text string | size int64 | token_count int64 |
|---|---|---|
import unittest
from licma.babel_fish.babel_fish import BabelFish
from licma.rules.python.pyrules_m2crypto.pyrule1_m2crypto import PyRule1M2Crypto
from licma.rules.python.pyrules_m2crypto.pyrule2_m2crypto import PyRule2M2Crypto
from licma.rules.python.pyrules_m2crypto.pyrule3_m2crypto import PyRule3M2Crypto
from licma.rules.python.pyrules_m2crypto.pyrule4_m2crypto import PyRule4M2Crypto
from licma.rules.python.pyrules_m2crypto.pyrule5_m2crypto import PyRule5M2Crypto
from licma.tests.test_utilities import UnitTestBase
class PyRuleM2CryptoTestOOP(UnitTestBase):
babel_fish = BabelFish()
def test_pyrule1_m2crypto_oop(self):
self.execute_test(PyRule1M2Crypto(),
"../resources/test_cases/python/M2Crypto/TestRule1c.py",
"../resources/test_cases/python/M2Crypto/TestRule1cExpectedMisuses.csv")
def test_pyrule2_m2crypto_oop(self):
self.execute_test(PyRule2M2Crypto(),
"../resources/test_cases/python/M2Crypto/TestRule2c.py",
"../resources/test_cases/python/M2Crypto/TestRule2cExpectedMisuses.csv")
def test_pyrule3_m2crypto_oop(self):
self.execute_test(PyRule3M2Crypto(),
"../resources/test_cases/python/M2Crypto/TestRule3c.py",
"../resources/test_cases/python/M2Crypto/TestRule3cExpectedMisuses.csv")
def test_pyrule4_m2crypto_oop(self):
self.execute_test(PyRule4M2Crypto(),
"../resources/test_cases/python/M2Crypto/TestRule4c.py",
"../resources/test_cases/python/M2Crypto/TestRule4cExpectedMisuses.csv")
def test_pyrule5_m2crypto_oop(self):
self.execute_test(PyRule5M2Crypto(),
"../resources/test_cases/python/M2Crypto/TestRule5c.py",
"../resources/test_cases/python/M2Crypto/TestRule5cExpectedMisuses.csv")
if __name__ == '__main__':
unittest.main()
| 1,992 | 703 |
from operator import add
import numpy as np
from pyspark.sql.types import StructField, StructType, IntegerType
from scipy.spatial.distance import euclidean
import sklearn.cluster as skc
from sparkdq.conf.Context import Context
from sparkdq.models.CommonUtils import DEFAULT_CLUSTER_COL, DEFAULT_INDEX_COL
from sparkdq.models.dbscan.ClusterAggregator import ClusterAggregator
from sparkdq.models.dbscan.KDPartitioner import KDPartitioner
class DBSCAN:
def __init__(self, eps=0.5, min_pts=5, dist_type="euclidean", max_partitions=5, prediction_col=DEFAULT_CLUSTER_COL):
self._eps = eps
self._min_pts = min_pts
self._dist_type = dist_type
self._max_partitions = max_partitions
self._prediction_col = prediction_col
def set_params(self, eps=0.5, min_pts=5, dist_type="euclidean", max_partitions=5,
prediction_col=DEFAULT_CLUSTER_COL):
self._eps = eps
self._min_pts = min_pts
self._dist_type = dist_type
self._max_partitions = max_partitions
self._prediction_col = prediction_col
def transform(self, data, columns, index_col=DEFAULT_INDEX_COL):
total_columns = [index_col] + columns
index_type = data.schema[index_col]
rdd = data.select(*total_columns).rdd.map(lambda row: (row[0], np.array(row[1:])))
partitioner = KDPartitioner(rdd, max_partitions=self._max_partitions)
bounding_boxes = partitioner.get_bounding_boxes()
expanded_boxes = {}
# create neighbors
neighbors = {}
new_data = rdd.context.emptyRDD()
for label, box in bounding_boxes.items():
expanded_box = box.expand(2 * self._eps)
expanded_boxes[label] = expanded_box
neighbors[label] = rdd.filter(lambda row: expanded_box.contains(row[1])) \
.map(lambda row: ((row[0], label), row[1]))
new_data = new_data.union(neighbors[label])
rdd = new_data
rdd = rdd.map(lambda row: (row[0][1], (row[0][0], row[1])))\
.partitionBy(len(partitioner.get_partitions()))\
.map(lambda row: ((row[1][0], row[0]), row[1][1]))
if self._dist_type == "euclidean":
params = {"eps": self._eps, "min_samples": self._min_pts, "metric": euclidean}
else:
raise Exception("unsupported metric type {}".format(self._dist_type))
rdd = rdd.mapPartitions(lambda iterable: dbscan_partition(iterable, params))
# remap cluster ids
labeled_points = rdd.groupByKey()
labeled_points.cache()
mapper = labeled_points.aggregate(ClusterAggregator(), add, add)
bc_forward_mapper = rdd.context.broadcast(mapper.forward)
rdd = labeled_points.map(lambda x: map_cluster_id(x, bc_forward_mapper)).sortByKey()
# convert rdd to df
tmp_schema = StructType([
index_type,
StructField(DEFAULT_CLUSTER_COL, IntegerType(), False)
])
tmp_df = Context().spark.createDataFrame(rdd, tmp_schema)
return data.join(tmp_df, on=index_col, how="inner")
def dbscan_partition(iterable, params):
"""
Perform a DBSCAN on a given partition
:param iterable:
:param params:
:return:
"""
data = []
for x in iterable:
data.append(x)
if len(data) > 0:
x = np.array([row[1] for row in data])
parts = [row[0][1] for row in data]
y = np.array([row[0][0] for row in data])
model = skc.DBSCAN(**params)
c = model.fit_predict(x)
for i in range(len(c)):
yield (y[i], (parts[i], c[i]))
def map_cluster_id(row_id_labels, bc_forward_mapper):
row_id = int(row_id_labels[0])
labels = []
for label in row_id_labels[1]:
labels.append(label)
cluster_id = next(iter(labels))
cluster_dict = bc_forward_mapper.value
if (cluster_id[1] != -1) and (cluster_id in cluster_dict):
return row_id, int(cluster_dict[cluster_id])
else:
return row_id, -1
if __name__ == "__main__":
pass
# spark = Context().spark
# rdd = spark.sparkContext.parallelize([
# (1, "A", 19, 181, 67),
# (2, "C", 17, 179, 67),
# (3, 'E', 18, 180, 68),
# (4, 'E', 29, 180, 68),
# (5, 'E', 18, 180, 68),
# (6, 'E', 18, 180, 68),
# (7, 'E', 18, 180, 68),
# (8, 'E', 18, -180, 68),
# (9, 'F', 28, 21, 7),
# (10, 'F', 28, 22, 8),
# (11, 'F', 28, 22, 8),
# (12, 'F', 28, 22, 8),
# (13, 'F', 28, 22, 8),
# (14, 'F', 28, 23, 7),
# ])
# from pyspark.sql.types import StructType, StructField, LongType, StringType, IntegerType
#
# schema = StructType([
# StructField("id", LongType(), True),
# StructField("name", StringType(), True),
# StructField("age", LongType(), True),
# StructField("height", IntegerType(), True),
# StructField("weight", IntegerType(), True)
# ])
# df = spark.createDataFrame(rdd, schema)
#
# db = DBSCAN(max_partitions=3)
# db.fit(df, ["height", "weight"], "id")
# print(db.detect())
| 5,161 | 1,864 |
import tensorflow as tf
import numpy as np
import sys
sys.path.append('../LeFlow/src')
import processMif as mif
# This creates a random 32-bit floating point input for your variables
def rand(dim):
#return np.random.random(dim).astype(np.float32)
return np.ones(dim).astype(np.float32)
nodes = 1
X = tf.placeholder(tf.float32, [1, 3])
test=rand([1,3])
dense1_w = tf.Variable(rand([3, nodes]))
dense1_b = tf.Variable(rand([nodes]))
with tf.Session() as sess:
# Generating circuit
with tf.device("device:XLA_CPU:0"):
dense1 = tf.add(tf.matmul(X, dense1_w)[0], dense1_b)
#y = tf.cast(dense1[0], tf.int32)
y = dense1[0]
# y = tf.cast(dense1, tf.int32)
#y = dense1
sess.run(tf.global_variables_initializer())
result = sess.run(y,{X: test})
# Print expected result
print("Result Calculated: "+ str(result))
# Creating .mif files to initialize the memories
# To do this, go to <your_file>_ir_7.ll and check the list of params (inputs) and their sizes
param5 = dense1_w.eval();
param6 = dense1_b.eval();
mif.createMem([param5,test,param6])
#mif.createMem([test,param5,param6])
| 1,160 | 447 |
import pickle
import numpy as np
pkl_file = open('D:\wangzhihui\\123\wzh\ResNext101_32_4d_faster_rcnn_merge_bn_scale_iter_60000\\detections.pkl', 'rb')
data1 = pickle.load(pkl_file)
def py_cpu_nms(dets, thresh):
"""Pure Python NMS baseline."""
x1 = dets[:, 0]
y1 = dets[:, 1]
x2 = dets[:, 2]
y2 = dets[:, 3]
scores = dets[:, 4]
areas = (x2 - x1 + 1) * (y2 - y1 + 1)
order = scores.argsort()[::-1]
keep = []
while order.size > 0:
i = order[0]
keep.append(i)
xx1 = np.maximum(x1[i], x1[order[1:]])
yy1 = np.maximum(y1[i], y1[order[1:]])
xx2 = np.minimum(x2[i], x2[order[1:]])
yy2 = np.minimum(y2[i], y2[order[1:]])
w = np.maximum(0.0, xx2 - xx1 + 1)
h = np.maximum(0.0, yy2 - yy1 + 1)
inter = w * h
ovr = inter / (areas[i] + areas[order[1:]] - inter)
inds = np.where(ovr <= thresh)[0]
order = order[inds + 1]
return keep
print type(data1)
print type(data1[1])
print len(data1[1])
aa = data1[1][26]
print aa
for i in xrange(4952):
if len(data1[1][i]) > 0:
aa = np.append(aa, data1[1][i], axis=0)
print len(aa)
keep = py_cpu_nms(aa, 0.3)
bb = aa[keep, :]
print len(bb)
print bb[45]
aa = np.append(aa, aa, axis=0)
aa = np.append(aa, aa, axis=0)
print len(aa)
keep = py_cpu_nms(aa, 0.3)
cc = aa[keep, :]
print len(cc)
print cc[45]
#################################################################################
import numpy as np
a = [[1, 2, 4, 5], [6, 8, 1, 0]]
b = [[1, 2, 4, 3], [6, 8, 1, 2]]
c = [[1, 2, 4, 9], [6, 8, 1, 5]]
aa = np.array(a)
print aa[:, 0::2]
aa = np.append(a, b, axis=0)
aa = np.append(aa, c, axis=0)
bb = aa[:, 0:3]
inds = np.where(aa[:, 1] > 1)[0]
print inds
cls_scores = aa[inds, 3]
print cls_scores
print aa
print bb
print bb[:, np.newaxis]
cls = np.hstack((aa, bb)).astype(np.float32, copy=False)
print cls
| 1,894 | 913 |
from calm.dsl.builtins import basic_cred, CalmTask, action
from calm.dsl.builtins import SimpleDeployment, SimpleBlueprint
from calm.dsl.builtins import read_provider_spec
from calm.dsl.builtins import CalmVariable
from calm.dsl.store import Secret
CENTOS = basic_cred('nutanix', 'nutanix/4u', name='CENTOS', default=True)
HYCU_CRED = basic_cred('admin', 'admin', name='HYCU_CRED', default=False)
class CentosDeployment(SimpleDeployment):
provider_spec = read_provider_spec('specs/centos-8.yaml')
os_type = 'Linux'
@action
def __create__(self):
CalmTask.Exec.escript(name='add_vm_to_hycu', filename='scripts/add_vm_to_hycu.py')
@action
def __install__(self):
# CalmTask.Exec.ssh(name='Update CentOS', script='sudo yum -y --quiet update')
CalmTask.Exec.ssh(name='Update CentOS', script='echo "hello world"')
class HYCUCentOS8(SimpleBlueprint):
credentials = [CENTOS, HYCU_CRED]
deployments = [CentosDeployment]
VM_NAME = CalmVariable.Simple.string('CentOS-VM', label='VM Name', runtime=True)
# HYCU IP address, assuming default port for API access (8443)
HYCU_IP = CalmVariable.Simple.string('10.21.21.100', runtime=False, is_hidden=True)
HYCU_PORT = CalmVariable.Simple.string('8443', runtime=False, is_hidden=True)
def main():
print(HYCUCentOS8.json_dumps(pprint=True))
if __name__ == '__main__':
main() | 1,398 | 510 |
# Definition for a binary tree node.
# class TreeNode:
# def __init__(self, val=0, left=None, right=None):
# self.val = val
# self.left = left
# self.right = right
# class Solution:
# def isSymmetric(self, root: TreeNode) -> bool:
# '''
# long, iterative solution
# '''
# if not root.left and not root.right:
# return True
# elif not root.left or not root.right:
# return False
# # BFS search left and right, compare the results
# from collections import deque
# q1 = deque()
# q2 = deque()
# q1.append(root.left)
# q2.append(root.right)
# while q1 and q2:
# n1 = q1.popleft()
# n2 = q2.popleft()
# if n1.val != n2.val or len(q1)!=len(q2):
# return False
# # add children
# if n1.right:
# q1.append(n1.right)
# elif n1.val != 200:
# q1.append(TreeNode(200))
# if n1.left:
# q1.append(n1.left)
# elif n1.val != 200:
# q1.append(TreeNode(200))
# if n2.left:
# q2.append(n2.left)
# elif n2.val != 200:
# q2.append(TreeNode(200))
# if n2.right:
# q2.append(n2.right)
# elif n2.val != 200:
# q2.append(TreeNode(200))
# if len(q1) != len(q2):
# return False
# return True
# class Solution:
# def isSymmetric(self, root: TreeNode) -> bool:
# '''
# recursive solution
# '''
# def mirror(n1, n2):
# if not n1 and not n2:
# return True
# if not n1 or not n2:
# return False
# return (n1.val == n2.val) & mirror(n1.right, n2.left) & mirror(n1.left, n2.right)
# return mirror(root, root)
class Solution:
def isSymmetric(self, root: TreeNode) -> bool:
'''
another iterative solution
'''
from collections import deque
q = deque()
q.append(root)
q.append(root)
while q:
n1 = q.popleft()
n2 = q.popleft()
if not n1 and not n2:
continue
if not n1 or not n2:
return False
if n1.val != n2.val:
return False
q.append(n1.left)
q.append(n2.right)
q.append(n1.right)
q.append(n2.left)
return True
| 2,695 | 836 |
#!/usr/bin/python2
from Tkinter import *
class Checkbar(Frame):
def __init__(self,parent=None,picks=[],side=LEFT,anchor=W):
Frame.__init__(self,parent)
self.vars = []
for pick in picks:
var = IntVar()
chk = Checkbutton(self,text=pick,variable=var)
chk.pack(side=side,anchor=anchor,expand=YES)
self.vars.append(var)
def state(self):
return map((lambda var:var.get()),self.vars)
if __name__ == '__main__':
root = Tk()
lng = Checkbar(root,['Python','Ruby','Perl','C++'])
tgl = Checkbar(root,['English','German'])
lng.pack(side=TOP,fill=X)
tgl.pack(side=TOP)
lng.config(relief=GROOVE,bd=2)
def allstates():
print(list(lng.state()),list(tgl.state()))
Button(root,text='Quit',command=root.quit).pack(side=RIGHT)
Button(root, text='Peek',command=allstates).pack(side=RIGHT)
root.mainloop() | 920 | 331 |
from draftHost.models import NflPosition, FantasyPosition
# path relative to manage.py
NFL_DATA_FILE = "draftHost/data/nfl_positions.txt"
FANTASY_DATA_FILE = "draftHost/data/fantasy_positions.txt"
class PositionImporter(object):
def add_positions(self):
try:
data = open(NFL_DATA_FILE, 'r')
for line in data:
parts = line.rstrip().split(',')
position, created = NflPosition.objects.get_or_create(**{
'id': parts[0],
'description': parts[1],
'abbreviation': parts[2],
})
if created:
print "added {p}".format(p=position)
data.close()
except IOError, e:
print "got error {e}".format(e=e)
def add_fantasy_positions(self):
try:
data = open(FANTASY_DATA_FILE, 'r')
for line in data:
abbrev = line.rstrip()
position = self.find_match(abbrev)
fantasy, created = FantasyPosition.objects.get_or_create(
position=position
)
if created:
print "created {f}".format(f=fantasy)
data.close()
except IOError, e:
print "got error {e}".format(e=e)
def find_match(self, abbreviation):
return NflPosition.objects.filter(abbreviation=abbreviation)[0]
| 1,447 | 422 |
# Network in Network CIFAR10 Model
# Original source: https://gist.github.com/mavenlin/e56253735ef32c3c296d
# License: unknown
# Download pretrained weights from:
# https://s3.amazonaws.com/lasagne/recipes/pretrained/cifar10/model.pkl
from lasagne.layers import InputLayer, DropoutLayer, FlattenLayer
from lasagne.layers.dnn import Conv2DDNNLayer as ConvLayer
from lasagne.layers import Pool2DLayer as PoolLayer
def build_model():
net = {}
net['input'] = InputLayer((None, 3, 32, 32))
net['conv1'] = ConvLayer(net['input'],
num_filters=192,
filter_size=5,
pad=2,
flip_filters=False)
net['cccp1'] = ConvLayer(
net['conv1'], num_filters=160, filter_size=1, flip_filters=False)
net['cccp2'] = ConvLayer(
net['cccp1'], num_filters=96, filter_size=1, flip_filters=False)
net['pool1'] = PoolLayer(net['cccp2'],
pool_size=3,
stride=2,
mode='max',
ignore_border=False)
net['drop3'] = DropoutLayer(net['pool1'], p=0.5)
net['conv2'] = ConvLayer(net['drop3'],
num_filters=192,
filter_size=5,
pad=2,
flip_filters=False)
net['cccp3'] = ConvLayer(
net['conv2'], num_filters=192, filter_size=1, flip_filters=False)
net['cccp4'] = ConvLayer(
net['cccp3'], num_filters=192, filter_size=1, flip_filters=False)
net['pool2'] = PoolLayer(net['cccp4'],
pool_size=3,
stride=2,
mode='average_exc_pad',
ignore_border=False)
net['drop6'] = DropoutLayer(net['pool2'], p=0.5)
net['conv3'] = ConvLayer(net['drop6'],
num_filters=192,
filter_size=3,
pad=1,
flip_filters=False)
net['cccp5'] = ConvLayer(
net['conv3'], num_filters=192, filter_size=1, flip_filters=False)
net['cccp6'] = ConvLayer(
net['cccp5'], num_filters=10, filter_size=1, flip_filters=False)
net['pool3'] = PoolLayer(net['cccp6'],
pool_size=8,
mode='average_exc_pad',
ignore_border=False)
net['output'] = FlattenLayer(net['pool3'])
return net
| 2,572 | 812 |
# -*- coding: utf-8 -*-
"""
Part of the OASIS ART PROJECT - https://github.com/orgs/oasis-art-project
Copyright (c) 2019-22 TEAM OASIS
License Artistic-2.0
"""
from marshmallow import fields, validate, post_dump
from sqlalchemy.types import ARRAY
from src.backend.extensions import db
from src.backend.models.model import SurrogatePK, BaseSchema
from src.backend.models.userModel import UserSchema, User
from src.backend.controllers.controller import build_image_list
class Place(SurrogatePK, db.Model):
__tablename__ = 'places'
host_id = db.Column(db.Integer, db.ForeignKey('users.id'))
name = db.Column(db.String(100), nullable=False)
description = db.Column(db.String(1000), nullable=True)
address = db.Column(db.String(300), nullable=False)
location = db.Column(db.String(12), nullable=True)
homepage = db.Column(db.String(100), nullable=True)
instagram = db.Column(db.String(30), nullable=True)
facebook = db.Column(db.String(30), nullable=True)
matterport_link = db.Column(db.String(15), nullable=True)
active = db.Column(db.Boolean, nullable=True)
creation_date = db.Column(db.TIMESTAMP, server_default=db.func.current_timestamp(), nullable=False)
host = db.relationship('User', backref=db.backref('places'))
def __init__(self, **kwargs):
super(Place, self).__init__(**kwargs)
class PlaceSchema(BaseSchema):
# Overwritten fields
host = fields.Nested(UserSchema, only=('id',), required=True)
name = fields.Str(required=True, validate=validate.Length(max=100))
description = fields.Str(validate=validate.Length(max=1000))
address = fields.Str(required=True, validate=validate.Length(max=300))
location = fields.Str(allow_none=True, validate=validate.Length(max=12))
homepage = fields.Str(allow_none=True, validate=validate.Length(max=100))
instagram = fields.Str(allow_none=True, validate=validate.Length(max=30))
facebook = fields.Str(allow_none=True, validate=validate.Length(max=30))
matterport_link = fields.Str(validate=validate.Length(max=15))
active = fields.Boolean(allow_none=True)
class Meta:
# BaseSchema automatically generates fields based on the model
model = Place
# Since according to Nested schema loading is only with ID,
# dump loads other non-sensitive data from DB, enumerated below
@post_dump
def get(self, data):
if 'host' in data:
host = User.get_by_id(data['host']['id'])
if not host:
raise ValueError
d = UserSchema(only=('id', 'tags', 'firstName', 'lastName', 'bio', 'files', 'homepage', 'instagram', 'youtube', 'showChat', 'confirmed', 'active')).dump(host).data
data['host'] = d
if 'files' in data:
data['fullImages'] = build_image_list('place', data['id'], data['files'], 'f')
data['prevImages'] = build_image_list('place', data['id'], data['files'], 'p')
return data
| 2,969 | 969 |
from einops import repeat, rearrange
from torch import nn
from torcharc import net_util
import math
import pydash as ps
import sys
import torch
def build_learned_pos_encoding(max_seq_len: int, embed_dim: int):
'''Build learned positional encoding with Deepmind's init'''
# learned position encoding
pos_encoding = nn.Parameter(torch.empty(max_seq_len, embed_dim))
nn.init.trunc_normal_(pos_encoding, mean=0.0, std=0.02) # Deepmind's init
return pos_encoding
class Identity(nn.Identity):
def __init__(self, in_shape: list):
super().__init__()
self.in_shape = in_shape
self.out_shape = in_shape
class TextPreprocessor(nn.Module):
'''Standard text preprocessing for transformer by embedding a tokenized tensor, then adding a learned position encoding.'''
def __init__(self, vocab_size: int, embed_dim: int, max_seq_len: int = 512, **_kwargs):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embed_dim)
# learned position encoding
self.pos_encoding = build_learned_pos_encoding(max_seq_len, embed_dim)
self.scale = embed_dim ** 0.5
self.out_shape = [max_seq_len, embed_dim]
def forward(self, x: torch.Tensor) -> torch.Tensor:
batch, seq_len = x.shape
pe = repeat(self.pos_encoding[:seq_len], '... -> b ...', b=batch) # repeat for batch
return self.embedding(x) * self.scale + pe
class FourierPreprocessor(nn.Module):
'''
Spatial input preprocessor for PerceiverEncoder using Fourier positional encoding for any dimensions of spatial tensor with channels, i.e. shape (x, y, ..., c)
This builds Fourier pos_encoding for coordinates of N-dimensional spatial data as a meshgrid
e.g. for image of shape (x, y, c) -> get a meshgrid of shape (x, y, d=2), where each slice at d is a meshgrid of a dimension
then generate the sin, cos frequencies, stack [pos, sin, cos],
then flatten the meshgrid's spatial dimension into 1D to get the final pos_encoding of shape (x*y*..., d*(2*num_freq_bands+1)).
When encoding, this flattens the spatial dimensions of input into 1D, e.g. (x, y, ..., c) into (x*y*..., c), then concat it with the pos_encoding, so the final output tensor is a stack of the [flattened input with channels, pos_encoding with d*(2*num_freq_bands+1).
The output shape is (x*y*..., out_dim), where out_dim = (c+d*(2*num_freq_bands+1))
@example
batch = 2
in_shape = [64, 3]
num_freq_bands = 32
x = torch.rand(batch, *in_shape)
module = FourierPreprocessor(in_shape, num_freq_bands)
out = module(x)
assert [math.prod(in_shape[:-1]), module.out_dim] == module.out_shape
assert list(out.shape) == [batch, *module.out_shape]
'''
def __init__(self, in_shape: list, num_freq_bands: int, max_reso: list = None, cat_pos: bool = True):
super().__init__()
*self.spatial_shape, num_c = self.in_shape = list(in_shape) # shape excluding batch
self.num_freq_bands = num_freq_bands
self.cat_pos = cat_pos
# create fourier positional encoding
pos = self.build_positions()
self.pos_encoding = self.build_pos_encoding(pos, max_reso=max_reso)
flat_dim = math.prod(in_shape[:-1])
self.out_dim = num_c + self.get_pos_encoding_dim() # in_dim to PerceiverEncoder; we stack pos_encoding with top of channels
self.out_shape = [flat_dim, self.out_dim]
def build_positions(self, start: float = -1.0, end: float = 1.0):
'''Build spatial coordinates as a meshgrid, i.e. coordinates laid out such that values along the channel is a point in coordinate, e.g. shape = (x, y, 2)'''
x_y = [torch.linspace(start, end, steps=s) for s in self.spatial_shape]
return torch.stack(torch.meshgrid(*x_y), dim=len(self.spatial_shape))
def build_pos_encoding(self, pos: torch.Tensor, max_reso: list = None) -> torch.Tensor:
'''
Generate a Fourier frequency position encoding with linear spacing.
@param pos: meshgrid position coordinates of shape (x, y, d=len(shape)), e.g. (x, y, 2), or (x, y, z, 3) etc. in general
@param max_reso: maximum resolution (pixels) per dimension. Useful when input such as picture varies in size
@param cat_pos: whether to concat pos before the fourier encoding
@return position encodings tensor of shape (x, y,... d*(2*num_freq_bands+1))
'''
max_reso = max_reso or pos.shape[:-1]
assert len(max_reso) == len(pos.shape[:-1]), f'max_reso len(shape) must match pos len(shape), but got {len(max_reso)} instead of {len(pos.shape[:-1])}'
freq_bands = torch.stack([torch.linspace(1.0, max_r / 2.0, steps=self.num_freq_bands) for max_r in max_reso])
pos_freqs = rearrange(torch.einsum('...d,df->d...f', pos, freq_bands), 'd ... f -> ... (d f)')
encodings = [pos] if self.cat_pos else []
encodings += [torch.sin(math.pi * pos_freqs), torch.cos(math.pi * pos_freqs)]
spatial_encoding = torch.cat(encodings, dim=-1) # shape (x, y,... d*(2*num_freq_bands+1))
# flatten spatial dimensions into 1D
pos_encoding = rearrange(spatial_encoding, '... c -> (...) c')
return nn.Parameter(pos_encoding)
def get_pos_encoding_dim(self) -> int:
return len(self.spatial_shape) * (2 * self.num_freq_bands + int(self.cat_pos))
def forward(self, x: torch.Tensor) -> torch.Tensor:
batch, *x_in_shape = x.shape
assert x_in_shape == self.in_shape, f'input shape {x_in_shape} != expected {self.in_shape}'
pos_encoding = repeat(self.pos_encoding, '... -> b ...', b=batch) # repeat for batch
x = rearrange(x, 'b ... c -> b (...) c') # flatten spatial dimensions into 1D
return torch.cat([x, pos_encoding], dim=-1) # stack 1D input with pos_encoding
class MultimodalPreprocessor(nn.Module):
'''
Multimodal preprocessor for multimodal input {mode: x}
This recursively builds a preprocessor for each mode, and applies them to the multimodal inputs in order.
To combine the multimodal preprocessed outputs,
first note that each output is a 2D array of (max_seq_len, channel) or (M, C) of Perceiver input array.
They are padded with trainable position encoding (1 position per mode, broadcasted) to have the same common_channels (max_channels + pad_channels), before getting concatenated along the sequences for transformer to attend to.
The output shape is [total_seq_len, common_channels]
'''
def __init__(self, in_shapes: dict, arc: dict, pad_channels: int = 2):
super().__init__()
self.preprocessors = nn.ModuleDict({
mode: net_util.build_component(arc, {'in_shape': in_shape}, mode, sys.modules[__name__])
for mode, in_shape in in_shapes.items()
})
self.out_shapes = {mode: preprocessor.out_shape for mode, preprocessor in self.preprocessors.items()}
total_seq_len = ps.sum_by(self.out_shapes, ps.head)
max_channels = ps.max_by(self.out_shapes, ps.last)[-1]
common_channels = max_channels + pad_channels
self.pos_encodings = nn.ParameterDict({
mode: build_learned_pos_encoding(1, common_channels - out_shape[-1])
for mode, out_shape in self.out_shapes.items()
})
self.out_shape = [total_seq_len, common_channels]
def pos_encoding_pad(self, mode: str, out: torch.Tensor) -> torch.Tensor:
'''
Pad output to ensure they result in shape [batch, seq_len, common_channels]
The padding channels ensured by pad_channels are used to stack learned pos_encoding of shape [1, common_channels - out_dim] (broadcasted) for each mode,
i.e. each mode has 1 encoded position for transformer to differentiate
'''
pos_encoding = self.pos_encodings[mode]
batch, seq_len, _channel = out.shape
padding = pos_encoding.broadcast_to((batch, seq_len, pos_encoding.shape[-1]))
return torch.cat([out, padding], dim=2) # concat along channel to result in common_channels
def forward(self, xs: dict) -> torch.Tensor:
outs = []
for mode, x in xs.items():
out = self.preprocessors[mode](x)
padded_out = self.pos_encoding_pad(mode, out)
outs.append(padded_out)
# NOTE concat along seq_len to result in [total_seq_len, common_channels] since transformer attention is along seq_len, not channel
return torch.cat(outs, dim=1)
| 8,509 | 2,803 |
import os
import random
import logging
import threading
base_path = os.path.dirname(os.path.dirname(__file__))
base_path = os.path.join(base_path, "datafield")
if not os.path.exists(base_path):
os.mkdir(base_path)
def get_path(fieldname):
return os.path.join(base_path, fieldname)
class DataField:
'''
操作datafield/中数据的对象
'''
def __init__(self):
self.fields = set()
self.field_cache = {}
self.field_sent = {}
self.load_field()
self.lock = threading.Lock()
def load_field(self):
'''
load需要读取的field
'''
for filename in os.listdir(base_path):
if os.path.isfile(get_path(filename)):
continue
self.fields.add(filename)
def new_field(self, fieldname):
'''
新建field
'''
if fieldname in self.fields:
return
abs_path = get_path(fieldname)
if not os.path.exists(abs_path):
os.mkdir(abs_path)
self.fields.add(fieldname)
def load_field_data(self, fieldname):
'''
读取field中的数据
注:本功能线程不安全,请不要用本函数读取一个会变化的field
'''
ret = []
for root, _, files in os.walk(get_path(fieldname)):
for file in files:
with open(os.path.join(root, file), "r", encoding="utf-8") as f:
s = f.read()
ret.append(s)
return ret
def get_field_data(self, fieldname):
'''
读取field中的数据
注:本函数会随机返回一行数据
'''
with self.lock:
if fieldname not in self.field_sent:
self.field_sent[fieldname] = set()
if fieldname not in self.field_cache or self.field_cache[fieldname] == []:
self.field_cache[fieldname] = "\n".join(self.load_field_data(fieldname)).split("\n")
random.shuffle(self.field_cache[fieldname])
self.field_cache[fieldname] = [i for i in self.field_cache[fieldname] if i != "" and i not in self.field_sent]
if self.field_cache[fieldname] == []:
return None
ret = self.field_cache[fieldname].pop()
self.field_sent[fieldname].add(ret)
return ret
def save_to_field(self, fieldname, s, filename=None, mode="w"):
'''
保存至field中的某个文件
注:请手动添加\\n
'''
if filename is None:
for i in range(16, len(s)):
filename = f"{s[:i].__hash__()}.txt"
path = os.path.join(get_path(fieldname), filename)
if not os.exist(path):
break
else:
path = os.path.join(get_path(fieldname), filename)
with open(path, mode, encoding="utf-8") as f:
f.write(s)
datafields = DataField()
if __name__ == '__main__':
print(datafields.fields)
print(datafields.get_field_data("up_mid"))
print(datafields.get_field_data("up_mid"))
print(datafields.get_field_data("up_mid")) | 3,027 | 996 |
# https://docs.microsoft.com/en-us/windows/python/beginners
# https://developers.google.com/identity/protocols/oauth2/service-account#python
from __future__ import print_function
from pathlib import Path
from googleapiclient.discovery import build
from google.oauth2 import service_account
SCOPES = ['https://www.googleapis.com/auth/gmail.readonly']
HOME_PATH = str(Path.home())
SERVICE_ACCOUNT_FILE = HOME_PATH + '/devkey/devhkmci-gmaildomainwide-1d7640a0c6d2.json'
def main():
DELEGATE='aaron.ko@dev.hkmci.com' # Service account will impersonate this user. Must have proper admin privileges in G Suite.
# TARGET='dev.hkmci.com' # Service account wants to access data from this.
credentials = service_account.Credentials.from_service_account_file(SERVICE_ACCOUNT_FILE, scopes=SCOPES)
credentials_delegated = credentials.with_subject(DELEGATE)
service = build('gmail', 'v1', credentials=credentials_delegated)
# Call the Gmail API
results = service.users().getProfile(userId='me').execute()
print(results)
results = service.users().labels().list(userId='me').execute()
print(results)
# labels = results.get('labels', [])
# for label in labels:
# print(label['name'])
# if not labels:
# print('No labels found.')
# else:
# print('Labels:')
# for label in labels:
# print(label['name'])
if __name__ == '__main__':
main()
# [END gmail_quickstart]
| 1,468 | 487 |
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.distributions as td
from torch.distributions import constraints
from dpp.nn import BaseModule, Hypernet
from dpp.utils import clamp_preserve_gradients
def inverse_sigmoid(x):
# Clamp tiny values (<1e-38 for float32)
finfo = torch.finfo(x.dtype)
x = x.clamp(min=finfo.tiny, max=1. - finfo.eps)
return torch.log(x) - torch.log1p(-x)
def logistic_sample(means, log_scales):
if means.shape != log_scales.shape:
raise ValueError("Shapes of means and scales don't match.")
z = torch.rand(means.shape)
return torch.exp(log_scales) * inverse_sigmoid(z) + means
# Credit: https://github.com/aravindsrinivas/flowpp/blob/master/flows/logistic.py
def logistic_logpdf(x, mean, log_scale):
z = (x - mean) * torch.exp(-log_scale)
return z - log_scale - 2 * F.softplus(z)
def logistic_logcdf(x, mean, log_scale):
z = (x - mean) * torch.exp(-log_scale)
return F.logsigmoid(z)
def mixlogistic_logpdf(x, prior_logits, means, log_scales):
log_prior = F.log_softmax(prior_logits, dim=-1)
return torch.logsumexp(
log_prior + logistic_logpdf(x.unsqueeze(-1), means, log_scales),
dim=-1
)
def mixlogistic_logcdf(x, prior_logits, means, log_scales):
log_prior = F.log_softmax(prior_logits, dim=-1)
return torch.logsumexp(
log_prior + logistic_logcdf(x.unsqueeze(-1), means, log_scales),
dim=-1
)
class LogisticMixtureDistribution(BaseModule):
def __init__(self, config, n_components=32, hypernet_hidden_sizes=[64], min_clip=-5., max_clip=3.):
super().__init__()
self.n_components = n_components
self.use_history(config.use_history)
self.use_embedding(config.use_embedding)
self.min_clip = min_clip
self.max_clip = max_clip
self.hypernet = Hypernet(config,
hidden_sizes=hypernet_hidden_sizes,
param_sizes=[n_components, n_components, n_components])
def get_params(self, h, emb):
"""Generate model parameters based on the history and embeddings.
Args:
h: history embedding, shape [*, rnn_hidden_size]
emb: sequence embedding, shape [*, embedding_size]
Returns:
prior_logits: shape [*, n_components]
means: shape [*, n_components]
log_scales: shape [*, n_components]
"""
if not self.using_history:
h = None
if not self.using_embedding:
emb = None
prior_logits, means, log_scales = self.hypernet(h, emb)
# Clamp values that go through exp for numerical stability
prior_logits = clamp_preserve_gradients(prior_logits, self.min_clip, self.max_clip)
log_scales = clamp_preserve_gradients(log_scales, self.min_clip, self.max_clip)
return prior_logits, means, log_scales
def log_prob(self, y, h=None, emb=None):
prior_logits, means, log_scales = self.get_params(h, emb)
return mixlogistic_logpdf(y, prior_logits, means, log_scales)
def log_cdf(self, y, h=None, emb=None):
prior_logits, means, log_scales = self.get_params(h, emb)
return mixlogistic_logcdf(y, prior_logits, means, log_scales)
def sample(self, n_samples, h=None, emb=None):
"""Draw samples from the model.
Args:
n_samples: number of samples to generate.
h: hidden state, shape [*, rnn_hidden_size]
emb: sequence embedding, shape [*, embedding_size]
Returns:
samples: shape [*, n_samples]
"""
with torch.no_grad():
prior_logits, means, log_scales = self.get_params(h, emb)
# model parameters should have two dimensions for bmm to work
# first dimensions will be restored later
prior_logits = prior_logits.view(-1, self.n_components)
means = means.view(-1, self.n_components)
log_scales = log_scales.view(-1, self.n_components)
categorical = td.Categorical(logits=prior_logits)
z = categorical.sample([n_samples])
# z has shape [n_samples, *], convert to [*, n_samples]
dim_order = np.arange(len(prior_logits.shape))
dim_order = tuple(np.concatenate([dim_order[1:], [0]]))
z = z.permute(dim_order).contiguous()
# z_oh has shape [*, n_samples, n_components]
# convert it to [*, n_components, n_samples] for bmm to work
z_oh = F.one_hot(z, num_classes=self.n_components).float().transpose(-2, -1)
# add extra dim to means and log_scales for bmm to work
means.unsqueeze_(-2)
log_scales.unsqueeze_(-2)
# select the correct component for each sample
means_select = torch.bmm(means, z_oh)
log_scales_select = torch.bmm(log_scales, z_oh)
means_select.squeeze_(-2)
log_scales_select.squeeze_(-2)
# means_select and log_scales_select have shape [*, n_samples]
samples = logistic_sample(means_select, log_scales_select)
# reshape the samples back to the original shape
if (h is not None):
first_dims = h.shape[:-1]
elif (emb is not None):
first_dims = emb.shape[:-1]
else:
first_dims = torch.Size()
shape = first_dims + torch.Size([n_samples])
return samples.reshape(shape)
| 5,575 | 1,840 |
from django.urls import path
from django.conf import settings
from django.conf.urls.static import static
from .views import *
urlpatterns = [
# Urls for authentication on noruh web
path('change_password/', RecoverPasswordByApi.as_view(), name='change_password'),
path('reset_passowrd/complete/', RecoverPasswordByApiComplete.as_view(), name='reset_password_complete'),
path('login/', Login.as_view(), name='login'),
path('user/logged/', ListUserLogged.as_view(), name='user_logged_detail'),
path('user/logged/alter/<int:pk>/', AlterUser.as_view(), name='user_logged_alter'),
path('', Home.as_view(), name='home'),
path('terms_and_conditions/', TermsAndContions.as_view(), name='terms_and_conditions'),
# Url's for Establishments and configurations
path('establishment/create/', CreateEstablishment.as_view(),
name='establishment_create'),
path('establishment/list/all/', ListAllEstablishment.as_view(),
name='establishment_list_all'),
path('establishment/search/list/', ListSearchEstablishment.as_view(),
name='establishment_search_list'),
path('establishment/configurations/<int:id>/', ConfigurationsEstablishment.as_view(),
name='establishment_configurations'),
path('establishment/base/<int:id>/', BaseEstablishment.as_view(),
name='establishment_base'),
path('establishment/update/location/<int:pk>/', UpdateEstablismentLocation.as_view(),
name='establishment_update_location'),
path('establishment/update/description/<int:pk>/', UpdateEstablismentDescription.as_view(),
name='establishment_update_description'),
path('establishment/update/amenities/<int:pk>/', UpdateEstablismentAmenities.as_view(),
name='establishment_update_amenities'),
path('establishment/update/gps_restriction/<int:pk>/', UpdateGPSRestrictionEstablishment.as_view(),
name='establishment_update_gps_restriction'),
path('establishment/update/featured/<int:pk>/', UpdateFeaturedEstablishment.as_view(),
name='establishment_update_featured'),
path('establishment/update/enable/<int:pk>/', DisableEstablishment.as_view(),
name='establishment_update_enable'),
path('establishment/update/taxes/<int:pk>/', UpdateEstablismentTaxes.as_view(),
name='establishment_update_taxes'),
path('establishment/update/pays_payment_tax/<int:pk>/', UpdateEstablishmentPaysPaymentTax.as_view(),
name='establishment_update_pays_payment_tax'),
path('establishment/update/couvert/<int:pk>/', UpdateEstablismentCouvert.as_view(),
name='establishment_update_couvert'),
path('establishment/update/offer_range_value/<int:pk>/', UpdateEstablismentOfferRangeValue.as_view(),
name='establishment_update_offer_range_value'),
path('establishment/update/open_close/<int:pk>/', UpdateOpenOrCloseEstablishment.as_view(),
name='establishment_update_open_close'),
path('establishment/delete/<int:pk>/', DeleteEstablishment.as_view(),
name='establishment_delete'),
path('establishment/create/photo/<int:establishment_id>/', AddPhotoOnEstablishment.as_view(),
name='establishment_add_photo'),
path('establishment/photo/delete/<int:pk>/', DeletePhotoFromEstablishment.as_view(),
name='establishment_delete_photo'),
path('dashboard/', DashboardAllEstablishments.as_view(), name='dashboard_all_establishments'),
path('establishment/dashboard/<int:establishment_id>/', DashboardEstablishment.as_view(),
name='establishment_dashboard'),
path('establishment/dashboard/items_more_requested/<int:establishment_id>/', ListAllItemsMoreRequested.as_view(),
name='establishment_items_more_requested'),
# Url's for when Request a Waiter
path('establishment/requests/list/<int:establishment_id>/',
RequestWaiter.as_view(), name='request_list'),
path('establishment/requests/accept/<int:pk>/',
AcceptRequestWaiter.as_view(), name='accept_request'),
path('establishment/requests/accept/all/<int:establishment_id>/',
AcceptAllRequestWaiter.as_view(), name='accept_all_requests'),
# Url's for Establishment Evaluations
path('establishment/evaluation/list/<int:establishment_id>/',
ListEvaluation.as_view(), name='evaluation_list'),
path('establishment/evaluation/answer/<int:evaluation_id>/',
CreateAnswerToEvaluation.as_view(), name='answer_evaluation'),
path('establishment/evaluation/delete/<int:pk>/',
DeleteEvaluation.as_view(), name='evaluation_delete'),
path('establishment/evaluation/answer/delete/<int:pk>/',
DeleteAnswerEvaluation.as_view(), name='evaluation_answer_delete'),
# Url's for Employees
path('establishment/employee/create/', CreateEmployee.as_view(),
name='employee_create'),
path('establishment/employee/list/<int:establishment_id>/',
ListEmployeeEstablishment.as_view(), name='employee_list_establishment'),
path('establishment/employee/list/<int:establishment_id>/search/',
ListSearchEmployeeEstablishment.as_view(), name='employee_list_search_establishment'),
path('establishment/employee/list/', ListEmployeeAll.as_view(),
name='employee_list_all'),
path('establishment/employee/list/search/', ListSearchEmployee.as_view(),
name='employee_search_list'),
path('establishment/employee/detail/<int:pk>/',
DetailEmployee.as_view(), name='employee_detail'),
path('establishment/employee/alter/<int:pk>/',
AlterEmployee.as_view(), name='employee_alter'),
path('establishment/employee/delete/<int:pk>/',
DeleteEmployee.as_view(), name='employee_delete'),
# Url's for Menu, MenuItem, ItemCategory, Observation
path('menu/list/<int:establishment_id>/', ListMenuFromEstablishment.as_view(),
name='menu_list_from_establishment'),
path('menu/list/<int:establishment_id>/search/', ListMenuSearchFromEstablishment.as_view(),
name='menu_list_search_from_establishment'),
# Items from Menu
path('menu/add/item/<int:establishment_id>/',
CreateItemOnMenu.as_view(), name='menu_create_item'),
path('menu/list/item/<int:establishment_id>/',
ListMenuItems.as_view(), name='menu_item_list'),
path('menu/list/item/<int:establishment_id>/search/',
ListMenuItemsSearch.as_view(), name='menu_item_list_search'),
path('menu/list/item/update/<int:pk>/',
UpdateItemOnMenu.as_view(), name='menu_item_update'),
path('menu/list/item/delete/<int:pk>/',
DeleteItemOnMenu.as_view(), name='menu_item_delete'),
# Category from Menu
path('menu/category/create/<int:establishment_id>/',
CreateCategory.as_view(), name='menu_category_create'),
path('menu/category/list/<int:establishment_id>/',
ListCategory.as_view(), name='menu_category_list'),
path('menu/category/update/<int:pk>/',
UpdateCategory.as_view(), name='menu_category_update'),
path('menu/category/delete<int:pk>/',
DeleteCategory.as_view(), name='menu_category_delete'),
# Observations from Menu
path('menu/observation/create/<int:establishment_id>',
CreateObservationItem.as_view(), name='menu_observation_item_create'),
path('menu/observation/list/<int:establishment_id>/',
ListObservationItem.as_view(), name='menu_observation_list'),
path('menu/observation/update/<int:pk>/',
UpdateObservationItem.as_view(), name='menu_observation_update'),
path('menu/observation/delete/<int:pk>/',
DeleteObservationItem.as_view(), name='menu_observation_delete'),
# Menu Offers
path('menu/offer/create/<int:establishment_id>',
CreateMenuOffer.as_view(), name='menu_offer_create'),
path('menu/offer/list/<int:establishment_id>/',
ListMenuOffers.as_view(), name='menu_offer_list'),
path('menu/offer/delete/<int:pk>/',
DeleteMenuOffer.as_view(), name='menu_offer_delete'),
path('menu/offer/update/<int:pk>/',
UpdateMenuOffer.as_view(), name='menu_offer_update'),
# Url's for Orders, Bills and Tables
path('orders/list/<int:establishment_id>/',
ListOrders.as_view(), name='orders_list'),
path('orders/list/kitchen/pending/<int:establishment_id>/',
ListOrdersPendingKitchen.as_view(), name='orders_list_kitchen_pending'),
path('orders/list/kitchen/preparing/<int:establishment_id>/',
ListOrdersPreparingKitchen.as_view(), name='orders_list_kitchen_preparing'),
path('orders/list/kitchen/done/<int:establishment_id>/',
ListOrdersDoneKitchen.as_view(), name='orders_list_kitchen_done'),
# Cancel Orders Button
path('order/cancel_from_list_orders/<int:order_id>/',
CancelOrderOnListOrders.as_view(), name='order_cancel_button_on_list_orders'),
path('order/cancel_from_bill/<int:order_id>/',
CancelOrderOnListBill.as_view(), name='order_cancel_button_on_list_bills'),
# Url's for Views for Kitchen List Orders
path('orders/list/kitchen/done/<int:establishment_id>/search/user/',
ListSearchDoneOrdersByUsers.as_view(), name='orders_kitchen_done_search_user'),
path('orders/list/kitchen/done/<int:establishment_id>/search/table/',
ListFilterOrdersByTableDone.as_view(), name='orders_kitchen_done_search_table'),
path('orders/list/<int:establishment_id>/search/',
ListSearchOrders.as_view(), name='orders_search_list'),
path('orders/list/filter/category/<int:establishment_id>/search/',
KitchenFilterOrdersByCategory.as_view(), name='orders_kitchen_category_filter'),
path('orders/list/<int:establishment_id>/filter_by_table/',
ListFilterOrdersByTable.as_view(), name='orders_filter_by_table'),
path('orders/list/items/to/order/<int:establishment_id>/',
ListItemsToOrder.as_view(), name='list_items_to_order'),
path('orders/create/<int:establishment_id>/',
CreateOrder.as_view(), name='order_create'),
path('orders/update/<int:pk>/',
UpdateOrder.as_view(), name='orders_update'),
path('orders/kitchen_accepted_at/<int:order_id>/',
KitchenAcceptOrder.as_view(), name='order_kitchen_accepted_at'),
path('orders/kitchen_done_order/<int:order_id>/',
KitchenDoneOrder.as_view(), name='order_kitchen_done'),
path('orders/kitchen_cancel_order/<int:order_id>/',
KitchenCancelOrder.as_view(), name='order_kitchen_cancel'),
# Url's for Bills and BillPayment
path('bill/list/<int:establishment_id>/',
ListBillsOpened.as_view(), name='bill_list'),
path('bill/list/closed/<int:establishment_id>/',
ListBillsClosed.as_view(), name='bill_list_closed'),
path('bill/list/<int:establishment_id>/search/',
ListSearchBills.as_view(), name='bill_search_list'),
path('bill/list/search/closed/<int:establishment_id>/search/',
ListSearchBillsClosed.as_view(), name='bill_search_list_closed'),
path('bill/payment/create/<int:bill_id>/',
CreatePaymentAllBill.as_view(), name='bill_payment_create'),
path('bill/payment/create/bill_member/<int:bill_member_id>/',
CreatePaymentOnBillMember.as_view(), name='bill_member_payment_create'),
path('bill/payment/aprove_or_reject/<int:bill_payment_id>/',
ApproveOrRejectPayment.as_view(), name='bill_payment_aprove_or_reject'),
path('bill/payment/reject/<int:bill_payment_id>/',
RejectPayment.as_view(), name='bill_payment_reject'),
path('bill/bill_members/list/<int:bill_id>/',
ListBillMembersOnBill.as_view(), name='bill_member_on_bill_list'),
path('ajax/load_bill_members/',
LoadBillMembers.as_view(), name='ajax_load_bill_members'),
path('bill/orders/list/<int:bill_id>/',
ListOrdersFromBill.as_view(), name='orders_from_bill'),
# Url's for Tables and TableZone
path('table_zone/create/<int:establishment_id>/',
CreateTableZone.as_view(), name='table_zone_create'),
path('table_zone/list/<int:establishment_id>/',
ListTableZone.as_view(), name='table_zone_list'),
path('table_zone/update/<int:pk>/',
UpdateTableZone.as_view(), name='table_zone_update'),
path('table_zone/delete/<int:pk>/',
DeleteTableZone.as_view(), name='table_zone_delete'),
path('table_zone/update/active_or_desactive/<int:pk>/',
DesactiveTableZone.as_view(), name='table_zone_active_or_desactive'),
path('table/create/<int:table_zone_id>/<int:establishment_id>/',
CreateTable.as_view(), name='table_create'),
path('table/update/<int:pk>/', UpdateTable.as_view(), name='table_update'),
path('table/update/enabled/<int:pk>/',
UpdateTableEnableOrDesable.as_view(), name='table_update_enabled'),
path('table/delete/<int:pk>/', DeleteTable.as_view(), name='table_delete'),
# Url's for Operating Hours
path('operating_hours/create/<int:establishment_id>/',
CreateOperatingHours.as_view(), name='operating_hour_create'),
path('operating_hours/delete/<int:pk>/',
DeleteOperatingHour.as_view(), name='operating_hour_delete'),
# Url's for Promocodes
path('promocode/create/<int:establishment_id>/',
CreatePromoCode.as_view(), name='promocode_create'),
path('promocode/update/<int:pk>/',
UpdatePromoCodes.as_view(), name='promocode_update'),
path('promocode/delete/<int:pk>/',
DeletePromocodes.as_view(), name='promocode_delete'),
# Url's for Events
path('events/create/<int:establishment_id>/',
CreateEvents.as_view(), name='events_create'),
path('events/update/<int:pk>/',
UpdateEvents.as_view(), name='events_update'),
path('events/delete/<int:pk>/',
DeleteEvents.as_view(), name='events_delete'),
# Url's for Wirecard Payment
path('wirecard/create/<int:establishment_id>/',
CreateWirecard.as_view(), name='wirecard_create'),
path('wirecard/company/create/<int:establishment_id>/',
CreateCompanyWirecard.as_view(), name='wirecard_company_create'),
path('wirecard/detail/<int:pk>/',
DetailWirecard.as_view(), name='wirecard_detail'),
# Url's for offline Compensations
path('offline/compensations/', ListCompensations.as_view(),
name='offline_compensations'),
path('offline/compensations/check_month/<int:month>/<int:year>/<int:establishment_id>/',
CreateCompensation.as_view(), name='offline_compensations_check_month'),
path('offline/compensations/generate_report/<int:month>/<int:year>/<int:establishment_id>/', GenerateCSVReport.as_view(),
name='offline_compensations_generate_report'),
]
if settings.DEBUG:
urlpatterns += static(settings.STATIC_URL, document_root=settings.STATIC_ROOT)
urlpatterns += static(settings.MEDIA_URL, document_root=settings.MEDIA_ROOT)
| 15,173 | 5,038 |
from yacs.config import CfgNode as CN
def model(cfg):
# For model
cfg.MODEL = CN()
cfg.MODEL.num_layers = 6
cfg.MODEL.num_heads = 10
cfg.MODEL.units = 500
cfg.MODEL.inner_size = 1000
cfg.MODEL.dropout = 0.1
cfg.MODEL.tie_embedding = True
cfg.MODEL.tie_proj = False
cfg.MODEL.attention_dropout = 0.1
cfg.MODEL.pre_lnorm = False
cfg.MODEL.clamp_len = -1
cfg.MODEL.same_length = False
return cfg
def train(cfg):
# For training
cfg.TRAIN = CN()
cfg.TRAIN.load_from_previous = "Null"
cfg.TRAIN.batch_size = 200
cfg.TRAIN.batch_chunk = 1
cfg.TRAIN.tgt_length = 500
cfg.TRAIN.mem_length = 50
cfg.TRAIN.seed = 1111
cfg.TRAIN.optim = "adam"
cfg.TRAIN.lr = 0.00025 / 4.0
cfg.TRAIN.lr_min = 0.0
cfg.TRAIN.scheduler = "cosine"
cfg.TRAIN.warmup_step = 0
cfg.TRAIN.decay_rate = 0.5
cfg.TRAIN.patience = 10
cfg.TRAIN.clip = 0.25
cfg.TRAIN.max_step = 200000
cfg.TRAIN.log_interval = 200
cfg.TRAIN.eval_interval = 4000
cfg.TRAIN.pad_type = "model" # model or anything else
cfg.TRAIN.use_mle = True
cfg.TRAIN.random_crop = False
cfg.TRAIN.replace_start_with_pad = False
cfg.TRAIN.weight_decay = 0.0 # Weight decay for adam or lamb
cfg.TRAIN.append_note_status = False # Append status to event representation
return cfg
def discriminator(cfg):
# For discriminator
# Discriminator related (used only if)
cfg.DISCRIMINATOR = CN()
cfg.DISCRIMINATOR.start_iter = 100 # To control when we start training critic
cfg.DISCRIMINATOR.dis_loss_freq = 50 # How often to use loss from discriminator
cfg.DISCRIMINATOR.gen_loss_freq = 10
cfg.DISCRIMINATOR.eval_loss_freq = 10 # How often to use loss from discriminator during eval
cfg.DISCRIMINATOR.freeze_discriminator = True
cfg.DISCRIMINATOR.truncate_backprop = False # while sampling do not propagate gradients beyond current token
cfg.DISCRIMINATOR.sample_chunks_mem = 1
cfg.DISCRIMINATOR.beta_max = 100. # TODO: temperature decay
cfg.DISCRIMINATOR.adapt = 'no'
cfg.DISCRIMINATOR.type = "Null" # or cnn or Null for no discriminator or 'bert' for BERT discriminator
cfg.DISCRIMINATOR.dis_steps = 1 # dis_step per gen_step (default 1 for bert and 5 for cnn)
cfg.DISCRIMINATOR.tgt_len = 64
cfg.DISCRIMINATOR.mem_len = 64
cfg.DISCRIMINATOR.gen_loss_factor = 30 # Multiplying factor for mmd/gan loss component in generator
cfg.DISCRIMINATOR.dis_loss_factor = 1 # Multiplying factor for mmd/gan loss component in discriminator
cfg.DISCRIMINATOR.batch_chunk = 1
cfg.DISCRIMINATOR.context_len = 5 # Randomly sample context length tokens from real data and use as context.
cfg.DISCRIMINATOR.backprop_outside = True
cfg.DISCRIMINATOR.src_mem_len = 200
# If 0 uses first token in real data
cfg.DISCRIMINATOR.gen_scheduler = "constant"
cfg.DISCRIMINATOR.gen_lr_min = 0.0
cfg.DISCRIMINATOR.gen_warmup_step = 0
cfg.DISCRIMINATOR.gen_decay_rate = 0.5
cfg.DISCRIMINATOR.gen_patience = 10
cfg.DISCRIMINATOR.gen_lr = 0.00025 / 4.0
cfg.DISCRIMINATOR.dis_scheduler = "constant"
cfg.DISCRIMINATOR.dis_lr_min = 0.0
cfg.DISCRIMINATOR.dis_warmup_step = 0
cfg.DISCRIMINATOR.dis_decay_rate = 0.5
cfg.DISCRIMINATOR.dis_patience = 10
cfg.DISCRIMINATOR.dis_lr = 0.00025 / 4.0
# Bert params
cfg.DISCRIMINATOR.BERT = CN()
cfg.DISCRIMINATOR.BERT.learning_rate = 1e-5 # Decrease learning rate since we're fine tuning
cfg.DISCRIMINATOR.BERT.weight_decay = 0.0
cfg.DISCRIMINATOR.BERT.adam_epsilon = 1e-8
cfg.DISCRIMINATOR.BERT.max_grad_norm = 1.0
cfg.DISCRIMINATOR.BERT.model_type = "bert_lm" # or "bert_cls"
cfg.DISCRIMINATOR.BERT.loss_type = "rsgan" # or 'standard’,'JS', 'KL', 'hinge', 'tv', 'rsgan', 'wgan-gp', "mmd", 'ppo', 'ppo-gp'
cfg.DISCRIMINATOR.BERT.model_path = "../BERT/checkpoint-1969000"
cfg.DISCRIMINATOR.BERT.freeze_layers = [] # Total layers ['0', '1', '2', '3', '4']
cfg.DISCRIMINATOR.BERT.random_weights = False # only implemented for bert_lm
# CNN params (Relgan)
cfg.DISCRIMINATOR.CNN = CN()
cfg.DISCRIMINATOR.CNN.learning_rate = 1e-4
cfg.DISCRIMINATOR.CNN.embed_dim = 64
cfg.DISCRIMINATOR.CNN.hidden_dim = 64
cfg.DISCRIMINATOR.CNN.num_rep = 64
cfg.DISCRIMINATOR.CNN.init = "uniform"
cfg.DISCRIMINATOR.CNN.loss_type = "rsgan" # or 'standard’,'JS', 'KL', 'hinge', 'tv', 'rsgan', 'wgan-gp', "mmd", "ppo-gp"
return cfg
def metric(cfg):
# Metrics
cfg.METRICS = CN()
cfg.METRICS.use_bleu = False # outdated
cfg.METRICS.use_self_bleu = False # outdated
cfg.METRICS.CLASSIFIER = CN()
cfg.METRICS.CLASSIFIER.use_classifier = False
cfg.METRICS.CLASSIFIER.gen_batch_size = 128
cfg.METRICS.CLASSIFIER.gen_seq_len = 2048
cfg.METRICS.CLASSIFIER.gen_num_samples = 256
cfg.METRICS.CLASSIFIER.block_size = 128 # For training classifier
cfg.METRICS.CLASSIFIER.bert_batch_size = 20 # For passing into bert
cfg.METRICS.CLASSIFIER.model_path = "../BERT/checkpoint-1969000"
return cfg
def init(cfg):
# For initialization
cfg.INITIALIZER = CN()
cfg.INITIALIZER.base_init = ["normal", 0.01]
cfg.INITIALIZER.embed_init = ["normal", 0.01]
# For evaluation
cfg.EVALUATE = CN()
cfg.EVALUATE.batch_size = 10
cfg.EVALUATE.tgt_length = 128
cfg.EVALUATE.mem_length = 128
# Event type related
cfg.DATASET = CN()
cfg.DATASET.event_type = "magenta" # or 'newevent'
cfg.DATASET.trim_padding = False
# Classifier related
cfg.PPO = CN() # For ppo loss type
cfg.PPO.dis_D_lr = 0.00025 / 4.0
cfg.PPO.dis_D_update_D0_freq = 20 # Should be multiple of gen_loss_freq
cfg.PPO.dis_D_type = "bert" # bert or cnn
cfg.PPO.clip_param = 0.4
cfg.PPO.dis_D_num_rep = 1
# For Problem Type
cfg.PROBLEM = CN()
cfg.PROBLEM.type = 'Null' # time extension: Null
cfg.PROBLEM.melody_len = 1024
return cfg
def get_default_cfg_training():
cfg = CN()
cfg = init(cfg)
cfg = model(cfg)
cfg = train(cfg)
cfg = discriminator(cfg)
cfg = metric(cfg)
cfg.freeze()
return cfg
| 6,220 | 2,834 |
import argbind
import typing
@argbind.bind()
def func(
arg1 : str = 'default',
arg2 : str = 'default',
arg3 : str = 'default',
arg4 : str = 'default',
arg5 : typing.List[str] = ['default'],
):
"""Dummy function for binding.
Parameters
----------
arg1 : str, optional
Argument 1, by default 'default'
arg2 : str, optional
Argument 2, by default 'default'
arg3 : str, optional
Argument 3, by default 'default'
arg4 : str, optional
Argument 4, by default 'default'
"""
print(
f"Argument 1: {arg1}\n"
f"Argument 2: {arg2}\n"
f"Argument 3: {arg3}\n"
f"Argument 4: {arg4}\n"
f"Argument 5: {arg5}"
)
if __name__ == "__main__":
args = argbind.parse_args()
with argbind.scope(args):
func()
| 832 | 287 |
from handlers import tables, helper, Handler
import time
class ArticleHandler(Handler):
def __init__(self, *args, **kwargs):
super(ArticleHandler, self).__init__(*args, **kwargs)
self.body_class = 'article-page'
def get(self, article_id):
if not article_id.isdigit():
self.page_redirect("/")
else:
article = tables.articles.get(article_id)
comments = tables.comments.get_comments(article_id)
self.render('article.jinja', handler=self,
article=article, comments=comments)
def post(self, article_id):
if not article_id.isdigit() or not self.is_loggedin():
self.page_redirect("/")
else:
like = self.request.get("like")
new_comment = self.request.get("new-comment")
delete_comment = self.request.get("delete-comment")
edit_comment = self.request.get("edit-comment")
if like:
username = self.get_cookie("username")
article = tables.articles.get(article_id)
if self.is_loggedin() and username != article.user:
if tables.likes.exist(article_id, username):
tables.likes.delete(article_id, username)
else:
tables.likes.add(article_id, username)
self.page_redirect("/article/%s/#like" % article_id)
elif new_comment:
new_comment = self.request.get("comment")
username = self.get_cookie("username")
if self.is_loggedin():
tables.comments.add(article_id, username, new_comment)
self.page_redirect("/article/%s/#comments" % article_id)
elif delete_comment:
comment_id = self.request.get("comment-id")
comment = tables.comments.get(comment_id)
if self.is_loggedin() == comment.user:
tables.comments.delete(comment_id)
self.page_redirect("/article/%s/#comments" % article_id)
elif edit_comment:
comment_id = self.request.get("comment-id")
comment = self.request.get("comment")
com = tables.comments.get(comment_id)
if self.is_loggedin() == com.user:
tables.comments.edit(comment_id, comment)
self.page_redirect("/article/%s/#comments" % article_id)
else:
self.page_redirect("/article/%s/" % article_id)
| 2,585 | 681 |
import FWCore.ParameterSet.Config as cms
MuonAlignmentPreFilter = cms.EDFilter("MuonAlignmentPreFilter",
tracksTag = cms.InputTag("ALCARECOMuAlCalIsolatedMu:GlobalMuon"),
minTrackPt = cms.double(20.),
minTrackP = cms.double(0.),
minTrackerHits = cms.int32(10),
minDTHits = cms.int32(6),
minCSCHits = cms.int32(4),
allowTIDTEC = cms.bool(True),
minTrackEta = cms.double(-2.4),
maxTrackEta = cms.double(2.4)
)
| 428 | 193 |
#!/usr/bin/env python
import wx
import wx.lib.mixins.listctrl as listmix
from six import unichr
#----------------------------------------------------------------------
keyMap = {
wx.WXK_BACK : "WXK_BACK",
wx.WXK_TAB : "WXK_TAB",
wx.WXK_RETURN : "WXK_RETURN",
wx.WXK_ESCAPE : "WXK_ESCAPE",
wx.WXK_SPACE : "WXK_SPACE",
wx.WXK_DELETE : "WXK_DELETE",
wx.WXK_START : "WXK_START",
wx.WXK_LBUTTON : "WXK_LBUTTON",
wx.WXK_RBUTTON : "WXK_RBUTTON",
wx.WXK_CANCEL : "WXK_CANCEL",
wx.WXK_MBUTTON : "WXK_MBUTTON",
wx.WXK_CLEAR : "WXK_CLEAR",
wx.WXK_SHIFT : "WXK_SHIFT",
wx.WXK_ALT : "WXK_ALT",
wx.WXK_MENU : "WXK_MENU",
wx.WXK_PAUSE : "WXK_PAUSE",
wx.WXK_CAPITAL : "WXK_CAPITAL",
#wx.WXK_PRIOR : "WXK_PRIOR",
#wx.WXK_NEXT : "WXK_NEXT",
wx.WXK_END : "WXK_END",
wx.WXK_HOME : "WXK_HOME",
wx.WXK_LEFT : "WXK_LEFT",
wx.WXK_UP : "WXK_UP",
wx.WXK_RIGHT : "WXK_RIGHT",
wx.WXK_DOWN : "WXK_DOWN",
wx.WXK_SELECT : "WXK_SELECT",
wx.WXK_PRINT : "WXK_PRINT",
wx.WXK_EXECUTE : "WXK_EXECUTE",
wx.WXK_SNAPSHOT : "WXK_SNAPSHOT",
wx.WXK_INSERT : "WXK_INSERT",
wx.WXK_HELP : "WXK_HELP",
wx.WXK_NUMPAD0 : "WXK_NUMPAD0",
wx.WXK_NUMPAD1 : "WXK_NUMPAD1",
wx.WXK_NUMPAD2 : "WXK_NUMPAD2",
wx.WXK_NUMPAD3 : "WXK_NUMPAD3",
wx.WXK_NUMPAD4 : "WXK_NUMPAD4",
wx.WXK_NUMPAD5 : "WXK_NUMPAD5",
wx.WXK_NUMPAD6 : "WXK_NUMPAD6",
wx.WXK_NUMPAD7 : "WXK_NUMPAD7",
wx.WXK_NUMPAD8 : "WXK_NUMPAD8",
wx.WXK_NUMPAD9 : "WXK_NUMPAD9",
wx.WXK_MULTIPLY : "WXK_MULTIPLY",
wx.WXK_ADD : "WXK_ADD",
wx.WXK_SEPARATOR : "WXK_SEPARATOR",
wx.WXK_SUBTRACT : "WXK_SUBTRACT",
wx.WXK_DECIMAL : "WXK_DECIMAL",
wx.WXK_DIVIDE : "WXK_DIVIDE",
wx.WXK_F1 : "WXK_F1",
wx.WXK_F2 : "WXK_F2",
wx.WXK_F3 : "WXK_F3",
wx.WXK_F4 : "WXK_F4",
wx.WXK_F5 : "WXK_F5",
wx.WXK_F6 : "WXK_F6",
wx.WXK_F7 : "WXK_F7",
wx.WXK_F8 : "WXK_F8",
wx.WXK_F9 : "WXK_F9",
wx.WXK_F10 : "WXK_F10",
wx.WXK_F11 : "WXK_F11",
wx.WXK_F12 : "WXK_F12",
wx.WXK_F13 : "WXK_F13",
wx.WXK_F14 : "WXK_F14",
wx.WXK_F15 : "WXK_F15",
wx.WXK_F16 : "WXK_F16",
wx.WXK_F17 : "WXK_F17",
wx.WXK_F18 : "WXK_F18",
wx.WXK_F19 : "WXK_F19",
wx.WXK_F20 : "WXK_F20",
wx.WXK_F21 : "WXK_F21",
wx.WXK_F22 : "WXK_F22",
wx.WXK_F23 : "WXK_F23",
wx.WXK_F24 : "WXK_F24",
wx.WXK_NUMLOCK : "WXK_NUMLOCK",
wx.WXK_SCROLL : "WXK_SCROLL",
wx.WXK_PAGEUP : "WXK_PAGEUP",
wx.WXK_PAGEDOWN : "WXK_PAGEDOWN",
wx.WXK_NUMPAD_SPACE : "WXK_NUMPAD_SPACE",
wx.WXK_NUMPAD_TAB : "WXK_NUMPAD_TAB",
wx.WXK_NUMPAD_ENTER : "WXK_NUMPAD_ENTER",
wx.WXK_NUMPAD_F1 : "WXK_NUMPAD_F1",
wx.WXK_NUMPAD_F2 : "WXK_NUMPAD_F2",
wx.WXK_NUMPAD_F3 : "WXK_NUMPAD_F3",
wx.WXK_NUMPAD_F4 : "WXK_NUMPAD_F4",
wx.WXK_NUMPAD_HOME : "WXK_NUMPAD_HOME",
wx.WXK_NUMPAD_LEFT : "WXK_NUMPAD_LEFT",
wx.WXK_NUMPAD_UP : "WXK_NUMPAD_UP",
wx.WXK_NUMPAD_RIGHT : "WXK_NUMPAD_RIGHT",
wx.WXK_NUMPAD_DOWN : "WXK_NUMPAD_DOWN",
#wx.WXK_NUMPAD_PRIOR : "WXK_NUMPAD_PRIOR",
wx.WXK_NUMPAD_PAGEUP : "WXK_NUMPAD_PAGEUP",
#wx.WXK_NUMPAD_NEXT : "WXK_NUMPAD_NEXT",
wx.WXK_NUMPAD_PAGEDOWN : "WXK_NUMPAD_PAGEDOWN",
wx.WXK_NUMPAD_END : "WXK_NUMPAD_END",
wx.WXK_NUMPAD_BEGIN : "WXK_NUMPAD_BEGIN",
wx.WXK_NUMPAD_INSERT : "WXK_NUMPAD_INSERT",
wx.WXK_NUMPAD_DELETE : "WXK_NUMPAD_DELETE",
wx.WXK_NUMPAD_EQUAL : "WXK_NUMPAD_EQUAL",
wx.WXK_NUMPAD_MULTIPLY : "WXK_NUMPAD_MULTIPLY",
wx.WXK_NUMPAD_ADD : "WXK_NUMPAD_ADD",
wx.WXK_NUMPAD_SEPARATOR : "WXK_NUMPAD_SEPARATOR",
wx.WXK_NUMPAD_SUBTRACT : "WXK_NUMPAD_SUBTRACT",
wx.WXK_NUMPAD_DECIMAL : "WXK_NUMPAD_DECIMAL",
wx.WXK_NUMPAD_DIVIDE : "WXK_NUMPAD_DIVIDE",
wx.WXK_WINDOWS_LEFT : "WXK_WINDOWS_LEFT",
wx.WXK_WINDOWS_RIGHT : "WXK_WINDOWS_RIGHT",
wx.WXK_WINDOWS_MENU : "WXK_WINDOWS_MENU",
wx.WXK_SPECIAL1 : "WXK_SPECIAL1",
wx.WXK_SPECIAL2 : "WXK_SPECIAL2",
wx.WXK_SPECIAL3 : "WXK_SPECIAL3",
wx.WXK_SPECIAL4 : "WXK_SPECIAL4",
wx.WXK_SPECIAL5 : "WXK_SPECIAL5",
wx.WXK_SPECIAL6 : "WXK_SPECIAL6",
wx.WXK_SPECIAL7 : "WXK_SPECIAL7",
wx.WXK_SPECIAL8 : "WXK_SPECIAL8",
wx.WXK_SPECIAL9 : "WXK_SPECIAL9",
wx.WXK_SPECIAL10 : "WXK_SPECIAL10",
wx.WXK_SPECIAL11 : "WXK_SPECIAL11",
wx.WXK_SPECIAL12 : "WXK_SPECIAL12",
wx.WXK_SPECIAL13 : "WXK_SPECIAL13",
wx.WXK_SPECIAL14 : "WXK_SPECIAL14",
wx.WXK_SPECIAL15 : "WXK_SPECIAL15",
wx.WXK_SPECIAL16 : "WXK_SPECIAL16",
wx.WXK_SPECIAL17 : "WXK_SPECIAL17",
wx.WXK_SPECIAL18 : "WXK_SPECIAL18",
wx.WXK_SPECIAL19 : "WXK_SPECIAL19",
}
if 'wxMac' in wx.PlatformInfo:
keyMap[wx.WXK_RAW_CONTROL] = 'WXK_RAW_CONTROL'
keyMap[wx.WXK_CONTROL] = "WXK_CONTROL"
keyMap[wx.WXK_COMMAND] = "WXK_COMMAND"
else:
keyMap[wx.WXK_COMMAND] = "WXK_COMMAND"
keyMap[wx.WXK_CONTROL] = "WXK_CONTROL"
#----------------------------------------------------------------------
class KeySink(wx.Window):
def __init__(self, parent):
wx.Window.__init__(self, parent, -1, style=wx.WANTS_CHARS
#| wx.RAISED_BORDER
#| wx.SUNKEN_BORDER
, name="sink")
self.SetBackgroundColour(wx.BLUE)
self.haveFocus = False
self.callSkip = True
self.logKeyDn = True
self.logKeyUp = True
self.logChar = True
self.Bind(wx.EVT_PAINT, self.OnPaint)
self.Bind(wx.EVT_SET_FOCUS, self.OnSetFocus)
self.Bind(wx.EVT_KILL_FOCUS, self.OnKillFocus)
self.Bind(wx.EVT_MOUSE_EVENTS, self.OnMouse)
self.Bind(wx.EVT_KEY_DOWN, self.OnKeyDown)
self.Bind(wx.EVT_KEY_UP, self.OnKeyUp)
self.Bind(wx.EVT_CHAR, self.OnChar)
def SetCallSkip(self, skip):
self.callSkip = skip
def SetLogKeyUp(self, val):
self.logKeyUp = val
def SetLogKeyDn(self, val):
self.logKeyDn = val
def SetLogChar(self, val):
self.logChar = val
def OnPaint(self, evt):
dc = wx.PaintDC(self)
rect = self.GetClientRect()
dc.SetTextForeground(wx.WHITE)
dc.DrawLabel("Click here and then press some keys",
rect, wx.ALIGN_CENTER | wx.ALIGN_TOP)
if self.haveFocus:
dc.SetTextForeground(wx.GREEN)
dc.DrawLabel("Have Focus", rect, wx.ALIGN_RIGHT | wx.ALIGN_BOTTOM)
else:
dc.SetTextForeground(wx.RED)
dc.DrawLabel("Need Focus!", rect, wx.ALIGN_RIGHT | wx.ALIGN_BOTTOM)
def OnSetFocus(self, evt):
self.haveFocus = True
self.Refresh()
def OnKillFocus(self, evt):
self.haveFocus = False
self.Refresh()
def OnMouse(self, evt):
if evt.ButtonDown():
self.SetFocus()
def OnKeyDown(self, evt):
if self.logKeyDn:
self.GetParent().keylog.LogKeyEvent("KeyDown", evt)
if self.callSkip:
evt.Skip()
def OnKeyUp(self, evt):
if self.logKeyUp:
self.GetParent().keylog.LogKeyEvent("KeyUp", evt)
if self.callSkip:
evt.Skip()
def OnChar(self, evt):
if self.logChar:
self.GetParent().keylog.LogKeyEvent("Char", evt)
#----------------------------------------------------------------------
class KeyLog(wx.ListCtrl, listmix.ListCtrlAutoWidthMixin):
colHeaders = [ "Event Type",
"Key Name",
"Key Code",
"Modifiers",
"Unicode",
"UniChr",
"RawKeyCode",
"RawKeyFlags",
]
def __init__(self, parent):
wx.ListCtrl.__init__(self, parent, -1,
style = wx.LC_REPORT|wx.LC_VRULES|wx.LC_HRULES)
listmix.ListCtrlAutoWidthMixin.__init__(self)
for idx, header in enumerate(self.colHeaders):
self.InsertColumn(idx, header)
idx += 1
self.InsertColumn(idx, "")
for x in range(idx):
self.SetColumnWidth(x, wx.LIST_AUTOSIZE_USEHEADER)
self.SetColumnWidth(1, 125)
def LogKeyEvent(self, evType, evt):
keycode = evt.GetKeyCode()
keyname = keyMap.get(keycode, None)
if keyname is None:
if keycode < 256:
if keycode == 0:
keyname = "NUL"
elif keycode < 27:
keyname = u"Ctrl-%s" % unichr(ord('A') + keycode-1)
else:
keyname = u"\"%s\"" % unichr(keycode)
else:
keyname = u"(%s)" % keycode
UniChr = ''
if "unicode" in wx.PlatformInfo:
UniChr = "\"" + unichr(evt.GetUnicodeKey()) + "\""
modifiers = ""
for mod, ch in [(evt.ControlDown(), 'C'),
(evt.AltDown(), 'A'),
(evt.ShiftDown(), 'S'),
(evt.MetaDown(), 'M'),
(evt.RawControlDown(), 'R'),]:
if mod:
modifiers += ch
else:
modifiers += '-'
id = self.InsertItem(self.GetItemCount(), evType)
self.SetItem(id, 1, keyname)
self.SetItem(id, 2, str(keycode))
self.SetItem(id, 3, modifiers)
self.SetItem(id, 4, str(evt.GetUnicodeKey()))
self.SetItem(id, 5, UniChr)
self.SetItem(id, 6, str(evt.GetRawKeyCode()))
self.SetItem(id, 7, str(evt.GetRawKeyFlags()))
self.EnsureVisible(id)
def ClearLog(self):
self.DeleteAllItems()
def CopyLog(self):
# build a newline and tab delimited string to put into the clipboard
if "unicode" in wx.PlatformInfo:
st = u""
else:
st = ""
for h in self.colHeaders:
st += h + "\t"
st += "\n"
for idx in range(self.GetItemCount()):
for col in range(self.GetColumnCount()):
item = self.GetItem(idx, col)
st += item.GetText() + "\t"
st += "\n"
data = wx.TextDataObject()
data.SetText(st)
if wx.TheClipboard.Open():
wx.TheClipboard.SetData(data)
wx.TheClipboard.Close()
else:
wx.MessageBox("Unable to open the clipboard", "Error")
#----------------------------------------------------------------------
class TestPanel(wx.Panel):
def __init__(self, parent, log):
self.log = log
wx.Panel.__init__(self, parent, -1, style=0)
self.keysink = KeySink(self)
self.keysink.SetMinSize((100, 65))
self.keylog = KeyLog(self)
btn = wx.Button(self, -1, "Clear", style=wx.BU_EXACTFIT)
self.Bind(wx.EVT_BUTTON, self.OnClearBtn, btn)
btn.SetToolTip(
"Clear the items from the log window")
btn2 = wx.Button(self, -1, "Copy", style=wx.BU_EXACTFIT)
self.Bind(wx.EVT_BUTTON, self.OnCopyBtn, btn2)
btn2.SetToolTip(
"Copy the contents of the log window to the clipboard")
cb1 = wx.CheckBox(self, -1, "Call evt.Skip in Key* events")
self.Bind(wx.EVT_CHECKBOX, self.OnSkipCB, cb1)
cb1.SetValue(True)
cb2 = wx.CheckBox(self, -1, "KEY_UP")
self.Bind(wx.EVT_CHECKBOX, self.OnKeyUpCB, cb2)
cb2.SetValue(True)
cb3 = wx.CheckBox(self, -1, "KEY_DOWN")
self.Bind(wx.EVT_CHECKBOX, self.OnKeyDnCB, cb3)
cb3.SetValue(True)
cb4 = wx.CheckBox(self, -1, "CHAR")
self.Bind(wx.EVT_CHECKBOX, self.OnCharCB, cb4)
cb4.SetValue(True)
buttons = wx.BoxSizer(wx.HORIZONTAL)
buttons.Add(btn, 0, wx.ALL, 4)
buttons.Add(btn2, 0, wx.ALL, 4)
buttons.Add(cb1, 0, wx.ALIGN_CENTER_VERTICAL|wx.LEFT|wx.RIGHT, 6)
buttons.Add(cb2, 0, wx.ALIGN_CENTER_VERTICAL|wx.LEFT, 6)
buttons.Add(cb3, 0, wx.ALIGN_CENTER_VERTICAL|wx.LEFT, 6)
buttons.Add(cb4, 0, wx.ALIGN_CENTER_VERTICAL|wx.LEFT, 6)
sizer = wx.BoxSizer(wx.VERTICAL)
sizer.Add(self.keysink, 0, wx.GROW)
sizer.Add(buttons)
sizer.Add(self.keylog, 1, wx.GROW)
self.SetSizer(sizer)
def OnClearBtn(self, evt):
self.keylog.ClearLog()
def OnCopyBtn(self, evt):
self.keylog.CopyLog()
def OnSkipCB(self, evt):
self.keysink.SetCallSkip(evt.GetInt())
def OnKeyUpCB(self, evt):
self.keysink.SetLogKeyUp(evt.GetInt())
def OnKeyDnCB(self, evt):
self.keysink.SetLogKeyDn(evt.GetInt())
def OnCharCB(self, evt):
self.keysink.SetLogChar(evt.GetInt())
#----------------------------------------------------------------------
def runTest(frame, nb, log):
win = TestPanel(nb, log)
return win
#----------------------------------------------------------------------
overview = """<html><body>
<h2><center>wxKeyEvents</center></h2>
This demo simply catches all key events and prints info about them.
It is meant to be used as a compatibility test for cross platform work.
</body></html>
"""
if __name__ == '__main__':
import sys,os
import run
run.main(['', os.path.basename(sys.argv[0])] + sys.argv[1:])
| 13,299 | 5,989 |
import time
import tensorflow
from tensorflow import keras
class TimeCallback(keras.callbacks.Callback):
def __init__ (self):
super(TimeCallback, self).__init__()
def on_epoch_begin(self, epoch, logs):
self.epoch_time_start = time.time()
def on_epoch_end(self, batch, logs):
logs['time'] = time.time() - self.epoch_time_start
| 373 | 124 |
from django.urls import reverse
from rest_framework.test import APITestCase, APIClient
from rest_framework.views import status
from movies.models import Movies
from movies.serializers import MoviesSerializer
from user.models import User
import json
class BaseViewTest(APITestCase):
client = APIClient()
def create_movie(self, title="", genre="", cast="", director=""):
"""create a movie"""
if title != "" and genre != "" and cast!= "" and director != "":
return Movies.objects.create(title=title, genre=genre, cast=cast, director=director)
else:
print("complete data")
def movie_request(self, kind="post", **kwargs):
"""Post create movie and put"""
if kind == "post":
return self.client.post(reverse("movies-all"),
data=json.dumps(kwargs["data"]),
content_type='application/json'
)
elif kind == "put":
return self.client.put(
reverse(
"movies-detail",
kwargs={"pk" : kwargs["id"]}
),
data=json.dumps(kwargs["data"]),
content_type='application/json'
)
else:
return None
def retrieve_movie(self, pk=0):
return self.client.get(
reverse(
"movies-detail",
kwargs={"pk" : pk}
)
)
def delete_movie(self, pk=0):
return self.client.delete(
reverse(
"movies-detail",
kwargs={"pk" : pk}
)
)
def setUp(self):
"""Add test data"""
self.movie_1 = self.create_movie(title="Fast_and_Furious", genre="Action", cast="Dwayne_Johnson", director="flata")
self.create_movie(title="The_lion_king", genre="Drama", cast="Donal_Glover", director='st')
self.create_movie(title="The_mummy", genre="Horror", cast="Brendan_Fraser", director='md')
self.valid_movie_id = self.movie_1.id
self.invalid_movie_id = 50
"""create a user"""
self.user = User.objects.create_superuser(
username="test",
email="test@gmail.com",
password="test123",
first_name="first name",
last_name="last name",
is_active=True,
)
url = reverse('user:login')
data = {
"email": "test@gmail.com",
"password": "test123",
}
res = self.client.post(url, data=data, format='json')
self.assertEqual(res.status_code, status.HTTP_200_OK, res.content)
token=res.json().get('token')
self.client.credentials(HTTP_AUTHORIZATION='Bearer {0}'.format(token))
class GetAllMoviesTest(BaseViewTest):
def test_get_all_movies(self):
"""
This test ensures that all movies added in the setUp method
exist when we make a GET request to the movies/ endpoint
"""
#self.login_client("test@gmail.com", "test123")
response = self.client.get(
reverse("movies-all")
)
expected = Movies.objects.all()
serialized = MoviesSerializer(expected, many=True)
self.assertEqual(response.data, serialized.data)
self.assertEqual(response.status_code, status.HTTP_200_OK)
class GetASingleMovieTest(BaseViewTest):
def test_get_a_movie(self):
"""Test movie with id exist"""
#self.login_client("test@gmail.com", "test123")
response = self.retrieve_movie(self.valid_movie_id)
expected = Movies.objects.get(pk=self.valid_movie_id)
serialized = MoviesSerializer(expected)
self.assertEqual(response.data, serialized.data)
self.assertEqual(response.status_code, status.HTTP_200_OK)
response = self.retrieve_movie(self.invalid_movie_id)
self.assertEqual(
response.data["message"],
"Movie with id: 50 does not exist"
)
self.assertEqual(response.status_code, status.HTTP_404_NOT_FOUND)
| 4,071 | 1,216 |
# %%
import numpy as np # numeric Python
from HARK.utilities import plotFuncs # basic plotting tools
from HARK.ConsumptionSaving.ConsMarkovModel import (
MarkovConsumerType,
) # An alternative, much longer way to solve the TBS model
from time import process_time # timing utility
from HARK.ConsumptionSaving.TractableBufferStockModel import TractableConsumerType
do_simulation = True
# %%
# Define the model primitives
base_primitives = {
"UnempPrb": 0.00625, # Probability of becoming unemployed
"DiscFac": 0.975, # Intertemporal discount factor
"Rfree": 1.01, # Risk-free interest factor on assets
"PermGroFac": 1.0025, # Permanent income growth factor (uncompensated)
"CRRA": 1.0,
} # Coefficient of relative risk aversion
# %%
# Define a dictionary to be used in case of simulation
simulation_values = {
"aLvlInitMean": 0.0, # Mean of log initial assets for new agents
"aLvlInitStd": 1.0, # Stdev of log initial assets for new agents
"AgentCount": 10000, # Number of agents to simulate
"T_sim": 120, # Number of periods to simulate
"T_cycle": 1,
} # Number of periods in the cycle
# %%
# Make and solve a tractable consumer type
ExampleType = TractableConsumerType(**base_primitives)
t_start = process_time()
ExampleType.solve()
t_end = process_time()
print(
"Solving a tractable consumption-savings model took "
+ str(t_end - t_start)
+ " seconds."
)
# %%
# Plot the consumption function and whatnot
m_upper = 1.5 * ExampleType.mTarg
conFunc_PF = lambda m: ExampleType.h * ExampleType.PFMPC + ExampleType.PFMPC * m
# plotFuncs([ExampleType.solution[0].cFunc,ExampleType.mSSfunc,ExampleType.cSSfunc],0,m_upper)
plotFuncs([ExampleType.solution[0].cFunc, ExampleType.solution[0].cFunc_U], 0, m_upper)
# %%
if do_simulation:
ExampleType(**simulation_values) # Set attributes needed for simulation
ExampleType.track_vars = ["mLvlNow"]
ExampleType.makeShockHistory()
ExampleType.initializeSim()
ExampleType.simulate()
# %%
# Now solve the same model using backward induction rather than the analytic method of TBS.
# The TBS model is equivalent to a Markov model with two states, one of them absorbing (permanent unemployment).
MrkvArray = np.array(
[[1.0 - base_primitives["UnempPrb"], base_primitives["UnempPrb"]], [0.0, 1.0]]
) # Define the two state, absorbing unemployment Markov array
init_consumer_objects = {
"CRRA": base_primitives["CRRA"],
"Rfree": np.array(
2 * [base_primitives["Rfree"]]
), # Interest factor (same in both states)
"PermGroFac": [
np.array(
2 * [base_primitives["PermGroFac"] / (1.0 - base_primitives["UnempPrb"])]
)
], # Unemployment-compensated permanent growth factor
"BoroCnstArt": None, # Artificial borrowing constraint
"PermShkStd": [0.0], # Permanent shock standard deviation
"PermShkCount": 1, # Number of shocks in discrete permanent shock distribution
"TranShkStd": [0.0], # Transitory shock standard deviation
"TranShkCount": 1, # Number of shocks in discrete permanent shock distribution
"T_cycle": 1, # Number of periods in cycle
"UnempPrb": 0.0, # Unemployment probability (not used, as the unemployment here is *permanent*, not transitory)
"UnempPrbRet": 0.0, # Unemployment probability when retired (irrelevant here)
"T_retire": 0, # Age at retirement (turned off)
"IncUnemp": 0.0, # Income when unemployed (irrelevant)
"IncUnempRet": 0.0, # Income when unemployed and retired (irrelevant)
"aXtraMin": 0.001, # Minimum value of assets above minimum in grid
"aXtraMax": ExampleType.mUpperBnd, # Maximum value of assets above minimum in grid
"aXtraCount": 48, # Number of points in assets grid
"aXtraExtra": [None], # Additional points to include in assets grid
"aXtraNestFac": 3, # Degree of exponential nesting when constructing assets grid
"LivPrb": [np.array([1.0, 1.0])], # Survival probability
"DiscFac": base_primitives["DiscFac"], # Intertemporal discount factor
"AgentCount": 1, # Number of agents in a simulation (irrelevant)
"tax_rate": 0.0, # Tax rate on labor income (irrelevant)
"vFuncBool": False, # Whether to calculate the value function
"CubicBool": True, # Whether to use cubic splines (False --> linear splines)
"MrkvArray": [MrkvArray], # State transition probabilities
}
MarkovType = MarkovConsumerType(**init_consumer_objects) # Make a basic consumer type
employed_income_dist = [
np.ones(1),
np.ones(1),
np.ones(1),
] # Income distribution when employed
unemployed_income_dist = [
np.ones(1),
np.ones(1),
np.zeros(1),
] # Income distribution when permanently unemployed
MarkovType.IncomeDstn = [
[employed_income_dist, unemployed_income_dist]
] # set the income distribution in each state
MarkovType.cycles = 0
# %%
# Solve the "Markov TBS" model
t_start = process_time()
MarkovType.solve()
t_end = process_time()
MarkovType.unpackcFunc()
# %%
print(
'Solving the same model "the long way" took ' + str(t_end - t_start) + " seconds."
)
# plotFuncs([ExampleType.solution[0].cFunc,ExampleType.solution[0].cFunc_U],0,m_upper)
plotFuncs(MarkovType.cFunc[0], 0, m_upper)
diffFunc = lambda m: ExampleType.solution[0].cFunc(m) - MarkovType.cFunc[0][0](m)
print("Difference between the (employed) consumption functions:")
plotFuncs(diffFunc, 0, m_upper)
| 5,426 | 1,895 |
"""Util functions
Extended from original PANet code
TODO: move part of dataset configurations to data_utils
"""
import random
import torch
import numpy as np
import operator
def set_seed(seed):
"""
Set the random seed
"""
random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
CLASS_LABELS = {
'SABS': {
'pa_all': set( [1,2,3,6] ),
0: set([1,6] ), # upper_abdomen: spleen + liver as training, kidneis are testing
1: set( [2,3] ), # lower_abdomen
},
'C0': {
'pa_all': set(range(1, 4)),
0: set([2,3]),
1: set([1,3]),
2: set([1,2]),
},
'CHAOST2': {
'pa_all': set(range(1, 5)),
0: set([1, 4]), # upper_abdomen, leaving kidneies as testing classes
1: set([2, 3]), # lower_abdomen
},
}
def get_bbox(fg_mask, inst_mask):
"""
Get the ground truth bounding boxes
"""
fg_bbox = torch.zeros_like(fg_mask, device=fg_mask.device)
bg_bbox = torch.ones_like(fg_mask, device=fg_mask.device)
inst_mask[fg_mask == 0] = 0
area = torch.bincount(inst_mask.view(-1))
cls_id = area[1:].argmax() + 1
cls_ids = np.unique(inst_mask)[1:]
mask_idx = np.where(inst_mask[0] == cls_id)
y_min = mask_idx[0].min()
y_max = mask_idx[0].max()
x_min = mask_idx[1].min()
x_max = mask_idx[1].max()
fg_bbox[0, y_min:y_max+1, x_min:x_max+1] = 1
for i in cls_ids:
mask_idx = np.where(inst_mask[0] == i)
y_min = max(mask_idx[0].min(), 0)
y_max = min(mask_idx[0].max(), fg_mask.shape[1] - 1)
x_min = max(mask_idx[1].min(), 0)
x_max = min(mask_idx[1].max(), fg_mask.shape[2] - 1)
bg_bbox[0, y_min:y_max+1, x_min:x_max+1] = 0
return fg_bbox, bg_bbox
def t2n(img_t):
"""
torch to numpy regardless of whether tensor is on gpu or memory
"""
if img_t.is_cuda:
return img_t.data.cpu().numpy()
else:
return img_t.data.numpy()
def to01(x_np):
"""
normalize a numpy to 0-1 for visualize
"""
return (x_np - x_np.min()) / (x_np.max() - x_np.min() + 1e-5)
def compose_wt_simple(is_wce, data_name):
"""
Weights for cross-entropy loss
"""
if is_wce:
if data_name in ['SABS', 'SABS_Superpix', 'C0', 'C0_Superpix', 'CHAOST2', 'CHAOST2_Superpix']:
return torch.FloatTensor([0.05, 1.0]).cuda()
else:
raise NotImplementedError
else:
return torch.FloatTensor([1.0, 1.0]).cuda()
class CircularList(list):
"""
Helper for spliting training and validation scans
Originally: https://stackoverflow.com/questions/8951020/pythonic-circular-list/8951224
"""
def __getitem__(self, x):
if isinstance(x, slice):
return [self[x] for x in self._rangeify(x)]
index = operator.index(x)
try:
return super().__getitem__(index % len(self))
except ZeroDivisionError:
raise IndexError('list index out of range')
def _rangeify(self, slice):
start, stop, step = slice.start, slice.stop, slice.step
if start is None:
start = 0
if stop is None:
stop = len(self)
if step is None:
step = 1
return range(start, stop, step)
| 3,299 | 1,290 |
from django.contrib import admin
from . models import Post,UserProfile,Notification,Comment,ThreadModel
# Register your models here.
admin.site.register(Post)
admin.site.register(UserProfile)
admin.site.register(Comment)
admin.site.register(Notification)
admin.site.register(ThreadModel) | 289 | 79 |
from typing import List
class Solution:
def finalPrices(self, prices: List[int]) -> List[int]:
# initialize variables
solution = list()
len_prices = len(prices)
flag = -1
for i in range(len_prices):
flag = -1
for j in range(i+1, len_prices):
if prices[j] <= prices[i]:
solution.append(prices[i]-prices[j])
flag = 1
break
if flag == 1: continue
else:
solution.append((prices[i]))
return solution
solution = Solution()
print(solution.finalPrices(prices = [1,2,3,4,5])) | 663 | 196 |
# Licensed under a 3-clause BSD style license - see LICENSE.rst
"""Functions to compute TS images."""
import functools
import logging
import warnings
import numpy as np
import scipy.optimize
from astropy.coordinates import Angle
from gammapy.datasets.map import MapEvaluator
from gammapy.maps import Map, WcsGeom
from gammapy.modeling.models import PointSpatialModel, PowerLawSpectralModel, SkyModel
from gammapy.stats import (
amplitude_bounds_cython,
cash,
cash_sum_cython,
f_cash_root_cython,
x_best_leastsq,
)
from gammapy.utils.array import shape_2N, symmetric_crop_pad_width
from .core import Estimator
__all__ = ["TSMapEstimator"]
log = logging.getLogger(__name__)
FLUX_FACTOR = 1e-12
MAX_NITER = 20
RTOL = 1e-3
def round_up_to_odd(f):
return int(np.ceil(f) // 2 * 2 + 1)
def _extract_array(array, shape, position):
"""Helper function to extract parts of a larger array.
Simple implementation of an array extract function , because
`~astropy.ndata.utils.extract_array` introduces too much overhead.`
Parameters
----------
array : `~numpy.ndarray`
The array from which to extract.
shape : tuple or int
The shape of the extracted array.
position : tuple of numbers or number
The position of the small array's center with respect to the
large array.
"""
x_width = shape[1] // 2
y_width = shape[0] // 2
y_lo = position[0] - y_width
y_hi = position[0] + y_width + 1
x_lo = position[1] - x_width
x_hi = position[1] + x_width + 1
return array[y_lo:y_hi, x_lo:x_hi]
def f_cash(x, counts, background, model):
"""Wrapper for cash statistics, that defines the model function.
Parameters
----------
x : float
Model amplitude.
counts : `~numpy.ndarray`
Count image slice, where model is defined.
background : `~numpy.ndarray`
Background image slice, where model is defined.
model : `~numpy.ndarray`
Source template (multiplied with exposure).
"""
return cash_sum_cython(
counts.ravel(), (background + x * FLUX_FACTOR * model).ravel()
)
class TSMapEstimator(Estimator):
r"""Compute TS map from a MapDataset using different optimization methods.
The map is computed fitting by a single parameter amplitude fit. The fit is
simplified by finding roots of the the derivative of the fit statistics using
various root finding algorithms. The approach is described in Appendix A
in Stewart (2009).
Parameters
----------
model : `~gammapy.modeling.model.SkyModel`
Source model kernel. If set to None, assume point source model, PointSpatialModel.
kernel_width : `~astropy.coordinates.Angle`
Width of the kernel to use: the kernel will be truncated at this size
downsampling_factor : int
Sample down the input maps to speed up the computation. Only integer
values that are a multiple of 2 are allowed. Note that the kernel is
not sampled down, but must be provided with the downsampled bin size.
method : str ('root')
The following options are available:
* ``'root brentq'`` (default)
Fit amplitude by finding the roots of the the derivative of the fit
statistics using the brentq method.
* ``'root newton'``
Fit amplitude by finding the roots of the the derivative of the fit
statistics using Newton's method.
* ``'leastsq iter'``
Fit the amplitude by an iterative least square fit, that can be solved
analytically.
error_method : ['covar', 'conf']
Error estimation method.
error_sigma : int (1)
Sigma for flux error.
ul_method : ['covar', 'conf']
Upper limit estimation method.
ul_sigma : int (2)
Sigma for flux upper limits.
threshold : float (None)
If the TS value corresponding to the initial flux estimate is not above
this threshold, the optimizing step is omitted to save computing time.
rtol : float (0.001)
Relative precision of the flux estimate. Used as a stopping criterion for
the amplitude fit.
Notes
-----
Negative :math:`TS` values are defined as following:
.. math::
TS = \left \{
\begin{array}{ll}
-TS \text{ if } F < 0 \\
TS \text{ else}
\end{array}
\right.
Where :math:`F` is the fitted flux amplitude.
References
----------
[Stewart2009]_
"""
tag = "TSMapEstimator"
def __init__(
self,
model=None,
kernel_width="0.2 deg",
downsampling_factor=None,
method="root brentq",
error_method="covar",
error_sigma=1,
ul_method="covar",
ul_sigma=2,
threshold=None,
rtol=0.001,
):
if method not in ["root brentq", "root newton", "leastsq iter"]:
raise ValueError(f"Not a valid method: '{method}'")
if error_method not in ["covar", "conf"]:
raise ValueError(f"Not a valid error method '{error_method}'")
self.kernel_width = Angle(kernel_width)
if model is None:
model = SkyModel(
spectral_model=PowerLawSpectralModel(),
spatial_model=PointSpatialModel(),
)
self.model = model
self.downsampling_factor = downsampling_factor
self.parameters = {
"method": method,
"error_method": error_method,
"error_sigma": error_sigma,
"ul_method": ul_method,
"ul_sigma": ul_sigma,
"threshold": threshold,
"rtol": rtol,
}
def get_kernel(self, dataset):
"""Set the convolution kernel for the input dataset.
Convolves the model with the PSFKernel at the center of the dataset.
If no PSFMap or PSFKernel is found the dataset, the model is used without convolution.
"""
# TODO: further simplify the code below
geom = dataset.counts.geom
if self.downsampling_factor:
geom = geom.downsample(self.downsampling_factor)
model = self.model.copy()
model.spatial_model.position = geom.center_skydir
binsz = np.mean(geom.pixel_scales)
width_pix = self.kernel_width / binsz
npix = round_up_to_odd(width_pix.to_value(""))
axis = dataset.exposure.geom.get_axis_by_name("energy_true")
geom = WcsGeom.create(
skydir=model.position, proj="TAN", npix=npix, axes=[axis], binsz=binsz
)
exposure = Map.from_geom(geom, unit="cm2 s1")
exposure.data += 1.0
# We use global evaluation mode to not modify the geometry
evaluator = MapEvaluator(model, evaluation_mode="global")
evaluator.update(exposure, dataset.psf, dataset.edisp, dataset.counts.geom)
kernel = evaluator.compute_npred().sum_over_axes()
kernel.data /= kernel.data.sum()
if (self.kernel_width > geom.width).any():
raise ValueError(
"Kernel shape larger than map shape, please adjust"
" size of the kernel"
)
return kernel
@staticmethod
def flux_default(dataset, kernel):
"""Estimate default flux map using a given kernel.
Parameters
----------
dataset : `~gammapy.cube.MapDataset`
Input dataset.
kernel : `~numpy.ndarray`
Source model kernel.
Returns
-------
flux_approx : `~gammapy.maps.WcsNDMap`
Approximate flux map (2D).
"""
flux = dataset.counts - dataset.npred()
flux = flux.sum_over_axes(keepdims=False)
flux /= dataset.exposure.sum_over_axes(keepdims=False)
flux /= np.sum(kernel ** 2)
return flux.convolve(kernel)
@staticmethod
def mask_default(exposure, background, kernel):
"""Compute default mask where to estimate TS values.
Parameters
----------
exposure : `~gammapy.maps.Map`
Input exposure map.
background : `~gammapy.maps.Map`
Input background map.
kernel : `~numpy.ndarray`
Source model kernel.
Returns
-------
mask : `gammapy.maps.WcsNDMap`
Mask map.
"""
mask = np.zeros(exposure.data.shape, dtype=int)
# mask boundary
slice_x = slice(kernel.shape[1] // 2, -kernel.shape[1] // 2 + 1)
slice_y = slice(kernel.shape[0] // 2, -kernel.shape[0] // 2 + 1)
mask[slice_y, slice_x] = 1
# positions where exposure == 0 are not processed
mask &= exposure.data > 0
# in some image there are pixels, which have exposure, but zero
# background, which doesn't make sense and causes the TS computation
# to fail, this is a temporary fix
mask[background.data == 0] = 0
return exposure.copy(data=mask.astype("int"), unit="")
@staticmethod
def sqrt_ts(map_ts):
r"""Compute sqrt(TS) map.
Compute sqrt(TS) as defined by:
.. math::
\sqrt{TS} = \left \{
\begin{array}{ll}
-\sqrt{-TS} & : \text{if} \ TS < 0 \\
\sqrt{TS} & : \text{else}
\end{array}
\right.
Parameters
----------
map_ts : `gammapy.maps.WcsNDMap`
Input TS map.
Returns
-------
sqrt_ts : `gammapy.maps.WcsNDMap`
Sqrt(TS) map.
"""
with np.errstate(invalid="ignore", divide="ignore"):
ts = map_ts.data
sqrt_ts = np.where(ts > 0, np.sqrt(ts), -np.sqrt(-ts))
return map_ts.copy(data=sqrt_ts)
def run(self, dataset, steps="all"):
"""
Run TS map estimation.
Requires a MapDataset with counts, exposure and background_model
properly set to run.
Parameters
----------
dataset : `~gammapy.datasets.MapDataset`
Input MapDataset.
steps : list of str or 'all'
Which maps to compute. Available options are:
* "ts": estimate delta TS and significance (sqrt_ts)
* "flux-err": estimate symmetric error on flux.
* "flux-ul": estimate upper limits on flux.
By default all steps are executed.
Returns
-------
maps : dict
Dictionary containing result maps. Keys are:
* ts : delta TS map
* sqrt_ts : sqrt(delta TS), or significance map
* flux : flux map
* flux_err : symmetric error map
* flux_ul : upper limit map
"""
p = self.parameters
# First create 2D map arrays
counts = dataset.counts.sum_over_axes(keepdims=False)
background = dataset.npred().sum_over_axes(keepdims=False)
exposure = dataset.exposure.sum_over_axes(keepdims=False)
kernel = self.get_kernel(dataset)
if dataset.mask is not None:
mask = counts.copy(data=(dataset.mask.sum(axis=0) > 0).astype("int"))
else:
mask = counts.copy(data=np.ones_like(counts).astype("int"))
if self.downsampling_factor:
shape = counts.data.shape
pad_width = symmetric_crop_pad_width(shape, shape_2N(shape))[0]
counts = counts.pad(pad_width).downsample(
self.downsampling_factor, preserve_counts=True
)
background = background.pad(pad_width).downsample(
self.downsampling_factor, preserve_counts=True
)
exposure = exposure.pad(pad_width).downsample(
self.downsampling_factor, preserve_counts=False
)
mask = mask.pad(pad_width).downsample(
self.downsampling_factor, preserve_counts=False
)
mask.data = mask.data.astype("int")
mask.data &= self.mask_default(exposure, background, kernel.data).data
if steps == "all":
steps = ["ts", "sqrt_ts", "flux", "flux_err", "flux_ul", "niter"]
result = {}
for name in steps:
data = np.nan * np.ones_like(counts.data)
result[name] = counts.copy(data=data)
flux_map = self.flux_default(dataset, kernel.data)
if p["threshold"] or p["method"] == "root newton":
flux = flux_map.data
else:
flux = None
# prepare dtype for cython methods
counts_array = counts.data.astype(float)
background_array = background.data.astype(float)
exposure_array = exposure.data.astype(float)
# Compute null statistics per pixel for the whole image
c_0 = cash(counts_array, background_array)
error_method = p["error_method"] if "flux_err" in steps else "none"
ul_method = p["ul_method"] if "flux_ul" in steps else "none"
wrap = functools.partial(
_ts_value,
counts=counts_array,
exposure=exposure_array,
background=background_array,
c_0=c_0,
kernel=kernel.data,
flux=flux,
method=p["method"],
error_method=error_method,
threshold=p["threshold"],
error_sigma=p["error_sigma"],
ul_method=ul_method,
ul_sigma=p["ul_sigma"],
rtol=p["rtol"],
)
x, y = np.where(np.squeeze(mask.data))
positions = list(zip(x, y))
results = list(map(wrap, positions))
# Set TS values at given positions
j, i = zip(*positions)
for name in ["ts", "flux", "niter"]:
result[name].data[j, i] = [_[name] for _ in results]
if "flux_err" in steps:
result["flux_err"].data[j, i] = [_["flux_err"] for _ in results]
if "flux_ul" in steps:
result["flux_ul"].data[j, i] = [_["flux_ul"] for _ in results]
# Compute sqrt(TS) values
if "sqrt_ts" in steps:
result["sqrt_ts"] = self.sqrt_ts(result["ts"])
if self.downsampling_factor:
for name in steps:
order = 0 if name == "niter" else 1
result[name] = result[name].upsample(
factor=self.downsampling_factor, preserve_counts=False, order=order
)
result[name] = result[name].crop(crop_width=pad_width)
# Set correct units
if "flux" in steps:
result["flux"].unit = flux_map.unit
if "flux_err" in steps:
result["flux_err"].unit = flux_map.unit
if "flux_ul" in steps:
result["flux_ul"].unit = flux_map.unit
return result
def __repr__(self):
p = self.parameters
info = self.__class__.__name__
info += "\n\nParameters:\n\n"
for key in p:
info += f"\t{key:13s}: {p[key]}\n"
return info
def _ts_value(
position,
counts,
exposure,
background,
c_0,
kernel,
flux,
method,
error_method,
error_sigma,
ul_method,
ul_sigma,
threshold,
rtol,
):
"""Compute TS value at a given pixel position.
Uses approach described in Stewart (2009).
Parameters
----------
position : tuple (i, j)
Pixel position.
counts : `~numpy.ndarray`
Counts image
background : `~numpy.ndarray`
Background image
exposure : `~numpy.ndarray`
Exposure image
kernel : `astropy.convolution.Kernel2D`
Source model kernel
flux : `~numpy.ndarray`
Flux image. The flux value at the given pixel position is used as
starting value for the minimization.
Returns
-------
TS : float
TS value at the given pixel position.
"""
# Get data slices
counts_ = _extract_array(counts, kernel.shape, position)
background_ = _extract_array(background, kernel.shape, position)
exposure_ = _extract_array(exposure, kernel.shape, position)
c_0_ = _extract_array(c_0, kernel.shape, position)
model = exposure_ * kernel
c_0 = c_0_.sum()
if threshold is not None:
with np.errstate(invalid="ignore", divide="ignore"):
amplitude = flux[position]
c_1 = f_cash(amplitude / FLUX_FACTOR, counts_, background_, model)
# Don't fit if pixel significance is low
if c_0 - c_1 < threshold:
result = {}
result["ts"] = (c_0 - c_1) * np.sign(amplitude)
result["flux"] = amplitude
result["niter"] = 0
result["flux_err"] = np.nan
result["flux_ul"] = np.nan
return result
if method == "root brentq":
amplitude, niter = _root_amplitude_brentq(
counts_, background_, model, rtol=rtol
)
elif method == "root newton":
amplitude, niter = _root_amplitude(
counts_, background_, model, flux[position], rtol=rtol
)
elif method == "leastsq iter":
amplitude, niter = _leastsq_iter_amplitude(
counts_, background_, model, rtol=rtol
)
else:
raise ValueError(f"Invalid method: {method}")
with np.errstate(invalid="ignore", divide="ignore"):
c_1 = f_cash(amplitude, counts_, background_, model)
result = {}
result["ts"] = (c_0 - c_1) * np.sign(amplitude)
result["flux"] = amplitude * FLUX_FACTOR
result["niter"] = niter
if error_method == "covar":
flux_err = _compute_flux_err_covar(amplitude, counts_, background_, model)
result["flux_err"] = flux_err * error_sigma
elif error_method == "conf":
flux_err = _compute_flux_err_conf(
amplitude, counts_, background_, model, c_1, error_sigma
)
result["flux_err"] = FLUX_FACTOR * flux_err
if ul_method == "covar":
result["flux_ul"] = result["flux"] + ul_sigma * result["flux_err"]
elif ul_method == "conf":
flux_ul = _compute_flux_err_conf(
amplitude, counts_, background_, model, c_1, ul_sigma
)
result["flux_ul"] = FLUX_FACTOR * flux_ul + result["flux"]
return result
def _leastsq_iter_amplitude(counts, background, model, maxiter=MAX_NITER, rtol=RTOL):
"""Fit amplitude using an iterative least squares algorithm.
Parameters
----------
counts : `~numpy.ndarray`
Slice of counts image
background : `~numpy.ndarray`
Slice of background image
model : `~numpy.ndarray`
Model template to fit.
maxiter : int
Maximum number of iterations.
rtol : float
Relative flux error.
Returns
-------
amplitude : float
Fitted flux amplitude.
niter : int
Number of function evaluations needed for the fit.
"""
bounds = amplitude_bounds_cython(counts, background, model)
amplitude_min, amplitude_max, amplitude_min_total = bounds
if not counts.sum() > 0:
return amplitude_min_total, 0
weights = np.ones(model.shape)
x_old = 0
for i in range(maxiter):
x = x_best_leastsq(counts, background, model, weights)
if abs((x - x_old) / x) < rtol:
return max(x / FLUX_FACTOR, amplitude_min_total), i + 1
else:
weights = x * model + background
x_old = x
return max(x / FLUX_FACTOR, amplitude_min_total), MAX_NITER
def _root_amplitude(counts, background, model, flux, rtol=RTOL):
"""Fit amplitude by finding roots using newton algorithm.
See Appendix A Stewart (2009).
Parameters
----------
counts : `~numpy.ndarray`
Slice of count image
background : `~numpy.ndarray`
Slice of background image
model : `~numpy.ndarray`
Model template to fit.
flux : float
Starting value for the fit.
Returns
-------
amplitude : float
Fitted flux amplitude.
niter : int
Number of function evaluations needed for the fit.
"""
args = (counts, background, model)
with warnings.catch_warnings():
warnings.simplefilter("ignore")
try:
return (
scipy.optimize.newton(
f_cash_root_cython, flux, args=args, maxiter=MAX_NITER, tol=rtol
),
0,
)
except RuntimeError:
# Where the root finding fails NaN is set as amplitude
return np.nan, MAX_NITER
def _root_amplitude_brentq(counts, background, model, rtol=RTOL):
"""Fit amplitude by finding roots using Brent algorithm.
See Appendix A Stewart (2009).
Parameters
----------
counts : `~numpy.ndarray`
Slice of count image
background : `~numpy.ndarray`
Slice of background image
model : `~numpy.ndarray`
Model template to fit.
Returns
-------
amplitude : float
Fitted flux amplitude.
niter : int
Number of function evaluations needed for the fit.
"""
# Compute amplitude bounds and assert counts > 0
bounds = amplitude_bounds_cython(counts, background, model)
amplitude_min, amplitude_max, amplitude_min_total = bounds
if not counts.sum() > 0:
return amplitude_min_total, 0
args = (counts, background, model)
with warnings.catch_warnings():
warnings.simplefilter("ignore")
try:
result = scipy.optimize.brentq(
f_cash_root_cython,
amplitude_min,
amplitude_max,
args=args,
maxiter=MAX_NITER,
full_output=True,
rtol=rtol,
)
return max(result[0], amplitude_min_total), result[1].iterations
except (RuntimeError, ValueError):
# Where the root finding fails NaN is set as amplitude
return np.nan, MAX_NITER
def _compute_flux_err_covar(x, counts, background, model):
"""
Compute amplitude errors using inverse 2nd derivative method.
"""
with np.errstate(invalid="ignore", divide="ignore"):
stat = (model ** 2 * counts) / (background + x * FLUX_FACTOR * model) ** 2
return np.sqrt(1.0 / stat.sum())
def _compute_flux_err_conf(amplitude, counts, background, model, c_1, error_sigma):
"""
Compute amplitude errors using likelihood profile method.
"""
def ts_diff(x, counts, background, model):
return (c_1 + error_sigma ** 2) - f_cash(x, counts, background, model)
args = (counts, background, model)
amplitude_max = amplitude + 1e4
with warnings.catch_warnings():
warnings.simplefilter("ignore")
try:
result = scipy.optimize.brentq(
ts_diff,
amplitude,
amplitude_max,
args=args,
maxiter=MAX_NITER,
rtol=1e-3,
)
return result - amplitude
except (RuntimeError, ValueError):
# Where the root finding fails NaN is set as amplitude
return np.nan
| 23,226 | 7,045 |
""" Test Secure Headers Midddleware """
from src.masonite.middleware import SecureHeadersMiddleware
from src.masonite.request import Request
from src.masonite.testing import TestCase, generate_wsgi
class TestSecureHeadersMiddleware(TestCase):
def setUp(self):
super().setUp()
self.request = Request(generate_wsgi())
self.middleware = SecureHeadersMiddleware(self.request)
self.container.bind('Request', self.request.load_app(self.container))
self.request = self.container.make('Request')
def test_secure_headers_middleware(self):
self.middleware.after()
self.assertEqual(self.request.header('Strict-Transport-Security'), 'max-age=63072000; includeSubdomains')
self.assertEqual(self.request.header('X-Frame-Options'), 'SAMEORIGIN')
def test_secure_headers_gets_middleware_from_the_config(self):
self.request = self.container.make('Request')
self.middleware.after()
self.assertEqual(self.request.header('X-Content-Type-Options'), 'sniff-test')
| 1,048 | 316 |
import string
from factory import Dict, DictFactory, Faker, List
from factory.fuzzy import FuzzyChoice, FuzzyText
from reporter.apps import __version__
class SectionButtonFactory(DictFactory):
"""A factory for a section with a button."""
type = 'section'
accessory = Dict({
'text': {
'emoji': True,
'text': 'Review Now',
'type': 'plain_text',
},
'type': 'button',
'url': FuzzyText(
prefix='https://bitbucket.org/example/example_repos/pull-requests/',
length=4,
chars=string.digits,
),
})
text = Dict({
'text': FuzzyText(prefix='<@', suffix='>', length=2, chars=string.digits),
'type': 'mrkdwn',
})
class SectionBlockFactory(DictFactory):
"""A factory for a section block."""
type = 'section'
text = Dict({
'text': Dict({
'text': Faker('sentence'),
'type': FuzzyChoice(['mrkdwn', 'plain_text']),
}),
})
class ContextBlockFactory(DictFactory):
"""A factory for a context block."""
type = 'context'
elements = List([
Dict({'text': '*Author:* dave', 'type': 'mrkdwn'}),
Dict({'text': f'*version:* {__version__}', 'type': 'mrkdwn'}),
])
class DividerBlockFactory(DictFactory):
"""A factory for a divider block."""
type = 'divider'
class BlockFactory(DictFactory):
"""A factory for a block used in slack messages."""
text = Dict({
'text': Dict({
'text': Faker('sentence'),
'type': FuzzyChoice(['mrkdwn', 'plain_text']),
'emoji': FuzzyChoice([True, False]),
}),
})
type = FuzzyChoice(['section', 'divider', 'context'])
class SlackMessageFactory(DictFactory):
"""
A factory for a slack message.
This message is built via block kits that is a UI framework designed for slack.
Support url:
https://api.slack.com/block-kit
"""
blocks = List([
SectionBlockFactory(),
ContextBlockFactory(),
DividerBlockFactory(),
SectionButtonFactory(),
SectionButtonFactory(),
DividerBlockFactory(),
])
| 2,195 | 683 |
from pathlib import Path
from fhirzeug.generator import generate
from fhirzeug.fhirspec import FHIRSpec
def test_write(spec: FHIRSpec, tmp_path: Path):
spec.generator_config.output_directory.destination = tmp_path
spec.generator_config.output_file.destination = Path("output.py")
generate(spec)
assert tmp_path.joinpath("output.py").is_file()
| 362 | 116 |
import os
import string
import math
from random import choices
from pprint import pprint
from urllib.parse import urlparse
from PIL import Image
from apng import APNG
from colorama import init, deinit
from hurry.filesize import size, alternative
from .config import IMG_EXTS, STATIC_IMG_EXTS, ANIMATED_IMG_EXTS
def _inspect_image(animage_path):
"""Returns information of an animted GIF/APNG"""
abspath = os.path.abspath(animage_path)
filename = str(os.path.basename(abspath))
ext = str.lower(os.path.splitext(filename)[1])
frame_count = 0
fps = 0
avg_delay = 0
fsize = size(os.stat(abspath).st_size, system=alternative)
# fsize = 0
width = height = 0
loop_duration = 0
extension = ''
if ext == '.gif':
try:
gif: Image = Image.open(abspath)
except Exception:
raise Exception(f'The chosen file ({filename}) is not a valid GIF image')
if gif.format != 'GIF' or not gif.is_animated:
raise Exception(f"The chosen GIF ({filename}) is not an animated GIF!")
width, height = gif.size
frame_count = gif.n_frames
# pprint(gif.info)
delays = []
for f in range(0, gif.n_frames):
gif.seek(f)
delays.append(gif.info['duration'])
avg_delay = sum(delays) / len(delays)
fps = round(1000.0 / avg_delay, 3)
loop_duration = round(frame_count / fps, 3)
extension = 'GIF'
elif ext == '.png':
try:
apng: APNG = APNG.open(abspath)
except Exception:
raise Exception(f'The chosen file ({filename}) is not a valid PNG image')
frames = apng.frames
frame_count = len(frames)
if frame_count <= 1:
raise Exception(f"The chosen PNG ({filename}) is not an animated PNG!")
png_one, controller_one = frames[0]
# pprint(png_one.__dict__)
# pprint(controller_one.__dict__)
extension = 'APNG'
width = png_one.width
height = png_one.height
avg_delay = sum([f[1].delay for f in frames]) / frame_count
fps = round(1000.0 / avg_delay, 3)
loop_duration = round(frame_count / fps, 3)
image_info = {
"name": filename,
"fps": fps,
"avg_delay": round(avg_delay / 1000, 3),
"fsize": fsize,
"extension": extension,
"frame_count": frame_count,
"absolute_url": abspath,
"width": width,
"height": height,
"loop_duration": loop_duration,
}
return image_info
def _inspect_sequence(image_paths):
"""Returns information of a selected sequence of images"""
abs_image_paths = [os.path.abspath(ip) for ip in image_paths if os.path.exists(ip)]
img_paths = [f for f in abs_image_paths if str.lower(os.path.splitext(f)[1][1:]) in STATIC_IMG_EXTS]
# raise Exception("imgs", imgs)
print("imgs count", len(img_paths))
# pprint(imgs)
if not img_paths:
raise Exception("No images selected. Make sure the path to them are correct")
first_img_name = os.path.splitext(img_paths[0])[0]
filename = os.path.basename(first_img_name.split('_')[0] if '_' in first_img_name else first_img_name)
# apngs = [apng for apng in (APNG.open(i) for i in imgs) if len(apng.frames) > 1]
# gifs = [gif for gif in (Image.open(i) for i in imgs) if gif.format == "GIF" and gif.is_animated]
static_imgs = [i for i in img_paths if len(APNG.open(i).frames) == 1 and Image.open(i).format != "GIF"]
sequence_size = size(sum([os.stat(i).st_size for i in static_imgs]), system=alternative)
print("statics count", len(static_imgs))
if not static_imgs:
raise Exception("The images choosen must be static images, not animted GIFs or PNGs!")
# pprint(apngs)
# pprint(gifs)
# if any(APNG.open(i) for i in imgs)):
sequence_info = {
"name": filename,
"total": len(static_imgs),
"sequences": static_imgs,
"size": sequence_size,
}
return sequence_info
| 4,040 | 1,399 |
# flake8: noqa - this is until we take care of the F401 violations with respect to __all__ & sphinx
from .data_types import ValueType, pd_schema_to_value_type, InferOptions
from .infer import DFDataInfer
class BaseDataInfer:
infer_schema = None
get_preview = None
get_stats = None
def get_infer_interface(df) -> BaseDataInfer:
if hasattr(df, "rdd"):
from .spark import SparkDataInfer
return SparkDataInfer
return DFDataInfer
| 468 | 158 |
N,M=map(int,input().split())
X=sorted(set(list(map(int,input().split()))))
s=[]
for i in range(M-1):
s.append(X[i+1]-X[i])
s=sorted(s)[::-1]
print(sum(s[N-1:])) | 164 | 85 |
# standard modules
import argparse
import os
# aliased standard modules
import pandas as pd
# modules of sanity checker
import add_exp_to_ref
import lib.paths as paths
import lib.utils as utils
import perform_test
import process_data
import lib.logger_config as logger_config
import lib.test_config as test_config
# aliased modules of sanity checker
import lib.plot_mean_std as plt
# standalone imports
from lib.logger_config import log
'''
Script to test sanity of climate models. It contains:
- main: process model output, perform tests and plot results,
each function called by main() can be called itself
as a main(). Prior to the execution, paths_init.py
needs to be executed.
Note that this script requires user input at some stages,
so it cannot be run as a batched job.
Help: python sanity_test.py --help
# C.Siegenthaler, 2019
# J.Jucker, 2020
'''
def main(new_exp,
p_raw_files,
raw_f_subfold,
p_stages,
p_ref_csv_files,
wrk_dir,
f_vars_to_extract,
f_pattern_ref,
tests,
spinup,
lclean,
ltestsuite,
lverbose):
# init logger
logger_config.init_logger(lverbose,__file__)
log.banner('Start sanity checker')
# make all paths from user to absolute paths
wrk_dir = utils.abs_path(wrk_dir)
p_stages = utils.abs_path(p_stages)
p_ref_csv_files = utils.abs_path(p_ref_csv_files)
f_pattern_ref = utils.abs_path(f_pattern_ref)
# create directories
os.makedirs(p_stages,exist_ok=True)
os.makedirs(wrk_dir,exist_ok=True)
# go to working directory
os.chdir((wrk_dir))
log.info('Working directory is {}'.format(wrk_dir))
# data processing takes a while, check that no step is done twice
actions = utils.determine_actions_for_data_processing(new_exp,
tests,
p_stages,
lclean)
# create dataframe out of raw data
results_data_processing = process_data.main(
new_exp,
actions,
tests,
spinup,
p_raw_files=p_raw_files,
p_stages=p_stages,
raw_f_subfold=raw_f_subfold,
f_vars_to_extract=f_vars_to_extract,
f_pattern_ref=f_pattern_ref)
results_test, references = perform_test.main(
new_exp,
results_data_processing=results_data_processing,
p_stages=p_stages,
tests=tests,
p_ref_csv_files=p_ref_csv_files,
ltestsuite=ltestsuite,
f_vars_to_extract=f_vars_to_extract)
if 'welch' in tests:
test = 'welch'
plt.plt_welchstest(
references[test].append(results_data_processing[test],
sort=False),
new_exp,
results_test[test],
p_stages=p_stages)
# Add experiment to the reference pool
#--------------------------------------------------------------------
log.banner('')
log.banner('Check results again before adding to reference pool')
log.banner('')
for test in tests:
test_cfg = test_config.get_config_of_current_test(test)
utils.print_warning_if_testresult_is_bad(
test,
results_test[test],
test_cfg.metric_threshold,test_cfg.metric)
if ltestsuite:
asw = 'YES'
else:
asw = input('If you are happy with this experiment, '
'do you want to add it to the reference pool ?'
'(yes/[No])\n')
if (asw.strip().upper() == 'YES') or (asw.strip().upper() == 'Y'):
add_exp_to_ref.main(new_exp,
tests,
p_stages=p_stages,
ltestsuite=ltestsuite,
p_ref_csv_files=p_ref_csv_files)
else:
args_for_manual_execution = \
utils.derive_arguments_for_add_exp_to_ref(new_exp,
tests,
p_stages,
p_ref_csv_files)
log.info('The experiment {} is NOT added to '
'the reference pool \n'.format(new_exp))
log.info('If you want to add the experiment {} '
'to the reference pool later on, type '
'the following line when you are ready:'
.format(new_exp, new_exp))
log.info('')
log.info('python add_exp_to_ref.py {}'
.format(args_for_manual_execution))
log.banner('')
log.banner('Sanity test finished')
log.banner('')
if __name__ == '__main__':
# parsing arguments
parser = argparse.ArgumentParser(
formatter_class=argparse.ArgumentDefaultsHelpFormatter)
parser.add_argument('--exp','-e', dest='exp',
required=True,
help='exp to proceed')
parser.add_argument('--p_raw_files', dest='p_raw_files',
default=paths.p_raw_files,
help='absolute path to raw files')
parser.add_argument('--p_stages', dest='p_stages',
default=paths.p_stages,
help='relative or absolute path '
'to write csv files of the testresults')
parser.add_argument('--raw_f_subfold', dest='raw_f_subfold',
default='',
help='Subfolder where the raw data are ')
parser.add_argument('--wrkdir','-w', dest='wrk_dir',
default=paths.p_wrkdir,
help='relative or absolute path to working directory')
parser.add_argument('--p_ref_csv_files', dest='p_ref_csv_files',
default=paths.p_ref_csv_files,
help='relative or absolute path to reference files')
parser.add_argument('--f_vars_to_extract',dest='f_vars_to_extract',
default='vars_echam-hammoz.csv',
help='File containing variables to anaylse')
parser.add_argument('--verbose','-v', dest='lverbose',
action='store_true',
help='Debug output')
parser.add_argument('--clean','-c', dest='lclean',
action='store_true',
help='Redo all processing steps')
parser.add_argument('--testsuite','-ts', dest='ltestsuite',
action='store_true',
help='Run of testsuite')
parser.add_argument('--spinup', dest='spinup',
type=int,
default=3,
help='Do not consider first month '
'of the data due to model spinup')
parser.add_argument('--tests','-t', dest='tests',
default=['welch','fldcor','rmse','emi'],
nargs='+',
help='Tests to apply on your data')
parser.add_argument('--f_pattern_ref', dest='f_pattern_ref',
default='',
help='Absolute or relative path to reference '
'netCDF for spatial correlation tests')
args = parser.parse_args()
main(new_exp=args.exp,
p_raw_files=args.p_raw_files,
raw_f_subfold=args.raw_f_subfold,
wrk_dir=args.wrk_dir,
p_stages=args.p_stages,
p_ref_csv_files=args.p_ref_csv_files,
f_vars_to_extract=args.f_vars_to_extract,
f_pattern_ref=args.f_pattern_ref,
tests=args.tests,
spinup=args.spinup,
lclean=args.lclean,
ltestsuite=args.ltestsuite,
lverbose=args.lverbose)
| 7,942 | 2,321 |
from Calculator.Addition import addition
from Calculator.Division import division
def mean(num):
values = len(num)
total = sum(num)
return division(values, total)
| 181 | 56 |
black = "\033[1;30m"
red = "\033[1;31m"
green = "\033[01;32m"
yellow = "\033[01;33m"
blue = "\033[01;34m"
white = "\033[01;37m"
end = "\033[0m"
| 144 | 100 |
import numpy as np
import pandas as pd
from ..master_equation import master_equation as meq
#import MSI.master_equation.master_equation as meq
import copy
import re
import cantera as ct
class OptMatrix(object):
def __init__(self):
self.S_matrix = None
self.s_matrix = None
self.Y_matrix = None
self.y_matrix = None
self.z_matrix = None
self.delta_X = None
self.X = None
self.sigma = None
# #loads one experiment into self.matrix. Decides padding based on previous matrix or handle based on total exp num?
def build_Z(self, exp_dict_list:list,
parsed_yaml_file_list:list,
loop_counter:int = 0,
reaction_uncertainty=None,
master_equation_uncertainty_df=None,
master_equation_reaction_list=[],
master_equation_flag = False):
'''
Builds the Z vector.
Arguments:
exp_dic_list -- the dictonary that is built after a simulation
that contains things like sensitivity coefficients
parsed_yaml_file_list -- a list of dictonaries that contain the
information stored in the yaml files.
Keyword Arguments:
loop_counter -- keeps track of the iteration number for the optimization (default 0)
reaction_uncertainty -- a csv file that contains all the reactions
in the cti file being used for optimization and their corresponding
A,n and Ea uncertainty values (default None)
master_equation_uncertainty_df -- a pandas dataframe that contains
the reactions being treated with theory paramters along with the
associated uncertainty values of those paramters (default None)
master_equation_reaction_list -- a list of the reactions being treated
with theory paramters (default [])
master_equation_flag -- a boolean that indicates if reactions being
represented by theory parameters are being used in the optimization (default False)
'''
Z = []
Z_data_Frame = []
sigma = []
def jsr_temp_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
temp_uncertainties = list(temp_uncertainties)
else:
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['Temperature'].values))
temp_uncertainties = list(temp_uncertainties)
return temp_uncertainties
def flow_reactor_time_shift_uncertainties(parsed_yaml_file_list,experiment_dict):
if len(parsed_yaml_file_list['timeShiftOriginal']) ==1:
time_shift_uncertainties = [experiment_dict['uncertainty']['time_shift_uncertainty']]
elif len(parsed_yaml_file_list['timeShiftOriginal']) >1:
time_shift_uncertainties = [experiment_dict['uncertainty']['time_shift_uncertainty']]*len(parsed_yaml_file_list['timeShiftOriginal'])
return time_shift_uncertainties
def flow_reactor_temp_uncertainties(experiment_dict):
if 'Temperature_Uncertainty' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['experimental_data'][0]['Temperature_Uncertainty'].values
temp_uncertainties = list(temp_uncertainties)
else:
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['Temperature'].values))
temp_uncertainties = list(temp_uncertainties)
return temp_uncertainties
def flame_speed_temp_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'Temperature' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
temp_uncertainties = list(temp_uncertainties)
elif 'Temperature' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['Temperature'].values))
temp_uncertainties = list(temp_uncertainties)
elif 'Pressure' in list(experiment_dict['experimental_data'][0].columns) or 'Phi' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']
temp_uncertainties = list(temp_uncertainties)
return temp_uncertainties
def flame_speed_press_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'Pressure' in list(experiment_dict['experimental_data'][0].columns):
press_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
press_uncertainties = list(press_uncertainties)
elif 'Pressure' in list(experiment_dict['experimental_data'][0].columns):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['Pressure'].values))
press_uncertainties = list(temp_uncertainties)
elif 'Temperature' in list(experiment_dict['experimental_data'][0].columns) or 'Phi' in list(experiment_dict['experimental_data'][0].columns):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']
press_uncertainties = list(temp_uncertainties)
return press_uncertainties
def igdelay_temp_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'pressure' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].temperatures) == len(experiment_dict['simulation'].pressures) and len(experiment_dict['simulation'].temperatures)>1 and len(experiment_dict['simulation'].pressures)>1:
temp_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
temp_uncertainties = list(temp_uncertainties)*len(experiment_dict['simulation'].temperatures)
elif 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'temperature' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
temp_uncertainties = list(temp_uncertainties)
elif 'temperature' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['temperature'].values))
temp_uncertainties = list(temp_uncertainties)
#stub this is where we are editing
elif 'pressure' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].temperatures) == len(experiment_dict['simulation'].pressures) and len(experiment_dict['simulation'].temperatures)>1 and len(experiment_dict['simulation'].pressures)>1 :
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['pressure'].values))
temp_uncertainties = list(temp_uncertainties)* len(experiment_dict['simulation'].temperatures)
elif 'pressure' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].temperatures) != len(experiment_dict['simulation'].pressures):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']
temp_uncertainties = list(temp_uncertainties)
elif len(experiment_dict['conditions_to_run'])>1 and len(experiment_dict['simulation'].temperatures)>1 and len(experiment_dict['simulation'].pressures)>1 and len(experiment_dict['simulation'].temperatures) == len(experiment_dict['simulation'].pressures):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']
temp_uncertainties = list(temp_uncertainties) * len(experiment_dict['simulation'].temperatures)
elif len(experiment_dict['conditions_to_run'])>1:
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']
temp_uncertainties = list(temp_uncertainties)
return temp_uncertainties
def igdelay_press_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'temperature' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].temperatures) == len(experiment_dict['simulation'].pressures) and len(experiment_dict['simulation'].temperatures)>1 and len(experiment_dict['simulation'].pressures)>1:
press_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
press_uncertainties = list(press_uncertainties)*len(experiment_dict['simulation'].temperatures)
elif 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'pressure' in list(experiment_dict['experimental_data'][0].columns):
press_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
press_uncertainties = list(press_uncertainties)
elif 'pressure' in list(experiment_dict['experimental_data'][0].columns):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['pressure'].values))
press_uncertainties = list(press_uncertainties)
elif 'temperature' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].temperatures) == len(experiment_dict['simulation'].pressures) and len(experiment_dict['simulation'].temperatures)>1 and len(experiment_dict['simulation'].pressures)>1:
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']
press_uncertainties = list(press_uncertainties) * len(experiment_dict['simulation'].temperatures)
#stub this is where editing is happening
elif 'temperature' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].temperatures) != len(experiment_dict['simulation'].pressures):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']
press_uncertainties = list(press_uncertainties)
elif len(experiment_dict['conditions_to_run'])>1 and len(experiment_dict['simulation'].temperatures)>1 and len(experiment_dict['simulation'].pressures)>1 and len(experiment_dict['simulation'].temperatures) == len(experiment_dict['simulation'].pressures):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']
press_uncertainties = list(press_uncertainties)* len(experiment_dict['simulation'].temperatures)
elif len(experiment_dict['conditions_to_run'])>1:
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']
press_uncertainties = list(press_uncertainties)
return press_uncertainties
def rcm_temp_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'temperature' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
temp_uncertainties = list(temp_uncertainties)
elif 'temperature' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].fullParsedYamlFile['temperatures'])==len(experiment_dict['simulation'].fullParsedYamlFile['pressures']):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['temperature'].values))
temp_uncertainties = list(temp_uncertainties)
elif 'pressure' in list(experiment_dict['experimental_data'][0].columns):
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']
temp_uncertainties = list(temp_uncertainties)
elif len(experiment_dict['conditions_to_run'])>1:
temp_uncertainties=experiment_dict['uncertainty']['temperature_relative_uncertainty']
temp_uncertainties = list(temp_uncertainties)
return temp_uncertainties
def rcm_press_uncertainties(experiment_dict):
if 'Relative_Uncertainty' in list(experiment_dict['experimental_data'][0].columns) and 'pressure' in list(experiment_dict['experimental_data'][0].columns):
press_uncertainties=experiment_dict['experimental_data'][0]['Relative_Uncertainty'].values
press_uncertainties = list(press_uncertainties)
elif 'pressure' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].fullParsedYamlFile['temperatures'])==len(experiment_dict['simulation'].fullParsedYamlFile['pressures']):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['pressure'].values))
press_uncertainties = list(press_uncertainties)
elif 'temperature' in list(experiment_dict['experimental_data'][0].columns) and len(experiment_dict['simulation'].fullParsedYamlFile['temperatures'])==len(experiment_dict['simulation'].fullParsedYamlFile['pressures']):
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']*np.ones(np.shape(experiment_dict['experimental_data'][0]['temperature'].values))
press_uncertainties = list(press_uncertainties)
elif len(experiment_dict['conditions_to_run'])>1:
press_uncertainties=experiment_dict['uncertainty']['pressure_relative_uncertainty']
press_uncertainties = list(press_uncertainties)
return press_uncertainties
#need to append to sigma
def uncertainty_calc(relative_uncertainty,absolute_uncertainty,data,experimental_data):
absolute_uncertainty=float(absolute_uncertainty)
length_of_data = data.shape[0]
if 'Relative_Uncertainty' in list(experimental_data.columns):
x_dependent_uncertainty = experimental_data['Relative_Uncertainty'].values
relative_uncertainty_array = copy.deepcopy(x_dependent_uncertainty)
relative_uncertainty_array = relative_uncertainty_array.reshape((relative_uncertainty_array.shape[0],1))
elif 'Relative_Uncertainty' not in list(experimental_data.columns):
relative_uncertainty_array = np.full((length_of_data,1),relative_uncertainty)
relative_uncertainty_array = relative_uncertainty_array.reshape((relative_uncertainty_array.shape[0],1))
if 'Absolute_Uncertainty' in list(experimental_data.columns):
x_dependent_a_uncertainty = experimental_data['Absolute_Uncertainty'].values
absolute_uncertainty_array = copy.deepcopy(x_dependent_a_uncertainty)
#Fix this to deal with 0 data.
absolute_uncertainty_array = np.divide(absolute_uncertainty_array,data)
absolute_uncertainty_array = absolute_uncertainty_array.reshape((absolute_uncertainty_array.shape[0],1))
elif 'Absolute_Uncertainty' not in list(experimental_data.columns):
absolute_uncertainty_array = np.divide(absolute_uncertainty,data)
absolute_uncertainty_array = absolute_uncertainty_array.reshape((absolute_uncertainty_array.shape[0],1))
total_uncertainty = np.sqrt(np.square(relative_uncertainty_array) + np.square(absolute_uncertainty_array))
un_weighted_uncertainty = copy.deepcopy(total_uncertainty)
if 'W' not in list(experimental_data.columns):
weighting_factor = (1/length_of_data**.5)
total_uncertainty = np.divide(total_uncertainty,weighting_factor)
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0],1))
elif 'W' in list(experimental_data.columns):
weighting_factor = experimental_data['W'].values
weighting_factor = weighting_factor.reshape((weighting_factor.shape[0],1))
total_uncertainty = np.divide(total_uncertainty,weighting_factor)
#total_uncertainty = total_uncertainty/weighting_factor
return total_uncertainty,un_weighted_uncertainty
#tab, start working here tomorrow with how we want to read in csv file
for i,exp_dic in enumerate(exp_dict_list):
counter = 0
#print(exp_dic)
for j,observable in enumerate(exp_dic['mole_fraction_observables']+
exp_dic['concentration_observables']+
exp_dic['flame_speed_observables']+
exp_dic['ignition_delay_observables']):
if observable == None:
pass
else:
if observable in exp_dic['mole_fraction_observables']:
## add ppm statment here ? check if it exists? and add concentration statment below just for parcing
total_uncertainty,un_weighted_uncertainty = uncertainty_calc(exp_dic['uncertainty']['mole_fraction_relative_uncertainty'][counter],
exp_dic['uncertainty']['mole_fraction_absolute_uncertainty'][counter],
exp_dic['experimental_data'][counter][observable].values,exp_dic['experimental_data'][counter])
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0],1))
un_weighted_uncertainty = un_weighted_uncertainty.reshape((un_weighted_uncertainty.shape[0], 1))
elif observable in exp_dic['concentration_observables'] and '_ppm' in exp_dic['experimental_data'][counter].columns[1]:
total_uncertainty,un_weighted_uncertainty = uncertainty_calc(exp_dic['uncertainty']['concentration_relative_uncertainty'][counter],
exp_dic['uncertainty']['concentration_absolute_uncertainty'][counter],
exp_dic['experimental_data'][counter][observable+'_ppm'].values,exp_dic['experimental_data'][counter])
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0], 1))
un_weighted_uncertainty = un_weighted_uncertainty.reshape((un_weighted_uncertainty.shape[0], 1))
elif observable in exp_dic['concentration_observables'] and '_mol/cm^3' in exp_dic['experimental_data'][counter].columns[1]:
total_uncertainty,un_weighted_uncertainty = uncertainty_calc(exp_dic['uncertainty']['concentration_relative_uncertainty'][counter],
exp_dic['uncertainty']['concentration_absolute_uncertainty'][counter],
exp_dic['experimental_data'][counter][observable+'_mol/cm^3'].values,exp_dic['experimental_data'][counter])
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0],1))
un_weighted_uncertainty = un_weighted_uncertainty.reshape((un_weighted_uncertainty.shape[0], 1))
elif observable in exp_dic['flame_speed_observables'] and '_cm/s' in exp_dic['experimental_data'][counter].columns[1]:
total_uncertainty,un_weighted_uncertainty = uncertainty_calc(exp_dic['uncertainty']['flame_speed_relative_uncertainty'][counter],
exp_dic['uncertainty']['flame_speed_absolute_uncertainty'][counter],
exp_dic['experimental_data'][counter][observable+'_cm/s'].values,exp_dic['experimental_data'][counter])
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0],1))
un_weighted_uncertainty = un_weighted_uncertainty.reshape((un_weighted_uncertainty.shape[0], 1))
elif observable in exp_dic['ignition_delay_observables'] and '_s'in exp_dic['experimental_data'][counter].columns[1]:
total_uncertainty,un_weighted_uncertainty = uncertainty_calc(exp_dic['uncertainty']['ignition_delay_relative_uncertainty'][counter],
exp_dic['uncertainty']['ignition_delay_absolute_uncertainty'][counter],
exp_dic['experimental_data'][counter][observable+'_s'].values,exp_dic['experimental_data'][counter])
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0],1))
un_weighted_uncertainty = un_weighted_uncertainty.reshape((un_weighted_uncertainty.shape[0], 1))
else:
raise Exception('We Do Not Have This Unit Installed, Please Use Mole Fraction, ppm, mol/cm^3 or cm/s')
Z.append(total_uncertainty)
sigma.append(un_weighted_uncertainty)
tempList = [observable+'_'+'experiment'+str(i)]*np.shape(total_uncertainty)[0]
Z_data_Frame.extend(tempList)
#print(Z_data_Frame)
counter+=1
if 'absorbance_observables' in list(exp_dic.keys()):
wavelengths = parsed_yaml_file_list[i]['absorbanceCsvWavelengths']
for k,wl in enumerate(wavelengths):
total_uncertainty,un_weighted_uncertainty = uncertainty_calc(exp_dic['uncertainty']['absorbance_relative_uncertainty'][k],
exp_dic['uncertainty']['absorbance_absolute_uncertainty'][k],
exp_dic['absorbance_experimental_data'][k]['Absorbance_'+str(wl)].values,exp_dic['absorbance_experimental_data'][k])
total_uncertainty = total_uncertainty.reshape((total_uncertainty.shape[0], 1))
un_weighted_uncertainty = un_weighted_uncertainty.reshape((un_weighted_uncertainty.shape[0], 1))
tempList = [str(wl)+'_'+'experiment'+'_'+str(i)]*np.shape(total_uncertainty)[0]
Z_data_Frame.extend(tempList)
Z.append(total_uncertainty)
sigma.append(un_weighted_uncertainty)
Z = np.vstack((Z))
sigma = np.vstack((sigma))
#Here we are adding A,n,and Ea uncertainty
#we go do not through an additional step to make sure that the A,N and Ea
#values are paired with the correct reactions as in the old code,
#because we wrote a function to make the excel sheet which will arrange things in the correct order
#We also need to decide if we want to put this in as ln values or not in the spreadsheet
active_parameters = []
reaction_uncertainty = pd.read_csv(reaction_uncertainty)
#Flatten master equation reaction list
flatten = lambda *n: (e for a in n
for e in (flatten(*a) if isinstance(a, (tuple, list)) else (a,)))
flattened_master_equation_reaction_list = list(flatten(master_equation_reaction_list))
if master_equation_flag:
for reaction in flattened_master_equation_reaction_list:
index = reaction_uncertainty.loc[reaction_uncertainty['Reaction'] == reaction].index[0]
reaction_uncertainty = reaction_uncertainty.drop([index])
#tab fix this correctly, this unit needs to be fixed when we make a decision what the spreadsheet looks like
uncertainty_As = reaction_uncertainty['Uncertainty A (unit)'].values
uncertainty_As = uncertainty_As.reshape((uncertainty_As.shape[0],
1))
#uncertainty_As = np.log(uncertainty_As)
Z = np.vstack((Z,uncertainty_As))
sigma = np.vstack((sigma,uncertainty_As))
for variable in range(uncertainty_As.shape[0]):
Z_data_Frame.append('A'+'_'+str(variable))
active_parameters.append('A'+'_'+str(variable))
uncertainty_ns = reaction_uncertainty['Uncertainty N (unit)'].values
uncertainty_ns = uncertainty_ns.reshape((uncertainty_ns.shape[0],
1))
Z = np.vstack((Z,uncertainty_ns))
sigma = np.vstack((sigma,uncertainty_ns))
for variable in range(uncertainty_ns.shape[0]):
Z_data_Frame.append('n'+'_'+str(variable))
active_parameters.append('n'+'_'+str(variable))
uncertainty_Eas = reaction_uncertainty['Uncertainty Ea (unit)'].values
uncertainty_Eas = uncertainty_Eas.reshape((uncertainty_Eas.shape[0],
1))
Z = np.vstack((Z,uncertainty_Eas))
sigma = np.vstack((sigma,uncertainty_Eas))
for variable in range(uncertainty_Eas.shape[0]):
Z_data_Frame.append('Ea'+'_'+str(variable))
active_parameters.append('Ea'+'_'+str(variable))
if master_equation_flag == True:
master_equation_uncertainty = []
for i,reaction in enumerate(master_equation_reaction_list):
if type(reaction)==str:
master_equation_uncertainty.append(list(master_equation_uncertainty_df[reaction].dropna().values))
elif type(reaction)==tuple:
column_headers = master_equation_uncertainty_df.columns.to_list()
for sub_reaction in reaction:
if sub_reaction in column_headers:
master_equation_uncertainty.append(list(master_equation_uncertainty_df[sub_reaction].dropna().values))
# if master_equation_flag ==True:
# master_equation_uncertainty = []
# for col in master_equation_uncertainty_df:
# master_equation_uncertainty.append(list(master_equation_uncertainty_df[col].dropna().values))
if master_equation_flag == True:
for i,reaction in enumerate(master_equation_reaction_list):
if type(reaction)==str:
for j,paramter in enumerate(master_equation_uncertainty_df[reaction].dropna()):
Z_data_Frame.append(str(reaction)+'_'+'P'+'_'+str(j))
active_parameters.append(master_equation_reaction_list[i]+'_P_'+str(j))
elif type(reaction)==tuple:
column_headers = master_equation_uncertainty_df.columns.to_list()
for sub_reaction in reaction:
if sub_reaction in column_headers:
for j,paramter in enumerate(master_equation_uncertainty_df[sub_reaction].dropna()):
Z_data_Frame.append(str(reaction)+'_'+'P'+'_'+str(j))
active_parameters.append(str(master_equation_reaction_list[i])+'_P_'+str(j))
# for i,reaction in enumerate(master_equation_uncertainty):
# for j,uncer in enumerate(reaction):
# Z_data_Frame.append('R'+'_'+str(i)+'_'+'P'+str(j))
# #This might not look right in the data frame but we can try
# #stub
# active_parameters.append(master_equation_reaction_list[i]+'_P_'+str(j))
##check this
master_equation_uncertainty = [item for sublist in master_equation_uncertainty for item in sublist]
master_equation_uncertainty = np.array(master_equation_uncertainty)
master_equation_uncertainty = master_equation_uncertainty.reshape((master_equation_uncertainty.shape[0],
1))
Z = np.vstack((Z,master_equation_uncertainty))
sigma = np.vstack((sigma,master_equation_uncertainty))
#This is going to have to be simulation specific
if exp_dict_list[0]['simulation'].physicalSens ==1:
for i, exp_dic in enumerate(exp_dict_list):
if re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']):
#for i,exp_dic in enumerate(exp_dict_list):
experiment_physical_uncertainty = []
#Temperature Uncertainty
experiment_physical_uncertainty.append(exp_dic['uncertainty']['temperature_relative_uncertainty'])
Z_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
active_parameters.append('T'+'_'+'experiment'+'_'+str(i))
#Pressure Uncertainty
experiment_physical_uncertainty.append(exp_dic['uncertainty']['pressure_relative_uncertainty'])
Z_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
active_parameters.append('P'+'_'+'experiment'+'_'+str(i))
#Species Uncertainty
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = exp_dic['uncertainty']['species_relative_uncertainty']['species']
dilluant = ['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']
for specie in species_to_loop:
if specie in dilluant:
continue
experiment_physical_uncertainty.append(species_uncertainties[specie])
Z_data_Frame.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
active_parameters.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty.append(exp_dic['uncertainty']['time_shift_absolute_uncertainty'])
Z_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
active_parameters.append('Time_shift'+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],
1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
elif re.match('[Jj][Ss][Rr]',exp_dict_list[i]['simulation_type']):
#ASK MARK WHAT TO ADD HERE
#for i,exp_dic in enumerate(exp_dict_list):
experiment_physical_uncertainty = []
#Temperature Uncertainty
temp_uncertainties=jsr_temp_uncertainties(exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+temp_uncertainties
#experiment_physical_uncertainty.append(exp_dic['uncertainty']['temperature_relative_uncertainty'])
Z_data_Frame=Z_data_Frame+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
active_parameters=active_parameters+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#Pressure Uncertainty
experiment_physical_uncertainty.append(exp_dic['uncertainty']['pressure_relative_uncertainty'])
Z_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
active_parameters.append('P'+'_'+'experiment'+'_'+str(i))
#Species Uncertainty
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = exp_dic['uncertainty']['species_relative_uncertainty']['species']
dilluant = ['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']
for specie in species_to_loop:
if specie in dilluant:
continue
experiment_physical_uncertainty.append(species_uncertainties[specie])
Z_data_Frame.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
active_parameters.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty.append(exp_dic['uncertainty']['restime_relative_uncertainty'])
Z_data_Frame.append('R_experiment_'+str(i))
active_parameters.append('R_experiment_'+str(i))
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
#print(Z_data_Frame)
elif re.match('[Ff]lame[- ][Ss]peed',exp_dict_list[i]['simulation_type']) and re.match('[Oo][Nn][Ee]|[1][ -][dD][ -][Ff]lame',exp_dict_list[i][['experiment_type']]):
#for i,exp_dic in enumerate(exp_dict_list):
experiment_physical_uncertainty = []
#Temperature Uncertainty
temp_uncertainties=flame_speed_temp_uncertainties(exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+temp_uncertainties
Z_data_Frame=Z_data_Frame+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
active_parameters=active_parameters+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#Pressure Uncertainty
press_uncertainties = flame_speed_press_uncertainties(exp_dic)
Z_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))*len(press_uncertainties)
active_parameters.append('P'+'_'+'experiment'+'_'+str(i))*len(press_uncertainties)
#Species Uncertainty
conditions = exp_dic['conditions']
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = exp_dic['uncertainty']['species_relative_uncertainty']['species']
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
if 'Diluant' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluant' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluant = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluant']
for nmbr_of_species_sets in range(max_species):
for specie in species_to_loop:
if specie in dilluant:
continue
experiment_physical_uncertainty.append(species_uncertainties[specie])
Z_data_Frame.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
active_parameters.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],
1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
elif re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
#ASK MARK WHAT TO ADD HERE
#for i,exp_dic in enumerate(exp_dict_list):
experiment_physical_uncertainty = []
#Temperature Uncertainty
temp_uncertainties=flow_reactor_temp_uncertainties(exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+temp_uncertainties
#experiment_physical_uncertainty.append(exp_dic['uncertainty']['temperature_relative_uncertainty'])
Z_data_Frame=Z_data_Frame+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
active_parameters=active_parameters+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#Pressure Uncertainty
experiment_physical_uncertainty.append(exp_dic['uncertainty']['pressure_relative_uncertainty'])
Z_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
active_parameters.append('P'+'_'+'experiment'+'_'+str(i))
#Species Uncertainty
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = exp_dic['uncertainty']['species_relative_uncertainty']['species']
dilluant = ['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']
for specie in species_to_loop:
if specie in dilluant:
continue
experiment_physical_uncertainty.append(species_uncertainties[specie])
Z_data_Frame.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
active_parameters.append('X'+'_'+str(specie)+'_'+'experiment'+'_'+str(i))
time_shift_uncertainties = flow_reactor_time_shift_uncertainties(parsed_yaml_file_list[i],exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+time_shift_uncertainties
Z_data_Frame=Z_data_Frame+['Time_Shift'+'_'+'experiment'+'_'+str(i)]*len(time_shift_uncertainties)
active_parameters=active_parameters+['Time_Shift'+'_'+'experiment'+'_'+str(i)]*len(time_shift_uncertainties)
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
elif re.match('[Ss]hock[- ][Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
#for i,exp_dic in enumerate(exp_dict_list):
if len(exp_dic['simulation'].temperatures) == len(exp_dic['simulation'].pressures) and len(exp_dic['simulation'].temperatures) >1 and len(exp_dic['simulation'].pressures) >1:
# print('inside z matrix')
experiment_physical_uncertainty = []
#Temperature Uncertainty
temp_uncertainties=igdelay_temp_uncertainties(exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+temp_uncertainties
for index in range(len(temp_uncertainties)):
Z_data_Frame.append('T'+str(index+1)+'_'+'experiment'+'_'+str(i))
active_parameters.append('T'+str(index+1)+'_'+'experiment'+'_'+str(i))
#Z_data_Frame=Z_data_Frame+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#active_parameters=active_parameters+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#Pressure Uncertainty
press_uncertainties = igdelay_press_uncertainties(exp_dic)
for index in range(len(press_uncertainties)):
Z_data_Frame.append('P'+str(index+1)+'_'+'experiment'+'_'+str(i))
active_parameters.append('P'+str(index+1)+'_'+'experiment'+'_'+str(i))
#Z_data_Frame=Z_data_Frame+['P'+'_'+'experiment'+'_'+str(i)]*len(press_uncertainties)
#active_parameters=active_parameters+['P'+'_'+'experiment'+'_'+str(i)]*len(press_uncertainties)
experiment_physical_uncertainty=experiment_physical_uncertainty+press_uncertainties
#print(len(press_uncertainties))
#Species Uncertainty
conditions = exp_dic['conditions_dict_list']
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Z_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
experiment_physical_uncertainty.append(species_uncertainties[specie])
active_parameters.append('X'+str(x+1)+'_'+species+'_'+'experiment'+'_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Z_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
experiment_physical_uncertainty.append(species_uncertainties[specie])
active_parameters.append('X'+str(x+1)+'_'+species+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty.append(exp_dic['uncertainty']['time_shift_absolute_uncertainty'])
Z_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
active_parameters.append('Time_shift'+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],
1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
else:
experiment_physical_uncertainty = []
#Temperature Uncertainty
temp_uncertainties=igdelay_temp_uncertainties(exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+temp_uncertainties
for index in range(len(temp_uncertainties)):
Z_data_Frame.append('T'+str(index+1)+'_'+'experiment'+'_'+str(i))
active_parameters.append('T'+str(index+1)+'_'+'experiment'+'_'+str(i))
#Z_data_Frame=Z_data_Frame+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#active_parameters=active_parameters+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#Pressure Uncertainty
press_uncertainties = igdelay_press_uncertainties(exp_dic)
for index in range(len(press_uncertainties)):
Z_data_Frame.append('P'+str(index+1)+'_'+'experiment'+'_'+str(i))
active_parameters.append('P'+str(index+1)+'_'+'experiment'+'_'+str(i))
#Z_data_Frame=Z_data_Frame+['P'+'_'+'experiment'+'_'+str(i)]*len(press_uncertainties)
#active_parameters=active_parameters+['P'+'_'+'experiment'+'_'+str(i)]*len(press_uncertainties)
experiment_physical_uncertainty=experiment_physical_uncertainty+press_uncertainties
#print(len(press_uncertainties))
#Species Uncertainty
conditions = exp_dic['conditions_dict_list']
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Z_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
experiment_physical_uncertainty.append(species_uncertainties[specie])
active_parameters.append('X'+str(x+1)+'_'+species+'_'+'experiment'+'_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Z_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
experiment_physical_uncertainty.append(species_uncertainties[specie])
active_parameters.append('X'+str(x+1)+'_'+species+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty.append(exp_dic['uncertainty']['time_shift_absolute_uncertainty'])
Z_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
active_parameters.append('Time_shift'+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],
1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
elif re.match('[Rr][Cc][Mm]',exp_dict_list[i]['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
#for i,exp_dic in enumerate(exp_dict_list):
experiment_physical_uncertainty = []
#Temperature Uncertainty
temp_uncertainties=rcm_temp_uncertainties(exp_dic)
experiment_physical_uncertainty=experiment_physical_uncertainty+temp_uncertainties
for index in range(len(temp_uncertainties)):
Z_data_Frame.append('T'+str(index+1)+'_'+'experiment'+'_'+str(i))
active_parameters.append('T'+str(index+1)+'_'+'experiment'+'_'+str(i))
#Z_data_Frame=Z_data_Frame+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#active_parameters=active_parameters+['T'+'_'+'experiment'+'_'+str(i)]*len(temp_uncertainties)
#Pressure Uncertainty
press_uncertainties = rcm_press_uncertainties(exp_dic)
for index in range(len(press_uncertainties)):
Z_data_Frame.append('P'+str(index+1)+'_'+'experiment'+'_'+str(i))
active_parameters.append('P'+str(index+1)+'_'+'experiment'+'_'+str(i))
#Z_data_Frame=Z_data_Frame+['P'+'_'+'experiment'+'_'+str(i)]*len(press_uncertainties)
#active_parameters=active_parameters+['P'+'_'+'experiment'+'_'+str(i)]*len(press_uncertainties)
experiment_physical_uncertainty=experiment_physical_uncertainty+press_uncertainties
#print(len(press_uncertainties))
#Species Uncertainty
conditions = exp_dic['conditions_dict_list']
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Z_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
experiment_physical_uncertainty.append(species_uncertainties[specie])
active_parameters.append('X'+str(x+1)+'_'+species+'_'+'experiment'+'_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Z_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
experiment_physical_uncertainty.append(species_uncertainties[specie])
active_parameters.append('X'+str(x+1)+'_'+species+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty.append(exp_dic['uncertainty']['time_shift_absolute_uncertainty'])
Z_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
active_parameters.append('Time_shift'+'_'+'experiment'+'_'+str(i))
experiment_physical_uncertainty = np.array(experiment_physical_uncertainty)
experiment_physical_uncertainty = experiment_physical_uncertainty.reshape((experiment_physical_uncertainty.shape[0],
1))
Z = np.vstack((Z,experiment_physical_uncertainty))
sigma = np.vstack((sigma,experiment_physical_uncertainty))
#print(exp_dict_list[i]['simulation_type'],exp_dict_list[i]['experiment_type'])
#building dictonary to keep track of independtend coupled coefficients
count = 0
coef_dict = {}
uncertainties_of_coefficents = []
for i,exp_dic in enumerate(exp_dict_list):
if 'perturbed_coef' not in exp_dic.keys():
continue
dictonary_of_coef_and_uncertainty = exp_dic['uncertainty']['coupled_coef_and_uncertainty']
for x in dictonary_of_coef_and_uncertainty:
if x not in coef_dict.keys():
coef_dict[x] = dictonary_of_coef_and_uncertainty[x]
for x in coef_dict:
for y in coef_dict[x]:
if y[0]!=0: #this might cause a problem in the future
count+=1
uncertainties_of_coefficents.append(y)
Z_data_Frame.append('Sigma'+'_'+str(count))
active_parameters.append('Sigma'+'_'+str(count))
uncertainties_of_coefficents = np.array(uncertainties_of_coefficents)
if uncertainties_of_coefficents.any() == True:
uncertainties_of_coefficents = uncertainties_of_coefficents.reshape((uncertainties_of_coefficents.shape[0],
1))
Z = np.vstack((Z,uncertainties_of_coefficents))
sigma = np.vstack((sigma,uncertainties_of_coefficents))
#return(Z,Z_data_Frame)
#print('THIS IS Z',Z_data_Frame)
Z_data_Frame = pd.DataFrame({'value': Z_data_Frame,'Uncertainty': Z.reshape((Z.shape[0],))})
self.z_matrix = Z
self.sigma = sigma
#print(Z.shape)
return Z,Z_data_Frame,sigma,active_parameters
def load_Y(self, exp_dict_list:list,parsed_yaml_file_list:list,
loop_counter:int = 0,
X:dict={},
master_equation_reactions = [],
master_equation_uncertainty_df = None,
master_equation_flag = False):
def natural_log_difference(experiment,model):
natural_log_diff = np.log(np.array(experiment)) - np.log(np.array(model))
return natural_log_diff
Y = []
Y_data_Frame = []
for i,exp_dic in enumerate(exp_dict_list):
counter = 0
for j,observable in enumerate((exp_dic['mole_fraction_observables']+
exp_dic['concentration_observables'] +
exp_dic['flame_speed_observables']+
exp_dic['ignition_delay_observables'])):
if observable == None:
pass
else:
#if you need to add something with concentration add it here
if 'ppm' in exp_dic['experimental_data'][counter].columns.tolist()[1]:
if re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_ppm'].values,
(exp_dic['simulation'].timeHistoryInterpToExperiment[observable].dropna().values)*1e6)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0],1))
if re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_ppm'].values,
(exp_dic['simulation'].timeHistories[0][observable].dropna().values)*1e6)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0],1))
if re.match('[Jj][Ss][Rr]',exp_dict_list[i]['simulation_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_ppm'].values,
(exp_dic['simulation'].timeHistories[0][observable].dropna().values)*1e6)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0],1))
elif 'mol/cm^3' in exp_dic['experimental_data'][counter].columns.tolist()[1]:
if re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']):
concentration = np.true_divide(1,exp_dic['simulation'].pressureAndTemperatureToExperiment[counter]['temperature'].to_numpy())*exp_dic['simulation'].pressureAndTemperatureToExperiment[counter]['pressure'].to_numpy()
concentration *= (1/(8.314e6))*exp_dic['simulation'].timeHistoryInterpToExperiment[observable].dropna().to_numpy()
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_mol/cm^3'].to_numpy(),concentration)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
if re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
concentration = np.true_divide(1,exp_dic['simulation'].timeHistories[0]['temperature'].to_numpy())*exp_dic['simulation'].timeHistories[0]['pressure'].to_numpy()
concentration *= (1/(8.314e6))*exp_dic['simulation'].timeHistories[0][observable].dropna().to_numpy()
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_mol/cm^3'].to_numpy(),concentration)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
if re.match('[Jj][Ss][Rr]',exp_dict_list[i]['simulation_type']):
concentration = np.true_divide(1.0,exp_dic['simulation'].pressure*ct.one_atm)*np.array(exp_dic['simulation'].temperatures)
concentration *= (1/(8.314e6))*exp_dic['simulation'].timeHistories[0][observable].dropna().to_numpy()
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_mol/cm^3'].to_numpy(),concentration)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
elif 'cm/s' in exp_dic['experimental_data'][counter].columns.tolist()[1]:
if re.match('[Ff]lame [Ss]peed',exp_dict_list[i]['simulation_type']) and re.match('[Oo][Nn][Ee]|[1][ -][dD][ -][Ff]lame',exp_dict_list[i]['experiment_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_cm/s'].to_numpy(),
exp_dic['simulation'].timeHistories[0][observable])
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
elif 's' in exp_dic['experimental_data'][counter].columns.tolist()[1]:
if re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
#check these units would be in seconds of ms?
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable+'_s'].to_numpy(),
exp_dic['simulation'].timeHistories[0]['delay'])
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
else:
if re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable].values,
exp_dic['simulation'].timeHistoryInterpToExperiment[observable].dropna().values)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
if re.match('[Jj][Ss][Rr]',exp_dict_list[i]['simulation_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable].values,
exp_dic['simulation'].timeHistories[0][observable].values)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
if re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
natural_log_diff = natural_log_difference(exp_dic['experimental_data'][counter][observable].values,
exp_dic['simulation'].timeHistories[0][observable].values)
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0], 1))
tempList = [observable+'_'+'experiment'+str(i)]*np.shape(natural_log_diff)[0]
Y_data_Frame.extend(tempList)
Y.append(natural_log_diff)
counter+=1
if 'absorbance_observables' in list(exp_dic.keys()):
wavelengths = parsed_yaml_file_list[i]['absorbanceCsvWavelengths']
for k,wl in enumerate(wavelengths):
natural_log_diff = natural_log_difference(exp_dic['absorbance_experimental_data'][k]['Absorbance_'+str(wl)].values,exp_dic['absorbance_model_data'][wl])
natural_log_diff = natural_log_diff.reshape((natural_log_diff.shape[0],
1))
tempList = [str(wl)+'_'+'experiment'+'_'+str(i)]*np.shape(natural_log_diff)[0]
Y_data_Frame.extend(tempList)
Y.append(natural_log_diff)
Y = np.vstack((Y))
#YdataFrame = pd.DataFrame({'value': YdataFrame,'ln_difference': Y})
reactions_in_cti_file = exp_dict_list[0]['simulation'].processor.solution.reaction_equations()
#assembling the target values portion of the Y matrix
#getting the size of the cti file from the first simulation because
#they all use the same cti file and it shouldn't matter
# add in a conditional statment for if there is master equation data
#which is getting included in the simulation
#Flatten master equation reaction list
flatten = lambda *n: (e for a in n
for e in (flatten(*a) if isinstance(a, (tuple, list)) else (a,)))
flattened_master_equation_reaction_list = list(flatten(master_equation_reactions))
if master_equation_flag ==True:
A_n_Ea_length = int((len(reactions_in_cti_file) - len(flattened_master_equation_reaction_list))*3)
number_of_molecular_parameters_list = []
for col in master_equation_uncertainty_df:
number_of_molecular_parameters_list.append(len(master_equation_uncertainty_df[col].dropna().values))
number_of_molecular_parameters = sum(number_of_molecular_parameters_list)
#print('we do not have master equation installed yet')
#subtract out the necessary target values and add the other ones in
else:
A_n_Ea_length = len(reactions_in_cti_file)*3
#addint the zeros to the Y array
#adding the strings to the dictonary
## making a,n and Ea zero list
A_n_Ea_zeros = np.zeros((A_n_Ea_length,1))
if master_equation_flag ==True:
molecular_paramter_zeros = np.zeros((number_of_molecular_parameters,1))
for variable in range(A_n_Ea_length//3):
Y_data_Frame.append('A'+'_'+str(variable))
for variable in range(A_n_Ea_length//3):
Y_data_Frame.append('n'+'_'+str(variable))
for variable in range(A_n_Ea_length//3):
Y_data_Frame.append('Ea'+'_'+str(variable))
#make this the order of master equation list
if master_equation_flag == True:
for i,reaction in enumerate(master_equation_reactions):
if type(reaction)==str:
for j,paramter in enumerate(master_equation_uncertainty_df[reaction].dropna()):
Y_data_Frame.append(str(reaction)+'_P'+'_'+str(j))
elif type(reaction)==tuple:
column_headers = master_equation_uncertainty_df.columns.to_list()
for sub_reaction in reaction:
if sub_reaction in column_headers:
for j,paramter in enumerate(master_equation_uncertainty_df[sub_reaction].dropna()):
Y_data_Frame.append(str(reaction)+'_P'+'_'+str(j))
# if master_equation_flag == True:
# for i,value in enumerate(number_of_molecular_parameters_list):
# for j,parameter in enumerate(range(value)):
# Y_data_Frame.append('R'+'_'+str(i)+'P'+'_'+str(j))
if loop_counter == 0:
Y = np.vstack((Y,A_n_Ea_zeros))
if master_equation_flag ==True:
Y = np.vstack((Y,molecular_paramter_zeros))
else:
#print('we do not have loop counter installed yet')
#need to check what we would need to do here
#should be tottal X ?
#clean this part of the code up here
temp_array = np.array(X['As_ns_Eas'])*-1
temp_array = temp_array.reshape((temp_array.shape[0],
1))
Y = np.vstack((Y, temp_array))
#clean this part of the code up here
#tab
if master_equation_flag == True:
temp_array = np.array(X['molecular_parameters'])*-1
temp_array = temp_array.reshape((temp_array.shape[0],
1))
Y = np.vstack((Y,temp_array))
#Assembling the phsycial portion of the Y matrix
if exp_dict_list[0]['simulation'].physicalSens ==1:
#print(exp_dict_list)
for i,exp_dic in enumerate(exp_dict_list):
if loop_counter ==0:
if re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ss]pecies[ -][Pp]rofile',exp_dict_list[i]['experiment_type']):
dic_of_conditions = exp_dic['simulation'].conditions
#subtract out the dilluant
species_in_simulation = len(set(dic_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
#add two for Temperature and Pressure
len_of_phsycial_observables_in_simulation = species_in_simulation + 2 + 1
temp_zeros = np.zeros((len_of_phsycial_observables_in_simulation,1))
#stacking the zeros onto the Y array
Y = np.vstack((Y,temp_zeros))
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
elif re.match('[Jj][Ss][Rr]',exp_dict_list[i]['simulation_type']) and re.match('[Ss]pecies[ -][Pp]rofile',exp_dict_list[i]['experiment_type']):
dict_of_conditions = exp_dic['simulation'].conditions
species_in_simulation = len(set(dict_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressure_in_simulation = 1
restime_in_simulation = 1
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressure_in_simulation+restime_in_simulation
temp_zeros = np.zeros((len_of_phsycial_observables_in_simulation,1))
Y = np.vstack((Y,temp_zeros))
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('R_experiment_'+str(i))
elif re.match('[Ff]lame [Ss]peed',exp_dict_list[i]['simulation_type']) and re.match('[Oo][Nn][Ee]|[1][ -][dD][ -][Ff]lame',exp_dict_list[i]['experimentType']):
conditions = exp_dic['conditions_dict_list']
species_to_loop = exp_dic['uncertainty']['species_relative_uncertainty']['species']
pressures_in_simulation = len(exp_dic['simulation'].pressures)
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
if 'Diluant' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluant' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluant = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluant']
# species_in_simulation = list(exp_dic['conditions_to_run'][0].keys())
species_in_simulation = len(set(species_in_simulation).difference(diluant)) * max_species
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressure_in_simulation = len(exp_dic['simulation'].pressures)
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressure_in_simulation
temp_zeros = np.zeros((len_of_phsycial_observables_in_simulation,1))
Y = np.vstack((Y,temp_zeros))
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
for value in range(pressures_in_simulation):
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
elif re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
conditions = exp_dic['conditions_dict_list']
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
#species_in_simulation = len(set(dict_of_conditions.keys()).difference(diluant)) * max_species
species = copy.deepcopy(species_to_loop)
species_in_simulation = int(len(singular_species)+((len(set(exp_dic['simulation'].fullParsedYamlFile['speciesNames']).difference(diluent))-len(singular_species))*len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])))
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressures_in_simulation = len(exp_dic['simulation'].pressures)
time_shift_length = 1
#print(species_in_simulation,temperatures_in_simulation,pressures_in_simulation)
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressures_in_simulation + time_shift_length
temp_zeros = np.zeros((len_of_phsycial_observables_in_simulation,1))
Y = np.vstack((Y,temp_zeros))
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+str(value+1)+'_'+'experiment'+'_'+str(i))
for value in range(pressures_in_simulation):
Y_data_Frame.append('P'+str(value+1)+'_'+'experiment'+'_'+str(i))
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
Y_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
elif re.match('[Rr][Cc][Mm]',exp_dict_list[i]['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
conditions = exp_dic['conditions_dict_list']
species_uncertainties = exp_dic['uncertainty']['species_relative_uncertainty']['dictonary_of_values']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
#species_in_simulation = len(set(dict_of_conditions.keys()).difference(diluant)) * max_species
species = copy.deepcopy(species_to_loop)
species_in_simulation = int(len(singular_species)+((len(set(exp_dic['simulation'].fullParsedYamlFile['speciesNames']).difference(diluent))-len(singular_species))*len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])))
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressures_in_simulation = len(exp_dic['simulation'].pressures)
time_shift_length = 1
#print(species_in_simulation,temperatures_in_simulation,pressures_in_simulation)
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressures_in_simulation + time_shift_length
temp_zeros = np.zeros((len_of_phsycial_observables_in_simulation,1))
Y = np.vstack((Y,temp_zeros))
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+str(value+1)+'_'+'experiment'+'_'+str(i))
for value in range(pressures_in_simulation):
Y_data_Frame.append('P'+str(value+1)+'_'+'experiment'+'_'+str(i))
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
Y_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
elif re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
dict_of_conditions = exp_dic['simulation'].conditions
species_in_simulation = len(set(dict_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
time_shift_in_simulation = len(parsed_yaml_file_list[i]['timeShiftOriginal'])
pressure_in_simulation = 1
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressure_in_simulation+time_shift_in_simulation
temp_zeros = np.zeros((len_of_phsycial_observables_in_simulation,1))
Y = np.vstack((Y,temp_zeros))
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
for variable in range(time_shift_in_simulation):
Y_data_Frame.append('Time_shift'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
else:
if re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']):
dic_of_conditions = exp_dic['simulation'].conditions
#subtract out the dilluant
species_in_simulation = len(set(dic_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
elif re.match('[Jj][Ss][Rr]',exp_dict_list[i]['simulation_type']):
dict_of_conditions = exp_dic['simulation'].conditions
species_in_simulation = len(set(dict_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressure_in_simulation = 1
restime_in_simulation = 1
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressure_in_simulation+restime_in_simulation
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('R_experiment_'+str(i))
elif re.match('[Ff]lame [Ss]peed',exp_dict_list[i]['simulation_type']) and re.match('[Oo][Nn][Ee]|[1][ -][dD][ -][Ff]lame',exp_dict_list[i]['experimentType']):
species_to_loop = exp_dic['uncertainty']['species_relative_uncertainty']['species']
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
if 'Diluant' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluant' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluant = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluant']
species_in_simulation = len(set(dict_of_conditions.keys()).difference(diluant)) * max_species
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressure_in_simulation = len(exp_dic['simulation'].pressures)
len_of_phsycial_observables_in_simulation = species_in_simulation+temperatures_in_simulation+pressure_in_simulation
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
for value in range(pressures_in_simulation):
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
elif re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
dict_of_conditions = exp_dic['simulation'].conditions
species_in_simulation = len(set(dict_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
time_shift_in_simulation = len(parsed_yaml_file_list[i]['timeShiftOriginal'])
pressure_in_simulation = 1
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+'_'+'experiment'+'_'+str(i))
Y_data_Frame.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
Y_data_Frame.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
for variable in range(time_shift_in_simulation):
Y_data_Frame.append('Time_shift'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
elif re.match('[Ss]hock [Tt]ube',exp_dict_list[i]['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
conditions = exp_dic['conditions_dict_list']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressures_in_simulation = len(exp_dic['simulation'].pressures)
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluant=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluant = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+str(value+1)+'_'+'experiment'+'_'+str(i))
for value in range(pressures_in_simulation):
Y_data_Frame.append('P'+str(value+1)+'_'+'experiment'+'_'+str(i))
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
Y_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
elif re.match('[Rr][Cc][Mm]',exp_dict_list[i]['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dict_list[i]['experiment_type']):
conditions = exp_dic['conditions_dict_list']
species_to_loop = list(exp_dic['conditions_dict_list'].keys())
temperatures_in_simulation = len(exp_dic['simulation'].temperatures)
pressures_in_simulation = len(exp_dic['simulation'].pressures)
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
max_species = max(list_with_most_species_in_them)
diluant=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluant = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
for value in range(temperatures_in_simulation):
Y_data_Frame.append('T'+str(value+1)+'_'+'experiment'+'_'+str(i))
for value in range(pressures_in_simulation):
Y_data_Frame.append('P'+str(value+1)+'_'+'experiment'+'_'+str(i))
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
Y_data_Frame.append('X'+str(x+1)+'_'+species+'_experiment_'+str(i))
Y_data_Frame.append('Time_shift'+'_'+'experiment'+'_'+str(i))
if i==len(exp_dict_list)-1:
temp_array = np.array(X['physical_observables'])*-1
temp_array = temp_array.reshape((temp_array.shape[0],
1))
Y = np.vstack((Y,temp_array))
#Assembling the portion of the Y matrix for the absorbance coefficient sensitiviteis
pert_coef = {} #build a dict matching pert_coef to their experiment and wavelength.
#length of the dict gives padding information
for exp in exp_dict_list:
if 'perturbed_coef' not in exp.keys():
continue
perturbed_for_exp = exp['perturbed_coef']
for x in perturbed_for_exp:
if x[0][2] not in pert_coef.keys():
pert_coef[x[0][2]] = [x[1]]
else:
pert_coef[x[0][2]].append(x[1])
num_ind_pert_coef = len(pert_coef)
temp_zeros = np.zeros((num_ind_pert_coef,1))
if loop_counter == 0:
Y = np.vstack((Y,temp_zeros))
else:
if 'absorbance_coefficent_observables' in X.keys():
#temp_array = np.array(X['absorbance_coefficent_observables'])
temp_array = X['absorbance_coefficent_observables']
temp_array = [a for a in temp_array if a != 'null']
#temp_array = temp_array[temp_array!=0]
#temp_array = temp_array[temp_array!=0]
temp_array = np.array(temp_array)
temp_array = np.array(temp_array)*-1
temp_array = temp_array.reshape((temp_array.shape[0],
1))
Y = np.vstack((Y,temp_array))
for x in range(num_ind_pert_coef):
Y_data_Frame.append('Sigma'+'_'+str(x))
Y_data_Frame = pd.DataFrame({'value': Y_data_Frame,'ln_difference': Y.reshape((Y.shape[0],))})
self.Y_matrix = Y
#print(Y.shape,'Y matrix without k targets')
return Y, Y_data_Frame
def load_S(self, exp_dict_list:list,parsed_yaml_list:list,
dk=.01,
master_equation_reactions = [],
mapped_master_equation_sensitivites=np.array(()),
master_equation_uncertainty_df = None,
master_equation_flag = False):
#preprocessing for padding
num_exp = len(exp_dict_list)
pert_coef = {} #build a dict matching pert_coef to their experiment and wavelength.
#length of the dict gives padding information
list_to_keep_order_of_coef = []
for exp in exp_dict_list:
if 'perturbed_coef' not in exp.keys():
continue
perturbed_for_exp = exp['perturbed_coef']
for x in perturbed_for_exp:
if x[0][2] not in pert_coef.keys():
pert_coef[x[0][2]] = [x[1]]
else:
pert_coef[x[0][2]].append(x[1])
if x[0][2] not in list_to_keep_order_of_coef:
list_to_keep_order_of_coef.append(x[0][2])
num_ind_pert_coef = len(pert_coef)
#print(pert_coef.keys())
#print(num_ind_pert_coef," sigmas")
#establish # of independent pert before hand, to proper pad the observables, put in list, make a dict of cc,
# values will be a list of tabs data?
# use the list to get the padding size
k_sens_for_whole_simulation = []
p_sens_for_whole_simulation = []
abs_coef_sens_for_whole_simulation = []
temps = []
for i,exp in enumerate(exp_dict_list):
ttl_kinetic_observables_for_exp = []
obs_counter =0
for j,observable in enumerate(exp['mole_fraction_observables'] + exp['concentration_observables']+ exp['ignition_delay_observables']):
if observable == None:
continue
#return exp['ksens']['A']
#print(np.shape(exp['ksens']['A'][obs_counter]))
#print(np.shape(exp['ksens']['N'][obs_counter]))
#print(np.shape(exp['ksens']['Ea'][obs_counter]))
single_obs_matrix = np.hstack((exp['ksens']['A'][obs_counter],
exp['ksens']['N'][obs_counter],
exp['ksens']['Ea'][obs_counter]))
#print(single_obs_matrix)
ttl_kinetic_observables_for_exp.append(single_obs_matrix)
obs_counter +=1
if 'perturbed_coef' in exp.keys():
wavelengths = parsed_yaml_list[i]['absorbanceCsvWavelengths']
for k,wl in enumerate(wavelengths):
single_obs_matrix = np.hstack((exp['absorbance_ksens'][wl][0],
exp['absorbance_ksens'][wl][1],
exp['absorbance_ksens'][wl][2]))
ttl_kinetic_observables_for_exp.append(single_obs_matrix)
ttl_kinetic_observables_for_exp = np.vstack((ttl_kinetic_observables_for_exp))
k_sens_for_whole_simulation.append(ttl_kinetic_observables_for_exp)
#print(np.shape(k_sens_for_whole_simulation))
####vstack ttl_kinetic_observables_for_exp and append somwehre else
if exp['simulation'].physicalSens ==1:
ttl_phsycal_obs_for_exp = []
for j,observable in enumerate(exp['mole_fraction_observables'] + exp['concentration_observables'] + exp['ignition_delay_observables']):
obs_counter = 0
if observable == None:
continue
if re.match('[Ss]hock [Tt]ube',exp['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp['experiment_type']):
temperature_sensitivity = exp['temperature'][observable].dropna().values
temperature_sensitivity = temperature_sensitivity.reshape((temperature_sensitivity.shape[0], 1))
time_shift_sensitivity = exp['time_shift'][observable].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
pressure_sensitivity = exp['pressure'][observable].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df[observable].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0]
,1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
elif re.match('[Jj][Ss][Rr]',exp['simulation_type']):
temperature_sensitivity=np.array(exp['temperature'][observable])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = exp['pressure'][observable].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
restime_sensitivity=exp['restime_sens'][observable].dropna().values
restime_sensitivity = restime_sensitivity.reshape((restime_sensitivity.shape[0],1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df[observable].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
restime_sensitivity=exp['restime_sens'][observable].dropna().values
restime_sensitivity = restime_sensitivity.reshape((restime_sensitivity.shape[0],1))
elif re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
temperature_sensitivity=np.array(exp['temperature'][observable])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = exp['pressure'][observable].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df[observable].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
if len(parsed_yaml_list[i]['timeShiftOriginal'])>1:
time_shift_sensitivity = np.array(exp['time_shift'][observable])*np.identity(len(exp['simulation'].temperatures))
else:
time_shift_sensitivity = np.array(exp['time_shift'][observable])
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif re.match('[Ii]gnition[- ][Dd]elay',exp['experiment_type']) and re.match('[Ss]hock[- ][Tt]ube',exp['simulation_type']):
#CHECK HOW MANY SPECIES THERE ARE.
conditions = exp['conditions_dict_list']
species_to_loop = list(exp['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
if len(exp['simulation'].temperatures)>1 and len(exp['simulation'].pressures)==1:
temperature_sensitivity=np.array(exp['temperature']['delay'])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = exp['pressure']['delay'].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
#print("INSIDE HERE")
elif len(exp['simulation'].pressures)>1 and len(exp['simulation'].temperatures)==1:
pressure_sensitivity = np.array(exp['pressure']['delay'])*np.identity(len(exp['simulation'].pressures))
temperature_sensitivity = exp['temperature']['delay'].dropna().values
temperature_sensitivity = temperature_sensitivity.reshape((temperature_sensitivity.shape[0], 1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif len(exp['simulation'].pressures)==1 and len(exp['simulation'].temperatures)==1 and len(list_with_most_species_in_them)>1:
pressure_sensitivity = exp['pressure']['delay'].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
temperature_sensitivity = exp['temperature']['delay'].dropna().values
temperature_sensitivity = temperature_sensitivity.reshape((temperature_sensitivity.shape[0], 1))
species_sensitivty=[]
conditions = exp['conditions_dict_list']
species_to_loop = list(exp['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
diluent=[]
if 'Diluent' in exp['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
single_species_sensitivty = exp['species'][x]['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
#print(single_species_sensitivty)
species_sensitivty.append(single_species_sensitivty)
elif species not in singular_species and species not in diluent:
single_species_sensitivty = np.array(exp['species'][x]['delay'])*np.identity(len(exp['species'][x]['delay']))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty=np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif len(exp['simulation'].pressures)>1 and len(exp['simulation'].temperatures)>1 and len(list_with_most_species_in_them)>1 and len(exp['simulation'].pressures)==len(exp['simulation'].temperatures):
temperature_sensitivity=np.array(exp['temperature']['delay'])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = np.array(exp['pressure']['delay'])*np.identity(len(exp['simulation'].pressures))
species_sensitivty=[]
conditions = exp['conditions_dict_list']
species_to_loop = list(exp['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
diluent=[]
if 'Diluent' in exp['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
single_species_sensitivty = exp['species'][x]['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
#print(single_species_sensitivty)
species_sensitivty.append(single_species_sensitivty)
elif species not in singular_species and species not in diluent:
single_species_sensitivty = np.array(exp['species'][x]['delay'])*np.identity(len(exp['species'][x]['delay']))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty=np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif len(exp['simulation'].pressures)>1 and len(exp['simulation'].temperatures)>1 and len(exp['simulation'].pressures) == len(exp['simulation'].temperatures):
temperature_sensitivity=np.array(exp['temperature']['delay'])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = np.array(exp['pressure']['delay'])*np.identity(len(exp['simulation'].pressures))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif re.match('[Ii]gnition[- ][Dd]elay',exp['experiment_type']) and re.match('[Rr][Cc][Mm]',exp['simulation_type']):
if len(exp['simulation'].temperatures)>1 and len(exp['simulation'].pressures)>1:
temperature_sensitivity=np.array(exp['temperature']['delay'])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = np.array(exp['pressure']['delay'])*np.identity(len(exp['simulation'].pressures))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif len(exp['simulation'].temperatures)>1:
temperature_sensitivity=np.array(exp['temperature']['delay'])*np.identity(len(exp['simulation'].temperatures))
pressure_sensitivity = exp['pressure']['delay'].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
#print("INSIDE HERE")
elif len(exp['simulation'].pressures)>1:
pressure_sensitivity = np.array(exp['pressure']['delay'])*np.identity(len(exp['simulation'].pressures))
temperature_sensitivity = exp['temperature']['delay'].dropna().values
temperature_sensitivity = temperature_sensitivity.reshape((temperature_sensitivity.shape[0], 1))
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif len(exp['simulation'].pressures)==1 and len(exp['simulation'].temperatures)==1:
pressure_sensitivity = exp['pressure']['delay'].dropna().values
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
temperature_sensitivity = exp['temperature']['delay'].dropna().values
temperature_sensitivity = temperature_sensitivity.reshape((temperature_sensitivity.shape[0], 1))
species_sensitivty=[]
conditions = exp['conditions_dict_list']
species_to_loop = list(exp['conditions_dict_list'].keys())
list_with_most_species_in_them = []
for specie in species_to_loop:
list_with_most_species_in_them.append(len(conditions[specie]))
diluent=[]
if 'Diluent' in exp['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
for x,species in enumerate(exp['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
single_species_sensitivty = exp['species'][x]['delay'].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0],1))
#print(single_species_sensitivty)
species_sensitivty.append(single_species_sensitivty)
elif species not in singular_species and species not in diluent:
single_species_sensitivty = np.array(exp['species'][x]['delay'])*np.identity(len(exp['species'][x]['delay']))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty=np.hstack((species_sensitivty))
time_shift_sensitivity = exp['time_shift']['delay'].dropna().values
time_shift_sensitivity = time_shift_sensitivity.reshape((time_shift_sensitivity.shape[0], 1))
elif re.match('[Ff]lame[- ][Ss]peed',exp['simulation_type']) and re.match('[Oo][Nn][Ee]|[1][ -][dD][ -][Ff]lame',exp['experiment_type']):
len_of_temperature_list = len(exp['simulation'].temperatures)
if len_of_temperature_list > 1:
temperature_sensitivity=np.array(exp['temperature'][observable])*np.identity(len(exp['simulation'].temperatures))
else:
temperature_sensitivity = np.array(exp['temperature'][observable])
temperature_sensitivity = temperature_sensitivity.reshape((temperature_sensitivity.shape[0], 1))
len_of_pressure_list = len(exp['simulation'].pressures)
if len_of_pressure_list >1:
pressure_sensitivity=np.array(exp['pressure'][observable])*np.identity(len(exp['simulation'].pressures))
else:
pressure_sensitivity=np.array(exp['pressure'][observable])
pressure_sensitivity = pressure_sensitivity.reshape((pressure_sensitivity.shape[0], 1))
#FIX THIS
#print('FIXXXX')
species_sensitivty = []
for df in exp['species']:
single_species_sensitivty = df[observable].dropna().values
single_species_sensitivty = single_species_sensitivty.reshape((single_species_sensitivty.shape[0]
,1))
species_sensitivty.append(single_species_sensitivty)
species_sensitivty = np.hstack((species_sensitivty))
if re.match('[Jj][Ss][Rr]',exp['simulation_type']):
single_obs_physical = np.hstack((temperature_sensitivity,pressure_sensitivity,species_sensitivty,restime_sensitivity))
elif re.match('[Ss]hock [Tt]ube',exp['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp['experiment_type']):
single_obs_physical = np.hstack((temperature_sensitivity,pressure_sensitivity,species_sensitivty,time_shift_sensitivity))
elif re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
single_obs_physical = np.hstack((temperature_sensitivity,pressure_sensitivity,species_sensitivty,time_shift_sensitivity))
elif re.match('[Ii]gnition[- ][Dd]elay',exp['experiment_type']) and re.match('[Ss]hock[- ][Tt]ube',exp['simulation_type']):
single_obs_physical = np.hstack((temperature_sensitivity,pressure_sensitivity,species_sensitivty,time_shift_sensitivity))
#print("INSIDE HERE")
elif re.match('[Ii]gnition[- ][Dd]elay',exp['experiment_type']) and re.match('[Rr][Cc][Mm]',exp['simulation_type']):
single_obs_physical = np.hstack((temperature_sensitivity,pressure_sensitivity,species_sensitivty,time_shift_sensitivity))
ttl_phsycal_obs_for_exp.append(single_obs_physical)
obs_counter +=1
if 'perturbed_coef' in exp.keys():
wavelengths = parsed_yaml_list[i]['absorbanceCsvWavelengths']
for k,wl in enumerate(wavelengths):
physical_sens = []
for p_sens in exp['absorbance_psens']:
array = p_sens[wl]
array = array.reshape((array.shape[0],1))
physical_sens.append(array)
for time_sens in exp['absorbance_time_shift']:
array2 = p_sens[wl]
array2 = array2.reshape((array2.shape[0],1))
physical_sens.append(array2)
physical_sens = np.hstack((physical_sens))
ttl_phsycal_obs_for_exp.append(physical_sens)
ttl_phsycal_obs_for_exp = np.vstack((ttl_phsycal_obs_for_exp))
p_sens_for_whole_simulation.append(ttl_phsycal_obs_for_exp)
#######################################################################################################################################################
if 'perturbed_coef' in exp.keys():
ttl_absorbance_obs_for_exp = []
wavelengths = parsed_yaml_list[i]['absorbanceCsvWavelengths']
for k,wl in enumerate(wavelengths):
perturbed_coefficeints = []
index_list = []
for xx in range(len(parsed_yaml_list[i]['coupledCoefficients'])):
for yy in range(len(parsed_yaml_list[i]['coupledCoefficients'][xx])):
ff = parsed_yaml_list[i]['functionalForm'][xx][yy]
#temp = list(parsed_yaml_list[i]['coupledCoefficients'][xx][yy])
for zz in range(len(parsed_yaml_list[i]['coupledCoefficients'][xx][yy])):
temp = list(parsed_yaml_list[i]['coupledCoefficients'][xx][yy])
coefficent = parsed_yaml_list[i]['coupledCoefficients'][xx][yy][zz]
if coefficent!=0:
perturbed_coefficent=coefficent+coefficent*dk
if zz==1 and ff =='F':
#change back tab
perturbed_coefficent = coefficent + .01*coefficent
temp[zz] = perturbed_coefficent
key = tuple(temp)
indx = list_to_keep_order_of_coef.index(key)
index_list.append(indx)
exp_index_sigma = temps.count(key)
temps.append(key)
array = pert_coef[key][exp_index_sigma][wl]
array = array.reshape((array.shape[0],1))
perturbed_coefficeints.append(array)
missing_sigmas = []
for indp_sigma in range(len(list_to_keep_order_of_coef)):
if indp_sigma not in index_list:
missing_sigmas.append(indp_sigma)
perturbed_coefficents_padded_with_zeros = []
count_sigma=0
for indp_sigma in range(len(list_to_keep_order_of_coef)):
if indp_sigma in missing_sigmas:
zero_array = np.zeros((perturbed_coefficeints[0].shape[0],1))
perturbed_coefficents_padded_with_zeros.append(zero_array)
else:
perturbed_coefficents_padded_with_zeros.append(perturbed_coefficeints[count_sigma])
count_sigma +=1
perturbed_coefficents_padded_with_zeros = np.hstack((perturbed_coefficents_padded_with_zeros))
ttl_absorbance_obs_for_exp.append(perturbed_coefficents_padded_with_zeros)
ttl_absorbance_obs_for_exp = np.vstack((ttl_absorbance_obs_for_exp))
abs_coef_sens_for_whole_simulation.append(ttl_absorbance_obs_for_exp)
#vstack ttl_absorbance_obs_for_exp and append somehwere else
else:
abs_coef_sens_for_whole_simulation.append(0)
######################################################################################################################################################
flatten = lambda *n: (e for a in n
for e in (flatten(*a) if isinstance(a, (tuple, list)) else (a,)))
flattened_master_equation_reaction_list = list(flatten(master_equation_reactions))
#assembling the S matrix from the individual experiments
#master_equation = False
if master_equation_flag == True:
S_ksens = np.vstack((k_sens_for_whole_simulation))
A_k = np.hsplit(S_ksens,3)[0]
N_k = np.hsplit(S_ksens,3)[1]
Ea_k = np.hsplit(S_ksens,3)[2]
number_of_master_equation_reactions = len(flattened_master_equation_reaction_list)
A_k = A_k[:,:-number_of_master_equation_reactions]
N_k = N_k[:,:-number_of_master_equation_reactions]
Ea_k = Ea_k[:,:-number_of_master_equation_reactions]
S_ksens = np.hstack((A_k,N_k,Ea_k))
#print(np.shape(S_ksens),'this is the shape of the S matrix before MP')
S_ksens = np.hstack((S_ksens,mapped_master_equation_sensitivites))
else:
S_ksens = np.vstack((k_sens_for_whole_simulation))
def sum_of_zeros(idx,array,column_list):
rows_behind = array.shape[0]
rows_infront = array.shape[0]
columns_behind = sum(column_list[:idx])
columns_infront = sum(column_list[idx+1:])
behind_tuple = (rows_behind,columns_behind)
infront_tuple = (rows_infront,columns_infront)
return (behind_tuple,infront_tuple)
if exp_dict_list[0]['simulation'].physicalSens ==1:
number_of_columns_in_psens_arrays = []
number_of_rows_in_psens_arrays=[]
for i,array in enumerate(p_sens_for_whole_simulation):
number_of_rows_in_psens_arrays.append(array.shape[0])
number_of_columns_in_psens_arrays.append(array.shape[1])
p_sens_whole_simulation_with_padding = []
for i,array in enumerate(p_sens_for_whole_simulation):
zero_array_behind = np.zeros(sum_of_zeros(i,array,number_of_columns_in_psens_arrays)[0])
if zero_array_behind.shape[1] != 0:
array = np.hstack((zero_array_behind,array))
zero_array_infront = np.zeros(sum_of_zeros(i,array,number_of_columns_in_psens_arrays)[1])
if zero_array_infront.shape[1] != 0:
array = np.hstack((array,zero_array_infront))
p_sens_whole_simulation_with_padding.append(array)
S_psens = np.vstack((p_sens_whole_simulation_with_padding))
##############################################################################################
absorb_coef_whole_simulation_with_padding = []
for i,exp in enumerate(exp_dict_list):
single_experiment_absorption = []
if exp['mole_fraction_observables'][0] != None or exp['concentration_observables'][0] != None or exp['ignition_delay_observables'][0] != None:
if 'perturbed_coef' not in exp.keys():
zero_array_for_observables_padding = np.zeros((number_of_rows_in_psens_arrays[i],
num_ind_pert_coef))
single_experiment_absorption.append(zero_array_for_observables_padding)
if 'perturbed_coef' in exp.keys():
zero_padded_aborption_coef_array = abs_coef_sens_for_whole_simulation[i]
combined = abs_coef_sens_for_whole_simulation[i]
if exp['mole_fraction_observables'][0] != None or exp['concentration_observables'][0] != None or exp['ignition_delay_observables'][0] != None:
zero_array_for_observables_padding = np.zeros((number_of_rows_in_psens_arrays[i]-zero_padded_aborption_coef_array.shape[0],
num_ind_pert_coef))
combined = np.vstack((zero_array_for_observables_padding,zero_padded_aborption_coef_array))
single_experiment_absorption.append(combined)
single_experiment_absorption = np.vstack((single_experiment_absorption))
absorb_coef_whole_simulation_with_padding.append(single_experiment_absorption)
absorb_coef_whole_simulation_with_padding = np.vstack((absorb_coef_whole_simulation_with_padding))
S_abs_coef = absorb_coef_whole_simulation_with_padding
#return((S_ksens,S_psens,S_abs_coef))
#print(np.shape(S_ksens),np.shape(S_psens),np.shape(S_abs_coef))
S_matrix = np.hstack((S_ksens,S_psens,S_abs_coef))
shape = np.shape(S_matrix)[1]
#append identy matrix
identity_matrix = np.identity(shape)
# identity_matrix[1,0]=.1
# identity_matrix[0,1]=.1
# identity_matrix[0,20]=.1
# identity_matrix[20,0]=.1
# identity_matrix[39,0]=.1
# identity_matrix[0,39]=.1
####making edits to this just for masten test
S_matrix = np.vstack((S_matrix,identity_matrix))
self.S_matrix = S_matrix
S_matrix_wo_k_targets = copy.deepcopy(self.S_matrix)
self.S_matrix_wo_k_targets = S_matrix_wo_k_targets
#print(S_matrix_wo_k_targets.shape,'S matrix without k targets')
S_matrix_df = pd.DataFrame(S_matrix)
return S_matrix
def grouping_physical_model_parameters(self,exp:list):
final_groups=[]
for i in exp['simulation'].fullParsedYamlFile['overallDict'].keys():
if not re.match('[dD]iluent',i['type']):
final_groups.append(i)
def breakup_X(self, X,
exp_dict_list:list,
exp_uncertainty_dict_list_original:list,
loop_counter:int = 0,
master_equation_uncertainty_df=None,
master_equation_reactions = [],
master_equation_flag = False):
X_to_subtract_from_Y = {}
reactions_in_cti_file = exp_dict_list[0]['simulation'].processor.solution.reaction_equations()
number_of_reactions = len(reactions_in_cti_file)
####Grab off updates directly for the CTI file
####need to add master equation reactions
##################################################################
if loop_counter !=0:
X_new = X
else:
X_new = X
##################################################################
#print('USING BURKE X VALUES')
#X = pd.read_csv('MSI/data/test_data/burke_X_values.csv')
#X= X['Burke_Value'].values
#X = X.reshape(X.shape[0],1)
################################################################
##################################################################
#print('RUNNING TEST')
#X_new = np.zeros(np.shape(X_new))
#X_new[79] = .01
# print(X_new)
# X_new[847] = -0.007258986471821074
# X_new[848] = -0.07160891432785314
# X_new[849] = -0.038747789992729584
# X_new[850] = -0.09184808671928052
# X_new[851] = -0.13343314153597205
# X_new[852] = 0.0046931837946472
# X_new[853] = -0.007191276020250346
#X= X['Burke_Value'].values
#X = X.reshape(X.shape[0],1)
#zeros = np.zeros((X_new.shape))
#X_new = zeros
# X_new[873,0] = .01
# print("X_NEW")
################################################################
flatten = lambda *n: (e for a in n
for e in (flatten(*a) if isinstance(a, (tuple, list)) else (a,)))
flattened_master_equation_reaction_list = list(flatten(master_equation_reactions))
X_new = list(X_new.flatten())
if exp_dict_list[0]['simulation'].kineticSens ==1:
value1 = 3*(number_of_reactions - len(flattened_master_equation_reaction_list))
AsNsEas = X_new[:value1]
X_to_subtract_from_Y['As_ns_Eas'] = AsNsEas
#### pickup here
dividedBy = int(len(AsNsEas) / 3)
def list_slice(S,step):
return [S[i::step] for i in range(step)]
resortedList = list_slice(AsNsEas,dividedBy)
innerDict = ['A','n','Ea']
l = [dict(zip(innerDict,resortedList[x])) for x in range(len(resortedList))]
Keys= []
for xx in range(int(value1/3)):
Keys.append('r'+str(xx))
deltaXAsNsEas = dict(zip(Keys,l))
innerDictNew = ['A_update','n_update','Ea_update']
ll = [dict(zip(innerDictNew,resortedList[x])) for x in range(len(resortedList))]
kinetic_paramter_dict = dict(zip(reactions_in_cti_file,ll))
#molecularParams = np.array([.1,.2,.3,.4,.2,.3,.4]).flatten().tolist()
# might need to fix this based on how lei is passing me information, check in notebook
if master_equation_flag == True:
# number_of_molecular_parameters_list = []
# for col in master_equation_uncertainty_df:
# number_of_molecular_parameters_list.append(len(master_equation_uncertainty_df[col].dropna().values))
number_of_molecular_parameters_list = []
for i,reaction in enumerate(master_equation_reactions):
if type(reaction)==str:
number_of_molecular_parameters_list.append(len(list(master_equation_uncertainty_df[reaction].dropna().values)))
elif type(reaction)==tuple:
column_headers = master_equation_uncertainty_df.columns.to_list()
for sub_reaction in reaction:
if sub_reaction in column_headers:
number_of_molecular_parameters_list.append(len(list(master_equation_uncertainty_df[sub_reaction].dropna().values)))
sum_of_moleular_paramters = sum(number_of_molecular_parameters_list)
value2 = sum_of_moleular_paramters
deltaXmolecularParams = X_new[value1:(value1+value2)]
X_to_subtract_from_Y['molecular_parameters'] = deltaXmolecularParams
molecular_paramters_by_reaction = []
reaction_numbers = []
start_mp = 0
for r,number in enumerate(number_of_molecular_parameters_list):
stop_mp = start_mp + number
molecular_paramters_by_reaction.append(deltaXmolecularParams[start_mp:stop_mp])
start_mp = stop_mp
reaction_numbers.append('R_'+str(r))
delta_x_molecular_params_by_reaction_dict = dict(zip(master_equation_reactions,molecular_paramters_by_reaction))
list_of_mp = []
for i,reaction in enumerate(molecular_paramters_by_reaction):
temp=[]
for j,value in enumerate(reaction):
temp.append('Paramter_'+str(j)+'_Update')
list_of_mp.append(temp)
inner_dict_temp = [dict(zip(list_of_mp[x],molecular_paramters_by_reaction[x])) for x in range(len(molecular_paramters_by_reaction))]
inner_dict_temp_2 = dict(zip(master_equation_reactions,inner_dict_temp))
kinetic_paramter_dict.update(inner_dict_temp_2)
#its possible this kinetic paramters dict might break
else:
value2 = 0
physical_observables = []
previous_value = 0
physical_observables_for_Y = []
if exp_dict_list[0]['simulation'].physicalSens ==1:
for i,exp_dic in enumerate(exp_dict_list):
if re.match('[Ss]hock [Tt]ube',exp_dic['simulation_type']) and re.match('[Ss]pecies[- ][Pp]rofile',exp_dic['experiment_type']):
dic_of_conditions = exp_dic['simulation'].conditions
#subtract out the dilluant
species_in_simulation = len(set(dic_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
#add two for Temperature and Pressure
len_of_phsycial_observables_in_simulation = species_in_simulation + 2 +1
new_value = previous_value + len_of_phsycial_observables_in_simulation
single_experiment_physical_observables = X_new[(value1+value2+previous_value):(value1+value2+new_value)]
physical_observables_for_Y.append(single_experiment_physical_observables)
temp_keys = []
#stacking the zeros onto the Y array
temp_keys.append('T'+'_'+'experiment'+'_'+str(i))
temp_keys.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
temp_keys.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
temp_keys.append('Time_shift'+'_'+'experiment'+'_'+str(i))
temp_dict = dict(zip(temp_keys,single_experiment_physical_observables))
physical_observables.append(temp_dict)
##come back to this and do a test on paper
previous_value = new_value
elif re.match('[Ss]hock [Tt]ube',exp_dic['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dic['experiment_type']):
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
species_in_simulation = int(len(singular_species)+((len(exp_dic['simulation'].fullParsedYamlFile['speciesNames'])-len(singular_species))*len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])))
len_of_phsycial_observables_in_simulation = species_in_simulation + len(exp_dic['simulation'].pressures)+len(exp_dic['simulation'].temperatures)+1
new_value = previous_value + len_of_phsycial_observables_in_simulation
single_experiment_physical_observables = X_new[(value1+value2+previous_value):(value1+value2+new_value)]
physical_observables_for_Y.append(single_experiment_physical_observables)
temp_keys = []
#stacking the zeros onto the Y array
for j in range(len(exp_dic['simulation'].temperatures)):
temp_keys.append('T'+str(j+1)+'_'+'experiment'+'_'+str(i))
#stacking the zeros onto the Y array
for j in range(len(exp_dic['simulation'].pressures)):
temp_keys.append('P'+str(j+1)+'_'+'experiment'+'_'+str(i))
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
temp_keys.append('X'+str(x+1)+'_cond'+str(0)+'_'+species+'_experiment_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
temp_keys.append('X'+str(x+1)+'_cond'+str(j)+'_'+species+'_experiment_'+str(i))
temp_keys.append('Time_shift'+'_'+'experiment'+'_'+str(i))
temp_dict = dict(zip(temp_keys,single_experiment_physical_observables))
physical_observables.append(temp_dict)
##come back to this and do a test on paper
previous_value = new_value
#print(temp_dict)
elif re.match('[Rc][Cc][Mm]',exp_dic['simulation_type']) and re.match('[Ii]gnition[- ][Dd]elay',exp_dic['experiment_type']):
diluent=[]
if 'Diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys() or 'diluent' in exp_dic['uncertainty']['species_relative_uncertainty']['type_dict'].keys():
diluent = exp_dic['uncertainty']['species_relative_uncertainty']['type_dict']['diluent']
singular_species=[]
for species in list(exp_dic['simulation'].fullParsedYamlFile['conditions'].keys()):
if len(exp_dic['simulation'].fullParsedYamlFile['conditions'][species])==1 and species not in diluent:
singular_species.append(species)
species_in_simulation = int(len(singular_species)+((len(exp_dic['simulation'].fullParsedYamlFile['speciesNames'])-len(singular_species))*len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])))
len_of_phsycial_observables_in_simulation = species_in_simulation + len(exp_dic['simulation'].pressures)+len(exp_dic['simulation'].temperatures)+1
new_value = previous_value + len_of_phsycial_observables_in_simulation
single_experiment_physical_observables = X_new[(value1+value2+previous_value):(value1+value2+new_value)]
physical_observables_for_Y.append(single_experiment_physical_observables)
temp_keys = []
#stacking the zeros onto the Y array
for j in range(len(exp_dic['simulation'].temperatures)):
temp_keys.append('T'+str(j+1)+'_'+'experiment'+'_'+str(i))
#stacking the zeros onto the Y array
for j in range(len(exp_dic['simulation'].pressures)):
temp_keys.append('P'+str(j+1)+'_'+'experiment'+'_'+str(i))
for x,species in enumerate(exp_dic['simulation'].fullParsedYamlFile['speciesNames']):
if species in singular_species and species not in diluent:
temp_keys.append('X'+str(x+1)+'_cond'+str(0)+'_'+species+'_experiment_'+str(i))
elif species not in singular_species and species not in diluent:
for j in range(len(exp_dic['simulation'].fullParsedYamlFile['conditions_to_run'])):
temp_keys.append('X'+str(x+1)+'_cond'+str(j)+'_'+species+'_experiment_'+str(i))
temp_keys.append('Time_shift'+'_'+'experiment'+'_'+str(i))
temp_dict = dict(zip(temp_keys,single_experiment_physical_observables))
physical_observables.append(temp_dict)
##come back to this and do a test on paper
previous_value = new_value
elif re.match('[Jj][Ss][Rr]',exp_dic['simulation_type']):
dic_of_conditions = exp_dic['simulation'].conditions
#subtract out the dilluant
species_in_simulation = len(set(dic_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
#add two for Temperature and Pressure
len_of_phsycial_observables_in_simulation = species_in_simulation + 1+len(exp_dic['simulation'].temperatures)+1
#print(len_of_phsycial_observables_in_simulation)
new_value = previous_value + len_of_phsycial_observables_in_simulation
single_experiment_physical_observables = X_new[(value1+value2+previous_value):(value1+value2+new_value)]
#print(len(single_experiment_physical_observables))
physical_observables_for_Y.append(single_experiment_physical_observables)
temp_keys = []
#stacking the zeros onto the Y array
for j in range(len(exp_dic['simulation'].temperatures)):
temp_keys.append('T'+str(j+1)+'_'+'experiment'+'_'+str(i))
temp_keys.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
temp_keys.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
temp_keys.append('R'+'_'+'experiment'+'_'+str(i))
temp_dict = dict(zip(temp_keys,single_experiment_physical_observables))
physical_observables.append(temp_dict)
##come back to this and do a test on paper
previous_value = new_value
elif re.match('[Ss]pecies[- ][Pp]rofile',exp_dict_list[i]['experiment_type']) and re.match('[Ff]low[ -][Rr]eactor',exp_dict_list[i]['simulation_type']):
dic_of_conditions = exp_dic['simulation'].conditions
#subtract out the dilluant
species_in_simulation = len(set(dic_of_conditions.keys()).difference(['Ar','AR','ar','HE','He','he','Kr','KR','kr','Xe','XE','xe','NE','Ne','ne']))
#add two for Temperature and Pressure
time_shift_length = len(exp_dic['simulation'].fullParsedYamlFile['timeShiftOriginal'])
len_of_phsycial_observables_in_simulation = species_in_simulation + 1+len(exp_dic['simulation'].temperatures)+time_shift_length
#print(len_of_phsycial_observables_in_simulation)
new_value = previous_value + len_of_phsycial_observables_in_simulation
single_experiment_physical_observables = X_new[(value1+value2+previous_value):(value1+value2+new_value)]
#print(len(single_experiment_physical_observables))
physical_observables_for_Y.append(single_experiment_physical_observables)
temp_keys = []
#stacking the zeros onto the Y array
for j in range(len(exp_dic['simulation'].temperatures)):
temp_keys.append('T'+str(j+1)+'_'+'experiment'+'_'+str(i))
temp_keys.append('P'+'_'+'experiment'+'_'+str(i))
for variable in range(species_in_simulation):
temp_keys.append('X'+'_'+str(variable)+'_'+'experiment'+'_'+str(i))
for j in range(time_shift_length):
temp_keys.append('Time_Shift'+str(j+1)+'_'+'experiment'+'_'+str(i))
temp_dict = dict(zip(temp_keys,single_experiment_physical_observables))
physical_observables.append(temp_dict)
##come back to this and do a test on paper
previous_value = new_value
physical_observables_for_Y = [item for sublist in physical_observables_for_Y for item in sublist]
X_to_subtract_from_Y['physical_observables'] = physical_observables_for_Y
test_abs = []
absorbance_coefficients_for_Y = []
coef_dict = {}
coef_dict_list = []
absorbance_coef_update_dict = {}
for i,exp_dic in enumerate(exp_uncertainty_dict_list_original):
if 'coupled_coef_and_uncertainty' not in exp_dic.keys():
continue
dictonary_of_coef_and_uncertainty = exp_dic['coupled_coef_and_uncertainty']
#tab start working here tomorrow, need to pass in the original version of this dict
#dictonary_of_coef_and_uncertainty = {(140000, 0.0): ([0.7], [0.0]), (1270000, 0.0): ([0.7], [0.0])}
for x in dictonary_of_coef_and_uncertainty:
if x not in coef_dict.keys():
coef_dict[x] = dictonary_of_coef_and_uncertainty[x]
if x not in coef_dict_list:
coef_dict_list.append(x)
start_abs = 0
stop_abs = 1
for i,cof in enumerate(coef_dict_list):
temp=[]
temp2=[]
# counter=1
for value in cof:
if value==0:
temp.append([0])
temp2.append(['null'])
else:
temp.append(X_new[(value1+value2+new_value+start_abs):(value1+value2+new_value+stop_abs)])
temp2.append(X_new[(value1+value2+new_value+start_abs):(value1+value2+new_value+stop_abs)])
start_abs = stop_abs
stop_abs +=1
temp = [item for sublist in temp for item in sublist]
temp2 = [item for sublist in temp2 for item in sublist]
absorbance_coef_update_dict[cof] = temp
absorbance_coefficients_for_Y.append(temp2)
test_abs.append(temp2)
# return everything in a dictonary??
absorbance_coefficients_for_Y = [item for sublist in absorbance_coefficients_for_Y for item in sublist]
X_to_subtract_from_Y['absorbance_coefficent_observables'] = absorbance_coefficients_for_Y
#
if master_equation_flag == False:
return deltaXAsNsEas,physical_observables,absorbance_coef_update_dict,X_to_subtract_from_Y,kinetic_paramter_dict
else:
return deltaXAsNsEas,physical_observables,absorbance_coef_update_dict,X_to_subtract_from_Y,delta_x_molecular_params_by_reaction_dict,kinetic_paramter_dict
def matrix_manipulation(self,runCounter,S_matrix,Y_matrix,z_matrix,XLastItteration = np.array(()),active_parameters=[]):
#RUnning test to link up to paramters
##################################################
#s_temp = np.zeros((1,S_matrix.shape[1]))
#s_temp[0,886]=1
#s_temp[0,888]=-1
#y_temp = np.zeros((1,1))
#y_temp[0,0]=0
#z_temp=np.zeros((1,1))
#z_temp[0,0]=.00001
#S_matrix=np.vstack((S_matrix,s_temp))
#Y_matrix = np.vstack((Y_matrix,y_temp))
#z_matrix = np.vstack((z_matrix,z_temp))
##################################################
# print("ONLY CONSIDERING RATE CONSTANT TARGETS")
# for value in np.arange(0,401):
# z_matrix[value,0] =1000000
##################################################
one_over_z = np.true_divide(1,z_matrix)
#print(Y_matrix)
y_matrix = Y_matrix * one_over_z
s_matrix = S_matrix * (one_over_z.flatten()[:,np.newaxis])
self.y_matrix = y_matrix
sTimesZ = S_matrix * (z_matrix.flatten())[:,np.newaxis]
#calculate covariance matrix
shape = np.shape(self.S_matrix_wo_k_targets)
s_wo_k_targets = s_matrix[:shape[0],:shape[1]]
identity_matrix = s_wo_k_targets[shape[0]-len(active_parameters):,:]
#try:
if runCounter==0:
c = np.dot(np.transpose(identity_matrix),identity_matrix)
c = np.linalg.inv(c)
prior_diag = np.diag(c)
prior_sigmas = np.sqrt(prior_diag)
covariance_prior_df = pd.DataFrame(c)
if active_parameters:
covariance_prior_df.columns = active_parameters
covariance_prior_df.reindex(labels = active_parameters)
prior_diag_df = pd.DataFrame({'parameter': active_parameters,'value': prior_diag.reshape((prior_diag.shape[0],))})
sorted_prior_diag = prior_diag_df.sort_values(by=['value'])
prior_sigmas_df = pd.DataFrame({'parameter': active_parameters,'value': prior_sigmas.reshape((prior_sigmas.shape[0],))})
else:
c = np.dot(np.transpose(s_matrix),s_matrix)
c = np.linalg.inv(c)
covariance_posterior_df = pd.DataFrame(c)
if active_parameters:
covariance_posterior_df.columns = active_parameters
covariance_posterior_df.reindex(labels = active_parameters)
posterior_diag = np.diag(c)
posterior_sigmas = np.sqrt(posterior_diag)
posterior_sigmas_df = pd.DataFrame({'parameter': active_parameters,'value': posterior_sigmas.reshape((posterior_sigmas.shape[0],))})
posterior_diag_df = pd.DataFrame({'parameter': active_parameters,'value': posterior_diag.reshape((posterior_diag.shape[0],))})
sorted_posterior_diag = posterior_diag_df.sort_values(by=['value'])
# except:
# #stub
# print('WE ARE IN THE EXCEPT STATMENT')
# if runCounter==0:
# c = -1
# c = -1
# prior_diag = -1
# prior_sigmas = -1
# covariance_prior_df = -1
# prior_diag_df = -1
# sorted_prior_diag = -1
# prior_sigmas_df = -1
# else:
# c = -1
# c =-1
# covariance_posterior_df = -1
# posterior_diag = -1
# posterior_sigmas = -1
# posterior_sigmas_df = -1
# posterior_diag_df = -1
# sorted_posterior_diag = -1
self.covariance = c
self.s_matrix = s_matrix
psudoInverse = np.linalg.pinv(s_matrix)
delta_X = np.dot(psudoInverse,y_matrix)
self.delta_X = delta_X
if runCounter == 0:
XlastItteration = np.zeros(np.shape(delta_X))
else:
XlastItteration = XLastItteration
X = XlastItteration + delta_X
#STUB THIS IS FOR A TESTING ITTERATION
#####################################################################
#X = np.zeros(np.shape(delta_X))
# X[564] = .01
#####################################################################
self.X = X
#STUB THIS
try:
X_data_frame = pd.DataFrame({'value': active_parameters,'Parameter': X.reshape((X.shape[0],))})
except:
X_data_frame = -1
if runCounter==0:
return X,c,s_matrix,y_matrix,delta_X,z_matrix,X_data_frame,prior_diag,prior_diag_df,sorted_prior_diag,covariance_prior_df,prior_sigmas_df
else:
return X,c,s_matrix,y_matrix,delta_X,z_matrix,X_data_frame,posterior_diag,posterior_diag_df,sorted_posterior_diag,covariance_posterior_df,posterior_sigmas_df
class Adding_Target_Values(meq.Master_Equation):
def __init__(self,S_matrix,Y_matrix,z_matrix,sigma,Y_data_Frame,z_data_Frame):
self.S_matrix = S_matrix
self.Y_matrix = Y_matrix
self.z_matrix = z_matrix
self.sigma = sigma
self.Y_data_Frame = Y_data_Frame
self.z_data_Frame = z_data_Frame
meq.Master_Equation.__init__(self)
def target_values_Y(self,target_value_csv,
exp_dict_list:list,
Y_data_Frame,
master_equation_reactions):
import cantera as ct
Y_df_list = []
Y_values = []
#make sure we put the reactions into the file in the units cantera uses
target_value_csv = pd.read_csv(target_value_csv)
target_reactions = target_value_csv['Reaction']
target_temp = target_value_csv['temperature']
target_press = target_value_csv['pressure']
target_k = target_value_csv['k']
bath_gas = target_value_csv['M']
reactions_in_cti_file = exp_dict_list[0]['simulation'].processor.solution.reaction_equations()
gas = ct.Solution(exp_dict_list[0]['simulation'].processor.cti_path)
diff_in_ks_for_Y = []
def check_if_M_in_reactants(list_to_append_to,
gas,
reactants_in_target_reactions,
reverse_reactants_in_target_reaction):
if reverse_reactants_in_target_reaction !=None:
for reaction_number_in_cti_file in range(gas.n_reactions):
if (gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' (+M)' or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction + ' (+M)' or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' + M' or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction + ' + M') :
list_to_append_to.append(reactions_in_cti_file[reaction_number_in_cti_file])
elif reverse_reactants_in_target_reaction==None:
for reaction_number_in_cti_file in range(gas.n_reactions):
if (gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' (+M)' or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' + M'):
list_to_append_to.append(reactions_in_cti_file[reaction_number_in_cti_file])
return list_to_append_to
for i,reaction in enumerate(target_reactions):
#ask about the mixture composition
#if reaction not in flattened_linked_channel_reactions:
if '*' not in reaction and reaction != 'More Complex Combination Rule' and '(+)' not in reaction:
index_in_cti_file = gas.reaction_equations().index(reaction)
units_reaction_types=['ElementaryReaction',
'PlogReaction',
'ChebyshevReaction',
'ThreeBodyReaction',
'FalloffReaction']
coeff_sum = sum(gas.reaction(index_in_cti_file).reactants.values())
if target_press[i] == 0:
pressure = 1e-9
else:
pressure = target_press[i]
if bath_gas[i] !=0:
gas.TPX = target_temp[i],pressure*101325,{'H2O':.013,'O2':.0099,'H':.0000007,'Ar':.9770993}
else:
gas.TPX = target_temp[i],pressure*101325,{'Ar':.99}
reaction_number_in_cti = reactions_in_cti_file.index(reaction)
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k = k
elif coeff_sum==2:
k=k*1000
elif coeff_sum==3:
k=k*1000000
#check and make sure we are subtracting in the correct order
difference = np.log(target_k[i]) - np.log(k)
diff_in_ks_for_Y.append(difference)
Y_df_list.append(reaction)
Y_values.append(difference)
#elif reaction in flattened_linked_channel_reactions:
elif '*' in reaction and reaction != 'More Complex Combination Rule' and '/' not in reaction:
reactions_in_cti_file_with_these_reactants = []
#might be a more comprehensive way to do this
reactants_in_target_reactions = reaction.split('<=>')[0].rstrip()
reverse_reactants_in_target_reaction=None
if len(reactants_in_target_reactions.split('+'))>1:
reverse_reactants_in_target_reaction = reactants_in_target_reactions.split('+')
temp = reverse_reactants_in_target_reaction[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction = temp
for reaction_number_in_cti_file in range(gas.n_reactions):
if gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction:
reactions_in_cti_file_with_these_reactants.append(reactions_in_cti_file[reaction_number_in_cti_file])
reactions_in_cti_file_with_these_reactants = check_if_M_in_reactants(reactions_in_cti_file_with_these_reactants,
gas,
reactants_in_target_reactions,
reactants_in_target_reactions)
if target_press[i] == 0:
pressure = 1e-9
else:
pressure = target_press[i]
if bath_gas[i] !=0:
gas.TPX = target_temp[i],pressure*101325,{'H2O':.013,'O2':.0099,'H':.0000007,'Ar':.9770993}
else:
gas.TPX = target_temp[i],pressure*101325,{'Ar':.99}
tottal_k = []
for secondary_reaction in reactions_in_cti_file_with_these_reactants:
reaction_number_in_cti = reactions_in_cti_file.index(secondary_reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k= k*1000000
tottal_k.append(k)
#check and make sure we are subtracting in the correct order
k=sum(tottal_k)
difference = np.log(target_k[i]) - np.log(k)
diff_in_ks_for_Y.append(difference)
#I guess i could append the tuple
Y_df_list.append(reaction)
Y_values.append(difference)
elif '/' in reaction:
reactants_in_numerator = reaction.split('/')[0].rstrip()
reactants_in_numerator = reactants_in_numerator.lstrip()
reactants_in_denominator = reaction.split('/')[1].rstrip()
reactants_in_denominator = reactants_in_denominator.lstrip()
reactions_in_cti_file_with_these_reactants_numerator = []
reactions_in_cti_file_with_these_reactants_denominator = []
#take back here
if '*' in reactants_in_numerator:
reactants_in_target_reactions_numerator = reactants_in_numerator.split('<=>')[0].rstrip()
reverse_reactants_in_target_reaction_in_numerator=None
if len(reactants_in_target_reactions_numerator.split('+'))>1:
reverse_reactants_in_target_reaction_in_numerator = reactants_in_target_reactions_numerator.split('+')
temp = reverse_reactants_in_target_reaction_in_numerator[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction_in_numerator[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction_in_numerator = temp
for reaction_number_in_cti_file in range(gas.n_reactions):
if gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions_numerator or gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction_in_numerator:
reactions_in_cti_file_with_these_reactants_numerator.append(reactions_in_cti_file[reaction_number_in_cti_file])
reactions_in_cti_file_with_these_reactants_numerator = check_if_M_in_reactants(reactions_in_cti_file_with_these_reactants_numerator,
gas,
reactants_in_target_reactions_numerator,
reverse_reactants_in_target_reaction_in_numerator)
else:
#need to figure out how to split addition of reactions
if '(+)' not in reactants_in_numerator:
reactions_in_cti_file_with_these_reactants_numerator.append(reactants_in_numerator)
else:
list_of_reactions_in_numerator = reactants_in_numerator.split('(+)')
list_of_reactions_in_numerator_cleaned=[]
for reaction in list_of_reactions_in_numerator:
reaction = reaction.rstrip()
reaction = reaction.lstrip()
list_of_reactions_in_numerator_cleaned.append(reaction)
reactions_in_cti_file_with_these_reactants_numerator = list_of_reactions_in_numerator_cleaned
if '*' in reactants_in_denominator:
reactants_in_target_reactions_denominator = reactants_in_denominator.split('<=>')[0].rstrip()
reverse_reactants_in_target_reaction_in_denominator=None
if len(reactants_in_target_reactions_denominator.split('+'))>1:
reverse_reactants_in_target_reaction_in_denominator = reactants_in_target_reactions_denominator.split('+')
temp = reverse_reactants_in_target_reaction_in_denominator[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction_in_denominator[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction_in_denominator = temp
for reaction_number_in_cti_file in range(gas.n_reactions):
if gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions_denominator or gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction_in_denominator:
reactions_in_cti_file_with_these_reactants_denominator.append(reactions_in_cti_file[reaction_number_in_cti_file])
reactions_in_cti_file_with_these_reactants_denominator = check_if_M_in_reactants(reactions_in_cti_file_with_these_reactants_denominator,
gas,
reactants_in_target_reactions_denominator,
reverse_reactants_in_target_reaction_in_denominator)
else:
#need to figure out how to split addition of reactions
if '(+)' not in reactants_in_denominator:
reactions_in_cti_file_with_these_reactants_denominator.append(reactants_in_denominator)
else:
list_of_reactions_in_denominator = reactants_in_denominator.split('(+)')
list_of_reactions_in_denominator_cleaned=[]
for reaction in list_of_reactions_in_denominator:
reaction = reaction.rstrip()
reaction = reaction.lstrip()
list_of_reactions_in_denominator_cleaned.append(reaction)
reactions_in_cti_file_with_these_reactants_denominator = list_of_reactions_in_denominator_cleaned
if target_press[i] == 0:
pressure = 1e-9
else:
pressure = target_press[i]
if bath_gas[i] !=0:
gas.TPX = target_temp[i],pressure*101325,{'H2O':.013,'O2':.0099,'H':.0000007,'Ar':.9770993}
else:
gas.TPX = target_temp[i],pressure*101325,{'Ar':.99}
tottal_k_numerator = []
for secondary_reaction in reactions_in_cti_file_with_these_reactants_numerator:
reaction_number_in_cti = reactions_in_cti_file.index(secondary_reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k = k*1000000
tottal_k_numerator.append(k)
#check and make sure we are subtracting in the correct order
k_numerator=sum(tottal_k_numerator)
tottal_k_denominator = []
for secondary_reaction in reactions_in_cti_file_with_these_reactants_denominator:
reaction_number_in_cti = reactions_in_cti_file.index(secondary_reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k = k*1000000
tottal_k_denominator.append(k)
k_denominator=sum(tottal_k_denominator)
k = k_numerator/k_denominator
difference = np.log(target_k[i]) - np.log(k)
#print(k_numerator,k_denominator)
##print(target_k[i],k)
diff_in_ks_for_Y.append(difference)
#I guess i could append the tuple
Y_df_list.append(reaction)
Y_values.append(difference)
elif '(+)' in reaction and '/' not in reaction and '*' not in reaction:
list_of_reactions = reaction.split('(+)')
list_of_reactions_cleaned=[]
for reaction in list_of_reactions:
reaction = reaction.rstrip()
reaction = reaction.lstrip()
list_of_reactions_cleaned.append(reaction)
reactions_in_cti_file_with_these_reactants = list_of_reactions_cleaned
if target_press[i] == 0:
pressure = 1e-9
else:
pressure = target_press[i]
if bath_gas[i] !=0:
gas.TPX = target_temp[i],pressure*101325,{'H2O':.013,'O2':.0099,'H':.0000007,'Ar':.9770993}
else:
gas.TPX = target_temp[i],pressure*101325,{'Ar':.99}
tottal_k = []
for secondary_reaction in reactions_in_cti_file_with_these_reactants:
reaction_number_in_cti = reactions_in_cti_file.index(secondary_reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k= k*1000000
tottal_k.append(k)
#check and make sure we are subtracting in the correct order
k=sum(tottal_k)
difference = np.log(target_k[i]) - np.log(k)
diff_in_ks_for_Y.append(difference)
#I guess i could append the tuple
Y_df_list.append(reaction)
Y_values.append(difference)
elif reaction == 'More Complex Combination Rule':
print('do someting else ')
k_targets_for_y = np.array(diff_in_ks_for_Y)
k_targets_for_y = k_targets_for_y.reshape((k_targets_for_y.shape[0],1))
Y_values = np.array(Y_values)
Y_df_temp = pd.DataFrame({'value': Y_df_list,'ln_difference': Y_values.reshape((Y_values.shape[0],))})
Y_data_Frame = Y_data_Frame.append(Y_df_temp, ignore_index=True)
#print(k_targets_for_y.shape,'k targets for y')
return k_targets_for_y,Y_data_Frame
def target_values_for_Z(self,target_value_csv,z_data_Frame):
z_over_w = []
sigma = []
target_value_csv = pd.read_csv(target_value_csv)
target_ln_uncertainty = target_value_csv['ln_unc_k']
target_W = target_value_csv['W']
target_reactions = target_value_csv['Reaction']
z_df_list=[]
z_values = []
for i,value in enumerate(target_ln_uncertainty):
temp = np.divide(value,target_W[i])
sigma.append(value)
z_over_w.append(temp)
z_values.append(temp)
z_df_list.append(target_reactions[i])
k_targets_for_z = np.array(z_over_w)
sigma = np.array(sigma)
sigma = sigma.reshape((sigma.shape[0],1))
z_values = np.array(z_values)
k_targets_for_z = k_targets_for_z.reshape((k_targets_for_z.shape[0],1))
Z_data_Frame_temp = pd.DataFrame({'value': z_df_list,'Uncertainty': z_values.reshape((z_values.shape[0],))})
z_data_Frame = z_data_Frame.append(Z_data_Frame_temp, ignore_index=True)
return k_targets_for_z,sigma,z_data_Frame
def target_values_for_S(self,target_value_csv,
exp_dict_list,
S_matrix,
master_equation_reaction_list = [],
master_equation_sensitivites = {}):
target_value_csv = pd.read_csv(target_value_csv)
target_reactions = target_value_csv['Reaction']
target_temp = target_value_csv['temperature']
target_press = target_value_csv['pressure']
target_k = target_value_csv['k']
bath_gas = target_value_csv['M']
reactions_in_cti_file = exp_dict_list[0]['simulation'].processor.solution.reaction_equations()
number_of_reactions_in_cti = len(reactions_in_cti_file)
gas = ct.Solution(exp_dict_list[0]['simulation'].processor.cti_path)
As = []
Ns = []
Eas = []
flatten = lambda *n: (e for a in n
for e in (flatten(*a) if isinstance(a, (tuple, list)) else (a,)))
flattened_master_equation_reaction_list = list(flatten(master_equation_reaction_list))
coupled_reaction_list = []
list_of_reaction_tuples = []
for reaction in master_equation_reaction_list:
if type(reaction)==tuple:
list_of_reaction_tuples.append(reaction)
for secondary_reaction in reaction:
coupled_reaction_list.append(secondary_reaction)
def reactants_in_master_equation_reactions(flattened_master_equation_reaction_list):
reactants = []
for me_reaction in flattened_master_equation_reaction_list:
reactants_in_master_equation_reaction = me_reaction.split('<=>')[0].rstrip()
reactants.append(reactants_in_master_equation_reaction)
if len(reactants_in_master_equation_reaction.split('+')) >1:
reverse_reactants_in_target_reaction = reactants_in_master_equation_reaction.split('+')
temp = reverse_reactants_in_target_reaction[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction = temp
reactants.append(reverse_reactants_in_target_reaction)
return reactants
master_equation_reactants_and_reverse_reactants = reactants_in_master_equation_reactions(flattened_master_equation_reaction_list)
#print(master_equation_reactants_and_reverse_reactants)
def calculate_weighting_factor_summation(rate_constant_list,gas,temperature,Press,bath_gas):
if Press == 0:
pressure = 1e-9
else:
pressure = Press
if bath_gas !=0:
gas.TPX = temperature,pressure*101325,{'H2O':.013,'O2':.0099,'H':.0000007,'Ar':.9770993}
else:
gas.TPX = temperature,pressure*101325,{'Ar':.99}
tottal_k = []
original_rc_dict = {}
for reaction in rate_constant_list:
reaction_number_in_cti = reactions_in_cti_file.index(reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k = k*1000000
original_rc_dict[reaction] = k
tottal_k.append(k)
#check and make sure we are subtracting in the correct order
k_summation=sum(tottal_k)
weighting_factor_dict = {}
for reaction in rate_constant_list:
weighting_factor_dict[reaction] = original_rc_dict[reaction] / k_summation
return weighting_factor_dict
def calculate_weighting_factor_summation_with_denominator(numerator_rate_constant_list,denominator_rate_constant_list,gas,temperature,Press,bath_gas):
if Press == 0:
pressure = 1e-9
else:
pressure = Press
if bath_gas !=0:
gas.TPX = temperature,pressure*101325,{'H2O':.013,'O2':.0099,'H':.0000007,'Ar':.9770993}
else:
gas.TPX = temperature,pressure*101325,{'Ar':.99}
tottal_k_numerator = []
original_rc_dict = {}
for reaction in numerator_rate_constant_list:
reaction_number_in_cti = reactions_in_cti_file.index(reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k = k*1000000
original_rc_dict[reaction] = k
tottal_k_numerator.append(k)
#check and make sure we are subtracting in the correct order
k_summation_numerator=sum(tottal_k_numerator)
weighting_factor_dict_numerator = {}
for reaction in numerator_rate_constant_list:
weighting_factor_dict_numerator[reaction] = original_rc_dict[reaction] / k_summation_numerator
tottal_k_denominator = []
original_rc_dict = {}
for reaction in denominator_rate_constant_list:
reaction_number_in_cti = reactions_in_cti_file.index(reaction)
coeff_sum = sum(gas.reaction(reaction_number_in_cti).reactants.values())
k = gas.forward_rate_constants[reaction_number_in_cti]
if coeff_sum==1:
k=k
elif coeff_sum==2:
k = k*1000
elif coeff_sum==3:
k = k*1000000
original_rc_dict[reaction] = k
tottal_k_denominator.append(k)
#check and make sure we are subtracting in the correct order
k_summation_denominator=sum(tottal_k_denominator)
weighting_factor_dict_denominator = {}
for reaction in denominator_rate_constant_list:
weighting_factor_dict_denominator[reaction] = -(original_rc_dict[reaction] / k_summation_denominator)
reactions_in_common = weighting_factor_dict_numerator.keys() & weighting_factor_dict_denominator.keys()
weighting_factor_dict = {}
for reaction in reactions_in_common:
weighting_factor_dict[reaction] = weighting_factor_dict_numerator[reaction] + weighting_factor_dict_denominator[reaction]
for reaction in weighting_factor_dict_numerator.keys():
if reaction in reactions_in_common:
pass
else:
weighting_factor_dict[reaction] = weighting_factor_dict_numerator[reaction]
for reaction in weighting_factor_dict_denominator.keys():
if reaction in reactions_in_common:
pass
else:
weighting_factor_dict[reaction] = weighting_factor_dict_denominator[reaction]
return weighting_factor_dict
def add_tuple_lists(nested_list,master_euqation_reactions_list):
if any(isinstance(x, tuple) for x in master_euqation_reactions_list) == False:
return nested_list
else:
all_tuple_summations = []
indexes_that_need_to_be_removed = []
indexes_to_replace_with = []
counter = 0
for i,reaction in enumerate(master_euqation_reactions_list):
if type(reaction) == str:
counter+=1
elif type(reaction) == tuple:
tuple_sublist=[]
indexes_to_replace_with.append(counter)
for j,secondary_reaction in enumerate(reaction):
tuple_sublist.append(np.array(nested_list[counter]))
if j!= 0:
indexes_that_need_to_be_removed.append(counter)
counter+=1
sum_of_tupe_sublist = list(sum(tuple_sublist))
all_tuple_summations.append(sum_of_tupe_sublist)
new_nested_list = copy.deepcopy(nested_list)
for i,replacment in enumerate(indexes_to_replace_with):
new_nested_list[replacment] = all_tuple_summations[i]
new_nested_list = [x for i,x in enumerate(new_nested_list) if i not in indexes_that_need_to_be_removed]
return new_nested_list
def create_empty_nested_reaction_list():
nested_reaction_list = [[] for x in range(len(flattened_master_equation_reaction_list))]
for reaction in flattened_master_equation_reaction_list:
for i,MP in enumerate(master_equation_sensitivites[reaction]):
nested_reaction_list[flattened_master_equation_reaction_list.index(reaction)].append(0)
return nested_reaction_list
def create_tuple_list(array_of_sensitivities):
tuple_list = []
for ix,iy in np.ndindex(array_of_sensitivities.shape):
tuple_list.append((ix,iy))
return tuple_list
def check_if_M_in_reactants(list_to_append_to,
gas,
reactants_in_target_reactions,
reverse_reactants_in_target_reaction):
if reverse_reactants_in_target_reaction !=None:
for reaction_number_in_cti_file in range(gas.n_reactions):
if (gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' (+M)' or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction + ' (+M)' or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' + M' or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction + ' + M') :
list_to_append_to.append(reactions_in_cti_file[reaction_number_in_cti_file])
elif reverse_reactants_in_target_reaction==None:
for reaction_number_in_cti_file in range(gas.n_reactions):
if (gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or
gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' (+M)' or
gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions + ' + M'):
list_to_append_to.append(reactions_in_cti_file[reaction_number_in_cti_file])
return list_to_append_to
def check_if_reaction_is_theory_or_not(reaction):
is_reaction_in_master_equation_list = False
is_reacton_in_normal_reaction_list = False
if '/' in reaction:
#check numerator and denominator
reactants_in_numerator = reaction.split('/')[0].rstrip()
reactants_in_numerator = reactants_in_numerator.lstrip()
reactants_in_denominator = reaction.split('/')[1].rstrip()
reactants_in_denominator = reactants_in_denominator.lstrip()
if '*' in reactants_in_numerator and '(+)' not in reactants_in_numerator:
reactions_in_numerator_with_these_reactants = []
#might be a more comprehensive way to do this
reactants_in_target_reactions = reaction.split('<=>')[0].rstrip()
reverse_reactants_in_target_reaction=None
if len(reactants_in_target_reactions.split('+'))>1:
reverse_reactants_in_target_reaction = reactants_in_target_reactions.split('+')
temp = reverse_reactants_in_target_reaction[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction = temp
for reaction_number_in_cti_file in range(gas.n_reactions):
if gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction:
reactions_in_numerator_with_these_reactants.append(reactions_in_cti_file[reaction_number_in_cti_file])
reactions_in_numerator_with_these_reactants = check_if_M_in_reactants(reactions_in_numerator_with_these_reactants,
gas,
reactants_in_target_reactions,
reverse_reactants_in_target_reaction)
elif '(+)' in reactants_in_numerator and '*' not in reactants_in_numerator:
list_of_reactions_in_numerator = reactants_in_numerator.split('(+)')
list_of_reactions_in_numerator_cleaned=[]
for reaction in list_of_reactions_in_numerator:
reaction = reaction.rstrip()
reaction = reaction.lstrip()
list_of_reactions_in_numerator_cleaned.append(reaction)
reactions_in_numerator_with_these_reactants = list_of_reactions_in_numerator_cleaned
elif '(+)' in reactants_in_numerator and '*' in reactants_in_numerator:
print('need to make rule')
else:
reactions_in_numerator_with_these_reactants = []
reactions_in_numerator_with_these_reactants.append(reactants_in_numerator)
#check reactants in numerator
if '*' in reactants_in_denominator and '(+)' not in reactants_in_denominator:
reactions_in_denominator_with_these_reactants = []
#might be a more comprehensive way to do this
reactants_in_target_reactions = reaction.split('<=>')[0].rstrip()
reverse_reactants_in_target_reaction=None
if len(reactants_in_target_reactions.split('+'))>1:
reverse_reactants_in_target_reaction = reactants_in_target_reactions.split('+')
temp = reverse_reactants_in_target_reaction[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction = temp
for reaction_number_in_cti_file in range(gas.n_reactions):
if gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction:
reactions_in_denominator_with_these_reactants.append(reactions_in_cti_file[reaction_number_in_cti_file])
reactions_in_denominator_with_these_reactants = check_if_M_in_reactants(reactions_in_denominator_with_these_reactants,
gas,
reactants_in_target_reactions,
reverse_reactants_in_target_reaction)
elif '(+)' in reactants_in_denominator and '*' not in reactants_in_denominator:
list_of_reactions_in_denominator = reactants_in_numerator.split('(+)')
list_of_reactions_in_denominator_cleaned=[]
for reaction in list_of_reactions_in_denominator:
reaction = reaction.rstrip()
reaction = reaction.lstrip()
list_of_reactions_in_denominator_cleaned.append(reaction)
reactions_in_denominator_with_these_reactants = list_of_reactions_in_numerator_cleaned
elif '(+)' in reactants_in_denominator and '*' in reactants_in_denominator:
print('need to make rule')
else:
reactions_in_denominator_with_these_reactants=[]
reactions_in_denominator_with_these_reactants.append(reactants_in_denominator)
reactions_in_numerator_and_denominator = reactions_in_numerator_with_these_reactants+reactions_in_denominator_with_these_reactants
for reaction_check in reactions_in_numerator_and_denominator:
if reaction_check in flattened_master_equation_reaction_list:
is_reaction_in_master_equation_list = True
else:
is_reacton_in_normal_reaction_list = True
if is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==False:
return 'master_equations_only', (reactions_in_numerator_with_these_reactants,reactions_in_denominator_with_these_reactants)
elif is_reaction_in_master_equation_list == False and is_reacton_in_normal_reaction_list==True:
return 'not_master_equations_only', (reactions_in_numerator_with_these_reactants,reactions_in_denominator_with_these_reactants)
elif is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==True:
return 'mixed', (reactions_in_numerator_with_these_reactants,reactions_in_denominator_with_these_reactants)
elif '(+)' in reaction and '/' not in reaction and '*' not in reaction:
list_of_reactions = reaction.split('(+)')
list_of_reactions_cleaned=[]
for reaction in list_of_reactions:
reaction = reaction.rstrip()
reaction = reaction.lstrip()
list_of_reactions_cleaned.append(reaction)
reactions_in_cti_file_with_these_reactants = list_of_reactions_cleaned
for reaction_check in reactions_in_cti_file_with_these_reactants:
if reaction_check in flattened_master_equation_reaction_list:
is_reaction_in_master_equation_list = True
else:
is_reacton_in_normal_reaction_list = True
if is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==False:
return 'master_equations_only', (reactions_in_cti_file_with_these_reactants,)
elif is_reaction_in_master_equation_list == False and is_reacton_in_normal_reaction_list==True:
return 'not_master_equations_only', (reactions_in_cti_file_with_these_reactants,)
elif is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==True:
return 'mixed', (reactions_in_cti_file_with_these_reactants,)
elif '*' in reaction and '/' not in reaction and '(+)' not in reaction:
reactions_in_cti_file_with_these_reactants = []
#might be a more comprehensive way to do this
reactants_in_target_reactions = reaction.split('<=>')[0].rstrip()
reverse_reactants_in_target_reaction=None
if len(reactants_in_target_reactions.split('+'))>1:
reverse_reactants_in_target_reaction = reactants_in_target_reactions.split('+')
temp = reverse_reactants_in_target_reaction[1] + ' '+ '+' +' '+ reverse_reactants_in_target_reaction[0]
temp = temp.lstrip()
temp = temp.rstrip()
reverse_reactants_in_target_reaction = temp
for reaction_number_in_cti_file in range(gas.n_reactions):
if gas.reactants(reaction_number_in_cti_file) == reactants_in_target_reactions or gas.reactants(reaction_number_in_cti_file) == reverse_reactants_in_target_reaction:
reactions_in_cti_file_with_these_reactants.append(reactions_in_cti_file[reaction_number_in_cti_file])
reactions_in_cti_file_with_these_reactants = check_if_M_in_reactants(reactions_in_cti_file_with_these_reactants,
gas,
reactants_in_target_reactions,
reverse_reactants_in_target_reaction)
for reaction_check in reactions_in_cti_file_with_these_reactants:
if reaction_check in flattened_master_equation_reaction_list:
is_reaction_in_master_equation_list = True
else:
is_reacton_in_normal_reaction_list = True
if is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==False:
return 'master_equations_only', (reactions_in_cti_file_with_these_reactants,)
elif is_reaction_in_master_equation_list == False and is_reacton_in_normal_reaction_list==True:
return 'not_master_equations_only', (reactions_in_cti_file_with_these_reactants,)
elif is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==True:
return 'mixed', (reactions_in_cti_file_with_these_reactants,)
else:
#normal reaction
reactions_in_cti_file_with_these_reactants=[]
for reaction_check in [reaction]:
if reaction_check in flattened_master_equation_reaction_list:
is_reaction_in_master_equation_list = True
else:
is_reacton_in_normal_reaction_list = True
reactions_in_cti_file_with_these_reactants.append(reaction)
if is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==False:
return 'master_equations_only', (reactions_in_cti_file_with_these_reactants,)
elif is_reaction_in_master_equation_list == False and is_reacton_in_normal_reaction_list==True:
return 'not_master_equations_only', (reactions_in_cti_file_with_these_reactants,)
elif is_reaction_in_master_equation_list == True and is_reacton_in_normal_reaction_list==True:
return 'mixed', (reactions_in_cti_file_with_these_reactants,)
MP_stack = []
target_values_to_stack = []
for i,reaction in enumerate(target_reactions):
type_of_reaction, reaction_tuple = check_if_reaction_is_theory_or_not(reaction)
if type_of_reaction== 'master_equations_only':
if len(reaction_tuple)==1:
if len(reaction_tuple[0])==1:
nested_reaction_list = create_empty_nested_reaction_list()
for j, MP_array in enumerate(master_equation_sensitivites[reaction]):
tuple_list = create_tuple_list(MP_array)
temp = []
counter = 0
for sensitivity in np.nditer(MP_array,order='C'):
k = tuple_list[counter][0]
l= tuple_list[counter][1]
counter +=1
#need to add reduced p and t, and check these units were using to map
#these might not work
t_alpha= meq.Master_Equation.chebyshev_specific_poly(self,k,meq.Master_Equation.calc_reduced_T(self,target_temp[i]))
if target_press[i] ==0:
target_press_new = 1e-9
else:
target_press_new=target_press[i]
p_alpha = meq.Master_Equation.chebyshev_specific_poly(self,l,meq.Master_Equation.calc_reduced_P(self,target_press_new*101325))
#these might nowt work
single_alpha_map = t_alpha*p_alpha*sensitivity
temp.append(single_alpha_map)
temp =sum(temp)
#should there be an = temp here
#nested_reaction_list[master_equation_reaction_list.index(reaction)][j]=temp
nested_reaction_list[flattened_master_equation_reaction_list.index(reaction)][j]=temp
temp2 = nested_reaction_list
temp2_summed = add_tuple_lists(temp2,master_equation_reaction_list)
flat_list = [item for sublist in temp2_summed for item in sublist]
#print(flat_list)
MP_stack.append(nested_reaction_list)
flat_list = np.array(flat_list)
flat_list = flat_list.reshape((1,flat_list.shape[0]))
target_values_to_stack.append(flat_list)
elif len(reaction_tuple[0])>1:
reactions_in_cti_file_with_these_reactants = reaction_tuple[0]
weighting_factor_dictonary = calculate_weighting_factor_summation(reactions_in_cti_file_with_these_reactants,
gas,
target_temp[i],
target_press[i],
bath_gas[i])
nested_reaction_list = create_empty_nested_reaction_list()
for secondary_reaction in reactions_in_cti_file_with_these_reactants:
for j, MP_array in enumerate(master_equation_sensitivites[secondary_reaction]):
tuple_list = create_tuple_list(MP_array)
temp = []
counter = 0
for sensitivity in np.nditer(MP_array,order='C'):
k = tuple_list[counter][0]
l= tuple_list[counter][1]
counter +=1
#need to add reduced p and t, and check these units were using to map
#these might not work
t_alpha= meq.Master_Equation.chebyshev_specific_poly(self,k,meq.Master_Equation.calc_reduced_T(self,target_temp[i]))
if target_press[i] ==0:
target_press_new = 1e-9
else:
target_press_new=target_press[i]
p_alpha = meq.Master_Equation.chebyshev_specific_poly(self,l,meq.Master_Equation.calc_reduced_P(self,target_press_new*101325))
#these might nowt work
single_alpha_map = t_alpha*p_alpha*sensitivity
temp.append(single_alpha_map)
temp =sum(temp)
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)][j]=temp
sub_array_to_apply_weighting_factor_to = list(np.array(nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)])*weighting_factor_dictonary[secondary_reaction])
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)] = sub_array_to_apply_weighting_factor_to
temp2 = nested_reaction_list
#print('THIS IS TEMP:',temp2)
temp2_summed = add_tuple_lists(temp2,master_equation_reaction_list)
#print('THIS IS TEMP SUMMED:',temp2_summed)
flat_list = [item for sublist in temp2_summed for item in sublist]
#print(flat_list)
MP_stack.append(nested_reaction_list)
flat_list = np.array(flat_list)
flat_list = flat_list.reshape((1,flat_list.shape[0]))
target_values_to_stack.append(flat_list)
elif len(reaction_tuple)==2:
reactions_in_cti_file_with_these_reactants_numerator = reaction_tuple[0]
reactions_in_cti_file_with_these_reactants_denominator= reaction_tuple[1]
weighting_factor_dictonary = calculate_weighting_factor_summation_with_denominator(reactions_in_cti_file_with_these_reactants_numerator,
reactions_in_cti_file_with_these_reactants_denominator,
gas,
target_temp[i],
target_press[i],
bath_gas[i])
#now need to add to S matrix
for secondary_reaction in (reactions_in_cti_file_with_these_reactants_numerator+reactions_in_cti_file_with_these_reactants_denominator):
for j, MP_array in enumerate(master_equation_sensitivites[secondary_reaction]):
tuple_list = create_tuple_list(MP_array)
temp = []
counter = 0
for sensitivity in np.nditer(MP_array,order='C'):
k = tuple_list[counter][0]
l= tuple_list[counter][1]
counter +=1
#need to add reduced p and t, and check these units were using to map
#these might not work
t_alpha= meq.Master_Equation.chebyshev_specific_poly(self,k,meq.Master_Equation.calc_reduced_T(self,target_temp[i]))
if target_press[i] ==0:
target_press_new = 1e-9
else:
target_press_new=target_press[i]
p_alpha = meq.Master_Equation.chebyshev_specific_poly(self,l,meq.Master_Equation.calc_reduced_P(self,target_press_new*101325))
#these might nowt work
single_alpha_map = t_alpha*p_alpha*sensitivity
temp.append(single_alpha_map)
temp =sum(temp)
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)][j]=temp
sub_array_to_apply_weighting_factor_to = list(np.array(nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)])*weighting_factor_dictonary[secondary_reaction])
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)] = sub_array_to_apply_weighting_factor_to
temp2 = nested_reaction_list
#print('THIS IS TEMP:',temp2)
temp2_summed = add_tuple_lists(temp2,master_equation_reaction_list)
#print('THIS IS TEMP SUMMED:',temp2_summed)
flat_list = [item for sublist in temp2_summed for item in sublist]
#print(flat_list)
MP_stack.append(nested_reaction_list)
flat_list = np.array(flat_list)
flat_list = flat_list.reshape((1,flat_list.shape[0]))
target_values_to_stack.append(flat_list)
elif type_of_reaction== 'not_master_equations_only':
if len(reaction_tuple)==1:
if len(reaction_tuple[0])==1:
A_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
N_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
Ea_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
#decide if this mapping is correct
A_temp[0,reactions_in_cti_file.index(reaction)] = 1
N_temp [0,reactions_in_cti_file.index(reaction)] = np.log(target_temp[i])
Ea_temp[0,reactions_in_cti_file.index(reaction)] = (-1/target_temp[i])
As.append(A_temp)
Ns.append(N_temp)
Eas.append(Ea_temp)
A_temp = A_temp.reshape((1,A_temp.shape[1]))
N_temp = N_temp.reshape((1,N_temp.shape[1]))
Ea_temp = Ea_temp.reshape((1,Ea_temp.shape[1]))
target_values_to_stack.append(np.hstack((A_temp,N_temp,Ea_temp)))
elif len(reaction_tuple[0])>1:
reactions_in_cti_file_with_these_reactants = reaction_tuple[0]
weighting_factor_dictonary = calculate_weighting_factor_summation(reactions_in_cti_file_with_these_reactants,
gas,
target_temp[i],
target_press[i],
bath_gas[i])
A_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
N_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
Ea_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
for secondary_reaction in reactions_in_cti_file_with_these_reactants:
#need to multiply by the weighting factor for the reaction
A_temp[0,reactions_in_cti_file.index(secondary_reaction)] = 1 * weighting_factor_dictonary[secondary_reaction]
N_temp [0,reactions_in_cti_file.index(secondary_reaction)] = np.log(target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
Ea_temp[0,reactions_in_cti_file.index(secondary_reaction)] = (-1/target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
As.append(A_temp)
Ns.append(N_temp)
Eas.append(Ea_temp)
A_temp = A_temp.reshape((1,A_temp.shape[1]))
N_temp = N_temp.reshape((1,N_temp.shape[1]))
Ea_temp = Ea_temp.reshape((1,Ea_temp.shape[1]))
target_values_to_stack.append(np.hstack((A_temp,N_temp,Ea_temp)))
elif len(reaction_tuple)==2:
reactions_in_cti_file_with_these_reactants_numerator = reaction_tuple[0]
reactions_in_cti_file_with_these_reactants_denominator= reaction_tuple[1]
weighting_factor_dictonary = calculate_weighting_factor_summation_with_denominator(reactions_in_cti_file_with_these_reactants_numerator,
reactions_in_cti_file_with_these_reactants_denominator,
gas,
target_temp[i],
target_press[i],
bath_gas[i])
A_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
N_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
Ea_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
for secondary_reaction in (reactions_in_cti_file_with_these_reactants_numerator+reactions_in_cti_file_with_these_reactants_denominator):
if reaction not in flattened_master_equation_reaction_list:
A_temp[0,reactions_in_cti_file.index(secondary_reaction)] = 1 * weighting_factor_dictonary[secondary_reaction]
N_temp [0,reactions_in_cti_file.index(secondary_reaction)] = np.log(target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
Ea_temp[0,reactions_in_cti_file.index(secondary_reaction)] = (-1/target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
As.append(A_temp)
Ns.append(N_temp)
Eas.append(Ea_temp)
A_temp = A_temp.reshape((1,A_temp.shape[1]))
N_temp = N_temp.reshape((1,N_temp.shape[1]))
Ea_temp = Ea_temp.reshape((1,Ea_temp.shape[1]))
target_values_to_stack.append(np.hstack((A_temp,N_temp,Ea_temp)))
elif type_of_reaction== 'mixed':
#need to figure out what is going in here
if len(reaction_tuple) == 1:
reactions_in_cti_file_with_these_reactants = reaction_tuple[0]
weighting_factor_dictonary = calculate_weighting_factor_summation(reactions_in_cti_file_with_these_reactants,
gas,
target_temp[i],
target_press[i],
bath_gas[i])
#fill in respective lists and figure out what to do with them?
A_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
N_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
Ea_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
nested_reaction_list = create_empty_nested_reaction_list()
for secondary_reaction in reactions_in_cti_file_with_these_reactants:
if secondary_reaction not in flattened_master_equation_reaction_list:
A_temp[0,reactions_in_cti_file.index(secondary_reaction)] = 1 * weighting_factor_dictonary[secondary_reaction]
N_temp [0,reactions_in_cti_file.index(secondary_reaction)] = np.log(target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
Ea_temp[0,reactions_in_cti_file.index(secondary_reaction)] = (-1/target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
elif secondary_reaction in flattened_master_equation_reaction_list:
for j, MP_array in enumerate(master_equation_sensitivites[secondary_reaction]):
tuple_list = create_tuple_list(MP_array)
temp = []
counter = 0
for sensitivity in np.nditer(MP_array,order='C'):
k = tuple_list[counter][0]
l= tuple_list[counter][1]
counter +=1
#need to add reduced p and t, and check these units were using to map
#these might not work
t_alpha= meq.Master_Equation.chebyshev_specific_poly(self,k,meq.Master_Equation.calc_reduced_T(self,target_temp[i]))
if target_press[i] ==0:
target_press_new = 1e-9
else:
target_press_new=target_press[i]
p_alpha = meq.Master_Equation.chebyshev_specific_poly(self,l,meq.Master_Equation.calc_reduced_P(self,target_press_new*101325))
#these might nowt work
single_alpha_map = t_alpha*p_alpha*sensitivity
temp.append(single_alpha_map)
temp =sum(temp)
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)][j]=temp
sub_array_to_apply_weighting_factor_to = list(np.array(nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)])*weighting_factor_dictonary[secondary_reaction])
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)] = sub_array_to_apply_weighting_factor_to
temp2 = nested_reaction_list
temp2_summed = add_tuple_lists(temp2,master_equation_reaction_list)
flat_list = [item for sublist in temp2_summed for item in sublist]
MP_stack.append(nested_reaction_list)
flat_list = np.array(flat_list)
flat_list = flat_list.reshape((1,flat_list.shape[0]))
master_equation_stacked = flat_list
As.append(A_temp)
Ns.append(N_temp)
Eas.append(Ea_temp)
A_temp = A_temp.reshape((1,A_temp.shape[1]))
N_temp = N_temp.reshape((1,N_temp.shape[1]))
Ea_temp = Ea_temp.reshape((1,Ea_temp.shape[1]))
A_n_Ea_stacked = (np.hstack((A_temp,N_temp,Ea_temp)))
combined_master_and_A_n_Ea= np.hstack((A_n_Ea_stacked,master_equation_stacked))
target_values_to_stack.append(combined_master_and_A_n_Ea)
elif len(reaction_tuple) == 2:
reactions_in_cti_file_with_these_reactants_numerator = reaction_tuple[0]
reactions_in_cti_file_with_these_reactants_denominator = reaction_tuple[1]
weighting_factor_dictonary = calculate_weighting_factor_summation_with_denominator(reactions_in_cti_file_with_these_reactants_numerator,
reactions_in_cti_file_with_these_reactants_denominator,
gas,
target_temp[i],
target_press[i],
bath_gas[i])
#fill in respective lists and figure out what to do with them?
A_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
N_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
Ea_temp = np.zeros((1,number_of_reactions_in_cti-len(flattened_master_equation_reaction_list)))
nested_reaction_list = create_empty_nested_reaction_list()
for secondary_reaction in (reactions_in_cti_file_with_these_reactants_numerator+reactions_in_cti_file_with_these_reactants_denominator):
if secondary_reaction not in flattened_master_equation_reaction_list:
A_temp[0,reactions_in_cti_file.index(secondary_reaction)] = 1 * weighting_factor_dictonary[secondary_reaction]
N_temp [0,reactions_in_cti_file.index(secondary_reaction)] = np.log(target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
Ea_temp[0,reactions_in_cti_file.index(secondary_reaction)] = (-1/target_temp[i]) * weighting_factor_dictonary[secondary_reaction]
elif secondary_reaction in flattened_master_equation_reaction_list:
for j, MP_array in enumerate(master_equation_sensitivites[secondary_reaction]):
tuple_list = create_tuple_list(MP_array)
temp = []
counter = 0
for sensitivity in np.nditer(MP_array,order='C'):
k = tuple_list[counter][0]
l= tuple_list[counter][1]
counter +=1
#need to add reduced p and t, and check these units were using to map
#these might not work
t_alpha= meq.Master_Equation.chebyshev_specific_poly(self,k,meq.Master_Equation.calc_reduced_T(self,target_temp[i]))
if target_press[i] ==0:
target_press_new = 1e-9
else:
target_press_new=target_press[i]
p_alpha = meq.Master_Equation.chebyshev_specific_poly(self,l,meq.Master_Equation.calc_reduced_P(self,target_press_new*101325))
#these might nowt work
single_alpha_map = t_alpha*p_alpha*sensitivity
temp.append(single_alpha_map)
temp =sum(temp)
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)][j]=temp
sub_array_to_apply_weighting_factor_to = list(np.array(nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)])*weighting_factor_dictonary[secondary_reaction])
nested_reaction_list[flattened_master_equation_reaction_list.index(secondary_reaction)] = sub_array_to_apply_weighting_factor_to
temp2 = nested_reaction_list
temp2_summed = add_tuple_lists(temp2,master_equation_reaction_list)
flat_list = [item for sublist in temp2_summed for item in sublist]
MP_stack.append(nested_reaction_list)
flat_list = np.array(flat_list)
flat_list = flat_list.reshape((1,flat_list.shape[0]))
master_equation_stacked = flat_list
As.append(A_temp)
Ns.append(N_temp)
Eas.append(Ea_temp)
A_temp = A_temp.reshape((1,A_temp.shape[1]))
N_temp = N_temp.reshape((1,N_temp.shape[1]))
Ea_temp = Ea_temp.reshape((1,Ea_temp.shape[1]))
A_n_Ea_stacked = (np.hstack((A_temp,N_temp,Ea_temp)))
combined_master_and_A_n_Ea= np.hstack((A_n_Ea_stacked,master_equation_stacked))
target_values_to_stack.append(combined_master_and_A_n_Ea)
S_matrix = S_matrix
shape_s = S_matrix.shape
S_target_values = []
#print(target_values_to_stack,target_values_to_stack[0].shape)
#this whole part needs to be edited
for i,row in enumerate(target_values_to_stack):
type_of_reaction, reaction_tuple = check_if_reaction_is_theory_or_not(target_reactions[i])
if type_of_reaction=='master_equations_only':
#zero_to_append_infront = np.zeros((1,((number_of_reactions_in_cti-len(master_equation_reaction_list))*3)))
zero_to_append_infront = np.zeros((1,((number_of_reactions_in_cti-len(flattened_master_equation_reaction_list))*3)))
zero_to_append_behind = np.zeros((1, shape_s[1] - ((number_of_reactions_in_cti-len(flattened_master_equation_reaction_list))*3) - np.shape(row)[1] ))
temp_array = np.hstack((zero_to_append_infront,row,zero_to_append_behind))
S_target_values.append(temp_array)
elif type_of_reaction=='not_master_equations_only':
zero_to_append_behind = np.zeros((1,shape_s[1]-np.shape(row)[1]))
temp_array = np.hstack((row,zero_to_append_behind))
S_target_values.append(temp_array)
elif type_of_reaction=='mixed':
zero_to_append_behind = np.zeros((1,shape_s[1]-np.shape(row)[1]))
temp_array = np.hstack((row,zero_to_append_behind))
S_target_values.append(temp_array)
S_target_values = np.vstack((S_target_values))
return S_target_values
def preprocessing_rate_constant_target_csv(self,target_value_csv,
master_equation_reactions):
#split up the master equations into multiple data frames
master_equation_df_list = []
master_equation_df_sorted_list = []
df_summation_list = []
for reaction in master_equation_reactions:
if type(reaction) == tuple:
master_equation_df_list.append([])
master_equation_df_sorted_list.append([])
df_ttl = pd.read_csv(target_value_csv)
counter = 0
for reaction in master_equation_reactions:
if type(reaction) == tuple:
for secondary_reaction in reaction:
temp = df_ttl.loc[df_ttl['Reaction'] == secondary_reaction]
if not temp.empty:
master_equation_df_list[counter].append(temp)
counter +=1
for i,lst in enumerate(master_equation_df_list):
for j,df in enumerate(lst):
df = df.sort_values(["temperature", "pressure"], ascending = (True, True))
master_equation_df_sorted_list[i].append(df)
for i,lst in enumerate(master_equation_df_sorted_list):
df_summation = pd.DataFrame()
df_summation['Reaction'] = lst[0]['Reaction']
df_summation['temperature'] = lst[0]['temperature']
df_summation['pressure'] = lst[0]['pressure']
df_summation['M'] = lst[0]['M']
df_summation['ln_unc_k'] = lst[0]['ln_unc_k']
df_summation['W'] = lst[0]['W']
k_summation_list=[]
for j,df in enumerate(lst):
k_summation_list.append(df['k'].to_numpy())
df_summation['k'] = sum(k_summation_list)
df_summation_list.append(df_summation)
reactions_to_remove = []
for reaction in master_equation_reactions:
if type(reaction) == tuple:
for secondary_reaction in reaction:
reactions_to_remove.append(secondary_reaction)
df_cleaned = df_ttl[~df_ttl['Reaction'].isin(reactions_to_remove)]
df_concat_list = [df_cleaned]+ df_summation_list
df_new_tottal = pd.concat(df_concat_list)
new_file_name = target_value_csv[:-4]
new_file_path = new_file_name+'_combined_channels.csv'
df_new_tottal.to_csv(new_file_path,
index=False)
return df_new_tottal,new_file_path
def appending_target_values(self,target_values_for_z,
target_values_for_Y,
target_values_for_S,
sigma_target_values,
S_matrix,
Y_matrix,
z_matrix,
sigma):
z_matrix = np.vstack((z_matrix ,target_values_for_z))
Y_matrix = np.vstack((Y_matrix,target_values_for_Y))
S_matrix = np.vstack((S_matrix,target_values_for_S))
sigma = np.vstack((sigma,sigma_target_values))
self.S_matrix = S_matrix
self.Y_matrix = Y_matrix
self.z_matrix = z_matrix
self.sigma = sigma
return S_matrix,Y_matrix,z_matrix,sigma
| 253,617 | 70,321 |
from django.contrib.auth.backends import ModelBackend
import re
from users.models import User
from django.db.models import Q
class MeiduoModelBackend(ModelBackend):
def authenticate(self, request, username=None, password=None, **kwargs):
if request is None:
try:
user = User.objects.get(Q(username=username) | Q(mobile=username), is_staff=True)
except:
return None
if user.check_password(password):
return user
else:
try:
user = User.objects.get(Q(username=username) | Q(mobile=username))
except:
return None
if user.check_password(password):
return user
| 745 | 188 |
#!/usr/bin/env python3
from flask import Flask, request
from datetime import datetime
from flask_opentracing import FlaskTracing
from jaeger_client import Config
app = Flask(__name__)
config = Config(config=
{
'sampler': {'type': 'const', 'param': 1},
'local_agent':
{'reporting_host': '172.2.1.5'}
},
service_name='api_rst_getter'
)
jaeger_tracer = config.initialize_tracer()
tracing = FlaskTracing(jaeger_tracer, True, app)
@app.route('/', methods=['GET', 'POST'])
def get_header():
now = datetime.now()
print(now)
file = open('./api_header.log', 'a')
req_header = request.headers.values()
time = '\n' + str(now) + '\n'
file.write(time)
req_body = request.values
for items in req_header:
file.write(' - ')
print(items)
file.write(items)
file.write('\n')
for items in req_body:
file.write(' - ')
print(items, ': ', req_body[items])
item = str(items)+': '
file.write(item)
file.write(req_body[items])
file.close()
return "it is what it is"
@app.route('/test')
def test():
print("This is a test method")
return ('Yooohooo, you\'re connected to backend\nv2')
if __name__ == '__main__':
app.run(host='0.0.0.0', debug=False)
| 1,278 | 466 |
from math import prod
def compute(n: int, k: int) -> float:
return round(7 * (1 - prod((i - k) / i for i in range(n // 7 * 6 + 1, n + 1))), 9)
| 149 | 66 |
import unittest
from collections.abc import (
Mapping,
)
from uuid import (
UUID,
uuid4,
)
from minos.common import (
DeclarativeModel,
Field,
Model,
)
from tests.model_classes import (
FooBar,
)
class TestModel(unittest.TestCase):
def test_base(self):
self.assertTrue(issubclass(Model, Mapping))
def test_fields(self):
uuid = uuid4()
model = FooBar(uuid)
self.assertEqual({"identifier": Field("identifier", UUID, uuid)}, model.fields)
def test_eq_reversing(self):
class _Fake(DeclarativeModel):
def __eq__(self, other):
return True
self.assertEqual(FooBar(uuid4()), _Fake())
self.assertEqual(_Fake(), FooBar(uuid4()))
if __name__ == "__main__":
unittest.main()
| 799 | 265 |
from tweepy.streaming import StreamListener
from tweepy import OAuthHandler
from tweepy import Stream
import json
import threading
import sys
import pygame
import os
if len(sys.argv) == 3:
search_strings = [sys.argv[1],sys.argv[2]]
else:
print("Usage: twitterbattlegame.py [TREND1_STRING] [TREND2_STRING]")
sys.exit(0)
# Go to http://apps.twitter.com and create an app.
# The consumer key and secret will be generated for you after
consumer_key=""
consumer_secret=""
# After the step above, you will be redirected to your app's page.
# Create an access token under the the "Your access token" section
access_token=""
access_token_secret=""
# This is the string to search in the twitter feed
# May be a word, an #hashtag or a @username
twitterText = ""
text_x = 30
color = 1
dwarfGo = False
gladiatorGo = False
finish = False
# final animation
dwarfdirection = -1
dwarfmove = 0
gladiatordirection = -1
gladiatormove = 0
def startTwitter():
l = StdOutListener()
auth = OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
stream = Stream(auth, l)
stream.filter(track=search_strings)
class StdOutListener(StreamListener):
def on_data(self, data):
global twitterText
global first
global text_x
global color
global dwarfGo
global gladiatorGo
data = json.loads(data)
twitterText = data['text'].lower()
if search_strings[0] in twitterText:
dwarfGo = True
if search_strings[1] in twitterText:
gladiatorGo = True
return True
def on_error(self, status):
return False
def get_sprite(image, x, y, width, height):
sprite = pygame.Surface([width, height], pygame.SRCALPHA, 32).convert_alpha()
sprite.blit(image, (0, 0), (x, y, width, height))
return sprite
def chunkstring(string, length):
return (string[0+i:length+i] for i in range(0, len(string), length))
twitterThread = threading.Thread(target = startTwitter)
twitterThread.start()
pygame.init()
clock = pygame.time.Clock()
size = width, height = 1056, 672
screen = pygame.display.set_mode(size)
# fonts
font = pygame.font.Font('./assets/PressStart2P-Regular.ttf', 16)
fontTitles = pygame.font.Font('./assets/PressStart2P-Regular.ttf', 32)
# default text from twitter
text = font.render(twitterText.encode('utf-8'), True, (0,0,0))
textRect = text.get_rect()
# info texts
textTile = fontTitles.render("Twitter #hashtags battle!", True, (100,250,80))
textTileRect = textTile.get_rect()
textTileRect.center = (520,40)
hashtagText = fontTitles.render(sys.argv[1]+" VS "+sys.argv[2], True, (250,50,250))
hashtagTextRect = hashtagText.get_rect()
hashtagTextRect.center = (520,100)
# set background
background = pygame.image.load("./assets/bulkhead-wallsx3.png")
backgroundRect = background.get_rect()
# set dwarf sprites
dwarfSpritesSheet = pygame.image.load("./assets/Dwarf_Sprite_Sheet1.2v-4x.png")
dwarfSprites = []
dwarfSpritesNumber = 4
for i in range(dwarfSpritesNumber):
dwarfSprites.append(get_sprite(dwarfSpritesSheet,150 * i,640,150,100))
dwarfRect = pygame.Rect(50,470,128,128)
dwarfSpritePos = 0
# set gladiator sprites
gladiatorSpritesSheet = pygame.image.load("./assets/Gladiator-Sprite Sheet-Left4x.png")
gladiatorSprites = []
gladiatorSpritesNumber = 5
for i in range(gladiatorSpritesNumber):
gladiatorSprites.append(get_sprite(gladiatorSpritesSheet,128 * i,0,128,128))
gladiatorRect = pygame.Rect(874,430,128,128)
gladiatorSpritePos = 0
# set key
collectablesSpritesSheet = pygame.image.load("./assets/Dungeon Collectables4x.png")
keySprites = []
keySpritesNumber = 12
for i in range(keySpritesNumber):
keySprites.append(get_sprite(collectablesSpritesSheet,64 * i,260,64,64))
keyRect = pygame.Rect(496,490,64,64)
keySpritePos = 0
# set box and money
box = pygame.image.load("./assets/box.png")
boxRect = box.get_rect()
boxRect.center = (523,510)
money = pygame.image.load("./assets/money.png")
moneyRect = money.get_rect()
moneyRect.center = (523,520)
while 1:
clock.tick(24)
for event in pygame.event.get():
if event.type == pygame.QUIT:
pygame.quit()
os._exit(1)
if event.type == pygame.KEYDOWN:
# key control (for testing)
if event.key == pygame.K_LEFT:
dwarfRect = dwarfRect.move(-10,0)
dwarfSpritePos -= 1
if dwarfSpritePos < 0:
dwarfSpritePos = dwarfSpritesNumber - 1
if event.key == pygame.K_RIGHT:
dwarfRect = dwarfRect.move(10,0)
dwarfSpritePos += 1
if dwarfSpritePos > dwarfSpritesNumber -1:
dwarfSpritePos = 0
if event.key == pygame.K_a:
gladiatorRect = gladiatorRect.move(-10,0)
gladiatorSpritePos -= 1
if gladiatorSpritePos < 0:
gladiatorSpritePos = gladiatorSpritesNumber - 1
if event.key == pygame.K_s:
gladiatorRect = gladiatorRect.move(10,0)
gladiatorSpritePos += 1
if gladiatorSpritePos > gladiatorSpritesNumber -1:
gladiatorSpritePos = 0
# draw background
screen.blit(background, backgroundRect)
# automated sprites movement
if dwarfGo == True and finish == False:
#print("ENTRAAAAA")
dwarfGo = False
dwarfRect = dwarfRect.move(10,0)
dwarfSpritePos += 1
if dwarfSpritePos > dwarfSpritesNumber -1:
dwarfSpritePos = 0
# render text
text = font.render(str(twitterText.encode('utf-8'))[:60]+"...", True, (255,0,0))
textRect = text.get_rect()
textRect.x = 20
textRect.y = dwarfRect.y - 200
if gladiatorGo == True and finish == False:
gladiatorGo = False
gladiatorRect = gladiatorRect.move(-10,0)
gladiatorSpritePos -= 1
if gladiatorSpritePos < 0:
gladiatorSpritePos = gladiatorSpritesNumber - 1
# render text
text = font.render(str(twitterText.encode('utf-8'))[:60]+"...", True, (0,0,255))
textRect = text.get_rect()
textRect.x = 20
textRect.y = gladiatorRect.y - 100
# draw tweet
if finish == False:
screen.blit(text,textRect)
# draw texts
screen.blit(textTile,textTileRect)
screen.blit(hashtagText,hashtagTextRect)
# draw box
screen.blit(box,boxRect)
# game ending
if dwarfRect.right > keyRect.left:
# draw money and box
finish = True
screen.blit(money,moneyRect)
dwarfRect = dwarfRect.move(0,dwarfdirection)
if dwarfmove > 10:
dwarfdirection = dwarfdirection * -1
dwarfmove = 0
dwarfmove += 1
winText = fontTitles.render(sys.argv[1]+" WINS!!", True, (0,0,255))
winTextRect = winText.get_rect()
winTextRect.center = (520,200)
screen.blit(winText,winTextRect)
if gladiatorRect.left < keyRect.right:
# draw money and box
finish = True
screen.blit(money,moneyRect)
gladiatorRect = gladiatorRect.move(0,gladiatordirection)
if gladiatormove > 10:
gladiatordirection = gladiatordirection * -1
gladiatormove = 0
gladiatormove += 1
winText = fontTitles.render(sys.argv[2]+" WINS!!", True, (0,0,255))
winTextRect = winText.get_rect()
winTextRect.center = (520,200)
screen.blit(winText,winTextRect)
# draw key
if finish == False:
screen.blit(keySprites[keySpritePos],keyRect)
keySpritePos += 1
if keySpritePos > keySpritesNumber -1:
keySpritePos = 0
# draw dwarf
screen.blit(dwarfSprites[dwarfSpritePos],dwarfRect)
# draw gladiator
screen.blit(gladiatorSprites[gladiatorSpritePos],gladiatorRect)
pygame.display.flip()
| 7,197 | 3,109 |
from rest_framework import permissions
class IsPictureCreatorOrSafeOnly(permissions.BasePermission):
def has_object_permission(self, request, view, obj):
if request.method in permissions.SAFE_METHODS or view.action == 'like':
return True
return obj.uploaded_by == request.user
class IsPictureCommentCreatorOrReadOnly(permissions.BasePermission):
def has_object_permission(self, request, view, obj):
if request.method in permissions.SAFE_METHODS:
return True
return obj.created_by == request.user
| 568 | 160 |
"""
TinyTim model PSF phase retrieval results.
HISTORY:
Created on March 3, 2011
:author: Sami-Matias Niemi
:contact: niemi@stsci.edu
:version: 0.1
"""
import matplotlib
matplotlib.use('PDF')
import pylab as P
import glob as G
import numpy as N
import os
from matplotlib import cm
import SamPy.focus.phaseretrievalresults as ph
__author__ = 'Sami-Matias Niemi'
__version__ = '0.1'
class TinyTimResults(ph.PhaseRetResults):
"""
Expansion to PhaseRetResults class
"""
def findRealFocus(self, file):
parms = './' + os.path.dirname(file) + '/complete_results/parameters.txt'
tmp = open(parms).readlines()
for line in tmp:
if 'Focus' in line:
fcs = float(line.split('|')[1].split()[0].strip())
return fcs
def plotFocusDifference(self, input, title, abs=False):
fig = P.figure()
ax = fig.add_subplot(111)
tmp1 = []
tmp2 = []
tmp3 = []
tmp4 = []
for key in input:
data = input[key]
if abs:
tmp1.append(N.abs(data[1][0] - data[0]))
tmp2.append(N.abs(data[1][1] - data[0]))
tmp3.append(N.abs(data[1][2] - data[0]))
tmp4.append(N.abs(data[1][3] - data[0]))
x1 = ax.plot(N.abs(data[1][0]), N.abs(data[0]), 'bo')
x2 = ax.plot(N.abs(data[1][1]), N.abs(data[0]), 'rs')
x3 = ax.plot(N.abs(data[1][2]), N.abs(data[0]), 'mD')
x4 = ax.plot(N.abs(data[1][2]), N.abs(data[0]), 'g*')
else:
tmp1.append(data[1][0] - data[0])
tmp2.append(data[1][1] - data[0])
tmp3.append(data[1][2] - data[0])
tmp4.append(data[1][3] - data[0])
x1 = ax.plot(data[1][0], data[0], 'bo')
x2 = ax.plot(data[1][1], data[0], 'rs')
x3 = ax.plot(data[1][2], data[0], 'mD')
x4 = ax.plot(data[1][2], data[0], 'g*')
ax.plot([-15, 15], [-15, 15], 'k:')
if abs:
ax.set_xlim(0, 7.5)
ax.set_ylim(0, 7.5)
else:
ax.set_xlim(-7.5, 7.5)
ax.set_ylim(-7.5, 7.5)
P.text(0.5, 1.05, title,
horizontalalignment='center',
verticalalignment='center',
transform=ax.transAxes)
ax.set_xlabel('Phase Retrieval Result [Secondary mirror despace]')
ax.set_ylabel('TinyTim Focus Offset [Secondary mirror despace]')
ax.legend([x1, x2, x3, x4],
['Nominal setup', 'Fixed Blur Kernel', 'Spherical = 0, fixed', 'sph = 0, fixed Blur'],
shadow=True, fancybox=True, numpoints=1,
loc='best')
if abs:
P.savefig('FocusAbsolute.pdf')
else:
P.savefig('Focus.pdf')
#print out statistics
tmp1 = N.array(tmp1)
tmp2 = N.array(tmp2)
tmp3 = N.array(tmp3)
tmp4 = N.array(tmp4)
print 'Average offset (PR - TT) for nominal method is %.2f' % N.mean(tmp1)
print 'while STD is %.3f' % N.std(tmp1)
print
print 'Average offset (PR - TT) for fixed blur kernel is %.2f' % N.mean(tmp2)
print 'while STD is %.3f' % N.std(tmp2)
print
print 'Average offset (PR - TT) for fixed spherical = 0 is %.2f' % N.mean(tmp3)
print 'while STD is %.3f' % N.std(tmp3)
print
print 'Average offset (PR - TT) for fixed blur and sph = 0 is %.2f' % N.mean(tmp4)
print 'while STD is %.3f' % N.std(tmp4)
def plotFocusFieldPosition(self, input, title):
fig = P.figure()
ax = fig.add_subplot(111)
c1, x1, y1 = [], [], []
for key in input:
data = input[key]
c1.append(data[0] - data[1][0][5])
x1.append(data[1][0][1])
y1.append(data[1][0][2])
x1 = ax.scatter(x1, y1, c=c1,
marker='o',
cmap=cm.get_cmap('jet'),
edgecolor='none')
c1 = fig.colorbar(x1, ax=ax, shrink=0.7, fraction=0.05)
c1.set_label('TinyTim Focus Offset - Phase Retrieval Measurement')
ax.set_xlim(0, 4096)
ax.set_ylim(0, 2051)
P.text(0.5, 1.05, title,
horizontalalignment='center',
verticalalignment='center',
transform=ax.transAxes)
P.savefig('FocusFieldPos.pdf')
if __name__ == '__main__':
#define some variables
str = {'file': 0,
'target': 0,
'mjd': 1,
'date': 2,
'time': 3,
'focus': 5}
cameras = ['WFC3']
t1 = 'UVIS1, PSF diameter = 3.0 arcsec, Filter F410M, G5V star, variable field positions'
t2 = 'UVIS1, PSF diameter = 3.0 arcsec, Filter F410M, G5V star'
PR = TinyTimResults(cameras, str)
#find all data files
results = G.glob('t*/results.txt')
#loop over the data files and find the focus info
out = {}
out2 = {}
for file in results:
print 'processing %s' % file
res = PR.readResults(file)
#get the observed values
tmp = []
for x in res:
tmp.append(x[str['focus']])
#find the true focus
trueFocus = PR.findRealFocus(file)
#output values
out[file] = [trueFocus, tmp]
out2[file] = [trueFocus, res]
#generate a plot
PR.plotFocusDifference(out, t1)
PR.plotFocusDifference(out, t1, abs=True)
PR.plotFocusFieldPosition(out2, t2)
| 5,597 | 2,021 |
"""This module defines the ReachyAudioAnswering class."""
import nltk
import json
import torch
import random
import pickle
from nltk.stem.lancaster import LancasterStemmer
stemmer = LancasterStemmer()
CONFIDENCE_THRESHOLD = 0.7
class ReachyAudioAnswering():
"""ReachyAudioAnswering class.
This class implements a small neural network allowing Reachy to answer to
simple questions. To make it flexible, it uses sentence tokenizing and word
stemming such that the network can provide answers to sentences different
to the one used for the training. These input sentences have to remain
close to the training sentences however.
"""
def __init__(self):
"""Train the model of the network or load it if it already exists."""
print("Initializing Reachy answering model...")
# Load the json file containing the training data
with open("utils/intents.json") as myFile:
self.data = json.load(myFile)
# Load the data necessary to the initialization
# of the network if the training has already been
# done before, create it otherwise
try:
with open("utils/data.pickle", "rb") as f:
self.words, self.labels,
train_input, train_target = pickle.load(f)
except:
# Contain all the different stemmed words constituing the patterns
self.words = []
# Contain all the different intents of the input sentences
self.labels = []
# Contain the training sentences of the network
docs_x = []
# Contain the corresponding intent of a tokenized pattern
docs_y = []
# Contain the training inputs of the network
train_input = []
# Contain the expected output for the training of the network
train_target = []
# Extract the data from the json file
for intent in self.data["intents"]:
for pattern in intent["patterns"]:
wrds = nltk.word_tokenize(pattern)
self.words.extend(wrds)
docs_x.append(pattern)
docs_y.append(intent["tag"])
if intent["tag"] not in self.labels:
self.labels.append(intent["tag"])
# Apply word stemming i.e. find the root of the word
# (ex: happened -> happen)
self.words = [stemmer.stem(w.lower()) for w in self.words
if w != "?"]
# transform to set to remove doublons
self.words = sorted(list(set(self.words)))
self.labels = sorted(self.labels)
out_empty = [0 for _ in range(len(self.labels))]
# Transform each training sentence into a bag of words (an input
# for the network) and compute the corresponding expected output
for x, doc in enumerate(docs_x):
bag = self.bag_of_words(doc)
# Expected output
output_row = out_empty[:]
output_row[self.labels.index(docs_y[x])] = 1
# We add the input and the expected output to the training set
train_input.append(bag)
train_target.append(output_row)
# We store the computed training set for future uses
with open("utils/data.pickle", "wb") as f:
pickle.dump((self.words, self.labels,
train_input, train_target), f)
# Load the model if it already exists, train it otherwise
try:
self.model = torch.load('utils/model.pth')
except:
self.model = torch.nn.Sequential(
torch.nn.Linear(len(train_input[0]), 8),
torch.nn.Linear(8, 8),
torch.nn.Linear(8, len(train_target[0])),
torch.nn.Softmax(dim=-1))
self.train_model(torch.Tensor(train_input),
torch.Tensor(train_target))
torch.save(self.model, 'utils/model.pth')
print("Done")
def train_model(self, train_input, train_target, nb_epochs=500,
show_metric=False):
"""Train the model of the network.
:param data_input: The inputs of the training set.
:param data_target: The corresponding outputs of the training set.
:param nb_epochs: The number of times that the learning algorithm will
work through the entire training dataset.
:param show_metric: Allow to show the performance of the model during
his training.
"""
criterion = torch.nn.MSELoss()
optimizer = torch.optim.Adam(self.model.parameters())
for e in range(nb_epochs):
# Compute the output of the model (forward pass)
output = self.model(train_input)
# Compute the error between the predicted output and the ground
# truth
loss = criterion(output, train_target)
# Reset the sum of the gradients (the previous epoch should not
# influence the current epoch)
self.model.zero_grad()
# Apply a backward pass
loss.backward()
# Update the parameters of the model with respect to the backward
# pass previously done
optimizer.step()
# Compute the error of the current state of the network's model
# with respect to the training set
if show_metric:
with torch.no_grad():
print("Epoch {} -> Train error = {:.02f} %".format(
e, self.compute_nb_errors(train_input, train_target) /
train_input.size(0) * 100))
def compute_nb_errors(self, data_input, data_target):
"""Compute the number of classification errors of our network's model.
:param data_input: The inputs of the testing set.
:param data_target: The corresponding outputs of the testing set.
:return: The number of classification errors made on the testing set.
"""
nb_data_errors = 0
# Compute the output of the model
output = self.model(data_input)
# Take the most confident output as the result
predicted_classes = torch.argmax(output, 1)
expected_classes = torch.argmax(data_target, 1)
# Compare the prediction of the model with the ground truth
for predicted_classe, expected_classe in zip(predicted_classes,
expected_classes):
if predicted_classe != expected_classe:
nb_data_errors = nb_data_errors + 1
return nb_data_errors
def bag_of_words(self, input_sentence):
"""Compute a bag of words that will be used as input for the network.
A bag of words is a vector whose length correspond to the "vocabulary"
known by the network (all the different words composing the sentences
of the training set). For each word of the vocabulary, if this word is
present in the input sentence, then the vector contains a 1, otherwise
it contains a 0.
:param input_sentence: The sentence to be answered.
:return: The bag of word corresponding to the input sentence.
"""
bag = []
# Tokenize the input sentence and apply word stemming
# on each of the tokenized words
sentence_words = nltk.word_tokenize(input_sentence)
stemmed_words = [stemmer.stem(word.lower()) for word in sentence_words]
# Fill the vector
for w in self.words:
if w in stemmed_words:
bag.append(1)
else:
bag.append(0)
return bag
def answer(self, input_sentence):
"""Allow Reachy to answer to a question.
:param input_sentence: The sentence to be answered.
:return: The detected intent of the input sentence
(None if the intent could not be detected).
:return: The answer to the input sentence.
"""
# Compute the output of the model with respect to the input sentence
results = self.model(torch.Tensor(self.bag_of_words(input_sentence)))
# Take the most confident output as the result
results_index = torch.argmax(results)
intent = self.labels[results_index]
# Provide an answer only if the network
# was confident enough about his output
if results[results_index] > CONFIDENCE_THRESHOLD:
for tg in self.data["intents"]:
if tg["tag"] == intent:
# The response is picked randomly among the ones
# related to the detected input sentence intent
responses = tg["responses"]
answer = random.choice(responses)
return intent, answer
return None, "I didn't get that, can you try again ?"
| 9,346 | 2,543 |
import numpy as np
def softmax(x):
ex = np.exp(-x)
return ex / np.sum(ex)
def relu(x):
return x * (x > 0.)
def relu_der(x):
return np.ones_like(x) * (x > 0.)
class MLP:
def __init__(self, lr, bs, momentum, verbose, max_iters, eps=0., hidden_dims=[10]):
self.layers = []
self.labels_ = []
self.lr = lr
self.batch_size = bs
self.momentum = momentum
self.verbose = verbose
self.max_iters = max_iters
self.eps = eps
assert len(hidden_dims) > 0
self.hidden_dims = hidden_dims
def _create_layer(self, num_inputs, num_outputs, activate=True):
return {'w':np.random.rand(num_inputs, num_outputs), 'b': np.random.rand(num_outputs), 'a':activate,
'batch_grad_w':np.zeros((num_inputs, num_outputs), dtype=np.float32),
'w_v':np.zeros((num_inputs, num_outputs), dtype=np.float32),
'batch_grad_b':np.zeros(num_outputs, dtype=np.float32),
'b_v':np.zeros(num_outputs, dtype=np.float32)}
def init_layers_(self, num_inputs, num_labels):
np.random.seed(0)
self.layers = []
self.layers.append(self._create_layer(num_inputs, self.hidden_dims[0], True))
for i in range(1, len(self.hidden_dims)):
self.layers.append(self._create_layer(self.hidden_dims[i - 1], self.hidden_dims[i], True))
self.layers.append(self._create_layer(self.hidden_dims[-1], num_labels, False))
def forward_(self, x, train=False):
signal = x
for layer in self.layers:
if train: layer['input'] = np.copy(signal)
signal = np.matmul(np.transpose(layer['w']), signal) + layer['b']
if layer['a']:
if train: layer['pre_output'] = signal
signal = relu(signal)
return signal
def backward_(self, expected, outputs):
for i in reversed(range(len(self.layers))):
current_layer = self.layers[i]
if i == len(self.layers) - 1: # handle the last layer
errors = expected - outputs
current_layer['delta'] = errors
if current_layer['a']:
current_layer['delta'] *= relu_der(current_layer['pre_output'])
else:
next_layer = self.layers[i + 1]
current_layer['delta'] = np.matmul(next_layer['w'], next_layer['delta']) * \
relu_der(current_layer['pre_output'])
current_layer['batch_grad_b'] += current_layer['delta']
current_layer['batch_grad_w'] += np.matmul(current_layer['input'].reshape(-1, 1),
current_layer['delta'].reshape(1, -1))
def update_weights_(self):
for i in reversed(range(len(self.layers))):
current_layer = self.layers[i]
current_layer['b_v'] = self.momentum * current_layer['b_v'] + (self.lr / self.batch_size) * current_layer['batch_grad_b']
current_layer['w_v'] = self.momentum * current_layer['w_v'] + (self.lr / self.batch_size) * current_layer['batch_grad_w']
current_layer['b'] -= current_layer['b_v']
current_layer['w'] -= current_layer['w_v']
def init_train_iter_(self):
for layer in self.layers:
layer['batch_grad_b'] *= 0.
layer['batch_grad_w'] *= 0.
def fit(self, x, y):
num_samples = len(x)
assert num_samples > 0
assert num_samples == len(y)
num_inputs = len(x[0])
assert num_inputs > 0
self.labels_ = np.unique(y)
num_labels = len(self.labels_)
assert num_labels > 0
x = np.array(x)
y = np.array(y)
self.init_layers_(num_inputs, num_labels)
np.random.seed(1)
for i in range(self.max_iters):
batch_indices = np.random.random_integers(0, num_samples - 1, self.batch_size)
batch_x = x[batch_indices]
batch_y = y[batch_indices]
self.init_train_iter_()
for j in range(self.batch_size):
outputs = softmax(self.forward_(batch_x[j], train=True))
idx = np.argmax(outputs)
label = self.labels_[idx]
expected = (self.labels_ == batch_y[j]).astype(np.int8)
self.backward_(expected, outputs)
self.update_weights_()
def predict(self, x):
predictions = np.zeros(len(x))
for i in range(len(x)):
probs = softmax(self.forward_(x[i]))
idx = np.argmax(probs)
predictions[i] = self.labels_[idx]
return predictions
def score(self, x, y):
assert len(x) == len(y)
y = np.array(y).reshape(-1)
predictions = self.predict(x)
num_correct = np.sum(predictions == y)
return float(num_correct) / y.shape[0]
| 4,937 | 1,623 |
import os
import subprocess
from munch import Munch
from .logutil import *
from .decorators import *
if __name__ == '__main__':
print('This module is not executable.')
exit(0)
FileSystems = [
'file',
'hdfs',
's3',
'gs',
'mysql',
'http',
'https'
]
# If a path could match more that one there is uncertainity in the outcome
extensionMapper = {
'.mt': ('mt', None),
'.ht': ('ht', None),
'.vcf': ('vcf', None),
'.vcf.gz': ('vcf', 'gz'),
'.vcf.bgz': ('vcf', 'bgz'),
'.tsv': ('tsv', None),
'.tsv.gz': ('tsv', 'gz'),
'.tsv.bgz': ('tsv', 'bgz'),
'.csv': ('csv', None),
'.csv.gz': ('csv', 'gz'),
'.csv.bgz': ('csv', 'bgz'),
'.json': ('json', None),
'.json.gz': ('json', 'gz'),
'.json.bgz': ('json', 'bgz'),
'.yaml': ('yaml', None),
'.yaml.gz': ('yaml', 'gz'),
'.yaml.bgz': ('yaml', 'bgz'),
'.bed': ('bed', None),
'.bim': ('bim', None),
'.fam': ('fam', None),
'.parquet': ('parquet', None)
}
class Path:
# If true, remove file system prefix (i.e. 'file://' or 'hdfs://') of the defaultFileSystem.
# For example, if 'defaultFileSystem=local' it removes the 'file://' from the path
__defaultMode = True
@classmethod
def SetDefaultMode(cls, defaultMode):
cls.__defaultMode = defaultMode
@classmethod
def GetDefaultMode(cls):
return cls.__defaultMode
# If the path does not have a file system prefix (i.e. 'file://' or 'hdfs://') adds the prefix based on the default file system
__defaultFileSystem = 'file'
@classmethod
def SetDefaultFileSystem(cls, defaultFileSystem):
if defaultFileSystem in FileSystems:
cls.__defaultFileSystem = defaultFileSystem
else:
LogException(f'File system `{defaultFileSystem}` is not supported')
@classmethod
def GetDefaultFileSystem(cls):
return cls.__defaultFileSystem
def __init__(self, path=None):
self.__path = None
self.__raw = None
if path:
self.path = path
def __repr__(self):
rep = dict()
for k in ['raw', 'path', 'fileSystem', 'format', 'compression']:
rep[k] = getattr(self,k)
return str(rep)
@property
def path(self):
if self.GetDefaultMode():
if self.__fileSystem == self.GetDefaultFileSystem():
return self.__path
return '://'.join([self.__fileSystem, self.__path])
@property
def local(self):
return self.__path
@property
def fileSystem(self):
return self.__fileSystem
@property
def raw(self):
return self.__raw
@property
def format(self):
return self.__format
@property
def compression(self):
return self.__compression
@path.setter
def path(self, path):
if isinstance(path, str):
self.__raw = str(path)
self.Processes()
else:
LogExceptionType(path, expectedType='str')
def Processes(self):
# Identify the file system and extract it from the path
rawPath = os.path.expandvars(self.__raw)
if ':' in rawPath:
parts = rawPath.split(':')
if len(parts) > 2:
LogException(f'Path `{rawPath}` has more than one `:`')
elif not parts[0]:
LogException(f'Path `{rawPath}` starts with `:`')
elif parts[0] not in FileSystems:
LogException(f'File system `{parts[0]}` in path `{rawPath}` not supported.')
else:
self.__fileSystem = parts[0]
path = self.Trim(parts[1])
else:
self.__fileSystem = self.GetDefaultFileSystem()
path = rawPath
self.__path = self.Trim(path)
self.Absolute()
self.InferFormat()
@classmethod
def Trim(cls, path, char='/'):
while True:
if path.startswith(char*2):
path = path[1:]
else:
break
return path
def Absolute(self):
fs = self.fileSystem
if fs not in ['file']:
LogException(f'File system `{fs}` is not supported')
elif fs == 'file':
self.__path = os.path.abspath(self.__path)
def InferFormat(self):
for ext in extensionMapper:
if self.local.endswith(ext):
self.__format, self.__compression = extensionMapper[ext]
break
def Exist(self):
fs = self.fileSystem
if fs not in ['file', 'hdfs']:
LogException(f'File system `{fs}` is not supported')
elif fs == 'file':
return os.path.exists(self.local)
elif fs == 'local':
return not subprocess.run(['hdfs', 'dfs', '-test', '-e', self.path]).returncode
| 4,858 | 1,481 |
# -*- coding: utf-8 -*-
from __future__ import absolute_import, unicode_literals, print_function
from markdown.treeprocessors import Treeprocessor
from markdown.extensions import Extension
class MediaTreeprocessor(Treeprocessor):
def __init__(self, media_url=None, **kwargs):
self.media_url = media_url
super(MediaTreeprocessor, self).__init__(**kwargs)
def run(self, root):
image_tags = root.findall('.//img')
if self.media_url is not None:
for image_tag in image_tags:
tag_src = image_tag.get('src')
if not tag_src.lower().startswith('http') and not tag_src.startswith('//'):
if tag_src.startswith('./'):
tag_src = tag_src[2:]
# TODO: relative image tag source starting with . like sequence
# diagrams?
# Make sure we don't create a url like http://example.org//something.html
# if media_url ends with / and tag_src starts with /
# example.com/ + /blah.html = example.com/blah.html
# example.com + /blah.html = example.com/blah.html
# example.com/ + blah.html = example.com/blah.html
# example.com + blah.html = example.com/blah.html
# example.com + ./blah.html = example.com/blah.html
image_tag.set('src', self.media_url.rstrip('/') + '/' + tag_src.lstrip('/'))
class MediaExtension(Extension):
def __init__(self, **kwargs):
self.config = {
'media_url': ['.', 'Path or URL base for the media'],
}
super(MediaExtension, self).__init__(**kwargs)
def extendMarkdown(self, md, md_globals):
""" Add MediaTreeprocessor to the Markdown instance. """
md.registerExtension(self)
media_url = self.getConfig('media_url')
md.treeprocessors.add('media', MediaTreeprocessor(media_url=media_url, markdown_instance=md), '>inline')
def makeExtension(*args, **kwargs):
return MediaExtension(*args, **kwargs)
| 2,115 | 600 |
import torch
from torch.autograd import Variable
from scatwave import Scattering1D
import matplotlib.pyplot as plt
import numpy as np
from sklearn.utils import check_random_state
def generate_harmonic_signal(T, num_intervals=4, gamma=0.9, random_state=42):
"""
Generates a harmonic signal, which is made of piecewise constant notes
(of random fundamental frequency), with half overlap
"""
rng = check_random_state(random_state)
num_notes = 2 * (num_intervals - 1) + 1
support = T // num_intervals
half_support = support // 2
base_freq = 0.1 * rng.rand(num_notes) + 0.05
phase = 2 * np.pi * rng.rand(num_notes)
window = np.hanning(support)
x = np.zeros(T, dtype='float32')
t = np.arange(0, support)
u = 2 * np.pi * t
for i in range(num_notes):
ind_start = i * half_support
note = np.zeros(support)
for k in range(1):
note += (np.power(gamma, k) *
np.cos(u * (k + 1) * base_freq[i] + phase[i]))
x[ind_start:ind_start + support] += note * window
# put x in a Variable
x = Variable(torch.from_numpy(x[np.newaxis, np.newaxis]))
return x
def show_signal(x, s, order0, order1, order2):
fig, axarr = plt.subplots(4, 1, figsize=(8, 16))
axarr[0].plot(x.data[0, 0])
axarr[0].set_title('Original signal')
axarr[1].plot(s[order0][0])
axarr[1].set_title('Scattering Order 0')
axarr[2].imshow(s[order1], aspect='auto')
axarr[2].set_title('Scattering Order 1')
axarr[3].imshow(s[order2], aspect='auto')
axarr[3].set_title('Scattering Order 2')
plt.show()
if __name__ == '__main__':
# Scattering definition
T = 2**13
J = 6
Q = 16
scattering = Scattering1D(T, J, Q)
# get the metadata on the coordinates of the scattering
coords = Scattering1D.compute_meta_scattering(J, Q, order2=True)
order0 = torch.LongTensor([0])
order1 = torch.LongTensor(
sorted([cc for cc in coords.keys() if coords[cc]['order'] == '1']))
order2 = torch.LongTensor(
sorted([cc for cc in coords.keys() if coords[cc]['order'] == '2']))
# harmonic signal
x = generate_harmonic_signal(T)
s = scattering.forward(x).data[0]
show_signal(x, s, order0, order1, order2)
| 2,275 | 851 |
from pyrogram import Client, filters
import os, shutil
from creds import my
from telegraph import upload_file
import logging
logging.basicConfig(level=logging.INFO)
TGraph = Client(
"Image upload bot",
bot_token = my.BOT_TOKEN,
api_id = my.API_ID,
api_hash = my.API_HASH
)
@TGraph.on_message(filters.command("start"))
async def start(client, message):
await message.reply_text(f"<b>Hello {message.from_user.first_name}, My Name Is MeG Telegraph Bot 🥳\n\nI'm A <u>Telegraph Uploader Bot.</u>\n\nSend Me Any <u>Image</u>& I'll Upload It To Telegra.ph & Send You Back A Link\n\n🙂 Join & Support Us Via 👉 @MeGLeech.\n\n 🌟 Powered By @MeGBots</b>", True)
@TGraph.on_message(filters.command("help"))
async def help(client, message):
await message.reply_text(f"<b> 💁 Hey Its Not Tough To Ise Me...!!!\n\n Just Follow These Steps\n\n ▪️ Send Me Any Image (or) GIF (or) MP4 Below 5MB \n ▪️ Wait For To Generate Link For U\n\n 🌟 Powered By @MeGBots || @MeGLeech</b>", True)
@TGraph.on_message(filters.photo)
async def getimage(client, message):
tmp = os.path.join("downloads",str(message.chat.id))
if not os.path.isdir(tmp):
os.makedirs(tmp)
imgdir = tmp + "/" + str(message.message_id) +".jpg"
dwn = await message.reply_text("Downloading Please Wait...🤗", True)
await client.download_media(
message=message,
file_name=imgdir
)
await dwn.edit_text("Starting Upload...🤗")
try:
response = upload_file(imgdir)
except Exception as error:
await dwn.edit_text(f"Oops something went wrong\n{error}")
return
await dwn.edit_text(f"https://telegra.ph{response[0]}")
shutil.rmtree(tmp,ignore_errors=True)
TGraph.run()
| 1,823 | 692 |
import os
from typing import Any, Dict, Optional
import googlemaps
import structlog
from geopy.point import Point
logger = structlog.get_logger()
DEFAULT_COUNTRY = "CA"
GOOGLE_API_KEY_ENV = "GOOGLE_GEOCODING_API_KEY"
GEOMETRY = "geometry"
LOCATION = "location"
LATITUDE = "lat"
LONGITUDE = "lng"
class Geocoding:
def __init__(self):
key = os.environ[GOOGLE_API_KEY_ENV]
self.client = googlemaps.Client(key=key)
def get_from_address(self, address: str) -> Optional[Point]:
request = {"address": address}
return self._get_geocode(request)
def get_from_postal_code(self, postal_code: str) -> Optional[Point]:
request = {
"components": {"postal_code": postal_code, "country": DEFAULT_COUNTRY}
}
return self._get_geocode(request)
def _get_geocode(self, request: Dict[str, Any]) -> Optional[Point]:
geocode_result = self.client.geocode(**request)
if (len(geocode_result)) == 0:
return None
location = geocode_result[0].get(GEOMETRY, {}).get(LOCATION, {})
if LATITUDE not in location or LONGITUDE not in location:
return None
return Point([location[LATITUDE], location[LONGITUDE]])
| 1,237 | 427 |
from functools import partial
from tkinter import *
#program by~ Welberthy Gustavo Developer
def calc(btn):
if btn['text'].isdigit() or btn['text'] == '.':
lbl['text'] += btn['text']
def soma():
global sinal
sinal = 'soma'
global valor1
valor1 = lbl['text']
lbl['text'] = ''
def sub():
global sinal
sinal = 'sub'
global valor1
valor1 = lbl['text']
lbl['text'] = ''
def mult():
global sinal
sinal = 'mult'
global valor1
valor1 = lbl['text']
lbl['text'] = ''
def div():
global sinal
sinal = 'div'
global valor1
valor1 = lbl['text']
lbl['text'] = ''
def raiz():
global sinal
sinal = 'raiz'
global valor1
valor1 = lbl['text']
lbl['text'] = '√'
def elev():
global sinal
sinal = 'elev'
global valor1
valor1 = lbl['text']
lbl['text'] = ''
def porc():
global sinal
sinal = 'porc'
global valor1
valor1 = lbl['text']
lbl['text'] = '%'
def ac():
lbl['text'] = ''
def igual():
if sinal == 'soma':
valor2 = lbl['text']
lbl['text'] = ''
soma = float(valor1) + float(valor2)
lbl['text'] = float(soma)
elif sinal == 'sub':
valor2 = lbl['text']
lbl['text'] = ''
subt = float(valor1) - float(valor2)
lbl['text'] = float(subt)
elif sinal == 'mult':
valor2 = lbl['text']
lbl['text'] = ''
multi = float(valor1) * float(valor2)
lbl['text'] = float(multi)
elif sinal == 'div':
valor2 = lbl['text']
lbl['text'] = ''
soma = float(valor1) / float(valor2)
lbl['text'] = float(soma)
elif sinal == 'raiz':
lbl['text'] = ''
rai = float(valor1) ** 0.5
lbl['text'] = float(rai)
elif sinal == 'elev':
valor2 = lbl['text']
lbl['text'] = ''
eleva = float(valor1) ** float(valor2)
lbl['text'] = float(eleva)
elif sinal == 'porc':
lbl['text'] = ''
porcen = float(valor1) / 100
lbl['text'] = float(porcen)
else:
lbl['text'] = 'Error!'
janela = Tk()
janela.title('Calculadora')
janela.iconbitmap('calculadoraProject/cal.ico')
janela['bg'] = 'gainsboro'
janela.geometry('400x450+400+100')
janela.resizable(0,0)
lbl = Label(janela,width=15, height=1, font='Arial 30', bd=1, relief='solid', justify=RIGHT, anchor=E, padx=15, pady=10)
lbl.place(x=100,y=100)
lbl.pack(side=TOP)
#Others buttons
btnab = Button(janela,width=8, height=2, font='Arial 11 bold', text='√', bg='gray80', command=raiz)
btnab.place(x=15,y=90)
btnfe = Button(janela,width=8, height=2, font='Arial 11 bold', text='x¹', bg='gray80', command=elev)
btnfe.place(x=110,y=90)
btnpor = Button(janela,width=8, height=2, font='Arial 11 bold', text='%', bg='gray80', command=porc)
btnpor.place(x=205,y=90)
btnac = Button(janela,width=8, height=2, font='Arial 11 bold', text='AC', bg='gray80', command=ac)
btnac.place(x=300,y=90)
#Numbers buttons
btn7 = Button(janela,width=8, height=2, font='Arial 12', text='7')
btn7['command'] = partial(calc, btn7)
btn7.place(x=15,y=160)
btn8 = Button(janela,width=8, height=2, font='Arial 12', text='8')
btn8['command'] = partial(calc, btn8)
btn8.place(x=110,y=160)
btn9 = Button(janela,width=8, height=2, font='Arial 12', text='9')
btn9['command'] = partial(calc, btn9)
btn9.place(x=205,y=160)
btn4 = Button(janela,width=8, height=2, font='Arial 12', text='4')
btn4['command'] = partial(calc, btn4)
btn4.place(x=15,y=230)
btn5 = Button(janela,width=8, height=2, font='Arial 12', text='5')
btn5['command'] = partial(calc, btn5)
btn5.place(x=110,y=230)
btn6 = Button(janela,width=8, height=2, font='Arial 12', text='6')
btn6['command'] = partial(calc, btn6)
btn6.place(x=205,y=230)
btn3 = Button(janela,width=8, height=2, font='Arial 12', text='3')
btn3['command'] = partial(calc, btn3)
btn3.place(x=15,y=300)
btn2 = Button(janela,width=8, height=2, font='Arial 12', text='2')
btn2['command'] = partial(calc, btn2)
btn2.place(x=110,y=300)
btn1 = Button(janela,width=8, height=2, font='Arial 12', text='1')
btn1['command'] = partial(calc, btn1)
btn1.place(x=205,y=300)
btn0 = Button(janela,width=8, height=2, font='Arial 12', text='0')
btn0['command'] = partial(calc, btn0)
btn0.place(x=15,y=370)
#Score button
btnp = Button(janela,width=8, height=2, font='Arial 11 bold', text='.')
btnp['command'] = partial(calc, btnp)
btnp.place(x=110,y=370)
#Equals button
btnig = Button(janela,width=8, height=2, font='Arial 11 bold', text='=', bg='blue2', fg='white', command=igual)
btnig.place(x=205,y=370)
#Operators button
btndiv = Button(janela,width=8, height=2, font='Arial 11 bold', text='÷', bg='gray80', command=div)
btndiv.place(x=300,y=160)
btnmul = Button(janela,width=8, height=2, font='Arial 11 bold', text='x',bg='gray80', command=mult)
btnmul.place(x=300,y=230)
btnsub = Button(janela,width=8, height=2, font='Arial 11 bold', text='-',bg='gray80', command=sub)
btnsub.place(x=300,y=300)
btnad = Button(janela,width=8, height=2, font='Arial 11 bold', text='+', bg='gray80', command=soma)
btnad.place(x=300,y=370)
janela.mainloop() | 5,319 | 2,323 |
# Copyright 2015 Rackspace US, Inc.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# pylint: disable=invalid-name
"""Fastfood Exceptions."""
import re
# python 2 vs. 3 string types
try:
basestring
except NameError:
basestring = str
_SPLITCASE_RE = re.compile(r'[A-Z][^A-Z]*')
class FastfoodError(Exception):
"""Base class for all exceptions raised by fastfood."""
class FastfoodStencilSetNotListed(FastfoodError):
"""Stencil set specified was not listed in the templatepack manifest."""
class FastfoodStencilSetInvalidPath(FastfoodError):
"""Specified path to stencil set does not exist."""
class FastfoodStencilSetMissingManifest(FastfoodError):
"""Stencil set is missing a manifest.json file."""
class FastfoodTemplatePackAttributeError(AttributeError, FastfoodError):
"""Invalid stencilset request from TemplatePack."""
def get_friendly_title(err):
"""Turn class, instance, or name (str) into an eyeball-friendly title.
E.g. FastfoodStencilSetNotListed --> 'Stencil Set Not Listed'
"""
if isinstance(err, basestring):
string = err
else:
try:
string = err.__name__
except AttributeError:
string = err.__class__.__name__
split = _SPLITCASE_RE.findall(string)
if not split:
split.append(string)
if len(split) > 1 and split[0] == 'Fastfood':
split.pop(0)
return " ".join(split)
| 1,928 | 626 |
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
from nb_comb import *
from sklearn.naive_bayes import *
from sklearn.tree import *
from sklearn.neural_network import *
from sklearn.model_selection import *
import pandas as pd
data = pd.read_csv('dataset.csv', index_col=0)
X, Y = data.iloc[:, :-1], data.iloc[:, -1].values
for i, y in enumerate(Y):
if y>600:
Y[i]=2
elif y>500:
Y[i]=1
else:
Y[i]=0
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.3)
import numpy as np
keys = data.columns
key1=['用户实名制是否通过核实', '是否大学生客户', '是否黑名单客户', '是否4G不健康客户', '缴费用户当前是否欠费缴费',
'是否经常逛商场的人', '当月是否逛过福州仓山万达', '当月是否到过福州山姆会员店', '当月是否看电影', '当月是否景点游览', '当月是否体育场馆消费']
key2 = ['用户年龄', '用户话费敏感度', '用户当月账户余额(元)', '近三个月月均商场出现次数',
'当月物流快递类应用使用次数', '当月飞机类应用使用次数', '当月火车类应用使用次数', '当月旅游资讯类应用使用次数',
'用户网龄(月)', '用户最近一次缴费距今时长(月)', '当月通话交往圈人数']
key3 = ['缴费用户最近一次缴费金额(元)', '用户近6个月平均消费值(元)', '用户账单当月总费用(元)',
'当月网购类应用使用次数', '当月金融理财类应用使用总次数', '当月视频播放类应用使用次数']
import time
estimators = [('bernoulli', BernoulliNB()), ('multinomial', MultinomialNB()), ('gauss', GaussianNB())]
nba1 = NBAdditive(estimators=estimators)
estimators = [('bernoulli', BernoulliNB()), ('tree', DecisionTreeClassifier()), ('gauss', GaussianNB())]
nba2 = NBAdditive(estimators=estimators)
estimators = [('bernoulli', BernoulliNB()), ('tree', DecisionTreeClassifier()), ('mlp', MLPClassifier(hidden_layer_sizes=(5,), max_iter=2000))]
nba3 = NBAdditive(estimators=estimators)
models = [('NB组合0(NB)', nba1), ('NB组合1(非NB)', nba2), ('NB组合2(非NB)', nba3),
('高斯NB', GaussianNB()), ('多项式NB', MultinomialNB()), ('决策树', DecisionTreeClassifier()), ('神经网络', MLPClassifier(hidden_layer_sizes=(8,), max_iter=2000))]
np.random.seed(1001)
perf = []
for name, model in models:
dts = []
for _ in range(2):
time1 = time.perf_counter()
if name.startswith('NB'):
model.fit(X_train, Y_train, inds=[key1, key2, key3])
else:
model.fit(X_train, Y_train)
time2 = time.perf_counter()
dt = time2 - time1
dts.append(dt)
perf.append([name, model.score(X_train, Y_train), model.score(X_test, Y_test), np.mean(dts)])
p = pd.DataFrame(data=perf, columns=('name', 'train-score', 'test-score', 'time'))
print(p)
| 2,320 | 1,227 |
name='avi'
print(name)
#name[0]='k' here this results in a error because strings are immutable
last_letters=name[1:]
print(last_letters)
print('k'+last_letters) #here is the string concateation
#here we replaced 'a' with 'k'
lname='koshal'
print("the concated string=",'Avinash'+ lname)
a= 'b';
print(a*10) # this will print the letter b 10 times
| 397 | 133 |
def func(x, y):
pass
| 25 | 12 |
# Copyright 2020 University of Illinois Board of Trustees. All Rights Reserved.
# Author: Beomyeol Jeon, DPRG (https://dprg.cs.uiuc.edu)
# This file is part of Baechi, which is released under specific terms. See file License.txt file for full license details.
# ==============================================================================
"""Unit tests for placer_lib module."""
# pylint: disable=missing-function-docstring
import unittest
import networkx as nx
from placer import placer_lib
class FusedOpPlacerTest(unittest.TestCase):
"""FusedOpPlacer test."""
def test_generate_fused_op_graph1(self):
op_graph = nx.DiGraph()
# op0 -> op1 -> op2
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 2,
'persistent_memory': 0, 'colocation_group': 'group0',
'output_memory': [2]}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 4,
'persistent_memory': 0, 'colocation_group': 'group0',
'output_memory': []}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_edge(
0, 1, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
1, 2, id=1, weight=2,
tensor=[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
# all ops are fused
self.assertEqual(fused_op_graph.number_of_nodes(), 1)
fused_op = fused_op_graph.nodes[0]
self.assertEqual(fused_op['weight'], 8)
self.assertEqual(fused_op['temporary_memory'], 4)
self.assertEqual(fused_op['persistent_memory'], 5)
self.assertEqual(len(fused_op['output_memory']), 0)
fused_op_names = [op['name'] for op in fused_op['fused_ops']]
self.assertIn('op1', fused_op_names)
self.assertIn('op2', fused_op_names)
def test_generate_fused_op_graph2(self):
op_graph = nx.DiGraph()
# op0 -> op1 -> op2
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 2,
'persistent_memory': 0, 'colocation_group': 'group0',
'output_memory': [2]}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 4,
'persistent_memory': 0, 'colocation_group': 'group1',
'output_memory': []}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_edge(
0, 1, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
1, 2, id=1, weight=2,
tensor=[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
# op0 and op1 are fused
self.assertEqual(fused_op_graph.number_of_nodes(), 2)
fused_op0 = fused_op_graph.nodes[0]
self.assertEqual(fused_op0['weight'], 5)
self.assertEqual(fused_op0['temporary_memory'], 2)
self.assertEqual(fused_op0['persistent_memory'], 5)
self.assertListEqual(fused_op0['output_memory'], [2])
fused_op1 = fused_op_graph.nodes[1]
self.assertEqual(fused_op1['weight'], 3)
self.assertEqual(fused_op1['temporary_memory'], 4)
self.assertEqual(fused_op1['persistent_memory'], 0)
self.assertListEqual(fused_op1['output_memory'], [])
self.assertEqual(fused_op_graph.number_of_edges(), 1)
edge = fused_op_graph[0][1]
self.assertEqual(edge['id'], 0)
self.assertEqual(edge['weight'], 2)
self.assertListEqual(edge['tensor'],
[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
def test_generate_fused_op_graph3(self):
op_graph = nx.DiGraph()
# op0 -> op2
# op1 /
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 0,
'persistent_memory': 3, 'colocation_group': 'group1',
'output_memory': [2]}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 4,
'persistent_memory': 2, 'colocation_group': 'group0',
'output_memory': []}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_edge(
0, 2, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
1, 2, id=1, weight=2,
tensor=[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
# op0 and op2 are fused.
self.assertEqual(fused_op_graph.number_of_nodes(), 2)
fused_op0 = fused_op_graph.nodes[0]
self.assertEqual(fused_op0['weight'], 5)
self.assertEqual(fused_op0['temporary_memory'], 4)
self.assertEqual(fused_op0['persistent_memory'], 7)
self.assertListEqual(fused_op0['output_memory'], [])
fused_op1 = fused_op_graph.nodes[1]
self.assertEqual(fused_op1['weight'], 3)
self.assertEqual(fused_op1['temporary_memory'], 0)
self.assertEqual(fused_op1['persistent_memory'], 3)
self.assertListEqual(fused_op1['output_memory'], [2])
self.assertEqual(fused_op_graph.number_of_edges(), 1)
edge = fused_op_graph[1][0]
self.assertEqual(edge['id'], 0)
self.assertEqual(edge['weight'], 2)
self.assertListEqual(edge['tensor'],
[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
def test_generate_fused_op_graph4(self):
op_graph = nx.DiGraph()
# op0 -> op2
# op1 /
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 0,
'persistent_memory': 3, 'colocation_group': 'group0',
'output_memory': [2]}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 4,
'persistent_memory': 2, 'colocation_group': 'group0',
'output_memory': []}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_edge(
0, 2, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
1, 2, id=1, weight=2,
tensor=[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
# all ops are fused
self.assertEqual(fused_op_graph.number_of_nodes(), 1)
fused_op = fused_op_graph.nodes[0]
self.assertEqual(fused_op['weight'], 8)
self.assertEqual(fused_op['temporary_memory'], 4)
self.assertEqual(fused_op['persistent_memory'], 10)
self.assertListEqual(fused_op['output_memory'], [])
def test_generate_fused_op_graph5(self):
op_graph = nx.DiGraph()
# op0 -> op2 -> op3 -> op4
# op1 /
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 0,
'persistent_memory': 3, 'colocation_group': 'group0',
'output_memory': [2]}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 4,
'persistent_memory': 0, 'colocation_group': 'group1',
'output_memory': [3]}
op3 = {'id': 3, 'name': 'op4', 'weight': 1, 'temporary_memory': 7,
'persistent_memory': 3, 'colocation_group': 'group1',
'output_memory': [4]}
op4 = {'id': 4, 'name': 'op4', 'weight': 5, 'temporary_memory': 2,
'persistent_memory': 0, 'colocation_group': 'group0',
'output_memory': [0]}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_node(3, **op3)
op_graph.add_node(4, **op4)
op_graph.add_edge(
0, 2, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
1, 2, id=1, weight=2,
tensor=[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
op_graph.add_edge(
2, 3, id=2, weight=3,
tensor=[{'name': 'op2:0', 'weight': 3, 'num_bytes': 3}])
op_graph.add_edge(
3, 4, id=3, weight=4,
tensor=[{'name': 'op3:0', 'weight': 4, 'num_bytes': 4}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
# op2 and op3 are fused.
self.assertEqual(fused_op_graph.number_of_nodes(), 4)
self.assertEqual(fused_op_graph.nodes[0], {**op0, 'old_id': 0})
self.assertEqual(fused_op_graph.nodes[1], {**op1, 'old_id': 1})
fused_op = fused_op_graph.nodes[2]
self.assertEqual(fused_op['weight'], 4)
self.assertEqual(fused_op['temporary_memory'], 7)
self.assertEqual(fused_op['persistent_memory'], 3)
self.assertListEqual(fused_op['output_memory'], [4])
expected_dict = {**op4, 'old_id': 4}
expected_dict['id'] = 3
self.assertEqual(fused_op_graph.nodes[3], expected_dict)
self.assertEqual(fused_op_graph.number_of_edges(), 3)
edge_ids = set()
edge_ids.add(fused_op_graph[0][2]['id'])
self.assertEqual(fused_op_graph[0][2]['weight'], 1)
self.assertListEqual(
fused_op_graph[0][2]['tensor'],
[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
edge_ids.add(fused_op_graph[1][2]['id'])
self.assertEqual(fused_op_graph[1][2]['weight'], 2)
self.assertListEqual(
fused_op_graph[1][2]['tensor'],
[{'name': 'op1:0', 'weight': 2, 'num_bytes': 2}])
edge_ids.add(fused_op_graph[2][3]['id'])
self.assertEqual(fused_op_graph[2][3]['weight'], 4)
self.assertListEqual(
fused_op_graph[2][3]['tensor'],
[{'name': 'op3:0', 'weight': 4, 'num_bytes': 4}])
self.assertSetEqual(edge_ids, set(list(range(3))))
def test_generate_fused_op_graph6(self):
op_graph = nx.DiGraph()
# -> op2 -> op3
# / /
# op0 -> op1 ->/
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1, 2]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 5,
'persistent_memory': 3, 'colocation_group': 'group0',
'output_memory': [3]}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 2,
'persistent_memory': 2, 'colocation_group': 'group1',
'output_memory': [4]}
op3 = {'id': 3, 'name': 'op4', 'weight': 1, 'temporary_memory': 7,
'persistent_memory': 0, 'colocation_group': 'group1',
'output_memory': []}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_node(3, **op3)
op_graph.add_edge(
0, 1, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
0, 2, id=1, weight=2,
tensor=[{'name': 'op0:1', 'weight': 2, 'num_bytes': 2}])
op_graph.add_edge(
1, 3, id=2, weight=3,
tensor=[{'name': 'op1:0', 'weight': 3, 'num_bytes': 3}])
op_graph.add_edge(
2, 3, id=3, weight=4,
tensor=[{'name': 'op2:0', 'weight': 4, 'num_bytes': 4}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
self.assertEqual(fused_op_graph.number_of_nodes(), 2)
fused_op0 = fused_op_graph.nodes[0] # op0, op1
self.assertEqual(fused_op0['weight'], 5)
self.assertEqual(fused_op0['temporary_memory'], 5)
self.assertEqual(fused_op0['persistent_memory'], 8)
self.assertEqual(sum(fused_op0['output_memory']), 5)
fused_op1 = fused_op_graph.nodes[1] # op2, op3
self.assertEqual(fused_op1['weight'], 4)
self.assertEqual(fused_op1['temporary_memory'], 7)
self.assertEqual(fused_op1['persistent_memory'], 2)
self.assertListEqual(fused_op1['output_memory'], [])
self.assertEqual(fused_op_graph.number_of_edges(), 1)
fused_edge = fused_op_graph[0][1]
self.assertEqual(fused_edge['id'], 0)
self.assertEqual(fused_edge['weight'], 5)
self.assertListEqual(
fused_edge['tensor'],
[{'name': 'op0:1', 'weight': 2, 'num_bytes': 2},
{'name': 'op1:0', 'weight': 3, 'num_bytes': 3}])
def test_generate_fused_op_graph7(self):
op_graph = nx.DiGraph()
# -> op2 -> op3
# / /
# op0 -> op1 <-/
op0 = {'id': 0, 'name': 'op0', 'weight': 2, 'temporary_memory': 0,
'persistent_memory': 5, 'colocation_group': 'group0',
'output_memory': [1, 2]}
op1 = {'id': 1, 'name': 'op1', 'weight': 3, 'temporary_memory': 5,
'persistent_memory': 3, 'colocation_group': 'group0',
'output_memory': []}
op2 = {'id': 2, 'name': 'op2', 'weight': 3, 'temporary_memory': 2,
'persistent_memory': 2, 'colocation_group': 'group1',
'output_memory': [4]}
op3 = {'id': 3, 'name': 'op4', 'weight': 1, 'temporary_memory': 7,
'persistent_memory': 0, 'colocation_group': 'group1',
'output_memory': [3]}
op_graph.add_node(0, **op0)
op_graph.add_node(1, **op1)
op_graph.add_node(2, **op2)
op_graph.add_node(3, **op3)
op_graph.add_edge(
0, 1, id=0, weight=1,
tensor=[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
op_graph.add_edge(
0, 2, id=1, weight=2,
tensor=[{'name': 'op0:1', 'weight': 2, 'num_bytes': 2}])
op_graph.add_edge(
3, 1, id=2, weight=3,
tensor=[{'name': 'op3:0', 'weight': 3, 'num_bytes': 3}])
op_graph.add_edge(
2, 3, id=3, weight=4,
tensor=[{'name': 'op2:0', 'weight': 4, 'num_bytes': 4}])
# pylint: disable=protected-access
fused_op_graph = placer_lib.FusedOpPlacer._generate_fused_op_graph(
op_graph, False)
self.assertEqual(fused_op_graph.number_of_nodes(), 3)
self.assertEqual(fused_op_graph.nodes[0], {**op0, 'old_id': 0})
self.assertEqual(fused_op_graph.nodes[1], {**op1, 'old_id': 1})
fused_op = fused_op_graph.nodes[2] # op2, op3
self.assertEqual(fused_op['weight'], 4)
self.assertEqual(fused_op['temporary_memory'], 7)
self.assertEqual(fused_op['persistent_memory'], 2)
self.assertEqual(sum(fused_op['output_memory']), 3)
self.assertEqual(fused_op_graph.number_of_edges(), 3)
self.assertEqual(fused_op_graph[0][1]['weight'], 1)
self.assertListEqual(
fused_op_graph[0][1]['tensor'],
[{'name': 'op0:0', 'weight': 1, 'num_bytes': 1}])
self.assertEqual(fused_op_graph[0][2]['weight'], 2)
self.assertListEqual(
fused_op_graph[0][2]['tensor'],
[{'name': 'op0:1', 'weight': 2, 'num_bytes': 2}])
self.assertEqual(fused_op_graph[2][1]['weight'], 3)
self.assertListEqual(
fused_op_graph[2][1]['tensor'],
[{'name': 'op3:0', 'weight': 3, 'num_bytes': 3}])
self.assertSetEqual(
{edge[-1] for edge in fused_op_graph.edges(data='id')},
set(range(3)))
if __name__ == "__main__":
unittest.main()
| 17,165 | 6,415 |
import os
import uuid
from io import BytesIO
from tempfile import NamedTemporaryFile
from zipfile import ZipFile
from django.test import TestCase
from corehq.apps.hqmedia.models import CommCareAudio, CommCareVideo, CommCareImage
from corehq.blobs import CODES, get_blob_db
from corehq.blobs.export import EXPORTERS
from corehq.blobs.tests.util import TemporaryFilesystemBlobDB, new_meta
class TestBlobExport(TestCase):
@classmethod
def setUpClass(cls):
super().setUpClass()
cls.db = TemporaryFilesystemBlobDB()
assert get_blob_db() is cls.db, (get_blob_db(), cls.db)
data = b'binary data not valid utf-8 \xe4\x94'
cls.blob_metas = []
cls.not_found = set()
cls.domain_name = str(uuid.uuid4)
for type_code in [CODES.form_xml, CODES.multimedia, CODES.data_export]:
for domain in (cls.domain_name, str(uuid.uuid4())):
meta = cls.db.put(BytesIO(data), meta=new_meta(domain=domain, type_code=type_code))
lost = new_meta(domain=domain, type_code=type_code, content_length=42)
cls.blob_metas.append(meta)
cls.blob_metas.append(lost)
lost.save()
cls.not_found.add(lost.key)
@classmethod
def tearDownClass(cls):
for blob in cls.blob_metas:
blob.delete()
cls.db.close()
super().tearDownClass()
def test_migrate_all(self):
expected = {
m.key for m in self.blob_metas
if m.domain == self.domain_name and m.key not in self.not_found
}
with NamedTemporaryFile() as out:
exporter = EXPORTERS['all_blobs'](self.domain_name)
exporter.migrate(out.name, force=True)
with ZipFile(out.name, 'r') as zip:
self.assertEqual(expected, set(zip.namelist()))
def test_migrate_multimedia(self):
image_path = os.path.join('corehq', 'apps', 'hqwebapp', 'static', 'hqwebapp', 'images',
'commcare-hq-logo.png')
with open(image_path, 'rb') as f:
image_data = f.read()
files = (
(CommCareImage, self.domain_name, image_data),
(CommCareAudio, self.domain_name, b'fake audio'),
(CommCareVideo, self.domain_name, b'fake video'),
(CommCareAudio, 'other_domain', b'fake audio 1'),
)
blob_keys = []
for doc_class, domain, data in files:
obj = doc_class.get_by_data(data)
obj.attach_data(data)
obj.add_domain(domain)
self.addCleanup(obj.delete)
self.assertEqual(data, obj.get_display_file(False))
blob_keys.append(obj.blobs[obj.attachment_id].key)
expected = set(blob_keys[:-1])
with NamedTemporaryFile() as out:
exporter = EXPORTERS['multimedia'](self.domain_name)
exporter.migrate(out.name, force=True)
with ZipFile(out.name, 'r') as zip:
self.assertEqual(expected, set(zip.namelist()))
| 3,073 | 1,015 |
# -*- coding: utf-8 -*-
from __future__ import print_function
from __future__ import absolute_import
__author__ = 'Matthew L. Bendall'
__copyright__ = "Copyright (C) 2019 Matthew L. Bendall"
def get_annotation_class(annotation_class_name):
""" Get Annotation class matching provided name
Args:
annotation_class_name (str): Name of annotation class.
Returns:
Annotation class with data structure and function(s) for finding
overlaps
"""
if annotation_class_name == 'htseq':
raise NotImplementedError('"htseq" is not compatible.')
# from ._annotation_htseq import _AnnotationHTSeq
# return _AnnotationHTSeq
elif annotation_class_name == 'intervaltree':
from ._annotation_intervaltree import _AnnotationIntervalTree
return _AnnotationIntervalTree
else:
raise NotImplementedError('Choices are "htseq" or "intervaltree".')
| 922 | 261 |
from backend.models import *
from django.views.generic import TemplateView
from django.shortcuts import render, redirect
import os
import hass_api.rest as hass_rest
from frontend.util import get_server, refresh_hass_token, \
get_device_names, get_activity_names, get_person_hass_names, \
get_person_names, input_is_empty
import frontend.experiment as experiment
LOCAL_URL_PROVIDED = 'server_local_url_provided'
INVALID_ADDRESS_PROVIDED = 'server_invalid_address_provided'
class ConfigView(TemplateView):
def get_context(self, add_to_context):
srv = get_server()
person_list = Person.objects.all()
act_list = Activity.objects.all()
url = 'config'
exp_active = experiment.is_active()
refresh_hass_token()
# get hass devices
hass_devices = hass_rest.get_device_list(
settings.HASS_API_URL , srv.hass_api_token)
dev_list = get_device_names()
hass_devices = list(set(hass_devices).difference(set(dev_list)))
# get hass users
hass_users = hass_rest.get_user_names(
settings.HASS_API_URL, srv.hass_api_token,)
hass_users = list(set(hass_users).difference(set(get_person_hass_names())))
context = {'server': srv,
'url': url,
'person_list':person_list,
'hass_dev_list' : hass_devices,
'aa_dev_list' : dev_list,
'activity_list' : act_list,
'hass_user_list' : hass_users,
'aa_user_list' : person_list,
'poll_int_list' : settings.POLL_INTERVAL_LST,
'experiment_active':exp_active,
}
context.update(add_to_context)
return context
def get(self, request, *args, **kwargs):
context = self.get_context({})
return render(request, 'config.html', context)
def post(self, request):
from_section = request.POST.get("from", "")
add_to_context = {}
assert from_section in ["conf_devices", "conf_persons",\
"conf_activities", "conf_server", "debug"]
if from_section == 'conf_devices':
conf_devices(request)
elif from_section == 'conf_persons':
conf_persons(request)
elif from_section == 'conf_activities':
conf_activities(request)
elif from_section == 'conf_server':
success, reason = conf_server(request)
if not success and reason:
add_to_context[reason] = True
if not success and reason:
add_to_context[reason] = True
elif from_section == 'debug':
debug(request)
context = self.get_context(add_to_context)
return render(request, 'config.html', context)
def debug(request):
from frontend.util import collect_data_from_hass
collect_data_from_hass()
def conf_server(request):
""" sets server related stuff
"""
logger.error('test')
srv = get_server()
try:
pol_int = request.POST.get("poll_interval", "")
srv.poll_interval = pol_int
except:
pass
srv.save()
try:
address = request.POST.get("address", "")
if input_is_valid_address(address):
if input_is_local_address(address):
return False, LOCAL_URL_PROVIDED
address = url_strip_appendix(address)
srv.server_address = address
srv.save()
return (True, None)
else:
return False, INVALID_ADDRESS_PROVIDED
except:
return (True, None)
def url_strip_appendix(url):
""" removes trailing stuff behind a url definition
"""
lst = url.split('/')
return lst[0] + '//' + lst[2]
def input_is_valid_address(address):
""" checks whether the given address is either a valid ipv4 or a valid url
"""
from django.core.validators import URLValidator
try:
URLValidator()(address)
return True
except:
return False
def input_is_local_address(address):
return '.local' in address
def conf_devices(request):
intent = request.POST.get("intent","")
assert intent in ['track', 'remove']
dev_lst = request.POST.getlist('devices')
if intent == 'track':
lst = request.POST.getlist('hass_select')
if len(lst) == 1 and input_is_empty(lst[0]):
return
for name in lst:
Device(name=name).save()
else:
lst = request.POST.getlist('act_assist_select')
if len(lst) == 1 and input_is_empty(lst[0]):
return
for name in lst:
Device.objects.get(name=name).delete()
def conf_activities(request):
intent = request.POST.get("intent", "")
assert intent in ['add', 'delete']
if intent == 'delete':
for name in request.POST.getlist('act_select'):
Activity.objects.get(name=name).delete()
else:
name = request.POST.get("name", "")
if name not in get_activity_names() and not input_is_empty(name):
Activity(name=name).save()
def conf_persons(request):
intent = request.POST.get("intent","")
assert intent in ['track', 'remove', 'add']
dev_lst = request.POST.getlist('devices')
if intent == 'track':
lst = request.POST.getlist('hass_select')
if len(lst) == 1 and input_is_empty(lst[0]):
return
for hass_name in lst:
name = hass_name.split('.')[1]
Person(name=name, hass_name=hass_name).save()
elif intent == 'remove':
lst = request.POST.getlist('act_assist_select')
if len(lst) == 1 and input_is_empty(lst[0]):
return
for col in lst:
name = col.split(' ')[0]
Person.objects.get(name=name).delete()
else:
name = request.POST.get("name", "")
if name not in get_person_names() and not input_is_empty(name):
Person(name=name, hass_name='person.' + name).save()
| 6,012 | 1,836 |
import os
from toolbox import *
import pickle
import logging
import commandresolve
def console(data:dict,logger):
'''
Main console program
'''
consoleobj=commandresolve.commandresolve(data,logger)
flag=True# to mark if it is time to exit
while (flag):
rawcommand=input(">")
flag=consoleobj.resolvecommand(rawcommand)
#Exit now
logger.info("Exit Successfully")
return #NOTE data should be saved in exit
if __name__=="__main__":
#Mainloop
#Search for file
filename="simdisk.bin"
'''
Setup logger
'''
logger = logging.getLogger()#创建对象
logger.setLevel(logging.INFO)#设定起始显示级别
# 创建Handler
# 终端Handler
consoleHandler = logging.StreamHandler()
consoleHandler.setLevel(logging.INFO)
# Formatter
formatter = logging.Formatter('%(asctime)s [%(levelname)s] \t %(message)s')
consoleHandler.setFormatter(formatter)
# 添加到Logger中
logger.addHandler(consoleHandler)
if not os.path.isfile(filename):
logger.warning("File not exist. Trying to create...")
createfile(filename,20000000)
with open(filename,'wb') as p:
pickle.dump({},p)
#Build simlink
data=dict()
with open(filename,"rb") as f:
data=pickle.load(f)
if data !={}:
logger.info("Get existed file data, trying to resolve...")
if type(data)!=dict:
print(data)
logger.error("File structure is unable to resolve")
else:
logger.info("File structure is resolved successfully")
logger.info("Jumping to command line...")
console(data,logger)
else:
logger.info("File structure is resolved successfully")
logger.info("Jumping to command line...")
console(data,logger)
| 1,843 | 559 |
import pandas as pd
from glob import glob
from utils import evaluate_augmented_dataset, evaluate_landmarks, evaluate_scene_detection
import os
from winnow.utils.config import resolve_config
import click
import numpy as np
import json
pd.options.mode.chained_assignment = None
@click.command()
@click.option("--benchmark", "-b", help="name of the benchmark to evaluated", default="augmented_dataset")
@click.option(
"--force-download",
"-fd",
help="Force download of the dataset (even if an existing directory for the dataset has been detected",
default=False,
is_flag=True,
)
@click.option(
"--overwrite",
"-o",
help="Force feature extraction, even if we detect that signatures have already been processed.",
default=False,
is_flag=True,
)
def main(benchmark, force_download, overwrite):
config_path = os.path.join("benchmarks", benchmark, "config.yml")
config = resolve_config(config_path)
if benchmark == "augmented_dataset":
evaluate_augmented_dataset(config, force_download, overwrite, config_path)
elif benchmark == "landmarks":
evaluate_landmarks(config, force_download, overwrite, config_path)
elif benchmark == "scene_detection":
evaluate_scene_detection(config, force_download, overwrite, config_path)
else:
print(f"Please review the dataset (@ {config.sources.root})")
if __name__ == "__main__":
main()
| 1,428 | 425 |
# generate a*z_n = {ax : x in z_n} for any a in z_n
# brupoon
n = int(input("Z_n (integer n)? "))
a = int(input("a for a*Z_n (integer a)?"))
z_n = []
az_n = []
for i in range(n):
# standard set z_n
z_n.append(i)
for x in range(n):
value = (a*x) % n
# find ax in z_n
az_n.append(value)
#az_n = sorted(az_n)
print("Z_n: {0}".format(z_n))
print("a*Z_n: {0}".format(az_n))
| 393 | 195 |
from itertools import combinations
class LFSR:
def __init__(self, register, taps):
self.register = register
self.taps = taps
def next(self):
new = 0
ret = self.register[0]
for i in self.taps:
new ^= self.register[i]
self.register = self.register[1:] + [new]
return ret
register = list(map(int, ('{:08b}'.format(i ^ j) for i, j in zip(b'flag', flag_enc))))
print('register: ', register)
for i in combinations(list(range(16)), 5):
lfsr = LFSR(register[:16], list(i))
if all(bit == lfsr.next() for bit in register):
taps = list(i)
break
print('taps: ', taps)
lfsr = LFSR(register[:16], taps)
flag = []
for char in flag_enc:
dec_char = 0
for binary in '{:08b}'.format(char):
dec_char <<= 1
dec_char += int(binary) ^ lfsr.next()
flag.append(dec_char)
print(bytes(flag).decode())
| 909 | 331 |
# SVM with RBF Kernel and Feature Preprocessing
*Hayley Boyce, April 28th 2021*
# Importing our libraries
import pandas as pd
import altair as alt
import numpy as np
from sklearn.tree import DecisionTreeClassifier
from sklearn.dummy import DummyClassifier, DummyRegressor
from sklearn.neighbors import KNeighborsClassifier, KNeighborsRegressor
from sklearn.model_selection import cross_validate, train_test_split
import sys
sys.path.append('code/')
from display_tree import display_tree
from plot_classifier import plot_classifier
import matplotlib.pyplot as plt
# Preprocessing and pipeline
from sklearn.impute import SimpleImputer
from sklearn.metrics.pairwise import euclidean_distances
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder, OrdinalEncoder, StandardScaler, MinMaxScaler
## House Keeping
- Assignment due today at 11:59pm!
- Course feedback!
- Assignment - things I should know?
- Assignment2 - before or after the weekend?
- Polls coming Monday!
- I hear you don't like breakout rooms, let's try this lecture without them!
- Per the announcement Monday, download the data for this lecture [here](https://www.kaggle.com/harrywang/housing) and include it in your `data` folder that resides in `lectures`.
## Lecture Learning Objectives
- Identify when to implement feature transformations such as imputation and scaling.
- Describe the difference between normalizing and standardizing and be able to use scikit-learn's `MinMaxScaler()` and `StandardScaler()` to pre-process numeric features.
- Apply `sklearn.pipeline.Pipeline` to build a machine learning pipeline.
- Use `sklearn` for applying numerical feature transformations to the data.
- Discuss the golden rule in the context of feature transformations.
## Five Minute Recap/ Lightning Questions
- When using a Dummy Regressor what value does the model predict for unseen data?
- When using a Dummy Classifier (the one we examined in lecture) what class does the model predict for unseen data?
- What is the name of the distance metric used in the $k$-nn model we looked at?
- If a dataset has 14 features and 1 target column, how many dimensions will the feature vector be?
- What is the hyperparameter name of the $k$-nn classifier we looked at last lecture?
### Some lingering questions
- How does a $k$-nn Regressor work?
- Are we ready to do machine learning on real-world datasets?
- We've looked at data with numeric features but what do we do if we have features with categories or string values?
- What happens if we are missing data in our features?
- Is there a cleaner way to do all the steps we need to do?
## Regression with $k$-NN
In $k$-nearest neighbour regression, we take the average of $k$-nearest neighbours instead of the majority vote.
Let's look at an example.
Here we are creating some synthetic data with fifty examples and only one feature.
We only have one feature of `length` and our goal is to predict `weight`.
Regression plots more naturally in 1D, classification in 2D, but of course we can do either for any $d$
Right now, do not worry about the code and only focus on data and our model.
np.random.seed(0)
n = 50
X_1 = np.linspace(0,2,n)+np.random.randn(n)*0.01
X = pd.DataFrame(X_1[:,None], columns=['length'])
X.head()
y = abs(np.random.randn(n,1))*2 + X_1[:,None]*5
y = pd.DataFrame(y, columns=['weight'])
y.head()
snake_X_train, snake_X_test, snake_y_train, snake_y_test = train_test_split(X, y, test_size=0.2, random_state=123)
Now let's visualize our training data.
source = pd.concat([snake_X_train, snake_y_train], axis=1)
scatter = alt.Chart(source, width=500, height=300).mark_point(filled=True, color='green').encode(
alt.X('length:Q'),
alt.Y('weight:Q'))
scatter
Now let's try the $k$-nearest neighbours regressor on this data.
Then we create our `KNeighborsRegressor` object with `n_neighbors=1` so we are only considering 1 neighbour and with `uniform` weights.
from sklearn.neighbors import KNeighborsRegressor
knnr_1 = KNeighborsRegressor(n_neighbors=1, weights="uniform")
knnr_1.fit(snake_X_train,snake_y_train);
predicted = knnr_1.predict(snake_X_train)
predicted
If we scored over regressors we get this perfect score of one since we have `n_neighbors=1` we are likely to overfit.
knnr_1.score(snake_X_train, snake_y_train)
Plotting this we can see our model is trying to get every example correct since n_neighbors=1. (the mean of 1 point is just going to be the point value)
plt.figure(figsize=(8, 5))
grid = np.linspace(np.min(snake_X_train), np.max(snake_X_train), 1000)
plt.plot(grid, knnr_1.predict(grid), color='orange', linewidth=1)
plt.plot(snake_X_train, snake_y_train, ".r", markersize=10, color='green')
plt.xticks(fontsize= 14);
plt.yticks(fontsize= 14);
plt.xlabel("length",fontsize= 14)
plt.ylabel("weight",fontsize= 14);
What happens when we use `n_neighbors=10`?
knnr_10 = KNeighborsRegressor(n_neighbors=10, weights="uniform")
knnr_10.fit(snake_X_train, snake_y_train)
knnr_10.score(snake_X_train, snake_y_train)
Now we can see we are getting a lower score over the training set. Our score decreased from 1.0 when to had `n_neighbors=1` to now having a score of 0.925.
When we plot our model, we can see that it no longer is trying to get every example correct.
plt.figure(figsize=(8, 5))
plt.plot(grid, knnr_10.predict(grid), color='orange', linewidth=1)
plt.plot(snake_X_train, snake_y_train, ".r", markersize=10, color='green')
plt.xticks(fontsize= 16);
plt.yticks(fontsize= 16);
plt.xlabel("length",fontsize= 16)
plt.ylabel("weight",fontsize= 16);
## Pros and Cons of 𝑘 -Nearest Neighbours
### Pros:
- Easy to understand, interpret.
- Simply hyperparameter $k$ (`n_neighbors`) controlling the fundamental tradeoff.
- Can learn very complex functions given enough data.
- Lazy learning: Takes no time to `fit`
<br>
### Cons:
- Can potentially be VERY slow during prediction time.
- Often not that great test accuracy compared to the modern approaches.
- Need to scale your features. We'll be looking into this in an upcoming lecture (lecture 4 I think?).
## Let's Practice
$$ X = \begin{bmatrix}5 & 2\\4 & 3\\ 2 & 2\\ 10 & 10\\ 9 & -1\\ 9& 9\end{bmatrix}, \quad y = \begin{bmatrix}0\\0\\1\\1\\1\\2\end{bmatrix}.$$
If $k=3$, what would you predict for $x=\begin{bmatrix} 0\\0\end{bmatrix}$ if we were doing regression rather than classification?
```{admonition} Solutions!
:class: dropdown
1. 1/3 ($\frac{0 + 0 + 0}{3}$)
```
## Support Vector Machines (SVMs) with RBF Kernel
Another popular similarity-based algorithm is Support Vector Machines (SVM).
SVMs use a different similarity metric which is called a “kernel” in "SVM land".
We are going to concentrate on the specific kernel called Radial Basis Functions (RBFs).
Back to the good ol' Canadian and USA cities data.
cities_df = pd.read_csv("data/canada_usa_cities.csv")
cities_train_df, cities_test_df = train_test_split(cities_df, test_size=0.2, random_state=123)
cities_train_df.head()
cities_X_train = cities_train_df.drop(columns=['country'])
cities_y_train = cities_train_df['country']
cities_X_test = cities_test_df.drop(columns=['country'])
cities_y_test = cities_test_df['country']
cities_X_train.head()
cities_y_train.head()
Unlike with $k$-nn, we are not going into detail about how support vector machine classifiers or regressor works but more so on how to use it with `sklearn`.
We can use our training feature table ($X$) and target ($y$) values by using this new SVM model with (RBF) but with the old set up with `.fit()` and `.score()` that we have seen time and time again.
We import the `SVC` tool from the `sklearn.svm` library (The "C" in SVC represents *Classifier*.
To import the regressor we import `SVR` - R for *Regressor*)
from sklearn.svm import SVC
from sklearn.svm import SVR
We can cross-validate and score exactly how we saw before.
(For now, ignore `gamma=0.01` we are addressing it coming up)
svm = SVC(gamma=0.01)
scores = cross_validate(svm, cities_X_train, cities_y_train, return_train_score=True)
pd.DataFrame(scores)
svm_cv_score = scores['test_score'].mean()
svm_cv_score
The biggest thing to know about support vector machines is that superficially, support vector machines are very similar to 𝑘-Nearest Neighbours.
You can think of SVM with RBF kernel as a "smoothed" version of the $k$-Nearest Neighbours.
svm.fit(cities_X_train, cities_y_train);
kn5_model = KNeighborsClassifier(n_neighbors=5)
kn5_model.fit(cities_X_train, cities_y_train);
plt.figure(figsize=(16, 8))
plt.subplot(1, 2, 1)
plt.title("SVC")
plot_classifier(cities_X_train, cities_y_train, svm, ax=plt.gca())
plt.subplot(1, 2, 2)
plt.title("KNN with k = 5")
plot_classifier(cities_X_train, cities_y_train, kn5_model, ax=plt.gca());
An observation is classified as a positive class if on average it looks more like positive examples. An observation is classified as a negative class if on average it looks more like negative examples.
The primary difference between 𝑘-NNs and SVMs is that:
- Unlike $k$-NNs, SVMs only remember the key examples (Those examples are called **support vectors**).
- When it comes to predicting a query point, we only consider the key examples from the data and only calculate the distance to these key examples. This makes it more efficient than 𝑘-NN.
### Hyperparameters of SVM
There are 2 main hyperparameters for support vector machines with an RBF kernel;
- `gamma`
- `C`
(told you we were coming back to it!)
We are not equipped to understand the meaning of these parameters at this point but you are expected to describe their relationship to the fundamental tradeoff.
(In short, `C` is the penalty the model accepts for wrongly classified examples, and `gamma` is the curvature (see [here](https://towardsdatascience.com/hyperparameter-tuning-for-support-vector-machines-c-and-gamma-parameters-6a5097416167) for more)
See [`scikit-learn`'s explanation of RBF SVM parameters](https://scikit-learn.org/stable/auto_examples/svm/plot_rbf_parameters.html)
#### `gamma` and the fundamental trade-off
`gamma` controls the complexity of a model, just like other hyperparameters we've seen.
- higher gamma, higher the complexity.
- lower gamma, lower the complexity.
plt.figure(figsize=(16, 4))
for i in range(4):
plt.subplot(1, 4, i + 1)
gamma = 10.0 ** (i - 3)
rbf_svm = SVC(gamma=gamma)
rbf_svm.fit(cities_X_train, cities_y_train)
plt.title("gamma = %s" % gamma);
plot_classifier(cities_X_train, cities_y_train, rbf_svm, ax=plt.gca(), show_data=False)
#### `C` and the fundamental trade-off
`C` also controls the complexity of a model and in turn the fundamental tradeoff.
- higher `C` values, higher the complexity.
- lower `C` values, lower the complexity.
plt.figure(figsize=(16, 4))
for i in range(4):
plt.subplot(1, 4, i + 1)
C = 10.0 ** (i - 1)
rbf_svm = SVC(C=C, gamma=0.01)
rbf_svm.fit(cities_X_train, cities_y_train)
plt.title("C = %s" % C);
plot_classifier(cities_X_train, cities_y_train, rbf_svm, ax=plt.gca(), show_data=False)
Obtaining optimal validation scores requires a hyperparameter search between both `gamma` and `C` to balance the fundamental trade-off.
We will learn how to search over multiple hyperparameters at a time in lecture 5.
## Let's Practice
**True or False**
1\.In Scikit Learn’s SVC classifier, large values of gamma tend to result in higher training scores but probably lower validation scores.
2\.If we increase both `gamma` and `C`, we can't be certain if the model becomes more complex or less complex.
```{admonition} Solutions!
:class: dropdown
1. True
2. False
```
## Let's Practice - Coding
Below is some starter code that creates your feature table and target column from the data from the `bball.csv` dataset (in the data folder).
bball_df = pd.read_csv('data/bball.csv')
bball_df = bball_df[(bball_df['position'] =='G') | (bball_df['position'] =='F')]
# Define X and y
X = bball_df.loc[:, ['height', 'weight', 'salary']]
y = bball_df['position']
1. Split the dataset into 4 objects: `X_train`, `X_test`, `y_train`, `y_test`. Make the test set 0.2 (or the train set 0.8) and make sure to use `random_state=7`.
2. Create an `SVM` model with `gamma` equal to 0.1 and `C` equal to 10.
3. Cross-validate using cross_validate() on the objects X_train and y_train specifying the model and making sure to use 5 fold cross-validation and `return_train_score=True`.
4. Calculate the mean training and cross-validation scores.
# 1. Split the dataset
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=7)
model = SVC(gamma=0.1, C=10)
# 3. Cross-validate
scores_df = pd.DataFrame(cross_validate(model,X_train,y_train, cv=5, return_train_score=True))
scores_df
# 4. Calculate the mean training and cross-validation scores.
scores_df.mean()
scores_df.mean()['test_score']
scores_df.mean()['train_score']
## Preprocessing
### The importance of Preprocessing - An Example of Why
So far we have seen:
- Models: Decision trees, 𝑘-NNs, SVMs with RBF kernel.
- Fundamentals: Train-validation-test split, cross-validation, the fundamental tradeoff, the golden rule.
Now ...
**Preprocessing**: Transforming input data into a format a machine learning model can use and understand.
#### Basketball dataset
Let's take a look at the `bball.csv` dataset we just used in practice.
- Let's look at the 3 feature columns `height`, `weight` and `salary`.
- Let's see if these features can help predict the `position` basketball players is.
bball_df = pd.read_csv('data/bball.csv')
bball_df = bball_df[(bball_df['position'] =='G') | (bball_df['position'] =='F')]
X = bball_df[['weight', 'height', 'salary']]
y =bball_df["position"]
X_train, X_test, y_train, y_test =train_test_split(X, y, test_size=0.20, random_state=123)
X_train.head()
y_train.head()
First, let's see what validations scores we get if we simply predict the most occurring target value in the dataset using the dummy classifier model we saw in the last lecture.
dummy = DummyClassifier(strategy="most_frequent")
scores = cross_validate(dummy, X_train, y_train, return_train_score=True)
print('Mean training score', scores['train_score'].mean().round(2))
print('Mean validation score', scores['test_score'].mean().round(2))
Here we get a mean validation score for our 5 fold cross_validation (5 is the default) of 57%. Let's now see how much better a $k$-nn model does on the data. We saw that it doesn't do to well on SVM, let's see if there is a difference with $k$-nn.
knn = KNeighborsClassifier()
scores = cross_validate(knn, X_train, y_train, return_train_score=True)
print('Mean training score', scores['train_score'].mean().round(2))
print('Mean validation score', scores['test_score'].mean().round(2))
Ok, not the score we were hoping for.
We are getting a worse score than the dummy classifier. This can't be right..... and it isn't and we are going to explain why!
Let's have a look at just 2 players.
We can see the values in each column.
two_players = X_train.sample(2, random_state=42)
two_players
- The values in the `weight` column are around 100.
- The values in the `height` column are around 2.
- The values in the `salary` column are much higher at around 2 million.
Let’s now calculate the distance between the two players.
euclidean_distances(two_players)
So the distance between the players is 117133.0018.
What happens if we only consider the salary column?
euclidean_distances(two_players[["salary"]])
It looks like it's almost the same distance!
The distance is completely dominated by the `salary` column, the feature with the largest values and the `weight` and `height` columns are being ignored in the distance calculation.
**Does it matter?**
Yes! The scale is based on how data was collected.
Features on a smaller scale can be highly informative and there is no good reason to ignore them.
We want our model to be robust and not sensitive to the scale.
**What about for decision trees? Did scale matter then?**
No. In decision trees we ask questions on one feature at a time and so the nodes are created independently without considering others.
We have to scale our columns before we use our $k$-nn algorithm (and many others) so they are all using a similar range of values!
And you guessed it - Sklearn has tools called transformers for this.
We'll be using `sklearn`'s [`StandardScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html) for this example.
We will talk about this type of preprocessing in more detail in a hot minute but for now, concentrate on the syntax.
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler() # Create feature transformer object, can accept hyperparameters like models can!
scaler.fit(X_train) # Fitting the transformer on the train split
X_train_scaled = scaler.transform(X_train) # Transforming the train split
X_test_scaled = scaler.transform(X_test) # Transforming the test split
`sklearn` uses `fit` and `transform` paradigms for feature transformations. (In model building it was `fit` and `predict` or `score`)
We `fit` the transformer on the train split and then `transform` the train split as well as the test split.
pd.DataFrame(X_train_scaled, columns=X_train.columns).head()
Now if we look at our features they are all within the same scales as opposed to what it was before:
X_train.head()
### Sklearn's *predict* vs *transform*
When we make models, we `fit` and `predict`(`score`) with the syntax:
```
model.fit(X_train, y_train)
X_train_predictions = model.predict(X_train)
```
With preprocessing, we replace the `.predict()` step with a `.transform()` step. We can pass `y_train` in `fit` but it's usually ignored. It allows us to pass it just to be consistent with the usual usage of `sklearn`'s `fit` method.
```
transformer.fit(X_train, [y_train])
X_train_transformed = transformer.transform(X_train)
```
We can also carry out fitting and transforming in one call using `.fit_transform()`, but we must be mindful to use it only on the train split and **NOT** on the test split.
```
X_train_transformed = transformer.fit_transform(X_train)
```
Let's scale our features for this basketball dataset and then compare the results with our original score without scaling.
knn_unscaled = KNeighborsClassifier()
knn_unscaled.fit(X_train, y_train);
print('Train score: ', (knn_unscaled.score(X_train, y_train).round(2)))
print('Test score: ', (knn_unscaled.score(X_test, y_test).round(2)))
knn_scaled = KNeighborsClassifier()
knn_scaled.fit(X_train_scaled, y_train);
print('Train score: ', (knn_scaled.score(X_train_scaled, y_train).round(2)))
print('Test score: ', (knn_scaled.score(X_test_scaled, y_test).round(2)))
The scores with scaled data are now much better compared to the unscaled data in the case of 𝑘-NNs.
We can see now that 𝑘-NN is doing better than the Dummy Classifier when we scaled our features.
We are not carrying out cross-validation here for a reason that we'll look into soon.
We are being a bit sloppy here by using the test set several times for teaching purposes.
But when we build any ML models, we should only assess the test set once.
### Common preprocessing techniques
Here are some commonly performed feature transformation techniques we will focus on in this lesson.
- Imputation
- Tackling missing values
- Scaling
- Scaling of numeric features
## Let's Practice
1\. Name a model that will still produce meaningful predictions with different scaled column values.
2\. Complete the following statement: Preprocessing is done ______.
a) To the model but before training
b) To the data before training the model
c) To the model after training
d) To the data after training the model
3\. `StandardScaler` is a type of what?
4\. What data splits does `StandardScaler` alter (Training, Testing, Validation, None, All)?
**True or False**
5\. Columns with lower magnitudes compared to columns with higher magnitudes are less important when making predictions.
6\. A model less sensitive to the scale of the data makes it more robust.
```{admonition} Solutions!
:class: dropdown
1. Decision Tree Algorithm
2. b) To the data before training the model
3. Transformer
4. All
5. False
6. True
```
## California housing data (A case study)
For the next few examples of preprocessing, we are going to be using a dataset exploring the prices of homes in California to demonstrate feature transformation techniques. The data can be downloaded from this site [here](https://www.kaggle.com/harrywang/housing). Please make sure that you include it in your `data` folder that resides in `lectures`.
This dataset is a modified version of the California Housing dataset available from [Luís Torgo's University of Porto website](https://www.dcc.fc.up.pt/~ltorgo/Regression/cal_housing.html)
The task is to predict median house values in California districts, given several features from these districts.
housing_df = pd.read_csv("data/housing.csv")
train_df, test_df = train_test_split(housing_df, test_size=0.1, random_state=123)
train_df.head()
Some column values are mean/median but some are not.
Before we use this data we need to do some **feature engineering**.
That means we are going to transform our data into features that may be more meaningful for our prediction.
Let's add some new features to the dataset which could help predict the target: `median_house_value`.
train_df = train_df.assign(rooms_per_household = train_df["total_rooms"]/train_df["households"],
bedrooms_per_household = train_df["total_bedrooms"]/train_df["households"],
population_per_household = train_df["population"]/train_df["households"])
test_df = test_df.assign(rooms_per_household = test_df["total_rooms"]/test_df["households"],
bedrooms_per_household = test_df["total_bedrooms"]/test_df["households"],
population_per_household = test_df["population"]/test_df["households"])
train_df = train_df.drop(columns=['total_rooms', 'total_bedrooms', 'population'])
test_df = test_df.drop(columns=['total_rooms', 'total_bedrooms', 'population'])
train_df.head()
### When is it OK to do things before splitting?
- Here it would have been OK to add new features before splitting because we are not using any global information in the data but only looking at one row at a time.
- But just to be safe and to avoid accidentally breaking the golden rule, it's better to do it after splitting.
## Preprocessing: Imputation
Imputation is handling missing values in our data so let's explore this a little.
We can `.info()` we can we all the different column dtypes and also all the number of null values.
train_df.info()
We see that we have all columns with dtype `float64` except for `ocean_proximity` which appears categorical.
We also notice that the `bedrooms_per_household` column appears to have some `Non-Null` rows.
train_df["bedrooms_per_household"].isnull().sum()
Knowing this information let's build a model.
When we create our feature table and target objects, we are going to drop the categorical variable `ocean_proximity`. Currently, we don't know how to build models with categorical data, but we will shortly. We will return to this column soon.
X_train = train_df.drop(columns=["median_house_value", "ocean_proximity"])
y_train = train_df["median_house_value"]
X_test = test_df.drop(columns=["median_house_value", "ocean_proximity"])
y_test = test_df["median_house_value"]
knn = KNeighborsRegressor()
What happens when we try to fit our model with this data?
knn.fit(X_train, y_train)
> `Input contains NaN, infinity or a value too large for dtype('float64').`
The classifier can't deal with missing values (NaNs).
How can we deal with this problem?
### Why we don't drop the rows
We could drop any rows that are missing information but that's problematic too.
Then we would need to do the same in our test set.
And what happens if we get missing values in our deployment data? what then?
Furthermore, what if the missing values don't occur at random and we're systematically dropping certain data?
Perhaps a certain type of house contributes to more missing values.
Dropping the rows is not a great solution, especially if there's a lot of missing values.
X_train.shape
X_train_no_nan = X_train.dropna()
y_train_no_nan = y_train.dropna()
X_train_no_nan.shape
### Why we don't drop the column
If we drop the column instead of the rows, we are throwing away, in this case, 18391 values just because we don't have 185 missing values out of a total of 18567.
We are throwing away 99% of the column’s data because we are missing 1%.
But perhaps if we were missing 99.9% of the column values, for example, it would make more sense to drop the column.
X_train.shape
X_train_no_col = X_train.dropna(axis=1)
X_train_no_col.shape
### Why we use imputation
With **Imputation**, we invent values for the missing data.
Using `sklearn`'s **transformer** `SimpleImputer`, we can impute the `NaN` values in the data with some value.
from sklearn.impute import SimpleImputer
We can impute missing values in:
- **Categorical columns**:
- with the most frequent value
- with a constant of our choosing.
- **Numeric columns**:
- with the mean of the column
- with the median of the column
- or a constant of our choosing.
If I sort the values by `bedrooms_per_household` and look at the end of the dataframe, we can see our missing values in the `bedrooms_per_household` column.
Pay close attention to index 7763 since we are going to look at this row after imputation.
X_train.sort_values('bedrooms_per_household').tail(10)
Using the same `fit` and `transform` syntax we saw earlier for transformers, we can impute the `NaN` values.
Here we specify `strategy="median"` which replaces all the missing values with the column median.
We fit on the training data and transform it on the train and test splits.
imputer = SimpleImputer(strategy="median")
imputer.fit(X_train);
X_train_imp = imputer.transform(X_train)
X_test_imp = imputer.transform(X_test)
X_train_imp
Ok, the output of this isn't a dataframe but a NumPy array!
I can do a bit of wrangling here to take a look at this new array with our previous column labels and as a dataframe.
If I search for our index 7763 which previously contained a `NaN` value, we can see that now I have the median value for the `bedrooms_per_household` column from the `X_train` dataframe.
X_train_imp_df = pd.DataFrame(X_train_imp, columns = X_train.columns, index = X_train.index)
X_train_imp_df.loc[[7763]]
X_train['bedrooms_per_household'].median()
X_train.loc[[7763]]
Now when we try and fit our model using `X_train_imp`, it works!
knn = KNeighborsRegressor();
knn.fit(X_train_imp, y_train)
knn.score(X_train_imp, y_train)
## Preprocessing: Scaling
So we've seen why scaling is important earlier but let's take a little bit of a closer look here.
There are many ways to scale your data but we are going to look at 2 of them.
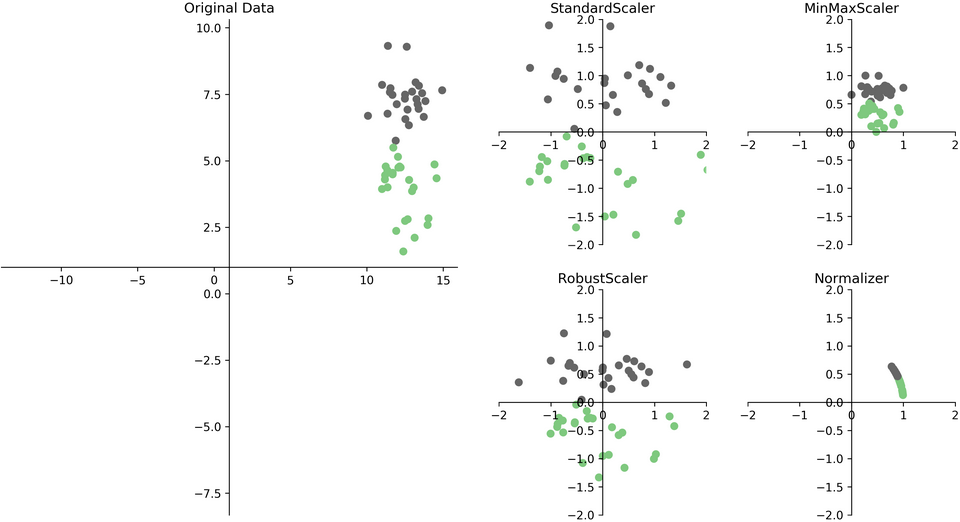
| Approach | What it does | How to update $X$ (but see below!) | sklearn implementation |
|---------|------------|-----------------------|----------------|
| normalization | sets range to $[0,1]$ | `X -= np.min(X,axis=0)`<br>`X /= np.max(X,axis=0)` | [`MinMaxScaler()`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
| standardization | sets sample mean to $0$, s.d. to $1$ | `X -= np.mean(X,axis=0)`<br>`X /= np.std(X,axis=0)` | [`StandardScaler()`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler) |
For more resources and articles on this, see [here](http://www.dataminingblog.com/standardization-vs-normalization/) and [here](https://medium.com/@rrfd/standardize-or-normalize-examples-in-python-e3f174b65dfc).
Let's see what happens when we use each of them.
from sklearn.preprocessing import MinMaxScaler, StandardScaler
First, let's see how standardization is done first.
scaler = StandardScaler()
X_train_scaled_std = scaler.fit_transform(X_train_imp)
X_test_scaled_std = scaler.transform(X_test_imp)
pd.DataFrame(X_train_scaled_std, columns=X_train.columns, index=X_train.index).head()
Here, any negative values represent values that are lower than the calculated feature mean and anything positive and greater than 0 are values greater than the original column mean.
knn = KNeighborsRegressor()
knn.fit(X_train_imp, y_train);
print('Unscaled training score :', knn.score(X_train_imp, y_train).round(3))
knn = KNeighborsRegressor()
knn.fit(X_train_scaled_std, y_train)
print('Scaled training score :',knn.score(X_train_scaled_std, y_train))
scaler = MinMaxScaler()
X_train_scaled_norm = scaler.fit_transform(X_train_imp)
X_test_scaled_norm = scaler.transform(X_test_imp)
pd.DataFrame(X_train_scaled_norm, columns=X_train.columns, index=X_train.index).head()
Looking at the data after normalizing it, we see this time there are no negative values and they all are between 0 and 1.
And the score now?
knn = KNeighborsRegressor()
knn.fit(X_train_scaled_norm, y_train)
print('Scaled training score :',knn.score(X_train_scaled_norm, y_train))
- Big difference in the KNN training performance after scaling the data.
- But we saw last week that the training score doesn't tell us much. We should look at the cross-validation score.
So let's see how we can do this but first.... let's practice!
## Let's Practice
1\. When/Why do we need to impute our data?
2\. If we have `NaN` values in our data, can we simply drop the column missing the data?
3\. Which scaling method will never produce negative values?
4\. Which scaling method will never produce values greater than 1?
5\. Which scaling method will produce values where the range depends on the values in the data?
**True or False**
6\. `SimpleImputer` is a type of transformer.
7\. Scaling is a form of transformation.
8\. We can use `SimpleImputer` to impute values that are missing from numerical and categorical columns.
```{admonition} Solutions!
:class: dropdown
1. When we have missing data so that sklearn doesn't give an error.
2. No but we can if the majority of the values are missing from the column.
3. Normalization (`MinMaxScaler`)
4. Normalization (`MinMaxScaler`)
5. Standardization (`StandardScaler`)
6. True
7. True
8. True
```
## Feature transformations and the golden rule
How to carry out cross-validation?
- Last week we saw that cross-validation is a better way to get a realistic assessment of the model.
- Let's try cross-validation with transformed data.
knn = KNeighborsRegressor()
scores = cross_validate(knn, X_train_scaled_std, y_train, return_train_score=True)
pd.DataFrame(scores)
- Do you see any problem here?
We are using our `X_train_scaled` in our `cross_validate()` function which already has all our preprocessing done.
<img src='imgs/cross-validation.png' width="80%">
That means that our validation set information is being used to calculate the mean and standard deviation (or min and max values for `MinMaxScaler`) for our training split!
We are allowing information from the validation set to **leak** into the training step.
What was our golden rule of machine learning again? Oh yeah -> ***Our test data should not influence our training data***.
This applies also to our validation data and that it also should not influence our training data.
With imputation and scaling, we are scaling and imputing values based on all the information in the data meaning the training data AND the validation data and so we are not adhering to the golden rule anymore.
Every row in our `x_train_scaled` has now been influenced in a minor way by every other row in `x_train_scaled`.
With scaling every row has been transformed based on all the data before splitting between training and validation.
We need to take care that we are keeping our validation data truly as unseen data.
Before we look at the right approach to this, let's look at the **WRONG** approaches.
### Bad methodology 1: Scaling the data separately
We make our transformer, we fit it on the training data and then transform the training data.
Then, we make a second transformer, fit it on the test data and then transform our test data.
scaler = StandardScaler();
scaler.fit(X_train_imp);
X_train_scaled = scaler.transform(X_train_imp)
# Creating a separate object for scaling test data - Not a good idea.
scaler = StandardScaler();
scaler.fit(X_test_imp); # Calling fit on the test data - Yikes!
X_test_scaled = scaler.transform(X_test_imp) # Transforming the test data using the scaler fit on test data ... Bad!
knn = KNeighborsRegressor()
knn.fit(X_train_scaled, y_train);
print("Training score: ", knn.score(X_train_scaled, y_train).round(2))
print("Test score: ", knn.score(X_test_scaled, y_test).round(2))
This is bad because we are using two different StandardScaler objects but we want to apply the same transformation on the training and test splits.
The test data will have different values than the training data producing a different transformation than the training data.
We should never fit on test data, whether it’s to build a model or with a transforming, test data should never be exposed to the fit function.
### Bad methodology 2: Scaling the data together
The next mistake is when we scale the data together. So instead of splitting our data, we are combining our training and testing and scaling it together.
X_train_imp.shape, X_test_imp.shape
# join the train and test sets back together
XX = np.vstack((X_train_imp, X_test_imp))## Don't do it!
XX.shape
scaler = StandardScaler()
scaler.fit(XX)
XX_scaled = scaler.transform(XX)
XX_train = XX_scaled[:18576]
XX_test = XX_scaled[18576:]
knn = KNeighborsRegressor()
knn.fit(XX_train, y_train);
print('Train score: ', (knn.score(XX_train, y_train).round(2))) # Misleading score
print('Test score: ', (knn.score(XX_test, y_test).round(2))) # Misleading score
Here we are scaling the train and test splits together.
The golden rule says that the test data shouldn’t influence the training in any way.
Information from the test split is now affecting the mean for standardization!
This is a clear violation of the golden rule.
So what do we do? Enter ....
## Pipelines
[Scikit-learn Pipeline](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html) is here to save the day!
A **pipeline** is a sklearn function that contains a sequence of steps.
Essentially we give it all the actions we want to do with our data such as transformers and models and the pipeline will execute them in steps.
from sklearn.pipeline import Pipeline
Let's combine the preprocessing and model with pipeline.
we will instruct the pipeline to:
1. Do imputation using `SimpleImputer()` using a strategy of “median”
2. Scale our data using `StandardScaler`
3. Build a `KNeighborsRegressor`.
(The last step should be a model and earlier steps should be transformers)
Note: The input for `Pipeline` is a list containing tuples (one for each step).
pipe = Pipeline([
("imputer", SimpleImputer(strategy="median")),
("scaler", StandardScaler()),
("reg", KNeighborsRegressor())
])
pipe.fit(X_train, y_train)
- Note that we are passing `X_train` and **NOT** the imputed or scaled data here.
When we call `fit` the pipeline is carrying out the following steps:
- Fit `SimpleImputer` on `X_train`.
- Transform `X_train` using the fit `SimpleImputer` to create `X_train_imp`.
- Fit `StandardScaler` on `X_train_imp`.
- Transform `X_train_imp` using the fit `StandardScaler` to create `X_train_imp_scaled`.
- Fit the model (`KNeighborsRegressor` in our case) on `X_train_imp_scaled`.
pipe.predict(X_train)
When we call `predict` on our data, the following steps are carrying out:
- Transform `X_train` using the fit `SimpleImputer` to create `X_train_imp`.
- Transform `X_train_imp` using the fit `StandardScaler` to create `X_train_imp_scaled`.
- Predict using the fit model (`KNeighborsRegressor` in our case) on `X_train_imp_scaled`.
It is not fitting any of the data this time.
<img src='https://amueller.github.io/COMS4995-s20/slides/aml-04-preprocessing/images/pipeline.png' width="50%">
[Source](https://amueller.github.io/COMS4995-s20/slides/aml-04-preprocessing/#18)
We can’t accidentally re-fit the preprocessor on the test data as we did before.
It automatically makes sure the same transformations are applied to train and test.
Now when we do cross-validation on the pipeline the transformers and the model are refit on each fold.
The pipeline applies the `fit_transform` on the train portion of the data and only `transform` on the validation portion in **each fold**.
This is how to avoid the Golden Rule violation!
scores_processed = cross_validate(pipe, X_train, y_train, return_train_score=True)
pd.DataFrame(scores_processed)
pd.DataFrame(scores_processed).mean()
dummy = DummyRegressor(strategy="median")
scores = cross_validate(dummy, X_train, y_train, return_train_score=True)
pd.DataFrame(scores).mean()
We can trust here now that the scores are not influenced but the training data and all our steps were done efficiently and easily too.
## Let's Practice
1\. Which of the following steps cannot be used in a pipeline?
a) Scaling
b) Model building
c) Imputation
d) Data Splitting
2\. Why can't we fit and transform the training and test data together?
**True or False**
3\. We have to be careful of the order we put each transformation and model in a pipeline.
4\. Pipelines will fit and transform on both the training and validation folds during cross-validation.
```{admonition} Solutions!
:class: dropdown
1. Data Splitting
2. It's violating the golden rule.
3. True
4. False
```
## Let's Practice - Coding
Let's bring in the basketball dataset again.
# Loading in the data
bball_df = pd.read_csv('data/bball.csv')
bball_df = bball_df[(bball_df['position'] =='G') | (bball_df['position'] =='F')]
# Define X and y
X = bball_df.loc[:, ['height', 'weight', 'salary']]
y = bball_df['position']
# Split the dataset
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=7)
Build a pipeline named `bb_pipe` that:
1. Imputes using "median" as a strategy,
2. scale using `StandardScaler`
3. builds a `KNeighborsClassifier`.
Next, do 5 fold cross-validation on the pipeline using `X_train` and `y_train` and save the results in a dataframe.
Take the mean of each column and assess your model.
## What We've Learned Today<a id="9"></a>
- How the $k$NN algorithm works for regression.
- How to build an SVM with RBF kernel model.
- How changing `gamma` and `C` hyperparameters affects the fundamental tradeoff.
- How to imputer values when we are missing data.
- Why it's important to scale our features.
- How to scales our features.
- How to build a pipeline that executes a number of steps without breaking the golden rule of ML.
| 39,571 | 12,625 |
# 1
import math
r = float(input('输入五边形顶点到中心的距离:'))
s = 2*r*(math.sin(math.pi/5))
Area = 5*s*s/(4*math.tan(math.pi/5))
print('The area of the pentagon is %.2f'%Area) | 169 | 97 |
# _*_ coding: utf-8 _*_
"""
本部分定义应用的所有数据库对象。
"""
from werkzeug.security import generate_password_hash, check_password_hash
from flask.ext.login import UserMixin, AnonymousUserMixin
from itsdangerous import TimedJSONWebSignatureSerializer as Serializer
from flask import current_app, url_for
import datetime
from markdown import markdown
import bleach
from . import db, login_manager
ALL_ALLOWED_TAGS = ['a', 'abbr', 'acronym', 'b', 'blockquote', 'code', 'em', \
'i', 'li', 'ol', 'pre', 'strong', 'ul', 'h1', 'h2', 'h3', \
'h4', 'h5', 'h6', 'h7', 'p']
class Permission:
"""
权限管理:
0b00000001: 关注
0b00000010: 评论
0b00000100: 写文章
0b00001000: 管理他人的评论
0b00010000: 管理员权限
"""
FOLLOW = 0x01
COMMIT = 0x02
WRITE_ARTICLES = 0x04
MODERATE_COMMITS = 0x08
ADMINISTER = 0x08
class Follow(db.Model):
__tablename__ = "follows"
follower_id = db.Column(db.Integer, db.ForeignKey('users.id'), primary_key=True)
followed_id = db.Column(db.Integer, db.ForeignKey('users.id'), primary_key=True)
time_stamp = db.Column(db.DateTime, default=datetime.datetime.utcnow)
class Attendor(db.Model):
__tablename__ = "attendors"
attendee_id = db.Column(db.Integer, db.ForeignKey('users.id'), primary_key=True)
atconference_id = db.Column(db.Integer, db.ForeignKey('conferences.id'), primary_key=True)
time_stamp = db.Column(db.DateTime, default=datetime.datetime.utcnow)
class User(UserMixin, db.Model):
"""
用户表
"""
__tablename__ = "users"
id = db.Column(db.Integer, primary_key=True)
email = db.Column(db.String(64), unique=True, index=True)
nickname = db.Column(db.String(64), unique=True, index=True)
password_hash = db.Column(db.String(128))
address = db.Column(db.String(128))
about_me = db.Column(db.Text)
about_me_html = db.Column(db.Text)
join_time = db.Column(db.DateTime(), default=datetime.datetime.utcnow)
last_seen = db.Column(db.DateTime(), default=datetime.datetime.utcnow)
confirmed = db.Column(db.Boolean, default=False)
role_id = db.Column(db.Integer, db.ForeignKey("roles.id"))
conferences = db.relationship("Conference", backref="organizer", lazy='dynamic')
comments = db.relationship("Comment", backref="author", lazy="dynamic")
portrait_addr = db.Column(db.String(128))
followed = db.relationship("Follow",
foreign_keys=[Follow.follower_id],
backref=db.backref("follower", lazy="joined"),
lazy="dynamic",
cascade='all, delete-orphan')
followers = db.relationship("Follow",
foreign_keys=[Follow.followed_id],
backref=db.backref("followed", lazy="joined"),
lazy="dynamic",
cascade='all, delete-orphan')
atconferences = db.relationship("Attendor",
foreign_keys=[Attendor.attendee_id],
backref=db.backref("attendee", lazy="joined"),
lazy="dynamic",
cascade="all, delete-orphan")
# user.atconferences[k].attendee 是其自身
def __init__(self, **kwargs):
super(User, self).__init__(**kwargs)
if self.role is None:
if self.email == current_app.config["XING_ADMIN"]:
self.role = Role.query.filter_by(permissions = 0xff).first()
else:
self.role = Role.query.filter_by(default=True).first()
def to_json(self):
json_user = {
"id": self.id,
"url": url_for("api.get_user", id=self.id, _external=True),
"nickname": self.nickname,
"email": self.email,
"address": self.address,
"about_me": self.about_me,
"join_time": self.join_time,
"last_seen": self.last_seen,
"confirmed": self.confirmed,
"portrait_addr": self.portrait_addr,
"conferences": url_for("api.get_user_conferences", id = self.id, _external=True),
"conferences_count": self.conferences.count()
}
return json_user
def attend(self, conference):
if (not self.is_attend(conference)):
a = Attendor(attendee = self, atconference = conference)
db.session.add(a)
def unattend(self, conference):
a = self.atconferences.filter_by(atconference_id=conference.id).first()
if a:
db.session.delete(a)
def is_attend(self, conference):
return self.atconferences.filter_by(atconference_id=conference.id).first() is not None
def follow(self, user):
if not self.is_following(user):
f = Follow(follower=self, followed=user)
db.session.add(f)
def unfollow(self, user):
f = self.followed.filter_by(follower_id=user.id).first()
if f:
db.session.delete(f)
@property
def followers_conferences(self):
return Conference.query.join(Follow, \
Follow.followed_id == Conference.organizer_id).filter(\
Follow.follower_id == self.id)
def is_following(self, user):
return self.followed.filter_by(followed_id=user.id).first() is not None
def is_followed_by(self, user):
return self.followers.filter_by(follower_id=user.id).first() is not None
def ping(self):
"""
刷新用户的访问时间
"""
self.last_seen = datetime.datetime.utcnow()
db.session.add(self)
db.session.commit()
def can(self, permission):
"""
Verify this user's role.
If this user has permission required, return true, vise vese.
"""
return (self.role is not None) and \
((self.role.permissions & permission) == permission)
def is_administrator(self):
return self.can(Permission.ADMINISTER)
@property
def passwd(self):
raise AttributeError("Password is not a readable attribute")
@passwd.setter
def passwd(self, passwd):
self.password_hash = generate_password_hash(passwd)
@property
def portrait(self):
if self.portrait_addr:
return self.portrait_addr
else:
return "https://avatars3.githubusercontent.com/u/3695283?v=3&u=9269932cb4ce7e9b4976b62b2abcfe27f5b6f0a6&s=140"
@portrait.setter
def portrait(self, pt):
self.portrait_addr = pt
def verify_passwd(self, passwd):
return check_password_hash(self.password_hash, passwd)
def generate_confirmation_token(self, expiration=3600):
s = Serializer(current_app.config["SECRET_KEY"], expiration)
return s.dumps({'confirm': self.id})
def confirm(self, token):
s = Serializer(current_app.config["SECRET_KEY"])
try:
data = s.loads(token)
except:
return False
if data.get("confirm") == self.id:
self.confirmed = True
db.session.add(self)
db.session.commit()
return True
return False
def generate_confirmation_token(self, expiration=3600):
s = Serializer(current_app.config['SECRET_KEY'], expiration)
return s.dumps({'confirm': self.id})
def generate_auth_token(self, expiration):
s = Serializer(current_app.config['SECRET_KEY'],
expires_in=expiration)
return s.dumps({'id': self.id}).decode('ascii')
@staticmethod
def verify_auth_token(token):
s = Serializer(current_app.config["SECRET_KEY"])
try:
data = s.loads(token)
except:
return None
return User.query.get(data['id'])
@staticmethod
def on_change_about_me(target, value, oldvalue, initiator):
target.about_me_html = bleach.linkify(bleach.clean(markdown(value, \
output_format="html"), tags=ALL_ALLOWED_TAGS, \
strip=True))
@staticmethod
def generate_fake(count=100):
"""
说明:
conferences留给Conference在fake时生成。
comments留给Comment在fake时生成
atconferences留给conference的fake来实现
"""
from sqlalchemy.exc import IntegrityError
from random import seed, randint
import forgery_py
seed()
role_count = Role.query.count()
u = User(
email = "lianyun08@126.com",
nickname = "admin",
passwd = "12",
confirmed = True,
address = forgery_py.address.street_address(),
about_me = "administrator",
portrait_addr = "https://avatars3.githubusercontent.com/u/3695283?v=3&s=260"
)
db.session.add(u)
try:
db.session.commit()
except IntegrityError:
db.session.rollback()
for i in xrange(count):
jt = forgery_py.date.date()
delta_time = datetime.timedelta(days=randint(1, 10))
lt = jt + delta_time
r = Role.query.offset(randint(0, role_count - 1)).first()
user_count = User.query.count()
u = User(
email = forgery_py.internet.email_address(),
nickname = forgery_py.name.full_name(),
passwd = forgery_py.lorem_ipsum.word(),
confirmed = True,
address = forgery_py.address.street_address(),
about_me = forgery_py.forgery.currency.description(),
join_time = jt,
last_seen = lt,
role = r,
portrait_addr = "http://www.ttoou.com/qqtouxiang/allimg/120918/co12091Q01643-6-lp.jpg"
)
db.session.add(u)
try:
db.session.commit()
except IntegrityError:
db.session.rollback()
user_count = User.query.count()
for i in xrange(count):
u = User.query.offset(i).first()
followed_n = randint(0, 20)
for j in xrange(followed_n):
fu = User.query.offset(randint(0, user_count - 1)).first()
if fu.id == u.id:
continue
u.follow(fu)
def __str__(self):
return "<User %s>" % self.nickname
__repr__ = __str__
db.event.listen(User.about_me, 'set', User.on_change_about_me)
class AnonymousUser(AnonymousUserMixin):
def can(self, permissions):
return False
def is_administrator(self):
return True
login_manager.anonymous_user = AnonymousUser
class Role(db.Model):
__tablename__ = 'roles'
id = db.Column(db.Integer, primary_key=True)
name = db.Column(db.String(64), unique=True)
default = db.Column(db.Boolean, default=False, index=True)
permissions = db.Column(db.Integer)
users = db.relationship("User", backref="role", lazy="dynamic")
def to_json(self):
json_role = {
"id": self.id,
"name": self.name
}
return json_role
@staticmethod
def insert_roles():
roles = {
"User" : (Permission.FOLLOW |
Permission.COMMIT |
Permission.WRITE_ARTICLES, True),
"Moderator" : ( Permission.FOLLOW |
Permission.COMMIT |
Permission.WRITE_ARTICLES |
Permission.MODERATE_COMMITS, False),
"Administrator" : (0xff, False)
}
for r in roles:
role = Role.query.filter_by(name=r).first()
if role is None:
role = Role(name=r)
role.permissions = roles[r][0]
role.default = roles[r][1]
db.session.add(role)
db.session.commit()
def __str__(self):
return "<Role %s>" % self.name
__repr__ = __str__
class City(db.Model):
__tablename__ = "cities"
id = db.Column(db.Integer, primary_key=True)
name = db.Column(db.String(128), unique=True)
conferences = db.relationship("Conference", backref="city", lazy="dynamic")
def to_json(self):
json_city = {
"id": self.id,
"name": self.name
}
return json_city
@staticmethod
def insert_cities():
cities = {
"default": "anywhere",
"xi_an": "Xi An",
"bei_jing": "Bei Jing",
"shang_hai": "Shang Hai"
}
for (value, name) in cities.items():
city = City.query.filter_by(name=name).first()
if city is None:
city = City(name=name)
db.session.add(city)
db.session.commit()
def __str__(self):
return "<City %s>" % self.name
__repr__ = __str__
class Topic(db.Model):
__tablename__ = "topics"
id = db.Column(db.Integer, primary_key=True)
name = db.Column(db.String(128), unique=True)
def to_json(self):
json_topic = {
"id": self.id,
"name": self.name
}
return json_topic
@staticmethod
def insert_topics():
topics = {
"programming": "Programming",
"web": "Web",
"movie": "Movie",
"health": "Health"
}
for (value, name) in topics.items():
topic = Topic.query.filter_by(name=name).first()
if topic is None:
topic = Topic(name=name)
db.session.add(topic)
db.session.commit()
def __str__(self):
return "<Topic %s>" % self.name
__repr__ = __str__
add_topics = db.Table('add_topics', \
db.Column("conference_id", db.Integer, db.ForeignKey("conferences.id")),
db.Column("topic_id", db.Integer, db.ForeignKey("topics.id")))
class Conference(db.Model):
__tablename__ = "conferences"
id = db.Column(db.Integer, primary_key=True)
organizer_id = db.Column(db.Integer, db.ForeignKey("users.id"))
title = db.Column(db.String(128), index=True, nullable=False)
city_id = db.Column(db.Integer, db.ForeignKey("cities.id"))
topics = db.relationship("Topic", secondary=add_topics, backref=db.backref("conferences", lazy="dynamic"), lazy="dynamic")
description = db.Column(db.Text)
description_html = db.Column(db.Text)
start_time = db.Column(db.DateTime)
end_time = db.Column(db.DateTime)
max_attendees = db.Column(db.Integer)
time_stamp = db.Column(db.DateTime, default=datetime.datetime.utcnow)
comments = db.relationship("Comment", backref="conference", lazy="dynamic")
attendees = db.relationship("Attendor",
foreign_keys=[Attendor.atconference_id],
backref=db.backref("atconference", lazy="joined"),
lazy="dynamic",
cascade="all, delete-orphan")
# 注意会议的attendees[k].atconference是其自身
def to_json(self):
json_conference = {
"id": self.id,
"url": url_for("api.get_conference", id = self.id, _external=True),
"title": self.title,
"city": url_for("api.get_city", id = self.city_id, _external=True),
"topics": url_for("api.get_conference_topics", id = self.id, _external=True),
"description": self.description,
"start_time": self.start_time,
"end_time": self.end_time,
"max_attendees": self.max_attendees,
"time_stamp": self.time_stamp,
"attendees": url_for("api.get_conference_attendees", id = self.id, _external=True),
"attendees_count": self.attendees.count(),
"comments": url_for("api.get_conference_comments", id = self.id, _external=True),
"comments_count": self.comments.count()
}
return json_conference
def __str__(self):
return "<Conference %s>" % self.title
__repr__ = __str__
@staticmethod
def on_change_description(target, value, oldvalue, initiator):
target.description_html = bleach.linkify(bleach.clean(markdown(value, \
output_format="html"), tags=ALL_ALLOWED_TAGS, \
strip=True))
@staticmethod
def generate_fake(count=100):
from random import seed, randint
import forgery_py
seed()
user_count = User.query.count()
city_count = City.query.count()
topic_count = Topic.query.count()
delta_time = datetime.timedelta(days=1)
for i in range(count):
u = User.query.offset(randint(0, user_count - 1)).first()
c = City.query.offset(randint(0, city_count - 1)).first()
t = forgery_py.lorem_ipsum.title()
tps = []
for ii in xrange(randint(0, topic_count-1)):
tps.append(Topic.query.offset(randint(0, topic_count-1)).first())
stime = forgery_py.date.date(True)
etime = stime + delta_time
des = forgery_py.forgery.basic.text(at_least=15, at_most=50, digits=True, spaces=True, punctuation=False)
max_attendees = randint(5, 25)
attendees = [User.query.offset(randint(0, user_count-1)).first() for i in xrange(randint(0,max_attendees))]
time_stamp = forgery_py.date.date(True)
conference = Conference(
organizer = u,
city = c,
title = t,
description = des,
start_time = stime,
end_time = etime,
max_attendees = max_attendees,
time_stamp = time_stamp
)
for t in tps:
conference.topics.append(t)
for u in attendees:
u.attend(conference)
db.session.add(conference)
try:
db.session.commit()
except IntegrityError:
db.session.rollback()
db.event.listen(Conference.description, 'set', Conference.on_change_description)
class Comment(db.Model):
__tablename__ = "comments"
id = db.Column(db.Integer, primary_key=True)
body = db.Column(db.Text)
body_html = db.Column(db.Text)
time_stamp = db.Column(db.DateTime, index=True, default=datetime.datetime.utcnow)
disabled = db.Column(db.Boolean)
author_id = db.Column(db.Integer, db.ForeignKey("users.id"))
conference_id = db.Column(db.Integer, db.ForeignKey("conferences.id"))
def to_json(self):
json_comment = {
"id": self.id,
"body": self.body,
"time_stamp": self.time_stamp,
"author": url_for("api.get_user", id=self.author_id, _external=True),
"conference": url_for("api.get_conference", id=self.conference_id, _external=True)
}
return json_comment
@staticmethod
def on_change_body(target, value, oldvalue, initiator):
target.body_html = bleach.linkify(bleach.clean(markdown(value, output_format="html"),\
tags=ALL_ALLOWED_TAGS, strip=True))
@staticmethod
def generate_fake(count=550):
from random import seed, randint
import forgery_py
seed()
user_count = User.query.count()
conference_count = Conference.query.count()
for i in xrange(count):
u = User.query.offset(randint(0, user_count-1)).first()
conf = Conference.query.offset(randint(0, conference_count-1)).first()
c = Comment(
body = forgery_py.forgery.basic.text(at_least=10, at_most=25),
time_stamp = forgery_py.forgery.date.date(),
disabled = False,
author = u,
conference = conf
)
db.session.add(c)
try:
db.session.commit()
except IntegrityError:
db.session.rollback()
db.event.listen(Comment.body, 'set', Comment.on_change_body)
@login_manager.user_loader
def load_user(user_id):
return User.query.get(int(user_id))
| 20,918 | 6,601 |
import random
al1 = input('Digite o nome do(a) primeiro(a) aluno(a): ')
al2 = input('Digite o nome do (a) segundo(a) aluno(a): ')
al3 = input('Digite o nome do(a) terceiro(a) aluno(a): ')
al4 = input('Digite o nome do(a) quarto(a) aluno(a): ')
lista = [al1, al2, al3, al4]
random.shuffle(lista)
print(f'A ordem dos alunos para a apresentação será: \n{lista}.')
#from random import shuffle
| 395 | 164 |
# RSA
# RSA Tool generate private public key
from Crypto.PublicKey import RSA
from Crypto.Cipher import PKCS1_OAEP
from Crypto import Random
from Crypto.Hash import SHA256
import base64
from Crypto.Signature import PKCS1_PSS
class CrptyoRSA:
PRIVATE_KEY_FILE = "private_key.pem"
PUBLIC_KEY_FILE = "public_key.pem"
def __init__(self):
return
def __save_file(self, contents, file_name):
f = open(file_name, "wb")
f.write(contents)
f.close()
def __read_file(self, file_name):
f = open(file_name, "rb")
contents = f.read()
f.close()
return contents
def __generate_random(self):
return Random.new().read()
def generate_keys(self):
keys = RSA.generate(4096)
private_key = keys.exportKey('PEM')
public_key = keys.publickey().exportKey("PEM")
self.__save_file(private_key, self.PRIVATE_KEY_FILE)
self.__save_file(public_key, self.PUBLIC_KEY_FILE)
print("public & private keys stored")
def encrypt(self, cleartext, public_keypath=None):
if public_keypath is None:
public_keypath = self.PUBLIC_KEY_FILE
public_key = RSA.importKey(self.__read_file(public_keypath))
cipher = PKCS1_OAEP.new(public_key)
encrypted_data = cipher.encrypt(cleartext)
return base64.b64encode(encrypted_data)
def decrypt(self, cleartext, private_keypath=None):
if private_keypath is None:
private_keypath = self.PRIVATE_KEY_FILE
cipher_text = base64.b64decode(cleartext)
private_key = RSA.importKey(self.__read_file(private_keypath))
cipher = PKCS1_OAEP.new(private_key)
return cipher.decrypt(cipher_text)
def __sha256(self, input):
sha256 = SHA256.new()
sha256.update(input)
return sha256
def sign(self, textmessage, private_key_path=None):
if private_key_path is None:
private_key_path = self.PRIVATE_KEY_FILE
# create public key object
private_key = RSA.importKey(self.__read_file(private_key_path))
# create the verifier
signature = PKCS1_PSS.new(private_key)
return signature.sign(self.__sha256(textmessage))
def verify(self, signed_signature, textmessage, public_key_path=None):
if public_key_path is None:
public_key_path = self.PUBLIC_KEY_FILE
# create public key object
public_key = RSA.importKey(self.__read_file(public_key_path))
# create the verifier
verifier = PKCS1_PSS.new(public_key)
verification = verifier.verify(
self.__sha256(textmessage), signed_signature)
print(verification)
# CrptyoRSA().generate_keys()
# encrypted_data = CrptyoRSA().encrypt("Hello World".encode('utf-8'))
# print(encrypted_data)
# decrypted_data = CrptyoRSA().decrypt(encrypted_data)
# print(decrypted_data)
signed_signature = CrptyoRSA().sign("Hello World".encode('utf-8'))
CrptyoRSA().verify(signed_signature, "Hello World".encode('utf-8'))
| 3,058 | 1,045 |
from utils.utils import bresenham_line
from math import hypot, cos, sin
from utils.position import Position
class Cartographer:
"""
Class that implements a Cartographer, used to update the map of the environment using the lasers echoes.
"""
def __init__(self, lasers_distance = 0.15, min_increment = 0.015, increment = 0.15, max_distance = 40, safe_distance_obstacle = 5, safe_distance_empty = 10):
"""
Instantiates a Cartographer.
:param lasers_distance: Offset of the lasers in regard of the robot.
:type lasers_distance: float
:param min_increment: Minimal increment for update of the cells of the map.
:type min_increment: float
:param increment: Increment for update of the cells of the map.
:type increment: float
:param max_distance: Maximum distance of the echoes.
:type max_distance: float
:param safe_distance_obstacle: Used to be more precise on echoes readings.
:type safe_distance_obstacle: float
:param safe_distance_obstacle: Used to be more precise on echoes readings.
:type safe_distance_obstacle: float
"""
self.__lasers_distance = lasers_distance
self.__max_distance = max_distance
self.__min_increment = min_increment
self.__increment = increment
self.__safe_distance_obstacle = safe_distance_obstacle
self.__safe_distance_empty = safe_distance_empty
def update(self, robot_map, robot_pos, lasers):
"""
Function used to update the map by analyzing the lasers echoes, it uses the Bresenham line algorithm (implemented in utils.utils) to update lines.
:param robot_map: The map to update.
:type robot_map: Map
:param robot_pos: Robot position in the real world.
:type robot_pos: Position
:param lasers: The lasers datas.
:type lasers: A list of Laser objects.
:return: The map updated.
:rtype: Map
"""
lasers_pos_x = robot_pos.x + self.__lasers_distance * cos(robot_pos.angle)
lasers_pos_y = robot_pos.y + self.__lasers_distance * sin(robot_pos.angle)
lasers_cell = robot_map.to_grid_pos(Position(lasers_pos_x, lasers_pos_y))
real_lasers_cell = robot_map.to_real_pos(lasers_cell)
for laser in lasers:
angle = robot_pos.angle + laser.angle
laser_hit = Position(lasers_pos_x + laser.echoe * cos(angle), lasers_pos_y + laser.echoe * sin(angle))
hit_cell = robot_map.to_grid_pos(laser_hit)
cells = bresenham_line(lasers_cell.x, lasers_cell.y, hit_cell.x, hit_cell.y)
for cell in cells:
if robot_map.is_in_bound(cell):
if cell.x == hit_cell.x and cell.y == hit_cell.y:
if laser.echoe < self.__max_distance - self.__safe_distance_obstacle:
inc_iro_certainty = self.__min_increment if robot_map.is_empty(cell) else self.__increment
inc_factor_iro_dist = (1.0 - (laser.echoe / self.__max_distance))
robot_map.grid[cell.x][cell.y] += inc_factor_iro_dist * inc_iro_certainty
if robot_map.grid[cell.x][cell.y] > 1.0:
robot_map.grid[cell.x][cell.y] = 1.0
else:
real_cell = robot_map.to_real_pos(cell)
distance = hypot(real_cell.x - real_lasers_cell.x, real_cell.y - real_lasers_cell.y)
if distance < self.__max_distance - self.__safe_distance_empty:
inc_iro_certainty = self.__min_increment if robot_map.is_obstacle(cell) else self.__increment
inc_factor_iro_dist = (1.0 - (distance / self.__max_distance))
robot_map.grid[cell.x][cell.y] -= inc_factor_iro_dist * inc_iro_certainty
if robot_map.grid[cell.x][cell.y] < 0.0:
robot_map.grid[cell.x][cell.y] = 0.0
return robot_map
| 4,122 | 1,244 |
"""
Draw methods for textures and depth textures
"""
import moderngl
from demosys import context, geometry
class TextureHelper:
"""Draw methods for textures and depth textures"""
_quad = None
_texture2d_shader = None # Type: moderngl.Program
_texture2d_sampler = None # Type: moderngl.Sampler
_depth_shader = None # Type: moderngl.Program
_depth_sampler = None # Type: moderngl.Sampler
def __init__(self):
self._init_texture2d_draw()
self._init_depth_texture_draw()
@property
def initialized(self):
return self._quad is not None
@property
def ctx(self):
return context.ctx()
def draw(self, texture, pos=(0.0, 0.0), scale=(1.0, 1.0)):
"""
Draw texture using a fullscreen quad.
By default this will conver the entire screen.
:param pos: (tuple) offset x, y
:param scale: (tuple) scale x, y
"""
if not self.initialized:
self.init()
self._texture2d_shader["offset"].value = (pos[0] - 1.0, pos[1] - 1.0)
self._texture2d_shader["scale"].value = (scale[0], scale[1])
texture.use(location=0)
self._texture2d_sampler.use(location=0)
self._texture2d_shader["texture0"].value = 0
self._quad.render(self._texture2d_shader)
self._texture2d_sampler.clear(location=0)
def draw_depth(self, texture, near, far, pos=(0.0, 0.0), scale=(1.0, 1.0)):
"""
Draw depth buffer linearized.
By default this will draw the texture as a full screen quad.
A sampler will be used to ensure the right conditions to draw the depth buffer.
:param near: Near plane in projection
:param far: Far plane in projection
:param pos: (tuple) offset x, y
:param scale: (tuple) scale x, y
"""
if not self.initialized:
self.init()
self._depth_shader["offset"].value = (pos[0] - 1.0, pos[1] - 1.0)
self._depth_shader["scale"].value = (scale[0], scale[1])
self._depth_shader["near"].value = near
self._depth_shader["far"].value = far
self._depth_sampler.use(location=0)
texture.use(location=0)
self._depth_shader["texture0"].value = 0
self._quad.render(self._depth_shader)
self._depth_sampler.clear(location=0)
def _init_texture2d_draw(self):
"""Initialize geometry and shader for drawing FBO layers"""
if not TextureHelper._quad:
TextureHelper._quad = geometry.quad_fs()
# Shader for drawing color layers
TextureHelper._texture2d_shader = context.ctx().program(
vertex_shader="""
#version 330
in vec3 in_position;
in vec2 in_uv;
out vec2 uv;
uniform vec2 offset;
uniform vec2 scale;
void main() {
uv = in_uv;
gl_Position = vec4((in_position.xy + vec2(1.0, 1.0)) * scale + offset, 0.0, 1.0);
}
""",
fragment_shader="""
#version 330
out vec4 out_color;
in vec2 uv;
uniform sampler2D texture0;
void main() {
out_color = texture(texture0, uv);
}
"""
)
TextureHelper._texture2d_sampler = self.ctx.sampler(
filter=(moderngl.LINEAR, moderngl.LINEAR),
)
def _init_depth_texture_draw(self):
"""Initialize geometry and shader for drawing FBO layers"""
from demosys import geometry
if not TextureHelper._quad:
TextureHelper._quad = geometry.quad_fs()
# Shader for drawing depth layers
TextureHelper._depth_shader = context.ctx().program(
vertex_shader="""
#version 330
in vec3 in_position;
in vec2 in_uv;
out vec2 uv;
uniform vec2 offset;
uniform vec2 scale;
void main() {
uv = in_uv;
gl_Position = vec4((in_position.xy + vec2(1.0, 1.0)) * scale + offset, 0.0, 1.0);
}
""",
fragment_shader="""
#version 330
out vec4 out_color;
in vec2 uv;
uniform sampler2D texture0;
uniform float near;
uniform float far;
void main() {
float z = texture(texture0, uv).r;
float d = (2.0 * near) / (far + near - z * (far - near));
out_color = vec4(d);
}
"""
)
TextureHelper._depth_sampler = self.ctx.sampler(
filter=(moderngl.LINEAR, moderngl.LINEAR),
compare_func='',
)
helper = TextureHelper()
| 5,106 | 1,579 |
class Meta(type):
def __new__(mcs, name, bases, namespace):
print(name)
if name == "A":
return type.__new__(mcs, name, bases, namespace)
if "attr_classe" in namespace:
print(f"{name} tentou sobrescrever o atributo attr_classe")
del namespace["attr_classe"] # excluindo attr_classe da classe B
print(namespace)
if "b_fala" not in namespace:
print(f"você precisa criar o metodo de fala em {name}")
else:
if not callable(namespace["b_fala"]):
print(f"b_fala precisa ser um metodo, não atributo em {name}")
return type.__new__(mcs, name, bases, namespace)
class A(metaclass=Meta):
def fala(self):
self.b_fala()
attr_classe = "valor A" # para não ser sobrescrito estou tratando na metaclasse
class B(A):
# b_fala = "olá"
def b_fala(self):
print("oi")
attr_classe = "valor B"
b = B()
b.b_fala()
print(b.attr_classe)
C = type("C", (), {"attr": "olá Mundo!"}) #nome da classe, de quem ela está herdando e o que tem nela.
c = C()
print(c.attr)
print(type(c))
| 1,136 | 407 |
from utils import *
from bot import telegram_chatbot
from bulb import *
bot = telegram_chatbot(CONFIG_LOCATION)
print('Initialized Bot')
bulb = bulb(IP_RANGE)
print('Connected to bulb. IP address: {}'.format(bulb.address))
while True:
updates = bot.get_updates(offset=update_id)
updates = updates["result"]
if updates:
for item in updates:
update_id = item["update_id"]
from_ = item["message"]["from"]["id"]
try:
message_type = item['message']['entities'][0]['type']
message = item['message']['text']
except:
message_type = None
message = None
if message_type=='bot_command':
if message=='/lighton':
# Turn light on
bulb.toggle('ON')
reply = 'Lights have been turned on'
elif message=='/lightoff':
# Turn light off
bulb.toggle('OFF')
reply = 'Lights have been turned off'
elif message=='/getstatus':
# display status of light
status = bulb.getStatus()
if status==True:
reply = 'Lights are on.'
else:
reply = 'Lights are off.'
else:
reply = 'This is not a valid bot command. Please reach out to the developer for assistance.'
bot.send_message(reply, from_)
print(item)
else:
reply = 'Input is not a valid bot command. Please retry'
bot.send_message(reply, from_)
| 1,787 | 478 |
# Futu Algo: Algorithmic High-Frequency Trading Framework
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# Written by Bill Chan <billpwchan@hotmail.com>, 2021
# Copyright (c) billpwchan - All Rights Reserved
# the first step is always the same: import all necessary components:
import smtplib
import ssl
from datetime import datetime
from email.mime.multipart import MIMEMultipart
from email.mime.text import MIMEText
from socket import gaierror
from util import logger
from util.global_vars import *
class Email:
def __init__(self):
"""
Email Engine Constructor
"""
self.config = config
self.port = self.config['Email'].get('Port')
self.smtp_server = self.config['Email'].get('SmtpServer')
self.sender = self.config['Email'].get('Sender')
self.login = self.config['Email'].get('Login')
self.password = self.config['Email'].get('Password')
# Create a secure SSL context
self.context = ssl.create_default_context()
self.default_logger = logger.get_logger("email")
def write_daily_stock_filter_email(self, receiver: str, filter_name: str, message_content: dict):
message = MIMEMultipart("alternative")
message["Subject"] = f"Daily Selected Stock List - {datetime.today().strftime('%Y-%m-%d')} - {filter_name}"
message["From"] = self.sender
message["To"] = receiver
text = "Please kindly review today's chosen stock list! "
html = """\
<style>
* {
font-family: sans-serif; /* Change your font family */
}
.content-table {
border-collapse: collapse;
margin: 25px 0;
font-size: 0.9em;
min-width: 400px;
border-radius: 5px 5px 0 0;
overflow: hidden;
box-shadow: 0 0 20px rgba(0, 0, 0, 0.15);
}
.content-table thead tr {
background-color: #009879;
color: #ffffff;
text-align: left;
font-weight: bold;
}
.content-table th,
.content-table td {
padding: 12px 15px;
}
.content-table tbody tr {
border-bottom: 1px solid #dddddd;
}
.content-table tbody tr:nth-of-type(even) {
background-color: #f3f3f3;
}
.content-table tbody tr:last-of-type {
border-bottom: 2px solid #009879;
}
.content-table tbody tr.active-row {
font-weight: bold;
color: #009879;
}
</style>
<table class="content-table">
<thead>
<tr>
<th>Stock Code</th>
<th>Company Name</th>
<th>Last Close</th>
<th>Day's Range</th>
<th>Market Cap</th>
<th>Beta (5Y Monthly)</th>
<th>PE (Trailing/Forward)</th>
<th>EPS (Trailing/Forward)</th>
<th>Volume</th>
</tr>
</thead>
<tbody>\n
"""
for equity, values in message_content.items():
html += f"""\
<tr>
<td>{equity}</td>
<td>{values['longName']}</td>
<td>{values['previousClose']}</td>
<td>{values['dayRange']}</td>
<td>{values['marketCap']}</td>
<td>{values['beta']}</td>
<td>{values['PE(Trailing/Forward)']}</td>
<td>{values['EPS(Trailing/Forward)']}</td>
<td>{values['volume']}</td>
</tr>\n
"""
html += """\
</tbody>
</table>
"""
# Turn these into plain/html MIMEText objects
part1 = MIMEText(text, "plain")
part2 = MIMEText(html, "html")
# Add HTML/plain-text parts to MIMEMultipart message
# The email client will try to render the last part first
message.attach(part1)
message.attach(part2)
try:
# send your message with credentials specified above
with smtplib.SMTP(self.smtp_server, self.port) as server:
server.starttls(context=self.context) # Secure the connection
server.login(self.login, self.password)
server.sendmail(self.sender, receiver, message.as_string())
self.default_logger.info(f'Email Sent: {receiver}')
except (gaierror, ConnectionRefusedError):
self.default_logger.info('Failed to connect to the server. Bad connection settings?')
except smtplib.SMTPServerDisconnected:
self.default_logger.info('Failed to connect to the server. Wrong user/password?')
except smtplib.SMTPException as e:
self.default_logger.info('SMTP error occurred: ' + str(e))
| 5,365 | 1,603 |
from os import system, name
system('cls' if name == 'nt' else 'clear')
dsc = ('''DESAFIO 066:
Crie um programa que leia vários números inteiros pelo teclado. O
programa só vai parar quando o usuário digitar o valor 999, que é a
condição de parada. No final, mostre quantos números foram
digitados e qual foi a soma entre eles (desconsiderando o flag).
''')
t = 0
c = 0
n = 0
while True:
n = int(input('Digite o número a ser somado: '))
if n == 999:
break
c += 1
t += n
print('você digitou {} números e a soma é {}'.format(c, t))
| 547 | 215 |
import string
import random
import redis
alpha = string.ascii_uppercase
l = []
while len(l) < 100:
res = ''
for i in range(16):
a = random.choice(alpha)
n = str(random.randrange(10))
rand = random.choice([a, n])
res += rand
if res not in l:
l.append(res)
# print(res)
print(len(l))
print(l)
r = redis.StrictRedis(host='localhost', port=6379, db=0)
for item in l:
r.set(item, True)
print("Showing data from redis:")
print(r.keys())
| 502 | 192 |
from setuptools import setup, find_packages
with open('requirements.txt') as f:
requirements = f.read()
setup(
name='World-Manager-CLI',
version='0.1.0',
packages=find_packages(),
include_package_data=True,
install_requires=requirements,
entry_points="""
[console_scripts]
world-manager=cli.cli:cli
""",
) | 355 | 116 |
import socket
import ure as re
import time
import machine
def run():
# Standard socket stuff:
host = ''
port = 80
sock = socket.socket(socket.AF_INET, socket.SOCK_STREAM)
sock.bind((host, port))
sock.listen(1) # don't queue up any requests
while True:
csock, caddr = sock.accept()
print("\nConnection from: " + str(caddr))
req = csock.recv(1024) # get the request, 1kB max
get_req = str(req).split('GET /')[1].split('HTTP')[0]
print('Req RAW:')
print(req)
output = parse_req(get_req)
csock.sendall("""HTTP/1.0 200 OK
Content-Type: text/html
<html>
<head>
</head>
<body>
<form action="" method="get">
<button name="pin1" value="True">P1-On</button>
</form>
<form action="" method="get">
<button name="pin1" value="False">P1-Off</button>
</form>
<br>
<form action="" method="get">
<button name="pin2" value="True">P2-On</button>
</form>
<form action="" method="get">
<button name="pin2" value="False">P2-Off</button>
</form>
<br>
OUTPUT:
{0}
</body>
</html>
""".format(str(output)))
csock.close()
def parse_req(get_req):
print('Get Req:')
print(get_req)
if 'favicon.ico' not in get_req:
get_req = get_req[1:]
data = get_req.split('=')
print(data)
return pin_logic(data)
def pin_logic(data):
import machine
if 'pin1' in data[0]:
machine.Pin(5, machine.Pin.OUT).on() if 'True' in data[1] else machine.Pin(5, machine.Pin.OUT).off()
if 'pin2' in data[0]:
machine.Pin(2, machine.Pin.OUT).on() if 'True' in data[1] else machine.Pin(2, machine.Pin.OUT).off()
try:
run()
except:
time.sleep(3)
machine.reset()
| 1,886 | 677 |
#!/usr/bin/env python
###############################################################################
#
# This script is to be used to restore and start pnfs from live backup
#
#
# $Id$
###############################################################################
import getopt
import os
import pwd
import re
import string
import subprocess
import sys
import time
import types
import configuration_client
import enstore_constants
import enstore_functions2
import e_errors
import file_utils
sys2host={'cms': ('cmspnfs1', 'psql-data'),
'd0en': ('d0ensrv1n', 'psql-data'),
'stken': ('stkensrv1n', 'psql-data'),
'cdfen': ('cdfensrv1n', 'psql-data'),
'dmsen': ('dmsen01', 'psql-data'),
}
PSQL_COMMAND = "psql -U enstore postgres -c 'select datname from pg_database' 2> /dev/null"
def copy(source,destination):
try:
s = open(source,"r")
d = open(destination,"w")
for l in s:
d.write(l)
return 0
except Exception, msg:
sys.stderr.write("Got exception %s copying %s to %s"%(str(msg),source,destination,))
sys.stderr.flush()
return 1
class PnfsSetup:
# find uncommented lines that look like
# " var = /value "
# there could be arbitrary number of white spaces between words
#
MATCH=re.compile("^[\s]*[^#]+[\s]*[\w]+[\s]*=[\s]*[\w\-/~]+")
# default pnfsSetup file name
NAME="pnfsSetup"
# default destination directory
DEFAULT_LOCATION="/usr/etc"
# mandatory keys:
REMOTE_BACKUP="remotebackup"
TRASH="trash"
DATABASE_POSTGRES="database_postgres"
DATABASE="database"
PNFS="pnfs"
def __init__(self,fname):
f=open(fname,"r")
self.contents={}
self.pnfs_dir = "/opt/pnfs"
for line in f:
if not line : continue
if PnfsSetup.MATCH.match(line):
data = line.split("=")
try:
ls=data[0].strip()
rs=string.join(data[1:],"=").strip()
if ls==PnfsSetup.REMOTE_BACKUP:
ls="#"+ls
self.remote_backup_host, self.remote_backup_dir = rs.split("#")[0].strip().split(":")
self.contents[ls]=rs
except Exception, msg:
sys.stderr.write("Got exception %s parsing line %s"%(str(msg),line))
sys.stderr.flush()
pass
f.close()
pgdb = self.contents[PnfsSetup.DATABASE_POSTGRES].split("#")[0].strip().rstrip("/")
self.backup_name = os.path.basename(pgdb.rstrip("/"))
def __repr__(self):
s=""
for k, v in self.contents.iteritems():
s+="%s=%s\n"%(k,v)
return s
def __str__(self):
return self.__repr__()
def write(self,directory=None):
if not directory :
directory = PnfsSetup.DEFAULT_LOCATION
f=open(os.path.join(directory,PnfsSetup.NAME),"w")
f.write(str(self))
f.close()
def __setitem__(self,key,value):
if self.contents.has_key(key):
self.contents[key]=value
else:
raise KeyError, "No such key %s"%(key,)
def __getitem__(self,key):
return self.contents[key]
def checkPostgres():
psqlFile = os.popen(PSQL_COMMAND)
psqlOutput = psqlFile.readlines()
psqlFile.close()
if not psqlOutput:
#sys.stderr.write('Postgres server is not running properly yet\n')
return False
return True
def getPostgresqlVersion():
ret = enstore_functions2.shell_command2("psql --version")
if not ret:
return None
rc = ret[0]
if rc:
return None
out = ret[1]
out = string.join(re.sub("[^0-9.]+","",out).split('.')[:2],'.')
return out
class PnfsDbRestore:
def __init__(self):
self.config = configuration_client.get_config_dict()
self.systems = self.config.get('known_config_servers',{})
self.pnfs_host = None
def get_configuration_client(self):
return self.config
def get_enstore_systems(self):
list = []
for s in sys2host.keys() :
list.append(s)
for s in self.systems.keys():
if s == "status" : continue
list.append(s)
return list
def usage(self,cmd):
print "Usage: %s -s [--system=] -t [backup_time=] "%(cmd,)
print "\t allowed systems: ", self.get_enstore_systems()
print "\t specify timestamp YYYY-MM-DD to get backup up to certain date"
def recover(self,name,backup_time=None):
postgresqlVersion = getPostgresqlVersion()
if not postgresqlVersion:
print "Failed to determine postgresql version"
return 1
pnfsSetupFile=os.path.join(os.getenv("ENSTORE_DIR"),"etc/%s-pnfsSetup"%(name))
if sys2host.has_key(name):
self.pnfs_host = sys2host[name][0]
for s in self.systems:
if s != name : continue
server_name,server_port = self.systems.get(s)
config_server_client = configuration_client.ConfigurationClient((server_name, server_port))
pnfs_server_dict=config_server_client.get('pnfs_server')
if pnfs_server_dict["status"][0] == e_errors.KEYERROR:
self.pnfs_host = sys2host[name][0]
elif pnfs_server_dict["status"][0] ==e_errors.OK:
self.pnfs_host=config_server_client.get('pnfs_server').get('host')
else:
print "Failed to communicate with config server ",pnfs_server_dict["status"]
return 1
#
# read configuration file
#
pnfsSetup = PnfsSetup(pnfsSetupFile)
#
# write modified configuration file in place (/usr/etc/pnfsSetup)
#
pnfsSetup.write()
for cmd in ["umount -f /pnfs/fs", \
"/sbin/service postgresql-%s stop"%(postgresqlVersion,)]:
print "Executing command ",cmd
rc=os.system(cmd)
if rc != 0 :
sys.stderr.write("Failed to execute command '%s' ignoring\n"%(cmd,))
sys.stderr.flush()
backup_file = get_backup(pnfsSetup)
pnfs_db = pnfsSetup[pnfsSetup.DATABASE]
if not os.path.exists(pnfs_db):
os.makedirs(pnfs_db)
else:
parent = pnfs_db
for i in range(2):
parent = os.path.dirname(parent)
os.rename(parent,"%s-%s.%d"%(parent,
time.strftime("%b-%d-%Y",time.localtime()),
os.getpid()))
os.makedirs(pnfs_db)
os.chdir(os.path.dirname(pnfs_db))
cwd=os.getcwd()
for d in ['%s/log'%(os.path.dirname(cwd),),\
pnfsSetup[pnfsSetup.TRASH]]:
if os.path.exists(d):
file_utils.rmdir(d)
print "Creating directory ",d
os.makedirs(d)
# copy backup file over
cmd = "/usr/bin/rsync -e rsh %s:%s . "%(pnfsSetup.remote_backup_host,
backup_file)
print 'copying: %s'% (cmd,)
err=os.system(cmd)
if err:
raise RuntimeError, '%s failed w/ exit code %d' % (cmd, err)
cmd='tar xzvf %s --preserve-permissions --same-owner'%(os.path.basename(backup_file),)
print 'Extracting %s with: %s'% (os.path.basename(backup_file), cmd)
os.system(cmd)
# fill in the database location in sysconfig file
psql_data=pnfsSetup[PnfsSetup.DATABASE_POSTGRES]
f=open("/etc/sysconfig/pgsql/postgresql-%s"%(postgresqlVersion,),"w")
f.write("PGDATA=%s\n"%(psql_data))
f.close()
# create recovery.conf
rdir = '%s:%s.xlogs'% (pnfsSetup.remote_backup_host,
os.path.dirname(backup_file))
cmd = "restore_command = '%s/sbin/enrcp %s/"% (os.getenv("ENSTORE_DIR"),
rdir) + "%f.Z %p.Z" + " && uncompress %p.Z'"
pgdb = pnfsSetup[PnfsSetup.DATABASE_POSTGRES]
print 'Creating recovery.conf: %s'% (cmd, )
f=open('%s/recovery.conf'%(pgdb,), 'w')
f.write('%s\n'%(cmd,))
if (backup_time != None) :
f.write("recovery_target_time='%s'\n"%(backup_time,))
f.write("recovery_target_inclusive='true'\n")
f.close()
os.system("cat %s/recovery.conf"%(pgdb))
for cmd in ['sed -i "s/archive_command/#archive_command/g" %s/postgresql.conf '% (pgdb,), \
'sed -i "s/^[ \t\r]*archive_mode[ \t\r]*=[ \t\r]*on/archive_mode = off/g" %s/postgresql.conf '% (pgdb,),\
'sed -i "s/^[ \t\r]*logging_collector[ \t\r]*=[ \t\r]*on/logging_collector = off/g" %s/postgresql.conf '% (pgdb,),\
"mkdir -p %s/pg_xlog/archive_status"% (pgdb,),\
"chown -R enstore.enstore %s/pg_xlog"% (pgdb,),\
"chown enstore.enstore %s/recovery.conf"%(pgdb)),\
"rm -f %s/postmaster.pid"%(pgdb),\
"/sbin/service postgresql-%s start"%(postgresqlVersion,)]:
rc = os.system(cmd)
if rc != 0 :
sys.stderr.write("Command %s failed, bailing out "%(cmd,))
sys.stderr.flush()
return rc
rc = checkPostgres()
print "Starting DB server"
while not rc:
print "Waiting for DB server"
time.sleep(60)
rc = checkPostgres()
print "DB server is ready"
cmd='dcache start'
print "Starting chimera: %s"% (cmd,)
rc=os.system(cmd)
if rc != 0 :
sys.stderr.write("Command %s failed, bailing out "%(cmd,))
sys.stderr.flush()
return rc
print "DONE"
return 0
def get_command_output(command):
child = os.popen(command)
data = child.read()
err = child.close()
if err:
raise RuntimeError, '%s failed w/ exit code %d' % (command, err)
return data[:-1] # (skip '\n' at the end)
#
# get next to the last backup
#
def get_backup(pnfsSetup):
cmd='rsh %s "ls -t %s/*.Z | grep -v xlog|head -n 2|tail -1"'%(pnfsSetup.remote_backup_host,
pnfsSetup.remote_backup_dir)
return get_command_output(cmd)
if __name__ == "__main__" :
r = PnfsDbRestore();
uname = pwd.getpwuid(os.getuid())[0]
backup_time = None
sysname = None
emergency = False
ourhost = string.split(os.uname()[1],'.')[0]
if uname != 'root':
print "Must be 'root' to run this program"
sys.exit(1)
try:
opts, args = getopt.getopt(sys.argv[1:], "hs:t:es:", ["help","system=","backup_time=","emergency"])
except getopt.GetoptError:
print "Failed to process arguments"
r.usage(sys.argv[0])
sys.exit(2)
for o, a in opts:
if o in ("-h", "--emergency") :
emergency=True
if o in ("-h", "--help"):
r.usage(sys.argv[0])
sys.exit(1)
if o in ("-s", "--system"):
sysname = a
if o in ("-t", "--backup_time"):
backup_time = a
for value in sys2host.values():
if ourhost == value[0] and emergency == False :
print "You are running on production system - ",ourhost
print "Re-run specifying --emergency switch to proceed"
r.usage(sys.argv[0])
sys.exit(1)
if (sysname == None or sysname=="") :
print "Error: Must specify enstore system name"
r.usage(sys.argv[0])
sys.exit(1)
else :
if sysname not in r.get_enstore_systems() and not sys2host.has_key(sysname):
print "Error: Unknown system specified ", sysname
print "Known systems : ", r.get_enstore_systems()
r.usage(sys.argv[0])
sys.exit(1)
if (backup_time == None or backup_time=="" ) :
backup_time=None
rc = r.recover(sysname,backup_time)
sys.exit(rc)
| 11,943 | 4,048 |
class Params:
################################
# dataset
################################
DATA_DIR = '../data/'
DATA_TRAIN_TITLE = 'train/train_title.npy'
DATA_TRAIN_BODY = 'train/train_body.npy'
DATA_TRAIN_LABEL = 'train/train_label.npy'
DATA_DEV_TITLE = 'dev/dev_title.npy'
DATA_DEV_BODY = 'dev/dev_body.npy'
DATA_DEV_LABEL = 'dev/dev_label.npy'
DATA_TEST_TITLE_BODY = 'test/data_para_test.pkl'
DATA_TEST_LABEL = 'test/test_label.npy'
DATA_DEBUG_TITLE_BODY = 'debug/data_para_debug.pkl'
VOCA_FILE_NAME = 'dic_mincutN.pkl'
GLOVE_FILE_NAME = 'W_embedding.npy'
################################
# train
################################
till_max_epoch = False
num_till_max_epoch = 8
CAL_ACCURACY_FROM = 0
MAX_EARLY_STOP_COUNT = 10
EPOCH_PER_VALID_FREQ = 0.3
is_embeddign_train = True # True is better
dr_text_in = 0.3 # 0.3 naacl-18
dr_text_out = 1.0
dr_con_in = 1.0 # 1.0 naacl-18
dr_con_out = 1.0
################################
# model
################################
reverse_bw = True
is_text_encoding_bidir = False
is_chunk_encoding_bidir = True
is_text_residual = False
is_chunk_residual = False
add_attention = True
add_LTC = False
LTC_topic_size = 3
LTC_memory_dim = 256
LTC_dr_prob = 0.8
################################
# etc
################################
IS_DEBUG = False # use short dataset
| 1,744 | 672 |
# encoding: utf-8
from baselayer.app.env import load_env
import subprocess
import sys
import time
import os
from pathlib import Path
env, cfg = load_env()
bundle = Path(os.path.dirname(__file__))/'../../static/build/bundle.js'
def run(cmd):
print("开始了")
p = subprocess.Popen(cmd, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
for line in p.stdout:
print(f'[service/webpack] {line.decode()}', end="")
sys.stdout.flush()
return p
if env.debug:
print("[service/webpack]: debug mode detected, launching webpack monitor")
p = run(['./node_modules/.bin/webpack', '--watch'])
sys.exit(p.returncode)
elif bundle.is_file():
print("[service/webpack]: bundle.js already built, exiting")
# Run for a few seconds so that supervisor knows the service was
# successful
time.sleep(3)
sys.exit(0)
else:
print("[service/webpack]: bundle.js not found, building")
p = run(['./node_modules/.bin/webpack'])
time.sleep(1)
sys.exit(p.returncode)
| 1,011 | 335 |
import imageio
import pytest
from subsurface.reader.read_netcdf import read_unstruct
import json
try:
import geopandas as gpd
GEOPANDAS_IMPORTED = True
except ImportError:
GEOPANDAS_IMPORTED = False
import pytest
import numpy as np
from subsurface import UnstructuredData, TriSurf, StructuredData
from subsurface.reader.profiles.profiles_core import create_mesh_from_trace
from subsurface.visualization import to_pyvista_mesh, pv_plot, \
to_pyvista_mesh_and_texture
@pytest.fixture(scope='module')
def unstruct(data_path):
us = read_unstruct(data_path + '/interpolator_meshes.nc')
return us
@pytest.fixture(scope='module')
def wells(data_path):
us = read_unstruct(data_path + '/wells.nc')
return us
def test_wells_to_binary(wells):
bytearray_le, header = wells.to_binary()
print(header)
with open('well_f.json', 'w') as outfile:
json.dump(header, outfile)
new_file = open("wells_f.le", "wb")
new_file.write(bytearray_le)
@pytest.mark.skipif(GEOPANDAS_IMPORTED is False, reason="Geopandas is not imported " )
def test_profile_to_binary(data_path):
traces = gpd.read_file(data_path + '/profiles/Traces.shp')
v, e = create_mesh_from_trace(traces.loc[0, 'geometry'], traces.loc[0, 'zmax'],
traces.loc[0, 'zmin'])
unstruct_temp = UnstructuredData.from_array(v, e)
cross = imageio.imread(data_path + '/profiles/Profil1_cropped.png')
struct = StructuredData.from_numpy(np.array(cross))
texture_binary, texture_header = struct.to_binary()
origin = [traces.loc[0, 'geometry'].xy[0][0],
traces.loc[0, 'geometry'].xy[1][0],
int(traces.loc[0, 'zmin'])]
point_u = [traces.loc[0, 'geometry'].xy[0][-1],
traces.loc[0, 'geometry'].xy[1][-1],
int(traces.loc[0, 'zmin'])]
point_v = [traces.loc[0, 'geometry'].xy[0][0],
traces.loc[0, 'geometry'].xy[1][0],
int(traces.loc[0, 'zmax'])]
texture_header['texture_origin'] = origin
texture_header['texture_point_u'] = point_u
texture_header['texture_point_v'] = point_v
ts = TriSurf(
mesh=unstruct_temp,
texture=struct,
texture_origin=origin,
texture_point_u=point_u,
texture_point_v=point_v
)
_, uv = to_pyvista_mesh_and_texture(ts)
import pandas as pd
unstruct = UnstructuredData.from_array(v, e, vertex_attr=pd.DataFrame(uv, columns=['u', 'v']))
mesh_binary, mesh_header = unstruct.to_binary()
with open('mesh_uv.json', 'w') as outfile:
import json
json.dump(mesh_header, outfile)
with open('texture.json', 'w') as outfile:
json.dump(texture_header, outfile)
new_file = open("mesh_uv_f.le", "wb")
new_file.write(mesh_binary)
new_file = open("texture_f.le", "wb")
new_file.write(texture_binary)
return mesh_binary
| 2,911 | 1,072 |
#!/usr/bin/env python
import sys
if len(sys.argv) < 3 or not sys.argv[2]:
print('/echo nick required.')
sys.exit()
nick = sys.argv[2]
print('/ctcp {} VERSION'.format(nick))
| 185 | 73 |
# Copyright 2021 Secureworks
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import base64
import logging
from os import urandom
from uuid import uuid4
from typing import Dict
from xml.etree import ElementTree
from ticketsplease.modules.adfs.envelope.utils import (
NAMESPACES,
send_envelope,
get_psha1,
derive_wstrustkey,
decrypt_wstrust_cipherdata,
create_soap_envelope,
)
class SCT_ENVELOPE:
def _create_sct_envelope(
self,
key: bytes,
clientSecret: bytes,
context: bytes,
keyIdentifier: bytes,
server: str,
):
"""Build a SCT enevlope.
Arguments:
key: security key from parsed RSTR
clientSecret: generated random bytes
context: security context from parsed RSTR
keyIdentifier: key identifier from parsed RSTR
server: ip_address|hostname of ADFS server
Returns:
SCT envelope
"""
# https://github.com/Gerenios/AADInternals/blob/master/ADFS_utils.ps1#L627
payload = f'<t:RequestSecurityToken xmlns:t="http://schemas.xmlsoap.org/ws/2005/02/trust"><t:TokenType>http://schemas.xmlsoap.org/ws/2005/02/sc/sct</t:TokenType><t:RequestType>http://schemas.xmlsoap.org/ws/2005/02/trust/Issue</t:RequestType><t:Entropy><t:BinarySecret Type="http://schemas.xmlsoap.org/ws/2005/02/trust/Nonce" u:Id="uuid-{uuid4()}">{base64.b64encode(clientSecret).decode()}</t:BinarySecret></t:Entropy><t:KeySize>256</t:KeySize></t:RequestSecurityToken>'
action = "http://schemas.xmlsoap.org/ws/2005/02/trust/RST/SCT"
envelope = create_soap_envelope(
key,
context,
keyIdentifier,
server,
payload,
action,
)
return envelope
def _parse_sct_envelope(
self,
envelope: bytes,
key: bytes,
clientSecret: bytes,
) -> str:
"""Parse the SCT response envelope.
Arguments:
envelope: SCT response envelope
cipher: KRB_TGT cipher object
sessionKey: KRB_TGT session key object
Returns:
parsed SCT envelope (context, key, key identifier)
"""
try:
tree = ElementTree.fromstring(envelope)
# https://github.com/Gerenios/AADInternals/blob/master/ADFS_utils.ps1#L709
# nonce0 = tree.findall(".//c:DerivedKeyToken", NAMESPACES["c"])[0][3].text
# cipher0 = tree.findall(".//e:EncryptedData", NAMESPACES["e"])[0][2][0].text
nonce1 = base64.b64decode(
tree.findall(".//c:DerivedKeyToken", NAMESPACES["c"])[1][1].text
)
cipher1 = base64.b64decode(
tree.findall(".//e:EncryptedData", NAMESPACES["e"])[1][2][0].text
)
except Exception as e:
logging.error(str(e))
raise TypeError("server responded with malformed SCT envelope") from e
derivedKey = derive_wstrustkey(key, nonce1, 32)
logging.debug(f"\tNonce: {base64.b64encode(nonce1)}")
logging.debug(f"\tDerived key: {base64.b64encode(derivedKey)}")
logging.info("\tDecrypting WSTrust Cipher Text")
# https://github.com/Gerenios/AADInternals/blob/master/ADFS_utils.ps1#L727
# Decrypt the cipher data
bPlainText = decrypt_wstrust_cipherdata(cipher1, derivedKey)
logging.debug(f"\tDecrypted SCT Data:\n{bPlainText.decode().strip()}\n")
# https://github.com/Gerenios/AADInternals/blob/master/ADFS_utils.ps1#L651
# Now parse the decrypted data from the outter SCT envelope
try:
tree = ElementTree.fromstring(bPlainText)
except Exception as e:
logging.error(str(e))
logging.error(f"invalid xml:\n{bPlainText}")
raise TypeError("failed to parse decrypted SCT envelope data") from e
token = tree.find(".//t:BinarySecret", NAMESPACES["t"]).text
# https://github.com/Gerenios/AADInternals/blob/master/ADFS_utils.ps1#L653
serverSecret = base64.b64decode(token)
computedKey = get_psha1(clientSecret, serverSecret, 32)
# fmt: off
# https://github.com/Gerenios/AADInternals/blob/c255cd66a3731c32cfbdf9fdb17f2b03c7665b72/ADFS_utils.ps1#L656
context = tree.find(".//t:RequestedSecurityToken", NAMESPACES["t"])[0]
context = context.attrib["{%s}Id" % NAMESPACES["u"]["u"]]
keyIdentifier = tree.find(".//t:RequestedSecurityToken", NAMESPACES["t"])[0][0].text.split(":")[2]
# fmt: on
logging.debug(f"\tServer secret: {base64.b64encode(serverSecret)}")
logging.debug(f"\tComputed key: {base64.b64encode(computedKey)}")
logging.debug(f"\tContext: {context}")
logging.debug(f"\tIdentifier: {keyIdentifier}")
# https://github.com/Gerenios/AADInternals/blob/master/ADFS_utils.ps1#L665
# Construct the return value
retVal = {
"Context": context,
"Key": computedKey,
"Identifier": keyIdentifier,
}
return retVal
@classmethod
def run(
cls,
adfs_host: str,
rstr: Dict[str, bytes],
):
"""Generate and send an SCT envelope to the target ADFS server.
Receive the SCT response and parse the message for the context,
key, and key identifier.
Arguments:
adfs_host: target ADFS server
rsts: parsed RST response object
Returns:
dictionary of parsed SCT response data (context, key,
key identifier)
"""
logging.info(f"[ * ] Building and sending SCT envelope to the ADFS server")
clientSecret = urandom(32)
# Build the SCT envelope to request the configuration
sct_envelope = cls._create_sct_envelope(
cls,
rstr["Key"],
clientSecret,
rstr["Context"],
rstr["Identifier"],
adfs_host,
)
logging.debug(f"\tSCT Envelope:\n{sct_envelope.strip()}")
# Send the SCT envelope
response = send_envelope(adfs_host, sct_envelope)
logging.debug(f"\tRST Response Status: {response}")
logging.debug(f"\tRST Response:\n{response.content}")
if response.status_code == 200:
logging.info(f"[ * ] Parsing SCT envelope response")
sct_data = cls._parse_sct_envelope(
cls,
response.content,
rstr["Key"],
clientSecret,
)
else:
raise ValueError(f"Bad response from ADFS server: {response.status_code}")
return sct_data
| 7,243 | 2,326 |
"""Geometry testing."""
import creopyson
from .fixtures import mk_creoson_post_dict, mk_creoson_post_None, mk_getactivefile
def test_geometry_bound_box(mk_creoson_post_dict, mk_getactivefile):
"""Test bound_box."""
c = creopyson.Client()
result = c.geometry_bound_box(file_="file")
assert isinstance(result, (dict))
result = c.geometry_bound_box()
assert isinstance(result, (dict))
def test_geometry_get_edges(mk_creoson_post_dict, mk_getactivefile):
"""Test get_edges."""
c = creopyson.Client()
result = c.geometry_get_edges(["12", "34"], file_="file")
assert isinstance(result, (list))
result = c.geometry_get_edges(["12", "34"])
assert isinstance(result, (list))
def test_geometry_get_surfaces(mk_creoson_post_dict, mk_getactivefile):
"""Test get_surfaces."""
c = creopyson.Client()
result = c.geometry_get_surfaces(file_="file")
assert isinstance(result, (list))
result = c.geometry_get_surfaces()
assert isinstance(result, (list))
| 1,014 | 367 |
#print prime factorisation of a number
n= int(input("enter the number \n"))
def prime(t, j):
if(j<t):
if(t%j!=0):
return prime(t, j=j + 1)
else:
return 0
else:
return 1
i=2
while(n>1):
if (prime(i, 2) == 1):
while n % i == 0:
print(i, ",",end="")
n=n/i
i+=1
| 356 | 142 |
import os
from backports.tempfile import TemporaryDirectory
from django.conf import settings
import pytest
from awx.api.versioning import reverse
@pytest.mark.django_db
class TestInsightsCredential:
def test_insights_credential(self, patch, insights_project, admin_user, insights_credential):
patch(insights_project.get_absolute_url(),
{'credential': insights_credential.id}, admin_user,
expect=200)
def test_non_insights_credential(self, patch, insights_project, admin_user, scm_credential):
patch(insights_project.get_absolute_url(),
{'credential': scm_credential.id}, admin_user,
expect=400)
@pytest.mark.django_db
def test_project_custom_virtualenv(get, patch, project, admin):
with TemporaryDirectory(dir=settings.BASE_VENV_PATH) as temp_dir:
os.makedirs(os.path.join(temp_dir, 'bin', 'activate'))
url = reverse('api:project_detail', kwargs={'pk': project.id})
patch(url, {'custom_virtualenv': temp_dir}, user=admin, expect=200)
assert get(url, user=admin).data['custom_virtualenv'] == os.path.join(temp_dir, '')
@pytest.mark.django_db
def test_project_invalid_custom_virtualenv(get, patch, project, admin):
url = reverse('api:project_detail', kwargs={'pk': project.id})
resp = patch(url, {'custom_virtualenv': '/foo/bar'}, user=admin, expect=400)
assert resp.data['custom_virtualenv'] == [
'/foo/bar is not a valid virtualenv in {}'.format(settings.BASE_VENV_PATH)
]
@pytest.mark.django_db
@pytest.mark.parametrize('value', ["", None])
def test_project_unset_custom_virtualenv(get, patch, project, admin, value):
url = reverse('api:project_detail', kwargs={'pk': project.id})
resp = patch(url, {'custom_virtualenv': value}, user=admin, expect=200)
assert resp.data['custom_virtualenv'] is None
| 1,865 | 609 |
'''
58-Melhore o jogo do desafio 028 onde o computador vai "pensar" em um numero
entre 0 e 10.Só que agora vai tentar advinhar até acertar mostrando no final
quantos palpites foram necessários até vencer.
'''
import random
palpite=0
print('\033[33m{:=^40}'.format('JOGO DA ADVINHAÇÂO 2.0'))
print('\033[m')
numerosorteado=random.randint(0,10)
print(numerosorteado)
acertou=False
while not acertou:
numero=int(input('Digite o valor entre 0 e 10: '))
palpite+=1
if numero == numerosorteado:
acertou=True
else:
if numero < numerosorteado:
print('É maior.Tente mais uma vez!')
elif numero > numerosorteado:
print('É menos.Tente mais uma vez!')
print('Você acertou com {} palpites.'.format(palpite))
| 761 | 293 |
# -*- coding: utf-8 -*-
"""
bandwidth
This file was automatically generated by APIMATIC v2.0 ( https://apimatic.io ).
"""
class DisconnectCauseEnum(object):
"""Implementation of the 'DisconnectCause' enum.
TODO: type enum description here.
Attributes:
BUSY: TODO: type description here.
CALLBACKERROR: TODO: type description here.
CANCEL: TODO: type description here.
ERROR: TODO: type description here.
HANGUP: TODO: type description here.
INVALIDBXML: TODO: type description here.
REJECTED: TODO: type description here.
TIMEOUT: TODO: type description here.
ACCOUNTLIMIT: TODO: type description here.
NODECAPACITYEXCEEDED: TODO: type description here.
UNKNOWN: TODO: type description here.
APPLICATIONERROR: TODO: type description here.
"""
BUSY = 'busy'
CALLBACKERROR = 'callback-error'
CANCEL = 'cancel'
ERROR = 'error'
HANGUP = 'hangup'
INVALIDBXML = 'invalid-bxml'
REJECTED = 'rejected'
TIMEOUT = 'timeout'
ACCOUNTLIMIT = 'account-limit'
NODECAPACITYEXCEEDED = 'node-capacity-exceeded'
UNKNOWN = 'unknown'
APPLICATIONERROR = 'application-error'
| 1,285 | 443 |
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import display
from pymongo import MongoClient
def get_database():
from pymongo import MongoClient
CONNECTION_STRING = "#"
from pymongo import MongoClient
client = MongoClient(CONNECTION_STRING) #conexão com o cliente
return client["socioeconomico"] #base de dados
dbname = get_database()
collection_name = dbname["venezuela2021"]
detalhes_itens = collection_name.find()
# consulta o db no Mongo, coloca todos os dados do database nessa variavel.
df = pd.DataFrame(list(detalhes_itens)) #criei um df com o banco de dados
perfil = df[["gender", "age", "geography", "financial_situation"]] # criando um df apenas com as chaves interessantes para o perfil
#########################################################################################
###Criar gráfico que mostra a quantidade de pessoas entrevistadas em cada faixa etária
def faixaetaria(perfil):
age_qtd = (perfil["age"]).value_counts() # crio uma variavel em que quantifica cada faixa etaria obtida na pesquisa
print(age_qtd) # exemplo: 1304 pessoas tem idade de 26 a 35 anos
plt.style.use("ggplot")
age_qtd.plot.barh() # defino o tipo de grafico
plt.title("Número de pessoas entrevistadas por faixa etária") # adiciona titulo
plt.xlabel("Número de pessoas") # nomeia eixo x
plt.ylabel("Faixa etária") # nomeia eixo y
plt.show() # exibe o grafico
###Criar grafico com gênero e idade no formato barras
def generoidade(df):
perfil2 = df[["gender", "age"]]
print(perfil2)
plt.style.use("ggplot")
graf = (perfil2).value_counts()
print(graf)
graf.plot.barh()
plt.title("Número de pessoas entrevistadas por gênero e faixa etária") # adiciona titulo
plt.xlabel("Número de pessoas") # nomeia eixo x
plt.ylabel("Gênero e Faixa etária") # nomeia eixo y
plt.show() # exibe o grafico
###Criar grafico de Gênero no formato Pizza:
def genero(perfil):
gen_qtd = (perfil["gender"]).value_counts() # crio uma variavel em que quantifica cada genero obtido na pesquisa
print(gen_qtd)
df1 = gen_qtd.iloc[[2,3,4]]
df2=gen_qtd.drop(gen_qtd.index[[2,3,4]]) #aqui estou eliminando essas linhas para colocar o resultado da soma em uma linha só
print(df2) #só tem genero female e male
df2.loc['Others: Non-Binary, Non Available, Prefer not to Answer'] = sum(df1) # acrescento uma nova linha com index others e o valor da soma.
plt.style.use("ggplot")
df2.plot.pie(ylabel='',autopct='%1.1f%%',startangle = 90) # defino o tipo de grafico
plt.title("Número de pessoas entrevistadas por gênero") # adiciona titulo
plt.show() # exibe o grafico
###Criar gráfico das situações financeiras da pessoas entrevistadas pela pesquisa
def sitfin(perfil):
sitfin = (perfil["financial_situation"]).value_counts() # crio uma variavel em que quantifica a sit financeira de cada pessoa da pesquisa
print(sitfin) # exemplo: 1445 só conseguem custear comida e nada mais
plt.style.use("ggplot")
sitfin.plot.pie(autopct = "%1.1f%%", ylabel='')
plt.title("Situação financeira das pessoas entrevistadas") # adiciona titulo
plt.show() # exibe o grafico
###Criar gráfico que mostra a quantidade de pessoas entrevistadas por cada região em que vivem
def geografia(perfil):
geography = (perfil["geography"]).value_counts() # crio uma variavel em que quantifica cada faixa etaria obtida na pesquisa
print(geography) # exemplo: 1304 pessoas tem idade de 26 a 35 anos
plt.style.use("ggplot")
geography.plot.barh(color = "lightsalmon") # defino o tipo de grafico
plt.title("Número de pessoas entrevistadas por região em que vivem") # adiciona titulo
plt.xlabel("Número de pessoas") # nomeia eixo x
plt.ylabel("Região") # nomeia eixo y
plt.show() # exibe o grafico
###Criar um grafico em que mostra a relação entre a região e as pessoas que são muito vulneraveis financeiramente
def relregiaositfin(df):
perfil3 = df[["geography", "financial_situation"]] #cria dataframe com as chaves de interesse
aux = perfil3[(perfil3['financial_situation'] == 'I cannot afford enough food for my family')] #crio uma variavel em que recebe a sit fin desejada
print(aux.groupby('geography').count()) #relaciona a sit financeira desejada com a geografia e faz a contagem do num de pessoas.
graf1 = aux.groupby('geography').count() #crio uma variavel que relaciona a auxiliar (sit fin) com a geografia e quantifica
graf1.plot()
plt.title("Região em que vivem as pessoas que não conseguem comprar comida suficiente para a família") #Não consigo comprar comida suficiente para a minha família.
plt.ylabel("Número de pessoas")
plt.xlabel("Geografia")
plt.show()
def favoVulne(df):
#perfil de pessoa na situação mais confortavel, universidade/faculdade/pos graduação completa ou nao e criança com acesso a internet.
df_fvvn = df[['_id', 'financial_situation', 'education', 'do_children_have_internet_connection']]
docTotais = 4436
favo1 = len(df_fvvn[(df_fvvn['financial_situation'] == "I can comfortably afford food, clothes, and furniture, and I have savings") & (df_fvvn['education'] == "University or college degree completed") & (df_fvvn['do_children_have_internet_connection'] == '1')])
favoPorcent1 = (favo1 * 100) / docTotais
favo2 = len(df_fvvn[(df_fvvn['financial_situation'] == "I can comfortably afford food, clothes, and furniture, and I have savings") & (df_fvvn['education'] == "Some university or college") & (df_fvvn['do_children_have_internet_connection'] == '1')])
favoPorcent2 = (favo2 * 100) / docTotais
favo3 = len(df_fvvn[(df_fvvn['financial_situation'] == "I can comfortably afford food, clothes, and furniture, and I have savings") & (df_fvvn['education'] == "Post-graduate education") & (df_fvvn['do_children_have_internet_connection'] == '1')])
favoPorcent3 = (favo3 * 100) / docTotais
favo4 = len(df_fvvn[(df_fvvn['financial_situation'] == "I can comfortably afford food, clothes, and furniture, and I have savings") & (df_fvvn['education'] == "Post graduate") & (df_fvvn['do_children_have_internet_connection'] == '1')])
favoPorcent4 = (favo4 * 100) / docTotais
pessoasFavoravel = favo1 + favo2 + favo3 + favo4
pessoasFavoravelPorc = favoPorcent1 + favoPorcent2 + favoPorcent3 + favoPorcent4
print(f"{pessoasFavoravel} documentos apontaram que tem condições financeiras confortaveis, alto nivel educacional e criança com acesso a internet \nIsso representa {pessoasFavoravelPorc} % da amostra total\n")
#perfil de pessoa na situação mais vulneravel, baixo nivel educacional e criança sem acesso a internet
vulne1 = len(df_fvvn[(df_fvvn['financial_situation'] == "I cannot afford enough food for my family") & (df_fvvn['education'] == "No formal education") & (df_fvvn['do_children_have_internet_connection'] == '0')])
vulnePorcent1 = (vulne1 * 100) / docTotais
vulne2 = len(df_fvvn[(df_fvvn['financial_situation'] == "I cannot afford enough food for my family") & (df_fvvn['education'] == "Some primary education") & (df_fvvn['do_children_have_internet_connection'] == '0')])
vulnePorcent2 = (vulne2 * 100) / docTotais
vulne3 = len(df_fvvn[(df_fvvn['financial_situation'] == "I cannot afford enough food for my family") & (df_fvvn['education'] == "Primary school completed") & (df_fvvn['do_children_have_internet_connection'] == '0')])
vulnePorcent3 = (vulne3 * 100) / docTotais
pessoasVulneraveis = vulne1 + vulne2 + vulne3
pessoasVulneraveisPorc = vulnePorcent1 + vulnePorcent2 + vulnePorcent3
print(f"{pessoasVulneraveis} documentos apontaram que não tem condições de custear alimentação suficiente, tem baixo nivel educacional e criança sem acesso a internet \nIsso representa {pessoasVulneraveisPorc} % da amostra total\n")
grupos = ['Condição Mais \n Favorável', 'Condição Menos \n Favorável']
valores = [pessoasFavoravel, pessoasVulneraveis]
plt.title('OS DOIS PERFIS EXTREMOS')
plt.ylabel('Numero de formularios')
plt.bar(grupos, valores)
plt.show()
def desfavoravel(df):
df_vul = df[['_id', 'financial_situation', 'education', 'do_children_have_internet_connection']]
docTotais = 4436
semAlimentacao = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family")])
#perfil de pessoa na situação mais vulneravel, universidade/faculdade/pos graduação completa ou nao e criança sem acesso a internet
alto1 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "University or college degree completed") & (df_vul['do_children_have_internet_connection'] == '0')])
altoPorcent1 = (alto1 * 100) / docTotais
alto2 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Some university or college") & (df_vul['do_children_have_internet_connection'] == '0')])
altoPorcent2 = (alto2 * 100) / docTotais
alto3 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Post-graduate education") & (df_vul['do_children_have_internet_connection'] == '0')])
altoPorcent3 = (alto3 * 100) / docTotais
alto4 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Post graduate") & (df_vul['do_children_have_internet_connection'] == '0')])
altoPorcent4 = (alto4 * 100) / docTotais
pessoasEducAlta = alto1 + alto2 + alto3 + alto4
educAlta = altoPorcent1 + altoPorcent2 + altoPorcent3 + altoPorcent4
print(f"{pessoasEducAlta} documentos apontaram não ter condições de custear alimentação suficiente, tem alto nivel educacional e criança sem acesso a internet \nIsso representa {educAlta} % da amostra total\n")
#perfil de pessoa na situação mais vulneravel, educação secundaria e criança sem acesso a internet
med1 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Secondary school/ high school completed") & (df_vul['do_children_have_internet_connection'] == '0')])
medPorcent1 = (med1 * 100) / docTotais
med2 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Some secondary school / high school") & (df_vul['do_children_have_internet_connection'] == '0')])
medPorcent2 = (med2 * 100) / docTotais
med3 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Secondary/high school") & (df_vul['do_children_have_internet_connection'] == '0')])
medPorcent3 = (med3 * 100) / docTotais
pessoasEducMedia = med1 + med2 + med3
educMedia = medPorcent1 + medPorcent2 + medPorcent3
print(f"{pessoasEducMedia} documentos apontaram não ter condições de custear alimentação suficiente, tem medio nivel educacional e criança sem acesso a internet \nIsso representa {educMedia} % da amostra total\n")
#perfil de pessoa na situação mais vulneravel, educação tecnica completa ou nao (agrupadas) e criança sem acesso a internet
tec1 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Technical school diploma or degree completed") & (df_vul['do_children_have_internet_connection'] == '0')])
tecPorcent1 = (tec1 * 100) / docTotais
tec2 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Some technical education (e.g polytechnic school") & (df_vul['do_children_have_internet_connection'] == '0')])
tecPorcent2 = (tec2 * 100) / docTotais
tec3 = len(df_vul[(df_vul['financial_situation'] == "I cannot afford enough food for my family") & (df_vul['education'] == "Technical school") & (df_vul['do_children_have_internet_connection'] == '0')])
tecPorcent3 = (tec3 * 100) / docTotais
pessoasEducTecnica = tec1 + tec2 + tec3
educTecnica = tecPorcent1 + tecPorcent2 + tecPorcent3
print(f"{pessoasEducTecnica} documentos apontaram não ter condições de custear alimentação suficiente para a familia, tem nivel educacional técnico e criança sem acesso a internet \nIsso representa {educTecnica} % da amostra total\n")
grupos = ['Não conseguem \nCustear alimentação', 'Ensino Superior', 'Ensino Médio', 'Ensino Tecnico']
valores = [semAlimentacao, pessoasEducAlta, pessoasEducMedia, pessoasEducTecnica]
plt.title('RELAÇÃO VULNERABILIDADE X NIVEL EDUCACIONAL')
plt.ylabel('Numero de formularios')
plt.bar(grupos, valores)
plt.show()
def intAcess1(df):
#se a criança tem acesso a internet e tem energia eletrica consistentes, se perde aula. se nao tem acesso, esta com aula presencial
df_vul = df[['_id', 'do_children_have_internet_connection', 'does_home_shows_severe_deficit_of_electricity', 'does_home_shows_severe_deficit_of_internet', 'do_children_3_to_17_yrs_miss_virtual_class_due_to_lack_of_electricity', 'are_children_attending_face_to_face_classes', 'are_children_being_teached_by_unqualified_people']]
docTotais = 4436
perfil1 = len(df_vul[(df_vul['does_home_shows_severe_deficit_of_electricity'] == '0') & (df_vul['does_home_shows_severe_deficit_of_internet'] == '0') & (df_vul['do_children_have_internet_connection'] == '1') & (df_vul['do_children_3_to_17_yrs_miss_virtual_class_due_to_lack_of_electricity'] == '0')])
porcentagem1 = (perfil1 * 100) / docTotais
print(f"{perfil1} documentos apontaram que há crianças sem problemas de conexão com internet ou falta de energia eletrica e não perdem aulas por estes motivos.\nIsso representa {porcentagem1} % da amostra total\n")
perfil2 = len(df_vul[(df_vul['does_home_shows_severe_deficit_of_electricity'] == '1') | (df_vul['does_home_shows_severe_deficit_of_internet'] == '1') & (df_vul['do_children_have_internet_connection'] == '1') & (df_vul['do_children_3_to_17_yrs_miss_virtual_class_due_to_lack_of_electricity'] == '1')])
porcentagem2 = (perfil2 * 100) / docTotais
print(f"{perfil2} documentos apontaram que há crianças com problemas de conexão com internet ou falta de energia eletrica e perdem aulas por estes motivos.\nIsso representa {porcentagem2} % da amostra total\n")
perfil3 = len(df_vul[(df_vul['are_children_attending_face_to_face_classes'] == '1') | (df_vul['does_home_shows_severe_deficit_of_internet'] == '1') & (df_vul['do_children_have_internet_connection'] == '0')])
porcentagem3 = (perfil3 * 100) / docTotais
print(f"{perfil3} documentos apontaram que há crianças sem acesso a internet ou tem problemas de conexão e estão tendo aulas presenciais.\nIsso representa {porcentagem3} % da amostra total\n")
perfil4 = len(df_vul[(df_vul['are_children_attending_face_to_face_classes'] == '0') & (df_vul['does_home_shows_severe_deficit_of_internet'] == '1') & (df_vul['do_children_have_internet_connection'] == '0')])
porcentagem4 = (perfil4 * 100) / docTotais
print(f"{perfil4} documentos apontaram que há crianças sem acesso a internet ou tem problemas de conexão e não estão tendo aulas presenciais.\nIsso representa {porcentagem4} % da amostra total\n")
perfil5 = len(df_vul[(df_vul['are_children_attending_face_to_face_classes'] == '0') & (df_vul['does_home_shows_severe_deficit_of_internet'] == '1') & (df_vul['do_children_have_internet_connection'] == '0') & (df_vul['are_children_being_teached_by_unqualified_people'] == '1')])
porcentagem5 = (perfil5 * 100) / docTotais
print(f"{perfil5} documentos apontaram que há crianças sem acesso a internet ou tem problemas de conexão e não estão tendo aulas presenciais \n e estão sendo ensinadas por pessoas sem qualificação.Isso representa {porcentagem5} % da amostra total\n")
grupos = ['Não perdem \naula virtual', 'Problemas técnicos \nPerdem aula virtual', 'Sem acesso \nAula presencial', 'Sem aula virtual\n nem presencial', 'Aula com pessoas\n não qualificadas']
valores = [perfil1, perfil2, perfil3, perfil4, perfil5]
plt.title('RELAÇÃO ACESSOS A INTERNET E ENERGIA x AULA VIRTUAL/PRESENCIAL')
plt.ylabel(' Numero de formularios')
plt.bar(grupos, valores)
plt.show()
def inseg(df):#Grafico barra alimentação
data=df[["financial_situation","do_children_3_and_17_yrs_receive_regular_school_meals"]]
aux = data[(data['do_children_3_and_17_yrs_receive_regular_school_meals'] == "No")]
graf = aux.groupby('financial_situation').count()
graf.plot.barh()
plt.title("Insegurança alimentar x Situação Financeira")
plt.ylabel("")
L=plt.legend(bbox_to_anchor=(1.1,1.1),\
bbox_transform=plt.gcf().transFigure)
L.get_texts()[0].set_text('Crianças que recebem comida na escola')
plt.savefig('temp.png')
plt.show()
def evesao(df): #Grafico barra para evasão escolar
data=df[["education","were_children_3_to_17_yrs_enrolled_and_did_not_return_to_school"]]
aux = data[(data['were_children_3_to_17_yrs_enrolled_and_did_not_return_to_school'] == "0")]
graf = aux.groupby('education').count()
graf.plot.barh()
plt.title("Evasão escolar x nível educacional do responsável")
plt.ylabel("")
L=plt.legend(bbox_to_anchor=(1.1,1.1),\
bbox_transform=plt.gcf().transFigure)
L.get_texts()[0].set_text('Crianças que não retornaram a escola')
plt.savefig('temp.png')
plt.show()
#retorna um gráfico do grau de escolaridade das pessoas que responderam o questionário
def educacao(df):
df_new = df[['education']]
#destaca a coluna com o maior valor
explode = (0.1, 0, 0, 0, 0, 0, 0, 0)
colors = ['#FFFF00', '#800080','#B22222','#483D8B','#FA8072','#CD853F','#2E8B57', '#FF4500']
labels = ['Graduação em faculdade completa','Segundo grau (Ensino médio) completo', 'Diploma de escola técnica ou algum título completo','Possui alguma educação universitária','Possui alguma educação técnica','Possui alguma educação secundária/ensino médio', 'Pós-graduação completa','Outros']
#gráfico de pizza da educação
graf = (df_new["education"]).value_counts()
# autopct = rotular as fatias com seu valor numérico
# shadow = sombra
graf2 = graf
soma = sum(graf2.iloc[[12, 13, 14, 15, 8, 11, 10, 7, 9]])
graf2 = graf2.drop(graf2.index[[12, 13, 14, 15, 8, 11, 10, 7, 9]])
graf2.loc['Others'] = soma
graf2.plot.pie(autopct='%1.1f%%', explode= explode, shadow=True, startangle = 90, labels=labels, ylabel='', colors = colors)
print(graf2)
plt.title('Educação na Venezuela')
plt.show()
#retorna um gráfico da situação financeira em relação a educação de pessoas que possuem algum diploma
def financial_situation_education(df):
colunas = ['financial_situation', 'education']
df_new = df.filter(items = colunas)
aux = df_new[(df_new['education'] == 'University or college degree completed') | (df_new['education'] == 'Secondary school/ high school completed') | (df_new['education'] == 'Technical school diploma or degree completed')]
graf = aux.groupby('financial_situation').count()
graf.plot.pie(autopct='%1.1f%%', shadow=True, startangle = 90, subplots=True, ylabel='')
L = plt.legend(bbox_to_anchor=(1.9, 1.1))
print(graf)
plt.savefig('temp.png')
plt.title('Situação finaceira X Educação')
plt.show()
#Chamando as funções
#faixaetaria(perfil) #gráfico que mostra a quantidade de pessoas entrevistadas em cada faixa etária
#generoidade(df) #grafico que exibe gênero e idade das pessoas entrevistadas no formato barras
#genero(perfil) #grafico que mostra a qtd de pessoas por gênero no formato Pizza
#sitfin(perfil) #gráfico das situações financeiras das pessoas entrevistadas na pesquisa
#geografia(perfil) #gráfico que mostra a quantidade de pessoas entrevistadas por cada região em que vivem
#relregiaositfin(df) #grafico mostra a relação entre a região e as pessoas que são vulneraveis financeiramente
#favoVulne(df) # apresenta perfis opostos: mais condições desfavoraveis e mais condições favoraveis
#intAcess1(df) #Relaciona acesso a internet e modalidade de aulas
#desfavoravel(df) # apresenta relação de pessoas que nao conseguem custear alimentação e nivel educacional alto e medio
#inseg(df) #grafico que compara a sitação financeira com alimentação na escola
#evesao(df) #grafico que compara evesão escolar com o nivel educacional do responsável
#educacao(df) #mostra o gráfico da educação
#financial_situation_education(df) #mostra o gráfico da situação financeira e educação
| 20,727 | 7,867 |
import os
import json
from tqdm import tqdm
def build_vocabulary(data_dir):
with open(os.path.join(data_dir, 'train_texts.txt'), 'rt') as fin:
train_texts = json.load(fin)
print("train text cuts load done")
with open(os.path.join(data_dir, "Telegram", 'train_key_words.dat'), 'rb') as fin:
train_key_words = pickle.load(fin)
print("train key_words load done")
words = set()
for train_text in tqdm(train_texts,miniters=1000):
for word in train_text:
words.add(word)
for key_word in tqdm(train_key_words,miniters=1000):
for word in key_word:
words.add(word)
with open(os.path.join(data_dir, "Telegram", "words.dat"), 'wb') as fout:
pickle.dump(words, fout)
print("Build Vocabulary Done!!!")
def get_word_embedding(data_home, word2vec_name):
with open(os.path.join(data_home, "Telegram", "words.dat"), 'rb') as fin:
words = pickle.load(fin)
telegram_word_embeddings = dict()
print("The number of words is {}".format(len(words)))
word2vec_path = os.path.join(data_home, "word_embedding", word2vec_name)
with open(word2vec_path, 'rt') as fin:
line = fin.readline()
words_num, embed_size = line.split()
print("The number of words is {}, the embedding size is {}".format(words_num, embed_size))
for line in tqdm(fin, miniters=5000):
word, embed = line.split(maxsplit=1)
if word in words:
try:
telegram_word_embeddings[word] = [float(vec) for vec in embed.split()]
except Exception as e:
print(e)
print(line)
vocab_size = len(telegram_word_embeddings)
with open(os.path.join(data_home, "word_embedding", "telegram_word_embedding.dat"), 'wb') as fout:
pickle.dump(telegram_word_embeddings, fout)
print("done!!!")
if __name__ == "__main__":
data_dir = r'/home/yaojq/data/text/reuters'
word2vec_path = "/home/yaojq/data/word_embedding/GoogleNews-vectors-negative300.bin"
print("build vocabulary")
build_vocabulary(data_dir)
get_word_embedding(data_dir, word2vec_path)
| 2,235 | 814 |
import requests
import json
import hashlib
import os
import argparse
def load_links(filename):
with open(filename) as f:
data = json.load(f)
return data
def get_story(session, uri):
""" Get mementos for html using Internet Archive """
try:
print(uri)
r = session.get(uri, verify=False)
# return entire response
return r
except Exception as e:
print("Failed with error", e)
return
if __name__ == "__main__":
# months to download stories from
months = ["2016_12", "2017_01"]
parser = argparse.ArgumentParser()
# parser.add_argument("links_json", type=str,
# help="Links per day JSON file to iterate upon")
parser.add_argument("--kval", type=str,
help="Links per day JSON file to iterate upon")
args = parser.parse_args()
session = requests.Session()
session.headers = headers = {
'user-agent': 'Web Science and Digital Libraries (@WebSciDL) '
'<gatki001@odu.edu>'}
session.max_redirects = 100
for mo in months:
print("Month {}".format(mo))
links_by_day = load_links(
"data/links_per_day/{}/links_per_day_{}.json".format(mo,
args.kval))
error_file = "data/errors/links_{}.txt".format(mo)
with open(error_file, 'w') as err_out:
for day in links_by_day:
links = links_by_day[day]
print("Day {}".format(day))
for uri in links:
directory = "./data/stories/if_/{}/{}/".format(mo, day)
link_hash = hashlib.md5(uri.encode()).hexdigest()
outfile = directory + link_hash + ".html"
if not os.path.exists(directory):
os.makedirs(directory)
if os.path.exists(outfile):
continue
resp = get_story(session, uri)
if not resp:
print("Error with response:", resp)
print("{}\nError with response: {}".format(
uri, resp), file=err_out)
continue
if resp.status_code == 200:
with open(outfile, "w") as out:
out.write(resp.text)
else:
print(resp.history)
print("ERR::{} response code".format(resp.status_code))
print("{}\nError with response code: {}".format(
uri, resp.status_code), file=err_out)
if os.path.getsize(error_file) == 0:
os.remove(error_file)
| 2,797 | 778 |
#!/usr/bin/env python
# -*- coding: utf-8 -*-
from __future__ import print_function
import os
import argparse
import rosbag
import rospy
from cv_bridge import CvBridge
import cv2
import play_images_in_rosbag
def decide_output_encoding(cv_img):
"""
For 16UC1 input image, we need to use mono16 as output encoding option.
see http://library.isr.ist.utl.pt/docs/roswiki/cv_bridge(2f)Tutorials(2f)
UsingCvBridgeToConvertBetweenROSImagesAndOpenCVImages.html
:param cv_img:
:return:
"""
coding = 'passthrough'
if cv_img.dtype == 'uint16':
coding = 'mono16'
return coding
def main():
parser = argparse.ArgumentParser(
description=("Downscale images and shift timestamps for sensor messages in "
"a ROS bag with topics '/cam0/image_raw', '/cam1/image_raw', '/imu0'."))
parser.add_argument("bag_file", help="Input ROS bag.")
parser.add_argument(
'--time_delay',
help="unit nanoseconds, time delay + original.header.stamp = "
"shifted.header.stamp. If not provided, time delay will set as "
"ros message time - message[0].header.stamp",
type=int,
default=None)
parser.add_argument("--out_bag_file",
help="Output ROS bag file.",
default=None)
args = parser.parse_args()
out_bag_file = args.out_bag_file
if args.out_bag_file is None:
out_bag_file = os.path.join(
os.path.splitext(args.bag_file)[0] + '_half.bag')
in_bag = rosbag.Bag(args.bag_file, "r")
out_bag = rosbag.Bag(out_bag_file, 'w')
time_shift = None
if args.time_delay is not None:
time_shift = rospy.Duration(args.time_delay // 1000000000,
args.time_delay % 1000000000)
print('Raw message time offset set to {}'.format(time_shift))
count = 0
for topic, msg, t in in_bag.read_messages(topics=['/imu0']):
if time_shift is None:
time_shift = t - msg.header.stamp
print('Raw message time offset set to {}'.format(time_shift))
msg.header.stamp = time_shift + msg.header.stamp
out_bag.write(topic, msg, msg.header.stamp)
count += 1
print('Saved {} messages on topic /imu0'.format(count))
bridge = CvBridge()
for k in range(2):
count = 0
image_topic = '/cam{}/image_raw'.format(k)
encoding = ''
for _, msg, t in in_bag.read_messages(topics=[image_topic]):
cv_img = bridge.imgmsg_to_cv2(msg, desired_encoding="passthrough")
h, w = cv_img.shape[:2]
cv_half_img = cv2.pyrDown(cv_img, dstsize=(w // 2, h // 2))
if count == 0:
print('Image info before and after half sampling:')
play_images_in_rosbag.print_image_info(cv_img)
play_images_in_rosbag.print_image_info(cv_half_img)
encoding = decide_output_encoding(cv_img)
cv2.imshow('Downsampled frame', cv_half_img)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
count += 1
rosimage = bridge.cv2_to_imgmsg(cv_half_img, encoding=encoding)
rosimage.header.stamp = time_shift + msg.header.stamp
out_bag.write(image_topic, rosimage, rosimage.header.stamp)
print('Saved {} images on topic {}'.format(count, image_topic))
cv2.destroyAllWindows()
out_bag.close()
in_bag.close()
print("Output bag: {}".format(out_bag_file))
if __name__ == '__main__':
main()
| 3,564 | 1,202 |
import json
import random
import sys
import time
sys.setrecursionlimit(2500)
memo = {}
def ks(capacity_left, n):
"""
capacity_left(int): remaining storage capacity of a bag
n(int): current item position
"""
if n == -1 or capacity_left == 0:
# No more items to add
return 0
# h = hash("%d_%d" % (capacity_left, n))
h = capacity_left * 2000 + n
if h in memo:
# print("memo", capacity_left, n)
return memo[h]
if weights[n] > capacity_left:
# Current item is too heavy for remaining capacity, ignore it and continue
return ks(capacity_left, n-1)
else:
# Do not add item, just move the pointer to the left
_without = ks(capacity_left, n-1)
# Add item into bag
_with = values[n] + ks(capacity_left-weights[n], n-1)
# Save value into memory
val = max(_with, _without)
memo[h] = val
return val
weights = []
values = []
capacities = []
bests = []
capacity = 0
for i in range(2001):
begin = time.time()
weights.append(random.randint(0, 100))
values.append(random.randint(0, 100))
capacity += random.randint(0, 25)
capacities.append(capacity)
best = ks(capacity, len(weights)-1)
bests.append(best)
memo = {}
end = time.time()
seconds = end - begin
print("Items", i)
# print(weights)
# print(values)
print("Capacity:", capacity)
print("Best:", best)
print("Seconds:", seconds)
print("*"*40)
with open("dataset.json", "w+") as f:
ds = {
"values": values,
"weights": weights,
"capacities": capacities,
"bests": bests
}
json.dump(ds, f, indent=4) | 1,770 | 616 |
''' CONFIDENTIAL
Copyright (c) 2021 Eugeniu Vezeteu,
Department of Remote Sensing and Photogrammetry,
Finnish Geospatial Research Institute (FGI), National Land Survey of Finland (NLS)
PERMISSION IS HEREBY LIMITED TO FGI'S INTERNAL USE ONLY. THE CODE
MAY BE RE-LICENSED, SHARED, OR TAKEN INTO OTHER USE ONLY WITH
A WRITTEN CONSENT FROM THE HEAD OF THE DEPARTMENT.
The software is provided "as is", without warranty of any kind, express or
implied, including but not limited to the warranties of merchantability,
fitness for a particular purpose and noninfringement. In no event shall the
authors or copyright holders be liable for any claim, damages or other
liability, whether in an action of contract, tort or otherwise, arising from,
out of or in connection with the software or the use or other dealings in the
software.
'''
import numpy as np
import cv2
import glob
import pickle
np.set_printoptions(suppress=True)
from sympy import *
class StereoChess_Calibrator(object):
def __init__(self, path):
self.term_criteria = (cv2.TERM_CRITERIA_EPS + cv2.TERM_CRITERIA_COUNT, 1000, 0.0001)
self.square = 0.1 # m (the size of each chessboard square is 10cm)
self.objp = np.zeros((10 * 7, 3), np.float32) #chessboard is 7x10
self.objp[:, :2] = np.mgrid[0:10, 0:7].T.reshape(-1, 2) * self.square
self.see = True
self.path = path
self.thermaImg, self.rgbImg, self.monoImg = [], [], []
self.axis = np.float32([[0,0,0], [9,0,0], [0,7,0], [0,0,-5]]).reshape(-1,3)*self.square
def draw(self, img, corners, imgpts):
corner = tuple(corners[0])
img = cv2.line(img, corner, tuple(imgpts[0]), (255, 0, 0), 5)
img = cv2.line(img, corner, tuple(imgpts[1]), (0, 255, 0), 5)
img = cv2.line(img, corner, tuple(imgpts[2]), (0, 0, 255), 5)
return img
def read_images(self):
'''
real all camera images (thermal, monochrome and rgb)
'''
thermal = glob.glob(self.path + '/themal_image_*.png')
rgb = glob.glob(self.path + '/rgb_image_*.png')
mono = glob.glob(self.path + '/monochrome_image_*.png')
thermal.sort()
rgb.sort()
mono.sort()
for i, fname in enumerate(thermal):
thermal_img = cv2.imread(thermal[i])
rgb_img = cv2.imread(rgb[i])
mono_img = cv2.imread(mono[i])
self.thermaImg.append(thermal_img)
self.rgbImg.append(rgb_img)
self.monoImg.append(mono_img)
self.thermaImg, self.rgbImg, self.monoImg = np.array(self.thermaImg), np.array(self.rgbImg), np.array(
self.monoImg)
print('read_images: thermaImg->{}, rgbImg->{}, monoImg->{} '.format(np.shape(self.thermaImg),
np.shape(self.rgbImg),
np.shape(self.monoImg)))
def read_points(self, camera=None): # camera in [mono,rgb,thermal]
'''
extract points from camera (thermal, monochrome and rgb)
'''
self.see = True
wait = 0
if camera == 'mono':
print('Mono camera calibration')
images = self.monoImg.copy()
elif camera == 'rgb':
print('RGB camera calibration')
images = self.rgbImg.copy()
elif camera == 'thermal':
print('Thermal camera calibration')
images = self.thermaImg.copy()
else:
print('Add right camera')
print('images -> {}'.format(np.shape(images)))
objpoints, imgpoints, img_shape = [], [], 0
# extract points
for i, fname in enumerate(images):
img = images[i]
if camera == 'thermal': # invert the thermal camera
img = np.array(256 - img, dtype='uint8')
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
ret, corners = cv2.findChessboardCorners(gray, (10, 7), flags=cv2.CALIB_CB_ADAPTIVE_THRESH)
if ret:
corners2 = cv2.cornerSubPix(gray, corners, (11, 11), (-1, -1), self.term_criteria)
cv2.drawChessboardCorners(img, (10, 7), corners2, ret)
objpoints.append(self.objp)
imgpoints.append(corners2)
# else:
# print('No board at {}'.format(i))
if self.see:
if camera == 'thermal':
cv2.imshow('Image', img)
else:
cv2.imshow('Image', cv2.resize(img, None, fx=.4, fy=.4))
k = cv2.waitKey(wait)
if k % 256 == 32: # pressed space
self.see = False
cv2.destroyAllWindows()
img_shape = gray.shape[::-1]
print('Camera {} objpoints->{},imgpoints->{}, img_shape->{}'.format(camera, np.shape(objpoints),
np.shape(imgpoints), img_shape))
return objpoints, imgpoints, img_shape
def calibrate(self, camera=None):
'''
perform internal calibration for given camera
'''
objpoints, imgpoints, img_shape = self.read_points(camera)
rms, K, D, _, _ = cv2.calibrateCamera(
objectPoints=objpoints,
imagePoints=imgpoints,
imageSize=img_shape,
cameraMatrix=None, distCoeffs=None,
flags=0, criteria=self.term_criteria)
print('{} camera calibration done with RMS:{}'.format(camera, rms))
print('K')
print(K)
print('D')
print(D)
return K, D
def stereoCalibrate(self, K_thermal, D_thermal,K,D, camera): # camera in [rgb,thermal]
'''
perform stereo calibration between thermal camera and given camera (mono or rgb)
'''
objpoints = [] # 3d point in real world space
imgpoints_l = [] # 2d points in image plane. - thermal camera
imgpoints_r = [] # 2d points in image plane. - mono or rgb camera
if camera == 'mono':
Second_images = self.monoImg.copy()
elif camera == 'rgb':
Second_images = self.rgbImg.copy()
images = self.thermaImg.copy()
# extract points
for i, fname in enumerate(images):
thermal_img = np.array(256 - images[i], dtype='uint8')
thermal_gray = cv2.cvtColor(thermal_img, cv2.COLOR_BGR2GRAY)
self.img_shape = thermal_gray.shape[::-1]
thermal_ret, thermal_corners = cv2.findChessboardCorners(thermal_gray, (10, 7),
flags=cv2.CALIB_CB_ADAPTIVE_THRESH)
img = Second_images[i]
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
self.second_img_shape = gray.shape[::-1]
ret, corners = cv2.findChessboardCorners(gray, (10, 7), flags=cv2.CALIB_CB_ADAPTIVE_THRESH)
if thermal_ret and ret:
objpoints.append(self.objp)
imgpoints_l.append(thermal_corners)
imgpoints_r.append(corners)
print('Thermal -> {} cam, {}-poses'.format(camera, len(objpoints)))
flags = cv2.CALIB_FIX_INTRINSIC
rms_stereo, _, _, _, _, R, T, E, F = cv2.stereoCalibrate(
objpoints, imgpoints_l, imgpoints_r, K_thermal, D_thermal, K, D, imageSize=None, criteria=self.term_criteria, flags=flags)
print('Stereo calibraion Therma-{} done'.format(camera))
print('rms_stereo:{}'.format(rms_stereo))
print('Rotation R')
print(R)
print('Translation T')
print(T)
return R,T,E,F
def doStuff(self):
'''
-Read all images for all cameras
-Do internal calibration for each cam
-Estimate R rotation and T translation between thermal cam and mono cam
-Estimate R rotation and T translation between thermal cam and rgb cam
-Save the data
'''
#Read all images
self.read_images()
#Calibrate mono camera
K_mono, D_mono = calib.calibrate(camera='mono')
#Calibrate rgb camera
K_rgb, D_rgb = calib.calibrate(camera='rgb')
#Calibrate thermal camera
K_thermal, D_thermal = calib.calibrate(camera='thermal')
#Stereo calibrate between Thermal and Mono camera
R_th_mono, T_th_mono, E_th_mono, F_th_mono = self.stereoCalibrate(K_thermal,D_thermal,K_mono,D_mono,camera='mono')
# Stereo calibrate between Thermal and Rgb camera
R_th_rgb, T_th_rgb, E_th_rgb, F_th_rgb = self.stereoCalibrate(K_thermal, D_thermal, K_rgb, D_rgb, camera='rgb')
calib_data = dict([('K_mono', K_mono), ('D_mono', D_mono),
('K_rgb', K_rgb),('D_rgb', D_rgb),
('K_thermal', K_thermal), ('D_thermal', D_thermal),
('R_th_mono', R_th_mono), ('T_th_mono', T_th_mono),('E_th_mono', E_th_mono), ('F_th_mono', F_th_mono),
('R_th_rgb', R_th_rgb), ('T_th_rgb', T_th_rgb), ('E_th_rgb', E_th_rgb),('F_th_rgb', F_th_rgb),
])
with open('calib_data.pkl', 'wb') as f:
pickle.dump(calib_data, f, protocol=2)
print('calib_data.pkl Object saved')
def testCalibration(self):
'''
-loads images
-load the calibration data
-check if patter is visible in all 3 images:
-Estimate the extrinsic R,T from world to thermal camera
-Use estimated R,T and reproject pixels from thermal camera to mono and rgb cam
'''
self.thermaImg, self.rgbImg, self.monoImg = [], [], []
# Read all images
self.read_images()
with open('calib_data.pkl', 'rb') as f:
calib_data = pickle.load(f)
K_mono = calib_data['K_mono']
D_mono = calib_data['D_mono']
K_rgb = calib_data['K_rgb']
D_rgb = calib_data['D_rgb']
K_thermal = calib_data['K_thermal']
D_thermal = calib_data['D_thermal']
R_th_mono = calib_data['R_th_mono']
T_th_mono = calib_data['T_th_mono']
R_th_rgb = calib_data['R_th_rgb']
T_th_rgb = calib_data['T_th_rgb']
F = calib_data['F_th_rgb']
# Define test the calibration-----------------------
for i, fname in enumerate(self.thermaImg):
thermal_img = np.array(256 - self.thermaImg[i], dtype='uint8')
thermal_gray = cv2.cvtColor(thermal_img, cv2.COLOR_BGR2GRAY)
thermal_ret, thermal_corners = cv2.findChessboardCorners(thermal_gray, (10, 7),
flags=cv2.CALIB_CB_ADAPTIVE_THRESH)
mono_img = self.monoImg[i]
mono_gray = cv2.cvtColor(mono_img, cv2.COLOR_BGR2GRAY)
mono_ret, mono_corners = cv2.findChessboardCorners(mono_gray, (10, 7), flags=cv2.CALIB_CB_ADAPTIVE_THRESH)
rgb_img = self.rgbImg[i]
rgb_gray = cv2.cvtColor(rgb_img, cv2.COLOR_BGR2GRAY)
rgb_ret, _ = cv2.findChessboardCorners(rgb_gray, (10, 7), flags=cv2.CALIB_CB_ADAPTIVE_THRESH)
if thermal_ret and rgb_ret and mono_ret:
thermal_corners2 = cv2.cornerSubPix(thermal_gray, thermal_corners, (11, 11), (-1, -1),
self.term_criteria)
# Find the rotation and translation vectors.
ret, rvecs, tvecs = cv2.solvePnP(self.objp, thermal_corners2, K_thermal, D_thermal)
# project 3D points to thermal image plane
imgpts_thermal, jac = cv2.projectPoints(self.axis[1:], rvecs, tvecs, K_thermal,
D_thermal) # thermal camera frame
thermaImg = self.draw(thermal_img, np.asarray(thermal_corners2).squeeze(),
np.asarray(imgpts_thermal).squeeze())
T_01 = np.vstack(
(np.hstack((cv2.Rodrigues(rvecs)[0], tvecs)), [0, 0, 0, 1])) # from world to thermal camera
# project thermal to rgb --------------------------------------------------------------------------------------
T_12 = np.vstack((np.hstack((R_th_rgb, T_th_rgb)), [0, 0, 0, 1])) # from thermal cam to rgb cam
T = np.dot(T_12, T_01) # world to rgb cam
rotation, translation = T[:3, :3], T[:3, -1]
imgpts_rgb, _ = cv2.projectPoints(self.axis, rotation, translation, K_rgb, D_rgb)
imgpts_rgb = np.array(imgpts_rgb).squeeze()
rgbImg = self.draw(rgb_img, [imgpts_rgb[0]], imgpts_rgb[1:])
# project thermal to mono ------------------------------------------------------------------------------------
'''T_12 = np.vstack((np.hstack((R_th_mono, T_th_mono)), [0, 0, 0, 1])) # from thermal cam to mono cam
T = np.dot(T_12, T_01) # world to mono cam
rotation, translation = T[:3, :3], T[:3, -1]
imgpts_mono, _ = cv2.projectPoints(self.axis, rotation, translation, K_mono, D_mono)
imgpts_mono = np.array(imgpts_mono).squeeze()
monoImg = self.draw(mono_img, [imgpts_mono[0]], imgpts_mono[1:])'''
thermal_corners2 = np.array(thermal_corners2).squeeze()
x_1 = thermal_corners2[0] #pixel in thermal camera
x_1 = np.array([x_1[0],x_1[1],1])
print(x_1)
'''Z = 1
Z = tvecs[-1]
print('tvecs -> {}, Z:{}'.format(tvecs,Z))
x_1 = x_1*Z
X_cam1 = np.linalg.inv(K_thermal).dot(x_1)
X_cam1 = np.array([X_cam1[0],X_cam1[1],X_cam1[2],1])
print('X_cam1 -> {}'.format(X_cam1))
P = np.hstack((R_th_rgb, T_th_rgb)) # from thermal cam to rgb cam
print(P)
x_2 = K_rgb.dot(P) @ X_cam1
print('x_2 -> {}'.format(x_2))
x_2 = np.array([x_2[0]/x_2[-1],x_2[1]/x_2[-1]]).astype(int)
print('x_2 -> {}'.format(x_2))
print('rgbImg -> {}'.format(np.shape(rgbImg)))
cv2.circle(rgbImg, (x_2[0], x_2[1]), 12, (0, 255, 0), 12)
cv2.circle(thermaImg, (thermal_corners2[0][0], thermal_corners2[0][1]), 6, (0, 255, 0), 6)'''
print('F')
print(F)
#x_1 * F * x_2 = 0
x1 = np.asarray(thermaImg).reshape(-1,3)
x2 = np.asarray(rgbImg).reshape(-1,3)
print('x1:{}, F:{}, x2:{}'.format(np.shape(x1), np.shape(F),np.shape(x2)))
x1F = x1 @ F
print('x1 * F = {}'.format(np.shape(x1F)))
x1Fx2 = x1F.dot(x2.T)
print('x1Fx2= {}'.format(np.shape(x1Fx2)))
cv2.imshow('thermal_img', thermaImg)
#cv2.imshow('monoImg', cv2.resize(monoImg, None, fx=.4, fy=.4))
cv2.imshow('rgbImg', cv2.resize(rgbImg, None, fx=.3, fy=.3))
cv2.waitKey(0)
cv2.destroyAllWindows()
if __name__ == '__main__':
path = '/home/eugeniu/cool'
calib = StereoChess_Calibrator(path)
#calib.doStuff() #this function load the data, does internal and stereo calibration - > save the data
calib.testCalibration()
| 15,629 | 5,372 |
# Generated by Django 2.2.5 on 2020-03-02 17:17
from django.db import migrations, models
import employee_management.models
class Migration(migrations.Migration):
dependencies = [
('employee_management', '0014_auto_20200302_1716'),
]
operations = [
migrations.AlterField(
model_name='education',
name='files',
field=models.FileField(upload_to=employee_management.models.education_file_path),
),
]
| 477 | 164 |
# When your package is installed in editable mode, you can call
# instances of that package from any directory. For example, this
# script may be run by calling
#
# python scripts/say-hello-world.py
#
# and it will call methods inside our python_ml_template project.
from math_so.utils import say_hello_world
if __name__ == '__main__':
say_hello_world()
| 364 | 105 |
#!/usr/bin/env python
"""
.. See the NOTICE file distributed with this work for additional information
regarding copyright ownership.
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
"""
from __future__ import print_function
import argparse
from basic_modules.workflow import Workflow
from utils import logger
from CHiC.tool.makeDesignFiles import makeDesignFilesTool
#####################################################
class process_design(Workflow):
"""
This class generates the Design files and chinput files,
imput for CHiCAGO. Starting from rmap and baitmap and capture
HiC BAM files.
"""
def __init__(self, configuration=None):
"""
Initiate the class
Parameters
----------
Configuration: dict
dictionary with parameters for different tools from the class
indicating how to run each of them.
"""
logger.info("Generating CHiCAGO input Design files")
if configuration is None:
configuration = {}
self.configuration.update(configuration)
def run(self, input_files, metadata, output_files):
"""
Main function to run the tools, MakeDesignFiles_Tool.py and
bam2chicago_Tool.py
Parameters
----------
input_files: dict
designDir: path to the folder with .rmap and .baitmap files
rmapFile: path to the .rmap file
baitmapFile: path to the .baitmap file
bamFile: path to the capture HiC bamfiles
metadata: dict
input metadata
output_files: dict
outPrefixDesign : Path and name of the output prefix,
recommend to be the same as rmap and baitmap files.
sample_name: Path and name of the .chinput file
Returns
-------
bool
output_metadata
"""
try:
design_caller = makeDesignFilesTool(self.configuration)
design_out, design_meta = design_caller.run(
{
"RMAP" : input_files["RMAP"],
"BAITMAP": input_files["BAITMAP"]
},
{
"RMAP" : metadata["RMAP"],
"BAITMAP" : metadata["BAITMAP"]
},
{
"nbpb" : output_files["nbpb"],
"npb" : output_files["npb"],
"poe" : output_files["poe"]
}
)
logger.info("design files succesfully generated =)")
return design_out, design_meta
except IOError:
logger.fatal("process_makeDesign failed to" +
"generate design files")
#############################################################
def main_json(config, in_metadata, out_metadata):
"""
Alternative main function
This function lauch the app using the configuration written
in two json files:
"""
#1.Instantiate and launch the app
print("Instantiate and launch the App")
from apps.jsonapp import JSONApp
app = JSONApp()
results = app.launch(process_design,
config,
in_metadata,
out_metadata)
#2. The App has finished
print("2. Execution finished: see " + out_metadata)
print(results)
return results
#########################################################
if __name__ == "__main__":
#set up the command line parameters
PARSER = argparse.ArgumentParser(
description="Pipeline to generate .baitmap file")
PARSER.add_argument("--config", help="Configuration file")
PARSER.add_argument(
"--in_metadata", help="Location of metadata file")
PARSER.add_argument(
"--out_metadata", help="Location of output metadata file")
PARSER.add_argument(
"--local", action="store_const", const=True, default=False)
#Get matching parameters from the command line
ARGS = PARSER.parse_args()
CONFIG = ARGS.config
IN_METADATA = ARGS.in_metadata
OUT_METADATA = ARGS.out_metadata
LOCAL = ARGS.local
if LOCAL:
import sys
sys._run_from_cmdl = True # pylint: disable=protected-access
RESULTS = main_json(CONFIG, IN_METADATA, OUT_METADATA)
print(RESULTS)
| 4,851 | 1,308 |
from balrogo import gaia
# Read HDU
print("Reading input...")
path = r"./samples/SCULPTOR.fits"
# Run sample code
print("Running code...")
final_data, results_sd, var_sd, results_pm, var_pm = gaia.extract_object(path)
print("results_pm:", results_pm)
print("var_pm:", var_pm)
print("results_sd:", results_sd)
print("var_sd:", var_sd)
| 353 | 141 |
import asyncio
import base64
import hashlib
import logging
import re
import time
from typing import Optional
import http_ece
import ujson
from aiohttp import ClientSession
from arq import ArqRedis
from atoolbox import JsonErrors, RequestError
from cryptography.hazmat.backends import default_backend
from cryptography.hazmat.primitives.asymmetric import ec
from py_vapid import Vapid02 as Vapid
from pydantic import BaseModel, HttpUrl
from em2.core import get_flag_counts
from em2.settings import Settings
from em2.utils.db import Connections
logger = logging.getLogger('em2.web_push')
def web_push_user_key_prefix(user_id):
return f'web-push-subs:{user_id}:'
class SubscriptionModel(BaseModel):
"""
Model as generated from PushSubscription.toJSON()
https://developer.mozilla.org/en-US/docs/Web/API/PushSubscription/toJSON
"""
endpoint: HttpUrl
expirationTime: Optional[int]
class SubKeys(BaseModel):
p256dh: bytes
auth: bytes
keys: SubKeys
def hash(self):
return hashlib.md5(b'|'.join([self.endpoint.encode(), self.keys.p256dh, self.keys.auth])).hexdigest()
async def subscribe(conns: Connections, client_session: ClientSession, sub: SubscriptionModel, user_id):
key = web_push_user_key_prefix(user_id) + sub.hash()
# we could use expirationTime here, but it seems to generally be null
await conns.redis.setex(key, 86400, sub.json())
msg = await conns.main.fetchval(
"""
select json_build_object('user_v', v, 'user_id', id)
from users where id=$1
""",
user_id,
)
if not msg:
raise JsonErrors.HTTPUnauthorized('user not found')
await _sub_post(conns, client_session, sub, user_id, msg)
async def unsubscribe(conns: Connections, sub: SubscriptionModel, user_id):
key = web_push_user_key_prefix(user_id) + sub.hash()
await conns.redis.delete(key)
async def web_push(ctx, actions_data: str):
settings: Settings = ctx['settings']
if not settings.vapid_private_key or not settings.vapid_sub_email:
return 'web push not configured'
session: ClientSession = ctx['client_session']
data = ujson.loads(actions_data)
participants = data.pop('participants')
# hack to avoid building json for every user, remove the ending "}" so extra json can be appended
msg_json_chunk = ujson.dumps(data)[:-1]
coros = [_user_web_push(ctx, session, p, msg_json_chunk) for p in participants]
pushes = await asyncio.gather(*coros)
return sum(pushes)
async def _user_web_push(ctx, session: ClientSession, participant: dict, msg_json_chunk: str):
user_id = participant['user_id']
match = web_push_user_key_prefix(user_id) + '*'
subs = []
redis: ArqRedis = ctx['redis']
with await redis as conn:
cur = b'0'
while cur:
cur, keys = await conn.scan(cur, match=match)
for key in keys:
subs.append(await conn.get(key))
if subs:
async with ctx['pg'].acquire() as conn:
conns = Connections(conn, redis, ctx['settings'])
participant['flags'] = await get_flag_counts(conns, user_id)
msg = msg_json_chunk + ',' + ujson.dumps(participant)[1:]
subs = [SubscriptionModel(**ujson.loads(s)) for s in subs]
await asyncio.gather(*[_sub_post(conns, session, s, user_id, msg) for s in subs])
return len(subs)
else:
return 0
async def _sub_post(conns: Connections, session: ClientSession, sub: SubscriptionModel, user_id: int, msg: str):
body = http_ece.encrypt(
msg.encode(),
private_key=ec.generate_private_key(ec.SECP256R1, default_backend()),
dh=_prepare_vapid_key(sub.keys.p256dh),
auth_secret=_prepare_vapid_key(sub.keys.auth),
version=vapid_encoding,
)
async with session.post(sub.endpoint, data=body, headers=_vapid_headers(sub, conns.settings)) as r:
text = await r.text()
if r.status == 410:
await unsubscribe(conns, sub, user_id)
elif r.status == 403 and text == 'invalid JWT provided\n':
# seems to happen with https://fcm.googleapis.com/fcm/send/...
await unsubscribe(conns, sub, user_id)
elif r.status != 201:
logger.error(
f'unexpected response from webpush %s: %s',
r.status,
repr(text[:100]),
extra={'headers': dict(r.headers), 'text': text, 'url': sub.endpoint},
)
raise RequestError(r.status, sub.endpoint, text=text)
vapid_encoding = 'aes128gcm'
aud_re = re.compile('https?://[^/]+')
def _vapid_headers(sub: SubscriptionModel, settings: Settings):
vapid_claims = {
'aud': aud_re.match(sub.endpoint).group(0),
'sub': 'mailto:' + settings.vapid_sub_email,
'ext': int(time.time()) + 300,
}
return {
'ttl': '60',
'content-encoding': vapid_encoding,
**Vapid.from_string(private_key=settings.vapid_private_key).sign(vapid_claims),
}
def _prepare_vapid_key(data: bytes) -> bytes:
return base64.urlsafe_b64decode(data + b'===='[: len(data) % 4])
| 5,136 | 1,739 |