdata = self._get_sorted_data()
return libreduction.apply_frame_axis0(sdata, f, names, starts, ends)
- def _chop(self, sdata, slice_obj: slice):
+ def _chop(self, sdata: DataFrame, slice_obj: slice) -> DataFrame:
if self.axis == 0:
return sdata.iloc[slice_obj]
else:
return sdata._slice(slice_obj, axis=1)
-def get_splitter(data: NDFrame, *args, **kwargs):
+def get_splitter(data: FrameOrSeries, *args, **kwargs) -> DataSplitter:
if isinstance(data, Series):
klass = SeriesSplitter # type: Type[DataSplitter]
else:
| @simonjayhawkins mypy is still giving a couple of complaints I could use your help sorting out:
```
pandas/core/groupby/ops.py:791: error: Signature of "groupings" incompatible with supertype "BaseGrouper"
pandas/core/groupby/ops.py:872: error: Argument 1 of "_chop" is incompatible with supertype "DataSplitter"; supertype defines the argument type as "NDFrame"
pandas/core/groupby/ops.py:884: error: Argument 1 of "_chop" is incompatible with supertype "DataSplitter"; supertype defines the argument type as "NDFrame"
```
For the groupings complaint, AFAICT the attribute has the same annotation, but in the subclass its a property instead of defined in `__init__`. For the other two, I annotated an argument with `NDFrame` in the base class and overrode with `Series` and `DataFrame` in the subclasses. What is the preferred idiom for this pattern? | https://api.github.com/repos/pandas-dev/pandas/pulls/29456 | 2019-11-07T05:18:19Z | 2019-11-13T00:39:19Z | 2019-11-13T00:39:19Z | 2019-11-13T12:12:10Z |
TST: add test for empty frame groupby dtypes consistency | diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 79c9fe2b60bd9..a535fcc511daa 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -1490,6 +1490,20 @@ def test_frame_dict_constructor_empty_series(self):
DataFrame({"foo": s1, "bar": s2, "baz": s3})
DataFrame.from_dict({"foo": s1, "baz": s3, "bar": s2})
+ @pytest.mark.parametrize("d", [4, "d"])
+ def test_empty_frame_groupby_dtypes_consistency(self, d):
+ # GH 20888
+ group_keys = ["a", "b", "c"]
+ df = DataFrame({"a": [1], "b": [2], "c": [3], "d": [d]})
+
+ g = df[df.a == 2].groupby(group_keys)
+ result = g.first().index
+ expected = MultiIndex(
+ levels=[[1], [2], [3]], codes=[[], [], []], names=["a", "b", "c"]
+ )
+
+ tm.assert_index_equal(result, expected)
+
def test_multiindex_na_repr(self):
# only an issue with long columns
df3 = DataFrame(
| - [x] closes #20888
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29455 | 2019-11-07T02:12:32Z | 2019-11-08T04:17:29Z | 2019-11-08T04:17:28Z | 2019-11-08T04:17:35Z |
TST: add test for df.where() with category dtype | diff --git a/pandas/tests/frame/test_dtypes.py b/pandas/tests/frame/test_dtypes.py
index 68844aeeb081e..c29f5e78b033f 100644
--- a/pandas/tests/frame/test_dtypes.py
+++ b/pandas/tests/frame/test_dtypes.py
@@ -815,6 +815,22 @@ def test_astype_extension_dtypes_duplicate_col(self, dtype):
expected = concat([a1.astype(dtype), a2.astype(dtype)], axis=1)
tm.assert_frame_equal(result, expected)
+ @pytest.mark.parametrize("kwargs", [dict(), dict(other=None)])
+ def test_df_where_with_category(self, kwargs):
+ # GH 16979
+ df = DataFrame(np.arange(2 * 3).reshape(2, 3), columns=list("ABC"))
+ mask = np.array([[True, False, True], [False, True, True]])
+
+ # change type to category
+ df.A = df.A.astype("category")
+ df.B = df.B.astype("category")
+ df.C = df.C.astype("category")
+
+ result = df.A.where(mask[:, 0], **kwargs)
+ expected = Series(pd.Categorical([0, np.nan], categories=[0, 3]), name="A")
+
+ tm.assert_series_equal(result, expected)
+
@pytest.mark.parametrize(
"dtype", [{100: "float64", 200: "uint64"}, "category", "float64"]
)
| - [x] xref #16979
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29454 | 2019-11-07T01:45:46Z | 2019-11-08T19:57:37Z | 2019-11-08T19:57:37Z | 2019-11-08T19:57:37Z |
CLN: remove is_datetimelike | diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 23ba0ac1c737e..26099a94834e8 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -29,7 +29,6 @@
is_complex_dtype,
is_datetime64_any_dtype,
is_datetime64_ns_dtype,
- is_datetimelike,
is_extension_array_dtype,
is_float_dtype,
is_integer,
@@ -834,7 +833,7 @@ def mode(values, dropna: bool = True) -> ABCSeries:
return Series(values.values.mode(dropna=dropna), name=values.name)
return values.mode(dropna=dropna)
- if dropna and is_datetimelike(values):
+ if dropna and needs_i8_conversion(values.dtype):
mask = values.isnull()
values = values[~mask]
diff --git a/pandas/core/arrays/categorical.py b/pandas/core/arrays/categorical.py
index ce174baa66a97..73716fdeb42bb 100644
--- a/pandas/core/arrays/categorical.py
+++ b/pandas/core/arrays/categorical.py
@@ -25,7 +25,6 @@
ensure_platform_int,
is_categorical_dtype,
is_datetime64_dtype,
- is_datetimelike,
is_dict_like,
is_dtype_equal,
is_extension_array_dtype,
@@ -37,6 +36,7 @@
is_scalar,
is_sequence,
is_timedelta64_dtype,
+ needs_i8_conversion,
)
from pandas.core.dtypes.dtypes import CategoricalDtype
from pandas.core.dtypes.generic import ABCDataFrame, ABCIndexClass, ABCSeries
@@ -1533,7 +1533,7 @@ def get_values(self):
def _internal_get_values(self):
# if we are a datetime and period index, return Index to keep metadata
- if is_datetimelike(self.categories):
+ if needs_i8_conversion(self.categories):
return self.categories.take(self._codes, fill_value=np.nan)
elif is_integer_dtype(self.categories) and -1 in self._codes:
return self.categories.astype("object").take(self._codes, fill_value=np.nan)
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 61dc5f35cadf7..eeb0b72e301dd 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -21,7 +21,6 @@
is_categorical_dtype,
is_datetime64_ns_dtype,
is_datetime64tz_dtype,
- is_datetimelike,
is_extension_array_dtype,
is_extension_type,
is_list_like,
@@ -1172,7 +1171,7 @@ def tolist(self):
--------
numpy.ndarray.tolist
"""
- if is_datetimelike(self._values):
+ if self.dtype.kind in ["m", "M"]:
return [com.maybe_box_datetimelike(x) for x in self._values]
elif is_extension_array_dtype(self._values):
return list(self._values)
@@ -1194,7 +1193,7 @@ def __iter__(self):
iterator
"""
# We are explicitly making element iterators.
- if is_datetimelike(self._values):
+ if self.dtype.kind in ["m", "M"]:
return map(com.maybe_box_datetimelike, self._values)
elif is_extension_array_dtype(self._values):
return iter(self._values)
diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index fad80d6bf5745..bbed3a545e478 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -27,7 +27,6 @@
is_datetime64_ns_dtype,
is_datetime64tz_dtype,
is_datetime_or_timedelta_dtype,
- is_datetimelike,
is_dtype_equal,
is_extension_array_dtype,
is_extension_type,
@@ -274,7 +273,7 @@ def maybe_upcast_putmask(result: np.ndarray, mask: np.ndarray, other):
# in np.place:
# NaN -> NaT
# integer or integer array -> date-like array
- if is_datetimelike(result.dtype):
+ if result.dtype.kind in ["m", "M"]:
if is_scalar(other):
if isna(other):
other = result.dtype.type("nat")
diff --git a/pandas/core/dtypes/common.py b/pandas/core/dtypes/common.py
index 2a46d335ff512..c3e98d4009135 100644
--- a/pandas/core/dtypes/common.py
+++ b/pandas/core/dtypes/common.py
@@ -799,54 +799,6 @@ def is_datetime_arraylike(arr):
return getattr(arr, "inferred_type", None) == "datetime"
-def is_datetimelike(arr):
- """
- Check whether an array-like is a datetime-like array-like.
-
- Acceptable datetime-like objects are (but not limited to) datetime
- indices, periodic indices, and timedelta indices.
-
- Parameters
- ----------
- arr : array-like
- The array-like to check.
-
- Returns
- -------
- boolean
- Whether or not the array-like is a datetime-like array-like.
-
- Examples
- --------
- >>> is_datetimelike([1, 2, 3])
- False
- >>> is_datetimelike(pd.Index([1, 2, 3]))
- False
- >>> is_datetimelike(pd.DatetimeIndex([1, 2, 3]))
- True
- >>> is_datetimelike(pd.DatetimeIndex([1, 2, 3], tz="US/Eastern"))
- True
- >>> is_datetimelike(pd.PeriodIndex([], freq="A"))
- True
- >>> is_datetimelike(np.array([], dtype=np.datetime64))
- True
- >>> is_datetimelike(pd.Series([], dtype="timedelta64[ns]"))
- True
- >>>
- >>> dtype = DatetimeTZDtype("ns", tz="US/Eastern")
- >>> s = pd.Series([], dtype=dtype)
- >>> is_datetimelike(s)
- True
- """
-
- return (
- is_datetime64_dtype(arr)
- or is_datetime64tz_dtype(arr)
- or is_timedelta64_dtype(arr)
- or isinstance(arr, ABCPeriodIndex)
- )
-
-
def is_dtype_equal(source, target):
"""
Check if two dtypes are equal.
@@ -1446,9 +1398,8 @@ def is_numeric(x):
"""
return is_integer_dtype(x) or is_float_dtype(x)
- is_datetimelike = needs_i8_conversion
- return (is_datetimelike(a) and is_numeric(b)) or (
- is_datetimelike(b) and is_numeric(a)
+ return (needs_i8_conversion(a) and is_numeric(b)) or (
+ needs_i8_conversion(b) and is_numeric(a)
)
diff --git a/pandas/core/dtypes/missing.py b/pandas/core/dtypes/missing.py
index 322011eb8e263..22e38a805f996 100644
--- a/pandas/core/dtypes/missing.py
+++ b/pandas/core/dtypes/missing.py
@@ -17,7 +17,6 @@
is_complex_dtype,
is_datetime64_dtype,
is_datetime64tz_dtype,
- is_datetimelike,
is_datetimelike_v_numeric,
is_dtype_equal,
is_extension_array_dtype,
@@ -494,7 +493,7 @@ def _infer_fill_value(val):
if not is_list_like(val):
val = [val]
val = np.array(val, copy=False)
- if is_datetimelike(val):
+ if needs_i8_conversion(val):
return np.array("NaT", dtype=val.dtype)
elif is_object_dtype(val.dtype):
dtype = lib.infer_dtype(ensure_object(val), skipna=False)
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 511b87dab087e..3b8c3148f5177 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -40,7 +40,6 @@
ensure_int64,
ensure_platform_int,
is_bool,
- is_datetimelike,
is_dict_like,
is_integer_dtype,
is_interval_dtype,
@@ -48,6 +47,7 @@
is_numeric_dtype,
is_object_dtype,
is_scalar,
+ needs_i8_conversion,
)
from pandas.core.dtypes.missing import _isna_ndarraylike, isna, notna
@@ -1287,7 +1287,7 @@ def first_not_none(values):
# if we have date/time like in the original, then coerce dates
# as we are stacking can easily have object dtypes here
so = self._selected_obj
- if so.ndim == 2 and so.dtypes.apply(is_datetimelike).any():
+ if so.ndim == 2 and so.dtypes.apply(needs_i8_conversion).any():
result = _recast_datetimelike_result(result)
else:
result = result._convert(datetime=True)
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index a189b2cd1ab84..30857a51debd1 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -24,7 +24,6 @@
is_bool_dtype,
is_categorical_dtype,
is_datetime64tz_dtype,
- is_datetimelike,
is_dtype_equal,
is_extension_array_dtype,
is_float_dtype,
@@ -1120,9 +1119,9 @@ def _maybe_coerce_merge_keys(self):
raise ValueError(msg)
# datetimelikes must match exactly
- elif is_datetimelike(lk) and not is_datetimelike(rk):
+ elif needs_i8_conversion(lk) and not needs_i8_conversion(rk):
raise ValueError(msg)
- elif not is_datetimelike(lk) and is_datetimelike(rk):
+ elif not needs_i8_conversion(lk) and needs_i8_conversion(rk):
raise ValueError(msg)
elif is_datetime64tz_dtype(lk) and not is_datetime64tz_dtype(rk):
raise ValueError(msg)
@@ -1637,7 +1636,7 @@ def _get_merge_keys(self):
)
)
- if is_datetimelike(lt):
+ if needs_i8_conversion(lt):
if not isinstance(self.tolerance, datetime.timedelta):
raise MergeError(msg)
if self.tolerance < Timedelta(0):
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 73a05b4cdfa66..6440d2f03cf1a 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -25,7 +25,6 @@
is_categorical,
is_categorical_dtype,
is_datetime64_dtype,
- is_datetimelike,
is_dict_like,
is_extension_array_dtype,
is_extension_type,
@@ -2886,7 +2885,7 @@ def combine_first(self, other):
new_index = self.index.union(other.index)
this = self.reindex(new_index, copy=False)
other = other.reindex(new_index, copy=False)
- if is_datetimelike(this) and not is_datetimelike(other):
+ if this.dtype.kind == "M" and other.dtype.kind != "M":
other = to_datetime(other)
return this.where(notna(this), other)
diff --git a/pandas/tests/dtypes/test_common.py b/pandas/tests/dtypes/test_common.py
index 894d6a40280b7..5e409b85049ae 100644
--- a/pandas/tests/dtypes/test_common.py
+++ b/pandas/tests/dtypes/test_common.py
@@ -309,21 +309,6 @@ def test_is_datetime_arraylike():
assert com.is_datetime_arraylike(pd.DatetimeIndex([1, 2, 3]))
-def test_is_datetimelike():
- assert not com.is_datetimelike([1, 2, 3])
- assert not com.is_datetimelike(pd.Index([1, 2, 3]))
-
- assert com.is_datetimelike(pd.DatetimeIndex([1, 2, 3]))
- assert com.is_datetimelike(pd.PeriodIndex([], freq="A"))
- assert com.is_datetimelike(np.array([], dtype=np.datetime64))
- assert com.is_datetimelike(pd.Series([], dtype="timedelta64[ns]"))
- assert com.is_datetimelike(pd.DatetimeIndex(["2000"], tz="US/Eastern"))
-
- dtype = DatetimeTZDtype("ns", tz="US/Eastern")
- s = pd.Series([], dtype=dtype)
- assert com.is_datetimelike(s)
-
-
integer_dtypes = [] # type: List
| - [x] closes #23914
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
AFAICT this is strictly dominated by needs_i8_conversion | https://api.github.com/repos/pandas-dev/pandas/pulls/29452 | 2019-11-07T00:28:50Z | 2019-11-07T21:23:32Z | 2019-11-07T21:23:31Z | 2019-11-07T21:56:05Z |
CLN: remove is_stringlike | diff --git a/pandas/core/dtypes/common.py b/pandas/core/dtypes/common.py
index 3f4ebc88c1c8a..2a46d335ff512 100644
--- a/pandas/core/dtypes/common.py
+++ b/pandas/core/dtypes/common.py
@@ -45,7 +45,6 @@
is_re_compilable,
is_scalar,
is_sequence,
- is_string_like,
)
from pandas._typing import ArrayLike
@@ -1383,8 +1382,8 @@ def is_numeric_v_string_like(a, b):
is_a_string_array = is_a_array and is_string_like_dtype(a)
is_b_string_array = is_b_array and is_string_like_dtype(b)
- is_a_scalar_string_like = not is_a_array and is_string_like(a)
- is_b_scalar_string_like = not is_b_array and is_string_like(b)
+ is_a_scalar_string_like = not is_a_array and isinstance(a, str)
+ is_b_scalar_string_like = not is_b_array and isinstance(b, str)
return (
(is_a_numeric_array and is_b_scalar_string_like)
diff --git a/pandas/core/dtypes/inference.py b/pandas/core/dtypes/inference.py
index e69e703f3a96c..61fa7940c1bce 100644
--- a/pandas/core/dtypes/inference.py
+++ b/pandas/core/dtypes/inference.py
@@ -67,30 +67,6 @@ def is_number(obj):
return isinstance(obj, (Number, np.number))
-def is_string_like(obj):
- """
- Check if the object is a string.
-
- Parameters
- ----------
- obj : The object to check
-
- Examples
- --------
- >>> is_string_like("foo")
- True
- >>> is_string_like(1)
- False
-
- Returns
- -------
- is_str_like : bool
- Whether `obj` is a string or not.
- """
-
- return isinstance(obj, str)
-
-
def _iterable_not_string(obj):
"""
Check if the object is an iterable but not a string.
diff --git a/pandas/core/indexes/datetimes.py b/pandas/core/indexes/datetimes.py
index e5cecd090e061..2d0ecf1b936da 100644
--- a/pandas/core/indexes/datetimes.py
+++ b/pandas/core/indexes/datetimes.py
@@ -16,7 +16,6 @@
is_integer,
is_list_like,
is_scalar,
- is_string_like,
)
from pandas.core.dtypes.concat import concat_compat
from pandas.core.dtypes.dtypes import DatetimeTZDtype
@@ -1659,7 +1658,7 @@ def bdate_range(
msg = "freq must be specified for bdate_range; use date_range instead"
raise TypeError(msg)
- if is_string_like(freq) and freq.startswith("C"):
+ if isinstance(freq, str) and freq.startswith("C"):
try:
weekmask = weekmask or "Mon Tue Wed Thu Fri"
freq = prefix_mapping[freq](holidays=holidays, weekmask=weekmask)
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 7b65816dc06b9..73a05b4cdfa66 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -34,7 +34,6 @@
is_list_like,
is_object_dtype,
is_scalar,
- is_string_like,
is_timedelta64_dtype,
)
from pandas.core.dtypes.generic import (
@@ -4539,7 +4538,7 @@ def to_csv(self, *args, **kwargs):
# passed as second argument (while the first is the same)
maybe_sep = args[1]
- if not (is_string_like(maybe_sep) and len(maybe_sep) == 1):
+ if not (isinstance(maybe_sep, str) and len(maybe_sep) == 1):
# old signature
warnings.warn(
"The signature of `Series.to_csv` was aligned "
diff --git a/pandas/core/strings.py b/pandas/core/strings.py
index f1a67d0892cad..7194d1cf08e4a 100644
--- a/pandas/core/strings.py
+++ b/pandas/core/strings.py
@@ -19,7 +19,6 @@
is_list_like,
is_re,
is_scalar,
- is_string_like,
)
from pandas.core.dtypes.generic import (
ABCDataFrame,
@@ -601,7 +600,7 @@ def str_replace(arr, pat, repl, n=-1, case=None, flags=0, regex=True):
"""
# Check whether repl is valid (GH 13438, GH 15055)
- if not (is_string_like(repl) or callable(repl)):
+ if not (isinstance(repl, str) or callable(repl)):
raise TypeError("repl must be a string or callable")
is_compiled_re = is_re(pat)
diff --git a/pandas/io/formats/style.py b/pandas/io/formats/style.py
index 9865087a26ae3..dce0afd8670b2 100644
--- a/pandas/io/formats/style.py
+++ b/pandas/io/formats/style.py
@@ -18,7 +18,7 @@
from pandas.compat._optional import import_optional_dependency
from pandas.util._decorators import Appender
-from pandas.core.dtypes.common import is_float, is_string_like
+from pandas.core.dtypes.common import is_float
import pandas as pd
from pandas.api.types import is_dict_like, is_list_like
@@ -1488,7 +1488,7 @@ def _get_level_lengths(index, hidden_elements=None):
def _maybe_wrap_formatter(formatter):
- if is_string_like(formatter):
+ if isinstance(formatter, str):
return lambda x: formatter.format(x)
elif callable(formatter):
return formatter
| its now just an alias for `isinstance(obj, str)` | https://api.github.com/repos/pandas-dev/pandas/pulls/29450 | 2019-11-06T22:23:54Z | 2019-11-07T07:34:28Z | 2019-11-07T07:34:28Z | 2019-11-07T17:18:30Z |
DOC: Improving (hopefully) the documintation | diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index 2d6c8e1008ce1..c2d6a3bc4906d 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -84,8 +84,8 @@ cpdef ndarray[int64_t, ndim=1] unique_deltas(const int64_t[:] arr):
Returns
-------
- result : ndarray[int64_t]
- result is sorted
+ ndarray[int64_t]
+ An ordered ndarray[int64_t]
"""
cdef:
Py_ssize_t i, n = len(arr)
@@ -150,9 +150,10 @@ def is_lexsorted(list_of_arrays: list) -> bint:
@cython.wraparound(False)
def groupsort_indexer(const int64_t[:] index, Py_ssize_t ngroups):
"""
- compute a 1-d indexer that is an ordering of the passed index,
- ordered by the groups. This is a reverse of the label
- factorization process.
+ Compute a 1-d indexer.
+
+ The indexer is an ordering of the passed index,
+ ordered by the groups.
Parameters
----------
@@ -161,7 +162,14 @@ def groupsort_indexer(const int64_t[:] index, Py_ssize_t ngroups):
ngroups: int64
number of groups
- return a tuple of (1-d indexer ordered by groups, group counts)
+ Returns
+ -------
+ tuple
+ 1-d indexer ordered by groups, group counts
+
+ Notes
+ -----
+ This is a reverse of the label factorization process.
"""
cdef:
@@ -391,6 +399,7 @@ def _validate_limit(nobs: int, limit=None) -> int:
Returns
-------
int
+ The limit.
"""
if limit is None:
lim = nobs
@@ -669,7 +678,8 @@ def is_monotonic(ndarray[algos_t, ndim=1] arr, bint timelike):
"""
Returns
-------
- is_monotonic_inc, is_monotonic_dec, is_unique
+ tuple
+ is_monotonic_inc, is_monotonic_dec, is_unique
"""
cdef:
Py_ssize_t i, n
| - [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` | https://api.github.com/repos/pandas-dev/pandas/pulls/29449 | 2019-11-06T22:13:18Z | 2019-11-07T16:10:39Z | 2019-11-07T16:10:39Z | 2019-11-07T20:52:15Z |
TST: add test for indexing with single/double tuples | diff --git a/pandas/tests/frame/indexing/test_indexing.py b/pandas/tests/frame/indexing/test_indexing.py
index 24a431fe42cf8..9a7cd4ace686f 100644
--- a/pandas/tests/frame/indexing/test_indexing.py
+++ b/pandas/tests/frame/indexing/test_indexing.py
@@ -2624,6 +2624,17 @@ def test_index_namedtuple(self):
result = df.loc[IndexType("foo", "bar")]["A"]
assert result == 1
+ @pytest.mark.parametrize("tpl", [tuple([1]), tuple([1, 2])])
+ def test_index_single_double_tuples(self, tpl):
+ # GH 20991
+ idx = pd.Index([tuple([1]), tuple([1, 2])], name="A", tupleize_cols=False)
+ df = DataFrame(index=idx)
+
+ result = df.loc[[tpl]]
+ idx = pd.Index([tpl], name="A", tupleize_cols=False)
+ expected = DataFrame(index=idx)
+ tm.assert_frame_equal(result, expected)
+
def test_boolean_indexing(self):
idx = list(range(3))
cols = ["A", "B", "C"]
| - [x] closes #20991
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` | https://api.github.com/repos/pandas-dev/pandas/pulls/29448 | 2019-11-06T21:56:29Z | 2019-11-20T17:14:29Z | 2019-11-20T17:14:29Z | 2019-11-20T17:15:58Z |
ENH: Add ORC reader | diff --git a/doc/source/getting_started/install.rst b/doc/source/getting_started/install.rst
index 9f3ab22496ae7..14530a9010a1c 100644
--- a/doc/source/getting_started/install.rst
+++ b/doc/source/getting_started/install.rst
@@ -258,7 +258,7 @@ matplotlib 2.2.2 Visualization
openpyxl 2.4.8 Reading / writing for xlsx files
pandas-gbq 0.8.0 Google Big Query access
psycopg2 PostgreSQL engine for sqlalchemy
-pyarrow 0.12.0 Parquet and feather reading / writing
+pyarrow 0.12.0 Parquet, ORC (requires 0.13.0), and feather reading / writing
pymysql 0.7.11 MySQL engine for sqlalchemy
pyreadstat SPSS files (.sav) reading
pytables 3.4.2 HDF5 reading / writing
diff --git a/doc/source/reference/io.rst b/doc/source/reference/io.rst
index 91f4942d03b0d..6d2d405a15850 100644
--- a/doc/source/reference/io.rst
+++ b/doc/source/reference/io.rst
@@ -98,6 +98,13 @@ Parquet
read_parquet
+ORC
+~~~
+.. autosummary::
+ :toctree: api/
+
+ read_orc
+
SAS
~~~
.. autosummary::
diff --git a/doc/source/user_guide/io.rst b/doc/source/user_guide/io.rst
index fa47a5944f7bf..972f36aecad24 100644
--- a/doc/source/user_guide/io.rst
+++ b/doc/source/user_guide/io.rst
@@ -28,6 +28,7 @@ The pandas I/O API is a set of top level ``reader`` functions accessed like
binary;`HDF5 Format <https://support.hdfgroup.org/HDF5/whatishdf5.html>`__;:ref:`read_hdf<io.hdf5>`;:ref:`to_hdf<io.hdf5>`
binary;`Feather Format <https://github.com/wesm/feather>`__;:ref:`read_feather<io.feather>`;:ref:`to_feather<io.feather>`
binary;`Parquet Format <https://parquet.apache.org/>`__;:ref:`read_parquet<io.parquet>`;:ref:`to_parquet<io.parquet>`
+ binary;`ORC Format <//https://orc.apache.org/>`__;:ref:`read_orc<io.orc>`;
binary;`Msgpack <https://msgpack.org/index.html>`__;:ref:`read_msgpack<io.msgpack>`;:ref:`to_msgpack<io.msgpack>`
binary;`Stata <https://en.wikipedia.org/wiki/Stata>`__;:ref:`read_stata<io.stata_reader>`;:ref:`to_stata<io.stata_writer>`
binary;`SAS <https://en.wikipedia.org/wiki/SAS_(software)>`__;:ref:`read_sas<io.sas_reader>`;
@@ -4858,6 +4859,17 @@ The above example creates a partitioned dataset that may look like:
except OSError:
pass
+.. _io.orc:
+
+ORC
+---
+
+.. versionadded:: 1.0.0
+
+Similar to the :ref:`parquet <io.parquet>` format, the `ORC Format <//https://orc.apache.org/>`__ is a binary columnar serialization
+for data frames. It is designed to make reading data frames efficient. Pandas provides *only* a reader for the
+ORC format, :func:`~pandas.read_orc`. This requires the `pyarrow <https://arrow.apache.org/docs/python/>`__ library.
+
.. _io.sql:
SQL queries
@@ -5761,6 +5773,3 @@ Space on disk (in bytes)
24009288 Oct 10 06:43 test_fixed_compress.hdf
24458940 Oct 10 06:44 test_table.hdf
24458940 Oct 10 06:44 test_table_compress.hdf
-
-
-
diff --git a/pandas/__init__.py b/pandas/__init__.py
index a60aa08b89f84..f72a12b58edcb 100644
--- a/pandas/__init__.py
+++ b/pandas/__init__.py
@@ -168,6 +168,7 @@
# misc
read_clipboard,
read_parquet,
+ read_orc,
read_feather,
read_gbq,
read_html,
diff --git a/pandas/io/api.py b/pandas/io/api.py
index 725e82604ca7f..e20aa18324a34 100644
--- a/pandas/io/api.py
+++ b/pandas/io/api.py
@@ -10,6 +10,7 @@
from pandas.io.gbq import read_gbq
from pandas.io.html import read_html
from pandas.io.json import read_json
+from pandas.io.orc import read_orc
from pandas.io.packers import read_msgpack, to_msgpack
from pandas.io.parquet import read_parquet
from pandas.io.parsers import read_csv, read_fwf, read_table
diff --git a/pandas/io/orc.py b/pandas/io/orc.py
new file mode 100644
index 0000000000000..bbefe447cb7fe
--- /dev/null
+++ b/pandas/io/orc.py
@@ -0,0 +1,57 @@
+""" orc compat """
+
+import distutils
+from typing import TYPE_CHECKING, List, Optional
+
+from pandas._typing import FilePathOrBuffer
+
+from pandas.io.common import get_filepath_or_buffer
+
+if TYPE_CHECKING:
+ from pandas import DataFrame
+
+
+def read_orc(
+ path: FilePathOrBuffer, columns: Optional[List[str]] = None, **kwargs,
+) -> "DataFrame":
+ """
+ Load an ORC object from the file path, returning a DataFrame.
+
+ .. versionadded:: 1.0.0
+
+ Parameters
+ ----------
+ path : str, path object or file-like object
+ Any valid string path is acceptable. The string could be a URL. Valid
+ URL schemes include http, ftp, s3, and file. For file URLs, a host is
+ expected. A local file could be:
+ ``file://localhost/path/to/table.orc``.
+
+ If you want to pass in a path object, pandas accepts any
+ ``os.PathLike``.
+
+ By file-like object, we refer to objects with a ``read()`` method,
+ such as a file handler (e.g. via builtin ``open`` function)
+ or ``StringIO``.
+ columns : list, default None
+ If not None, only these columns will be read from the file.
+ **kwargs
+ Any additional kwargs are passed to pyarrow.
+
+ Returns
+ -------
+ DataFrame
+ """
+
+ # we require a newer version of pyarrow than we support for parquet
+ import pyarrow
+
+ if distutils.version.LooseVersion(pyarrow.__version__) < "0.13.0":
+ raise ImportError("pyarrow must be >= 0.13.0 for read_orc")
+
+ import pyarrow.orc
+
+ path, _, _, _ = get_filepath_or_buffer(path)
+ orc_file = pyarrow.orc.ORCFile(path)
+ result = orc_file.read(columns=columns, **kwargs).to_pandas()
+ return result
diff --git a/pandas/tests/api/test_api.py b/pandas/tests/api/test_api.py
index 76141dceae930..870d7fd6e44c1 100644
--- a/pandas/tests/api/test_api.py
+++ b/pandas/tests/api/test_api.py
@@ -167,6 +167,7 @@ class TestPDApi(Base):
"read_table",
"read_feather",
"read_parquet",
+ "read_orc",
"read_spss",
]
diff --git a/pandas/tests/io/data/orc/TestOrcFile.decimal.orc b/pandas/tests/io/data/orc/TestOrcFile.decimal.orc
new file mode 100644
index 0000000000000..cb0f7b9d767a3
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.decimal.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.emptyFile.orc b/pandas/tests/io/data/orc/TestOrcFile.emptyFile.orc
new file mode 100644
index 0000000000000..ecdadcbff1346
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.emptyFile.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.test1.orc b/pandas/tests/io/data/orc/TestOrcFile.test1.orc
new file mode 100644
index 0000000000000..4fb0beff86897
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.test1.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.testDate1900.orc b/pandas/tests/io/data/orc/TestOrcFile.testDate1900.orc
new file mode 100644
index 0000000000000..f51ffdbd03a43
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.testDate1900.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.testDate2038.orc b/pandas/tests/io/data/orc/TestOrcFile.testDate2038.orc
new file mode 100644
index 0000000000000..cd11fa8a4e91d
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.testDate2038.orc differ
diff --git a/pandas/tests/io/data/orc/TestOrcFile.testSnappy.orc b/pandas/tests/io/data/orc/TestOrcFile.testSnappy.orc
new file mode 100644
index 0000000000000..aa6cc9c9ba1a7
Binary files /dev/null and b/pandas/tests/io/data/orc/TestOrcFile.testSnappy.orc differ
diff --git a/pandas/tests/io/test_orc.py b/pandas/tests/io/test_orc.py
new file mode 100644
index 0000000000000..9f3ec274007d0
--- /dev/null
+++ b/pandas/tests/io/test_orc.py
@@ -0,0 +1,227 @@
+""" test orc compat """
+import datetime
+import os
+
+import numpy as np
+import pytest
+
+import pandas as pd
+from pandas import read_orc
+import pandas.util.testing as tm
+
+pytest.importorskip("pyarrow", minversion="0.13.0")
+pytest.importorskip("pyarrow.orc")
+
+pytestmark = pytest.mark.filterwarnings(
+ "ignore:RangeIndex.* is deprecated:DeprecationWarning"
+)
+
+
+@pytest.fixture
+def dirpath(datapath):
+ return datapath("io", "data", "orc")
+
+
+def test_orc_reader_empty(dirpath):
+ columns = [
+ "boolean1",
+ "byte1",
+ "short1",
+ "int1",
+ "long1",
+ "float1",
+ "double1",
+ "bytes1",
+ "string1",
+ ]
+ dtypes = [
+ "bool",
+ "int8",
+ "int16",
+ "int32",
+ "int64",
+ "float32",
+ "float64",
+ "object",
+ "object",
+ ]
+ expected = pd.DataFrame(index=pd.RangeIndex(0))
+ for colname, dtype in zip(columns, dtypes):
+ expected[colname] = pd.Series(dtype=dtype)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.emptyFile.orc")
+ got = read_orc(inputfile, columns=columns)
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_basic(dirpath):
+ data = {
+ "boolean1": np.array([False, True], dtype="bool"),
+ "byte1": np.array([1, 100], dtype="int8"),
+ "short1": np.array([1024, 2048], dtype="int16"),
+ "int1": np.array([65536, 65536], dtype="int32"),
+ "long1": np.array([9223372036854775807, 9223372036854775807], dtype="int64"),
+ "float1": np.array([1.0, 2.0], dtype="float32"),
+ "double1": np.array([-15.0, -5.0], dtype="float64"),
+ "bytes1": np.array([b"\x00\x01\x02\x03\x04", b""], dtype="object"),
+ "string1": np.array(["hi", "bye"], dtype="object"),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.test1.orc")
+ got = read_orc(inputfile, columns=data.keys())
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_decimal(dirpath):
+ from decimal import Decimal
+
+ # Only testing the first 10 rows of data
+ data = {
+ "_col0": np.array(
+ [
+ Decimal("-1000.50000"),
+ Decimal("-999.60000"),
+ Decimal("-998.70000"),
+ Decimal("-997.80000"),
+ Decimal("-996.90000"),
+ Decimal("-995.10000"),
+ Decimal("-994.11000"),
+ Decimal("-993.12000"),
+ Decimal("-992.13000"),
+ Decimal("-991.14000"),
+ ],
+ dtype="object",
+ )
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.decimal.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_date_low(dirpath):
+ data = {
+ "time": np.array(
+ [
+ "1900-05-05 12:34:56.100000",
+ "1900-05-05 12:34:56.100100",
+ "1900-05-05 12:34:56.100200",
+ "1900-05-05 12:34:56.100300",
+ "1900-05-05 12:34:56.100400",
+ "1900-05-05 12:34:56.100500",
+ "1900-05-05 12:34:56.100600",
+ "1900-05-05 12:34:56.100700",
+ "1900-05-05 12:34:56.100800",
+ "1900-05-05 12:34:56.100900",
+ ],
+ dtype="datetime64[ns]",
+ ),
+ "date": np.array(
+ [
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ datetime.date(1900, 12, 25),
+ ],
+ dtype="object",
+ ),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.testDate1900.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_date_high(dirpath):
+ data = {
+ "time": np.array(
+ [
+ "2038-05-05 12:34:56.100000",
+ "2038-05-05 12:34:56.100100",
+ "2038-05-05 12:34:56.100200",
+ "2038-05-05 12:34:56.100300",
+ "2038-05-05 12:34:56.100400",
+ "2038-05-05 12:34:56.100500",
+ "2038-05-05 12:34:56.100600",
+ "2038-05-05 12:34:56.100700",
+ "2038-05-05 12:34:56.100800",
+ "2038-05-05 12:34:56.100900",
+ ],
+ dtype="datetime64[ns]",
+ ),
+ "date": np.array(
+ [
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ datetime.date(2038, 12, 25),
+ ],
+ dtype="object",
+ ),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.testDate2038.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
+
+
+def test_orc_reader_snappy_compressed(dirpath):
+ data = {
+ "int1": np.array(
+ [
+ -1160101563,
+ 1181413113,
+ 2065821249,
+ -267157795,
+ 172111193,
+ 1752363137,
+ 1406072123,
+ 1911809390,
+ -1308542224,
+ -467100286,
+ ],
+ dtype="int32",
+ ),
+ "string1": np.array(
+ [
+ "f50dcb8",
+ "382fdaaa",
+ "90758c6",
+ "9e8caf3f",
+ "ee97332b",
+ "d634da1",
+ "2bea4396",
+ "d67d89e8",
+ "ad71007e",
+ "e8c82066",
+ ],
+ dtype="object",
+ ),
+ }
+ expected = pd.DataFrame.from_dict(data)
+
+ inputfile = os.path.join(dirpath, "TestOrcFile.testSnappy.orc")
+ got = read_orc(inputfile).iloc[:10]
+
+ tm.assert_equal(expected, got)
| - [x] closes #25229
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
Added an ORC reader following the `read_parquet` API. Still need to give some additional love to the docstrings but this is at least ready for some discussion and eyes on it. | https://api.github.com/repos/pandas-dev/pandas/pulls/29447 | 2019-11-06T21:54:14Z | 2019-12-11T08:11:52Z | 2019-12-11T08:11:52Z | 2019-12-23T20:57:03Z |
DOC: Remove errant backslashes from the Ecosystem tab on new website. | diff --git a/web/pandas/community/ecosystem.md b/web/pandas/community/ecosystem.md
index cf242e86f879f..af6fd1ac77605 100644
--- a/web/pandas/community/ecosystem.md
+++ b/web/pandas/community/ecosystem.md
@@ -86,12 +86,12 @@ models to emphasize patterns in a dataset.
### [yhat/ggpy](https://github.com/yhat/ggpy)
-Hadley Wickham\'s [ggplot2](https://ggplot2.tidyverse.org/) is a
+Hadley Wickham's [ggplot2](https://ggplot2.tidyverse.org/) is a
foundational exploratory visualization package for the R language. Based
-on [\"The Grammar of
-Graphics\"](https://www.cs.uic.edu/~wilkinson/TheGrammarOfGraphics/GOG.html)
+on ["The Grammar of
+Graphics"](https://www.cs.uic.edu/~wilkinson/TheGrammarOfGraphics/GOG.html)
it provides a powerful, declarative and extremely general way to
-generate bespoke plots of any kind of data. It\'s really quite
+generate bespoke plots of any kind of data. It's really quite
incredible. Various implementations to other languages are available,
but a faithful implementation for Python users has long been missing.
Although still young (as of Jan-2014), the
@@ -100,9 +100,7 @@ quickly in that direction.
### [IPython Vega](https://github.com/vega/ipyvega)
-[IPython Vega](https://github.com/vega/ipyvega) leverages [Vega
-\<https://github.com/trifacta/vega\>]\_\_ to create plots
-within Jupyter Notebook.
+[IPython Vega](https://github.com/vega/ipyvega) leverages [Vega](https://github.com/vega/vega) to create plots within Jupyter Notebook.
### [Plotly](https://plot.ly/python)
@@ -158,8 +156,8 @@ for pandas `display.` settings.
### [quantopian/qgrid](https://github.com/quantopian/qgrid)
-qgrid is \"an interactive grid for sorting and filtering DataFrames in
-IPython Notebook\" built with SlickGrid.
+qgrid is "an interactive grid for sorting and filtering DataFrames in
+IPython Notebook" built with SlickGrid.
### [Spyder](https://www.spyder-ide.org/)
@@ -172,8 +170,8 @@ environment like MATLAB or Rstudio.
Its [Variable
Explorer](https://docs.spyder-ide.org/variableexplorer.html) allows
users to view, manipulate and edit pandas `Index`, `Series`, and
-`DataFrame` objects like a \"spreadsheet\", including copying and
-modifying values, sorting, displaying a \"heatmap\", converting data
+`DataFrame` objects like a "spreadsheet", including copying and
+modifying values, sorting, displaying a "heatmap", converting data
types and more. Pandas objects can also be renamed, duplicated, new
columns added, copyed/pasted to/from the clipboard (as TSV), and
saved/loaded to/from a file. Spyder can also import data from a variety
@@ -181,8 +179,8 @@ of plain text and binary files or the clipboard into a new pandas
DataFrame via a sophisticated import wizard.
Most pandas classes, methods and data attributes can be autocompleted in
-Spyder\'s [Editor](https://docs.spyder-ide.org/editor.html) and [IPython
-Console](https://docs.spyder-ide.org/ipythonconsole.html), and Spyder\'s
+Spyder's [Editor](https://docs.spyder-ide.org/editor.html) and [IPython
+Console](https://docs.spyder-ide.org/ipythonconsole.html), and Spyder's
[Help pane](https://docs.spyder-ide.org/help.html) can retrieve and
render Numpydoc documentation on pandas objects in rich text with Sphinx
both automatically and on-demand.
@@ -355,7 +353,7 @@ which work well with pandas' data containers.
### [cyberpandas](https://cyberpandas.readthedocs.io/en/latest)
Cyberpandas provides an extension type for storing arrays of IP
-Addresses. These arrays can be stored inside pandas\' Series and
+Addresses. These arrays can be stored inside pandas' Series and
DataFrame.
## Accessors
@@ -364,7 +362,7 @@ A directory of projects providing
`extension accessors <extending.register-accessors>`. This is for users to discover new accessors and for library
authors to coordinate on the namespace.
- Library Accessor Classes
- ------------------------------------------------------------- ---------- -----------------------
- [cyberpandas](https://cyberpandas.readthedocs.io/en/latest) `ip` `Series`
- [pdvega](https://altair-viz.github.io/pdvega/) `vgplot` `Series`, `DataFrame`
+ | Library | Accessor | Classes |
+ | ------------------------------------------------------------|----------|-----------------------|
+ | [cyberpandas](https://cyberpandas.readthedocs.io/en/latest) | `ip` | `Series` |
+ | [pdvega](https://altair-viz.github.io/pdvega/) | `vgplot` | `Series`, `DataFrame` |
| Pandas Sprint at PyData NYC :-)
Fixed at the direction of @datapythonista
The only file touched is a markdown file.
- [ ] closes #xxxx
- [ ] tests added / passed
- [ ] passes `black pandas`
- [ ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29446 | 2019-11-06T21:11:17Z | 2019-11-07T16:31:50Z | 2019-11-07T16:31:50Z | 2019-11-07T16:31:52Z |
fixup pip env | diff --git a/requirements-dev.txt b/requirements-dev.txt
index e7df704925485..13e2c95126f0c 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -1,5 +1,4 @@
numpy>=1.15
-python==3.7
python-dateutil>=2.6.1
pytz
asv
diff --git a/scripts/generate_pip_deps_from_conda.py b/scripts/generate_pip_deps_from_conda.py
index f1c7c3298fb26..6f809669d917f 100755
--- a/scripts/generate_pip_deps_from_conda.py
+++ b/scripts/generate_pip_deps_from_conda.py
@@ -19,7 +19,7 @@
import yaml
-EXCLUDE = {"python=3"}
+EXCLUDE = {"python"}
RENAME = {"pytables": "tables", "pyqt": "pyqt5", "dask-core": "dask"}
@@ -33,15 +33,15 @@ def conda_package_to_pip(package):
- A package requiring a specific version, in conda is defined with a single
equal (e.g. ``pandas=1.0``) and in pip with two (e.g. ``pandas==1.0``)
"""
- if package in EXCLUDE:
- return
-
package = re.sub("(?<=[^<>])=", "==", package).strip()
+
for compare in ("<=", ">=", "=="):
if compare not in package:
continue
pkg, version = package.split(compare)
+ if pkg in EXCLUDE:
+ return
if pkg in RENAME:
return "".join((RENAME[pkg], compare, version))
| Closes https://github.com/pandas-dev/pandas/issues/29443
cc @MomIsBestFriend | https://api.github.com/repos/pandas-dev/pandas/pulls/29445 | 2019-11-06T21:07:27Z | 2019-11-07T14:28:18Z | 2019-11-07T14:28:17Z | 2019-11-07T14:28:22Z |
Adding more documentation for upsampling with replacement and error m… | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index b40a64420a0be..1cb1f745fb61b 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -333,6 +333,7 @@ Numeric
- :class:`DataFrame` flex inequality comparisons methods (:meth:`DataFrame.lt`, :meth:`DataFrame.le`, :meth:`DataFrame.gt`, :meth: `DataFrame.ge`) with object-dtype and ``complex`` entries failing to raise ``TypeError`` like their :class:`Series` counterparts (:issue:`28079`)
- Bug in :class:`DataFrame` logical operations (`&`, `|`, `^`) not matching :class:`Series` behavior by filling NA values (:issue:`28741`)
- Bug in :meth:`DataFrame.interpolate` where specifying axis by name references variable before it is assigned (:issue:`29142`)
+- Improved error message when using `frac` > 1 and `replace` = False (:issue:`27451`)
-
Conversion
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index bafc37d478fdb..ffe8e794a03ea 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -4934,6 +4934,10 @@ def sample(
numpy.random.choice: Generates a random sample from a given 1-D numpy
array.
+ Notes
+ -----
+ If `frac` > 1, `replacement` should be set to `True`.
+
Examples
--------
>>> df = pd.DataFrame({'num_legs': [2, 4, 8, 0],
@@ -4964,6 +4968,20 @@ def sample(
dog 4 0 2
fish 0 0 8
+ An upsample sample of the ``DataFrame`` with replacement:
+ Note that `replace` parameter has to be `True` for `frac` parameter > 1.
+
+ >>> df.sample(frac=2, replace=True, random_state=1)
+ num_legs num_wings num_specimen_seen
+ dog 4 0 2
+ fish 0 0 8
+ falcon 2 2 10
+ falcon 2 2 10
+ fish 0 0 8
+ dog 4 0 2
+ fish 0 0 8
+ dog 4 0 2
+
Using a DataFrame column as weights. Rows with larger value in the
`num_specimen_seen` column are more likely to be sampled.
@@ -5039,6 +5057,11 @@ def sample(
# If no frac or n, default to n=1.
if n is None and frac is None:
n = 1
+ elif frac is not None and frac > 1 and not replace:
+ raise ValueError(
+ "Replace has to be set to `True` when "
+ "upsampling the population `frac` > 1."
+ )
elif n is not None and frac is None and n % 1 != 0:
raise ValueError("Only integers accepted as `n` values")
elif n is None and frac is not None:
diff --git a/pandas/tests/generic/test_generic.py b/pandas/tests/generic/test_generic.py
index a7506f3d60b3c..c180511e31619 100644
--- a/pandas/tests/generic/test_generic.py
+++ b/pandas/tests/generic/test_generic.py
@@ -322,6 +322,7 @@ def test_sample(self):
self._compare(
o.sample(n=4, random_state=seed), o.sample(n=4, random_state=seed)
)
+
self._compare(
o.sample(frac=0.7, random_state=seed),
o.sample(frac=0.7, random_state=seed),
@@ -337,6 +338,15 @@ def test_sample(self):
o.sample(frac=0.7, random_state=np.random.RandomState(test)),
)
+ self._compare(
+ o.sample(
+ frac=2, replace=True, random_state=np.random.RandomState(test)
+ ),
+ o.sample(
+ frac=2, replace=True, random_state=np.random.RandomState(test)
+ ),
+ )
+
os1, os2 = [], []
for _ in range(2):
np.random.seed(test)
@@ -424,6 +434,17 @@ def test_sample(self):
weights_with_None[5] = 0.5
self._compare(o.sample(n=1, axis=0, weights=weights_with_None), o.iloc[5:6])
+ def test_sample_upsampling_without_replacement(self):
+ # GH27451
+
+ df = pd.DataFrame({"A": list("abc")})
+ msg = (
+ "Replace has to be set to `True` when "
+ "upsampling the population `frac` > 1."
+ )
+ with pytest.raises(ValueError, match=msg):
+ df.sample(frac=2, replace=False)
+
def test_size_compat(self):
# GH8846
# size property should be defined
| …essage in case replacement is set to False
- [X] closes #27451
- [x] tests added / passed
- [X] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29444 | 2019-11-06T20:41:21Z | 2019-11-08T01:03:24Z | 2019-11-08T01:03:24Z | 2019-11-08T01:03:38Z |
TST: Add docstrings to arithmetic fixtures | diff --git a/pandas/tests/arithmetic/conftest.py b/pandas/tests/arithmetic/conftest.py
index 774ff14398bdb..1f8fdfd671856 100644
--- a/pandas/tests/arithmetic/conftest.py
+++ b/pandas/tests/arithmetic/conftest.py
@@ -21,7 +21,24 @@ def id_func(x):
@pytest.fixture(params=[1, np.array(1, dtype=np.int64)])
def one(request):
- # zero-dim integer array behaves like an integer
+ """
+ Several variants of integer value 1. The zero-dim integer array
+ behaves like an integer.
+
+ This fixture can be used to check that datetimelike indexes handle
+ addition and subtraction of integers and zero-dimensional arrays
+ of integers.
+
+ Examples
+ --------
+ >>> dti = pd.date_range('2016-01-01', periods=2, freq='H')
+ >>> dti
+ DatetimeIndex(['2016-01-01 00:00:00', '2016-01-01 01:00:00'],
+ dtype='datetime64[ns]', freq='H')
+ >>> dti + one
+ DatetimeIndex(['2016-01-01 01:00:00', '2016-01-01 02:00:00'],
+ dtype='datetime64[ns]', freq='H')
+ """
return request.param
@@ -40,8 +57,21 @@ def one(request):
@pytest.fixture(params=zeros)
def zero(request):
- # For testing division by (or of) zero for Index with length 5, this
- # gives several scalar-zeros and length-5 vector-zeros
+ """
+ Several types of scalar zeros and length 5 vectors of zeros.
+
+ This fixture can be used to check that numeric-dtype indexes handle
+ division by any zero numeric-dtype.
+
+ Uses vector of length 5 for broadcasting with `numeric_idx` fixture,
+ which creates numeric-dtype vectors also of length 5.
+
+ Examples
+ --------
+ >>> arr = pd.RangeIndex(5)
+ >>> arr / zeros
+ Float64Index([nan, inf, inf, inf, inf], dtype='float64')
+ """
return request.param
| Relates to #19159
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29441 | 2019-11-06T20:27:23Z | 2019-11-18T01:40:48Z | 2019-11-18T01:40:48Z | 2019-11-18T01:40:59Z |
add unit tests for issue #19351 | diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 79c9fe2b60bd9..a1f58922ea0ca 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -358,6 +358,49 @@ def test_unstack(self):
# test that int32 work
self.ymd.astype(np.int32).unstack()
+ @pytest.mark.parametrize(
+ "result_rows,result_columns,index_product,expected_row",
+ [
+ (
+ [[1, 1, None, None, 30.0, None], [2, 2, None, None, 30.0, None]],
+ [u"ix1", u"ix2", u"col1", u"col2", u"col3", u"col4"],
+ 2,
+ [None, None, 30.0, None],
+ ),
+ (
+ [[1, 1, None, None, 30.0], [2, 2, None, None, 30.0]],
+ [u"ix1", u"ix2", u"col1", u"col2", u"col3"],
+ 2,
+ [None, None, 30.0],
+ ),
+ (
+ [[1, 1, None, None, 30.0], [2, None, None, None, 30.0]],
+ [u"ix1", u"ix2", u"col1", u"col2", u"col3"],
+ None,
+ [None, None, 30.0],
+ ),
+ ],
+ )
+ def test_unstack_partial(
+ self, result_rows, result_columns, index_product, expected_row
+ ):
+ # check for regressions on this issue:
+ # https://github.com/pandas-dev/pandas/issues/19351
+ # make sure DataFrame.unstack() works when its run on a subset of the DataFrame
+ # and the Index levels contain values that are not present in the subset
+ result = pd.DataFrame(result_rows, columns=result_columns).set_index(
+ [u"ix1", "ix2"]
+ )
+ result = result.iloc[1:2].unstack("ix2")
+ expected = pd.DataFrame(
+ [expected_row],
+ columns=pd.MultiIndex.from_product(
+ [result_columns[2:], [index_product]], names=[None, "ix2"]
+ ),
+ index=pd.Index([2], name="ix1"),
+ )
+ tm.assert_frame_equal(result, expected)
+
def test_unstack_multiple_no_empty_columns(self):
index = MultiIndex.from_tuples(
[(0, "foo", 0), (0, "bar", 0), (1, "baz", 1), (1, "qux", 1)]
| - [x] closes #19351
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
This is a unit test for an already fixed problem.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29438 | 2019-11-06T19:33:12Z | 2019-11-06T21:23:27Z | 2019-11-06T21:23:27Z | 2019-11-06T21:23:33Z |
CLN: remove unnecessary check in MultiIndex | diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index 2e3f440573a0f..fe91a588c7dde 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -2164,6 +2164,7 @@ def drop(self, codes, level=None, errors="raise"):
codes : array-like
Must be a list of tuples
level : int or level name, default None
+ errors : str, default 'raise'
Returns
-------
@@ -2172,18 +2173,11 @@ def drop(self, codes, level=None, errors="raise"):
if level is not None:
return self._drop_from_level(codes, level)
- try:
- if not isinstance(codes, (np.ndarray, Index)):
+ if not isinstance(codes, (np.ndarray, Index)):
+ try:
codes = com.index_labels_to_array(codes)
- indexer = self.get_indexer(codes)
- mask = indexer == -1
- if mask.any():
- if errors != "ignore":
- raise ValueError(
- "codes {codes} not contained in axis".format(codes=codes[mask])
- )
- except Exception:
- pass
+ except ValueError:
+ pass
inds = []
for level_codes in codes:
| We raise a ValueError and immediately ignore it. | https://api.github.com/repos/pandas-dev/pandas/pulls/29437 | 2019-11-06T19:06:28Z | 2019-11-06T21:22:46Z | 2019-11-06T21:22:46Z | 2019-11-06T21:30:26Z |
Pr09 batch 3 | diff --git a/pandas/_libs/lib.pyx b/pandas/_libs/lib.pyx
index 328b67b6722f1..a14efd3313eaf 100644
--- a/pandas/_libs/lib.pyx
+++ b/pandas/_libs/lib.pyx
@@ -125,7 +125,7 @@ def is_scalar(val: object) -> bool:
- Interval
- DateOffset
- Fraction
- - Number
+ - Number.
Returns
-------
@@ -867,9 +867,10 @@ def is_list_like(obj: object, allow_sets: bool = True):
Parameters
----------
- obj : The object to check
- allow_sets : boolean, default True
- If this parameter is False, sets will not be considered list-like
+ obj : object
+ The object to check.
+ allow_sets : bool, default True
+ If this parameter is False, sets will not be considered list-like.
.. versionadded:: 0.24.0
diff --git a/pandas/core/arrays/interval.py b/pandas/core/arrays/interval.py
index 4039cc91fb554..869019cd3d222 100644
--- a/pandas/core/arrays/interval.py
+++ b/pandas/core/arrays/interval.py
@@ -260,9 +260,9 @@ def _from_factorized(cls, values, original):
Whether the intervals are closed on the left-side, right-side, both
or neither.
copy : bool, default False
- copy the data
+ Copy the data.
dtype : dtype or None, default None
- If None, dtype will be inferred
+ If None, dtype will be inferred.
.. versionadded:: 0.23.0
@@ -383,16 +383,16 @@ def from_arrays(cls, left, right, closed="right", copy=False, dtype=None):
Parameters
----------
data : array-like (1-dimensional)
- Array of tuples
+ Array of tuples.
closed : {'left', 'right', 'both', 'neither'}, default 'right'
Whether the intervals are closed on the left-side, right-side, both
or neither.
copy : bool, default False
- by-default copy the data, this is compat only and ignored
+ By-default copy the data, this is compat only and ignored.
dtype : dtype or None, default None
- If None, dtype will be inferred
+ If None, dtype will be inferred.
- ..versionadded:: 0.23.0
+ .. versionadded:: 0.23.0
Returns
-------
diff --git a/pandas/core/dtypes/concat.py b/pandas/core/dtypes/concat.py
index f2176f573207c..a62d3d0f4e65b 100644
--- a/pandas/core/dtypes/concat.py
+++ b/pandas/core/dtypes/concat.py
@@ -185,13 +185,14 @@ def concat_categorical(to_concat, axis=0):
def union_categoricals(to_union, sort_categories=False, ignore_order=False):
"""
- Combine list-like of Categorical-like, unioning categories. All
- categories must have the same dtype.
+ Combine list-like of Categorical-like, unioning categories.
+
+ All categories must have the same dtype.
Parameters
----------
- to_union : list-like of Categorical, CategoricalIndex,
- or Series with dtype='category'
+ to_union : list-like
+ Categorical, CategoricalIndex, or Series with dtype='category'.
sort_categories : bool, default False
If true, resulting categories will be lexsorted, otherwise
they will be ordered as they appear in the data.
@@ -201,7 +202,7 @@ def union_categoricals(to_union, sort_categories=False, ignore_order=False):
Returns
-------
- result : Categorical
+ Categorical
Raises
------
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index 9d7ddcf3c7727..e418461883e6c 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -441,7 +441,7 @@ def pad(self, limit=None):
Parameters
----------
limit : int, optional
- limit of how many values to fill
+ Limit of how many values to fill.
Returns
-------
@@ -856,7 +856,7 @@ def var(self, ddof=1, *args, **kwargs):
Parameters
----------
ddof : int, default 1
- degrees of freedom
+ Degrees of freedom.
Returns
-------
@@ -1237,11 +1237,11 @@ def _upsample(self, method, limit=None, fill_value=None):
Parameters
----------
method : string {'backfill', 'bfill', 'pad', 'ffill'}
- method for upsampling
+ Method for upsampling.
limit : int, default None
- Maximum size gap to fill when reindexing
+ Maximum size gap to fill when reindexing.
fill_value : scalar, default None
- Value to use for missing values
+ Value to use for missing values.
See Also
--------
diff --git a/pandas/io/html.py b/pandas/io/html.py
index 7da7a819f81e8..9a368907b65aa 100644
--- a/pandas/io/html.py
+++ b/pandas/io/html.py
@@ -960,7 +960,7 @@ def read_html(
This value is converted to a regular expression so that there is
consistent behavior between Beautiful Soup and lxml.
- flavor : str or None, container of strings
+ flavor : str or None
The parsing engine to use. 'bs4' and 'html5lib' are synonymous with
each other, they are both there for backwards compatibility. The
default of ``None`` tries to use ``lxml`` to parse and if that fails it
@@ -974,7 +974,7 @@ def read_html(
The column (or list of columns) to use to create the index.
skiprows : int or list-like or slice or None, optional
- 0-based. Number of rows to skip after parsing the column integer. If a
+ Number of rows to skip after parsing the column integer. 0-based. If a
sequence of integers or a slice is given, will skip the rows indexed by
that sequence. Note that a single element sequence means 'skip the nth
row' whereas an integer means 'skip n rows'.
@@ -1024,18 +1024,19 @@ def read_html(
transformed content.
na_values : iterable, default None
- Custom NA values
+ Custom NA values.
keep_default_na : bool, default True
If na_values are specified and keep_default_na is False the default NaN
- values are overridden, otherwise they're appended to
+ values are overridden, otherwise they're appended to.
displayed_only : bool, default True
- Whether elements with "display: none" should be parsed
+ Whether elements with "display: none" should be parsed.
Returns
-------
- dfs : list of DataFrames
+ dfs
+ A list of DataFrames.
See Also
--------
diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index f3b0226547c78..5a2f189ad8d10 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -593,14 +593,14 @@ def assert_index_equal(
check_less_precise : bool or int, default False
Specify comparison precision. Only used when check_exact is False.
5 digits (False) or 3 digits (True) after decimal points are compared.
- If int, then specify the digits to compare
+ If int, then specify the digits to compare.
check_exact : bool, default True
Whether to compare number exactly.
check_categorical : bool, default True
Whether to compare internal Categorical exactly.
obj : str, default 'Index'
Specify object name being compared, internally used to show appropriate
- assertion message
+ assertion message.
"""
__tracebackhide__ = True
@@ -1273,10 +1273,7 @@ def assert_frame_equal(
check whether it is equivalent to 1 within the specified precision.
check_names : bool, default True
Whether to check that the `names` attribute for both the `index`
- and `column` attributes of the DataFrame is identical, i.e.
-
- * left.index.names == right.index.names
- * left.columns.names == right.columns.names
+ and `column` attributes of the DataFrame is identical.
by_blocks : bool, default False
Specify how to compare internal data. If False, compare by columns.
If True, compare by blocks.
| part of #28602 | https://api.github.com/repos/pandas-dev/pandas/pulls/29434 | 2019-11-06T17:03:42Z | 2019-11-06T20:36:34Z | 2019-11-06T20:36:34Z | 2020-01-06T16:47:00Z |
CI: workaround numpydev bug | diff --git a/ci/azure/posix.yml b/ci/azure/posix.yml
index 281107559a38c..62b15bae6d2ce 100644
--- a/ci/azure/posix.yml
+++ b/ci/azure/posix.yml
@@ -45,13 +45,16 @@ jobs:
PATTERN: "not slow and not network"
LOCALE_OVERRIDE: "zh_CN.UTF-8"
- py37_np_dev:
- ENV_FILE: ci/deps/azure-37-numpydev.yaml
- CONDA_PY: "37"
- PATTERN: "not slow and not network"
- TEST_ARGS: "-W error"
- PANDAS_TESTING_MODE: "deprecate"
- EXTRA_APT: "xsel"
+ # https://github.com/pandas-dev/pandas/issues/29432
+ # py37_np_dev:
+ # ENV_FILE: ci/deps/azure-37-numpydev.yaml
+ # CONDA_PY: "37"
+ # PATTERN: "not slow and not network"
+ # TEST_ARGS: "-W error"
+ # PANDAS_TESTING_MODE: "deprecate"
+ # EXTRA_APT: "xsel"
+ # # TODO:
+ # continueOnError: true
steps:
- script: |
| We don't want this long-term. But there's no easy way
to skip this for numpydev, since it errors in setup.
xref #29432 (keep open till numpydev is fixed) | https://api.github.com/repos/pandas-dev/pandas/pulls/29433 | 2019-11-06T16:31:35Z | 2019-11-06T19:10:04Z | 2019-11-06T19:10:03Z | 2019-11-06T21:21:01Z |
Cleanup env | diff --git a/environment.yml b/environment.yml
index 443dc483aedf8..e9ac76f5bc52c 100644
--- a/environment.yml
+++ b/environment.yml
@@ -1,11 +1,10 @@
name: pandas-dev
channels:
- - defaults
- conda-forge
dependencies:
# required
- numpy>=1.15
- - python=3
+ - python=3.7
- python-dateutil>=2.6.1
- pytz
@@ -22,7 +21,7 @@ dependencies:
- flake8-comprehensions # used by flake8, linting of unnecessary comprehensions
- flake8-rst>=0.6.0,<=0.7.0 # linting of code blocks in rst files
- isort # check that imports are in the right order
- - mypy
+ - mypy=0.720
- pycodestyle # used by flake8
# documentation
@@ -54,7 +53,6 @@ dependencies:
- moto # mock S3
- pytest>=4.0.2
- pytest-cov
- - pytest-mock
- pytest-xdist
- seaborn
- statsmodels
diff --git a/requirements-dev.txt b/requirements-dev.txt
index 7a378cd2f2697..e7df704925485 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -1,4 +1,5 @@
numpy>=1.15
+python==3.7
python-dateutil>=2.6.1
pytz
asv
@@ -9,7 +10,7 @@ flake8
flake8-comprehensions
flake8-rst>=0.6.0,<=0.7.0
isort
-mypy
+mypy==0.720
pycodestyle
gitpython
sphinx
@@ -32,7 +33,6 @@ hypothesis>=3.82
moto
pytest>=4.0.2
pytest-cov
-pytest-mock
pytest-xdist
seaborn
statsmodels
| Closes #29330
This was most likely due to inconsistent constraints between conda-forge & defaults.
Also, pinning to 3.7 for now until the 3.8 buildout is done to make the solver's life a bit easier. | https://api.github.com/repos/pandas-dev/pandas/pulls/29431 | 2019-11-06T16:09:15Z | 2019-11-06T18:12:52Z | 2019-11-06T18:12:51Z | 2019-11-06T18:13:05Z |
BUG: Styling user guide points to a wrong nbviewer link | diff --git a/doc/source/user_guide/style.ipynb b/doc/source/user_guide/style.ipynb
index 006f928c037bd..e0dc2e734e660 100644
--- a/doc/source/user_guide/style.ipynb
+++ b/doc/source/user_guide/style.ipynb
@@ -6,7 +6,7 @@
"source": [
"# Styling\n",
"\n",
- "This document is written as a Jupyter Notebook, and can be viewed or downloaded [here](http://nbviewer.ipython.org/github/pandas-dev/pandas/blob/master/doc/source/style.ipynb).\n",
+ "This document is written as a Jupyter Notebook, and can be viewed or downloaded [here](http://nbviewer.ipython.org/github/pandas-dev/pandas/blob/master/doc/source/user_guide/style.ipynb).\n",
"\n",
"You can apply **conditional formatting**, the visual styling of a DataFrame\n",
"depending on the data within, by using the ``DataFrame.style`` property.\n",
| Just missing the 'user_guide' part => one line change.
- [ ] closes #xxxx (NOT THAT I KNOW)
- [ ] tests added / passed (NO CODE CHANGE)
- [ ] passes `black pandas` (NO CODE CHANGE)
- [ ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff` (NO CODE CHANGE)
- [ ] whatsnew entry (NO CHANGE)
| https://api.github.com/repos/pandas-dev/pandas/pulls/29429 | 2019-11-06T09:13:30Z | 2019-11-06T16:45:13Z | 2019-11-06T16:45:13Z | 2019-11-06T16:45:18Z |
REF: Separate window bounds calculation from aggregation functions | diff --git a/pandas/_libs/window.pyx b/pandas/_libs/window.pyx
index d6bad0f20d760..303b4f6f24eac 100644
--- a/pandas/_libs/window.pyx
+++ b/pandas/_libs/window.pyx
@@ -96,280 +96,20 @@ def _check_minp(win, minp, N, floor=None) -> int:
# Physical description: 366 p.
# Series: Prentice-Hall Series in Automatic Computation
-# ----------------------------------------------------------------------
-# The indexer objects for rolling
-# These define start/end indexers to compute offsets
-
-
-cdef class WindowIndexer:
-
- cdef:
- ndarray start, end
- int64_t N, minp, win
- bint is_variable
-
- def get_data(self):
- return (self.start, self.end, <int64_t>self.N,
- <int64_t>self.win, <int64_t>self.minp,
- self.is_variable)
-
-
-cdef class MockFixedWindowIndexer(WindowIndexer):
- """
-
- We are just checking parameters of the indexer,
- and returning a consistent API with fixed/variable
- indexers.
-
- Parameters
- ----------
- values: ndarray
- values data array
- win: int64_t
- window size
- minp: int64_t
- min number of obs in a window to consider non-NaN
- index: object
- index of the values
- floor: optional
- unit for flooring
- left_closed: bint
- left endpoint closedness
- right_closed: bint
- right endpoint closedness
-
- """
- def __init__(self, ndarray values, int64_t win, int64_t minp,
- bint left_closed, bint right_closed,
- object index=None, object floor=None):
-
- assert index is None
- self.is_variable = 0
- self.N = len(values)
- self.minp = _check_minp(win, minp, self.N, floor=floor)
- self.start = np.empty(0, dtype='int64')
- self.end = np.empty(0, dtype='int64')
- self.win = win
-
-
-cdef class FixedWindowIndexer(WindowIndexer):
- """
- create a fixed length window indexer object
- that has start & end, that point to offsets in
- the index object; these are defined based on the win
- arguments
-
- Parameters
- ----------
- values: ndarray
- values data array
- win: int64_t
- window size
- minp: int64_t
- min number of obs in a window to consider non-NaN
- index: object
- index of the values
- floor: optional
- unit for flooring the unit
- left_closed: bint
- left endpoint closedness
- right_closed: bint
- right endpoint closedness
-
- """
- def __init__(self, ndarray values, int64_t win, int64_t minp,
- bint left_closed, bint right_closed,
- object index=None, object floor=None):
- cdef:
- ndarray[int64_t] start_s, start_e, end_s, end_e
-
- assert index is None
- self.is_variable = 0
- self.N = len(values)
- self.minp = _check_minp(win, minp, self.N, floor=floor)
-
- start_s = np.zeros(win, dtype='int64')
- start_e = np.arange(win, self.N, dtype='int64') - win + 1
- self.start = np.concatenate([start_s, start_e])
-
- end_s = np.arange(win, dtype='int64') + 1
- end_e = start_e + win
- self.end = np.concatenate([end_s, end_e])
- self.win = win
-
-
-cdef class VariableWindowIndexer(WindowIndexer):
- """
- create a variable length window indexer object
- that has start & end, that point to offsets in
- the index object; these are defined based on the win
- arguments
-
- Parameters
- ----------
- values: ndarray
- values data array
- win: int64_t
- window size
- minp: int64_t
- min number of obs in a window to consider non-NaN
- index: ndarray
- index of the values
- left_closed: bint
- left endpoint closedness
- True if the left endpoint is closed, False if open
- right_closed: bint
- right endpoint closedness
- True if the right endpoint is closed, False if open
- floor: optional
- unit for flooring the unit
- """
- def __init__(self, ndarray values, int64_t win, int64_t minp,
- bint left_closed, bint right_closed, ndarray index,
- object floor=None):
-
- self.is_variable = 1
- self.N = len(index)
- self.minp = _check_minp(win, minp, self.N, floor=floor)
-
- self.start = np.empty(self.N, dtype='int64')
- self.start.fill(-1)
-
- self.end = np.empty(self.N, dtype='int64')
- self.end.fill(-1)
-
- self.build(index, win, left_closed, right_closed)
-
- # max window size
- self.win = (self.end - self.start).max()
-
- def build(self, const int64_t[:] index, int64_t win, bint left_closed,
- bint right_closed):
-
- cdef:
- ndarray[int64_t] start, end
- int64_t start_bound, end_bound, N
- Py_ssize_t i, j
-
- start = self.start
- end = self.end
- N = self.N
-
- start[0] = 0
-
- # right endpoint is closed
- if right_closed:
- end[0] = 1
- # right endpoint is open
- else:
- end[0] = 0
-
- with nogil:
-
- # start is start of slice interval (including)
- # end is end of slice interval (not including)
- for i in range(1, N):
- end_bound = index[i]
- start_bound = index[i] - win
-
- # left endpoint is closed
- if left_closed:
- start_bound -= 1
-
- # advance the start bound until we are
- # within the constraint
- start[i] = i
- for j in range(start[i - 1], i):
- if index[j] > start_bound:
- start[i] = j
- break
-
- # end bound is previous end
- # or current index
- if index[end[i - 1]] <= end_bound:
- end[i] = i + 1
- else:
- end[i] = end[i - 1]
-
- # right endpoint is open
- if not right_closed:
- end[i] -= 1
-
-
-def get_window_indexer(values, win, minp, index, closed,
- floor=None, use_mock=True):
- """
- Return the correct window indexer for the computation.
-
- Parameters
- ----------
- values: 1d ndarray
- win: integer, window size
- minp: integer, minimum periods
- index: 1d ndarray, optional
- index to the values array
- closed: string, default None
- {'right', 'left', 'both', 'neither'}
- window endpoint closedness. Defaults to 'right' in
- VariableWindowIndexer and to 'both' in FixedWindowIndexer
- floor: optional
- unit for flooring the unit
- use_mock: boolean, default True
- if we are a fixed indexer, return a mock indexer
- instead of the FixedWindow Indexer. This is a type
- compat Indexer that allows us to use a standard
- code path with all of the indexers.
-
- Returns
- -------
- tuple of 1d int64 ndarrays of the offsets & data about the window
-
- """
-
- cdef:
- bint left_closed = False
- bint right_closed = False
-
- assert closed is None or closed in ['right', 'left', 'both', 'neither']
-
- # if windows is variable, default is 'right', otherwise default is 'both'
- if closed is None:
- closed = 'right' if index is not None else 'both'
-
- if closed in ['right', 'both']:
- right_closed = True
-
- if closed in ['left', 'both']:
- left_closed = True
-
- if index is not None:
- indexer = VariableWindowIndexer(values, win, minp, left_closed,
- right_closed, index, floor)
- elif use_mock:
- indexer = MockFixedWindowIndexer(values, win, minp, left_closed,
- right_closed, index, floor)
- else:
- indexer = FixedWindowIndexer(values, win, minp, left_closed,
- right_closed, index, floor)
- return indexer.get_data()
-
# ----------------------------------------------------------------------
# Rolling count
# this is only an impl for index not None, IOW, freq aware
-def roll_count(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_count(ndarray[float64_t] values, ndarray[int64_t] start, ndarray[int64_t] end,
+ int64_t minp):
cdef:
float64_t val, count_x = 0.0
- int64_t s, e, nobs, N
+ int64_t s, e, nobs, N = len(values)
Py_ssize_t i, j
- int64_t[:] start, end
ndarray[float64_t] output
- start, end, N, win, minp, _ = get_window_indexer(values, win,
- minp, index, closed)
output = np.empty(N, dtype=float)
with nogil:
@@ -442,80 +182,75 @@ cdef inline void remove_sum(float64_t val, int64_t *nobs, float64_t *sum_x) nogi
sum_x[0] = sum_x[0] - val
-def roll_sum(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_sum_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
cdef:
- float64_t val, prev_x, sum_x = 0
- int64_t s, e, range_endpoint
- int64_t nobs = 0, i, j, N
- bint is_variable
- int64_t[:] start, end
+ float64_t sum_x = 0
+ int64_t s, e
+ int64_t nobs = 0, i, j, N = len(values)
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed,
- floor=0)
output = np.empty(N, dtype=float)
- # for performance we are going to iterate
- # fixed windows separately, makes the code more complex as we have 2 paths
- # but is faster
+ with nogil:
- if is_variable:
+ for i in range(0, N):
+ s = start[i]
+ e = end[i]
- # variable window
- with nogil:
+ if i == 0:
- for i in range(0, N):
- s = start[i]
- e = end[i]
+ # setup
+ sum_x = 0.0
+ nobs = 0
+ for j in range(s, e):
+ add_sum(values[j], &nobs, &sum_x)
- if i == 0:
+ else:
- # setup
- sum_x = 0.0
- nobs = 0
- for j in range(s, e):
- add_sum(values[j], &nobs, &sum_x)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ remove_sum(values[j], &nobs, &sum_x)
- else:
+ # calculate adds
+ for j in range(end[i - 1], e):
+ add_sum(values[j], &nobs, &sum_x)
- # calculate deletes
- for j in range(start[i - 1], s):
- remove_sum(values[j], &nobs, &sum_x)
+ output[i] = calc_sum(minp, nobs, sum_x)
- # calculate adds
- for j in range(end[i - 1], e):
- add_sum(values[j], &nobs, &sum_x)
+ return output
- output[i] = calc_sum(minp, nobs, sum_x)
- else:
+def roll_sum_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
+ cdef:
+ float64_t val, prev_x, sum_x = 0
+ int64_t range_endpoint
+ int64_t nobs = 0, i, N = len(values)
+ ndarray[float64_t] output
- # fixed window
+ output = np.empty(N, dtype=float)
- range_endpoint = int_max(minp, 1) - 1
+ range_endpoint = int_max(minp, 1) - 1
- with nogil:
+ with nogil:
- for i in range(0, range_endpoint):
- add_sum(values[i], &nobs, &sum_x)
- output[i] = NaN
+ for i in range(0, range_endpoint):
+ add_sum(values[i], &nobs, &sum_x)
+ output[i] = NaN
- for i in range(range_endpoint, N):
- val = values[i]
- add_sum(val, &nobs, &sum_x)
+ for i in range(range_endpoint, N):
+ val = values[i]
+ add_sum(val, &nobs, &sum_x)
- if i > win - 1:
- prev_x = values[i - win]
- remove_sum(prev_x, &nobs, &sum_x)
+ if i > win - 1:
+ prev_x = values[i - win]
+ remove_sum(prev_x, &nobs, &sum_x)
- output[i] = calc_sum(minp, nobs, sum_x)
+ output[i] = calc_sum(minp, nobs, sum_x)
return output
-
# ----------------------------------------------------------------------
# Rolling mean
@@ -563,77 +298,75 @@ cdef inline void remove_mean(float64_t val, Py_ssize_t *nobs, float64_t *sum_x,
neg_ct[0] = neg_ct[0] - 1
-def roll_mean(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_mean_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
- float64_t val, prev_x, result, sum_x = 0
- int64_t s, e
- bint is_variable
- Py_ssize_t nobs = 0, i, j, neg_ct = 0, N
- int64_t[:] start, end
+ float64_t val, prev_x, sum_x = 0
+ Py_ssize_t nobs = 0, i, neg_ct = 0, N = len(values)
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
- # for performance we are going to iterate
- # fixed windows separately, makes the code more complex as we have 2 paths
- # but is faster
+ with nogil:
+ for i in range(minp - 1):
+ val = values[i]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
+ output[i] = NaN
+
+ for i in range(minp - 1, N):
+ val = values[i]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
- if is_variable:
+ if i > win - 1:
+ prev_x = values[i - win]
+ remove_mean(prev_x, &nobs, &sum_x, &neg_ct)
- with nogil:
+ output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
- for i in range(0, N):
- s = start[i]
- e = end[i]
+ return output
- if i == 0:
- # setup
- sum_x = 0.0
- nobs = 0
- for j in range(s, e):
- val = values[j]
- add_mean(val, &nobs, &sum_x, &neg_ct)
+def roll_mean_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ cdef:
+ float64_t val, sum_x = 0
+ int64_t s, e
+ Py_ssize_t nobs = 0, i, j, neg_ct = 0, N = len(values)
+ ndarray[float64_t] output
- else:
+ output = np.empty(N, dtype=float)
- # calculate deletes
- for j in range(start[i - 1], s):
- val = values[j]
- remove_mean(val, &nobs, &sum_x, &neg_ct)
+ with nogil:
- # calculate adds
- for j in range(end[i - 1], e):
- val = values[j]
- add_mean(val, &nobs, &sum_x, &neg_ct)
+ for i in range(0, N):
+ s = start[i]
+ e = end[i]
- output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
+ if i == 0:
- else:
+ # setup
+ sum_x = 0.0
+ nobs = 0
+ for j in range(s, e):
+ val = values[j]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
- with nogil:
- for i in range(minp - 1):
- val = values[i]
- add_mean(val, &nobs, &sum_x, &neg_ct)
- output[i] = NaN
+ else:
- for i in range(minp - 1, N):
- val = values[i]
- add_mean(val, &nobs, &sum_x, &neg_ct)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ val = values[j]
+ remove_mean(val, &nobs, &sum_x, &neg_ct)
- if i > win - 1:
- prev_x = values[i - win]
- remove_mean(prev_x, &nobs, &sum_x, &neg_ct)
+ # calculate adds
+ for j in range(end[i - 1], e):
+ val = values[j]
+ add_mean(val, &nobs, &sum_x, &neg_ct)
- output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
+ output[i] = calc_mean(minp, nobs, neg_ct, sum_x)
return output
-
# ----------------------------------------------------------------------
# Rolling variance
@@ -696,8 +429,8 @@ cdef inline void remove_var(float64_t val, float64_t *nobs, float64_t *mean_x,
ssqdm_x[0] = 0
-def roll_var(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed, int ddof=1):
+def roll_var_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win, int ddof=1):
"""
Numerically stable implementation using Welford's method.
"""
@@ -705,98 +438,102 @@ def roll_var(ndarray[float64_t] values, int64_t win, int64_t minp,
float64_t mean_x = 0, ssqdm_x = 0, nobs = 0,
float64_t val, prev, delta, mean_x_old
int64_t s, e
- bint is_variable
- Py_ssize_t i, j, N
- int64_t[:] start, end
+ Py_ssize_t i, j, N = len(values)
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
# Check for windows larger than array, addresses #7297
win = min(win, N)
- # for performance we are going to iterate
- # fixed windows separately, makes the code more complex as we
- # have 2 paths but is faster
+ with nogil:
- if is_variable:
+ # Over the first window, observations can only be added, never
+ # removed
+ for i in range(win):
+ add_var(values[i], &nobs, &mean_x, &ssqdm_x)
+ output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
- with nogil:
+ # a part of Welford's method for the online variance-calculation
+ # https://en.wikipedia.org/wiki/Algorithms_for_calculating_variance
- for i in range(0, N):
+ # After the first window, observations can both be added and
+ # removed
+ for i in range(win, N):
+ val = values[i]
+ prev = values[i - win]
- s = start[i]
- e = end[i]
+ if notnan(val):
+ if prev == prev:
- # Over the first window, observations can only be added
- # never removed
- if i == 0:
+ # Adding one observation and removing another one
+ delta = val - prev
+ mean_x_old = mean_x
- for j in range(s, e):
- add_var(values[j], &nobs, &mean_x, &ssqdm_x)
+ mean_x += delta / nobs
+ ssqdm_x += ((nobs - 1) * val
+ + (nobs + 1) * prev
+ - 2 * nobs * mean_x_old) * delta / nobs
else:
+ add_var(val, &nobs, &mean_x, &ssqdm_x)
+ elif prev == prev:
+ remove_var(prev, &nobs, &mean_x, &ssqdm_x)
- # After the first window, observations can both be added
- # and removed
+ output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+
+ return output
- # calculate adds
- for j in range(end[i - 1], e):
- add_var(values[j], &nobs, &mean_x, &ssqdm_x)
- # calculate deletes
- for j in range(start[i - 1], s):
- remove_var(values[j], &nobs, &mean_x, &ssqdm_x)
+def roll_var_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int ddof=1):
+ """
+ Numerically stable implementation using Welford's method.
+ """
+ cdef:
+ float64_t mean_x = 0, ssqdm_x = 0, nobs = 0,
+ float64_t val, prev, delta, mean_x_old
+ int64_t s, e
+ Py_ssize_t i, j, N = len(values)
+ ndarray[float64_t] output
- output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+ output = np.empty(N, dtype=float)
- else:
+ with nogil:
- with nogil:
+ for i in range(0, N):
- # Over the first window, observations can only be added, never
- # removed
- for i in range(win):
- add_var(values[i], &nobs, &mean_x, &ssqdm_x)
- output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+ s = start[i]
+ e = end[i]
- # a part of Welford's method for the online variance-calculation
- # https://en.wikipedia.org/wiki/Algorithms_for_calculating_variance
+ # Over the first window, observations can only be added
+ # never removed
+ if i == 0:
- # After the first window, observations can both be added and
- # removed
- for i in range(win, N):
- val = values[i]
- prev = values[i - win]
+ for j in range(s, e):
+ add_var(values[j], &nobs, &mean_x, &ssqdm_x)
- if notnan(val):
- if prev == prev:
+ else:
- # Adding one observation and removing another one
- delta = val - prev
- mean_x_old = mean_x
+ # After the first window, observations can both be added
+ # and removed
- mean_x += delta / nobs
- ssqdm_x += ((nobs - 1) * val
- + (nobs + 1) * prev
- - 2 * nobs * mean_x_old) * delta / nobs
+ # calculate adds
+ for j in range(end[i - 1], e):
+ add_var(values[j], &nobs, &mean_x, &ssqdm_x)
- else:
- add_var(val, &nobs, &mean_x, &ssqdm_x)
- elif prev == prev:
- remove_var(prev, &nobs, &mean_x, &ssqdm_x)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ remove_var(values[j], &nobs, &mean_x, &ssqdm_x)
- output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
+ output[i] = calc_var(minp, ddof, nobs, ssqdm_x)
return output
-
# ----------------------------------------------------------------------
# Rolling skewness
+
cdef inline float64_t calc_skew(int64_t minp, int64_t nobs,
float64_t x, float64_t xx,
float64_t xxx) nogil:
@@ -861,76 +598,80 @@ cdef inline void remove_skew(float64_t val, int64_t *nobs,
xxx[0] = xxx[0] - val * val * val
-def roll_skew(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_skew_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
float64_t val, prev
float64_t x = 0, xx = 0, xxx = 0
- int64_t nobs = 0, i, j, N
+ int64_t nobs = 0, i, j, N = len(values)
int64_t s, e
- bint is_variable
- int64_t[:] start, end
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
- if is_variable:
+ with nogil:
+ for i in range(minp - 1):
+ val = values[i]
+ add_skew(val, &nobs, &x, &xx, &xxx)
+ output[i] = NaN
- with nogil:
+ for i in range(minp - 1, N):
+ val = values[i]
+ add_skew(val, &nobs, &x, &xx, &xxx)
- for i in range(0, N):
+ if i > win - 1:
+ prev = values[i - win]
+ remove_skew(prev, &nobs, &x, &xx, &xxx)
- s = start[i]
- e = end[i]
+ output[i] = calc_skew(minp, nobs, x, xx, xxx)
- # Over the first window, observations can only be added
- # never removed
- if i == 0:
+ return output
- for j in range(s, e):
- val = values[j]
- add_skew(val, &nobs, &x, &xx, &xxx)
- else:
+def roll_skew_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ cdef:
+ float64_t val, prev
+ float64_t x = 0, xx = 0, xxx = 0
+ int64_t nobs = 0, i, j, N = len(values)
+ int64_t s, e
+ ndarray[float64_t] output
+
+ output = np.empty(N, dtype=float)
- # After the first window, observations can both be added
- # and removed
+ with nogil:
- # calculate adds
- for j in range(end[i - 1], e):
- val = values[j]
- add_skew(val, &nobs, &x, &xx, &xxx)
+ for i in range(0, N):
- # calculate deletes
- for j in range(start[i - 1], s):
- val = values[j]
- remove_skew(val, &nobs, &x, &xx, &xxx)
+ s = start[i]
+ e = end[i]
- output[i] = calc_skew(minp, nobs, x, xx, xxx)
+ # Over the first window, observations can only be added
+ # never removed
+ if i == 0:
- else:
+ for j in range(s, e):
+ val = values[j]
+ add_skew(val, &nobs, &x, &xx, &xxx)
- with nogil:
- for i in range(minp - 1):
- val = values[i]
- add_skew(val, &nobs, &x, &xx, &xxx)
- output[i] = NaN
+ else:
- for i in range(minp - 1, N):
- val = values[i]
- add_skew(val, &nobs, &x, &xx, &xxx)
+ # After the first window, observations can both be added
+ # and removed
- if i > win - 1:
- prev = values[i - win]
- remove_skew(prev, &nobs, &x, &xx, &xxx)
+ # calculate adds
+ for j in range(end[i - 1], e):
+ val = values[j]
+ add_skew(val, &nobs, &x, &xx, &xxx)
- output[i] = calc_skew(minp, nobs, x, xx, xxx)
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ val = values[j]
+ remove_skew(val, &nobs, &x, &xx, &xxx)
- return output
+ output[i] = calc_skew(minp, nobs, x, xx, xxx)
+ return output
# ----------------------------------------------------------------------
# Rolling kurtosis
@@ -1005,69 +746,73 @@ cdef inline void remove_kurt(float64_t val, int64_t *nobs,
xxxx[0] = xxxx[0] - val * val * val * val
-def roll_kurt(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_kurt_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
float64_t val, prev
float64_t x = 0, xx = 0, xxx = 0, xxxx = 0
- int64_t nobs = 0, i, j, N
+ int64_t nobs = 0, i, j, N = len(values)
int64_t s, e
- bint is_variable
- int64_t[:] start, end
ndarray[float64_t] output
- start, end, N, win, minp, is_variable = get_window_indexer(values, win,
- minp, index,
- closed)
output = np.empty(N, dtype=float)
- if is_variable:
+ with nogil:
- with nogil:
+ for i in range(minp - 1):
+ add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
+ output[i] = NaN
- for i in range(0, N):
+ for i in range(minp - 1, N):
+ add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
- s = start[i]
- e = end[i]
+ if i > win - 1:
+ prev = values[i - win]
+ remove_kurt(prev, &nobs, &x, &xx, &xxx, &xxxx)
- # Over the first window, observations can only be added
- # never removed
- if i == 0:
+ output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
- for j in range(s, e):
- add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+ return output
- else:
- # After the first window, observations can both be added
- # and removed
+def roll_kurt_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ cdef:
+ float64_t val, prev
+ float64_t x = 0, xx = 0, xxx = 0, xxxx = 0
+ int64_t nobs = 0, i, j, s, e, N = len(values)
+ ndarray[float64_t] output
- # calculate adds
- for j in range(end[i - 1], e):
- add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+ output = np.empty(N, dtype=float)
- # calculate deletes
- for j in range(start[i - 1], s):
- remove_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+ with nogil:
- output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
+ for i in range(0, N):
- else:
+ s = start[i]
+ e = end[i]
- with nogil:
+ # Over the first window, observations can only be added
+ # never removed
+ if i == 0:
- for i in range(minp - 1):
- add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
- output[i] = NaN
+ for j in range(s, e):
+ add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
- for i in range(minp - 1, N):
- add_kurt(values[i], &nobs, &x, &xx, &xxx, &xxxx)
+ else:
- if i > win - 1:
- prev = values[i - win]
- remove_kurt(prev, &nobs, &x, &xx, &xxx, &xxxx)
+ # After the first window, observations can both be added
+ # and removed
- output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
+ # calculate adds
+ for j in range(end[i - 1], e):
+ add_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+
+ # calculate deletes
+ for j in range(start[i - 1], s):
+ remove_kurt(values[j], &nobs, &x, &xx, &xxx, &xxxx)
+
+ output[i] = calc_kurt(minp, nobs, x, xx, xxx, xxxx)
return output
@@ -1076,31 +821,26 @@ def roll_kurt(ndarray[float64_t] values, int64_t win, int64_t minp,
# Rolling median, min, max
-def roll_median_c(ndarray[float64_t] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_median_c(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
cdef:
float64_t val, res, prev
- bint err = 0, is_variable
+ bint err = 0
int ret = 0
skiplist_t *sl
Py_ssize_t i, j
- int64_t nobs = 0, N, s, e
+ int64_t nobs = 0, N = len(values), s, e
int midpoint
- int64_t[:] start, end
ndarray[float64_t] output
# we use the Fixed/Variable Indexer here as the
# actual skiplist ops outweigh any window computation costs
- start, end, N, win, minp, is_variable = get_window_indexer(
- values, win,
- minp, index, closed,
- use_mock=False)
output = np.empty(N, dtype=float)
- if win == 0:
+ if win == 0 or (end - start).max() == 0:
output[:] = NaN
return output
-
+ win = (end - start).max()
sl = skiplist_init(<int>win)
if sl == NULL:
raise MemoryError("skiplist_init failed")
@@ -1209,76 +949,89 @@ cdef inline numeric calc_mm(int64_t minp, Py_ssize_t nobs,
return result
-def roll_max(ndarray[numeric] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_max_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
"""
Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
Parameters
----------
- values: numpy array
- window: int, size of rolling window
- minp: if number of observations in window
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
is below this, output a NaN
- index: ndarray, optional
+ index : ndarray, optional
index for window computation
- closed: 'right', 'left', 'both', 'neither'
+ closed : 'right', 'left', 'both', 'neither'
make the interval closed on the right, left,
both or neither endpoints
"""
- return _roll_min_max(values, win, minp, index, closed=closed, is_max=1)
+ return _roll_min_max_fixed(values, start, end, minp, win, is_max=1)
-def roll_min(ndarray[numeric] values, int64_t win, int64_t minp,
- object index, object closed):
+def roll_max_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
"""
Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
Parameters
----------
- values: numpy array
- window: int, size of rolling window
- minp: if number of observations in window
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
is below this, output a NaN
- index: ndarray, optional
+ index : ndarray, optional
index for window computation
+ closed : 'right', 'left', 'both', 'neither'
+ make the interval closed on the right, left,
+ both or neither endpoints
"""
- return _roll_min_max(values, win, minp, index, is_max=0, closed=closed)
+ return _roll_min_max_variable(values, start, end, minp, is_max=1)
-cdef _roll_min_max(ndarray[numeric] values, int64_t win, int64_t minp,
- object index, object closed, bint is_max):
+def roll_min_fixed(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win):
"""
- Moving min/max of 1d array of any numeric type along axis=0
- ignoring NaNs.
+ Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
+
+ Parameters
+ ----------
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
+ is below this, output a NaN
+ index : ndarray, optional
+ index for window computation
"""
- cdef:
- ndarray[int64_t] starti, endi
- int64_t N
- bint is_variable
+ return _roll_min_max_fixed(values, start, end, minp, win, is_max=0)
- starti, endi, N, win, minp, is_variable = get_window_indexer(
- values, win,
- minp, index, closed)
- if is_variable:
- return _roll_min_max_variable(values, starti, endi, N, win, minp,
- is_max)
- else:
- return _roll_min_max_fixed(values, N, win, minp, is_max)
+def roll_min_variable(ndarray[float64_t] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp):
+ """
+ Moving max of 1d array of any numeric type along axis=0 ignoring NaNs.
+
+ Parameters
+ ----------
+ values : np.ndarray[np.float64]
+ window : int, size of rolling window
+ minp : if number of observations in window
+ is below this, output a NaN
+ index : ndarray, optional
+ index for window computation
+ """
+ return _roll_min_max_variable(values, start, end, minp, is_max=0)
cdef _roll_min_max_variable(ndarray[numeric] values,
ndarray[int64_t] starti,
ndarray[int64_t] endi,
- int64_t N,
- int64_t win,
int64_t minp,
bint is_max):
cdef:
numeric ai
int64_t i, close_offset, curr_win_size
- Py_ssize_t nobs = 0
+ Py_ssize_t nobs = 0, N = len(values)
deque Q[int64_t] # min/max always the front
deque W[int64_t] # track the whole window for nobs compute
ndarray[float64_t, ndim=1] output
@@ -1353,15 +1106,16 @@ cdef _roll_min_max_variable(ndarray[numeric] values,
cdef _roll_min_max_fixed(ndarray[numeric] values,
- int64_t N,
- int64_t win,
+ ndarray[int64_t] starti,
+ ndarray[int64_t] endi,
int64_t minp,
+ int64_t win,
bint is_max):
cdef:
numeric ai
bint should_replace
int64_t i, removed, window_i,
- Py_ssize_t nobs = 0
+ Py_ssize_t nobs = 0, N = len(values)
int64_t* death
numeric* ring
numeric* minvalue
@@ -1457,8 +1211,8 @@ interpolation_types = {
}
-def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
- int64_t minp, object index, object closed,
+def roll_quantile(ndarray[float64_t, cast=True] values, ndarray[int64_t] start,
+ ndarray[int64_t] end, int64_t minp, int64_t win,
float64_t quantile, str interpolation):
"""
O(N log(window)) implementation using skip list
@@ -1466,10 +1220,8 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
cdef:
float64_t val, prev, midpoint, idx_with_fraction
skiplist_t *skiplist
- int64_t nobs = 0, i, j, s, e, N
+ int64_t nobs = 0, i, j, s, e, N = len(values)
Py_ssize_t idx
- bint is_variable
- int64_t[:] start, end
ndarray[float64_t] output
float64_t vlow, vhigh
InterpolationType interpolation_type
@@ -1485,16 +1237,12 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
# we use the Fixed/Variable Indexer here as the
# actual skiplist ops outweigh any window computation costs
- start, end, N, win, minp, is_variable = get_window_indexer(
- values, win,
- minp, index, closed,
- use_mock=False)
output = np.empty(N, dtype=float)
- if win == 0:
+ if win == 0 or (end - start).max() == 0:
output[:] = NaN
return output
-
+ win = (end - start).max()
skiplist = skiplist_init(<int>win)
if skiplist == NULL:
raise MemoryError("skiplist_init failed")
@@ -1575,18 +1323,17 @@ def roll_quantile(ndarray[float64_t, cast=True] values, int64_t win,
return output
-def roll_generic(object obj,
- int64_t win, int64_t minp, object index, object closed,
- int offset, object func, bint raw,
- object args, object kwargs):
+def roll_generic_fixed(object obj,
+ ndarray[int64_t] start, ndarray[int64_t] end,
+ int64_t minp, int64_t win,
+ int offset, object func, bint raw,
+ object args, object kwargs):
cdef:
ndarray[float64_t] output, counts, bufarr
ndarray[float64_t, cast=True] arr
float64_t *buf
float64_t *oldbuf
- int64_t nobs = 0, i, j, s, e, N
- bint is_variable
- int64_t[:] start, end
+ int64_t nobs = 0, i, j, s, e, N = len(start)
n = len(obj)
if n == 0:
@@ -1599,36 +1346,13 @@ def roll_generic(object obj,
if not arr.flags.c_contiguous:
arr = arr.copy('C')
- counts = roll_sum(np.concatenate([np.isfinite(arr).astype(float),
- np.array([0.] * offset)]),
- win, minp, index, closed)[offset:]
-
- start, end, N, win, minp, is_variable = get_window_indexer(arr, win,
- minp, index,
- closed,
- floor=0)
+ counts = roll_sum_fixed(np.concatenate([np.isfinite(arr).astype(float),
+ np.array([0.] * offset)]),
+ start, end, minp, win)[offset:]
output = np.empty(N, dtype=float)
- if is_variable:
- # variable window arr or series
-
- if offset != 0:
- raise ValueError("unable to roll_generic with a non-zero offset")
-
- for i in range(0, N):
- s = start[i]
- e = end[i]
-
- if counts[i] >= minp:
- if raw:
- output[i] = func(arr[s:e], *args, **kwargs)
- else:
- output[i] = func(obj.iloc[s:e], *args, **kwargs)
- else:
- output[i] = NaN
-
- elif not raw:
+ if not raw:
# series
for i in range(N):
if counts[i] >= minp:
@@ -1672,6 +1396,53 @@ def roll_generic(object obj,
return output
+def roll_generic_variable(object obj,
+ ndarray[int64_t] start, ndarray[int64_t] end,
+ int64_t minp,
+ int offset, object func, bint raw,
+ object args, object kwargs):
+ cdef:
+ ndarray[float64_t] output, counts, bufarr
+ ndarray[float64_t, cast=True] arr
+ float64_t *buf
+ float64_t *oldbuf
+ int64_t nobs = 0, i, j, s, e, N = len(start)
+
+ n = len(obj)
+ if n == 0:
+ return obj
+
+ arr = np.asarray(obj)
+
+ # ndarray input
+ if raw:
+ if not arr.flags.c_contiguous:
+ arr = arr.copy('C')
+
+ counts = roll_sum_variable(np.concatenate([np.isfinite(arr).astype(float),
+ np.array([0.] * offset)]),
+ start, end, minp)[offset:]
+
+ output = np.empty(N, dtype=float)
+
+ if offset != 0:
+ raise ValueError("unable to roll_generic with a non-zero offset")
+
+ for i in range(0, N):
+ s = start[i]
+ e = end[i]
+
+ if counts[i] >= minp:
+ if raw:
+ output[i] = func(arr[s:e], *args, **kwargs)
+ else:
+ output[i] = func(obj.iloc[s:e], *args, **kwargs)
+ else:
+ output[i] = NaN
+
+ return output
+
+
# ----------------------------------------------------------------------
# Rolling sum and mean for weighted window
diff --git a/pandas/_libs/window_indexer.pyx b/pandas/_libs/window_indexer.pyx
new file mode 100644
index 0000000000000..8f49a8b9462d3
--- /dev/null
+++ b/pandas/_libs/window_indexer.pyx
@@ -0,0 +1,165 @@
+# cython: boundscheck=False, wraparound=False, cdivision=True
+
+import numpy as np
+from numpy cimport ndarray, int64_t
+
+# ----------------------------------------------------------------------
+# The indexer objects for rolling
+# These define start/end indexers to compute offsets
+
+
+class MockFixedWindowIndexer:
+ """
+
+ We are just checking parameters of the indexer,
+ and returning a consistent API with fixed/variable
+ indexers.
+
+ Parameters
+ ----------
+ values: ndarray
+ values data array
+ win: int64_t
+ window size
+ index: object
+ index of the values
+ closed: string
+ closed behavior
+ """
+ def __init__(self, ndarray values, int64_t win, object closed, object index=None):
+
+ self.start = np.empty(0, dtype='int64')
+ self.end = np.empty(0, dtype='int64')
+
+ def get_window_bounds(self):
+ return self.start, self.end
+
+
+class FixedWindowIndexer:
+ """
+ create a fixed length window indexer object
+ that has start & end, that point to offsets in
+ the index object; these are defined based on the win
+ arguments
+
+ Parameters
+ ----------
+ values: ndarray
+ values data array
+ win: int64_t
+ window size
+ index: object
+ index of the values
+ closed: string
+ closed behavior
+ """
+ def __init__(self, ndarray values, int64_t win, object closed, object index=None):
+ cdef:
+ ndarray[int64_t, ndim=1] start_s, start_e, end_s, end_e
+ int64_t N = len(values)
+
+ start_s = np.zeros(win, dtype='int64')
+ start_e = np.arange(win, N, dtype='int64') - win + 1
+ self.start = np.concatenate([start_s, start_e])[:N]
+
+ end_s = np.arange(win, dtype='int64') + 1
+ end_e = start_e + win
+ self.end = np.concatenate([end_s, end_e])[:N]
+
+ def get_window_bounds(self):
+ return self.start, self.end
+
+
+class VariableWindowIndexer:
+ """
+ create a variable length window indexer object
+ that has start & end, that point to offsets in
+ the index object; these are defined based on the win
+ arguments
+
+ Parameters
+ ----------
+ values: ndarray
+ values data array
+ win: int64_t
+ window size
+ index: ndarray
+ index of the values
+ closed: string
+ closed behavior
+ """
+ def __init__(self, ndarray values, int64_t win, object closed, ndarray index):
+ cdef:
+ bint left_closed = False
+ bint right_closed = False
+ int64_t N = len(index)
+
+ # if windows is variable, default is 'right', otherwise default is 'both'
+ if closed is None:
+ closed = 'right' if index is not None else 'both'
+
+ if closed in ['right', 'both']:
+ right_closed = True
+
+ if closed in ['left', 'both']:
+ left_closed = True
+
+ self.start, self.end = self.build(index, win, left_closed, right_closed, N)
+
+ @staticmethod
+ def build(const int64_t[:] index, int64_t win, bint left_closed,
+ bint right_closed, int64_t N):
+
+ cdef:
+ ndarray[int64_t] start, end
+ int64_t start_bound, end_bound
+ Py_ssize_t i, j
+
+ start = np.empty(N, dtype='int64')
+ start.fill(-1)
+ end = np.empty(N, dtype='int64')
+ end.fill(-1)
+
+ start[0] = 0
+
+ # right endpoint is closed
+ if right_closed:
+ end[0] = 1
+ # right endpoint is open
+ else:
+ end[0] = 0
+
+ with nogil:
+
+ # start is start of slice interval (including)
+ # end is end of slice interval (not including)
+ for i in range(1, N):
+ end_bound = index[i]
+ start_bound = index[i] - win
+
+ # left endpoint is closed
+ if left_closed:
+ start_bound -= 1
+
+ # advance the start bound until we are
+ # within the constraint
+ start[i] = i
+ for j in range(start[i - 1], i):
+ if index[j] > start_bound:
+ start[i] = j
+ break
+
+ # end bound is previous end
+ # or current index
+ if index[end[i - 1]] <= end_bound:
+ end[i] = i + 1
+ else:
+ end[i] = end[i - 1]
+
+ # right endpoint is open
+ if not right_closed:
+ end[i] -= 1
+ return start, end
+
+ def get_window_bounds(self):
+ return self.start, self.end
diff --git a/pandas/core/window/common.py b/pandas/core/window/common.py
index 3fd567f97edae..453fd12495543 100644
--- a/pandas/core/window/common.py
+++ b/pandas/core/window/common.py
@@ -1,5 +1,6 @@
"""Common utility functions for rolling operations"""
from collections import defaultdict
+from typing import Callable, Optional
import warnings
import numpy as np
@@ -62,12 +63,20 @@ def __init__(self, obj, *args, **kwargs):
cov = _dispatch("cov", other=None, pairwise=None)
def _apply(
- self, func, name=None, window=None, center=None, check_minp=None, **kwargs
+ self,
+ func: Callable,
+ center: bool,
+ require_min_periods: int = 0,
+ floor: int = 1,
+ is_weighted: bool = False,
+ name: Optional[str] = None,
+ **kwargs,
):
"""
Dispatch to apply; we are stripping all of the _apply kwargs and
performing the original function call on the grouped object.
"""
+ kwargs.pop("floor", None)
# TODO: can we de-duplicate with _dispatch?
def f(x, name=name, *args):
@@ -267,6 +276,44 @@ def _use_window(minp, window):
return minp
+def calculate_min_periods(
+ window: int,
+ min_periods: Optional[int],
+ num_values: int,
+ required_min_periods: int,
+ floor: int,
+) -> int:
+ """
+ Calculates final minimum periods value for rolling aggregations.
+
+ Parameters
+ ----------
+ window : passed window value
+ min_periods : passed min periods value
+ num_values : total number of values
+ required_min_periods : required min periods per aggregation function
+ floor : required min periods per aggregation function
+
+ Returns
+ -------
+ min_periods : int
+ """
+ if min_periods is None:
+ min_periods = window
+ else:
+ min_periods = max(required_min_periods, min_periods)
+ if min_periods > window:
+ raise ValueError(
+ "min_periods {min_periods} must be <= "
+ "window {window}".format(min_periods=min_periods, window=window)
+ )
+ elif min_periods > num_values:
+ min_periods = num_values + 1
+ elif min_periods < 0:
+ raise ValueError("min_periods must be >= 0")
+ return max(min_periods, floor)
+
+
def _zsqrt(x):
with np.errstate(all="ignore"):
result = np.sqrt(x)
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index bec350f6b7d8b..fd2e8aa2ad02f 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -3,6 +3,7 @@
similar to how we have a Groupby object.
"""
from datetime import timedelta
+from functools import partial
from textwrap import dedent
from typing import Callable, Dict, List, Optional, Set, Tuple, Union
import warnings
@@ -10,6 +11,7 @@
import numpy as np
import pandas._libs.window as libwindow
+import pandas._libs.window_indexer as libwindow_indexer
from pandas.compat._optional import import_optional_dependency
from pandas.compat.numpy import function as nv
from pandas.util._decorators import Appender, Substitution, cache_readonly
@@ -43,10 +45,10 @@
_doc_template,
_flex_binary_moment,
_offset,
- _require_min_periods,
_shared_docs,
_use_window,
_zsqrt,
+ calculate_min_periods,
)
@@ -366,39 +368,55 @@ def _center_window(self, result, window) -> np.ndarray:
result = np.copy(result[tuple(lead_indexer)])
return result
- def _get_roll_func(
- self, cfunc: Callable, check_minp: Callable, index: np.ndarray, **kwargs
- ) -> Callable:
+ def _get_roll_func(self, func_name: str) -> Callable:
"""
Wrap rolling function to check values passed.
Parameters
----------
- cfunc : callable
+ func_name : str
Cython function used to calculate rolling statistics
- check_minp : callable
- function to check minimum period parameter
- index : ndarray
- used for variable window
Returns
-------
func : callable
"""
+ window_func = getattr(libwindow, func_name, None)
+ if window_func is None:
+ raise ValueError(
+ "we do not support this function "
+ "in libwindow.{func_name}".format(func_name=func_name)
+ )
+ return window_func
- def func(arg, window, min_periods=None, closed=None):
- minp = check_minp(min_periods, window)
- return cfunc(arg, window, minp, index, closed, **kwargs)
+ def _get_cython_func_type(self, func):
+ """
+ Return a variable or fixed cython function type.
- return func
+ Variable algorithms do not use window while fixed do.
+ """
+ if self.is_freq_type:
+ return self._get_roll_func("{}_variable".format(func))
+ return partial(
+ self._get_roll_func("{}_fixed".format(func)), win=self._get_window()
+ )
+
+ def _get_window_indexer(self):
+ """
+ Return an indexer class that will compute the window start and end bounds
+ """
+ if self.is_freq_type:
+ return libwindow_indexer.VariableWindowIndexer
+ return libwindow_indexer.FixedWindowIndexer
def _apply(
self,
- func: Union[str, Callable],
+ func: Callable,
+ center: bool,
+ require_min_periods: int = 0,
+ floor: int = 1,
+ is_weighted: bool = False,
name: Optional[str] = None,
- window: Optional[Union[int, str]] = None,
- center: Optional[bool] = None,
- check_minp: Optional[Callable] = None,
**kwargs,
):
"""
@@ -408,13 +426,13 @@ def _apply(
Parameters
----------
- func : str/callable to apply
- name : str, optional
- name of this function
- window : int/str, default to _get_window()
- window length or offset
- center : bool, default to self.center
- check_minp : function, default to _use_window
+ func : callable function to apply
+ center : bool
+ require_min_periods : int
+ floor: int
+ is_weighted
+ name: str,
+ compatibility with groupby.rolling
**kwargs
additional arguments for rolling function and window function
@@ -422,20 +440,13 @@ def _apply(
-------
y : type of input
"""
-
- if center is None:
- center = self.center
-
- if check_minp is None:
- check_minp = _use_window
-
- if window is None:
- win_type = self._get_win_type(kwargs)
- window = self._get_window(win_type=win_type)
+ win_type = self._get_win_type(kwargs)
+ window = self._get_window(win_type=win_type)
blocks, obj = self._create_blocks()
block_list = list(blocks)
index_as_array = self._get_index()
+ window_indexer = self._get_window_indexer()
results = []
exclude = [] # type: List[Scalar]
@@ -455,36 +466,27 @@ def _apply(
results.append(values.copy())
continue
- # if we have a string function name, wrap it
- if isinstance(func, str):
- cfunc = getattr(libwindow, func, None)
- if cfunc is None:
- raise ValueError(
- "we do not support this function "
- "in libwindow.{func}".format(func=func)
- )
-
- func = self._get_roll_func(cfunc, check_minp, index_as_array, **kwargs)
-
# calculation function
- if center:
- offset = _offset(window, center)
- additional_nans = np.array([np.NaN] * offset)
+ offset = _offset(window, center) if center else 0
+ additional_nans = np.array([np.nan] * offset)
+
+ if not is_weighted:
def calc(x):
- return func(
- np.concatenate((x, additional_nans)),
- window,
- min_periods=self.min_periods,
- closed=self.closed,
+ x = np.concatenate((x, additional_nans))
+ min_periods = calculate_min_periods(
+ window, self.min_periods, len(x), require_min_periods, floor
)
+ start, end = window_indexer(
+ x, window, self.closed, index_as_array
+ ).get_window_bounds()
+ return func(x, start, end, min_periods)
else:
def calc(x):
- return func(
- x, window, min_periods=self.min_periods, closed=self.closed
- )
+ x = np.concatenate((x, additional_nans))
+ return func(x, window, self.min_periods)
with np.errstate(all="ignore"):
if values.ndim > 1:
@@ -995,8 +997,8 @@ def _get_window(
# GH #15662. `False` makes symmetric window, rather than periodic.
return sig.get_window(win_type, window, False).astype(float)
- def _get_roll_func(
- self, cfunc: Callable, check_minp: Callable, index: np.ndarray, **kwargs
+ def _get_weighted_roll_func(
+ self, cfunc: Callable, check_minp: Callable, **kwargs
) -> Callable:
def func(arg, window, min_periods=None, closed=None):
minp = check_minp(min_periods, len(window))
@@ -1070,25 +1072,38 @@ def aggregate(self, func, *args, **kwargs):
@Appender(_shared_docs["sum"])
def sum(self, *args, **kwargs):
nv.validate_window_func("sum", args, kwargs)
- return self._apply("roll_weighted_sum", **kwargs)
+ window_func = self._get_roll_func("roll_weighted_sum")
+ window_func = self._get_weighted_roll_func(window_func, _use_window)
+ return self._apply(
+ window_func, center=self.center, is_weighted=True, name="sum", **kwargs
+ )
@Substitution(name="window")
@Appender(_shared_docs["mean"])
def mean(self, *args, **kwargs):
nv.validate_window_func("mean", args, kwargs)
- return self._apply("roll_weighted_mean", **kwargs)
+ window_func = self._get_roll_func("roll_weighted_mean")
+ window_func = self._get_weighted_roll_func(window_func, _use_window)
+ return self._apply(
+ window_func, center=self.center, is_weighted=True, name="mean", **kwargs
+ )
@Substitution(name="window", versionadded="\n.. versionadded:: 1.0.0\n")
@Appender(_shared_docs["var"])
def var(self, ddof=1, *args, **kwargs):
nv.validate_window_func("var", args, kwargs)
- return self._apply("roll_weighted_var", ddof=ddof, **kwargs)
+ window_func = partial(self._get_roll_func("roll_weighted_var"), ddof=ddof)
+ window_func = self._get_weighted_roll_func(window_func, _use_window)
+ kwargs.pop("name", None)
+ return self._apply(
+ window_func, center=self.center, is_weighted=True, name="var", **kwargs
+ )
@Substitution(name="window", versionadded="\n.. versionadded:: 1.0.0\n")
@Appender(_shared_docs["std"])
def std(self, ddof=1, *args, **kwargs):
nv.validate_window_func("std", args, kwargs)
- return _zsqrt(self.var(ddof=ddof, **kwargs))
+ return _zsqrt(self.var(ddof=ddof, name="std", **kwargs))
class _Rolling(_Window):
@@ -1203,9 +1218,9 @@ def apply(self, func, raw=None, args=(), kwargs={}):
from pandas import Series
kwargs.pop("_level", None)
+ kwargs.pop("floor", None)
window = self._get_window()
offset = _offset(window, self.center)
- index_as_array = self._get_index()
# TODO: default is for backward compat
# change to False in the future
@@ -1221,28 +1236,31 @@ def apply(self, func, raw=None, args=(), kwargs={}):
)
raw = True
- def f(arg, window, min_periods, closed):
- minp = _use_window(min_periods, window)
+ window_func = partial(
+ self._get_cython_func_type("roll_generic"),
+ args=args,
+ kwargs=kwargs,
+ raw=raw,
+ offset=offset,
+ func=func,
+ )
+
+ def apply_func(values, begin, end, min_periods, raw=raw):
if not raw:
- arg = Series(arg, index=self.obj.index)
- return libwindow.roll_generic(
- arg,
- window,
- minp,
- index_as_array,
- closed,
- offset,
- func,
- raw,
- args,
- kwargs,
- )
+ values = Series(values, index=self.obj.index)
+ return window_func(values, begin, end, min_periods)
- return self._apply(f, func, args=args, kwargs=kwargs, center=False, raw=raw)
+ # TODO: Why do we always pass center=False?
+ # name=func for WindowGroupByMixin._apply
+ return self._apply(apply_func, center=False, floor=0, name=func)
def sum(self, *args, **kwargs):
nv.validate_window_func("sum", args, kwargs)
- return self._apply("roll_sum", "sum", **kwargs)
+ window_func = self._get_cython_func_type("roll_sum")
+ kwargs.pop("floor", None)
+ return self._apply(
+ window_func, center=self.center, floor=0, name="sum", **kwargs
+ )
_shared_docs["max"] = dedent(
"""
@@ -1257,7 +1275,8 @@ def sum(self, *args, **kwargs):
def max(self, *args, **kwargs):
nv.validate_window_func("max", args, kwargs)
- return self._apply("roll_max", "max", **kwargs)
+ window_func = self._get_cython_func_type("roll_max")
+ return self._apply(window_func, center=self.center, name="max", **kwargs)
_shared_docs["min"] = dedent(
"""
@@ -1298,11 +1317,13 @@ def max(self, *args, **kwargs):
def min(self, *args, **kwargs):
nv.validate_window_func("min", args, kwargs)
- return self._apply("roll_min", "min", **kwargs)
+ window_func = self._get_cython_func_type("roll_min")
+ return self._apply(window_func, center=self.center, name="min", **kwargs)
def mean(self, *args, **kwargs):
nv.validate_window_func("mean", args, kwargs)
- return self._apply("roll_mean", "mean", **kwargs)
+ window_func = self._get_cython_func_type("roll_mean")
+ return self._apply(window_func, center=self.center, name="mean", **kwargs)
_shared_docs["median"] = dedent(
"""
@@ -1342,27 +1363,40 @@ def mean(self, *args, **kwargs):
)
def median(self, **kwargs):
- return self._apply("roll_median_c", "median", **kwargs)
+ window_func = self._get_roll_func("roll_median_c")
+ window_func = partial(window_func, win=self._get_window())
+ return self._apply(window_func, center=self.center, name="median", **kwargs)
def std(self, ddof=1, *args, **kwargs):
nv.validate_window_func("std", args, kwargs)
- window = self._get_window()
- index_as_array = self._get_index()
+ kwargs.pop("require_min_periods", None)
+ window_func = self._get_cython_func_type("roll_var")
- def f(arg, *args, **kwargs):
- minp = _require_min_periods(1)(self.min_periods, window)
- return _zsqrt(
- libwindow.roll_var(arg, window, minp, index_as_array, self.closed, ddof)
- )
+ def zsqrt_func(values, begin, end, min_periods):
+ return _zsqrt(window_func(values, begin, end, min_periods, ddof=ddof))
+ # ddof passed again for compat with groupby.rolling
return self._apply(
- f, "std", check_minp=_require_min_periods(1), ddof=ddof, **kwargs
+ zsqrt_func,
+ center=self.center,
+ require_min_periods=1,
+ name="std",
+ ddof=ddof,
+ **kwargs,
)
def var(self, ddof=1, *args, **kwargs):
nv.validate_window_func("var", args, kwargs)
+ kwargs.pop("require_min_periods", None)
+ window_func = partial(self._get_cython_func_type("roll_var"), ddof=ddof)
+ # ddof passed again for compat with groupby.rolling
return self._apply(
- "roll_var", "var", check_minp=_require_min_periods(1), ddof=ddof, **kwargs
+ window_func,
+ center=self.center,
+ require_min_periods=1,
+ name="var",
+ ddof=ddof,
+ **kwargs,
)
_shared_docs[
@@ -1377,8 +1411,14 @@ def var(self, ddof=1, *args, **kwargs):
"""
def skew(self, **kwargs):
+ window_func = self._get_cython_func_type("roll_skew")
+ kwargs.pop("require_min_periods", None)
return self._apply(
- "roll_skew", "skew", check_minp=_require_min_periods(3), **kwargs
+ window_func,
+ center=self.center,
+ require_min_periods=3,
+ name="skew",
+ **kwargs,
)
_shared_docs["kurt"] = dedent(
@@ -1414,8 +1454,14 @@ def skew(self, **kwargs):
)
def kurt(self, **kwargs):
+ window_func = self._get_cython_func_type("roll_kurt")
+ kwargs.pop("require_min_periods", None)
return self._apply(
- "roll_kurt", "kurt", check_minp=_require_min_periods(4), **kwargs
+ window_func,
+ center=self.center,
+ require_min_periods=4,
+ name="kurt",
+ **kwargs,
)
_shared_docs["quantile"] = dedent(
@@ -1475,33 +1521,22 @@ def kurt(self, **kwargs):
)
def quantile(self, quantile, interpolation="linear", **kwargs):
- window = self._get_window()
- index_as_array = self._get_index()
-
- def f(arg, *args, **kwargs):
- minp = _use_window(self.min_periods, window)
- if quantile == 1.0:
- return libwindow.roll_max(
- arg, window, minp, index_as_array, self.closed
- )
- elif quantile == 0.0:
- return libwindow.roll_min(
- arg, window, minp, index_as_array, self.closed
- )
- else:
- return libwindow.roll_quantile(
- arg,
- window,
- minp,
- index_as_array,
- self.closed,
- quantile,
- interpolation,
- )
+ if quantile == 1.0:
+ window_func = self._get_cython_func_type("roll_max")
+ elif quantile == 0.0:
+ window_func = self._get_cython_func_type("roll_min")
+ else:
+ window_func = partial(
+ self._get_roll_func("roll_quantile"),
+ win=self._get_window(),
+ quantile=quantile,
+ interpolation=interpolation,
+ )
- return self._apply(
- f, "quantile", quantile=quantile, interpolation=interpolation, **kwargs
- )
+ # Pass through for groupby.rolling
+ kwargs["quantile"] = quantile
+ kwargs["interpolation"] = interpolation
+ return self._apply(window_func, center=self.center, name="quantile", **kwargs)
_shared_docs[
"cov"
@@ -1856,7 +1891,8 @@ def count(self):
# different impl for freq counting
if self.is_freq_type:
- return self._apply("roll_count", "count")
+ window_func = self._get_roll_func("roll_count")
+ return self._apply(window_func, center=self.center, name="count")
return super().count()
diff --git a/setup.py b/setup.py
index 545765ecb114d..0915b6aba113a 100755
--- a/setup.py
+++ b/setup.py
@@ -344,6 +344,7 @@ class CheckSDist(sdist_class):
"pandas/_libs/tslibs/resolution.pyx",
"pandas/_libs/tslibs/parsing.pyx",
"pandas/_libs/tslibs/tzconversion.pyx",
+ "pandas/_libs/window_indexer.pyx",
"pandas/_libs/writers.pyx",
"pandas/io/sas/sas.pyx",
]
@@ -683,6 +684,7 @@ def srcpath(name=None, suffix=".pyx", subdir="src"):
},
"_libs.testing": {"pyxfile": "_libs/testing"},
"_libs.window": {"pyxfile": "_libs/window", "language": "c++", "suffix": ".cpp"},
+ "_libs.window_indexer": {"pyxfile": "_libs/window_indexer"},
"_libs.writers": {"pyxfile": "_libs/writers"},
"io.sas._sas": {"pyxfile": "io/sas/sas"},
"io.msgpack._packer": {
| - [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Pre-req for https://github.com/pandas-dev/pandas/issues/28987
Currently many of the aggregation functions in `window.pyx` follow the form:
```
def roll_func(values, window, minp, N, closed):
# calculate window bounds _and_ validate arguments
start, end, ... = get_window_bounds(values, window, minp, N, ...)
for i in range(values):
s = start[i]
....
```
This PR refactors out the window bound calculation into `window_indexer.pyx` and validation so the aggregation functions can be of the form:
```
def roll_func(values, start, end, minp):
for i in range(values):
s = start[i]
....
```
The methods therefore in `rolling.py` now have the following pattern:
1. Fetch the correct cython aggregation function (whether the window is fixed or variable), and prep it with kwargs if needed
2. Compute the `start` and `end` window bounds from functionality in `window_indexer.pyx`
3. Pass in the `values`, `start`, `end`, `min periods` into the aggregation function. | https://api.github.com/repos/pandas-dev/pandas/pulls/29428 | 2019-11-06T07:30:05Z | 2019-11-21T12:59:31Z | 2019-11-21T12:59:31Z | 2019-11-26T14:22:18Z |
REF: separate out ShallowMixin | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 1a2f906f97152..65e531f96614a 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -4,7 +4,7 @@
import builtins
from collections import OrderedDict
import textwrap
-from typing import Dict, FrozenSet, Optional
+from typing import Dict, FrozenSet, List, Optional
import warnings
import numpy as np
@@ -569,7 +569,7 @@ def _aggregate_multiple_funcs(self, arg, _level, _axis):
try:
new_res = colg.aggregate(a)
- except (TypeError, DataError):
+ except TypeError:
pass
else:
results.append(new_res)
@@ -618,6 +618,23 @@ def _aggregate_multiple_funcs(self, arg, _level, _axis):
raise ValueError("cannot combine transform and aggregation operations")
return result
+ def _get_cython_func(self, arg: str) -> Optional[str]:
+ """
+ if we define an internal function for this argument, return it
+ """
+ return self._cython_table.get(arg)
+
+ def _is_builtin_func(self, arg):
+ """
+ if we define an builtin function for this argument, return it,
+ otherwise return the arg
+ """
+ return self._builtin_table.get(arg, arg)
+
+
+class ShallowMixin:
+ _attributes = [] # type: List[str]
+
def _shallow_copy(self, obj=None, obj_type=None, **kwargs):
"""
return a new object with the replacement attributes
@@ -633,19 +650,6 @@ def _shallow_copy(self, obj=None, obj_type=None, **kwargs):
kwargs[attr] = getattr(self, attr)
return obj_type(obj, **kwargs)
- def _get_cython_func(self, arg: str) -> Optional[str]:
- """
- if we define an internal function for this argument, return it
- """
- return self._cython_table.get(arg)
-
- def _is_builtin_func(self, arg):
- """
- if we define an builtin function for this argument, return it,
- otherwise return the arg
- """
- return self._builtin_table.get(arg, arg)
-
class IndexOpsMixin:
"""
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index e68a2efc3f4e6..9d7ddcf3c7727 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -17,7 +17,7 @@
from pandas.core.dtypes.generic import ABCDataFrame, ABCSeries
import pandas.core.algorithms as algos
-from pandas.core.base import DataError
+from pandas.core.base import DataError, ShallowMixin
from pandas.core.generic import _shared_docs
from pandas.core.groupby.base import GroupByMixin
from pandas.core.groupby.generic import SeriesGroupBy
@@ -34,7 +34,7 @@
_shared_docs_kwargs = dict() # type: Dict[str, str]
-class Resampler(_GroupBy):
+class Resampler(_GroupBy, ShallowMixin):
"""
Class for resampling datetimelike data, a groupby-like operation.
See aggregate, transform, and apply functions on this object.
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index 68eb1f630bfc3..0718acd6360bf 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -35,7 +35,7 @@
)
from pandas._typing import Axis, FrameOrSeries, Scalar
-from pandas.core.base import DataError, PandasObject, SelectionMixin
+from pandas.core.base import DataError, PandasObject, SelectionMixin, ShallowMixin
import pandas.core.common as com
from pandas.core.index import Index, ensure_index
from pandas.core.window.common import (
@@ -50,7 +50,7 @@
)
-class _Window(PandasObject, SelectionMixin):
+class _Window(PandasObject, ShallowMixin, SelectionMixin):
_attributes = [
"window",
"min_periods",
| - [x] closes #28938
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Stops catching DataError in the 1D case for _aggregate_multiple_funcs. This change is mostly unrelated, but shares the process of reasoning about what cases need _shallow_copy/DataError. | https://api.github.com/repos/pandas-dev/pandas/pulls/29427 | 2019-11-06T03:11:19Z | 2019-11-06T19:11:31Z | 2019-11-06T19:11:30Z | 2019-11-06T19:24:46Z |
BUG: fix TypeErrors raised within _python_agg_general | diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 873a31e658625..fa4a184e8f7a4 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -899,10 +899,21 @@ def _python_agg_general(self, func, *args, **kwargs):
output = {}
for name, obj in self._iterate_slices():
try:
- result, counts = self.grouper.agg_series(obj, f)
+ # if this function is invalid for this dtype, we will ignore it.
+ func(obj[:0])
except TypeError:
continue
- else:
+ except AssertionError:
+ raise
+ except Exception:
+ # Our function depends on having a non-empty argument
+ # See test_groupby_agg_err_catching
+ pass
+
+ result, counts = self.grouper.agg_series(obj, f)
+ if result is not None:
+ # TODO: only 3 test cases get None here, do something
+ # in those cases
output[name] = self._try_cast(result, obj, numeric_only=True)
if len(output) == 0:
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 2cc0e5fde2290..5bad73bf40ff5 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -61,8 +61,7 @@ class BaseGrouper:
Parameters
----------
- axis : int
- the axis to group
+ axis : Index
groupings : array of grouping
all the grouping instances to handle in this grouper
for example for grouper list to groupby, need to pass the list
@@ -78,8 +77,15 @@ class BaseGrouper:
"""
def __init__(
- self, axis, groupings, sort=True, group_keys=True, mutated=False, indexer=None
+ self,
+ axis: Index,
+ groupings,
+ sort=True,
+ group_keys=True,
+ mutated=False,
+ indexer=None,
):
+ assert isinstance(axis, Index), axis
self._filter_empty_groups = self.compressed = len(groupings) != 1
self.axis = axis
self.groupings = groupings
@@ -623,7 +629,7 @@ def _aggregate_series_pure_python(self, obj, func):
counts = np.zeros(ngroups, dtype=int)
result = None
- splitter = get_splitter(obj, group_index, ngroups, axis=self.axis)
+ splitter = get_splitter(obj, group_index, ngroups, axis=0)
for label, group in splitter:
res = func(group)
@@ -635,8 +641,12 @@ def _aggregate_series_pure_python(self, obj, func):
counts[label] = group.shape[0]
result[label] = res
- result = lib.maybe_convert_objects(result, try_float=0)
- # TODO: try_cast back to EA?
+ if result is not None:
+ # if splitter is empty, result can be None, in which case
+ # maybe_convert_objects would raise TypeError
+ result = lib.maybe_convert_objects(result, try_float=0)
+ # TODO: try_cast back to EA?
+
return result, counts
@@ -781,6 +791,11 @@ def groupings(self):
]
def agg_series(self, obj: Series, func):
+ if is_extension_array_dtype(obj.dtype):
+ # pre-empty SeriesBinGrouper from raising TypeError
+ # TODO: watch out, this can return None
+ return self._aggregate_series_pure_python(obj, func)
+
dummy = obj[:0]
grouper = libreduction.SeriesBinGrouper(obj, func, self.bins, dummy)
return grouper.get_result()
@@ -809,12 +824,13 @@ def _is_indexed_like(obj, axes) -> bool:
class DataSplitter:
- def __init__(self, data, labels, ngroups, axis=0):
+ def __init__(self, data, labels, ngroups, axis: int = 0):
self.data = data
self.labels = ensure_int64(labels)
self.ngroups = ngroups
self.axis = axis
+ assert isinstance(axis, int), axis
@cache_readonly
def slabels(self):
@@ -837,12 +853,6 @@ def __iter__(self):
starts, ends = lib.generate_slices(self.slabels, self.ngroups)
for i, (start, end) in enumerate(zip(starts, ends)):
- # Since I'm now compressing the group ids, it's now not "possible"
- # to produce empty slices because such groups would not be observed
- # in the data
- # if start >= end:
- # raise AssertionError('Start %s must be less than end %s'
- # % (str(start), str(end)))
yield i, self._chop(sdata, slice(start, end))
def _get_sorted_data(self):
diff --git a/pandas/tests/groupby/aggregate/test_other.py b/pandas/tests/groupby/aggregate/test_other.py
index 5dad868c8c3aa..1c297f3e2ada3 100644
--- a/pandas/tests/groupby/aggregate/test_other.py
+++ b/pandas/tests/groupby/aggregate/test_other.py
@@ -602,3 +602,41 @@ def test_agg_lambda_with_timezone():
columns=["date"],
)
tm.assert_frame_equal(result, expected)
+
+
+@pytest.mark.parametrize(
+ "err_cls",
+ [
+ NotImplementedError,
+ RuntimeError,
+ KeyError,
+ IndexError,
+ OSError,
+ ValueError,
+ ArithmeticError,
+ AttributeError,
+ ],
+)
+def test_groupby_agg_err_catching(err_cls):
+ # make sure we suppress anything other than TypeError or AssertionError
+ # in _python_agg_general
+
+ # Use a non-standard EA to make sure we don't go down ndarray paths
+ from pandas.tests.extension.decimal.array import DecimalArray, make_data, to_decimal
+
+ data = make_data()[:5]
+ df = pd.DataFrame(
+ {"id1": [0, 0, 0, 1, 1], "id2": [0, 1, 0, 1, 1], "decimals": DecimalArray(data)}
+ )
+
+ expected = pd.Series(to_decimal([data[0], data[3]]))
+
+ def weird_func(x):
+ # weird function that raise something other than TypeError or IndexError
+ # in _python_agg_general
+ if len(x) == 0:
+ raise err_cls
+ return x.iloc[0]
+
+ result = df["decimals"].groupby(df["id1"]).agg(weird_func)
+ tm.assert_series_equal(result, expected, check_names=False)
| cc @jreback @WillAyd
There are a few ways in which we incorrectly raise TypeError within _python_agg_general that this fixes.
A lot of the complexity in this code comes from the fact that we drop columns on which a function is invalid instead of requiring the user to subset columns. | https://api.github.com/repos/pandas-dev/pandas/pulls/29425 | 2019-11-06T01:24:04Z | 2019-11-06T21:25:07Z | 2019-11-06T21:25:07Z | 2019-11-06T21:33:09Z |
TST: consistent result in dropping NA from CSV | diff --git a/pandas/tests/io/parser/test_na_values.py b/pandas/tests/io/parser/test_na_values.py
index f154d09358dc1..f52c6b8858fd3 100644
--- a/pandas/tests/io/parser/test_na_values.py
+++ b/pandas/tests/io/parser/test_na_values.py
@@ -536,3 +536,31 @@ def test_cast_NA_to_bool_raises_error(all_parsers, data, na_values):
dtype={"a": "bool"},
na_values=na_values,
)
+
+
+def test_str_nan_dropped(all_parsers):
+ # see gh-21131
+ parser = all_parsers
+
+ data = """File: small.csv,,
+10010010233,0123,654
+foo,,bar
+01001000155,4530,898"""
+
+ result = parser.read_csv(
+ StringIO(data),
+ header=None,
+ names=["col1", "col2", "col3"],
+ dtype={"col1": str, "col2": str, "col3": str},
+ ).dropna()
+
+ expected = DataFrame(
+ {
+ "col1": ["10010010233", "01001000155"],
+ "col2": ["0123", "4530"],
+ "col3": ["654", "898"],
+ },
+ index=[1, 3],
+ )
+
+ tm.assert_frame_equal(result, expected)
| Closes https://github.com/pandas-dev/pandas/issues/21131 | https://api.github.com/repos/pandas-dev/pandas/pulls/29424 | 2019-11-05T23:10:16Z | 2019-11-06T21:11:44Z | 2019-11-06T21:11:44Z | 2019-11-06T21:43:39Z |
TST: Test nLargest with MI grouper | diff --git a/pandas/tests/groupby/test_function.py b/pandas/tests/groupby/test_function.py
index 2d7dfe49dc038..18c4d7ceddc65 100644
--- a/pandas/tests/groupby/test_function.py
+++ b/pandas/tests/groupby/test_function.py
@@ -607,6 +607,51 @@ def test_nlargest():
tm.assert_series_equal(gb.nlargest(3, keep="last"), e)
+def test_nlargest_mi_grouper():
+ # see gh-21411
+ npr = np.random.RandomState(123456789)
+
+ dts = date_range("20180101", periods=10)
+ iterables = [dts, ["one", "two"]]
+
+ idx = MultiIndex.from_product(iterables, names=["first", "second"])
+ s = Series(npr.randn(20), index=idx)
+
+ result = s.groupby("first").nlargest(1)
+
+ exp_idx = MultiIndex.from_tuples(
+ [
+ (dts[0], dts[0], "one"),
+ (dts[1], dts[1], "one"),
+ (dts[2], dts[2], "one"),
+ (dts[3], dts[3], "two"),
+ (dts[4], dts[4], "one"),
+ (dts[5], dts[5], "one"),
+ (dts[6], dts[6], "one"),
+ (dts[7], dts[7], "one"),
+ (dts[8], dts[8], "two"),
+ (dts[9], dts[9], "one"),
+ ],
+ names=["first", "first", "second"],
+ )
+
+ exp_values = [
+ 2.2129019979039612,
+ 1.8417114045748335,
+ 0.858963679564603,
+ 1.3759151378258088,
+ 0.9430284594687134,
+ 0.5296914208183142,
+ 0.8318045593815487,
+ -0.8476703342910327,
+ 0.3804446884133735,
+ -0.8028845810770998,
+ ]
+
+ expected = Series(exp_values, index=exp_idx)
+ tm.assert_series_equal(result, expected, check_exact=False, check_less_precise=True)
+
+
def test_nsmallest():
a = Series([1, 3, 5, 7, 2, 9, 0, 4, 6, 10])
b = Series(list("a" * 5 + "b" * 5))
| Closes https://github.com/pandas-dev/pandas/issues/21411 | https://api.github.com/repos/pandas-dev/pandas/pulls/29423 | 2019-11-05T22:36:02Z | 2019-11-06T19:10:35Z | 2019-11-06T19:10:35Z | 2019-11-06T19:28:04Z |
TST: ignore _version.py | diff --git a/setup.cfg b/setup.cfg
index d4657100c1291..c7a71222ac91f 100644
--- a/setup.cfg
+++ b/setup.cfg
@@ -80,6 +80,7 @@ branch = False
omit =
*/tests/*
pandas/_typing.py
+ pandas/_version.py
plugins = Cython.Coverage
[coverage:report]
| - [x] closes #26877
The file is auto-generated, not something for us to worry about | https://api.github.com/repos/pandas-dev/pandas/pulls/29421 | 2019-11-05T22:14:11Z | 2019-11-06T19:27:15Z | 2019-11-06T19:27:15Z | 2019-11-06T20:03:02Z |
Correct type inference for UInt64Index during access | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 30a828064f812..950b8db373eef 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -345,7 +345,8 @@ Numeric
- Improved error message when using `frac` > 1 and `replace` = False (:issue:`27451`)
- Bug in numeric indexes resulted in it being possible to instantiate an :class:`Int64Index`, :class:`UInt64Index`, or :class:`Float64Index` with an invalid dtype (e.g. datetime-like) (:issue:`29539`)
- Bug in :class:`UInt64Index` precision loss while constructing from a list with values in the ``np.uint64`` range (:issue:`29526`)
--
+- Bug in :class:`NumericIndex` construction that caused indexing to fail when integers in the ``np.uint64`` range were used (:issue:`28023`)
+- Bug in :class:`NumericIndex` construction that caused :class:`UInt64Index` to be casted to :class:`Float64Index` when integers in the ``np.uint64`` range were used to index a :class:`DataFrame` (:issue:`28279`)
Conversion
^^^^^^^^^^
diff --git a/pandas/core/indexes/numeric.py b/pandas/core/indexes/numeric.py
index 29f56259dac79..747a9f75a3e00 100644
--- a/pandas/core/indexes/numeric.py
+++ b/pandas/core/indexes/numeric.py
@@ -2,7 +2,7 @@
import numpy as np
-from pandas._libs import index as libindex
+from pandas._libs import index as libindex, lib
from pandas.util._decorators import Appender, cache_readonly
from pandas.core.dtypes.cast import astype_nansafe
@@ -331,13 +331,15 @@ def _convert_scalar_indexer(self, key, kind=None):
@Appender(_index_shared_docs["_convert_arr_indexer"])
def _convert_arr_indexer(self, keyarr):
- # Cast the indexer to uint64 if possible so
- # that the values returned from indexing are
- # also uint64.
- keyarr = com.asarray_tuplesafe(keyarr)
- if is_integer_dtype(keyarr):
- return com.asarray_tuplesafe(keyarr, dtype=np.uint64)
- return keyarr
+ # Cast the indexer to uint64 if possible so that the values returned
+ # from indexing are also uint64.
+ dtype = None
+ if is_integer_dtype(keyarr) or (
+ lib.infer_dtype(keyarr, skipna=False) == "integer"
+ ):
+ dtype = np.uint64
+
+ return com.asarray_tuplesafe(keyarr, dtype=dtype)
@Appender(_index_shared_docs["_convert_index_indexer"])
def _convert_index_indexer(self, keyarr):
diff --git a/pandas/tests/indexes/test_numeric.py b/pandas/tests/indexes/test_numeric.py
index 6ee1ce5c4f2ad..37976d89ecba4 100644
--- a/pandas/tests/indexes/test_numeric.py
+++ b/pandas/tests/indexes/test_numeric.py
@@ -1209,3 +1209,29 @@ def test_range_float_union_dtype():
result = other.union(index)
tm.assert_index_equal(result, expected)
+
+
+def test_uint_index_does_not_convert_to_float64():
+ # https://github.com/pandas-dev/pandas/issues/28279
+ # https://github.com/pandas-dev/pandas/issues/28023
+ series = pd.Series(
+ [0, 1, 2, 3, 4, 5],
+ index=[
+ 7606741985629028552,
+ 17876870360202815256,
+ 17876870360202815256,
+ 13106359306506049338,
+ 8991270399732411471,
+ 8991270399732411472,
+ ],
+ )
+
+ result = series.loc[[7606741985629028552, 17876870360202815256]]
+
+ expected = UInt64Index(
+ [7606741985629028552, 17876870360202815256, 17876870360202815256],
+ dtype="uint64",
+ )
+ tm.assert_index_equal(result.index, expected)
+
+ tm.assert_equal(result, series[:3])
| - [x] closes #28023 and closes #28279
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29420 | 2019-11-05T21:28:23Z | 2019-11-27T20:47:44Z | 2019-11-27T20:47:43Z | 2019-11-28T12:21:18Z |
CLN: assorted, mostly typing | diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index a08ae66865e20..2d6c8e1008ce1 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -1150,6 +1150,77 @@ def rank_2d(rank_t[:, :] in_arr, axis=0, ties_method='average',
return ranks
+ctypedef fused diff_t:
+ float64_t
+ float32_t
+ int8_t
+ int16_t
+ int32_t
+ int64_t
+
+ctypedef fused out_t:
+ float32_t
+ float64_t
+
+
+@cython.boundscheck(False)
+@cython.wraparound(False)
+def diff_2d(ndarray[diff_t, ndim=2] arr,
+ ndarray[out_t, ndim=2] out,
+ Py_ssize_t periods, int axis):
+ cdef:
+ Py_ssize_t i, j, sx, sy, start, stop
+ bint f_contig = arr.flags.f_contiguous
+
+ # Disable for unsupported dtype combinations,
+ # see https://github.com/cython/cython/issues/2646
+ if (out_t is float32_t
+ and not (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
+ raise NotImplementedError
+ elif (out_t is float64_t
+ and (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
+ raise NotImplementedError
+ else:
+ # We put this inside an indented else block to avoid cython build
+ # warnings about unreachable code
+ sx, sy = (<object>arr).shape
+ with nogil:
+ if f_contig:
+ if axis == 0:
+ if periods >= 0:
+ start, stop = periods, sx
+ else:
+ start, stop = 0, sx + periods
+ for j in range(sy):
+ for i in range(start, stop):
+ out[i, j] = arr[i, j] - arr[i - periods, j]
+ else:
+ if periods >= 0:
+ start, stop = periods, sy
+ else:
+ start, stop = 0, sy + periods
+ for j in range(start, stop):
+ for i in range(sx):
+ out[i, j] = arr[i, j] - arr[i, j - periods]
+ else:
+ if axis == 0:
+ if periods >= 0:
+ start, stop = periods, sx
+ else:
+ start, stop = 0, sx + periods
+ for i in range(start, stop):
+ for j in range(sy):
+ out[i, j] = arr[i, j] - arr[i - periods, j]
+ else:
+ if periods >= 0:
+ start, stop = periods, sy
+ else:
+ start, stop = 0, sy + periods
+ for i in range(sx):
+ for j in range(start, stop):
+ out[i, j] = arr[i, j] - arr[i, j - periods]
+
+
# generated from template
include "algos_common_helper.pxi"
include "algos_take_helper.pxi"
diff --git a/pandas/_libs/algos_common_helper.pxi.in b/pandas/_libs/algos_common_helper.pxi.in
index ea05c4afc8fce..5bfc594602dd8 100644
--- a/pandas/_libs/algos_common_helper.pxi.in
+++ b/pandas/_libs/algos_common_helper.pxi.in
@@ -4,77 +4,6 @@ Template for each `dtype` helper function using 1-d template
WARNING: DO NOT edit .pxi FILE directly, .pxi is generated from .pxi.in
"""
-ctypedef fused diff_t:
- float64_t
- float32_t
- int8_t
- int16_t
- int32_t
- int64_t
-
-ctypedef fused out_t:
- float32_t
- float64_t
-
-
-@cython.boundscheck(False)
-@cython.wraparound(False)
-def diff_2d(ndarray[diff_t, ndim=2] arr,
- ndarray[out_t, ndim=2] out,
- Py_ssize_t periods, int axis):
- cdef:
- Py_ssize_t i, j, sx, sy, start, stop
- bint f_contig = arr.flags.f_contiguous
-
- # Disable for unsupported dtype combinations,
- # see https://github.com/cython/cython/issues/2646
- if (out_t is float32_t
- and not (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
- raise NotImplementedError
- elif (out_t is float64_t
- and (diff_t is float32_t or diff_t is int8_t or diff_t is int16_t)):
- raise NotImplementedError
- else:
- # We put this inside an indented else block to avoid cython build
- # warnings about unreachable code
- sx, sy = (<object>arr).shape
- with nogil:
- if f_contig:
- if axis == 0:
- if periods >= 0:
- start, stop = periods, sx
- else:
- start, stop = 0, sx + periods
- for j in range(sy):
- for i in range(start, stop):
- out[i, j] = arr[i, j] - arr[i - periods, j]
- else:
- if periods >= 0:
- start, stop = periods, sy
- else:
- start, stop = 0, sy + periods
- for j in range(start, stop):
- for i in range(sx):
- out[i, j] = arr[i, j] - arr[i, j - periods]
- else:
- if axis == 0:
- if periods >= 0:
- start, stop = periods, sx
- else:
- start, stop = 0, sx + periods
- for i in range(start, stop):
- for j in range(sy):
- out[i, j] = arr[i, j] - arr[i - periods, j]
- else:
- if periods >= 0:
- start, stop = periods, sy
- else:
- start, stop = 0, sy + periods
- for i in range(sx):
- for j in range(start, stop):
- out[i, j] = arr[i, j] - arr[i, j - periods]
-
-
# ----------------------------------------------------------------------
# ensure_dtype
# ----------------------------------------------------------------------
diff --git a/pandas/_libs/missing.pyx b/pandas/_libs/missing.pyx
index 052b081988c9e..9568ddb7fe53f 100644
--- a/pandas/_libs/missing.pyx
+++ b/pandas/_libs/missing.pyx
@@ -121,7 +121,7 @@ cpdef ndarray[uint8_t] isnaobj(ndarray arr):
@cython.wraparound(False)
@cython.boundscheck(False)
-def isnaobj_old(ndarray arr):
+def isnaobj_old(arr: ndarray) -> ndarray:
"""
Return boolean mask denoting which elements of a 1-D array are na-like,
defined as being any of:
@@ -156,7 +156,7 @@ def isnaobj_old(ndarray arr):
@cython.wraparound(False)
@cython.boundscheck(False)
-def isnaobj2d(ndarray arr):
+def isnaobj2d(arr: ndarray) -> ndarray:
"""
Return boolean mask denoting which elements of a 2-D array are na-like,
according to the criteria defined in `checknull`:
@@ -198,7 +198,7 @@ def isnaobj2d(ndarray arr):
@cython.wraparound(False)
@cython.boundscheck(False)
-def isnaobj2d_old(ndarray arr):
+def isnaobj2d_old(arr: ndarray) -> ndarray:
"""
Return boolean mask denoting which elements of a 2-D array are na-like,
according to the criteria defined in `checknull_old`:
diff --git a/pandas/_libs/window.pyx b/pandas/_libs/window.pyx
index d1adc7789a7a3..b51d61d05ce98 100644
--- a/pandas/_libs/window.pyx
+++ b/pandas/_libs/window.pyx
@@ -69,8 +69,8 @@ def _check_minp(win, minp, N, floor=None) -> int:
if not util.is_integer_object(minp):
raise ValueError("min_periods must be an integer")
if minp > win:
- raise ValueError("min_periods (%d) must be <= "
- "window (%d)" % (minp, win))
+ raise ValueError("min_periods (minp) must be <= "
+ "window (win)".format(minp=minp, win=win))
elif minp > N:
minp = N + 1
elif minp < 0:
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 1a2f906f97152..0e088a381e964 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -207,7 +207,7 @@ def _selected_obj(self):
return self.obj[self._selection]
@cache_readonly
- def ndim(self):
+ def ndim(self) -> int:
return self._selected_obj.ndim
@cache_readonly
@@ -339,7 +339,7 @@ def _aggregate(self, arg, *args, **kwargs):
obj = self._selected_obj
- def nested_renaming_depr(level=4):
+ def nested_renaming_depr(level: int = 4):
# deprecation of nested renaming
# GH 15931
msg = textwrap.dedent(
@@ -488,11 +488,11 @@ def _agg(arg, func):
# combine results
- def is_any_series():
+ def is_any_series() -> bool:
# return a boolean if we have *any* nested series
return any(isinstance(r, ABCSeries) for r in result.values())
- def is_any_frame():
+ def is_any_frame() -> bool:
# return a boolean if we have *any* nested series
return any(isinstance(r, ABCDataFrame) for r in result.values())
diff --git a/pandas/core/groupby/categorical.py b/pandas/core/groupby/categorical.py
index fcf52ecfcbbcd..399ed9ddc9ba1 100644
--- a/pandas/core/groupby/categorical.py
+++ b/pandas/core/groupby/categorical.py
@@ -8,7 +8,7 @@
)
-def recode_for_groupby(c, sort, observed):
+def recode_for_groupby(c: Categorical, sort: bool, observed: bool):
"""
Code the categories to ensure we can groupby for categoricals.
@@ -74,7 +74,7 @@ def recode_for_groupby(c, sort, observed):
return c.reorder_categories(cat.categories), None
-def recode_from_groupby(c, sort, ci):
+def recode_from_groupby(c: Categorical, sort: bool, ci):
"""
Reverse the codes_to_groupby to account for sort / observed.
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 1e38dde2096ba..8512b6c3ae530 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -21,6 +21,7 @@
Tuple,
Type,
Union,
+ cast,
)
import warnings
@@ -369,7 +370,7 @@ def _wrap_applied_output(self, keys, values, not_indexed_same=False):
# GH #6265
return Series([], name=self._selection_name, index=keys)
- def _get_index():
+ def _get_index() -> Index:
if self.grouper.nkeys > 1:
index = MultiIndex.from_tuples(keys, names=self.grouper.names)
else:
@@ -462,7 +463,7 @@ def transform(self, func, *args, **kwargs):
result.index = self._selected_obj.index
return result
- def _transform_fast(self, func, func_nm):
+ def _transform_fast(self, func, func_nm) -> Series:
"""
fast version of transform, only applicable to
builtin/cythonizable functions
@@ -512,7 +513,7 @@ def filter(self, func, dropna=True, *args, **kwargs):
wrapper = lambda x: func(x, *args, **kwargs)
# Interpret np.nan as False.
- def true_and_notna(x, *args, **kwargs):
+ def true_and_notna(x, *args, **kwargs) -> bool:
b = wrapper(x, *args, **kwargs)
return b and notna(b)
@@ -526,7 +527,7 @@ def true_and_notna(x, *args, **kwargs):
filtered = self._apply_filter(indices, dropna)
return filtered
- def nunique(self, dropna=True):
+ def nunique(self, dropna: bool = True) -> Series:
"""
Return number of unique elements in the group.
@@ -719,7 +720,7 @@ def value_counts(
out = ensure_int64(out)
return Series(out, index=mi, name=self._selection_name)
- def count(self):
+ def count(self) -> Series:
"""
Compute count of group, excluding missing values.
@@ -768,8 +769,6 @@ class DataFrameGroupBy(GroupBy):
_apply_whitelist = base.dataframe_apply_whitelist
- _block_agg_axis = 1
-
_agg_see_also_doc = dedent(
"""
See Also
@@ -944,19 +943,21 @@ def _iterate_slices(self) -> Iterable[Tuple[Optional[Hashable], Series]]:
yield label, values
- def _cython_agg_general(self, how, alt=None, numeric_only=True, min_count=-1):
+ def _cython_agg_general(
+ self, how: str, alt=None, numeric_only: bool = True, min_count: int = -1
+ ):
new_items, new_blocks = self._cython_agg_blocks(
how, alt=alt, numeric_only=numeric_only, min_count=min_count
)
return self._wrap_agged_blocks(new_items, new_blocks)
- _block_agg_axis = 0
-
- def _cython_agg_blocks(self, how, alt=None, numeric_only=True, min_count=-1):
+ def _cython_agg_blocks(
+ self, how: str, alt=None, numeric_only: bool = True, min_count: int = -1
+ ):
# TODO: the actual managing of mgr_locs is a PITA
# here, it should happen via BlockManager.combine
- data, agg_axis = self._get_data_to_aggregate()
+ data = self._get_data_to_aggregate()
if numeric_only:
data = data.get_numeric_data(copy=False)
@@ -971,7 +972,7 @@ def _cython_agg_blocks(self, how, alt=None, numeric_only=True, min_count=-1):
locs = block.mgr_locs.as_array
try:
result, _ = self.grouper.aggregate(
- block.values, how, axis=agg_axis, min_count=min_count
+ block.values, how, axis=1, min_count=min_count
)
except NotImplementedError:
# generally if we have numeric_only=False
@@ -1000,12 +1001,13 @@ def _cython_agg_blocks(self, how, alt=None, numeric_only=True, min_count=-1):
# continue and exclude the block
deleted_items.append(locs)
continue
-
- # unwrap DataFrame to get array
- assert len(result._data.blocks) == 1
- result = result._data.blocks[0].values
- if result.ndim == 1 and isinstance(result, np.ndarray):
- result = result.reshape(1, -1)
+ else:
+ result = cast(DataFrame, result)
+ # unwrap DataFrame to get array
+ assert len(result._data.blocks) == 1
+ result = result._data.blocks[0].values
+ if isinstance(result, np.ndarray) and result.ndim == 1:
+ result = result.reshape(1, -1)
finally:
assert not isinstance(result, DataFrame)
@@ -1081,11 +1083,11 @@ def _aggregate_frame(self, func, *args, **kwargs):
return self._wrap_frame_output(result, obj)
- def _aggregate_item_by_item(self, func, *args, **kwargs):
+ def _aggregate_item_by_item(self, func, *args, **kwargs) -> DataFrame:
# only for axis==0
obj = self._obj_with_exclusions
- result = OrderedDict()
+ result = OrderedDict() # type: dict
cannot_agg = []
errors = None
for item in obj:
@@ -1291,12 +1293,12 @@ def first_not_none(values):
# values are not series or array-like but scalars
else:
# only coerce dates if we find at least 1 datetime
- coerce = any(isinstance(x, Timestamp) for x in values)
+ should_coerce = any(isinstance(x, Timestamp) for x in values)
# self._selection_name not passed through to Series as the
# result should not take the name of original selection
# of columns
return Series(values, index=key_index)._convert(
- datetime=True, coerce=coerce
+ datetime=True, coerce=should_coerce
)
else:
@@ -1391,7 +1393,7 @@ def transform(self, func, *args, **kwargs):
return self._transform_fast(result, obj, func)
- def _transform_fast(self, result, obj, func_nm):
+ def _transform_fast(self, result: DataFrame, obj: DataFrame, func_nm) -> DataFrame:
"""
Fast transform path for aggregations
"""
@@ -1451,7 +1453,7 @@ def _choose_path(self, fast_path, slow_path, group):
return path, res
- def _transform_item_by_item(self, obj, wrapper):
+ def _transform_item_by_item(self, obj: DataFrame, wrapper) -> DataFrame:
# iterate through columns
output = {}
inds = []
@@ -1536,7 +1538,7 @@ def filter(self, func, dropna=True, *args, **kwargs):
return self._apply_filter(indices, dropna)
- def _gotitem(self, key, ndim, subset=None):
+ def _gotitem(self, key, ndim: int, subset=None):
"""
sub-classes to define
return a sliced object
@@ -1571,7 +1573,7 @@ def _gotitem(self, key, ndim, subset=None):
raise AssertionError("invalid ndim for _gotitem")
- def _wrap_frame_output(self, result, obj):
+ def _wrap_frame_output(self, result, obj) -> DataFrame:
result_index = self.grouper.levels[0]
if self.axis == 0:
@@ -1582,9 +1584,9 @@ def _wrap_frame_output(self, result, obj):
def _get_data_to_aggregate(self):
obj = self._obj_with_exclusions
if self.axis == 1:
- return obj.T._data, 1
+ return obj.T._data
else:
- return obj._data, 1
+ return obj._data
def _insert_inaxis_grouper_inplace(self, result):
# zip in reverse so we can always insert at loc 0
@@ -1622,7 +1624,7 @@ def _wrap_aggregated_output(self, output, names=None):
return self._reindex_output(result)._convert(datetime=True)
- def _wrap_transformed_output(self, output, names=None):
+ def _wrap_transformed_output(self, output, names=None) -> DataFrame:
return DataFrame(output, index=self.obj.index)
def _wrap_agged_blocks(self, items, blocks):
@@ -1670,7 +1672,7 @@ def count(self):
DataFrame
Count of values within each group.
"""
- data, _ = self._get_data_to_aggregate()
+ data = self._get_data_to_aggregate()
ids, _, ngroups = self.grouper.group_info
mask = ids != -1
@@ -1687,7 +1689,7 @@ def count(self):
return self._wrap_agged_blocks(data.items, list(blk))
- def nunique(self, dropna=True):
+ def nunique(self, dropna: bool = True):
"""
Return DataFrame with number of distinct observations per group for
each column.
@@ -1756,7 +1758,7 @@ def groupby_series(obj, col=None):
boxplot = boxplot_frame_groupby
-def _is_multi_agg_with_relabel(**kwargs):
+def _is_multi_agg_with_relabel(**kwargs) -> bool:
"""
Check whether kwargs passed to .agg look like multi-agg with relabeling.
@@ -1778,7 +1780,9 @@ def _is_multi_agg_with_relabel(**kwargs):
>>> _is_multi_agg_with_relabel()
False
"""
- return all(isinstance(v, tuple) and len(v) == 2 for v in kwargs.values()) and kwargs
+ return all(isinstance(v, tuple) and len(v) == 2 for v in kwargs.values()) and (
+ len(kwargs) > 0
+ )
def _normalize_keyword_aggregation(kwargs):
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 59b118431cfc9..873a31e658625 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -756,7 +756,7 @@ def _iterate_slices(self) -> Iterable[Tuple[Optional[Hashable], Series]]:
def transform(self, func, *args, **kwargs):
raise AbstractMethodError(self)
- def _cumcount_array(self, ascending=True):
+ def _cumcount_array(self, ascending: bool = True):
"""
Parameters
----------
@@ -788,7 +788,7 @@ def _cumcount_array(self, ascending=True):
rev[sorter] = np.arange(count, dtype=np.intp)
return out[rev].astype(np.int64, copy=False)
- def _try_cast(self, result, obj, numeric_only=False):
+ def _try_cast(self, result, obj, numeric_only: bool = False):
"""
Try to cast the result to our obj original type,
we may have roundtripped through object in the mean-time.
@@ -828,7 +828,7 @@ def _try_cast(self, result, obj, numeric_only=False):
return result
- def _transform_should_cast(self, func_nm):
+ def _transform_should_cast(self, func_nm: str) -> bool:
"""
Parameters
----------
@@ -844,8 +844,8 @@ def _transform_should_cast(self, func_nm):
func_nm not in base.cython_cast_blacklist
)
- def _cython_transform(self, how, numeric_only=True, **kwargs):
- output = collections.OrderedDict()
+ def _cython_transform(self, how: str, numeric_only: bool = True, **kwargs):
+ output = collections.OrderedDict() # type: dict
for name, obj in self._iterate_slices():
is_numeric = is_numeric_dtype(obj.dtype)
if numeric_only and not is_numeric:
@@ -871,10 +871,12 @@ def _wrap_aggregated_output(self, output, names=None):
def _wrap_transformed_output(self, output, names=None):
raise AbstractMethodError(self)
- def _wrap_applied_output(self, keys, values, not_indexed_same=False):
+ def _wrap_applied_output(self, keys, values, not_indexed_same: bool = False):
raise AbstractMethodError(self)
- def _cython_agg_general(self, how, alt=None, numeric_only=True, min_count=-1):
+ def _cython_agg_general(
+ self, how: str, alt=None, numeric_only: bool = True, min_count: int = -1
+ ):
output = {}
for name, obj in self._iterate_slices():
is_numeric = is_numeric_dtype(obj.dtype)
@@ -920,7 +922,7 @@ def _python_agg_general(self, func, *args, **kwargs):
return self._wrap_aggregated_output(output)
- def _concat_objects(self, keys, values, not_indexed_same=False):
+ def _concat_objects(self, keys, values, not_indexed_same: bool = False):
from pandas.core.reshape.concat import concat
def reset_identity(values):
@@ -980,10 +982,7 @@ def reset_identity(values):
values = reset_identity(values)
result = concat(values, axis=self.axis)
- if (
- isinstance(result, Series)
- and getattr(self, "_selection_name", None) is not None
- ):
+ if isinstance(result, Series) and self._selection_name is not None:
result.name = self._selection_name
@@ -1104,7 +1103,7 @@ def result_to_bool(result: np.ndarray, inference: Type) -> np.ndarray:
@Substitution(name="groupby")
@Appender(_common_see_also)
- def any(self, skipna=True):
+ def any(self, skipna: bool = True):
"""
Return True if any value in the group is truthful, else False.
@@ -1121,7 +1120,7 @@ def any(self, skipna=True):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def all(self, skipna=True):
+ def all(self, skipna: bool = True):
"""
Return True if all values in the group are truthful, else False.
@@ -1221,7 +1220,7 @@ def median(self, **kwargs):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def std(self, ddof=1, *args, **kwargs):
+ def std(self, ddof: int = 1, *args, **kwargs):
"""
Compute standard deviation of groups, excluding missing values.
@@ -1244,7 +1243,7 @@ def std(self, ddof=1, *args, **kwargs):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def var(self, ddof=1, *args, **kwargs):
+ def var(self, ddof: int = 1, *args, **kwargs):
"""
Compute variance of groups, excluding missing values.
@@ -1272,7 +1271,7 @@ def var(self, ddof=1, *args, **kwargs):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def sem(self, ddof=1):
+ def sem(self, ddof: int = 1):
"""
Compute standard error of the mean of groups, excluding missing values.
@@ -1313,7 +1312,13 @@ def _add_numeric_operations(cls):
Add numeric operations to the GroupBy generically.
"""
- def groupby_function(name, alias, npfunc, numeric_only=True, min_count=-1):
+ def groupby_function(
+ name: str,
+ alias: str,
+ npfunc,
+ numeric_only: bool = True,
+ min_count: int = -1,
+ ):
_local_template = """
Compute %(f)s of group values.
@@ -1403,7 +1408,7 @@ def last(x):
@Substitution(name="groupby")
@Appender(_common_see_also)
- def ohlc(self):
+ def ohlc(self) -> DataFrame:
"""
Compute sum of values, excluding missing values.
@@ -1815,7 +1820,7 @@ def nth(self, n: Union[int, List[int]], dropna: Optional[str] = None) -> DataFra
return result
- def quantile(self, q=0.5, interpolation="linear"):
+ def quantile(self, q=0.5, interpolation: str = "linear"):
"""
Return group values at the given quantile, a la numpy.percentile.
@@ -1928,7 +1933,7 @@ def post_processor(vals: np.ndarray, inference: Optional[Type]) -> np.ndarray:
return result.take(indices)
@Substitution(name="groupby")
- def ngroup(self, ascending=True):
+ def ngroup(self, ascending: bool = True):
"""
Number each group from 0 to the number of groups - 1.
@@ -1997,7 +2002,7 @@ def ngroup(self, ascending=True):
return result
@Substitution(name="groupby")
- def cumcount(self, ascending=True):
+ def cumcount(self, ascending: bool = True):
"""
Number each item in each group from 0 to the length of that group - 1.
@@ -2058,7 +2063,12 @@ def cumcount(self, ascending=True):
@Substitution(name="groupby")
@Appender(_common_see_also)
def rank(
- self, method="average", ascending=True, na_option="keep", pct=False, axis=0
+ self,
+ method: str = "average",
+ ascending: bool = True,
+ na_option: str = "keep",
+ pct: bool = False,
+ axis: int = 0,
):
"""
Provide the rank of values within each group.
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 9bbe73c1851b5..2cc0e5fde2290 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,7 +7,7 @@
"""
import collections
-from typing import List, Optional
+from typing import List, Optional, Type
import numpy as np
@@ -96,7 +96,7 @@ def __iter__(self):
return iter(self.indices)
@property
- def nkeys(self):
+ def nkeys(self) -> int:
return len(self.groupings)
def get_iterator(self, data, axis=0):
@@ -135,7 +135,7 @@ def _get_group_keys(self):
# provide "flattened" iterator for multi-group setting
return get_flattened_iterator(comp_ids, ngroups, self.levels, self.labels)
- def apply(self, f, data, axis=0):
+ def apply(self, f, data, axis: int = 0):
mutated = self.mutated
splitter = self._get_splitter(data, axis=axis)
group_keys = self._get_group_keys()
@@ -220,7 +220,7 @@ def levels(self):
def names(self):
return [ping.name for ping in self.groupings]
- def size(self):
+ def size(self) -> Series:
"""
Compute group sizes
@@ -244,7 +244,7 @@ def groups(self):
return self.axis.groupby(to_groupby)
@cache_readonly
- def is_monotonic(self):
+ def is_monotonic(self) -> bool:
# return if my group orderings are monotonic
return Index(self.group_info[0]).is_monotonic
@@ -275,7 +275,7 @@ def _get_compressed_labels(self):
return ping.labels, np.arange(len(ping.group_index))
@cache_readonly
- def ngroups(self):
+ def ngroups(self) -> int:
return len(self.result_index)
@property
@@ -345,7 +345,7 @@ def _is_builtin_func(self, arg):
"""
return SelectionMixin._builtin_table.get(arg, arg)
- def _get_cython_function(self, kind, how, values, is_numeric):
+ def _get_cython_function(self, kind: str, how: str, values, is_numeric: bool):
dtype_str = values.dtype.name
@@ -386,7 +386,9 @@ def get_func(fname):
return func
- def _cython_operation(self, kind: str, values, how, axis, min_count=-1, **kwargs):
+ def _cython_operation(
+ self, kind: str, values, how: str, axis: int, min_count: int = -1, **kwargs
+ ):
assert kind in ["transform", "aggregate"]
orig_values = values
@@ -530,16 +532,23 @@ def _cython_operation(self, kind: str, values, how, axis, min_count=-1, **kwargs
return result, names
- def aggregate(self, values, how, axis=0, min_count=-1):
+ def aggregate(self, values, how: str, axis: int = 0, min_count: int = -1):
return self._cython_operation(
"aggregate", values, how, axis, min_count=min_count
)
- def transform(self, values, how, axis=0, **kwargs):
+ def transform(self, values, how: str, axis: int = 0, **kwargs):
return self._cython_operation("transform", values, how, axis, **kwargs)
def _aggregate(
- self, result, counts, values, comp_ids, agg_func, is_datetimelike, min_count=-1
+ self,
+ result,
+ counts,
+ values,
+ comp_ids,
+ agg_func,
+ is_datetimelike: bool,
+ min_count: int = -1,
):
if values.ndim > 2:
# punting for now
@@ -554,7 +563,7 @@ def _aggregate(
return result
def _transform(
- self, result, values, comp_ids, transform_func, is_datetimelike, **kwargs
+ self, result, values, comp_ids, transform_func, is_datetimelike: bool, **kwargs
):
comp_ids, _, ngroups = self.group_info
@@ -566,7 +575,7 @@ def _transform(
return result
- def agg_series(self, obj, func):
+ def agg_series(self, obj: Series, func):
if is_extension_array_dtype(obj.dtype) and obj.dtype.kind != "M":
# _aggregate_series_fast would raise TypeError when
# calling libreduction.Slider
@@ -684,7 +693,7 @@ def groups(self):
return result
@property
- def nkeys(self):
+ def nkeys(self) -> int:
return 1
def _get_grouper(self):
@@ -771,7 +780,7 @@ def groupings(self):
for lvl, name in zip(self.levels, self.names)
]
- def agg_series(self, obj, func):
+ def agg_series(self, obj: Series, func):
dummy = obj[:0]
grouper = libreduction.SeriesBinGrouper(obj, func, self.bins, dummy)
return grouper.get_result()
@@ -863,10 +872,11 @@ def _chop(self, sdata, slice_obj: slice):
return sdata._slice(slice_obj, axis=1)
-def get_splitter(data, *args, **kwargs):
+def get_splitter(data: NDFrame, *args, **kwargs):
if isinstance(data, Series):
- klass = SeriesSplitter
- elif isinstance(data, DataFrame):
+ klass = SeriesSplitter # type: Type[DataSplitter]
+ else:
+ # i.e. DataFrame
klass = FrameSplitter
return klass(data, *args, **kwargs)
diff --git a/pandas/core/internals/concat.py b/pandas/core/internals/concat.py
index 36e1b06230d7e..4ba485c85d8ba 100644
--- a/pandas/core/internals/concat.py
+++ b/pandas/core/internals/concat.py
@@ -244,7 +244,7 @@ def concatenate_join_units(join_units, concat_axis, copy):
# Concatenating join units along ax0 is handled in _merge_blocks.
raise AssertionError("Concatenating join units along axis0")
- empty_dtype, upcasted_na = get_empty_dtype_and_na(join_units)
+ empty_dtype, upcasted_na = _get_empty_dtype_and_na(join_units)
to_concat = [
ju.get_reindexed_values(empty_dtype=empty_dtype, upcasted_na=upcasted_na)
@@ -268,7 +268,7 @@ def concatenate_join_units(join_units, concat_axis, copy):
return concat_values
-def get_empty_dtype_and_na(join_units):
+def _get_empty_dtype_and_na(join_units):
"""
Return dtype and N/A values to use when concatenating specified units.
@@ -284,7 +284,7 @@ def get_empty_dtype_and_na(join_units):
if blk is None:
return np.float64, np.nan
- if is_uniform_reindex(join_units):
+ if _is_uniform_reindex(join_units):
# FIXME: integrate property
empty_dtype = join_units[0].block.dtype
upcasted_na = join_units[0].block.fill_value
@@ -398,7 +398,7 @@ def is_uniform_join_units(join_units):
)
-def is_uniform_reindex(join_units):
+def _is_uniform_reindex(join_units) -> bool:
return (
# TODO: should this be ju.block._can_hold_na?
all(ju.block and ju.block.is_extension for ju in join_units)
@@ -406,7 +406,7 @@ def is_uniform_reindex(join_units):
)
-def trim_join_unit(join_unit, length):
+def _trim_join_unit(join_unit, length):
"""
Reduce join_unit's shape along item axis to length.
@@ -486,9 +486,9 @@ def _next_or_none(seq):
for i, (plc, unit) in enumerate(next_items):
yielded_units[i] = unit
if len(plc) > min_len:
- # trim_join_unit updates unit in place, so only
+ # _trim_join_unit updates unit in place, so only
# placement needs to be sliced to skip min_len.
- next_items[i] = (plc[min_len:], trim_join_unit(unit, min_len))
+ next_items[i] = (plc[min_len:], _trim_join_unit(unit, min_len))
else:
yielded_placement = plc
next_items[i] = _next_or_none(plans[i])
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index 39e00047ea968..772ac1cd93059 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -29,15 +29,15 @@
def concat(
objs,
axis=0,
- join="outer",
+ join: str = "outer",
join_axes=None,
- ignore_index=False,
+ ignore_index: bool = False,
keys=None,
levels=None,
names=None,
- verify_integrity=False,
+ verify_integrity: bool = False,
sort=None,
- copy=True,
+ copy: bool = True,
):
"""
Concatenate pandas objects along a particular axis with optional set logic
@@ -265,14 +265,14 @@ def __init__(
self,
objs,
axis=0,
- join="outer",
+ join: str = "outer",
join_axes=None,
keys=None,
levels=None,
names=None,
- ignore_index=False,
- verify_integrity=False,
- copy=True,
+ ignore_index: bool = False,
+ verify_integrity: bool = False,
+ copy: bool = True,
sort=False,
):
if isinstance(objs, (NDFrame, str)):
@@ -324,8 +324,8 @@ def __init__(
for obj in objs:
if not isinstance(obj, (Series, DataFrame)):
msg = (
- "cannot concatenate object of type '{}';"
- " only Series and DataFrame objs are valid".format(type(obj))
+ "cannot concatenate object of type '{typ}';"
+ " only Series and DataFrame objs are valid".format(typ=type(obj))
)
raise TypeError(msg)
@@ -580,7 +580,7 @@ def _get_concat_axis(self):
return concat_axis
- def _maybe_check_integrity(self, concat_index):
+ def _maybe_check_integrity(self, concat_index: Index):
if self.verify_integrity:
if not concat_index.is_unique:
overlap = concat_index[concat_index.duplicated()].unique()
@@ -590,11 +590,11 @@ def _maybe_check_integrity(self, concat_index):
)
-def _concat_indexes(indexes):
+def _concat_indexes(indexes) -> Index:
return indexes[0].append(indexes[1:])
-def _make_concat_multiindex(indexes, keys, levels=None, names=None):
+def _make_concat_multiindex(indexes, keys, levels=None, names=None) -> MultiIndex:
if (levels is None and isinstance(keys[0], tuple)) or (
levels is not None and len(levels) > 1
@@ -715,7 +715,6 @@ def _get_series_result_type(result, objs=None):
"""
# TODO: See if we can just inline with _constructor_expanddim
# now that sparse is removed.
- from pandas import DataFrame
# concat Series with axis 1
if isinstance(result, dict):
diff --git a/pandas/core/reshape/melt.py b/pandas/core/reshape/melt.py
index c85050bc4232b..98fee491e0a73 100644
--- a/pandas/core/reshape/melt.py
+++ b/pandas/core/reshape/melt.py
@@ -188,7 +188,7 @@ def lreshape(data, groups, dropna=True, label=None):
return data._constructor(mdata, columns=id_cols + pivot_cols)
-def wide_to_long(df, stubnames, i, j, sep="", suffix=r"\d+"):
+def wide_to_long(df, stubnames, i, j, sep: str = "", suffix: str = r"\d+"):
r"""
Wide panel to long format. Less flexible but more user-friendly than melt.
@@ -419,7 +419,7 @@ def get_var_names(df, stub, sep, suffix):
pattern = re.compile(regex)
return [col for col in df.columns if pattern.match(col)]
- def melt_stub(df, stub, i, j, value_vars, sep):
+ def melt_stub(df, stub, i, j, value_vars, sep: str):
newdf = melt(
df,
id_vars=i,
@@ -456,8 +456,8 @@ def melt_stub(df, stub, i, j, value_vars, sep):
value_vars_flattened = [e for sublist in value_vars for e in sublist]
id_vars = list(set(df.columns.tolist()).difference(value_vars_flattened))
- melted = [melt_stub(df, s, i, j, v, sep) for s, v in zip(stubnames, value_vars)]
- melted = melted[0].join(melted[1:], how="outer")
+ _melted = [melt_stub(df, s, i, j, v, sep) for s, v in zip(stubnames, value_vars)]
+ melted = _melted[0].join(_melted[1:], how="outer")
if len(i) == 1:
new = df[id_vars].set_index(i).join(melted)
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index 6ef13a62ee366..a189b2cd1ab84 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -10,7 +10,7 @@
import numpy as np
-from pandas._libs import hashtable as libhashtable, lib
+from pandas._libs import Timedelta, hashtable as libhashtable, lib
import pandas._libs.join as libjoin
from pandas.errors import MergeError
from pandas.util._decorators import Appender, Substitution
@@ -36,9 +36,10 @@
is_object_dtype,
needs_i8_conversion,
)
-from pandas.core.dtypes.missing import isnull, na_value_for_dtype
+from pandas.core.dtypes.generic import ABCDataFrame, ABCSeries
+from pandas.core.dtypes.missing import isna, na_value_for_dtype
-from pandas import Categorical, DataFrame, Index, MultiIndex, Series, Timedelta
+from pandas import Categorical, Index, MultiIndex
import pandas.core.algorithms as algos
from pandas.core.arrays.categorical import _recode_for_categories
import pandas.core.common as com
@@ -1204,7 +1205,7 @@ def _validate_specification(self):
if len(self.right_on) != len(self.left_on):
raise ValueError("len(right_on) must equal len(left_on)")
- def _validate(self, validate):
+ def _validate(self, validate: str):
# Check uniqueness of each
if self.left_index:
@@ -1300,7 +1301,12 @@ def _get_join_indexers(left_keys, right_keys, sort=False, how="inner", **kwargs)
def _restore_dropped_levels_multijoin(
- left, right, dropped_level_names, join_index, lindexer, rindexer
+ left: MultiIndex,
+ right: MultiIndex,
+ dropped_level_names,
+ join_index,
+ lindexer,
+ rindexer,
):
"""
*this is an internal non-public method*
@@ -1338,7 +1344,7 @@ def _restore_dropped_levels_multijoin(
"""
- def _convert_to_mulitindex(index):
+ def _convert_to_mulitindex(index) -> MultiIndex:
if isinstance(index, MultiIndex):
return index
else:
@@ -1686,13 +1692,13 @@ def flip(xs):
msg_missings = "Merge keys contain null values on {side} side"
if not Index(left_values).is_monotonic:
- if isnull(left_values).any():
+ if isna(left_values).any():
raise ValueError(msg_missings.format(side="left"))
else:
raise ValueError(msg_sorted.format(side="left"))
if not Index(right_values).is_monotonic:
- if isnull(right_values).any():
+ if isna(right_values).any():
raise ValueError(msg_missings.format(side="right"))
else:
raise ValueError(msg_sorted.format(side="right"))
@@ -1959,9 +1965,9 @@ def _any(x) -> bool:
def validate_operand(obj):
- if isinstance(obj, DataFrame):
+ if isinstance(obj, ABCDataFrame):
return obj
- elif isinstance(obj, Series):
+ elif isinstance(obj, ABCSeries):
if obj.name is None:
raise ValueError("Cannot merge a Series without a name")
else:
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index 7537dd0ac2065..a8dcc995e48da 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -958,7 +958,7 @@ def _get_dummies_1d(
if is_object_dtype(dtype):
raise ValueError("dtype=object is not a valid dtype for get_dummies")
- def get_empty_frame(data):
+ def get_empty_frame(data) -> DataFrame:
if isinstance(data, Series):
index = data.index
else:
diff --git a/pandas/core/window/common.py b/pandas/core/window/common.py
index 0f2920b3558c9..2ad5a1eb6faed 100644
--- a/pandas/core/window/common.py
+++ b/pandas/core/window/common.py
@@ -32,7 +32,7 @@ class _GroupByMixin(GroupByMixin):
"""
def __init__(self, obj, *args, **kwargs):
- parent = kwargs.pop("parent", None) # noqa
+ kwargs.pop("parent", None)
groupby = kwargs.pop("groupby", None)
if groupby is None:
groupby, obj = obj, obj.obj
diff --git a/pandas/core/window/rolling.py b/pandas/core/window/rolling.py
index 68eb1f630bfc3..f6d27de132ad9 100644
--- a/pandas/core/window/rolling.py
+++ b/pandas/core/window/rolling.py
@@ -1642,17 +1642,18 @@ def _get_corr(a, b):
class Rolling(_Rolling_and_Expanding):
@cache_readonly
- def is_datetimelike(self):
+ def is_datetimelike(self) -> bool:
return isinstance(
self._on, (ABCDatetimeIndex, ABCTimedeltaIndex, ABCPeriodIndex)
)
@cache_readonly
- def _on(self):
+ def _on(self) -> Index:
if self.on is None:
if self.axis == 0:
return self.obj.index
- elif self.axis == 1:
+ else:
+ # i.e. self.axis == 1
return self.obj.columns
elif isinstance(self.on, Index):
return self.on
@@ -1660,9 +1661,9 @@ def _on(self):
return Index(self.obj[self.on])
else:
raise ValueError(
- "invalid on specified as {0}, "
+ "invalid on specified as {on}, "
"must be a column (of DataFrame), an Index "
- "or None".format(self.on)
+ "or None".format(on=self.on)
)
def validate(self):
@@ -1711,7 +1712,9 @@ def _validate_monotonic(self):
formatted = self.on
if self.on is None:
formatted = "index"
- raise ValueError("{0} must be monotonic".format(formatted))
+ raise ValueError(
+ "{formatted} must be monotonic".format(formatted=formatted)
+ )
def _validate_freq(self):
"""
@@ -1723,9 +1726,9 @@ def _validate_freq(self):
return to_offset(self.window)
except (TypeError, ValueError):
raise ValueError(
- "passed window {0} is not "
+ "passed window {window} is not "
"compatible with a datetimelike "
- "index".format(self.window)
+ "index".format(window=self.window)
)
_agg_see_also_doc = dedent(
diff --git a/pandas/tseries/offsets.py b/pandas/tseries/offsets.py
index 1e3f5c1ed870e..f5e40e712642e 100644
--- a/pandas/tseries/offsets.py
+++ b/pandas/tseries/offsets.py
@@ -36,8 +36,6 @@
from pandas.core.dtypes.inference import is_list_like
-from pandas.core.tools.datetimes import to_datetime
-
__all__ = [
"Day",
"BusinessDay",
@@ -2752,8 +2750,10 @@ def generate_range(start=None, end=None, periods=None, offset=BDay()):
offset = to_offset(offset)
- start = to_datetime(start)
- end = to_datetime(end)
+ start = Timestamp(start)
+ start = start if start is not NaT else None
+ end = Timestamp(end)
+ end = end if end is not NaT else None
if start and not offset.onOffset(start):
start = offset.rollforward(start)
| diff_2d no longer needs to be in the pxi.in file, so moved it to the pyx
A couple of recently-identified bugs in the groupby code are caused by passing incorrect types, so im getting more motivated to add annotations in/around the affected code. | https://api.github.com/repos/pandas-dev/pandas/pulls/29419 | 2019-11-05T19:46:33Z | 2019-11-06T18:11:04Z | 2019-11-06T18:11:04Z | 2019-11-06T18:19:32Z |
maybe_promote: Restrict fill_value to scalar for non-object dtype | diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index 542618e332f7b..fad80d6bf5745 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -339,6 +339,11 @@ def changeit():
def maybe_promote(dtype, fill_value=np.nan):
+ if not is_scalar(fill_value) and not is_object_dtype(dtype):
+ # with object dtype there is nothing to promote, and the user can
+ # pass pretty much any weird fill_value they like
+ raise ValueError("fill_value must be a scalar")
+
# if we passed an array here, determine the fill value by dtype
if isinstance(fill_value, np.ndarray):
if issubclass(fill_value.dtype.type, (np.datetime64, np.timedelta64)):
@@ -686,7 +691,8 @@ def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
dtype : if None, then use the dtype of the values, else coerce to this type
copy : if True always make a copy even if no upcast is required
"""
- if not is_scalar(fill_value):
+ if not is_scalar(fill_value) and not is_object_dtype(values.dtype):
+ # We allow arbitrary fill values for object dtype
raise ValueError("fill_value must be a scalar")
if is_extension_type(values):
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index c792460add429..448d2faf8b85f 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -1283,10 +1283,6 @@ def diff(self, n: int, axis: int = 1) -> List["Block"]:
def shift(self, periods, axis=0, fill_value=None):
""" shift the block by periods, possibly upcast """
- if not lib.is_scalar(fill_value):
- # We could go further and require e.g. self._can_hold_element(fv)
- raise ValueError("fill_value must be a scalar")
-
# convert integer to float if necessary. need to do a lot more than
# that, handle boolean etc also
new_values, fill_value = maybe_upcast(self.values, fill_value)
diff --git a/pandas/tests/dtypes/cast/test_promote.py b/pandas/tests/dtypes/cast/test_promote.py
index 5c61574eddb50..0939e35bd64fa 100644
--- a/pandas/tests/dtypes/cast/test_promote.py
+++ b/pandas/tests/dtypes/cast/test_promote.py
@@ -19,7 +19,6 @@
is_integer_dtype,
is_object_dtype,
is_scalar,
- is_string_dtype,
is_timedelta64_dtype,
)
from pandas.core.dtypes.dtypes import DatetimeTZDtype
@@ -65,42 +64,7 @@ def any_numpy_dtype_reduced(request):
return request.param
-@pytest.fixture(
- params=[(True, None), (True, object), (False, None)],
- ids=["True-None", "True-object", "False-None"],
-)
-def box(request):
- """
- Parametrized fixture determining whether/how to transform fill_value.
-
- Since fill_value is defined on a per-test basis, the actual transformation
- (based on this fixture) is executed in _check_promote.
-
- Returns
- -------
- boxed : Boolean
- Whether fill_value should be wrapped in an np.array.
- box_dtype : dtype
- The dtype to pass to np.array([fill_value], dtype=box_dtype). If None,
- then this is passed on unmodified, and corresponds to the numpy default
- dtype for the given fill_value.
-
- * (True, None) # fill_value wrapped in array with default dtype
- * (True, object) # fill_value wrapped in array with object dtype
- * (False, None) # fill_value passed on as scalar
- """
- return request.param
-
-
-def _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar=None,
- exp_val_for_array=None,
-):
+def _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar=None):
"""
Auxiliary function to unify testing of scalar/array promotion.
@@ -109,13 +73,8 @@ def _check_promote(
dtype : dtype
The value to pass on as the first argument to maybe_promote.
fill_value : scalar
- The value to pass on as the second argument to maybe_promote, either as
- a scalar, or boxed into an array (depending on the parameter `boxed`).
- boxed : Boolean
- Parameter whether fill_value should be passed to maybe_promote
- directly, or wrapped in an array (of dtype box_dtype).
- box_dtype : dtype
- The dtype to enforce when wrapping fill_value into an np.array.
+ The value to pass on as the second argument to maybe_promote as
+ a scalar.
expected_dtype : dtype
The expected dtype returned by maybe_promote (by design this is the
same regardless of whether fill_value was passed as a scalar or in an
@@ -123,25 +82,14 @@ def _check_promote(
exp_val_for_scalar : scalar
The expected value for the (potentially upcast) fill_value returned by
maybe_promote.
- exp_val_for_array : scalar
- The expected missing value marker for the expected_dtype (which is
- returned by maybe_promote when it receives an array).
"""
assert is_scalar(fill_value)
- if boxed:
- # in this case, we pass on fill_value wrapped in an array of specified
- # box_dtype; the expected value returned from maybe_promote is the
- # missing value marker for the returned dtype.
- fill_array = np.array([fill_value], dtype=box_dtype)
- result_dtype, result_fill_value = maybe_promote(dtype, fill_array)
- expected_fill_value = exp_val_for_array
- else:
- # here, we pass on fill_value as a scalar directly; the expected value
- # returned from maybe_promote is fill_value, potentially upcast to the
- # returned dtype.
- result_dtype, result_fill_value = maybe_promote(dtype, fill_value)
- expected_fill_value = exp_val_for_scalar
+ # here, we pass on fill_value as a scalar directly; the expected value
+ # returned from maybe_promote is fill_value, potentially upcast to the
+ # returned dtype.
+ result_dtype, result_fill_value = maybe_promote(dtype, fill_value)
+ expected_fill_value = exp_val_for_scalar
assert result_dtype == expected_dtype
_assert_match(result_fill_value, expected_fill_value)
@@ -280,41 +228,19 @@ def _assert_match(result_fill_value, expected_fill_value):
("uint64", np.iinfo("int64").min - 1, "object"),
],
)
-def test_maybe_promote_int_with_int(dtype, fill_value, expected_dtype, box):
+def test_maybe_promote_int_with_int(dtype, fill_value, expected_dtype):
dtype = np.dtype(dtype)
expected_dtype = np.dtype(expected_dtype)
- boxed, box_dtype = box # read from parametrized fixture
-
- if boxed:
- if expected_dtype != object:
- pytest.xfail("falsely casts to object")
- if box_dtype is None and (
- fill_value > np.iinfo("int64").max or np.iinfo("int64").min < fill_value < 0
- ):
- pytest.xfail("falsely casts to float instead of object")
# output is not a generic int, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- # no missing value marker for integers
- exp_val_for_array = None if expected_dtype != "object" else np.nan
-
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
-
-
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, None), (False, None)])
-def test_maybe_promote_int_with_float(any_int_dtype, float_dtype, box):
+
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
+
+
+def test_maybe_promote_int_with_float(any_int_dtype, float_dtype):
dtype = np.dtype(any_int_dtype)
fill_dtype = np.dtype(float_dtype)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -323,26 +249,14 @@ def test_maybe_promote_int_with_float(any_int_dtype, float_dtype, box):
expected_dtype = np.float64
# fill_value can be different float type
exp_val_for_scalar = np.float64(fill_value)
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, None), (False, None)])
-def test_maybe_promote_float_with_int(float_dtype, any_int_dtype, box):
+def test_maybe_promote_float_with_int(float_dtype, any_int_dtype):
dtype = np.dtype(float_dtype)
fill_dtype = np.dtype(any_int_dtype)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -352,17 +266,8 @@ def test_maybe_promote_float_with_int(float_dtype, any_int_dtype, box):
expected_dtype = dtype
# output is not a generic float, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize(
@@ -390,49 +295,20 @@ def test_maybe_promote_float_with_int(float_dtype, any_int_dtype, box):
("complex128", np.finfo("float32").max * (1.1 + 1j), "complex128"),
],
)
-def test_maybe_promote_float_with_float(dtype, fill_value, expected_dtype, box):
+def test_maybe_promote_float_with_float(dtype, fill_value, expected_dtype):
dtype = np.dtype(dtype)
expected_dtype = np.dtype(expected_dtype)
- boxed, box_dtype = box # read from parametrized fixture
-
- if box_dtype == object:
- pytest.xfail("falsely upcasts to object")
- elif boxed and is_float_dtype(dtype) and is_complex_dtype(expected_dtype):
- pytest.xfail("does not upcast to complex")
- elif boxed and (dtype, expected_dtype) in [
- ("float32", "float64"),
- ("float32", "complex64"),
- ("complex64", "complex128"),
- ]:
- pytest.xfail("does not upcast correctly depending on value")
# output is not a generic float, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_bool_with_any(any_numpy_dtype_reduced, box):
+def test_maybe_promote_bool_with_any(any_numpy_dtype_reduced):
dtype = np.dtype(bool)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if boxed and fill_dtype == bool:
- pytest.xfail("falsely upcasts to object")
- if boxed and box_dtype is None and fill_dtype.kind == "M":
- pytest.xfail("wrongly casts fill_value")
- if boxed and box_dtype is None and fill_dtype.kind == "m":
- pytest.xfail("wrongly casts fill_value")
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -440,50 +316,25 @@ def test_maybe_promote_bool_with_any(any_numpy_dtype_reduced, box):
# filling bool with anything but bool casts to object
expected_dtype = np.dtype(object) if fill_dtype != bool else fill_dtype
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan if fill_dtype != bool else None
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_any_with_bool(any_numpy_dtype_reduced, box):
+def test_maybe_promote_any_with_bool(any_numpy_dtype_reduced):
dtype = np.dtype(any_numpy_dtype_reduced)
fill_value = True
- boxed, box_dtype = box # read from parametrized fixture
-
- if boxed and dtype == bool:
- pytest.xfail("falsely upcasts to object")
- if boxed and dtype not in (str, object) and box_dtype is None:
- pytest.xfail("falsely upcasts to object")
# filling anything but bool with bool casts to object
expected_dtype = np.dtype(object) if dtype != bool else dtype
# output is not a generic bool, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan if dtype != bool else None
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_bytes_with_any(bytes_dtype, any_numpy_dtype_reduced, box):
+def test_maybe_promote_bytes_with_any(bytes_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(bytes_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -491,78 +342,27 @@ def test_maybe_promote_bytes_with_any(bytes_dtype, any_numpy_dtype_reduced, box)
# we never use bytes dtype internally, always promote to object
expected_dtype = np.dtype(np.object_)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization of box to add special case for bytes
-@pytest.mark.parametrize(
- "box",
- [
- (True, None), # fill_value wrapped in array with auto-dtype (fixed len)
- (True, "bytes"), # fill_value wrapped in array with generic bytes-dtype
- (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value directly
- ],
-)
-def test_maybe_promote_any_with_bytes(any_numpy_dtype_reduced, bytes_dtype, box):
+def test_maybe_promote_any_with_bytes(any_numpy_dtype_reduced, bytes_dtype):
dtype = np.dtype(any_numpy_dtype_reduced)
- fill_dtype = np.dtype(bytes_dtype)
- boxed, box_dtype = box # read from parametrized fixture
-
- if not issubclass(dtype.type, np.bytes_):
- if (
- boxed
- and (box_dtype == "bytes" or box_dtype is None)
- and not (is_string_dtype(dtype) or dtype == bool)
- ):
- pytest.xfail("does not upcast to object")
# create array of given dtype
fill_value = b"abc"
- # special case for box_dtype (cannot use fixture in parametrization)
- box_dtype = fill_dtype if box_dtype == "bytes" else box_dtype
-
# we never use bytes dtype internally, always promote to object
expected_dtype = np.dtype(np.object_)
# output is not a generic bytes, but corresponds to expected_dtype
exp_val_for_scalar = np.array([fill_value], dtype=expected_dtype)[0]
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_datetime64_with_any(
- datetime64_dtype, any_numpy_dtype_reduced, box
-):
+def test_maybe_promote_datetime64_with_any(datetime64_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(datetime64_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_datetime64_dtype(fill_dtype):
- if box_dtype == object:
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and box_dtype is None:
- pytest.xfail("does not upcast to object")
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -572,34 +372,13 @@ def test_maybe_promote_datetime64_with_any(
expected_dtype = dtype
# for datetime dtypes, scalar values get cast to to_datetime64
exp_val_for_scalar = pd.Timestamp(fill_value).to_datetime64()
- exp_val_for_array = np.datetime64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization of box to add special case for dt_dtype
-@pytest.mark.parametrize(
- "box",
- [
- (True, None), # fill_value wrapped in array with default dtype
- # disabled due to too many xfails; see GH 23982 / 25425
- # (True, 'dt_dtype'), # fill_value in array with explicit datetime dtype
- # (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value passed on as scalar
- ],
-)
@pytest.mark.parametrize(
"fill_value",
[
@@ -611,57 +390,28 @@ def test_maybe_promote_datetime64_with_any(
ids=["pd.Timestamp", "np.datetime64", "datetime.datetime", "datetime.date"],
)
def test_maybe_promote_any_with_datetime64(
- any_numpy_dtype_reduced, datetime64_dtype, fill_value, box
+ any_numpy_dtype_reduced, datetime64_dtype, fill_value
):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_datetime64_dtype(dtype):
- if boxed and (
- box_dtype == object
- or (box_dtype is None and not is_datetime64_dtype(type(fill_value)))
- ):
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and (
- box_dtype == "dt_dtype"
- or (box_dtype is None and is_datetime64_dtype(type(fill_value)))
- ):
- pytest.xfail("mix of lack of upcasting, resp. wrong missing value")
-
- # special case for box_dtype
- box_dtype = np.dtype(datetime64_dtype) if box_dtype == "dt_dtype" else box_dtype
# filling datetime with anything but datetime casts to object
if is_datetime64_dtype(dtype):
expected_dtype = dtype
# for datetime dtypes, scalar values get cast to pd.Timestamp.value
exp_val_for_scalar = pd.Timestamp(fill_value).to_datetime64()
- exp_val_for_array = np.datetime64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, object)])
+@pytest.mark.xfail(reason="Fails to upcast to object")
def test_maybe_promote_datetimetz_with_any_numpy_dtype(
- tz_aware_fixture, any_numpy_dtype_reduced, box
+ tz_aware_fixture, any_numpy_dtype_reduced
):
dtype = DatetimeTZDtype(tz=tz_aware_fixture)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -669,34 +419,18 @@ def test_maybe_promote_datetimetz_with_any_numpy_dtype(
# filling datetimetz with any numpy dtype casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
-
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
-
-
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(True, None), (True, object)])
-def test_maybe_promote_datetimetz_with_datetimetz(
- tz_aware_fixture, tz_aware_fixture2, box
-):
+
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
+
+
+def test_maybe_promote_datetimetz_with_datetimetz(tz_aware_fixture, tz_aware_fixture2):
dtype = DatetimeTZDtype(tz=tz_aware_fixture)
fill_dtype = DatetimeTZDtype(tz=tz_aware_fixture2)
- boxed, box_dtype = box # read from parametrized fixture
from dateutil.tz import tzlocal
if is_platform_windows() and tz_aware_fixture2 == tzlocal():
pytest.xfail("Cannot process fill_value with this dtype, see GH 24310")
- if dtype.tz == fill_dtype.tz and boxed:
- pytest.xfail("falsely upcasts")
# create array of given dtype; casts "1" to correct dtype
fill_value = pd.Series([10 ** 9], dtype=fill_dtype)[0]
@@ -705,43 +439,22 @@ def test_maybe_promote_datetimetz_with_datetimetz(
exp_val_for_scalar = fill_value
if dtype.tz == fill_dtype.tz:
expected_dtype = dtype
- exp_val_for_array = NaT
else:
expected_dtype = np.dtype(object)
- exp_val_for_array = np.nan
+ pytest.xfail("fails to cast to object")
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize("fill_value", [None, np.nan, NaT])
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(False, None)])
-def test_maybe_promote_datetimetz_with_na(tz_aware_fixture, fill_value, box):
+def test_maybe_promote_datetimetz_with_na(tz_aware_fixture, fill_value):
dtype = DatetimeTZDtype(tz=tz_aware_fixture)
- boxed, box_dtype = box # read from parametrized fixture
expected_dtype = dtype
exp_val_for_scalar = NaT
- exp_val_for_array = NaT
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize(
@@ -755,43 +468,23 @@ def test_maybe_promote_datetimetz_with_na(tz_aware_fixture, fill_value, box):
ids=["pd.Timestamp", "np.datetime64", "datetime.datetime", "datetime.date"],
)
def test_maybe_promote_any_numpy_dtype_with_datetimetz(
- any_numpy_dtype_reduced, tz_aware_fixture, fill_value, box
+ any_numpy_dtype_reduced, tz_aware_fixture, fill_value
):
dtype = np.dtype(any_numpy_dtype_reduced)
fill_dtype = DatetimeTZDtype(tz=tz_aware_fixture)
- boxed, box_dtype = box # read from parametrized fixture
fill_value = pd.Series([fill_value], dtype=fill_dtype)[0]
# filling any numpy dtype with datetimetz casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_timedelta64_with_any(
- timedelta64_dtype, any_numpy_dtype_reduced, box
-):
+def test_maybe_promote_timedelta64_with_any(timedelta64_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(timedelta64_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_timedelta64_dtype(fill_dtype):
- if box_dtype == object:
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and box_dtype is None:
- pytest.xfail("does not upcast to object")
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -801,21 +494,11 @@ def test_maybe_promote_timedelta64_with_any(
expected_dtype = dtype
# for timedelta dtypes, scalar values get cast to pd.Timedelta.value
exp_val_for_scalar = pd.Timedelta(fill_value).to_timedelta64()
- exp_val_for_array = np.timedelta64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize(
@@ -823,62 +506,26 @@ def test_maybe_promote_timedelta64_with_any(
[pd.Timedelta(days=1), np.timedelta64(24, "h"), datetime.timedelta(1)],
ids=["pd.Timedelta", "np.timedelta64", "datetime.timedelta"],
)
-# override parametrization of box to add special case for td_dtype
-@pytest.mark.parametrize(
- "box",
- [
- (True, None), # fill_value wrapped in array with default dtype
- # disabled due to too many xfails; see GH 23982 / 25425
- # (True, 'td_dtype'), # fill_value in array with explicit timedelta dtype
- (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value passed on as scalar
- ],
-)
def test_maybe_promote_any_with_timedelta64(
- any_numpy_dtype_reduced, timedelta64_dtype, fill_value, box
+ any_numpy_dtype_reduced, timedelta64_dtype, fill_value
):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
-
- if is_timedelta64_dtype(dtype):
- if boxed and (
- box_dtype == object
- or (box_dtype is None and not is_timedelta64_dtype(type(fill_value)))
- ):
- pytest.xfail("falsely upcasts to object")
- else:
- if boxed and box_dtype is None and is_timedelta64_dtype(type(fill_value)):
- pytest.xfail("does not upcast correctly")
-
- # special case for box_dtype
- box_dtype = np.dtype(timedelta64_dtype) if box_dtype == "td_dtype" else box_dtype
# filling anything but timedelta with timedelta casts to object
if is_timedelta64_dtype(dtype):
expected_dtype = dtype
# for timedelta dtypes, scalar values get cast to pd.Timedelta.value
exp_val_for_scalar = pd.Timedelta(fill_value).to_timedelta64()
- exp_val_for_array = np.timedelta64("NaT", "ns")
else:
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_string_with_any(string_dtype, any_numpy_dtype_reduced, box):
+def test_maybe_promote_string_with_any(string_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(string_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -886,61 +533,26 @@ def test_maybe_promote_string_with_any(string_dtype, any_numpy_dtype_reduced, bo
# filling string with anything casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-# override parametrization of box to add special case for str
-@pytest.mark.parametrize(
- "box",
- [
- # disabled due to too many xfails; see GH 23982 / 25425
- # (True, None), # fill_value wrapped in array with default dtype
- # (True, 'str'), # fill_value wrapped in array with generic string-dtype
- (True, object), # fill_value wrapped in array with object dtype
- (False, None), # fill_value passed on as scalar
- ],
-)
-def test_maybe_promote_any_with_string(any_numpy_dtype_reduced, string_dtype, box):
+def test_maybe_promote_any_with_string(any_numpy_dtype_reduced, string_dtype):
dtype = np.dtype(any_numpy_dtype_reduced)
- fill_dtype = np.dtype(string_dtype)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype
fill_value = "abc"
- # special case for box_dtype (cannot use fixture in parametrization)
- box_dtype = fill_dtype if box_dtype == "str" else box_dtype
-
# filling anything with a string casts to object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_object_with_any(object_dtype, any_numpy_dtype_reduced, box):
+def test_maybe_promote_object_with_any(object_dtype, any_numpy_dtype_reduced):
dtype = np.dtype(object_dtype)
fill_dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of given dtype; casts "1" to correct dtype
fill_value = np.array([1], dtype=fill_dtype)[0]
@@ -948,22 +560,12 @@ def test_maybe_promote_object_with_any(object_dtype, any_numpy_dtype_reduced, bo
# filling object with anything stays object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
-def test_maybe_promote_any_with_object(any_numpy_dtype_reduced, object_dtype, box):
+def test_maybe_promote_any_with_object(any_numpy_dtype_reduced, object_dtype):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
# create array of object dtype from a scalar value (i.e. passing
# dtypes.common.is_scalar), which can however not be cast to int/float etc.
@@ -972,27 +574,13 @@ def test_maybe_promote_any_with_object(any_numpy_dtype_reduced, object_dtype, bo
# filling object with anything stays object
expected_dtype = np.dtype(object)
exp_val_for_scalar = fill_value
- exp_val_for_array = np.nan
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize("fill_value", [None, np.nan, NaT])
-# override parametrization due to to many xfails; see GH 23982 / 25425
-@pytest.mark.parametrize("box", [(False, None)])
-def test_maybe_promote_any_numpy_dtype_with_na(
- any_numpy_dtype_reduced, fill_value, box
-):
+def test_maybe_promote_any_numpy_dtype_with_na(any_numpy_dtype_reduced, fill_value):
dtype = np.dtype(any_numpy_dtype_reduced)
- boxed, box_dtype = box # read from parametrized fixture
if is_integer_dtype(dtype) and fill_value is not NaT:
# integer + other missing value (np.nan / None) casts to float
@@ -1020,24 +608,7 @@ def test_maybe_promote_any_numpy_dtype_with_na(
expected_dtype = np.dtype(object)
exp_val_for_scalar = np.nan
- # array case has same expected_dtype; but returns corresponding na-marker
- if is_integer_dtype(expected_dtype):
- # integers cannot hold NaNs; maybe_promote_with_array returns None
- exp_val_for_array = None
- elif is_datetime_or_timedelta_dtype(expected_dtype):
- exp_val_for_array = expected_dtype.type("NaT", "ns")
- else: # expected_dtype = float / complex / object
- exp_val_for_array = np.nan
-
- _check_promote(
- dtype,
- fill_value,
- boxed,
- box_dtype,
- expected_dtype,
- exp_val_for_scalar,
- exp_val_for_array,
- )
+ _check_promote(dtype, fill_value, expected_dtype, exp_val_for_scalar)
@pytest.mark.parametrize("dim", [0, 2, 3])
@@ -1051,12 +622,18 @@ def test_maybe_promote_dimensions(any_numpy_dtype_reduced, dim):
for _ in range(dim):
fill_array = np.expand_dims(fill_array, 0)
- # test against 1-dimensional case
- expected_dtype, expected_missing_value = maybe_promote(
- dtype, np.array([1], dtype=dtype)
- )
+ if dtype != object:
+ # test against 1-dimensional case
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ maybe_promote(dtype, np.array([1], dtype=dtype))
- result_dtype, result_missing_value = maybe_promote(dtype, fill_array)
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ maybe_promote(dtype, fill_array)
- assert result_dtype == expected_dtype
- _assert_match(result_missing_value, expected_missing_value)
+ else:
+ expected_dtype, expected_missing_value = maybe_promote(
+ dtype, np.array([1], dtype=dtype)
+ )
+ result_dtype, result_missing_value = maybe_promote(dtype, fill_array)
+ assert result_dtype == expected_dtype
+ _assert_match(result_missing_value, expected_missing_value)
diff --git a/pandas/tests/series/test_analytics.py b/pandas/tests/series/test_analytics.py
index 457c976137c11..79eaeaf051d2e 100644
--- a/pandas/tests/series/test_analytics.py
+++ b/pandas/tests/series/test_analytics.py
@@ -1028,6 +1028,24 @@ def test_shift_int(self, datetime_series):
expected = ts.astype(float).shift(1)
tm.assert_series_equal(shifted, expected)
+ def test_shift_object_non_scalar_fill(self):
+ # shift requires scalar fill_value except for object dtype
+ ser = Series(range(3))
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ ser.shift(1, fill_value=[])
+
+ df = ser.to_frame()
+ with pytest.raises(ValueError, match="fill_value must be a scalar"):
+ df.shift(1, fill_value=np.arange(3))
+
+ obj_ser = ser.astype(object)
+ result = obj_ser.shift(1, fill_value={})
+ assert result[0] == {}
+
+ obj_df = obj_ser.to_frame()
+ result = obj_df.shift(1, fill_value={})
+ assert result.iloc[0, 0] == {}
+
def test_shift_categorical(self):
# GH 9416
s = pd.Series(["a", "b", "c", "d"], dtype="category")
| Partially reverts #29362 by allowing non-scalar fill_value for _object_ dtypes. i.e. in 0.25.3 `pd.Series(range(3), dtype=object).shift(1, fill_value={})` would work, #29362 broke that, and this restores it. Added `test_shift_object_non_scalar_fill` for this.
With the new restriction on `maybe_promote` in place, we can get rid of all the `box` tests and simplify test_promote a _ton_. This removes about 2500 tests. This also uncovers the fact that we were failing to run some of the non-box cases, which are now xfailed. | https://api.github.com/repos/pandas-dev/pandas/pulls/29416 | 2019-11-05T17:45:37Z | 2019-11-06T19:29:03Z | 2019-11-06T19:29:03Z | 2020-04-05T17:44:46Z |
Fixed SS03 errors | diff --git a/pandas/_libs/interval.pyx b/pandas/_libs/interval.pyx
index 1a712d0c4efa8..b13ce7c294f37 100644
--- a/pandas/_libs/interval.pyx
+++ b/pandas/_libs/interval.pyx
@@ -94,7 +94,7 @@ cdef class IntervalMixin:
@property
def mid(self):
"""
- Return the midpoint of the Interval
+ Return the midpoint of the Interval.
"""
try:
return 0.5 * (self.left + self.right)
@@ -104,7 +104,9 @@ cdef class IntervalMixin:
@property
def length(self):
- """Return the length of the Interval"""
+ """
+ Return the length of the Interval.
+ """
return self.right - self.left
@property
@@ -283,15 +285,19 @@ cdef class Interval(IntervalMixin):
_typ = "interval"
cdef readonly object left
- """Left bound for the interval"""
+ """
+ Left bound for the interval.
+ """
cdef readonly object right
- """Right bound for the interval"""
+ """
+ Right bound for the interval.
+ """
cdef readonly str closed
"""
Whether the interval is closed on the left-side, right-side, both or
- neither
+ neither.
"""
def __init__(self, left, right, str closed='right'):
diff --git a/pandas/_libs/tslibs/nattype.pyx b/pandas/_libs/tslibs/nattype.pyx
index 0bd4b78d51e4e..241aff0e19112 100644
--- a/pandas/_libs/tslibs/nattype.pyx
+++ b/pandas/_libs/tslibs/nattype.pyx
@@ -464,7 +464,7 @@ class NaTType(_NaT):
"""
Timestamp.combine(date, time)
- date, time -> datetime with same date and time fields
+ date, time -> datetime with same date and time fields.
"""
)
utcnow = _make_error_func('utcnow', # noqa:E128
@@ -503,8 +503,8 @@ class NaTType(_NaT):
"""
Timestamp.fromordinal(ordinal, freq=None, tz=None)
- passed an ordinal, translate and convert to a ts
- note: by definition there cannot be any tz info on the ordinal itself
+ Passed an ordinal, translate and convert to a ts.
+ Note: by definition there cannot be any tz info on the ordinal itself.
Parameters
----------
diff --git a/pandas/_libs/tslibs/period.pyx b/pandas/_libs/tslibs/period.pyx
index aed64aff14e0a..e297d11c5144d 100644
--- a/pandas/_libs/tslibs/period.pyx
+++ b/pandas/_libs/tslibs/period.pyx
@@ -2244,7 +2244,7 @@ cdef class _Period:
containing one or several directives. The method recognizes the same
directives as the :func:`time.strftime` function of the standard Python
distribution, as well as the specific additional directives ``%f``,
- ``%F``, ``%q``. (formatting & docs originally from scikits.timeries)
+ ``%F``, ``%q``. (formatting & docs originally from scikits.timeries).
+-----------+--------------------------------+-------+
| Directive | Meaning | Notes |
diff --git a/pandas/_libs/tslibs/timestamps.pyx b/pandas/_libs/tslibs/timestamps.pyx
index 50a71d062c63f..317dc769636fb 100644
--- a/pandas/_libs/tslibs/timestamps.pyx
+++ b/pandas/_libs/tslibs/timestamps.pyx
@@ -242,8 +242,8 @@ class Timestamp(_Timestamp):
"""
Timestamp.fromordinal(ordinal, freq=None, tz=None)
- passed an ordinal, translate and convert to a ts
- note: by definition there cannot be any tz info on the ordinal itself
+ Passed an ordinal, translate and convert to a ts.
+ Note: by definition there cannot be any tz info on the ordinal itself.
Parameters
----------
@@ -333,7 +333,7 @@ class Timestamp(_Timestamp):
"""
Timestamp.combine(date, time)
- date, time -> datetime with same date and time fields
+ date, time -> datetime with same date and time fields.
"""
return cls(datetime.combine(date, time))
@@ -601,7 +601,7 @@ timedelta}, default 'raise'
@property
def dayofweek(self):
"""
- Return day of whe week.
+ Return day of the week.
"""
return self.weekday()
| Fixed SS03 errors for:
`pandas.Timestamp.combine`; `pandas.Timestamp.fromordinal`; `pandas.Period.strftime`; `pandas.Interval.closed`; `pandas.Interval.left`; `pandas.Interval.length`; `pandas.Interval.mid`; `pandas.Interval.right`.
- [x] xref to #29315
- [ ] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
Similar to #28053 I could not find methods for `pandas.Timestamp.isoweekday` or `pandas.Timestamp.weekday`. This was not resolved in #28053. I think this may be due to docstrings in the original python datetime file.
Timestamp import _Timestamp which imports datetime. Neither Timestamp nor _Timestamp have `isoweekday` or `weekday` methods.
@datapythonista | https://api.github.com/repos/pandas-dev/pandas/pulls/29410 | 2019-11-05T08:24:51Z | 2019-11-05T15:19:21Z | 2019-11-05T15:19:21Z | 2019-11-06T06:37:57Z |
CLN: assorted cleanups | diff --git a/pandas/_libs/sparse.pyx b/pandas/_libs/sparse.pyx
index 4906e45c884e9..6abaaca010b00 100644
--- a/pandas/_libs/sparse.pyx
+++ b/pandas/_libs/sparse.pyx
@@ -597,7 +597,7 @@ cdef class BlockIndex(SparseIndex):
result = np.empty(other.npoints, dtype=np.float64)
- for 0 <= i < other.nblocks:
+ for i in range(other.nblocks):
ocur = olocs[i]
ocurlen = olens[i]
@@ -746,9 +746,6 @@ cdef class BlockUnion(BlockMerge):
nend = xend[xi]
- # print 'here xi=%d, yi=%d, mode=%d, nend=%d' % (self.xi, self.yi,
- # mode, nend)
-
# done with y?
if yi == ynblocks:
self._set_current_indices(xi + 1, yi, mode)
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 009e83b861523..1e38dde2096ba 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -1124,10 +1124,6 @@ def _decide_output_index(self, output, labels):
output_keys = labels
else:
output_keys = sorted(output)
- try:
- output_keys.sort()
- except TypeError:
- pass
if isinstance(labels, MultiIndex):
output_keys = MultiIndex.from_tuples(output_keys, names=labels.names)
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 642b1e93a057a..59b118431cfc9 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -1092,9 +1092,8 @@ def result_to_bool(result: np.ndarray, inference: Type) -> np.ndarray:
return self._get_cythonized_result(
"group_any_all",
- self.grouper,
aggregate=True,
- cython_dtype=np.uint8,
+ cython_dtype=np.dtype(np.uint8),
needs_values=True,
needs_mask=True,
pre_processing=objs_to_bool,
@@ -1305,7 +1304,7 @@ def size(self):
result = self.grouper.size()
if isinstance(self.obj, Series):
- result.name = getattr(self.obj, "name", None)
+ result.name = self.obj.name
return result
@classmethod
@@ -1586,9 +1585,8 @@ def _fill(self, direction, limit=None):
return self._get_cythonized_result(
"group_fillna_indexer",
- self.grouper,
needs_mask=True,
- cython_dtype=np.int64,
+ cython_dtype=np.dtype(np.int64),
result_is_index=True,
direction=direction,
limit=limit,
@@ -1882,11 +1880,10 @@ def post_processor(vals: np.ndarray, inference: Optional[Type]) -> np.ndarray:
if is_scalar(q):
return self._get_cythonized_result(
"group_quantile",
- self.grouper,
aggregate=True,
needs_values=True,
needs_mask=True,
- cython_dtype=np.float64,
+ cython_dtype=np.dtype(np.float64),
pre_processing=pre_processor,
post_processing=post_processor,
q=q,
@@ -1896,11 +1893,10 @@ def post_processor(vals: np.ndarray, inference: Optional[Type]) -> np.ndarray:
results = [
self._get_cythonized_result(
"group_quantile",
- self.grouper,
aggregate=True,
needs_values=True,
needs_mask=True,
- cython_dtype=np.float64,
+ cython_dtype=np.dtype(np.float64),
pre_processing=pre_processor,
post_processing=post_processor,
q=qi,
@@ -2167,14 +2163,13 @@ def cummax(self, axis=0, **kwargs):
def _get_cythonized_result(
self,
- how,
- grouper,
- aggregate=False,
- cython_dtype=None,
- needs_values=False,
- needs_mask=False,
- needs_ngroups=False,
- result_is_index=False,
+ how: str,
+ cython_dtype: np.dtype,
+ aggregate: bool = False,
+ needs_values: bool = False,
+ needs_mask: bool = False,
+ needs_ngroups: bool = False,
+ result_is_index: bool = False,
pre_processing=None,
post_processing=None,
**kwargs
@@ -2185,13 +2180,11 @@ def _get_cythonized_result(
Parameters
----------
how : str, Cythonized function name to be called
- grouper : Grouper object containing pertinent group info
+ cython_dtype : np.dtype
+ Type of the array that will be modified by the Cython call.
aggregate : bool, default False
Whether the result should be aggregated to match the number of
groups
- cython_dtype : default None
- Type of the array that will be modified by the Cython call. If
- `None`, the type will be inferred from the values of each slice
needs_values : bool, default False
Whether the values should be a part of the Cython call
signature
@@ -2234,8 +2227,10 @@ def _get_cythonized_result(
"Cannot use 'pre_processing' without specifying 'needs_values'!"
)
+ grouper = self.grouper
+
labels, _, ngroups = grouper.group_info
- output = collections.OrderedDict()
+ output = collections.OrderedDict() # type: dict
base_func = getattr(libgroupby, how)
for name, obj in self._iterate_slices():
@@ -2246,9 +2241,6 @@ def _get_cythonized_result(
else:
result_sz = len(values)
- if not cython_dtype:
- cython_dtype = values.dtype
-
result = np.zeros(result_sz, dtype=cython_dtype)
func = partial(base_func, result, labels)
inferences = None
@@ -2308,8 +2300,7 @@ def shift(self, periods=1, freq=None, axis=0, fill_value=None):
return self._get_cythonized_result(
"group_shift_indexer",
- self.grouper,
- cython_dtype=np.int64,
+ cython_dtype=np.dtype(np.int64),
needs_ngroups=True,
result_is_index=True,
periods=periods,
@@ -2478,11 +2469,13 @@ def _reindex_output(self, output):
@Appender(GroupBy.__doc__)
-def groupby(obj, by, **kwds):
+def groupby(obj: NDFrame, by, **kwds):
if isinstance(obj, Series):
from pandas.core.groupby.generic import SeriesGroupBy
- klass = SeriesGroupBy
+ klass = (
+ SeriesGroupBy
+ ) # type: Union[Type["SeriesGroupBy"], Type["DataFrameGroupBy"]]
elif isinstance(obj, DataFrame):
from pandas.core.groupby.generic import DataFrameGroupBy
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 7918e463c73ac..9bbe73c1851b5 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -592,13 +592,10 @@ def agg_series(self, obj, func):
return self._aggregate_series_pure_python(obj, func)
def _aggregate_series_fast(self, obj, func):
+ # At this point we have already checked that obj.index is not a MultiIndex
+ # and that obj is backed by an ndarray, not ExtensionArray
func = self._is_builtin_func(func)
- # TODO: pre-empt this, also pre-empt get_result raising TypError if we pass a EA
- # for EAs backed by ndarray we may have a performant workaround
- if obj.index._has_complex_internals:
- raise TypeError("Incompatible index for Cython grouper")
-
group_index, _, ngroups = self.group_info
# avoids object / Series creation overhead
@@ -842,15 +839,12 @@ def __iter__(self):
def _get_sorted_data(self):
return self.data.take(self.sort_idx, axis=self.axis)
- def _chop(self, sdata, slice_obj):
- raise AbstractMethodError(self)
-
- def apply(self, f):
+ def _chop(self, sdata, slice_obj: slice):
raise AbstractMethodError(self)
class SeriesSplitter(DataSplitter):
- def _chop(self, sdata, slice_obj):
+ def _chop(self, sdata, slice_obj: slice):
return sdata._get_values(slice_obj)
@@ -862,7 +856,7 @@ def fast_apply(self, f, names):
sdata = self._get_sorted_data()
return libreduction.apply_frame_axis0(sdata, f, names, starts, ends)
- def _chop(self, sdata, slice_obj):
+ def _chop(self, sdata, slice_obj: slice):
if self.axis == 0:
return sdata.iloc[slice_obj]
else:
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 5751ce6ea730e..c9697c530628a 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -4747,10 +4747,9 @@ def get_indexer_for(self, target, **kwargs):
def _maybe_promote(self, other):
# A hack, but it works
- from pandas import DatetimeIndex
- if self.inferred_type == "date" and isinstance(other, DatetimeIndex):
- return DatetimeIndex(self), other
+ if self.inferred_type == "date" and isinstance(other, ABCDatetimeIndex):
+ return type(other)(self), other
elif self.inferred_type == "boolean":
if not is_object_dtype(self.dtype):
return self.astype("object"), other.astype("object")
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index caaf55546189c..2e3f440573a0f 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -2179,7 +2179,9 @@ def drop(self, codes, level=None, errors="raise"):
mask = indexer == -1
if mask.any():
if errors != "ignore":
- raise ValueError("codes %s not contained in axis" % codes[mask])
+ raise ValueError(
+ "codes {codes} not contained in axis".format(codes=codes[mask])
+ )
except Exception:
pass
diff --git a/pandas/core/internals/construction.py b/pandas/core/internals/construction.py
index 4a8216cc73264..05a2803b3fc2f 100644
--- a/pandas/core/internals/construction.py
+++ b/pandas/core/internals/construction.py
@@ -167,6 +167,7 @@ def init_ndarray(values, index, columns, dtype=None, copy=False):
try:
values = values.astype(dtype)
except Exception as orig:
+ # e.g. ValueError when trying to cast object dtype to float64
raise ValueError(
"failed to cast to '{dtype}' (Exception "
"was: {orig})".format(dtype=dtype, orig=orig)
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index c11915c00c59d..39e00047ea968 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -478,7 +478,7 @@ def get_result(self):
self, method="concat"
)
- def _get_result_dim(self):
+ def _get_result_dim(self) -> int:
if self._is_series and self.axis == 1:
return 2
else:
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index 9845c570ca704..6ef13a62ee366 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -1948,13 +1948,13 @@ def _get_join_keys(llab, rlab, shape, sort):
return _get_join_keys(llab, rlab, shape, sort)
-def _should_fill(lname, rname):
+def _should_fill(lname, rname) -> bool:
if not isinstance(lname, str) or not isinstance(rname, str):
return True
return lname == rname
-def _any(x):
+def _any(x) -> bool:
return x is not None and com.any_not_none(*x)
diff --git a/pandas/core/reshape/pivot.py b/pandas/core/reshape/pivot.py
index d653dd87308cf..404292fe4d539 100644
--- a/pandas/core/reshape/pivot.py
+++ b/pandas/core/reshape/pivot.py
@@ -620,7 +620,9 @@ def _normalize(table, normalize, margins, margins_name="All"):
if (margins_name not in table.iloc[-1, :].name) | (
margins_name != table.iloc[:, -1].name
):
- raise ValueError("{} not in pivoted DataFrame".format(margins_name))
+ raise ValueError(
+ "{mname} not in pivoted DataFrame".format(mname=margins_name)
+ )
column_margin = table.iloc[:-1, -1]
index_margin = table.iloc[-1, :-1]
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index d7eae1c543804..7537dd0ac2065 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -88,7 +88,7 @@ class _Unstacker:
def __init__(
self,
- values,
+ values: np.ndarray,
index,
level=-1,
value_columns=None,
@@ -985,7 +985,7 @@ def get_empty_frame(data):
else:
# PY2 embedded unicode, gh-22084
- def _make_col_name(prefix, prefix_sep, level):
+ def _make_col_name(prefix, prefix_sep, level) -> str:
fstr = "{prefix}{prefix_sep}{level}"
return fstr.format(prefix=prefix, prefix_sep=prefix_sep, level=level)
| https://api.github.com/repos/pandas-dev/pandas/pulls/29406 | 2019-11-04T23:26:44Z | 2019-11-05T15:14:55Z | 2019-11-05T15:14:54Z | 2019-11-07T18:36:53Z | |
API: Use object dtype for empty Series | diff --git a/doc/source/user_guide/missing_data.rst b/doc/source/user_guide/missing_data.rst
index 11957cfa265f5..1cc485a229123 100644
--- a/doc/source/user_guide/missing_data.rst
+++ b/doc/source/user_guide/missing_data.rst
@@ -190,7 +190,7 @@ The sum of an empty or all-NA Series or column of a DataFrame is 0.
pd.Series([np.nan]).sum()
- pd.Series([]).sum()
+ pd.Series([], dtype="float64").sum()
The product of an empty or all-NA Series or column of a DataFrame is 1.
@@ -198,7 +198,7 @@ The product of an empty or all-NA Series or column of a DataFrame is 1.
pd.Series([np.nan]).prod()
- pd.Series([]).prod()
+ pd.Series([], dtype="float64").prod()
NA values in GroupBy
diff --git a/doc/source/user_guide/scale.rst b/doc/source/user_guide/scale.rst
index ba213864ec469..0611c6334937f 100644
--- a/doc/source/user_guide/scale.rst
+++ b/doc/source/user_guide/scale.rst
@@ -358,6 +358,7 @@ results will fit in memory, so we can safely call ``compute`` without running
out of memory. At that point it's just a regular pandas object.
.. ipython:: python
+ :okwarning:
@savefig dask_resample.png
ddf[['x', 'y']].resample("1D").mean().cumsum().compute().plot()
diff --git a/doc/source/whatsnew/v0.19.0.rst b/doc/source/whatsnew/v0.19.0.rst
index 61a65415f6b57..6f6446c3f74e1 100644
--- a/doc/source/whatsnew/v0.19.0.rst
+++ b/doc/source/whatsnew/v0.19.0.rst
@@ -707,6 +707,7 @@ A ``Series`` will now correctly promote its dtype for assignment with incompat v
.. ipython:: python
+ :okwarning:
s = pd.Series()
diff --git a/doc/source/whatsnew/v0.21.0.rst b/doc/source/whatsnew/v0.21.0.rst
index a9c7937308204..f33943e423b25 100644
--- a/doc/source/whatsnew/v0.21.0.rst
+++ b/doc/source/whatsnew/v0.21.0.rst
@@ -428,6 +428,7 @@ Note that this also changes the sum of an empty ``Series``. Previously this alwa
but for consistency with the all-NaN case, this was changed to return NaN as well:
.. ipython:: python
+ :okwarning:
pd.Series([]).sum()
diff --git a/doc/source/whatsnew/v0.22.0.rst b/doc/source/whatsnew/v0.22.0.rst
index ea36b35d61740..75949a90d09a6 100644
--- a/doc/source/whatsnew/v0.22.0.rst
+++ b/doc/source/whatsnew/v0.22.0.rst
@@ -55,6 +55,7 @@ The default sum for empty or all-*NA* ``Series`` is now ``0``.
*pandas 0.22.0*
.. ipython:: python
+ :okwarning:
pd.Series([]).sum()
pd.Series([np.nan]).sum()
@@ -67,6 +68,7 @@ pandas 0.20.3 without bottleneck, or pandas 0.21.x), use the ``min_count``
keyword.
.. ipython:: python
+ :okwarning:
pd.Series([]).sum(min_count=1)
@@ -85,6 +87,7 @@ required for a non-NA sum or product.
returning ``1`` instead.
.. ipython:: python
+ :okwarning:
pd.Series([]).prod()
pd.Series([np.nan]).prod()
diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 4ce4c12483b36..771b3e484f67c 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -366,6 +366,23 @@ When :class:`Categorical` contains ``np.nan``,
pd.Categorical([1, 2, np.nan], ordered=True).min()
+
+Default dtype of empty :class:`pandas.Series`
+^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
+
+Initialising an empty :class:`pandas.Series` without specifying a dtype will raise a `DeprecationWarning` now
+(:issue:`17261`). The default dtype will change from ``float64`` to ``object`` in future releases so that it is
+consistent with the behaviour of :class:`DataFrame` and :class:`Index`.
+
+*pandas 1.0.0*
+
+.. code-block:: ipython
+
+ In [1]: pd.Series()
+ Out[2]:
+ DeprecationWarning: The default dtype for empty Series will be 'object' instead of 'float64' in a future version. Specify a dtype explicitly to silence this warning.
+ Series([], dtype: float64)
+
.. _whatsnew_1000.api_breaking.deps:
Increased minimum versions for dependencies
@@ -494,7 +511,7 @@ Removal of prior version deprecations/changes
Previously, pandas would register converters with matplotlib as a side effect of importing pandas (:issue:`18720`).
This changed the output of plots made via matplotlib plots after pandas was imported, even if you were using
-matplotlib directly rather than rather than :meth:`~DataFrame.plot`.
+matplotlib directly rather than :meth:`~DataFrame.plot`.
To use pandas formatters with a matplotlib plot, specify
diff --git a/pandas/compat/pickle_compat.py b/pandas/compat/pickle_compat.py
index aeec5e8a0400a..7dfed94482a05 100644
--- a/pandas/compat/pickle_compat.py
+++ b/pandas/compat/pickle_compat.py
@@ -64,7 +64,7 @@ def __new__(cls) -> "Series": # type: ignore
stacklevel=6,
)
- return Series()
+ return Series(dtype=object)
class _LoadSparseFrame:
diff --git a/pandas/core/apply.py b/pandas/core/apply.py
index 8c49b2b803241..ef3d8cd53596b 100644
--- a/pandas/core/apply.py
+++ b/pandas/core/apply.py
@@ -15,6 +15,8 @@
)
from pandas.core.dtypes.generic import ABCMultiIndex, ABCSeries
+from pandas.core.construction import create_series_with_explicit_dtype
+
if TYPE_CHECKING:
from pandas import DataFrame, Series, Index
@@ -203,7 +205,7 @@ def apply_empty_result(self):
if not should_reduce:
try:
- r = self.f(Series([]))
+ r = self.f(Series([], dtype=np.float64))
except Exception:
pass
else:
@@ -211,7 +213,7 @@ def apply_empty_result(self):
if should_reduce:
if len(self.agg_axis):
- r = self.f(Series([]))
+ r = self.f(Series([], dtype=np.float64))
else:
r = np.nan
@@ -346,6 +348,7 @@ def apply_series_generator(self) -> Tuple[ResType, "Index"]:
def wrap_results(
self, results: ResType, res_index: "Index"
) -> Union["Series", "DataFrame"]:
+ from pandas import Series
# see if we can infer the results
if len(results) > 0 and 0 in results and is_sequence(results[0]):
@@ -353,7 +356,17 @@ def wrap_results(
return self.wrap_results_for_axis(results, res_index)
# dict of scalars
- result = self.obj._constructor_sliced(results)
+
+ # the default dtype of an empty Series will be `object`, but this
+ # code can be hit by df.mean() where the result should have dtype
+ # float64 even if it's an empty Series.
+ constructor_sliced = self.obj._constructor_sliced
+ if constructor_sliced is Series:
+ result = create_series_with_explicit_dtype(
+ results, dtype_if_empty=np.float64
+ )
+ else:
+ result = constructor_sliced(results)
result.index = res_index
return result
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 5e613849ba8d5..b7216d2a70ee6 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -34,6 +34,7 @@
from pandas.core.accessor import DirNamesMixin
from pandas.core.algorithms import duplicated, unique1d, value_counts
from pandas.core.arrays import ExtensionArray
+from pandas.core.construction import create_series_with_explicit_dtype
import pandas.core.nanops as nanops
_shared_docs: Dict[str, str] = dict()
@@ -1132,9 +1133,14 @@ def _map_values(self, mapper, na_action=None):
# convert to an Series for efficiency.
# we specify the keys here to handle the
# possibility that they are tuples
- from pandas import Series
- mapper = Series(mapper)
+ # The return value of mapping with an empty mapper is
+ # expected to be pd.Series(np.nan, ...). As np.nan is
+ # of dtype float64 the return value of this method should
+ # be float64 as well
+ mapper = create_series_with_explicit_dtype(
+ mapper, dtype_if_empty=np.float64
+ )
if isinstance(mapper, ABCSeries):
# Since values were input this means we came from either
diff --git a/pandas/core/construction.py b/pandas/core/construction.py
index dc537d50b3419..b03c69d865301 100644
--- a/pandas/core/construction.py
+++ b/pandas/core/construction.py
@@ -4,7 +4,7 @@
These should not depend on core.internals.
"""
-from typing import Optional, Sequence, Union, cast
+from typing import TYPE_CHECKING, Any, Optional, Sequence, Union, cast
import numpy as np
import numpy.ma as ma
@@ -44,8 +44,13 @@
)
from pandas.core.dtypes.missing import isna
+from pandas._typing import ArrayLike, Dtype
import pandas.core.common as com
+if TYPE_CHECKING:
+ from pandas.core.series import Series # noqa: F401
+ from pandas.core.index import Index # noqa: F401
+
def array(
data: Sequence[object],
@@ -565,3 +570,62 @@ def _try_cast(
else:
subarr = np.array(arr, dtype=object, copy=copy)
return subarr
+
+
+def is_empty_data(data: Any) -> bool:
+ """
+ Utility to check if a Series is instantiated with empty data,
+ which does not contain dtype information.
+
+ Parameters
+ ----------
+ data : array-like, Iterable, dict, or scalar value
+ Contains data stored in Series.
+
+ Returns
+ -------
+ bool
+ """
+ is_none = data is None
+ is_list_like_without_dtype = is_list_like(data) and not hasattr(data, "dtype")
+ is_simple_empty = is_list_like_without_dtype and not data
+ return is_none or is_simple_empty
+
+
+def create_series_with_explicit_dtype(
+ data: Any = None,
+ index: Optional[Union[ArrayLike, "Index"]] = None,
+ dtype: Optional[Dtype] = None,
+ name: Optional[str] = None,
+ copy: bool = False,
+ fastpath: bool = False,
+ dtype_if_empty: Dtype = object,
+) -> "Series":
+ """
+ Helper to pass an explicit dtype when instantiating an empty Series.
+
+ This silences a DeprecationWarning described in GitHub-17261.
+
+ Parameters
+ ----------
+ data : Mirrored from Series.__init__
+ index : Mirrored from Series.__init__
+ dtype : Mirrored from Series.__init__
+ name : Mirrored from Series.__init__
+ copy : Mirrored from Series.__init__
+ fastpath : Mirrored from Series.__init__
+ dtype_if_empty : str, numpy.dtype, or ExtensionDtype
+ This dtype will be passed explicitly if an empty Series will
+ be instantiated.
+
+ Returns
+ -------
+ Series
+ """
+ from pandas.core.series import Series
+
+ if is_empty_data(data) and dtype is None:
+ dtype = dtype_if_empty
+ return Series(
+ data=data, index=index, dtype=dtype, name=name, copy=copy, fastpath=fastpath
+ )
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 601dac3a1208b..c1616efabcdba 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -7956,7 +7956,7 @@ def quantile(self, q=0.5, axis=0, numeric_only=True, interpolation="linear"):
cols = Index([], name=self.columns.name)
if is_list_like(q):
return self._constructor([], index=q, columns=cols)
- return self._constructor_sliced([], index=cols, name=q)
+ return self._constructor_sliced([], index=cols, name=q, dtype=np.float64)
result = data._data.quantile(
qs=q, axis=1, interpolation=interpolation, transposed=is_transposed
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index 9aecd97194aad..efdcfa7edbba3 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -72,6 +72,7 @@
import pandas.core.algorithms as algos
from pandas.core.base import PandasObject, SelectionMixin
import pandas.core.common as com
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.core.index import (
Index,
InvalidIndexError,
@@ -6042,9 +6043,9 @@ def fillna(
if self.ndim == 1:
if isinstance(value, (dict, ABCSeries)):
- from pandas import Series
-
- value = Series(value)
+ value = create_series_with_explicit_dtype(
+ value, dtype_if_empty=object
+ )
elif not is_list_like(value):
pass
else:
@@ -6996,7 +6997,7 @@ def asof(self, where, subset=None):
if not is_series:
from pandas import Series
- return Series(index=self.columns, name=where)
+ return Series(index=self.columns, name=where, dtype=np.float64)
return np.nan
# It's always much faster to use a *while* loop here for
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 4726cdfb05a70..9bb0b8de9ba71 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -51,6 +51,7 @@
import pandas.core.algorithms as algorithms
from pandas.core.base import DataError, SpecificationError
import pandas.core.common as com
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.core.frame import DataFrame
from pandas.core.generic import ABCDataFrame, ABCSeries, NDFrame, _shared_docs
from pandas.core.groupby import base
@@ -259,7 +260,9 @@ def aggregate(self, func=None, *args, **kwargs):
result = self._aggregate_named(func, *args, **kwargs)
index = Index(sorted(result), name=self.grouper.names[0])
- ret = Series(result, index=index)
+ ret = create_series_with_explicit_dtype(
+ result, index=index, dtype_if_empty=object
+ )
if not self.as_index: # pragma: no cover
print("Warning, ignoring as_index=True")
@@ -407,7 +410,7 @@ def _wrap_transformed_output(
def _wrap_applied_output(self, keys, values, not_indexed_same=False):
if len(keys) == 0:
# GH #6265
- return Series([], name=self._selection_name, index=keys)
+ return Series([], name=self._selection_name, index=keys, dtype=np.float64)
def _get_index() -> Index:
if self.grouper.nkeys > 1:
@@ -493,7 +496,7 @@ def _transform_general(self, func, *args, **kwargs):
result = concat(results).sort_index()
else:
- result = Series()
+ result = Series(dtype=np.float64)
# we will only try to coerce the result type if
# we have a numeric dtype, as these are *always* user-defined funcs
@@ -1205,10 +1208,18 @@ def first_not_none(values):
if v is None:
return DataFrame()
elif isinstance(v, NDFrame):
- values = [
- x if x is not None else v._constructor(**v._construct_axes_dict())
- for x in values
- ]
+
+ # this is to silence a DeprecationWarning
+ # TODO: Remove when default dtype of empty Series is object
+ kwargs = v._construct_axes_dict()
+ if v._constructor is Series:
+ backup = create_series_with_explicit_dtype(
+ **kwargs, dtype_if_empty=object
+ )
+ else:
+ backup = v._constructor(**kwargs)
+
+ values = [x if (x is not None) else backup for x in values]
v = values[0]
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 537a960f7d463..efa3d33a2a79a 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -54,7 +54,12 @@
from pandas.core.arrays.categorical import Categorical, CategoricalAccessor
from pandas.core.arrays.sparse import SparseAccessor
import pandas.core.common as com
-from pandas.core.construction import extract_array, sanitize_array
+from pandas.core.construction import (
+ create_series_with_explicit_dtype,
+ extract_array,
+ is_empty_data,
+ sanitize_array,
+)
from pandas.core.index import (
Float64Index,
Index,
@@ -177,7 +182,6 @@ class Series(base.IndexOpsMixin, generic.NDFrame):
def __init__(
self, data=None, index=None, dtype=None, name=None, copy=False, fastpath=False
):
-
# we are called internally, so short-circuit
if fastpath:
@@ -191,6 +195,18 @@ def __init__(
else:
+ if is_empty_data(data) and dtype is None:
+ # gh-17261
+ warnings.warn(
+ "The default dtype for empty Series will be 'object' instead"
+ " of 'float64' in a future version. Specify a dtype explicitly"
+ " to silence this warning.",
+ DeprecationWarning,
+ stacklevel=2,
+ )
+ # uncomment the line below when removing the DeprecationWarning
+ # dtype = np.dtype(object)
+
if index is not None:
index = ensure_index(index)
@@ -330,7 +346,11 @@ def _init_dict(self, data, index=None, dtype=None):
keys, values = [], []
# Input is now list-like, so rely on "standard" construction:
- s = Series(values, index=keys, dtype=dtype)
+
+ # TODO: passing np.float64 to not break anything yet. See GH-17261
+ s = create_series_with_explicit_dtype(
+ values, index=keys, dtype=dtype, dtype_if_empty=np.float64
+ )
# Now we just make sure the order is respected, if any
if data and index is not None:
diff --git a/pandas/core/tools/datetimes.py b/pandas/core/tools/datetimes.py
index 453d1cca2e085..3dfafd04dff0a 100644
--- a/pandas/core/tools/datetimes.py
+++ b/pandas/core/tools/datetimes.py
@@ -145,7 +145,8 @@ def _maybe_cache(arg, format, cache, convert_listlike):
"""
from pandas import Series
- cache_array = Series()
+ cache_array = Series(dtype=object)
+
if cache:
# Perform a quicker unique check
if not should_cache(arg):
diff --git a/pandas/io/html.py b/pandas/io/html.py
index b8cb6679a9562..c629c0bab7779 100644
--- a/pandas/io/html.py
+++ b/pandas/io/html.py
@@ -14,7 +14,7 @@
from pandas.core.dtypes.common import is_list_like
-from pandas import Series
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.io.common import _is_url, _validate_header_arg, urlopen
from pandas.io.formats.printing import pprint_thing
@@ -762,7 +762,8 @@ def _parse_tfoot_tr(self, table):
def _expand_elements(body):
- lens = Series([len(elem) for elem in body])
+ data = [len(elem) for elem in body]
+ lens = create_series_with_explicit_dtype(data, dtype_if_empty=object)
lens_max = lens.max()
not_max = lens[lens != lens_max]
diff --git a/pandas/io/json/_json.py b/pandas/io/json/_json.py
index 89d5b52ffbf1e..30c1c2d59e983 100644
--- a/pandas/io/json/_json.py
+++ b/pandas/io/json/_json.py
@@ -1,4 +1,5 @@
from collections import OrderedDict
+import functools
from io import StringIO
from itertools import islice
import os
@@ -14,6 +15,7 @@
from pandas import DataFrame, MultiIndex, Series, isna, to_datetime
from pandas._typing import JSONSerializable
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.core.reshape.concat import concat
from pandas.io.common import (
@@ -1006,44 +1008,34 @@ class SeriesParser(Parser):
_split_keys = ("name", "index", "data")
def _parse_no_numpy(self):
+ data = loads(self.json, precise_float=self.precise_float)
- json = self.json
- orient = self.orient
- if orient == "split":
- decoded = {
- str(k): v
- for k, v in loads(json, precise_float=self.precise_float).items()
- }
+ if self.orient == "split":
+ decoded = {str(k): v for k, v in data.items()}
self.check_keys_split(decoded)
- self.obj = Series(dtype=None, **decoded)
+ self.obj = create_series_with_explicit_dtype(**decoded)
else:
- self.obj = Series(loads(json, precise_float=self.precise_float), dtype=None)
+ self.obj = create_series_with_explicit_dtype(data, dtype_if_empty=object)
def _parse_numpy(self):
+ load_kwargs = {
+ "dtype": None,
+ "numpy": True,
+ "precise_float": self.precise_float,
+ }
+ if self.orient in ["columns", "index"]:
+ load_kwargs["labelled"] = True
+ loads_ = functools.partial(loads, **load_kwargs)
+ data = loads_(self.json)
- json = self.json
- orient = self.orient
- if orient == "split":
- decoded = loads(
- json, dtype=None, numpy=True, precise_float=self.precise_float
- )
- decoded = {str(k): v for k, v in decoded.items()}
+ if self.orient == "split":
+ decoded = {str(k): v for k, v in data.items()}
self.check_keys_split(decoded)
- self.obj = Series(**decoded)
- elif orient == "columns" or orient == "index":
- self.obj = Series(
- *loads(
- json,
- dtype=None,
- numpy=True,
- labelled=True,
- precise_float=self.precise_float,
- )
- )
+ self.obj = create_series_with_explicit_dtype(**decoded)
+ elif self.orient in ["columns", "index"]:
+ self.obj = create_series_with_explicit_dtype(*data, dtype_if_empty=object)
else:
- self.obj = Series(
- loads(json, dtype=None, numpy=True, precise_float=self.precise_float)
- )
+ self.obj = create_series_with_explicit_dtype(data, dtype_if_empty=object)
def _try_convert_types(self):
if self.obj is None:
diff --git a/pandas/plotting/_matplotlib/boxplot.py b/pandas/plotting/_matplotlib/boxplot.py
index 7bcca659ee3f6..deeeb0016142c 100644
--- a/pandas/plotting/_matplotlib/boxplot.py
+++ b/pandas/plotting/_matplotlib/boxplot.py
@@ -114,7 +114,7 @@ def maybe_color_bp(self, bp):
def _make_plot(self):
if self.subplots:
- self._return_obj = pd.Series()
+ self._return_obj = pd.Series(dtype=object)
for i, (label, y) in enumerate(self._iter_data()):
ax = self._get_ax(i)
@@ -405,7 +405,8 @@ def boxplot_frame_groupby(
)
axes = _flatten(axes)
- ret = pd.Series()
+ ret = pd.Series(dtype=object)
+
for (key, group), ax in zip(grouped, axes):
d = group.boxplot(
ax=ax, column=column, fontsize=fontsize, rot=rot, grid=grid, **kwds
diff --git a/pandas/tests/arrays/categorical/test_algos.py b/pandas/tests/arrays/categorical/test_algos.py
index dce3c4e4d5e98..da142fa0bd63c 100644
--- a/pandas/tests/arrays/categorical/test_algos.py
+++ b/pandas/tests/arrays/categorical/test_algos.py
@@ -77,7 +77,7 @@ def test_replace(to_replace, value, result):
tm.assert_categorical_equal(cat, expected)
-@pytest.mark.parametrize("empty", [[], pd.Series(), np.array([])])
+@pytest.mark.parametrize("empty", [[], pd.Series(dtype=object), np.array([])])
def test_isin_empty(empty):
s = pd.Categorical(["a", "b"])
expected = np.array([False, False], dtype=bool)
diff --git a/pandas/tests/dtypes/test_inference.py b/pandas/tests/dtypes/test_inference.py
index 75e86a2ee7ecc..3fb4e291d7d91 100644
--- a/pandas/tests/dtypes/test_inference.py
+++ b/pandas/tests/dtypes/test_inference.py
@@ -78,7 +78,7 @@ def coerce(request):
((x for x in [1, 2]), True, "generator"),
((_ for _ in []), True, "generator-empty"),
(Series([1]), True, "Series"),
- (Series([]), True, "Series-empty"),
+ (Series([], dtype=object), True, "Series-empty"),
(Series(["a"]).str, True, "StringMethods"),
(Series([], dtype="O").str, True, "StringMethods-empty"),
(Index([1]), True, "Index"),
@@ -139,7 +139,7 @@ def __getitem__(self):
def test_is_array_like():
- assert inference.is_array_like(Series([]))
+ assert inference.is_array_like(Series([], dtype=object))
assert inference.is_array_like(Series([1, 2]))
assert inference.is_array_like(np.array(["a", "b"]))
assert inference.is_array_like(Index(["2016-01-01"]))
@@ -165,7 +165,7 @@ class DtypeList(list):
{"a": 1},
{1, "a"},
Series([1]),
- Series([]),
+ Series([], dtype=object),
Series(["a"]).str,
(x for x in range(5)),
],
@@ -1404,7 +1404,7 @@ def test_is_scalar_pandas_scalars(self):
assert is_scalar(DateOffset(days=1))
def test_is_scalar_pandas_containers(self):
- assert not is_scalar(Series())
+ assert not is_scalar(Series(dtype=object))
assert not is_scalar(Series([1]))
assert not is_scalar(DataFrame())
assert not is_scalar(DataFrame([[1]]))
diff --git a/pandas/tests/dtypes/test_missing.py b/pandas/tests/dtypes/test_missing.py
index 89474cf8fa953..5e7c6e4b48682 100644
--- a/pandas/tests/dtypes/test_missing.py
+++ b/pandas/tests/dtypes/test_missing.py
@@ -90,7 +90,8 @@ def test_isna_isnull(self, isna_f):
assert not isna_f(-np.inf)
# type
- assert not isna_f(type(pd.Series()))
+ assert not isna_f(type(pd.Series(dtype=object)))
+ assert not isna_f(type(pd.Series(dtype=np.float64)))
assert not isna_f(type(pd.DataFrame()))
# series
diff --git a/pandas/tests/frame/indexing/test_indexing.py b/pandas/tests/frame/indexing/test_indexing.py
index 9a7cd4ace686f..716be92ebca3f 100644
--- a/pandas/tests/frame/indexing/test_indexing.py
+++ b/pandas/tests/frame/indexing/test_indexing.py
@@ -2572,7 +2572,7 @@ def test_xs_corner(self):
# no columns but Index(dtype=object)
df = DataFrame(index=["a", "b", "c"])
result = df.xs("a")
- expected = Series([], name="a", index=pd.Index([], dtype=object))
+ expected = Series([], name="a", index=pd.Index([]), dtype=np.float64)
tm.assert_series_equal(result, expected)
def test_xs_duplicates(self):
diff --git a/pandas/tests/frame/test_analytics.py b/pandas/tests/frame/test_analytics.py
index 005ca8d95182e..5c14c3cd2a2b5 100644
--- a/pandas/tests/frame/test_analytics.py
+++ b/pandas/tests/frame/test_analytics.py
@@ -1067,13 +1067,13 @@ def test_mean_mixed_datetime_numeric(self, tz):
tm.assert_series_equal(result, expected)
@pytest.mark.parametrize("tz", [None, "UTC"])
- def test_mean_excludeds_datetimes(self, tz):
+ def test_mean_excludes_datetimes(self, tz):
# https://github.com/pandas-dev/pandas/issues/24752
# Our long-term desired behavior is unclear, but the behavior in
# 0.24.0rc1 was buggy.
df = pd.DataFrame({"A": [pd.Timestamp("2000", tz=tz)] * 2})
result = df.mean()
- expected = pd.Series()
+ expected = pd.Series(dtype=np.float64)
tm.assert_series_equal(result, expected)
def test_mean_mixed_string_decimal(self):
@@ -1907,7 +1907,7 @@ def test_isin(self):
expected = DataFrame([df.loc[s].isin(other) for s in df.index])
tm.assert_frame_equal(result, expected)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_isin_empty(self, empty):
# GH 16991
df = DataFrame({"A": ["a", "b", "c"], "B": ["a", "e", "f"]})
diff --git a/pandas/tests/frame/test_apply.py b/pandas/tests/frame/test_apply.py
index 26a3c738750ca..eb98bdc49f976 100644
--- a/pandas/tests/frame/test_apply.py
+++ b/pandas/tests/frame/test_apply.py
@@ -105,13 +105,15 @@ def test_apply_with_reduce_empty(self):
result = empty_frame.apply(x.append, axis=1, result_type="expand")
tm.assert_frame_equal(result, empty_frame)
result = empty_frame.apply(x.append, axis=1, result_type="reduce")
- tm.assert_series_equal(result, Series([], index=pd.Index([], dtype=object)))
+ expected = Series([], index=pd.Index([], dtype=object), dtype=np.float64)
+ tm.assert_series_equal(result, expected)
empty_with_cols = DataFrame(columns=["a", "b", "c"])
result = empty_with_cols.apply(x.append, axis=1, result_type="expand")
tm.assert_frame_equal(result, empty_with_cols)
result = empty_with_cols.apply(x.append, axis=1, result_type="reduce")
- tm.assert_series_equal(result, Series([], index=pd.Index([], dtype=object)))
+ expected = Series([], index=pd.Index([], dtype=object), dtype=np.float64)
+ tm.assert_series_equal(result, expected)
# Ensure that x.append hasn't been called
assert x == []
@@ -134,7 +136,7 @@ def test_nunique_empty(self):
tm.assert_series_equal(result, expected)
result = df.T.nunique()
- expected = Series([], index=pd.Index([]))
+ expected = Series([], index=pd.Index([]), dtype=np.float64)
tm.assert_series_equal(result, expected)
def test_apply_standard_nonunique(self):
@@ -1284,16 +1286,16 @@ def func(group_col):
_get_cython_table_params(
DataFrame(),
[
- ("sum", Series()),
- ("max", Series()),
- ("min", Series()),
+ ("sum", Series(dtype="float64")),
+ ("max", Series(dtype="float64")),
+ ("min", Series(dtype="float64")),
("all", Series(dtype=bool)),
("any", Series(dtype=bool)),
- ("mean", Series()),
- ("prod", Series()),
- ("std", Series()),
- ("var", Series()),
- ("median", Series()),
+ ("mean", Series(dtype="float64")),
+ ("prod", Series(dtype="float64")),
+ ("std", Series(dtype="float64")),
+ ("var", Series(dtype="float64")),
+ ("median", Series(dtype="float64")),
],
),
_get_cython_table_params(
diff --git a/pandas/tests/frame/test_arithmetic.py b/pandas/tests/frame/test_arithmetic.py
index 88bd5a4fedfae..f6e203afb0898 100644
--- a/pandas/tests/frame/test_arithmetic.py
+++ b/pandas/tests/frame/test_arithmetic.py
@@ -470,7 +470,7 @@ def test_arith_flex_series(self, simple_frame):
def test_arith_flex_zero_len_raises(self):
# GH 19522 passing fill_value to frame flex arith methods should
# raise even in the zero-length special cases
- ser_len0 = pd.Series([])
+ ser_len0 = pd.Series([], dtype=object)
df_len0 = pd.DataFrame(columns=["A", "B"])
df = pd.DataFrame([[1, 2], [3, 4]], columns=["A", "B"])
diff --git a/pandas/tests/frame/test_asof.py b/pandas/tests/frame/test_asof.py
index 9a7d806c79dc3..89be3779e5748 100644
--- a/pandas/tests/frame/test_asof.py
+++ b/pandas/tests/frame/test_asof.py
@@ -67,7 +67,9 @@ def test_missing(self, date_range_frame):
df = date_range_frame.iloc[:N].copy()
result = df.asof("1989-12-31")
- expected = Series(index=["A", "B"], name=Timestamp("1989-12-31"))
+ expected = Series(
+ index=["A", "B"], name=Timestamp("1989-12-31"), dtype=np.float64
+ )
tm.assert_series_equal(result, expected)
result = df.asof(to_datetime(["1989-12-31"]))
diff --git a/pandas/tests/frame/test_constructors.py b/pandas/tests/frame/test_constructors.py
index ce0ebdbe56354..08dbeb9e585f1 100644
--- a/pandas/tests/frame/test_constructors.py
+++ b/pandas/tests/frame/test_constructors.py
@@ -25,6 +25,7 @@
date_range,
isna,
)
+from pandas.core.construction import create_series_with_explicit_dtype
import pandas.util.testing as tm
MIXED_FLOAT_DTYPES = ["float16", "float32", "float64"]
@@ -1216,7 +1217,9 @@ def test_constructor_list_of_series(self):
OrderedDict([["a", 1.5], ["b", 3], ["c", 4]]),
OrderedDict([["b", 3], ["c", 4], ["d", 6]]),
]
- data = [Series(d) for d in data]
+ data = [
+ create_series_with_explicit_dtype(d, dtype_if_empty=object) for d in data
+ ]
result = DataFrame(data)
sdict = OrderedDict(zip(range(len(data)), data))
@@ -1226,7 +1229,7 @@ def test_constructor_list_of_series(self):
result2 = DataFrame(data, index=np.arange(6))
tm.assert_frame_equal(result, result2)
- result = DataFrame([Series()])
+ result = DataFrame([Series(dtype=object)])
expected = DataFrame(index=[0])
tm.assert_frame_equal(result, expected)
@@ -1450,7 +1453,7 @@ def test_constructor_Series_named(self):
DataFrame(s, columns=[1, 2])
# #2234
- a = Series([], name="x")
+ a = Series([], name="x", dtype=object)
df = DataFrame(a)
assert df.columns[0] == "x"
@@ -2356,11 +2359,11 @@ def test_from_records_series_list_dict(self):
def test_to_frame_with_falsey_names(self):
# GH 16114
- result = Series(name=0).to_frame().dtypes
- expected = Series({0: np.float64})
+ result = Series(name=0, dtype=object).to_frame().dtypes
+ expected = Series({0: object})
tm.assert_series_equal(result, expected)
- result = DataFrame(Series(name=0)).dtypes
+ result = DataFrame(Series(name=0, dtype=object)).dtypes
tm.assert_series_equal(result, expected)
@pytest.mark.parametrize("dtype", [None, "uint8", "category"])
diff --git a/pandas/tests/frame/test_dtypes.py b/pandas/tests/frame/test_dtypes.py
index 6709cdcb1eebf..d8d56e90a2f31 100644
--- a/pandas/tests/frame/test_dtypes.py
+++ b/pandas/tests/frame/test_dtypes.py
@@ -656,8 +656,8 @@ def test_astype_dict_like(self, dtype_class):
# GH 16717
# if dtypes provided is empty, the resulting DataFrame
# should be the same as the original DataFrame
- dt7 = dtype_class({})
- result = df.astype(dt7)
+ dt7 = dtype_class({}) if dtype_class is dict else dtype_class({}, dtype=object)
+ equiv = df.astype(dt7)
tm.assert_frame_equal(df, equiv)
tm.assert_frame_equal(df, original)
diff --git a/pandas/tests/frame/test_quantile.py b/pandas/tests/frame/test_quantile.py
index 78953d43677fc..5ca7dd32200ee 100644
--- a/pandas/tests/frame/test_quantile.py
+++ b/pandas/tests/frame/test_quantile.py
@@ -472,7 +472,7 @@ def test_quantile_empty_no_columns(self):
df = pd.DataFrame(pd.date_range("1/1/18", periods=5))
df.columns.name = "captain tightpants"
result = df.quantile(0.5)
- expected = pd.Series([], index=[], name=0.5)
+ expected = pd.Series([], index=[], name=0.5, dtype=np.float64)
expected.index.name = "captain tightpants"
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/frame/test_replace.py b/pandas/tests/frame/test_replace.py
index 434ea6ea7b4f0..3b01ae0c3c2e8 100644
--- a/pandas/tests/frame/test_replace.py
+++ b/pandas/tests/frame/test_replace.py
@@ -1251,7 +1251,7 @@ def test_replace_with_empty_dictlike(self, mix_abc):
# GH 15289
df = DataFrame(mix_abc)
tm.assert_frame_equal(df, df.replace({}))
- tm.assert_frame_equal(df, df.replace(Series([])))
+ tm.assert_frame_equal(df, df.replace(Series([], dtype=object)))
tm.assert_frame_equal(df, df.replace({"b": {}}))
tm.assert_frame_equal(df, df.replace(Series({"b": {}})))
diff --git a/pandas/tests/generic/test_generic.py b/pandas/tests/generic/test_generic.py
index 0912a8901dc6a..0ff9d7fcdb209 100644
--- a/pandas/tests/generic/test_generic.py
+++ b/pandas/tests/generic/test_generic.py
@@ -33,6 +33,7 @@ def _construct(self, shape, value=None, dtype=None, **kwargs):
if is_scalar(value):
if value == "empty":
arr = None
+ dtype = np.float64
# remove the info axis
kwargs.pop(self._typ._info_axis_name, None)
@@ -732,13 +733,10 @@ def test_squeeze(self):
tm.assert_series_equal(df.squeeze(), df["A"])
# don't fail with 0 length dimensions GH11229 & GH8999
- empty_series = Series([], name="five")
+ empty_series = Series([], name="five", dtype=np.float64)
empty_frame = DataFrame([empty_series])
-
- [
- tm.assert_series_equal(empty_series, higher_dim.squeeze())
- for higher_dim in [empty_series, empty_frame]
- ]
+ tm.assert_series_equal(empty_series, empty_series.squeeze())
+ tm.assert_series_equal(empty_series, empty_frame.squeeze())
# axis argument
df = tm.makeTimeDataFrame(nper=1).iloc[:, :1]
@@ -898,10 +896,10 @@ def test_equals(self):
# GH 8437
a = pd.Series([False, np.nan])
b = pd.Series([False, np.nan])
- c = pd.Series(index=range(2))
- d = pd.Series(index=range(2))
- e = pd.Series(index=range(2))
- f = pd.Series(index=range(2))
+ c = pd.Series(index=range(2), dtype=object)
+ d = c.copy()
+ e = c.copy()
+ f = c.copy()
c[:-1] = d[:-1] = e[0] = f[0] = False
assert a.equals(a)
assert a.equals(b)
@@ -940,7 +938,7 @@ def test_pipe_tuple_error(self):
@pytest.mark.parametrize("box", [pd.Series, pd.DataFrame])
def test_axis_classmethods(self, box):
- obj = box()
+ obj = box(dtype=object)
values = (
list(box._AXIS_NAMES.keys())
+ list(box._AXIS_NUMBERS.keys())
diff --git a/pandas/tests/generic/test_series.py b/pandas/tests/generic/test_series.py
index 096a5aa99bd80..aaf523956aaed 100644
--- a/pandas/tests/generic/test_series.py
+++ b/pandas/tests/generic/test_series.py
@@ -224,7 +224,7 @@ def test_to_xarray_index_types(self, index):
def test_to_xarray(self):
from xarray import DataArray
- s = Series([])
+ s = Series([], dtype=object)
s.index.name = "foo"
result = s.to_xarray()
assert len(result) == 0
diff --git a/pandas/tests/groupby/test_counting.py b/pandas/tests/groupby/test_counting.py
index 9882f12714d2d..8e9554085b9ee 100644
--- a/pandas/tests/groupby/test_counting.py
+++ b/pandas/tests/groupby/test_counting.py
@@ -20,7 +20,7 @@ def test_cumcount(self):
def test_cumcount_empty(self):
ge = DataFrame().groupby(level=0)
- se = Series().groupby(level=0)
+ se = Series(dtype=object).groupby(level=0)
# edge case, as this is usually considered float
e = Series(dtype="int64")
@@ -95,7 +95,7 @@ def test_ngroup_one_group(self):
def test_ngroup_empty(self):
ge = DataFrame().groupby(level=0)
- se = Series().groupby(level=0)
+ se = Series(dtype=object).groupby(level=0)
# edge case, as this is usually considered float
e = Series(dtype="int64")
diff --git a/pandas/tests/groupby/test_filters.py b/pandas/tests/groupby/test_filters.py
index 2ce04fc774083..b3ee12b6691d7 100644
--- a/pandas/tests/groupby/test_filters.py
+++ b/pandas/tests/groupby/test_filters.py
@@ -593,5 +593,5 @@ def test_filter_dropna_with_empty_groups():
tm.assert_series_equal(result_false, expected_false)
result_true = groupped.filter(lambda x: x.mean() > 1, dropna=True)
- expected_true = pd.Series(index=pd.Index([], dtype=int))
+ expected_true = pd.Series(index=pd.Index([], dtype=int), dtype=np.float64)
tm.assert_series_equal(result_true, expected_true)
diff --git a/pandas/tests/groupby/test_function.py b/pandas/tests/groupby/test_function.py
index c41f762e9128d..4ca23c61ba920 100644
--- a/pandas/tests/groupby/test_function.py
+++ b/pandas/tests/groupby/test_function.py
@@ -1047,7 +1047,7 @@ def test_nunique_with_object():
def test_nunique_with_empty_series():
# GH 12553
- data = pd.Series(name="name")
+ data = pd.Series(name="name", dtype=object)
result = data.groupby(level=0).nunique()
expected = pd.Series(name="name", dtype="int64")
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/groupby/test_grouping.py b/pandas/tests/groupby/test_grouping.py
index e4edc64016567..2c84c2f034fc6 100644
--- a/pandas/tests/groupby/test_grouping.py
+++ b/pandas/tests/groupby/test_grouping.py
@@ -585,9 +585,18 @@ def test_list_grouper_with_nat(self):
@pytest.mark.parametrize(
"func,expected",
[
- ("transform", pd.Series(name=2, index=pd.RangeIndex(0, 0, 1))),
- ("agg", pd.Series(name=2, index=pd.Float64Index([], name=1))),
- ("apply", pd.Series(name=2, index=pd.Float64Index([], name=1))),
+ (
+ "transform",
+ pd.Series(name=2, dtype=np.float64, index=pd.RangeIndex(0, 0, 1)),
+ ),
+ (
+ "agg",
+ pd.Series(name=2, dtype=np.float64, index=pd.Float64Index([], name=1)),
+ ),
+ (
+ "apply",
+ pd.Series(name=2, dtype=np.float64, index=pd.Float64Index([], name=1)),
+ ),
],
)
def test_evaluate_with_empty_groups(self, func, expected):
@@ -602,7 +611,7 @@ def test_evaluate_with_empty_groups(self, func, expected):
def test_groupby_empty(self):
# https://github.com/pandas-dev/pandas/issues/27190
- s = pd.Series([], name="name")
+ s = pd.Series([], name="name", dtype="float64")
gr = s.groupby([])
result = gr.mean()
@@ -731,7 +740,7 @@ def test_get_group_grouped_by_tuple(self):
def test_groupby_with_empty(self):
index = pd.DatetimeIndex(())
data = ()
- series = pd.Series(data, index)
+ series = pd.Series(data, index, dtype=object)
grouper = pd.Grouper(freq="D")
grouped = series.groupby(grouper)
assert next(iter(grouped), None) is None
diff --git a/pandas/tests/indexes/datetimelike.py b/pandas/tests/indexes/datetimelike.py
index 42244626749b9..6eedfca129856 100644
--- a/pandas/tests/indexes/datetimelike.py
+++ b/pandas/tests/indexes/datetimelike.py
@@ -72,7 +72,7 @@ def test_map_callable(self):
"mapper",
[
lambda values, index: {i: e for e, i in zip(values, index)},
- lambda values, index: pd.Series(values, index),
+ lambda values, index: pd.Series(values, index, dtype=object),
],
)
def test_map_dictlike(self, mapper):
diff --git a/pandas/tests/indexes/datetimes/test_arithmetic.py b/pandas/tests/indexes/datetimes/test_arithmetic.py
index 4851dd5a55c1e..2bcaa973acd6b 100644
--- a/pandas/tests/indexes/datetimes/test_arithmetic.py
+++ b/pandas/tests/indexes/datetimes/test_arithmetic.py
@@ -100,9 +100,9 @@ def test_dti_shift_localized(self, tzstr):
def test_dti_shift_across_dst(self):
# GH 8616
idx = date_range("2013-11-03", tz="America/Chicago", periods=7, freq="H")
- s = Series(index=idx[:-1])
+ s = Series(index=idx[:-1], dtype=object)
result = s.shift(freq="H")
- expected = Series(index=idx[1:])
+ expected = Series(index=idx[1:], dtype=object)
tm.assert_series_equal(result, expected)
@pytest.mark.parametrize(
diff --git a/pandas/tests/indexes/test_base.py b/pandas/tests/indexes/test_base.py
index c0c677b076e2c..e62d50f64d8ff 100644
--- a/pandas/tests/indexes/test_base.py
+++ b/pandas/tests/indexes/test_base.py
@@ -2001,7 +2001,7 @@ def test_isin_level_kwarg_bad_label_raises(self, label, indices):
with pytest.raises(KeyError, match=msg):
index.isin([], level=label)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_isin_empty(self, empty):
# see gh-16991
index = Index(["a", "b"])
diff --git a/pandas/tests/indexing/common.py b/pandas/tests/indexing/common.py
index db6dddfdca11b..e5b2c83f29030 100644
--- a/pandas/tests/indexing/common.py
+++ b/pandas/tests/indexing/common.py
@@ -93,7 +93,7 @@ def setup_method(self, method):
self.frame_ts_rev = DataFrame(np.random.randn(4, 4), index=dates_rev)
self.frame_empty = DataFrame()
- self.series_empty = Series()
+ self.series_empty = Series(dtype=object)
# form agglomerates
for kind in self._kinds:
diff --git a/pandas/tests/indexing/multiindex/test_loc.py b/pandas/tests/indexing/multiindex/test_loc.py
index 76425c72ce4f9..b6b9f7f205394 100644
--- a/pandas/tests/indexing/multiindex/test_loc.py
+++ b/pandas/tests/indexing/multiindex/test_loc.py
@@ -48,7 +48,9 @@ def test_loc_getitem_series(self):
empty = Series(data=[], dtype=np.float64)
expected = Series(
- [], index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64)
+ [],
+ index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64),
+ dtype=np.float64,
)
result = x.loc[empty]
tm.assert_series_equal(result, expected)
@@ -70,7 +72,9 @@ def test_loc_getitem_array(self):
# empty array:
empty = np.array([])
expected = Series(
- [], index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64)
+ [],
+ index=MultiIndex(levels=index.levels, codes=[[], []], dtype=np.float64),
+ dtype="float64",
)
result = x.loc[empty]
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/indexing/test_iloc.py b/pandas/tests/indexing/test_iloc.py
index e4d387fd3ac38..f9bded5b266f1 100644
--- a/pandas/tests/indexing/test_iloc.py
+++ b/pandas/tests/indexing/test_iloc.py
@@ -286,7 +286,7 @@ def test_iloc_getitem_dups(self):
def test_iloc_getitem_array(self):
# array like
- s = Series(index=range(1, 4))
+ s = Series(index=range(1, 4), dtype=object)
self.check_result(
"iloc",
s.index,
@@ -499,7 +499,7 @@ def test_iloc_getitem_frame(self):
tm.assert_frame_equal(result, expected)
# with index-like
- s = Series(index=range(1, 5))
+ s = Series(index=range(1, 5), dtype=object)
result = df.iloc[s.index]
with catch_warnings(record=True):
filterwarnings("ignore", "\\n.ix", FutureWarning)
diff --git a/pandas/tests/indexing/test_indexing.py b/pandas/tests/indexing/test_indexing.py
index 25b8713eb0307..d75afd1540f22 100644
--- a/pandas/tests/indexing/test_indexing.py
+++ b/pandas/tests/indexing/test_indexing.py
@@ -895,7 +895,7 @@ def test_range_in_series_indexing(self):
# range can cause an indexing error
# GH 11652
for x in [5, 999999, 1000000]:
- s = Series(index=range(x))
+ s = Series(index=range(x), dtype=np.float64)
s.loc[range(1)] = 42
tm.assert_series_equal(s.loc[range(1)], Series(42.0, index=[0]))
diff --git a/pandas/tests/indexing/test_loc.py b/pandas/tests/indexing/test_loc.py
index cb523efb78cf4..e5e899bfb7f0d 100644
--- a/pandas/tests/indexing/test_loc.py
+++ b/pandas/tests/indexing/test_loc.py
@@ -217,7 +217,7 @@ def test_loc_getitem_label_array_like(self):
# array like
self.check_result(
"loc",
- Series(index=[0, 2, 4]).index,
+ Series(index=[0, 2, 4], dtype=object).index,
"ix",
[0, 2, 4],
typs=["ints", "uints"],
@@ -225,7 +225,7 @@ def test_loc_getitem_label_array_like(self):
)
self.check_result(
"loc",
- Series(index=[3, 6, 9]).index,
+ Series(index=[3, 6, 9], dtype=object).index,
"ix",
[3, 6, 9],
typs=["ints", "uints"],
@@ -282,7 +282,7 @@ def test_loc_to_fail(self):
# GH 7496
# loc should not fallback
- s = Series()
+ s = Series(dtype=object)
s.loc[1] = 1
s.loc["a"] = 2
@@ -794,13 +794,13 @@ def test_setitem_new_key_tz(self):
]
expected = pd.Series(vals, index=["foo", "bar"])
- ser = pd.Series()
+ ser = pd.Series(dtype=object)
ser["foo"] = vals[0]
ser["bar"] = vals[1]
tm.assert_series_equal(ser, expected)
- ser = pd.Series()
+ ser = pd.Series(dtype=object)
ser.loc["foo"] = vals[0]
ser.loc["bar"] = vals[1]
@@ -1016,7 +1016,7 @@ def test_loc_reverse_assignment(self):
data = [1, 2, 3, 4, 5, 6] + [None] * 4
expected = Series(data, index=range(2010, 2020))
- result = pd.Series(index=range(2010, 2020))
+ result = pd.Series(index=range(2010, 2020), dtype=np.float64)
result.loc[2015:2010:-1] = [6, 5, 4, 3, 2, 1]
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/indexing/test_partial.py b/pandas/tests/indexing/test_partial.py
index aa49edd51aa39..3adc206335e6f 100644
--- a/pandas/tests/indexing/test_partial.py
+++ b/pandas/tests/indexing/test_partial.py
@@ -368,19 +368,19 @@ def test_partial_set_empty_series(self):
# GH5226
# partially set with an empty object series
- s = Series()
+ s = Series(dtype=object)
s.loc[1] = 1
tm.assert_series_equal(s, Series([1], index=[1]))
s.loc[3] = 3
tm.assert_series_equal(s, Series([1, 3], index=[1, 3]))
- s = Series()
+ s = Series(dtype=object)
s.loc[1] = 1.0
tm.assert_series_equal(s, Series([1.0], index=[1]))
s.loc[3] = 3.0
tm.assert_series_equal(s, Series([1.0, 3.0], index=[1, 3]))
- s = Series()
+ s = Series(dtype=object)
s.loc["foo"] = 1
tm.assert_series_equal(s, Series([1], index=["foo"]))
s.loc["bar"] = 3
@@ -512,11 +512,11 @@ def test_partial_set_empty_frame_row(self):
def test_partial_set_empty_frame_set_series(self):
# GH 5756
# setting with empty Series
- df = DataFrame(Series())
- tm.assert_frame_equal(df, DataFrame({0: Series()}))
+ df = DataFrame(Series(dtype=object))
+ tm.assert_frame_equal(df, DataFrame({0: Series(dtype=object)}))
- df = DataFrame(Series(name="foo"))
- tm.assert_frame_equal(df, DataFrame({"foo": Series()}))
+ df = DataFrame(Series(name="foo", dtype=object))
+ tm.assert_frame_equal(df, DataFrame({"foo": Series(dtype=object)}))
def test_partial_set_empty_frame_empty_copy_assignment(self):
# GH 5932
diff --git a/pandas/tests/io/formats/test_format.py b/pandas/tests/io/formats/test_format.py
index 004a1d184537d..e875a6f137d80 100644
--- a/pandas/tests/io/formats/test_format.py
+++ b/pandas/tests/io/formats/test_format.py
@@ -1017,7 +1017,7 @@ def test_east_asian_unicode_true(self):
def test_to_string_buffer_all_unicode(self):
buf = StringIO()
- empty = DataFrame({"c/\u03c3": Series()})
+ empty = DataFrame({"c/\u03c3": Series(dtype=object)})
nonempty = DataFrame({"c/\u03c3": Series([1, 2, 3])})
print(empty, file=buf)
@@ -2765,7 +2765,7 @@ def test_to_string_length(self):
assert res == exp
def test_to_string_na_rep(self):
- s = pd.Series(index=range(100))
+ s = pd.Series(index=range(100), dtype=np.float64)
res = s.to_string(na_rep="foo", max_rows=2)
exp = "0 foo\n ..\n99 foo"
assert res == exp
diff --git a/pandas/tests/io/json/test_pandas.py b/pandas/tests/io/json/test_pandas.py
index d31aa04b223e8..bce3d1de849aa 100644
--- a/pandas/tests/io/json/test_pandas.py
+++ b/pandas/tests/io/json/test_pandas.py
@@ -53,7 +53,7 @@ def setup(self, datapath):
self.objSeries = tm.makeObjectSeries()
self.objSeries.name = "objects"
- self.empty_series = Series([], index=[])
+ self.empty_series = Series([], index=[], dtype=np.float64)
self.empty_frame = DataFrame()
self.frame = _frame.copy()
diff --git a/pandas/tests/io/pytables/test_store.py b/pandas/tests/io/pytables/test_store.py
index d79280f9ea494..d9a76fe97f813 100644
--- a/pandas/tests/io/pytables/test_store.py
+++ b/pandas/tests/io/pytables/test_store.py
@@ -2376,8 +2376,8 @@ def test_frame(self, compression, setup_path):
@td.xfail_non_writeable
def test_empty_series_frame(self, setup_path):
- s0 = Series()
- s1 = Series(name="myseries")
+ s0 = Series(dtype=object)
+ s1 = Series(name="myseries", dtype=object)
df0 = DataFrame()
df1 = DataFrame(index=["a", "b", "c"])
df2 = DataFrame(columns=["d", "e", "f"])
diff --git a/pandas/tests/io/test_html.py b/pandas/tests/io/test_html.py
index 353946a311c1a..c34f2ebace683 100644
--- a/pandas/tests/io/test_html.py
+++ b/pandas/tests/io/test_html.py
@@ -395,8 +395,7 @@ def test_empty_tables(self):
"""
Make sure that read_html ignores empty tables.
"""
- result = self.read_html(
- """
+ html = """
<table>
<thead>
<tr>
@@ -416,8 +415,7 @@ def test_empty_tables(self):
</tbody>
</table>
"""
- )
-
+ result = self.read_html(html)
assert len(result) == 1
def test_multiple_tbody(self):
diff --git a/pandas/tests/plotting/test_misc.py b/pandas/tests/plotting/test_misc.py
index 1e59fbf928876..9e947d4ba878a 100644
--- a/pandas/tests/plotting/test_misc.py
+++ b/pandas/tests/plotting/test_misc.py
@@ -34,7 +34,7 @@ def test_get_accessor_args():
msg = "should not be called with positional arguments"
with pytest.raises(TypeError, match=msg):
- func(backend_name="", data=Series(), args=["line", None], kwargs={})
+ func(backend_name="", data=Series(dtype=object), args=["line", None], kwargs={})
x, y, kind, kwargs = func(
backend_name="",
@@ -48,7 +48,10 @@ def test_get_accessor_args():
assert kwargs == {"grid": False}
x, y, kind, kwargs = func(
- backend_name="pandas.plotting._matplotlib", data=Series(), args=[], kwargs={}
+ backend_name="pandas.plotting._matplotlib",
+ data=Series(dtype=object),
+ args=[],
+ kwargs={},
)
assert x is None
assert y is None
diff --git a/pandas/tests/reductions/test_reductions.py b/pandas/tests/reductions/test_reductions.py
index 80d148c919ab2..3f78a6ac4a778 100644
--- a/pandas/tests/reductions/test_reductions.py
+++ b/pandas/tests/reductions/test_reductions.py
@@ -79,7 +79,7 @@ def test_nanops(self):
assert pd.isna(getattr(obj, opname)())
assert pd.isna(getattr(obj, opname)(skipna=False))
- obj = klass([])
+ obj = klass([], dtype=object)
assert pd.isna(getattr(obj, opname)())
assert pd.isna(getattr(obj, opname)(skipna=False))
@@ -528,7 +528,7 @@ def test_empty(self, method, unit, use_bottleneck):
with pd.option_context("use_bottleneck", use_bottleneck):
# GH#9422 / GH#18921
# Entirely empty
- s = Series([])
+ s = Series([], dtype=object)
# NA by default
result = getattr(s, method)()
assert result == unit
@@ -900,7 +900,7 @@ def test_timedelta64_analytics(self):
@pytest.mark.parametrize(
"test_input,error_type",
[
- (pd.Series([]), ValueError),
+ (pd.Series([], dtype="float64"), ValueError),
# For strings, or any Series with dtype 'O'
(pd.Series(["foo", "bar", "baz"]), TypeError),
(pd.Series([(1,), (2,)]), TypeError),
diff --git a/pandas/tests/resample/test_base.py b/pandas/tests/resample/test_base.py
index 161581e16b6fe..622b85f2a398c 100644
--- a/pandas/tests/resample/test_base.py
+++ b/pandas/tests/resample/test_base.py
@@ -139,7 +139,7 @@ def test_resample_empty_dataframe(empty_frame, freq, resample_method):
expected = df.copy()
else:
# GH14962
- expected = Series([])
+ expected = Series([], dtype=object)
if isinstance(df.index, PeriodIndex):
expected.index = df.index.asfreq(freq=freq)
diff --git a/pandas/tests/resample/test_datetime_index.py b/pandas/tests/resample/test_datetime_index.py
index f9229e8066be4..5837d526e3978 100644
--- a/pandas/tests/resample/test_datetime_index.py
+++ b/pandas/tests/resample/test_datetime_index.py
@@ -1429,10 +1429,11 @@ def test_downsample_across_dst_weekly():
tm.assert_frame_equal(result, expected)
idx = pd.date_range("2013-04-01", "2013-05-01", tz="Europe/London", freq="H")
- s = Series(index=idx)
+ s = Series(index=idx, dtype=np.float64)
result = s.resample("W").mean()
expected = Series(
- index=pd.date_range("2013-04-07", freq="W", periods=5, tz="Europe/London")
+ index=pd.date_range("2013-04-07", freq="W", periods=5, tz="Europe/London"),
+ dtype=np.float64,
)
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/resample/test_period_index.py b/pandas/tests/resample/test_period_index.py
index 93ce7a9480b35..219491367d292 100644
--- a/pandas/tests/resample/test_period_index.py
+++ b/pandas/tests/resample/test_period_index.py
@@ -594,7 +594,7 @@ def test_resample_with_dst_time_change(self):
def test_resample_bms_2752(self):
# GH2753
- foo = Series(index=pd.bdate_range("20000101", "20000201"))
+ foo = Series(index=pd.bdate_range("20000101", "20000201"), dtype=np.float64)
res1 = foo.resample("BMS").mean()
res2 = foo.resample("BMS").mean().resample("B").mean()
assert res1.index[0] == Timestamp("20000103")
diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index 63f1ef7595f31..8ef35882dcc12 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -27,6 +27,7 @@
isna,
read_csv,
)
+from pandas.core.construction import create_series_with_explicit_dtype
from pandas.tests.extension.decimal import to_decimal
import pandas.util.testing as tm
@@ -2177,7 +2178,7 @@ def test_concat_period_other_series(self):
def test_concat_empty_series(self):
# GH 11082
s1 = pd.Series([1, 2, 3], name="x")
- s2 = pd.Series(name="y")
+ s2 = pd.Series(name="y", dtype="float64")
res = pd.concat([s1, s2], axis=1)
exp = pd.DataFrame(
{"x": [1, 2, 3], "y": [np.nan, np.nan, np.nan]},
@@ -2186,7 +2187,7 @@ def test_concat_empty_series(self):
tm.assert_frame_equal(res, exp)
s1 = pd.Series([1, 2, 3], name="x")
- s2 = pd.Series(name="y")
+ s2 = pd.Series(name="y", dtype="float64")
res = pd.concat([s1, s2], axis=0)
# name will be reset
exp = pd.Series([1, 2, 3])
@@ -2194,7 +2195,7 @@ def test_concat_empty_series(self):
# empty Series with no name
s1 = pd.Series([1, 2, 3], name="x")
- s2 = pd.Series(name=None)
+ s2 = pd.Series(name=None, dtype="float64")
res = pd.concat([s1, s2], axis=1)
exp = pd.DataFrame(
{"x": [1, 2, 3], 0: [np.nan, np.nan, np.nan]},
@@ -2209,7 +2210,9 @@ def test_concat_empty_series_timelike(self, tz, values):
# GH 18447
first = Series([], dtype="M8[ns]").dt.tz_localize(tz)
- second = Series(values)
+ dtype = None if values else np.float64
+ second = Series(values, dtype=dtype)
+
expected = DataFrame(
{
0: pd.Series([pd.NaT] * len(values), dtype="M8[ns]").dt.tz_localize(tz),
@@ -2569,7 +2572,8 @@ def test_concat_odered_dict(self):
@pytest.mark.parametrize("dt", np.sctypes["float"])
def test_concat_no_unnecessary_upcast(dt, pdt):
# GH 13247
- dims = pdt().ndim
+ dims = pdt(dtype=object).ndim
+
dfs = [
pdt(np.array([1], dtype=dt, ndmin=dims)),
pdt(np.array([np.nan], dtype=dt, ndmin=dims)),
@@ -2579,7 +2583,7 @@ def test_concat_no_unnecessary_upcast(dt, pdt):
assert x.values.dtype == dt
-@pytest.mark.parametrize("pdt", [pd.Series, pd.DataFrame])
+@pytest.mark.parametrize("pdt", [create_series_with_explicit_dtype, pd.DataFrame])
@pytest.mark.parametrize("dt", np.sctypes["int"])
def test_concat_will_upcast(dt, pdt):
with catch_warnings(record=True):
@@ -2605,7 +2609,8 @@ def test_concat_empty_and_non_empty_frame_regression():
def test_concat_empty_and_non_empty_series_regression():
# GH 18187 regression test
s1 = pd.Series([1])
- s2 = pd.Series([])
+ s2 = pd.Series([], dtype=object)
+
expected = s1
result = pd.concat([s1, s2])
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/series/indexing/test_alter_index.py b/pandas/tests/series/indexing/test_alter_index.py
index 7509d21b8832f..c47b99fa38989 100644
--- a/pandas/tests/series/indexing/test_alter_index.py
+++ b/pandas/tests/series/indexing/test_alter_index.py
@@ -230,7 +230,7 @@ def test_reindex_with_datetimes():
def test_reindex_corner(datetime_series):
# (don't forget to fix this) I think it's fixed
- empty = Series()
+ empty = Series(dtype=object)
empty.reindex(datetime_series.index, method="pad") # it works
# corner case: pad empty series
@@ -539,8 +539,9 @@ def test_drop_with_ignore_errors():
def test_drop_empty_list(index, drop_labels):
# GH 21494
expected_index = [i for i in index if i not in drop_labels]
- series = pd.Series(index=index).drop(drop_labels)
- tm.assert_series_equal(series, pd.Series(index=expected_index))
+ series = pd.Series(index=index, dtype=object).drop(drop_labels)
+ expected = pd.Series(index=expected_index, dtype=object)
+ tm.assert_series_equal(series, expected)
@pytest.mark.parametrize(
@@ -554,4 +555,5 @@ def test_drop_empty_list(index, drop_labels):
def test_drop_non_empty_list(data, index, drop_labels):
# GH 21494 and GH 16877
with pytest.raises(KeyError, match="not found in axis"):
- pd.Series(data=data, index=index).drop(drop_labels)
+ dtype = object if data is None else None
+ pd.Series(data=data, index=index, dtype=dtype).drop(drop_labels)
diff --git a/pandas/tests/series/indexing/test_datetime.py b/pandas/tests/series/indexing/test_datetime.py
index fab3310fa3dfe..83c1c0ff16f4c 100644
--- a/pandas/tests/series/indexing/test_datetime.py
+++ b/pandas/tests/series/indexing/test_datetime.py
@@ -105,7 +105,7 @@ def test_series_set_value():
dates = [datetime(2001, 1, 1), datetime(2001, 1, 2)]
index = DatetimeIndex(dates)
- s = Series()._set_value(dates[0], 1.0)
+ s = Series(dtype=object)._set_value(dates[0], 1.0)
s2 = s._set_value(dates[1], np.nan)
expected = Series([1.0, np.nan], index=index)
diff --git a/pandas/tests/series/indexing/test_indexing.py b/pandas/tests/series/indexing/test_indexing.py
index 173bc9d9d6409..5bebd480ce8d4 100644
--- a/pandas/tests/series/indexing/test_indexing.py
+++ b/pandas/tests/series/indexing/test_indexing.py
@@ -105,7 +105,9 @@ def test_getitem_get(datetime_series, string_series, object_series):
# None
# GH 5652
- for s in [Series(), Series(index=list("abc"))]:
+ s1 = Series(dtype=object)
+ s2 = Series(dtype=object, index=list("abc"))
+ for s in [s1, s2]:
result = s.get(None)
assert result is None
@@ -130,7 +132,7 @@ def test_getitem_generator(string_series):
def test_type_promotion():
# GH12599
- s = pd.Series()
+ s = pd.Series(dtype=object)
s["a"] = pd.Timestamp("2016-01-01")
s["b"] = 3.0
s["c"] = "foo"
@@ -168,7 +170,7 @@ def test_getitem_out_of_bounds(datetime_series):
datetime_series[len(datetime_series)]
# GH #917
- s = Series([])
+ s = Series([], dtype=object)
with pytest.raises(IndexError, match=msg):
s[-1]
@@ -324,12 +326,12 @@ def test_setitem(datetime_series, string_series):
# Test for issue #10193
key = pd.Timestamp("2012-01-01")
- series = pd.Series()
+ series = pd.Series(dtype=object)
series[key] = 47
expected = pd.Series(47, [key])
tm.assert_series_equal(series, expected)
- series = pd.Series([], pd.DatetimeIndex([], freq="D"))
+ series = pd.Series([], pd.DatetimeIndex([], freq="D"), dtype=object)
series[key] = 47
expected = pd.Series(47, pd.DatetimeIndex([key], freq="D"))
tm.assert_series_equal(series, expected)
@@ -637,7 +639,7 @@ def test_setitem_na():
def test_timedelta_assignment():
# GH 8209
- s = Series([])
+ s = Series([], dtype=object)
s.loc["B"] = timedelta(1)
tm.assert_series_equal(s, Series(Timedelta("1 days"), index=["B"]))
diff --git a/pandas/tests/series/indexing/test_numeric.py b/pandas/tests/series/indexing/test_numeric.py
index 426a98b00827e..a641b47f2e690 100644
--- a/pandas/tests/series/indexing/test_numeric.py
+++ b/pandas/tests/series/indexing/test_numeric.py
@@ -150,7 +150,7 @@ def test_delitem():
tm.assert_series_equal(s, expected)
# empty
- s = Series()
+ s = Series(dtype=object)
with pytest.raises(KeyError, match=r"^0$"):
del s[0]
diff --git a/pandas/tests/series/test_analytics.py b/pandas/tests/series/test_analytics.py
index fe9306a06efc7..71b4819bb4da8 100644
--- a/pandas/tests/series/test_analytics.py
+++ b/pandas/tests/series/test_analytics.py
@@ -843,7 +843,7 @@ def test_isin_with_i8(self):
result = s.isin(s[0:2])
tm.assert_series_equal(result, expected)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_isin_empty(self, empty):
# see gh-16991
s = Series(["a", "b"])
diff --git a/pandas/tests/series/test_api.py b/pandas/tests/series/test_api.py
index 8acab3fa2541d..5da0ee9b5b1c0 100644
--- a/pandas/tests/series/test_api.py
+++ b/pandas/tests/series/test_api.py
@@ -266,7 +266,7 @@ def get_dir(s):
)
def test_index_tab_completion(self, index):
# dir contains string-like values of the Index.
- s = pd.Series(index=index)
+ s = pd.Series(index=index, dtype=object)
dir_s = dir(s)
for i, x in enumerate(s.index.unique(level=0)):
if i < 100:
@@ -275,7 +275,7 @@ def test_index_tab_completion(self, index):
assert x not in dir_s
def test_not_hashable(self):
- s_empty = Series()
+ s_empty = Series(dtype=object)
s = Series([1])
msg = "'Series' objects are mutable, thus they cannot be hashed"
with pytest.raises(TypeError, match=msg):
@@ -474,10 +474,11 @@ def test_str_attribute(self):
s.str.repeat(2)
def test_empty_method(self):
- s_empty = pd.Series()
+ s_empty = pd.Series(dtype=object)
assert s_empty.empty
- for full_series in [pd.Series([1]), pd.Series(index=[1])]:
+ s2 = pd.Series(index=[1], dtype=object)
+ for full_series in [pd.Series([1]), s2]:
assert not full_series.empty
def test_tab_complete_warning(self, ip):
diff --git a/pandas/tests/series/test_apply.py b/pandas/tests/series/test_apply.py
index eb4f3273f8713..8956b8b0b2d20 100644
--- a/pandas/tests/series/test_apply.py
+++ b/pandas/tests/series/test_apply.py
@@ -37,7 +37,7 @@ def test_apply(self, datetime_series):
assert s.name == rs.name
# index but no data
- s = Series(index=[1, 2, 3])
+ s = Series(index=[1, 2, 3], dtype=np.float64)
rs = s.apply(lambda x: x)
tm.assert_series_equal(s, rs)
@@ -340,7 +340,7 @@ def test_non_callable_aggregates(self):
"series, func, expected",
chain(
_get_cython_table_params(
- Series(),
+ Series(dtype=np.float64),
[
("sum", 0),
("max", np.nan),
@@ -395,8 +395,11 @@ def test_agg_cython_table(self, series, func, expected):
"series, func, expected",
chain(
_get_cython_table_params(
- Series(),
- [("cumprod", Series([], Index([]))), ("cumsum", Series([], Index([])))],
+ Series(dtype=np.float64),
+ [
+ ("cumprod", Series([], Index([]), dtype=np.float64)),
+ ("cumsum", Series([], Index([]), dtype=np.float64)),
+ ],
),
_get_cython_table_params(
Series([np.nan, 1, 2, 3]),
diff --git a/pandas/tests/series/test_combine_concat.py b/pandas/tests/series/test_combine_concat.py
index 9d02c1bdc2d9c..c6f4ce364f328 100644
--- a/pandas/tests/series/test_combine_concat.py
+++ b/pandas/tests/series/test_combine_concat.py
@@ -107,7 +107,8 @@ def test_combine_first(self):
# corner case
s = Series([1.0, 2, 3], index=[0, 1, 2])
- result = s.combine_first(Series([], index=[]))
+ empty = Series([], index=[], dtype=object)
+ result = s.combine_first(empty)
s.index = s.index.astype("O")
tm.assert_series_equal(s, result)
diff --git a/pandas/tests/series/test_constructors.py b/pandas/tests/series/test_constructors.py
index 34b11a0d008aa..293ec9580436e 100644
--- a/pandas/tests/series/test_constructors.py
+++ b/pandas/tests/series/test_constructors.py
@@ -52,8 +52,10 @@ class TestSeriesConstructors:
],
)
def test_empty_constructor(self, constructor, check_index_type):
- expected = Series()
- result = constructor()
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ expected = Series()
+ result = constructor()
+
assert len(result.index) == 0
tm.assert_series_equal(result, expected, check_index_type=check_index_type)
@@ -76,8 +78,8 @@ def test_scalar_conversion(self):
assert int(Series([1.0])) == 1
def test_constructor(self, datetime_series):
- empty_series = Series()
-
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty_series = Series()
assert datetime_series.index.is_all_dates
# Pass in Series
@@ -94,7 +96,8 @@ def test_constructor(self, datetime_series):
assert mixed[1] is np.NaN
assert not empty_series.index.is_all_dates
- assert not Series().index.is_all_dates
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ assert not Series().index.is_all_dates
# exception raised is of type Exception
with pytest.raises(Exception, match="Data must be 1-dimensional"):
@@ -113,8 +116,9 @@ def test_constructor(self, datetime_series):
@pytest.mark.parametrize("input_class", [list, dict, OrderedDict])
def test_constructor_empty(self, input_class):
- empty = Series()
- empty2 = Series(input_class())
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty = Series()
+ empty2 = Series(input_class())
# these are Index() and RangeIndex() which don't compare type equal
# but are just .equals
@@ -132,8 +136,9 @@ def test_constructor_empty(self, input_class):
if input_class is not list:
# With index:
- empty = Series(index=range(10))
- empty2 = Series(input_class(), index=range(10))
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty = Series(index=range(10))
+ empty2 = Series(input_class(), index=range(10))
tm.assert_series_equal(empty, empty2)
# With index and dtype float64:
@@ -165,7 +170,8 @@ def test_constructor_dtype_only(self, dtype, index):
assert len(result) == 0
def test_constructor_no_data_index_order(self):
- result = pd.Series(index=["b", "a", "c"])
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ result = pd.Series(index=["b", "a", "c"])
assert result.index.tolist() == ["b", "a", "c"]
def test_constructor_no_data_string_type(self):
@@ -631,7 +637,8 @@ def test_constructor_limit_copies(self, index):
assert s._data.blocks[0].values is not index
def test_constructor_pass_none(self):
- s = Series(None, index=range(5))
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ s = Series(None, index=range(5))
assert s.dtype == np.float64
s = Series(None, index=range(5), dtype=object)
@@ -639,8 +646,9 @@ def test_constructor_pass_none(self):
# GH 7431
# inference on the index
- s = Series(index=np.array([None]))
- expected = Series(index=Index([None]))
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ s = Series(index=np.array([None]))
+ expected = Series(index=Index([None]))
tm.assert_series_equal(s, expected)
def test_constructor_pass_nan_nat(self):
@@ -1029,7 +1037,7 @@ def test_constructor_dict(self):
pidx = tm.makePeriodIndex(100)
d = {pidx[0]: 0, pidx[1]: 1}
result = Series(d, index=pidx)
- expected = Series(np.nan, pidx)
+ expected = Series(np.nan, pidx, dtype=np.float64)
expected.iloc[0] = 0
expected.iloc[1] = 1
tm.assert_series_equal(result, expected)
@@ -1135,7 +1143,7 @@ def test_fromDict(self):
def test_fromValue(self, datetime_series):
- nans = Series(np.NaN, index=datetime_series.index)
+ nans = Series(np.NaN, index=datetime_series.index, dtype=np.float64)
assert nans.dtype == np.float_
assert len(nans) == len(datetime_series)
diff --git a/pandas/tests/series/test_dtypes.py b/pandas/tests/series/test_dtypes.py
index 065be966efa49..22b00425abb6b 100644
--- a/pandas/tests/series/test_dtypes.py
+++ b/pandas/tests/series/test_dtypes.py
@@ -205,7 +205,11 @@ def test_astype_dict_like(self, dtype_class):
# GH16717
# if dtypes provided is empty, it should error
- dt5 = dtype_class({})
+ if dtype_class is Series:
+ dt5 = dtype_class({}, dtype=object)
+ else:
+ dt5 = dtype_class({})
+
with pytest.raises(KeyError, match=msg):
s.astype(dt5)
@@ -408,7 +412,8 @@ def test_astype_empty_constructor_equality(self, dtype):
"m", # Generic timestamps raise a ValueError. Already tested.
):
init_empty = Series([], dtype=dtype)
- as_type_empty = Series([]).astype(dtype)
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ as_type_empty = Series([]).astype(dtype)
tm.assert_series_equal(init_empty, as_type_empty)
def test_arg_for_errors_in_astype(self):
@@ -472,7 +477,9 @@ def test_infer_objects_series(self):
tm.assert_series_equal(actual, expected)
def test_is_homogeneous_type(self):
- assert Series()._is_homogeneous_type
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty = Series()
+ assert empty._is_homogeneous_type
assert Series([1, 2])._is_homogeneous_type
assert Series(pd.Categorical([1, 2]))._is_homogeneous_type
diff --git a/pandas/tests/series/test_duplicates.py b/pandas/tests/series/test_duplicates.py
index 0f7e3e307ed19..666354e70bdd4 100644
--- a/pandas/tests/series/test_duplicates.py
+++ b/pandas/tests/series/test_duplicates.py
@@ -2,6 +2,7 @@
import pytest
from pandas import Categorical, Series
+from pandas.core.construction import create_series_with_explicit_dtype
import pandas.util.testing as tm
@@ -70,7 +71,7 @@ def test_unique_data_ownership():
)
def test_is_unique(data, expected):
# GH11946 / GH25180
- s = Series(data)
+ s = create_series_with_explicit_dtype(data, dtype_if_empty=object)
assert s.is_unique is expected
diff --git a/pandas/tests/series/test_explode.py b/pandas/tests/series/test_explode.py
index 6262da6bdfabf..e79d3c0556cf1 100644
--- a/pandas/tests/series/test_explode.py
+++ b/pandas/tests/series/test_explode.py
@@ -29,7 +29,7 @@ def test_mixed_type():
def test_empty():
- s = pd.Series()
+ s = pd.Series(dtype=object)
result = s.explode()
expected = s.copy()
tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/series/test_missing.py b/pandas/tests/series/test_missing.py
index c5ce125d10ac2..72f08876e71ae 100644
--- a/pandas/tests/series/test_missing.py
+++ b/pandas/tests/series/test_missing.py
@@ -710,7 +710,7 @@ def test_fillna(self, datetime_series):
tm.assert_series_equal(result, expected)
result = s1.fillna({})
tm.assert_series_equal(result, s1)
- result = s1.fillna(Series(()))
+ result = s1.fillna(Series((), dtype=object))
tm.assert_series_equal(result, s1)
result = s2.fillna(s1)
tm.assert_series_equal(result, s2)
@@ -834,7 +834,8 @@ def test_timedelta64_nan(self):
# tm.assert_series_equal(selector, expected)
def test_dropna_empty(self):
- s = Series([])
+ s = Series([], dtype=object)
+
assert len(s.dropna()) == 0
s.dropna(inplace=True)
assert len(s) == 0
@@ -1163,7 +1164,7 @@ def test_interpolate_corners(self, kwargs):
s = Series([np.nan, np.nan])
tm.assert_series_equal(s.interpolate(**kwargs), s)
- s = Series([]).interpolate()
+ s = Series([], dtype=object).interpolate()
tm.assert_series_equal(s.interpolate(**kwargs), s)
def test_interpolate_index_values(self):
diff --git a/pandas/tests/series/test_operators.py b/pandas/tests/series/test_operators.py
index 983560d68c28c..06fe64d69fb6b 100644
--- a/pandas/tests/series/test_operators.py
+++ b/pandas/tests/series/test_operators.py
@@ -33,7 +33,7 @@ def test_logical_operators_bool_dtype_with_empty(self):
s_tft = Series([True, False, True], index=index)
s_fff = Series([False, False, False], index=index)
- s_empty = Series([])
+ s_empty = Series([], dtype=object)
res = s_tft & s_empty
expected = s_fff
@@ -408,11 +408,13 @@ def test_logical_ops_label_based(self):
# filling
# vs empty
- result = a & Series([])
+ empty = Series([], dtype=object)
+
+ result = a & empty.copy()
expected = Series([False, False, False], list("bca"))
tm.assert_series_equal(result, expected)
- result = a | Series([])
+ result = a | empty.copy()
expected = Series([True, False, True], list("bca"))
tm.assert_series_equal(result, expected)
@@ -428,7 +430,7 @@ def test_logical_ops_label_based(self):
# identity
# we would like s[s|e] == s to hold for any e, whether empty or not
for e in [
- Series([]),
+ empty.copy(),
Series([1], ["z"]),
Series(np.nan, b.index),
Series(np.nan, a.index),
@@ -797,12 +799,12 @@ def test_ops_datetimelike_align(self):
tm.assert_series_equal(result, expected)
def test_operators_corner(self, datetime_series):
- empty = Series([], index=Index([]))
+ empty = Series([], index=Index([]), dtype=np.float64)
result = datetime_series + empty
assert np.isnan(result).all()
- result = empty + Series([], index=Index([]))
+ result = empty + empty.copy()
assert len(result) == 0
# TODO: this returned NotImplemented earlier, what to do?
diff --git a/pandas/tests/series/test_quantile.py b/pandas/tests/series/test_quantile.py
index 1a4a3f523cbbe..4eb275d63e878 100644
--- a/pandas/tests/series/test_quantile.py
+++ b/pandas/tests/series/test_quantile.py
@@ -67,7 +67,7 @@ def test_quantile_multi(self, datetime_series):
result = datetime_series.quantile([])
expected = pd.Series(
- [], name=datetime_series.name, index=Index([], dtype=float)
+ [], name=datetime_series.name, index=Index([], dtype=float), dtype="float64"
)
tm.assert_series_equal(result, expected)
@@ -104,7 +104,8 @@ def test_quantile_nan(self):
assert result == expected
# all nan/empty
- cases = [Series([]), Series([np.nan, np.nan])]
+ s1 = Series([], dtype=object)
+ cases = [s1, Series([np.nan, np.nan])]
for s in cases:
res = s.quantile(0.5)
diff --git a/pandas/tests/series/test_replace.py b/pandas/tests/series/test_replace.py
index 8018ecf03960c..4125b5816422a 100644
--- a/pandas/tests/series/test_replace.py
+++ b/pandas/tests/series/test_replace.py
@@ -245,7 +245,10 @@ def test_replace_with_empty_dictlike(self):
# GH 15289
s = pd.Series(list("abcd"))
tm.assert_series_equal(s, s.replace(dict()))
- tm.assert_series_equal(s, s.replace(pd.Series([])))
+
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ empty_series = pd.Series([])
+ tm.assert_series_equal(s, s.replace(empty_series))
def test_replace_string_with_number(self):
# GH 15743
diff --git a/pandas/tests/series/test_repr.py b/pandas/tests/series/test_repr.py
index f1661ad034e4c..b687179f176c3 100644
--- a/pandas/tests/series/test_repr.py
+++ b/pandas/tests/series/test_repr.py
@@ -62,7 +62,7 @@ def test_name_printing(self):
s.name = None
assert "Name:" not in repr(s)
- s = Series(index=date_range("20010101", "20020101"), name="test")
+ s = Series(index=date_range("20010101", "20020101"), name="test", dtype=object)
assert "Name: test" in repr(s)
def test_repr(self, datetime_series, string_series, object_series):
@@ -75,7 +75,7 @@ def test_repr(self, datetime_series, string_series, object_series):
str(Series(tm.randn(1000), index=np.arange(1000, 0, step=-1)))
# empty
- str(Series())
+ str(Series(dtype=object))
# with NaNs
string_series[5:7] = np.NaN
diff --git a/pandas/tests/series/test_sorting.py b/pandas/tests/series/test_sorting.py
index 8039b133cae10..fd3445e271699 100644
--- a/pandas/tests/series/test_sorting.py
+++ b/pandas/tests/series/test_sorting.py
@@ -157,8 +157,8 @@ def test_sort_index_multiindex(self, level):
def test_sort_index_kind(self):
# GH #14444 & #13589: Add support for sort algo choosing
- series = Series(index=[3, 2, 1, 4, 3])
- expected_series = Series(index=[1, 2, 3, 3, 4])
+ series = Series(index=[3, 2, 1, 4, 3], dtype=object)
+ expected_series = Series(index=[1, 2, 3, 3, 4], dtype=object)
index_sorted_series = series.sort_index(kind="mergesort")
tm.assert_series_equal(expected_series, index_sorted_series)
@@ -170,13 +170,14 @@ def test_sort_index_kind(self):
tm.assert_series_equal(expected_series, index_sorted_series)
def test_sort_index_na_position(self):
- series = Series(index=[3, 2, 1, 4, 3, np.nan])
+ series = Series(index=[3, 2, 1, 4, 3, np.nan], dtype=object)
+ expected_series_first = Series(index=[np.nan, 1, 2, 3, 3, 4], dtype=object)
- expected_series_first = Series(index=[np.nan, 1, 2, 3, 3, 4])
index_sorted_series = series.sort_index(na_position="first")
tm.assert_series_equal(expected_series_first, index_sorted_series)
- expected_series_last = Series(index=[1, 2, 3, 3, 4, np.nan])
+ expected_series_last = Series(index=[1, 2, 3, 3, 4, np.nan], dtype=object)
+
index_sorted_series = series.sort_index(na_position="last")
tm.assert_series_equal(expected_series_last, index_sorted_series)
diff --git a/pandas/tests/series/test_subclass.py b/pandas/tests/series/test_subclass.py
index 6b82f890e974b..5e2d23a70e5be 100644
--- a/pandas/tests/series/test_subclass.py
+++ b/pandas/tests/series/test_subclass.py
@@ -32,4 +32,6 @@ def test_subclass_unstack(self):
tm.assert_frame_equal(res, exp)
def test_subclass_empty_repr(self):
- assert "SubclassedSeries" in repr(tm.SubclassedSeries())
+ with tm.assert_produces_warning(DeprecationWarning, check_stacklevel=False):
+ sub_series = tm.SubclassedSeries()
+ assert "SubclassedSeries" in repr(sub_series)
diff --git a/pandas/tests/series/test_timeseries.py b/pandas/tests/series/test_timeseries.py
index 1587ae5eb7d07..6d00b9f2b09df 100644
--- a/pandas/tests/series/test_timeseries.py
+++ b/pandas/tests/series/test_timeseries.py
@@ -346,10 +346,9 @@ def test_asfreq(self):
def test_asfreq_datetimeindex_empty_series(self):
# GH 14320
- expected = Series(index=pd.DatetimeIndex(["2016-09-29 11:00"])).asfreq("H")
- result = Series(index=pd.DatetimeIndex(["2016-09-29 11:00"]), data=[3]).asfreq(
- "H"
- )
+ index = pd.DatetimeIndex(["2016-09-29 11:00"])
+ expected = Series(index=index, dtype=object).asfreq("H")
+ result = Series([3], index=index.copy()).asfreq("H")
tm.assert_index_equal(expected.index, result.index)
def test_pct_change(self, datetime_series):
@@ -410,7 +409,7 @@ def test_pct_change_periods_freq(
)
tm.assert_series_equal(rs_freq, rs_periods)
- empty_ts = Series(index=datetime_series.index)
+ empty_ts = Series(index=datetime_series.index, dtype=object)
rs_freq = empty_ts.pct_change(freq=freq, fill_method=fill_method, limit=limit)
rs_periods = empty_ts.pct_change(periods, fill_method=fill_method, limit=limit)
tm.assert_series_equal(rs_freq, rs_periods)
@@ -457,12 +456,12 @@ def test_first_last_valid(self, datetime_series):
assert ts.last_valid_index() is None
assert ts.first_valid_index() is None
- ser = Series([], index=[])
+ ser = Series([], index=[], dtype=object)
assert ser.last_valid_index() is None
assert ser.first_valid_index() is None
# GH12800
- empty = Series()
+ empty = Series(dtype=object)
assert empty.last_valid_index() is None
assert empty.first_valid_index() is None
diff --git a/pandas/tests/series/test_timezones.py b/pandas/tests/series/test_timezones.py
index c03101265f7e7..5e255e7cd5dcd 100644
--- a/pandas/tests/series/test_timezones.py
+++ b/pandas/tests/series/test_timezones.py
@@ -89,7 +89,7 @@ def test_series_tz_localize_nonexistent(self, tz, method, exp):
@pytest.mark.parametrize("tzstr", ["US/Eastern", "dateutil/US/Eastern"])
def test_series_tz_localize_empty(self, tzstr):
# GH#2248
- ser = Series()
+ ser = Series(dtype=object)
ser2 = ser.tz_localize("utc")
assert ser2.index.tz == pytz.utc
diff --git a/pandas/tests/test_algos.py b/pandas/tests/test_algos.py
index 02b50d84c6eca..e0e4beffe113a 100644
--- a/pandas/tests/test_algos.py
+++ b/pandas/tests/test_algos.py
@@ -812,7 +812,7 @@ def test_no_cast(self):
result = algos.isin(comps, values)
tm.assert_numpy_array_equal(expected, result)
- @pytest.mark.parametrize("empty", [[], Series(), np.array([])])
+ @pytest.mark.parametrize("empty", [[], Series(dtype=object), np.array([])])
def test_empty(self, empty):
# see gh-16991
vals = Index(["a", "b"])
diff --git a/pandas/tests/test_base.py b/pandas/tests/test_base.py
index d515a015cdbec..5c9a119400319 100644
--- a/pandas/tests/test_base.py
+++ b/pandas/tests/test_base.py
@@ -589,7 +589,7 @@ def test_value_counts_bins(self, index_or_series):
tm.assert_numpy_array_equal(s.unique(), exp)
assert s.nunique() == 3
- s = klass({})
+ s = klass({}) if klass is dict else klass({}, dtype=object)
expected = Series([], dtype=np.int64)
tm.assert_series_equal(s.value_counts(), expected, check_index_type=False)
# returned dtype differs depending on original
diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 44829423be1bb..204cdee2d9e1f 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -1538,7 +1538,7 @@ def test_frame_dict_constructor_empty_series(self):
s2 = Series(
[1, 2, 3, 4], index=MultiIndex.from_tuples([(1, 2), (1, 3), (3, 2), (3, 4)])
)
- s3 = Series()
+ s3 = Series(dtype=object)
# it works!
DataFrame({"foo": s1, "bar": s2, "baz": s3})
diff --git a/pandas/tests/test_register_accessor.py b/pandas/tests/test_register_accessor.py
index 97086f8ab1e85..6b40ff8b3fa1e 100644
--- a/pandas/tests/test_register_accessor.py
+++ b/pandas/tests/test_register_accessor.py
@@ -45,7 +45,8 @@ def test_register(obj, registrar):
with ensure_removed(obj, "mine"):
before = set(dir(obj))
registrar("mine")(MyAccessor)
- assert obj([]).mine.prop == "item"
+ o = obj([]) if obj is not pd.Series else obj([], dtype=object)
+ assert o.mine.prop == "item"
after = set(dir(obj))
assert (before ^ after) == {"mine"}
assert "mine" in obj._accessors
@@ -88,4 +89,4 @@ def __init__(self, data):
raise AttributeError("whoops")
with pytest.raises(AttributeError, match="whoops"):
- pd.Series([]).bad
+ pd.Series([], dtype=object).bad
diff --git a/pandas/tests/util/test_hashing.py b/pandas/tests/util/test_hashing.py
index df3c7fe9c9936..ebbdbd6c29842 100644
--- a/pandas/tests/util/test_hashing.py
+++ b/pandas/tests/util/test_hashing.py
@@ -207,7 +207,7 @@ def test_multiindex_objects():
Series(["a", np.nan, "c"]),
Series(["a", None, "c"]),
Series([True, False, True]),
- Series(),
+ Series(dtype=object),
Index([1, 2, 3]),
Index([True, False, True]),
DataFrame({"x": ["a", "b", "c"], "y": [1, 2, 3]}),
diff --git a/pandas/tests/window/test_moments.py b/pandas/tests/window/test_moments.py
index f1c89d3c6c1b4..2c65c9e2ac82c 100644
--- a/pandas/tests/window/test_moments.py
+++ b/pandas/tests/window/test_moments.py
@@ -108,7 +108,7 @@ def test_cmov_window_corner(self):
assert np.isnan(result).all()
# empty
- vals = pd.Series([])
+ vals = pd.Series([], dtype=object)
result = vals.rolling(5, center=True, win_type="boxcar").mean()
assert len(result) == 0
@@ -674,7 +674,7 @@ def f(x):
self._check_moment_func(np.mean, name="apply", func=f, raw=raw)
- expected = Series([])
+ expected = Series([], dtype="float64")
result = expected.rolling(10).apply(lambda x: x.mean(), raw=raw)
tm.assert_series_equal(result, expected)
@@ -1193,8 +1193,10 @@ def _check_ew(self, name=None, preserve_nan=False):
assert not result[11:].isna().any()
# check series of length 0
- result = getattr(Series().ewm(com=50, min_periods=min_periods), name)()
- tm.assert_series_equal(result, Series())
+ result = getattr(
+ Series(dtype=object).ewm(com=50, min_periods=min_periods), name
+ )()
+ tm.assert_series_equal(result, Series(dtype="float64"))
# check series of length 1
result = getattr(Series([1.0]).ewm(50, min_periods=min_periods), name)()
@@ -1214,7 +1216,7 @@ def _check_ew(self, name=None, preserve_nan=False):
def _create_consistency_data():
def create_series():
return [
- Series(),
+ Series(dtype=object),
Series([np.nan]),
Series([np.nan, np.nan]),
Series([3.0]),
@@ -1989,8 +1991,9 @@ def func(A, B, com, **kwargs):
assert not np.isnan(result.values[11:]).any()
# check series of length 0
- result = func(Series([]), Series([]), 50, min_periods=min_periods)
- tm.assert_series_equal(result, Series([]))
+ empty = Series([], dtype=np.float64)
+ result = func(empty, empty, 50, min_periods=min_periods)
+ tm.assert_series_equal(result, empty)
# check series of length 1
result = func(Series([1.0]), Series([1.0]), 50, min_periods=min_periods)
@@ -2190,7 +2193,7 @@ def test_rolling_functions_window_non_shrinkage_binary(self):
def test_moment_functions_zero_length(self):
# GH 8056
- s = Series()
+ s = Series(dtype=np.float64)
s_expected = s
df1 = DataFrame()
df1_expected = df1
@@ -2409,7 +2412,7 @@ def expanding_mean(x, min_periods=1):
# here to make this pass
self._check_expanding(expanding_mean, np.mean, preserve_nan=False)
- ser = Series([])
+ ser = Series([], dtype=np.float64)
tm.assert_series_equal(ser, ser.expanding().apply(lambda x: x.mean(), raw=raw))
# GH 8080
| - [x] closes #17261
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29405 | 2019-11-04T23:20:57Z | 2019-12-05T19:00:16Z | 2019-12-05T19:00:15Z | 2019-12-05T22:29:50Z |
API: rename labels to codes in core/groupby | diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 8512b6c3ae530..511b87dab087e 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -655,16 +655,17 @@ def value_counts(
rep = partial(np.repeat, repeats=np.add.reduceat(inc, idx))
# multi-index components
- labels = list(map(rep, self.grouper.recons_labels)) + [llab(lab, inc)]
+ codes = self.grouper.recons_codes
+ codes = [rep(level_codes) for level_codes in codes] + [llab(lab, inc)]
levels = [ping.group_index for ping in self.grouper.groupings] + [lev]
names = self.grouper.names + [self._selection_name]
if dropna:
- mask = labels[-1] != -1
+ mask = codes[-1] != -1
if mask.all():
dropna = False
else:
- out, labels = out[mask], [label[mask] for label in labels]
+ out, codes = out[mask], [level_codes[mask] for level_codes in codes]
if normalize:
out = out.astype("float")
@@ -680,11 +681,11 @@ def value_counts(
if sort and bins is None:
cat = ids[inc][mask] if dropna else ids[inc]
sorter = np.lexsort((out if ascending else -out, cat))
- out, labels[-1] = out[sorter], labels[-1][sorter]
+ out, codes[-1] = out[sorter], codes[-1][sorter]
if bins is None:
mi = MultiIndex(
- levels=levels, codes=labels, names=names, verify_integrity=False
+ levels=levels, codes=codes, names=names, verify_integrity=False
)
if is_integer_dtype(out):
@@ -694,14 +695,14 @@ def value_counts(
# for compat. with libgroupby.value_counts need to ensure every
# bin is present at every index level, null filled with zeros
diff = np.zeros(len(out), dtype="bool")
- for lab in labels[:-1]:
- diff |= np.r_[True, lab[1:] != lab[:-1]]
+ for level_codes in codes[:-1]:
+ diff |= np.r_[True, level_codes[1:] != level_codes[:-1]]
ncat, nbin = diff.sum(), len(levels[-1])
left = [np.repeat(np.arange(ncat), nbin), np.tile(np.arange(nbin), ncat)]
- right = [diff.cumsum() - 1, labels[-1]]
+ right = [diff.cumsum() - 1, codes[-1]]
_, idx = _get_join_indexers(left, right, sort=False, how="left")
out = np.where(idx != -1, out[idx], 0)
@@ -711,7 +712,10 @@ def value_counts(
out, left[-1] = out[sorter], left[-1][sorter]
# build the multi-index w/ full levels
- codes = list(map(lambda lab: np.repeat(lab[diff], nbin), labels[:-1]))
+ def build_codes(lev_codes: np.ndarray) -> np.ndarray:
+ return np.repeat(lev_codes[diff], nbin)
+
+ codes = [build_codes(lev_codes) for lev_codes in codes[:-1]]
codes.append(left[-1])
mi = MultiIndex(levels=levels, codes=codes, names=names, verify_integrity=False)
@@ -758,7 +762,7 @@ def pct_change(self, periods=1, fill_method="pad", limit=None, freq=None):
)
)
filled = getattr(self, fill_method)(limit=limit)
- fill_grp = filled.groupby(self.grouper.labels)
+ fill_grp = filled.groupby(self.grouper.codes)
shifted = fill_grp.shift(periods=periods, freq=freq)
return (filled / shifted) - 1
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index fa4a184e8f7a4..81ba594c97391 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -2349,7 +2349,7 @@ def pct_change(self, periods=1, fill_method="pad", limit=None, freq=None, axis=0
)
)
filled = getattr(self, fill_method)(limit=limit)
- fill_grp = filled.groupby(self.grouper.labels)
+ fill_grp = filled.groupby(self.grouper.codes)
shifted = fill_grp.shift(periods=periods, freq=freq)
return (filled / shifted) - 1
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index 45d2a819ae5ad..dc6336b17ac1e 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -3,7 +3,7 @@
split-apply-combine paradigm.
"""
-from typing import Tuple
+from typing import Optional, Tuple
import warnings
import numpy as np
@@ -21,6 +21,7 @@
)
from pandas.core.dtypes.generic import ABCSeries
+from pandas._typing import FrameOrSeries
import pandas.core.algorithms as algorithms
from pandas.core.arrays import Categorical, ExtensionArray
import pandas.core.common as com
@@ -228,10 +229,10 @@ class Grouping:
----------
index : Index
grouper :
- obj :
+ obj Union[DataFrame, Series]:
name :
level :
- observed : boolean, default False
+ observed : bool, default False
If we are a Categorical, use the observed values
in_axis : if the Grouping is a column in self.obj and hence among
Groupby.exclusions list
@@ -240,25 +241,22 @@ class Grouping:
-------
**Attributes**:
* indices : dict of {group -> index_list}
- * labels : ndarray, group labels
- * ids : mapping of label -> group
- * counts : array of group counts
+ * codes : ndarray, group codes
* group_index : unique groups
* groups : dict of {group -> label_list}
"""
def __init__(
self,
- index,
+ index: Index,
grouper=None,
- obj=None,
+ obj: Optional[FrameOrSeries] = None,
name=None,
level=None,
- sort=True,
- observed=False,
- in_axis=False,
+ sort: bool = True,
+ observed: bool = False,
+ in_axis: bool = False,
):
-
self.name = name
self.level = level
self.grouper = _convert_grouper(index, grouper)
@@ -290,12 +288,12 @@ def __init__(
if self.name is None:
self.name = index.names[level]
- self.grouper, self._labels, self._group_index = index._get_grouper_for_level( # noqa: E501
+ self.grouper, self._codes, self._group_index = index._get_grouper_for_level( # noqa: E501
self.grouper, level
)
# a passed Grouper like, directly get the grouper in the same way
- # as single grouper groupby, use the group_info to get labels
+ # as single grouper groupby, use the group_info to get codes
elif isinstance(self.grouper, Grouper):
# get the new grouper; we already have disambiguated
# what key/level refer to exactly, don't need to
@@ -308,7 +306,7 @@ def __init__(
self.grouper = grouper._get_grouper()
else:
- if self.grouper is None and self.name is not None:
+ if self.grouper is None and self.name is not None and self.obj is not None:
self.grouper = self.obj[self.name]
elif isinstance(self.grouper, (list, tuple)):
@@ -324,7 +322,7 @@ def __init__(
# we make a CategoricalIndex out of the cat grouper
# preserving the categories / ordered attributes
- self._labels = self.grouper.codes
+ self._codes = self.grouper.codes
if observed:
codes = algorithms.unique1d(self.grouper.codes)
codes = codes[codes != -1]
@@ -380,11 +378,11 @@ def __repr__(self):
def __iter__(self):
return iter(self.indices)
- _labels = None
- _group_index = None
+ _codes = None # type: np.ndarray
+ _group_index = None # type: Index
@property
- def ngroups(self):
+ def ngroups(self) -> int:
return len(self.group_index)
@cache_readonly
@@ -397,38 +395,38 @@ def indices(self):
return values._reverse_indexer()
@property
- def labels(self):
- if self._labels is None:
- self._make_labels()
- return self._labels
+ def codes(self) -> np.ndarray:
+ if self._codes is None:
+ self._make_codes()
+ return self._codes
@cache_readonly
- def result_index(self):
+ def result_index(self) -> Index:
if self.all_grouper is not None:
return recode_from_groupby(self.all_grouper, self.sort, self.group_index)
return self.group_index
@property
- def group_index(self):
+ def group_index(self) -> Index:
if self._group_index is None:
- self._make_labels()
+ self._make_codes()
return self._group_index
- def _make_labels(self):
- if self._labels is None or self._group_index is None:
+ def _make_codes(self) -> None:
+ if self._codes is None or self._group_index is None:
# we have a list of groupers
if isinstance(self.grouper, BaseGrouper):
- labels = self.grouper.label_info
+ codes = self.grouper.codes_info
uniques = self.grouper.result_index
else:
- labels, uniques = algorithms.factorize(self.grouper, sort=self.sort)
+ codes, uniques = algorithms.factorize(self.grouper, sort=self.sort)
uniques = Index(uniques, name=self.name)
- self._labels = labels
+ self._codes = codes
self._group_index = uniques
@cache_readonly
- def groups(self):
- return self.index.groupby(Categorical.from_codes(self.labels, self.group_index))
+ def groups(self) -> dict:
+ return self.index.groupby(Categorical.from_codes(self.codes, self.group_index))
def _get_grouper(
@@ -678,7 +676,7 @@ def _is_label_like(val):
return isinstance(val, (str, tuple)) or (val is not None and is_scalar(val))
-def _convert_grouper(axis, grouper):
+def _convert_grouper(axis: Index, grouper):
if isinstance(grouper, dict):
return grouper.get
elif isinstance(grouper, Series):
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 5bad73bf40ff5..2c8aa1294451d 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,7 +7,7 @@
"""
import collections
-from typing import List, Optional, Type
+from typing import List, Optional, Sequence, Type
import numpy as np
@@ -41,7 +41,7 @@
import pandas.core.common as com
from pandas.core.frame import DataFrame
from pandas.core.generic import NDFrame
-from pandas.core.groupby import base
+from pandas.core.groupby import base, grouper
from pandas.core.index import Index, MultiIndex, ensure_index
from pandas.core.series import Series
from pandas.core.sorting import (
@@ -62,13 +62,13 @@ class BaseGrouper:
Parameters
----------
axis : Index
- groupings : array of grouping
+ groupings : Sequence[Grouping]
all the grouping instances to handle in this grouper
for example for grouper list to groupby, need to pass the list
- sort : boolean, default True
+ sort : bool, default True
whether this grouper will give sorted result or not
- group_keys : boolean, default True
- mutated : boolean, default False
+ group_keys : bool, default True
+ mutated : bool, default False
indexer : intp array, optional
the indexer created by Grouper
some groupers (TimeGrouper) will sort its axis and its
@@ -79,16 +79,17 @@ class BaseGrouper:
def __init__(
self,
axis: Index,
- groupings,
- sort=True,
- group_keys=True,
- mutated=False,
- indexer=None,
+ groupings: "Sequence[grouper.Grouping]",
+ sort: bool = True,
+ group_keys: bool = True,
+ mutated: bool = False,
+ indexer: Optional[np.ndarray] = None,
):
assert isinstance(axis, Index), axis
+
self._filter_empty_groups = self.compressed = len(groupings) != 1
self.axis = axis
- self.groupings = groupings
+ self.groupings = groupings # type: Sequence[grouper.Grouping]
self.sort = sort
self.group_keys = group_keys
self.mutated = mutated
@@ -139,7 +140,7 @@ def _get_group_keys(self):
comp_ids, _, ngroups = self.group_info
# provide "flattened" iterator for multi-group setting
- return get_flattened_iterator(comp_ids, ngroups, self.levels, self.labels)
+ return get_flattened_iterator(comp_ids, ngroups, self.levels, self.codes)
def apply(self, f, data, axis: int = 0):
mutated = self.mutated
@@ -210,13 +211,13 @@ def indices(self):
if len(self.groupings) == 1:
return self.groupings[0].indices
else:
- label_list = [ping.labels for ping in self.groupings]
+ codes_list = [ping.codes for ping in self.groupings]
keys = [com.values_from_object(ping.group_index) for ping in self.groupings]
- return get_indexer_dict(label_list, keys)
+ return get_indexer_dict(codes_list, keys)
@property
- def labels(self):
- return [ping.labels for ping in self.groupings]
+ def codes(self):
+ return [ping.codes for ping in self.groupings]
@property
def levels(self):
@@ -256,46 +257,46 @@ def is_monotonic(self) -> bool:
@cache_readonly
def group_info(self):
- comp_ids, obs_group_ids = self._get_compressed_labels()
+ comp_ids, obs_group_ids = self._get_compressed_codes()
ngroups = len(obs_group_ids)
comp_ids = ensure_int64(comp_ids)
return comp_ids, obs_group_ids, ngroups
@cache_readonly
- def label_info(self):
- # return the labels of items in original grouped axis
- labels, _, _ = self.group_info
+ def codes_info(self):
+ # return the codes of items in original grouped axis
+ codes, _, _ = self.group_info
if self.indexer is not None:
- sorter = np.lexsort((labels, self.indexer))
- labels = labels[sorter]
- return labels
-
- def _get_compressed_labels(self):
- all_labels = [ping.labels for ping in self.groupings]
- if len(all_labels) > 1:
- group_index = get_group_index(all_labels, self.shape, sort=True, xnull=True)
+ sorter = np.lexsort((codes, self.indexer))
+ codes = codes[sorter]
+ return codes
+
+ def _get_compressed_codes(self):
+ all_codes = [ping.codes for ping in self.groupings]
+ if len(all_codes) > 1:
+ group_index = get_group_index(all_codes, self.shape, sort=True, xnull=True)
return compress_group_index(group_index, sort=self.sort)
ping = self.groupings[0]
- return ping.labels, np.arange(len(ping.group_index))
+ return ping.codes, np.arange(len(ping.group_index))
@cache_readonly
def ngroups(self) -> int:
return len(self.result_index)
@property
- def recons_labels(self):
+ def recons_codes(self):
comp_ids, obs_ids, _ = self.group_info
- labels = (ping.labels for ping in self.groupings)
- return decons_obs_group_ids(comp_ids, obs_ids, self.shape, labels, xnull=True)
+ codes = (ping.codes for ping in self.groupings)
+ return decons_obs_group_ids(comp_ids, obs_ids, self.shape, codes, xnull=True)
@cache_readonly
def result_index(self):
if not self.compressed and len(self.groupings) == 1:
return self.groupings[0].result_index.rename(self.names[0])
- codes = self.recons_labels
+ codes = self.recons_codes
levels = [ping.result_index for ping in self.groupings]
result = MultiIndex(
levels=levels, codes=codes, verify_integrity=False, names=self.names
@@ -307,9 +308,9 @@ def get_group_levels(self):
return [self.groupings[0].result_index]
name_list = []
- for ping, labels in zip(self.groupings, self.recons_labels):
- labels = ensure_platform_int(labels)
- levels = ping.result_index.take(labels)
+ for ping, codes in zip(self.groupings, self.recons_codes):
+ codes = ensure_platform_int(codes)
+ levels = ping.result_index.take(codes)
name_list.append(levels)
@@ -490,7 +491,7 @@ def _cython_operation(
else:
out_dtype = "object"
- labels, _, _ = self.group_info
+ codes, _, _ = self.group_info
if kind == "aggregate":
result = _maybe_fill(
@@ -498,7 +499,7 @@ def _cython_operation(
)
counts = np.zeros(self.ngroups, dtype=np.int64)
result = self._aggregate(
- result, counts, values, labels, func, is_datetimelike, min_count
+ result, counts, values, codes, func, is_datetimelike, min_count
)
elif kind == "transform":
result = _maybe_fill(
@@ -507,7 +508,7 @@ def _cython_operation(
# TODO: min_count
result = self._transform(
- result, values, labels, func, is_datetimelike, **kwargs
+ result, values, codes, func, is_datetimelike, **kwargs
)
if is_integer_dtype(result) and not is_datetimelike:
diff --git a/pandas/tests/groupby/test_grouping.py b/pandas/tests/groupby/test_grouping.py
index e1fd8d7da6833..e4edc64016567 100644
--- a/pandas/tests/groupby/test_grouping.py
+++ b/pandas/tests/groupby/test_grouping.py
@@ -559,12 +559,12 @@ def test_level_preserve_order(self, sort, labels, mframe):
# GH 17537
grouped = mframe.groupby(level=0, sort=sort)
exp_labels = np.array(labels, np.intp)
- tm.assert_almost_equal(grouped.grouper.labels[0], exp_labels)
+ tm.assert_almost_equal(grouped.grouper.codes[0], exp_labels)
def test_grouping_labels(self, mframe):
grouped = mframe.groupby(mframe.index.get_level_values(0))
exp_labels = np.array([2, 2, 2, 0, 0, 1, 1, 3, 3, 3], dtype=np.intp)
- tm.assert_almost_equal(grouped.grouper.labels[0], exp_labels)
+ tm.assert_almost_equal(grouped.grouper.codes[0], exp_labels)
def test_list_grouper_with_nat(self):
# GH 14715
diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index 5a2f189ad8d10..4ba32c377a345 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -621,8 +621,8 @@ def _check_types(l, r, obj="Index"):
def _get_ilevel_values(index, level):
# accept level number only
unique = index.levels[level]
- labels = index.codes[level]
- filled = take_1d(unique.values, labels, fill_value=unique._na_value)
+ level_codes = index.codes[level]
+ filled = take_1d(unique.values, level_codes, fill_value=unique._na_value)
values = unique._shallow_copy(filled, name=index.names[level])
return values
| This PR renames the various ``*label*`` names in core/groupby to like-named ``*codes*``.
I think the name ``label`` can be confused by the single values in a index, and ``codes`` sound smore like an array of ints, so by renaming we get a cleaner nomenclature, IMO.
All these attributes/methods are internal, so no deprecations needed. | https://api.github.com/repos/pandas-dev/pandas/pulls/29402 | 2019-11-04T22:24:21Z | 2019-11-07T01:34:47Z | 2019-11-07T01:34:47Z | 2019-11-07T01:34:51Z |
removing kendall tests | diff --git a/asv_bench/benchmarks/stat_ops.py b/asv_bench/benchmarks/stat_ops.py
index ed5ebfa61594e..ec67394e55a1e 100644
--- a/asv_bench/benchmarks/stat_ops.py
+++ b/asv_bench/benchmarks/stat_ops.py
@@ -7,20 +7,14 @@
class FrameOps:
- params = [ops, ["float", "int"], [0, 1], [True, False]]
- param_names = ["op", "dtype", "axis", "use_bottleneck"]
+ params = [ops, ["float", "int"], [0, 1]]
+ param_names = ["op", "dtype", "axis"]
- def setup(self, op, dtype, axis, use_bottleneck):
+ def setup(self, op, dtype, axis):
df = pd.DataFrame(np.random.randn(100000, 4)).astype(dtype)
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
-
- nanops._USE_BOTTLENECK = use_bottleneck
self.df_func = getattr(df, op)
- def time_op(self, op, dtype, axis, use_bottleneck):
+ def time_op(self, op, dtype, axis):
self.df_func(axis=axis)
@@ -46,20 +40,14 @@ def time_op(self, level, op):
class SeriesOps:
- params = [ops, ["float", "int"], [True, False]]
- param_names = ["op", "dtype", "use_bottleneck"]
+ params = [ops, ["float", "int"]]
+ param_names = ["op", "dtype"]
- def setup(self, op, dtype, use_bottleneck):
+ def setup(self, op, dtype):
s = pd.Series(np.random.randn(100000)).astype(dtype)
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
-
- nanops._USE_BOTTLENECK = use_bottleneck
self.s_func = getattr(s, op)
- def time_op(self, op, dtype, use_bottleneck):
+ def time_op(self, op, dtype):
self.s_func()
@@ -101,61 +89,49 @@ def time_average_old(self, constructor, pct):
class Correlation:
- params = [["spearman", "kendall", "pearson"], [True, False]]
- param_names = ["method", "use_bottleneck"]
+ params = [["spearman", "kendall", "pearson"]]
+ param_names = ["method"]
- def setup(self, method, use_bottleneck):
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
+ def setup(self, method):
+ self.df = pd.DataFrame(np.random.randn(500, 15))
+ self.df2 = pd.DataFrame(np.random.randn(500, 15))
+ self.df_wide = pd.DataFrame(np.random.randn(500, 100))
+ self.df_wide_nans = self.df_wide.where(np.random.random((500, 100)) < 0.9)
+ self.s = pd.Series(np.random.randn(500))
+ self.s2 = pd.Series(np.random.randn(500))
- nanops._USE_BOTTLENECK = use_bottleneck
- self.df = pd.DataFrame(np.random.randn(1000, 30))
- self.df2 = pd.DataFrame(np.random.randn(1000, 30))
- self.df_wide = pd.DataFrame(np.random.randn(1000, 200))
- self.df_wide_nans = self.df_wide.where(np.random.random((1000, 200)) < 0.9)
- self.s = pd.Series(np.random.randn(1000))
- self.s2 = pd.Series(np.random.randn(1000))
-
- def time_corr(self, method, use_bottleneck):
+ def time_corr(self, method):
self.df.corr(method=method)
- def time_corr_wide(self, method, use_bottleneck):
+ def time_corr_wide(self, method):
self.df_wide.corr(method=method)
- def time_corr_wide_nans(self, method, use_bottleneck):
+ def time_corr_wide_nans(self, method):
self.df_wide_nans.corr(method=method)
- def peakmem_corr_wide(self, method, use_bottleneck):
+ def peakmem_corr_wide(self, method):
self.df_wide.corr(method=method)
- def time_corr_series(self, method, use_bottleneck):
+ def time_corr_series(self, method):
self.s.corr(self.s2, method=method)
- def time_corrwith_cols(self, method, use_bottleneck):
+ def time_corrwith_cols(self, method):
self.df.corrwith(self.df2, method=method)
- def time_corrwith_rows(self, method, use_bottleneck):
+ def time_corrwith_rows(self, method):
self.df.corrwith(self.df2, axis=1, method=method)
class Covariance:
- params = [[True, False]]
- param_names = ["use_bottleneck"]
-
- def setup(self, use_bottleneck):
- try:
- pd.options.compute.use_bottleneck = use_bottleneck
- except TypeError:
- from pandas.core import nanops
+ params = []
+ param_names = []
- nanops._USE_BOTTLENECK = use_bottleneck
+ def setup(self):
self.s = pd.Series(np.random.randn(100000))
self.s2 = pd.Series(np.random.randn(100000))
- def time_cov_series(self, use_bottleneck):
+ def time_cov_series(self):
self.s.cov(self.s2)
| closes #29270
Following is the output after removing "kendall"
```
· Creating environments
· Discovering benchmarks
·· Uninstalling from conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
·· Building cd59acf5 <fix-kendall-issues> for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt................................................
·· Installing cd59acf5 <fix-kendall-issues> into conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt..
· Running 14 total benchmarks (2 commits * 1 environments * 7 benchmarks)
[ 0.00%] · For pandas commit 165d5ee4 <master> (round 1/2):
[ 0.00%] ·· Building for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt...
[ 0.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 7.14%] ··· Running (stat_ops.Correlation.time_corr--)....
[ 21.43%] ··· Running (stat_ops.Correlation.time_corrwith_cols--)..
[ 25.00%] · For pandas commit cd59acf5 <fix-kendall-issues> (round 1/2):
[ 25.00%] ·· Building for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt...
[ 25.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 32.14%] ··· Running (stat_ops.Correlation.time_corr--)....
[ 46.43%] ··· Running (stat_ops.Correlation.time_corrwith_cols--)..
[ 50.00%] · For pandas commit cd59acf5 <fix-kendall-issues> (round 2/2):
[ 50.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 53.57%] ··· stat_ops.Correlation.peakmem_corr_wide ok
[ 53.57%] ··· ========== ====== =======
-- use_bottleneck
---------- --------------
method True False
========== ====== =======
spearman 108M 108M
pearson 105M 105M
========== ====== =======
[ 57.14%] ··· stat_ops.Correlation.time_corr ok
[ 57.14%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 8.74±1ms 8.71±2ms
pearson 2.88±0.6ms 2.84±0.4ms
========== ============ ============
[ 60.71%] ··· stat_ops.Correlation.time_corr_series ok
[ 60.71%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 1.34±0.2ms 1.35±0.3ms
pearson 317±60μs 319±70μs
========== ============ ============
[ 64.29%] ··· stat_ops.Correlation.time_corr_wide ok
[ 64.29%] ··· ========== ========== ==========
-- use_bottleneck
---------- ---------------------
method True False
========== ========== ==========
spearman 286±40ms 273±10ms
pearson 176±10ms 178±10ms
========== ========== ==========
[ 67.86%] ··· stat_ops.Correlation.time_corr_wide_nans ok
[ 67.86%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 3.23±0.02s 3.20±0.01s
pearson 196±7ms 192±10ms
========== ============ ============
[ 71.43%] ··· stat_ops.Correlation.time_corrwith_cols ok
[ 71.43%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 24.2±1ms 25.5±1ms
pearson 36.1±0.3ms 36.2±0.4ms
========== ============ ============
[ 75.00%] ··· stat_ops.Correlation.time_corrwith_rows ok
[ 75.00%] ··· ========== ========= ==========
-- use_bottleneck
---------- --------------------
method True False
========== ========= ==========
spearman 549±4ms 551±3ms
pearson 884±7ms 890±20ms
========== ========= ==========
[ 75.00%] · For pandas commit 165d5ee4 <master> (round 2/2):
[ 75.00%] ·· Building for conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt...
[ 75.00%] ·· Benchmarking conda-py3.6-Cython-matplotlib-numexpr-numpy-odfpy-openpyxl-pytables-pytest-scipy-sqlalchemy-xlrd-xlsxwriter-xlwt
[ 78.57%] ··· stat_ops.Correlation.peakmem_corr_wide ok
[ 78.57%] ··· ========== ====== =======
-- use_bottleneck
---------- --------------
method True False
========== ====== =======
spearman 108M 108M
pearson 106M 106M
========== ====== =======
[ 82.14%] ··· stat_ops.Correlation.time_corr ok
[ 82.14%] ··· ========== ============ =============
-- use_bottleneck
---------- --------------------------
method True False
========== ============ =============
spearman 10.2±0.2ms 10.1±0.2ms
pearson 3.51±0.2ms 3.54±0.09ms
========== ============ =============
[ 85.71%] ··· stat_ops.Correlation.time_corr_series ok
[ 85.71%] ··· ========== ============= =============
-- use_bottleneck
---------- ---------------------------
method True False
========== ============= =============
spearman 1.52±0.03ms 1.26±0.01ms
pearson 316±10μs 389±0.6μs
========== ============= =============
[ 89.29%] ··· stat_ops.Correlation.time_corr_wide ok
[ 89.29%] ··· ========== ========== ==========
-- use_bottleneck
---------- ---------------------
method True False
========== ========== ==========
spearman 359±20ms 288±10ms
pearson 186±6ms 184±7ms
========== ========== ==========
[ 92.86%] ··· stat_ops.Correlation.time_corr_wide_nans ok
[ 92.86%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 3.29±0.01s 3.26±0.03s
pearson 195±6ms 203±5ms
========== ============ ============
[ 96.43%] ··· stat_ops.Correlation.time_corrwith_cols ok
[ 96.43%] ··· ========== ============ ============
-- use_bottleneck
---------- -------------------------
method True False
========== ============ ============
spearman 24.4±2ms 24.8±0.9ms
pearson 35.8±0.5ms 36.1±0.5ms
========== ============ ============
[100.00%] ··· stat_ops.Correlation.time_corrwith_rows ok
[100.00%] ··· ========== ========== ==========
-- use_bottleneck
---------- ---------------------
method True False
========== ========== ==========
spearman 551±6ms 555±7ms
pearson 894±10ms 899±10ms
========== ========== ==========
before after ratio
[165d5ee4] [cd59acf5]
<master> <fix-kendall-issues>
- 3.54±0.09ms 2.84±0.4ms 0.80 stat_ops.Correlation.time_corr('pearson', False)
SOME BENCHMARKS HAVE CHANGED SIGNIFICANTLY.
PERFORMANCE INCREASED.
``` | https://api.github.com/repos/pandas-dev/pandas/pulls/29401 | 2019-11-04T22:02:52Z | 2019-12-05T16:35:46Z | 2019-12-05T16:35:45Z | 2019-12-05T16:35:52Z |
REF: implement first_valid_index in core.missing | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index cbeee88d75b51..bafc37d478fdb 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -83,6 +83,7 @@
from pandas.core.indexes.period import Period, PeriodIndex
import pandas.core.indexing as indexing
from pandas.core.internals import BlockManager
+from pandas.core.missing import find_valid_index
from pandas.core.ops import _align_method_FRAME
from pandas.io.formats import format as fmt
@@ -10870,27 +10871,11 @@ def _find_valid_index(self, how: str):
-------
idx_first_valid : type of index
"""
- assert how in ["first", "last"]
- if len(self) == 0: # early stop
+ idxpos = find_valid_index(self._values, how)
+ if idxpos is None:
return None
- is_valid = ~self.isna()
-
- if self.ndim == 2:
- is_valid = is_valid.any(1) # reduce axis 1
-
- if how == "first":
- idxpos = is_valid.values[::].argmax()
-
- if how == "last":
- idxpos = len(self) - 1 - is_valid.values[::-1].argmax()
-
- chk_notna = is_valid.iat[idxpos]
- idx = self.index[idxpos]
-
- if not chk_notna:
- return None
- return idx
+ return self.index[idxpos]
@Appender(
_shared_docs["valid_index"] % {"position": "first", "klass": "Series/DataFrame"}
diff --git a/pandas/core/missing.py b/pandas/core/missing.py
index 5a1bf6d37b081..c1e63a49a0f0a 100644
--- a/pandas/core/missing.py
+++ b/pandas/core/missing.py
@@ -128,6 +128,43 @@ def clean_interp_method(method, **kwargs):
return method
+def find_valid_index(values, how: str):
+ """
+ Retrieves the index of the first valid value.
+
+ Parameters
+ ----------
+ values : ndarray or ExtensionArray
+ how : {'first', 'last'}
+ Use this parameter to change between the first or last valid index.
+
+ Returns
+ -------
+ int or None
+ """
+ assert how in ["first", "last"]
+
+ if len(values) == 0: # early stop
+ return None
+
+ is_valid = ~isna(values)
+
+ if values.ndim == 2:
+ is_valid = is_valid.any(1) # reduce axis 1
+
+ if how == "first":
+ idxpos = is_valid[::].argmax()
+
+ if how == "last":
+ idxpos = len(values) - 1 - is_valid[::-1].argmax()
+
+ chk_notna = is_valid[idxpos]
+
+ if not chk_notna:
+ return None
+ return idxpos
+
+
def interpolate_1d(
xvalues,
yvalues,
@@ -192,14 +229,10 @@ def interpolate_1d(
# default limit is unlimited GH #16282
limit = algos._validate_limit(nobs=None, limit=limit)
- from pandas import Series
-
- ys = Series(yvalues)
-
# These are sets of index pointers to invalid values... i.e. {0, 1, etc...
all_nans = set(np.flatnonzero(invalid))
- start_nans = set(range(ys.first_valid_index()))
- end_nans = set(range(1 + ys.last_valid_index(), len(valid)))
+ start_nans = set(range(find_valid_index(yvalues, "first")))
+ end_nans = set(range(1 + find_valid_index(yvalues, "last"), len(valid)))
mid_nans = all_nans - start_nans - end_nans
# Like the sets above, preserve_nans contains indices of invalid values,
| The implementation here operates on the values (ndarray or EA) instead of on the Series/DataFrame.
This lets us avoid a runtime import of Series, so core.missing joins the Simple Dependencies Club. | https://api.github.com/repos/pandas-dev/pandas/pulls/29400 | 2019-11-04T21:43:44Z | 2019-11-05T17:13:17Z | 2019-11-05T17:13:16Z | 2019-11-05T17:47:01Z |
PR09 Batch 2 | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 2fb552af717fc..1a2f906f97152 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -1073,7 +1073,7 @@ def argmax(self, axis=None, skipna=True, *args, **kwargs):
Parameters
----------
axis : {None}
- Dummy argument for consistency with Series
+ Dummy argument for consistency with Series.
skipna : bool, default True
Returns
@@ -1096,7 +1096,7 @@ def min(self, axis=None, skipna=True, *args, **kwargs):
Parameters
----------
axis : {None}
- Dummy argument for consistency with Series
+ Dummy argument for consistency with Series.
skipna : bool, default True
Returns
@@ -1137,7 +1137,7 @@ def argmin(self, axis=None, skipna=True, *args, **kwargs):
Parameters
----------
axis : {None}
- Dummy argument for consistency with Series
+ Dummy argument for consistency with Series.
skipna : bool, default True
Returns
@@ -1486,7 +1486,7 @@ def memory_usage(self, deep=False):
----------
deep : bool
Introspect the data deeply, interrogate
- `object` dtypes for system-level memory consumption
+ `object` dtypes for system-level memory consumption.
Returns
-------
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 301426d237d19..feee6dca23ac8 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -176,11 +176,11 @@ class Index(IndexOpsMixin, PandasObject):
If an actual dtype is provided, we coerce to that dtype if it's safe.
Otherwise, an error will be raised.
copy : bool
- Make a copy of input ndarray
+ Make a copy of input ndarray.
name : object
- Name to be stored in the index
+ Name to be stored in the index.
tupleize_cols : bool (default: True)
- When True, attempt to create a MultiIndex if possible
+ When True, attempt to create a MultiIndex if possible.
See Also
--------
@@ -791,13 +791,13 @@ def astype(self, dtype, copy=True):
Parameters
----------
indices : list
- Indices to be taken
+ Indices to be taken.
axis : int, optional
The axis over which to select values, always 0.
allow_fill : bool, default True
fill_value : bool, default None
If allow_fill=True and fill_value is not None, indices specified by
- -1 is regarded as NA. If Index doesn't hold NA, raise ValueError
+ -1 is regarded as NA. If Index doesn't hold NA, raise ValueError.
Returns
-------
@@ -1077,7 +1077,7 @@ def to_native_types(self, slicer=None, **kwargs):
2) quoting : bool or None
Whether or not there are quoted values in `self`
3) date_format : str
- The format used to represent date-like values
+ The format used to represent date-like values.
Returns
-------
@@ -2001,7 +2001,7 @@ def notna(self):
downcast : dict, default is None
a dict of item->dtype of what to downcast if possible,
or the string 'infer' which will try to downcast to an appropriate
- equal type (e.g. float64 to int64 if possible)
+ equal type (e.g. float64 to int64 if possible).
Returns
-------
@@ -2056,7 +2056,7 @@ def dropna(self, how="any"):
Parameters
----------
level : int or str, optional, default None
- Only return values from specified level (for MultiIndex)
+ Only return values from specified level (for MultiIndex).
.. versionadded:: 0.23.0
@@ -3413,7 +3413,7 @@ def _reindex_non_unique(self, target):
return_indexers : bool, default False
sort : bool, default False
Sort the join keys lexicographically in the result Index. If False,
- the order of the join keys depends on the join type (how keyword)
+ the order of the join keys depends on the join type (how keyword).
Returns
-------
@@ -4923,9 +4923,9 @@ def slice_indexer(self, start=None, end=None, step=None, kind=None):
Parameters
----------
start : label, default None
- If None, defaults to the beginning
+ If None, defaults to the beginning.
end : label, default None
- If None, defaults to the end
+ If None, defaults to the end.
step : int, default None
kind : str, default None
@@ -5122,11 +5122,11 @@ def slice_locs(self, start=None, end=None, step=None, kind=None):
Parameters
----------
start : label, default None
- If None, defaults to the beginning
+ If None, defaults to the beginning.
end : label, default None
- If None, defaults to the end
+ If None, defaults to the end.
step : int, defaults None
- If None, defaults to 1
+ If None, defaults to 1.
kind : {'ix', 'loc', 'getitem'} or None
Returns
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index 66deacac37789..caaf55546189c 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -737,19 +737,18 @@ def _set_levels(
def set_levels(self, levels, level=None, inplace=False, verify_integrity=True):
"""
- Set new levels on MultiIndex. Defaults to returning
- new index.
+ Set new levels on MultiIndex. Defaults to returning new index.
Parameters
----------
levels : sequence or list of sequence
- new level(s) to apply
+ New level(s) to apply.
level : int, level name, or sequence of int/level names (default None)
- level(s) to set (None for all levels)
+ Level(s) to set (None for all levels).
inplace : bool
- if True, mutates in place
+ If True, mutates in place.
verify_integrity : bool (default True)
- if True, checks that levels and codes are compatible
+ If True, checks that levels and codes are compatible.
Returns
-------
diff --git a/pandas/core/indexes/numeric.py b/pandas/core/indexes/numeric.py
index e83360dc701f3..46bb8eafee3b9 100644
--- a/pandas/core/indexes/numeric.py
+++ b/pandas/core/indexes/numeric.py
@@ -176,9 +176,9 @@ def _union(self, other, sort):
data : array-like (1-dimensional)
dtype : NumPy dtype (default: %(dtype)s)
copy : bool
- Make a copy of input ndarray
+ Make a copy of input ndarray.
name : object
- Name to be stored in the index
+ Name to be stored in the index.
Attributes
----------
diff --git a/pandas/core/indexes/range.py b/pandas/core/indexes/range.py
index 6e2d500f4c5ab..5fa3431fc97c0 100644
--- a/pandas/core/indexes/range.py
+++ b/pandas/core/indexes/range.py
@@ -51,7 +51,7 @@ class RangeIndex(Int64Index):
stop : int (default: 0)
step : int (default: 1)
name : object, optional
- Name to be stored in the index
+ Name to be stored in the index.
copy : bool, default False
Unused, accepted for homogeneity with other index types.
diff --git a/pandas/core/reshape/tile.py b/pandas/core/reshape/tile.py
index 95534755b8beb..073bb4707f890 100644
--- a/pandas/core/reshape/tile.py
+++ b/pandas/core/reshape/tile.py
@@ -275,17 +275,18 @@ def qcut(
duplicates: str = "raise",
):
"""
- Quantile-based discretization function. Discretize variable into
- equal-sized buckets based on rank or based on sample quantiles. For example
- 1000 values for 10 quantiles would produce a Categorical object indicating
- quantile membership for each data point.
+ Quantile-based discretization function.
+
+ Discretize variable into equal-sized buckets based on rank or based
+ on sample quantiles. For example 1000 values for 10 quantiles would
+ produce a Categorical object indicating quantile membership for each data point.
Parameters
----------
x : 1d ndarray or Series
q : int or list-like of int
Number of quantiles. 10 for deciles, 4 for quartiles, etc. Alternately
- array of quantiles, e.g. [0, .25, .5, .75, 1.] for quartiles
+ array of quantiles, e.g. [0, .25, .5, .75, 1.] for quartiles.
labels : array or bool, default None
Used as labels for the resulting bins. Must be of the same length as
the resulting bins. If False, return only integer indicators of the
| Another batch of commits for #28602. Also fixes a few summary formatting errors and PR08 capitalization errors. | https://api.github.com/repos/pandas-dev/pandas/pulls/29396 | 2019-11-04T14:58:19Z | 2019-11-04T16:11:23Z | 2019-11-04T16:11:23Z | 2020-01-06T16:47:02Z |
DOC: remove okwarning once pyarrow 0.12 is released | diff --git a/doc/source/user_guide/io.rst b/doc/source/user_guide/io.rst
index 173bcf7537154..f9fbc33cba966 100644
--- a/doc/source/user_guide/io.rst
+++ b/doc/source/user_guide/io.rst
@@ -4685,7 +4685,6 @@ Write to a feather file.
Read from a feather file.
.. ipython:: python
- :okwarning:
result = pd.read_feather('example.feather')
result
@@ -4764,7 +4763,6 @@ Write to a parquet file.
Read from a parquet file.
.. ipython:: python
- :okwarning:
result = pd.read_parquet('example_fp.parquet', engine='fastparquet')
result = pd.read_parquet('example_pa.parquet', engine='pyarrow')
@@ -4839,7 +4837,6 @@ Partitioning Parquet files
Parquet supports partitioning of data based on the values of one or more columns.
.. ipython:: python
- :okwarning:
df = pd.DataFrame({'a': [0, 0, 1, 1], 'b': [0, 1, 0, 1]})
df.to_parquet(fname='test', engine='pyarrow',
diff --git a/environment.yml b/environment.yml
index 4c96ab815dc90..443dc483aedf8 100644
--- a/environment.yml
+++ b/environment.yml
@@ -81,7 +81,7 @@ dependencies:
- html5lib # pandas.read_html
- lxml # pandas.read_html
- openpyxl # pandas.read_excel, DataFrame.to_excel, pandas.ExcelWriter, pandas.ExcelFile
- - pyarrow>=0.13.1 # pandas.read_paquet, DataFrame.to_parquet, pandas.read_feather, DataFrame.to_feather
+ - pyarrow>=0.13.1 # pandas.read_parquet, DataFrame.to_parquet, pandas.read_feather, DataFrame.to_feather
- pyqt>=5.9.2 # pandas.read_clipboard
- pytables>=3.4.2 # pandas.read_hdf, DataFrame.to_hdf
- python-snappy # required by pyarrow
| - [x] closes #24617
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29395 | 2019-11-04T14:05:01Z | 2019-11-04T14:53:10Z | 2019-11-04T14:53:09Z | 2019-11-04T14:53:10Z |
BUG: GH25495 incorrect dtype when using .loc to set Categorical value for column in 1-row DataFrame | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index f0ba1250b7f8d..c9e2e7e133133 100755
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -1086,6 +1086,7 @@ Indexing
- Bug when indexing with ``.loc`` where the index was a :class:`CategoricalIndex` with non-string categories didn't work (:issue:`17569`, :issue:`30225`)
- :meth:`Index.get_indexer_non_unique` could fail with ``TypeError`` in some cases, such as when searching for ints in a string index (:issue:`28257`)
- Bug in :meth:`Float64Index.get_loc` incorrectly raising ``TypeError`` instead of ``KeyError`` (:issue:`29189`)
+- Bug in :meth:`DataFrame.loc` with incorrect dtype when setting Categorical value in 1-row DataFrame (:issue:`25495`)
- :meth:`MultiIndex.get_loc` can't find missing values when input includes missing values (:issue:`19132`)
- Bug in :meth:`Series.__setitem__` incorrectly assigning values with boolean indexer when the length of new data matches the number of ``True`` values and new data is not a ``Series`` or an ``np.array`` (:issue:`30567`)
- Bug in indexing with a :class:`PeriodIndex` incorrectly accepting integers representing years, use e.g. ``ser.loc["2007"]`` instead of ``ser.loc[2007]`` (:issue:`30763`)
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index a93211edf162b..43edc246da6dd 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -876,7 +876,11 @@ def setitem(self, indexer, value):
# length checking
check_setitem_lengths(indexer, value, values)
-
+ exact_match = (
+ len(arr_value.shape)
+ and arr_value.shape[0] == values.shape[0]
+ and arr_value.size == values.size
+ )
if is_empty_indexer(indexer, arr_value):
# GH#8669 empty indexers
pass
@@ -886,14 +890,21 @@ def setitem(self, indexer, value):
# be e.g. a list; see GH#6043
values[indexer] = value
- # if we are an exact match (ex-broadcasting),
- # then use the resultant dtype
elif (
- len(arr_value.shape)
- and arr_value.shape[0] == values.shape[0]
- and arr_value.size == values.size
+ exact_match
+ and is_categorical_dtype(arr_value.dtype)
+ and not is_categorical_dtype(values)
):
+ # GH25495 - If the current dtype is not categorical,
+ # we need to create a new categorical block
values[indexer] = value
+ return self.make_block(Categorical(self.values, dtype=arr_value.dtype))
+
+ # if we are an exact match (ex-broadcasting),
+ # then use the resultant dtype
+ elif exact_match:
+ values[indexer] = value
+
try:
values = values.astype(arr_value.dtype)
except ValueError:
diff --git a/pandas/tests/frame/indexing/test_categorical.py b/pandas/tests/frame/indexing/test_categorical.py
index 5de38915f04c1..a29c193676db2 100644
--- a/pandas/tests/frame/indexing/test_categorical.py
+++ b/pandas/tests/frame/indexing/test_categorical.py
@@ -354,6 +354,16 @@ def test_functions_no_warnings(self):
df.value, range(0, 105, 10), right=False, labels=labels
)
+ def test_setitem_single_row_categorical(self):
+ # GH 25495
+ df = DataFrame({"Alpha": ["a"], "Numeric": [0]})
+ categories = pd.Categorical(df["Alpha"], categories=["a", "b", "c"])
+ df.loc[:, "Alpha"] = categories
+
+ result = df["Alpha"]
+ expected = Series(categories, index=df.index, name="Alpha")
+ tm.assert_series_equal(result, expected)
+
def test_loc_indexing_preserves_index_category_dtype(self):
# GH 15166
df = DataFrame(
| - [x] closes https://github.com/pandas-dev/pandas/issues/25495
- [x] 1 test added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29393 | 2019-11-04T09:25:45Z | 2020-01-27T12:34:43Z | 2020-01-27T12:34:43Z | 2020-01-28T15:29:45Z |
TST: Test DataFrame.rolling with window as string | diff --git a/pandas/tests/window/test_rolling.py b/pandas/tests/window/test_rolling.py
index 70ba85120af3c..72b72b31d8faa 100644
--- a/pandas/tests/window/test_rolling.py
+++ b/pandas/tests/window/test_rolling.py
@@ -1,4 +1,4 @@
-from datetime import timedelta
+from datetime import datetime, timedelta
import numpy as np
import pytest
@@ -7,7 +7,7 @@
import pandas.util._test_decorators as td
import pandas as pd
-from pandas import DataFrame, Series
+from pandas import DataFrame, Index, Series
from pandas.core.window import Rolling
from pandas.tests.window.common import Base
import pandas.util.testing as tm
@@ -361,3 +361,60 @@ def test_rolling_datetime(self, axis_frame, tz_naive_fixture):
}
)
tm.assert_frame_equal(result, expected)
+
+
+def test_rolling_window_as_string():
+ # see gh-22590
+ date_today = datetime.now()
+ days = pd.date_range(date_today, date_today + timedelta(365), freq="D")
+
+ npr = np.random.RandomState(seed=421)
+
+ data = npr.randint(1, high=100, size=len(days))
+ df = DataFrame({"DateCol": days, "metric": data})
+
+ df.set_index("DateCol", inplace=True)
+ result = df.rolling(window="21D", min_periods=2, closed="left")["metric"].agg("max")
+
+ expData = (
+ [np.nan] * 2
+ + [88.0] * 16
+ + [97.0] * 9
+ + [98.0]
+ + [99.0] * 21
+ + [95.0] * 16
+ + [93.0] * 5
+ + [89.0] * 5
+ + [96.0] * 21
+ + [94.0] * 14
+ + [90.0] * 13
+ + [88.0] * 2
+ + [90.0] * 9
+ + [96.0] * 21
+ + [95.0] * 6
+ + [91.0]
+ + [87.0] * 6
+ + [92.0] * 21
+ + [83.0] * 2
+ + [86.0] * 10
+ + [87.0] * 5
+ + [98.0] * 21
+ + [97.0] * 14
+ + [93.0] * 7
+ + [87.0] * 4
+ + [86.0] * 4
+ + [95.0] * 21
+ + [85.0] * 14
+ + [83.0] * 2
+ + [76.0] * 5
+ + [81.0] * 2
+ + [98.0] * 21
+ + [95.0] * 14
+ + [91.0] * 7
+ + [86.0]
+ + [93.0] * 3
+ + [95.0] * 20
+ )
+
+ expected = Series(expData, index=Index(days, name="DateCol"), name="metric")
+ tm.assert_series_equal(result, expected)
| Closes https://github.com/pandas-dev/pandas/issues/22590 | https://api.github.com/repos/pandas-dev/pandas/pulls/29392 | 2019-11-04T02:51:56Z | 2019-11-04T22:10:32Z | 2019-11-04T22:10:32Z | 2019-11-04T23:56:07Z |
changing docstring for to_csv compression to 1.0.0 | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index f88c26c7bc782..cbeee88d75b51 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -3221,7 +3221,7 @@ def to_csv(
and mode is 'zip' or inferred as 'zip', other entries passed as
additional compression options.
- .. versionchanged:: 0.25.0
+ .. versionchanged:: 1.0.0
May now be a dict with key 'method' as compression mode
and other entries as additional compression options if
| - closes #29328
- Just updating doc-string for to_csv() compression.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29390 | 2019-11-04T00:06:07Z | 2019-11-04T13:50:14Z | 2019-11-04T13:50:14Z | 2019-11-04T13:50:14Z |
CLN core.groupby | diff --git a/pandas/core/base.py b/pandas/core/base.py
index 9586d49c555ff..2fb552af717fc 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -586,9 +586,16 @@ def _aggregate_multiple_funcs(self, arg, _level, _axis):
new_res = colg.aggregate(arg)
except (TypeError, DataError):
pass
- except ValueError:
+ except ValueError as err:
# cannot aggregate
- continue
+ if "Must produce aggregated value" in str(err):
+ # raised directly in _aggregate_named
+ pass
+ elif "no results" in str(err):
+ # raised direcly in _aggregate_multiple_funcs
+ pass
+ else:
+ raise
else:
results.append(new_res)
keys.append(col)
diff --git a/pandas/core/groupby/generic.py b/pandas/core/groupby/generic.py
index 996c178bd7feb..009e83b861523 100644
--- a/pandas/core/groupby/generic.py
+++ b/pandas/core/groupby/generic.py
@@ -244,7 +244,7 @@ def aggregate(self, func=None, *args, **kwargs):
if isinstance(func, str):
return getattr(self, func)(*args, **kwargs)
- if isinstance(func, abc.Iterable):
+ elif isinstance(func, abc.Iterable):
# Catch instances of lists / tuples
# but not the class list / tuple itself.
func = _maybe_mangle_lambdas(func)
@@ -261,8 +261,6 @@ def aggregate(self, func=None, *args, **kwargs):
try:
return self._python_agg_general(func, *args, **kwargs)
- except (AssertionError, TypeError):
- raise
except (ValueError, KeyError, AttributeError, IndexError):
# TODO: IndexError can be removed here following GH#29106
# TODO: AttributeError is caused by _index_data hijinx in
@@ -325,7 +323,7 @@ def _aggregate_multiple_funcs(self, arg, _level):
if name in results:
raise SpecificationError(
"Function names must be unique, found multiple named "
- "{}".format(name)
+ "{name}".format(name=name)
)
# reset the cache so that we
@@ -1464,8 +1462,6 @@ def _transform_item_by_item(self, obj, wrapper):
for i, col in enumerate(obj):
try:
output[col] = self[col].transform(wrapper)
- except AssertionError:
- raise
except TypeError:
# e.g. trying to call nanmean with string values
pass
@@ -1538,8 +1534,8 @@ def filter(self, func, dropna=True, *args, **kwargs):
else:
# non scalars aren't allowed
raise TypeError(
- "filter function returned a %s, "
- "but expected a scalar bool" % type(res).__name__
+ "filter function returned a {typ}, "
+ "but expected a scalar bool".format(typ=type(res).__name__)
)
return self._apply_filter(indices, dropna)
diff --git a/pandas/core/groupby/groupby.py b/pandas/core/groupby/groupby.py
index 404da096d8535..642b1e93a057a 100644
--- a/pandas/core/groupby/groupby.py
+++ b/pandas/core/groupby/groupby.py
@@ -344,7 +344,7 @@ def __init__(
self,
obj: NDFrame,
keys=None,
- axis=0,
+ axis: int = 0,
level=None,
grouper=None,
exclusions=None,
@@ -561,7 +561,9 @@ def __getattr__(self, attr):
return self[attr]
raise AttributeError(
- "%r object has no attribute %r" % (type(self).__name__, attr)
+ "'{typ}' object has no attribute '{attr}'".format(
+ typ=type(self).__name__, attr=attr
+ )
)
@Substitution(
@@ -2486,6 +2488,6 @@ def groupby(obj, by, **kwds):
klass = DataFrameGroupBy
else:
- raise TypeError("invalid type: {}".format(obj))
+ raise TypeError("invalid type: {obj}".format(obj=obj))
return klass(obj, by, **kwds)
diff --git a/pandas/core/groupby/grouper.py b/pandas/core/groupby/grouper.py
index d7eaaca5ac83a..45d2a819ae5ad 100644
--- a/pandas/core/groupby/grouper.py
+++ b/pandas/core/groupby/grouper.py
@@ -172,7 +172,9 @@ def _set_grouper(self, obj, sort=False):
ax = self._grouper.take(obj.index)
else:
if key not in obj._info_axis:
- raise KeyError("The grouper name {0} is not found".format(key))
+ raise KeyError(
+ "The grouper name {key} is not found".format(key=key)
+ )
ax = Index(obj[key], name=key)
else:
@@ -188,7 +190,9 @@ def _set_grouper(self, obj, sort=False):
else:
if level not in (0, ax.name):
- raise ValueError("The level {0} is not valid".format(level))
+ raise ValueError(
+ "The level {level} is not valid".format(level=level)
+ )
# possibly sort
if (self.sort or sort) and not ax.is_monotonic:
@@ -278,7 +282,9 @@ def __init__(
if level is not None:
if not isinstance(level, int):
if level not in index.names:
- raise AssertionError("Level {} not in index".format(level))
+ raise AssertionError(
+ "Level {level} not in index".format(level=level)
+ )
level = index.names.index(level)
if self.name is None:
@@ -344,7 +350,7 @@ def __init__(
):
if getattr(self.grouper, "ndim", 1) != 1:
t = self.name or str(type(self.grouper))
- raise ValueError("Grouper for '{}' not 1-dimensional".format(t))
+ raise ValueError("Grouper for '{t}' not 1-dimensional".format(t=t))
self.grouper = self.index.map(self.grouper)
if not (
hasattr(self.grouper, "__len__")
@@ -352,7 +358,9 @@ def __init__(
):
errmsg = (
"Grouper result violates len(labels) == "
- "len(data)\nresult: %s" % pprint_thing(self.grouper)
+ "len(data)\nresult: {grper}".format(
+ grper=pprint_thing(self.grouper)
+ )
)
self.grouper = None # Try for sanity
raise AssertionError(errmsg)
@@ -426,7 +434,7 @@ def groups(self):
def _get_grouper(
obj: NDFrame,
key=None,
- axis=0,
+ axis: int = 0,
level=None,
sort=True,
observed=False,
@@ -493,7 +501,9 @@ def _get_grouper(
if isinstance(level, str):
if obj.index.name != level:
raise ValueError(
- "level name {} is not the name of the index".format(level)
+ "level name {level} is not the name of the index".format(
+ level=level
+ )
)
elif level > 0 or level < -1:
raise ValueError("level > 0 or level < -1 only valid with MultiIndex")
@@ -582,7 +592,7 @@ def _get_grouper(
exclusions = []
# if the actual grouper should be obj[key]
- def is_in_axis(key):
+ def is_in_axis(key) -> bool:
if not _is_label_like(key):
items = obj._data.items
try:
@@ -594,7 +604,7 @@ def is_in_axis(key):
return True
# if the grouper is obj[name]
- def is_in_obj(gpr):
+ def is_in_obj(gpr) -> bool:
if not hasattr(gpr, "name"):
return False
try:
diff --git a/pandas/core/groupby/ops.py b/pandas/core/groupby/ops.py
index 8d13c37270d7a..7918e463c73ac 100644
--- a/pandas/core/groupby/ops.py
+++ b/pandas/core/groupby/ops.py
@@ -7,6 +7,7 @@
"""
import collections
+from typing import List, Optional
import numpy as np
@@ -385,7 +386,7 @@ def get_func(fname):
return func
- def _cython_operation(self, kind, values, how, axis, min_count=-1, **kwargs):
+ def _cython_operation(self, kind: str, values, how, axis, min_count=-1, **kwargs):
assert kind in ["transform", "aggregate"]
orig_values = values
@@ -398,16 +399,18 @@ def _cython_operation(self, kind, values, how, axis, min_count=-1, **kwargs):
# categoricals are only 1d, so we
# are not setup for dim transforming
if is_categorical_dtype(values) or is_sparse(values):
- raise NotImplementedError("{} dtype not supported".format(values.dtype))
+ raise NotImplementedError(
+ "{dtype} dtype not supported".format(dtype=values.dtype)
+ )
elif is_datetime64_any_dtype(values):
if how in ["add", "prod", "cumsum", "cumprod"]:
raise NotImplementedError(
- "datetime64 type does not support {} operations".format(how)
+ "datetime64 type does not support {how} operations".format(how=how)
)
elif is_timedelta64_dtype(values):
if how in ["prod", "cumprod"]:
raise NotImplementedError(
- "timedelta64 type does not support {} operations".format(how)
+ "timedelta64 type does not support {how} operations".format(how=how)
)
if is_datetime64tz_dtype(values.dtype):
@@ -513,7 +516,7 @@ def _cython_operation(self, kind, values, how, axis, min_count=-1, **kwargs):
result = result[:, 0]
if how in self._name_functions:
- names = self._name_functions[how]()
+ names = self._name_functions[how]() # type: Optional[List[str]]
else:
names = None
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index 13cb0f9aed303..e68a2efc3f4e6 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -361,8 +361,6 @@ def _groupby_and_aggregate(self, how, grouper=None, *args, **kwargs):
result = grouped._aggregate_item_by_item(how, *args, **kwargs)
else:
result = grouped.aggregate(how, *args, **kwargs)
- except AssertionError:
- raise
except DataError:
# we have a non-reducing function; try to evaluate
result = grouped.apply(how, *args, **kwargs)
@@ -1450,7 +1448,7 @@ def _get_resampler(self, obj, kind=None):
raise TypeError(
"Only valid with DatetimeIndex, "
"TimedeltaIndex or PeriodIndex, "
- "but got an instance of %r" % type(ax).__name__
+ "but got an instance of '{typ}'".format(typ=type(ax).__name__)
)
def _get_grouper(self, obj, validate=True):
@@ -1463,7 +1461,7 @@ def _get_time_bins(self, ax):
if not isinstance(ax, DatetimeIndex):
raise TypeError(
"axis must be a DatetimeIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
if len(ax) == 0:
@@ -1539,7 +1537,7 @@ def _get_time_delta_bins(self, ax):
if not isinstance(ax, TimedeltaIndex):
raise TypeError(
"axis must be a TimedeltaIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
if not len(ax):
@@ -1564,7 +1562,7 @@ def _get_time_period_bins(self, ax):
if not isinstance(ax, DatetimeIndex):
raise TypeError(
"axis must be a DatetimeIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
freq = self.freq
@@ -1586,7 +1584,7 @@ def _get_period_bins(self, ax):
if not isinstance(ax, PeriodIndex):
raise TypeError(
"axis must be a PeriodIndex, but got "
- "an instance of %r" % type(ax).__name__
+ "an instance of {typ}".format(typ=type(ax).__name__)
)
memb = ax.asfreq(self.freq, how=self.convention)
| Broken off from local branches doing non-CLN work.
Foreshadowing: some of the TypeErrors we are catching are being caused by `self.axis` not being an Index instead of an int in some cases. | https://api.github.com/repos/pandas-dev/pandas/pulls/29389 | 2019-11-03T22:18:33Z | 2019-11-04T13:38:04Z | 2019-11-04T13:38:04Z | 2019-11-04T14:55:19Z |
API: drop kwargs from Series.dropna, add explicit `how` parameter | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 101c5ec9137fc..8a481f194d408 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -192,6 +192,8 @@ Other API changes
Now, pandas custom formatters will only be applied to plots created by pandas, through :meth:`~DataFrame.plot`.
Previously, pandas' formatters would be applied to all plots created *after* a :meth:`~DataFrame.plot`.
See :ref:`units registration <whatsnew_1000.matplotlib_units>` for more.
+- :meth:`Series.dropna` has dropped its ``**kwargs`` argument in favor of a single ``how`` parameter.
+ Supplying anything else than ``how`` to ``**kwargs`` raised a ``TypeError`` previously (:issue:`29388`)
-
diff --git a/pandas/core/series.py b/pandas/core/series.py
index e57de0e69b366..7b65816dc06b9 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -4595,7 +4595,7 @@ def notna(self):
def notnull(self):
return super().notnull()
- def dropna(self, axis=0, inplace=False, **kwargs):
+ def dropna(self, axis=0, inplace=False, how=None):
"""
Return a new Series with missing values removed.
@@ -4608,8 +4608,8 @@ def dropna(self, axis=0, inplace=False, **kwargs):
There is only one axis to drop values from.
inplace : bool, default False
If True, do operation inplace and return None.
- **kwargs
- Not in use.
+ how : str, optional
+ Not in use. Kept for compatibility.
Returns
-------
@@ -4667,12 +4667,6 @@ def dropna(self, axis=0, inplace=False, **kwargs):
dtype: object
"""
inplace = validate_bool_kwarg(inplace, "inplace")
- kwargs.pop("how", None)
- if kwargs:
- raise TypeError(
- "dropna() got an unexpected keyword "
- 'argument "{0}"'.format(list(kwargs.keys())[0])
- )
# Validate the axis parameter
self._get_axis_number(axis or 0)
| Using ``**kwargs`` gave false type hints on what the dropna method could take for arguments, and supplying anything but ``how`` raised a TypeError already.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29388 | 2019-11-03T21:54:23Z | 2019-11-04T19:58:48Z | 2019-11-04T19:58:48Z | 2019-11-04T19:58:52Z |
DOC: Added the flag "--no-use-pep517" to contrib guide. | diff --git a/README.md b/README.md
index c299241722b7e..158d48898a7bd 100644
--- a/README.md
+++ b/README.md
@@ -190,7 +190,7 @@ or for installing in [development mode](https://pip.pypa.io/en/latest/reference/
```sh
-python -m pip install --no-build-isolation -e .
+python -m pip install -e . --no-build-isolation --no-use-pep517
```
If you have `make`, you can also use `make develop` to run the same command.
diff --git a/doc/source/development/contributing.rst b/doc/source/development/contributing.rst
index 56fac1cb6852a..eed4a7862cc5f 100644
--- a/doc/source/development/contributing.rst
+++ b/doc/source/development/contributing.rst
@@ -208,7 +208,7 @@ We'll now kick off a three-step process:
# Build and install pandas
python setup.py build_ext --inplace -j 4
- python -m pip install -e . --no-build-isolation
+ python -m pip install -e . --no-build-isolation --no-use-pep517
At this point you should be able to import pandas from your locally built version::
@@ -255,7 +255,7 @@ You'll need to have at least python3.5 installed on your system.
# Build and install pandas
python setup.py build_ext --inplace -j 0
- python -m pip install -e . --no-build-isolation
+ python -m pip install -e . --no-build-isolation --no-use-pep517
**Windows**
| - [x] closes #28633
| https://api.github.com/repos/pandas-dev/pandas/pulls/29387 | 2019-11-03T21:19:14Z | 2019-11-04T13:42:42Z | 2019-11-04T13:42:42Z | 2019-11-04T13:42:42Z |
REF: simplify core.algorithms, reshape.cut | diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 06ba2a7e0ccfb..ceec0652b7ce2 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -8,7 +8,7 @@
import numpy as np
-from pandas._libs import algos, hashtable as htable, lib
+from pandas._libs import Timestamp, algos, hashtable as htable, lib
from pandas._libs.tslib import iNaT
from pandas.util._decorators import Appender, Substitution, deprecate_kwarg
@@ -1440,7 +1440,9 @@ def _take_nd_object(arr, indexer, out, axis: int, fill_value, mask_info):
}
-def _get_take_nd_function(ndim, arr_dtype, out_dtype, axis: int = 0, mask_info=None):
+def _get_take_nd_function(
+ ndim: int, arr_dtype, out_dtype, axis: int = 0, mask_info=None
+):
if ndim <= 2:
tup = (arr_dtype.name, out_dtype.name)
if ndim == 1:
@@ -1474,7 +1476,7 @@ def func2(arr, indexer, out, fill_value=np.nan):
return func2
-def take(arr, indices, axis=0, allow_fill: bool = False, fill_value=None):
+def take(arr, indices, axis: int = 0, allow_fill: bool = False, fill_value=None):
"""
Take elements from an array.
@@ -1568,13 +1570,7 @@ def take(arr, indices, axis=0, allow_fill: bool = False, fill_value=None):
def take_nd(
- arr,
- indexer,
- axis=0,
- out=None,
- fill_value=np.nan,
- mask_info=None,
- allow_fill: bool = True,
+ arr, indexer, axis: int = 0, out=None, fill_value=np.nan, allow_fill: bool = True
):
"""
Specialized Cython take which sets NaN values in one pass
@@ -1597,10 +1593,6 @@ def take_nd(
maybe_promote to determine this type for any fill_value
fill_value : any, default np.nan
Fill value to replace -1 values with
- mask_info : tuple of (ndarray, boolean)
- If provided, value should correspond to:
- (indexer != -1, (indexer != -1).any())
- If not provided, it will be computed internally if necessary
allow_fill : boolean, default True
If False, indexer is assumed to contain no -1 values so no filling
will be done. This short-circuits computation of a mask. Result is
@@ -1611,6 +1603,7 @@ def take_nd(
subarray : array-like
May be the same type as the input, or cast to an ndarray.
"""
+ mask_info = None
if is_extension_array_dtype(arr):
return arr.take(indexer, fill_value=fill_value, allow_fill=allow_fill)
@@ -1632,12 +1625,9 @@ def take_nd(
dtype, fill_value = maybe_promote(arr.dtype, fill_value)
if dtype != arr.dtype and (out is None or out.dtype != dtype):
# check if promotion is actually required based on indexer
- if mask_info is not None:
- mask, needs_masking = mask_info
- else:
- mask = indexer == -1
- needs_masking = mask.any()
- mask_info = mask, needs_masking
+ mask = indexer == -1
+ needs_masking = mask.any()
+ mask_info = mask, needs_masking
if needs_masking:
if out is not None and out.dtype != dtype:
raise TypeError("Incompatible type for fill_value")
@@ -1818,12 +1808,12 @@ def searchsorted(arr, value, side="left", sorter=None):
elif not (
is_object_dtype(arr) or is_numeric_dtype(arr) or is_categorical_dtype(arr)
):
- from pandas.core.series import Series
-
# E.g. if `arr` is an array with dtype='datetime64[ns]'
# and `value` is a pd.Timestamp, we may need to convert value
- value_ser = Series(value)._values
+ value_ser = array([value]) if is_scalar(value) else array(value)
value = value_ser[0] if is_scalar(value) else value_ser
+ if isinstance(value, Timestamp) and value.tzinfo is None:
+ value = value.to_datetime64()
result = arr.searchsorted(value, side=side, sorter=sorter)
return result
diff --git a/pandas/core/reshape/tile.py b/pandas/core/reshape/tile.py
index 09db840ca4db0..95534755b8beb 100644
--- a/pandas/core/reshape/tile.py
+++ b/pandas/core/reshape/tile.py
@@ -4,6 +4,7 @@
import numpy as np
from pandas._libs import Timedelta, Timestamp
+from pandas._libs.interval import Interval
from pandas._libs.lib import infer_dtype
from pandas.core.dtypes.common import (
@@ -18,17 +19,10 @@
is_scalar,
is_timedelta64_dtype,
)
+from pandas.core.dtypes.generic import ABCSeries
from pandas.core.dtypes.missing import isna
-from pandas import (
- Categorical,
- Index,
- Interval,
- IntervalIndex,
- Series,
- to_datetime,
- to_timedelta,
-)
+from pandas import Categorical, Index, IntervalIndex, to_datetime, to_timedelta
import pandas.core.algorithms as algos
import pandas.core.nanops as nanops
@@ -206,7 +200,8 @@ def cut(
# NOTE: this binning code is changed a bit from histogram for var(x) == 0
# for handling the cut for datetime and timedelta objects
- x_is_series, series_index, name, x = _preprocess_for_cut(x)
+ original = x
+ x = _preprocess_for_cut(x)
x, dtype = _coerce_to_type(x)
if not np.iterable(bins):
@@ -268,9 +263,7 @@ def cut(
duplicates=duplicates,
)
- return _postprocess_for_cut(
- fac, bins, retbins, x_is_series, series_index, name, dtype
- )
+ return _postprocess_for_cut(fac, bins, retbins, dtype, original)
def qcut(
@@ -333,8 +326,8 @@ def qcut(
>>> pd.qcut(range(5), 4, labels=False)
array([0, 0, 1, 2, 3])
"""
- x_is_series, series_index, name, x = _preprocess_for_cut(x)
-
+ original = x
+ x = _preprocess_for_cut(x)
x, dtype = _coerce_to_type(x)
if is_integer(q):
@@ -352,9 +345,7 @@ def qcut(
duplicates=duplicates,
)
- return _postprocess_for_cut(
- fac, bins, retbins, x_is_series, series_index, name, dtype
- )
+ return _postprocess_for_cut(fac, bins, retbins, dtype, original)
def _bins_to_cuts(
@@ -544,13 +535,6 @@ def _preprocess_for_cut(x):
input to array, strip the index information and store it
separately
"""
- x_is_series = isinstance(x, Series)
- series_index = None
- name = None
-
- if x_is_series:
- series_index = x.index
- name = x.name
# Check that the passed array is a Pandas or Numpy object
# We don't want to strip away a Pandas data-type here (e.g. datetimetz)
@@ -560,19 +544,17 @@ def _preprocess_for_cut(x):
if x.ndim != 1:
raise ValueError("Input array must be 1 dimensional")
- return x_is_series, series_index, name, x
+ return x
-def _postprocess_for_cut(
- fac, bins, retbins: bool, x_is_series, series_index, name, dtype
-):
+def _postprocess_for_cut(fac, bins, retbins: bool, dtype, original):
"""
handles post processing for the cut method where
we combine the index information if the originally passed
datatype was a series
"""
- if x_is_series:
- fac = Series(fac, index=series_index, name=name)
+ if isinstance(original, ABCSeries):
+ fac = original._constructor(fac, index=original.index, name=original.name)
if not retbins:
return fac
| https://api.github.com/repos/pandas-dev/pandas/pulls/29385 | 2019-11-03T19:34:03Z | 2019-11-04T13:40:47Z | 2019-11-04T13:40:47Z | 2019-11-04T14:59:54Z | |
REF: move safe_sort to algos to avoid private/circular dependencies | diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 06ba2a7e0ccfb..fc55bfbae0900 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -14,6 +14,7 @@
from pandas.core.dtypes.cast import (
construct_1d_object_array_from_listlike,
+ infer_dtype_from_array,
maybe_promote,
)
from pandas.core.dtypes.common import (
@@ -639,8 +640,6 @@ def factorize(values, sort: bool = False, order=None, na_sentinel=-1, size_hint=
)
if sort and len(uniques) > 0:
- from pandas.core.sorting import safe_sort
-
uniques, labels = safe_sort(
uniques, labels, na_sentinel=na_sentinel, assume_unique=True, verify=False
)
@@ -1920,3 +1919,138 @@ def diff(arr, n: int, axis: int = 0):
out_arr = out_arr.astype("int64").view("timedelta64[ns]")
return out_arr
+
+
+# --------------------------------------------------------------------
+# Helper functions
+
+# Note: safe_sort is in algorithms.py instead of sorting.py because it is
+# low-dependency, is used in this module, and used private methods from
+# this module.
+def safe_sort(
+ values,
+ labels=None,
+ na_sentinel: int = -1,
+ assume_unique: bool = False,
+ verify: bool = True,
+):
+ """
+ Sort ``values`` and reorder corresponding ``labels``.
+ ``values`` should be unique if ``labels`` is not None.
+ Safe for use with mixed types (int, str), orders ints before strs.
+
+ Parameters
+ ----------
+ values : list-like
+ Sequence; must be unique if ``labels`` is not None.
+ labels : list_like
+ Indices to ``values``. All out of bound indices are treated as
+ "not found" and will be masked with ``na_sentinel``.
+ na_sentinel : int, default -1
+ Value in ``labels`` to mark "not found".
+ Ignored when ``labels`` is None.
+ assume_unique : bool, default False
+ When True, ``values`` are assumed to be unique, which can speed up
+ the calculation. Ignored when ``labels`` is None.
+ verify : bool, default True
+ Check if labels are out of bound for the values and put out of bound
+ labels equal to na_sentinel. If ``verify=False``, it is assumed there
+ are no out of bound labels. Ignored when ``labels`` is None.
+
+ .. versionadded:: 0.25.0
+
+ Returns
+ -------
+ ordered : ndarray
+ Sorted ``values``
+ new_labels : ndarray
+ Reordered ``labels``; returned when ``labels`` is not None.
+
+ Raises
+ ------
+ TypeError
+ * If ``values`` is not list-like or if ``labels`` is neither None
+ nor list-like
+ * If ``values`` cannot be sorted
+ ValueError
+ * If ``labels`` is not None and ``values`` contain duplicates.
+ """
+ if not is_list_like(values):
+ raise TypeError(
+ "Only list-like objects are allowed to be passed to safe_sort as values"
+ )
+
+ if not isinstance(values, np.ndarray) and not is_extension_array_dtype(values):
+ # don't convert to string types
+ dtype, _ = infer_dtype_from_array(values)
+ values = np.asarray(values, dtype=dtype)
+
+ def sort_mixed(values):
+ # order ints before strings, safe in py3
+ str_pos = np.array([isinstance(x, str) for x in values], dtype=bool)
+ nums = np.sort(values[~str_pos])
+ strs = np.sort(values[str_pos])
+ return np.concatenate([nums, np.asarray(strs, dtype=object)])
+
+ sorter = None
+ if (
+ not is_extension_array_dtype(values)
+ and lib.infer_dtype(values, skipna=False) == "mixed-integer"
+ ):
+ # unorderable in py3 if mixed str/int
+ ordered = sort_mixed(values)
+ else:
+ try:
+ sorter = values.argsort()
+ ordered = values.take(sorter)
+ except TypeError:
+ # try this anyway
+ ordered = sort_mixed(values)
+
+ # labels:
+
+ if labels is None:
+ return ordered
+
+ if not is_list_like(labels):
+ raise TypeError(
+ "Only list-like objects or None are allowed to be"
+ "passed to safe_sort as labels"
+ )
+ labels = ensure_platform_int(np.asarray(labels))
+
+ from pandas import Index
+
+ if not assume_unique and not Index(values).is_unique:
+ raise ValueError("values should be unique if labels is not None")
+
+ if sorter is None:
+ # mixed types
+ hash_klass, values = _get_data_algo(values)
+ t = hash_klass(len(values))
+ t.map_locations(values)
+ sorter = ensure_platform_int(t.lookup(ordered))
+
+ if na_sentinel == -1:
+ # take_1d is faster, but only works for na_sentinels of -1
+ order2 = sorter.argsort()
+ new_labels = take_1d(order2, labels, fill_value=-1)
+ if verify:
+ mask = (labels < -len(values)) | (labels >= len(values))
+ else:
+ mask = None
+ else:
+ reverse_indexer = np.empty(len(sorter), dtype=np.int_)
+ reverse_indexer.put(sorter, np.arange(len(sorter)))
+ # Out of bound indices will be masked with `na_sentinel` next, so we
+ # may deal with them here without performance loss using `mode='wrap'`
+ new_labels = reverse_indexer.take(labels, mode="wrap")
+
+ mask = labels == na_sentinel
+ if verify:
+ mask = mask | (labels < -len(values)) | (labels >= len(values))
+
+ if mask is not None:
+ np.putmask(new_labels, mask, na_sentinel)
+
+ return ordered, ensure_platform_int(new_labels)
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 187c7e2f3a7f7..4c5b7442337fb 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -73,7 +73,6 @@
import pandas.core.missing as missing
from pandas.core.ops import get_op_result_name
from pandas.core.ops.invalid import make_invalid_op
-import pandas.core.sorting as sorting
from pandas.core.strings import StringMethods
from pandas.io.formats.printing import (
@@ -2507,7 +2506,7 @@ def _union(self, other, sort):
if sort is None:
try:
- result = sorting.safe_sort(result)
+ result = algos.safe_sort(result)
except TypeError as e:
warnings.warn(
"{}, sort order is undefined for "
@@ -2603,7 +2602,7 @@ def intersection(self, other, sort=False):
taken = other.take(indexer)
if sort is None:
- taken = sorting.safe_sort(taken.values)
+ taken = algos.safe_sort(taken.values)
if self.name != other.name:
name = None
else:
@@ -2673,7 +2672,7 @@ def difference(self, other, sort=None):
the_diff = this.values.take(label_diff)
if sort is None:
try:
- the_diff = sorting.safe_sort(the_diff)
+ the_diff = algos.safe_sort(the_diff)
except TypeError:
pass
@@ -2750,7 +2749,7 @@ def symmetric_difference(self, other, result_name=None, sort=None):
the_diff = concat_compat([left_diff, right_diff])
if sort is None:
try:
- the_diff = sorting.safe_sort(the_diff)
+ the_diff = algos.safe_sort(the_diff)
except TypeError:
pass
diff --git a/pandas/core/reshape/merge.py b/pandas/core/reshape/merge.py
index ea334503a4302..9845c570ca704 100644
--- a/pandas/core/reshape/merge.py
+++ b/pandas/core/reshape/merge.py
@@ -44,7 +44,6 @@
import pandas.core.common as com
from pandas.core.frame import _merge_doc
from pandas.core.internals import _transform_index, concatenate_block_managers
-import pandas.core.sorting as sorting
from pandas.core.sorting import is_int64_overflow_possible
@@ -1912,7 +1911,7 @@ def _sort_labels(uniques, left, right):
llength = len(left)
labels = np.concatenate([left, right])
- _, new_labels = sorting.safe_sort(uniques, labels, na_sentinel=-1)
+ _, new_labels = algos.safe_sort(uniques, labels, na_sentinel=-1)
new_labels = ensure_int64(new_labels)
new_left, new_right = new_labels[:llength], new_labels[llength:]
diff --git a/pandas/core/sorting.py b/pandas/core/sorting.py
index 9b8a1a76e419c..82eb93dd4c879 100644
--- a/pandas/core/sorting.py
+++ b/pandas/core/sorting.py
@@ -4,13 +4,11 @@
from pandas._libs import algos, hashtable, lib
from pandas._libs.hashtable import unique_label_indices
-from pandas.core.dtypes.cast import infer_dtype_from_array
from pandas.core.dtypes.common import (
ensure_int64,
ensure_platform_int,
is_categorical_dtype,
is_extension_array_dtype,
- is_list_like,
)
from pandas.core.dtypes.missing import isna
@@ -389,132 +387,3 @@ def _reorder_by_uniques(uniques, labels):
uniques = algorithms.take_nd(uniques, sorter, allow_fill=False)
return uniques, labels
-
-
-def safe_sort(
- values,
- labels=None,
- na_sentinel: int = -1,
- assume_unique: bool = False,
- verify: bool = True,
-):
- """
- Sort ``values`` and reorder corresponding ``labels``.
- ``values`` should be unique if ``labels`` is not None.
- Safe for use with mixed types (int, str), orders ints before strs.
-
- Parameters
- ----------
- values : list-like
- Sequence; must be unique if ``labels`` is not None.
- labels : list_like
- Indices to ``values``. All out of bound indices are treated as
- "not found" and will be masked with ``na_sentinel``.
- na_sentinel : int, default -1
- Value in ``labels`` to mark "not found".
- Ignored when ``labels`` is None.
- assume_unique : bool, default False
- When True, ``values`` are assumed to be unique, which can speed up
- the calculation. Ignored when ``labels`` is None.
- verify : bool, default True
- Check if labels are out of bound for the values and put out of bound
- labels equal to na_sentinel. If ``verify=False``, it is assumed there
- are no out of bound labels. Ignored when ``labels`` is None.
-
- .. versionadded:: 0.25.0
-
- Returns
- -------
- ordered : ndarray
- Sorted ``values``
- new_labels : ndarray
- Reordered ``labels``; returned when ``labels`` is not None.
-
- Raises
- ------
- TypeError
- * If ``values`` is not list-like or if ``labels`` is neither None
- nor list-like
- * If ``values`` cannot be sorted
- ValueError
- * If ``labels`` is not None and ``values`` contain duplicates.
- """
- if not is_list_like(values):
- raise TypeError(
- "Only list-like objects are allowed to be passed to safe_sort as values"
- )
-
- if not isinstance(values, np.ndarray) and not is_extension_array_dtype(values):
- # don't convert to string types
- dtype, _ = infer_dtype_from_array(values)
- values = np.asarray(values, dtype=dtype)
-
- def sort_mixed(values):
- # order ints before strings, safe in py3
- str_pos = np.array([isinstance(x, str) for x in values], dtype=bool)
- nums = np.sort(values[~str_pos])
- strs = np.sort(values[str_pos])
- return np.concatenate([nums, np.asarray(strs, dtype=object)])
-
- sorter = None
- if (
- not is_extension_array_dtype(values)
- and lib.infer_dtype(values, skipna=False) == "mixed-integer"
- ):
- # unorderable in py3 if mixed str/int
- ordered = sort_mixed(values)
- else:
- try:
- sorter = values.argsort()
- ordered = values.take(sorter)
- except TypeError:
- # try this anyway
- ordered = sort_mixed(values)
-
- # labels:
-
- if labels is None:
- return ordered
-
- if not is_list_like(labels):
- raise TypeError(
- "Only list-like objects or None are allowed to be"
- "passed to safe_sort as labels"
- )
- labels = ensure_platform_int(np.asarray(labels))
-
- from pandas import Index
-
- if not assume_unique and not Index(values).is_unique:
- raise ValueError("values should be unique if labels is not None")
-
- if sorter is None:
- # mixed types
- hash_klass, values = algorithms._get_data_algo(values)
- t = hash_klass(len(values))
- t.map_locations(values)
- sorter = ensure_platform_int(t.lookup(ordered))
-
- if na_sentinel == -1:
- # take_1d is faster, but only works for na_sentinels of -1
- order2 = sorter.argsort()
- new_labels = algorithms.take_1d(order2, labels, fill_value=-1)
- if verify:
- mask = (labels < -len(values)) | (labels >= len(values))
- else:
- mask = None
- else:
- reverse_indexer = np.empty(len(sorter), dtype=np.int_)
- reverse_indexer.put(sorter, np.arange(len(sorter)))
- # Out of bound indices will be masked with `na_sentinel` next, so we
- # may deal with them here without performance loss using `mode='wrap'`
- new_labels = reverse_indexer.take(labels, mode="wrap")
-
- mask = labels == na_sentinel
- if verify:
- mask = mask | (labels < -len(values)) | (labels >= len(values))
-
- if mask is not None:
- np.putmask(new_labels, mask, na_sentinel)
-
- return ordered, ensure_platform_int(new_labels)
diff --git a/pandas/tests/indexes/test_base.py b/pandas/tests/indexes/test_base.py
index 8d0cb0edf51df..e43d340a46d9f 100644
--- a/pandas/tests/indexes/test_base.py
+++ b/pandas/tests/indexes/test_base.py
@@ -33,13 +33,13 @@
isna,
period_range,
)
+from pandas.core.algorithms import safe_sort
from pandas.core.index import (
_get_combined_index,
ensure_index,
ensure_index_from_sequences,
)
from pandas.core.indexes.api import Index, MultiIndex
-from pandas.core.sorting import safe_sort
from pandas.tests.indexes.common import Base
from pandas.tests.indexes.conftest import indices_dict
import pandas.util.testing as tm
diff --git a/pandas/tests/test_algos.py b/pandas/tests/test_algos.py
index 9dd88fd5dd25b..a64501040442d 100644
--- a/pandas/tests/test_algos.py
+++ b/pandas/tests/test_algos.py
@@ -26,7 +26,6 @@
import pandas.core.algorithms as algos
from pandas.core.arrays import DatetimeArray
import pandas.core.common as com
-from pandas.core.sorting import safe_sort
import pandas.util.testing as tm
@@ -309,7 +308,7 @@ def test_factorize_na_sentinel(self, sort, na_sentinel, data, uniques):
labels, uniques = algos.factorize(data, sort=sort, na_sentinel=na_sentinel)
if sort:
expected_labels = np.array([1, 0, na_sentinel, 1], dtype=np.intp)
- expected_uniques = safe_sort(uniques)
+ expected_uniques = algos.safe_sort(uniques)
else:
expected_labels = np.array([0, 1, na_sentinel, 0], dtype=np.intp)
expected_uniques = uniques
diff --git a/pandas/tests/test_sorting.py b/pandas/tests/test_sorting.py
index b86aaa0ed7e1f..5d7eb70817a11 100644
--- a/pandas/tests/test_sorting.py
+++ b/pandas/tests/test_sorting.py
@@ -6,6 +6,7 @@
import pytest
from pandas import DataFrame, MultiIndex, Series, array, concat, merge
+from pandas.core.algorithms import safe_sort
import pandas.core.common as com
from pandas.core.sorting import (
decons_group_index,
@@ -13,7 +14,6 @@
is_int64_overflow_possible,
lexsort_indexer,
nargsort,
- safe_sort,
)
import pandas.util.testing as tm
diff --git a/pandas/tests/window/test_pairwise.py b/pandas/tests/window/test_pairwise.py
index 56d89e15c418c..6f6d4c09526ff 100644
--- a/pandas/tests/window/test_pairwise.py
+++ b/pandas/tests/window/test_pairwise.py
@@ -3,7 +3,7 @@
import pytest
from pandas import DataFrame, Series
-from pandas.core.sorting import safe_sort
+from pandas.core.algorithms import safe_sort
import pandas.util.testing as tm
| safe_sort uses private functions from core.algorithms and is runtime-imported into core.algorithms. It also doesn't use _anything_ else defined in core.sorting. This move cleans up the dependency structure, in particular is a step towards getting #29133 working. | https://api.github.com/repos/pandas-dev/pandas/pulls/29384 | 2019-11-03T17:22:08Z | 2019-11-04T16:21:26Z | 2019-11-04T16:21:25Z | 2019-11-04T16:46:38Z |
CLN: Remove unused _isfinite, make_axis_dummies | diff --git a/pandas/core/nanops.py b/pandas/core/nanops.py
index 070891c4acb5e..7e50348962fc5 100644
--- a/pandas/core/nanops.py
+++ b/pandas/core/nanops.py
@@ -16,7 +16,6 @@
is_any_int_dtype,
is_bool_dtype,
is_complex,
- is_complex_dtype,
is_datetime64_dtype,
is_datetime64tz_dtype,
is_datetime_or_timedelta_dtype,
@@ -325,19 +324,6 @@ def _get_values(
return values, mask, dtype, dtype_max, fill_value
-def _isfinite(values):
- if is_datetime_or_timedelta_dtype(values):
- return isna(values)
- if (
- is_complex_dtype(values)
- or is_float_dtype(values)
- or is_integer_dtype(values)
- or is_bool_dtype(values)
- ):
- return ~np.isfinite(values)
- return ~np.isfinite(values.astype("float64"))
-
-
def _na_ok_dtype(dtype):
# TODO: what about datetime64tz? PeriodDtype?
return not issubclass(dtype.type, (np.integer, np.timedelta64, np.datetime64))
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index ad7081fb17703..949d8f1bfb09c 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -1046,43 +1046,7 @@ def _make_col_name(prefix, prefix_sep, level):
return DataFrame(dummy_mat, index=index, columns=dummy_cols)
-def make_axis_dummies(frame, axis="minor", transform=None):
- """
- Construct 1-0 dummy variables corresponding to designated axis
- labels
-
- Parameters
- ----------
- frame : DataFrame
- axis : {'major', 'minor'}, default 'minor'
- transform : function, default None
- Function to apply to axis labels first. For example, to
- get "day of week" dummies in a time series regression
- you might call::
-
- make_axis_dummies(panel, axis='major',
- transform=lambda d: d.weekday())
- Returns
- -------
- dummies : DataFrame
- Column names taken from chosen axis
- """
- numbers = {"major": 0, "minor": 1}
- num = numbers.get(axis, axis)
-
- items = frame.index.levels[num]
- codes = frame.index.codes[num]
- if transform is not None:
- mapped_items = items.map(transform)
- codes, items = _factorize_from_iterable(mapped_items.take(codes))
-
- values = np.eye(len(items), dtype=float)
- values = values.take(codes, axis=0)
-
- return DataFrame(values, columns=items, index=frame.index)
-
-
-def _reorder_for_extension_array_stack(arr, n_rows, n_columns):
+def _reorder_for_extension_array_stack(arr, n_rows: int, n_columns: int):
"""
Re-orders the values when stacking multiple extension-arrays.
diff --git a/pandas/tests/reshape/test_reshape.py b/pandas/tests/reshape/test_reshape.py
index 2e94eeba1d05b..b695b05c7c7db 100644
--- a/pandas/tests/reshape/test_reshape.py
+++ b/pandas/tests/reshape/test_reshape.py
@@ -645,24 +645,3 @@ def test_reshaping_multi_index_categorical(self):
index=dti.rename("major"),
)
tm.assert_frame_equal(result, expected)
-
-
-class TestMakeAxisDummies:
- def test_preserve_categorical_dtype(self):
- # GH13854
- for ordered in [False, True]:
- cidx = pd.CategoricalIndex(list("xyz"), ordered=ordered)
- midx = pd.MultiIndex(levels=[["a"], cidx], codes=[[0, 0], [0, 1]])
- df = DataFrame([[10, 11]], index=midx)
-
- expected = DataFrame(
- [[1.0, 0.0, 0.0], [0.0, 1.0, 0.0]], index=midx, columns=cidx
- )
-
- from pandas.core.reshape.reshape import make_axis_dummies
-
- result = make_axis_dummies(df)
- tm.assert_frame_equal(result, expected)
-
- result = make_axis_dummies(df, transform=lambda x: x)
- tm.assert_frame_equal(result, expected)
diff --git a/pandas/tests/test_nanops.py b/pandas/tests/test_nanops.py
index 49d1777df0751..52ad56967220f 100644
--- a/pandas/tests/test_nanops.py
+++ b/pandas/tests/test_nanops.py
@@ -704,46 +704,6 @@ def test__has_infs(self):
self.check_bool(nanops._has_infs, val.astype("f4"), correct)
self.check_bool(nanops._has_infs, val.astype("f2"), correct)
- def test__isfinite(self):
- pairs = [
- ("arr_complex", False),
- ("arr_int", False),
- ("arr_bool", False),
- ("arr_str", False),
- ("arr_utf", False),
- ("arr_complex", False),
- ("arr_complex_nan", True),
- ("arr_nan_nanj", True),
- ("arr_nan_infj", True),
- ("arr_complex_nan_infj", True),
- ]
- pairs_float = [
- ("arr_float", False),
- ("arr_nan", True),
- ("arr_float_nan", True),
- ("arr_nan_nan", True),
- ("arr_float_inf", True),
- ("arr_inf", True),
- ("arr_nan_inf", True),
- ("arr_float_nan_inf", True),
- ("arr_nan_nan_inf", True),
- ]
-
- func1 = lambda x: np.any(nanops._isfinite(x).ravel())
-
- # TODO: unused?
- # func2 = lambda x: np.any(nanops._isfinite(x).values.ravel())
-
- for arr, correct in pairs:
- val = getattr(self, arr)
- self.check_bool(func1, val, correct)
-
- for arr, correct in pairs_float:
- val = getattr(self, arr)
- self.check_bool(func1, val, correct)
- self.check_bool(func1, val.astype("f4"), correct)
- self.check_bool(func1, val.astype("f2"), correct)
-
def test__bn_ok_dtype(self):
assert nanops._bn_ok_dtype(self.arr_float.dtype, "test")
assert nanops._bn_ok_dtype(self.arr_complex.dtype, "test")
| https://api.github.com/repos/pandas-dev/pandas/pulls/29380 | 2019-11-03T04:35:44Z | 2019-11-03T14:54:47Z | 2019-11-03T14:54:47Z | 2019-11-03T15:34:09Z | |
Fixes typo | diff --git a/web/pandas/getting_started.md b/web/pandas/getting_started.md
index 9682cf90cad6f..4195cc00b2419 100644
--- a/web/pandas/getting_started.md
+++ b/web/pandas/getting_started.md
@@ -9,7 +9,7 @@ the [advanced installation page]({{ base_url}}/docs/getting_started/install.html
1. Download [Anaconda](https://www.anaconda.com/distribution/) for your operating system and
the latest Python version, run the installer, and follow the steps. Detailed instructions
on how to install Anaconda can be found in the
- [Anaconda documentation](https://docs.anaconda.com/anaconda/install/)).
+ [Anaconda documentation](https://docs.anaconda.com/anaconda/install/).
2. In the Anaconda prompt (or terminal in Linux or MacOS), start JupyterLab:
| https://api.github.com/repos/pandas-dev/pandas/pulls/29379 | 2019-11-03T03:33:15Z | 2019-11-03T14:53:47Z | 2019-11-03T14:53:47Z | 2019-11-03T14:53:51Z | |
TST: new test for incorrect series assignment | diff --git a/pandas/tests/series/indexing/test_indexing.py b/pandas/tests/series/indexing/test_indexing.py
index 4673dabca811b..5aba2920999d5 100644
--- a/pandas/tests/series/indexing/test_indexing.py
+++ b/pandas/tests/series/indexing/test_indexing.py
@@ -391,6 +391,22 @@ def test_setslice(datetime_series):
assert sl.index.is_unique is True
+def test_2d_to_1d_assignment_raises():
+ x = np.random.randn(2, 2)
+ y = pd.Series(range(2))
+
+ msg = (
+ r"shape mismatch: value array of shape \(2,2\) could not be"
+ r" broadcast to indexing result of shape \(2,\)"
+ )
+ with pytest.raises(ValueError, match=msg):
+ y.loc[range(2)] = x
+
+ msg = r"could not broadcast input array from shape \(2,2\) into shape \(2\)"
+ with pytest.raises(ValueError, match=msg):
+ y.loc[:] = x
+
+
# FutureWarning from NumPy about [slice(None, 5).
@pytest.mark.filterwarnings("ignore:Using a non-tuple:FutureWarning")
def test_basic_getitem_setitem_corner(datetime_series):
| - [x] closes #14525
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29378 | 2019-11-03T00:41:52Z | 2019-11-07T22:13:28Z | 2019-11-07T22:13:28Z | 2019-11-07T22:13:43Z |
CLN: deprivatize factorize_from_iterable | diff --git a/pandas/core/arrays/categorical.py b/pandas/core/arrays/categorical.py
index 612e54ba426f3..ce174baa66a97 100644
--- a/pandas/core/arrays/categorical.py
+++ b/pandas/core/arrays/categorical.py
@@ -2678,7 +2678,7 @@ def _convert_to_list_like(list_like):
return [list_like]
-def _factorize_from_iterable(values):
+def factorize_from_iterable(values):
"""
Factorize an input `values` into `categories` and `codes`. Preserves
categorical dtype in `categories`.
@@ -2716,9 +2716,9 @@ def _factorize_from_iterable(values):
return codes, categories
-def _factorize_from_iterables(iterables):
+def factorize_from_iterables(iterables):
"""
- A higher-level wrapper over `_factorize_from_iterable`.
+ A higher-level wrapper over `factorize_from_iterable`.
*This is an internal function*
@@ -2733,9 +2733,9 @@ def _factorize_from_iterables(iterables):
Notes
-----
- See `_factorize_from_iterable` for more info.
+ See `factorize_from_iterable` for more info.
"""
if len(iterables) == 0:
# For consistency, it should return a list of 2 lists.
return [[], []]
- return map(list, zip(*(_factorize_from_iterable(it) for it in iterables)))
+ return map(list, zip(*(factorize_from_iterable(it) for it in iterables)))
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index f968a9eb4103c..66deacac37789 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -31,7 +31,7 @@
import pandas.core.algorithms as algos
from pandas.core.arrays import Categorical
-from pandas.core.arrays.categorical import _factorize_from_iterables
+from pandas.core.arrays.categorical import factorize_from_iterables
import pandas.core.common as com
import pandas.core.indexes.base as ibase
from pandas.core.indexes.base import (
@@ -440,7 +440,7 @@ def from_arrays(cls, arrays, sortorder=None, names=_no_default_names):
if len(arrays[i]) != len(arrays[i - 1]):
raise ValueError("all arrays must be same length")
- codes, levels = _factorize_from_iterables(arrays)
+ codes, levels = factorize_from_iterables(arrays)
if names is _no_default_names:
names = [getattr(arr, "name", None) for arr in arrays]
@@ -562,7 +562,7 @@ def from_product(cls, iterables, sortorder=None, names=_no_default_names):
elif is_iterator(iterables):
iterables = list(iterables)
- codes, levels = _factorize_from_iterables(iterables)
+ codes, levels = factorize_from_iterables(iterables)
if names is _no_default_names:
names = [getattr(it, "name", None) for it in iterables]
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index bbf41fc28e9d2..c11915c00c59d 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -8,8 +8,8 @@
from pandas import DataFrame, Index, MultiIndex, Series
from pandas.core.arrays.categorical import (
- _factorize_from_iterable,
- _factorize_from_iterables,
+ factorize_from_iterable,
+ factorize_from_iterables,
)
import pandas.core.common as com
from pandas.core.generic import NDFrame
@@ -604,7 +604,7 @@ def _make_concat_multiindex(indexes, keys, levels=None, names=None):
names = [None] * len(zipped)
if levels is None:
- _, levels = _factorize_from_iterables(zipped)
+ _, levels = factorize_from_iterables(zipped)
else:
levels = [ensure_index(x) for x in levels]
else:
@@ -645,7 +645,7 @@ def _make_concat_multiindex(indexes, keys, levels=None, names=None):
levels.extend(concat_index.levels)
codes_list.extend(concat_index.codes)
else:
- codes, categories = _factorize_from_iterable(concat_index)
+ codes, categories = factorize_from_iterable(concat_index)
levels.append(categories)
codes_list.append(codes)
diff --git a/pandas/core/reshape/reshape.py b/pandas/core/reshape/reshape.py
index 949d8f1bfb09c..d7eae1c543804 100644
--- a/pandas/core/reshape/reshape.py
+++ b/pandas/core/reshape/reshape.py
@@ -22,7 +22,7 @@
import pandas.core.algorithms as algos
from pandas.core.arrays import SparseArray
-from pandas.core.arrays.categorical import _factorize_from_iterable
+from pandas.core.arrays.categorical import factorize_from_iterable
from pandas.core.construction import extract_array
from pandas.core.frame import DataFrame
from pandas.core.index import Index, MultiIndex
@@ -504,7 +504,7 @@ def stack(frame, level=-1, dropna=True):
def factorize(index):
if index.is_unique:
return index, np.arange(len(index))
- codes, categories = _factorize_from_iterable(index)
+ codes, categories = factorize_from_iterable(index)
return categories, codes
N, K = frame.shape
@@ -725,7 +725,7 @@ def _convert_level_number(level_num, columns):
new_names = list(this.index.names)
new_codes = [lab.repeat(levsize) for lab in this.index.codes]
else:
- old_codes, old_levels = _factorize_from_iterable(this.index)
+ old_codes, old_levels = factorize_from_iterable(this.index)
new_levels = [old_levels]
new_codes = [old_codes.repeat(levsize)]
new_names = [this.index.name] # something better?
@@ -949,7 +949,7 @@ def _get_dummies_1d(
from pandas.core.reshape.concat import concat
# Series avoids inconsistent NaN handling
- codes, levels = _factorize_from_iterable(Series(data))
+ codes, levels = factorize_from_iterable(Series(data))
if dtype is None:
dtype = np.uint8
| Deprivatize ``_factorize_from_iterable`` and ``_factorize_from_iterables`` | https://api.github.com/repos/pandas-dev/pandas/pulls/29377 | 2019-11-02T21:26:57Z | 2019-11-03T22:11:43Z | 2019-11-03T22:11:43Z | 2019-11-03T22:11:47Z |
Updated index links for 0.25.3 | diff --git a/doc/source/index.rst.template b/doc/source/index.rst.template
index 09d18d6f96197..cbdbbda7e530d 100644
--- a/doc/source/index.rst.template
+++ b/doc/source/index.rst.template
@@ -39,7 +39,7 @@ See the :ref:`overview` for more detail about what's in the library.
:hidden:
{% endif %}
{% if not single_doc %}
- What's New in 0.25.2 <whatsnew/v0.25.2>
+ What's New in 0.25.3 <whatsnew/v0.25.3>
install
getting_started/index
user_guide/index
@@ -53,7 +53,7 @@ See the :ref:`overview` for more detail about what's in the library.
whatsnew/index
{% endif %}
-* :doc:`whatsnew/v0.25.2`
+* :doc:`whatsnew/v0.25.3`
* :doc:`install`
* :doc:`getting_started/index`
| I think this was supposed to be updated for the release. Might need to retag after repush docs after this
@TomAugspurger | https://api.github.com/repos/pandas-dev/pandas/pulls/29374 | 2019-11-02T19:44:11Z | 2019-11-08T16:27:25Z | 2019-11-08T16:27:25Z | 2019-11-14T16:20:20Z |
Add documentation linking to sqlalchemy | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index f88c26c7bc782..cfbd125b7445e 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -2645,7 +2645,11 @@ def to_sql(
Name of SQL table.
con : sqlalchemy.engine.Engine or sqlite3.Connection
Using SQLAlchemy makes it possible to use any DB supported by that
- library. Legacy support is provided for sqlite3.Connection objects.
+ library. Legacy support is provided for sqlite3.Connection objects. The user
+ is responsible for engine disposal and connection closure for the SQLAlchemy
+ connectable See `here \
+ <https://docs.sqlalchemy.org/en/13/core/connections.html>`_
+
schema : str, optional
Specify the schema (if database flavor supports this). If None, use
default schema.
diff --git a/pandas/io/sql.py b/pandas/io/sql.py
index e90e19649f645..684e602f06d12 100644
--- a/pandas/io/sql.py
+++ b/pandas/io/sql.py
@@ -361,7 +361,9 @@ def read_sql(
or DBAPI2 connection (fallback mode)
Using SQLAlchemy makes it possible to use any DB supported by that
- library. If a DBAPI2 object, only sqlite3 is supported.
+ library. If a DBAPI2 object, only sqlite3 is supported. The user is responsible
+ for engine disposal and connection closure for the SQLAlchemy connectable. See
+ `here <https://docs.sqlalchemy.org/en/13/core/connections.html>`_
index_col : string or list of strings, optional, default: None
Column(s) to set as index(MultiIndex).
coerce_float : boolean, default True
| - [x] closes #23086
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29373 | 2019-11-02T19:16:28Z | 2019-11-22T16:19:28Z | 2019-11-22T16:19:27Z | 2019-11-22T16:19:34Z |
TST: new test for str to small float conversion dtype | diff --git a/pandas/tests/frame/test_dtypes.py b/pandas/tests/frame/test_dtypes.py
index 68844aeeb081e..07accdb47d252 100644
--- a/pandas/tests/frame/test_dtypes.py
+++ b/pandas/tests/frame/test_dtypes.py
@@ -1063,6 +1063,18 @@ def test_asarray_homogenous(self):
expected = np.array([[1, 1], [2, 2]], dtype="object")
tm.assert_numpy_array_equal(result, expected)
+ def test_str_to_small_float_conversion_type(self):
+ # GH 20388
+ np.random.seed(13)
+ col_data = [str(np.random.random() * 1e-12) for _ in range(5)]
+ result = pd.DataFrame(col_data, columns=["A"])
+ expected = pd.DataFrame(col_data, columns=["A"], dtype=object)
+ tm.assert_frame_equal(result, expected)
+ # change the dtype of the elements from object to float one by one
+ result.loc[result.index, "A"] = [float(x) for x in col_data]
+ expected = pd.DataFrame(col_data, columns=["A"], dtype=float)
+ tm.assert_frame_equal(result, expected)
+
class TestDataFrameDatetimeWithTZ:
def test_interleave(self, timezone_frame):
| - [x] closes #20388
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29372 | 2019-11-02T18:40:16Z | 2019-11-03T21:22:19Z | 2019-11-03T21:22:19Z | 2019-11-03T21:22:30Z |
Update setup.py to https | diff --git a/setup.py b/setup.py
index c75ad5896a439..3dd38bdb6adbb 100755
--- a/setup.py
+++ b/setup.py
@@ -165,7 +165,7 @@ def build_extensions(self):
(2-dimensional), handle the vast majority of typical use cases in finance,
statistics, social science, and many areas of engineering. For R users,
DataFrame provides everything that R's ``data.frame`` provides and much
-more. pandas is built on top of `NumPy <http://www.numpy.org>`__ and is
+more. pandas is built on top of `NumPy <https://www.numpy.org>`__ and is
intended to integrate well within a scientific computing environment with many
other 3rd party libraries.
@@ -209,11 +209,11 @@ def build_extensions(self):
LICENSE = "BSD"
AUTHOR = "The PyData Development Team"
EMAIL = "pydata@googlegroups.com"
-URL = "http://pandas.pydata.org"
+URL = "https://pandas.pydata.org"
DOWNLOAD_URL = ""
PROJECT_URLS = {
"Bug Tracker": "https://github.com/pandas-dev/pandas/issues",
- "Documentation": "http://pandas.pydata.org/pandas-docs/stable/",
+ "Documentation": "https://pandas.pydata.org/pandas-docs/stable/",
"Source Code": "https://github.com/pandas-dev/pandas",
}
CLASSIFIERS = [
| Avoids a redirect in browser when clicking these links.
| https://api.github.com/repos/pandas-dev/pandas/pulls/29371 | 2019-11-02T18:08:06Z | 2019-11-02T19:49:01Z | 2019-11-02T19:49:01Z | 2019-11-02T19:49:06Z |
TST: Adding merge test for non-string columns [Ref 17962] | diff --git a/pandas/tests/reshape/merge/test_merge.py b/pandas/tests/reshape/merge/test_merge.py
index 37c0b57bc7581..dd51a1a6c8359 100644
--- a/pandas/tests/reshape/merge/test_merge.py
+++ b/pandas/tests/reshape/merge/test_merge.py
@@ -134,6 +134,18 @@ def test_merge_common(self):
exp = merge(self.df, self.df2, on=["key1", "key2"])
tm.assert_frame_equal(joined, exp)
+ def test_merge_non_string_columns(self):
+ # https://github.com/pandas-dev/pandas/issues/17962
+ # Checks that method runs for non string column names
+ left = pd.DataFrame(
+ {0: [1, 0, 1, 0], 1: [0, 1, 0, 0], 2: [0, 0, 2, 0], 3: [1, 0, 0, 3]}
+ )
+
+ right = left.astype(float)
+ expected = left
+ result = pd.merge(left, right)
+ tm.assert_frame_equal(expected, result)
+
def test_merge_index_as_on_arg(self):
# GH14355
| - [x] closes #17962
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29370 | 2019-11-02T17:55:39Z | 2019-11-03T21:23:01Z | 2019-11-03T21:23:01Z | 2019-11-03T21:23:12Z |
GH:11670: possible bug when calculating mean of DataFrame? | diff --git a/pandas/tests/frame/test_analytics.py b/pandas/tests/frame/test_analytics.py
index e99208ac78e15..f694689fa9dfb 100644
--- a/pandas/tests/frame/test_analytics.py
+++ b/pandas/tests/frame/test_analytics.py
@@ -1,4 +1,5 @@
from datetime import timedelta
+from decimal import Decimal
import operator
from string import ascii_lowercase
import warnings
@@ -1075,6 +1076,29 @@ def test_mean_excludeds_datetimes(self, tz):
expected = pd.Series()
tm.assert_series_equal(result, expected)
+ def test_mean_mixed_string_decimal(self):
+ # GH 11670
+ # possible bug when calculating mean of DataFrame?
+
+ d = [
+ {"A": 2, "B": None, "C": Decimal("628.00")},
+ {"A": 1, "B": None, "C": Decimal("383.00")},
+ {"A": 3, "B": None, "C": Decimal("651.00")},
+ {"A": 2, "B": None, "C": Decimal("575.00")},
+ {"A": 4, "B": None, "C": Decimal("1114.00")},
+ {"A": 1, "B": "TEST", "C": Decimal("241.00")},
+ {"A": 2, "B": None, "C": Decimal("572.00")},
+ {"A": 4, "B": None, "C": Decimal("609.00")},
+ {"A": 3, "B": None, "C": Decimal("820.00")},
+ {"A": 5, "B": None, "C": Decimal("1223.00")},
+ ]
+
+ df = pd.DataFrame(d)
+
+ result = df.mean()
+ expected = pd.Series([2.7, 681.6], index=["A", "C"])
+ tm.assert_series_equal(result, expected)
+
def test_var_std(self, datetime_frame):
result = datetime_frame.std(ddof=4)
expected = datetime_frame.apply(lambda x: x.std(ddof=4))
| - [x] closes #11670
- [x] tests added / passed
- [x] passes `black pandas`
- [x] added test test_mean_mixed_string_float | https://api.github.com/repos/pandas-dev/pandas/pulls/29369 | 2019-11-02T17:39:28Z | 2019-11-03T14:59:06Z | 2019-11-03T14:59:06Z | 2019-11-03T14:59:10Z |
Fix pipe docs | diff --git a/doc/source/getting_started/basics.rst b/doc/source/getting_started/basics.rst
index 9b97aa25a9240..125990f7cadcd 100644
--- a/doc/source/getting_started/basics.rst
+++ b/doc/source/getting_started/basics.rst
@@ -753,28 +753,51 @@ on an entire ``DataFrame`` or ``Series``, row- or column-wise, or elementwise.
Tablewise function application
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-``DataFrames`` and ``Series`` can of course just be passed into functions.
+``DataFrames`` and ``Series`` can be passed into functions.
However, if the function needs to be called in a chain, consider using the :meth:`~DataFrame.pipe` method.
-Compare the following
-.. code-block:: python
+First some setup:
+
+.. ipython:: python
- # f, g, and h are functions taking and returning ``DataFrames``
- >>> f(g(h(df), arg1=1), arg2=2, arg3=3)
+ def extract_city_name(df):
+ """
+ Chicago, IL -> Chicago for city_name column
+ """
+ df['city_name'] = df['city_and_code'].str.split(",").str.get(0)
+ return df
-with the equivalent
+ def add_country_name(df, country_name=None):
+ """
+ Chicago -> Chicago-US for city_name column
+ """
+ col = 'city_name'
+ df['city_and_country'] = df[col] + country_name
+ return df
-.. code-block:: python
+ df_p = pd.DataFrame({'city_and_code': ['Chicago, IL']})
+
+
+``extract_city_name`` and ``add_country_name`` are functions taking and returning ``DataFrames``.
+
+Now compare the following:
+
+.. ipython:: python
+
+ add_country_name(extract_city_name(df_p), country_name='US')
+
+Is equivalent to:
+
+.. ipython:: python
- >>> (df.pipe(h)
- ... .pipe(g, arg1=1)
- ... .pipe(f, arg2=2, arg3=3))
+ (df_p.pipe(extract_city_name)
+ .pipe(add_country_name, country_name="US"))
Pandas encourages the second style, which is known as method chaining.
``pipe`` makes it easy to use your own or another library's functions
in method chains, alongside pandas' methods.
-In the example above, the functions ``f``, ``g``, and ``h`` each expected the ``DataFrame`` as the first positional argument.
+In the example above, the functions ``extract_city_name`` and ``add_country_name`` each expected a ``DataFrame`` as the first positional argument.
What if the function you wish to apply takes its data as, say, the second argument?
In this case, provide ``pipe`` with a tuple of ``(callable, data_keyword)``.
``.pipe`` will route the ``DataFrame`` to the argument specified in the tuple.
diff --git a/setup.cfg b/setup.cfg
index d4657100c1291..2dcb46584f19e 100644
--- a/setup.cfg
+++ b/setup.cfg
@@ -47,7 +47,6 @@ ignore = E402, # module level import not at top of file
E711, # comparison to none should be 'if cond is none:'
exclude =
- doc/source/getting_started/basics.rst
doc/source/development/contributing_docstring.rst
| - [x] closes #27054
See the image below which highlights what i've done here
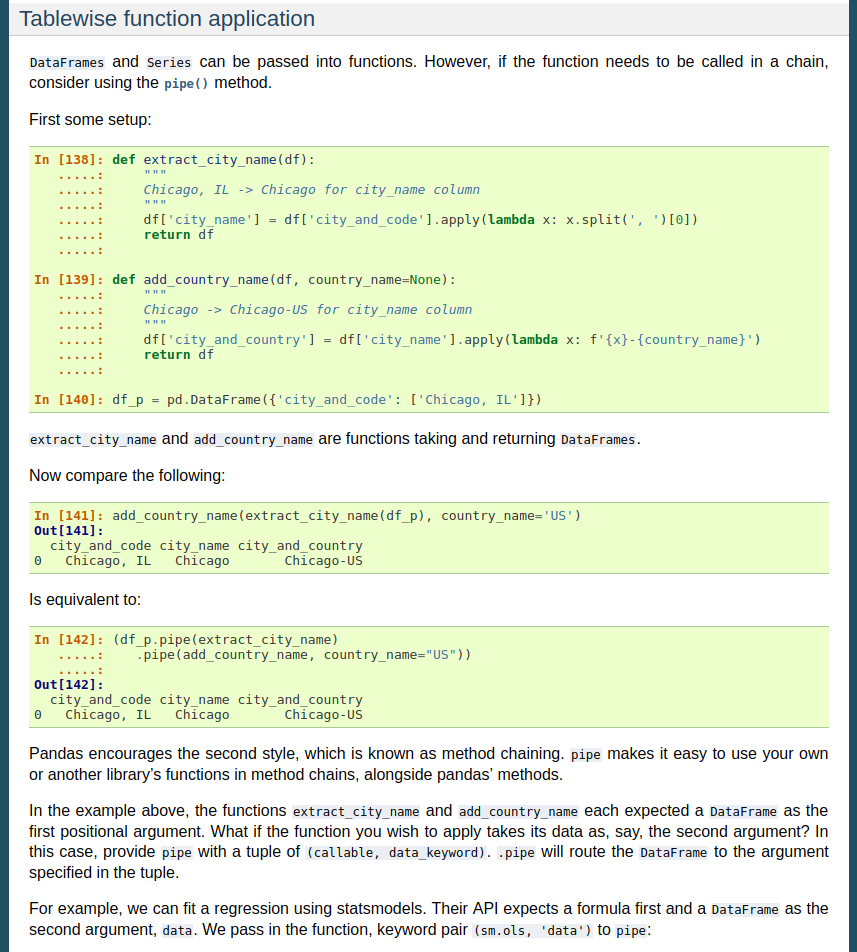
`flake8-rst doc/source --filename=/getting_started/basics.rst` runs clean
cc. @datapythonista to review | https://api.github.com/repos/pandas-dev/pandas/pulls/29368 | 2019-11-02T17:30:12Z | 2019-11-06T17:51:05Z | 2019-11-06T17:51:05Z | 2019-11-07T01:48:17Z |
TST: Adding map test for dict with np.nan key [Ref 17648] | diff --git a/pandas/tests/series/test_apply.py b/pandas/tests/series/test_apply.py
index e56294669a546..971ce5b18c323 100644
--- a/pandas/tests/series/test_apply.py
+++ b/pandas/tests/series/test_apply.py
@@ -581,6 +581,14 @@ def test_map_defaultdict(self):
expected = Series(["stuff", "blank", "blank"], index=["a", "b", "c"])
tm.assert_series_equal(result, expected)
+ def test_map_dict_na_key(self):
+ # https://github.com/pandas-dev/pandas/issues/17648
+ # Checks that np.nan key is appropriately mapped
+ s = Series([1, 2, np.nan])
+ expected = Series(["a", "b", "c"])
+ result = s.map({1: "a", 2: "b", np.nan: "c"})
+ tm.assert_series_equal(result, expected)
+
def test_map_dict_subclass_with_missing(self):
"""
Test Series.map with a dictionary subclass that defines __missing__,
| - [x] closes #17648
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29367 | 2019-11-02T17:18:17Z | 2019-11-03T01:57:00Z | 2019-11-03T01:57:00Z | 2019-11-03T01:57:09Z |
Test added: uint64 multicolumn sort | diff --git a/pandas/tests/frame/test_sorting.py b/pandas/tests/frame/test_sorting.py
index 9ea78b974fcbb..6ed245b6ebb98 100644
--- a/pandas/tests/frame/test_sorting.py
+++ b/pandas/tests/frame/test_sorting.py
@@ -735,3 +735,26 @@ def test_sort_index_na_position_with_categories_raises(self):
with pytest.raises(ValueError):
df.sort_values(by="c", ascending=False, na_position="bad_position")
+
+ def test_sort_multicolumn_uint64(self):
+ # GH9918
+ # uint64 multicolumn sort
+
+ df = pd.DataFrame(
+ {
+ "a": pd.Series([18446637057563306014, 1162265347240853609]),
+ "b": pd.Series([1, 2]),
+ }
+ )
+ df["a"] = df["a"].astype(np.uint64)
+ result = df.sort_values(["a", "b"])
+
+ expected = pd.DataFrame(
+ {
+ "a": pd.Series([18446637057563306014, 1162265347240853609]),
+ "b": pd.Series([1, 2]),
+ },
+ index=pd.Index([1, 0]),
+ )
+
+ tm.assert_frame_equal(result, expected)
| - [x] closes #9918
- [x] tests added / passed
- [x] passes `black pandas`
- [x] added test test_sort_multicolumn_uint64 | https://api.github.com/repos/pandas-dev/pandas/pulls/29365 | 2019-11-02T17:02:53Z | 2019-11-03T15:05:47Z | 2019-11-03T15:05:46Z | 2019-11-03T15:05:51Z |
Disallow non-scalar fill_value in maybe_upcast | diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index 304eeac87f64d..69c2e7fef365f 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -686,6 +686,8 @@ def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
dtype : if None, then use the dtype of the values, else coerce to this type
copy : if True always make a copy even if no upcast is required
"""
+ if not is_scalar(fill_value):
+ raise ValueError("fill_value must be a scalar")
if is_extension_type(values):
if copy:
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 51108d9a5a573..1f5a14a41e6a3 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -1286,6 +1286,10 @@ def diff(self, n: int, axis: int = 1) -> List["Block"]:
def shift(self, periods, axis=0, fill_value=None):
""" shift the block by periods, possibly upcast """
+ if not lib.is_scalar(fill_value):
+ # We could go further and require e.g. self._can_hold_element(fv)
+ raise ValueError("fill_value must be a scalar")
+
# convert integer to float if necessary. need to do a lot more than
# that, handle boolean etc also
new_values, fill_value = maybe_upcast(self.values, fill_value)
diff --git a/pandas/core/internals/construction.py b/pandas/core/internals/construction.py
index 176f4acd113fe..4a8216cc73264 100644
--- a/pandas/core/internals/construction.py
+++ b/pandas/core/internals/construction.py
@@ -97,6 +97,9 @@ def masked_rec_array_to_mgr(data, index, columns, dtype, copy):
# fill if needed
new_arrays = []
for fv, arr, col in zip(fill_value, arrays, arr_columns):
+ # TODO: numpy docs suggest fv must be scalar, but could it be
+ # non-scalar for object dtype?
+ assert lib.is_scalar(fv), fv
mask = ma.getmaskarray(data[col])
if mask.any():
arr, fv = maybe_upcast(arr, fill_value=fv, copy=True)
| Along with #29331 and #29332 this allows us to rule out non-scalar fill_value being passed to maybe_promote for _almost_ all cases, the last few of which we'll have to address individually.
The restriction in `Block.shift` could be an API change if you squint and tilt your head.
The assertion in masked_rec_array_to_mgr is based on my read of the numpy docs, but it'd be worth double-checking. | https://api.github.com/repos/pandas-dev/pandas/pulls/29362 | 2019-11-02T16:27:26Z | 2019-11-04T14:41:06Z | 2019-11-04T14:41:06Z | 2019-11-04T14:59:25Z |
CLN: requested follow-ups | diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index e3c7fef6f048f..a08ae66865e20 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -380,6 +380,18 @@ ctypedef fused algos_t:
def _validate_limit(nobs: int, limit=None) -> int:
+ """
+ Check that the `limit` argument is a positive integer.
+
+ Parameters
+ ----------
+ nobs : int
+ limit : object
+
+ Returns
+ -------
+ int
+ """
if limit is None:
lim = nobs
else:
diff --git a/pandas/core/internals/blocks.py b/pandas/core/internals/blocks.py
index 51108d9a5a573..448d2faf8b85f 100644
--- a/pandas/core/internals/blocks.py
+++ b/pandas/core/internals/blocks.py
@@ -7,7 +7,7 @@
import numpy as np
-from pandas._libs import NaT, lib, tslib, writers
+from pandas._libs import NaT, algos as libalgos, lib, tslib, writers
from pandas._libs.index import convert_scalar
import pandas._libs.internals as libinternals
from pandas._libs.tslibs import Timedelta, conversion
@@ -393,10 +393,7 @@ def fillna(self, value, limit=None, inplace=False, downcast=None):
mask = isna(self.values)
if limit is not None:
- if not is_integer(limit):
- raise ValueError("Limit must be an integer")
- if limit < 1:
- raise ValueError("Limit must be greater than 0")
+ limit = libalgos._validate_limit(None, limit=limit)
mask[mask.cumsum(self.ndim - 1) > limit] = False
if not self._can_hold_na:
diff --git a/pandas/core/missing.py b/pandas/core/missing.py
index f2655c126b9e5..5a1bf6d37b081 100644
--- a/pandas/core/missing.py
+++ b/pandas/core/missing.py
@@ -11,7 +11,6 @@
ensure_float64,
is_datetime64_dtype,
is_datetime64tz_dtype,
- is_integer,
is_integer_dtype,
is_numeric_v_string_like,
is_scalar,
@@ -191,13 +190,7 @@ def interpolate_1d(
)
# default limit is unlimited GH #16282
- if limit is None:
- # limit = len(xvalues)
- pass
- elif not is_integer(limit):
- raise ValueError("Limit must be an integer")
- elif limit < 1:
- raise ValueError("Limit must be greater than 0")
+ limit = algos._validate_limit(nobs=None, limit=limit)
from pandas import Series
diff --git a/pandas/tests/reductions/test_reductions.py b/pandas/tests/reductions/test_reductions.py
index a04f8f0df1151..4dfe561831ced 100644
--- a/pandas/tests/reductions/test_reductions.py
+++ b/pandas/tests/reductions/test_reductions.py
@@ -299,15 +299,6 @@ def test_timedelta_ops(self):
result = td.to_frame().std()
assert result[0] == expected
- # invalid ops
- for op in ["skew", "kurt", "sem", "prod", "var"]:
- msg = "reduction operation '{}' not allowed for this dtype"
- with pytest.raises(TypeError, match=msg.format(op)):
- getattr(td, op)()
-
- with pytest.raises(TypeError, match=msg.format(op)):
- getattr(td.to_frame(), op)(numeric_only=False)
-
# GH#10040
# make sure NaT is properly handled by median()
s = Series([Timestamp("2015-02-03"), Timestamp("2015-02-07")])
@@ -318,6 +309,22 @@ def test_timedelta_ops(self):
)
assert s.diff().median() == timedelta(days=6)
+ @pytest.mark.parametrize("opname", ["skew", "kurt", "sem", "prod", "var"])
+ def test_invalid_td64_reductions(self, opname):
+ s = Series(
+ [Timestamp("20130101") + timedelta(seconds=i * i) for i in range(10)]
+ )
+ td = s.diff()
+
+ msg = "reduction operation '{op}' not allowed for this dtype"
+ msg = msg.format(op=opname)
+
+ with pytest.raises(TypeError, match=msg):
+ getattr(td, opname)()
+
+ with pytest.raises(TypeError, match=msg):
+ getattr(td.to_frame(), opname)(numeric_only=False)
+
def test_minmax_tz(self, tz_naive_fixture):
tz = tz_naive_fixture
# monotonic
| Also use the new _validate_limit in two places in the non-cython code | https://api.github.com/repos/pandas-dev/pandas/pulls/29360 | 2019-11-02T16:10:56Z | 2019-11-02T19:48:23Z | 2019-11-02T19:48:23Z | 2019-11-02T20:58:25Z |
GH 16051: DataFrame.replace() overwrites when values are non-numeric | diff --git a/pandas/tests/frame/test_replace.py b/pandas/tests/frame/test_replace.py
index 5eb2416d0dcd7..c30efa121262f 100644
--- a/pandas/tests/frame/test_replace.py
+++ b/pandas/tests/frame/test_replace.py
@@ -1295,3 +1295,30 @@ def test_replace_method(self, to_replace, method, expected):
result = df.replace(to_replace=to_replace, value=None, method=method)
expected = DataFrame(expected)
tm.assert_frame_equal(result, expected)
+
+ @pytest.mark.parametrize(
+ "df, to_replace, exp",
+ [
+ (
+ {"col1": [1, 2, 3], "col2": [4, 5, 6]},
+ {4: 5, 5: 6, 6: 7},
+ {"col1": [1, 2, 3], "col2": [5, 6, 7]},
+ ),
+ (
+ {"col1": [1, 2, 3], "col2": ["4", "5", "6"]},
+ {"4": "5", "5": "6", "6": "7"},
+ {"col1": [1, 2, 3], "col2": ["5", "6", "7"]},
+ ),
+ ],
+ )
+ def test_replace_commutative(self, df, to_replace, exp):
+ # GH 16051
+ # DataFrame.replace() overwrites when values are non-numeric
+ # also added to data frame whilst issue was for series
+
+ df = pd.DataFrame(df)
+
+ expected = pd.DataFrame(exp)
+ result = df.replace(to_replace)
+
+ tm.assert_frame_equal(result, expected)
diff --git a/pandas/tests/series/test_replace.py b/pandas/tests/series/test_replace.py
index 86a54922fcf86..ebfd468e034f9 100644
--- a/pandas/tests/series/test_replace.py
+++ b/pandas/tests/series/test_replace.py
@@ -306,6 +306,24 @@ def test_replace_with_no_overflowerror(self):
expected = pd.Series([0, 1, "100000000000000000001"])
tm.assert_series_equal(result, expected)
+ @pytest.mark.parametrize(
+ "ser, to_replace, exp",
+ [
+ ([1, 2, 3], {1: 2, 2: 3, 3: 4}, [2, 3, 4]),
+ (["1", "2", "3"], {"1": "2", "2": "3", "3": "4"}, ["2", "3", "4"]),
+ ],
+ )
+ def test_replace_commutative(self, ser, to_replace, exp):
+ # GH 16051
+ # DataFrame.replace() overwrites when values are non-numeric
+
+ series = pd.Series(ser)
+
+ expected = pd.Series(exp)
+ result = series.replace(to_replace)
+
+ tm.assert_series_equal(result, expected)
+
@pytest.mark.parametrize(
"ser, exp", [([1, 2, 3], [1, True, 3]), (["x", 2, 3], ["x", True, 3])]
)
@@ -316,4 +334,5 @@ def test_replace_no_cast(self, ser, exp):
series = pd.Series(ser)
result = series.replace(2, True)
expected = pd.Series(exp)
+
tm.assert_series_equal(result, expected)
| - [x] closes #16051
- [x] tests added and passed
- [x] passes `black pandas`
- [x] added new test test_replace_commutative | https://api.github.com/repos/pandas-dev/pandas/pulls/29359 | 2019-11-02T15:36:33Z | 2019-11-04T16:54:55Z | 2019-11-04T16:54:55Z | 2019-11-04T16:55:03Z |
TST: Apply method broken for empty integer series with datetime index | diff --git a/pandas/tests/series/test_apply.py b/pandas/tests/series/test_apply.py
index e56294669a546..8eac79ae826c3 100644
--- a/pandas/tests/series/test_apply.py
+++ b/pandas/tests/series/test_apply.py
@@ -170,6 +170,12 @@ def test_apply_categorical_with_nan_values(self, series):
expected = expected.astype(object)
tm.assert_series_equal(result, expected)
+ def test_apply_empty_integer_series_with_datetime_index(self):
+ # GH 21245
+ s = pd.Series([], index=pd.date_range(start="2018-01-01", periods=0), dtype=int)
+ result = s.apply(lambda x: x)
+ tm.assert_series_equal(result, s)
+
class TestSeriesAggregate:
def test_transform(self, string_series):
| I added a unit test for an edge case that was failing. Using the apply method on an empty integer series with a datetime index would throw an error.
- [x] closes #21245
- [x] 1 test added
- [x] passes pandas and flake8
| https://api.github.com/repos/pandas-dev/pandas/pulls/29358 | 2019-11-02T15:23:05Z | 2019-11-02T19:51:08Z | 2019-11-02T19:51:08Z | 2019-11-02T19:51:13Z |
TST: new test for subset of a MultiIndex dtype | diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 79c9fe2b60bd9..4430628ce3d92 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -1932,6 +1932,15 @@ def test_repeat(self):
m_df = Series(data, index=m_idx)
assert m_df.repeat(3).shape == (3 * len(data),)
+ def test_subsets_multiindex_dtype(self):
+ # GH 20757
+ data = [["x", 1]]
+ columns = [("a", "b", np.nan), ("a", "c", 0.0)]
+ df = DataFrame(data, columns=pd.MultiIndex.from_tuples(columns))
+ expected = df.dtypes.a.b
+ result = df.a.b.dtypes
+ tm.assert_series_equal(result, expected)
+
class TestSorted(Base):
""" everything you wanted to test about sorting """
| Tried unsuccessfully reproducing Issue #20757 on pandas: 0.23.0 and 0.26.0.dev0+734.g0de99558b.dirty. That's why I decided to keep the input data from Issue #20757 unchanged.
- [x] closes #20757
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/29356 | 2019-11-02T15:11:47Z | 2019-11-16T22:06:10Z | 2019-11-16T22:06:09Z | 2019-11-16T22:06:14Z |
TST: Test type issue fix in empty groupby from DataFrame with categorical | diff --git a/pandas/tests/groupby/test_categorical.py b/pandas/tests/groupby/test_categorical.py
index 22a23407b2521..a187781ea214c 100644
--- a/pandas/tests/groupby/test_categorical.py
+++ b/pandas/tests/groupby/test_categorical.py
@@ -781,6 +781,22 @@ def test_categorical_no_compress():
tm.assert_numpy_array_equal(result, exp)
+def test_groupby_empty_with_category():
+ # GH-9614
+ # test fix for when group by on None resulted in
+ # coercion of dtype categorical -> float
+ df = pd.DataFrame(
+ {"A": [None] * 3, "B": pd.Categorical(["train", "train", "test"])}
+ )
+ result = df.groupby("A").first()["B"]
+ expected = pd.Series(
+ pd.Categorical([], categories=["test", "train"]),
+ index=pd.Series([], dtype="object", name="A"),
+ name="B",
+ )
+ tm.assert_series_equal(result, expected)
+
+
def test_sort():
# http://stackoverflow.com/questions/23814368/sorting-pandas-
| TST: Test type issue fix in empty groupby from DataFrame with categorical
closes #9614 | https://api.github.com/repos/pandas-dev/pandas/pulls/29355 | 2019-11-02T14:58:13Z | 2019-11-13T01:57:20Z | 2019-11-13T01:57:19Z | 2019-11-13T01:57:24Z |
Pv feature2 | diff --git a/pandas/tests/series/test_replace.py b/pandas/tests/series/test_replace.py
index e9d5a4b105a35..86a54922fcf86 100644
--- a/pandas/tests/series/test_replace.py
+++ b/pandas/tests/series/test_replace.py
@@ -305,3 +305,15 @@ def test_replace_with_no_overflowerror(self):
result = s.replace(["100000000000000000000"], [1])
expected = pd.Series([0, 1, "100000000000000000001"])
tm.assert_series_equal(result, expected)
+
+ @pytest.mark.parametrize(
+ "ser, exp", [([1, 2, 3], [1, True, 3]), (["x", 2, 3], ["x", True, 3])]
+ )
+ def test_replace_no_cast(self, ser, exp):
+ # GH 9113
+ # BUG: replace int64 dtype with bool coerces to int64
+
+ series = pd.Series(ser)
+ result = series.replace(2, True)
+ expected = pd.Series(exp)
+ tm.assert_series_equal(result, expected)
| - [x] closes #9113
- [x] tests added / passed
- [x] passes `black pandas`
- [x] new test added: test_replace_no_cast() | https://api.github.com/repos/pandas-dev/pandas/pulls/29354 | 2019-11-02T14:50:34Z | 2019-11-03T01:42:18Z | 2019-11-03T01:42:18Z | 2019-11-03T01:42:36Z |
BUG: Issue #29128 Series.var not returning the correct result | diff --git a/doc/source/whatsnew/v1.0.0.rst b/doc/source/whatsnew/v1.0.0.rst
index 664fcc91bacc4..2ae3379a6a23c 100644
--- a/doc/source/whatsnew/v1.0.0.rst
+++ b/doc/source/whatsnew/v1.0.0.rst
@@ -335,6 +335,7 @@ Numeric
- :class:`DataFrame` flex inequality comparisons methods (:meth:`DataFrame.lt`, :meth:`DataFrame.le`, :meth:`DataFrame.gt`, :meth: `DataFrame.ge`) with object-dtype and ``complex`` entries failing to raise ``TypeError`` like their :class:`Series` counterparts (:issue:`28079`)
- Bug in :class:`DataFrame` logical operations (`&`, `|`, `^`) not matching :class:`Series` behavior by filling NA values (:issue:`28741`)
- Bug in :meth:`DataFrame.interpolate` where specifying axis by name references variable before it is assigned (:issue:`29142`)
+- Bug in :meth:`Series.var` not computing the right value with a nullable integer dtype series not passing through ddof argument (:issue:`29128`)
- Improved error message when using `frac` > 1 and `replace` = False (:issue:`27451`)
-
diff --git a/pandas/core/arrays/integer.py b/pandas/core/arrays/integer.py
index 08b53e54b91ef..86e19508f2adc 100644
--- a/pandas/core/arrays/integer.py
+++ b/pandas/core/arrays/integer.py
@@ -652,7 +652,7 @@ def _reduce(self, name, skipna=True, **kwargs):
data[mask] = self._na_value
op = getattr(nanops, "nan" + name)
- result = op(data, axis=0, skipna=skipna, mask=mask)
+ result = op(data, axis=0, skipna=skipna, mask=mask, **kwargs)
# if we have a boolean op, don't coerce
if name in ["any", "all"]:
diff --git a/pandas/tests/arrays/test_integer.py b/pandas/tests/arrays/test_integer.py
index 793de66767cc3..025366e5b210b 100644
--- a/pandas/tests/arrays/test_integer.py
+++ b/pandas/tests/arrays/test_integer.py
@@ -829,6 +829,26 @@ def test_arrow_array(data):
assert arr.equals(expected)
+@pytest.mark.parametrize(
+ "pandasmethname, kwargs",
+ [
+ ("var", {"ddof": 0}),
+ ("var", {"ddof": 1}),
+ ("kurtosis", {}),
+ ("skew", {}),
+ ("sem", {}),
+ ],
+)
+def test_stat_method(pandasmethname, kwargs):
+ s = pd.Series(data=[1, 2, 3, 4, 5, 6, np.nan, np.nan], dtype="Int64")
+ pandasmeth = getattr(s, pandasmethname)
+ result = pandasmeth(**kwargs)
+ s2 = pd.Series(data=[1, 2, 3, 4, 5, 6], dtype="Int64")
+ pandasmeth = getattr(s2, pandasmethname)
+ expected = pandasmeth(**kwargs)
+ assert expected == result
+
+
# TODO(jreback) - these need testing / are broken
# shift
| - [ x ] closes #29128
- [ x ] tests added / passed
- [ x ] passes `black pandas`
- [ x ] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ x ] whatsnew entry
This PR closes https://github.com/pandas-dev/pandas/issues/29128 | https://api.github.com/repos/pandas-dev/pandas/pulls/29353 | 2019-11-02T14:48:42Z | 2019-11-19T04:23:55Z | 2019-11-19T04:23:55Z | 2019-11-19T04:24:04Z |
TST: Add test to check category dtype remains unchanged after concat. | diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index b537200dd7664..46dafbc4e1ec8 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -2747,6 +2747,22 @@ def test_concat_categorical_tz():
tm.assert_series_equal(result, expected)
+def test_concat_categorical_unchanged():
+ # GH-12007
+ # test fix for when concat on categorical and float
+ # coerces dtype categorical -> float
+ df = pd.DataFrame(pd.Series(["a", "b", "c"], dtype="category", name="A"))
+ ser = pd.Series([0, 1, 2], index=[0, 1, 3], name="B")
+ result = pd.concat([df, ser], axis=1)
+ expected = pd.DataFrame(
+ {
+ "A": pd.Series(["a", "b", "c", np.nan], dtype="category"),
+ "B": pd.Series([0, 1, np.nan, 2], dtype="float"),
+ }
+ )
+ tm.assert_equal(result, expected)
+
+
def test_concat_datetimeindex_freq():
# GH 3232
# Monotonic index result
| - closes #12007
- 1 tests added / passed | https://api.github.com/repos/pandas-dev/pandas/pulls/29352 | 2019-11-02T14:22:46Z | 2019-11-12T23:07:55Z | 2019-11-12T23:07:55Z | 2019-11-12T23:07:59Z |
pin black, xref gh-29341 | diff --git a/environment.yml b/environment.yml
index 163bd08b93c9e..4c96ab815dc90 100644
--- a/environment.yml
+++ b/environment.yml
@@ -16,7 +16,7 @@ dependencies:
- cython>=0.29.13
# code checks
- - black
+ - black<=19.3b0
- cpplint
- flake8
- flake8-comprehensions # used by flake8, linting of unnecessary comprehensions
diff --git a/requirements-dev.txt b/requirements-dev.txt
index 8a9974d393297..7a378cd2f2697 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -3,7 +3,7 @@ python-dateutil>=2.6.1
pytz
asv
cython>=0.29.13
-black
+black<=19.3b0
cpplint
flake8
flake8-comprehensions
| xref #29341
pinning until we can update the codebase | https://api.github.com/repos/pandas-dev/pandas/pulls/29351 | 2019-11-02T13:57:20Z | 2019-11-02T15:02:06Z | 2019-11-02T15:02:06Z | 2019-11-04T16:28:05Z |
Stable python 3.8.0 | diff --git a/.travis.yml b/.travis.yml
index b9fa06304d387..398dd07089ef9 100644
--- a/.travis.yml
+++ b/.travis.yml
@@ -32,7 +32,7 @@ matrix:
include:
- dist: bionic
# 18.04
- python: 3.8-dev
+ python: 3.8.0
env:
- JOB="3.8-dev" PATTERN="(not slow and not network)"
| Python 3.8.0 stable release is now available on travis https://travis-ci.community/t/add-python-3-8-support/5463 we can use it?
The 3.8-dev snapshot seemed to cause some issues here:
https://travis-ci.org/pandas-dev/pandas/jobs/606411398?utm_medium=notification&utm_source=github_status
related - https://github.com/pandas-dev/pandas/issues/26626 | https://api.github.com/repos/pandas-dev/pandas/pulls/29350 | 2019-11-02T13:40:57Z | 2019-11-02T15:51:27Z | 2019-11-02T15:51:27Z | 2019-11-02T16:09:16Z |
GH14422: BUG: to_numeric doesn't work uint64 numbers | diff --git a/pandas/tests/tools/test_numeric.py b/pandas/tests/tools/test_numeric.py
index 55f83e492e2cc..082277796e602 100644
--- a/pandas/tests/tools/test_numeric.py
+++ b/pandas/tests/tools/test_numeric.py
@@ -567,6 +567,24 @@ def test_downcast_limits(dtype, downcast, min_max):
assert series.dtype == dtype
+@pytest.mark.parametrize(
+ "ser,expected",
+ [
+ (
+ pd.Series([0, 9223372036854775808]),
+ pd.Series([0, 9223372036854775808], dtype=np.uint64),
+ )
+ ],
+)
+def test_downcast_uint64(ser, expected):
+ # see gh-14422:
+ # BUG: to_numeric doesn't work uint64 numbers
+
+ result = pd.to_numeric(ser, downcast="unsigned")
+
+ tm.assert_series_equal(result, expected)
+
+
@pytest.mark.parametrize(
"data,exp_data",
[
| -closes #14422
-passes `black pandas`
-added test test_downcast_uint64_exception() | https://api.github.com/repos/pandas-dev/pandas/pulls/29348 | 2019-11-02T13:07:14Z | 2019-11-05T17:19:53Z | 2019-11-05T17:19:53Z | 2019-11-05T17:19:56Z |
Update contributing.rst | diff --git a/doc/source/development/contributing.rst b/doc/source/development/contributing.rst
index 1f77c19f02301..56fac1cb6852a 100644
--- a/doc/source/development/contributing.rst
+++ b/doc/source/development/contributing.rst
@@ -482,7 +482,7 @@ reducing the turn-around time for checking your changes.
python make.py --no-api
# compile the docs with only a single section, relative to the "source" folder.
- # For example, compiling only this guide (docs/source/development/contributing.rst)
+ # For example, compiling only this guide (doc/source/development/contributing.rst)
python make.py clean
python make.py --single development/contributing.rst
| Correct Docs path
| https://api.github.com/repos/pandas-dev/pandas/pulls/29347 | 2019-11-02T13:06:48Z | 2019-11-02T14:15:51Z | 2019-11-02T14:15:51Z | 2019-11-02T14:15:55Z |
TST: Adding styler applymap multindex & code test [Ref: #25858] | diff --git a/pandas/tests/io/formats/test_style.py b/pandas/tests/io/formats/test_style.py
index 0f1402d7da389..0e88f5433c33b 100644
--- a/pandas/tests/io/formats/test_style.py
+++ b/pandas/tests/io/formats/test_style.py
@@ -376,6 +376,25 @@ def color_negative_red(val):
(df.style.applymap(color_negative_red, subset=idx[:, idx["b", "d"]]).render())
+ def test_applymap_subset_multiindex_code(self):
+ # https://github.com/pandas-dev/pandas/issues/25858
+ # Checks styler.applymap works with multindex when codes are provided
+ codes = np.array([[0, 0, 1, 1], [0, 1, 0, 1]])
+ columns = pd.MultiIndex(
+ levels=[["a", "b"], ["%", "#"]], codes=codes, names=["", ""]
+ )
+ df = DataFrame(
+ [[1, -1, 1, 1], [-1, 1, 1, 1]], index=["hello", "world"], columns=columns
+ )
+ pct_subset = pd.IndexSlice[:, pd.IndexSlice[:, "%":"%"]]
+
+ def color_negative_red(val):
+ color = "red" if val < 0 else "black"
+ return "color: %s" % color
+
+ df.loc[pct_subset]
+ df.style.applymap(color_negative_red, subset=pct_subset)
+
def test_where_with_one_style(self):
# GH 17474
def f(x):
| - [x] closes #25858
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29346 | 2019-11-02T12:44:13Z | 2019-11-05T20:57:40Z | 2019-11-05T20:57:39Z | 2019-11-05T20:57:49Z |
ensure consistency between columns aggregates with missing values present | diff --git a/pandas/tests/frame/test_apply.py b/pandas/tests/frame/test_apply.py
index 16d17b04423b7..a1172610b847e 100644
--- a/pandas/tests/frame/test_apply.py
+++ b/pandas/tests/frame/test_apply.py
@@ -1359,3 +1359,14 @@ def test_apply_datetime_tz_issue(self):
expected = pd.Series(index=timestamps, data=timestamps)
tm.assert_series_equal(result, expected)
+
+ @pytest.mark.parametrize("df", [pd.DataFrame({"A": ["a", None], "B": ["c", "d"]})])
+ @pytest.mark.parametrize("method", ["min", "max", "sum"])
+ def test_consistency_of_aggregates_of_columns_with_missing_values(self, df, method):
+ # GH 16832
+ none_in_first_column_result = getattr(df[["A", "B"]], method)()
+ none_in_second_column_result = getattr(df[["B", "A"]], method)()
+
+ tm.assert_series_equal(
+ none_in_first_column_result, none_in_second_column_result
+ )
| - [x] closes #16832
- [x] tests added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29345 | 2019-11-02T12:35:00Z | 2019-11-02T14:09:40Z | 2019-11-02T14:09:40Z | 2019-11-02T14:09:44Z |
TST: Test for Boolean Series with missing to Categorical dtype | diff --git a/pandas/tests/series/test_dtypes.py b/pandas/tests/series/test_dtypes.py
index 6ee120f3bec64..8f628d045a7f4 100644
--- a/pandas/tests/series/test_dtypes.py
+++ b/pandas/tests/series/test_dtypes.py
@@ -377,6 +377,15 @@ def test_astype_categorical_to_categorical(
result = s.astype("category")
tm.assert_series_equal(result, expected)
+ def test_astype_bool_missing_to_categorical(self):
+ # GH-19182
+ s = Series([True, False, np.nan])
+ assert s.dtypes == np.object_
+
+ result = s.astype(CategoricalDtype(categories=[True, False]))
+ expected = Series(Categorical([True, False, np.nan], categories=[True, False]))
+ tm.assert_series_equal(result, expected)
+
def test_astype_categoricaldtype(self):
s = Series(["a", "b", "a"])
result = s.astype(CategoricalDtype(["a", "b"], ordered=True))
| - [x] closes #19182
- [x] 1 test added / passed
- [x] passes `black pandas`
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
| https://api.github.com/repos/pandas-dev/pandas/pulls/29344 | 2019-11-02T12:30:06Z | 2019-11-02T20:12:28Z | 2019-11-02T20:12:28Z | 2019-11-04T01:00:44Z |
CI: fix conda version | diff --git a/ci/install_travis.sh b/ci/install_travis.sh
index 272e7f2e05d14..c92da8d4774e1 100755
--- a/ci/install_travis.sh
+++ b/ci/install_travis.sh
@@ -48,7 +48,12 @@ echo
echo "[update conda]"
conda config --set ssl_verify false || exit 1
conda config --set quiet true --set always_yes true --set changeps1 false || exit 1
-conda update -q conda
+
+# TODO(jreback), fix conoda version
+echo
+echo "[conda version]"
+conda install conda=4.4.4
+# conda update -q conda
if [ "$CONDA_BUILD_TEST" ]; then
echo
| https://api.github.com/repos/pandas-dev/pandas/pulls/19025 | 2018-01-01T13:43:53Z | 2018-01-01T14:15:38Z | 2018-01-01T14:15:38Z | 2018-01-01T14:15:38Z | |
dispatch Series[datetime64] ops to DatetimeIndex | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index bd3bee507baa3..a182f8bd9c9a4 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -208,6 +208,9 @@ Other API Changes
- In :func:`read_excel`, the ``comment`` argument is now exposed as a named parameter (:issue:`18735`)
- Rearranged the order of keyword arguments in :func:`read_excel()` to align with :func:`read_csv()` (:issue:`16672`)
- The options ``html.border`` and ``mode.use_inf_as_null`` were deprecated in prior versions, these will now show ``FutureWarning`` rather than a ``DeprecationWarning`` (:issue:`19003`)
+- Subtracting ``NaT`` from a :class:`Series` with ``dtype='datetime64[ns]'`` returns a ``Series`` with ``dtype='timedelta64[ns]'`` instead of ``dtype='datetime64[ns]'``(:issue:`18808`)
+- Operations between a :class:`Series` with dtype ``dtype='datetime64[ns]'`` and a :class:`PeriodIndex` will correctly raises ``TypeError`` (:issue:`18850`)
+- Subtraction of :class:`Series` with timezone-aware ``dtype='datetime64[ns]'`` with mis-matched timezones will raise ``TypeError`` instead of ``ValueError`` (issue:`18817`)
.. _whatsnew_0230.deprecations:
diff --git a/pandas/core/ops.py b/pandas/core/ops.py
index 0229f7c256464..554f0cb3803e9 100644
--- a/pandas/core/ops.py
+++ b/pandas/core/ops.py
@@ -341,10 +341,8 @@ def get_op(cls, left, right, name, na_op):
normal numpy path.
"""
is_timedelta_lhs = is_timedelta64_dtype(left)
- is_datetime_lhs = (is_datetime64_dtype(left) or
- is_datetime64tz_dtype(left))
- if not (is_datetime_lhs or is_timedelta_lhs):
+ if not is_timedelta_lhs:
return _Op(left, right, name, na_op)
else:
return _TimeOp(left, right, name, na_op)
@@ -364,14 +362,8 @@ def __init__(self, left, right, name, na_op):
rvalues = self._convert_to_array(right, name=name, other=lvalues)
# left
- self.is_offset_lhs = is_offsetlike(left)
self.is_timedelta_lhs = is_timedelta64_dtype(lvalues)
- self.is_datetime64_lhs = is_datetime64_dtype(lvalues)
- self.is_datetime64tz_lhs = is_datetime64tz_dtype(lvalues)
- self.is_datetime_lhs = (self.is_datetime64_lhs or
- self.is_datetime64tz_lhs)
- self.is_integer_lhs = left.dtype.kind in ['i', 'u']
- self.is_floating_lhs = left.dtype.kind == 'f'
+ assert self.is_timedelta_lhs
# right
self.is_offset_rhs = is_offsetlike(right)
@@ -387,34 +379,6 @@ def __init__(self, left, right, name, na_op):
self.lvalues, self.rvalues = self._convert_for_datetime(lvalues,
rvalues)
- def _validate_datetime(self, lvalues, rvalues, name):
- # assumes self.is_datetime_lhs
-
- if (self.is_timedelta_rhs or self.is_offset_rhs):
- # datetime and timedelta/DateOffset
- if name not in ('__add__', '__radd__', '__sub__'):
- raise TypeError("can only operate on a datetime with a rhs of "
- "a timedelta/DateOffset for addition and "
- "subtraction, but the operator [{name}] was "
- "passed".format(name=name))
-
- elif self.is_datetime_rhs:
- # 2 datetimes
- if name not in ('__sub__', '__rsub__'):
- raise TypeError("can only operate on a datetimes for"
- " subtraction, but the operator [{name}] was"
- " passed".format(name=name))
-
- # if tz's must be equal (same or None)
- if getattr(lvalues, 'tz', None) != getattr(rvalues, 'tz', None):
- raise ValueError("Incompatible tz's on datetime subtraction "
- "ops")
-
- else:
- raise TypeError('cannot operate on a series without a rhs '
- 'of a series/ndarray of type datetime64[ns] '
- 'or a timedelta')
-
def _validate_timedelta(self, name):
# assumes self.is_timedelta_lhs
@@ -440,44 +404,8 @@ def _validate_timedelta(self, name):
'of a series/ndarray of type datetime64[ns] '
'or a timedelta')
- def _validate_offset(self, name):
- # assumes self.is_offset_lhs
-
- if self.is_timedelta_rhs:
- # 2 timedeltas
- if name not in ('__div__', '__rdiv__', '__truediv__',
- '__rtruediv__', '__add__', '__radd__', '__sub__',
- '__rsub__'):
- raise TypeError("can only operate on a timedeltas for addition"
- ", subtraction, and division, but the operator"
- " [{name}] was passed".format(name=name))
-
- elif self.is_datetime_rhs:
- if name not in ('__add__', '__radd__'):
- raise TypeError("can only operate on a timedelta/DateOffset "
- "and a datetime for addition, but the operator"
- " [{name}] was passed".format(name=name))
-
- else:
- raise TypeError('cannot operate on a series without a rhs '
- 'of a series/ndarray of type datetime64[ns] '
- 'or a timedelta')
-
def _validate(self, lvalues, rvalues, name):
- if self.is_datetime_lhs:
- return self._validate_datetime(lvalues, rvalues, name)
- elif self.is_timedelta_lhs:
- return self._validate_timedelta(name)
- elif self.is_offset_lhs:
- return self._validate_offset(name)
-
- if ((self.is_integer_lhs or self.is_floating_lhs) and
- self.is_timedelta_rhs):
- self._check_timedelta_with_numeric(name)
- else:
- raise TypeError('cannot operate on a series without a rhs '
- 'of a series/ndarray of type datetime64[ns] '
- 'or a timedelta')
+ return self._validate_timedelta(name)
def _check_timedelta_with_numeric(self, name):
if name not in ('__div__', '__truediv__', '__mul__', '__rmul__'):
@@ -498,7 +426,7 @@ def _convert_to_array(self, values, name=None, other=None):
# if this is a Series that contains relevant dtype info, then use this
# instead of the inferred type; this avoids coercing Series([NaT],
# dtype='datetime64[ns]') to Series([NaT], dtype='timedelta64[ns]')
- elif (isinstance(values, pd.Series) and
+ elif (isinstance(values, (pd.Series, ABCDatetimeIndex)) and
(is_timedelta64_dtype(values) or is_datetime64_dtype(values))):
supplied_dtype = values.dtype
@@ -513,13 +441,11 @@ def _convert_to_array(self, values, name=None, other=None):
values = np.empty(values.shape, dtype='timedelta64[ns]')
values[:] = iNaT
- # a datelike
elif isinstance(values, ABCDatetimeIndex):
- # TODO: why are we casting to_series in the first place?
- values = values.to_series(keep_tz=True)
- # datetime with tz
- elif (isinstance(ovalues, datetime.datetime) and
- hasattr(ovalues, 'tzinfo')):
+ # a datelike
+ pass
+ elif isinstance(ovalues, datetime.datetime):
+ # datetime scalar
values = pd.DatetimeIndex(values)
# datetime array with tz
elif is_datetimetz(values):
@@ -571,17 +497,10 @@ def _convert_for_datetime(self, lvalues, rvalues):
mask = isna(lvalues) | isna(rvalues)
# datetimes require views
- if self.is_datetime_lhs or self.is_datetime_rhs:
+ if self.is_datetime_rhs:
# datetime subtraction means timedelta
- if self.is_datetime_lhs and self.is_datetime_rhs:
- if self.name in ('__sub__', '__rsub__'):
- self.dtype = 'timedelta64[ns]'
- else:
- self.dtype = 'datetime64[ns]'
- elif self.is_datetime64tz_lhs:
- self.dtype = lvalues.dtype
- elif self.is_datetime64tz_rhs:
+ if self.is_datetime64tz_rhs:
self.dtype = rvalues.dtype
else:
self.dtype = 'datetime64[ns]'
@@ -601,15 +520,11 @@ def _offset(lvalues, rvalues):
self.na_op = lambda x, y: getattr(x, self.name)(y)
return lvalues, rvalues
- if self.is_offset_lhs:
- lvalues, rvalues = _offset(lvalues, rvalues)
- elif self.is_offset_rhs:
+ if self.is_offset_rhs:
rvalues, lvalues = _offset(rvalues, lvalues)
else:
# with tz, convert to UTC
- if self.is_datetime64tz_lhs:
- lvalues = lvalues.tz_convert('UTC').tz_localize(None)
if self.is_datetime64tz_rhs:
rvalues = rvalues.tz_convert('UTC').tz_localize(None)
@@ -622,8 +537,6 @@ def _offset(lvalues, rvalues):
self.dtype = 'timedelta64[ns]'
# convert Tick DateOffset to underlying delta
- if self.is_offset_lhs:
- lvalues = to_timedelta(lvalues, box=False)
if self.is_offset_rhs:
rvalues = to_timedelta(rvalues, box=False)
@@ -634,7 +547,7 @@ def _offset(lvalues, rvalues):
# time delta division -> unit less
# integer gets converted to timedelta in np < 1.6
if ((self.is_timedelta_lhs and self.is_timedelta_rhs) and
- not self.is_integer_rhs and not self.is_integer_lhs and
+ not self.is_integer_rhs and
self.name in ('__div__', '__rdiv__',
'__truediv__', '__rtruediv__',
'__floordiv__', '__rfloordiv__')):
@@ -750,10 +663,16 @@ def wrapper(left, right, name=name, na_op=na_op):
return NotImplemented
left, right = _align_method_SERIES(left, right)
+ if is_datetime64_dtype(left) or is_datetime64tz_dtype(left):
+ result = op(pd.DatetimeIndex(left), right)
+ res_name = _get_series_op_result_name(left, right)
+ result.name = res_name # needs to be overriden if None
+ return construct_result(left, result,
+ index=left.index, name=res_name,
+ dtype=result.dtype)
converted = _Op.get_op(left, right, name, na_op)
- left, right = converted.left, converted.right
lvalues, rvalues = converted.lvalues, converted.rvalues
dtype = converted.dtype
wrap_results = converted.wrap_results
@@ -775,6 +694,7 @@ def wrapper(left, right, name=name, na_op=na_op):
res_name = left.name
result = wrap_results(safe_na_op(lvalues, rvalues))
+ res_name = _get_series_op_result_name(left, right)
return construct_result(
left,
result,
@@ -786,6 +706,15 @@ def wrapper(left, right, name=name, na_op=na_op):
return wrapper
+def _get_series_op_result_name(left, right):
+ # `left` is always a pd.Series
+ if isinstance(right, (ABCSeries, pd.Index)):
+ name = _maybe_match_name(left, right)
+ else:
+ name = left.name
+ return name
+
+
def _comp_method_OBJECT_ARRAY(op, x, y):
if isinstance(y, list):
y = construct_1d_object_array_from_listlike(y)
@@ -1388,23 +1317,6 @@ def f(self, other):
def _arith_method_PANEL(op, name, str_rep=None, fill_zeros=None,
default_axis=None, **eval_kwargs):
- # copied from Series na_op above, but without unnecessary branch for
- # non-scalar
- def na_op(x, y):
- import pandas.core.computation.expressions as expressions
-
- try:
- result = expressions.evaluate(op, str_rep, x, y, **eval_kwargs)
- except TypeError:
-
- # TODO: might need to find_common_type here?
- result = np.empty(len(x), dtype=x.dtype)
- mask = notna(x)
- result[mask] = op(x[mask], y)
- result, changed = maybe_upcast_putmask(result, ~mask, np.nan)
-
- result = missing.fill_zeros(result, x, y, name, fill_zeros)
- return result
# work only for scalars
def f(self, other):
diff --git a/pandas/tests/series/test_operators.py b/pandas/tests/series/test_operators.py
index a421f2cb15bba..c1e9a62d98fd3 100644
--- a/pandas/tests/series/test_operators.py
+++ b/pandas/tests/series/test_operators.py
@@ -960,6 +960,13 @@ def test_timedelta64_ops_nat(self):
assert_series_equal(timedelta_series / nan,
nat_series_dtype_timedelta)
+ def test_td64_sub_NaT(self):
+ # GH#18808
+ ser = Series([NaT, Timedelta('1s')])
+ res = ser - NaT
+ expected = Series([NaT, NaT], dtype='timedelta64[ns]')
+ tm.assert_series_equal(res, expected)
+
@pytest.mark.parametrize('scalar_td', [timedelta(minutes=5, seconds=4),
Timedelta(minutes=5, seconds=4),
Timedelta('5m4s').to_timedelta64()])
@@ -1076,7 +1083,7 @@ def run_ops(ops, get_ser, test_ser):
# defined
for op_str in ops:
op = getattr(get_ser, op_str, None)
- with tm.assert_raises_regex(TypeError, 'operate'):
+ with tm.assert_raises_regex(TypeError, 'operate|cannot'):
op(test_ser)
# ## timedelta64 ###
@@ -1253,6 +1260,20 @@ def test_datetime_series_with_DateOffset(self):
s + op(5)
op(5) + s
+ def test_dt64_sub_NaT(self):
+ # GH#18808
+ dti = pd.DatetimeIndex([pd.NaT, pd.Timestamp('19900315')])
+ ser = pd.Series(dti)
+ res = ser - pd.NaT
+ expected = pd.Series([pd.NaT, pd.NaT], dtype='timedelta64[ns]')
+ tm.assert_series_equal(res, expected)
+
+ dti_tz = dti.tz_localize('Asia/Tokyo')
+ ser_tz = pd.Series(dti_tz)
+ res = ser_tz - pd.NaT
+ expected = pd.Series([pd.NaT, pd.NaT], dtype='timedelta64[ns]')
+ tm.assert_series_equal(res, expected)
+
def test_datetime64_ops_nat(self):
# GH 11349
datetime_series = Series([NaT, Timestamp('19900315')])
@@ -1260,13 +1281,10 @@ def test_datetime64_ops_nat(self):
single_nat_dtype_datetime = Series([NaT], dtype='datetime64[ns]')
# subtraction
- assert_series_equal(datetime_series - NaT, nat_series_dtype_timestamp)
assert_series_equal(-NaT + datetime_series, nat_series_dtype_timestamp)
with pytest.raises(TypeError):
-single_nat_dtype_datetime + datetime_series
- assert_series_equal(nat_series_dtype_timestamp - NaT,
- nat_series_dtype_timestamp)
assert_series_equal(-NaT + nat_series_dtype_timestamp,
nat_series_dtype_timestamp)
with pytest.raises(TypeError):
@@ -2036,8 +2054,9 @@ def test_datetime64_with_index(self):
result = s - s.index
assert_series_equal(result, expected)
- result = s - s.index.to_period()
- assert_series_equal(result, expected)
+ with pytest.raises(TypeError):
+ # GH#18850
+ result = s - s.index.to_period()
df = DataFrame(np.random.randn(5, 2),
index=date_range('20130101', periods=5))
diff --git a/pandas/tests/series/test_timeseries.py b/pandas/tests/series/test_timeseries.py
index 2e3a7a6c28a11..6e711abf4491b 100644
--- a/pandas/tests/series/test_timeseries.py
+++ b/pandas/tests/series/test_timeseries.py
@@ -107,7 +107,7 @@ def test_shift(self):
# incompat tz
s2 = Series(date_range('2000-01-01 09:00:00', periods=5,
tz='CET'), name='foo')
- pytest.raises(ValueError, lambda: s - s2)
+ pytest.raises(TypeError, lambda: s - s2)
def test_shift2(self):
ts = Series(np.random.randn(5),
| This is the culmination of a bunch of recent work.
- [x] closes #18850, closes #18808, closes #17837
- [x] tests added / passed
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
If merged, this will subsume #18960 and will obviate parts of #18964. This will also fix (but not test, so we'll leave the issue open for now) the lack of overflow checks #12534. Also in a follow-up we'll be able to remove a bunch of _TimeOp. | https://api.github.com/repos/pandas-dev/pandas/pulls/19024 | 2018-01-01T02:18:52Z | 2018-01-04T00:27:50Z | 2018-01-04T00:27:49Z | 2018-01-23T04:40:43Z |
CLN: Remove tseries v0.19.0 deprecations | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 5fd7c3e217928..f0ed3ebf6e192 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -247,6 +247,8 @@ Removal of prior version deprecations/changes
- :func:`read_csv` has dropped the ``compact_ints`` and ``use_unsigned`` parameters (:issue:`13323`)
- The ``Timestamp`` class has dropped the ``offset`` attribute in favor of ``freq`` (:issue:`13593`)
- The ``Series``, ``Categorical``, and ``Index`` classes have dropped the ``reshape`` method (:issue:`13012`)
+- ``pandas.tseries.frequencies.get_standard_freq`` has been removed in favor of ``pandas.tseries.frequencies.to_offset(freq).rule_code`` (:issue:`13874`)
+- The ``freqstr`` keyword has been removed from ``pandas.tseries.frequencies.to_offset`` in favor of ``freq`` (:issue:`13874`)
.. _whatsnew_0230.performance:
diff --git a/pandas/tests/tseries/offsets/test_offsets.py b/pandas/tests/tseries/offsets/test_offsets.py
index e1a6463e7c351..7c7e5c4a5a35c 100644
--- a/pandas/tests/tseries/offsets/test_offsets.py
+++ b/pandas/tests/tseries/offsets/test_offsets.py
@@ -11,9 +11,8 @@
from pandas.compat.numpy import np_datetime64_compat
from pandas.core.series import Series
-from pandas.tseries.frequencies import (_offset_map, get_freq_code,
- _get_freq_str, _INVALID_FREQ_ERROR,
- get_offset, get_standard_freq)
+from pandas.tseries.frequencies import (_offset_map, get_freq_code, get_offset,
+ _get_freq_str, _INVALID_FREQ_ERROR)
from pandas.core.indexes.datetimes import (
_to_m8, DatetimeIndex, _daterange_cache)
import pandas._libs.tslibs.offsets as liboffsets
@@ -2786,33 +2785,6 @@ def test_get_offset_legacy():
get_offset(name)
-def test_get_standard_freq():
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- fstr = get_standard_freq('W')
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- assert fstr == get_standard_freq('w')
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- assert fstr == get_standard_freq('1w')
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- assert fstr == get_standard_freq(('W', 1))
-
- with tm.assert_raises_regex(ValueError, _INVALID_FREQ_ERROR):
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- get_standard_freq('WeEk')
-
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- fstr = get_standard_freq('5Q')
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- assert fstr == get_standard_freq('5q')
-
- with tm.assert_raises_regex(ValueError, _INVALID_FREQ_ERROR):
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- get_standard_freq('5QuarTer')
-
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- assert fstr == get_standard_freq(('q', 5))
-
-
class TestOffsetAliases(object):
def setup_method(self, method):
diff --git a/pandas/tests/tseries/test_frequencies.py b/pandas/tests/tseries/test_frequencies.py
index beea6df086b72..2486895086b2f 100644
--- a/pandas/tests/tseries/test_frequencies.py
+++ b/pandas/tests/tseries/test_frequencies.py
@@ -551,10 +551,6 @@ def test_frequency_misc(self):
with tm.assert_raises_regex(ValueError, 'Could not evaluate'):
frequencies.to_offset(('', ''))
- with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
- result = frequencies.get_standard_freq(offsets.Hour())
- assert result == 'H'
-
_dti = DatetimeIndex
diff --git a/pandas/tseries/frequencies.py b/pandas/tseries/frequencies.py
index f6e3d1f271036..4d1dd422be946 100644
--- a/pandas/tseries/frequencies.py
+++ b/pandas/tseries/frequencies.py
@@ -3,7 +3,6 @@
from pandas.compat import zip
from pandas import compat
import re
-import warnings
import numpy as np
@@ -14,7 +13,6 @@
is_datetime64_dtype)
from pandas.tseries.offsets import DateOffset
-from pandas.util._decorators import deprecate_kwarg
import pandas.tseries.offsets as offsets
from pandas._libs.tslib import Timedelta
@@ -143,7 +141,6 @@ def get_period_alias(offset_str):
'nanoseconds': Nano(1)}
-@deprecate_kwarg(old_arg_name='freqstr', new_arg_name='freq')
def to_offset(freq):
"""
Return DateOffset object from string or tuple representation
@@ -294,18 +291,6 @@ def get_offset(name):
getOffset = get_offset
-
-def get_standard_freq(freq):
- """
- Return the standardized frequency string
- """
-
- msg = ("get_standard_freq is deprecated. Use to_offset(freq).rule_code "
- "instead.")
- warnings.warn(msg, FutureWarning, stacklevel=2)
- return to_offset(freq).rule_code
-
-
# ---------------------------------------------------------------------
# Period codes
| * Remove frequencies.get_standard_freq
* Drop the "freqstr" keyword from frequencies.to_offset
Deprecated in v0.19.0
xref #13874 | https://api.github.com/repos/pandas-dev/pandas/pulls/19023 | 2018-01-01T01:25:30Z | 2018-01-02T11:20:59Z | 2018-01-02T11:20:59Z | 2018-01-02T17:11:30Z |
API: Prohibit non-numeric dtypes in IntervalIndex | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 4d806f1f05a16..a62a737fbba31 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -211,6 +211,7 @@ Other API Changes
- Subtracting ``NaT`` from a :class:`Series` with ``dtype='datetime64[ns]'`` returns a ``Series`` with ``dtype='timedelta64[ns]'`` instead of ``dtype='datetime64[ns]'``(:issue:`18808`)
- Operations between a :class:`Series` with dtype ``dtype='datetime64[ns]'`` and a :class:`PeriodIndex` will correctly raises ``TypeError`` (:issue:`18850`)
- Subtraction of :class:`Series` with timezone-aware ``dtype='datetime64[ns]'`` with mis-matched timezones will raise ``TypeError`` instead of ``ValueError`` (issue:`18817`)
+- :class:`IntervalIndex` and ``IntervalDtype`` no longer support categorical, object, and string subtypes (:issue:`19016`)
- The default ``Timedelta`` constructor now accepts an ``ISO 8601 Duration`` string as an argument (:issue:`19040`)
.. _whatsnew_0230.deprecations:
@@ -279,11 +280,11 @@ Performance Improvements
Documentation Changes
~~~~~~~~~~~~~~~~~~~~~
-- Changed spelling of "numpy" to "NumPy", and "python" to "Python". (:issue:`19017`)
+- Changed spelling of "numpy" to "NumPy", and "python" to "Python". (:issue:`19017`)
- Consistency when introducing code samples, using either colon or period.
Rewrote some sentences for greater clarity, added more dynamic references
to functions, methods and classes.
- (:issue:`18941`, :issue:`18948`, :issue:`18973`, :issue:`19017`)
+ (:issue:`18941`, :issue:`18948`, :issue:`18973`, :issue:`19017`)
-
.. _whatsnew_0230.bug_fixes:
@@ -310,7 +311,7 @@ Conversion
- Bug in :class:`DatetimeIndex` where adding or subtracting an array-like of ``DateOffset`` objects either raised (``np.array``, ``pd.Index``) or broadcast incorrectly (``pd.Series``) (:issue:`18849`)
- Bug in :class:`Series` floor-division where operating on a scalar ``timedelta`` raises an exception (:issue:`18846`)
- Bug in :class:`FY5253Quarter`, :class:`LastWeekOfMonth` where rollback and rollforward behavior was inconsistent with addition and subtraction behavior (:issue:`18854`)
-- Bug in :class:`Index` constructor with ``dtype=CategoricalDtype(...)`` where ``categories`` and ``ordered`` are not maintained (issue:`19032`)
+- Bug in :class:`Index` constructor with ``dtype=CategoricalDtype(...)`` where ``categories`` and ``ordered`` are not maintained (issue:`19032`)
Indexing
diff --git a/pandas/core/dtypes/dtypes.py b/pandas/core/dtypes/dtypes.py
index d1637873eb6e1..08773354d44d8 100644
--- a/pandas/core/dtypes/dtypes.py
+++ b/pandas/core/dtypes/dtypes.py
@@ -641,6 +641,8 @@ def __new__(cls, subtype=None):
----------
subtype : the dtype of the Interval
"""
+ from pandas.core.dtypes.common import (
+ is_categorical_dtype, is_string_dtype, pandas_dtype)
if isinstance(subtype, IntervalDtype):
return subtype
@@ -659,7 +661,6 @@ def __new__(cls, subtype=None):
if m is not None:
subtype = m.group('subtype')
- from pandas.core.dtypes.common import pandas_dtype
try:
subtype = pandas_dtype(subtype)
except TypeError:
@@ -670,6 +671,12 @@ def __new__(cls, subtype=None):
u.subtype = None
return u
+ if is_categorical_dtype(subtype) or is_string_dtype(subtype):
+ # GH 19016
+ msg = ('category, object, and string subtypes are not supported '
+ 'for IntervalDtype')
+ raise TypeError(msg)
+
try:
return cls._cache[str(subtype)]
except KeyError:
diff --git a/pandas/core/indexes/interval.py b/pandas/core/indexes/interval.py
index def9b151f5c91..fd1980f9ab429 100644
--- a/pandas/core/indexes/interval.py
+++ b/pandas/core/indexes/interval.py
@@ -11,6 +11,8 @@
is_list_like,
is_datetime_or_timedelta_dtype,
is_datetime64tz_dtype,
+ is_categorical_dtype,
+ is_string_dtype,
is_integer_dtype,
is_float_dtype,
is_interval_dtype,
@@ -92,6 +94,30 @@ def _get_interval_closed_bounds(interval):
return left, right
+def maybe_convert_platform_interval(values):
+ """
+ Try to do platform conversion, with special casing for IntervalIndex.
+ Wrapper around maybe_convert_platform that alters the default return
+ dtype in certain cases to be compatible with IntervalIndex. For example,
+ empty lists return with integer dtype instead of object dtype, which is
+ prohibited for IntervalIndex.
+
+ Parameters
+ ----------
+ values : array-like
+
+ Returns
+ -------
+ array
+ """
+ if isinstance(values, (list, tuple)) and len(values) == 0:
+ # GH 19016
+ # empty lists/tuples get object dtype by default, but this is not
+ # prohibited for IntervalIndex, so coerce to integer instead
+ return np.array([], dtype=np.intp)
+ return maybe_convert_platform(values)
+
+
def _new_IntervalIndex(cls, d):
"""
This is called upon unpickling, rather than the default which doesn't have
@@ -206,7 +232,7 @@ def __new__(cls, data, closed=None,
if is_scalar(data):
cls._scalar_data_error(data)
- data = maybe_convert_platform(data)
+ data = maybe_convert_platform_interval(data)
left, right, infer_closed = intervals_to_interval_bounds(data)
if _all_not_none(closed, infer_closed) and closed != infer_closed:
@@ -242,6 +268,11 @@ def _simple_new(cls, left, right, closed=None, name=None,
'[{rtype}] types')
raise ValueError(msg.format(ltype=type(left).__name__,
rtype=type(right).__name__))
+ elif is_categorical_dtype(left.dtype) or is_string_dtype(left.dtype):
+ # GH 19016
+ msg = ('category, object, and string subtypes are not supported '
+ 'for IntervalIndex')
+ raise TypeError(msg)
elif isinstance(left, ABCPeriodIndex):
msg = 'Period dtypes are not supported, use a PeriodIndex instead'
raise ValueError(msg)
@@ -403,7 +434,7 @@ def from_breaks(cls, breaks, closed='right', name=None, copy=False):
IntervalIndex.from_tuples : Construct an IntervalIndex from a
list/array of tuples
"""
- breaks = maybe_convert_platform(breaks)
+ breaks = maybe_convert_platform_interval(breaks)
return cls.from_arrays(breaks[:-1], breaks[1:], closed,
name=name, copy=copy)
@@ -444,8 +475,8 @@ def from_arrays(cls, left, right, closed='right', name=None, copy=False):
IntervalIndex.from_tuples : Construct an IntervalIndex from a
list/array of tuples
"""
- left = maybe_convert_platform(left)
- right = maybe_convert_platform(right)
+ left = maybe_convert_platform_interval(left)
+ right = maybe_convert_platform_interval(right)
return cls._simple_new(left, right, closed, name=name,
copy=copy, verify_integrity=True)
@@ -493,7 +524,7 @@ def from_intervals(cls, data, name=None, copy=False):
left, right, closed = data.left, data.right, data.closed
name = name or data.name
else:
- data = maybe_convert_platform(data)
+ data = maybe_convert_platform_interval(data)
left, right, closed = intervals_to_interval_bounds(data)
return cls.from_arrays(left, right, closed, name=name, copy=False)
diff --git a/pandas/tests/dtypes/test_dtypes.py b/pandas/tests/dtypes/test_dtypes.py
index d8e16482a414e..6a3715fd66159 100644
--- a/pandas/tests/dtypes/test_dtypes.py
+++ b/pandas/tests/dtypes/test_dtypes.py
@@ -152,7 +152,7 @@ def test_update_dtype(self, dtype, new_dtype):
assert result.ordered is expected_ordered
@pytest.mark.parametrize('bad_dtype', [
- 'foo', object, np.int64, PeriodDtype('Q'), IntervalDtype(object)])
+ 'foo', object, np.int64, PeriodDtype('Q')])
def test_update_dtype_errors(self, bad_dtype):
dtype = CategoricalDtype(list('abc'), False)
msg = 'a CategoricalDtype must be passed to perform an update, '
@@ -460,6 +460,17 @@ def test_construction(self):
assert i.subtype == np.dtype('int64')
assert is_interval_dtype(i)
+ @pytest.mark.parametrize('subtype', [
+ CategoricalDtype(list('abc'), False),
+ CategoricalDtype(list('wxyz'), True),
+ object, str, '<U10', 'interval[category]', 'interval[object]'])
+ def test_construction_not_supported(self, subtype):
+ # GH 19016
+ msg = ('category, object, and string subtypes are not supported '
+ 'for IntervalDtype')
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalDtype(subtype)
+
def test_construction_generic(self):
# generic
i = IntervalDtype('interval')
diff --git a/pandas/tests/indexes/interval/test_interval.py b/pandas/tests/indexes/interval/test_interval.py
index e2f48f40e9b7a..dd673294b128f 100644
--- a/pandas/tests/indexes/interval/test_interval.py
+++ b/pandas/tests/indexes/interval/test_interval.py
@@ -4,7 +4,7 @@
import numpy as np
from pandas import (
Interval, IntervalIndex, Index, isna, notna, interval_range, Timestamp,
- Timedelta, date_range, timedelta_range)
+ Timedelta, date_range, timedelta_range, Categorical)
from pandas.compat import lzip
from pandas.core.common import _asarray_tuplesafe
from pandas.tests.indexes.common import Base
@@ -42,7 +42,6 @@ def create_index_with_nan(self, closed='right'):
@pytest.mark.parametrize('data', [
Index([0, 1, 2, 3, 4]),
- Index(list('abcde')),
date_range('2017-01-01', periods=5),
date_range('2017-01-01', periods=5, tz='US/Eastern'),
timedelta_range('1 day', periods=5)])
@@ -138,10 +137,10 @@ def test_constructors_nan(self, closed, data):
[],
np.array([], dtype='int64'),
np.array([], dtype='float64'),
- np.array([], dtype=object)])
+ np.array([], dtype='datetime64[ns]')])
def test_constructors_empty(self, data, closed):
# GH 18421
- expected_dtype = data.dtype if isinstance(data, np.ndarray) else object
+ expected_dtype = getattr(data, 'dtype', np.intp)
expected_values = np.array([], dtype=object)
expected_index = IntervalIndex(data, closed=closed)
@@ -223,6 +222,48 @@ def test_constructors_errors(self):
with tm.assert_raises_regex(ValueError, msg):
IntervalIndex.from_arrays(range(10, -1, -1), range(9, -2, -1))
+ # GH 19016: categorical data
+ data = Categorical(list('01234abcde'), ordered=True)
+ msg = ('category, object, and string subtypes are not supported '
+ 'for IntervalIndex')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex.from_breaks(data)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex.from_arrays(data[:-1], data[1:])
+
+ @pytest.mark.parametrize('data', [
+ tuple('0123456789'),
+ list('abcdefghij'),
+ np.array(list('abcdefghij'), dtype=object),
+ np.array(list('abcdefghij'), dtype='<U1')])
+ def test_constructors_errors_string(self, data):
+ # GH 19016
+ left, right = data[:-1], data[1:]
+ tuples = lzip(left, right)
+ ivs = [Interval(l, r) for l, r in tuples] or data
+ msg = ('category, object, and string subtypes are not supported '
+ 'for IntervalIndex')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex(ivs)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ Index(ivs)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex.from_intervals(ivs)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex.from_breaks(data)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex.from_arrays(left, right)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ IntervalIndex.from_tuples(tuples)
+
@pytest.mark.parametrize('tz_left, tz_right', [
(None, 'UTC'), ('UTC', None), ('UTC', 'US/Eastern')])
def test_constructors_errors_tz(self, tz_left, tz_right):
@@ -298,18 +339,6 @@ def test_length(self, closed, breaks):
expected = Index(iv.length if notna(iv) else iv for iv in index)
tm.assert_index_equal(result, expected)
- @pytest.mark.parametrize('breaks', [
- list('abcdefgh'),
- lzip(range(10), range(1, 11)),
- [['A', 'B'], ['a', 'b'], ['c', 'd'], ['e', 'f']],
- [Interval(0, 1), Interval(1, 2), Interval(3, 4), Interval(4, 5)]])
- def test_length_errors(self, closed, breaks):
- # GH 18789
- index = IntervalIndex.from_breaks(breaks)
- msg = 'IntervalIndex contains Intervals without defined length'
- with tm.assert_raises_regex(TypeError, msg):
- index.length
-
def test_with_nans(self, closed):
index = self.create_index(closed=closed)
assert not index.hasnans
@@ -428,9 +457,7 @@ def test_delete(self, closed):
interval_range(0, periods=10, closed='neither'),
interval_range(1.7, periods=8, freq=2.5, closed='both'),
interval_range(Timestamp('20170101'), periods=12, closed='left'),
- interval_range(Timedelta('1 day'), periods=6, closed='right'),
- IntervalIndex.from_tuples([('a', 'd'), ('e', 'j'), ('w', 'z')]),
- IntervalIndex.from_tuples([(1, 2), ('a', 'z'), (3.14, 6.28)])])
+ interval_range(Timedelta('1 day'), periods=6, closed='right')])
def test_insert(self, data):
item = data[0]
idx_item = IntervalIndex([item])
@@ -504,15 +531,6 @@ def test_unique(self, closed):
[(0, 1), (0, 1), (2, 3)], closed=closed)
assert not idx.is_unique
- # unique mixed
- idx = IntervalIndex.from_tuples([(0, 1), ('a', 'b')], closed=closed)
- assert idx.is_unique
-
- # duplicate mixed
- idx = IntervalIndex.from_tuples(
- [(0, 1), ('a', 'b'), (0, 1)], closed=closed)
- assert not idx.is_unique
-
# empty
idx = IntervalIndex([], closed=closed)
assert idx.is_unique
| - [X] closes #19016
- [X] tests added / passed
- [X] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [X] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/19022 | 2018-01-01T00:56:09Z | 2018-01-05T14:15:45Z | 2018-01-05T14:15:45Z | 2018-01-05T17:44:28Z |
ENH: DataFrame.append preserves columns dtype if possible | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 408a52e0526ee..14146b9e455b4 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -380,6 +380,7 @@ Other Enhancements
- :class:`IntervalIndex` now supports time zone aware ``Interval`` objects (:issue:`18537`, :issue:`18538`)
- :func:`Series` / :func:`DataFrame` tab completion also returns identifiers in the first level of a :func:`MultiIndex`. (:issue:`16326`)
- :func:`read_excel()` has gained the ``nrows`` parameter (:issue:`16645`)
+- :meth:`DataFrame.append` can now in more cases preserve the type of the calling dataframe's columns (e.g. if both are ``CategoricalIndex``) (:issue:`18359`)
- :func:``DataFrame.to_json`` and ``Series.to_json`` now accept an ``index`` argument which allows the user to exclude the index from the JSON output (:issue:`17394`)
- ``IntervalIndex.to_tuples()`` has gained the ``na_tuple`` parameter to control whether NA is returned as a tuple of NA, or NA itself (:issue:`18756`)
- ``Categorical.rename_categories``, ``CategoricalIndex.rename_categories`` and :attr:`Series.cat.rename_categories`
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 9e57579ddfc05..ca20def643c2b 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -6113,8 +6113,11 @@ def append(self, other, ignore_index=False, verify_integrity=False):
# index name will be reset
index = Index([other.name], name=self.index.name)
- combined_columns = self.columns.tolist() + self.columns.union(
- other.index).difference(self.columns).tolist()
+ idx_diff = other.index.difference(self.columns)
+ try:
+ combined_columns = self.columns.append(idx_diff)
+ except TypeError:
+ combined_columns = self.columns.astype(object).append(idx_diff)
other = other.reindex(combined_columns, copy=False)
other = DataFrame(other.values.reshape((1, len(other))),
index=index,
diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index ffd37dc4b2f59..640d09f3587fb 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -1,5 +1,7 @@
from warnings import catch_warnings
+from itertools import combinations, product
+import datetime as dt
import dateutil
import numpy as np
from numpy.random import randn
@@ -829,12 +831,102 @@ def test_append_preserve_index_name(self):
result = df1.append(df2)
assert result.index.name == 'A'
+ indexes_can_append = [
+ pd.RangeIndex(3),
+ pd.Index([4, 5, 6]),
+ pd.Index([4.5, 5.5, 6.5]),
+ pd.Index(list('abc')),
+ pd.CategoricalIndex('A B C'.split()),
+ pd.CategoricalIndex('D E F'.split(), ordered=True),
+ pd.DatetimeIndex([dt.datetime(2013, 1, 3, 0, 0),
+ dt.datetime(2013, 1, 3, 6, 10),
+ dt.datetime(2013, 1, 3, 7, 12)]),
+ ]
+
+ indexes_cannot_append_with_other = [
+ pd.IntervalIndex.from_breaks([0, 1, 2, 3]),
+ pd.MultiIndex.from_arrays(['A B C'.split(), 'D E F'.split()]),
+ ]
+
+ all_indexes = indexes_can_append + indexes_cannot_append_with_other
+
+ @pytest.mark.parametrize("index",
+ all_indexes,
+ ids=lambda x: x.__class__.__name__)
+ def test_append_same_columns_type(self, index):
+ # GH18359
+
+ # df wider than ser
+ df = pd.DataFrame([[1, 2, 3], [4, 5, 6]], columns=index)
+ ser_index = index[:2]
+ ser = pd.Series([7, 8], index=ser_index, name=2)
+ result = df.append(ser)
+ expected = pd.DataFrame([[1., 2., 3.], [4, 5, 6], [7, 8, np.nan]],
+ index=[0, 1, 2],
+ columns=index)
+ assert_frame_equal(result, expected)
+
+ # ser wider than df
+ ser_index = index
+ index = index[:2]
+ df = pd.DataFrame([[1, 2], [4, 5]], columns=index)
+ ser = pd.Series([7, 8, 9], index=ser_index, name=2)
+ result = df.append(ser)
+ expected = pd.DataFrame([[1, 2, np.nan], [4, 5, np.nan], [7, 8, 9]],
+ index=[0, 1, 2],
+ columns=ser_index)
+ assert_frame_equal(result, expected)
+
+ @pytest.mark.parametrize("df_columns, series_index",
+ combinations(indexes_can_append, r=2),
+ ids=lambda x: x.__class__.__name__)
+ def test_append_different_columns_types(self, df_columns, series_index):
+ # GH18359
+ # See also test 'test_append_different_columns_types_raises' below
+ # for errors raised when appending
+
+ df = pd.DataFrame([[1, 2, 3], [4, 5, 6]], columns=df_columns)
+ ser = pd.Series([7, 8, 9], index=series_index, name=2)
+
+ result = df.append(ser)
+ idx_diff = ser.index.difference(df_columns)
+ combined_columns = Index(df_columns.tolist()).append(idx_diff)
+ expected = pd.DataFrame([[1., 2., 3., np.nan, np.nan, np.nan],
+ [4, 5, 6, np.nan, np.nan, np.nan],
+ [np.nan, np.nan, np.nan, 7, 8, 9]],
+ index=[0, 1, 2],
+ columns=combined_columns)
+ assert_frame_equal(result, expected)
+
+ @pytest.mark.parametrize(
+ "index_can_append, index_cannot_append_with_other",
+ product(indexes_can_append, indexes_cannot_append_with_other),
+ ids=lambda x: x.__class__.__name__)
+ def test_append_different_columns_types_raises(
+ self, index_can_append, index_cannot_append_with_other):
+ # GH18359
+ # Dataframe.append will raise if IntervalIndex/MultiIndex appends
+ # or is appended to a different index type
+ #
+ # See also test 'test_append_different_columns_types' above for
+ # appending without raising.
+
+ df = pd.DataFrame([[1, 2, 3], [4, 5, 6]], columns=index_can_append)
+ ser = pd.Series([7, 8, 9], index=index_cannot_append_with_other,
+ name=2)
+ with pytest.raises(TypeError):
+ df.append(ser)
+
+ df = pd.DataFrame([[1, 2, 3], [4, 5, 6]],
+ columns=index_cannot_append_with_other)
+ ser = pd.Series([7, 8, 9], index=index_can_append, name=2)
+ with pytest.raises(TypeError):
+ df.append(ser)
+
def test_append_dtype_coerce(self):
# GH 4993
# appending with datetime will incorrectly convert datetime64
- import datetime as dt
- from pandas import NaT
df1 = DataFrame(index=[1, 2], data=[dt.datetime(2013, 1, 1, 0, 0),
dt.datetime(2013, 1, 2, 0, 0)],
@@ -845,7 +937,9 @@ def test_append_dtype_coerce(self):
dt.datetime(2013, 1, 4, 7, 10)]],
columns=['start_time', 'end_time'])
- expected = concat([Series([NaT, NaT, dt.datetime(2013, 1, 3, 6, 10),
+ expected = concat([Series([pd.NaT,
+ pd.NaT,
+ dt.datetime(2013, 1, 3, 6, 10),
dt.datetime(2013, 1, 4, 7, 10)],
name='end_time'),
Series([dt.datetime(2013, 1, 1, 0, 0),
diff --git a/pandas/tests/reshape/test_pivot.py b/pandas/tests/reshape/test_pivot.py
index 92bedbabdf2f1..1004b40bfb4c1 100644
--- a/pandas/tests/reshape/test_pivot.py
+++ b/pandas/tests/reshape/test_pivot.py
@@ -1540,12 +1540,14 @@ def test_crosstab_normalize(self):
index=pd.Index([1, 2, 'All'],
name='a',
dtype='object'),
- columns=pd.Index([3, 4], name='b'))
+ columns=pd.Index([3, 4], name='b',
+ dtype='object'))
col_normal_margins = pd.DataFrame([[0.5, 0, 0.2], [0.5, 1.0, 0.8]],
index=pd.Index([1, 2], name='a',
dtype='object'),
columns=pd.Index([3, 4, 'All'],
- name='b'))
+ name='b',
+ dtype='object'))
all_normal_margins = pd.DataFrame([[0.2, 0, 0.2],
[0.2, 0.6, 0.8],
@@ -1554,7 +1556,8 @@ def test_crosstab_normalize(self):
name='a',
dtype='object'),
columns=pd.Index([3, 4, 'All'],
- name='b'))
+ name='b',
+ dtype='object'))
tm.assert_frame_equal(pd.crosstab(df.a, df.b, normalize='index',
margins=True), row_normal_margins)
tm.assert_frame_equal(pd.crosstab(df.a, df.b, normalize='columns',
| - [x] closes #18359
- [x] tests added / passed
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
This PR makes ``DataFrame.append`` preserve columns dtype, e.g.:
```python
>>> idx = pd.CategoricalIndex('a b'.split())
>>> df = pd.DataFrame([[1, 2]], columns=idx)
>>> ser = pd.Series([3, 4], index=idx, name=1)
>>> df.append(ser).columns
CategoricalIndex(['a', 'b'], categories=['a', 'b'], ordered=False, dtype='category')
```
Previously, the above returned ``Index(['a', 'b'], dtype='object')``, i.e. the index type information was lost when using ``append``. | https://api.github.com/repos/pandas-dev/pandas/pulls/19021 | 2017-12-31T23:14:24Z | 2018-04-20T00:35:14Z | 2018-04-20T00:35:14Z | 2021-05-22T23:20:06Z |
Spellcheck | diff --git a/doc/source/10min.rst b/doc/source/10min.rst
index 46c3ffef58228..da7679d8a3f54 100644
--- a/doc/source/10min.rst
+++ b/doc/source/10min.rst
@@ -48,7 +48,7 @@ a default integer index:
s = pd.Series([1,3,5,np.nan,6,8])
s
-Creating a :class:`DataFrame` by passing a numpy array, with a datetime index
+Creating a :class:`DataFrame` by passing a NumPy array, with a datetime index
and labeled columns:
.. ipython:: python
@@ -114,7 +114,7 @@ Here is how to view the top and bottom rows of the frame:
df.head()
df.tail(3)
-Display the index, columns, and the underlying numpy data:
+Display the index, columns, and the underlying NumPy data:
.. ipython:: python
@@ -311,7 +311,7 @@ Setting values by position:
df.iat[0,1] = 0
-Setting by assigning with a numpy array:
+Setting by assigning with a NumPy array:
.. ipython:: python
diff --git a/doc/source/advanced.rst b/doc/source/advanced.rst
index be749dfc1f594..25f7c5a3ad948 100644
--- a/doc/source/advanced.rst
+++ b/doc/source/advanced.rst
@@ -316,7 +316,9 @@ Basic multi-index slicing using slices, lists, and labels.
dfmi.loc[(slice('A1','A3'), slice(None), ['C1', 'C3']), :]
-You can use :class:`pandas.IndexSlice` to facilitate a more natural syntax using ``:``, rather than using ``slice(None)``.
+
+You can use :class:`pandas.IndexSlice` to facilitate a more natural syntax
+using ``:``, rather than using ``slice(None)``.
.. ipython:: python
@@ -557,7 +559,7 @@ Take Methods
.. _advanced.take:
-Similar to numpy ndarrays, pandas Index, Series, and DataFrame also provides
+Similar to NumPy ndarrays, pandas Index, Series, and DataFrame also provides
the ``take`` method that retrieves elements along a given axis at the given
indices. The given indices must be either a list or an ndarray of integer
index positions. ``take`` will also accept negative integers as relative positions to the end of the object.
@@ -729,7 +731,7 @@ This is an Immutable array implementing an ordered, sliceable set.
Prior to 0.18.0, the ``Int64Index`` would provide the default index for all ``NDFrame`` objects.
``RangeIndex`` is a sub-class of ``Int64Index`` added in version 0.18.0, now providing the default index for all ``NDFrame`` objects.
-``RangeIndex`` is an optimized version of ``Int64Index`` that can represent a monotonic ordered set. These are analogous to python `range types <https://docs.python.org/3/library/stdtypes.html#typesseq-range>`__.
+``RangeIndex`` is an optimized version of ``Int64Index`` that can represent a monotonic ordered set. These are analogous to Python `range types <https://docs.python.org/3/library/stdtypes.html#typesseq-range>`__.
.. _indexing.float64index:
@@ -763,7 +765,6 @@ The only positional indexing is via ``iloc``.
sf.iloc[3]
A scalar index that is not found will raise a ``KeyError``.
-
Slicing is primarily on the values of the index when using ``[],ix,loc``, and
**always** positional when using ``iloc``. The exception is when the slice is
boolean, in which case it will always be positional.
diff --git a/doc/source/api.rst b/doc/source/api.rst
index 17f6b8df0170d..02f729c89295b 100644
--- a/doc/source/api.rst
+++ b/doc/source/api.rst
@@ -730,7 +730,7 @@ The dtype information is available on the ``Categorical``
Categorical.codes
``np.asarray(categorical)`` works by implementing the array interface. Be aware, that this converts
-the Categorical back to a numpy array, so categories and order information is not preserved!
+the Categorical back to a NumPy array, so categories and order information is not preserved!
.. autosummary::
:toctree: generated/
diff --git a/doc/source/basics.rst b/doc/source/basics.rst
index 74b3dbb83ea91..bd49b5b7c9b32 100644
--- a/doc/source/basics.rst
+++ b/doc/source/basics.rst
@@ -395,7 +395,7 @@ raise a ValueError:
In [56]: pd.Series(['foo', 'bar', 'baz']) == pd.Series(['foo'])
ValueError: Series lengths must match to compare
-Note that this is different from the numpy behavior where a comparison can
+Note that this is different from the NumPy behavior where a comparison can
be broadcast:
.. ipython:: python
@@ -1000,7 +1000,7 @@ We create a frame similar to the one used in the above sections.
tsdf.iloc[3:7] = np.nan
tsdf
-Transform the entire frame. ``.transform()`` allows input functions as: a numpy function, a string
+Transform the entire frame. ``.transform()`` allows input functions as: a NumPy function, a string
function name or a user defined function.
.. ipython:: python
@@ -1510,7 +1510,7 @@ To iterate over the rows of a DataFrame, you can use the following methods:
one of the following approaches:
* Look for a *vectorized* solution: many operations can be performed using
- built-in methods or numpy functions, (boolean) indexing, ...
+ built-in methods or NumPy functions, (boolean) indexing, ...
* When you have a function that cannot work on the full DataFrame/Series
at once, it is better to use :meth:`~DataFrame.apply` instead of iterating
@@ -1971,7 +1971,7 @@ from the current type (e.g. ``int`` to ``float``).
df3.dtypes
The ``values`` attribute on a DataFrame return the *lower-common-denominator* of the dtypes, meaning
-the dtype that can accommodate **ALL** of the types in the resulting homogeneous dtyped numpy array. This can
+the dtype that can accommodate **ALL** of the types in the resulting homogeneous dtyped NumPy array. This can
force some *upcasting*.
.. ipython:: python
@@ -2253,7 +2253,7 @@ can define a function that returns a tree of child dtypes:
return dtype
return [dtype, [subdtypes(dt) for dt in subs]]
-All numpy dtypes are subclasses of ``numpy.generic``:
+All NumPy dtypes are subclasses of ``numpy.generic``:
.. ipython:: python
@@ -2262,4 +2262,4 @@ All numpy dtypes are subclasses of ``numpy.generic``:
.. note::
Pandas also defines the types ``category``, and ``datetime64[ns, tz]``, which are not integrated into the normal
- numpy hierarchy and wont show up with the above function.
+ NumPy hierarchy and wont show up with the above function.
diff --git a/doc/source/categorical.rst b/doc/source/categorical.rst
index 2acc919d1fbdf..7364167611730 100644
--- a/doc/source/categorical.rst
+++ b/doc/source/categorical.rst
@@ -40,7 +40,7 @@ The categorical data type is useful in the following cases:
* The lexical order of a variable is not the same as the logical order ("one", "two", "three").
By converting to a categorical and specifying an order on the categories, sorting and
min/max will use the logical order instead of the lexical order, see :ref:`here <categorical.sort>`.
-* As a signal to other python libraries that this column should be treated as a categorical
+* As a signal to other Python libraries that this column should be treated as a categorical
variable (e.g. to use suitable statistical methods or plot types).
See also the :ref:`API docs on categoricals<api.categorical>`.
@@ -366,7 +366,7 @@ or simply set the categories to a predefined scale, use :func:`Categorical.set_c
.. note::
Be aware that :func:`Categorical.set_categories` cannot know whether some category is omitted
intentionally or because it is misspelled or (under Python3) due to a type difference (e.g.,
- numpys S1 dtype and python strings). This can result in surprising behaviour!
+ numpys S1 dtype and Python strings). This can result in surprising behaviour!
Sorting and Order
-----------------
diff --git a/doc/source/comparison_with_sas.rst b/doc/source/comparison_with_sas.rst
index 1f2424d8a22f3..e9e0d7716af3a 100644
--- a/doc/source/comparison_with_sas.rst
+++ b/doc/source/comparison_with_sas.rst
@@ -10,7 +10,7 @@ performed in pandas.
If you're new to pandas, you might want to first read through :ref:`10 Minutes to pandas<10min>`
to familiarize yourself with the library.
-As is customary, we import pandas and numpy as follows:
+As is customary, we import pandas and NumPy as follows:
.. ipython:: python
@@ -100,7 +100,7 @@ specifying the column names.
A pandas ``DataFrame`` can be constructed in many different ways,
but for a small number of values, it is often convenient to specify it as
-a python dictionary, where the keys are the column names
+a Python dictionary, where the keys are the column names
and the values are the data.
.. ipython:: python
diff --git a/doc/source/comparison_with_sql.rst b/doc/source/comparison_with_sql.rst
index 2112c7de8c897..ba069b5a44c72 100644
--- a/doc/source/comparison_with_sql.rst
+++ b/doc/source/comparison_with_sql.rst
@@ -10,7 +10,7 @@ various SQL operations would be performed using pandas.
If you're new to pandas, you might want to first read through :ref:`10 Minutes to pandas<10min>`
to familiarize yourself with the library.
-As is customary, we import pandas and numpy as follows:
+As is customary, we import pandas and NumPy as follows:
.. ipython:: python
diff --git a/doc/source/computation.rst b/doc/source/computation.rst
index 30071c6c5b83c..06afa440aa26c 100644
--- a/doc/source/computation.rst
+++ b/doc/source/computation.rst
@@ -57,9 +57,8 @@ Covariance
s2 = pd.Series(np.random.randn(1000))
s1.cov(s2)
-Analogously, :meth:`DataFrame.cov` to compute
-pairwise covariances among the series in the DataFrame, also excluding
-NA/null values.
+Analogously, :meth:`DataFrame.cov` to compute pairwise covariances among the
+series in the DataFrame, also excluding NA/null values.
.. _computation.covariance.caveats:
diff --git a/doc/source/contributing.rst b/doc/source/contributing.rst
index b25f9779d3636..83437022563d5 100644
--- a/doc/source/contributing.rst
+++ b/doc/source/contributing.rst
@@ -118,7 +118,7 @@ Creating a development environment
----------------------------------
To test out code changes, you'll need to build pandas from source, which
-requires a C compiler and python environment. If you're making documentation
+requires a C compiler and Python environment. If you're making documentation
changes, you can skip to :ref:`contributing.documentation` but you won't be able
to build the documentation locally before pushing your changes.
@@ -187,7 +187,7 @@ At this point you should be able to import pandas from your locally built versio
0.22.0.dev0+29.g4ad6d4d74
This will create the new environment, and not touch any of your existing environments,
-nor any existing python installation.
+nor any existing Python installation.
To view your environments::
diff --git a/doc/source/cookbook.rst b/doc/source/cookbook.rst
index f13e5e67de07e..da54a6a5f5c02 100644
--- a/doc/source/cookbook.rst
+++ b/doc/source/cookbook.rst
@@ -41,7 +41,7 @@ above what the in-line examples offer.
Pandas (pd) and Numpy (np) are the only two abbreviated imported modules. The rest are kept
explicitly imported for newer users.
-These examples are written for python 3.4. Minor tweaks might be necessary for earlier python
+These examples are written for Python 3. Minor tweaks might be necessary for earlier python
versions.
Idioms
@@ -750,7 +750,7 @@ Timeseries
<http://nipunbatra.github.io/2015/06/timeseries/>`__
Turn a matrix with hours in columns and days in rows into a continuous row sequence in the form of a time series.
-`How to rearrange a python pandas DataFrame?
+`How to rearrange a Python pandas DataFrame?
<http://stackoverflow.com/questions/15432659/how-to-rearrange-a-python-pandas-dataframe>`__
`Dealing with duplicates when reindexing a timeseries to a specified frequency
@@ -1152,7 +1152,7 @@ Storing Attributes to a group node
store = pd.HDFStore('test.h5')
store.put('df',df)
- # you can store an arbitrary python object via pickle
+ # you can store an arbitrary Python object via pickle
store.get_storer('df').attrs.my_attribute = dict(A = 10)
store.get_storer('df').attrs.my_attribute
@@ -1167,7 +1167,7 @@ Storing Attributes to a group node
Binary Files
************
-pandas readily accepts numpy record arrays, if you need to read in a binary
+pandas readily accepts NumPy record arrays, if you need to read in a binary
file consisting of an array of C structs. For example, given this C program
in a file called ``main.c`` compiled with ``gcc main.c -std=gnu99`` on a
64-bit machine,
diff --git a/doc/source/dsintro.rst b/doc/source/dsintro.rst
index da9d2123bd1ca..7237dc5f1200b 100644
--- a/doc/source/dsintro.rst
+++ b/doc/source/dsintro.rst
@@ -23,7 +23,7 @@ Intro to Data Structures
We'll start with a quick, non-comprehensive overview of the fundamental data
structures in pandas to get you started. The fundamental behavior about data
types, indexing, and axis labeling / alignment apply across all of the
-objects. To get started, import numpy and load pandas into your namespace:
+objects. To get started, import NumPy and load pandas into your namespace:
.. ipython:: python
@@ -877,7 +877,7 @@ of DataFrames:
wp['Item3'] = wp['Item1'] / wp['Item2']
The API for insertion and deletion is the same as for DataFrame. And as with
-DataFrame, if the item is a valid python identifier, you can access it as an
+DataFrame, if the item is a valid Python identifier, you can access it as an
attribute and tab-complete it in IPython.
Transposing
diff --git a/doc/source/ecosystem.rst b/doc/source/ecosystem.rst
index 8ed647c2a19bc..c770bf2851643 100644
--- a/doc/source/ecosystem.rst
+++ b/doc/source/ecosystem.rst
@@ -27,7 +27,7 @@ Statistics and Machine Learning
`Statsmodels <http://www.statsmodels.org/>`__
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-Statsmodels is the prominent python "statistics and econometrics library" and it has
+Statsmodels is the prominent Python "statistics and econometrics library" and it has
a long-standing special relationship with pandas. Statsmodels provides powerful statistics,
econometrics, analysis and modeling functionality that is out of pandas' scope.
Statsmodels leverages pandas objects as the underlying data container for computation.
@@ -72,7 +72,7 @@ Hadley Wickham's `ggplot2 <http://ggplot2.org/>`__ is a foundational exploratory
Based on `"The Grammar of Graphics" <http://www.cs.uic.edu/~wilkinson/TheGrammarOfGraphics/GOG.html>`__ it
provides a powerful, declarative and extremely general way to generate bespoke plots of any kind of data.
It's really quite incredible. Various implementations to other languages are available,
-but a faithful implementation for python users has long been missing. Although still young
+but a faithful implementation for Python users has long been missing. Although still young
(as of Jan-2014), the `yhat/ggplot <https://github.com/yhat/ggplot>`__ project has been
progressing quickly in that direction.
@@ -192,7 +192,7 @@ or multi-indexed DataFrames.
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
fredapi is a Python interface to the `Federal Reserve Economic Data (FRED) <http://research.stlouisfed.org/fred2/>`__
provided by the Federal Reserve Bank of St. Louis. It works with both the FRED database and ALFRED database that
-contains point-in-time data (i.e. historic data revisions). fredapi provides a wrapper in python to the FRED
+contains point-in-time data (i.e. historic data revisions). fredapi provides a wrapper in Python to the FRED
HTTP API, and also provides several convenient methods for parsing and analyzing point-in-time data from ALFRED.
fredapi makes use of pandas and returns data in a Series or DataFrame. This module requires a FRED API key that
you can obtain for free on the FRED website.
diff --git a/doc/source/enhancingperf.rst b/doc/source/enhancingperf.rst
index 362c998493ae8..57f07a41afbc3 100644
--- a/doc/source/enhancingperf.rst
+++ b/doc/source/enhancingperf.rst
@@ -24,13 +24,13 @@ Enhancing Performance
Cython (Writing C extensions for pandas)
----------------------------------------
-For many use cases writing pandas in pure python and numpy is sufficient. In some
+For many use cases writing pandas in pure Python and NumPy is sufficient. In some
computationally heavy applications however, it can be possible to achieve sizeable
speed-ups by offloading work to `cython <http://cython.org/>`__.
This tutorial assumes you have refactored as much as possible in python, for example
-trying to remove for loops and making use of numpy vectorization, it's always worth
-optimising in python first.
+trying to remove for loops and making use of NumPy vectorization, it's always worth
+optimising in Python first.
This tutorial walks through a "typical" process of cythonizing a slow computation.
We use an `example from the cython documentation <http://docs.cython.org/src/quickstart/cythonize.html>`__
@@ -86,8 +86,8 @@ hence we'll concentrate our efforts cythonizing these two functions.
.. note::
- In python 2 replacing the ``range`` with its generator counterpart (``xrange``)
- would mean the ``range`` line would vanish. In python 3 ``range`` is already a generator.
+ In Python 2 replacing the ``range`` with its generator counterpart (``xrange``)
+ would mean the ``range`` line would vanish. In Python 3 ``range`` is already a generator.
.. _enhancingperf.plain:
@@ -232,7 +232,7 @@ the rows, applying our ``integrate_f_typed``, and putting this in the zeros arra
.. note::
Loops like this would be *extremely* slow in python, but in Cython looping
- over numpy arrays is *fast*.
+ over NumPy arrays is *fast*.
.. code-block:: ipython
@@ -315,7 +315,7 @@ Numba works by generating optimized machine code using the LLVM compiler infrast
Jit
~~~
-Using ``numba`` to just-in-time compile your code. We simply take the plain python code from above and annotate with the ``@jit`` decorator.
+Using ``numba`` to just-in-time compile your code. We simply take the plain Python code from above and annotate with the ``@jit`` decorator.
.. code-block:: python
@@ -391,7 +391,7 @@ Caveats
``numba`` will execute on any function, but can only accelerate certain classes of functions.
-``numba`` is best at accelerating functions that apply numerical functions to numpy arrays. When passed a function that only uses operations it knows how to accelerate, it will execute in ``nopython`` mode.
+``numba`` is best at accelerating functions that apply numerical functions to NumPy arrays. When passed a function that only uses operations it knows how to accelerate, it will execute in ``nopython`` mode.
If ``numba`` is passed a function that includes something it doesn't know how to work with -- a category that currently includes sets, lists, dictionaries, or string functions -- it will revert to ``object mode``. In ``object mode``, numba will execute but your code will not speed up significantly. If you would prefer that ``numba`` throw an error if it cannot compile a function in a way that speeds up your code, pass numba the argument ``nopython=True`` (e.g. ``@numba.jit(nopython=True)``). For more on troubleshooting ``numba`` modes, see the `numba troubleshooting page <http://numba.pydata.org/numba-doc/0.20.0/user/troubleshoot.html#the-compiled-code-is-too-slow>`__.
@@ -779,7 +779,7 @@ Technical Minutia Regarding Expression Evaluation
Expressions that would result in an object dtype or involve datetime operations
(because of ``NaT``) must be evaluated in Python space. The main reason for
-this behavior is to maintain backwards compatibility with versions of numpy <
+this behavior is to maintain backwards compatibility with versions of NumPy <
1.7. In those versions of ``numpy`` a call to ``ndarray.astype(str)`` will
truncate any strings that are more than 60 characters in length. Second, we
can't pass ``object`` arrays to ``numexpr`` thus string comparisons must be
diff --git a/doc/source/gotchas.rst b/doc/source/gotchas.rst
index 5da0f4fd07819..bc490877e190d 100644
--- a/doc/source/gotchas.rst
+++ b/doc/source/gotchas.rst
@@ -91,7 +91,7 @@ See also :ref:`Categorical Memory Usage <categorical.memory>`.
Using If/Truth Statements with pandas
-------------------------------------
-pandas follows the numpy convention of raising an error when you try to convert something to a ``bool``.
+pandas follows the NumPy convention of raising an error when you try to convert something to a ``bool``.
This happens in a ``if`` or when using the boolean operations, ``and``, ``or``, or ``not``. It is not clear
what the result of
diff --git a/doc/source/groupby.rst b/doc/source/groupby.rst
index 552ddabb7359a..413138b1e52fc 100644
--- a/doc/source/groupby.rst
+++ b/doc/source/groupby.rst
@@ -20,38 +20,38 @@ Group By: split-apply-combine
*****************************
By "group by" we are referring to a process involving one or more of the following
-steps
+steps:
- - **Splitting** the data into groups based on some criteria
- - **Applying** a function to each group independently
- - **Combining** the results into a data structure
+ - **Splitting** the data into groups based on some criteria.
+ - **Applying** a function to each group independently.
+ - **Combining** the results into a data structure.
-Of these, the split step is the most straightforward. In fact, in many
-situations you may wish to split the data set into groups and do something with
-those groups yourself. In the apply step, we might wish to one of the
+Out of these, the split step is the most straightforward. In fact, in many
+situations we may wish to split the data set into groups and do something with
+those groups. In the apply step, we might wish to one of the
following:
- - **Aggregation**: computing a summary statistic (or statistics) about each
+ - **Aggregation**: compute a summary statistic (or statistics) for each
group. Some examples:
- - Compute group sums or means
- - Compute group sizes / counts
+ - Compute group sums or means.
+ - Compute group sizes / counts.
- **Transformation**: perform some group-specific computations and return a
- like-indexed. Some examples:
+ like-indexed object. Some examples:
- - Standardizing data (zscore) within group
- - Filling NAs within groups with a value derived from each group
+ - Standardize data (zscore) within a group.
+ - Filling NAs within groups with a value derived from each group.
- **Filtration**: discard some groups, according to a group-wise computation
that evaluates True or False. Some examples:
- - Discarding data that belongs to groups with only a few members
- - Filtering out data based on the group sum or mean
+ - Discard data that belongs to groups with only a few members.
+ - Filter out data based on the group sum or mean.
- Some combination of the above: GroupBy will examine the results of the apply
step and try to return a sensibly combined result if it doesn't fit into
- either of the above two categories
+ either of the above two categories.
Since the set of object instance methods on pandas data structures are generally
rich and expressive, we often simply want to invoke, say, a DataFrame function
@@ -68,7 +68,7 @@ We aim to make operations like this natural and easy to express using
pandas. We'll address each area of GroupBy functionality then provide some
non-trivial examples / use cases.
-See the :ref:`cookbook<cookbook.grouping>` for some advanced strategies
+See the :ref:`cookbook<cookbook.grouping>` for some advanced strategies.
.. _groupby.split:
@@ -77,7 +77,7 @@ Splitting an object into groups
pandas objects can be split on any of their axes. The abstract definition of
grouping is to provide a mapping of labels to group names. To create a GroupBy
-object (more on what the GroupBy object is later), you do the following:
+object (more on what the GroupBy object is later), you may do the following:
.. code-block:: ipython
@@ -88,17 +88,18 @@ object (more on what the GroupBy object is later), you do the following:
The mapping can be specified many different ways:
- - A Python function, to be called on each of the axis labels
- - A list or NumPy array of the same length as the selected axis
- - A dict or Series, providing a ``label -> group name`` mapping
- - For DataFrame objects, a string indicating a column to be used to group. Of
- course ``df.groupby('A')`` is just syntactic sugar for
- ``df.groupby(df['A'])``, but it makes life simpler
- - For DataFrame objects, a string indicating an index level to be used to group.
- - A list of any of the above things
+ - A Python function, to be called on each of the axis labels.
+ - A list or NumPy array of the same length as the selected axis.
+ - A dict or ``Series``, providing a ``label -> group name`` mapping.
+ - For ``DataFrame`` objects, a string indicating a column to be used to group.
+ Of course ``df.groupby('A')`` is just syntactic sugar for
+ ``df.groupby(df['A'])``, but it makes life simpler.
+ - For ``DataFrame`` objects, a string indicating an index level to be used to
+ group.
+ - A list of any of the above things.
Collectively we refer to the grouping objects as the **keys**. For example,
-consider the following DataFrame:
+consider the following ``DataFrame``:
.. note::
@@ -119,7 +120,8 @@ consider the following DataFrame:
'D' : np.random.randn(8)})
df
-We could naturally group by either the ``A`` or ``B`` columns or both:
+On a DataFrame, we obtain a GroupBy object by calling :meth:`~DataFrame.groupby`.
+We could naturally group by either the ``A`` or ``B`` columns, or both:
.. ipython:: python
@@ -140,7 +142,7 @@ columns:
In [5]: grouped = df.groupby(get_letter_type, axis=1)
-pandas Index objects support duplicate values. If a
+pandas :class:`~pandas.Index` objects support duplicate values. If a
non-unique index is used as the group key in a groupby operation, all values
for the same index value will be considered to be in one group and thus the
output of aggregation functions will only contain unique index values:
@@ -220,7 +222,7 @@ the length of the ``groups`` dict, so it is largely just a convenience:
.. _groupby.tabcompletion:
-``GroupBy`` will tab complete column names (and other attributes)
+``GroupBy`` will tab complete column names (and other attributes):
.. ipython:: python
:suppress:
@@ -358,9 +360,9 @@ Index level names may be specified as keys directly to ``groupby``.
DataFrame column selection in GroupBy
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-Once you have created the GroupBy object from a DataFrame, for example, you
-might want to do something different for each of the columns. Thus, using
-``[]`` similar to getting a column from a DataFrame, you can do:
+Once you have created the GroupBy object from a DataFrame, you might want to do
+something different for each of the columns. Thus, using ``[]`` similar to
+getting a column from a DataFrame, you can do:
.. ipython:: python
:suppress:
@@ -393,7 +395,7 @@ Iterating through groups
------------------------
With the GroupBy object in hand, iterating through the grouped data is very
-natural and functions similarly to ``itertools.groupby``:
+natural and functions similarly to :py:func:`itertools.groupby`:
.. ipython::
@@ -419,7 +421,8 @@ statement if you wish: ``for (k1, k2), group in grouped:``.
Selecting a group
-----------------
-A single group can be selected using ``GroupBy.get_group()``:
+A single group can be selected using
+:meth:`~pandas.core.groupby.DataFrameGroupBy.get_group`:
.. ipython:: python
@@ -441,7 +444,9 @@ perform a computation on the grouped data. These operations are similar to the
:ref:`aggregating API <basics.aggregate>`, :ref:`window functions API <stats.aggregate>`,
and :ref:`resample API <timeseries.aggregate>`.
-An obvious one is aggregation via the ``aggregate`` or equivalently ``agg`` method:
+An obvious one is aggregation via the
+:meth:`~pandas.core.groupby.DataFrameGroupBy.aggregate` or equivalently
+:meth:`~pandas.core.groupby.DataFrameGroupBy.agg` method:
.. ipython:: python
@@ -491,11 +496,34 @@ index are the group names and whose values are the sizes of each group.
Passing ``as_index=False`` **will** return the groups that you are aggregating over, if they are
named *columns*.
- Aggregating functions are ones that reduce the dimension of the returned objects,
- for example: ``mean, sum, size, count, std, var, sem, describe, first, last, nth, min, max``. This is
- what happens when you do for example ``DataFrame.sum()`` and get back a ``Series``.
-
- ``nth`` can act as a reducer *or* a filter, see :ref:`here <groupby.nth>`
+Aggregating functions are the ones that reduce the dimension of the returned objects.
+Some common aggregating functions are tabulated below:
+
+.. csv-table::
+ :header: "Function", "Description"
+ :widths: 20, 80
+ :delim: ;
+
+ :meth:`~pd.core.groupby.DataFrameGroupBy.mean`;Compute mean of groups
+ :meth:`~pd.core.groupby.DataFrameGroupBy.sum`;Compute sum of group values
+ :meth:`~pd.core.groupby.DataFrameGroupBy.size`;Compute group sizes
+ :meth:`~pd.core.groupby.DataFrameGroupBy.count`;Compute count of group
+ :meth:`~pd.core.groupby.DataFrameGroupBy.std`;Standard deviation of groups
+ :meth:`~pd.core.groupby.DataFrameGroupBy.var`;Compute variance of groups
+ :meth:`~pd.core.groupby.DataFrameGroupBy.sem`;Standard error of the mean of groups
+ :meth:`~pd.core.groupby.DataFrameGroupBy.describe`;Generates descriptive statistics
+ :meth:`~pd.core.groupby.DataFrameGroupBy.first`;Compute first of group values
+ :meth:`~pd.core.groupby.DataFrameGroupBy.last`;Compute last of group values
+ :meth:`~pd.core.groupby.DataFrameGroupBy.nth`;Take nth value, or a subset if n is a list
+ :meth:`~pd.core.groupby.DataFrameGroupBy.min`;Compute min of group values
+ :meth:`~pd.core.groupby.DataFrameGroupBy.max`;Compute max of group values
+
+
+The aggregating functions above will exclude NA values. Any function which
+reduces a :class:`Series` to a scalar value is an aggregation function and will work,
+a trivial example is ``df.groupby('A').agg(lambda ser: 1)``. Note that
+:meth:`~pd.core.groupby.DataFrameGroupBy.nth` can act as a reducer *or* a
+filter, see :ref:`here <groupby.nth>`.
.. _groupby.aggregate.multifunc:
@@ -703,11 +731,11 @@ and that the transformed data contains no NAs.
.. note::
- Some functions when applied to a groupby object will automatically transform
- the input, returning an object of the same shape as the original. Passing
- ``as_index=False`` will not affect these transformation methods.
+ Some functions will automatically transform the input when applied to a
+ GroupBy object, but returning an object of the same shape as the original.
+ Passing ``as_index=False`` will not affect these transformation methods.
- For example: ``fillna, ffill, bfill, shift``.
+ For example: ``fillna, ffill, bfill, shift.``.
.. ipython:: python
@@ -898,7 +926,8 @@ The dimension of the returned result can also change:
In [11]: grouped.apply(f)
-``apply`` on a Series can operate on a returned value from the applied function, that is itself a series, and possibly upcast the result to a DataFrame
+``apply`` on a Series can operate on a returned value from the applied function,
+that is itself a series, and possibly upcast the result to a DataFrame:
.. ipython:: python
@@ -955,15 +984,21 @@ will be (silently) dropped. Thus, this does not pose any problems:
df.groupby('A').std()
+Note that ``df.groupby('A').colname.std().`` is more efficient than
+``df.groupby('A').std().colname``, so if the result of an aggregation function
+is only interesting over one column (here ``colname``), it may be filtered
+*before* applying the aggregation function.
+
.. _groupby.missing:
NA and NaT group handling
~~~~~~~~~~~~~~~~~~~~~~~~~
-If there are any NaN or NaT values in the grouping key, these will be automatically
-excluded. So there will never be an "NA group" or "NaT group". This was not the case in older
-versions of pandas, but users were generally discarding the NA group anyway
-(and supporting it was an implementation headache).
+If there are any NaN or NaT values in the grouping key, these will be
+automatically excluded. In other words, there will never be an "NA group" or
+"NaT group". This was not the case in older versions of pandas, but users were
+generally discarding the NA group anyway (and supporting it was an
+implementation headache).
Grouping with ordered factors
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@@ -1049,7 +1084,9 @@ This shows the first or last n rows from each group.
Taking the nth row of each group
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-To select from a DataFrame or Series the nth item, use the nth method. This is a reduction method, and will return a single row (or no row) per group if you pass an int for n:
+To select from a DataFrame or Series the nth item, use
+:meth:`~pd.core.groupby.DataFrameGroupBy.nth`. This is a reduction method, and
+will return a single row (or no row) per group if you pass an int for n:
.. ipython:: python
@@ -1116,8 +1153,10 @@ Enumerate groups
.. versionadded:: 0.20.2
To see the ordering of the groups (as opposed to the order of rows
-within a group given by ``cumcount``) you can use the ``ngroup``
-method.
+within a group given by ``cumcount``) you can use
+:meth:`~pandas.core.groupby.DataFrameGroupBy.ngroup`.
+
+
Note that the numbers given to the groups match the order in which the
groups would be seen when iterating over the groupby object, not the
@@ -1178,7 +1217,7 @@ allow for a cleaner, more readable syntax. To read about ``.pipe`` in general te
see :ref:`here <basics.pipe>`.
Combining ``.groupby`` and ``.pipe`` is often useful when you need to reuse
-GroupB objects.
+GroupBy objects.
For an example, imagine having a DataFrame with columns for stores, products,
revenue and sold quantity. We'd like to do a groupwise calculation of *prices*
@@ -1233,9 +1272,9 @@ Regroup columns of a DataFrame according to their sum, and sum the aggregated on
Multi-column factorization
~~~~~~~~~~~~~~~~~~~~~~~~~~
-By using ``.ngroup()``, we can extract information about the groups in
-a way similar to :func:`factorize` (as described further in the
-:ref:`reshaping API <reshaping.factorize>`) but which applies
+By using :meth:`~pandas.core.groupby.DataFrameGroupBy.ngroup`, we can extract
+information about the groups in a way similar to :func:`factorize` (as described
+further in the :ref:`reshaping API <reshaping.factorize>`) but which applies
naturally to multiple columns of mixed type and different
sources. This can be useful as an intermediate categorical-like step
in processing, when the relationships between the group rows are more
diff --git a/doc/source/indexing.rst b/doc/source/indexing.rst
index 355be5039f146..0467ac225585b 100644
--- a/doc/source/indexing.rst
+++ b/doc/source/indexing.rst
@@ -228,7 +228,7 @@ as an attribute:
.. warning::
- - You can use this access only if the index element is a valid python identifier, e.g. ``s.1`` is not allowed.
+ - You can use this access only if the index element is a valid Python identifier, e.g. ``s.1`` is not allowed.
See `here for an explanation of valid identifiers
<https://docs.python.org/3/reference/lexical_analysis.html#identifiers>`__.
@@ -441,7 +441,7 @@ Selection By Position
This is sometimes called ``chained assignment`` and should be avoided.
See :ref:`Returning a View versus Copy <indexing.view_versus_copy>`.
-Pandas provides a suite of methods in order to get **purely integer based indexing**. The semantics follow closely python and numpy slicing. These are ``0-based`` indexing. When slicing, the start bounds is *included*, while the upper bound is *excluded*. Trying to use a non-integer, even a **valid** label will raise an ``IndexError``.
+Pandas provides a suite of methods in order to get **purely integer based indexing**. The semantics follow closely Python and NumPy slicing. These are ``0-based`` indexing. When slicing, the start bounds is *included*, while the upper bound is *excluded*. Trying to use a non-integer, even a **valid** label will raise an ``IndexError``.
The ``.iloc`` attribute is the primary access method. The following are valid inputs:
@@ -777,7 +777,7 @@ using the ``replace`` option:
By default, each row has an equal probability of being selected, but if you want rows
to have different probabilities, you can pass the ``sample`` function sampling weights as
-``weights``. These weights can be a list, a numpy array, or a Series, but they must be of the same length as the object you are sampling. Missing values will be treated as a weight of zero, and inf values are not allowed. If weights do not sum to 1, they will be re-normalized by dividing all weights by the sum of the weights. For example:
+``weights``. These weights can be a list, a NumPy array, or a Series, but they must be of the same length as the object you are sampling. Missing values will be treated as a weight of zero, and inf values are not allowed. If weights do not sum to 1, they will be re-normalized by dividing all weights by the sum of the weights. For example:
.. ipython :: python
@@ -805,7 +805,7 @@ as a string.
df3 = pd.DataFrame({'col1':[1,2,3], 'col2':[2,3,4]})
df3.sample(n=1, axis=1)
-Finally, one can also set a seed for ``sample``'s random number generator using the ``random_state`` argument, which will accept either an integer (as a seed) or a numpy RandomState object.
+Finally, one can also set a seed for ``sample``'s random number generator using the ``random_state`` argument, which will accept either an integer (as a seed) or a NumPy RandomState object.
.. ipython :: python
@@ -893,7 +893,7 @@ evaluate an expression such as ``df.A > 2 & df.B < 3`` as
``df.A > (2 & df.B) < 3``, while the desired evaluation order is
``(df.A > 2) & (df.B < 3)``.
-Using a boolean vector to index a Series works exactly as in a numpy ndarray:
+Using a boolean vector to index a Series works exactly as in a NumPy ndarray:
.. ipython:: python
@@ -1125,7 +1125,6 @@ as condition and ``other`` argument.
'C': [7, 8, 9]})
df3.where(lambda x: x > 4, lambda x: x + 10)
-
Mask
~~~~
@@ -1712,6 +1711,7 @@ As a convenience, there is a new function on DataFrame called
DataFrame's columns and sets a simple integer index.
This is the inverse operation of :meth:`~DataFrame.set_index`.
+
.. ipython:: python
data
@@ -1772,7 +1772,7 @@ These both yield the same results, so which should you use? It is instructive to
of operations on these and why method 2 (``.loc``) is much preferred over method 1 (chained ``[]``).
``dfmi['one']`` selects the first level of the columns and returns a DataFrame that is singly-indexed.
-Then another python operation ``dfmi_with_one['second']`` selects the series indexed by ``'second'`` happens.
+Then another Python operation ``dfmi_with_one['second']`` selects the series indexed by ``'second'`` happens.
This is indicated by the variable ``dfmi_with_one`` because pandas sees these operations as separate events.
e.g. separate calls to ``__getitem__``, so it has to treat them as linear operations, they happen one after another.
diff --git a/doc/source/io.rst b/doc/source/io.rst
index 49d742d9905d7..5878272a3da42 100644
--- a/doc/source/io.rst
+++ b/doc/source/io.rst
@@ -164,7 +164,7 @@ dtype : Type name or dict of column -> type, default ``None``
.. versionadded:: 0.20.0 support for the Python parser.
engine : {``'c'``, ``'python'``}
- Parser engine to use. The C engine is faster while the python engine is
+ Parser engine to use. The C engine is faster while the Python engine is
currently more feature-complete.
converters : dict, default ``None``
Dict of functions for converting values in certain columns. Keys can either be
@@ -1529,9 +1529,9 @@ Specifying the parser engine
''''''''''''''''''''''''''''
Under the hood pandas uses a fast and efficient parser implemented in C as well
-as a python implementation which is currently more feature-complete. Where
+as a Python implementation which is currently more feature-complete. Where
possible pandas uses the C parser (specified as ``engine='c'``), but may fall
-back to python if C-unsupported options are specified. Currently, C-unsupported
+back to Python if C-unsupported options are specified. Currently, C-unsupported
options include:
- ``sep`` other than a single character (e.g. regex separators)
@@ -1582,7 +1582,7 @@ function takes a number of arguments. Only the first is required.
used. (A sequence should be given if the DataFrame uses MultiIndex).
- ``mode`` : Python write mode, default 'w'
- ``encoding``: a string representing the encoding to use if the contents are
- non-ASCII, for python versions prior to 3
+ non-ASCII, for Python versions prior to 3
- ``line_terminator``: Character sequence denoting line end (default '\\n')
- ``quoting``: Set quoting rules as in csv module (default csv.QUOTE_MINIMAL). Note that if you have set a `float_format` then floats are converted to strings and csv.QUOTE_NONNUMERIC will treat them as non-numeric
- ``quotechar``: Character used to quote fields (default '"')
@@ -1851,7 +1851,7 @@ is ``None``. To explicitly force ``Series`` parsing, pass ``typ=series``
- ``convert_axes`` : boolean, try to convert the axes to the proper dtypes, default is True
- ``convert_dates`` : a list of columns to parse for dates; If True, then try to parse date-like columns, default is True
- ``keep_default_dates`` : boolean, default True. If parsing dates, then parse the default date-like columns
-- ``numpy`` : direct decoding to numpy arrays. default is False;
+- ``numpy`` : direct decoding to NumPy arrays. default is False;
Supports numeric data only, although labels may be non-numeric. Also note that the JSON ordering **MUST** be the same for each term if ``numpy=True``
- ``precise_float`` : boolean, default ``False``. Set to enable usage of higher precision (strtod) function when decoding string to double values. Default (``False``) is to use fast but less precise builtin functionality
- ``date_unit`` : string, the timestamp unit to detect if converting dates. Default
@@ -1962,7 +1962,7 @@ The Numpy Parameter
If ``numpy=True`` is passed to ``read_json`` an attempt will be made to sniff
an appropriate dtype during deserialization and to subsequently decode directly
-to numpy arrays, bypassing the need for intermediate Python objects.
+to NumPy arrays, bypassing the need for intermediate Python objects.
This can provide speedups if you are deserialising a large amount of numeric
data:
@@ -1999,7 +1999,7 @@ The speedup is less noticeable for smaller datasets:
.. warning::
- Direct numpy decoding makes a number of assumptions and may fail or produce
+ Direct NumPy decoding makes a number of assumptions and may fail or produce
unexpected output if these assumptions are not satisfied:
- data is numeric.
@@ -3187,7 +3187,7 @@ You can pass ``append=True`` to the writer to append to an existing pack
Unlike other io methods, ``to_msgpack`` is available on both a per-object basis,
``df.to_msgpack()`` and using the top-level ``pd.to_msgpack(...)`` where you
-can pack arbitrary collections of python lists, dicts, scalars, while intermixing
+can pack arbitrary collections of Python lists, dicts, scalars, while intermixing
pandas objects.
.. ipython:: python
@@ -4411,7 +4411,7 @@ Several caveats.
can ``.reset_index()`` to store the index or ``.reset_index(drop=True)`` to
ignore it.
- Duplicate column names and non-string columns names are not supported
-- Non supported types include ``Period`` and actual python object types. These will raise a helpful error message
+- Non supported types include ``Period`` and actual Python object types. These will raise a helpful error message
on an attempt at serialization.
See the `Full Documentation <https://github.com/wesm/feather>`__
@@ -4475,7 +4475,7 @@ Several caveats.
- Duplicate column names and non-string columns names are not supported
- Index level names, if specified, must be strings
- Categorical dtypes can be serialized to parquet, but will de-serialize as ``object`` dtype.
-- Non supported types include ``Period`` and actual python object types. These will raise a helpful error message
+- Non supported types include ``Period`` and actual Python object types. These will raise a helpful error message
on an attempt at serialization.
You can specify an ``engine`` to direct the serialization. This can be one of ``pyarrow``, or ``fastparquet``, or ``auto``.
diff --git a/doc/source/missing_data.rst b/doc/source/missing_data.rst
index e20537efc0e71..d2250ae7b2116 100644
--- a/doc/source/missing_data.rst
+++ b/doc/source/missing_data.rst
@@ -27,7 +27,7 @@ pandas.
NumPy will soon be able to provide a native NA type solution (similar to R)
performant enough to be used in pandas.
-See the :ref:`cookbook<cookbook.missing_data>` for some advanced strategies
+See the :ref:`cookbook<cookbook.missing_data>` for some advanced strategies.
Missing data basics
-------------------
@@ -43,7 +43,7 @@ series might start on different dates. Thus, values prior to the start date
would generally be marked as missing.
In pandas, one of the most common ways that missing data is **introduced** into
-a data set is by reindexing. For example
+a data set is by reindexing. For example:
.. ipython:: python
@@ -86,7 +86,7 @@ pandas provides the :func:`isna` and
.. warning::
- One has to be mindful that in python (and numpy), the ``nan's`` don't compare equal, but ``None's`` **do**.
+ One has to be mindful that in Python (and numpy), the ``nan's`` don't compare equal, but ``None's`` **do**.
Note that Pandas/numpy uses the fact that ``np.nan != np.nan``, and treats ``None`` like ``np.nan``.
.. ipython:: python
@@ -104,7 +104,7 @@ Datetimes
---------
For datetime64[ns] types, ``NaT`` represents missing values. This is a pseudo-native
-sentinel value that can be represented by numpy in a singular dtype (datetime64[ns]).
+sentinel value that can be represented by NumPy in a singular dtype (datetime64[ns]).
pandas objects provide intercompatibility between ``NaT`` and ``NaN``.
.. ipython:: python
@@ -169,10 +169,10 @@ The descriptive statistics and computational methods discussed in the
<api.series.stats>` and :ref:`here <api.dataframe.stats>`) are all written to
account for missing data. For example:
-* When summing data, NA (missing) values will be treated as zero
-* If the data are all NA, the result will be NA
+* When summing data, NA (missing) values will be treated as zero.
+* If the data are all NA, the result will be NA.
* Methods like **cumsum** and **cumprod** ignore NA values, but preserve them
- in the resulting arrays
+ in the resulting arrays.
.. ipython:: python
@@ -190,7 +190,8 @@ Sum/Prod of Empties/Nans
.. warning::
This behavior is now standard as of v0.21.0; previously sum/prod would give different
- results if the ``bottleneck`` package was installed. See the :ref:`here <whatsnew_0210.api_breaking.bottleneck>`.
+ results if the ``bottleneck`` package was installed.
+ See the :ref:`v0.21.0 whatsnew <whatsnew_0210.api_breaking.bottleneck>`.
With ``sum`` or ``prod`` on an empty or all-``NaN`` ``Series``, or columns of a ``DataFrame``, the result will be all-``NaN``.
@@ -200,7 +201,7 @@ With ``sum`` or ``prod`` on an empty or all-``NaN`` ``Series``, or columns of a
s.sum()
-Summing of an empty ``Series``
+Summing over an empty ``Series`` will return ``NaN``:
.. ipython:: python
@@ -250,7 +251,7 @@ of ways, which we illustrate:
df2
df2.fillna(0)
- df2['four'].fillna('missing')
+ df2['one'].fillna('missing')
**Fill gaps forward or backward**
@@ -328,7 +329,7 @@ Dropping axis labels with missing data: dropna
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
You may wish to simply exclude labels from a data set which refer to missing
-data. To do this, use the **dropna** method:
+data. To do this, use the :meth:`~DataFrame.dropna` method:
.. ipython:: python
:suppress:
@@ -343,7 +344,7 @@ data. To do this, use the **dropna** method:
df.dropna(axis=1)
df['one'].dropna()
-Series.dropna is a simpler method as it only has one axis to consider.
+An equivalent :meth:`~Series.dropna` method is available for Series.
DataFrame.dropna has considerably more options than Series.dropna, which can be
examined :ref:`in the API <api.dataframe.missing>`.
@@ -352,8 +353,8 @@ examined :ref:`in the API <api.dataframe.missing>`.
Interpolation
~~~~~~~~~~~~~
-Both Series and DataFrame objects have an ``interpolate`` method that, by default,
-performs linear interpolation at missing datapoints.
+Both Series and DataFrame objects have an :meth:`~DataFrame.interpolate` method
+that, by default, performs linear interpolation at missing datapoints.
.. ipython:: python
:suppress:
@@ -411,7 +412,7 @@ You can also interpolate with a DataFrame:
df.interpolate()
The ``method`` argument gives access to fancier interpolation methods.
-If you have scipy_ installed, you can set pass the name of a 1-d interpolation routine to ``method``.
+If you have scipy_ installed, you can pass the name of a 1-d interpolation routine to ``method``.
You'll want to consult the full scipy interpolation documentation_ and reference guide_ for details.
The appropriate interpolation method will depend on the type of data you are working with.
@@ -419,7 +420,7 @@ The appropriate interpolation method will depend on the type of data you are wor
``method='quadratic'`` may be appropriate.
* If you have values approximating a cumulative distribution function,
then ``method='pchip'`` should work well.
-* To fill missing values with goal of smooth plotting, use ``method='akima'``.
+* To fill missing values with goal of smooth plotting, consider ``method='akima'``.
.. warning::
@@ -562,7 +563,7 @@ String/Regular Expression Replacement
<https://docs.python.org/3/reference/lexical_analysis.html#string-literals>`__
if this is unclear.
-Replace the '.' with ``NaN`` (str -> str)
+Replace the '.' with ``NaN`` (str -> str):
.. ipython:: python
@@ -571,58 +572,58 @@ Replace the '.' with ``NaN`` (str -> str)
df.replace('.', np.nan)
Now do it with a regular expression that removes surrounding whitespace
-(regex -> regex)
+(regex -> regex):
.. ipython:: python
df.replace(r'\s*\.\s*', np.nan, regex=True)
-Replace a few different values (list -> list)
+Replace a few different values (list -> list):
.. ipython:: python
df.replace(['a', '.'], ['b', np.nan])
-list of regex -> list of regex
+list of regex -> list of regex:
.. ipython:: python
df.replace([r'\.', r'(a)'], ['dot', '\1stuff'], regex=True)
-Only search in column ``'b'`` (dict -> dict)
+Only search in column ``'b'`` (dict -> dict):
.. ipython:: python
df.replace({'b': '.'}, {'b': np.nan})
Same as the previous example, but use a regular expression for
-searching instead (dict of regex -> dict)
+searching instead (dict of regex -> dict):
.. ipython:: python
df.replace({'b': r'\s*\.\s*'}, {'b': np.nan}, regex=True)
-You can pass nested dictionaries of regular expressions that use ``regex=True``
+You can pass nested dictionaries of regular expressions that use ``regex=True``:
.. ipython:: python
df.replace({'b': {'b': r''}}, regex=True)
-or you can pass the nested dictionary like so
+Alternatively, you can pass the nested dictionary like so:
.. ipython:: python
df.replace(regex={'b': {r'\s*\.\s*': np.nan}})
You can also use the group of a regular expression match when replacing (dict
-of regex -> dict of regex), this works for lists as well
+of regex -> dict of regex), this works for lists as well.
.. ipython:: python
df.replace({'b': r'\s*(\.)\s*'}, {'b': r'\1ty'}, regex=True)
You can pass a list of regular expressions, of which those that match
-will be replaced with a scalar (list of regex -> regex)
+will be replaced with a scalar (list of regex -> regex).
.. ipython:: python
@@ -631,7 +632,7 @@ will be replaced with a scalar (list of regex -> regex)
All of the regular expression examples can also be passed with the
``to_replace`` argument as the ``regex`` argument. In this case the ``value``
argument must be passed explicitly by name or ``regex`` must be a nested
-dictionary. The previous example, in this case, would then be
+dictionary. The previous example, in this case, would then be:
.. ipython:: python
@@ -648,7 +649,7 @@ want to use a regular expression.
Numeric Replacement
~~~~~~~~~~~~~~~~~~~
-Similar to ``DataFrame.fillna``
+The :meth:`~DataFrame.replace` method is similar to :meth:`~DataFrame.fillna`.
.. ipython:: python
@@ -656,7 +657,7 @@ Similar to ``DataFrame.fillna``
df[np.random.rand(df.shape[0]) > 0.5] = 1.5
df.replace(1.5, np.nan)
-Replacing more than one value via lists works as well
+Replacing more than one value is possible by passing a list.
.. ipython:: python
@@ -664,7 +665,7 @@ Replacing more than one value via lists works as well
df.replace([1.5, df00], [np.nan, 'a'])
df[1].dtype
-You can also operate on the DataFrame in place
+You can also operate on the DataFrame in place:
.. ipython:: python
@@ -674,7 +675,7 @@ You can also operate on the DataFrame in place
When replacing multiple ``bool`` or ``datetime64`` objects, the first
argument to ``replace`` (``to_replace``) must match the type of the value
- being replaced type. For example,
+ being replaced. For example,
.. code-block:: python
@@ -702,9 +703,9 @@ Missing data casting rules and indexing
While pandas supports storing arrays of integer and boolean type, these types
are not capable of storing missing data. Until we can switch to using a native
-NA type in NumPy, we've established some "casting rules" when reindexing will
-cause missing data to be introduced into, say, a Series or DataFrame. Here they
-are:
+NA type in NumPy, we've established some "casting rules". When a reindexing
+operation introduces missing data, the Series will be cast according to the
+rules introduced in the table below.
.. csv-table::
:header: "data type", "Cast to"
diff --git a/doc/source/release.rst b/doc/source/release.rst
index 12932d9fcee4f..de045c426cf7b 100644
--- a/doc/source/release.rst
+++ b/doc/source/release.rst
@@ -2894,7 +2894,7 @@ Improvements to existing features
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
- Fixed various issues with internal pprinting code, the repr() for various objects
- including TimeStamp and Index now produces valid python code strings and
+ including TimeStamp and Index now produces valid Python code strings and
can be used to recreate the object, (:issue:`3038`, :issue:`3379`, :issue:`3251`, :issue:`3460`)
- ``convert_objects`` now accepts a ``copy`` parameter (defaults to ``True``)
- ``HDFStore``
diff --git a/doc/source/timedeltas.rst b/doc/source/timedeltas.rst
index 778db17a56b58..6bbfb54629c4d 100644
--- a/doc/source/timedeltas.rst
+++ b/doc/source/timedeltas.rst
@@ -238,7 +238,7 @@ Frequency Conversion
Timedelta Series, ``TimedeltaIndex``, and ``Timedelta`` scalars can be converted to other 'frequencies' by dividing by another timedelta,
or by astyping to a specific timedelta type. These operations yield Series and propagate ``NaT`` -> ``nan``.
-Note that division by the numpy scalar is true division, while astyping is equivalent of floor division.
+Note that division by the NumPy scalar is true division, while astyping is equivalent of floor division.
.. ipython:: python
diff --git a/doc/source/timeseries.rst b/doc/source/timeseries.rst
index 201af3c7d5355..fa21cc997d4f4 100644
--- a/doc/source/timeseries.rst
+++ b/doc/source/timeseries.rst
@@ -2016,7 +2016,7 @@ Pandas provides rich support for working with timestamps in different time
zones using ``pytz`` and ``dateutil`` libraries. ``dateutil`` currently is only
supported for fixed offset and tzfile zones. The default library is ``pytz``.
Support for ``dateutil`` is provided for compatibility with other
-applications e.g. if you use ``dateutil`` in other python packages.
+applications e.g. if you use ``dateutil`` in other Python packages.
Working with Time Zones
~~~~~~~~~~~~~~~~~~~~~~~
@@ -2264,15 +2264,15 @@ a convert on an aware stamp.
.. note::
- Using the ``.values`` accessor on a ``Series``, returns an numpy array of the data.
- These values are converted to UTC, as numpy does not currently support timezones (even though it is *printing* in the local timezone!).
+ Using the ``.values`` accessor on a ``Series``, returns an NumPy array of the data.
+ These values are converted to UTC, as NumPy does not currently support timezones (even though it is *printing* in the local timezone!).
.. ipython:: python
s_naive.values
s_aware.values
- Further note that once converted to a numpy array these would lose the tz tenor.
+ Further note that once converted to a NumPy array these would lose the tz tenor.
.. ipython:: python
diff --git a/doc/source/visualization.rst b/doc/source/visualization.rst
index 2c1d54c27caab..cbd17493beb7e 100644
--- a/doc/source/visualization.rst
+++ b/doc/source/visualization.rst
@@ -1270,7 +1270,7 @@ The layout of subplots can be specified by ``layout`` keyword. It can accept
The number of axes which can be contained by rows x columns specified by ``layout`` must be
larger than the number of required subplots. If layout can contain more axes than required,
-blank axes are not drawn. Similar to a numpy array's ``reshape`` method, you
+blank axes are not drawn. Similar to a NumPy array's ``reshape`` method, you
can use ``-1`` for one dimension to automatically calculate the number of rows
or columns needed, given the other.
diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index bd3bee507baa3..1280634aa6c1a 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -273,8 +273,11 @@ Performance Improvements
Documentation Changes
~~~~~~~~~~~~~~~~~~~~~
--
--
+- Changed spelling of "numpy" to "NumPy", and "python" to "Python". (:issue:`19017`)
+- Consistency when introducing code samples, using either colon or period.
+ Rewrote some sentences for greater clarity, added more dynamic references
+ to functions, methods and classes.
+ (:issue:`18941`, :issue:`18948`, :issue:`18973`, :issue:`19017`)
-
.. _whatsnew_0230.bug_fixes:
| This PR is a continuation of my read-through of the docs, the earlier PRs are #18941, #18973 and #18948.
**The changes included in this PR include**
- Changed spelling of "python" to "Python" for consistency, same with "numpy" to "NumPy". This change was made globally, i.e. for most documents in `/docs/source/`.
- A review of `groupby.rst` and `missing_data.rst`
- Added punctuation where missing.
- Added more function references (i.e. `:meth:...`) where appropriate.
- Rewrote a few sentences.
Feedback is very welcome.
| https://api.github.com/repos/pandas-dev/pandas/pulls/19017 | 2017-12-31T16:19:22Z | 2018-01-03T12:25:22Z | 2018-01-03T12:25:22Z | 2018-01-03T12:25:53Z |
DOC: Update some outdated information | diff --git a/README.md b/README.md
index ac043f5586498..4b9c9505e320a 100644
--- a/README.md
+++ b/README.md
@@ -160,10 +160,9 @@ pip install pandas
```
## Dependencies
-- [NumPy](http://www.numpy.org): 1.7.0 or higher
-- [python-dateutil](https://labix.org/python-dateutil): 1.5 or higher
-- [pytz](https://pythonhosted.org/pytz)
- - Needed for time zone support with ``pandas.date_range``
+- [NumPy](http://www.numpy.org): 1.9.0 or higher
+- [python-dateutil](https://labix.org/python-dateutil): 2.5.0 or higher
+- [pytz](https://pythonhosted.org/pytz): 2011k or higher
See the [full installation instructions](https://pandas.pydata.org/pandas-docs/stable/install.html#dependencies)
for recommended and optional dependencies.
@@ -205,9 +204,6 @@ See the full instructions for [installing from source](https://pandas.pydata.org
## Documentation
The official documentation is hosted on PyData.org: https://pandas.pydata.org/pandas-docs/stable
-The Sphinx documentation should provide a good starting point for learning how
-to use the library. Expect the docs to continue to expand as time goes on.
-
## Background
Work on ``pandas`` started at AQR (a quantitative hedge fund) in 2008 and
has been under active development since then.
diff --git a/setup.py b/setup.py
index 443f3eba69b4d..7dbf6c84a0451 100755
--- a/setup.py
+++ b/setup.py
@@ -198,10 +198,6 @@ def build_extensions(self):
munging and cleaning data, analyzing / modeling it, then organizing the results
of the analysis into a form suitable for plotting or tabular display. pandas is
the ideal tool for all of these tasks.
-
-Notes
------
-Windows binaries built against NumPy 1.8.1
"""
DISTNAME = 'pandas'
| https://api.github.com/repos/pandas-dev/pandas/pulls/19015 | 2017-12-31T12:56:45Z | 2017-12-31T14:43:54Z | 2017-12-31T14:43:54Z | 2018-05-02T13:09:34Z | |
datetimelike indexes add/sub zero-dim integer arrays | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 6407a33c442d0..b169d8600169e 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -368,7 +368,7 @@ Numeric
^^^^^^^
- Bug in :func:`Series.__sub__` subtracting a non-nanosecond ``np.datetime64`` object from a ``Series`` gave incorrect results (:issue:`7996`)
--
+- Bug in :class:`DatetimeIndex`, :class:`TimedeltaIndex` addition and subtraction of zero-dimensional integer arrays gave incorrect results (:issue:`19012`)
-
Categorical
diff --git a/pandas/core/indexes/datetimelike.py b/pandas/core/indexes/datetimelike.py
index 10c9e8e7dd18f..2a77a23c2cfa1 100644
--- a/pandas/core/indexes/datetimelike.py
+++ b/pandas/core/indexes/datetimelike.py
@@ -669,6 +669,8 @@ def __add__(self, other):
from pandas.core.index import Index
from pandas.core.indexes.timedeltas import TimedeltaIndex
from pandas.tseries.offsets import DateOffset
+
+ other = lib.item_from_zerodim(other)
if is_timedelta64_dtype(other):
return self._add_delta(other)
elif isinstance(self, TimedeltaIndex) and isinstance(other, Index):
@@ -689,6 +691,7 @@ def __add__(self, other):
return self._add_datelike(other)
else: # pragma: no cover
return NotImplemented
+
cls.__add__ = __add__
cls.__radd__ = __add__
@@ -697,6 +700,8 @@ def __sub__(self, other):
from pandas.core.indexes.datetimes import DatetimeIndex
from pandas.core.indexes.timedeltas import TimedeltaIndex
from pandas.tseries.offsets import DateOffset
+
+ other = lib.item_from_zerodim(other)
if is_timedelta64_dtype(other):
return self._add_delta(-other)
elif isinstance(self, TimedeltaIndex) and isinstance(other, Index):
@@ -724,6 +729,7 @@ def __sub__(self, other):
else: # pragma: no cover
return NotImplemented
+
cls.__sub__ = __sub__
def __rsub__(self, other):
@@ -737,8 +743,10 @@ def _add_delta(self, other):
return NotImplemented
def _add_delta_td(self, other):
- # add a delta of a timedeltalike
- # return the i8 result view
+ """
+ Add a delta of a timedeltalike
+ return the i8 result view
+ """
inc = delta_to_nanoseconds(other)
new_values = checked_add_with_arr(self.asi8, inc,
@@ -748,8 +756,10 @@ def _add_delta_td(self, other):
return new_values.view('i8')
def _add_delta_tdi(self, other):
- # add a delta of a TimedeltaIndex
- # return the i8 result view
+ """
+ Add a delta of a TimedeltaIndex
+ return the i8 result view
+ """
# delta operation
if not len(self) == len(other):
diff --git a/pandas/tests/indexes/conftest.py b/pandas/tests/indexes/conftest.py
index a0ee3e511ef37..217ee07affa84 100644
--- a/pandas/tests/indexes/conftest.py
+++ b/pandas/tests/indexes/conftest.py
@@ -1,4 +1,5 @@
import pytest
+import numpy as np
import pandas.util.testing as tm
from pandas.core.indexes.api import Index, MultiIndex
@@ -22,3 +23,9 @@
ids=lambda x: type(x).__name__)
def indices(request):
return request.param
+
+
+@pytest.fixture(params=[1, np.array(1, dtype=np.int64)])
+def one(request):
+ # zero-dim integer array behaves like an integer
+ return request.param
diff --git a/pandas/tests/indexes/datetimes/test_arithmetic.py b/pandas/tests/indexes/datetimes/test_arithmetic.py
index 11a52267ed1b4..4684eb89557bf 100644
--- a/pandas/tests/indexes/datetimes/test_arithmetic.py
+++ b/pandas/tests/indexes/datetimes/test_arithmetic.py
@@ -58,36 +58,37 @@ def test_dti_radd_timestamp_raises(self):
# -------------------------------------------------------------
# Binary operations DatetimeIndex and int
- def test_dti_add_int(self, tz):
+ def test_dti_add_int(self, tz, one):
+ # Variants of `one` for #19012
rng = pd.date_range('2000-01-01 09:00', freq='H',
periods=10, tz=tz)
- result = rng + 1
+ result = rng + one
expected = pd.date_range('2000-01-01 10:00', freq='H',
periods=10, tz=tz)
tm.assert_index_equal(result, expected)
- def test_dti_iadd_int(self, tz):
+ def test_dti_iadd_int(self, tz, one):
rng = pd.date_range('2000-01-01 09:00', freq='H',
periods=10, tz=tz)
expected = pd.date_range('2000-01-01 10:00', freq='H',
periods=10, tz=tz)
- rng += 1
+ rng += one
tm.assert_index_equal(rng, expected)
- def test_dti_sub_int(self, tz):
+ def test_dti_sub_int(self, tz, one):
rng = pd.date_range('2000-01-01 09:00', freq='H',
periods=10, tz=tz)
- result = rng - 1
+ result = rng - one
expected = pd.date_range('2000-01-01 08:00', freq='H',
periods=10, tz=tz)
tm.assert_index_equal(result, expected)
- def test_dti_isub_int(self, tz):
+ def test_dti_isub_int(self, tz, one):
rng = pd.date_range('2000-01-01 09:00', freq='H',
periods=10, tz=tz)
expected = pd.date_range('2000-01-01 08:00', freq='H',
periods=10, tz=tz)
- rng -= 1
+ rng -= one
tm.assert_index_equal(rng, expected)
# -------------------------------------------------------------
diff --git a/pandas/tests/indexes/period/test_arithmetic.py b/pandas/tests/indexes/period/test_arithmetic.py
index b64f9074c3cf0..356ea5fc656de 100644
--- a/pandas/tests/indexes/period/test_arithmetic.py
+++ b/pandas/tests/indexes/period/test_arithmetic.py
@@ -131,19 +131,21 @@ def test_add_iadd(self):
period.IncompatibleFrequency, msg):
rng += delta
- # int
+ def test_pi_add_int(self, one):
+ # Variants of `one` for #19012
rng = pd.period_range('2000-01-01 09:00', freq='H', periods=10)
- result = rng + 1
+ result = rng + one
expected = pd.period_range('2000-01-01 10:00', freq='H', periods=10)
tm.assert_index_equal(result, expected)
- rng += 1
+ rng += one
tm.assert_index_equal(rng, expected)
- def test_sub(self):
+ @pytest.mark.parametrize('five', [5, np.array(5, dtype=np.int64)])
+ def test_sub(self, five):
rng = period_range('2007-01', periods=50)
- result = rng - 5
- exp = rng + (-5)
+ result = rng - five
+ exp = rng + (-five)
tm.assert_index_equal(result, exp)
def test_sub_isub(self):
diff --git a/pandas/tests/indexes/timedeltas/test_arithmetic.py b/pandas/tests/indexes/timedeltas/test_arithmetic.py
index 3c567e52cccb5..3ecfcaff63bc5 100644
--- a/pandas/tests/indexes/timedeltas/test_arithmetic.py
+++ b/pandas/tests/indexes/timedeltas/test_arithmetic.py
@@ -121,28 +121,29 @@ def test_ufunc_coercions(self):
# -------------------------------------------------------------
# Binary operations TimedeltaIndex and integer
- def test_tdi_add_int(self):
+ def test_tdi_add_int(self, one):
+ # Variants of `one` for #19012
rng = timedelta_range('1 days 09:00:00', freq='H', periods=10)
- result = rng + 1
+ result = rng + one
expected = timedelta_range('1 days 10:00:00', freq='H', periods=10)
tm.assert_index_equal(result, expected)
- def test_tdi_iadd_int(self):
+ def test_tdi_iadd_int(self, one):
rng = timedelta_range('1 days 09:00:00', freq='H', periods=10)
expected = timedelta_range('1 days 10:00:00', freq='H', periods=10)
- rng += 1
+ rng += one
tm.assert_index_equal(rng, expected)
- def test_tdi_sub_int(self):
+ def test_tdi_sub_int(self, one):
rng = timedelta_range('1 days 09:00:00', freq='H', periods=10)
- result = rng - 1
+ result = rng - one
expected = timedelta_range('1 days 08:00:00', freq='H', periods=10)
tm.assert_index_equal(result, expected)
- def test_tdi_isub_int(self):
+ def test_tdi_isub_int(self, one):
rng = timedelta_range('1 days 09:00:00', freq='H', periods=10)
expected = timedelta_range('1 days 08:00:00', freq='H', periods=10)
- rng -= 1
+ rng -= one
tm.assert_index_equal(rng, expected)
# -------------------------------------------------------------
| Setup:
```
dti = pd.date_range('2016-01-01', periods=3, freq='H')
one = np.array(1)
```
0.21.1:
```
>>> dti + one
DatetimeIndex(['2016-01-01 00:00:00.000000001',
'2016-01-01 01:00:00.000000001',
'2016-01-01 02:00:00.000000001'],
dtype='datetime64[ns]', freq='H')
>>> dti.freq = None
>>> dti + one
DatetimeIndex(['2016-01-01 00:00:00.000000001',
'2016-01-01 01:00:00.000000001',
'2016-01-01 02:00:00.000000001'],
dtype='datetime64[ns]', freq=None)
```
Master: (See #19011)
```
>>> dti + one
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "pandas/core/indexes/datetimelike.py", line 685, in __add__
elif is_offsetlike(other):
File "pandas/core/dtypes/common.py", line 294, in is_offsetlike
elif (is_list_like(arr_or_obj) and len(arr_or_obj) and
TypeError: len() of unsized object
```
After
```
>>> dti + one
DatetimeIndex(['2016-01-01 01:00:00', '2016-01-01 02:00:00',
'2016-01-01 03:00:00'],
dtype='datetime64[ns]', freq='H')
>>> dti.freq = None
>>> dti + one
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "pandas/core/indexes/datetimelike.py", line 683, in __add__
return self.shift(other)
File "pandas/core/indexes/datetimelike.py", line 821, in shift
raise ValueError("Cannot shift with no freq")
ValueError: Cannot shift with no freq
```
- [x] closes #19012
- [x] tests added / passed
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/19013 | 2017-12-31T01:18:48Z | 2017-12-31T14:49:25Z | 2017-12-31T14:49:25Z | 2018-01-23T04:40:46Z |
TST: Split tests/indexes/interval/test_interval.py into separate files | diff --git a/pandas/tests/indexes/interval/test_interval.py b/pandas/tests/indexes/interval/test_interval.py
index 3ca4c31b7f059..73520e984ae12 100644
--- a/pandas/tests/indexes/interval/test_interval.py
+++ b/pandas/tests/indexes/interval/test_interval.py
@@ -2,14 +2,11 @@
import pytest
import numpy as np
-from datetime import timedelta
from pandas import (
Interval, IntervalIndex, Index, isna, notna, interval_range, Timestamp,
- Timedelta, compat, date_range, timedelta_range, DateOffset)
+ Timedelta, date_range, timedelta_range)
from pandas.compat import lzip
from pandas.core.common import _asarray_tuplesafe
-from pandas.tseries.offsets import Day
-from pandas._libs.interval import IntervalTree
from pandas.tests.indexes.common import Base
import pandas.util.testing as tm
import pandas as pd
@@ -1158,367 +1155,3 @@ def test_to_tuples_na(self, tuples, na_tuple):
assert all(isna(x) for x in result_na)
else:
assert isna(result_na)
-
-
-class TestIntervalRange(object):
-
- def test_construction_from_numeric(self, closed, name):
- # combinations of start/end/periods without freq
- expected = IntervalIndex.from_breaks(
- np.arange(0, 6), name=name, closed=closed)
-
- result = interval_range(start=0, end=5, name=name, closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=0, periods=5, name=name, closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=5, periods=5, name=name, closed=closed)
- tm.assert_index_equal(result, expected)
-
- # combinations of start/end/periods with freq
- expected = IntervalIndex.from_tuples([(0, 2), (2, 4), (4, 6)],
- name=name, closed=closed)
-
- result = interval_range(start=0, end=6, freq=2, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=0, periods=3, freq=2, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=6, periods=3, freq=2, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # output truncates early if freq causes end to be skipped.
- expected = IntervalIndex.from_tuples([(0.0, 1.5), (1.5, 3.0)],
- name=name, closed=closed)
- result = interval_range(start=0, end=4, freq=1.5, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- @pytest.mark.parametrize('tz', [None, 'US/Eastern'])
- def test_construction_from_timestamp(self, closed, name, tz):
- # combinations of start/end/periods without freq
- start = Timestamp('2017-01-01', tz=tz)
- end = Timestamp('2017-01-06', tz=tz)
- breaks = date_range(start=start, end=end)
- expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
-
- result = interval_range(start=start, end=end, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=start, periods=5, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=end, periods=5, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # combinations of start/end/periods with fixed freq
- freq = '2D'
- start = Timestamp('2017-01-01', tz=tz)
- end = Timestamp('2017-01-07', tz=tz)
- breaks = date_range(start=start, end=end, freq=freq)
- expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
-
- result = interval_range(start=start, end=end, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=start, periods=3, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=end, periods=3, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # output truncates early if freq causes end to be skipped.
- end = Timestamp('2017-01-08', tz=tz)
- result = interval_range(start=start, end=end, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # combinations of start/end/periods with non-fixed freq
- freq = 'M'
- start = Timestamp('2017-01-01', tz=tz)
- end = Timestamp('2017-12-31', tz=tz)
- breaks = date_range(start=start, end=end, freq=freq)
- expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
-
- result = interval_range(start=start, end=end, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=start, periods=11, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=end, periods=11, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # output truncates early if freq causes end to be skipped.
- end = Timestamp('2018-01-15', tz=tz)
- result = interval_range(start=start, end=end, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- def test_construction_from_timedelta(self, closed, name):
- # combinations of start/end/periods without freq
- start, end = Timedelta('1 day'), Timedelta('6 days')
- breaks = timedelta_range(start=start, end=end)
- expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
-
- result = interval_range(start=start, end=end, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=start, periods=5, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=end, periods=5, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # combinations of start/end/periods with fixed freq
- freq = '2D'
- start, end = Timedelta('1 day'), Timedelta('7 days')
- breaks = timedelta_range(start=start, end=end, freq=freq)
- expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
-
- result = interval_range(start=start, end=end, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(start=start, periods=3, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- result = interval_range(end=end, periods=3, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- # output truncates early if freq causes end to be skipped.
- end = Timedelta('7 days 1 hour')
- result = interval_range(start=start, end=end, freq=freq, name=name,
- closed=closed)
- tm.assert_index_equal(result, expected)
-
- def test_constructor_coverage(self):
- # float value for periods
- expected = pd.interval_range(start=0, periods=10)
- result = pd.interval_range(start=0, periods=10.5)
- tm.assert_index_equal(result, expected)
-
- # equivalent timestamp-like start/end
- start, end = Timestamp('2017-01-01'), Timestamp('2017-01-15')
- expected = pd.interval_range(start=start, end=end)
-
- result = pd.interval_range(start=start.to_pydatetime(),
- end=end.to_pydatetime())
- tm.assert_index_equal(result, expected)
-
- result = pd.interval_range(start=start.asm8, end=end.asm8)
- tm.assert_index_equal(result, expected)
-
- # equivalent freq with timestamp
- equiv_freq = ['D', Day(), Timedelta(days=1), timedelta(days=1),
- DateOffset(days=1)]
- for freq in equiv_freq:
- result = pd.interval_range(start=start, end=end, freq=freq)
- tm.assert_index_equal(result, expected)
-
- # equivalent timedelta-like start/end
- start, end = Timedelta(days=1), Timedelta(days=10)
- expected = pd.interval_range(start=start, end=end)
-
- result = pd.interval_range(start=start.to_pytimedelta(),
- end=end.to_pytimedelta())
- tm.assert_index_equal(result, expected)
-
- result = pd.interval_range(start=start.asm8, end=end.asm8)
- tm.assert_index_equal(result, expected)
-
- # equivalent freq with timedelta
- equiv_freq = ['D', Day(), Timedelta(days=1), timedelta(days=1)]
- for freq in equiv_freq:
- result = pd.interval_range(start=start, end=end, freq=freq)
- tm.assert_index_equal(result, expected)
-
- def test_errors(self):
- # not enough params
- msg = ('Of the three parameters: start, end, and periods, '
- 'exactly two must be specified')
-
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(start=0)
-
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(end=5)
-
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(periods=2)
-
- with tm.assert_raises_regex(ValueError, msg):
- interval_range()
-
- # too many params
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(start=0, end=5, periods=6)
-
- # mixed units
- msg = 'start, end, freq need to be type compatible'
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=0, end=Timestamp('20130101'), freq=2)
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=0, end=Timedelta('1 day'), freq=2)
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=0, end=10, freq='D')
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=Timestamp('20130101'), end=10, freq='D')
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=Timestamp('20130101'),
- end=Timedelta('1 day'), freq='D')
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=Timestamp('20130101'),
- end=Timestamp('20130110'), freq=2)
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=Timedelta('1 day'), end=10, freq='D')
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=Timedelta('1 day'),
- end=Timestamp('20130110'), freq='D')
-
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=Timedelta('1 day'),
- end=Timedelta('10 days'), freq=2)
-
- # invalid periods
- msg = 'periods must be a number, got foo'
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=0, periods='foo')
-
- # invalid start
- msg = 'start must be numeric or datetime-like, got foo'
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(start='foo', periods=10)
-
- # invalid end
- msg = r'end must be numeric or datetime-like, got \(0, 1\]'
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(end=Interval(0, 1), periods=10)
-
- # invalid freq for datetime-like
- msg = 'freq must be numeric or convertible to DateOffset, got foo'
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(start=0, end=10, freq='foo')
-
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(start=Timestamp('20130101'), periods=10, freq='foo')
-
- with tm.assert_raises_regex(ValueError, msg):
- interval_range(end=Timedelta('1 day'), periods=10, freq='foo')
-
- # mixed tz
- start = Timestamp('2017-01-01', tz='US/Eastern')
- end = Timestamp('2017-01-07', tz='US/Pacific')
- msg = 'Start and end cannot both be tz-aware with different timezones'
- with tm.assert_raises_regex(TypeError, msg):
- interval_range(start=start, end=end)
-
-
-class TestIntervalTree(object):
- def setup_method(self, method):
- gentree = lambda dtype: IntervalTree(np.arange(5, dtype=dtype),
- np.arange(5, dtype=dtype) + 2)
- self.tree = gentree('int64')
- self.trees = {dtype: gentree(dtype)
- for dtype in ['int32', 'int64', 'float32', 'float64']}
-
- def test_get_loc(self):
- for dtype, tree in self.trees.items():
- tm.assert_numpy_array_equal(tree.get_loc(1),
- np.array([0], dtype='int64'))
- tm.assert_numpy_array_equal(np.sort(tree.get_loc(2)),
- np.array([0, 1], dtype='int64'))
- with pytest.raises(KeyError):
- tree.get_loc(-1)
-
- def test_get_indexer(self):
- for dtype, tree in self.trees.items():
- tm.assert_numpy_array_equal(
- tree.get_indexer(np.array([1.0, 5.5, 6.5])),
- np.array([0, 4, -1], dtype='int64'))
- with pytest.raises(KeyError):
- tree.get_indexer(np.array([3.0]))
-
- def test_get_indexer_non_unique(self):
- indexer, missing = self.tree.get_indexer_non_unique(
- np.array([1.0, 2.0, 6.5]))
- tm.assert_numpy_array_equal(indexer[:1],
- np.array([0], dtype='int64'))
- tm.assert_numpy_array_equal(np.sort(indexer[1:3]),
- np.array([0, 1], dtype='int64'))
- tm.assert_numpy_array_equal(np.sort(indexer[3:]),
- np.array([-1], dtype='int64'))
- tm.assert_numpy_array_equal(missing, np.array([2], dtype='int64'))
-
- def test_duplicates(self):
- tree = IntervalTree([0, 0, 0], [1, 1, 1])
- tm.assert_numpy_array_equal(np.sort(tree.get_loc(0.5)),
- np.array([0, 1, 2], dtype='int64'))
-
- with pytest.raises(KeyError):
- tree.get_indexer(np.array([0.5]))
-
- indexer, missing = tree.get_indexer_non_unique(np.array([0.5]))
- tm.assert_numpy_array_equal(np.sort(indexer),
- np.array([0, 1, 2], dtype='int64'))
- tm.assert_numpy_array_equal(missing, np.array([], dtype='int64'))
-
- def test_get_loc_closed(self):
- for closed in ['left', 'right', 'both', 'neither']:
- tree = IntervalTree([0], [1], closed=closed)
- for p, errors in [(0, tree.open_left),
- (1, tree.open_right)]:
- if errors:
- with pytest.raises(KeyError):
- tree.get_loc(p)
- else:
- tm.assert_numpy_array_equal(tree.get_loc(p),
- np.array([0], dtype='int64'))
-
- @pytest.mark.skipif(compat.is_platform_32bit(),
- reason="int type mismatch on 32bit")
- def test_get_indexer_closed(self):
- x = np.arange(1000, dtype='float64')
- found = x.astype('intp')
- not_found = (-1 * np.ones(1000)).astype('intp')
-
- for leaf_size in [1, 10, 100, 10000]:
- for closed in ['left', 'right', 'both', 'neither']:
- tree = IntervalTree(x, x + 0.5, closed=closed,
- leaf_size=leaf_size)
- tm.assert_numpy_array_equal(found,
- tree.get_indexer(x + 0.25))
-
- expected = found if tree.closed_left else not_found
- tm.assert_numpy_array_equal(expected,
- tree.get_indexer(x + 0.0))
-
- expected = found if tree.closed_right else not_found
- tm.assert_numpy_array_equal(expected,
- tree.get_indexer(x + 0.5))
diff --git a/pandas/tests/indexes/interval/test_interval_range.py b/pandas/tests/indexes/interval/test_interval_range.py
new file mode 100644
index 0000000000000..203e8e3128edc
--- /dev/null
+++ b/pandas/tests/indexes/interval/test_interval_range.py
@@ -0,0 +1,301 @@
+from __future__ import division
+
+import pytest
+import numpy as np
+from datetime import timedelta
+from pandas import (
+ Interval, IntervalIndex, Timestamp, Timedelta, DateOffset,
+ interval_range, date_range, timedelta_range)
+from pandas.tseries.offsets import Day
+import pandas.util.testing as tm
+import pandas as pd
+
+
+@pytest.fixture(scope='class', params=['left', 'right', 'both', 'neither'])
+def closed(request):
+ return request.param
+
+
+@pytest.fixture(scope='class', params=[None, 'foo'])
+def name(request):
+ return request.param
+
+
+class TestIntervalRange(object):
+
+ def test_construction_from_numeric(self, closed, name):
+ # combinations of start/end/periods without freq
+ expected = IntervalIndex.from_breaks(
+ np.arange(0, 6), name=name, closed=closed)
+
+ result = interval_range(start=0, end=5, name=name, closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=0, periods=5, name=name, closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=5, periods=5, name=name, closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # combinations of start/end/periods with freq
+ expected = IntervalIndex.from_tuples([(0, 2), (2, 4), (4, 6)],
+ name=name, closed=closed)
+
+ result = interval_range(start=0, end=6, freq=2, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=0, periods=3, freq=2, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=6, periods=3, freq=2, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # output truncates early if freq causes end to be skipped.
+ expected = IntervalIndex.from_tuples([(0.0, 1.5), (1.5, 3.0)],
+ name=name, closed=closed)
+ result = interval_range(start=0, end=4, freq=1.5, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ @pytest.mark.parametrize('tz', [None, 'US/Eastern'])
+ def test_construction_from_timestamp(self, closed, name, tz):
+ # combinations of start/end/periods without freq
+ start = Timestamp('2017-01-01', tz=tz)
+ end = Timestamp('2017-01-06', tz=tz)
+ breaks = date_range(start=start, end=end)
+ expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
+
+ result = interval_range(start=start, end=end, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=start, periods=5, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=end, periods=5, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # combinations of start/end/periods with fixed freq
+ freq = '2D'
+ start = Timestamp('2017-01-01', tz=tz)
+ end = Timestamp('2017-01-07', tz=tz)
+ breaks = date_range(start=start, end=end, freq=freq)
+ expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
+
+ result = interval_range(start=start, end=end, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=start, periods=3, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=end, periods=3, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # output truncates early if freq causes end to be skipped.
+ end = Timestamp('2017-01-08', tz=tz)
+ result = interval_range(start=start, end=end, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # combinations of start/end/periods with non-fixed freq
+ freq = 'M'
+ start = Timestamp('2017-01-01', tz=tz)
+ end = Timestamp('2017-12-31', tz=tz)
+ breaks = date_range(start=start, end=end, freq=freq)
+ expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
+
+ result = interval_range(start=start, end=end, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=start, periods=11, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=end, periods=11, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # output truncates early if freq causes end to be skipped.
+ end = Timestamp('2018-01-15', tz=tz)
+ result = interval_range(start=start, end=end, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ def test_construction_from_timedelta(self, closed, name):
+ # combinations of start/end/periods without freq
+ start, end = Timedelta('1 day'), Timedelta('6 days')
+ breaks = timedelta_range(start=start, end=end)
+ expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
+
+ result = interval_range(start=start, end=end, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=start, periods=5, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=end, periods=5, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # combinations of start/end/periods with fixed freq
+ freq = '2D'
+ start, end = Timedelta('1 day'), Timedelta('7 days')
+ breaks = timedelta_range(start=start, end=end, freq=freq)
+ expected = IntervalIndex.from_breaks(breaks, name=name, closed=closed)
+
+ result = interval_range(start=start, end=end, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(start=start, periods=3, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ result = interval_range(end=end, periods=3, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ # output truncates early if freq causes end to be skipped.
+ end = Timedelta('7 days 1 hour')
+ result = interval_range(start=start, end=end, freq=freq, name=name,
+ closed=closed)
+ tm.assert_index_equal(result, expected)
+
+ def test_constructor_coverage(self):
+ # float value for periods
+ expected = pd.interval_range(start=0, periods=10)
+ result = pd.interval_range(start=0, periods=10.5)
+ tm.assert_index_equal(result, expected)
+
+ # equivalent timestamp-like start/end
+ start, end = Timestamp('2017-01-01'), Timestamp('2017-01-15')
+ expected = pd.interval_range(start=start, end=end)
+
+ result = pd.interval_range(start=start.to_pydatetime(),
+ end=end.to_pydatetime())
+ tm.assert_index_equal(result, expected)
+
+ result = pd.interval_range(start=start.asm8, end=end.asm8)
+ tm.assert_index_equal(result, expected)
+
+ # equivalent freq with timestamp
+ equiv_freq = ['D', Day(), Timedelta(days=1), timedelta(days=1),
+ DateOffset(days=1)]
+ for freq in equiv_freq:
+ result = pd.interval_range(start=start, end=end, freq=freq)
+ tm.assert_index_equal(result, expected)
+
+ # equivalent timedelta-like start/end
+ start, end = Timedelta(days=1), Timedelta(days=10)
+ expected = pd.interval_range(start=start, end=end)
+
+ result = pd.interval_range(start=start.to_pytimedelta(),
+ end=end.to_pytimedelta())
+ tm.assert_index_equal(result, expected)
+
+ result = pd.interval_range(start=start.asm8, end=end.asm8)
+ tm.assert_index_equal(result, expected)
+
+ # equivalent freq with timedelta
+ equiv_freq = ['D', Day(), Timedelta(days=1), timedelta(days=1)]
+ for freq in equiv_freq:
+ result = pd.interval_range(start=start, end=end, freq=freq)
+ tm.assert_index_equal(result, expected)
+
+ def test_errors(self):
+ # not enough params
+ msg = ('Of the three parameters: start, end, and periods, '
+ 'exactly two must be specified')
+
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(start=0)
+
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(end=5)
+
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(periods=2)
+
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range()
+
+ # too many params
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(start=0, end=5, periods=6)
+
+ # mixed units
+ msg = 'start, end, freq need to be type compatible'
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=0, end=Timestamp('20130101'), freq=2)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=0, end=Timedelta('1 day'), freq=2)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=0, end=10, freq='D')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=Timestamp('20130101'), end=10, freq='D')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=Timestamp('20130101'),
+ end=Timedelta('1 day'), freq='D')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=Timestamp('20130101'),
+ end=Timestamp('20130110'), freq=2)
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=Timedelta('1 day'), end=10, freq='D')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=Timedelta('1 day'),
+ end=Timestamp('20130110'), freq='D')
+
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=Timedelta('1 day'),
+ end=Timedelta('10 days'), freq=2)
+
+ # invalid periods
+ msg = 'periods must be a number, got foo'
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=0, periods='foo')
+
+ # invalid start
+ msg = 'start must be numeric or datetime-like, got foo'
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(start='foo', periods=10)
+
+ # invalid end
+ msg = r'end must be numeric or datetime-like, got \(0, 1\]'
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(end=Interval(0, 1), periods=10)
+
+ # invalid freq for datetime-like
+ msg = 'freq must be numeric or convertible to DateOffset, got foo'
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(start=0, end=10, freq='foo')
+
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(start=Timestamp('20130101'), periods=10, freq='foo')
+
+ with tm.assert_raises_regex(ValueError, msg):
+ interval_range(end=Timedelta('1 day'), periods=10, freq='foo')
+
+ # mixed tz
+ start = Timestamp('2017-01-01', tz='US/Eastern')
+ end = Timestamp('2017-01-07', tz='US/Pacific')
+ msg = 'Start and end cannot both be tz-aware with different timezones'
+ with tm.assert_raises_regex(TypeError, msg):
+ interval_range(start=start, end=end)
diff --git a/pandas/tests/indexes/interval/test_interval_tree.py b/pandas/tests/indexes/interval/test_interval_tree.py
new file mode 100644
index 0000000000000..343131125f640
--- /dev/null
+++ b/pandas/tests/indexes/interval/test_interval_tree.py
@@ -0,0 +1,93 @@
+from __future__ import division
+
+import pytest
+import numpy as np
+from pandas import compat
+from pandas._libs.interval import IntervalTree
+import pandas.util.testing as tm
+
+
+@pytest.fixture(scope='class', params=['left', 'right', 'both', 'neither'])
+def closed(request):
+ return request.param
+
+
+class TestIntervalTree(object):
+ def setup_method(self, method):
+ def gentree(dtype):
+ left = np.arange(5, dtype=dtype)
+ right = left + 2
+ return IntervalTree(left, right)
+
+ self.tree = gentree('int64')
+ self.trees = {dtype: gentree(dtype)
+ for dtype in ['int32', 'int64', 'float32', 'float64']}
+
+ def test_get_loc(self):
+ for dtype, tree in self.trees.items():
+ tm.assert_numpy_array_equal(tree.get_loc(1),
+ np.array([0], dtype='int64'))
+ tm.assert_numpy_array_equal(np.sort(tree.get_loc(2)),
+ np.array([0, 1], dtype='int64'))
+ with pytest.raises(KeyError):
+ tree.get_loc(-1)
+
+ def test_get_indexer(self):
+ for dtype, tree in self.trees.items():
+ tm.assert_numpy_array_equal(
+ tree.get_indexer(np.array([1.0, 5.5, 6.5])),
+ np.array([0, 4, -1], dtype='int64'))
+ with pytest.raises(KeyError):
+ tree.get_indexer(np.array([3.0]))
+
+ def test_get_indexer_non_unique(self):
+ indexer, missing = self.tree.get_indexer_non_unique(
+ np.array([1.0, 2.0, 6.5]))
+ tm.assert_numpy_array_equal(indexer[:1],
+ np.array([0], dtype='int64'))
+ tm.assert_numpy_array_equal(np.sort(indexer[1:3]),
+ np.array([0, 1], dtype='int64'))
+ tm.assert_numpy_array_equal(np.sort(indexer[3:]),
+ np.array([-1], dtype='int64'))
+ tm.assert_numpy_array_equal(missing, np.array([2], dtype='int64'))
+
+ def test_duplicates(self):
+ tree = IntervalTree([0, 0, 0], [1, 1, 1])
+ tm.assert_numpy_array_equal(np.sort(tree.get_loc(0.5)),
+ np.array([0, 1, 2], dtype='int64'))
+
+ with pytest.raises(KeyError):
+ tree.get_indexer(np.array([0.5]))
+
+ indexer, missing = tree.get_indexer_non_unique(np.array([0.5]))
+ tm.assert_numpy_array_equal(np.sort(indexer),
+ np.array([0, 1, 2], dtype='int64'))
+ tm.assert_numpy_array_equal(missing, np.array([], dtype='int64'))
+
+ def test_get_loc_closed(self, closed):
+ tree = IntervalTree([0], [1], closed=closed)
+ for p, errors in [(0, tree.open_left),
+ (1, tree.open_right)]:
+ if errors:
+ with pytest.raises(KeyError):
+ tree.get_loc(p)
+ else:
+ tm.assert_numpy_array_equal(tree.get_loc(p),
+ np.array([0], dtype='int64'))
+
+ @pytest.mark.skipif(compat.is_platform_32bit(),
+ reason="int type mismatch on 32bit")
+ @pytest.mark.parametrize('leaf_size', [1, 10, 100, 10000])
+ def test_get_indexer_closed(self, closed, leaf_size):
+ x = np.arange(1000, dtype='float64')
+ found = x.astype('intp')
+ not_found = (-1 * np.ones(1000)).astype('intp')
+
+ tree = IntervalTree(x, x + 0.5, closed=closed, leaf_size=leaf_size)
+ tm.assert_numpy_array_equal(found, tree.get_indexer(x + 0.25))
+
+ expected = found if tree.closed_left else not_found
+ tm.assert_numpy_array_equal(expected, tree.get_indexer(x + 0.0))
+
+ expected = found if tree.closed_right else not_found
+ tm.assert_numpy_array_equal(expected, tree.get_indexer(x + 0.5))
| - [X] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Since we now have an interval subdirectory, seems logical to split `test_interval.py` into separate files, similar to what has been done for `DatetimeIndex`, `PeriodIndex`, and `TimedeltaIndex`.
Just made very basic changes here, moving things at the class level, and didn't break apart any classes. All of the tests should functionally be the same.
Summary:
- Moved `TestIntervalRange` class from `test_interval.py` to `test_interval_range.py`
- Moved `TestIntervalTree` class from `test_interval.py` to `test_interval_tree.py`
- Changed a few `for` loops to `@pytest.mark.parametrize`
- Converted a `lambda` to an actual function (my linter was complaining about PEP8 E731)
- No changes to `test_interval_new.py`
- Should still only cover tests in `test_interval.py`
Down the road we could probably split `test_interval.py` into smaller components, much like what was done for `DatetimeIndex`, `PeriodIndex`, and `TimedeltaIndex`. Might want to wait until the `_new.py` files are fully addressed though. Didn't look like there were any high level changes like this that could be made in `tests/indexing/interval/`.
| https://api.github.com/repos/pandas-dev/pandas/pulls/19009 | 2017-12-30T22:37:00Z | 2017-12-30T23:49:23Z | 2017-12-30T23:49:23Z | 2017-12-31T20:47:18Z |
CI: move 3.5 build back to required on travis | diff --git a/.travis.yml b/.travis.yml
index e56435faeec19..5cc6547968b7d 100644
--- a/.travis.yml
+++ b/.travis.yml
@@ -49,7 +49,6 @@ matrix:
apt:
packages:
- python-gtk2
- # In allow_failures
- dist: trusty
env:
- JOB="3.5_CONDA_BUILD_TEST" TEST_ARGS="--skip-slow --skip-network" CONDA_BUILD_TEST=true
@@ -77,10 +76,6 @@ matrix:
env:
- JOB="3.6_DOC" DOC=true
allow_failures:
- # TODO(jreback)
- - dist: trusty
- env:
- - JOB="3.5_CONDA_BUILD_TEST" TEST_ARGS="--skip-slow --skip-network" CONDA_BUILD_TEST=true
- dist: trusty
env:
- JOB="2.7_SLOW" SLOW=true
| https://api.github.com/repos/pandas-dev/pandas/pulls/19007 | 2017-12-30T22:23:18Z | 2017-12-30T23:20:59Z | 2017-12-30T23:20:59Z | 2017-12-30T23:20:59Z | |
COMPAT: Drop reference to deprecated dateutil.zoneinfo.gettz | diff --git a/pandas/_libs/tslibs/timezones.pyx b/pandas/_libs/tslibs/timezones.pyx
index de9f75344b2bf..fdcf40337fab9 100644
--- a/pandas/_libs/tslibs/timezones.pyx
+++ b/pandas/_libs/tslibs/timezones.pyx
@@ -10,13 +10,7 @@ from dateutil.tz import (
tzlocal as _dateutil_tzlocal,
tzfile as _dateutil_tzfile)
-import sys
-if sys.platform == 'win32' or sys.platform == 'cygwin':
- # equiv pd.compat.is_platform_windows()
- from dateutil.zoneinfo import gettz as dateutil_gettz
-else:
- from dateutil.tz import gettz as dateutil_gettz
-
+from dateutil.tz import gettz as dateutil_gettz
from pytz.tzinfo import BaseTzInfo as _pytz_BaseTzInfo
import pytz
| - [X] closes #19004
I'm assuming these will be handled by CI:
- [ ] tests added / passed
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
Not sure this is necessary, it's a very "behind the scenes" fix, replacing `zoneinfo.gettz` with the superior `tz.gettz()`:
- [ ] whatsnew entry
| https://api.github.com/repos/pandas-dev/pandas/pulls/19006 | 2017-12-30T20:02:30Z | 2017-12-30T22:12:05Z | 2017-12-30T22:12:05Z | 2017-12-30T22:14:38Z |
DOC: Fix min_count docstring | diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index 06d82578cb9ef..84799d12df0c4 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -7664,7 +7664,7 @@ def _doc_parms(cls):
_min_count_stub = """\
-min_count : int, default 1
+min_count : int, default 0
The required number of valid values to perform the operation. If fewer than
``min_count`` non-NA values are present the result will be NA.
| [ci skip]
missed this earlier. | https://api.github.com/repos/pandas-dev/pandas/pulls/19005 | 2017-12-30T19:55:04Z | 2017-12-30T19:55:37Z | 2017-12-30T19:55:37Z | 2017-12-30T19:55:47Z |
COMPAT: clean up warnings | diff --git a/appveyor.yml b/appveyor.yml
index 0aaac322c4ac7..ba001208864a8 100644
--- a/appveyor.yml
+++ b/appveyor.yml
@@ -15,6 +15,7 @@ environment:
# See: http://stackoverflow.com/a/13751649/163740
CMD_IN_ENV: "cmd /E:ON /V:ON /C .\\ci\\run_with_env.cmd"
clone_folder: C:\projects\pandas
+ PANDAS_TESTING_MODE: "deprecate"
matrix:
diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 92564285bb36a..6407a33c442d0 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -207,6 +207,7 @@ Other API Changes
- :func:`wide_to_long` previously kept numeric-like suffixes as ``object`` dtype. Now they are cast to numeric if possible (:issue:`17627`)
- In :func:`read_excel`, the ``comment`` argument is now exposed as a named parameter (:issue:`18735`)
- Rearranged the order of keyword arguments in :func:`read_excel()` to align with :func:`read_csv()` (:issue:`16672`)
+- The options ``html.border`` and ``mode.use_inf_as_null`` were deprecated in prior versions, these will now show ``FutureWarning`` rather than a ``DeprecationWarning`` (:issue:`19003`)
.. _whatsnew_0230.deprecations:
diff --git a/pandas/core/config.py b/pandas/core/config.py
index d10e2d19be665..692aed178719d 100644
--- a/pandas/core/config.py
+++ b/pandas/core/config.py
@@ -613,7 +613,7 @@ def _warn_if_deprecated(key):
if d:
if d.msg:
print(d.msg)
- warnings.warn(d.msg, DeprecationWarning)
+ warnings.warn(d.msg, FutureWarning)
else:
msg = "'{key}' is deprecated".format(key=key)
if d.removal_ver:
@@ -624,7 +624,7 @@ def _warn_if_deprecated(key):
else:
msg += ', please refrain from using it.'
- warnings.warn(msg, DeprecationWarning)
+ warnings.warn(msg, FutureWarning)
return True
return False
diff --git a/pandas/core/dtypes/missing.py b/pandas/core/dtypes/missing.py
index d208c72ffee19..ffac702476af1 100644
--- a/pandas/core/dtypes/missing.py
+++ b/pandas/core/dtypes/missing.py
@@ -316,6 +316,10 @@ def array_equivalent(left, right, strict_nan=False):
# NaNs can occur in float and complex arrays.
if is_float_dtype(left) or is_complex_dtype(left):
+
+ # empty
+ if not (np.prod(left.shape) and np.prod(right.shape)):
+ return True
return ((left == right) | (isna(left) & isna(right))).all()
# numpy will will not allow this type of datetimelike vs integer comparison
diff --git a/pandas/tests/frame/test_constructors.py b/pandas/tests/frame/test_constructors.py
index 8be6c4875ae24..b7d3a60ecf6e4 100644
--- a/pandas/tests/frame/test_constructors.py
+++ b/pandas/tests/frame/test_constructors.py
@@ -328,13 +328,13 @@ def test_constructor_error_msgs(self):
# wrong size axis labels
with tm.assert_raises_regex(ValueError, "Shape of passed values "
- "is \(3, 2\), indices "
- "imply \(3, 1\)"):
+ r"is \(3, 2\), indices "
+ r"imply \(3, 1\)"):
DataFrame(np.random.rand(2, 3), columns=['A', 'B', 'C'], index=[1])
with tm.assert_raises_regex(ValueError, "Shape of passed values "
- "is \(3, 2\), indices "
- "imply \(2, 2\)"):
+ r"is \(3, 2\), indices "
+ r"imply \(2, 2\)"):
DataFrame(np.random.rand(2, 3), columns=['A', 'B'], index=[1, 2])
with tm.assert_raises_regex(ValueError, "If using all scalar "
@@ -1220,12 +1220,12 @@ def test_constructor_from_items(self):
def test_constructor_from_items_scalars(self):
# GH 17312
with tm.assert_raises_regex(ValueError,
- 'The value in each \(key, value\) '
+ r'The value in each \(key, value\) '
'pair must be an array, Series, or dict'):
DataFrame.from_items([('A', 1), ('B', 4)])
with tm.assert_raises_regex(ValueError,
- 'The value in each \(key, value\) '
+ r'The value in each \(key, value\) '
'pair must be an array, Series, or dict'):
DataFrame.from_items([('A', 1), ('B', 2)], columns=['col1'],
orient='index')
diff --git a/pandas/tests/frame/test_query_eval.py b/pandas/tests/frame/test_query_eval.py
index 22066d59cf14d..55aeaf6e77be1 100644
--- a/pandas/tests/frame/test_query_eval.py
+++ b/pandas/tests/frame/test_query_eval.py
@@ -1040,6 +1040,6 @@ def test_invalid_type_for_operator_raises(self, parser, engine):
ops = '+', '-', '*', '/'
for op in ops:
with tm.assert_raises_regex(TypeError,
- "unsupported operand type\(s\) "
+ r"unsupported operand type\(s\) "
"for .+: '.+' and '.+'"):
df.eval('a {0} b'.format(op), engine=engine, parser=parser)
diff --git a/pandas/tests/groupby/test_nth.py b/pandas/tests/groupby/test_nth.py
index 2a408b85f0ed1..ccde545b5b8e9 100644
--- a/pandas/tests/groupby/test_nth.py
+++ b/pandas/tests/groupby/test_nth.py
@@ -175,7 +175,7 @@ def test_nth(self):
df = DataFrame([[1, np.nan], [1, 4], [5, 6]], columns=['A', 'B'])
g = df.groupby('A')
# PR 17493, related to issue 11038
- # test Series.nth with True for dropna produces DeprecationWarning
+ # test Series.nth with True for dropna produces FutureWarning
with assert_produces_warning(FutureWarning):
result = g.B.nth(0, dropna=True)
expected = g.B.first()
diff --git a/pandas/tests/groupby/test_transform.py b/pandas/tests/groupby/test_transform.py
index c0ea968ab0819..8f72da293a50c 100644
--- a/pandas/tests/groupby/test_transform.py
+++ b/pandas/tests/groupby/test_transform.py
@@ -261,7 +261,7 @@ def test_transform_casting(self):
9 B-053 b76cd912ff "2014-10-08 19:17:48"
10 B-065 b76cd912ff "2014-10-08 19:21:38"
"""
- df = pd.read_csv(StringIO(data), sep='\s+',
+ df = pd.read_csv(StringIO(data), sep=r'\s+',
index_col=[0], parse_dates=['DATETIME'])
result = df.groupby('ID3')['DATETIME'].transform(lambda x: x.diff())
diff --git a/pandas/tests/indexes/datetimes/test_date_range.py b/pandas/tests/indexes/datetimes/test_date_range.py
index 3ce51983c111d..3738398d017f8 100644
--- a/pandas/tests/indexes/datetimes/test_date_range.py
+++ b/pandas/tests/indexes/datetimes/test_date_range.py
@@ -402,7 +402,7 @@ def test_daterange_bug_456(self):
assert isinstance(result, DatetimeIndex)
def test_error_with_zero_monthends(self):
- msg = 'Offset <0 \* MonthEnds> did not increment date'
+ msg = r'Offset <0 \* MonthEnds> did not increment date'
with tm.assert_raises_regex(ValueError, msg):
date_range('1/1/2000', '1/1/2001', freq=MonthEnd(0))
diff --git a/pandas/tests/indexes/datetimes/test_tools.py b/pandas/tests/indexes/datetimes/test_tools.py
index f94a438fcdaa5..44f3c21d23e62 100644
--- a/pandas/tests/indexes/datetimes/test_tools.py
+++ b/pandas/tests/indexes/datetimes/test_tools.py
@@ -709,7 +709,7 @@ def test_dataframe(self, cache):
'day': [4, 5]})
msg = ("cannot assemble the datetimes: time data .+ does not "
- "match format '%Y%m%d' \(match\)")
+ r"match format '%Y%m%d' \(match\)")
with tm.assert_raises_regex(ValueError, msg):
to_datetime(df2, cache=cache)
result = to_datetime(df2, errors='coerce', cache=cache)
@@ -719,15 +719,15 @@ def test_dataframe(self, cache):
# extra columns
msg = ("extra keys have been passed to the datetime assemblage: "
- "\[foo\]")
+ r"\[foo\]")
with tm.assert_raises_regex(ValueError, msg):
df2 = df.copy()
df2['foo'] = 1
to_datetime(df2, cache=cache)
# not enough
- msg = ('to assemble mappings requires at least that \[year, month, '
- 'day\] be specified: \[.+\] is missing')
+ msg = (r'to assemble mappings requires at least that \[year, month, '
+ r'day\] be specified: \[.+\] is missing')
for c in [['year'],
['year', 'month'],
['year', 'month', 'second'],
diff --git a/pandas/tests/indexes/interval/test_interval.py b/pandas/tests/indexes/interval/test_interval.py
index 3ca4c31b7f059..4805c957907e6 100644
--- a/pandas/tests/indexes/interval/test_interval.py
+++ b/pandas/tests/indexes/interval/test_interval.py
@@ -175,7 +175,7 @@ def test_constructors_empty(self, data, closed):
def test_constructors_errors(self):
# scalar
- msg = ('IntervalIndex\(...\) must be called with a collection of '
+ msg = (r'IntervalIndex\(...\) must be called with a collection of '
'some kind, 5 was passed')
with tm.assert_raises_regex(TypeError, msg):
IntervalIndex(5)
diff --git a/pandas/tests/indexing/test_multiindex.py b/pandas/tests/indexing/test_multiindex.py
index d89c64fc5b9f8..f69b9d98143b0 100644
--- a/pandas/tests/indexing/test_multiindex.py
+++ b/pandas/tests/indexing/test_multiindex.py
@@ -299,7 +299,7 @@ def test_getitem_partial_int(self):
# missing item:
with tm.assert_raises_regex(KeyError, '1'):
df[1]
- with tm.assert_raises_regex(KeyError, "'\[1\] not in index'"):
+ with tm.assert_raises_regex(KeyError, r"'\[1\] not in index'"):
df[[1]]
def test_loc_multiindex_indexer_none(self):
diff --git a/pandas/tests/io/formats/test_to_html.py b/pandas/tests/io/formats/test_to_html.py
index b263d368f41f5..9e063c2d176e1 100644
--- a/pandas/tests/io/formats/test_to_html.py
+++ b/pandas/tests/io/formats/test_to_html.py
@@ -1411,8 +1411,9 @@ def test_to_html_border_zero(self):
result = df.to_html(border=0)
assert 'border="0"' in result
+ @tm.capture_stdout
def test_display_option_warning(self):
- with tm.assert_produces_warning(DeprecationWarning,
+ with tm.assert_produces_warning(FutureWarning,
check_stacklevel=False):
pd.options.html.border
diff --git a/pandas/tests/io/formats/test_to_latex.py b/pandas/tests/io/formats/test_to_latex.py
index 2f8ef32722051..f266a8b3a3268 100644
--- a/pandas/tests/io/formats/test_to_latex.py
+++ b/pandas/tests/io/formats/test_to_latex.py
@@ -424,11 +424,11 @@ def test_to_latex_longtable(self, frame):
df = DataFrame({'a': [1, 2]})
with1column_result = df.to_latex(index=False, longtable=True)
- assert "\multicolumn{1}" in with1column_result
+ assert r"\multicolumn{1}" in with1column_result
df = DataFrame({'a': [1, 2], 'b': [3, 4], 'c': [5, 6]})
with3columns_result = df.to_latex(index=False, longtable=True)
- assert "\multicolumn{3}" in with3columns_result
+ assert r"\multicolumn{3}" in with3columns_result
def test_to_latex_escape_special_chars(self):
special_characters = ['&', '%', '$', '#', '_', '{', '}', '~', '^',
diff --git a/pandas/tests/io/test_parquet.py b/pandas/tests/io/test_parquet.py
index c59acbd946f91..31c2ded49b7a0 100644
--- a/pandas/tests/io/test_parquet.py
+++ b/pandas/tests/io/test_parquet.py
@@ -7,7 +7,7 @@
import numpy as np
import pandas as pd
-from pandas.compat import PY3, is_platform_windows
+from pandas.compat import PY3, is_platform_windows, is_platform_mac
from pandas.io.parquet import (to_parquet, read_parquet, get_engine,
PyArrowImpl, FastParquetImpl)
from pandas.util import testing as tm
@@ -174,8 +174,8 @@ def test_options_get_engine(fp, pa):
assert isinstance(get_engine('fastparquet'), FastParquetImpl)
-@pytest.mark.xfail(is_platform_windows(),
- reason="reading pa metadata failing on Windows")
+@pytest.mark.xfail(is_platform_windows() or is_platform_mac(),
+ reason="reading pa metadata failing on Windows/mac")
def test_cross_engine_pa_fp(df_cross_compat, pa, fp):
# cross-compat with differing reading/writing engines
diff --git a/pandas/tests/plotting/test_misc.py b/pandas/tests/plotting/test_misc.py
index 60ed280bc050e..9e538ae130a85 100644
--- a/pandas/tests/plotting/test_misc.py
+++ b/pandas/tests/plotting/test_misc.py
@@ -206,6 +206,7 @@ def test_parallel_coordinates(self):
def test_parallel_coordinates_with_sorted_labels(self):
""" For #15908 """
from pandas.plotting import parallel_coordinates
+
df = DataFrame({"feat": [i for i in range(30)],
"class": [2 for _ in range(10)] +
[3 for _ in range(10)] +
diff --git a/pandas/tests/reshape/merge/test_merge_asof.py b/pandas/tests/reshape/merge/test_merge_asof.py
index 4b2680b9be592..2f48aef1894a9 100644
--- a/pandas/tests/reshape/merge/test_merge_asof.py
+++ b/pandas/tests/reshape/merge/test_merge_asof.py
@@ -976,7 +976,7 @@ def test_on_float_by_int(self):
def test_merge_datatype_error(self):
""" Tests merge datatype mismatch error """
- msg = 'merge keys \[0\] object and int64, must be the same type'
+ msg = r'merge keys \[0\] object and int64, must be the same type'
left = pd.DataFrame({'left_val': [1, 5, 10],
'a': ['a', 'b', 'c']})
diff --git a/pandas/tests/reshape/test_concat.py b/pandas/tests/reshape/test_concat.py
index bdbf2a0ee2f68..f66cb12b11210 100644
--- a/pandas/tests/reshape/test_concat.py
+++ b/pandas/tests/reshape/test_concat.py
@@ -178,9 +178,9 @@ def test_concatlike_same_dtypes(self):
tm.assert_series_equal(res, exp, check_index_type=True)
# cannot append non-index
- msg = ('cannot concatenate object of type \"(.+?)\";'
+ msg = (r'cannot concatenate object of type \"(.+?)\";'
' only pd.Series, pd.DataFrame, and pd.Panel'
- ' \(deprecated\) objs are valid')
+ r' \(deprecated\) objs are valid')
with tm.assert_raises_regex(TypeError, msg):
pd.Series(vals1).append(vals2)
diff --git a/pandas/tests/scalar/test_interval.py b/pandas/tests/scalar/test_interval.py
index 3db474e32c4dd..23dad9736dac5 100644
--- a/pandas/tests/scalar/test_interval.py
+++ b/pandas/tests/scalar/test_interval.py
@@ -122,7 +122,7 @@ def test_math_add(self, interval):
actual += 1
assert expected == actual
- msg = "unsupported operand type\(s\) for \+"
+ msg = r"unsupported operand type\(s\) for \+"
with tm.assert_raises_regex(TypeError, msg):
interval + Interval(1, 2)
@@ -138,7 +138,7 @@ def test_math_sub(self, interval):
actual -= 1
assert expected == actual
- msg = "unsupported operand type\(s\) for -"
+ msg = r"unsupported operand type\(s\) for -"
with tm.assert_raises_regex(TypeError, msg):
interval - Interval(1, 2)
@@ -158,11 +158,11 @@ def test_math_mult(self, interval):
actual *= 2
assert expected == actual
- msg = "unsupported operand type\(s\) for \*"
+ msg = r"unsupported operand type\(s\) for \*"
with tm.assert_raises_regex(TypeError, msg):
interval * Interval(1, 2)
- msg = "can\'t multiply sequence by non-int"
+ msg = r"can\'t multiply sequence by non-int"
with tm.assert_raises_regex(TypeError, msg):
interval * 'foo'
@@ -175,7 +175,7 @@ def test_math_div(self, interval):
actual /= 2.0
assert expected == actual
- msg = "unsupported operand type\(s\) for /"
+ msg = r"unsupported operand type\(s\) for /"
with tm.assert_raises_regex(TypeError, msg):
interval / Interval(1, 2)
diff --git a/pandas/tests/series/test_missing.py b/pandas/tests/series/test_missing.py
index 8c6a4fcf4b1d4..0dc5e23184af7 100644
--- a/pandas/tests/series/test_missing.py
+++ b/pandas/tests/series/test_missing.py
@@ -480,12 +480,9 @@ def test_isna_for_inf(self):
def test_isnull_for_inf_deprecated(self):
# gh-17115
s = Series(['a', np.inf, np.nan, 1.0])
- with tm.assert_produces_warning(DeprecationWarning,
- check_stacklevel=False):
- pd.set_option('mode.use_inf_as_null', True)
+ with pd.option_context('mode.use_inf_as_null', True):
r = s.isna()
dr = s.dropna()
- pd.reset_option('mode.use_inf_as_null')
e = Series([False, True, True, False])
de = Series(['a', 1.0], index=[0, 3])
diff --git a/pandas/tests/sparse/frame/test_frame.py b/pandas/tests/sparse/frame/test_frame.py
index cf002ff046c2e..058892e3b85ff 100644
--- a/pandas/tests/sparse/frame/test_frame.py
+++ b/pandas/tests/sparse/frame/test_frame.py
@@ -7,21 +7,14 @@
from numpy import nan
import numpy as np
import pandas as pd
-from distutils.version import LooseVersion
from pandas import Series, DataFrame, bdate_range, Panel
-from pandas.core.dtypes.common import (
- is_bool_dtype,
- is_float_dtype,
- is_object_dtype,
- is_float)
from pandas.core.indexes.datetimes import DatetimeIndex
from pandas.tseries.offsets import BDay
from pandas.util import testing as tm
from pandas.compat import lrange
from pandas import compat
from pandas.core.sparse import frame as spf
-import pandas.util._test_decorators as td
from pandas._libs.sparse import BlockIndex, IntIndex
from pandas.core.sparse.api import SparseSeries, SparseDataFrame, SparseArray
@@ -1171,163 +1164,6 @@ def test_notna(self):
tm.assert_frame_equal(res.to_dense(), exp)
-@td.skip_if_no_scipy
-@pytest.mark.parametrize('index', [None, list('abc')]) # noqa: F811
-@pytest.mark.parametrize('columns', [None, list('def')])
-@pytest.mark.parametrize('fill_value', [None, 0, np.nan])
-@pytest.mark.parametrize('dtype', [bool, int, float, np.uint16])
-def test_from_to_scipy(spmatrix, index, columns, fill_value, dtype):
- # GH 4343
- # Make one ndarray and from it one sparse matrix, both to be used for
- # constructing frames and comparing results
- arr = np.eye(3, dtype=dtype)
- # GH 16179
- arr[0, 1] = dtype(2)
- try:
- spm = spmatrix(arr)
- assert spm.dtype == arr.dtype
- except (TypeError, AssertionError):
- # If conversion to sparse fails for this spmatrix type and arr.dtype,
- # then the combination is not currently supported in NumPy, so we
- # can just skip testing it thoroughly
- return
-
- sdf = pd.SparseDataFrame(spm, index=index, columns=columns,
- default_fill_value=fill_value)
-
- # Expected result construction is kind of tricky for all
- # dtype-fill_value combinations; easiest to cast to something generic
- # and except later on
- rarr = arr.astype(object)
- rarr[arr == 0] = np.nan
- expected = pd.SparseDataFrame(rarr, index=index, columns=columns).fillna(
- fill_value if fill_value is not None else np.nan)
-
- # Assert frame is as expected
- sdf_obj = sdf.astype(object)
- tm.assert_sp_frame_equal(sdf_obj, expected)
- tm.assert_frame_equal(sdf_obj.to_dense(), expected.to_dense())
-
- # Assert spmatrices equal
- assert dict(sdf.to_coo().todok()) == dict(spm.todok())
-
- # Ensure dtype is preserved if possible
- was_upcast = ((fill_value is None or is_float(fill_value)) and
- not is_object_dtype(dtype) and
- not is_float_dtype(dtype))
- res_dtype = (bool if is_bool_dtype(dtype) else
- float if was_upcast else
- dtype)
- tm.assert_contains_all(sdf.dtypes, {np.dtype(res_dtype)})
- assert sdf.to_coo().dtype == res_dtype
-
- # However, adding a str column results in an upcast to object
- sdf['strings'] = np.arange(len(sdf)).astype(str)
- assert sdf.to_coo().dtype == np.object_
-
-
-@td.skip_if_no_scipy
-@pytest.mark.parametrize('fill_value', [None, 0, np.nan]) # noqa: F811
-def test_from_to_scipy_object(spmatrix, fill_value):
- # GH 4343
- dtype = object
- columns = list('cd')
- index = list('ab')
- import scipy
- if (spmatrix is scipy.sparse.dok_matrix and LooseVersion(
- scipy.__version__) >= LooseVersion('0.19.0')):
- pytest.skip("dok_matrix from object does not work in SciPy >= 0.19")
-
- # Make one ndarray and from it one sparse matrix, both to be used for
- # constructing frames and comparing results
- arr = np.eye(2, dtype=dtype)
- try:
- spm = spmatrix(arr)
- assert spm.dtype == arr.dtype
- except (TypeError, AssertionError):
- # If conversion to sparse fails for this spmatrix type and arr.dtype,
- # then the combination is not currently supported in NumPy, so we
- # can just skip testing it thoroughly
- return
-
- sdf = pd.SparseDataFrame(spm, index=index, columns=columns,
- default_fill_value=fill_value)
-
- # Expected result construction is kind of tricky for all
- # dtype-fill_value combinations; easiest to cast to something generic
- # and except later on
- rarr = arr.astype(object)
- rarr[arr == 0] = np.nan
- expected = pd.SparseDataFrame(rarr, index=index, columns=columns).fillna(
- fill_value if fill_value is not None else np.nan)
-
- # Assert frame is as expected
- sdf_obj = sdf.astype(object)
- tm.assert_sp_frame_equal(sdf_obj, expected)
- tm.assert_frame_equal(sdf_obj.to_dense(), expected.to_dense())
-
- # Assert spmatrices equal
- assert dict(sdf.to_coo().todok()) == dict(spm.todok())
-
- # Ensure dtype is preserved if possible
- res_dtype = object
- tm.assert_contains_all(sdf.dtypes, {np.dtype(res_dtype)})
- assert sdf.to_coo().dtype == res_dtype
-
-
-@td.skip_if_no_scipy
-def test_from_scipy_correct_ordering(spmatrix):
- # GH 16179
- arr = np.arange(1, 5).reshape(2, 2)
- try:
- spm = spmatrix(arr)
- assert spm.dtype == arr.dtype
- except (TypeError, AssertionError):
- # If conversion to sparse fails for this spmatrix type and arr.dtype,
- # then the combination is not currently supported in NumPy, so we
- # can just skip testing it thoroughly
- return
-
- sdf = pd.SparseDataFrame(spm)
- expected = pd.SparseDataFrame(arr)
- tm.assert_sp_frame_equal(sdf, expected)
- tm.assert_frame_equal(sdf.to_dense(), expected.to_dense())
-
-
-@td.skip_if_no_scipy
-def test_from_scipy_fillna(spmatrix):
- # GH 16112
- arr = np.eye(3)
- arr[1:, 0] = np.nan
-
- try:
- spm = spmatrix(arr)
- assert spm.dtype == arr.dtype
- except (TypeError, AssertionError):
- # If conversion to sparse fails for this spmatrix type and arr.dtype,
- # then the combination is not currently supported in NumPy, so we
- # can just skip testing it thoroughly
- return
-
- sdf = pd.SparseDataFrame(spm).fillna(-1.0)
-
- # Returning frame should fill all nan values with -1.0
- expected = pd.SparseDataFrame({
- 0: pd.SparseSeries([1., -1, -1]),
- 1: pd.SparseSeries([np.nan, 1, np.nan]),
- 2: pd.SparseSeries([np.nan, np.nan, 1]),
- }, default_fill_value=-1)
-
- # fill_value is expected to be what .fillna() above was called with
- # We don't use -1 as initial fill_value in expected SparseSeries
- # construction because this way we obtain "compressed" SparseArrays,
- # avoiding having to construct them ourselves
- for col in expected:
- expected[col].fill_value = -1
-
- tm.assert_sp_frame_equal(sdf, expected)
-
-
class TestSparseDataFrameArithmetic(object):
def test_numeric_op_scalar(self):
diff --git a/pandas/tests/sparse/frame/test_to_from_scipy.py b/pandas/tests/sparse/frame/test_to_from_scipy.py
new file mode 100644
index 0000000000000..aef49c84fc2ad
--- /dev/null
+++ b/pandas/tests/sparse/frame/test_to_from_scipy.py
@@ -0,0 +1,168 @@
+import pytest
+import numpy as np
+from warnings import catch_warnings
+from pandas.util import testing as tm
+from pandas import SparseDataFrame, SparseSeries
+from distutils.version import LooseVersion
+from pandas.core.dtypes.common import (
+ is_bool_dtype,
+ is_float_dtype,
+ is_object_dtype,
+ is_float)
+
+
+scipy = pytest.importorskip('scipy')
+
+
+@pytest.mark.parametrize('index', [None, list('abc')]) # noqa: F811
+@pytest.mark.parametrize('columns', [None, list('def')])
+@pytest.mark.parametrize('fill_value', [None, 0, np.nan])
+@pytest.mark.parametrize('dtype', [bool, int, float, np.uint16])
+def test_from_to_scipy(spmatrix, index, columns, fill_value, dtype):
+ # GH 4343
+ # Make one ndarray and from it one sparse matrix, both to be used for
+ # constructing frames and comparing results
+ arr = np.eye(3, dtype=dtype)
+ # GH 16179
+ arr[0, 1] = dtype(2)
+ try:
+ spm = spmatrix(arr)
+ assert spm.dtype == arr.dtype
+ except (TypeError, AssertionError):
+ # If conversion to sparse fails for this spmatrix type and arr.dtype,
+ # then the combination is not currently supported in NumPy, so we
+ # can just skip testing it thoroughly
+ return
+
+ sdf = SparseDataFrame(spm, index=index, columns=columns,
+ default_fill_value=fill_value)
+
+ # Expected result construction is kind of tricky for all
+ # dtype-fill_value combinations; easiest to cast to something generic
+ # and except later on
+ rarr = arr.astype(object)
+ rarr[arr == 0] = np.nan
+ expected = SparseDataFrame(rarr, index=index, columns=columns).fillna(
+ fill_value if fill_value is not None else np.nan)
+
+ # Assert frame is as expected
+ sdf_obj = sdf.astype(object)
+ tm.assert_sp_frame_equal(sdf_obj, expected)
+ tm.assert_frame_equal(sdf_obj.to_dense(), expected.to_dense())
+
+ # Assert spmatrices equal
+ assert dict(sdf.to_coo().todok()) == dict(spm.todok())
+
+ # Ensure dtype is preserved if possible
+ was_upcast = ((fill_value is None or is_float(fill_value)) and
+ not is_object_dtype(dtype) and
+ not is_float_dtype(dtype))
+ res_dtype = (bool if is_bool_dtype(dtype) else
+ float if was_upcast else
+ dtype)
+ tm.assert_contains_all(sdf.dtypes, {np.dtype(res_dtype)})
+ assert sdf.to_coo().dtype == res_dtype
+
+ # However, adding a str column results in an upcast to object
+ sdf['strings'] = np.arange(len(sdf)).astype(str)
+ assert sdf.to_coo().dtype == np.object_
+
+
+@pytest.mark.parametrize('fill_value', [None, 0, np.nan]) # noqa: F811
+def test_from_to_scipy_object(spmatrix, fill_value):
+ # GH 4343
+ dtype = object
+ columns = list('cd')
+ index = list('ab')
+
+ if (spmatrix is scipy.sparse.dok_matrix and LooseVersion(
+ scipy.__version__) >= LooseVersion('0.19.0')):
+ pytest.skip("dok_matrix from object does not work in SciPy >= 0.19")
+
+ # Make one ndarray and from it one sparse matrix, both to be used for
+ # constructing frames and comparing results
+ arr = np.eye(2, dtype=dtype)
+ try:
+ spm = spmatrix(arr)
+ assert spm.dtype == arr.dtype
+ except (TypeError, AssertionError):
+ # If conversion to sparse fails for this spmatrix type and arr.dtype,
+ # then the combination is not currently supported in NumPy, so we
+ # can just skip testing it thoroughly
+ return
+
+ sdf = SparseDataFrame(spm, index=index, columns=columns,
+ default_fill_value=fill_value)
+
+ # Expected result construction is kind of tricky for all
+ # dtype-fill_value combinations; easiest to cast to something generic
+ # and except later on
+ rarr = arr.astype(object)
+ rarr[arr == 0] = np.nan
+ expected = SparseDataFrame(rarr, index=index, columns=columns).fillna(
+ fill_value if fill_value is not None else np.nan)
+
+ # Assert frame is as expected
+ sdf_obj = sdf.astype(object)
+ tm.assert_sp_frame_equal(sdf_obj, expected)
+ tm.assert_frame_equal(sdf_obj.to_dense(), expected.to_dense())
+
+ # Assert spmatrices equal
+ with catch_warnings(record=True):
+ assert dict(sdf.to_coo().todok()) == dict(spm.todok())
+
+ # Ensure dtype is preserved if possible
+ res_dtype = object
+ tm.assert_contains_all(sdf.dtypes, {np.dtype(res_dtype)})
+ assert sdf.to_coo().dtype == res_dtype
+
+
+def test_from_scipy_correct_ordering(spmatrix):
+ # GH 16179
+ arr = np.arange(1, 5).reshape(2, 2)
+ try:
+ spm = spmatrix(arr)
+ assert spm.dtype == arr.dtype
+ except (TypeError, AssertionError):
+ # If conversion to sparse fails for this spmatrix type and arr.dtype,
+ # then the combination is not currently supported in NumPy, so we
+ # can just skip testing it thoroughly
+ return
+
+ sdf = SparseDataFrame(spm)
+ expected = SparseDataFrame(arr)
+ tm.assert_sp_frame_equal(sdf, expected)
+ tm.assert_frame_equal(sdf.to_dense(), expected.to_dense())
+
+
+def test_from_scipy_fillna(spmatrix):
+ # GH 16112
+ arr = np.eye(3)
+ arr[1:, 0] = np.nan
+
+ try:
+ spm = spmatrix(arr)
+ assert spm.dtype == arr.dtype
+ except (TypeError, AssertionError):
+ # If conversion to sparse fails for this spmatrix type and arr.dtype,
+ # then the combination is not currently supported in NumPy, so we
+ # can just skip testing it thoroughly
+ return
+
+ sdf = SparseDataFrame(spm).fillna(-1.0)
+
+ # Returning frame should fill all nan values with -1.0
+ expected = SparseDataFrame({
+ 0: SparseSeries([1., -1, -1]),
+ 1: SparseSeries([np.nan, 1, np.nan]),
+ 2: SparseSeries([np.nan, np.nan, 1]),
+ }, default_fill_value=-1)
+
+ # fill_value is expected to be what .fillna() above was called with
+ # We don't use -1 as initial fill_value in expected SparseSeries
+ # construction because this way we obtain "compressed" SparseArrays,
+ # avoiding having to construct them ourselves
+ for col in expected:
+ expected[col].fill_value = -1
+
+ tm.assert_sp_frame_equal(sdf, expected)
diff --git a/pandas/tests/test_algos.py b/pandas/tests/test_algos.py
index d7fc5033bab90..6b3b519d49f7f 100644
--- a/pandas/tests/test_algos.py
+++ b/pandas/tests/test_algos.py
@@ -788,10 +788,10 @@ def test_duplicated_with_nas(self):
2, 4, 1, 5, 6]),
np.array([1.1, 2.2, 1.1, np.nan, 3.3,
2.2, 4.4, 1.1, np.nan, 6.6]),
- pytest.mark.xfail(reason="Complex bug. GH 16399")(
- np.array([1 + 1j, 2 + 2j, 1 + 1j, 5 + 5j, 3 + 3j,
- 2 + 2j, 4 + 4j, 1 + 1j, 5 + 5j, 6 + 6j])
- ),
+ pytest.param(np.array([1 + 1j, 2 + 2j, 1 + 1j, 5 + 5j, 3 + 3j,
+ 2 + 2j, 4 + 4j, 1 + 1j, 5 + 5j, 6 + 6j]),
+ marks=pytest.mark.xfail(reason="Complex bug. GH 16399")
+ ),
np.array(['a', 'b', 'a', 'e', 'c',
'b', 'd', 'a', 'e', 'f'], dtype=object),
np.array([1, 2**63, 1, 3**5, 10, 2**63, 39, 1, 3**5, 7],
diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 86d9a9fa91e47..424ba6aab9a56 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -2289,7 +2289,7 @@ def test_reset_index_multiindex_columns(self):
# gh-16120: already existing column
with tm.assert_raises_regex(ValueError,
- ("cannot insert \('A', ''\), "
+ (r"cannot insert \('A', ''\), "
"already exists")):
df.rename_axis('A').reset_index()
@@ -2323,7 +2323,7 @@ def test_reset_index_multiindex_columns(self):
# ... which is incompatible with col_fill=None
with tm.assert_raises_regex(ValueError,
("col_fill=None is incompatible with "
- "incomplete column name \('C', 'c'\)")):
+ r"incomplete column name \('C', 'c'\)")):
df2.rename_axis([('C', 'c')]).reset_index(col_fill=None)
# with col_level != 0
diff --git a/pandas/tests/util/test_testing.py b/pandas/tests/util/test_testing.py
index 31580bc9eab57..1c878604b11a2 100644
--- a/pandas/tests/util/test_testing.py
+++ b/pandas/tests/util/test_testing.py
@@ -48,12 +48,18 @@ def test_assert_almost_equal_numbers_with_mixed(self):
self._assert_not_almost_equal_both(1, [1, ])
self._assert_not_almost_equal_both(1, object())
- def test_assert_almost_equal_edge_case_ndarrays(self):
- self._assert_almost_equal_both(np.array([], dtype='M8[ns]'),
- np.array([], dtype='float64'),
- check_dtype=False)
- self._assert_almost_equal_both(np.array([], dtype=str),
- np.array([], dtype='int64'),
+ @pytest.mark.parametrize(
+ "left_dtype",
+ ['M8[ns]', 'm8[ns]', 'float64', 'int64', 'object'])
+ @pytest.mark.parametrize(
+ "right_dtype",
+ ['M8[ns]', 'm8[ns]', 'float64', 'int64', 'object'])
+ def test_assert_almost_equal_edge_case_ndarrays(
+ self, left_dtype, right_dtype):
+
+ # empty compare
+ self._assert_almost_equal_both(np.array([], dtype=left_dtype),
+ np.array([], dtype=right_dtype),
check_dtype=False)
def test_assert_almost_equal_dicts(self):
| https://api.github.com/repos/pandas-dev/pandas/pulls/19003 | 2017-12-30T17:42:27Z | 2017-12-30T22:48:01Z | 2017-12-30T22:48:01Z | 2017-12-30T22:48:02Z | |
DOC: More 0.22.0 updates | diff --git a/doc/source/whatsnew/v0.21.0.txt b/doc/source/whatsnew/v0.21.0.txt
index 89e2d3006696c..3e673bd4cbc28 100644
--- a/doc/source/whatsnew/v0.21.0.txt
+++ b/doc/source/whatsnew/v0.21.0.txt
@@ -372,6 +372,12 @@ Additionally, support has been dropped for Python 3.4 (:issue:`15251`).
Sum/Prod of all-NaN or empty Series/DataFrames is now consistently NaN
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
+.. note::
+
+ The changes described here have been partially reverted. See
+ the :ref:`v0.22.0 Whatsnew <whatsnew_0220>` for more.
+
+
The behavior of ``sum`` and ``prod`` on all-NaN Series/DataFrames no longer depends on
whether `bottleneck <http://berkeleyanalytics.com/bottleneck>`__ is installed, and return value of ``sum`` and ``prod`` on an empty Series has changed (:issue:`9422`, :issue:`15507`).
diff --git a/doc/source/whatsnew/v0.22.0.txt b/doc/source/whatsnew/v0.22.0.txt
index da4acd99e3873..d165339cb0de9 100644
--- a/doc/source/whatsnew/v0.22.0.txt
+++ b/doc/source/whatsnew/v0.22.0.txt
@@ -218,3 +218,26 @@ returns ``0``.
The default behavior of ``min_periods=None``, implying that ``min_periods``
equals the window size, is unchanged.
+
+Compatibility
+~~~~~~~~~~~~~
+
+If you maintain a library that should work across pandas versions, it
+may be easiest to exclude pandas 0.21 from your requirements. Otherwise, all your
+``sum()`` calls would need to check if the ``Series`` is empty before summing.
+
+With setuptools, in your ``setup.py`` use::
+
+ install_requires=['pandas!=0.21.*', ...]
+
+With conda, use
+
+.. code-block:: yaml
+
+ requirements:
+ run:
+ - pandas !=0.21.0,!=0.21.1
+
+Note that the inconsistency in the return value for all-*NA* series is still
+there for pandas 0.20.3 and earlier. Avoiding pandas 0.21 will only help with
+the empty case.
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index c5359ba2c5ea1..7a0e1fe361c59 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -7534,8 +7534,7 @@ def _doc_parms(cls):
----------
axis : %(axis_descr)s
skipna : boolean, default True
- Exclude NA/null values. If an entire row/column is NA or empty, the result
- will be NA
+ Exclude NA/null values when computing the result.
level : int or level name, default None
If the axis is a MultiIndex (hierarchical), count along a
particular level, collapsing into a %(name1)s
@@ -7669,7 +7668,7 @@ def _doc_parms(cls):
The required number of valid values to perform the operation. If fewer than
``min_count`` non-NA values are present the result will be NA.
- .. versionadded :: 0.21.2
+ .. versionadded :: 0.22.0
Added with the default being 1. This means the sum or product
of an all-NA or empty series is ``NaN``.
| [ci skip]
cc @jorisvandenbossche
xref https://github.com/pandas-dev/pandas/issues/18985#issuecomment-354525713 and https://github.com/pandas-dev/pandas/pull/18983#issuecomment-354525079 | https://api.github.com/repos/pandas-dev/pandas/pulls/19002 | 2017-12-30T13:37:57Z | 2017-12-30T19:30:40Z | 2017-12-30T19:30:40Z | 2017-12-30T19:31:11Z |
TST: limit printing of xfail cases & catch Performance Warnings | diff --git a/pandas/tests/indexes/datetimes/test_arithmetic.py b/pandas/tests/indexes/datetimes/test_arithmetic.py
index 6cfa083172921..11a52267ed1b4 100644
--- a/pandas/tests/indexes/datetimes/test_arithmetic.py
+++ b/pandas/tests/indexes/datetimes/test_arithmetic.py
@@ -368,12 +368,15 @@ def test_dti_add_offset_array(self, tz, box):
# GH#18849
dti = pd.date_range('2017-01-01', periods=2, tz=tz)
other = box([pd.offsets.MonthEnd(), pd.offsets.Day(n=2)])
- res = dti + other
+
+ with tm.assert_produces_warning(PerformanceWarning):
+ res = dti + other
expected = DatetimeIndex([dti[n] + other[n] for n in range(len(dti))],
name=dti.name, freq='infer')
tm.assert_index_equal(res, expected)
- res2 = other + dti
+ with tm.assert_produces_warning(PerformanceWarning):
+ res2 = other + dti
tm.assert_index_equal(res2, expected)
@pytest.mark.parametrize('box', [np.array, pd.Index])
@@ -381,7 +384,9 @@ def test_dti_sub_offset_array(self, tz, box):
# GH#18824
dti = pd.date_range('2017-01-01', periods=2, tz=tz)
other = box([pd.offsets.MonthEnd(), pd.offsets.Day(n=2)])
- res = dti - other
+
+ with tm.assert_produces_warning(PerformanceWarning):
+ res = dti - other
expected = DatetimeIndex([dti[n] - other[n] for n in range(len(dti))],
name=dti.name, freq='infer')
tm.assert_index_equal(res, expected)
@@ -392,20 +397,25 @@ def test_dti_sub_offset_array(self, tz, box):
def test_dti_with_offset_series(self, tz, names):
# GH#18849
dti = pd.date_range('2017-01-01', periods=2, tz=tz, name=names[0])
- other = pd.Series([pd.offsets.MonthEnd(), pd.offsets.Day(n=2)],
- name=names[1])
+ other = Series([pd.offsets.MonthEnd(), pd.offsets.Day(n=2)],
+ name=names[1])
- expected_add = pd.Series([dti[n] + other[n] for n in range(len(dti))],
- name=names[2])
- res = dti + other
+ expected_add = Series([dti[n] + other[n] for n in range(len(dti))],
+ name=names[2])
+
+ with tm.assert_produces_warning(PerformanceWarning):
+ res = dti + other
tm.assert_series_equal(res, expected_add)
- res2 = other + dti
+
+ with tm.assert_produces_warning(PerformanceWarning):
+ res2 = other + dti
tm.assert_series_equal(res2, expected_add)
- expected_sub = pd.Series([dti[n] - other[n] for n in range(len(dti))],
- name=names[2])
+ expected_sub = Series([dti[n] - other[n] for n in range(len(dti))],
+ name=names[2])
- res3 = dti - other
+ with tm.assert_produces_warning(PerformanceWarning):
+ res3 = dti - other
tm.assert_series_equal(res3, expected_sub)
diff --git a/pandas/tests/indexing/test_coercion.py b/pandas/tests/indexing/test_coercion.py
index 619a8ca3bf112..52b2d7205c849 100644
--- a/pandas/tests/indexing/test_coercion.py
+++ b/pandas/tests/indexing/test_coercion.py
@@ -819,8 +819,8 @@ def test_replace_series(self, how, to_key, from_key):
assert obj.dtype == from_key
if (from_key.startswith('datetime') and to_key.startswith('datetime')):
- pytest.xfail("different tz, currently mask_missing "
- "raises SystemError")
+ # tested below
+ return
if how == 'dict':
replacer = dict(zip(self.rep[from_key], self.rep[to_key]))
@@ -849,5 +849,38 @@ def test_replace_series(self, how, to_key, from_key):
tm.assert_series_equal(result, exp)
+ # TODO(jreback) commented out to only have a single xfail printed
+ @pytest.mark.xfail(reason="different tz, "
+ "currently mask_missing raises SystemError")
+ # @pytest.mark.parametrize('how', ['dict', 'series'])
+ # @pytest.mark.parametrize('to_key', [
+ # 'datetime64[ns]', 'datetime64[ns, UTC]',
+ # 'datetime64[ns, US/Eastern]'])
+ # @pytest.mark.parametrize('from_key', [
+ # 'datetime64[ns]', 'datetime64[ns, UTC]',
+ # 'datetime64[ns, US/Eastern]'])
+ # def test_replace_series_datetime_datetime(self, how, to_key, from_key):
+ def test_replace_series_datetime_datetime(self):
+ how = 'dict'
+ to_key = 'datetime64[ns]'
+ from_key = 'datetime64[ns]'
+
+ index = pd.Index([3, 4], name='xxx')
+ obj = pd.Series(self.rep[from_key], index=index, name='yyy')
+ assert obj.dtype == from_key
+
+ if how == 'dict':
+ replacer = dict(zip(self.rep[from_key], self.rep[to_key]))
+ elif how == 'series':
+ replacer = pd.Series(self.rep[to_key], index=self.rep[from_key])
+ else:
+ raise ValueError
+
+ result = obj.replace(replacer)
+ exp = pd.Series(self.rep[to_key], index=index, name='yyy')
+ assert exp.dtype == to_key
+
+ tm.assert_series_equal(result, exp)
+
def test_replace_series_period(self):
pass
diff --git a/pandas/tests/series/test_rank.py b/pandas/tests/series/test_rank.py
index bccc46f1e0ca8..6220ce8ff7669 100644
--- a/pandas/tests/series/test_rank.py
+++ b/pandas/tests/series/test_rank.py
@@ -209,11 +209,9 @@ def test_rank_signature(self):
pytest.param([np.iinfo(np.int64).min, -100, 0, 1, 9999, 100000,
1e10, np.iinfo(np.int64).max],
'int64',
- marks=pytest.mark.xfail(reason='''iNaT is equivalent to
- minimum value of dtype
- int64 pending issue
- #16674'''),
- ),
+ marks=pytest.mark.xfail(
+ reason="iNaT is equivalent to minimum value of dtype"
+ "int64 pending issue #16674")),
([NegInfinity(), '1', 'A', 'BA', 'Ba', 'C', Infinity()],
'object')
])
diff --git a/pandas/tests/tseries/offsets/test_offsets.py b/pandas/tests/tseries/offsets/test_offsets.py
index b304ebff55b6e..edabf4a7ccc99 100644
--- a/pandas/tests/tseries/offsets/test_offsets.py
+++ b/pandas/tests/tseries/offsets/test_offsets.py
@@ -113,7 +113,7 @@ def _get_offset(self, klass, value=1, normalize=False):
else:
try:
klass = klass(value, normalize=normalize)
- except:
+ except Exception:
klass = klass(normalize=normalize)
return klass
@@ -143,10 +143,10 @@ def test_apply_out_of_range(self, tz):
except tslib.OutOfBoundsDatetime:
raise
- except (ValueError, KeyError) as e:
- pytest.skip(
- "cannot create out_of_range offset: {0} {1}".format(
- str(self).split('.')[-1], e))
+ except (ValueError, KeyError):
+ # we are creating an invalid offset
+ # so ignore
+ pass
class TestCommon(Base):
| TST: limit printing of xfail cases
no need to report invalid offset creation
STYLE/DEPR: catch PerformanceWarnings
closes #18989
| https://api.github.com/repos/pandas-dev/pandas/pulls/19001 | 2017-12-30T13:21:53Z | 2017-12-30T15:10:23Z | 2017-12-30T15:10:23Z | 2017-12-30T15:10:23Z |
CLN: rename lib.isscalar to lib.is_scalar | diff --git a/pandas/_libs/lib.pyx b/pandas/_libs/lib.pyx
index 3898f7499e85e..bfcf0c6e69a2f 100644
--- a/pandas/_libs/lib.pyx
+++ b/pandas/_libs/lib.pyx
@@ -98,7 +98,7 @@ def memory_usage_of_objects(ndarray[object, ndim=1] arr):
# ----------------------------------------------------------------------
-cpdef bint isscalar(object val):
+cpdef bint is_scalar(object val):
"""
Return True if given value is scalar.
diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 167f215b6c0ac..1f1e47a6c54d6 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -750,7 +750,7 @@ def _broadcast(arr_or_scalar, shape):
Helper function to broadcast arrays / scalars to the desired shape.
"""
if _np_version_under1p10:
- if lib.isscalar(arr_or_scalar):
+ if is_scalar(arr_or_scalar):
out = np.empty(shape)
out.fill(arr_or_scalar)
else:
diff --git a/pandas/core/computation/align.py b/pandas/core/computation/align.py
index 0e7ae0cbe7c87..2e912b0075bfd 100644
--- a/pandas/core/computation/align.py
+++ b/pandas/core/computation/align.py
@@ -126,7 +126,7 @@ def _align(terms):
return np.result_type(terms.type), None
# if all resolved variables are numeric scalars
- if all(term.isscalar for term in terms):
+ if all(term.is_scalar for term in terms):
return _result_type_many(*(term.value for term in terms)).type, None
# perform the main alignment
diff --git a/pandas/core/computation/expr.py b/pandas/core/computation/expr.py
index 23abfa8b3fca1..b68b6970a89cc 100644
--- a/pandas/core/computation/expr.py
+++ b/pandas/core/computation/expr.py
@@ -368,11 +368,11 @@ def _maybe_transform_eq_ne(self, node, left=None, right=None):
def _maybe_downcast_constants(self, left, right):
f32 = np.dtype(np.float32)
- if left.isscalar and not right.isscalar and right.return_type == f32:
+ if left.is_scalar and not right.is_scalar and right.return_type == f32:
# right is a float32 array, left is a scalar
name = self.env.add_tmp(np.float32(left.value))
left = self.term_type(name, self.env)
- if right.isscalar and not left.isscalar and left.return_type == f32:
+ if right.is_scalar and not left.is_scalar and left.return_type == f32:
# left is a float32 array, right is a scalar
name = self.env.add_tmp(np.float32(right.value))
right = self.term_type(name, self.env)
diff --git a/pandas/core/computation/ops.py b/pandas/core/computation/ops.py
index 7ba2c16530cad..ca0c4db4947c4 100644
--- a/pandas/core/computation/ops.py
+++ b/pandas/core/computation/ops.py
@@ -99,7 +99,7 @@ def update(self, value):
self.value = value
@property
- def isscalar(self):
+ def is_scalar(self):
return is_scalar(self._value)
@property
@@ -214,8 +214,8 @@ def operand_types(self):
return frozenset(term.type for term in com.flatten(self))
@property
- def isscalar(self):
- return all(operand.isscalar for operand in self.operands)
+ def is_scalar(self):
+ return all(operand.is_scalar for operand in self.operands)
@property
def is_datetime(self):
@@ -412,7 +412,7 @@ def stringify(value):
lhs, rhs = self.lhs, self.rhs
- if is_term(lhs) and lhs.is_datetime and is_term(rhs) and rhs.isscalar:
+ if is_term(lhs) and lhs.is_datetime and is_term(rhs) and rhs.is_scalar:
v = rhs.value
if isinstance(v, (int, float)):
v = stringify(v)
@@ -421,7 +421,7 @@ def stringify(value):
v = v.tz_convert('UTC')
self.rhs.update(v)
- if is_term(rhs) and rhs.is_datetime and is_term(lhs) and lhs.isscalar:
+ if is_term(rhs) and rhs.is_datetime and is_term(lhs) and lhs.is_scalar:
v = lhs.value
if isinstance(v, (int, float)):
v = stringify(v)
@@ -431,7 +431,7 @@ def stringify(value):
self.lhs.update(v)
def _disallow_scalar_only_bool_ops(self):
- if ((self.lhs.isscalar or self.rhs.isscalar) and
+ if ((self.lhs.is_scalar or self.rhs.is_scalar) and
self.op in _bool_ops_dict and
(not (issubclass(self.rhs.return_type, (bool, np.bool_)) and
issubclass(self.lhs.return_type, (bool, np.bool_))))):
diff --git a/pandas/core/dtypes/inference.py b/pandas/core/dtypes/inference.py
index de769c69f44fd..8010a213efaf0 100644
--- a/pandas/core/dtypes/inference.py
+++ b/pandas/core/dtypes/inference.py
@@ -17,7 +17,7 @@
is_complex = lib.is_complex
-is_scalar = lib.isscalar
+is_scalar = lib.is_scalar
is_decimal = lib.is_decimal
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 04b8ade7e5253..94fbf290900b4 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -1101,7 +1101,7 @@ def _convert_for_op(self, value):
def _assert_can_do_op(self, value):
""" Check value is valid for scalar op """
- if not lib.isscalar(value):
+ if not is_scalar(value):
msg = "'value' must be a scalar, passed: {0}"
raise TypeError(msg.format(type(value).__name__))
diff --git a/pandas/core/indexes/datetimelike.py b/pandas/core/indexes/datetimelike.py
index 40c07376d2522..e40a3ba742609 100644
--- a/pandas/core/indexes/datetimelike.py
+++ b/pandas/core/indexes/datetimelike.py
@@ -905,7 +905,7 @@ def astype(self, dtype, copy=True):
def _ensure_datetimelike_to_i8(other):
""" helper for coercing an input scalar or array to i8 """
- if lib.isscalar(other) and isna(other):
+ if is_scalar(other) and isna(other):
other = iNaT
elif isinstance(other, ABCIndexClass):
# convert tz if needed
diff --git a/pandas/core/indexes/numeric.py b/pandas/core/indexes/numeric.py
index 5995b9fc7674c..6337c2f73d5ec 100644
--- a/pandas/core/indexes/numeric.py
+++ b/pandas/core/indexes/numeric.py
@@ -38,7 +38,7 @@ def __new__(cls, data=None, dtype=None, copy=False, name=None,
if fastpath:
return cls._simple_new(data, name=name)
- # isscalar, generators handled in coerce_to_ndarray
+ # is_scalar, generators handled in coerce_to_ndarray
data = cls._coerce_to_ndarray(data)
if issubclass(data.dtype.type, compat.string_types):
diff --git a/pandas/tests/dtypes/test_inference.py b/pandas/tests/dtypes/test_inference.py
index 219d1b2852938..33c570a814e7d 100644
--- a/pandas/tests/dtypes/test_inference.py
+++ b/pandas/tests/dtypes/test_inference.py
@@ -1098,9 +1098,9 @@ def test_is_timedelta(self):
assert not is_timedelta64_ns_dtype(tdi.astype('timedelta64[h]'))
-class Testisscalar(object):
+class TestIsScalar(object):
- def test_isscalar_builtin_scalars(self):
+ def test_is_scalar_builtin_scalars(self):
assert is_scalar(None)
assert is_scalar(True)
assert is_scalar(False)
@@ -1115,7 +1115,7 @@ def test_isscalar_builtin_scalars(self):
assert is_scalar(timedelta(hours=1))
assert is_scalar(pd.NaT)
- def test_isscalar_builtin_nonscalars(self):
+ def test_is_scalar_builtin_nonscalars(self):
assert not is_scalar({})
assert not is_scalar([])
assert not is_scalar([1])
@@ -1124,7 +1124,7 @@ def test_isscalar_builtin_nonscalars(self):
assert not is_scalar(slice(None))
assert not is_scalar(Ellipsis)
- def test_isscalar_numpy_array_scalars(self):
+ def test_is_scalar_numpy_array_scalars(self):
assert is_scalar(np.int64(1))
assert is_scalar(np.float64(1.))
assert is_scalar(np.int32(1))
@@ -1135,7 +1135,7 @@ def test_isscalar_numpy_array_scalars(self):
assert is_scalar(np.datetime64('2014-01-01'))
assert is_scalar(np.timedelta64(1, 'h'))
- def test_isscalar_numpy_zerodim_arrays(self):
+ def test_is_scalar_numpy_zerodim_arrays(self):
for zerodim in [np.array(1), np.array('foobar'),
np.array(np.datetime64('2014-01-01')),
np.array(np.timedelta64(1, 'h')),
@@ -1143,19 +1143,19 @@ def test_isscalar_numpy_zerodim_arrays(self):
assert not is_scalar(zerodim)
assert is_scalar(lib.item_from_zerodim(zerodim))
- def test_isscalar_numpy_arrays(self):
+ def test_is_scalar_numpy_arrays(self):
assert not is_scalar(np.array([]))
assert not is_scalar(np.array([[]]))
assert not is_scalar(np.matrix('1; 2'))
- def test_isscalar_pandas_scalars(self):
+ def test_is_scalar_pandas_scalars(self):
assert is_scalar(Timestamp('2014-01-01'))
assert is_scalar(Timedelta(hours=1))
assert is_scalar(Period('2014-01-01'))
assert is_scalar(Interval(left=0, right=1))
assert is_scalar(DateOffset(days=1))
- def test_lisscalar_pandas_containers(self):
+ def test_is_scalar_pandas_containers(self):
assert not is_scalar(Series())
assert not is_scalar(Series([1]))
assert not is_scalar(DataFrame())
| - [x] closes #18987
- [ ] tests added / passed
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [ ] whatsnew entry
There's also a handful of methods which call `is_scalar` that are named `isscalar`, e.g.:
https://github.com/pandas-dev/pandas/blob/84335621ad0a0d83302a80b6911d3985c00b5cee/pandas/core/computation/ops.py#L101-L103
should these be changed as well?
Also does a CLN require an entry in whatsnew?
| https://api.github.com/repos/pandas-dev/pandas/pulls/19000 | 2017-12-30T05:37:45Z | 2017-12-30T22:41:23Z | 2017-12-30T22:41:23Z | 2017-12-30T22:41:27Z |
Fix IntervalDtype Bugs and Inconsistencies | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index d7a3f0d077302..783de6569a542 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -270,6 +270,7 @@ Other API Changes
- Subtraction of :class:`Series` with timezone-aware ``dtype='datetime64[ns]'`` with mis-matched timezones will raise ``TypeError`` instead of ``ValueError`` (issue:`18817`)
- :class:`IntervalIndex` and ``IntervalDtype`` no longer support categorical, object, and string subtypes (:issue:`19016`)
- The default ``Timedelta`` constructor now accepts an ``ISO 8601 Duration`` string as an argument (:issue:`19040`)
+- ``IntervalDtype`` now returns ``True`` when compared against ``'interval'`` regardless of subtype, and ``IntervalDtype.name`` now returns ``'interval'`` regardless of subtype (:issue:`18980`)
.. _whatsnew_0230.deprecations:
diff --git a/pandas/core/dtypes/dtypes.py b/pandas/core/dtypes/dtypes.py
index 08773354d44d8..2ec35889d6a7a 100644
--- a/pandas/core/dtypes/dtypes.py
+++ b/pandas/core/dtypes/dtypes.py
@@ -626,6 +626,7 @@ class IntervalDtype(ExtensionDtype):
THIS IS NOT A REAL NUMPY DTYPE
"""
+ name = 'interval'
type = IntervalDtypeType
kind = None
str = '|O08'
@@ -653,8 +654,8 @@ def __new__(cls, subtype=None):
u.subtype = None
return u
elif (isinstance(subtype, compat.string_types) and
- subtype == 'interval'):
- subtype = ''
+ subtype.lower() == 'interval'):
+ subtype = None
else:
if isinstance(subtype, compat.string_types):
m = cls._match.search(subtype)
@@ -666,11 +667,6 @@ def __new__(cls, subtype=None):
except TypeError:
raise ValueError("could not construct IntervalDtype")
- if subtype is None:
- u = object.__new__(cls)
- u.subtype = None
- return u
-
if is_categorical_dtype(subtype) or is_string_dtype(subtype):
# GH 19016
msg = ('category, object, and string subtypes are not supported '
@@ -692,31 +688,29 @@ def construct_from_string(cls, string):
if its not possible
"""
if isinstance(string, compat.string_types):
- try:
- return cls(string)
- except ValueError:
- pass
- raise TypeError("could not construct IntervalDtype")
+ return cls(string)
+ msg = "a string needs to be passed, got type {typ}"
+ raise TypeError(msg.format(typ=type(string)))
def __unicode__(self):
if self.subtype is None:
return "interval"
return "interval[{subtype}]".format(subtype=self.subtype)
- @property
- def name(self):
- return str(self)
-
def __hash__(self):
# make myself hashable
return hash(str(self))
def __eq__(self, other):
if isinstance(other, compat.string_types):
- return other == self.name or other == self.name.title()
-
- return (isinstance(other, IntervalDtype) and
- self.subtype == other.subtype)
+ return other.lower() in (self.name.lower(), str(self).lower())
+ elif not isinstance(other, IntervalDtype):
+ return False
+ elif self.subtype is None or other.subtype is None:
+ # None should match any subtype
+ return True
+ else:
+ return self.subtype == other.subtype
@classmethod
def is_dtype(cls, dtype):
diff --git a/pandas/tests/dtypes/test_dtypes.py b/pandas/tests/dtypes/test_dtypes.py
index 6a3715fd66159..692fb3271cfda 100644
--- a/pandas/tests/dtypes/test_dtypes.py
+++ b/pandas/tests/dtypes/test_dtypes.py
@@ -433,7 +433,7 @@ def test_hash_vs_equality(self):
assert dtype2 == dtype
assert dtype3 == dtype
assert dtype is dtype2
- assert dtype2 is dtype
+ assert dtype2 is dtype3
assert dtype3 is dtype
assert hash(dtype) == hash(dtype2)
assert hash(dtype) == hash(dtype3)
@@ -451,14 +451,19 @@ def test_hash_vs_equality(self):
assert hash(dtype2) == hash(dtype2)
assert hash(dtype2) == hash(dtype3)
- def test_construction(self):
- with pytest.raises(ValueError):
- IntervalDtype('xx')
+ @pytest.mark.parametrize('subtype', [
+ 'interval[int64]', 'Interval[int64]', 'int64', np.dtype('int64')])
+ def test_construction(self, subtype):
+ i = IntervalDtype(subtype)
+ assert i.subtype == np.dtype('int64')
+ assert is_interval_dtype(i)
- for s in ['interval[int64]', 'Interval[int64]', 'int64']:
- i = IntervalDtype(s)
- assert i.subtype == np.dtype('int64')
- assert is_interval_dtype(i)
+ @pytest.mark.parametrize('subtype', [None, 'interval', 'Interval'])
+ def test_construction_generic(self, subtype):
+ # generic
+ i = IntervalDtype(subtype)
+ assert i.subtype is None
+ assert is_interval_dtype(i)
@pytest.mark.parametrize('subtype', [
CategoricalDtype(list('abc'), False),
@@ -471,17 +476,27 @@ def test_construction_not_supported(self, subtype):
with tm.assert_raises_regex(TypeError, msg):
IntervalDtype(subtype)
- def test_construction_generic(self):
- # generic
- i = IntervalDtype('interval')
- assert i.subtype == ''
- assert is_interval_dtype(i)
- assert str(i) == 'interval[]'
+ def test_construction_errors(self):
+ msg = 'could not construct IntervalDtype'
+ with tm.assert_raises_regex(ValueError, msg):
+ IntervalDtype('xx')
- i = IntervalDtype()
- assert i.subtype is None
- assert is_interval_dtype(i)
- assert str(i) == 'interval'
+ def test_construction_from_string(self):
+ result = IntervalDtype('interval[int64]')
+ assert is_dtype_equal(self.dtype, result)
+ result = IntervalDtype.construct_from_string('interval[int64]')
+ assert is_dtype_equal(self.dtype, result)
+
+ @pytest.mark.parametrize('string', [
+ 'foo', 'interval[foo]', 'foo[int64]', 0, 3.14, ('a', 'b'), None])
+ def test_construction_from_string_errors(self, string):
+ if isinstance(string, string_types):
+ error, msg = ValueError, 'could not construct IntervalDtype'
+ else:
+ error, msg = TypeError, 'a string needs to be passed, got type'
+
+ with tm.assert_raises_regex(error, msg):
+ IntervalDtype.construct_from_string(string)
def test_subclass(self):
a = IntervalDtype('interval[int64]')
@@ -506,36 +521,45 @@ def test_is_dtype(self):
assert not IntervalDtype.is_dtype(np.int64)
assert not IntervalDtype.is_dtype(np.float64)
- def test_identity(self):
- assert (IntervalDtype('interval[int64]') ==
- IntervalDtype('interval[int64]'))
-
def test_coerce_to_dtype(self):
assert (_coerce_to_dtype('interval[int64]') ==
IntervalDtype('interval[int64]'))
- def test_construction_from_string(self):
- result = IntervalDtype('interval[int64]')
- assert is_dtype_equal(self.dtype, result)
- result = IntervalDtype.construct_from_string('interval[int64]')
- assert is_dtype_equal(self.dtype, result)
- with pytest.raises(TypeError):
- IntervalDtype.construct_from_string('foo')
- with pytest.raises(TypeError):
- IntervalDtype.construct_from_string('interval[foo]')
- with pytest.raises(TypeError):
- IntervalDtype.construct_from_string('foo[int64]')
-
def test_equality(self):
assert is_dtype_equal(self.dtype, 'interval[int64]')
assert is_dtype_equal(self.dtype, IntervalDtype('int64'))
- assert is_dtype_equal(self.dtype, IntervalDtype('int64'))
assert is_dtype_equal(IntervalDtype('int64'), IntervalDtype('int64'))
assert not is_dtype_equal(self.dtype, 'int64')
assert not is_dtype_equal(IntervalDtype('int64'),
IntervalDtype('float64'))
+ @pytest.mark.parametrize('subtype', [
+ None, 'interval', 'Interval', 'int64', 'uint64', 'float64',
+ 'complex128', 'datetime64', 'timedelta64', PeriodDtype('Q')])
+ def test_equality_generic(self, subtype):
+ # GH 18980
+ dtype = IntervalDtype(subtype)
+ assert is_dtype_equal(dtype, 'interval')
+ assert is_dtype_equal(dtype, IntervalDtype())
+
+ @pytest.mark.parametrize('subtype', [
+ 'int64', 'uint64', 'float64', 'complex128', 'datetime64',
+ 'timedelta64', PeriodDtype('Q')])
+ def test_name_repr(self, subtype):
+ # GH 18980
+ dtype = IntervalDtype(subtype)
+ expected = 'interval[{subtype}]'.format(subtype=subtype)
+ assert str(dtype) == expected
+ assert dtype.name == 'interval'
+
+ @pytest.mark.parametrize('subtype', [None, 'interval', 'Interval'])
+ def test_name_repr_generic(self, subtype):
+ # GH 18980
+ dtype = IntervalDtype(subtype)
+ assert str(dtype) == 'interval'
+ assert dtype.name == 'interval'
+
def test_basic(self):
assert is_interval_dtype(self.dtype)
| - [X] closes #18980
- [X] tests added / passed
- [X] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [X] whatsnew entry
Summary:
- `IntervalDtype(*)` now returns `True` when compared against `'interval'` and `IntervalDtype()` regardless of subtype
- `IntervalDtype(*)` now returns `'interval'` regardless of subtype
- `str(IntervalDtype(*))` still displays subtype information, e.g. `'interval[int64]'`
- Cleaned up miscellaneous tests related to `IntervalDtype`
| https://api.github.com/repos/pandas-dev/pandas/pulls/18997 | 2017-12-29T21:09:36Z | 2018-01-10T21:28:03Z | 2018-01-10T21:28:03Z | 2018-09-24T17:26:29Z |
TST: Remove pow test in expressions | diff --git a/ci/install_travis.sh b/ci/install_travis.sh
index ab45b5113802c..272e7f2e05d14 100755
--- a/ci/install_travis.sh
+++ b/ci/install_travis.sh
@@ -56,11 +56,6 @@ if [ "$CONDA_BUILD_TEST" ]; then
conda install conda-build
fi
-# TODO(jreback)
-echo
-echo "[fix conda version]"
-conda install conda=4.3.30
-
echo
echo "[add channels]"
conda config --remove channels defaults || exit 1
diff --git a/pandas/tests/test_expressions.py b/pandas/tests/test_expressions.py
index 6d2607962dfb0..aebc9cd3deaac 100644
--- a/pandas/tests/test_expressions.py
+++ b/pandas/tests/test_expressions.py
@@ -73,17 +73,11 @@ def teardown_method(self, method):
def run_arithmetic(self, df, other, assert_func, check_dtype=False,
test_flex=True):
expr._MIN_ELEMENTS = 0
- operations = ['add', 'sub', 'mul', 'mod', 'truediv', 'floordiv', 'pow']
+ operations = ['add', 'sub', 'mul', 'mod', 'truediv', 'floordiv']
if not compat.PY3:
operations.append('div')
for arith in operations:
- # numpy >= 1.11 doesn't handle integers
- # raised to integer powers
- # https://github.com/pandas-dev/pandas/issues/15363
- if arith == 'pow' and not _np_version_under1p11:
- continue
-
operator_name = arith
if arith == 'div':
operator_name = 'truediv'
| These are already skipped for NumPy>=1.12, and buggy for NumPy
1.10.4
cc @jreback
This will be backported.
closes #18992 | https://api.github.com/repos/pandas-dev/pandas/pulls/18995 | 2017-12-29T19:15:27Z | 2017-12-29T20:21:29Z | 2017-12-29T20:21:29Z | 2018-06-29T08:42:01Z |
CI: fix pip install | diff --git a/ci/install_travis.sh b/ci/install_travis.sh
index 693a2fe1fd6a6..ab45b5113802c 100755
--- a/ci/install_travis.sh
+++ b/ci/install_travis.sh
@@ -106,6 +106,9 @@ time conda create -n pandas --file=${REQ} || exit 1
source activate pandas
+# https://github.com/travis-ci/travis-ci/issues/8920#issuecomment-352661024
+python -c "import fcntl; fcntl.fcntl(1, fcntl.F_SETFL, 0)"
+
# may have addtl installation instructions for this build
echo
echo "[build addtl installs]"
| https://api.github.com/repos/pandas-dev/pandas/pulls/18990 | 2017-12-29T16:49:42Z | 2017-12-29T17:42:28Z | 2017-12-29T17:42:28Z | 2017-12-29T17:42:28Z | |
0.22.0 backports | diff --git a/doc/source/release.rst b/doc/source/release.rst
index 0298eda2c78ab..aea6280a490d6 100644
--- a/doc/source/release.rst
+++ b/doc/source/release.rst
@@ -37,6 +37,27 @@ analysis / manipulation tool available in any language.
* Binary installers on PyPI: http://pypi.python.org/pypi/pandas
* Documentation: http://pandas.pydata.org
+pandas 0.22.0
+-------------
+
+**Release date:** December 29, 2017
+
+This is a major release from 0.21.1 and includes a single, API-breaking change.
+We recommend that all users upgrade to this version after carefully reading the
+release note.
+
+The only changes are:
+
+- The sum of an empty or all-*NA* ``Series`` is now ``0``
+- The product of an empty or all-*NA* ``Series`` is now ``1``
+- We've added a ``min_count`` parameter to ``.sum()`` and ``.prod()`` controlling
+ the minimum number of valid values for the result to be valid. If fewer than
+ ``min_count`` non-*NA* values are present, the result is *NA*. The default is
+ ``0``. To return ``NaN``, the 0.21 behavior, use ``min_count=1``.
+
+See the :ref:`v0.22.0 Whatsnew <whatsnew_0220>` overview for further explanation
+of all the places in the library this affects.
+
pandas 0.21.1
-------------
diff --git a/doc/source/whatsnew.rst b/doc/source/whatsnew.rst
index 8f779e01a6be5..64cbe0b050a61 100644
--- a/doc/source/whatsnew.rst
+++ b/doc/source/whatsnew.rst
@@ -18,6 +18,8 @@ What's New
These are new features and improvements of note in each release.
+.. include:: whatsnew/v0.22.0.txt
+
.. include:: whatsnew/v0.21.1.txt
.. include:: whatsnew/v0.21.0.txt
diff --git a/doc/source/whatsnew/v0.22.0.txt b/doc/source/whatsnew/v0.22.0.txt
index 53b052a955b45..da4acd99e3873 100644
--- a/doc/source/whatsnew/v0.22.0.txt
+++ b/doc/source/whatsnew/v0.22.0.txt
@@ -1,156 +1,220 @@
.. _whatsnew_0220:
-v0.22.0
--------
+v0.22.0 (December 29, 2017)
+---------------------------
-This is a major release from 0.21.1 and includes a number of API changes,
-deprecations, new features, enhancements, and performance improvements along
-with a large number of bug fixes. We recommend that all users upgrade to this
-version.
+This is a major release from 0.21.1 and includes a single, API-breaking change.
+We recommend that all users upgrade to this version after carefully reading the
+release note (singular!).
-.. _whatsnew_0220.enhancements:
+.. _whatsnew_0220.api_breaking:
-New features
-~~~~~~~~~~~~
+Backwards incompatible API changes
+~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
--
--
--
+Pandas 0.22.0 changes the handling of empty and all-*NA* sums and products. The
+summary is that
-.. _whatsnew_0220.enhancements.other:
+* The sum of an empty or all-*NA* ``Series`` is now ``0``
+* The product of an empty or all-*NA* ``Series`` is now ``1``
+* We've added a ``min_count`` parameter to ``.sum()`` and ``.prod()`` controlling
+ the minimum number of valid values for the result to be valid. If fewer than
+ ``min_count`` non-*NA* values are present, the result is *NA*. The default is
+ ``0``. To return ``NaN``, the 0.21 behavior, use ``min_count=1``.
-Other Enhancements
-^^^^^^^^^^^^^^^^^^
+Some background: In pandas 0.21, we fixed a long-standing inconsistency
+in the return value of all-*NA* series depending on whether or not bottleneck
+was installed. See :ref:`whatsnew_0210.api_breaking.bottleneck`. At the same
+time, we changed the sum and prod of an empty ``Series`` to also be ``NaN``.
--
--
--
+Based on feedback, we've partially reverted those changes.
-.. _whatsnew_0220.api_breaking:
+Arithmetic Operations
+^^^^^^^^^^^^^^^^^^^^^
-Backwards incompatible API changes
-~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
+The default sum for empty or all-*NA* ``Series`` is now ``0``.
--
--
--
+*pandas 0.21.x*
-.. _whatsnew_0220.api:
+.. code-block:: ipython
-Other API Changes
-^^^^^^^^^^^^^^^^^
+ In [1]: pd.Series([]).sum()
+ Out[1]: nan
--
--
--
+ In [2]: pd.Series([np.nan]).sum()
+ Out[2]: nan
-.. _whatsnew_0220.deprecations:
+*pandas 0.22.0*
-Deprecations
-~~~~~~~~~~~~
+.. ipython:: python
--
--
--
+ pd.Series([]).sum()
+ pd.Series([np.nan]).sum()
-.. _whatsnew_0220.prior_deprecations:
+The default behavior is the same as pandas 0.20.3 with bottleneck installed. It
+also matches the behavior of NumPy's ``np.nansum`` on empty and all-*NA* arrays.
-Removal of prior version deprecations/changes
-~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
+To have the sum of an empty series return ``NaN`` (the default behavior of
+pandas 0.20.3 without bottleneck, or pandas 0.21.x), use the ``min_count``
+keyword.
--
--
--
+.. ipython:: python
-.. _whatsnew_0220.performance:
+ pd.Series([]).sum(min_count=1)
-Performance Improvements
-~~~~~~~~~~~~~~~~~~~~~~~~
+Thanks to the ``skipna`` parameter, the ``.sum`` on an all-*NA*
+series is conceptually the same as the ``.sum`` of an empty one with
+``skipna=True`` (the default).
--
--
--
+.. ipython:: python
-.. _whatsnew_0220.docs:
+ pd.Series([np.nan]).sum(min_count=1) # skipna=True by default
-Documentation Changes
-~~~~~~~~~~~~~~~~~~~~~
+The ``min_count`` parameter refers to the minimum number of *non-null* values
+required for a non-NA sum or product.
--
--
--
+:meth:`Series.prod` has been updated to behave the same as :meth:`Series.sum`,
+returning ``1`` instead.
-.. _whatsnew_0220.bug_fixes:
+.. ipython:: python
-Bug Fixes
-~~~~~~~~~
+ pd.Series([]).prod()
+ pd.Series([np.nan]).prod()
+ pd.Series([]).prod(min_count=1)
-Conversion
-^^^^^^^^^^
+These changes affect :meth:`DataFrame.sum` and :meth:`DataFrame.prod` as well.
+Finally, a few less obvious places in pandas are affected by this change.
--
--
--
+Grouping by a Categorical
+^^^^^^^^^^^^^^^^^^^^^^^^^
-Indexing
-^^^^^^^^
+Grouping by a ``Categorical`` and summing now returns ``0`` instead of
+``NaN`` for categories with no observations. The product now returns ``1``
+instead of ``NaN``.
+
+*pandas 0.21.x*
+
+.. code-block:: ipython
--
--
--
+ In [8]: grouper = pd.Categorical(['a', 'a'], categories=['a', 'b'])
-I/O
-^^^
+ In [9]: pd.Series([1, 2]).groupby(grouper).sum()
+ Out[9]:
+ a 3.0
+ b NaN
+ dtype: float64
--
--
--
+*pandas 0.22*
-Plotting
+.. ipython:: python
+
+ grouper = pd.Categorical(['a', 'a'], categories=['a', 'b'])
+ pd.Series([1, 2]).groupby(grouper).sum()
+
+To restore the 0.21 behavior of returning ``NaN`` for unobserved groups,
+use ``min_count>=1``.
+
+.. ipython:: python
+
+ pd.Series([1, 2]).groupby(grouper).sum(min_count=1)
+
+Resample
^^^^^^^^
--
--
--
+The sum and product of all-*NA* bins has changed from ``NaN`` to ``0`` for
+sum and ``1`` for product.
+
+*pandas 0.21.x*
+
+.. code-block:: ipython
+
+ In [11]: s = pd.Series([1, 1, np.nan, np.nan],
+ ...: index=pd.date_range('2017', periods=4))
+ ...: s
+ Out[11]:
+ 2017-01-01 1.0
+ 2017-01-02 1.0
+ 2017-01-03 NaN
+ 2017-01-04 NaN
+ Freq: D, dtype: float64
+
+ In [12]: s.resample('2d').sum()
+ Out[12]:
+ 2017-01-01 2.0
+ 2017-01-03 NaN
+ Freq: 2D, dtype: float64
+
+*pandas 0.22.0*
+
+.. ipython:: python
+
+ s = pd.Series([1, 1, np.nan, np.nan],
+ index=pd.date_range('2017', periods=4))
+ s.resample('2d').sum()
+
+To restore the 0.21 behavior of returning ``NaN``, use ``min_count>=1``.
+
+.. ipython:: python
+
+ s.resample('2d').sum(min_count=1)
+
+In particular, upsampling and taking the sum or product is affected, as
+upsampling introduces missing values even if the original series was
+entirely valid.
+
+*pandas 0.21.x*
+
+.. code-block:: ipython
+
+ In [14]: idx = pd.DatetimeIndex(['2017-01-01', '2017-01-02'])
+
+ In [15]: pd.Series([1, 2], index=idx).resample('12H').sum()
+ Out[15]:
+ 2017-01-01 00:00:00 1.0
+ 2017-01-01 12:00:00 NaN
+ 2017-01-02 00:00:00 2.0
+ Freq: 12H, dtype: float64
+
+*pandas 0.22.0*
+
+.. ipython:: python
+
+ idx = pd.DatetimeIndex(['2017-01-01', '2017-01-02'])
+ pd.Series([1, 2], index=idx).resample("12H").sum()
+
+Once again, the ``min_count`` keyword is available to restore the 0.21 behavior.
-Groupby/Resample/Rolling
-^^^^^^^^^^^^^^^^^^^^^^^^
+.. ipython:: python
--
--
--
+ pd.Series([1, 2], index=idx).resample("12H").sum(min_count=1)
-Sparse
-^^^^^^
+Rolling and Expanding
+^^^^^^^^^^^^^^^^^^^^^
--
--
--
+Rolling and expanding already have a ``min_periods`` keyword that behaves
+similar to ``min_count``. The only case that changes is when doing a rolling
+or expanding sum with ``min_periods=0``. Previously this returned ``NaN``,
+when fewer than ``min_periods`` non-*NA* values were in the window. Now it
+returns ``0``.
-Reshaping
-^^^^^^^^^
+*pandas 0.21.1*
--
--
--
+.. code-block:: ipython
-Numeric
-^^^^^^^
+ In [17]: s = pd.Series([np.nan, np.nan])
--
--
--
+ In [18]: s.rolling(2, min_periods=0).sum()
+ Out[18]:
+ 0 NaN
+ 1 NaN
+ dtype: float64
-Categorical
-^^^^^^^^^^^
+*pandas 0.22.0*
--
--
--
+.. ipython:: python
-Other
-^^^^^
+ s = pd.Series([np.nan, np.nan])
+ s.rolling(2, min_periods=0).sum()
--
--
--
+The default behavior of ``min_periods=None``, implying that ``min_periods``
+equals the window size, is unchanged.
diff --git a/pandas/_libs/groupby_helper.pxi.in b/pandas/_libs/groupby_helper.pxi.in
index d38b677df321c..14d47398ac1df 100644
--- a/pandas/_libs/groupby_helper.pxi.in
+++ b/pandas/_libs/groupby_helper.pxi.in
@@ -36,7 +36,8 @@ def get_dispatch(dtypes):
def group_add_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{c_type}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=0):
"""
Only aggregates on axis=0
"""
@@ -88,7 +89,7 @@ def group_add_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
for i in range(ncounts):
for j in range(K):
- if nobs[i, j] == 0:
+ if nobs[i, j] < min_count:
out[i, j] = NAN
else:
out[i, j] = sumx[i, j]
@@ -99,7 +100,8 @@ def group_add_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_prod_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{c_type}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=0):
"""
Only aggregates on axis=0
"""
@@ -147,7 +149,7 @@ def group_prod_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
for i in range(ncounts):
for j in range(K):
- if nobs[i, j] == 0:
+ if nobs[i, j] < min_count:
out[i, j] = NAN
else:
out[i, j] = prodx[i, j]
@@ -159,12 +161,15 @@ def group_prod_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_var_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{dest_type2}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
cdef:
Py_ssize_t i, j, N, K, lab, ncounts = len(counts)
{{dest_type2}} val, ct, oldmean
ndarray[{{dest_type2}}, ndim=2] nobs, mean
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if not len(values) == len(labels):
raise AssertionError("len(index) != len(labels)")
@@ -208,12 +213,15 @@ def group_var_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_mean_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{dest_type2}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
cdef:
Py_ssize_t i, j, N, K, lab, ncounts = len(counts)
{{dest_type2}} val, count
ndarray[{{dest_type2}}, ndim=2] sumx, nobs
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if not len(values) == len(labels):
raise AssertionError("len(index) != len(labels)")
@@ -263,7 +271,8 @@ def group_mean_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_ohlc_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{dest_type2}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
"""
Only aggregates on axis=0
"""
@@ -272,6 +281,8 @@ def group_ohlc_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
{{dest_type2}} val, count
Py_ssize_t ngroups = len(counts)
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if len(labels) == 0:
return
@@ -332,7 +343,8 @@ def get_dispatch(dtypes):
def group_last_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{c_type}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
"""
Only aggregates on axis=0
"""
@@ -342,6 +354,8 @@ def group_last_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[{{dest_type2}}, ndim=2] resx
ndarray[int64_t, ndim=2] nobs
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if not len(values) == len(labels):
raise AssertionError("len(index) != len(labels)")
@@ -382,7 +396,8 @@ def group_last_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_nth_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{c_type}}, ndim=2] values,
- ndarray[int64_t] labels, int64_t rank):
+ ndarray[int64_t] labels, int64_t rank,
+ Py_ssize_t min_count=-1):
"""
Only aggregates on axis=0
"""
@@ -392,6 +407,8 @@ def group_nth_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[{{dest_type2}}, ndim=2] resx
ndarray[int64_t, ndim=2] nobs
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if not len(values) == len(labels):
raise AssertionError("len(index) != len(labels)")
@@ -455,7 +472,8 @@ def get_dispatch(dtypes):
def group_max_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{dest_type2}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
"""
Only aggregates on axis=0
"""
@@ -464,6 +482,8 @@ def group_max_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
{{dest_type2}} val, count
ndarray[{{dest_type2}}, ndim=2] maxx, nobs
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if not len(values) == len(labels):
raise AssertionError("len(index) != len(labels)")
@@ -526,7 +546,8 @@ def group_max_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_min_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
ndarray[int64_t] counts,
ndarray[{{dest_type2}}, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
"""
Only aggregates on axis=0
"""
@@ -535,6 +556,8 @@ def group_min_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
{{dest_type2}} val, count
ndarray[{{dest_type2}}, ndim=2] minx, nobs
+ assert min_count == -1, "'min_count' only used in add and prod"
+
if not len(values) == len(labels):
raise AssertionError("len(index) != len(labels)")
@@ -686,7 +709,8 @@ def group_cummax_{{name}}(ndarray[{{dest_type2}}, ndim=2] out,
def group_median_float64(ndarray[float64_t, ndim=2] out,
ndarray[int64_t] counts,
ndarray[float64_t, ndim=2] values,
- ndarray[int64_t] labels):
+ ndarray[int64_t] labels,
+ Py_ssize_t min_count=-1):
"""
Only aggregates on axis=0
"""
@@ -695,6 +719,9 @@ def group_median_float64(ndarray[float64_t, ndim=2] out,
ndarray[int64_t] _counts
ndarray data
float64_t* ptr
+
+ assert min_count == -1, "'min_count' only used in add and prod"
+
ngroups = len(counts)
N, K = (<object> values).shape
diff --git a/pandas/_libs/window.pyx b/pandas/_libs/window.pyx
index a1c4ddbc8d0b0..3a7a6d54d3851 100644
--- a/pandas/_libs/window.pyx
+++ b/pandas/_libs/window.pyx
@@ -225,14 +225,16 @@ cdef class VariableWindowIndexer(WindowIndexer):
right_closed: bint
right endpoint closedness
True if the right endpoint is closed, False if open
-
+ floor: optional
+ unit for flooring the unit
"""
def __init__(self, ndarray input, int64_t win, int64_t minp,
- bint left_closed, bint right_closed, ndarray index):
+ bint left_closed, bint right_closed, ndarray index,
+ object floor=None):
self.is_variable = 1
self.N = len(index)
- self.minp = _check_minp(win, minp, self.N)
+ self.minp = _check_minp(win, minp, self.N, floor=floor)
self.start = np.empty(self.N, dtype='int64')
self.start.fill(-1)
@@ -347,7 +349,7 @@ def get_window_indexer(input, win, minp, index, closed,
if index is not None:
indexer = VariableWindowIndexer(input, win, minp, left_closed,
- right_closed, index)
+ right_closed, index, floor)
elif use_mock:
indexer = MockFixedWindowIndexer(input, win, minp, left_closed,
right_closed, index, floor)
@@ -446,7 +448,7 @@ def roll_sum(ndarray[double_t] input, int64_t win, int64_t minp,
object index, object closed):
cdef:
double val, prev_x, sum_x = 0
- int64_t s, e
+ int64_t s, e, range_endpoint
int64_t nobs = 0, i, j, N
bint is_variable
ndarray[int64_t] start, end
@@ -454,7 +456,8 @@ def roll_sum(ndarray[double_t] input, int64_t win, int64_t minp,
start, end, N, win, minp, is_variable = get_window_indexer(input, win,
minp, index,
- closed)
+ closed,
+ floor=0)
output = np.empty(N, dtype=float)
# for performance we are going to iterate
@@ -494,13 +497,15 @@ def roll_sum(ndarray[double_t] input, int64_t win, int64_t minp,
# fixed window
+ range_endpoint = int_max(minp, 1) - 1
+
with nogil:
- for i in range(0, minp - 1):
+ for i in range(0, range_endpoint):
add_sum(input[i], &nobs, &sum_x)
output[i] = NaN
- for i in range(minp - 1, N):
+ for i in range(range_endpoint, N):
val = input[i]
add_sum(val, &nobs, &sum_x)
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index 58d86251a4a62..9a2a763cf6def 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -6921,7 +6921,8 @@ def _add_numeric_operations(cls):
@Substitution(outname='mad',
desc="Return the mean absolute deviation of the values "
"for the requested axis",
- name1=name, name2=name2, axis_descr=axis_descr)
+ name1=name, name2=name2, axis_descr=axis_descr,
+ min_count='', examples='')
@Appender(_num_doc)
def mad(self, axis=None, skipna=None, level=None):
if skipna is None:
@@ -6962,7 +6963,8 @@ def mad(self, axis=None, skipna=None, level=None):
@Substitution(outname='compounded',
desc="Return the compound percentage of the values for "
"the requested axis", name1=name, name2=name2,
- axis_descr=axis_descr)
+ axis_descr=axis_descr,
+ min_count='', examples='')
@Appender(_num_doc)
def compound(self, axis=None, skipna=None, level=None):
if skipna is None:
@@ -6986,10 +6988,10 @@ def compound(self, axis=None, skipna=None, level=None):
lambda y, axis: np.maximum.accumulate(y, axis), "max",
-np.inf, np.nan)
- cls.sum = _make_stat_function(
+ cls.sum = _make_min_count_stat_function(
cls, 'sum', name, name2, axis_descr,
'Return the sum of the values for the requested axis',
- nanops.nansum)
+ nanops.nansum, _sum_examples)
cls.mean = _make_stat_function(
cls, 'mean', name, name2, axis_descr,
'Return the mean of the values for the requested axis',
@@ -7005,10 +7007,10 @@ def compound(self, axis=None, skipna=None, level=None):
"by N-1\n",
nanops.nankurt)
cls.kurtosis = cls.kurt
- cls.prod = _make_stat_function(
+ cls.prod = _make_min_count_stat_function(
cls, 'prod', name, name2, axis_descr,
'Return the product of the values for the requested axis',
- nanops.nanprod)
+ nanops.nanprod, _prod_examples)
cls.product = cls.prod
cls.median = _make_stat_function(
cls, 'median', name, name2, axis_descr,
@@ -7139,10 +7141,13 @@ def _doc_parms(cls):
numeric_only : boolean, default None
Include only float, int, boolean columns. If None, will attempt to use
everything, then use only numeric data. Not implemented for Series.
+%(min_count)s\
Returns
-------
-%(outname)s : %(name1)s or %(name2)s (if level specified)\n"""
+%(outname)s : %(name1)s or %(name2)s (if level specified)
+
+%(examples)s"""
_num_ddof_doc = """
@@ -7210,9 +7215,92 @@ def _doc_parms(cls):
"""
+_sum_examples = """\
+Examples
+--------
+By default, the sum of an empty or all-NA Series is ``0``.
+
+>>> pd.Series([]).sum() # min_count=0 is the default
+0.0
+
+This can be controlled with the ``min_count`` parameter. For example, if
+you'd like the sum of an empty series to be NaN, pass ``min_count=1``.
+
+>>> pd.Series([]).sum(min_count=1)
+nan
+
+Thanks to the ``skipna`` parameter, ``min_count`` handles all-NA and
+empty series identically.
+
+>>> pd.Series([np.nan]).sum()
+0.0
+
+>>> pd.Series([np.nan]).sum(min_count=1)
+nan
+"""
+
+_prod_examples = """\
+Examples
+--------
+By default, the product of an empty or all-NA Series is ``1``
+
+>>> pd.Series([]).prod()
+1.0
+
+This can be controlled with the ``min_count`` parameter
+
+>>> pd.Series([]).prod(min_count=1)
+nan
+
+Thanks to the ``skipna`` parameter, ``min_count`` handles all-NA and
+empty series identically.
+
+>>> pd.Series([np.nan]).prod()
+1.0
+
+>>> pd.Series([np.nan]).sum(min_count=1)
+nan
+"""
+
+
+_min_count_stub = """\
+min_count : int, default 1
+ The required number of valid values to perform the operation. If fewer than
+ ``min_count`` non-NA values are present the result will be NA.
+
+ .. versionadded :: 0.21.2
+
+ Added with the default being 1. This means the sum or product
+ of an all-NA or empty series is ``NaN``.
+"""
+
+
+def _make_min_count_stat_function(cls, name, name1, name2, axis_descr, desc,
+ f, examples):
+ @Substitution(outname=name, desc=desc, name1=name1, name2=name2,
+ axis_descr=axis_descr, min_count=_min_count_stub,
+ examples=examples)
+ @Appender(_num_doc)
+ def stat_func(self, axis=None, skipna=None, level=None, numeric_only=None,
+ min_count=0,
+ **kwargs):
+ nv.validate_stat_func(tuple(), kwargs, fname=name)
+ if skipna is None:
+ skipna = True
+ if axis is None:
+ axis = self._stat_axis_number
+ if level is not None:
+ return self._agg_by_level(name, axis=axis, level=level,
+ skipna=skipna, min_count=min_count)
+ return self._reduce(f, name, axis=axis, skipna=skipna,
+ numeric_only=numeric_only, min_count=min_count)
+
+ return set_function_name(stat_func, name, cls)
+
+
def _make_stat_function(cls, name, name1, name2, axis_descr, desc, f):
@Substitution(outname=name, desc=desc, name1=name1, name2=name2,
- axis_descr=axis_descr)
+ axis_descr=axis_descr, min_count='', examples='')
@Appender(_num_doc)
def stat_func(self, axis=None, skipna=None, level=None, numeric_only=None,
**kwargs):
diff --git a/pandas/core/groupby.py b/pandas/core/groupby.py
index 5931f6e009dab..aef5ff7ba64d3 100644
--- a/pandas/core/groupby.py
+++ b/pandas/core/groupby.py
@@ -908,7 +908,8 @@ def _cython_transform(self, how, numeric_only=True):
return self._wrap_transformed_output(output, names)
- def _cython_agg_general(self, how, alt=None, numeric_only=True):
+ def _cython_agg_general(self, how, alt=None, numeric_only=True,
+ min_count=-1):
output = {}
for name, obj in self._iterate_slices():
is_numeric = is_numeric_dtype(obj.dtype)
@@ -916,7 +917,8 @@ def _cython_agg_general(self, how, alt=None, numeric_only=True):
continue
try:
- result, names = self.grouper.aggregate(obj.values, how)
+ result, names = self.grouper.aggregate(obj.values, how,
+ min_count=min_count)
except AssertionError as e:
raise GroupByError(str(e))
output[name] = self._try_cast(result, obj)
@@ -1223,7 +1225,8 @@ def _add_numeric_operations(cls):
""" add numeric operations to the GroupBy generically """
def groupby_function(name, alias, npfunc,
- numeric_only=True, _convert=False):
+ numeric_only=True, _convert=False,
+ min_count=-1):
_local_template = "Compute %(f)s of group values"
@@ -1233,6 +1236,8 @@ def groupby_function(name, alias, npfunc,
def f(self, **kwargs):
if 'numeric_only' not in kwargs:
kwargs['numeric_only'] = numeric_only
+ if 'min_count' not in kwargs:
+ kwargs['min_count'] = min_count
self._set_group_selection()
try:
return self._cython_agg_general(
@@ -1280,8 +1285,8 @@ def last(x):
else:
return last(x)
- cls.sum = groupby_function('sum', 'add', np.sum)
- cls.prod = groupby_function('prod', 'prod', np.prod)
+ cls.sum = groupby_function('sum', 'add', np.sum, min_count=0)
+ cls.prod = groupby_function('prod', 'prod', np.prod, min_count=0)
cls.min = groupby_function('min', 'min', np.min, numeric_only=False)
cls.max = groupby_function('max', 'max', np.max, numeric_only=False)
cls.first = groupby_function('first', 'first', first_compat,
@@ -2107,7 +2112,7 @@ def get_group_levels(self):
'var': 'group_var',
'first': {
'name': 'group_nth',
- 'f': lambda func, a, b, c, d: func(a, b, c, d, 1)
+ 'f': lambda func, a, b, c, d, e: func(a, b, c, d, 1, -1)
},
'last': 'group_last',
'ohlc': 'group_ohlc',
@@ -2177,7 +2182,7 @@ def wrapper(*args, **kwargs):
(how, dtype_str))
return func, dtype_str
- def _cython_operation(self, kind, values, how, axis):
+ def _cython_operation(self, kind, values, how, axis, min_count=-1):
assert kind in ['transform', 'aggregate']
# can we do this operation with our cython functions
@@ -2262,11 +2267,12 @@ def _cython_operation(self, kind, values, how, axis):
counts = np.zeros(self.ngroups, dtype=np.int64)
result = self._aggregate(
result, counts, values, labels, func, is_numeric,
- is_datetimelike)
+ is_datetimelike, min_count)
elif kind == 'transform':
result = _maybe_fill(np.empty_like(values, dtype=out_dtype),
fill_value=np.nan)
+ # TODO: min_count
result = self._transform(
result, values, labels, func, is_numeric, is_datetimelike)
@@ -2303,14 +2309,15 @@ def _cython_operation(self, kind, values, how, axis):
return result, names
- def aggregate(self, values, how, axis=0):
- return self._cython_operation('aggregate', values, how, axis)
+ def aggregate(self, values, how, axis=0, min_count=-1):
+ return self._cython_operation('aggregate', values, how, axis,
+ min_count=min_count)
def transform(self, values, how, axis=0):
return self._cython_operation('transform', values, how, axis)
def _aggregate(self, result, counts, values, comp_ids, agg_func,
- is_numeric, is_datetimelike):
+ is_numeric, is_datetimelike, min_count=-1):
if values.ndim > 3:
# punting for now
raise NotImplementedError("number of dimensions is currently "
@@ -2319,9 +2326,10 @@ def _aggregate(self, result, counts, values, comp_ids, agg_func,
for i, chunk in enumerate(values.transpose(2, 0, 1)):
chunk = chunk.squeeze()
- agg_func(result[:, :, i], counts, chunk, comp_ids)
+ agg_func(result[:, :, i], counts, chunk, comp_ids,
+ min_count)
else:
- agg_func(result, counts, values, comp_ids)
+ agg_func(result, counts, values, comp_ids, min_count)
return result
@@ -3595,9 +3603,10 @@ def _iterate_slices(self):
continue
yield val, slicer(val)
- def _cython_agg_general(self, how, alt=None, numeric_only=True):
+ def _cython_agg_general(self, how, alt=None, numeric_only=True,
+ min_count=-1):
new_items, new_blocks = self._cython_agg_blocks(
- how, alt=alt, numeric_only=numeric_only)
+ how, alt=alt, numeric_only=numeric_only, min_count=min_count)
return self._wrap_agged_blocks(new_items, new_blocks)
def _wrap_agged_blocks(self, items, blocks):
@@ -3623,7 +3632,8 @@ def _wrap_agged_blocks(self, items, blocks):
_block_agg_axis = 0
- def _cython_agg_blocks(self, how, alt=None, numeric_only=True):
+ def _cython_agg_blocks(self, how, alt=None, numeric_only=True,
+ min_count=-1):
# TODO: the actual managing of mgr_locs is a PITA
# here, it should happen via BlockManager.combine
@@ -3640,7 +3650,7 @@ def _cython_agg_blocks(self, how, alt=None, numeric_only=True):
locs = block.mgr_locs.as_array
try:
result, _ = self.grouper.aggregate(
- block.values, how, axis=agg_axis)
+ block.values, how, axis=agg_axis, min_count=min_count)
except NotImplementedError:
# generally if we have numeric_only=False
# and non-applicable functions
diff --git a/pandas/core/nanops.py b/pandas/core/nanops.py
index e1c09947ac0b4..d1a355021f388 100644
--- a/pandas/core/nanops.py
+++ b/pandas/core/nanops.py
@@ -107,21 +107,14 @@ def f(values, axis=None, skipna=True, **kwds):
if k not in kwds:
kwds[k] = v
try:
- if values.size == 0:
-
- # we either return np.nan or pd.NaT
- if is_numeric_dtype(values):
- values = values.astype('float64')
- fill_value = na_value_for_dtype(values.dtype)
-
- if values.ndim == 1:
- return fill_value
- else:
- result_shape = (values.shape[:axis] +
- values.shape[axis + 1:])
- result = np.empty(result_shape, dtype=values.dtype)
- result.fill(fill_value)
- return result
+ if values.size == 0 and kwds.get('min_count') is None:
+ # We are empty, returning NA for our type
+ # Only applies for the default `min_count` of None
+ # since that affects how empty arrays are handled.
+ # TODO(GH-18976) update all the nanops methods to
+ # correctly handle empty inputs and remove this check.
+ # It *may* just be `var`
+ return _na_for_min_count(values, axis)
if (_USE_BOTTLENECK and skipna and
_bn_ok_dtype(values.dtype, bn_name)):
@@ -292,6 +285,36 @@ def _wrap_results(result, dtype):
return result
+def _na_for_min_count(values, axis):
+ """Return the missing value for `values`
+
+ Parameters
+ ----------
+ values : ndarray
+ axis : int or None
+ axis for the reduction
+
+ Returns
+ -------
+ result : scalar or ndarray
+ For 1-D values, returns a scalar of the correct missing type.
+ For 2-D values, returns a 1-D array where each element is missing.
+ """
+ # we either return np.nan or pd.NaT
+ if is_numeric_dtype(values):
+ values = values.astype('float64')
+ fill_value = na_value_for_dtype(values.dtype)
+
+ if values.ndim == 1:
+ return fill_value
+ else:
+ result_shape = (values.shape[:axis] +
+ values.shape[axis + 1:])
+ result = np.empty(result_shape, dtype=values.dtype)
+ result.fill(fill_value)
+ return result
+
+
def nanany(values, axis=None, skipna=True):
values, mask, dtype, _ = _get_values(values, skipna, False, copy=skipna)
return values.any(axis)
@@ -304,7 +327,7 @@ def nanall(values, axis=None, skipna=True):
@disallow('M8')
@bottleneck_switch()
-def nansum(values, axis=None, skipna=True):
+def nansum(values, axis=None, skipna=True, min_count=0):
values, mask, dtype, dtype_max = _get_values(values, skipna, 0)
dtype_sum = dtype_max
if is_float_dtype(dtype):
@@ -312,7 +335,7 @@ def nansum(values, axis=None, skipna=True):
elif is_timedelta64_dtype(dtype):
dtype_sum = np.float64
the_sum = values.sum(axis, dtype=dtype_sum)
- the_sum = _maybe_null_out(the_sum, axis, mask)
+ the_sum = _maybe_null_out(the_sum, axis, mask, min_count=min_count)
return _wrap_results(the_sum, dtype)
@@ -641,13 +664,13 @@ def nankurt(values, axis=None, skipna=True):
@disallow('M8', 'm8')
-def nanprod(values, axis=None, skipna=True):
+def nanprod(values, axis=None, skipna=True, min_count=0):
mask = isna(values)
if skipna and not is_any_int_dtype(values):
values = values.copy()
values[mask] = 1
result = values.prod(axis)
- return _maybe_null_out(result, axis, mask)
+ return _maybe_null_out(result, axis, mask, min_count=min_count)
def _maybe_arg_null_out(result, axis, mask, skipna):
@@ -683,9 +706,9 @@ def _get_counts(mask, axis, dtype=float):
return np.array(count, dtype=dtype)
-def _maybe_null_out(result, axis, mask):
+def _maybe_null_out(result, axis, mask, min_count=1):
if axis is not None and getattr(result, 'ndim', False):
- null_mask = (mask.shape[axis] - mask.sum(axis)) == 0
+ null_mask = (mask.shape[axis] - mask.sum(axis) - min_count) < 0
if np.any(null_mask):
if is_numeric_dtype(result):
if np.iscomplexobj(result):
@@ -698,7 +721,7 @@ def _maybe_null_out(result, axis, mask):
result[null_mask] = None
elif result is not tslib.NaT:
null_mask = mask.size - mask.sum()
- if null_mask == 0:
+ if null_mask < min_count:
result = np.nan
return result
diff --git a/pandas/core/resample.py b/pandas/core/resample.py
index 1adb3a078cca3..db1d3d4c5e31b 100644
--- a/pandas/core/resample.py
+++ b/pandas/core/resample.py
@@ -601,9 +601,20 @@ def size(self):
Resampler._deprecated_valids += dir(Resampler)
+
+# downsample methods
+for method in ['sum', 'prod']:
+
+ def f(self, _method=method, min_count=0, *args, **kwargs):
+ nv.validate_resampler_func(_method, args, kwargs)
+ return self._downsample(_method, min_count=min_count)
+ f.__doc__ = getattr(GroupBy, method).__doc__
+ setattr(Resampler, method, f)
+
+
# downsample methods
-for method in ['min', 'max', 'first', 'last', 'sum', 'mean', 'sem',
- 'median', 'prod', 'ohlc']:
+for method in ['min', 'max', 'first', 'last', 'mean', 'sem',
+ 'median', 'ohlc']:
def f(self, _method=method, *args, **kwargs):
nv.validate_resampler_func(_method, args, kwargs)
diff --git a/pandas/tests/frame/test_analytics.py b/pandas/tests/frame/test_analytics.py
index 1bac4037e99c9..97ab0deb50d50 100644
--- a/pandas/tests/frame/test_analytics.py
+++ b/pandas/tests/frame/test_analytics.py
@@ -440,7 +440,8 @@ def test_nunique(self):
Series({0: 1, 1: 3, 2: 2}))
def test_sum(self):
- self._check_stat_op('sum', np.sum, has_numeric_only=True)
+ self._check_stat_op('sum', np.sum, has_numeric_only=True,
+ skipna_alternative=np.nansum)
# mixed types (with upcasting happening)
self._check_stat_op('sum', np.sum,
@@ -716,7 +717,8 @@ def alt(x):
def _check_stat_op(self, name, alternative, frame=None, has_skipna=True,
has_numeric_only=False, check_dtype=True,
- check_dates=False, check_less_precise=False):
+ check_dates=False, check_less_precise=False,
+ skipna_alternative=None):
if frame is None:
frame = self.frame
# set some NAs
@@ -737,15 +739,11 @@ def _check_stat_op(self, name, alternative, frame=None, has_skipna=True,
assert len(result)
if has_skipna:
- def skipna_wrapper(x):
- nona = x.dropna()
- if len(nona) == 0:
- return np.nan
- return alternative(nona)
-
def wrapper(x):
return alternative(x.values)
+ skipna_wrapper = tm._make_skipna_wrapper(alternative,
+ skipna_alternative)
result0 = f(axis=0, skipna=False)
result1 = f(axis=1, skipna=False)
tm.assert_series_equal(result0, frame.apply(wrapper),
@@ -797,8 +795,11 @@ def wrapper(x):
r0 = getattr(all_na, name)(axis=0)
r1 = getattr(all_na, name)(axis=1)
if name in ['sum', 'prod']:
- assert np.isnan(r0).all()
- assert np.isnan(r1).all()
+ unit = int(name == 'prod')
+ expected = pd.Series(unit, index=r0.index, dtype=r0.dtype)
+ tm.assert_series_equal(r0, expected)
+ expected = pd.Series(unit, index=r1.index, dtype=r1.dtype)
+ tm.assert_series_equal(r1, expected)
def test_mode(self):
df = pd.DataFrame({"A": [12, 12, 11, 12, 19, 11],
@@ -936,6 +937,66 @@ def test_sum_corner(self):
assert len(axis0) == 0
assert len(axis1) == 0
+ @pytest.mark.parametrize('method, unit', [
+ ('sum', 0),
+ ('prod', 1),
+ ])
+ def test_sum_prod_nanops(self, method, unit):
+ idx = ['a', 'b', 'c']
+ df = pd.DataFrame({"a": [unit, unit],
+ "b": [unit, np.nan],
+ "c": [np.nan, np.nan]})
+ # The default
+ result = getattr(df, method)
+ expected = pd.Series([unit, unit, unit], index=idx, dtype='float64')
+
+ # min_count=1
+ result = getattr(df, method)(min_count=1)
+ expected = pd.Series([unit, unit, np.nan], index=idx)
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = getattr(df, method)(min_count=0)
+ expected = pd.Series([unit, unit, unit], index=idx, dtype='float64')
+ tm.assert_series_equal(result, expected)
+
+ result = getattr(df.iloc[1:], method)(min_count=1)
+ expected = pd.Series([unit, np.nan, np.nan], index=idx)
+ tm.assert_series_equal(result, expected)
+
+ # min_count > 1
+ df = pd.DataFrame({"A": [unit] * 10, "B": [unit] * 5 + [np.nan] * 5})
+ result = getattr(df, method)(min_count=5)
+ expected = pd.Series(result, index=['A', 'B'])
+ tm.assert_series_equal(result, expected)
+
+ result = getattr(df, method)(min_count=6)
+ expected = pd.Series(result, index=['A', 'B'])
+ tm.assert_series_equal(result, expected)
+
+ def test_sum_nanops_timedelta(self):
+ # prod isn't defined on timedeltas
+ idx = ['a', 'b', 'c']
+ df = pd.DataFrame({"a": [0, 0],
+ "b": [0, np.nan],
+ "c": [np.nan, np.nan]})
+
+ df2 = df.apply(pd.to_timedelta)
+
+ # 0 by default
+ result = df2.sum()
+ expected = pd.Series([0, 0, 0], dtype='m8[ns]', index=idx)
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = df2.sum(min_count=0)
+ tm.assert_series_equal(result, expected)
+
+ # min_count=1
+ result = df2.sum(min_count=1)
+ expected = pd.Series([0, 0, np.nan], dtype='m8[ns]', index=idx)
+ tm.assert_series_equal(result, expected)
+
def test_sum_object(self):
values = self.frame.values.astype(int)
frame = DataFrame(values, index=self.frame.index,
diff --git a/pandas/tests/groupby/test_aggregate.py b/pandas/tests/groupby/test_aggregate.py
index 913d3bcc09869..ad1a322fdaae9 100644
--- a/pandas/tests/groupby/test_aggregate.py
+++ b/pandas/tests/groupby/test_aggregate.py
@@ -809,26 +809,60 @@ def test__cython_agg_general(self):
exc.args += ('operation: %s' % op, )
raise
- def test_cython_agg_empty_buckets(self):
- ops = [('mean', np.mean),
- ('median', lambda x: np.median(x) if len(x) > 0 else np.nan),
- ('var', lambda x: np.var(x, ddof=1)),
- ('add', lambda x: np.sum(x) if len(x) > 0 else np.nan),
- ('prod', np.prod),
- ('min', np.min),
- ('max', np.max), ]
-
+ @pytest.mark.parametrize('op, targop', [
+ ('mean', np.mean),
+ ('median', lambda x: np.median(x) if len(x) > 0 else np.nan),
+ ('var', lambda x: np.var(x, ddof=1)),
+ ('min', np.min),
+ ('max', np.max), ]
+ )
+ def test_cython_agg_empty_buckets(self, op, targop):
df = pd.DataFrame([11, 12, 13])
grps = range(0, 55, 5)
- for op, targop in ops:
- result = df.groupby(pd.cut(df[0], grps))._cython_agg_general(op)
- expected = df.groupby(pd.cut(df[0], grps)).agg(lambda x: targop(x))
- try:
- tm.assert_frame_equal(result, expected)
- except BaseException as exc:
- exc.args += ('operation: %s' % op,)
- raise
+ # calling _cython_agg_general directly, instead of via the user API
+ # which sets different values for min_count, so do that here.
+ result = df.groupby(pd.cut(df[0], grps))._cython_agg_general(op)
+ expected = df.groupby(pd.cut(df[0], grps)).agg(lambda x: targop(x))
+ try:
+ tm.assert_frame_equal(result, expected)
+ except BaseException as exc:
+ exc.args += ('operation: %s' % op,)
+ raise
+
+ def test_cython_agg_empty_buckets_nanops(self):
+ # GH-18869 can't call nanops on empty groups, so hardcode expected
+ # for these
+ df = pd.DataFrame([11, 12, 13], columns=['a'])
+ grps = range(0, 25, 5)
+ # add / sum
+ result = df.groupby(pd.cut(df['a'], grps))._cython_agg_general('add')
+ intervals = pd.interval_range(0, 20, freq=5)
+ expected = pd.DataFrame(
+ {"a": [0, 0, 36, 0]},
+ index=pd.CategoricalIndex(intervals, name='a', ordered=True))
+ tm.assert_frame_equal(result, expected)
+
+ # prod
+ result = df.groupby(pd.cut(df['a'], grps))._cython_agg_general('prod')
+ expected = pd.DataFrame(
+ {"a": [1, 1, 1716, 1]},
+ index=pd.CategoricalIndex(intervals, name='a', ordered=True))
+ tm.assert_frame_equal(result, expected)
+
+ @pytest.mark.xfail(reason="GH-18869: agg func not called on empty groups.")
+ def test_agg_category_nansum(self):
+ categories = ['a', 'b', 'c']
+ df = pd.DataFrame({"A": pd.Categorical(['a', 'a', 'b'],
+ categories=categories),
+ 'B': [1, 2, 3]})
+ result = df.groupby("A").B.agg(np.nansum)
+ expected = pd.Series([3, 3, 0],
+ index=pd.CategoricalIndex(['a', 'b', 'c'],
+ categories=categories,
+ name='A'),
+ name='B')
+ tm.assert_series_equal(result, expected)
def test_agg_over_numpy_arrays(self):
# GH 3788
diff --git a/pandas/tests/groupby/test_categorical.py b/pandas/tests/groupby/test_categorical.py
index fdc03acd3e931..d4f35aa8755d1 100644
--- a/pandas/tests/groupby/test_categorical.py
+++ b/pandas/tests/groupby/test_categorical.py
@@ -17,6 +17,142 @@
class TestGroupByCategorical(MixIn):
+ def test_groupby(self):
+
+ cats = Categorical(["a", "a", "a", "b", "b", "b", "c", "c", "c"],
+ categories=["a", "b", "c", "d"], ordered=True)
+ data = DataFrame({"a": [1, 1, 1, 2, 2, 2, 3, 4, 5], "b": cats})
+
+ exp_index = CategoricalIndex(list('abcd'), name='b', ordered=True)
+ expected = DataFrame({'a': [1, 2, 4, np.nan]}, index=exp_index)
+ result = data.groupby("b").mean()
+ tm.assert_frame_equal(result, expected)
+
+ raw_cat1 = Categorical(["a", "a", "b", "b"],
+ categories=["a", "b", "z"], ordered=True)
+ raw_cat2 = Categorical(["c", "d", "c", "d"],
+ categories=["c", "d", "y"], ordered=True)
+ df = DataFrame({"A": raw_cat1, "B": raw_cat2, "values": [1, 2, 3, 4]})
+
+ # single grouper
+ gb = df.groupby("A")
+ exp_idx = CategoricalIndex(['a', 'b', 'z'], name='A', ordered=True)
+ expected = DataFrame({'values': Series([3, 7, 0], index=exp_idx)})
+ result = gb.sum()
+ tm.assert_frame_equal(result, expected)
+
+ # multiple groupers
+ gb = df.groupby(['A', 'B'])
+ exp_index = pd.MultiIndex.from_product(
+ [Categorical(["a", "b", "z"], ordered=True),
+ Categorical(["c", "d", "y"], ordered=True)],
+ names=['A', 'B'])
+ expected = DataFrame({'values': [1, 2, np.nan, 3, 4, np.nan,
+ np.nan, np.nan, np.nan]},
+ index=exp_index)
+ result = gb.sum()
+ tm.assert_frame_equal(result, expected)
+
+ # multiple groupers with a non-cat
+ df = df.copy()
+ df['C'] = ['foo', 'bar'] * 2
+ gb = df.groupby(['A', 'B', 'C'])
+ exp_index = pd.MultiIndex.from_product(
+ [Categorical(["a", "b", "z"], ordered=True),
+ Categorical(["c", "d", "y"], ordered=True),
+ ['foo', 'bar']],
+ names=['A', 'B', 'C'])
+ expected = DataFrame({'values': Series(
+ np.nan, index=exp_index)}).sort_index()
+ expected.iloc[[1, 2, 7, 8], 0] = [1, 2, 3, 4]
+ result = gb.sum()
+ tm.assert_frame_equal(result, expected)
+
+ # GH 8623
+ x = DataFrame([[1, 'John P. Doe'], [2, 'Jane Dove'],
+ [1, 'John P. Doe']],
+ columns=['person_id', 'person_name'])
+ x['person_name'] = Categorical(x.person_name)
+
+ g = x.groupby(['person_id'])
+ result = g.transform(lambda x: x)
+ tm.assert_frame_equal(result, x[['person_name']])
+
+ result = x.drop_duplicates('person_name')
+ expected = x.iloc[[0, 1]]
+ tm.assert_frame_equal(result, expected)
+
+ def f(x):
+ return x.drop_duplicates('person_name').iloc[0]
+
+ result = g.apply(f)
+ expected = x.iloc[[0, 1]].copy()
+ expected.index = Index([1, 2], name='person_id')
+ expected['person_name'] = expected['person_name'].astype('object')
+ tm.assert_frame_equal(result, expected)
+
+ # GH 9921
+ # Monotonic
+ df = DataFrame({"a": [5, 15, 25]})
+ c = pd.cut(df.a, bins=[0, 10, 20, 30, 40])
+
+ result = df.a.groupby(c).transform(sum)
+ tm.assert_series_equal(result, df['a'])
+
+ tm.assert_series_equal(
+ df.a.groupby(c).transform(lambda xs: np.sum(xs)), df['a'])
+ tm.assert_frame_equal(df.groupby(c).transform(sum), df[['a']])
+ tm.assert_frame_equal(
+ df.groupby(c).transform(lambda xs: np.max(xs)), df[['a']])
+
+ # Filter
+ tm.assert_series_equal(df.a.groupby(c).filter(np.all), df['a'])
+ tm.assert_frame_equal(df.groupby(c).filter(np.all), df)
+
+ # Non-monotonic
+ df = DataFrame({"a": [5, 15, 25, -5]})
+ c = pd.cut(df.a, bins=[-10, 0, 10, 20, 30, 40])
+
+ result = df.a.groupby(c).transform(sum)
+ tm.assert_series_equal(result, df['a'])
+
+ tm.assert_series_equal(
+ df.a.groupby(c).transform(lambda xs: np.sum(xs)), df['a'])
+ tm.assert_frame_equal(df.groupby(c).transform(sum), df[['a']])
+ tm.assert_frame_equal(
+ df.groupby(c).transform(lambda xs: np.sum(xs)), df[['a']])
+
+ # GH 9603
+ df = DataFrame({'a': [1, 0, 0, 0]})
+ c = pd.cut(df.a, [0, 1, 2, 3, 4], labels=Categorical(list('abcd')))
+ result = df.groupby(c).apply(len)
+
+ exp_index = CategoricalIndex(
+ c.values.categories, ordered=c.values.ordered)
+ expected = Series([1, 0, 0, 0], index=exp_index)
+ expected.index.name = 'a'
+ tm.assert_series_equal(result, expected)
+
+ def test_groupby_sort(self):
+
+ # http://stackoverflow.com/questions/23814368/sorting-pandas-categorical-labels-after-groupby
+ # This should result in a properly sorted Series so that the plot
+ # has a sorted x axis
+ # self.cat.groupby(['value_group'])['value_group'].count().plot(kind='bar')
+
+ df = DataFrame({'value': np.random.randint(0, 10000, 100)})
+ labels = ["{0} - {1}".format(i, i + 499) for i in range(0, 10000, 500)]
+ cat_labels = Categorical(labels, labels)
+
+ df = df.sort_values(by=['value'], ascending=True)
+ df['value_group'] = pd.cut(df.value, range(0, 10500, 500),
+ right=False, labels=cat_labels)
+
+ res = df.groupby(['value_group'])['value_group'].count()
+ exp = res[sorted(res.index, key=lambda x: float(x.split()[0]))]
+ exp.index = CategoricalIndex(exp.index, name=exp.index.name)
+ tm.assert_series_equal(res, exp)
+
def test_level_groupby_get_group(self):
# GH15155
df = DataFrame(data=np.arange(2, 22, 2),
@@ -526,3 +662,53 @@ def test_groupby_categorical_two_columns(self):
"C3": [nan, nan, nan, nan, 10, 100,
nan, nan, nan, nan, 200, 34]}, index=idx)
tm.assert_frame_equal(res, exp)
+
+ def test_empty_sum(self):
+ # https://github.com/pandas-dev/pandas/issues/18678
+ df = pd.DataFrame({"A": pd.Categorical(['a', 'a', 'b'],
+ categories=['a', 'b', 'c']),
+ 'B': [1, 2, 1]})
+ expected_idx = pd.CategoricalIndex(['a', 'b', 'c'], name='A')
+
+ # 0 by default
+ result = df.groupby("A").B.sum()
+ expected = pd.Series([3, 1, 0], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = df.groupby("A").B.sum(min_count=0)
+ expected = pd.Series([3, 1, 0], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
+
+ # min_count=1
+ result = df.groupby("A").B.sum(min_count=1)
+ expected = pd.Series([3, 1, np.nan], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
+
+ # min_count>1
+ result = df.groupby("A").B.sum(min_count=2)
+ expected = pd.Series([3, np.nan, np.nan], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
+
+ def test_empty_prod(self):
+ # https://github.com/pandas-dev/pandas/issues/18678
+ df = pd.DataFrame({"A": pd.Categorical(['a', 'a', 'b'],
+ categories=['a', 'b', 'c']),
+ 'B': [1, 2, 1]})
+
+ expected_idx = pd.CategoricalIndex(['a', 'b', 'c'], name='A')
+
+ # 1 by default
+ result = df.groupby("A").B.prod()
+ expected = pd.Series([2, 1, 1], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = df.groupby("A").B.prod(min_count=0)
+ expected = pd.Series([2, 1, 1], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
+
+ # min_count=1
+ result = df.groupby("A").B.prod(min_count=1)
+ expected = pd.Series([2, 1, np.nan], expected_idx, name='B')
+ tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/groupby/test_groupby.py b/pandas/tests/groupby/test_groupby.py
index 675f8d6413b2a..7a5581c897231 100644
--- a/pandas/tests/groupby/test_groupby.py
+++ b/pandas/tests/groupby/test_groupby.py
@@ -3850,7 +3850,7 @@ def h(df, arg3):
# Assert the results here
index = pd.Index(['A', 'B', 'C'], name='group')
- expected = pd.Series([-79.5160891089, -78.4839108911, None],
+ expected = pd.Series([-79.5160891089, -78.4839108911, -80],
index=index)
assert_series_equal(expected, result)
diff --git a/pandas/tests/groupby/test_timegrouper.py b/pandas/tests/groupby/test_timegrouper.py
index c8503b16a0e16..d359bfa5351a9 100644
--- a/pandas/tests/groupby/test_timegrouper.py
+++ b/pandas/tests/groupby/test_timegrouper.py
@@ -41,12 +41,11 @@ def test_groupby_with_timegrouper(self):
df = df.set_index(['Date'])
expected = DataFrame(
- {'Quantity': np.nan},
+ {'Quantity': 0},
index=date_range('20130901 13:00:00',
'20131205 13:00:00', freq='5D',
name='Date', closed='left'))
- expected.iloc[[0, 6, 18], 0] = np.array(
- [24., 6., 9.], dtype='float64')
+ expected.iloc[[0, 6, 18], 0] = np.array([24, 6, 9], dtype='int64')
result1 = df.resample('5D') .sum()
assert_frame_equal(result1, expected)
@@ -245,6 +244,8 @@ def test_timegrouper_with_reg_groups(self):
result = df.groupby([pd.Grouper(freq='1M', key='Date')]).sum()
assert_frame_equal(result, expected)
+ @pytest.mark.parametrize('freq', ['D', 'M', 'A', 'Q-APR'])
+ def test_timegrouper_with_reg_groups_freq(self, freq):
# GH 6764 multiple grouping with/without sort
df = DataFrame({
'date': pd.to_datetime([
@@ -258,20 +259,24 @@ def test_timegrouper_with_reg_groups(self):
'cost1': [12, 15, 10, 24, 39, 1, 0, 90, 45, 34, 1, 12]
}).set_index('date')
- for freq in ['D', 'M', 'A', 'Q-APR']:
- expected = df.groupby('user_id')[
- 'whole_cost'].resample(
- freq).sum().dropna().reorder_levels(
- ['date', 'user_id']).sort_index().astype('int64')
- expected.name = 'whole_cost'
-
- result1 = df.sort_index().groupby([pd.Grouper(freq=freq),
- 'user_id'])['whole_cost'].sum()
- assert_series_equal(result1, expected)
-
- result2 = df.groupby([pd.Grouper(freq=freq), 'user_id'])[
- 'whole_cost'].sum()
- assert_series_equal(result2, expected)
+ expected = (
+ df.groupby('user_id')['whole_cost']
+ .resample(freq)
+ .sum(min_count=1) # XXX
+ .dropna()
+ .reorder_levels(['date', 'user_id'])
+ .sort_index()
+ .astype('int64')
+ )
+ expected.name = 'whole_cost'
+
+ result1 = df.sort_index().groupby([pd.Grouper(freq=freq),
+ 'user_id'])['whole_cost'].sum()
+ assert_series_equal(result1, expected)
+
+ result2 = df.groupby([pd.Grouper(freq=freq), 'user_id'])[
+ 'whole_cost'].sum()
+ assert_series_equal(result2, expected)
def test_timegrouper_get_group(self):
# GH 6914
diff --git a/pandas/tests/series/test_analytics.py b/pandas/tests/series/test_analytics.py
index 2ee404ab5fe0d..d6db2ab83098b 100644
--- a/pandas/tests/series/test_analytics.py
+++ b/pandas/tests/series/test_analytics.py
@@ -28,40 +28,124 @@
class TestSeriesAnalytics(TestData):
@pytest.mark.parametrize("use_bottleneck", [True, False])
- @pytest.mark.parametrize("method", ["sum", "prod"])
- def test_empty(self, method, use_bottleneck):
-
+ @pytest.mark.parametrize("method, unit", [
+ ("sum", 0.0),
+ ("prod", 1.0)
+ ])
+ def test_empty(self, method, unit, use_bottleneck):
with pd.option_context("use_bottleneck", use_bottleneck):
- # GH 9422
- # treat all missing as NaN
+ # GH 9422 / 18921
+ # Entirely empty
s = Series([])
+ # NA by default
result = getattr(s, method)()
+ assert result == unit
+
+ # Explict
+ result = getattr(s, method)(min_count=0)
+ assert result == unit
+
+ result = getattr(s, method)(min_count=1)
assert isna(result)
+ # Skipna, default
result = getattr(s, method)(skipna=True)
+ result == unit
+
+ # Skipna, explicit
+ result = getattr(s, method)(skipna=True, min_count=0)
+ assert result == unit
+
+ result = getattr(s, method)(skipna=True, min_count=1)
assert isna(result)
+ # All-NA
s = Series([np.nan])
+ # NA by default
result = getattr(s, method)()
+ assert result == unit
+
+ # Explicit
+ result = getattr(s, method)(min_count=0)
+ assert result == unit
+
+ result = getattr(s, method)(min_count=1)
assert isna(result)
+ # Skipna, default
result = getattr(s, method)(skipna=True)
+ result == unit
+
+ # skipna, explicit
+ result = getattr(s, method)(skipna=True, min_count=0)
+ assert result == unit
+
+ result = getattr(s, method)(skipna=True, min_count=1)
assert isna(result)
+ # Mix of valid, empty
s = Series([np.nan, 1])
+ # Default
result = getattr(s, method)()
assert result == 1.0
- s = Series([np.nan, 1])
+ # Explicit
+ result = getattr(s, method)(min_count=0)
+ assert result == 1.0
+
+ result = getattr(s, method)(min_count=1)
+ assert result == 1.0
+
+ # Skipna
result = getattr(s, method)(skipna=True)
assert result == 1.0
+ result = getattr(s, method)(skipna=True, min_count=0)
+ assert result == 1.0
+
+ result = getattr(s, method)(skipna=True, min_count=1)
+ assert result == 1.0
+
# GH #844 (changed in 9422)
df = DataFrame(np.empty((10, 0)))
- assert (df.sum(1).isnull()).all()
+ assert (getattr(df, method)(1) == unit).all()
+
+ s = pd.Series([1])
+ result = getattr(s, method)(min_count=2)
+ assert isna(result)
+
+ s = pd.Series([np.nan])
+ result = getattr(s, method)(min_count=2)
+ assert isna(result)
+
+ s = pd.Series([np.nan, 1])
+ result = getattr(s, method)(min_count=2)
+ assert isna(result)
+
+ @pytest.mark.parametrize('method, unit', [
+ ('sum', 0.0),
+ ('prod', 1.0),
+ ])
+ def test_empty_multi(self, method, unit):
+ s = pd.Series([1, np.nan, np.nan, np.nan],
+ index=pd.MultiIndex.from_product([('a', 'b'), (0, 1)]))
+ # 1 / 0 by default
+ result = getattr(s, method)(level=0)
+ expected = pd.Series([1, unit], index=['a', 'b'])
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = getattr(s, method)(level=0, min_count=0)
+ expected = pd.Series([1, unit], index=['a', 'b'])
+ tm.assert_series_equal(result, expected)
+
+ # min_count=1
+ result = getattr(s, method)(level=0, min_count=1)
+ expected = pd.Series([1, np.nan], index=['a', 'b'])
+ tm.assert_series_equal(result, expected)
@pytest.mark.parametrize(
- "method", ['sum', 'mean', 'median', 'std', 'var'])
+ "method", ['mean', 'median', 'std', 'var'])
def test_ops_consistency_on_empty(self, method):
# GH 7869
@@ -109,7 +193,7 @@ def test_sum_overflow(self, use_bottleneck):
assert np.allclose(float(result), v[-1])
def test_sum(self):
- self._check_stat_op('sum', np.sum, check_allna=True)
+ self._check_stat_op('sum', np.sum, check_allna=False)
def test_sum_inf(self):
s = Series(np.random.randn(10))
diff --git a/pandas/tests/series/test_quantile.py b/pandas/tests/series/test_quantile.py
index cf5e3fe4f29b0..255367523a3d8 100644
--- a/pandas/tests/series/test_quantile.py
+++ b/pandas/tests/series/test_quantile.py
@@ -38,7 +38,7 @@ def test_quantile(self):
# GH7661
result = Series([np.timedelta64('NaT')]).sum()
- assert result is pd.NaT
+ assert result == pd.Timedelta(0)
msg = 'percentiles should all be in the interval \\[0, 1\\]'
for invalid in [-1, 2, [0.5, -1], [0.5, 2]]:
diff --git a/pandas/tests/test_categorical.py b/pandas/tests/test_categorical.py
index 6366aae8ccdf6..48c1622aa0c4e 100644
--- a/pandas/tests/test_categorical.py
+++ b/pandas/tests/test_categorical.py
@@ -3163,18 +3163,6 @@ def test_info(self):
buf = compat.StringIO()
df2.info(buf=buf)
- def test_groupby_sort(self):
-
- # http://stackoverflow.com/questions/23814368/sorting-pandas-categorical-labels-after-groupby
- # This should result in a properly sorted Series so that the plot
- # has a sorted x axis
- # self.cat.groupby(['value_group'])['value_group'].count().plot(kind='bar')
-
- res = self.cat.groupby(['value_group'])['value_group'].count()
- exp = res[sorted(res.index, key=lambda x: float(x.split()[0]))]
- exp.index = pd.CategoricalIndex(exp.index, name=exp.index.name)
- tm.assert_series_equal(res, exp)
-
def test_min_max(self):
# unordered cats have no min/max
cat = Series(Categorical(["a", "b", "c", "d"], ordered=False))
@@ -3294,123 +3282,6 @@ def test_value_counts_with_nan(self):
res = s.value_counts(dropna=False, sort=False)
tm.assert_series_equal(res, exp)
- def test_groupby(self):
-
- cats = Categorical(["a", "a", "a", "b", "b", "b", "c", "c", "c"],
- categories=["a", "b", "c", "d"], ordered=True)
- data = DataFrame({"a": [1, 1, 1, 2, 2, 2, 3, 4, 5], "b": cats})
-
- exp_index = pd.CategoricalIndex(['a', 'b', 'c', 'd'], name='b',
- ordered=True)
- expected = DataFrame({'a': [1, 2, 4, np.nan]}, index=exp_index)
- result = data.groupby("b").mean()
- tm.assert_frame_equal(result, expected)
-
- raw_cat1 = Categorical(["a", "a", "b", "b"],
- categories=["a", "b", "z"], ordered=True)
- raw_cat2 = Categorical(["c", "d", "c", "d"],
- categories=["c", "d", "y"], ordered=True)
- df = DataFrame({"A": raw_cat1, "B": raw_cat2, "values": [1, 2, 3, 4]})
-
- # single grouper
- gb = df.groupby("A")
- exp_idx = pd.CategoricalIndex(['a', 'b', 'z'], name='A', ordered=True)
- expected = DataFrame({'values': Series([3, 7, np.nan], index=exp_idx)})
- result = gb.sum()
- tm.assert_frame_equal(result, expected)
-
- # multiple groupers
- gb = df.groupby(['A', 'B'])
- exp_index = pd.MultiIndex.from_product(
- [Categorical(["a", "b", "z"], ordered=True),
- Categorical(["c", "d", "y"], ordered=True)],
- names=['A', 'B'])
- expected = DataFrame({'values': [1, 2, np.nan, 3, 4, np.nan,
- np.nan, np.nan, np.nan]},
- index=exp_index)
- result = gb.sum()
- tm.assert_frame_equal(result, expected)
-
- # multiple groupers with a non-cat
- df = df.copy()
- df['C'] = ['foo', 'bar'] * 2
- gb = df.groupby(['A', 'B', 'C'])
- exp_index = pd.MultiIndex.from_product(
- [Categorical(["a", "b", "z"], ordered=True),
- Categorical(["c", "d", "y"], ordered=True),
- ['foo', 'bar']],
- names=['A', 'B', 'C'])
- expected = DataFrame({'values': Series(
- np.nan, index=exp_index)}).sort_index()
- expected.iloc[[1, 2, 7, 8], 0] = [1, 2, 3, 4]
- result = gb.sum()
- tm.assert_frame_equal(result, expected)
-
- # GH 8623
- x = pd.DataFrame([[1, 'John P. Doe'], [2, 'Jane Dove'],
- [1, 'John P. Doe']],
- columns=['person_id', 'person_name'])
- x['person_name'] = pd.Categorical(x.person_name)
-
- g = x.groupby(['person_id'])
- result = g.transform(lambda x: x)
- tm.assert_frame_equal(result, x[['person_name']])
-
- result = x.drop_duplicates('person_name')
- expected = x.iloc[[0, 1]]
- tm.assert_frame_equal(result, expected)
-
- def f(x):
- return x.drop_duplicates('person_name').iloc[0]
-
- result = g.apply(f)
- expected = x.iloc[[0, 1]].copy()
- expected.index = Index([1, 2], name='person_id')
- expected['person_name'] = expected['person_name'].astype('object')
- tm.assert_frame_equal(result, expected)
-
- # GH 9921
- # Monotonic
- df = DataFrame({"a": [5, 15, 25]})
- c = pd.cut(df.a, bins=[0, 10, 20, 30, 40])
-
- result = df.a.groupby(c).transform(sum)
- tm.assert_series_equal(result, df['a'])
-
- tm.assert_series_equal(
- df.a.groupby(c).transform(lambda xs: np.sum(xs)), df['a'])
- tm.assert_frame_equal(df.groupby(c).transform(sum), df[['a']])
- tm.assert_frame_equal(
- df.groupby(c).transform(lambda xs: np.max(xs)), df[['a']])
-
- # Filter
- tm.assert_series_equal(df.a.groupby(c).filter(np.all), df['a'])
- tm.assert_frame_equal(df.groupby(c).filter(np.all), df)
-
- # Non-monotonic
- df = DataFrame({"a": [5, 15, 25, -5]})
- c = pd.cut(df.a, bins=[-10, 0, 10, 20, 30, 40])
-
- result = df.a.groupby(c).transform(sum)
- tm.assert_series_equal(result, df['a'])
-
- tm.assert_series_equal(
- df.a.groupby(c).transform(lambda xs: np.sum(xs)), df['a'])
- tm.assert_frame_equal(df.groupby(c).transform(sum), df[['a']])
- tm.assert_frame_equal(
- df.groupby(c).transform(lambda xs: np.sum(xs)), df[['a']])
-
- # GH 9603
- df = pd.DataFrame({'a': [1, 0, 0, 0]})
- c = pd.cut(df.a, [0, 1, 2, 3, 4], labels=pd.Categorical(list('abcd')))
- result = df.groupby(c).apply(len)
-
- exp_index = pd.CategoricalIndex(c.values.categories,
- ordered=c.values.ordered)
- expected = pd.Series([1, 0, 0, 0], index=exp_index)
- expected.index.name = 'a'
- tm.assert_series_equal(result, expected)
-
def test_pivot_table(self):
raw_cat1 = Categorical(["a", "a", "b", "b"],
diff --git a/pandas/tests/test_expressions.py b/pandas/tests/test_expressions.py
index 6d2607962dfb0..aebc9cd3deaac 100644
--- a/pandas/tests/test_expressions.py
+++ b/pandas/tests/test_expressions.py
@@ -73,17 +73,11 @@ def teardown_method(self, method):
def run_arithmetic(self, df, other, assert_func, check_dtype=False,
test_flex=True):
expr._MIN_ELEMENTS = 0
- operations = ['add', 'sub', 'mul', 'mod', 'truediv', 'floordiv', 'pow']
+ operations = ['add', 'sub', 'mul', 'mod', 'truediv', 'floordiv']
if not compat.PY3:
operations.append('div')
for arith in operations:
- # numpy >= 1.11 doesn't handle integers
- # raised to integer powers
- # https://github.com/pandas-dev/pandas/issues/15363
- if arith == 'pow' and not _np_version_under1p11:
- continue
-
operator_name = arith
if arith == 'div':
operator_name = 'truediv'
diff --git a/pandas/tests/test_nanops.py b/pandas/tests/test_nanops.py
index 9305504f8d5e3..5d56088193d30 100644
--- a/pandas/tests/test_nanops.py
+++ b/pandas/tests/test_nanops.py
@@ -1,6 +1,7 @@
# -*- coding: utf-8 -*-
from __future__ import division, print_function
+from distutils.version import LooseVersion
from functools import partial
import pytest
@@ -181,12 +182,17 @@ def _coerce_tds(targ, res):
check_dtype=check_dtype)
def check_fun_data(self, testfunc, targfunc, testarval, targarval,
- targarnanval, check_dtype=True, **kwargs):
+ targarnanval, check_dtype=True, empty_targfunc=None,
+ **kwargs):
for axis in list(range(targarval.ndim)) + [None]:
for skipna in [False, True]:
targartempval = targarval if skipna else targarnanval
- try:
+ if skipna and empty_targfunc and isna(targartempval).all():
+ targ = empty_targfunc(targartempval, axis=axis, **kwargs)
+ else:
targ = targfunc(targartempval, axis=axis, **kwargs)
+
+ try:
res = testfunc(testarval, axis=axis, skipna=skipna,
**kwargs)
self.check_results(targ, res, axis,
@@ -218,10 +224,11 @@ def check_fun_data(self, testfunc, targfunc, testarval, targarval,
except ValueError:
return
self.check_fun_data(testfunc, targfunc, testarval2, targarval2,
- targarnanval2, check_dtype=check_dtype, **kwargs)
+ targarnanval2, check_dtype=check_dtype,
+ empty_targfunc=empty_targfunc, **kwargs)
def check_fun(self, testfunc, targfunc, testar, targar=None,
- targarnan=None, **kwargs):
+ targarnan=None, empty_targfunc=None, **kwargs):
if targar is None:
targar = testar
if targarnan is None:
@@ -231,7 +238,8 @@ def check_fun(self, testfunc, targfunc, testar, targar=None,
targarnanval = getattr(self, targarnan)
try:
self.check_fun_data(testfunc, targfunc, testarval, targarval,
- targarnanval, **kwargs)
+ targarnanval, empty_targfunc=empty_targfunc,
+ **kwargs)
except BaseException as exc:
exc.args += ('testar: %s' % testar, 'targar: %s' % targar,
'targarnan: %s' % targarnan)
@@ -328,7 +336,8 @@ def test_nanall(self):
def test_nansum(self):
self.check_funs(nanops.nansum, np.sum, allow_str=False,
- allow_date=False, allow_tdelta=True, check_dtype=False)
+ allow_date=False, allow_tdelta=True, check_dtype=False,
+ empty_targfunc=np.nansum)
def test_nanmean(self):
self.check_funs(nanops.nanmean, np.mean, allow_complex=False,
@@ -461,8 +470,12 @@ def test_nankurt(self):
allow_tdelta=False)
def test_nanprod(self):
+ if LooseVersion(np.__version__) < LooseVersion("1.10.0"):
+ raise pytest.skip("np.nanprod added in 1.10.0")
+
self.check_funs(nanops.nanprod, np.prod, allow_str=False,
- allow_date=False, allow_tdelta=False)
+ allow_date=False, allow_tdelta=False,
+ empty_targfunc=np.nanprod)
def check_nancorr_nancov_2d(self, checkfun, targ0, targ1, **kwargs):
res00 = checkfun(self.arr_float_2d, self.arr_float1_2d, **kwargs)
diff --git a/pandas/tests/test_panel.py b/pandas/tests/test_panel.py
index 33fb6f1108bf2..7e442fcc2fc8b 100644
--- a/pandas/tests/test_panel.py
+++ b/pandas/tests/test_panel.py
@@ -3,6 +3,7 @@
from warnings import catch_warnings
from datetime import datetime
+from distutils.version import LooseVersion
import operator
import pytest
@@ -10,7 +11,6 @@
import pandas as pd
from pandas.core.dtypes.common import is_float_dtype
-from pandas.core.dtypes.missing import remove_na_arraylike
from pandas import (Series, DataFrame, Index, date_range, isna, notna,
pivot, MultiIndex)
from pandas.core.nanops import nanall, nanany
@@ -83,13 +83,16 @@ def test_count(self):
self._check_stat_op('count', f, obj=self.panel, has_skipna=False)
def test_sum(self):
- self._check_stat_op('sum', np.sum)
+ self._check_stat_op('sum', np.sum, skipna_alternative=np.nansum)
def test_mean(self):
self._check_stat_op('mean', np.mean)
def test_prod(self):
- self._check_stat_op('prod', np.prod)
+ if LooseVersion(np.__version__) < LooseVersion("1.10.0"):
+ raise pytest.skip("np.nanprod added in 1.10.0")
+
+ self._check_stat_op('prod', np.prod, skipna_alternative=np.nanprod)
def test_median(self):
def wrapper(x):
@@ -142,7 +145,8 @@ def alt(x):
self._check_stat_op('sem', alt)
- def _check_stat_op(self, name, alternative, obj=None, has_skipna=True):
+ def _check_stat_op(self, name, alternative, obj=None, has_skipna=True,
+ skipna_alternative=None):
if obj is None:
obj = self.panel
@@ -154,11 +158,8 @@ def _check_stat_op(self, name, alternative, obj=None, has_skipna=True):
if has_skipna:
- def skipna_wrapper(x):
- nona = remove_na_arraylike(x)
- if len(nona) == 0:
- return np.nan
- return alternative(nona)
+ skipna_wrapper = tm._make_skipna_wrapper(alternative,
+ skipna_alternative)
def wrapper(x):
return alternative(np.asarray(x))
diff --git a/pandas/tests/test_panel4d.py b/pandas/tests/test_panel4d.py
index c0e8770dff8b8..ef19f11499e00 100644
--- a/pandas/tests/test_panel4d.py
+++ b/pandas/tests/test_panel4d.py
@@ -4,11 +4,11 @@
import operator
import pytest
from warnings import catch_warnings
+from distutils.version import LooseVersion
import numpy as np
from pandas import Series, Index, isna, notna
from pandas.core.dtypes.common import is_float_dtype
-from pandas.core.dtypes.missing import remove_na_arraylike
from pandas.core.panel import Panel
from pandas.core.panel4d import Panel4D
from pandas.tseries.offsets import BDay
@@ -37,13 +37,16 @@ def test_count(self):
self._check_stat_op('count', f, obj=self.panel4d, has_skipna=False)
def test_sum(self):
- self._check_stat_op('sum', np.sum)
+ self._check_stat_op('sum', np.sum, skipna_alternative=np.nansum)
def test_mean(self):
self._check_stat_op('mean', np.mean)
def test_prod(self):
- self._check_stat_op('prod', np.prod)
+ if LooseVersion(np.__version__) < LooseVersion("1.10.0"):
+ raise pytest.skip("np.nanprod added in 1.10.0")
+
+ self._check_stat_op('prod', np.prod, skipna_alternative=np.nanprod)
def test_median(self):
def wrapper(x):
@@ -106,7 +109,8 @@ def alt(x):
# self._check_stat_op('skew', alt)
- def _check_stat_op(self, name, alternative, obj=None, has_skipna=True):
+ def _check_stat_op(self, name, alternative, obj=None, has_skipna=True,
+ skipna_alternative=None):
if obj is None:
obj = self.panel4d
@@ -117,11 +121,9 @@ def _check_stat_op(self, name, alternative, obj=None, has_skipna=True):
f = getattr(obj, name)
if has_skipna:
- def skipna_wrapper(x):
- nona = remove_na_arraylike(x)
- if len(nona) == 0:
- return np.nan
- return alternative(nona)
+
+ skipna_wrapper = tm._make_skipna_wrapper(alternative,
+ skipna_alternative)
def wrapper(x):
return alternative(np.asarray(x))
diff --git a/pandas/tests/test_resample.py b/pandas/tests/test_resample.py
index e64bf2217e717..04e702644913f 100644
--- a/pandas/tests/test_resample.py
+++ b/pandas/tests/test_resample.py
@@ -4,6 +4,7 @@
from datetime import datetime, timedelta
from functools import partial
from textwrap import dedent
+from operator import methodcaller
import pytz
import pytest
@@ -3377,8 +3378,45 @@ def test_aggregate_normal(self):
assert_frame_equal(expected, dt_result)
"""
- def test_aggregate_with_nat(self):
+ @pytest.mark.parametrize('method, unit', [
+ ('sum', 0),
+ ('prod', 1),
+ ])
+ def test_resample_entirly_nat_window(self, method, unit):
+ s = pd.Series([0] * 2 + [np.nan] * 2,
+ index=pd.date_range('2017', periods=4))
+ # 0 / 1 by default
+ result = methodcaller(method)(s.resample("2d"))
+ expected = pd.Series([0.0, unit],
+ index=pd.to_datetime(['2017-01-01',
+ '2017-01-03']))
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = methodcaller(method, min_count=0)(s.resample("2d"))
+ expected = pd.Series([0.0, unit],
+ index=pd.to_datetime(['2017-01-01',
+ '2017-01-03']))
+ tm.assert_series_equal(result, expected)
+
+ # min_count=1
+ result = methodcaller(method, min_count=1)(s.resample("2d"))
+ expected = pd.Series([0.0, np.nan],
+ index=pd.to_datetime(['2017-01-01',
+ '2017-01-03']))
+ tm.assert_series_equal(result, expected)
+
+ @pytest.mark.parametrize('func, fill_value', [
+ ('min', np.nan),
+ ('max', np.nan),
+ ('sum', 0),
+ ('prod', 1),
+ ('count', 0),
+ ])
+ def test_aggregate_with_nat(self, func, fill_value):
# check TimeGrouper's aggregation is identical as normal groupby
+ # if NaT is included, 'var', 'std', 'mean', 'first','last'
+ # and 'nth' doesn't work yet
n = 20
data = np.random.randn(n, 4).astype('int64')
@@ -3392,42 +3430,42 @@ def test_aggregate_with_nat(self):
normal_grouped = normal_df.groupby('key')
dt_grouped = dt_df.groupby(TimeGrouper(key='key', freq='D'))
- for func in ['min', 'max', 'sum', 'prod']:
- normal_result = getattr(normal_grouped, func)()
- dt_result = getattr(dt_grouped, func)()
- pad = DataFrame([[np.nan, np.nan, np.nan, np.nan]], index=[3],
- columns=['A', 'B', 'C', 'D'])
- expected = normal_result.append(pad)
- expected = expected.sort_index()
- expected.index = date_range(start='2013-01-01', freq='D',
- periods=5, name='key')
- assert_frame_equal(expected, dt_result)
+ normal_result = getattr(normal_grouped, func)()
+ dt_result = getattr(dt_grouped, func)()
- for func in ['count']:
- normal_result = getattr(normal_grouped, func)()
- pad = DataFrame([[0, 0, 0, 0]], index=[3],
- columns=['A', 'B', 'C', 'D'])
- expected = normal_result.append(pad)
- expected = expected.sort_index()
- expected.index = date_range(start='2013-01-01', freq='D',
- periods=5, name='key')
- dt_result = getattr(dt_grouped, func)()
- assert_frame_equal(expected, dt_result)
+ pad = DataFrame([[fill_value] * 4], index=[3],
+ columns=['A', 'B', 'C', 'D'])
+ expected = normal_result.append(pad)
+ expected = expected.sort_index()
+ expected.index = date_range(start='2013-01-01', freq='D',
+ periods=5, name='key')
+ assert_frame_equal(expected, dt_result)
+ assert dt_result.index.name == 'key'
- for func in ['size']:
- normal_result = getattr(normal_grouped, func)()
- pad = Series([0], index=[3])
- expected = normal_result.append(pad)
- expected = expected.sort_index()
- expected.index = date_range(start='2013-01-01', freq='D',
- periods=5, name='key')
- dt_result = getattr(dt_grouped, func)()
- assert_series_equal(expected, dt_result)
- # GH 9925
- assert dt_result.index.name == 'key'
+ def test_aggregate_with_nat_size(self):
+ # GH 9925
+ n = 20
+ data = np.random.randn(n, 4).astype('int64')
+ normal_df = DataFrame(data, columns=['A', 'B', 'C', 'D'])
+ normal_df['key'] = [1, 2, np.nan, 4, 5] * 4
+
+ dt_df = DataFrame(data, columns=['A', 'B', 'C', 'D'])
+ dt_df['key'] = [datetime(2013, 1, 1), datetime(2013, 1, 2), pd.NaT,
+ datetime(2013, 1, 4), datetime(2013, 1, 5)] * 4
+
+ normal_grouped = normal_df.groupby('key')
+ dt_grouped = dt_df.groupby(TimeGrouper(key='key', freq='D'))
- # if NaT is included, 'var', 'std', 'mean', 'first','last'
- # and 'nth' doesn't work yet
+ normal_result = normal_grouped.size()
+ dt_result = dt_grouped.size()
+
+ pad = Series([0], index=[3])
+ expected = normal_result.append(pad)
+ expected = expected.sort_index()
+ expected.index = date_range(start='2013-01-01', freq='D',
+ periods=5, name='key')
+ assert_series_equal(expected, dt_result)
+ assert dt_result.index.name == 'key'
def test_repr(self):
# GH18203
@@ -3436,3 +3474,34 @@ def test_repr(self):
"closed='left', label='left', how='mean', "
"convention='e', base=0)")
assert result == expected
+
+ @pytest.mark.parametrize('method, unit', [
+ ('sum', 0),
+ ('prod', 1),
+ ])
+ def test_upsample_sum(self, method, unit):
+ s = pd.Series(1, index=pd.date_range("2017", periods=2, freq="H"))
+ resampled = s.resample("30T")
+ index = pd.to_datetime(['2017-01-01T00:00:00',
+ '2017-01-01T00:30:00',
+ '2017-01-01T01:00:00'])
+
+ # 0 / 1 by default
+ result = methodcaller(method)(resampled)
+ expected = pd.Series([1, unit, 1], index=index)
+ tm.assert_series_equal(result, expected)
+
+ # min_count=0
+ result = methodcaller(method, min_count=0)(resampled)
+ expected = pd.Series([1, unit, 1], index=index)
+ tm.assert_series_equal(result, expected)
+
+ # min_count=1
+ result = methodcaller(method, min_count=1)(resampled)
+ expected = pd.Series([1, np.nan, 1], index=index)
+ tm.assert_series_equal(result, expected)
+
+ # min_count>1
+ result = methodcaller(method, min_count=2)(resampled)
+ expected = pd.Series([np.nan, np.nan, np.nan], index=index)
+ tm.assert_series_equal(result, expected)
diff --git a/pandas/tests/test_window.py b/pandas/tests/test_window.py
index 35ae4ad4d5db4..e65de10c51300 100644
--- a/pandas/tests/test_window.py
+++ b/pandas/tests/test_window.py
@@ -475,6 +475,28 @@ def tests_empty_df_rolling(self, roller):
result = DataFrame(index=pd.DatetimeIndex([])).rolling(roller).sum()
tm.assert_frame_equal(result, expected)
+ def test_missing_minp_zero(self):
+ # https://github.com/pandas-dev/pandas/pull/18921
+ # minp=0
+ x = pd.Series([np.nan])
+ result = x.rolling(1, min_periods=0).sum()
+ expected = pd.Series([0.0])
+ tm.assert_series_equal(result, expected)
+
+ # minp=1
+ result = x.rolling(1, min_periods=1).sum()
+ expected = pd.Series([np.nan])
+ tm.assert_series_equal(result, expected)
+
+ def test_missing_minp_zero_variable(self):
+ # https://github.com/pandas-dev/pandas/pull/18921
+ x = pd.Series([np.nan] * 4,
+ index=pd.DatetimeIndex(['2017-01-01', '2017-01-04',
+ '2017-01-06', '2017-01-07']))
+ result = x.rolling(pd.Timedelta("2d"), min_periods=0).sum()
+ expected = pd.Series(0.0, index=x.index)
+ tm.assert_series_equal(result, expected)
+
def test_multi_index_names(self):
# GH 16789, 16825
@@ -548,6 +570,19 @@ def test_empty_df_expanding(self, expander):
index=pd.DatetimeIndex([])).expanding(expander).sum()
tm.assert_frame_equal(result, expected)
+ def test_missing_minp_zero(self):
+ # https://github.com/pandas-dev/pandas/pull/18921
+ # minp=0
+ x = pd.Series([np.nan])
+ result = x.expanding(min_periods=0).sum()
+ expected = pd.Series([0.0])
+ tm.assert_series_equal(result, expected)
+
+ # minp=1
+ result = x.expanding(min_periods=1).sum()
+ expected = pd.Series([np.nan])
+ tm.assert_series_equal(result, expected)
+
class TestEWM(Base):
@@ -864,7 +899,8 @@ def test_centered_axis_validation(self):
.rolling(window=3, center=True, axis=2).mean())
def test_rolling_sum(self):
- self._check_moment_func(mom.rolling_sum, np.sum, name='sum')
+ self._check_moment_func(mom.rolling_sum, np.nansum, name='sum',
+ zero_min_periods_equal=False)
def test_rolling_count(self):
counter = lambda x: np.isfinite(x).astype(float).sum()
@@ -1349,14 +1385,18 @@ def test_fperr_robustness(self):
def _check_moment_func(self, f, static_comp, name=None, window=50,
has_min_periods=True, has_center=True,
has_time_rule=True, preserve_nan=True,
- fill_value=None, test_stable=False, **kwargs):
+ fill_value=None, test_stable=False,
+ zero_min_periods_equal=True,
+ **kwargs):
with warnings.catch_warnings(record=True):
self._check_ndarray(f, static_comp, window=window,
has_min_periods=has_min_periods,
preserve_nan=preserve_nan,
has_center=has_center, fill_value=fill_value,
- test_stable=test_stable, **kwargs)
+ test_stable=test_stable,
+ zero_min_periods_equal=zero_min_periods_equal,
+ **kwargs)
with warnings.catch_warnings(record=True):
self._check_structures(f, static_comp,
@@ -1375,7 +1415,8 @@ def _check_moment_func(self, f, static_comp, name=None, window=50,
def _check_ndarray(self, f, static_comp, window=50, has_min_periods=True,
preserve_nan=True, has_center=True, fill_value=None,
- test_stable=False, test_window=True, **kwargs):
+ test_stable=False, test_window=True,
+ zero_min_periods_equal=True, **kwargs):
def get_result(arr, window, min_periods=None, center=False):
return f(arr, window, min_periods=min_periods, center=center, **
kwargs)
@@ -1408,10 +1449,11 @@ def get_result(arr, window, min_periods=None, center=False):
assert isna(result[3])
assert notna(result[4])
- # min_periods=0
- result0 = get_result(arr, 20, min_periods=0)
- result1 = get_result(arr, 20, min_periods=1)
- tm.assert_almost_equal(result0, result1)
+ if zero_min_periods_equal:
+ # min_periods=0 may be equivalent to min_periods=1
+ result0 = get_result(arr, 20, min_periods=0)
+ result1 = get_result(arr, 20, min_periods=1)
+ tm.assert_almost_equal(result0, result1)
else:
result = get_result(arr, 50)
tm.assert_almost_equal(result[-1], static_comp(arr[10:-10]))
diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index dec67bbea854f..b6fc9c78d6476 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -2861,3 +2861,31 @@ def setTZ(tz):
yield
finally:
setTZ(orig_tz)
+
+
+def _make_skipna_wrapper(alternative, skipna_alternative=None):
+ """Create a function for calling on an array.
+
+ Parameters
+ ----------
+ alternative : function
+ The function to be called on the array with no NaNs.
+ Only used when 'skipna_alternative' is None.
+ skipna_alternative : function
+ The function to be called on the original array
+
+ Returns
+ -------
+ skipna_wrapper : function
+ """
+ if skipna_alternative:
+ def skipna_wrapper(x):
+ return skipna_alternative(x.values)
+ else:
+ def skipna_wrapper(x):
+ nona = x.dropna()
+ if len(nona) == 0:
+ return np.nan
+ return alternative(nona)
+
+ return skipna_wrapper
| https://api.github.com/repos/pandas-dev/pandas/pulls/18986 | 2017-12-29T14:29:30Z | 2017-12-30T01:22:59Z | 2017-12-30T01:22:59Z | 2017-12-30T01:42:09Z | |
revert geopandas xfail | diff --git a/pandas/tests/test_downstream.py b/pandas/tests/test_downstream.py
index 6407bee49ad15..0f0abd8cd3400 100644
--- a/pandas/tests/test_downstream.py
+++ b/pandas/tests/test_downstream.py
@@ -92,7 +92,6 @@ def test_pandas_datareader():
pandas_datareader.get_data_google('AAPL')
-@pytest.mark.xfail(reason="install not working, gh-18780")
def test_geopandas():
geopandas = import_module('geopandas') # noqa
| closes #18780
| https://api.github.com/repos/pandas-dev/pandas/pulls/18984 | 2017-12-29T14:22:17Z | 2017-12-29T15:12:42Z | 2017-12-29T15:12:41Z | 2017-12-29T15:12:42Z |
DOC: 0.22.0 release docs | diff --git a/doc/source/release.rst b/doc/source/release.rst
index 0298eda2c78ab..aea6280a490d6 100644
--- a/doc/source/release.rst
+++ b/doc/source/release.rst
@@ -37,6 +37,27 @@ analysis / manipulation tool available in any language.
* Binary installers on PyPI: http://pypi.python.org/pypi/pandas
* Documentation: http://pandas.pydata.org
+pandas 0.22.0
+-------------
+
+**Release date:** December 29, 2017
+
+This is a major release from 0.21.1 and includes a single, API-breaking change.
+We recommend that all users upgrade to this version after carefully reading the
+release note.
+
+The only changes are:
+
+- The sum of an empty or all-*NA* ``Series`` is now ``0``
+- The product of an empty or all-*NA* ``Series`` is now ``1``
+- We've added a ``min_count`` parameter to ``.sum()`` and ``.prod()`` controlling
+ the minimum number of valid values for the result to be valid. If fewer than
+ ``min_count`` non-*NA* values are present, the result is *NA*. The default is
+ ``0``. To return ``NaN``, the 0.21 behavior, use ``min_count=1``.
+
+See the :ref:`v0.22.0 Whatsnew <whatsnew_0220>` overview for further explanation
+of all the places in the library this affects.
+
pandas 0.21.1
-------------
diff --git a/doc/source/whatsnew/v0.22.0.txt b/doc/source/whatsnew/v0.22.0.txt
index 8617aa6c03e1f..da4acd99e3873 100644
--- a/doc/source/whatsnew/v0.22.0.txt
+++ b/doc/source/whatsnew/v0.22.0.txt
@@ -1,7 +1,7 @@
.. _whatsnew_0220:
-v0.22.0
--------
+v0.22.0 (December 29, 2017)
+---------------------------
This is a major release from 0.21.1 and includes a single, API-breaking change.
We recommend that all users upgrade to this version after carefully reading the
| [ci skip] | https://api.github.com/repos/pandas-dev/pandas/pulls/18983 | 2017-12-29T13:19:20Z | 2017-12-29T14:27:47Z | 2017-12-29T14:27:47Z | 2017-12-30T03:36:09Z |
ENH: is_scalar returns True for DateOffset objects | diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 24f3e4433411e..0061a636cafb6 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -144,6 +144,7 @@ Other Enhancements
- :class:`Interval` and :class:`IntervalIndex` have gained a ``length`` attribute (:issue:`18789`)
- ``Resampler`` objects now have a functioning :attr:`~pandas.core.resample.Resampler.pipe` method.
Previously, calls to ``pipe`` were diverted to the ``mean`` method (:issue:`17905`).
+- :func:`~pandas.api.types.is_scalar` now returns ``True`` for ``DateOffset`` objects (:issue:`18943`).
.. _whatsnew_0230.api_breaking:
diff --git a/pandas/_libs/lib.pyx b/pandas/_libs/lib.pyx
index 5a62203f79642..3898f7499e85e 100644
--- a/pandas/_libs/lib.pyx
+++ b/pandas/_libs/lib.pyx
@@ -112,6 +112,7 @@ cpdef bint isscalar(object val):
- Period
- instances of decimal.Decimal
- Interval
+ - DateOffset
"""
@@ -126,7 +127,8 @@ cpdef bint isscalar(object val):
or PyTime_Check(val)
or util.is_period_object(val)
or is_decimal(val)
- or is_interval(val))
+ or is_interval(val)
+ or is_offset(val))
def item_from_zerodim(object val):
diff --git a/pandas/_libs/src/inference.pyx b/pandas/_libs/src/inference.pyx
index 5ed8828a0f122..b74b3a79fd69a 100644
--- a/pandas/_libs/src/inference.pyx
+++ b/pandas/_libs/src/inference.pyx
@@ -45,6 +45,8 @@ cpdef bint is_period(object val):
""" Return a boolean if this is a Period object """
return util.is_period_object(val)
+cdef inline bint is_offset(object val):
+ return getattr(val, '_typ', '_typ') == 'dateoffset'
_TYPE_MAP = {
'categorical': 'categorical',
diff --git a/pandas/tests/dtypes/test_inference.py b/pandas/tests/dtypes/test_inference.py
index e8bdd2a551a34..219d1b2852938 100644
--- a/pandas/tests/dtypes/test_inference.py
+++ b/pandas/tests/dtypes/test_inference.py
@@ -18,7 +18,8 @@
from pandas._libs import tslib, lib, missing as libmissing
from pandas import (Series, Index, DataFrame, Timedelta,
DatetimeIndex, TimedeltaIndex, Timestamp,
- Panel, Period, Categorical, isna)
+ Panel, Period, Categorical, isna, Interval,
+ DateOffset)
from pandas.compat import u, PY2, PY3, StringIO, lrange
from pandas.core.dtypes import inference
from pandas.core.dtypes.common import (
@@ -1151,6 +1152,8 @@ def test_isscalar_pandas_scalars(self):
assert is_scalar(Timestamp('2014-01-01'))
assert is_scalar(Timedelta(hours=1))
assert is_scalar(Period('2014-01-01'))
+ assert is_scalar(Interval(left=0, right=1))
+ assert is_scalar(DateOffset(days=1))
def test_lisscalar_pandas_containers(self):
assert not is_scalar(Series())
| - [x] closes #18943
- [x] tests added / passed
- [x] passes `git diff upstream/master -u -- "*.py" | flake8 --diff`
- [x] whatsnew entry
I've implemented `is_offset` identically to `is_period_object` and added it to `isscalar` and `is_scalar` is now returning true for DateOffsets.
But not sure how to go about this exactly:
> `is_offset` should also be imported / tested in `pandas/core/dtypes/common.py`
The other "is" functions that are explicitly imported from inference `is_string_like` and `is_list_like` are used for testing but don't look like they themselves are being tested so I'm not sure what kind of test is needed. The rest of the "is" functions are imported with a wildcard but pylint is telling me they are not used (should I go through and make the required imports explicit?).
| https://api.github.com/repos/pandas-dev/pandas/pulls/18982 | 2017-12-29T05:50:01Z | 2017-12-29T14:24:39Z | 2017-12-29T14:24:39Z | 2017-12-29T14:28:41Z |
Spellcheck of docs, a few minor changes | diff --git a/doc/source/advanced.rst b/doc/source/advanced.rst
index e591825cec748..be749dfc1f594 100644
--- a/doc/source/advanced.rst
+++ b/doc/source/advanced.rst
@@ -24,9 +24,9 @@ See the :ref:`Indexing and Selecting Data <indexing>` for general indexing docum
Whether a copy or a reference is returned for a setting operation, may
depend on the context. This is sometimes called ``chained assignment`` and
should be avoided. See :ref:`Returning a View versus Copy
- <indexing.view_versus_copy>`
+ <indexing.view_versus_copy>`.
-See the :ref:`cookbook<cookbook.selection>` for some advanced strategies
+See the :ref:`cookbook<cookbook.selection>` for some advanced strategies.
.. _advanced.hierarchical:
@@ -46,7 +46,7 @@ described above and in prior sections. Later, when discussing :ref:`group by
non-trivial applications to illustrate how it aids in structuring data for
analysis.
-See the :ref:`cookbook<cookbook.multi_index>` for some advanced strategies
+See the :ref:`cookbook<cookbook.multi_index>` for some advanced strategies.
Creating a MultiIndex (hierarchical index) object
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@@ -59,7 +59,7 @@ can think of ``MultiIndex`` as an array of tuples where each tuple is unique. A
``MultiIndex.from_tuples``), or a crossed set of iterables (using
``MultiIndex.from_product``). The ``Index`` constructor will attempt to return
a ``MultiIndex`` when it is passed a list of tuples. The following examples
-demo different ways to initialize MultiIndexes.
+demonstrate different ways to initialize MultiIndexes.
.. ipython:: python
@@ -196,7 +196,8 @@ highly performant. If you want to see the actual used levels.
# for a specific level
df[['foo','qux']].columns.get_level_values(0)
-To reconstruct the ``MultiIndex`` with only the used levels
+To reconstruct the ``MultiIndex`` with only the used levels, the
+``remove_unused_levels`` method may be used.
.. versionadded:: 0.20.0
@@ -216,7 +217,7 @@ tuples:
s + s[:-2]
s + s[::2]
-``reindex`` can be called with another ``MultiIndex`` or even a list or array
+``reindex`` can be called with another ``MultiIndex``, or even a list or array
of tuples:
.. ipython:: python
@@ -230,7 +231,7 @@ Advanced indexing with hierarchical index
-----------------------------------------
Syntactically integrating ``MultiIndex`` in advanced indexing with ``.loc`` is a
-bit challenging, but we've made every effort to do so. for example the
+bit challenging, but we've made every effort to do so. For example the
following works as you would expect:
.. ipython:: python
@@ -286,7 +287,7 @@ As usual, **both sides** of the slicers are included as this is label indexing.
df.loc[(slice('A1','A3'),.....), :]
- rather than this:
+ You should **not** do this:
.. code-block:: python
@@ -315,7 +316,7 @@ Basic multi-index slicing using slices, lists, and labels.
dfmi.loc[(slice('A1','A3'), slice(None), ['C1', 'C3']), :]
-You can use a ``pd.IndexSlice`` to have a more natural syntax using ``:`` rather than using ``slice(None)``
+You can use :class:`pandas.IndexSlice` to facilitate a more natural syntax using ``:``, rather than using ``slice(None)``.
.. ipython:: python
@@ -344,7 +345,7 @@ slicers on a single axis.
dfmi.loc(axis=0)[:, :, ['C1', 'C3']]
-Furthermore you can *set* the values using these methods
+Furthermore you can *set* the values using the following methods.
.. ipython:: python
@@ -379,7 +380,7 @@ selecting data at a particular level of a MultiIndex easier.
df.loc[(slice(None),'one'),:]
You can also select on the columns with :meth:`~pandas.MultiIndex.xs`, by
-providing the axis argument
+providing the axis argument.
.. ipython:: python
@@ -391,7 +392,7 @@ providing the axis argument
# using the slicers
df.loc[:,(slice(None),'one')]
-:meth:`~pandas.MultiIndex.xs` also allows selection with multiple keys
+:meth:`~pandas.MultiIndex.xs` also allows selection with multiple keys.
.. ipython:: python
@@ -403,13 +404,13 @@ providing the axis argument
df.loc[:,('bar','one')]
You can pass ``drop_level=False`` to :meth:`~pandas.MultiIndex.xs` to retain
-the level that was selected
+the level that was selected.
.. ipython:: python
df.xs('one', level='second', axis=1, drop_level=False)
-versus the result with ``drop_level=True`` (the default value)
+Compare the above with the result using ``drop_level=True`` (the default value).
.. ipython:: python
@@ -470,7 +471,7 @@ allowing you to permute the hierarchical index levels in one step:
Sorting a :class:`~pandas.MultiIndex`
-------------------------------------
-For MultiIndex-ed objects to be indexed & sliced effectively, they need
+For MultiIndex-ed objects to be indexed and sliced effectively, they need
to be sorted. As with any index, you can use ``sort_index``.
.. ipython:: python
@@ -623,7 +624,8 @@ Index Types
-----------
We have discussed ``MultiIndex`` in the previous sections pretty extensively. ``DatetimeIndex`` and ``PeriodIndex``
-are shown :ref:`here <timeseries.overview>`. ``TimedeltaIndex`` are :ref:`here <timedeltas.timedeltas>`.
+are shown :ref:`here <timeseries.overview>`, and information about
+`TimedeltaIndex`` is found :ref:`here <timedeltas.timedeltas>`.
In the following sub-sections we will highlight some other index types.
@@ -647,7 +649,7 @@ and allows efficient indexing and storage of an index with a large number of dup
df.dtypes
df.B.cat.categories
-Setting the index, will create a ``CategoricalIndex``
+Setting the index will create a ``CategoricalIndex``.
.. ipython:: python
@@ -655,36 +657,38 @@ Setting the index, will create a ``CategoricalIndex``
df2.index
Indexing with ``__getitem__/.iloc/.loc`` works similarly to an ``Index`` with duplicates.
-The indexers MUST be in the category or the operation will raise.
+The indexers **must** be in the category or the operation will raise a ``KeyError``.
.. ipython:: python
df2.loc['a']
-These PRESERVE the ``CategoricalIndex``
+The ``CategoricalIndex`` is **preserved** after indexing:
.. ipython:: python
df2.loc['a'].index
-Sorting will order by the order of the categories
+Sorting the index will sort by the order of the categories (Recall that we
+created the index with with ``CategoricalDtype(list('cab'))``, so the sorted
+order is ``cab``.).
.. ipython:: python
df2.sort_index()
-Groupby operations on the index will preserve the index nature as well
+Groupby operations on the index will preserve the index nature as well.
.. ipython:: python
df2.groupby(level=0).sum()
df2.groupby(level=0).sum().index
-Reindexing operations, will return a resulting index based on the type of the passed
-indexer, meaning that passing a list will return a plain-old-``Index``; indexing with
+Reindexing operations will return a resulting index based on the type of the passed
+indexer. Passing a list will return a plain-old ``Index``; indexing with
a ``Categorical`` will return a ``CategoricalIndex``, indexed according to the categories
-of the PASSED ``Categorical`` dtype. This allows one to arbitrarily index these even with
-values NOT in the categories, similarly to how you can reindex ANY pandas index.
+of the **passed** ``Categorical`` dtype. This allows one to arbitrarily index these even with
+values **not** in the categories, similarly to how you can reindex **any** pandas index.
.. ipython :: python
@@ -720,7 +724,8 @@ Int64Index and RangeIndex
Indexing on an integer-based Index with floats has been clarified in 0.18.0, for a summary of the changes, see :ref:`here <whatsnew_0180.float_indexers>`.
-``Int64Index`` is a fundamental basic index in *pandas*. This is an Immutable array implementing an ordered, sliceable set.
+``Int64Index`` is a fundamental basic index in pandas.
+This is an Immutable array implementing an ordered, sliceable set.
Prior to 0.18.0, the ``Int64Index`` would provide the default index for all ``NDFrame`` objects.
``RangeIndex`` is a sub-class of ``Int64Index`` added in version 0.18.0, now providing the default index for all ``NDFrame`` objects.
@@ -742,7 +747,7 @@ same.
sf = pd.Series(range(5), index=indexf)
sf
-Scalar selection for ``[],.loc`` will always be label based. An integer will match an equal float index (e.g. ``3`` is equivalent to ``3.0``)
+Scalar selection for ``[],.loc`` will always be label based. An integer will match an equal float index (e.g. ``3`` is equivalent to ``3.0``).
.. ipython:: python
@@ -751,15 +756,17 @@ Scalar selection for ``[],.loc`` will always be label based. An integer will mat
sf.loc[3]
sf.loc[3.0]
-The only positional indexing is via ``iloc``
+The only positional indexing is via ``iloc``.
.. ipython:: python
sf.iloc[3]
-A scalar index that is not found will raise ``KeyError``
+A scalar index that is not found will raise a ``KeyError``.
-Slicing is ALWAYS on the values of the index, for ``[],ix,loc`` and ALWAYS positional with ``iloc``
+Slicing is primarily on the values of the index when using ``[],ix,loc``, and
+**always** positional when using ``iloc``. The exception is when the slice is
+boolean, in which case it will always be positional.
.. ipython:: python
@@ -767,14 +774,14 @@ Slicing is ALWAYS on the values of the index, for ``[],ix,loc`` and ALWAYS posit
sf.loc[2:4]
sf.iloc[2:4]
-In float indexes, slicing using floats is allowed
+In float indexes, slicing using floats is allowed.
.. ipython:: python
sf[2.1:4.6]
sf.loc[2.1:4.6]
-In non-float indexes, slicing using floats will raise a ``TypeError``
+In non-float indexes, slicing using floats will raise a ``TypeError``.
.. code-block:: ipython
@@ -786,7 +793,7 @@ In non-float indexes, slicing using floats will raise a ``TypeError``
.. warning::
- Using a scalar float indexer for ``.iloc`` has been removed in 0.18.0, so the following will raise a ``TypeError``
+ Using a scalar float indexer for ``.iloc`` has been removed in 0.18.0, so the following will raise a ``TypeError``:
.. code-block:: ipython
@@ -816,13 +823,13 @@ Selection operations then will always work on a value basis, for all selection o
dfir.loc[0:1001,'A']
dfir.loc[1000.4]
-You could then easily pick out the first 1 second (1000 ms) of data then.
+You could retrieve the first 1 second (1000 ms) of data as such:
.. ipython:: python
dfir[0:1000]
-Of course if you need integer based selection, then use ``iloc``
+If you need integer based selection, you should use ``iloc``:
.. ipython:: python
@@ -975,6 +982,7 @@ consider the following Series:
s
Suppose we wished to slice from ``c`` to ``e``, using integers this would be
+accomplished as such:
.. ipython:: python
diff --git a/doc/source/basics.rst b/doc/source/basics.rst
index f9995472866ed..da82f56d315e6 100644
--- a/doc/source/basics.rst
+++ b/doc/source/basics.rst
@@ -436,7 +436,7 @@ General DataFrame Combine
~~~~~~~~~~~~~~~~~~~~~~~~~
The :meth:`~DataFrame.combine_first` method above calls the more general
-DataFrame method :meth:`~DataFrame.combine`. This method takes another DataFrame
+:meth:`DataFrame.combine`. This method takes another DataFrame
and a combiner function, aligns the input DataFrame and then passes the combiner
function pairs of Series (i.e., columns whose names are the same).
@@ -540,8 +540,8 @@ will exclude NAs on Series input by default:
np.mean(df['one'])
np.mean(df['one'].values)
-``Series`` also has a method :meth:`~Series.nunique` which will return the
-number of unique non-NA values:
+:meth:`Series.nunique` will return the number of unique non-NA values in a
+Series:
.. ipython:: python
@@ -852,7 +852,8 @@ Aggregation API
The aggregation API allows one to express possibly multiple aggregation operations in a single concise way.
This API is similar across pandas objects, see :ref:`groupby API <groupby.aggregate>`, the
:ref:`window functions API <stats.aggregate>`, and the :ref:`resample API <timeseries.aggregate>`.
-The entry point for aggregation is the method :meth:`~DataFrame.aggregate`, or the alias :meth:`~DataFrame.agg`.
+The entry point for aggregation is :meth:`DataFrame.aggregate`, or the alias
+:meth:`DataFrame.agg`.
We will use a similar starting frame from above:
@@ -1913,8 +1914,8 @@ dtype of the column will be chosen to accommodate all of the data types
# string data forces an ``object`` dtype
pd.Series([1, 2, 3, 6., 'foo'])
-The method :meth:`~DataFrame.get_dtype_counts` will return the number of columns of
-each type in a ``DataFrame``:
+The number of columns of each type in a ``DataFrame`` can be found by calling
+:meth:`~DataFrame.get_dtype_counts`.
.. ipython:: python
diff --git a/doc/source/computation.rst b/doc/source/computation.rst
index a6bc9431d3bcc..0994d35999191 100644
--- a/doc/source/computation.rst
+++ b/doc/source/computation.rst
@@ -26,9 +26,10 @@ Statistical Functions
Percent Change
~~~~~~~~~~~~~~
-``Series``, ``DataFrame``, and ``Panel`` all have a method ``pct_change`` to compute the
-percent change over a given number of periods (using ``fill_method`` to fill
-NA/null values *before* computing the percent change).
+``Series``, ``DataFrame``, and ``Panel`` all have a method
+:meth:`~DataFrame.pct_change` to compute the percent change over a given number
+of periods (using ``fill_method`` to fill NA/null values *before* computing
+the percent change).
.. ipython:: python
@@ -47,8 +48,8 @@ NA/null values *before* computing the percent change).
Covariance
~~~~~~~~~~
-The ``Series`` object has a method ``cov`` to compute covariance between series
-(excluding NA/null values).
+:meth:`Series.cov` can be used to compute covariance between series
+(excluding missing values).
.. ipython:: python
@@ -56,8 +57,9 @@ The ``Series`` object has a method ``cov`` to compute covariance between series
s2 = pd.Series(np.random.randn(1000))
s1.cov(s2)
-Analogously, ``DataFrame`` has a method ``cov`` to compute pairwise covariances
-among the series in the DataFrame, also excluding NA/null values.
+Analogously, :meth:`DataFrame.cov` to compute
+pairwise covariances among the series in the DataFrame, also excluding
+NA/null values.
.. _computation.covariance.caveats:
@@ -97,7 +99,9 @@ in order to have a valid result.
Correlation
~~~~~~~~~~~
-Several methods for computing correlations are provided:
+Correlation may be computed using the :meth:`~DataFrame.corr` method.
+Using the ``method`` parameter, several methods for computing correlations are
+provided:
.. csv-table::
:header: "Method name", "Description"
@@ -110,6 +114,11 @@ Several methods for computing correlations are provided:
.. \rho = \cov(x, y) / \sigma_x \sigma_y
All of these are currently computed using pairwise complete observations.
+Wikipedia has articles covering the above correlation coefficients:
+
+* `Pearson correlation coefficient <https://en.wikipedia.org/wiki/Pearson_correlation_coefficient>`_
+* `Kendall rank correlation coefficient <https://en.wikipedia.org/wiki/Kendall_rank_correlation_coefficient>`_
+* `Spearman's rank correlation coefficient <https://en.wikipedia.org/wiki/Spearman%27s_rank_correlation_coefficient>`_
.. note::
@@ -145,9 +154,9 @@ Like ``cov``, ``corr`` also supports the optional ``min_periods`` keyword:
frame.corr(min_periods=12)
-A related method ``corrwith`` is implemented on DataFrame to compute the
-correlation between like-labeled Series contained in different DataFrame
-objects.
+A related method :meth:`~DataFrame.corrwith` is implemented on DataFrame to
+compute the correlation between like-labeled Series contained in different
+DataFrame objects.
.. ipython:: python
@@ -163,8 +172,8 @@ objects.
Data ranking
~~~~~~~~~~~~
-The ``rank`` method produces a data ranking with ties being assigned the mean
-of the ranks (by default) for the group:
+The :meth:`~Series.rank` method produces a data ranking with ties being
+assigned the mean of the ranks (by default) for the group:
.. ipython:: python
@@ -172,8 +181,9 @@ of the ranks (by default) for the group:
s['d'] = s['b'] # so there's a tie
s.rank()
-``rank`` is also a DataFrame method and can rank either the rows (``axis=0``)
-or the columns (``axis=1``). ``NaN`` values are excluded from the ranking.
+:meth:`~DataFrame.rank` is also a DataFrame method and can rank either the rows
+(``axis=0``) or the columns (``axis=1``). ``NaN`` values are excluded from the
+ranking.
.. ipython:: python
@@ -205,7 +215,7 @@ Window Functions
Prior to version 0.18.0, ``pd.rolling_*``, ``pd.expanding_*``, and ``pd.ewm*`` were module level
functions and are now deprecated. These are replaced by using the :class:`~pandas.core.window.Rolling`, :class:`~pandas.core.window.Expanding` and :class:`~pandas.core.window.EWM`. objects and a corresponding method call.
- The deprecation warning will show the new syntax, see an example :ref:`here <whatsnew_0180.window_deprecations>`
+ The deprecation warning will show the new syntax, see an example :ref:`here <whatsnew_0180.window_deprecations>`.
For working with data, a number of windows functions are provided for
computing common *window* or *rolling* statistics. Among these are count, sum,
@@ -219,7 +229,7 @@ see the :ref:`groupby docs <groupby.transform.window_resample>`.
.. note::
- The API for window statistics is quite similar to the way one works with ``GroupBy`` objects, see the documentation :ref:`here <groupby>`
+ The API for window statistics is quite similar to the way one works with ``GroupBy`` objects, see the documentation :ref:`here <groupby>`.
We work with ``rolling``, ``expanding`` and ``exponentially weighted`` data through the corresponding
objects, :class:`~pandas.core.window.Rolling`, :class:`~pandas.core.window.Expanding` and :class:`~pandas.core.window.EWM`.
@@ -289,7 +299,7 @@ sugar for applying the moving window operator to all of the DataFrame's columns:
Method Summary
~~~~~~~~~~~~~~
-We provide a number of the common statistical functions:
+We provide a number of common statistical functions:
.. currentmodule:: pandas.core.window
@@ -564,7 +574,7 @@ Computing rolling pairwise covariances and correlations
.. warning::
Prior to version 0.20.0 if ``pairwise=True`` was passed, a ``Panel`` would be returned.
- This will now return a 2-level MultiIndexed DataFrame, see the whatsnew :ref:`here <whatsnew_0200.api_breaking.rolling_pairwise>`
+ This will now return a 2-level MultiIndexed DataFrame, see the whatsnew :ref:`here <whatsnew_0200.api_breaking.rolling_pairwise>`.
In financial data analysis and other fields it's common to compute covariance
and correlation matrices for a collection of time series. Often one is also
@@ -623,7 +633,8 @@ perform multiple computations on the data. These operations are similar to the :
r = dfa.rolling(window=60,min_periods=1)
r
-We can aggregate by passing a function to the entire DataFrame, or select a Series (or multiple Series) via standard getitem.
+We can aggregate by passing a function to the entire DataFrame, or select a
+Series (or multiple Series) via standard ``__getitem__``.
.. ipython:: python
@@ -741,14 +752,14 @@ all accept are:
- ``min_periods``: threshold of non-null data points to require. Defaults to
minimum needed to compute statistic. No ``NaNs`` will be output once
``min_periods`` non-null data points have been seen.
-- ``center``: boolean, whether to set the labels at the center (default is False)
+- ``center``: boolean, whether to set the labels at the center (default is False).
.. _stats.moments.expanding.note:
.. note::
The output of the ``.rolling`` and ``.expanding`` methods do not return a
``NaN`` if there are at least ``min_periods`` non-null values in the current
- window. For example,
+ window. For example:
.. ipython:: python
@@ -818,7 +829,8 @@ In general, a weighted moving average is calculated as
y_t = \frac{\sum_{i=0}^t w_i x_{t-i}}{\sum_{i=0}^t w_i},
-where :math:`x_t` is the input and :math:`y_t` is the result.
+where :math:`x_t` is the input, :math:`y_t` is the result and the :math:`w_i`
+are the weights.
The EW functions support two variants of exponential weights.
The default, ``adjust=True``, uses the weights :math:`w_i = (1 - \alpha)^i`
@@ -931,7 +943,7 @@ average of ``3, NaN, 5`` would be calculated as
.. math::
- \frac{(1-\alpha)^2 \cdot 3 + 1 \cdot 5}{(1-\alpha)^2 + 1}
+ \frac{(1-\alpha)^2 \cdot 3 + 1 \cdot 5}{(1-\alpha)^2 + 1}.
Whereas if ``ignore_na=True``, the weighted average would be calculated as
@@ -953,4 +965,4 @@ are scaled by debiasing factors
(For :math:`w_i = 1`, this reduces to the usual :math:`N / (N - 1)` factor,
with :math:`N = t + 1`.)
See `Weighted Sample Variance <http://en.wikipedia.org/wiki/Weighted_arithmetic_mean#Weighted_sample_variance>`__
-for further details.
+on Wikipedia for further details.
diff --git a/doc/source/indexing.rst b/doc/source/indexing.rst
index 3c2fd4d959d63..b9223c6ad9f7a 100644
--- a/doc/source/indexing.rst
+++ b/doc/source/indexing.rst
@@ -18,16 +18,14 @@ Indexing and Selecting Data
The axis labeling information in pandas objects serves many purposes:
- Identifies data (i.e. provides *metadata*) using known indicators,
- important for analysis, visualization, and interactive console display
- - Enables automatic and explicit data alignment
- - Allows intuitive getting and setting of subsets of the data set
+ important for analysis, visualization, and interactive console display.
+ - Enables automatic and explicit data alignment.
+ - Allows intuitive getting and setting of subsets of the data set.
In this section, we will focus on the final point: namely, how to slice, dice,
and generally get and set subsets of pandas objects. The primary focus will be
on Series and DataFrame as they have received more development attention in
-this area. Expect more work to be invested in higher-dimensional data
-structures (including ``Panel``) in the future, especially in label-based
-advanced indexing.
+this area.
.. note::
@@ -43,9 +41,9 @@ advanced indexing.
.. warning::
Whether a copy or a reference is returned for a setting operation, may
- depend on the context. This is sometimes called ``chained assignment`` and
- should be avoided. See :ref:`Returning a View versus Copy
- <indexing.view_versus_copy>`
+ depend on the context. This is sometimes called ``chained assignment`` and
+ should be avoided. See :ref:`Returning a View versus Copy
+ <indexing.view_versus_copy>`.
.. warning::
@@ -53,7 +51,7 @@ advanced indexing.
See the :ref:`MultiIndex / Advanced Indexing <advanced>` for ``MultiIndex`` and more advanced indexing documentation.
-See the :ref:`cookbook<cookbook.selection>` for some advanced strategies
+See the :ref:`cookbook<cookbook.selection>` for some advanced strategies.
.. _indexing.choice:
@@ -66,21 +64,21 @@ of multi-axis indexing.
- ``.loc`` is primarily label based, but may also be used with a boolean array. ``.loc`` will raise ``KeyError`` when the items are not found. Allowed inputs are:
- - A single label, e.g. ``5`` or ``'a'``, (note that ``5`` is interpreted as a
+ - A single label, e.g. ``5`` or ``'a'`` (Note that ``5`` is interpreted as a
*label* of the index. This use is **not** an integer position along the
- index)
- - A list or array of labels ``['a', 'b', 'c']``
- - A slice object with labels ``'a':'f'`` (note that contrary to usual python
+ index.).
+ - A list or array of labels ``['a', 'b', 'c']``.
+ - A slice object with labels ``'a':'f'`` (Note that contrary to usual python
slices, **both** the start and the stop are included, when present in the
- index! - also see :ref:`Slicing with labels
- <indexing.slicing_with_labels>`)
+ index! See :ref:`Slicing with labels
+ <indexing.slicing_with_labels>`.).
- A boolean array
- A ``callable`` function with one argument (the calling Series, DataFrame or Panel) and
- that returns valid output for indexing (one of the above)
+ that returns valid output for indexing (one of the above).
.. versionadded:: 0.18.1
- See more at :ref:`Selection by Label <indexing.label>`
+ See more at :ref:`Selection by Label <indexing.label>`.
- ``.iloc`` is primarily integer position based (from ``0`` to
``length-1`` of the axis), but may also be used with a boolean
@@ -89,27 +87,26 @@ of multi-axis indexing.
out-of-bounds indexing. (this conforms with python/numpy *slice*
semantics). Allowed inputs are:
- - An integer e.g. ``5``
- - A list or array of integers ``[4, 3, 0]``
- - A slice object with ints ``1:7``
- - A boolean array
+ - An integer e.g. ``5``.
+ - A list or array of integers ``[4, 3, 0]``.
+ - A slice object with ints ``1:7``.
+ - A boolean array.
- A ``callable`` function with one argument (the calling Series, DataFrame or Panel) and
- that returns valid output for indexing (one of the above)
+ that returns valid output for indexing (one of the above).
.. versionadded:: 0.18.1
- See more at :ref:`Selection by Position <indexing.integer>`
-
- See more at :ref:`Advanced Indexing <advanced>` and :ref:`Advanced
+ See more at :ref:`Selection by Position <indexing.integer>`,
+ :ref:`Advanced Indexing <advanced>` and :ref:`Advanced
Hierarchical <advanced.advanced_hierarchical>`.
- ``.loc``, ``.iloc``, and also ``[]`` indexing can accept a ``callable`` as indexer. See more at :ref:`Selection By Callable <indexing.callable>`.
Getting values from an object with multi-axes selection uses the following
-notation (using ``.loc`` as an example, but applies to ``.iloc`` as
+notation (using ``.loc`` as an example, but the following applies to ``.iloc`` as
well). Any of the axes accessors may be the null slice ``:``. Axes left out of
-the specification are assumed to be ``:``. (e.g. ``p.loc['a']`` is equiv to
-``p.loc['a', :, :]``)
+the specification are assumed to be ``:``, e.g. ``p.loc['a']`` is equivalent to
+``p.loc['a', :, :]``.
.. csv-table::
:header: "Object Type", "Indexers"
@@ -128,7 +125,8 @@ Basics
As mentioned when introducing the data structures in the :ref:`last section
<basics>`, the primary function of indexing with ``[]`` (a.k.a. ``__getitem__``
for those familiar with implementing class behavior in Python) is selecting out
-lower-dimensional slices. Thus,
+lower-dimensional slices. The following table shows return type values when
+indexing pandas objects with ``[]``:
.. csv-table::
:header: "Object Type", "Selection", "Return Value Type"
@@ -188,7 +186,7 @@ columns.
df.loc[:,['B', 'A']] = df[['A', 'B']]
df[['A', 'B']]
- The correct way is to use raw values
+ The correct way to swap column values is by using raw values:
.. ipython:: python
@@ -310,7 +308,7 @@ Selection By Label
Whether a copy or a reference is returned for a setting operation, may depend on the context.
This is sometimes called ``chained assignment`` and should be avoided.
- See :ref:`Returning a View versus Copy <indexing.view_versus_copy>`
+ See :ref:`Returning a View versus Copy <indexing.view_versus_copy>`.
.. warning::
@@ -336,23 +334,23 @@ Selection By Label
.. warning::
Starting in 0.21.0, pandas will show a ``FutureWarning`` if indexing with a list with missing labels. In the future
- this will raise a ``KeyError``. See :ref:`list-like Using loc with missing keys in a list is Deprecated <indexing.deprecate_loc_reindex_listlike>`
+ this will raise a ``KeyError``. See :ref:`list-like Using loc with missing keys in a list is Deprecated <indexing.deprecate_loc_reindex_listlike>`.
pandas provides a suite of methods in order to have **purely label based indexing**. This is a strict inclusion based protocol.
-All of the labels for which you ask, must be in the index or a ``KeyError`` will be raised!
+Every label asked for must be in the index, or a ``KeyError`` will be raised.
When slicing, both the start bound **AND** the stop bound are *included*, if present in the index.
Integers are valid labels, but they refer to the label **and not the position**.
The ``.loc`` attribute is the primary access method. The following are valid inputs:
-- A single label, e.g. ``5`` or ``'a'``, (note that ``5`` is interpreted as a *label* of the index. This use is **not** an integer position along the index)
-- A list or array of labels ``['a', 'b', 'c']``
-- A slice object with labels ``'a':'f'`` (note that contrary to usual python
+- A single label, e.g. ``5`` or ``'a'`` (Note that ``5`` is interpreted as a *label* of the index. This use is **not** an integer position along the index.).
+- A list or array of labels ``['a', 'b', 'c']``.
+- A slice object with labels ``'a':'f'`` (Note that contrary to usual python
slices, **both** the start and the stop are included, when present in the
- index! - also See :ref:`Slicing with labels
- <indexing.slicing_with_labels>`)
-- A boolean array
-- A ``callable``, see :ref:`Selection By Callable <indexing.callable>`
+ index! See :ref:`Slicing with labels
+ <indexing.slicing_with_labels>`.).
+- A boolean array.
+- A ``callable``, see :ref:`Selection By Callable <indexing.callable>`.
.. ipython:: python
@@ -368,7 +366,7 @@ Note that setting works as well:
s1.loc['c':] = 0
s1
-With a DataFrame
+With a DataFrame:
.. ipython:: python
@@ -378,26 +376,26 @@ With a DataFrame
df1
df1.loc[['a', 'b', 'd'], :]
-Accessing via label slices
+Accessing via label slices:
.. ipython:: python
df1.loc['d':, 'A':'C']
-For getting a cross section using a label (equiv to ``df.xs('a')``)
+For getting a cross section using a label (equivalent to ``df.xs('a')``):
.. ipython:: python
df1.loc['a']
-For getting values with a boolean array
+For getting values with a boolean array:
.. ipython:: python
df1.loc['a'] > 0
df1.loc[:, df1.loc['a'] > 0]
-For getting a value explicitly (equiv to deprecated ``df.get_value('a','A')``)
+For getting a value explicitly (equivalent to deprecated ``df.get_value('a','A')``):
.. ipython:: python
@@ -441,17 +439,17 @@ Selection By Position
Whether a copy or a reference is returned for a setting operation, may depend on the context.
This is sometimes called ``chained assignment`` and should be avoided.
- See :ref:`Returning a View versus Copy <indexing.view_versus_copy>`
+ See :ref:`Returning a View versus Copy <indexing.view_versus_copy>`.
Pandas provides a suite of methods in order to get **purely integer based indexing**. The semantics follow closely python and numpy slicing. These are ``0-based`` indexing. When slicing, the start bounds is *included*, while the upper bound is *excluded*. Trying to use a non-integer, even a **valid** label will raise an ``IndexError``.
The ``.iloc`` attribute is the primary access method. The following are valid inputs:
-- An integer e.g. ``5``
-- A list or array of integers ``[4, 3, 0]``
-- A slice object with ints ``1:7``
-- A boolean array
-- A ``callable``, see :ref:`Selection By Callable <indexing.callable>`
+- An integer e.g. ``5``.
+- A list or array of integers ``[4, 3, 0]``.
+- A slice object with ints ``1:7``.
+- A boolean array.
+- A ``callable``, see :ref:`Selection By Callable <indexing.callable>`.
.. ipython:: python
@@ -467,7 +465,7 @@ Note that setting works as well:
s1.iloc[:3] = 0
s1
-With a DataFrame
+With a DataFrame:
.. ipython:: python
@@ -476,14 +474,14 @@ With a DataFrame
columns=list(range(0,8,2)))
df1
-Select via integer slicing
+Select via integer slicing:
.. ipython:: python
df1.iloc[:3]
df1.iloc[1:5, 2:4]
-Select via integer list
+Select via integer list:
.. ipython:: python
@@ -502,7 +500,7 @@ Select via integer list
# this is also equivalent to ``df1.iat[1,1]``
df1.iloc[1, 1]
-For getting a cross section using an integer position (equiv to ``df.xs(1)``)
+For getting a cross section using an integer position (equiv to ``df.xs(1)``):
.. ipython:: python
@@ -523,7 +521,7 @@ Out of range slice indexes are handled gracefully just as in Python/Numpy.
s.iloc[8:10]
Note that using slices that go out of bounds can result in
-an empty axis (e.g. an empty DataFrame being returned)
+an empty axis (e.g. an empty DataFrame being returned).
.. ipython:: python
@@ -535,7 +533,7 @@ an empty axis (e.g. an empty DataFrame being returned)
A single indexer that is out of bounds will raise an ``IndexError``.
A list of indexers where any element is out of bounds will raise an
-``IndexError``
+``IndexError``.
.. code-block:: python
@@ -601,7 +599,7 @@ bit of user confusion over the years.
The recommended methods of indexing are:
-- ``.loc`` if you want to *label* index
+- ``.loc`` if you want to *label* index.
- ``.iloc`` if you want to *positionally* index.
.. ipython:: python
@@ -612,7 +610,7 @@ The recommended methods of indexing are:
dfd
-Previous Behavior, where you wish to get the 0th and the 2nd elements from the index in the 'A' column.
+Previous behavior, where you wish to get the 0th and the 2nd elements from the index in the 'A' column.
.. code-block:: ipython
@@ -635,7 +633,7 @@ This can also be expressed using ``.iloc``, by explicitly getting locations on t
dfd.iloc[[0, 2], dfd.columns.get_loc('A')]
-For getting *multiple* indexers, using ``.get_indexer``
+For getting *multiple* indexers, using ``.get_indexer``:
.. ipython:: python
@@ -824,7 +822,7 @@ Setting With Enlargement
The ``.loc/[]`` operations can perform enlargement when setting a non-existent key for that axis.
-In the ``Series`` case this is effectively an appending operation
+In the ``Series`` case this is effectively an appending operation.
.. ipython:: python
@@ -833,7 +831,7 @@ In the ``Series`` case this is effectively an appending operation
se[5] = 5.
se
-A ``DataFrame`` can be enlarged on either axis via ``.loc``
+A ``DataFrame`` can be enlarged on either axis via ``.loc``.
.. ipython:: python
@@ -889,7 +887,11 @@ Boolean indexing
.. _indexing.boolean:
Another common operation is the use of boolean vectors to filter the data.
-The operators are: ``|`` for ``or``, ``&`` for ``and``, and ``~`` for ``not``. These **must** be grouped by using parentheses.
+The operators are: ``|`` for ``or``, ``&`` for ``and``, and ``~`` for ``not``.
+These **must** be grouped by using parentheses, since by default Python will
+evaluate an expression such as ``df.A > 2 & df.B < 3`` as
+``df.A > (2 & df.B) < 3``, while the desired evaluation order is
+``(df.A > 2) & (df.B < 3)``.
Using a boolean vector to index a Series works exactly as in a numpy ndarray:
@@ -929,7 +931,7 @@ more complex criteria:
# Multiple criteria
df2[criterion & (df2['b'] == 'x')]
-Note, with the choice methods :ref:`Selection by Label <indexing.label>`, :ref:`Selection by Position <indexing.integer>`,
+With the choice methods :ref:`Selection by Label <indexing.label>`, :ref:`Selection by Position <indexing.integer>`,
and :ref:`Advanced Indexing <advanced>` you may select along more than one axis using boolean vectors combined with other indexing expressions.
.. ipython:: python
@@ -941,9 +943,9 @@ and :ref:`Advanced Indexing <advanced>` you may select along more than one axis
Indexing with isin
------------------
-Consider the ``isin`` method of Series, which returns a boolean vector that is
-true wherever the Series elements exist in the passed list. This allows you to
-select rows where one or more columns have values you want:
+Consider the :meth:`~Series.isin` method of ``Series``, which returns a boolean
+vector that is true wherever the ``Series`` elements exist in the passed list.
+This allows you to select rows where one or more columns have values you want:
.. ipython:: python
@@ -973,7 +975,7 @@ in the membership check:
s_mi.iloc[s_mi.index.isin([(1, 'a'), (2, 'b'), (0, 'c')])]
s_mi.iloc[s_mi.index.isin(['a', 'c', 'e'], level=1)]
-DataFrame also has an ``isin`` method. When calling ``isin``, pass a set of
+DataFrame also has an :meth:`~DataFrame.isin` method. When calling ``isin``, pass a set of
values as either an array or dict. If values is an array, ``isin`` returns
a DataFrame of booleans that is the same shape as the original DataFrame, with True
wherever the element is in the sequence of values.
@@ -1018,13 +1020,13 @@ Selecting values from a Series with a boolean vector generally returns a
subset of the data. To guarantee that selection output has the same shape as
the original data, you can use the ``where`` method in ``Series`` and ``DataFrame``.
-To return only the selected rows
+To return only the selected rows:
.. ipython:: python
s[s > 0]
-To return a Series of the same shape as the original
+To return a Series of the same shape as the original:
.. ipython:: python
@@ -1032,7 +1034,7 @@ To return a Series of the same shape as the original
Selecting values from a DataFrame with a boolean criterion now also preserves
input data shape. ``where`` is used under the hood as the implementation.
-Equivalent is ``df.where(df < 0)``
+The code below is equivalent to ``df.where(df < 0)``.
.. ipython:: python
:suppress:
@@ -1087,12 +1089,12 @@ without creating a copy:
Furthermore, ``where`` aligns the input boolean condition (ndarray or DataFrame),
such that partial selection with setting is possible. This is analogous to
-partial setting via ``.loc`` (but on the contents rather than the axis labels)
+partial setting via ``.loc`` (but on the contents rather than the axis labels).
.. ipython:: python
df2 = df.copy()
- df2[ df2[1:4] > 0 ] = 3
+ df2[ df2[1:4] > 0] = 3
df2
Where can also accept ``axis`` and ``level`` parameters to align the input when
@@ -1103,7 +1105,7 @@ performing the ``where``.
df2 = df.copy()
df2.where(df2>0,df2['A'],axis='index')
-This is equivalent (but faster than) the following.
+This is equivalent to (but faster than) the following.
.. ipython:: python
@@ -1123,9 +1125,11 @@ as condition and ``other`` argument.
'C': [7, 8, 9]})
df3.where(lambda x: x > 4, lambda x: x + 10)
-**mask**
-``mask`` is the inverse boolean operation of ``where``.
+Mask
+~~~~
+
+:meth:`~pandas.DataFrame.mask` is the inverse boolean operation of ``where``.
.. ipython:: python
@@ -1134,8 +1138,8 @@ as condition and ``other`` argument.
.. _indexing.query:
-The :meth:`~pandas.DataFrame.query` Method (Experimental)
----------------------------------------------------------
+The :meth:`~pandas.DataFrame.query` Method
+------------------------------------------
:class:`~pandas.DataFrame` objects have a :meth:`~pandas.DataFrame.query`
method that allows selection using an expression.
@@ -1263,7 +1267,7 @@ having to specify which frame you're interested in querying
:meth:`~pandas.DataFrame.query` Python versus pandas Syntax Comparison
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-Full numpy-like syntax
+Full numpy-like syntax:
.. ipython:: python
@@ -1273,19 +1277,19 @@ Full numpy-like syntax
df[(df.a < df.b) & (df.b < df.c)]
Slightly nicer by removing the parentheses (by binding making comparison
-operators bind tighter than ``&``/``|``)
+operators bind tighter than ``&`` and ``|``).
.. ipython:: python
df.query('a < b & b < c')
-Use English instead of symbols
+Use English instead of symbols:
.. ipython:: python
df.query('a < b and b < c')
-Pretty close to how you might write it on paper
+Pretty close to how you might write it on paper:
.. ipython:: python
@@ -1356,7 +1360,7 @@ Special use of the ``==`` operator with ``list`` objects
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Comparing a ``list`` of values to a column using ``==``/``!=`` works similarly
-to ``in``/``not in``
+to ``in``/``not in``.
.. ipython:: python
@@ -1391,7 +1395,7 @@ You can negate boolean expressions with the word ``not`` or the ``~`` operator.
df.query('not bools')
df.query('not bools') == df[~df.bools]
-Of course, expressions can be arbitrarily complex too
+Of course, expressions can be arbitrarily complex too:
.. ipython:: python
@@ -1420,7 +1424,7 @@ Performance of :meth:`~pandas.DataFrame.query`
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
``DataFrame.query()`` using ``numexpr`` is slightly faster than Python for
-large frames
+large frames.
.. image:: _static/query-perf.png
@@ -1428,7 +1432,7 @@ large frames
You will only see the performance benefits of using the ``numexpr`` engine
with ``DataFrame.query()`` if your frame has more than approximately 200,000
- rows
+ rows.
.. image:: _static/query-perf-small.png
@@ -1482,7 +1486,7 @@ Also, you can pass a list of columns to identify duplications.
df2.drop_duplicates(['a', 'b'])
To drop duplicates by index value, use ``Index.duplicated`` then perform slicing.
-Same options are available in ``keep`` parameter.
+The same set of options are available for the ``keep`` parameter.
.. ipython:: python
@@ -1514,7 +1518,7 @@ The :meth:`~pandas.DataFrame.lookup` Method
Sometimes you want to extract a set of values given a sequence of row labels
and column labels, and the ``lookup`` method allows for this and returns a
-numpy array. For instance,
+numpy array. For instance:
.. ipython:: python
@@ -1599,7 +1603,7 @@ Set operations on Index objects
.. _indexing.set_ops:
-The two main operations are ``union (|)``, ``intersection (&)``
+The two main operations are ``union (|)`` and ``intersection (&)``.
These can be directly called as instance methods or used via overloaded
operators. Difference is provided via the ``.difference()`` method.
@@ -1612,7 +1616,7 @@ operators. Difference is provided via the ``.difference()`` method.
a.difference(b)
Also available is the ``symmetric_difference (^)`` operation, which returns elements
-that appear in either ``idx1`` or ``idx2`` but not both. This is
+that appear in either ``idx1`` or ``idx2``, but not in both. This is
equivalent to the Index created by ``idx1.difference(idx2).union(idx2.difference(idx1))``,
with duplicates dropped.
@@ -1662,9 +1666,9 @@ Set an index
.. _indexing.set_index:
-DataFrame has a ``set_index`` method which takes a column name (for a regular
-``Index``) or a list of column names (for a ``MultiIndex``), to create a new,
-indexed DataFrame:
+DataFrame has a :meth:`~DataFrame.set_index` method which takes a column name
+(for a regular ``Index``) or a list of column names (for a ``MultiIndex``).
+To create a new, re-indexed DataFrame:
.. ipython:: python
:suppress:
@@ -1703,9 +1707,10 @@ the index in-place (without creating a new object):
Reset the index
~~~~~~~~~~~~~~~
-As a convenience, there is a new function on DataFrame called ``reset_index``
-which transfers the index values into the DataFrame's columns and sets a simple
-integer index. This is the inverse operation to ``set_index``
+As a convenience, there is a new function on DataFrame called
+:meth:`~DataFrame.reset_index` which transfers the index values into the
+DataFrame's columns and sets a simple integer index.
+This is the inverse operation of :meth:`~DataFrame.set_index`.
.. ipython:: python
@@ -1726,11 +1731,6 @@ You can use the ``level`` keyword to remove only a portion of the index:
``reset_index`` takes an optional parameter ``drop`` which if true simply
discards the index, instead of putting index values in the DataFrame's columns.
-.. note::
-
- The ``reset_index`` method used to be called ``delevel`` which is now
- deprecated.
-
Adding an ad hoc index
~~~~~~~~~~~~~~~~~~~~~~
@@ -1769,7 +1769,7 @@ Compare these two access methods:
dfmi.loc[:,('one','second')]
These both yield the same results, so which should you use? It is instructive to understand the order
-of operations on these and why method 2 (``.loc``) is much preferred over method 1 (chained ``[]``)
+of operations on these and why method 2 (``.loc``) is much preferred over method 1 (chained ``[]``).
``dfmi['one']`` selects the first level of the columns and returns a DataFrame that is singly-indexed.
Then another python operation ``dfmi_with_one['second']`` selects the series indexed by ``'second'`` happens.
@@ -1807,7 +1807,7 @@ But this code is handled differently:
See that ``__getitem__`` in there? Outside of simple cases, it's very hard to
predict whether it will return a view or a copy (it depends on the memory layout
-of the array, about which *pandas* makes no guarantees), and therefore whether
+of the array, about which pandas makes no guarantees), and therefore whether
the ``__setitem__`` will modify ``dfmi`` or a temporary object that gets thrown
out immediately afterward. **That's** what ``SettingWithCopy`` is warning you
about!
@@ -1882,9 +1882,9 @@ A chained assignment can also crop up in setting in a mixed dtype frame.
.. note::
- These setting rules apply to all of ``.loc/.iloc``
+ These setting rules apply to all of ``.loc/.iloc``.
-This is the correct access method
+This is the correct access method:
.. ipython:: python
@@ -1892,7 +1892,7 @@ This is the correct access method
dfc.loc[0,'A'] = 11
dfc
-This *can* work at times, but is not guaranteed, and so should be avoided
+This *can* work at times, but it is not guaranteed to, and therefore should be avoided:
.. ipython:: python
:okwarning:
@@ -1901,7 +1901,7 @@ This *can* work at times, but is not guaranteed, and so should be avoided
dfc['A'][0] = 111
dfc
-This will **not** work at all, and so should be avoided
+This will **not** work at all, and so should be avoided:
::
diff --git a/doc/source/options.rst b/doc/source/options.rst
index 505a5ade68de0..5641b2628fe40 100644
--- a/doc/source/options.rst
+++ b/doc/source/options.rst
@@ -37,7 +37,7 @@ namespace:
- :func:`~pandas.option_context` - execute a codeblock with a set of options
that revert to prior settings after execution.
-**Note:** developers can check out pandas/core/config.py for more info.
+**Note:** Developers can check out `pandas/core/config.py <https://github.com/pandas-dev/pandas/blob/master/pandas/core/config.py>`_ for more information.
All of the functions above accept a regexp pattern (``re.search`` style) as an argument,
and so passing in a substring will work - as long as it is unambiguous:
@@ -78,8 +78,9 @@ with no argument ``describe_option`` will print out the descriptions for all ava
Getting and Setting Options
---------------------------
-As described above, ``get_option()`` and ``set_option()`` are available from the
-pandas namespace. To change an option, call ``set_option('option regex', new_value)``
+As described above, :func:`~pandas.get_option` and :func:`~pandas.set_option`
+are available from the pandas namespace. To change an option, call
+``set_option('option regex', new_value)``.
.. ipython:: python
@@ -87,7 +88,7 @@ pandas namespace. To change an option, call ``set_option('option regex', new_va
pd.set_option('mode.sim_interactive', True)
pd.get_option('mode.sim_interactive')
-**Note:** that the option 'mode.sim_interactive' is mostly used for debugging purposes.
+**Note:** The option 'mode.sim_interactive' is mostly used for debugging purposes.
All options also have a default value, and you can use ``reset_option`` to do just that:
@@ -221,7 +222,7 @@ can specify the option ``df.info(null_counts=True)`` to override on showing a pa
.. ipython:: python
- df =pd.DataFrame(np.random.choice([0,1,np.nan], size=(10,10)))
+ df = pd.DataFrame(np.random.choice([0,1,np.nan], size=(10,10)))
df
pd.set_option('max_info_rows', 11)
df.info()
@@ -229,8 +230,8 @@ can specify the option ``df.info(null_counts=True)`` to override on showing a pa
df.info()
pd.reset_option('max_info_rows')
-``display.precision`` sets the output display precision in terms of decimal places. This is only a
-suggestion.
+``display.precision`` sets the output display precision in terms of decimal places.
+This is only a suggestion.
.. ipython:: python
@@ -241,7 +242,7 @@ suggestion.
df
``display.chop_threshold`` sets at what level pandas rounds to zero when
-it displays a Series of DataFrame. Note, this does not effect the
+it displays a Series of DataFrame. This setting does not change the
precision at which the number is stored.
.. ipython:: python
@@ -254,7 +255,7 @@ precision at which the number is stored.
pd.reset_option('chop_threshold')
``display.colheader_justify`` controls the justification of the headers.
-Options are 'right', and 'left'.
+The options are 'right', and 'left'.
.. ipython:: python
diff --git a/doc/source/text.rst b/doc/source/text.rst
index 2a86d92978043..2b6459b581c1e 100644
--- a/doc/source/text.rst
+++ b/doc/source/text.rst
@@ -99,7 +99,7 @@ Elements in the split lists can be accessed using ``get`` or ``[]`` notation:
s2.str.split('_').str.get(1)
s2.str.split('_').str[1]
-Easy to expand this to return a DataFrame using ``expand``.
+It is easy to expand this to return a DataFrame using ``expand``.
.. ipython:: python
@@ -268,7 +268,7 @@ It returns a Series if ``expand=False``.
pd.Series(['a1', 'b2', 'c3']).str.extract('[ab](\d)', expand=False)
Calling on an ``Index`` with a regex with exactly one capture group
-returns a ``DataFrame`` with one column if ``expand=True``,
+returns a ``DataFrame`` with one column if ``expand=True``.
.. ipython:: python
@@ -373,7 +373,7 @@ You can check whether elements contain a pattern:
pattern = r'[0-9][a-z]'
pd.Series(['1', '2', '3a', '3b', '03c']).str.contains(pattern)
-or match a pattern:
+Or whether elements match a pattern:
.. ipython:: python
| This PR continues my read-through of the docs, the previous PRs submitted are #18941 and #18948.
The following edits have been made:
- Missing periods and colons added before introducing code examples.
- Increased number of function references (clickable links).
- Cleared up a few sentences which I found unclear.
Feedback is welcome. | https://api.github.com/repos/pandas-dev/pandas/pulls/18973 | 2017-12-28T16:54:15Z | 2017-12-29T21:48:57Z | 2017-12-29T21:48:57Z | 2017-12-29T21:49:01Z |
TST: split out some sparse tests | diff --git a/ci/install_travis.sh b/ci/install_travis.sh
index 800a20aa94b8f..693a2fe1fd6a6 100755
--- a/ci/install_travis.sh
+++ b/ci/install_travis.sh
@@ -178,15 +178,15 @@ if [ "$PIP_BUILD_TEST" ]; then
# build & install testing
echo "[building release]"
- bash scripts/build_dist_for_release.sh
+ time bash scripts/build_dist_for_release.sh || exit 1
conda uninstall -y cython
- time pip install dist/*tar.gz --quiet || exit 1
+ time pip install dist/*tar.gz || exit 1
elif [ "$CONDA_BUILD_TEST" ]; then
# build & install testing
echo "[building conda recipe]"
- conda build ./conda.recipe --numpy 1.13 --python 3.5 -q --no-test
+ time conda build ./conda.recipe --numpy 1.13 --python 3.5 -q --no-test
echo "[installing]"
conda install pandas --use-local
diff --git a/pandas/core/dtypes/dtypes.py b/pandas/core/dtypes/dtypes.py
index 3a8edf9f066ee..a47f2c0d4ab13 100644
--- a/pandas/core/dtypes/dtypes.py
+++ b/pandas/core/dtypes/dtypes.py
@@ -402,7 +402,7 @@ class DatetimeTZDtype(ExtensionDtype):
num = 101
base = np.dtype('M8[ns]')
_metadata = ['unit', 'tz']
- _match = re.compile("(datetime64|M8)\[(?P<unit>.+), (?P<tz>.+)\]")
+ _match = re.compile(r"(datetime64|M8)\[(?P<unit>.+), (?P<tz>.+)\]")
_cache = {}
def __new__(cls, unit=None, tz=None):
@@ -514,7 +514,7 @@ class PeriodDtype(ExtensionDtype):
base = np.dtype('O')
num = 102
_metadata = ['freq']
- _match = re.compile("(P|p)eriod\[(?P<freq>.+)\]")
+ _match = re.compile(r"(P|p)eriod\[(?P<freq>.+)\]")
_cache = {}
def __new__(cls, freq=None):
@@ -632,7 +632,7 @@ class IntervalDtype(ExtensionDtype):
base = np.dtype('O')
num = 103
_metadata = ['subtype']
- _match = re.compile("(I|i)nterval\[(?P<subtype>.+)\]")
+ _match = re.compile(r"(I|i)nterval\[(?P<subtype>.+)\]")
_cache = {}
def __new__(cls, subtype=None):
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 3243baa0008ae..12a4a7fdaedad 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -2635,7 +2635,7 @@ def insert(self, loc, column, value, allow_duplicates=False):
allow_duplicates=allow_duplicates)
def assign(self, **kwargs):
- """
+ r"""
Assign new columns to a DataFrame, returning a new object
(a copy) with all the original columns in addition to the new ones.
diff --git a/pandas/core/strings.py b/pandas/core/strings.py
index 9614641aa1abf..99c7563d5b249 100644
--- a/pandas/core/strings.py
+++ b/pandas/core/strings.py
@@ -306,7 +306,7 @@ def str_endswith(arr, pat, na=np.nan):
def str_replace(arr, pat, repl, n=-1, case=None, flags=0):
- """
+ r"""
Replace occurrences of pattern/regex in the Series/Index with
some other string. Equivalent to :meth:`str.replace` or
:func:`re.sub`.
@@ -598,7 +598,7 @@ def _str_extract_frame(arr, pat, flags=0):
def str_extract(arr, pat, flags=0, expand=None):
- """
+ r"""
For each subject string in the Series, extract groups from the
first match of regular expression pat.
@@ -635,7 +635,7 @@ def str_extract(arr, pat, flags=0, expand=None):
Non-matches will be NaN.
>>> s = Series(['a1', 'b2', 'c3'])
- >>> s.str.extract('([ab])(\d)')
+ >>> s.str.extract(r'([ab])(\d)')
0 1
0 a 1
1 b 2
@@ -643,7 +643,7 @@ def str_extract(arr, pat, flags=0, expand=None):
A pattern may contain optional groups.
- >>> s.str.extract('([ab])?(\d)')
+ >>> s.str.extract(r'([ab])?(\d)')
0 1
0 a 1
1 b 2
@@ -651,7 +651,7 @@ def str_extract(arr, pat, flags=0, expand=None):
Named groups will become column names in the result.
- >>> s.str.extract('(?P<letter>[ab])(?P<digit>\d)')
+ >>> s.str.extract(r'(?P<letter>[ab])(?P<digit>\d)')
letter digit
0 a 1
1 b 2
@@ -660,7 +660,7 @@ def str_extract(arr, pat, flags=0, expand=None):
A pattern with one group will return a DataFrame with one column
if expand=True.
- >>> s.str.extract('[ab](\d)', expand=True)
+ >>> s.str.extract(r'[ab](\d)', expand=True)
0
0 1
1 2
@@ -668,7 +668,7 @@ def str_extract(arr, pat, flags=0, expand=None):
A pattern with one group will return a Series if expand=False.
- >>> s.str.extract('[ab](\d)', expand=False)
+ >>> s.str.extract(r'[ab](\d)', expand=False)
0 1
1 2
2 NaN
@@ -694,7 +694,7 @@ def str_extract(arr, pat, flags=0, expand=None):
def str_extractall(arr, pat, flags=0):
- """
+ r"""
For each subject string in the Series, extract groups from all
matches of regular expression pat. When each subject string in the
Series has exactly one match, extractall(pat).xs(0, level='match')
@@ -728,7 +728,7 @@ def str_extractall(arr, pat, flags=0):
Indices with no matches will not appear in the result.
>>> s = Series(["a1a2", "b1", "c1"], index=["A", "B", "C"])
- >>> s.str.extractall("[ab](\d)")
+ >>> s.str.extractall(r"[ab](\d)")
0
match
A 0 1
@@ -737,7 +737,7 @@ def str_extractall(arr, pat, flags=0):
Capture group names are used for column names of the result.
- >>> s.str.extractall("[ab](?P<digit>\d)")
+ >>> s.str.extractall(r"[ab](?P<digit>\d)")
digit
match
A 0 1
@@ -746,7 +746,7 @@ def str_extractall(arr, pat, flags=0):
A pattern with two groups will return a DataFrame with two columns.
- >>> s.str.extractall("(?P<letter>[ab])(?P<digit>\d)")
+ >>> s.str.extractall(r"(?P<letter>[ab])(?P<digit>\d)")
letter digit
match
A 0 a 1
@@ -755,7 +755,7 @@ def str_extractall(arr, pat, flags=0):
Optional groups that do not match are NaN in the result.
- >>> s.str.extractall("(?P<letter>[ab])?(?P<digit>\d)")
+ >>> s.str.extractall(r"(?P<letter>[ab])?(?P<digit>\d)")
letter digit
match
A 0 a 1
diff --git a/pandas/io/clipboards.py b/pandas/io/clipboards.py
index 8e9b5497083f6..347ec41baf0e1 100644
--- a/pandas/io/clipboards.py
+++ b/pandas/io/clipboards.py
@@ -3,7 +3,7 @@
from pandas.compat import StringIO, PY2
-def read_clipboard(sep='\s+', **kwargs): # pragma: no cover
+def read_clipboard(sep=r'\s+', **kwargs): # pragma: no cover
r"""
Read text from clipboard and pass to read_table. See read_table for the
full argument list
@@ -55,10 +55,10 @@ def read_clipboard(sep='\s+', **kwargs): # pragma: no cover
counts = {x.lstrip().count('\t') for x in lines}
if len(lines) > 1 and len(counts) == 1 and counts.pop() != 0:
- sep = '\t'
+ sep = r'\t'
if sep is None and kwargs.get('delim_whitespace') is None:
- sep = '\s+'
+ sep = r'\s+'
return read_table(StringIO(text), sep=sep, **kwargs)
@@ -99,7 +99,7 @@ def to_clipboard(obj, excel=None, sep=None, **kwargs): # pragma: no cover
if excel:
try:
if sep is None:
- sep = '\t'
+ sep = r'\t'
buf = StringIO()
# clipboard_set (pyperclip) expects unicode
obj.to_csv(buf, sep=sep, encoding='utf-8', **kwargs)
diff --git a/pandas/io/formats/format.py b/pandas/io/formats/format.py
index 95b3a9162db45..a4678e5b40849 100644
--- a/pandas/io/formats/format.py
+++ b/pandas/io/formats/format.py
@@ -1002,7 +1002,7 @@ def get_col_type(dtype):
buf.write('\\end{longtable}\n')
def _format_multicolumn(self, row, ilevels):
- """
+ r"""
Combine columns belonging to a group to a single multicolumn entry
according to self.multicolumn_format
@@ -1040,7 +1040,7 @@ def append_col():
return row2
def _format_multirow(self, row, ilevels, i, rows):
- """
+ r"""
Check following rows, whether row should be a multirow
e.g.: becomes:
@@ -1071,7 +1071,7 @@ def _print_cline(self, buf, i, icol):
"""
for cl in self.clinebuf:
if cl[0] == i:
- buf.write('\cline{{{cl:d}-{icol:d}}}\n'
+ buf.write('\\cline{{{cl:d}-{icol:d}}}\n'
.format(cl=cl[1], icol=icol))
# remove entries that have been written to buffer
self.clinebuf = [x for x in self.clinebuf if x[0] != i]
diff --git a/pandas/io/parsers.py b/pandas/io/parsers.py
index dca26d028d8a4..e053af17667c4 100755
--- a/pandas/io/parsers.py
+++ b/pandas/io/parsers.py
@@ -51,7 +51,7 @@
# so we need to remove it if we see it.
_BOM = u('\ufeff')
-_parser_params = """Also supports optionally iterating or breaking of the file
+_parser_params = r"""Also supports optionally iterating or breaking of the file
into chunks.
Additional help can be found in the `online docs for IO Tools
@@ -842,19 +842,19 @@ def _clean_options(self, options, engine):
" sep=None with delim_whitespace=False"
engine = 'python'
elif sep is not None and len(sep) > 1:
- if engine == 'c' and sep == '\s+':
+ if engine == 'c' and sep == r'\s+':
result['delim_whitespace'] = True
del result['delimiter']
elif engine not in ('python', 'python-fwf'):
# wait until regex engine integrated
fallback_reason = "the 'c' engine does not support"\
" regex separators (separators > 1 char and"\
- " different from '\s+' are"\
+ r" different from '\s+' are"\
" interpreted as regex)"
engine = 'python'
elif delim_whitespace:
if 'python' in engine:
- result['delimiter'] = '\s+'
+ result['delimiter'] = r'\s+'
elif sep is not None:
encodeable = True
try:
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index d73417f7b0c95..c428000d73593 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -1792,7 +1792,7 @@ def create_for_block(
# name values_0
try:
if version[0] == 0 and version[1] <= 10 and version[2] == 0:
- m = re.search("values_block_(\d+)", name)
+ m = re.search(r"values_block_(\d+)", name)
if m:
name = "values_%s" % m.groups()[0]
except:
@@ -4297,7 +4297,7 @@ class AppendableMultiFrameTable(AppendableFrameTable):
table_type = u('appendable_multiframe')
obj_type = DataFrame
ndim = 2
- _re_levels = re.compile("^level_\d+$")
+ _re_levels = re.compile(r"^level_\d+$")
@property
def table_type_short(self):
diff --git a/pandas/io/sql.py b/pandas/io/sql.py
index 0d398ad3135a6..c7bbbf9940ba1 100644
--- a/pandas/io/sql.py
+++ b/pandas/io/sql.py
@@ -1306,7 +1306,7 @@ def _create_table_setup(self):
column_names_and_types = \
self._get_column_names_and_types(self._sql_type_name)
- pat = re.compile('\s+')
+ pat = re.compile(r'\s+')
column_names = [col_name for col_name, _, _ in column_names_and_types]
if any(map(pat.search, column_names)):
warnings.warn(_SAFE_NAMES_WARNING, stacklevel=6)
diff --git a/pandas/tests/indexes/interval/test_interval.py b/pandas/tests/indexes/interval/test_interval.py
index 8ce3c74fe6a31..6fc5526e63e59 100644
--- a/pandas/tests/indexes/interval/test_interval.py
+++ b/pandas/tests/indexes/interval/test_interval.py
@@ -599,7 +599,7 @@ def test_monotonic(self, closed):
assert idx.is_monotonic_decreasing
assert idx._is_strictly_monotonic_decreasing
- @pytest.mark.xfail(reason='not a valid repr as we use interval notation')
+ @pytest.mark.skip(reason='not a valid repr as we use interval notation')
def test_repr(self):
i = IntervalIndex.from_tuples([(0, 1), (1, 2)], closed='right')
expected = ("IntervalIndex(left=[0, 1],"
@@ -619,11 +619,11 @@ def test_repr(self):
"\n dtype='interval[datetime64[ns]]')")
assert repr(i) == expected
- @pytest.mark.xfail(reason='not a valid repr as we use interval notation')
+ @pytest.mark.skip(reason='not a valid repr as we use interval notation')
def test_repr_max_seq_item_setting(self):
super(TestIntervalIndex, self).test_repr_max_seq_item_setting()
- @pytest.mark.xfail(reason='not a valid repr as we use interval notation')
+ @pytest.mark.skip(reason='not a valid repr as we use interval notation')
def test_repr_roundtrip(self):
super(TestIntervalIndex, self).test_repr_roundtrip()
diff --git a/pandas/tests/sparse/frame/__init__.py b/pandas/tests/sparse/frame/__init__.py
new file mode 100644
index 0000000000000..e69de29bb2d1d
diff --git a/pandas/tests/sparse/frame/test_analytics.py b/pandas/tests/sparse/frame/test_analytics.py
new file mode 100644
index 0000000000000..ccb30502b862e
--- /dev/null
+++ b/pandas/tests/sparse/frame/test_analytics.py
@@ -0,0 +1,40 @@
+import pytest
+import numpy as np
+from pandas import SparseDataFrame, DataFrame, SparseSeries
+from pandas.util import testing as tm
+
+
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_quantile():
+ # GH 17386
+ data = [[1, 1], [2, 10], [3, 100], [np.nan, np.nan]]
+ q = 0.1
+
+ sparse_df = SparseDataFrame(data)
+ result = sparse_df.quantile(q)
+
+ dense_df = DataFrame(data)
+ dense_expected = dense_df.quantile(q)
+ sparse_expected = SparseSeries(dense_expected)
+
+ tm.assert_series_equal(result, dense_expected)
+ tm.assert_sp_series_equal(result, sparse_expected)
+
+
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_quantile_multi():
+ # GH 17386
+ data = [[1, 1], [2, 10], [3, 100], [np.nan, np.nan]]
+ q = [0.1, 0.5]
+
+ sparse_df = SparseDataFrame(data)
+ result = sparse_df.quantile(q)
+
+ dense_df = DataFrame(data)
+ dense_expected = dense_df.quantile(q)
+ sparse_expected = SparseDataFrame(dense_expected)
+
+ tm.assert_frame_equal(result, dense_expected)
+ tm.assert_sp_frame_equal(result, sparse_expected)
diff --git a/pandas/tests/sparse/test_frame.py b/pandas/tests/sparse/frame/test_frame.py
similarity index 94%
rename from pandas/tests/sparse/test_frame.py
rename to pandas/tests/sparse/frame/test_frame.py
index 4b9d6621a20fb..cf002ff046c2e 100644
--- a/pandas/tests/sparse/test_frame.py
+++ b/pandas/tests/sparse/frame/test_frame.py
@@ -1402,108 +1402,6 @@ def test_numpy_func_call(self):
for func in funcs:
getattr(np, func)(self.frame)
- @pytest.mark.parametrize('data', [
- [[1, 1], [2, 2], [3, 3], [4, 4], [0, 0]],
- [[1.0, 1.0], [2.0, 2.0], [3.0, 3.0], [4.0, 4.0], [nan, nan]],
- [
- [1.0, 1.0 + 1.0j],
- [2.0 + 2.0j, 2.0],
- [3.0, 3.0 + 3.0j],
- [4.0 + 4.0j, 4.0],
- [nan, nan]
- ]
- ])
- @pytest.mark.xfail(reason='Wrong SparseBlock initialization '
- '(GH 17386)')
- def test_where_with_numeric_data(self, data):
- # GH 17386
- lower_bound = 1.5
-
- sparse = SparseDataFrame(data)
- result = sparse.where(sparse > lower_bound)
-
- dense = DataFrame(data)
- dense_expected = dense.where(dense > lower_bound)
- sparse_expected = SparseDataFrame(dense_expected)
-
- tm.assert_frame_equal(result, dense_expected)
- tm.assert_sp_frame_equal(result, sparse_expected)
-
- @pytest.mark.parametrize('data', [
- [[1, 1], [2, 2], [3, 3], [4, 4], [0, 0]],
- [[1.0, 1.0], [2.0, 2.0], [3.0, 3.0], [4.0, 4.0], [nan, nan]],
- [
- [1.0, 1.0 + 1.0j],
- [2.0 + 2.0j, 2.0],
- [3.0, 3.0 + 3.0j],
- [4.0 + 4.0j, 4.0],
- [nan, nan]
- ]
- ])
- @pytest.mark.parametrize('other', [
- True,
- -100,
- 0.1,
- 100.0 + 100.0j
- ])
- @pytest.mark.xfail(reason='Wrong SparseBlock initialization '
- '(GH 17386)')
- def test_where_with_numeric_data_and_other(self, data, other):
- # GH 17386
- lower_bound = 1.5
-
- sparse = SparseDataFrame(data)
- result = sparse.where(sparse > lower_bound, other)
-
- dense = DataFrame(data)
- dense_expected = dense.where(dense > lower_bound, other)
- sparse_expected = SparseDataFrame(dense_expected,
- default_fill_value=other)
-
- tm.assert_frame_equal(result, dense_expected)
- tm.assert_sp_frame_equal(result, sparse_expected)
-
- @pytest.mark.xfail(reason='Wrong SparseBlock initialization '
- '(GH 17386)')
- def test_where_with_bool_data(self):
- # GH 17386
- data = [[False, False], [True, True], [False, False]]
- cond = True
-
- sparse = SparseDataFrame(data)
- result = sparse.where(sparse == cond)
-
- dense = DataFrame(data)
- dense_expected = dense.where(dense == cond)
- sparse_expected = SparseDataFrame(dense_expected)
-
- tm.assert_frame_equal(result, dense_expected)
- tm.assert_sp_frame_equal(result, sparse_expected)
-
- @pytest.mark.parametrize('other', [
- True,
- 0,
- 0.1,
- 100.0 + 100.0j
- ])
- @pytest.mark.xfail(reason='Wrong SparseBlock initialization '
- '(GH 17386)')
- def test_where_with_bool_data_and_other(self, other):
- # GH 17386
- data = [[False, False], [True, True], [False, False]]
- cond = True
-
- sparse = SparseDataFrame(data)
- result = sparse.where(sparse == cond, other)
-
- dense = DataFrame(data)
- dense_expected = dense.where(dense == cond, other)
- sparse_expected = SparseDataFrame(dense_expected,
- default_fill_value=other)
-
- tm.assert_frame_equal(result, dense_expected)
- tm.assert_sp_frame_equal(result, sparse_expected)
-
@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
'(GH 17386)')
def test_quantile(self):
diff --git a/pandas/tests/sparse/frame/test_indexing.py b/pandas/tests/sparse/frame/test_indexing.py
new file mode 100644
index 0000000000000..1c27d44015c2b
--- /dev/null
+++ b/pandas/tests/sparse/frame/test_indexing.py
@@ -0,0 +1,113 @@
+import pytest
+import numpy as np
+from pandas import SparseDataFrame, DataFrame
+from pandas.util import testing as tm
+
+
+pytestmark = pytest.mark.skip("Wrong SparseBlock initialization (GH 17386)")
+
+
+@pytest.mark.parametrize('data', [
+ [[1, 1], [2, 2], [3, 3], [4, 4], [0, 0]],
+ [[1.0, 1.0], [2.0, 2.0], [3.0, 3.0], [4.0, 4.0], [np.nan, np.nan]],
+ [
+ [1.0, 1.0 + 1.0j],
+ [2.0 + 2.0j, 2.0],
+ [3.0, 3.0 + 3.0j],
+ [4.0 + 4.0j, 4.0],
+ [np.nan, np.nan]
+ ]
+])
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_where_with_numeric_data(data):
+ # GH 17386
+ lower_bound = 1.5
+
+ sparse = SparseDataFrame(data)
+ result = sparse.where(sparse > lower_bound)
+
+ dense = DataFrame(data)
+ dense_expected = dense.where(dense > lower_bound)
+ sparse_expected = SparseDataFrame(dense_expected)
+
+ tm.assert_frame_equal(result, dense_expected)
+ tm.assert_sp_frame_equal(result, sparse_expected)
+
+
+@pytest.mark.parametrize('data', [
+ [[1, 1], [2, 2], [3, 3], [4, 4], [0, 0]],
+ [[1.0, 1.0], [2.0, 2.0], [3.0, 3.0], [4.0, 4.0], [np.nan, np.nan]],
+ [
+ [1.0, 1.0 + 1.0j],
+ [2.0 + 2.0j, 2.0],
+ [3.0, 3.0 + 3.0j],
+ [4.0 + 4.0j, 4.0],
+ [np.nan, np.nan]
+ ]
+])
+@pytest.mark.parametrize('other', [
+ True,
+ -100,
+ 0.1,
+ 100.0 + 100.0j
+])
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_where_with_numeric_data_and_other(data, other):
+ # GH 17386
+ lower_bound = 1.5
+
+ sparse = SparseDataFrame(data)
+ result = sparse.where(sparse > lower_bound, other)
+
+ dense = DataFrame(data)
+ dense_expected = dense.where(dense > lower_bound, other)
+ sparse_expected = SparseDataFrame(dense_expected,
+ default_fill_value=other)
+
+ tm.assert_frame_equal(result, dense_expected)
+ tm.assert_sp_frame_equal(result, sparse_expected)
+
+
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_where_with_bool_data():
+ # GH 17386
+ data = [[False, False], [True, True], [False, False]]
+ cond = True
+
+ sparse = SparseDataFrame(data)
+ result = sparse.where(sparse == cond)
+
+ dense = DataFrame(data)
+ dense_expected = dense.where(dense == cond)
+ sparse_expected = SparseDataFrame(dense_expected)
+
+ tm.assert_frame_equal(result, dense_expected)
+ tm.assert_sp_frame_equal(result, sparse_expected)
+
+
+@pytest.mark.parametrize('other', [
+ True,
+ 0,
+ 0.1,
+ 100.0 + 100.0j
+])
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_where_with_bool_data_and_other(other):
+ # GH 17386
+ data = [[False, False], [True, True], [False, False]]
+ cond = True
+
+ sparse = SparseDataFrame(data)
+ result = sparse.where(sparse == cond, other)
+
+ dense = DataFrame(data)
+ dense_expected = dense.where(dense == cond, other)
+ sparse_expected = SparseDataFrame(dense_expected,
+ default_fill_value=other)
+
+ tm.assert_frame_equal(result, dense_expected)
+ tm.assert_sp_frame_equal(result, sparse_expected)
diff --git a/pandas/tests/sparse/series/__init__.py b/pandas/tests/sparse/series/__init__.py
new file mode 100644
index 0000000000000..e69de29bb2d1d
diff --git a/pandas/tests/sparse/series/test_indexing.py b/pandas/tests/sparse/series/test_indexing.py
new file mode 100644
index 0000000000000..de01b065a9fa0
--- /dev/null
+++ b/pandas/tests/sparse/series/test_indexing.py
@@ -0,0 +1,113 @@
+import pytest
+import numpy as np
+from pandas import SparseSeries, Series
+from pandas.util import testing as tm
+
+
+pytestmark = pytest.mark.skip("Wrong SparseBlock initialization (GH 17386)")
+
+
+@pytest.mark.parametrize('data', [
+ [1, 1, 2, 2, 3, 3, 4, 4, 0, 0],
+ [1.0, 1.0, 2.0, 2.0, 3.0, 3.0, 4.0, 4.0, np.nan, np.nan],
+ [
+ 1.0, 1.0 + 1.0j,
+ 2.0 + 2.0j, 2.0,
+ 3.0, 3.0 + 3.0j,
+ 4.0 + 4.0j, 4.0,
+ np.nan, np.nan
+ ]
+])
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_where_with_numeric_data(data):
+ # GH 17386
+ lower_bound = 1.5
+
+ sparse = SparseSeries(data)
+ result = sparse.where(sparse > lower_bound)
+
+ dense = Series(data)
+ dense_expected = dense.where(dense > lower_bound)
+ sparse_expected = SparseSeries(dense_expected)
+
+ tm.assert_series_equal(result, dense_expected)
+ tm.assert_sp_series_equal(result, sparse_expected)
+
+
+@pytest.mark.parametrize('data', [
+ [1, 1, 2, 2, 3, 3, 4, 4, 0, 0],
+ [1.0, 1.0, 2.0, 2.0, 3.0, 3.0, 4.0, 4.0, np.nan, np.nan],
+ [
+ 1.0, 1.0 + 1.0j,
+ 2.0 + 2.0j, 2.0,
+ 3.0, 3.0 + 3.0j,
+ 4.0 + 4.0j, 4.0,
+ np.nan, np.nan
+ ]
+])
+@pytest.mark.parametrize('other', [
+ True,
+ -100,
+ 0.1,
+ 100.0 + 100.0j
+])
+@pytest.mark.skip(reason='Wrong SparseBlock initialization '
+ '(Segfault) '
+ '(GH 17386)')
+def test_where_with_numeric_data_and_other(data, other):
+ # GH 17386
+ lower_bound = 1.5
+
+ sparse = SparseSeries(data)
+ result = sparse.where(sparse > lower_bound, other)
+
+ dense = Series(data)
+ dense_expected = dense.where(dense > lower_bound, other)
+ sparse_expected = SparseSeries(dense_expected, fill_value=other)
+
+ tm.assert_series_equal(result, dense_expected)
+ tm.assert_sp_series_equal(result, sparse_expected)
+
+
+@pytest.mark.xfail(reason='Wrong SparseBlock initialization '
+ '(GH 17386)')
+def test_where_with_bool_data():
+ # GH 17386
+ data = [False, False, True, True, False, False]
+ cond = True
+
+ sparse = SparseSeries(data)
+ result = sparse.where(sparse == cond)
+
+ dense = Series(data)
+ dense_expected = dense.where(dense == cond)
+ sparse_expected = SparseSeries(dense_expected)
+
+ tm.assert_series_equal(result, dense_expected)
+ tm.assert_sp_series_equal(result, sparse_expected)
+
+
+@pytest.mark.parametrize('other', [
+ True,
+ 0,
+ 0.1,
+ 100.0 + 100.0j
+])
+@pytest.mark.skip(reason='Wrong SparseBlock initialization '
+ '(Segfault) '
+ '(GH 17386)')
+def test_where_with_bool_data_and_other(other):
+ # GH 17386
+ data = [False, False, True, True, False, False]
+ cond = True
+
+ sparse = SparseSeries(data)
+ result = sparse.where(sparse == cond, other)
+
+ dense = Series(data)
+ dense_expected = dense.where(dense == cond, other)
+ sparse_expected = SparseSeries(dense_expected, fill_value=other)
+
+ tm.assert_series_equal(result, dense_expected)
+ tm.assert_sp_series_equal(result, sparse_expected)
diff --git a/pandas/tests/sparse/test_series.py b/pandas/tests/sparse/series/test_series.py
similarity index 94%
rename from pandas/tests/sparse/test_series.py
rename to pandas/tests/sparse/series/test_series.py
index 438e32b16f676..2ea1e63433520 100644
--- a/pandas/tests/sparse/test_series.py
+++ b/pandas/tests/sparse/series/test_series.py
@@ -1419,108 +1419,6 @@ def test_deprecated_reindex_axis(self):
self.bseries.reindex_axis([0, 1, 2])
assert 'reindex' in str(m[0].message)
- @pytest.mark.parametrize('data', [
- [1, 1, 2, 2, 3, 3, 4, 4, 0, 0],
- [1.0, 1.0, 2.0, 2.0, 3.0, 3.0, 4.0, 4.0, nan, nan],
- [
- 1.0, 1.0 + 1.0j,
- 2.0 + 2.0j, 2.0,
- 3.0, 3.0 + 3.0j,
- 4.0 + 4.0j, 4.0,
- nan, nan
- ]
- ])
- @pytest.mark.xfail(reason='Wrong SparseBlock initialization '
- '(GH 17386)')
- def test_where_with_numeric_data(self, data):
- # GH 17386
- lower_bound = 1.5
-
- sparse = SparseSeries(data)
- result = sparse.where(sparse > lower_bound)
-
- dense = Series(data)
- dense_expected = dense.where(dense > lower_bound)
- sparse_expected = SparseSeries(dense_expected)
-
- tm.assert_series_equal(result, dense_expected)
- tm.assert_sp_series_equal(result, sparse_expected)
-
- @pytest.mark.parametrize('data', [
- [1, 1, 2, 2, 3, 3, 4, 4, 0, 0],
- [1.0, 1.0, 2.0, 2.0, 3.0, 3.0, 4.0, 4.0, nan, nan],
- [
- 1.0, 1.0 + 1.0j,
- 2.0 + 2.0j, 2.0,
- 3.0, 3.0 + 3.0j,
- 4.0 + 4.0j, 4.0,
- nan, nan
- ]
- ])
- @pytest.mark.parametrize('other', [
- True,
- -100,
- 0.1,
- 100.0 + 100.0j
- ])
- @pytest.mark.skip(reason='Wrong SparseBlock initialization '
- '(Segfault) '
- '(GH 17386)')
- def test_where_with_numeric_data_and_other(self, data, other):
- # GH 17386
- lower_bound = 1.5
-
- sparse = SparseSeries(data)
- result = sparse.where(sparse > lower_bound, other)
-
- dense = Series(data)
- dense_expected = dense.where(dense > lower_bound, other)
- sparse_expected = SparseSeries(dense_expected, fill_value=other)
-
- tm.assert_series_equal(result, dense_expected)
- tm.assert_sp_series_equal(result, sparse_expected)
-
- @pytest.mark.xfail(reason='Wrong SparseBlock initialization '
- '(GH 17386)')
- def test_where_with_bool_data(self):
- # GH 17386
- data = [False, False, True, True, False, False]
- cond = True
-
- sparse = SparseSeries(data)
- result = sparse.where(sparse == cond)
-
- dense = Series(data)
- dense_expected = dense.where(dense == cond)
- sparse_expected = SparseSeries(dense_expected)
-
- tm.assert_series_equal(result, dense_expected)
- tm.assert_sp_series_equal(result, sparse_expected)
-
- @pytest.mark.parametrize('other', [
- True,
- 0,
- 0.1,
- 100.0 + 100.0j
- ])
- @pytest.mark.skip(reason='Wrong SparseBlock initialization '
- '(Segfault) '
- '(GH 17386)')
- def test_where_with_bool_data_and_other(self, other):
- # GH 17386
- data = [False, False, True, True, False, False]
- cond = True
-
- sparse = SparseSeries(data)
- result = sparse.where(sparse == cond, other)
-
- dense = Series(data)
- dense_expected = dense.where(dense == cond, other)
- sparse_expected = SparseSeries(dense_expected, fill_value=other)
-
- tm.assert_series_equal(result, dense_expected)
- tm.assert_sp_series_equal(result, sparse_expected)
-
@pytest.mark.parametrize(
'datetime_type', (np.datetime64,
| https://api.github.com/repos/pandas-dev/pandas/pulls/18968 | 2017-12-28T13:17:20Z | 2017-12-28T17:22:02Z | 2017-12-28T17:22:02Z | 2017-12-28T17:22:28Z | |
Misc. Typo fixes | diff --git a/appveyor.yml b/appveyor.yml
index 44af73b498aa8..0aaac322c4ac7 100644
--- a/appveyor.yml
+++ b/appveyor.yml
@@ -11,7 +11,7 @@ matrix:
environment:
global:
# SDK v7.0 MSVC Express 2008's SetEnv.cmd script will fail if the
- # /E:ON and /V:ON options are not enabled in the batch script intepreter
+ # /E:ON and /V:ON options are not enabled in the batch script interpreter
# See: http://stackoverflow.com/a/13751649/163740
CMD_IN_ENV: "cmd /E:ON /V:ON /C .\\ci\\run_with_env.cmd"
clone_folder: C:\projects\pandas
diff --git a/asv_bench/benchmarks/frame_ctor.py b/asv_bench/benchmarks/frame_ctor.py
index 391a209cb2a89..21b20cb123ed6 100644
--- a/asv_bench/benchmarks/frame_ctor.py
+++ b/asv_bench/benchmarks/frame_ctor.py
@@ -4,7 +4,7 @@
try:
from pandas.tseries.offsets import Nano, Hour
except ImportError:
- # For compatability with older versions
+ # For compatibility with older versions
from pandas.core.datetools import * # noqa
from .pandas_vb_common import setup # noqa
diff --git a/asv_bench/benchmarks/pandas_vb_common.py b/asv_bench/benchmarks/pandas_vb_common.py
index 4de87ddcb0683..7b4fec0090701 100644
--- a/asv_bench/benchmarks/pandas_vb_common.py
+++ b/asv_bench/benchmarks/pandas_vb_common.py
@@ -18,8 +18,8 @@
np.float64, np.int16, np.int8, np.uint16, np.uint8]
datetime_dtypes = [np.datetime64, np.timedelta64]
-# This function just needs to be imported into each benchmark file in order to
-# sets up the random seed before each function.
+# This function just needs to be imported into each benchmark file in order to
+# sets up the random seed before each function.
# http://asv.readthedocs.io/en/latest/writing_benchmarks.html
def setup(*args, **kwargs):
np.random.seed(1234)
@@ -36,14 +36,14 @@ def remove(self, f):
try:
os.remove(f)
except:
- # On Windows, attempting to remove a file that is in use
+ # On Windows, attempting to remove a file that is in use
# causes an exception to be raised
pass
def teardown(self, *args, **kwargs):
self.remove(self.fname)
-# Compatability import for lib
+# Compatibility import for lib
for imp in ['pandas._libs.lib', 'pandas.lib', 'pandas_tseries']:
try:
lib = import_module(imp)
diff --git a/doc/source/_static/banklist.html b/doc/source/_static/banklist.html
index 8ec1561f8c394..cbcce5a2d49ff 100644
--- a/doc/source/_static/banklist.html
+++ b/doc/source/_static/banklist.html
@@ -7,7 +7,7 @@
<meta charset="UTF-8">
<!-- Unicode character encoding -->
<meta http-equiv="X-UA-Compatible" content="IE=edge">
-<!-- Turns off IE Compatiblity Mode -->
+<!-- Turns off IE Compatibility Mode -->
<meta name="viewport" content="width=device-width, initial-scale=1, maximum-scale=1">
<!-- Makes it so phones don't auto zoom out. -->
<meta name="author" content="DRR">
@@ -4849,7 +4849,7 @@ <h1 class="page_title">Failed Bank List</h1>
<ul>
<li><a href="/about/freedom/" title="Freedom of Information Act (FOIA) Service Center">Freedom of Information Act (FOIA) Service Center</a></li>
<li><a href="/open/" title="FDIC Open Government Webpage">FDIC Open Government Webpage</a></li>
- <li><a href="/about/diversity/nofear/" title="No FEAR Act Data">No FEAR Act Data</a></li>
+ <li><a href="/about/diversity/nofear/" title="No FEAR Act Data">No FEAR Act Data</a></li>
</ul>
</div>
<div id="responsive_footer-small">
diff --git a/doc/source/api.rst b/doc/source/api.rst
index 68721b76eed7e..17f6b8df0170d 100644
--- a/doc/source/api.rst
+++ b/doc/source/api.rst
@@ -10,7 +10,7 @@ methods. In general, all classes and functions exposed in the top-level
``pandas.*`` namespace are regarded as public.
Further some of the subpackages are public, including ``pandas.errors``,
-``pandas.plotting``, and ``pandas.testing``. Certain functions in the the
+``pandas.plotting``, and ``pandas.testing``. Certain functions in the
``pandas.io`` and ``pandas.tseries`` submodules are public as well (those
mentioned in the documentation). Further, the ``pandas.api.types`` subpackage
holds some public functions related to data types in pandas.
diff --git a/doc/source/basics.rst b/doc/source/basics.rst
index da82f56d315e6..74b3dbb83ea91 100644
--- a/doc/source/basics.rst
+++ b/doc/source/basics.rst
@@ -947,7 +947,7 @@ Mixed Dtypes
++++++++++++
When presented with mixed dtypes that cannot aggregate, ``.agg`` will only take the valid
-aggregations. This is similiar to how groupby ``.agg`` works.
+aggregations. This is similar to how groupby ``.agg`` works.
.. ipython:: python
diff --git a/doc/source/computation.rst b/doc/source/computation.rst
index 0994d35999191..30071c6c5b83c 100644
--- a/doc/source/computation.rst
+++ b/doc/source/computation.rst
@@ -247,7 +247,7 @@ These are created from methods on ``Series`` and ``DataFrame``.
r = s.rolling(window=60)
r
-These object provide tab-completion of the avaible methods and properties.
+These object provide tab-completion of the available methods and properties.
.. code-block:: ipython
diff --git a/doc/source/conf.py b/doc/source/conf.py
index bcb83d5699d7e..c188f83f80250 100644
--- a/doc/source/conf.py
+++ b/doc/source/conf.py
@@ -380,7 +380,7 @@
'import pandas as pd',
# This ensures correct rendering on system with console encoding != utf8
# (windows). It forces pandas to encode its output reprs using utf8
- # whereever the docs are built. The docs' target is the browser, not
+ # wherever the docs are built. The docs' target is the browser, not
# the console, so this is fine.
'pd.options.display.encoding="utf8"'
]
diff --git a/doc/source/contributing.rst b/doc/source/contributing.rst
index 0e5d701353d78..b25f9779d3636 100644
--- a/doc/source/contributing.rst
+++ b/doc/source/contributing.rst
@@ -124,16 +124,16 @@ to build the documentation locally before pushing your changes.
.. _contributiong.dev_c:
-Installing a C Complier
+Installing a C Compiler
~~~~~~~~~~~~~~~~~~~~~~~
Pandas uses C extensions (mostly written using Cython) to speed up certain
operations. To install pandas from source, you need to compile these C
-extensions, which means you need a C complier. This process depends on which
+extensions, which means you need a C compiler. This process depends on which
platform you're using. Follow the `CPython contributing guidelines
<https://docs.python.org/devguide/setup.html#build-dependencies>`_ for getting a
-complier installed. You don't need to do any of the ``./configure`` or ``make``
-steps; you only need to install the complier.
+compiler installed. You don't need to do any of the ``./configure`` or ``make``
+steps; you only need to install the compiler.
For Windows developers, the following links may be helpful.
@@ -151,7 +151,7 @@ Let us know if you have any difficulties by opening an issue or reaching out on
Creating a Python Environment
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
-Now that you have a C complier, create an isolated pandas development
+Now that you have a C compiler, create an isolated pandas development
environment:
- Install either `Anaconda <https://www.anaconda.com/download/>`_ or `miniconda
diff --git a/doc/source/developer.rst b/doc/source/developer.rst
index 9c214020ab43d..b8bb2b2fcbe2f 100644
--- a/doc/source/developer.rst
+++ b/doc/source/developer.rst
@@ -40,7 +40,7 @@ where ``KeyValue`` is
}
So that a ``pandas.DataFrame`` can be faithfully reconstructed, we store a
-``pandas`` metadata key in the ``FileMetaData`` with the the value stored as :
+``pandas`` metadata key in the ``FileMetaData`` with the value stored as :
.. code-block:: text
diff --git a/doc/source/dsintro.rst b/doc/source/dsintro.rst
index c8018c8e66f72..da9d2123bd1ca 100644
--- a/doc/source/dsintro.rst
+++ b/doc/source/dsintro.rst
@@ -470,7 +470,7 @@ derived from existing columns.
.head())
In the example above, we inserted a precomputed value. We can also pass in
-a function of one argument to be evalutated on the DataFrame being assigned to.
+a function of one argument to be evaluated on the DataFrame being assigned to.
.. ipython:: python
@@ -957,7 +957,7 @@ pandas to focus on these areas exclusively.
Oftentimes, one can simply use a MultiIndex ``DataFrame`` for easily working with higher dimensional data.
-In additon, the ``xarray`` package was built from the ground up, specifically in order to
+In addition, the ``xarray`` package was built from the ground up, specifically in order to
support the multi-dimensional analysis that is one of ``Panel`` s main usecases.
`Here is a link to the xarray panel-transition documentation <http://xarray.pydata.org/en/stable/pandas.html#panel-transition>`__.
diff --git a/doc/source/enhancingperf.rst b/doc/source/enhancingperf.rst
index d2ca76713ba3b..362c998493ae8 100644
--- a/doc/source/enhancingperf.rst
+++ b/doc/source/enhancingperf.rst
@@ -173,7 +173,7 @@ Using ndarray
It's calling series... a lot! It's creating a Series from each row, and get-ting from both
the index and the series (three times for each row). Function calls are expensive
-in python, so maybe we could minimise these by cythonizing the apply part.
+in python, so maybe we could minimize these by cythonizing the apply part.
.. note::
@@ -578,7 +578,7 @@ on the original ``DataFrame`` or return a copy with the new column.
.. warning::
- For backwards compatability, ``inplace`` defaults to ``True`` if not
+ For backwards compatibility, ``inplace`` defaults to ``True`` if not
specified. This will change in a future version of pandas - if your
code depends on an inplace assignment you should update to explicitly
set ``inplace=True``
diff --git a/doc/source/indexing.rst b/doc/source/indexing.rst
index b9223c6ad9f7a..355be5039f146 100644
--- a/doc/source/indexing.rst
+++ b/doc/source/indexing.rst
@@ -651,7 +651,7 @@ Indexing with list with missing labels is Deprecated
In prior versions, using ``.loc[list-of-labels]`` would work as long as *at least 1* of the keys was found (otherwise it
would raise a ``KeyError``). This behavior is deprecated and will show a warning message pointing to this section. The
-recommeded alternative is to use ``.reindex()``.
+recommended alternative is to use ``.reindex()``.
For example.
@@ -724,7 +724,7 @@ Having a duplicated index will raise for a ``.reindex()``:
In [17]: s.reindex(labels)
ValueError: cannot reindex from a duplicate axis
-Generally, you can interesect the desired labels with the current
+Generally, you can intersect the desired labels with the current
axis, and then reindex.
.. ipython:: python
diff --git a/doc/source/install.rst b/doc/source/install.rst
index 6133da220aa8d..c4e331d64e721 100644
--- a/doc/source/install.rst
+++ b/doc/source/install.rst
@@ -152,7 +152,7 @@ To install pandas for Python 2 you may need to use the package ``python-pandas``
Debian, stable, `official Debian repository <http://packages.debian.org/search?keywords=pandas&searchon=names&suite=all§ion=all>`__ , ``sudo apt-get install python3-pandas``
Debian & Ubuntu, unstable (latest packages), `NeuroDebian <http://neuro.debian.net/index.html#how-to-use-this-repository>`__ , ``sudo apt-get install python3-pandas``
Ubuntu, stable, `official Ubuntu repository <http://packages.ubuntu.com/search?keywords=pandas&searchon=names&suite=all§ion=all>`__ , ``sudo apt-get install python3-pandas``
- OpenSuse, stable, `OpenSuse Repository <http://software.opensuse.org/package/python-pandas?search_term=pandas>`__ , ``zypper in python3-pandas``
+ OpenSuse, stable, `OpenSuse Repository <http://software.opensuse.org/package/python-pandas?search_term=pandas>`__ , ``zypper in python3-pandas``
Fedora, stable, `official Fedora repository <https://admin.fedoraproject.org/pkgdb/package/rpms/python-pandas/>`__ , ``dnf install python3-pandas``
Centos/RHEL, stable, `EPEL repository <https://admin.fedoraproject.org/pkgdb/package/rpms/python-pandas/>`__ , ``yum install python3-pandas``
diff --git a/doc/source/io.rst b/doc/source/io.rst
index a5a0a41147a6b..49d742d9905d7 100644
--- a/doc/source/io.rst
+++ b/doc/source/io.rst
@@ -1048,7 +1048,7 @@ The ``thousands`` keyword allows integers to be parsed correctly
NA Values
'''''''''
-To control which values are parsed as missing values (which are signified by ``NaN``), specifiy a
+To control which values are parsed as missing values (which are signified by ``NaN``), specify a
string in ``na_values``. If you specify a list of strings, then all values in
it are considered to be missing values. If you specify a number (a ``float``, like ``5.0`` or an ``integer`` like ``5``),
the corresponding equivalent values will also imply a missing value (in this case effectively
@@ -4153,7 +4153,7 @@ Caveats
.. warning::
- ``PyTables`` will show a ``NaturalNameWarning`` if a column name
+ ``PyTables`` will show a ``NaturalNameWarning`` if a column name
cannot be used as an attribute selector.
*Natural* identifiers contain only letters, numbers, and underscores,
and may not begin with a number.
@@ -4478,7 +4478,7 @@ Several caveats.
- Non supported types include ``Period`` and actual python object types. These will raise a helpful error message
on an attempt at serialization.
-You can specifiy an ``engine`` to direct the serialization. This can be one of ``pyarrow``, or ``fastparquet``, or ``auto``.
+You can specify an ``engine`` to direct the serialization. This can be one of ``pyarrow``, or ``fastparquet``, or ``auto``.
If the engine is NOT specified, then the ``pd.options.io.parquet.engine`` option is checked; if this is also ``auto``, then
then ``pyarrow`` is tried, and falling back to ``fastparquet``.
diff --git a/doc/source/options.rst b/doc/source/options.rst
index 5641b2628fe40..cce16a5396377 100644
--- a/doc/source/options.rst
+++ b/doc/source/options.rst
@@ -164,7 +164,7 @@ lines are replaced by an ellipsis.
df
pd.reset_option('max_rows')
-``display.expand_frame_repr`` allows for the the representation of
+``display.expand_frame_repr`` allows for the representation of
dataframes to stretch across pages, wrapped over the full column vs row-wise.
.. ipython:: python
diff --git a/doc/source/overview.rst b/doc/source/overview.rst
index 73e7704b43be6..4443428ca6c9b 100644
--- a/doc/source/overview.rst
+++ b/doc/source/overview.rst
@@ -109,7 +109,7 @@ Wes McKinney is the Benevolent Dictator for Life (BDFL).
Development Team
-----------------
-The list of the Core Team members and more detailed information can be found on the `people’s page <https://github.com/pandas-dev/pandas-governance/blob/master/people.md>`__ of the governance repo.
+The list of the Core Team members and more detailed information can be found on the `people’s page <https://github.com/pandas-dev/pandas-governance/blob/master/people.md>`__ of the governance repo.
Institutional Partners
diff --git a/doc/source/release.rst b/doc/source/release.rst
index aea6280a490d6..12932d9fcee4f 100644
--- a/doc/source/release.rst
+++ b/doc/source/release.rst
@@ -2043,7 +2043,7 @@ Bug Fixes
- Fixed missing arg validation in get_options_data (:issue:`6105`)
- Bug in assignment with duplicate columns in a frame where the locations
are a slice (e.g. next to each other) (:issue:`6120`)
-- Bug in propogating _ref_locs during construction of a DataFrame with dups
+- Bug in propagating _ref_locs during construction of a DataFrame with dups
index/columns (:issue:`6121`)
- Bug in ``DataFrame.apply`` when using mixed datelike reductions (:issue:`6125`)
- Bug in ``DataFrame.append`` when appending a row with different columns (:issue:`6129`)
@@ -2056,7 +2056,7 @@ Bug Fixes
- Bug in ``HDFStore`` on appending a dataframe with multi-indexed columns to
an existing table (:issue:`6167`)
- Consistency with dtypes in setting an empty DataFrame (:issue:`6171`)
-- Bug in selecting on a multi-index ``HDFStore`` even in the presence of under
+- Bug in selecting on a multi-index ``HDFStore`` even in the presence of under
specified column spec (:issue:`6169`)
- Bug in ``nanops.var`` with ``ddof=1`` and 1 elements would sometimes return ``inf``
rather than ``nan`` on some platforms (:issue:`6136`)
@@ -2437,7 +2437,7 @@ API Changes
- The refactoring involving``Series`` deriving from ``NDFrame`` breaks ``rpy2<=2.3.8``. an Issue
has been opened against rpy2 and a workaround is detailed in :issue:`5698`. Thanks @JanSchulz.
- ``Series.argmin`` and ``Series.argmax`` are now aliased to ``Series.idxmin`` and ``Series.idxmax``.
- These return the *index* of the min or max element respectively. Prior to 0.13.0 these would return
+ These return the *index* of the min or max element respectively. Prior to 0.13.0 these would return
the position of the min / max element (:issue:`6214`)
Internal Refactoring
@@ -3097,7 +3097,7 @@ Bug Fixes
- Fixed bug where a time-series was being selected in preference to an actual column name
in a frame (:issue:`3594`)
- Make secondary_y work properly for bar plots (:issue:`3598`)
-- Fix modulo and integer division on Series,DataFrames to act similary to ``float`` dtypes to return
+- Fix modulo and integer division on Series,DataFrames to act similarly to ``float`` dtypes to return
``np.nan`` or ``np.inf`` as appropriate (:issue:`3590`)
- Fix incorrect dtype on groupby with ``as_index=False`` (:issue:`3610`)
- Fix ``read_csv/read_excel`` to correctly encode identical na_values, e.g. ``na_values=[-999.0,-999]``
@@ -3400,11 +3400,11 @@ Bug Fixes
- Fixed bug in reshape if not passed correct input, now raises TypeError (:issue:`2719`)
- Fixed a bug where Series ctor did not respect ordering if OrderedDict passed in (:issue:`3282`)
- Fix NameError issue on RESO_US (:issue:`2787`)
-- Allow selection in an *unordered* timeseries to work similary
+- Allow selection in an *unordered* timeseries to work similarly
to an *ordered* timeseries (:issue:`2437`).
- Fix implemented ``.xs`` when called with ``axes=1`` and a level parameter (:issue:`2903`)
- Timestamp now supports the class method fromordinal similar to datetimes (:issue:`3042`)
-- Fix issue with indexing a series with a boolean key and specifiying a 1-len list on the rhs (:issue:`2745`)
+- Fix issue with indexing a series with a boolean key and specifying a 1-len list on the rhs (:issue:`2745`)
or a list on the rhs (:issue:`3235`)
- Fixed bug in groupby apply when kernel generate list of arrays having unequal len (:issue:`1738`)
- fixed handling of rolling_corr with center=True which could produce corr>1 (:issue:`3155`)
@@ -3555,7 +3555,7 @@ Bug Fixes
- Upconvert datetime + datetime64 values when concatenating frames (:issue:`2624`)
- Raise a more helpful error message in merge operations when one DataFrame
has duplicate columns (:issue:`2649`)
-- Fix partial date parsing issue occuring only when code is run at EOM
+- Fix partial date parsing issue occurring only when code is run at EOM
(:issue:`2618`)
- Prevent MemoryError when using counting sort in sortlevel with
high-cardinality MultiIndex objects (:issue:`2684`)
@@ -3973,7 +3973,7 @@ Bug Fixes
- Don't lose tzinfo when passing DatetimeIndex as DataFrame column (:issue:`1682`)
- Fix tz conversion with time zones that haven't had any DST transitions since
first date in the array (:issue:`1673`)
-- Fix field access with UTC->local conversion on unsorted arrays (:issue:`1756`)
+- Fix field access with UTC->local conversion on unsorted arrays (:issue:`1756`)
- Fix isnull handling of array-like (list) inputs (:issue:`1755`)
- Fix regression in handling of Series in Series constructor (:issue:`1671`)
- Fix comparison of Int64Index with DatetimeIndex (:issue:`1681`)
@@ -4525,7 +4525,7 @@ Bug Fixes
- Fix na-filling handling in mixed-type DataFrame (:issue:`910`)
- Fix to DataFrame.set_value with non-existant row/col (:issue:`911`)
- Fix malformed block in groupby when excluding nuisance columns (:issue:`916`)
-- Fix inconsistant NA handling in dtype=object arrays (:issue:`925`)
+- Fix inconsistent NA handling in dtype=object arrays (:issue:`925`)
- Fix missing center-of-mass computation in ewmcov (:issue:`862`)
- Don't raise exception when opening read-only HDF5 file (:issue:`847`)
- Fix possible out-of-bounds memory access in 0-length Series (:issue:`917`)
@@ -5395,9 +5395,9 @@ pandas 0.4.3
**Release date:** 10/9/2011
-is is largely a bugfix release from 0.4.2 but also includes a handful of new
-d enhanced features. Also, pandas can now be installed and used on Python 3
-hanks Thomas Kluyver!).
+This is largely a bugfix release from 0.4.2 but also includes a handful of new
+and enhanced features. Also, pandas can now be installed and used on Python 3
+(thanks Thomas Kluyver!).
New Features
~~~~~~~~~~~~
@@ -5460,9 +5460,9 @@ pandas 0.4.2
**Release date:** 10/3/2011
-is is a performance optimization release with several bug fixes. The new
-t64Index and new merging / joining Cython code and related Python
-frastructure are the main new additions
+This is a performance optimization release with several bug fixes. The new
+Int64Index and new merging / joining Cython code and related Python
+infrastructure are the main new additions
New Features
~~~~~~~~~~~~
@@ -5537,7 +5537,7 @@ pandas 0.4.1
**Release date:** 9/25/2011
-is is primarily a bug fix release but includes some new features and
+This is primarily a bug fix release but includes some new features and
improvements
New Features
diff --git a/doc/source/style.ipynb b/doc/source/style.ipynb
index 20f7c2a93b9e6..152ca90049bf1 100644
--- a/doc/source/style.ipynb
+++ b/doc/source/style.ipynb
@@ -318,7 +318,7 @@
"Both `Styler.apply`, and `Styler.applymap` accept a `subset` keyword.\n",
"This allows you to apply styles to specific rows or columns, without having to code that logic into your `style` function.\n",
"\n",
- "The value passed to `subset` behaves simlar to slicing a DataFrame.\n",
+ "The value passed to `subset` behaves similar to slicing a DataFrame.\n",
"\n",
"- A scalar is treated as a column label\n",
"- A list (or series or numpy array)\n",
diff --git a/doc/source/timeseries.rst b/doc/source/timeseries.rst
index 26e701d008b3f..201af3c7d5355 100644
--- a/doc/source/timeseries.rst
+++ b/doc/source/timeseries.rst
@@ -1580,7 +1580,7 @@ We can instead only resample those groups where we have points as follows:
Aggregation
~~~~~~~~~~~
-Similar to the :ref:`aggregating API <basics.aggregate>`, :ref:`groupby API <groupby.aggregate>`, and the :ref:`window functions API <stats.aggregate>`,
+Similar to the :ref:`aggregating API <basics.aggregate>`, :ref:`groupby API <groupby.aggregate>`, and the :ref:`window functions API <stats.aggregate>`,
a ``Resampler`` can be selectively resampled.
Resampling a ``DataFrame``, the default will be to act on all columns with the same function.
@@ -2108,7 +2108,7 @@ tz-aware data to another time zone:
It is incorrect to pass a timezone directly into the ``datetime.datetime`` constructor (e.g.,
``datetime.datetime(2011, 1, 1, tz=timezone('US/Eastern'))``. Instead, the datetime
- needs to be localized using the the localize method on the timezone.
+ needs to be localized using the localize method on the timezone.
Under the hood, all timestamps are stored in UTC. Scalar values from a
``DatetimeIndex`` with a time zone will have their fields (day, hour, minute)
diff --git a/doc/source/visualization.rst b/doc/source/visualization.rst
index ea720a9ae4ed0..2c1d54c27caab 100644
--- a/doc/source/visualization.rst
+++ b/doc/source/visualization.rst
@@ -140,7 +140,7 @@ You can also create these other plots using the methods ``DataFrame.plot.<kind>`
df.plot.area df.plot.barh df.plot.density df.plot.hist df.plot.line df.plot.scatter
df.plot.bar df.plot.box df.plot.hexbin df.plot.kde df.plot.pie
-In addition to these ``kind`` s, there are the :ref:`DataFrame.hist() <visualization.hist>`,
+In addition to these ``kind`` s, there are the :ref:`DataFrame.hist() <visualization.hist>`,
and :ref:`DataFrame.boxplot() <visualization.box>` methods, which use a separate interface.
Finally, there are several :ref:`plotting functions <visualization.tools>` in ``pandas.plotting``
@@ -716,7 +716,7 @@ You can use the ``labels`` and ``colors`` keywords to specify the labels and col
.. warning::
- Most pandas plots use the the ``label`` and ``color`` arguments (note the lack of "s" on those).
+ Most pandas plots use the ``label`` and ``color`` arguments (note the lack of "s" on those).
To be consistent with :func:`matplotlib.pyplot.pie` you must use ``labels`` and ``colors``.
If you want to hide wedge labels, specify ``labels=None``.
@@ -1187,7 +1187,7 @@ time-series data. For limited cases where pandas cannot infer the frequency
information (e.g., in an externally created ``twinx``), you can choose to
suppress this behavior for alignment purposes.
-Here is the default behavior, notice how the x-axis tick labelling is performed:
+Here is the default behavior, notice how the x-axis tick labeling is performed:
.. ipython:: python
diff --git a/doc/source/whatsnew/v0.10.0.txt b/doc/source/whatsnew/v0.10.0.txt
index f0db1d82252c1..a0c4a3e0073f9 100644
--- a/doc/source/whatsnew/v0.10.0.txt
+++ b/doc/source/whatsnew/v0.10.0.txt
@@ -369,7 +369,7 @@ Updated PyTables Support
df1
df1.get_dtype_counts()
-- performance improvments on table writing
+- performance improvements on table writing
- support for arbitrarily indexed dimensions
- ``SparseSeries`` now has a ``density`` property (:issue:`2384`)
- enable ``Series.str.strip/lstrip/rstrip`` methods to take an input argument
diff --git a/doc/source/whatsnew/v0.10.1.txt b/doc/source/whatsnew/v0.10.1.txt
index d5880e44e46c6..2d5843101dec2 100644
--- a/doc/source/whatsnew/v0.10.1.txt
+++ b/doc/source/whatsnew/v0.10.1.txt
@@ -153,7 +153,7 @@ combined result, by using ``where`` on a selector table.
table
- You can pass ``chunksize=an integer`` to ``append``, to change the writing
- chunksize (default is 50000). This will signficantly lower your memory usage
+ chunksize (default is 50000). This will significantly lower your memory usage
on writing.
- You can pass ``expectedrows=an integer`` to the first ``append``, to set the
diff --git a/doc/source/whatsnew/v0.11.0.txt b/doc/source/whatsnew/v0.11.0.txt
index ea149595e681f..b90a597815ec5 100644
--- a/doc/source/whatsnew/v0.11.0.txt
+++ b/doc/source/whatsnew/v0.11.0.txt
@@ -88,7 +88,7 @@ Numeric dtypes will propagate and can coexist in DataFrames. If a dtype is passe
Dtype Conversion
~~~~~~~~~~~~~~~~
-This is lower-common-denomicator upcasting, meaning you get the dtype which can accomodate all of the types
+This is lower-common-denominator upcasting, meaning you get the dtype which can accommodate all of the types
.. ipython:: python
@@ -193,7 +193,7 @@ Furthermore ``datetime64[ns]`` columns are created by default, when passed datet
df.loc[df.index[2:4], ['A','timestamp']] = np.nan
df
-Astype conversion on ``datetime64[ns]`` to ``object``, implicity converts ``NaT`` to ``np.nan``
+Astype conversion on ``datetime64[ns]`` to ``object``, implicitly converts ``NaT`` to ``np.nan``
.. ipython:: python
diff --git a/doc/source/whatsnew/v0.12.0.txt b/doc/source/whatsnew/v0.12.0.txt
index 27aa47a6bb097..ad33c49792d9f 100644
--- a/doc/source/whatsnew/v0.12.0.txt
+++ b/doc/source/whatsnew/v0.12.0.txt
@@ -38,7 +38,7 @@ API changes
* ``to_clipboard``
- - Fix modulo and integer division on Series,DataFrames to act similary to ``float`` dtypes to return
+ - Fix modulo and integer division on Series,DataFrames to act similarly to ``float`` dtypes to return
``np.nan`` or ``np.inf`` as appropriate (:issue:`3590`). This correct a numpy bug that treats ``integer``
and ``float`` dtypes differently.
@@ -154,7 +154,7 @@ API changes
- The behavior of ``datetime64`` dtypes has changed with respect to certain
so-called reduction operations (:issue:`3726`). The following operations now
- raise a ``TypeError`` when perfomed on a ``Series`` and return an *empty*
+ raise a ``TypeError`` when performed on a ``Series`` and return an *empty*
``Series`` when performed on a ``DataFrame`` similar to performing these
operations on, for example, a ``DataFrame`` of ``slice`` objects:
@@ -206,11 +206,11 @@ I/O Enhancements
:ref:`See the installation docs<install.optional_dependencies>`
- Added module for reading and writing Stata files: ``pandas.io.stata`` (:issue:`1512`)
- accessable via ``read_stata`` top-level function for reading,
+ accessible via ``read_stata`` top-level function for reading,
and ``to_stata`` DataFrame method for writing, :ref:`See the docs<io.stata>`
- Added module for reading and writing json format files: ``pandas.io.json``
- accessable via ``read_json`` top-level function for reading,
+ accessible via ``read_json`` top-level function for reading,
and ``to_json`` DataFrame method for writing, :ref:`See the docs<io.json>`
various issues (:issue:`1226`, :issue:`3804`, :issue:`3876`, :issue:`3867`, :issue:`1305`)
@@ -220,7 +220,7 @@ I/O Enhancements
list of the rows from which to read the index.
- The option, ``tupleize_cols`` can now be specified in both ``to_csv`` and
- ``read_csv``, to provide compatiblity for the pre 0.12 behavior of
+ ``read_csv``, to provide compatibility for the pre 0.12 behavior of
writing and reading ``MultIndex`` columns via a list of tuples. The default in
0.12 is to write lists of tuples and *not* interpret list of tuples as a
``MultiIndex`` column.
diff --git a/doc/source/whatsnew/v0.14.0.txt b/doc/source/whatsnew/v0.14.0.txt
index f1feab4b909dc..be962ceb181ff 100644
--- a/doc/source/whatsnew/v0.14.0.txt
+++ b/doc/source/whatsnew/v0.14.0.txt
@@ -83,7 +83,7 @@ API changes
been removed, instead a header with the column names is returned (:issue:`6062`).
- ``Series`` and ``Index`` now internall share more common operations, e.g. ``factorize(),nunique(),value_counts()`` are
now supported on ``Index`` types as well. The ``Series.weekday`` property from is removed
- from Series for API consistency. Using a ``DatetimeIndex/PeriodIndex`` method on a Series will now raise a ``TypeError``.
+ from Series for API consistency. Using a ``DatetimeIndex/PeriodIndex`` method on a Series will now raise a ``TypeError``.
(:issue:`4551`, :issue:`4056`, :issue:`5519`, :issue:`6380`, :issue:`7206`).
- Add ``is_month_start``, ``is_month_end``, ``is_quarter_start``, ``is_quarter_end``, ``is_year_start``, ``is_year_end`` accessors for ``DateTimeIndex`` / ``Timestamp`` which return a boolean array of whether the timestamp(s) are at the start/end of the month/quarter/year defined by the frequency of the ``DateTimeIndex`` / ``Timestamp`` (:issue:`4565`, :issue:`6998`)
@@ -284,7 +284,7 @@ Display Changes
`large_repr` set to 'info' (:issue:`7105`)
- The `verbose` keyword in ``DataFrame.info()``, which controls whether to shorten the ``info``
representation, is now ``None`` by default. This will follow the global setting in
- ``display.max_info_columns``. The global setting can be overriden with ``verbose=True`` or
+ ``display.max_info_columns``. The global setting can be overridden with ``verbose=True`` or
``verbose=False``.
- Fixed a bug with the `info` repr not honoring the `display.max_info_columns` setting (:issue:`6939`)
- Offset/freq info now in Timestamp __repr__ (:issue:`4553`)
@@ -446,7 +446,7 @@ Some other enhancements to the sql functions include:
- support for writing the index. This can be controlled with the ``index``
keyword (default is True).
- specify the column label to use when writing the index with ``index_label``.
-- specify string columns to parse as datetimes withh the ``parse_dates``
+- specify string columns to parse as datetimes with the ``parse_dates``
keyword in :func:`~pandas.read_sql_query` and :func:`~pandas.read_sql_table`.
.. warning::
@@ -596,15 +596,15 @@ Plotting
- `align`: Specify the bar alignment. Default is `center` (different from matplotlib). In previous versions, pandas passes `align='edge'` to matplotlib and adjust the location to `center` by itself, and it results `align` keyword is not applied as expected. (:issue:`4525`)
- `position`: Specify relative alignments for bar plot layout. From 0 (left/bottom-end) to 1(right/top-end). Default is 0.5 (center). (:issue:`6604`)
- Because of the default `align` value changes, coordinates of bar plots are now located on integer values (0.0, 1.0, 2.0 ...). This is intended to make bar plot be located on the same coodinates as line plot. However, bar plot may differs unexpectedly when you manually adjust the bar location or drawing area, such as using `set_xlim`, `set_ylim`, etc. In this cases, please modify your script to meet with new coordinates.
+ Because of the default `align` value changes, coordinates of bar plots are now located on integer values (0.0, 1.0, 2.0 ...). This is intended to make bar plot be located on the same coordinates as line plot. However, bar plot may differs unexpectedly when you manually adjust the bar location or drawing area, such as using `set_xlim`, `set_ylim`, etc. In this cases, please modify your script to meet with new coordinates.
- The :func:`parallel_coordinates` function now takes argument ``color``
- instead of ``colors``. A ``FutureWarning`` is raised to alert that
+ instead of ``colors``. A ``FutureWarning`` is raised to alert that
the old ``colors`` argument will not be supported in a future release. (:issue:`6956`)
- The :func:`parallel_coordinates` and :func:`andrews_curves` functions now take
positional argument ``frame`` instead of ``data``. A ``FutureWarning`` is
- raised if the old ``data`` argument is used by name. (:issue:`6956`)
+ raised if the old ``data`` argument is used by name. (:issue:`6956`)
- :meth:`DataFrame.boxplot` now supports ``layout`` keyword (:issue:`6769`)
- :meth:`DataFrame.boxplot` has a new keyword argument, `return_type`. It accepts ``'dict'``,
@@ -645,17 +645,17 @@ Deprecations
- The :func:`pivot_table`/:meth:`DataFrame.pivot_table` and :func:`crosstab` functions
now take arguments ``index`` and ``columns`` instead of ``rows`` and ``cols``. A
- ``FutureWarning`` is raised to alert that the old ``rows`` and ``cols`` arguments
+ ``FutureWarning`` is raised to alert that the old ``rows`` and ``cols`` arguments
will not be supported in a future release (:issue:`5505`)
- The :meth:`DataFrame.drop_duplicates` and :meth:`DataFrame.duplicated` methods
now take argument ``subset`` instead of ``cols`` to better align with
- :meth:`DataFrame.dropna`. A ``FutureWarning`` is raised to alert that the old
+ :meth:`DataFrame.dropna`. A ``FutureWarning`` is raised to alert that the old
``cols`` arguments will not be supported in a future release (:issue:`6680`)
- The :meth:`DataFrame.to_csv` and :meth:`DataFrame.to_excel` functions
now takes argument ``columns`` instead of ``cols``. A
- ``FutureWarning`` is raised to alert that the old ``cols`` arguments
+ ``FutureWarning`` is raised to alert that the old ``cols`` arguments
will not be supported in a future release (:issue:`6645`)
- Indexers will warn ``FutureWarning`` when used with a scalar indexer and
@@ -698,12 +698,12 @@ Deprecations
ALWAYS return a view. (:issue:`6894`)
- The :func:`parallel_coordinates` function now takes argument ``color``
- instead of ``colors``. A ``FutureWarning`` is raised to alert that
+ instead of ``colors``. A ``FutureWarning`` is raised to alert that
the old ``colors`` argument will not be supported in a future release. (:issue:`6956`)
- The :func:`parallel_coordinates` and :func:`andrews_curves` functions now take
positional argument ``frame`` instead of ``data``. A ``FutureWarning`` is
- raised if the old ``data`` argument is used by name. (:issue:`6956`)
+ raised if the old ``data`` argument is used by name. (:issue:`6956`)
- The support for the 'mysql' flavor when using DBAPI connection objects has been deprecated.
MySQL will be further supported with SQLAlchemy engines (:issue:`6900`).
@@ -899,7 +899,7 @@ Bug Fixes
- Raise when trying to align on different levels of a multi-index assignment (:issue:`3738`)
- Bug in setting complex dtypes via boolean indexing (:issue:`6345`)
- Bug in TimeGrouper/resample when presented with a non-monotonic DatetimeIndex that would return invalid results. (:issue:`4161`)
-- Bug in index name propogation in TimeGrouper/resample (:issue:`4161`)
+- Bug in index name propagation in TimeGrouper/resample (:issue:`4161`)
- TimeGrouper has a more compatible API to the rest of the groupers (e.g. ``groups`` was missing) (:issue:`3881`)
- Bug in multiple grouping with a TimeGrouper depending on target column order (:issue:`6764`)
- Bug in ``pd.eval`` when parsing strings with possible tokens like ``'&'``
@@ -976,7 +976,7 @@ Bug Fixes
clean`` (:issue:`6768`)
- Bug with numpy < 1.7.2 when reading long strings from ``HDFStore`` (:issue:`6166`)
- Bug in ``DataFrame._reduce`` where non bool-like (0/1) integers were being
- coverted into bools. (:issue:`6806`)
+ converted into bools. (:issue:`6806`)
- Regression from 0.13 with ``fillna`` and a Series on datetime-like (:issue:`6344`)
- Bug in adding ``np.timedelta64`` to ``DatetimeIndex`` with timezone outputs incorrect results (:issue:`6818`)
- Bug in ``DataFrame.replace()`` where changing a dtype through replacement
diff --git a/doc/source/whatsnew/v0.14.1.txt b/doc/source/whatsnew/v0.14.1.txt
index 239d6c9c6e0d4..d8a6dc1793612 100644
--- a/doc/source/whatsnew/v0.14.1.txt
+++ b/doc/source/whatsnew/v0.14.1.txt
@@ -75,7 +75,7 @@ API changes
Note that for the other offsets the default behaviour did not change.
-- Add back ``#N/A N/A`` as a default NA value in text parsing, (regresion from 0.12) (:issue:`5521`)
+- Add back ``#N/A N/A`` as a default NA value in text parsing, (regression from 0.12) (:issue:`5521`)
- Raise a ``TypeError`` on inplace-setting with a ``.where`` and a non ``np.nan`` value as this is inconsistent
with a set-item expression like ``df[mask] = None`` (:issue:`7656`)
@@ -88,7 +88,7 @@ Enhancements
- Add ``dropna`` argument to ``value_counts`` and ``nunique`` (:issue:`5569`).
- Add :meth:`~pandas.DataFrame.select_dtypes` method to allow selection of
columns based on dtype (:issue:`7316`). See :ref:`the docs <basics.selectdtypes>`.
-- All ``offsets`` suppports the ``normalize`` keyword to specify whether
+- All ``offsets`` supports the ``normalize`` keyword to specify whether
``offsets.apply``, ``rollforward`` and ``rollback`` resets the time (hour,
minute, etc) or not (default ``False``, preserves time) (:issue:`7156`):
diff --git a/doc/source/whatsnew/v0.15.0.txt b/doc/source/whatsnew/v0.15.0.txt
index e44bc6e9e91e0..ef17904d5ab1a 100644
--- a/doc/source/whatsnew/v0.15.0.txt
+++ b/doc/source/whatsnew/v0.15.0.txt
@@ -22,7 +22,7 @@ users upgrade to this version.
- ``read_csv`` will now by default ignore blank lines when parsing, see :ref:`here <whatsnew_0150.blanklines>`
- API change in using Indexes in set operations, see :ref:`here <whatsnew_0150.index_set_ops>`
- Enhancements in the handling of timezones, see :ref:`here <whatsnew_0150.tz>`
- - A lot of improvements to the rolling and expanding moment funtions, see :ref:`here <whatsnew_0150.roll>`
+ - A lot of improvements to the rolling and expanding moment functions, see :ref:`here <whatsnew_0150.roll>`
- Internal refactoring of the ``Index`` class to no longer sub-class ``ndarray``, see :ref:`Internal Refactoring <whatsnew_0150.refactoring>`
- dropping support for ``PyTables`` less than version 3.0.0, and ``numexpr`` less than version 2.1 (:issue:`7990`)
- Split indexing documentation into :ref:`Indexing and Selecting Data <indexing>` and :ref:`MultiIndex / Advanced Indexing <advanced>`
@@ -326,7 +326,7 @@ Timezone handling improvements
- ``Timestamp.tz_localize`` and ``Timestamp.tz_convert`` now raise ``TypeError`` in error cases, rather than ``Exception`` (:issue:`8025`)
-- a timeseries/index localized to UTC when inserted into a Series/DataFrame will preserve the UTC timezone (rather than being a naive ``datetime64[ns]``) as ``object`` dtype (:issue:`8411`)
+- a timeseries/index localized to UTC when inserted into a Series/DataFrame will preserve the UTC timezone (rather than being a naive ``datetime64[ns]``) as ``object`` dtype (:issue:`8411`)
- ``Timestamp.__repr__`` displays ``dateutil.tz.tzoffset`` info (:issue:`7907`)
@@ -837,7 +837,7 @@ Other notable API changes:
A value is trying to be set on a copy of a slice from a DataFrame.
Try using .loc[row_indexer,col_indexer] = value instead
- See the the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/indexing.html#indexing-view-versus-copy
+ See the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/indexing.html#indexing-view-versus-copy
- ``merge``, ``DataFrame.merge``, and ``ordered_merge`` now return the same type
as the ``left`` argument (:issue:`7737`).
@@ -878,7 +878,7 @@ a transparent change with only very limited API implications (:issue:`5080`, :is
- you may need to unpickle pandas version < 0.15.0 pickles using ``pd.read_pickle`` rather than ``pickle.load``. See :ref:`pickle docs <io.pickle>`
- when plotting with a ``PeriodIndex``, the matplotlib internal axes will now be arrays of ``Period`` rather than a ``PeriodIndex`` (this is similar to how a ``DatetimeIndex`` passes arrays of ``datetimes`` now)
-- MultiIndexes will now raise similary to other pandas objects w.r.t. truth testing, see :ref:`here <gotchas.truth>` (:issue:`7897`).
+- MultiIndexes will now raise similarly to other pandas objects w.r.t. truth testing, see :ref:`here <gotchas.truth>` (:issue:`7897`).
- When plotting a DatetimeIndex directly with matplotlib's `plot` function,
the axis labels will no longer be formatted as dates but as integers (the
internal representation of a ``datetime64``). **UPDATE** This is fixed
@@ -1118,7 +1118,7 @@ Bug Fixes
- Bug in multi-index slicing with various edge cases (:issue:`8132`)
- Regression in multi-index indexing with a non-scalar type object (:issue:`7914`)
- Bug in ``Timestamp`` comparisons with ``==`` and ``int64`` dtype (:issue:`8058`)
-- Bug in pickles contains ``DateOffset`` may raise ``AttributeError`` when ``normalize`` attribute is reffered internally (:issue:`7748`)
+- Bug in pickles contains ``DateOffset`` may raise ``AttributeError`` when ``normalize`` attribute is referred internally (:issue:`7748`)
- Bug in ``Panel`` when using ``major_xs`` and ``copy=False`` is passed (deprecation warning fails because of missing ``warnings``) (:issue:`8152`).
- Bug in pickle deserialization that failed for pre-0.14.1 containers with dup items trying to avoid ambiguity
when matching block and manager items, when there's only one block there's no ambiguity (:issue:`7794`)
diff --git a/doc/source/whatsnew/v0.15.1.txt b/doc/source/whatsnew/v0.15.1.txt
index cd9298c74539a..f84f25d3e906c 100644
--- a/doc/source/whatsnew/v0.15.1.txt
+++ b/doc/source/whatsnew/v0.15.1.txt
@@ -274,7 +274,7 @@ Enhancements
Bug Fixes
~~~~~~~~~
-- Bug in unpickling of a ``CustomBusinessDay`` object (:issue:`8591`)
+- Bug in unpickling of a ``CustomBusinessDay`` object (:issue:`8591`)
- Bug in coercing ``Categorical`` to a records array, e.g. ``df.to_records()`` (:issue:`8626`)
- Bug in ``Categorical`` not created properly with ``Series.to_frame()`` (:issue:`8626`)
- Bug in coercing in astype of a ``Categorical`` of a passed ``pd.Categorical`` (this now raises ``TypeError`` correctly), (:issue:`8626`)
diff --git a/doc/source/whatsnew/v0.15.2.txt b/doc/source/whatsnew/v0.15.2.txt
index b908b60334f4c..f1dfab0f57ed3 100644
--- a/doc/source/whatsnew/v0.15.2.txt
+++ b/doc/source/whatsnew/v0.15.2.txt
@@ -215,7 +215,7 @@ Bug Fixes
- ``io.data.Options`` now raises ``RemoteDataError`` when no expiry dates are available from Yahoo and when it receives no data from Yahoo (:issue:`8761`), (:issue:`8783`).
- Fix: The font size was only set on x axis if vertical or the y axis if horizontal. (:issue:`8765`)
- Fixed division by 0 when reading big csv files in python 3 (:issue:`8621`)
-- Bug in outputing a Multindex with ``to_html,index=False`` which would add an extra column (:issue:`8452`)
+- Bug in outputting a Multindex with ``to_html,index=False`` which would add an extra column (:issue:`8452`)
- Imported categorical variables from Stata files retain the ordinal information in the underlying data (:issue:`8836`).
- Defined ``.size`` attribute across ``NDFrame`` objects to provide compat with numpy >= 1.9.1; buggy with ``np.array_split`` (:issue:`8846`)
- Skip testing of histogram plots for matplotlib <= 1.2 (:issue:`8648`).
diff --git a/doc/source/whatsnew/v0.16.0.txt b/doc/source/whatsnew/v0.16.0.txt
index 8238cc32d7bb0..48af06d124f2e 100644
--- a/doc/source/whatsnew/v0.16.0.txt
+++ b/doc/source/whatsnew/v0.16.0.txt
@@ -56,7 +56,7 @@ and the entire DataFrame (with all original and new columns) is returned.
iris.assign(sepal_ratio=iris['SepalWidth'] / iris['SepalLength']).head()
Above was an example of inserting a precomputed value. We can also pass in
-a function to be evalutated.
+a function to be evaluated.
.. ipython :: python
@@ -595,7 +595,7 @@ Bug Fixes
- Bug in ``unstack`` with ``TimedeltaIndex`` or ``DatetimeIndex`` and nulls (:issue:`9491`).
- Bug in ``rank`` where comparing floats with tolerance will cause inconsistent behaviour (:issue:`8365`).
- Fixed character encoding bug in ``read_stata`` and ``StataReader`` when loading data from a URL (:issue:`9231`).
-- Bug in adding ``offsets.Nano`` to other offets raises ``TypeError`` (:issue:`9284`)
+- Bug in adding ``offsets.Nano`` to other offsets raises ``TypeError`` (:issue:`9284`)
- Bug in ``DatetimeIndex`` iteration, related to (:issue:`8890`), fixed in (:issue:`9100`)
- Bugs in ``resample`` around DST transitions. This required fixing offset classes so they behave correctly on DST transitions. (:issue:`5172`, :issue:`8744`, :issue:`8653`, :issue:`9173`, :issue:`9468`).
- Bug in binary operator method (eg ``.mul()``) alignment with integer levels (:issue:`9463`).
@@ -611,7 +611,7 @@ Bug Fixes
- Accessing ``Series.str`` methods on with non-string values now raises ``TypeError`` instead of producing incorrect results (:issue:`9184`)
- Bug in ``DatetimeIndex.__contains__`` when index has duplicates and is not monotonic increasing (:issue:`9512`)
- Fixed division by zero error for ``Series.kurt()`` when all values are equal (:issue:`9197`)
-- Fixed issue in the ``xlsxwriter`` engine where it added a default 'General' format to cells if no other format wass applied. This prevented other row or column formatting being applied. (:issue:`9167`)
+- Fixed issue in the ``xlsxwriter`` engine where it added a default 'General' format to cells if no other format was applied. This prevented other row or column formatting being applied. (:issue:`9167`)
- Fixes issue with ``index_col=False`` when ``usecols`` is also specified in ``read_csv``. (:issue:`9082`)
- Bug where ``wide_to_long`` would modify the input stubnames list (:issue:`9204`)
- Bug in ``to_sql`` not storing float64 values using double precision. (:issue:`9009`)
diff --git a/doc/source/whatsnew/v0.17.0.txt b/doc/source/whatsnew/v0.17.0.txt
index a3bbaf73c01ca..239b2ba96404c 100644
--- a/doc/source/whatsnew/v0.17.0.txt
+++ b/doc/source/whatsnew/v0.17.0.txt
@@ -1157,7 +1157,7 @@ Bug Fixes
- Bug in ``.var()`` causing roundoff errors for highly similar values (:issue:`10242`)
- Bug in ``DataFrame.plot(subplots=True)`` with duplicated columns outputs incorrect result (:issue:`10962`)
- Bug in ``Index`` arithmetic may result in incorrect class (:issue:`10638`)
-- Bug in ``date_range`` results in empty if freq is negative annualy, quarterly and monthly (:issue:`11018`)
+- Bug in ``date_range`` results in empty if freq is negative annually, quarterly and monthly (:issue:`11018`)
- Bug in ``DatetimeIndex`` cannot infer negative freq (:issue:`11018`)
- Remove use of some deprecated numpy comparison operations, mainly in tests. (:issue:`10569`)
- Bug in ``Index`` dtype may not applied properly (:issue:`11017`)
diff --git a/doc/source/whatsnew/v0.17.1.txt b/doc/source/whatsnew/v0.17.1.txt
index 1ad7279ea79f7..d5ed0503d9ee3 100644
--- a/doc/source/whatsnew/v0.17.1.txt
+++ b/doc/source/whatsnew/v0.17.1.txt
@@ -157,11 +157,11 @@ Bug Fixes
- ``Series.sort_index()`` now correctly handles the ``inplace`` option (:issue:`11402`)
- Incorrectly distributed .c file in the build on ``PyPi`` when reading a csv of floats and passing ``na_values=<a scalar>`` would show an exception (:issue:`11374`)
- Bug in ``.to_latex()`` output broken when the index has a name (:issue:`10660`)
-- Bug in ``HDFStore.append`` with strings whose encoded length exceded the max unencoded length (:issue:`11234`)
+- Bug in ``HDFStore.append`` with strings whose encoded length exceeded the max unencoded length (:issue:`11234`)
- Bug in merging ``datetime64[ns, tz]`` dtypes (:issue:`11405`)
- Bug in ``HDFStore.select`` when comparing with a numpy scalar in a where clause (:issue:`11283`)
- Bug in using ``DataFrame.ix`` with a multi-index indexer (:issue:`11372`)
-- Bug in ``date_range`` with ambigous endpoints (:issue:`11626`)
+- Bug in ``date_range`` with ambiguous endpoints (:issue:`11626`)
- Prevent adding new attributes to the accessors ``.str``, ``.dt`` and ``.cat``. Retrieving such
a value was not possible, so error out on setting it. (:issue:`10673`)
- Bug in tz-conversions with an ambiguous time and ``.dt`` accessors (:issue:`11295`)
diff --git a/doc/source/whatsnew/v0.18.0.txt b/doc/source/whatsnew/v0.18.0.txt
index 4b27cf706f9b2..bfd314639aa60 100644
--- a/doc/source/whatsnew/v0.18.0.txt
+++ b/doc/source/whatsnew/v0.18.0.txt
@@ -217,7 +217,7 @@ It returns a ``DataFrame`` with one column if ``expand=True``.
pd.Series(['a1', 'b2', 'c3']).str.extract('[ab](\d)', expand=True)
Calling on an ``Index`` with a regex with exactly one capture group
-returns an ``Index`` if ``expand=False``.
+returns an ``Index`` if ``expand=False``.
.. ipython:: python
@@ -944,7 +944,7 @@ assignment should be done inplace or return a copy.
.. warning::
- For backwards compatability, ``inplace`` defaults to ``True`` if not specified.
+ For backwards compatibility, ``inplace`` defaults to ``True`` if not specified.
This will change in a future version of pandas. If your code depends on an
inplace assignment you should update to explicitly set ``inplace=True``
@@ -1039,7 +1039,7 @@ Deprecations
2 0.5
dtype: float64
-- The the ``freq`` and ``how`` arguments to the ``.rolling``, ``.expanding``, and ``.ewm`` (new) functions are deprecated, and will be removed in a future version. You can simply resample the input prior to creating a window function. (:issue:`11603`).
+- The ``freq`` and ``how`` arguments to the ``.rolling``, ``.expanding``, and ``.ewm`` (new) functions are deprecated, and will be removed in a future version. You can simply resample the input prior to creating a window function. (:issue:`11603`).
For example, instead of ``s.rolling(window=5,freq='D').max()`` to get the max value on a rolling 5 Day window, one could use ``s.resample('D').mean().rolling(window=5).max()``, which first resamples the data to daily data, then provides a rolling 5 day window.
diff --git a/doc/source/whatsnew/v0.18.1.txt b/doc/source/whatsnew/v0.18.1.txt
index ca386da03295d..de9a5d5d8afae 100644
--- a/doc/source/whatsnew/v0.18.1.txt
+++ b/doc/source/whatsnew/v0.18.1.txt
@@ -226,7 +226,7 @@ Other Enhancements
^^^^^^^^^^^^^^^^^^
- ``pd.read_csv()`` now supports ``delim_whitespace=True`` for the Python engine (:issue:`12958`)
-- ``pd.read_csv()`` now supports opening ZIP files that contains a single CSV, via extension inference or explict ``compression='zip'`` (:issue:`12175`)
+- ``pd.read_csv()`` now supports opening ZIP files that contains a single CSV, via extension inference or explicit ``compression='zip'`` (:issue:`12175`)
- ``pd.read_csv()`` now supports opening files using xz compression, via extension inference or explicit ``compression='xz'`` is specified; ``xz`` compressions is also supported by ``DataFrame.to_csv`` in the same way (:issue:`11852`)
- ``pd.read_msgpack()`` now always gives writeable ndarrays even when compression is used (:issue:`12359`).
- ``pd.read_msgpack()`` now supports serializing and de-serializing categoricals with msgpack (:issue:`12573`)
diff --git a/doc/source/whatsnew/v0.19.0.txt b/doc/source/whatsnew/v0.19.0.txt
index 6093e53029cb6..302105c1e653c 100644
--- a/doc/source/whatsnew/v0.19.0.txt
+++ b/doc/source/whatsnew/v0.19.0.txt
@@ -1413,7 +1413,7 @@ Performance Improvements
- Improved performance of ``factorize`` of datetime with timezone (:issue:`13750`)
- Improved performance of by lazily creating indexing hashtables on larger Indexes (:issue:`14266`)
- Improved performance of ``groupby.groups`` (:issue:`14293`)
-- Unecessary materializing of a MultiIndex when introspecting for memory usage (:issue:`14308`)
+- Unnecessary materializing of a MultiIndex when introspecting for memory usage (:issue:`14308`)
.. _whatsnew_0190.bug_fixes:
@@ -1454,7 +1454,7 @@ Bug Fixes
- Bug in ``.tz_localize`` with ``dateutil.tz.tzlocal`` may return incorrect result (:issue:`13583`)
- Bug in ``DatetimeTZDtype`` dtype with ``dateutil.tz.tzlocal`` cannot be regarded as valid dtype (:issue:`13583`)
- Bug in ``pd.read_hdf()`` where attempting to load an HDF file with a single dataset, that had one or more categorical columns, failed unless the key argument was set to the name of the dataset. (:issue:`13231`)
-- Bug in ``.rolling()`` that allowed a negative integer window in contruction of the ``Rolling()`` object, but would later fail on aggregation (:issue:`13383`)
+- Bug in ``.rolling()`` that allowed a negative integer window in construction of the ``Rolling()`` object, but would later fail on aggregation (:issue:`13383`)
- Bug in ``Series`` indexing with tuple-valued data and a numeric index (:issue:`13509`)
- Bug in printing ``pd.DataFrame`` where unusual elements with the ``object`` dtype were causing segfaults (:issue:`13717`)
- Bug in ranking ``Series`` which could result in segfaults (:issue:`13445`)
diff --git a/doc/source/whatsnew/v0.19.2.txt b/doc/source/whatsnew/v0.19.2.txt
index 722e494c9e614..171d97b76de75 100644
--- a/doc/source/whatsnew/v0.19.2.txt
+++ b/doc/source/whatsnew/v0.19.2.txt
@@ -26,7 +26,7 @@ Enhancements
The ``pd.merge_asof()``, added in 0.19.0, gained some improvements:
- ``pd.merge_asof()`` gained ``left_index``/``right_index`` and ``left_by``/``right_by`` arguments (:issue:`14253`)
-- ``pd.merge_asof()`` can take multiple columns in ``by`` parameter and has specialized dtypes for better performace (:issue:`13936`)
+- ``pd.merge_asof()`` can take multiple columns in ``by`` parameter and has specialized dtypes for better performance (:issue:`13936`)
.. _whatsnew_0192.performance:
@@ -62,7 +62,7 @@ Bug Fixes
- Bug in ``pd.to_numeric`` where a 0 was not unsigned on a ``downcast='unsigned'`` argument (:issue:`14401`)
- Bug in plotting regular and irregular timeseries using shared axes
(``sharex=True`` or ``ax.twinx()``) (:issue:`13341`, :issue:`14322`).
-- Bug in not propogating exceptions in parsing invalid datetimes, noted in python 3.6 (:issue:`14561`)
+- Bug in not propagating exceptions in parsing invalid datetimes, noted in python 3.6 (:issue:`14561`)
- Bug in resampling a ``DatetimeIndex`` in local TZ, covering a DST change, which would raise ``AmbiguousTimeError`` (:issue:`14682`)
- Bug in indexing that transformed ``RecursionError`` into ``KeyError`` or ``IndexingError`` (:issue:`14554`)
- Bug in ``HDFStore`` when writing a ``MultiIndex`` when using ``data_columns=True`` (:issue:`14435`)
diff --git a/doc/source/whatsnew/v0.20.0.txt b/doc/source/whatsnew/v0.20.0.txt
index fc869956c820e..d04a34f7a44d6 100644
--- a/doc/source/whatsnew/v0.20.0.txt
+++ b/doc/source/whatsnew/v0.20.0.txt
@@ -97,7 +97,7 @@ The API also supports a ``.transform()`` function for broadcasting results.
df.transform(['abs', lambda x: x - x.min()])
When presented with mixed dtypes that cannot be aggregated, ``.agg()`` will only take the valid
-aggregations. This is similiar to how groupby ``.agg()`` works. (:issue:`15015`)
+aggregations. This is similar to how groupby ``.agg()`` works. (:issue:`15015`)
.. ipython:: python
@@ -1504,7 +1504,7 @@ Other Deprecations
- ``TimedeltaIndex.searchsorted()``, ``DatetimeIndex.searchsorted()``, and ``PeriodIndex.searchsorted()`` have deprecated the ``key`` parameter in favor of ``value`` (:issue:`12662`)
- ``DataFrame.astype()`` has deprecated the ``raise_on_error`` parameter in favor of ``errors`` (:issue:`14878`)
- ``Series.sortlevel`` and ``DataFrame.sortlevel`` have been deprecated in favor of ``Series.sort_index`` and ``DataFrame.sort_index`` (:issue:`15099`)
-- importing ``concat`` from ``pandas.tools.merge`` has been deprecated in favor of imports from the ``pandas`` namespace. This should only affect explict imports (:issue:`15358`)
+- importing ``concat`` from ``pandas.tools.merge`` has been deprecated in favor of imports from the ``pandas`` namespace. This should only affect explicit imports (:issue:`15358`)
- ``Series/DataFrame/Panel.consolidate()`` been deprecated as a public method. (:issue:`15483`)
- The ``as_indexer`` keyword of ``Series.str.match()`` has been deprecated (ignored keyword) (:issue:`15257`).
- The following top-level pandas functions have been deprecated and will be removed in a future version (:issue:`13790`, :issue:`15940`)
diff --git a/doc/source/whatsnew/v0.21.1.txt b/doc/source/whatsnew/v0.21.1.txt
index a7dde5d6ee410..67c52dac6128d 100644
--- a/doc/source/whatsnew/v0.21.1.txt
+++ b/doc/source/whatsnew/v0.21.1.txt
@@ -168,4 +168,4 @@ Categorical
String
^^^^^^
-- :meth:`Series.str.split()` will now propogate ``NaN`` values across all expanded columns instead of ``None`` (:issue:`18450`)
+- :meth:`Series.str.split()` will now propagate ``NaN`` values across all expanded columns instead of ``None`` (:issue:`18450`)
diff --git a/doc/source/whatsnew/v0.23.0.txt b/doc/source/whatsnew/v0.23.0.txt
index 0f6660d2f4125..d6766afdf9d4a 100644
--- a/doc/source/whatsnew/v0.23.0.txt
+++ b/doc/source/whatsnew/v0.23.0.txt
@@ -127,7 +127,7 @@ Other Enhancements
- Better support for :func:`Dataframe.style.to_excel` output with the ``xlsxwriter`` engine. (:issue:`16149`)
- :func:`pandas.tseries.frequencies.to_offset` now accepts leading '+' signs e.g. '+1h'. (:issue:`18171`)
- :func:`MultiIndex.unique` now supports the ``level=`` argument, to get unique values from a specific index level (:issue:`17896`)
-- :class:`pandas.io.formats.style.Styler` now has method ``hide_index()`` to determine whether the index will be rendered in ouptut (:issue:`14194`)
+- :class:`pandas.io.formats.style.Styler` now has method ``hide_index()`` to determine whether the index will be rendered in output (:issue:`14194`)
- :class:`pandas.io.formats.style.Styler` now has method ``hide_columns()`` to determine whether columns will be hidden in output (:issue:`14194`)
- Improved wording of ``ValueError`` raised in :func:`to_datetime` when ``unit=`` is passed with a non-convertible value (:issue:`14350`)
- :func:`Series.fillna` now accepts a Series or a dict as a ``value`` for a categorical dtype (:issue:`17033`)
@@ -172,7 +172,7 @@ Build Changes
^^^^^^^^^^^^^
- Building pandas for development now requires ``cython >= 0.24`` (:issue:`18613`)
-- Building from source now explicity requires ``setuptools`` in ``setup.py`` (:issue:`18113`)
+- Building from source now explicitly requires ``setuptools`` in ``setup.py`` (:issue:`18113`)
- Updated conda recipe to be in compliance with conda-build 3.0+ (:issue:`18002`)
.. _whatsnew_0230.api:
@@ -231,7 +231,7 @@ Removal of prior version deprecations/changes
- Warnings against the obsolete usage ``Categorical(codes, categories)``, which were emitted for instance when the first two arguments to ``Categorical()`` had different dtypes, and recommended the use of ``Categorical.from_codes``, have now been removed (:issue:`8074`)
- The ``levels`` and ``labels`` attributes of a ``MultiIndex`` can no longer be set directly (:issue:`4039`).
-- ``pd.tseries.util.pivot_annual`` has been removed (deprecated since v0.19). Use ``pivot_table`` instead (:issue:`18370`)
+- ``pd.tseries.util.pivot_annual`` has been removed (deprecated since v0.19). Use ``pivot_table`` instead (:issue:`18370`)
- ``pd.tseries.util.isleapyear`` has been removed (deprecated since v0.19). Use ``.is_leap_year`` property in Datetime-likes instead (:issue:`18370`)
- ``pd.ordered_merge`` has been removed (deprecated since v0.19). Use ``pd.merge_ordered`` instead (:issue:`18459`)
- The ``SparseList`` class has been removed (:issue:`14007`)
@@ -257,7 +257,7 @@ Performance Improvements
- :class`DateOffset` arithmetic performance is improved (:issue:`18218`)
- Converting a ``Series`` of ``Timedelta`` objects to days, seconds, etc... sped up through vectorization of underlying methods (:issue:`18092`)
- Improved performance of ``.map()`` with a ``Series/dict`` input (:issue:`15081`)
-- The overriden ``Timedelta`` properties of days, seconds and microseconds have been removed, leveraging their built-in Python versions instead (:issue:`18242`)
+- The overridden ``Timedelta`` properties of days, seconds and microseconds have been removed, leveraging their built-in Python versions instead (:issue:`18242`)
- ``Series`` construction will reduce the number of copies made of the input data in certain cases (:issue:`17449`)
- Improved performance of :func:`Series.dt.date` and :func:`DatetimeIndex.date` (:issue:`18058`)
- Improved performance of :func:`Series.dt.time` and :func:`DatetimeIndex.time` (:issue:`18461`)
diff --git a/doc/source/whatsnew/v0.8.1.txt b/doc/source/whatsnew/v0.8.1.txt
index 8227bc6bc9c66..add96bec9d1dd 100644
--- a/doc/source/whatsnew/v0.8.1.txt
+++ b/doc/source/whatsnew/v0.8.1.txt
@@ -32,5 +32,5 @@ Performance improvements
strings with ``DatetimeIndex`` or ``to_datetime`` (:issue:`1571`)
- Improve the performance of GroupBy on single-key aggregations and use with
Categorical types
- - Significant datetime parsing performance improvments
+ - Significant datetime parsing performance improvements
diff --git a/doc/source/whatsnew/v0.9.1.txt b/doc/source/whatsnew/v0.9.1.txt
index 4faf38219ebee..e2d6d7a275086 100644
--- a/doc/source/whatsnew/v0.9.1.txt
+++ b/doc/source/whatsnew/v0.9.1.txt
@@ -80,7 +80,7 @@ New features
df.where(df>0,-df)
Furthermore, `where` now aligns the input boolean condition (ndarray or DataFrame), such that partial selection
- with setting is possible. This is analagous to partial setting via `.ix` (but on the contents rather than the axis labels)
+ with setting is possible. This is analogous to partial setting via `.ix` (but on the contents rather than the axis labels)
.. ipython:: python
diff --git a/doc/sphinxext/README.rst b/doc/sphinxext/README.rst
index e39cf8daac036..2be5372bc0216 100644
--- a/doc/sphinxext/README.rst
+++ b/doc/sphinxext/README.rst
@@ -14,4 +14,4 @@ pandas documentation. These copies originate from other projects:
These copies are maintained at the respective projects, so fixes should,
to the extent possible, be pushed upstream instead of only adapting our
- local copy to avoid divergence between the the local and upstream version.
+ local copy to avoid divergence between the local and upstream version.
diff --git a/doc/sphinxext/ipython_sphinxext/ipython_console_highlighting.py b/doc/sphinxext/ipython_sphinxext/ipython_console_highlighting.py
index dfb489e49394d..c5ec26aefd442 100644
--- a/doc/sphinxext/ipython_sphinxext/ipython_console_highlighting.py
+++ b/doc/sphinxext/ipython_sphinxext/ipython_console_highlighting.py
@@ -83,7 +83,7 @@ def get_tokens_unprocessed(self, text):
curcode += line[continue_prompt.end():]
elif output_prompt is not None:
# Use the 'error' token for output. We should probably make
- # our own token, but error is typicaly in a bright color like
+ # our own token, but error is typically in a bright color like
# red, so it works fine for our output prompts.
insertions.append((len(curcode),
[(0, Generic.Error, output_prompt.group())]))
diff --git a/doc/sphinxext/ipython_sphinxext/ipython_directive.py b/doc/sphinxext/ipython_sphinxext/ipython_directive.py
index 4f7b32840680d..5616d732eb1c6 100644
--- a/doc/sphinxext/ipython_sphinxext/ipython_directive.py
+++ b/doc/sphinxext/ipython_sphinxext/ipython_directive.py
@@ -93,7 +93,7 @@
Authors
-------
-- John D Hunter: orignal author.
+- John D Hunter: original author.
- Fernando Perez: refactoring, documentation, cleanups, port to 0.11.
- VáclavŠmilauer <eudoxos-AT-arcig.cz>: Prompt generalizations.
- Skipper Seabold, refactoring, cleanups, pure python addition
@@ -154,7 +154,7 @@
def block_parser(part, rgxin, rgxout, fmtin, fmtout):
"""
part is a string of ipython text, comprised of at most one
- input, one ouput, comments, and blank lines. The block parser
+ input, one output, comments, and blank lines. The block parser
parses the text into a list of::
blocks = [ (TOKEN0, data0), (TOKEN1, data1), ...]
@@ -268,7 +268,7 @@ def write(self,data):
return super(DecodingStringIO, self).write(data)
except :
pass
- # default to brute utf8 if no encoding succeded
+ # default to brute utf8 if no encoding succeeded
return super(DecodingStringIO, self).write(data.decode('utf8', 'replace'))
diff --git a/doc/sphinxext/numpydoc/docscrape_sphinx.py b/doc/sphinxext/numpydoc/docscrape_sphinx.py
index 9017480c9ab76..127ed49c106ad 100755
--- a/doc/sphinxext/numpydoc/docscrape_sphinx.py
+++ b/doc/sphinxext/numpydoc/docscrape_sphinx.py
@@ -115,7 +115,7 @@ def _str_member_list(self, name):
or inspect.isgetsetdescriptor(param_obj)):
param_obj = None
- # pandas HACK - do not exclude attributes wich are None
+ # pandas HACK - do not exclude attributes which are None
# if param_obj and (pydoc.getdoc(param_obj) or not desc):
# # Referenced object has a docstring
# autosum += [" %s%s" % (prefix, param)]
diff --git a/pandas/__init__.py b/pandas/__init__.py
index 861c8e7d622fc..93c5b6484b840 100644
--- a/pandas/__init__.py
+++ b/pandas/__init__.py
@@ -106,8 +106,8 @@
- Easy handling of missing data in floating point as well as non-floating
point data.
- Size mutability: columns can be inserted and deleted from DataFrame and
- higher dimensional objects.
- - Automatic and explicit data alignment: objects can be explicitly aligned
+ higher dimensional objects
+ - Automatic and explicit data alignment: objects can be explicitly aligned
to a set of labels, or the user can simply ignore the labels and let
`Series`, `DataFrame`, etc. automatically align the data for you in
computations.
diff --git a/pandas/_libs/algos.pyx b/pandas/_libs/algos.pyx
index 3710ddc33c7c5..7b61cd22f45d1 100644
--- a/pandas/_libs/algos.pyx
+++ b/pandas/_libs/algos.pyx
@@ -61,7 +61,7 @@ cdef inline are_diff(object left, object right):
class Infinity(object):
- """ provide a positive Infinity comparision method for ranking """
+ """ provide a positive Infinity comparison method for ranking """
__lt__ = lambda self, other: False
__le__ = lambda self, other: isinstance(other, Infinity)
@@ -73,7 +73,7 @@ class Infinity(object):
class NegInfinity(object):
- """ provide a negative Infinity comparision method for ranking """
+ """ provide a negative Infinity comparison method for ranking """
__lt__ = lambda self, other: (not isinstance(other, NegInfinity) and
not missing.checknull(other))
diff --git a/pandas/_libs/hashing.pyx b/pandas/_libs/hashing.pyx
index aa7aa4b528194..c6f182ac5003f 100644
--- a/pandas/_libs/hashing.pyx
+++ b/pandas/_libs/hashing.pyx
@@ -80,7 +80,7 @@ def hash_object_array(ndarray[object] arr, object key, object encoding='utf8'):
lens[i] = l
cdata = data
- # keep the refernce alive thru the end of the
+ # keep the references alive thru the end of the
# function
datas.append(data)
vecs[i] = cdata
diff --git a/pandas/_libs/hashtable.pyx b/pandas/_libs/hashtable.pyx
index 4bbe8c654ea0e..72c2834b0bd57 100644
--- a/pandas/_libs/hashtable.pyx
+++ b/pandas/_libs/hashtable.pyx
@@ -148,7 +148,7 @@ cdef class Int64Factorizer:
def unique_label_indices(ndarray[int64_t, ndim=1] labels):
"""
indices of the first occurrences of the unique labels
- *excluding* -1. equivelent to:
+ *excluding* -1. equivalent to:
np.unique(labels, return_index=True)[1]
"""
cdef:
diff --git a/pandas/_libs/lib.pyx b/pandas/_libs/lib.pyx
index 3898f7499e85e..788d3c4ac80ad 100644
--- a/pandas/_libs/lib.pyx
+++ b/pandas/_libs/lib.pyx
@@ -929,7 +929,7 @@ def is_lexsorted(list list_of_arrays):
# TODO: could do even better if we know something about the data. eg, index has
-# 1-min data, binner has 5-min data, then bins are just strides in index. This
+# 1-min data, binner has 5-min data, then bins are just strides in index. This
# is a general, O(max(len(values), len(binner))) method.
@cython.boundscheck(False)
@cython.wraparound(False)
diff --git a/pandas/_libs/parsers.pyx b/pandas/_libs/parsers.pyx
index 1f7c359b519a5..cf63b5083885e 100644
--- a/pandas/_libs/parsers.pyx
+++ b/pandas/_libs/parsers.pyx
@@ -424,7 +424,7 @@ cdef class TextReader:
if escapechar is not None:
if len(escapechar) != 1:
- raise ValueError('Only length-1 escapes supported')
+ raise ValueError('Only length-1 escapes supported')
self.parser.escapechar = ord(escapechar)
self._set_quoting(quotechar, quoting)
@@ -523,7 +523,7 @@ cdef class TextReader:
else:
if isinstance(header, list):
if len(header) > 1:
- # need to artifically skip the final line
+ # need to artificially skip the final line
# which is still a header line
header = list(header)
header.append(header[-1] + 1)
diff --git a/pandas/_libs/src/datetime/np_datetime.c b/pandas/_libs/src/datetime/np_datetime.c
index fd76f3328c05b..89753ccf7d773 100644
--- a/pandas/_libs/src/datetime/np_datetime.c
+++ b/pandas/_libs/src/datetime/np_datetime.c
@@ -327,7 +327,7 @@ int cmp_pandas_datetimestruct(const pandas_datetimestruct *a,
* this style of access anyway.
*
* Returns -1 on error, 0 on success, and 1 (with no error set)
- * if obj doesn't have the neeeded date or datetime attributes.
+ * if obj doesn't have the needed date or datetime attributes.
*/
int convert_pydatetime_to_datetimestruct(PyObject *obj,
pandas_datetimestruct *out) {
diff --git a/pandas/_libs/src/numpy.pxd b/pandas/_libs/src/numpy.pxd
index 6fa2bc6af9d1f..8ce398ce218a8 100644
--- a/pandas/_libs/src/numpy.pxd
+++ b/pandas/_libs/src/numpy.pxd
@@ -196,7 +196,7 @@ cdef extern from "numpy/arrayobject.h":
# -- the details of this may change.
def __getbuffer__(ndarray self, Py_buffer* info, int flags):
# This implementation of getbuffer is geared towards Cython
- # requirements, and does not yet fullfill the PEP.
+ # requirements, and does not yet fulfill the PEP.
# In particular strided access is always provided regardless
# of flags
diff --git a/pandas/_libs/src/period_helper.c b/pandas/_libs/src/period_helper.c
index 19f810eb54ea7..01fc46481d5b4 100644
--- a/pandas/_libs/src/period_helper.c
+++ b/pandas/_libs/src/period_helper.c
@@ -247,7 +247,7 @@ static int dInfoCalc_SetFromAbsDate(register struct date_info *dinfo,
///////////////////////////////////////////////
-// frequency specifc conversion routines
+// frequency specific conversion routines
// each function must take an integer fromDate and
// a char relation ('S' or 'E' for 'START' or 'END')
///////////////////////////////////////////////////////////////////////
diff --git a/pandas/_libs/src/ujson/lib/ultrajson.h b/pandas/_libs/src/ujson/lib/ultrajson.h
index 159645b4007e1..0470fef450dde 100644
--- a/pandas/_libs/src/ujson/lib/ultrajson.h
+++ b/pandas/_libs/src/ujson/lib/ultrajson.h
@@ -140,7 +140,7 @@ typedef int64_t JSLONG;
#endif
#if !defined(__LITTLE_ENDIAN__) && !defined(__BIG_ENDIAN__)
-#error "Endianess not supported"
+#error "Endianness not supported"
#endif
enum JSTYPES {
@@ -245,7 +245,7 @@ typedef struct __JSONObjectEncoder {
int encodeHTMLChars;
/*
- Set to an error message if error occured */
+ Set to an error message if error occurred */
const char *errorMsg;
JSOBJ errorObj;
diff --git a/pandas/_libs/src/ujson/python/ujson.c b/pandas/_libs/src/ujson/python/ujson.c
index a0c2146c30eed..da19afab030b1 100644
--- a/pandas/_libs/src/ujson/python/ujson.c
+++ b/pandas/_libs/src/ujson/python/ujson.c
@@ -58,12 +58,12 @@ PyObject *JSONFileToObj(PyObject *self, PyObject *args, PyObject *kwargs);
static PyMethodDef ujsonMethods[] = {
{"encode", (PyCFunction)objToJSON, METH_VARARGS | METH_KEYWORDS,
- "Converts arbitrary object recursivly into JSON. " ENCODER_HELP_TEXT},
+ "Converts arbitrary object recursively into JSON. " ENCODER_HELP_TEXT},
{"decode", (PyCFunction)JSONToObj, METH_VARARGS | METH_KEYWORDS,
"Converts JSON as string to dict object structure. Use precise_float=True "
"to use high precision float decoder."},
{"dumps", (PyCFunction)objToJSON, METH_VARARGS | METH_KEYWORDS,
- "Converts arbitrary object recursivly into JSON. " ENCODER_HELP_TEXT},
+ "Converts arbitrary object recursively into JSON. " ENCODER_HELP_TEXT},
{"loads", (PyCFunction)JSONToObj, METH_VARARGS | METH_KEYWORDS,
"Converts JSON as string to dict object structure. Use precise_float=True "
"to use high precision float decoder."},
diff --git a/pandas/_libs/tslibs/offsets.pyx b/pandas/_libs/tslibs/offsets.pyx
index d3278e42e413f..585c904a601ed 100644
--- a/pandas/_libs/tslibs/offsets.pyx
+++ b/pandas/_libs/tslibs/offsets.pyx
@@ -104,7 +104,7 @@ cpdef bint _is_normalized(dt):
def apply_index_wraps(func):
# Note: normally we would use `@functools.wraps(func)`, but this does
- # not play nicely wtih cython class methods
+ # not play nicely with cython class methods
def wrapper(self, other):
result = func(self, other)
if self.normalize:
@@ -316,7 +316,7 @@ class EndMixin(object):
class _BaseOffset(object):
"""
- Base class for DateOffset methods that are not overriden by subclasses
+ Base class for DateOffset methods that are not overridden by subclasses
and will (after pickle errors are resolved) go into a cdef class.
"""
_typ = "dateoffset"
@@ -783,7 +783,7 @@ cpdef int get_day_of_month(datetime other, day_opt) except? -1:
other : datetime or Timestamp
day_opt : 'start', 'end'
'start': returns 1
- 'end': returns last day of the month
+ 'end': returns last day of the month
Returns
-------
@@ -924,7 +924,7 @@ cpdef int roll_yearday(datetime other, int n, int month,
month : reference month giving the first month of the year
day_opt : 'start', 'end'
'start': returns 1
- 'end': returns last day of the month
+ 'end': returns last day of the month
Returns
-------
diff --git a/pandas/_libs/tslibs/strptime.pyx b/pandas/_libs/tslibs/strptime.pyx
index 65594de586bac..2921291973373 100644
--- a/pandas/_libs/tslibs/strptime.pyx
+++ b/pandas/_libs/tslibs/strptime.pyx
@@ -557,7 +557,7 @@ class TimeRE(dict):
"""Convert a list to a regex string for matching a directive.
Want possible matching values to be from longest to shortest. This
- prevents the possibility of a match occuring for a value that also
+ prevents the possibility of a match occurring for a value that also
a substring of a larger value that should have matched (e.g., 'abc'
matching when 'abcdef' should have been the match).
diff --git a/pandas/_libs/tslibs/timestamps.pyx b/pandas/_libs/tslibs/timestamps.pyx
index 1792f852c9e1e..c7744bf9db58e 100644
--- a/pandas/_libs/tslibs/timestamps.pyx
+++ b/pandas/_libs/tslibs/timestamps.pyx
@@ -373,7 +373,7 @@ class Timestamp(_Timestamp):
"""Pandas replacement for datetime.datetime
Timestamp is the pandas equivalent of python's Datetime
- and is interchangable with it in most cases. It's the type used
+ and is interchangeable with it in most cases. It's the type used
for the entries that make up a DatetimeIndex, and other timeseries
oriented data structures in pandas.
diff --git a/pandas/core/accessor.py b/pandas/core/accessor.py
index 53ead5e8f74a3..73e01fbf17205 100644
--- a/pandas/core/accessor.py
+++ b/pandas/core/accessor.py
@@ -17,7 +17,7 @@ def _dir_deletions(self):
return self._accessors | self._deprecations
def _dir_additions(self):
- """ add addtional __dir__ for this object """
+ """ add additional __dir__ for this object """
rv = set()
for accessor in self._accessors:
try:
diff --git a/pandas/core/algorithms.py b/pandas/core/algorithms.py
index 167f215b6c0ac..571db40537cfc 100644
--- a/pandas/core/algorithms.py
+++ b/pandas/core/algorithms.py
@@ -68,7 +68,7 @@ def _ensure_data(values, dtype=None):
return _ensure_object(np.asarray(values)), 'object', 'object'
if is_bool_dtype(values) or is_bool_dtype(dtype):
# we are actually coercing to uint64
- # until our algos suppport uint8 directly (see TODO)
+ # until our algos support uint8 directly (see TODO)
return np.asarray(values).astype('uint64'), 'bool', 'uint64'
elif is_signed_integer_dtype(values) or is_signed_integer_dtype(dtype):
return _ensure_int64(values), 'int64', 'int64'
@@ -120,7 +120,7 @@ def _ensure_data(values, dtype=None):
dtype = 'category'
# we are actually coercing to int64
- # until our algos suppport int* directly (not all do)
+ # until our algos support int* directly (not all do)
values = _ensure_int64(values)
return values, dtype, 'int64'
diff --git a/pandas/core/base.py b/pandas/core/base.py
index 72acd0052202b..e90794c6c2e1a 100644
--- a/pandas/core/base.py
+++ b/pandas/core/base.py
@@ -685,7 +685,7 @@ def _gotitem(self, key, ndim, subset=None):
class IndexOpsMixin(object):
- """ common ops mixin to support a unified inteface / docs for Series /
+ """ common ops mixin to support a unified interface / docs for Series /
Index
"""
diff --git a/pandas/core/categorical.py b/pandas/core/categorical.py
index baf15b3ca5bc4..d47cb0762447b 100644
--- a/pandas/core/categorical.py
+++ b/pandas/core/categorical.py
@@ -648,7 +648,7 @@ def _codes_for_groupby(self, sort):
Parameters
----------
sort : boolean
- The value of the sort paramter groupby was called with.
+ The value of the sort parameter groupby was called with.
Returns
-------
@@ -770,7 +770,7 @@ def set_categories(self, new_categories, ordered=None, rename=False,
If not given, do not change the ordered information.
rename : boolean (default: False)
Whether or not the new_categories should be considered as a rename
- of the old categories or as reordered categories.
+ of the old categories or as reordered categories.
inplace : boolean (default: False)
Whether or not to reorder the categories inplace or return a copy of
this categorical with reordered categories.
@@ -1139,7 +1139,7 @@ def shift(self, periods):
shifted : Categorical
"""
# since categoricals always have ndim == 1, an axis parameter
- # doesnt make any sense here.
+ # doesn't make any sense here.
codes = self.codes
if codes.ndim > 1:
raise NotImplementedError("Categorical with ndim > 1.")
diff --git a/pandas/core/common.py b/pandas/core/common.py
index 775ecc32b0f3c..e606be3cc2a23 100644
--- a/pandas/core/common.py
+++ b/pandas/core/common.py
@@ -305,7 +305,7 @@ def split_ranges(mask):
ranges = [(0, len(mask))]
for pos, val in enumerate(mask):
- if not val: # this pos should be ommited, split off the prefix range
+ if not val: # this pos should be omitted, split off the prefix range
r = ranges.pop()
if pos > r[0]: # yield non-zero range
yield (r[0], pos)
diff --git a/pandas/core/computation/expressions.py b/pandas/core/computation/expressions.py
index c74da6379e32f..1dc19d33f3365 100644
--- a/pandas/core/computation/expressions.py
+++ b/pandas/core/computation/expressions.py
@@ -71,7 +71,7 @@ def _can_use_numexpr(op, op_str, a, b, dtype_check):
# required min elements (otherwise we are adding overhead)
if np.prod(a.shape) > _MIN_ELEMENTS:
- # check for dtype compatiblity
+ # check for dtype compatibility
dtypes = set()
for o in [a, b]:
if hasattr(o, 'get_dtype_counts'):
@@ -224,7 +224,7 @@ def where(cond, a, b, use_numexpr=True):
def set_test_mode(v=True):
"""
- Keeps track of whether numexpr was used. Stores an additional ``True``
+ Keeps track of whether numexpr was used. Stores an additional ``True``
for every successful use of evaluate with numexpr since the last
``get_test_result``
"""
diff --git a/pandas/core/computation/pytables.py b/pandas/core/computation/pytables.py
index 4b3c608a88be8..26eefa75b2675 100644
--- a/pandas/core/computation/pytables.py
+++ b/pandas/core/computation/pytables.py
@@ -439,7 +439,7 @@ def visit_Attribute(self, node, **kwargs):
return self.term_type(getattr(resolved, attr), self.env)
except AttributeError:
- # something like datetime.datetime where scope is overriden
+ # something like datetime.datetime where scope is overridden
if isinstance(value, ast.Name) and value.id == attr:
return resolved
diff --git a/pandas/core/config.py b/pandas/core/config.py
index e71c3b6f58562..d10e2d19be665 100644
--- a/pandas/core/config.py
+++ b/pandas/core/config.py
@@ -23,7 +23,7 @@
- all options in a certain sub - namespace can be reset at once.
- the user can set / get / reset or ask for the description of an option.
- a developer can register and mark an option as deprecated.
-- you can register a callback to be invoked when the the option value
+- you can register a callback to be invoked when the option value
is set or reset. Changing the stored value is considered misuse, but
is not verboten.
@@ -33,8 +33,8 @@
- Data is stored using nested dictionaries, and should be accessed
through the provided API.
-- "Registered options" and "Deprecated options" have metadata associcated
- with them, which are stored in auxilary dictionaries keyed on the
+- "Registered options" and "Deprecated options" have metadata associated
+ with them, which are stored in auxiliary dictionaries keyed on the
fully-qualified key, e.g. "x.y.z.option".
- the config_init module is imported by the package's __init__.py file.
@@ -209,7 +209,7 @@ def __dir__(self):
# in the docstring. For dev convenience we'd like to generate the docstrings
# dynamically instead of maintaining them by hand. To this, we use the
# class below which wraps functions inside a callable, and converts
-# __doc__ into a propery function. The doctsrings below are templates
+# __doc__ into a property function. The doctsrings below are templates
# using the py2.6+ advanced formatting syntax to plug in a concise list
# of options, and option descriptions.
@@ -691,7 +691,7 @@ def pp(name, ks):
@contextmanager
def config_prefix(prefix):
- """contextmanager for multiple invocations of API with a common prefix
+ """contextmanager for multiple invocations of API with a common prefix
supported API functions: (register / get / set )__option
diff --git a/pandas/core/dtypes/cast.py b/pandas/core/dtypes/cast.py
index 87c6fb69f33bf..5fcb5f09dfae7 100644
--- a/pandas/core/dtypes/cast.py
+++ b/pandas/core/dtypes/cast.py
@@ -520,7 +520,7 @@ def maybe_infer_dtype_type(element):
def maybe_upcast(values, fill_value=np.nan, dtype=None, copy=False):
- """ provide explict type promotion and coercion
+ """ provide explicit type promotion and coercion
Parameters
----------
diff --git a/pandas/core/dtypes/dtypes.py b/pandas/core/dtypes/dtypes.py
index a47f2c0d4ab13..d1637873eb6e1 100644
--- a/pandas/core/dtypes/dtypes.py
+++ b/pandas/core/dtypes/dtypes.py
@@ -597,8 +597,8 @@ def is_dtype(cls, dtype):
"""
if isinstance(dtype, compat.string_types):
- # PeriodDtype can be instanciated from freq string like "U",
- # but dosn't regard freq str like "U" as dtype.
+ # PeriodDtype can be instantiated from freq string like "U",
+ # but doesn't regard freq str like "U" as dtype.
if dtype.startswith('period[') or dtype.startswith('Period['):
try:
if cls._parse_dtype_strict(dtype) is not None:
diff --git a/pandas/core/dtypes/missing.py b/pandas/core/dtypes/missing.py
index ce57b544d9d66..d208c72ffee19 100644
--- a/pandas/core/dtypes/missing.py
+++ b/pandas/core/dtypes/missing.py
@@ -362,7 +362,7 @@ def _infer_fill_value(val):
def _maybe_fill(arr, fill_value=np.nan):
"""
- if we have a compatiable fill_value and arr dtype, then fill
+ if we have a compatible fill_value and arr dtype, then fill
"""
if _isna_compat(arr, fill_value):
arr.fill(fill_value)
diff --git a/pandas/core/frame.py b/pandas/core/frame.py
index 12a4a7fdaedad..d85f3bf552617 100644
--- a/pandas/core/frame.py
+++ b/pandas/core/frame.py
@@ -591,7 +591,7 @@ def _repr_fits_horizontal_(self, ignore_width=False):
max_rows = get_option("display.max_rows")
# when auto-detecting, so width=None and not in ipython front end
- # check whether repr fits horizontal by actualy checking
+ # check whether repr fits horizontal by actually checking
# the width of the rendered repr
buf = StringIO()
@@ -1578,7 +1578,7 @@ def to_stata(self, fname, convert_dates=None, write_index=True,
String path of file-like object
convert_dates : dict
Dictionary mapping columns containing datetime types to stata
- internal format to use when wirting the dates. Options are 'tc',
+ internal format to use when writing the dates. Options are 'tc',
'td', 'tm', 'tw', 'th', 'tq', 'ty'. Column can be either an integer
or a name. Datetime columns that do not have a conversion type
specified will be converted to 'tc'. Raises NotImplementedError if
@@ -1606,7 +1606,7 @@ def to_stata(self, fname, convert_dates=None, write_index=True,
* If datetimes contain timezone information
* Column dtype is not representable in Stata
ValueError
- * Columns listed in convert_dates are noth either datetime64[ns]
+ * Columns listed in convert_dates are neither datetime64[ns]
or datetime.datetime
* Column listed in convert_dates is not in DataFrame
* Categorical label contains more than 32,000 characters
@@ -5736,7 +5736,7 @@ def idxmax(self, axis=0, skipna=True):
return Series(result, index=self._get_agg_axis(axis))
def _get_agg_axis(self, axis_num):
- """ let's be explict about this """
+ """ let's be explicit about this """
if axis_num == 0:
return self.columns
elif axis_num == 1:
diff --git a/pandas/core/generic.py b/pandas/core/generic.py
index c5359ba2c5ea1..004cfce6769c8 100644
--- a/pandas/core/generic.py
+++ b/pandas/core/generic.py
@@ -2341,7 +2341,7 @@ def _check_setitem_copy(self, stacklevel=4, t='setting', force=False):
if value is None:
return
- # see if the copy is not actually refererd; if so, then disolve
+ # see if the copy is not actually referred; if so, then dissolve
# the copy weakref
try:
gc.collect(2)
@@ -3109,7 +3109,7 @@ def sort_index(self, axis=0, level=None, ascending=True, inplace=False,
%(optional_axis)s
method : {None, 'backfill'/'bfill', 'pad'/'ffill', 'nearest'}, optional
method to use for filling holes in reindexed DataFrame.
- Please note: this is only applicable to DataFrames/Series with a
+ Please note: this is only applicable to DataFrames/Series with a
monotonically increasing/decreasing index.
* default: don't fill gaps
diff --git a/pandas/core/groupby.py b/pandas/core/groupby.py
index 06b7dbb4ecf7b..285a347153a82 100644
--- a/pandas/core/groupby.py
+++ b/pandas/core/groupby.py
@@ -2956,7 +2956,7 @@ def is_in_axis(key):
return True
- # if the the grouper is obj[name]
+ # if the grouper is obj[name]
def is_in_obj(gpr):
try:
return id(gpr) == id(obj[gpr.name])
diff --git a/pandas/core/indexes/base.py b/pandas/core/indexes/base.py
index 04b8ade7e5253..99ee28c84365f 100644
--- a/pandas/core/indexes/base.py
+++ b/pandas/core/indexes/base.py
@@ -319,7 +319,7 @@ def __new__(cls, data=None, dtype=None, copy=False, name=None,
return IntervalIndex.from_intervals(subarr, name=name,
copy=copy)
elif inferred == 'boolean':
- # don't support boolean explicity ATM
+ # don't support boolean explicitly ATM
pass
elif inferred != 'string':
if inferred.startswith('datetime'):
@@ -887,7 +887,7 @@ def _format_data(self, name=None):
# are we a truncated display
is_truncated = n > max_seq_items
- # adj can optionaly handle unicode eastern asian width
+ # adj can optionally handle unicode eastern asian width
adj = _get_adjustment()
def _extend_line(s, line, value, display_width, next_line_prefix):
@@ -1788,7 +1788,7 @@ def _concat_same_dtype(self, to_concat, name):
"""
Concatenate to_concat which has the same class
"""
- # must be overrided in specific classes
+ # must be overridden in specific classes
return _concat._concat_index_asobject(to_concat, name)
_index_shared_docs['take'] = """
@@ -3276,7 +3276,7 @@ def _get_leaf_sorter(labels):
sorter, _ = libalgos.groupsort_indexer(lab, 1 + lab.max())
return sorter
- # find indexers of begining of each set of
+ # find indexers of beginning of each set of
# same-key labels w.r.t all but last level
tic = labels[0][:-1] != labels[0][1:]
for lab in labels[1:-1]:
@@ -3573,7 +3573,7 @@ def _searchsorted_monotonic(self, label, side='left'):
def _get_loc_only_exact_matches(self, key):
"""
- This is overriden on subclasses (namely, IntervalIndex) to control
+ This is overridden on subclasses (namely, IntervalIndex) to control
get_slice_bound.
"""
return self.get_loc(key)
diff --git a/pandas/core/indexes/category.py b/pandas/core/indexes/category.py
index 241907a54f393..ac7cb30fa823d 100644
--- a/pandas/core/indexes/category.py
+++ b/pandas/core/indexes/category.py
@@ -522,7 +522,7 @@ def reindex(self, target, method=None, level=None, limit=None,
# we always want to return an Index type here
# to be consistent with .reindex for other index types (e.g. they don't
# coerce based on the actual values, only on the dtype)
- # unless we had an inital Categorical to begin with
+ # unless we had an initial Categorical to begin with
# in which case we are going to conform to the passed Categorical
new_target = np.asarray(new_target)
if is_categorical_dtype(target):
@@ -746,7 +746,7 @@ def _evaluate_compare(self, other):
if isinstance(other, ABCCategorical):
if not self.values.is_dtype_equal(other):
- raise TypeError("categorical index comparisions must "
+ raise TypeError("categorical index comparisons must "
"have the same categories and ordered "
"attributes")
diff --git a/pandas/core/indexes/datetimelike.py b/pandas/core/indexes/datetimelike.py
index 40c07376d2522..3fca40562899a 100644
--- a/pandas/core/indexes/datetimelike.py
+++ b/pandas/core/indexes/datetimelike.py
@@ -132,7 +132,7 @@ def ceil(self, freq):
class DatetimeIndexOpsMixin(object):
- """ common ops mixin to support a unified inteface datetimelike Index """
+ """ common ops mixin to support a unified interface datetimelike Index """
def equals(self, other):
"""
diff --git a/pandas/core/indexes/datetimes.py b/pandas/core/indexes/datetimes.py
index af901440d8abd..b17682b6c3448 100644
--- a/pandas/core/indexes/datetimes.py
+++ b/pandas/core/indexes/datetimes.py
@@ -355,7 +355,7 @@ def __new__(cls, data=None,
raise ValueError("Must provide freq argument if no data is "
"supplied")
- # if dtype has an embeded tz, capture it
+ # if dtype has an embedded tz, capture it
if dtype is not None:
try:
dtype = DatetimeTZDtype.construct_from_string(dtype)
diff --git a/pandas/core/indexes/interval.py b/pandas/core/indexes/interval.py
index 2a132f683c519..def9b151f5c91 100644
--- a/pandas/core/indexes/interval.py
+++ b/pandas/core/indexes/interval.py
@@ -683,7 +683,7 @@ def inferred_type(self):
@Appender(Index.memory_usage.__doc__)
def memory_usage(self, deep=False):
- # we don't use an explict engine
+ # we don't use an explicit engine
# so return the bytes here
return (self.left.memory_usage(deep=deep) +
self.right.memory_usage(deep=deep))
diff --git a/pandas/core/indexes/multi.py b/pandas/core/indexes/multi.py
index f4c4f91d2cc57..7107378671ba5 100644
--- a/pandas/core/indexes/multi.py
+++ b/pandas/core/indexes/multi.py
@@ -799,8 +799,8 @@ def _hashed_indexing_key(self, key):
*this is internal for use for the cython routines*
- Paramters
- ---------
+ Parameters
+ ----------
key : string or tuple
Returns
diff --git a/pandas/core/indexing.py b/pandas/core/indexing.py
index de6713249a7c7..fa6614d27cd19 100755
--- a/pandas/core/indexing.py
+++ b/pandas/core/indexing.py
@@ -1308,7 +1308,7 @@ class _IXIndexer(_NDFrameIndexer):
``.ix`` is the most general indexer and will support any of the
inputs in ``.loc`` and ``.iloc``. ``.ix`` also supports floating
point label schemes. ``.ix`` is exceptionally useful when dealing
- with mixed positional and label based hierachical indexes.
+ with mixed positional and label based hierarchical indexes.
However, when an axis is integer based, ONLY label based access
and not positional access is supported. Thus, in such cases, it's
@@ -1441,8 +1441,8 @@ def _has_valid_type(self, key, axis):
ax = self.obj._get_axis(axis)
# valid for a label where all labels are in the index
- # slice of lables (where start-end in labels)
- # slice of integers (only if in the lables)
+ # slice of labels (where start-end in labels)
+ # slice of integers (only if in the labels)
# boolean
if isinstance(key, slice):
@@ -1929,7 +1929,7 @@ def _has_valid_setitem_indexer(self, indexer):
self._has_valid_positional_setitem_indexer(indexer)
def _convert_key(self, key, is_setter=False):
- """ require integer args (and convert to label arguments) """
+ """ require integer args (and convert to label arguments) """
for a, i in zip(self.obj.axes, key):
if not is_integer(i):
raise ValueError("iAt based indexing can only have integer "
@@ -2118,7 +2118,7 @@ def maybe_convert_ix(*args):
def is_nested_tuple(tup, labels):
- # check for a compatiable nested tuple and multiindexes among the axes
+ # check for a compatible nested tuple and multiindexes among the axes
if not isinstance(tup, tuple):
return False
diff --git a/pandas/core/internals.py b/pandas/core/internals.py
index 3a64a0ef84e3d..ba90503e3bf40 100644
--- a/pandas/core/internals.py
+++ b/pandas/core/internals.py
@@ -170,7 +170,7 @@ def formatting_values(self):
def get_values(self, dtype=None):
"""
return an internal format, currently just the ndarray
- this is often overriden to handle to_dense like operations
+ this is often overridden to handle to_dense like operations
"""
if is_object_dtype(dtype):
return self.values.astype(object)
@@ -954,7 +954,7 @@ def putmask(self, mask, new, align=True, inplace=False, axis=0,
new_values = new_values.T
# If the default repeat behavior in np.putmask would go in the
- # wrong direction, then explictly repeat and reshape new instead
+ # wrong direction, then explicitly repeat and reshape new instead
if getattr(new, 'ndim', 0) >= 1:
if self.ndim - 1 == new.ndim and axis == 1:
new = np.repeat(
@@ -1455,7 +1455,7 @@ def where(self, other, cond, align=True, errors='raise',
cond = cond.values
# If the default broadcasting would go in the wrong direction, then
- # explictly reshape other instead
+ # explicitly reshape other instead
if getattr(other, 'ndim', 0) >= 1:
if values.ndim - 1 == other.ndim and axis == 1:
other = other.reshape(tuple(other.shape + (1, )))
@@ -1493,7 +1493,7 @@ def func(cond, values, other):
except TypeError:
# we cannot coerce, return a compat dtype
- # we are explicity ignoring errors
+ # we are explicitly ignoring errors
block = self.coerce_to_target_dtype(other)
blocks = block.where(orig_other, cond, align=align,
errors=errors,
@@ -4939,7 +4939,7 @@ def _maybe_compare(a, b, op):
is_a_array = isinstance(a, np.ndarray)
is_b_array = isinstance(b, np.ndarray)
- # numpy deprecation warning to have i8 vs integer comparisions
+ # numpy deprecation warning to have i8 vs integer comparisons
if is_datetimelike_v_numeric(a, b):
result = False
diff --git a/pandas/core/missing.py b/pandas/core/missing.py
index c3e72d6c31bf5..74fa21fa4b53d 100644
--- a/pandas/core/missing.py
+++ b/pandas/core/missing.py
@@ -127,7 +127,7 @@ def interpolate_1d(xvalues, yvalues, method='linear', limit=None,
if not valid.any():
# have to call np.asarray(xvalues) since xvalues could be an Index
- # which cant be mutated
+ # which can't be mutated
result = np.empty_like(np.asarray(xvalues), dtype=np.float64)
result.fill(np.nan)
return result
diff --git a/pandas/core/ops.py b/pandas/core/ops.py
index 3a7a5e44d5a88..faac8ab312d6b 100644
--- a/pandas/core/ops.py
+++ b/pandas/core/ops.py
@@ -1357,7 +1357,7 @@ def f(self, other):
return self._combine_series_infer(other, func, try_cast=False)
else:
- # straight boolean comparisions we want to allow all columns
+ # straight boolean comparisons we want to allow all columns
# (regardless of dtype to pass thru) See #4537 for discussion.
res = self._combine_const(other, func,
errors='ignore',
diff --git a/pandas/core/panel.py b/pandas/core/panel.py
index 6d85e5bf7c7f9..7ec177b03aeb1 100644
--- a/pandas/core/panel.py
+++ b/pandas/core/panel.py
@@ -860,7 +860,7 @@ def xs(self, key, axis=1):
xs is only for getting, not setting values.
MultiIndex Slicers is a generic way to get/set values on any level or
- levels and is a superset of xs functionality, see
+ levels and is a superset of xs functionality, see
:ref:`MultiIndex Slicers <advanced.mi_slicers>`
"""
diff --git a/pandas/core/reshape/concat.py b/pandas/core/reshape/concat.py
index 9bd5abb2cd476..aaadf6d3ca32f 100644
--- a/pandas/core/reshape/concat.py
+++ b/pandas/core/reshape/concat.py
@@ -276,7 +276,7 @@ def __init__(self, objs, axis=0, join='outer', join_axes=None,
ndims.add(obj.ndim)
# get the sample
- # want the higest ndim that we have, and must be non-empty
+ # want the highest ndim that we have, and must be non-empty
# unless all objs are empty
sample = None
if len(ndims) > 1:
diff --git a/pandas/core/reshape/melt.py b/pandas/core/reshape/melt.py
index c2804c8f8e63e..b648c426a877f 100644
--- a/pandas/core/reshape/melt.py
+++ b/pandas/core/reshape/melt.py
@@ -186,7 +186,7 @@ def wide_to_long(df, stubnames, i, j, sep="", suffix=r'\d+'):
A character indicating the separation of the variable names
in the wide format, to be stripped from the names in the long format.
For example, if your column names are A-suffix1, A-suffix2, you
- can strip the hypen by specifying `sep='-'`
+ can strip the hyphen by specifying `sep='-'`
.. versionadded:: 0.20.0
diff --git a/pandas/core/series.py b/pandas/core/series.py
index 360095c386e8b..5d8092fd30496 100644
--- a/pandas/core/series.py
+++ b/pandas/core/series.py
@@ -345,7 +345,7 @@ def _set_axis(self, axis, labels, fastpath=False):
(DatetimeIndex, PeriodIndex, TimedeltaIndex)):
try:
labels = DatetimeIndex(labels)
- # need to set here becuase we changed the index
+ # need to set here because we changed the index
if fastpath:
self._data.set_axis(axis, labels)
except (libts.OutOfBoundsDatetime, ValueError):
@@ -487,7 +487,7 @@ def nonzero(self):
Return the indices of the elements that are non-zero
This method is equivalent to calling `numpy.nonzero` on the
- series data. For compatability with NumPy, the return value is
+ series data. For compatibility with NumPy, the return value is
the same (a tuple with an array of indices for each dimension),
but it will always be a one-item tuple because series only have
one dimension.
@@ -2388,7 +2388,7 @@ def aggregate(self, func, axis=0, *args, **kwargs):
# expression, e.g.: lambda x: x-x.quantile(0.25)
# this will fail, so we can try a vectorized evaluation
- # we cannot FIRST try the vectorized evaluation, becuase
+ # we cannot FIRST try the vectorized evaluation, because
# then .agg and .apply would have different semantics if the
# operation is actually defined on the Series, e.g. str
try:
diff --git a/pandas/core/sparse/array.py b/pandas/core/sparse/array.py
index 0424ac8703e25..9b2650359bf68 100644
--- a/pandas/core/sparse/array.py
+++ b/pandas/core/sparse/array.py
@@ -525,7 +525,7 @@ def __setitem__(self, key, value):
# if is_integer(key):
# self.values[key] = value
# else:
- # raise Exception("SparseArray does not support seting non-scalars
+ # raise Exception("SparseArray does not support setting non-scalars
# via setitem")
raise TypeError(
"SparseArray does not support item assignment via setitem")
@@ -538,7 +538,7 @@ def __setslice__(self, i, j, value):
slobj = slice(i, j) # noqa
# if not is_scalar(value):
- # raise Exception("SparseArray does not support seting non-scalars
+ # raise Exception("SparseArray does not support setting non-scalars
# via slices")
# x = self.values
diff --git a/pandas/core/strings.py b/pandas/core/strings.py
index 99c7563d5b249..fab4e77ce4467 100644
--- a/pandas/core/strings.py
+++ b/pandas/core/strings.py
@@ -1414,7 +1414,7 @@ def _wrap_result(self, result, use_codes=True,
elif expand is True and not isinstance(self._orig, Index):
# required when expand=True is explicitly specified
- # not needed when infered
+ # not needed when inferred
def cons_row(x):
if is_list_like(x):
@@ -1424,7 +1424,7 @@ def cons_row(x):
result = [cons_row(x) for x in result]
if result:
- # propogate nan values to match longest sequence (GH 18450)
+ # propagate nan values to match longest sequence (GH 18450)
max_len = max(len(x) for x in result)
result = [x * max_len if x[0] is np.nan else x for x in result]
diff --git a/pandas/core/tools/datetimes.py b/pandas/core/tools/datetimes.py
index 6b8edbb146e4b..1de43116d0b49 100644
--- a/pandas/core/tools/datetimes.py
+++ b/pandas/core/tools/datetimes.py
@@ -197,7 +197,8 @@ def to_datetime(arg, errors='raise', dayfirst=False, yearfirst=False,
In case when it is not possible to return designated types (e.g. when
any element of input is before Timestamp.min or after Timestamp.max)
- return will have datetime.datetime type (or correspoding array/Series).
+ return will have datetime.datetime type (or corresponding
+ array/Series).
Examples
--------
@@ -497,7 +498,7 @@ def _convert_listlike(arg, box, format, name=None, tz=tz):
def _assemble_from_unit_mappings(arg, errors):
"""
- assemble the unit specifed fields from the arg (DataFrame)
+ assemble the unit specified fields from the arg (DataFrame)
Return a Series for actual parsing
Parameters
diff --git a/pandas/core/window.py b/pandas/core/window.py
index 5ad8d20cc03e2..76ba76b7a9da9 100644
--- a/pandas/core/window.py
+++ b/pandas/core/window.py
@@ -253,8 +253,8 @@ def _wrap_results(self, results, blocks, obj):
"""
wrap the results
- Paramters
- ---------
+ Parameters
+ ----------
results : list of ndarrays
blocks : list of blocks
obj : conformed data (may be resampled)
@@ -403,7 +403,7 @@ class Window(_Window):
3 NaN
4 NaN
- Same as above, but explicity set the min_periods
+ Same as above, but explicitly set the min_periods
>>> df.rolling(2, min_periods=1).sum()
B
diff --git a/pandas/errors/__init__.py b/pandas/errors/__init__.py
index 42b3bdd4991a9..b3d1ce31d66ae 100644
--- a/pandas/errors/__init__.py
+++ b/pandas/errors/__init__.py
@@ -38,7 +38,7 @@ class ParserError(ValueError):
class DtypeWarning(Warning):
"""
- Warning that is raised for a dtype incompatiblity. This
+ Warning that is raised for a dtype incompatibility. This
can happen whenever `pd.read_csv` encounters non-
uniform dtypes in a column(s) of a given CSV file.
"""
@@ -56,7 +56,7 @@ class ParserWarning(Warning):
Warning that is raised in `pd.read_csv` whenever it is necessary
to change parsers (generally from 'c' to 'python') contrary to the
one specified by the user due to lack of support or functionality for
- parsing particular attributes of a CSV file with the requsted engine.
+ parsing particular attributes of a CSV file with the requested engine.
"""
diff --git a/pandas/io/common.py b/pandas/io/common.py
index 534c1e0671150..da60698fe529f 100644
--- a/pandas/io/common.py
+++ b/pandas/io/common.py
@@ -312,7 +312,7 @@ def _get_handle(path_or_buf, mode, encoding=None, compression=None,
f : file-like
A file-like object
handles : list of file-like objects
- A list of file-like object that were openned in this function.
+ A list of file-like object that were opened in this function.
"""
try:
from s3fs import S3File
@@ -533,7 +533,7 @@ def _check_as_is(x):
# Fetch UTF-8 output from the queue ...
data = self.queue.getvalue()
data = data.decode("utf-8")
- # ... and reencode it into the target encoding
+ # ... and re-encode it into the target encoding
data = self.encoder.encode(data)
# write to the target stream
self.stream.write(data)
@@ -553,7 +553,7 @@ def _check_as_is(x):
# Fetch UTF-8 output from the queue ...
data = self.queue.getvalue()
data = data.decode("utf-8")
- # ... and reencode it into the target encoding
+ # ... and re-encode it into the target encoding
data = self.encoder.encode(data)
# write to the target stream
self.stream.write(data)
diff --git a/pandas/io/excel.py b/pandas/io/excel.py
index 4f0655cff9b57..92b29c8da7e3f 100644
--- a/pandas/io/excel.py
+++ b/pandas/io/excel.py
@@ -561,7 +561,7 @@ def _parse_cell(cell_contents, cell_typ):
cell_contents = bool(cell_contents)
elif convert_float and cell_typ == XL_CELL_NUMBER:
# GH5394 - Excel 'numbers' are always floats
- # it's a minimal perf hit and less suprising
+ # it's a minimal perf hit and less surprising
val = int(cell_contents)
if val == cell_contents:
cell_contents = val
@@ -881,12 +881,12 @@ def engine(self):
def write_cells(self, cells, sheet_name=None, startrow=0, startcol=0,
freeze_panes=None):
"""
- Write given formated cells into Excel an excel sheet
+ Write given formatted cells into Excel an excel sheet
Parameters
----------
cells : generator
- cell of formated data to save to Excel sheet
+ cell of formatted data to save to Excel sheet
sheet_name : string, default None
Name of Excel sheet, if None, then use self.cur_sheet
startrow: upper left cell row to dump data frame
diff --git a/pandas/io/formats/console.py b/pandas/io/formats/console.py
index bdff59939a4de..36eac8dd57fbd 100644
--- a/pandas/io/formats/console.py
+++ b/pandas/io/formats/console.py
@@ -14,7 +14,7 @@
def detect_console_encoding():
"""
Try to find the most capable encoding supported by the console.
- slighly modified from the way IPython handles the same issue.
+ slightly modified from the way IPython handles the same issue.
"""
global _initial_defencoding
diff --git a/pandas/io/formats/excel.py b/pandas/io/formats/excel.py
index a36e82edf6e57..aff3e35861434 100644
--- a/pandas/io/formats/excel.py
+++ b/pandas/io/formats/excel.py
@@ -492,7 +492,7 @@ def _format_regular_rows(self):
# output index and index_label?
if self.index:
- # chek aliases
+ # check aliases
# if list only take first as this is not a MultiIndex
if (self.index_label and
isinstance(self.index_label, (list, tuple, np.ndarray,
diff --git a/pandas/io/formats/style.py b/pandas/io/formats/style.py
index 3af9e78a5aac4..2c3d92cea0ad8 100644
--- a/pandas/io/formats/style.py
+++ b/pandas/io/formats/style.py
@@ -420,7 +420,7 @@ def render(self, **kwargs):
the rendered HTML in the notebook.
Pandas uses the following keys in render. Arguments passed
- in ``**kwargs`` take precedence, so think carefuly if you want
+ in ``**kwargs`` take precedence, so think carefully if you want
to override them:
* head
@@ -1201,7 +1201,7 @@ def _is_visible(idx_row, idx_col, lengths):
def _get_level_lengths(index, hidden_elements=None):
"""
- Given an index, find the level lenght for each element.
+ Given an index, find the level length for each element.
Optional argument is a list of index positions which
should not be visible.
@@ -1229,7 +1229,7 @@ def _get_level_lengths(index, hidden_elements=None):
lengths[(i, last_label)] = 1
elif (row != sentinel):
# even if its hidden, keep track of it in case
- # length >1 and later elemens are visible
+ # length >1 and later elements are visible
last_label = j
lengths[(i, last_label)] = 0
elif(j not in hidden_elements):
diff --git a/pandas/io/html.py b/pandas/io/html.py
index 67a48198adc27..e7794864ccb3e 100644
--- a/pandas/io/html.py
+++ b/pandas/io/html.py
@@ -263,7 +263,7 @@ def _parse_tables(self, doc, match, attrs):
attrs : dict
A dictionary of table attributes that can be used to disambiguate
- mutliple tables on a page.
+ multiple tables on a page.
Raises
------
diff --git a/pandas/io/json/json.py b/pandas/io/json/json.py
index bb435c625ff35..72ec5c59c90af 100644
--- a/pandas/io/json/json.py
+++ b/pandas/io/json/json.py
@@ -162,7 +162,7 @@ class JSONTableWriter(FrameWriter):
def __init__(self, obj, orient, date_format, double_precision,
ensure_ascii, date_unit, index, default_handler=None):
"""
- Adds a `schema` attribut with the Table Schema, resets
+ Adds a `schema` attribute with the Table Schema, resets
the index (can't do in caller, because the schema inference needs
to know what the index is, forces orient to records, and forces
date_format to 'iso'.
@@ -534,7 +534,7 @@ def _get_object_parser(self, json):
def close(self):
"""
- If we opened a stream earlier, in _get_data_from_filepath, we should
+ If we opened a stream earlier, in _get_data_from_filepath, we should
close it. If an open stream or file was passed, we leave it open.
"""
if self.should_close:
diff --git a/pandas/io/msgpack/_unpacker.pyx b/pandas/io/msgpack/_unpacker.pyx
index 05dfaad8b2058..04bb330e595dd 100644
--- a/pandas/io/msgpack/_unpacker.pyx
+++ b/pandas/io/msgpack/_unpacker.pyx
@@ -202,7 +202,7 @@ cdef class Unpacker(object):
:param int max_buffer_size:
Limits size of data waiting unpacked. 0 means system's
INT_MAX (default). Raises `BufferFull` exception when it
- is insufficient. You shoud set this parameter when unpacking
+ is insufficient. You should set this parameter when unpacking
data from untrasted source.
:param int max_str_len:
diff --git a/pandas/io/packers.py b/pandas/io/packers.py
index ef65a3275060b..9289853a1bbfd 100644
--- a/pandas/io/packers.py
+++ b/pandas/io/packers.py
@@ -70,7 +70,7 @@
move_into_mutable_buffer as _move_into_mutable_buffer,
)
-# check whcih compression libs we have installed
+# check which compression libs we have installed
try:
import zlib
diff --git a/pandas/io/parsers.py b/pandas/io/parsers.py
index e053af17667c4..acb7d00284693 100755
--- a/pandas/io/parsers.py
+++ b/pandas/io/parsers.py
@@ -1541,7 +1541,7 @@ def _convert_to_ndarrays(self, dct, na_values, na_fvalues, verbose=False,
values, set(col_na_values) | col_na_fvalues,
try_num_bool)
- # type specificed in dtype param
+ # type specified in dtype param
if cast_type and not is_dtype_equal(cvals, cast_type):
cvals = self._cast_types(cvals, cast_type, c)
@@ -2054,7 +2054,7 @@ def __init__(self, f, **kwds):
self.data = f
# Get columns in two steps: infer from data, then
- # infer column indices from self.usecols if is is specified.
+ # infer column indices from self.usecols if it is specified.
self._col_indices = None
self.columns, self.num_original_columns = self._infer_columns()
diff --git a/pandas/io/pytables.py b/pandas/io/pytables.py
index c428000d73593..efe6ab6c18868 100644
--- a/pandas/io/pytables.py
+++ b/pandas/io/pytables.py
@@ -97,7 +97,7 @@ def _ensure_term(where, scope_level):
create the terms here with a frame_level=2 (we are 2 levels down)
"""
- # only consider list/tuple here as an ndarray is automaticaly a coordinate
+ # only consider list/tuple here as an ndarray is automatically a coordinate
# list
level = scope_level + 1
if isinstance(where, (list, tuple)):
@@ -301,7 +301,7 @@ def read_hdf(path_or_buf, key=None, mode='r', **kwargs):
contains a single pandas object.
mode : string, {'r', 'r+', 'a'}, default 'r'. Mode to use when opening
the file. Ignored if path_or_buf is a pd.HDFStore.
- where : list of Term (or convertable) objects, optional
+ where : list of Term (or convertible) objects, optional
start : optional, integer (defaults to None), row number to start
selection
stop : optional, integer (defaults to None), row number to stop
@@ -498,7 +498,7 @@ def __getattr__(self, name):
(type(self).__name__, name))
def __contains__(self, key):
- """ check for existance of this key
+ """ check for existence of this key
can match the exact pathname or the pathnm w/o the leading '/'
"""
node = self.get_node(key)
@@ -679,7 +679,7 @@ def select(self, key, where=None, start=None, stop=None, columns=None,
Parameters
----------
key : object
- where : list of Term (or convertable) objects, optional
+ where : list of Term (or convertible) objects, optional
start : integer (defaults to None), row number to start selection
stop : integer (defaults to None), row number to stop selection
columns : a list of columns that if not None, will limit the return
@@ -724,7 +724,7 @@ def select_as_coordinates(
Parameters
----------
key : object
- where : list of Term (or convertable) objects, optional
+ where : list of Term (or convertible) objects, optional
start : integer (defaults to None), row number to start selection
stop : integer (defaults to None), row number to stop selection
"""
@@ -873,7 +873,7 @@ def remove(self, key, where=None, start=None, stop=None):
----------
key : string
Node to remove or delete rows from
- where : list of Term (or convertable) objects, optional
+ where : list of Term (or convertible) objects, optional
start : integer (defaults to None), row number to start selection
stop : integer (defaults to None), row number to stop selection
@@ -1250,7 +1250,7 @@ def error(t):
# existing node (and must be a table)
if tt is None:
- # if we are a writer, determin the tt
+ # if we are a writer, determine the tt
if value is not None:
if pt == u('series_table'):
@@ -1370,7 +1370,7 @@ class TableIterator(object):
----------
store : the reference store
- s : the refered storer
+ s : the referred storer
func : the function to execute the query
where : the where of the query
nrows : the rows to iterate on
@@ -4653,7 +4653,7 @@ class Selection(object):
Parameters
----------
table : a Table object
- where : list of Terms (or convertable to)
+ where : list of Terms (or convertible to)
start, stop: indicies to start and/or stop selection
"""
@@ -4718,7 +4718,7 @@ def generate(self, where):
raise ValueError(
"The passed where expression: {0}\n"
" contains an invalid variable reference\n"
- " all of the variable refrences must be a "
+ " all of the variable references must be a "
"reference to\n"
" an axis (e.g. 'index' or 'columns'), or a "
"data_column\n"
diff --git a/pandas/io/sql.py b/pandas/io/sql.py
index c7bbbf9940ba1..e2f3033c580a5 100644
--- a/pandas/io/sql.py
+++ b/pandas/io/sql.py
@@ -1484,7 +1484,7 @@ def to_sql(self, frame, name, if_exists='fail', index=True,
`index` is True, then the index names are used.
A sequence should be given if the DataFrame uses MultiIndex.
schema : string, default None
- Ignored parameter included for compatability with SQLAlchemy
+ Ignored parameter included for compatibility with SQLAlchemy
version of ``to_sql``.
chunksize : int, default None
If not None, then rows will be written in batches of this
diff --git a/pandas/io/stata.py b/pandas/io/stata.py
index aafe5f2ce76bd..2b97b447921bb 100644
--- a/pandas/io/stata.py
+++ b/pandas/io/stata.py
@@ -645,7 +645,7 @@ def __init__(self, catarray):
def _encode(self, s):
"""
- Python 3 compatability shim
+ Python 3 compatibility shim
"""
if compat.PY3:
return s.encode(self._encoding)
@@ -968,7 +968,7 @@ def __init__(self, path_or_buf, convert_dates=True,
self._order_categoricals = order_categoricals
if encoding is not None:
if encoding not in VALID_ENCODINGS:
- raise ValueError('Unknown encoding. Only latin-1 and ascii '
+ raise ValueError('Unknown encoding. Only latin-1 and ascii '
'supported.')
self._encoding = encoding
self._chunksize = chunksize
@@ -1881,7 +1881,7 @@ class StataWriter(StataParser):
Input to save
convert_dates : dict
Dictionary mapping columns containing datetime types to stata internal
- format to use when wirting the dates. Options are 'tc', 'td', 'tm',
+ format to use when writing the dates. Options are 'tc', 'td', 'tm',
'tw', 'th', 'tq', 'ty'. Column can be either an integer or a name.
Datetime columns that do not have a conversion type specified will be
converted to 'tc'. Raises NotImplementedError if a datetime column has
@@ -1913,7 +1913,7 @@ class StataWriter(StataParser):
NotImplementedError
* If datetimes contain timezone information
ValueError
- * Columns listed in convert_dates are noth either datetime64[ns]
+ * Columns listed in convert_dates are neither datetime64[ns]
or datetime.datetime
* Column dtype is not representable in Stata
* Column listed in convert_dates is not in DataFrame
diff --git a/pandas/plotting/_core.py b/pandas/plotting/_core.py
index 9d74a308f79c8..3094d7d0ab1c6 100644
--- a/pandas/plotting/_core.py
+++ b/pandas/plotting/_core.py
@@ -1951,7 +1951,7 @@ def plot_series(data, kind='line', ax=None, # Series unique
return_type : {None, 'axes', 'dict', 'both'}, default None
The kind of object to return. The default is ``axes``
'axes' returns the matplotlib axes the boxplot is drawn on;
- 'dict' returns a dictionary whose values are the matplotlib
+ 'dict' returns a dictionary whose values are the matplotlib
Lines of the boxplot;
'both' returns a namedtuple with the axes and dict.
diff --git a/pandas/plotting/_style.py b/pandas/plotting/_style.py
index 145597e52ae14..887202e22b4e0 100644
--- a/pandas/plotting/_style.py
+++ b/pandas/plotting/_style.py
@@ -67,9 +67,9 @@ def _maybe_valid_colors(colors):
except ValueError:
return False
- # check whether the string can be convertable to single color
+ # check whether the string can be convertible to single color
maybe_single_color = _maybe_valid_colors([colors])
- # check whether each character can be convertable to colors
+ # check whether each character can be convertible to colors
maybe_color_cycle = _maybe_valid_colors(list(colors))
if maybe_single_color and maybe_color_cycle and len(colors) > 1:
# Special case for single str 'CN' match and convert to hex
diff --git a/pandas/tests/frame/test_alter_axes.py b/pandas/tests/frame/test_alter_axes.py
index 6e3b7a059fd49..c824f0026af50 100644
--- a/pandas/tests/frame/test_alter_axes.py
+++ b/pandas/tests/frame/test_alter_axes.py
@@ -480,7 +480,7 @@ def test_rename_multiindex(self):
df = DataFrame([(0, 0), (1, 1)], index=index, columns=columns)
#
- # without specifying level -> accross all levels
+ # without specifying level -> across all levels
renamed = df.rename(index={'foo1': 'foo3', 'bar2': 'bar3'},
columns={'fizz1': 'fizz3', 'buzz2': 'buzz3'})
diff --git a/pandas/tests/frame/test_analytics.py b/pandas/tests/frame/test_analytics.py
index 69f1aeddc43e9..b9275fc69e7ff 100644
--- a/pandas/tests/frame/test_analytics.py
+++ b/pandas/tests/frame/test_analytics.py
@@ -1907,7 +1907,7 @@ def test_round_issue(self):
def test_built_in_round(self):
if not compat.PY3:
- pytest.skip("build in round cannot be overriden "
+ pytest.skip("build in round cannot be overridden "
"prior to Python 3")
# GH11763
diff --git a/pandas/tests/frame/test_operators.py b/pandas/tests/frame/test_operators.py
index 430562ce727da..fd1eb23643c2b 100644
--- a/pandas/tests/frame/test_operators.py
+++ b/pandas/tests/frame/test_operators.py
@@ -865,7 +865,7 @@ def test_combineSeries(self):
# 10890
# we no longer allow auto timeseries broadcasting
- # and require explict broadcasting
+ # and require explicit broadcasting
added = self.tsframe.add(ts, axis='index')
for key, col in compat.iteritems(self.tsframe):
diff --git a/pandas/tests/groupby/test_groupby.py b/pandas/tests/groupby/test_groupby.py
index a13d985ab6974..5172efe25d697 100644
--- a/pandas/tests/groupby/test_groupby.py
+++ b/pandas/tests/groupby/test_groupby.py
@@ -303,7 +303,7 @@ def test_with_na_groups(self):
# assert issubclass(agged.dtype.type, np.integer)
- # explicity return a float from my function
+ # explicitly return a float from my function
def f(x):
return float(len(x))
diff --git a/pandas/tests/groupby/test_whitelist.py b/pandas/tests/groupby/test_whitelist.py
index de0deb442e516..3117525d899f6 100644
--- a/pandas/tests/groupby/test_whitelist.py
+++ b/pandas/tests/groupby/test_whitelist.py
@@ -184,7 +184,7 @@ def test_regression_whitelist_methods(
axis, skipna, sort):
# GH6944
# GH 17537
- # explicity test the whitelest methods
+ # explicitly test the whitelest methods
if axis == 0:
frame = raw_frame
diff --git a/pandas/tests/indexes/datetimes/test_astype.py b/pandas/tests/indexes/datetimes/test_astype.py
index 1d72ca609b1d3..4b989eb35e900 100644
--- a/pandas/tests/indexes/datetimes/test_astype.py
+++ b/pandas/tests/indexes/datetimes/test_astype.py
@@ -71,7 +71,7 @@ def test_astype_with_tz(self):
def test_astype_str_compat(self):
# GH 13149, GH 13209
- # verify that we are returing NaT as a string (and not unicode)
+ # verify that we are returning NaT as a string (and not unicode)
idx = DatetimeIndex(['2016-05-16', 'NaT', NaT, np.NaN])
result = idx.astype(str)
diff --git a/pandas/tests/indexes/datetimes/test_tools.py b/pandas/tests/indexes/datetimes/test_tools.py
index c89e3ddbfc5d0..f94a438fcdaa5 100644
--- a/pandas/tests/indexes/datetimes/test_tools.py
+++ b/pandas/tests/indexes/datetimes/test_tools.py
@@ -1464,7 +1464,7 @@ def test_parsers_iso8601(self):
actual = tslib._test_parse_iso8601(date_str)
assert actual == exp
- # seperators must all match - YYYYMM not valid
+ # separators must all match - YYYYMM not valid
invalid_cases = ['2011-01/02', '2011^11^11',
'201401', '201111', '200101',
# mixed separated and unseparated
diff --git a/pandas/tests/indexes/interval/test_interval.py b/pandas/tests/indexes/interval/test_interval.py
index 6fc5526e63e59..3ca4c31b7f059 100644
--- a/pandas/tests/indexes/interval/test_interval.py
+++ b/pandas/tests/indexes/interval/test_interval.py
@@ -1112,7 +1112,7 @@ def test_is_non_overlapping_monotonic(self, closed):
idx = IntervalIndex.from_tuples(tpls[::-1], closed=closed)
assert idx.is_non_overlapping_monotonic is False
- # Should be False for closed='both', overwise True (GH16560)
+ # Should be False for closed='both', otherwise True (GH16560)
if closed == 'both':
idx = IntervalIndex.from_breaks(range(4), closed=closed)
assert idx.is_non_overlapping_monotonic is False
diff --git a/pandas/tests/indexes/test_base.py b/pandas/tests/indexes/test_base.py
index e33fd1e0f4c1e..5109542403b43 100644
--- a/pandas/tests/indexes/test_base.py
+++ b/pandas/tests/indexes/test_base.py
@@ -1067,7 +1067,7 @@ def test_format(self):
# GH 14626
# windows has different precision on datetime.datetime.now (it doesn't
# include us since the default for Timestamp shows these but Index
- # formating does not we are skipping)
+ # formatting does not we are skipping)
now = datetime.now()
if not str(now).endswith("000"):
index = Index([now])
diff --git a/pandas/tests/indexes/test_numeric.py b/pandas/tests/indexes/test_numeric.py
index cbd819fa9cfb7..dcd592345b91c 100644
--- a/pandas/tests/indexes/test_numeric.py
+++ b/pandas/tests/indexes/test_numeric.py
@@ -117,7 +117,7 @@ def test_numeric_compat(self):
def test_explicit_conversions(self):
# GH 8608
- # add/sub are overriden explicity for Float/Int Index
+ # add/sub are overridden explicitly for Float/Int Index
idx = self._holder(np.arange(5, dtype='int64'))
# float conversions
diff --git a/pandas/tests/indexes/test_range.py b/pandas/tests/indexes/test_range.py
index 55c06e8854333..1ebeef072fdc5 100644
--- a/pandas/tests/indexes/test_range.py
+++ b/pandas/tests/indexes/test_range.py
@@ -779,7 +779,7 @@ def test_slice_keep_name(self):
def test_explicit_conversions(self):
# GH 8608
- # add/sub are overriden explicity for Float/Int Index
+ # add/sub are overridden explicitly for Float/Int Index
idx = RangeIndex(5)
# float conversions
diff --git a/pandas/tests/indexing/common.py b/pandas/tests/indexing/common.py
index c5fb2580f0a15..ded16224aedf2 100644
--- a/pandas/tests/indexing/common.py
+++ b/pandas/tests/indexing/common.py
@@ -120,8 +120,8 @@ def get_result(self, obj, method, key, axis):
if isinstance(key, dict):
key = key[axis]
- # use an artifical conversion to map the key as integers to the labels
- # so ix can work for comparisions
+ # use an artificial conversion to map the key as integers to the labels
+ # so ix can work for comparisons
if method == 'indexer':
method = 'ix'
key = obj._get_axis(axis)[key]
@@ -138,7 +138,7 @@ def get_result(self, obj, method, key, axis):
def get_value(self, f, i, values=False):
""" return the value for the location i """
- # check agains values
+ # check against values
if values:
return f.values[i]
@@ -160,7 +160,7 @@ def check_values(self, f, func, values=False):
for i in indicies:
result = getattr(f, func)[i]
- # check agains values
+ # check against values
if values:
expected = f.values[i]
else:
diff --git a/pandas/tests/indexing/test_floats.py b/pandas/tests/indexing/test_floats.py
index 6c5af84f0ce02..d2692c7dc302e 100644
--- a/pandas/tests/indexing/test_floats.py
+++ b/pandas/tests/indexing/test_floats.py
@@ -360,7 +360,7 @@ def test_slice_integer(self):
# same as above, but for Integer based indexes
# these coerce to a like integer
- # oob indiciates if we are out of bounds
+ # oob indicates if we are out of bounds
# of positional indexing
for index, oob in [(tm.makeIntIndex(5), False),
(tm.makeRangeIndex(5), False),
diff --git a/pandas/tests/indexing/test_indexing.py b/pandas/tests/indexing/test_indexing.py
index 0e66c15760653..c66310d10ebdc 100644
--- a/pandas/tests/indexing/test_indexing.py
+++ b/pandas/tests/indexing/test_indexing.py
@@ -362,7 +362,7 @@ def test_multi_nan_indexing(self):
def test_multi_assign(self):
- # GH 3626, an assignement of a sub-df to a df
+ # GH 3626, an assignment of a sub-df to a df
df = DataFrame({'FC': ['a', 'b', 'a', 'b', 'a', 'b'],
'PF': [0, 0, 0, 0, 1, 1],
'col1': lrange(6),
diff --git a/pandas/tests/indexing/test_ix.py b/pandas/tests/indexing/test_ix.py
index 568dd7cec5ecb..3f71e673a4ffe 100644
--- a/pandas/tests/indexing/test_ix.py
+++ b/pandas/tests/indexing/test_ix.py
@@ -235,7 +235,7 @@ def test_ix_assign_column_mixed(self):
tm.assert_frame_equal(df, expected)
# ok, but chained assignments are dangerous
- # if we turn off chained assignement it will work
+ # if we turn off chained assignment it will work
with option_context('chained_assignment', None):
df = DataFrame({'a': lrange(4)})
df['b'] = np.nan
diff --git a/pandas/tests/indexing/test_loc.py b/pandas/tests/indexing/test_loc.py
index 6f0d8b1f29b77..fb5f094f9462b 100644
--- a/pandas/tests/indexing/test_loc.py
+++ b/pandas/tests/indexing/test_loc.py
@@ -17,7 +17,7 @@ class TestLoc(Base):
def test_loc_getitem_dups(self):
# GH 5678
- # repeated gettitems on a dup index returing a ndarray
+ # repeated gettitems on a dup index returning a ndarray
df = DataFrame(
np.random.random_sample((20, 5)),
index=['ABCDE' [x % 5] for x in range(20)])
@@ -385,7 +385,7 @@ def test_loc_general(self):
def test_loc_setitem_consistency(self):
# GH 6149
- # coerce similary for setitem and loc when rows have a null-slice
+ # coerce similarly for setitem and loc when rows have a null-slice
expected = DataFrame({'date': Series(0, index=range(5),
dtype=np.int64),
'val': Series(range(5), dtype=np.int64)})
@@ -588,7 +588,7 @@ def test_loc_non_unique(self):
# non-unique indexer with loc slice
# https://groups.google.com/forum/?fromgroups#!topic/pydata/zTm2No0crYs
- # these are going to raise becuase the we are non monotonic
+ # these are going to raise because the we are non monotonic
df = DataFrame({'A': [1, 2, 3, 4, 5, 6],
'B': [3, 4, 5, 6, 7, 8]}, index=[0, 1, 0, 1, 2, 3])
pytest.raises(KeyError, df.loc.__getitem__,
diff --git a/pandas/tests/io/data/banklist.html b/pandas/tests/io/data/banklist.html
index 8ec1561f8c394..cbcce5a2d49ff 100644
--- a/pandas/tests/io/data/banklist.html
+++ b/pandas/tests/io/data/banklist.html
@@ -7,7 +7,7 @@
<meta charset="UTF-8">
<!-- Unicode character encoding -->
<meta http-equiv="X-UA-Compatible" content="IE=edge">
-<!-- Turns off IE Compatiblity Mode -->
+<!-- Turns off IE Compatibility Mode -->
<meta name="viewport" content="width=device-width, initial-scale=1, maximum-scale=1">
<!-- Makes it so phones don't auto zoom out. -->
<meta name="author" content="DRR">
@@ -4849,7 +4849,7 @@ <h1 class="page_title">Failed Bank List</h1>
<ul>
<li><a href="/about/freedom/" title="Freedom of Information Act (FOIA) Service Center">Freedom of Information Act (FOIA) Service Center</a></li>
<li><a href="/open/" title="FDIC Open Government Webpage">FDIC Open Government Webpage</a></li>
- <li><a href="/about/diversity/nofear/" title="No FEAR Act Data">No FEAR Act Data</a></li>
+ <li><a href="/about/diversity/nofear/" title="No FEAR Act Data">No FEAR Act Data</a></li>
</ul>
</div>
<div id="responsive_footer-small">
diff --git a/pandas/tests/io/data/macau.html b/pandas/tests/io/data/macau.html
index be62b3221518d..cfd1a0702460a 100644
--- a/pandas/tests/io/data/macau.html
+++ b/pandas/tests/io/data/macau.html
@@ -476,7 +476,7 @@ <h4>這個頁面上的內容需要較新版本的 Adobe Flash Player。</h4>
toggleclass: ["", "selected"], //Two CSS classes to be applied to the header when it's collapsed and expanded, respectively ["class1", "class2"]
togglehtml: ["", "", ""], //Additional HTML added to the header when it's collapsed and expanded, respectively ["position", "html1", "html2"] (see docs)
animatespeed: "normal", //speed of animation: integer in milliseconds (ie: 200), or keywords "fast", "normal", or "slow"
- oninit:function(headers, expandedindices){ //custom code to run when headers have initalized
+ oninit:function(headers, expandedindices){ //custom code to run when headers have initialized
//do nothing
},
onopenclose:function(header, index, state, isuseractivated){ //custom code to run whenever a header is opened or closed
diff --git a/pandas/tests/io/data/spam.html b/pandas/tests/io/data/spam.html
index 935b39f6d6011..e4fadab6eafd2 100644
--- a/pandas/tests/io/data/spam.html
+++ b/pandas/tests/io/data/spam.html
@@ -208,7 +208,7 @@ <h1>Nutrient data for 07908, Luncheon meat, pork with ham, minced, canned, inclu
<table>
<thead>
- <tr><td colspan="6" style="vertical-align:middle;text-align:center;height:2em;" class="buttons"><input type="submit" name="_action_show" value="Apply Changes" class="calc" title="Click to recalculate measures" id="1732" /><a href="/ndb/help/contextHelp/measures" onclick="jQuery.ajax({type:'POST', url:'/ndb/help/contextHelp/measures',success:function(data,textStatus){jQuery('#helpDiv').html(data);},error:function(XMLHttpRequest,textStatus,errorThrown){},complete:function(XMLHttpRequest,textStatus){GRAILSUI.measuresHelpDialog.show();}});return false;" controller="help" action="contextHelp" id="measures"><img title="Click for more information on calculating household measures" src="/ndb/static/images/skin/help.png" alt="Help" border="0" style="vertical-align:middle"/></a></span></td></tr>
+ <tr><td colspan="6" style="vertical-align:middle;text-align:center;height:2em;" class="buttons"><input type="submit" name="_action_show" value="Apply Changes" class="calc" title="Click to recalculate measures" id="1732" /><a href="/ndb/help/contextHelp/measures" onclick="jQuery.ajax({type:'POST', url:'/ndb/help/contextHelp/measures',success:function(data,textStatus){jQuery('#helpDiv').html(data);},error:function(XMLHttpRequest,textStatus,errorThrown){},complete:function(XMLHttpRequest,textStatus){GRAILSUI.measuresHelpDialog.show();}});return false;" controller="help" action="contextHelp" id="measures"><img title="Click for more information on calculating household measures" src="/ndb/static/images/skin/help.png" alt="Help" border="0" style="vertical-align:middle"/></a></span></td></tr>
<th style="vertical-align:middle">Nutrient</th>
<th style="vertical-align:middle" >Unit</th>
<th style="vertical-align:middle"><input type="text" name="Qv" style="width:30px;text-align:right;border-style:inset;height:15px" maxlength="5" value="1" id="Qv" /><br/>Value per 100.0g</th>
diff --git a/pandas/tests/io/formats/test_format.py b/pandas/tests/io/formats/test_format.py
index 1fefec6035a20..23b42b612dace 100644
--- a/pandas/tests/io/formats/test_format.py
+++ b/pandas/tests/io/formats/test_format.py
@@ -369,7 +369,7 @@ def test_str_max_colwidth(self):
def test_auto_detect(self):
term_width, term_height = get_terminal_size()
- fac = 1.05 # Arbitrary large factor to exceed term widht
+ fac = 1.05 # Arbitrary large factor to exceed term width
cols = range(int(term_width * fac))
index = range(10)
df = DataFrame(index=index, columns=cols)
diff --git a/pandas/tests/io/formats/test_style.py b/pandas/tests/io/formats/test_style.py
index 4b0ca872da326..bedb11d4fc4ae 100644
--- a/pandas/tests/io/formats/test_style.py
+++ b/pandas/tests/io/formats/test_style.py
@@ -650,7 +650,7 @@ def test_highlight_max(self):
(0, 0): [''], (1, 0): ['']}
assert result == expected
- # separate since we cant negate the strs
+ # separate since we can't negate the strs
df['C'] = ['a', 'b']
result = df.style.highlight_max()._compute().ctx
expected = {(1, 1): ['background-color: yellow']}
diff --git a/pandas/tests/io/msgpack/test_extension.py b/pandas/tests/io/msgpack/test_extension.py
index 26a611bea224c..2ee72c8a55cb4 100644
--- a/pandas/tests/io/msgpack/test_extension.py
+++ b/pandas/tests/io/msgpack/test_extension.py
@@ -46,7 +46,7 @@ def default(obj):
typecode = 123 # application specific typecode
data = tobytes(obj)
return ExtType(typecode, data)
- raise TypeError("Unknwon type object %r" % (obj, ))
+ raise TypeError("Unknown type object %r" % (obj, ))
def ext_hook(code, data):
print('ext_hook called', code, data)
diff --git a/pandas/tests/io/msgpack/test_seq.py b/pandas/tests/io/msgpack/test_seq.py
index 5f203e8997ccb..06e9872a22777 100644
--- a/pandas/tests/io/msgpack/test_seq.py
+++ b/pandas/tests/io/msgpack/test_seq.py
@@ -25,7 +25,7 @@ def test_exceeding_unpacker_read_size():
# double free or corruption (!prev)
# 40 ok for read_size=1024, while 50 introduces errors
- # 7000 ok for read_size=1024*1024, while 8000 leads to glibc detected ***
+ # 7000 ok for read_size=1024*1024, while 8000 leads to glibc detected ***
# python: double free or corruption (!prev):
for idx in range(NUMBER_OF_STRINGS):
diff --git a/pandas/tests/io/parser/parse_dates.py b/pandas/tests/io/parser/parse_dates.py
index 7ff2ac9ff1305..b7d0dd1a3484f 100644
--- a/pandas/tests/io/parser/parse_dates.py
+++ b/pandas/tests/io/parser/parse_dates.py
@@ -217,8 +217,8 @@ def test_nat_parse(self):
tm.assert_series_equal(expected, result.dtypes)
# test with NaT for the nan_rep
- # we don't have a method to specif the Datetime na_rep (it defaults
- # to '')
+ # we don't have a method to specify the Datetime na_rep
+ # (it defaults to '')
df.to_csv(path)
result = self.read_csv(path, index_col=0, parse_dates=['B'])
tm.assert_frame_equal(result, df)
diff --git a/pandas/tests/io/test_excel.py b/pandas/tests/io/test_excel.py
index 168144d78b3be..3263f71dea3c3 100644
--- a/pandas/tests/io/test_excel.py
+++ b/pandas/tests/io/test_excel.py
@@ -861,8 +861,8 @@ def test_excel_multindex_roundtrip(self):
if (c_idx_levels == 1 and c_idx_names):
continue
- # empty name case current read in as unamed levels,
- # not Nones
+ # empty name case current read in as unnamed
+ # levels, not Nones
check_names = True
if not r_idx_names and r_idx_levels > 1:
check_names = False
diff --git a/pandas/tests/io/test_packers.py b/pandas/tests/io/test_packers.py
index b9d66426c9dcb..c343e0105eb4f 100644
--- a/pandas/tests/io/test_packers.py
+++ b/pandas/tests/io/test_packers.py
@@ -205,7 +205,7 @@ def test_list_numpy_float(self):
def test_list_numpy_float_complex(self):
if not hasattr(np, 'complex128'):
- pytest.skip('numpy cant handle complex128')
+ pytest.skip('numpy can not handle complex128')
x = [np.float32(np.random.rand()) for i in range(5)] + \
[np.complex128(np.random.rand() + 1j * np.random.rand())
diff --git a/pandas/tests/io/test_pickle.py b/pandas/tests/io/test_pickle.py
index d5bcf72488d09..5d2ba8e4fa712 100644
--- a/pandas/tests/io/test_pickle.py
+++ b/pandas/tests/io/test_pickle.py
@@ -38,7 +38,7 @@ def current_pickle_data():
# ---------------------
-# comparision functions
+# comparison functions
# ---------------------
def compare_element(result, expected, typ, version=None):
if isinstance(expected, Index):
diff --git a/pandas/tests/io/test_pytables.py b/pandas/tests/io/test_pytables.py
index 305c1ebcedc6f..b40350ada546c 100644
--- a/pandas/tests/io/test_pytables.py
+++ b/pandas/tests/io/test_pytables.py
@@ -903,7 +903,7 @@ def test_append(self):
'items', 'major_axis', 'minor_axis'])
assert_panel4d_equal(store['p4d'], p4d)
- # test using differnt number of items on each axis
+ # test using different number of items on each axis
p4d2 = p4d.copy()
p4d2['l4'] = p4d['l1']
p4d2['l5'] = p4d['l1']
@@ -1300,11 +1300,11 @@ def test_append_with_different_block_ordering(self):
df['int16'] = Series([1] * len(df), dtype='int16')
store.append('df', df)
- # store additonal fields in different blocks
+ # store additional fields in different blocks
df['int16_2'] = Series([1] * len(df), dtype='int16')
pytest.raises(ValueError, store.append, 'df', df)
- # store multile additonal fields in different blocks
+ # store multile additional fields in different blocks
df['float_3'] = Series([1.] * len(df), dtype='float64')
pytest.raises(ValueError, store.append, 'df', df)
@@ -1330,7 +1330,7 @@ def check_indexers(key, indexers):
assert_panel4d_equal(store.select('p4d'), p4d)
check_indexers('p4d', indexers)
- # same as above, but try to append with differnt axes
+ # same as above, but try to append with different axes
_maybe_remove(store, 'p4d')
store.append('p4d', p4d.iloc[:, :, :10, :], axes=indexers)
store.append('p4d', p4d.iloc[:, :, 10:, :], axes=[
@@ -2083,7 +2083,7 @@ def test_append_raise(self):
assert df.dtypes['invalid'] == np.object_
pytest.raises(TypeError, store.append, 'df', df)
- # directy ndarray
+ # directly ndarray
pytest.raises(TypeError, store.append, 'df', np.arange(10))
# series directly
@@ -3066,7 +3066,7 @@ def test_select_with_dups(self):
expected = df.loc[:, ['A']]
assert_frame_equal(result, expected)
- # dups accross dtypes
+ # dups across dtypes
df = concat([DataFrame(np.random.randn(10, 4),
columns=['A', 'A', 'B', 'B']),
DataFrame(np.random.randint(0, 10, size=20)
@@ -5410,7 +5410,7 @@ def _compare_with_tz(self, a, b):
b_e = b.loc[i, c]
if not (a_e == b_e and a_e.tz == b_e.tz):
raise AssertionError(
- "invalid tz comparsion [%s] [%s]" % (a_e, b_e))
+ "invalid tz comparison [%s] [%s]" % (a_e, b_e))
def test_append_with_timezones_dateutil(self):
diff --git a/pandas/tests/plotting/test_frame.py b/pandas/tests/plotting/test_frame.py
index d61b0a40380f3..3d25b0b51e052 100644
--- a/pandas/tests/plotting/test_frame.py
+++ b/pandas/tests/plotting/test_frame.py
@@ -675,7 +675,7 @@ def test_negative_log(self):
def _compare_stacked_y_cood(self, normal_lines, stacked_lines):
base = np.zeros(len(normal_lines[0].get_data()[1]))
for nl, sl in zip(normal_lines, stacked_lines):
- base += nl.get_data()[1] # get y coodinates
+ base += nl.get_data()[1] # get y coordinates
sy = sl.get_data()[1]
tm.assert_numpy_array_equal(base, sy)
diff --git a/pandas/tests/plotting/test_misc.py b/pandas/tests/plotting/test_misc.py
index bb590d5232b62..60ed280bc050e 100644
--- a/pandas/tests/plotting/test_misc.py
+++ b/pandas/tests/plotting/test_misc.py
@@ -219,7 +219,7 @@ def test_parallel_coordinates_with_sorted_labels(self):
prev_next_tupels = zip([i for i in ordered_color_label_tuples[0:-1]],
[i for i in ordered_color_label_tuples[1:]])
for prev, nxt in prev_next_tupels:
- # lables and colors are ordered strictly increasing
+ # labels and colors are ordered strictly increasing
assert prev[1] < nxt[1] and prev[0] < nxt[0]
@pytest.mark.slow
diff --git a/pandas/tests/reshape/test_reshape.py b/pandas/tests/reshape/test_reshape.py
index 0312af12e0715..22925cceb30d1 100644
--- a/pandas/tests/reshape/test_reshape.py
+++ b/pandas/tests/reshape/test_reshape.py
@@ -341,7 +341,7 @@ def test_basic_drop_first_one_level(self, sparse):
assert_frame_equal(result, expected)
def test_basic_drop_first_NA(self, sparse):
- # Test NA hadling together with drop_first
+ # Test NA handling together with drop_first
s_NA = ['a', 'b', np.nan]
res = get_dummies(s_NA, drop_first=True, sparse=sparse)
exp = DataFrame({'b': [0, 1, 0]}, dtype=np.uint8)
diff --git a/pandas/tests/series/test_analytics.py b/pandas/tests/series/test_analytics.py
index 14bf194ba5ee4..f2b7c20b774b0 100644
--- a/pandas/tests/series/test_analytics.py
+++ b/pandas/tests/series/test_analytics.py
@@ -704,7 +704,7 @@ def test_numpy_round(self):
def test_built_in_round(self):
if not compat.PY3:
pytest.skip(
- 'build in round cannot be overriden prior to Python 3')
+ 'build in round cannot be overridden prior to Python 3')
s = Series([1.123, 2.123, 3.123], index=lrange(3))
result = round(s)
@@ -1338,7 +1338,7 @@ def test_numpy_argmin_deprecated(self):
with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
# The deprecation of Series.argmin also causes a deprecation
# warning when calling np.argmin. This behavior is temporary
- # until the implemention of Series.argmin is corrected.
+ # until the implementation of Series.argmin is corrected.
result = np.argmin(s)
assert result == 1
@@ -1408,7 +1408,7 @@ def test_numpy_argmax_deprecated(self):
with tm.assert_produces_warning(FutureWarning, check_stacklevel=False):
# The deprecation of Series.argmax also causes a deprecation
# warning when calling np.argmax. This behavior is temporary
- # until the implemention of Series.argmax is corrected.
+ # until the implementation of Series.argmax is corrected.
result = np.argmax(s)
assert result == 10
diff --git a/pandas/tests/series/test_api.py b/pandas/tests/series/test_api.py
index a2838f803421c..8ae7feab451f9 100644
--- a/pandas/tests/series/test_api.py
+++ b/pandas/tests/series/test_api.py
@@ -216,7 +216,7 @@ def test_tab_completion(self):
assert 'dt' not in dir(s)
assert 'cat' not in dir(s)
- # similiarly for .dt
+ # similarly for .dt
s = Series(date_range('1/1/2015', periods=5))
assert 'dt' in dir(s)
assert 'str' not in dir(s)
diff --git a/pandas/tests/series/test_constructors.py b/pandas/tests/series/test_constructors.py
index 08416fe34efcc..5de5f1f0584f4 100644
--- a/pandas/tests/series/test_constructors.py
+++ b/pandas/tests/series/test_constructors.py
@@ -487,7 +487,7 @@ def test_constructor_dtype_nocast(self):
def test_constructor_datelike_coercion(self):
# GH 9477
- # incorrectly infering on dateimelike looking when object dtype is
+ # incorrectly inferring on dateimelike looking when object dtype is
# specified
s = Series([Timestamp('20130101'), 'NOV'], dtype=object)
assert s.iloc[0] == Timestamp('20130101')
diff --git a/pandas/tests/series/test_indexing.py b/pandas/tests/series/test_indexing.py
index 00fa980d9a139..0503a7b30e91c 100644
--- a/pandas/tests/series/test_indexing.py
+++ b/pandas/tests/series/test_indexing.py
@@ -1616,7 +1616,7 @@ def test_where_numeric_with_string(self):
def test_setitem_boolean(self):
mask = self.series > self.series.median()
- # similiar indexed series
+ # similar indexed series
result = self.series.copy()
result[mask] = self.series * 2
expected = self.series * 2
@@ -1668,7 +1668,7 @@ def test_setitem_na(self):
s[::2] = np.nan
assert_series_equal(s, expected)
- # get's coerced to float, right?
+ # gets coerced to float, right?
expected = Series([np.nan, 1, np.nan, 0])
s = Series([True, True, False, False])
s[::2] = np.nan
@@ -2113,7 +2113,7 @@ def test_reindex_pad(self):
result = s.reindex(new_index, method='ffill')
assert_series_equal(result, expected)
- # inferrence of new dtype
+ # inference of new dtype
s = Series([True, False, False, True], index=list('abcd'))
new_index = 'agc'
result = s.reindex(list(new_index)).ffill()
diff --git a/pandas/tests/series/test_operators.py b/pandas/tests/series/test_operators.py
index 433e3cf440cbd..ce4e388bc6f39 100644
--- a/pandas/tests/series/test_operators.py
+++ b/pandas/tests/series/test_operators.py
@@ -1359,7 +1359,7 @@ def check(series, other):
expecteds = divmod(series.values, np.asarray(other_np))
for result, expected in zip(results, expecteds):
- # check the values, name, and index separatly
+ # check the values, name, and index separately
assert_almost_equal(np.asarray(result), expected)
assert result.name == series.name
@@ -1449,7 +1449,7 @@ def timedelta64(*args):
assert_series_equal(lhs, rhs)
except:
raise AssertionError(
- "invalid comparsion [op->{0},d->{1},h->{2},m->{3},"
+ "invalid comparison [op->{0},d->{1},h->{2},m->{3},"
"s->{4},us->{5}]\n{6}\n{7}\n".format(op, d, h, m, s,
us, lhs, rhs))
diff --git a/pandas/tests/test_base.py b/pandas/tests/test_base.py
index df76390d7ce7a..cb905d8186ea9 100644
--- a/pandas/tests/test_base.py
+++ b/pandas/tests/test_base.py
@@ -116,7 +116,8 @@ def setup_method(self, method):
def test_invalida_delgation(self):
# these show that in order for the delegation to work
- # the _delegate_* methods need to be overriden to not raise a TypeError
+ # the _delegate_* methods need to be overridden to not raise
+ # a TypeError
self.Delegate._add_delegate_accessors(
delegate=self.Delegator,
diff --git a/pandas/tests/test_multilevel.py b/pandas/tests/test_multilevel.py
index 592b069ef8bac..86d9a9fa91e47 100644
--- a/pandas/tests/test_multilevel.py
+++ b/pandas/tests/test_multilevel.py
@@ -2383,7 +2383,7 @@ def test_iloc_mi(self):
class TestSorted(Base):
- """ everthing you wanted to test about sorting """
+ """ everything you wanted to test about sorting """
def test_sort_index_preserve_levels(self):
result = self.frame.sort_index()
diff --git a/pandas/tests/test_panel.py b/pandas/tests/test_panel.py
index d772dba25868e..770560134d8d6 100644
--- a/pandas/tests/test_panel.py
+++ b/pandas/tests/test_panel.py
@@ -2582,19 +2582,19 @@ def test_truncate(self):
trunced = self.panel.truncate(start, end).to_panel()
expected = self.panel.to_panel()['ItemA'].truncate(start, end)
- # TODO trucate drops index.names
+ # TODO truncate drops index.names
assert_frame_equal(trunced['ItemA'], expected, check_names=False)
trunced = self.panel.truncate(before=start).to_panel()
expected = self.panel.to_panel()['ItemA'].truncate(before=start)
- # TODO trucate drops index.names
+ # TODO truncate drops index.names
assert_frame_equal(trunced['ItemA'], expected, check_names=False)
trunced = self.panel.truncate(after=end).to_panel()
expected = self.panel.to_panel()['ItemA'].truncate(after=end)
- # TODO trucate drops index.names
+ # TODO truncate drops index.names
assert_frame_equal(trunced['ItemA'], expected, check_names=False)
# truncate on dates that aren't in there
diff --git a/pandas/tests/test_sorting.py b/pandas/tests/test_sorting.py
index 57bd5e7b62fdf..d0350ba252329 100644
--- a/pandas/tests/test_sorting.py
+++ b/pandas/tests/test_sorting.py
@@ -82,7 +82,7 @@ def test_int64_overflow_moar(self):
# verify this is testing what it is supposed to test!
assert is_int64_overflow_possible(gr.grouper.shape)
- # mannually compute groupings
+ # manually compute groupings
jim, joe = defaultdict(list), defaultdict(list)
for key, a, b in zip(map(tuple, arr), df['jim'], df['joe']):
jim[key].append(a)
diff --git a/pandas/tests/test_window.py b/pandas/tests/test_window.py
index ccffc554e00c7..6f9e872526d0a 100644
--- a/pandas/tests/test_window.py
+++ b/pandas/tests/test_window.py
@@ -3417,7 +3417,7 @@ def test_frame_on(self):
# test as a frame
# we should be ignoring the 'on' as an aggregation column
- # note that the expected is setting, computing, and reseting
+ # note that the expected is setting, computing, and resetting
# so the columns need to be switched compared
# to the actual result where they are ordered as in the
# original
@@ -3815,7 +3815,7 @@ def test_ragged_apply(self):
def test_all(self):
- # simple comparision of integer vs time-based windowing
+ # simple comparison of integer vs time-based windowing
df = self.regular * 2
er = df.rolling(window=1)
r = df.rolling(window='1s')
@@ -3837,7 +3837,7 @@ def test_all(self):
def test_all2(self):
- # more sophisticated comparision of integer vs.
+ # more sophisticated comparison of integer vs.
# time-based windowing
df = DataFrame({'B': np.arange(50)},
index=pd.date_range('20130101',
diff --git a/pandas/tests/tseries/offsets/test_offsets.py b/pandas/tests/tseries/offsets/test_offsets.py
index edabf4a7ccc99..e1a6463e7c351 100644
--- a/pandas/tests/tseries/offsets/test_offsets.py
+++ b/pandas/tests/tseries/offsets/test_offsets.py
@@ -3016,7 +3016,7 @@ def _test_offset(self, offset_name, offset_n, tstart, expected_utc_offset):
t.minute == tstart.minute and
t.second == tstart.second)
elif offset_name in self.valid_date_offsets_singular:
- # expect the signular offset value to match between tstart and t
+ # expect the singular offset value to match between tstart and t
datepart_offset = getattr(t, offset_name
if offset_name != 'weekday' else
'dayofweek')
@@ -3063,7 +3063,7 @@ def test_springforward_plural(self):
expected_utc_offset=hrs_post)
def test_fallback_singular(self):
- # in the case of signular offsets, we dont neccesarily know which utc
+ # in the case of singular offsets, we don't necessarily know which utc
# offset the new Timestamp will wind up in (the tz for 1 month may be
# different from 1 second) so we don't specify an expected_utc_offset
for tz, utc_offsets in self.timezone_utc_offsets.items():
diff --git a/pandas/tests/tseries/test_timezones.py b/pandas/tests/tseries/test_timezones.py
index 9530cd5ac3f43..b3813d03532fb 100644
--- a/pandas/tests/tseries/test_timezones.py
+++ b/pandas/tests/tseries/test_timezones.py
@@ -577,7 +577,7 @@ def test_ambiguous_nat(self):
'11/06/2011 03:00']
di_test = DatetimeIndex(times, tz='US/Eastern')
- # left dtype is datetime64[ns, US/Eastern]
+ # left dtype is datetime64[ns, US/Eastern]
# right is datetime64[ns, tzfile('/usr/share/zoneinfo/US/Eastern')]
tm.assert_numpy_array_equal(di_test.values, localized.values)
diff --git a/pandas/tseries/holiday.py b/pandas/tseries/holiday.py
index 0e6cbea21493c..4e874eac9e6c6 100644
--- a/pandas/tseries/holiday.py
+++ b/pandas/tseries/holiday.py
@@ -133,7 +133,7 @@ def __init__(self, name, year=None, month=None, day=None, offset=None,
Name of the holiday , defaults to class name
offset : array of pandas.tseries.offsets or
class from pandas.tseries.offsets
- computes offset from date
+ computes offset from date
observance: function
computes when holiday is given a pandas Timestamp
days_of_week:
diff --git a/pandas/util/_validators.py b/pandas/util/_validators.py
index 728db6af5558b..b30ffc7416f92 100644
--- a/pandas/util/_validators.py
+++ b/pandas/util/_validators.py
@@ -40,7 +40,7 @@ def _check_for_default_values(fname, arg_val_dict, compat_args):
"""
for key in arg_val_dict:
# try checking equality directly with '=' operator,
- # as comparison may have been overriden for the left
+ # as comparison may have been overridden for the left
# hand object
try:
v1 = arg_val_dict[key]
@@ -292,7 +292,7 @@ def validate_axis_style_args(data, args, kwargs, arg_name, method_name):
out[ax] = v
# All user-provided kwargs have been handled now.
- # Now we supplement with positional arguments, emmitting warnings
+ # Now we supplement with positional arguments, emitting warnings
# when there's ambiguity and raising when there's conflicts
if len(args) == 0:
@@ -307,7 +307,7 @@ def validate_axis_style_args(data, args, kwargs, arg_name, method_name):
"or 'columns'")
raise TypeError(msg)
- msg = ("Intepreting call\n\t'.{method_name}(a, b)' as "
+ msg = ("Interpreting call\n\t'.{method_name}(a, b)' as "
"\n\t'.{method_name}(index=a, columns=b)'.\nUse named "
"arguments to remove any ambiguity. In the future, using "
"positional arguments for 'index' or 'columns' will raise "
diff --git a/pandas/util/testing.py b/pandas/util/testing.py
index 8acf16536f1de..8dc0aa1e85ef4 100644
--- a/pandas/util/testing.py
+++ b/pandas/util/testing.py
@@ -107,7 +107,7 @@ def round_trip_pickle(obj, path=None):
def round_trip_pathlib(writer, reader, path=None):
"""
- Write an object to file specifed by a pathlib.Path and read it back
+ Write an object to file specified by a pathlib.Path and read it back
Parameters
----------
@@ -136,7 +136,7 @@ def round_trip_pathlib(writer, reader, path=None):
def round_trip_localpath(writer, reader, path=None):
"""
- Write an object to file specifed by a py.path LocalPath and read it back
+ Write an object to file specified by a py.path LocalPath and read it back
Parameters
----------
@@ -1784,8 +1784,8 @@ def makeCustomDataframe(nrows, ncols, c_idx_names=True, r_idx_names=True,
"""
nrows, ncols - number of data rows/cols
c_idx_names, idx_names - False/True/list of strings, yields No names ,
- default names or uses the provided names for the levels of the
- corresponding index. You can provide a single string when
+ default names or uses the provided names for the levels of the
+ corresponding index. You can provide a single string when
c_idx_nlevels ==1.
c_idx_nlevels - number of levels in columns index. > 1 will yield MultiIndex
r_idx_nlevels - number of levels in rows index. > 1 will yield MultiIndex
@@ -2081,7 +2081,7 @@ def network(t, url="http://www.google.com",
_skip_on_messages: iterable of string
any exception e for which one of the strings is
a substring of str(e) will be skipped with an appropriate
- message. Intended to supress errors where an errno isn't available.
+ message. Intended to suppress errors where an errno isn't available.
Notes
-----
diff --git a/scripts/find_commits_touching_func.py b/scripts/find_commits_touching_func.py
index 4f3b519775c39..0dd609417d7ba 100755
--- a/scripts/find_commits_touching_func.py
+++ b/scripts/find_commits_touching_func.py
@@ -26,21 +26,21 @@
import argparse
desc = """
-Find all commits touching a sepcified function across the codebase.
+Find all commits touching a specified function across the codebase.
""".strip()
argparser = argparse.ArgumentParser(description=desc)
argparser.add_argument('funcname', metavar='FUNCNAME',
help='Name of function/method to search for changes on.')
argparser.add_argument('-f', '--file-masks', metavar='f_re(,f_re)*',
default=["\.py.?$"],
- help='comma seperated list of regexes to match filenames against\n'+
+ help='comma separated list of regexes to match filenames against\n'+
'defaults all .py? files')
argparser.add_argument('-d', '--dir-masks', metavar='d_re(,d_re)*',
default=[],
- help='comma seperated list of regexes to match base path against')
+ help='comma separated list of regexes to match base path against')
argparser.add_argument('-p', '--path-masks', metavar='p_re(,p_re)*',
default=[],
- help='comma seperated list of regexes to match full file path against')
+ help='comma separated list of regexes to match full file path against')
argparser.add_argument('-y', '--saw-the-warning',
action='store_true',default=False,
help='must specify this to run, acknowledge you realize this will erase untracked files')
diff --git a/scripts/find_undoc_args.py b/scripts/find_undoc_args.py
index 49273bacccf98..32b23a67b187f 100755
--- a/scripts/find_undoc_args.py
+++ b/scripts/find_undoc_args.py
@@ -19,7 +19,7 @@
parser.add_argument('-m', '--module', metavar='MODULE', type=str,required=True,
help='name of package to import and examine',action='store')
parser.add_argument('-G', '--github_repo', metavar='REPO', type=str,required=False,
- help='github project where the the code lives, e.g. "pandas-dev/pandas"',
+ help='github project where the code lives, e.g. "pandas-dev/pandas"',
default=None,action='store')
args = parser.parse_args()
diff --git a/setup.py b/setup.py
index 515e1660fa6de..443f3eba69b4d 100755
--- a/setup.py
+++ b/setup.py
@@ -109,7 +109,7 @@ def build_extensions(self):
# generate template output
if cython:
for pxifile in _pxifiles:
- # build pxifiles first, template extention must be .pxi.in
+ # build pxifiles first, template extension must be .pxi.in
assert pxifile.endswith('.pxi.in')
outfile = pxifile[:-3]
| Both user and non-user facing. Found via `codespell -q 3 -I ../pandas-whitelist.txt` whereby the whitelist consisted of:
```
behaviour
indicies
initialise
initialised
initialising
resetted
thru
writeable
``` | https://api.github.com/repos/pandas-dev/pandas/pulls/18966 | 2017-12-28T10:49:32Z | 2017-12-30T16:15:09Z | 2017-12-30T16:15:09Z | 2017-12-30T17:11:53Z |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.