markdown stringlengths 0 1.02M | code stringlengths 0 832k | output stringlengths 0 1.02M | license stringlengths 3 36 | path stringlengths 6 265 | repo_name stringlengths 6 127 |
|---|---|---|---|---|---|
Performance Analysis One of the advantages of theano is the posibility to create a full profile of the function. This has to be included in at the time of the creation of the function. At the moment it should be active (the downside is larger compilation time and I think also a bit in the computation so be careful if you need a fast call) CPU The following profile is with a 2 core laptop. Nothing spectacular. Looking at the profile we can see that most of time is in pow operation (exponential). This probably is that the extent is huge and we are doing it with too much precision. I am working on it GPU | %%timeit
# Compute the block
GeMpy.compute_block_model(geo_data, [0,1,2], verbose = 0)
geo_data.interpolator._interpolate.profile.summary() | Function profiling
==================
Message: ../GeMpy/DataManagement.py:994
Time in 3 calls to Function.__call__: 8.400567e-01s
Time in Function.fn.__call__: 8.395956e-01s (99.945%)
Time in thunks: 8.275988e-01s (98.517%)
Total compile time: 3.540267e+00s
Number of Apply nodes: 342
Theano Optimizer time: 2.592782e+00s
Theano validate time: 1.640296e-01s
Theano Linker time (includes C, CUDA code generation/compiling): 8.665011e-01s
Import time 1.915064e-01s
Time in all call to theano.grad() 0.000000e+00s
Time since theano import 72.847s
Class
---
<% time> <sum %> <apply time> <time per call> <type> <#call> <#apply> <Class name>
57.3% 57.3% 0.474s 2.87e-03s C 165 55 theano.tensor.elemwise.Elemwise
10.1% 67.4% 0.084s 2.79e-03s C 30 10 theano.tensor.blas.Dot22Scalar
9.6% 77.0% 0.079s 9.81e-04s C 81 27 theano.sandbox.cuda.basic_ops.HostFromGpu
6.4% 83.4% 0.053s 8.89e-03s Py 6 2 theano.tensor.basic.Nonzero
6.4% 89.8% 0.053s 1.77e-02s Py 3 1 theano.tensor.nlinalg.MatrixInverse
5.1% 95.0% 0.042s 2.01e-03s C 21 7 theano.tensor.elemwise.Sum
2.3% 97.2% 0.019s 3.13e-03s C 6 2 theano.sandbox.cuda.basic_ops.GpuAdvancedSubtensor1
0.9% 98.1% 0.007s 5.00e-04s C 15 5 theano.tensor.basic.Alloc
0.5% 98.6% 0.004s 2.34e-04s C 18 6 theano.sandbox.cuda.basic_ops.GpuAlloc
0.5% 99.1% 0.004s 1.43e-04s C 27 9 theano.sandbox.cuda.basic_ops.GpuJoin
0.4% 99.5% 0.004s 3.59e-05s C 102 34 theano.sandbox.cuda.basic_ops.GpuElemwise
0.2% 99.7% 0.001s 5.48e-05s C 27 9 theano.sandbox.cuda.basic_ops.GpuFromHost
0.1% 99.8% 0.001s 1.49e-05s C 66 22 theano.sandbox.cuda.basic_ops.GpuReshape
0.0% 99.9% 0.000s 4.41e-05s C 6 2 theano.compile.ops.DeepCopyOp
0.0% 99.9% 0.000s 2.63e-06s C 72 24 theano.tensor.subtensor.IncSubtensor
0.0% 99.9% 0.000s 2.80e-06s C 48 16 theano.sandbox.cuda.basic_ops.GpuSubtensor
0.0% 99.9% 0.000s 1.13e-06s C 114 38 theano.sandbox.cuda.basic_ops.GpuDimShuffle
0.0% 99.9% 0.000s 3.96e-05s C 3 1 theano.sandbox.cuda.basic_ops.GpuAllocEmpty
0.0% 100.0% 0.000s 3.20e-05s Py 3 1 theano.tensor.extra_ops.FillDiagonal
0.0% 100.0% 0.000s 1.23e-06s C 69 23 theano.tensor.elemwise.DimShuffle
... (remaining 9 Classes account for 0.03%(0.00s) of the runtime)
Ops
---
<% time> <sum %> <apply time> <time per call> <type> <#call> <#apply> <Op name>
36.2% 36.2% 0.300s 9.98e-02s C 3 1 Elemwise{Composite{(i0 * i1 * LT(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4), i5) * (((i6 + ((i7 * Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4)) / i8)) - ((i9 * Composite{(sqr(i0) * i0)}(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4))) / i10)) + ((i11 * Composite{(sqr(sqr(i0)) * i0)}(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4))) / i12)))}}[(0, 4)]
19.2% 55.4% 0.159s 5.30e-02s C 3 1 Elemwise{Composite{(i0 * ((LT(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3), i4) * ((i5 + (i6 * Composite{(sqr(i0) * i0)}((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4))) + (i7 * Composite{((sqr(sqr(i0)) * sqr(i0)) * i0)}((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4)))) - ((i8 * sqr((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4))) + (i9 * Composite{(sqr(sqr(i0)
10.1% 65.5% 0.084s 2.79e-03s C 30 10 Dot22Scalar
9.6% 75.1% 0.079s 9.81e-04s C 81 27 HostFromGpu
6.4% 81.5% 0.053s 8.89e-03s Py 6 2 Nonzero
6.4% 88.0% 0.053s 1.77e-02s Py 3 1 MatrixInverse
5.0% 92.9% 0.041s 4.58e-03s C 9 3 Sum{axis=[0], acc_dtype=float64}
2.3% 95.2% 0.019s 3.13e-03s C 6 2 GpuAdvancedSubtensor1
0.9% 96.1% 0.008s 1.27e-03s C 6 2 Elemwise{Mul}[(0, 1)]
0.9% 97.0% 0.007s 5.00e-04s C 15 5 Alloc
0.6% 97.6% 0.005s 1.72e-03s C 3 1 Elemwise{Composite{((i0 / i1) + ((i2 * i3) / i1) + ((i4 * i2 * i5) / i6))}}[(0, 0)]
0.5% 98.1% 0.004s 2.76e-04s C 15 5 GpuAlloc
0.5% 98.6% 0.004s 1.43e-04s C 27 9 GpuJoin
0.3% 98.9% 0.002s 4.95e-05s C 45 15 GpuElemwise{sub,no_inplace}
0.2% 99.1% 0.001s 5.48e-05s C 27 9 GpuFromHost
0.2% 99.2% 0.001s 1.08e-04s C 12 4 Elemwise{Cast{float64}}
0.1% 99.3% 0.001s 9.13e-05s C 12 4 Sum{axis=[1], acc_dtype=float64}
0.1% 99.5% 0.001s 6.88e-05s C 15 5 GpuElemwise{mul,no_inplace}
0.1% 99.6% 0.001s 6.47e-05s C 15 5 Elemwise{Sqr}[(0, 0)]
0.1% 99.7% 0.001s 1.61e-05s C 60 20 GpuReshape{2}
... (remaining 75 Ops account for 0.29%(0.00s) of the runtime)
Apply
------
<% time> <sum %> <apply time> <time per call> <#call> <id> <Apply name>
36.2% 36.2% 0.300s 9.98e-02s 3 332 Elemwise{Composite{(i0 * i1 * LT(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4), i5) * (((i6 + ((i7 * Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4)) / i8)) - ((i9 * Composite{(sqr(i0) * i0)}(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4))) / i10)) + ((i11 * Composite{(sqr(sqr(i0)) * i0)}(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4))) / i12)))}}[(0, 4)](HostFromGpu.0, HostFromGpu.0, Reshape{2
19.2% 55.4% 0.159s 5.30e-02s 3 331 Elemwise{Composite{(i0 * ((LT(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3), i4) * ((i5 + (i6 * Composite{(sqr(i0) * i0)}((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4))) + (i7 * Composite{((sqr(sqr(i0)) * sqr(i0)) * i0)}((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4)))) - ((i8 * sqr((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4))) + (i9 * Composite{(sqr(sqr(i0)) * i0)}((C
6.4% 61.8% 0.053s 1.77e-02s 3 318 MatrixInverse(IncSubtensor{InplaceSet;int64::, int64:int64:}.0)
5.3% 67.1% 0.044s 1.46e-02s 3 180 Nonzero(HostFromGpu.0)
5.0% 72.1% 0.042s 1.39e-02s 3 269 Dot22Scalar(Elemwise{Cast{float64}}.0, InplaceDimShuffle{1,0}.0, TensorConstant{2.0})
3.8% 75.9% 0.031s 1.04e-02s 3 335 Sum{axis=[0], acc_dtype=float64}(Elemwise{Composite{(i0 * i1 * LT(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4), i5) * (((i6 + ((i7 * Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4)) / i8)) - ((i9 * Composite{(sqr(i0) * i0)}(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4))) / i10)) + ((i11 * Composite{(sqr(sqr(i0)) * i0)}(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i2, i3, i4))) / i12)))}}[(0, 4)].0)
3.5% 79.4% 0.029s 9.55e-03s 3 286 HostFromGpu(GpuJoin.0)
3.0% 82.3% 0.025s 8.18e-03s 3 329 HostFromGpu(GpuSubtensor{:int64:}.0)
2.5% 84.8% 0.021s 6.96e-03s 3 268 Dot22Scalar(Elemwise{Cast{float64}}.0, InplaceDimShuffle{1,0}.0, TensorConstant{2.0})
2.5% 87.4% 0.021s 6.93e-03s 3 267 Dot22Scalar(Elemwise{Cast{float64}}.0, InplaceDimShuffle{1,0}.0, TensorConstant{2.0})
1.6% 89.0% 0.013s 4.46e-03s 3 216 GpuAdvancedSubtensor1(GpuReshape{1}.0, Subtensor{int64}.0)
1.3% 90.3% 0.011s 3.56e-03s 3 328 HostFromGpu(GpuSubtensor{int64:int64:}.0)
1.2% 91.4% 0.010s 3.22e-03s 3 200 Nonzero(HostFromGpu.0)
0.9% 92.4% 0.008s 2.53e-03s 3 333 Elemwise{Mul}[(0, 1)](HostFromGpu.0, InplaceDimShuffle{1,0}.0, HostFromGpu.0)
0.9% 93.3% 0.007s 2.47e-03s 3 235 Alloc(Subtensor{:int64:}.0, Elemwise{Composite{((i0 // i1) + i2)}}[(0, 0)].0, TensorConstant{1}, TensorConstant{1}, Elemwise{Composite{Switch(LT(i0, i1), Switch(LT((i2 + i0), i1), i1, (i2 + i0)), Switch(LT(i0, i2), i0, i2))}}.0)
0.9% 94.1% 0.007s 2.36e-03s 3 334 Sum{axis=[0], acc_dtype=float64}(Elemwise{Composite{(i0 * ((LT(Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3), i4) * ((i5 + (i6 * Composite{(sqr(i0) * i0)}((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4))) + (i7 * Composite{((sqr(sqr(i0)) * sqr(i0)) * i0)}((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4)))) - ((i8 * sqr((Composite{Cast{float32}(sqrt(((i0 + i1) - i2)))}(i1, i2, i3) / i4))) + (i9 *
0.6% 94.8% 0.005s 1.79e-03s 3 226 GpuAdvancedSubtensor1(GpuReshape{1}.0, Subtensor{int64}.0)
0.6% 95.4% 0.005s 1.72e-03s 3 337 Elemwise{Composite{((i0 / i1) + ((i2 * i3) / i1) + ((i4 * i2 * i5) / i6))}}[(0, 0)](Sum{axis=[0], acc_dtype=float64}.0, InplaceDimShuffle{x}.0, InplaceDimShuffle{x}.0, Sum{axis=[0], acc_dtype=float64}.0, TensorConstant{(1,) of -1.0}, Sum{axis=[0], acc_dtype=float64}.0, InplaceDimShuffle{x}.0)
0.6% 96.0% 0.005s 1.69e-03s 3 330 HostFromGpu(GpuSubtensor{int64::}.0)
0.5% 96.5% 0.004s 1.37e-03s 3 153 HostFromGpu(GpuReshape{1}.0)
... (remaining 322 Apply instances account for 3.51%(0.03s) of the runtime)
Here are tips to potentially make your code run faster
(if you think of new ones, suggest them on the mailing list).
Test them first, as they are not guaranteed to always provide a speedup.
- Try installing amdlibm and set the Theano flag lib.amdlibm=True. This speeds up only some Elemwise operation.
| MIT | Prototype Notebook/Example_1_Sandstone.ipynb | nre-aachen/gempy |
Ungraded Lab: Activation in Custom LayersIn this lab, we extend our knowledge of building custom layers by adding an activation parameter. The implementation is pretty straightforward as you'll see below. Imports | try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow.keras.layers import Layer | _____no_output_____ | MIT | C1W3_L3_CustomLayerWithActivation.ipynb | 100rab-S/TensorFlow-Advanced-Techniques |
Adding an activation layerTo use the built-in activations in Keras, we can specify an `activation` parameter in the `__init__()` method of our custom layer class. From there, we can initialize it by using the `tf.keras.activations.get()` method. This takes in a string identifier that corresponds to one of the [available activations](https://keras.io/api/layers/activations/available-activations) in Keras. Next, you can now pass in the forward computation to this activation in the `call()` method. | class SimpleDense(Layer):
# add an activation parameter
def __init__(self, units=32, activation=None):
super(SimpleDense, self).__init__()
self.units = units
# define the activation to get from the built-in activation layers in Keras
self.activation = tf.keras.activations.get(activation)
def build(self, input_shape): # we don't need to change anything in this method to add activation to our custom layer
w_init = tf.random_normal_initializer()
self.w = tf.Variable(name="kernel",
initial_value=w_init(shape=(input_shape[-1], self.units),
dtype='float32'),
trainable=True)
b_init = tf.zeros_initializer()
self.b = tf.Variable(name="bias",
initial_value=b_init(shape=(self.units,), dtype='float32'),
trainable=True)
#super().build(input_shape)
def call(self, inputs):
# pass the computation to the activation layer
return self.activation(tf.matmul(inputs, self.w) + self.b) | _____no_output_____ | MIT | C1W3_L3_CustomLayerWithActivation.ipynb | 100rab-S/TensorFlow-Advanced-Techniques |
We can now pass in an activation parameter to our custom layer. The string identifier is mostly the same as the function name so 'relu' below will get `tf.keras.activations.relu`. | mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
SimpleDense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(x_train, y_train, epochs=5)
model.evaluate(x_test, y_test) | Downloading data from https://storage.googleapis.com/tensorflow/tf-keras-datasets/mnist.npz
11493376/11490434 [==============================] - 0s 0us/step
Epoch 1/5
1875/1875 [==============================] - 5s 2ms/step - loss: 0.4861 - accuracy: 0.8560
Epoch 2/5
1875/1875 [==============================] - 4s 2ms/step - loss: 0.1496 - accuracy: 0.9553
Epoch 3/5
1875/1875 [==============================] - 4s 2ms/step - loss: 0.1051 - accuracy: 0.9682
Epoch 4/5
1875/1875 [==============================] - 4s 2ms/step - loss: 0.0864 - accuracy: 0.9735
Epoch 5/5
1875/1875 [==============================] - 4s 2ms/step - loss: 0.0727 - accuracy: 0.9771
313/313 [==============================] - 1s 1ms/step - loss: 0.0716 - accuracy: 0.9778
| MIT | C1W3_L3_CustomLayerWithActivation.ipynb | 100rab-S/TensorFlow-Advanced-Techniques |
Using Google Cloud Functions to support event-based triggering of Cloud AI Platform Pipelines> This post shows how you can run a Cloud AI Platform Pipeline from a Google Cloud Function, providing a way for Pipeline runs to be triggered by events.- toc: true - badges: true- comments: true- categories: [ml, pipelines, mlops, kfp, gcf] This example shows how you can run a [Cloud AI Platform Pipeline](https://cloud.google.com/blog/products/ai-machine-learning/introducing-cloud-ai-platform-pipelines) from a [Google Cloud Function](https://cloud.google.com/functions/docs/), thus providing a way for Pipeline runs to be triggered by events (in the interim before this is supported by Pipelines itself). In this example, the function is triggered by the addition of or update to a file in a [Google Cloud Storage](https://cloud.google.com/storage/) (GCS) bucket, but Cloud Functions can have other triggers too (including [Pub/Sub](https://cloud.google.com/pubsub/docs/)-based triggers).The example is Google Cloud Platform (GCP)-specific, and requires a [Cloud AI Platform Pipelines](https://cloud.google.com/ai-platform/pipelines/docs) installation using Pipelines version >= 0.4. To run this example as a notebook, click on one of the badges at the top of the page or see [here](https://github.com/amygdala/code-snippets/blob/master/ml/notebook_examples/functions/hosted_kfp_gcf.ipynb).(If you are instead interested in how to do this with a Kubeflow-based pipelines installation, see [this notebook](https://github.com/amygdala/kubeflow-examples/blob/cookbook/cookbook/pipelines/notebooks/gcf_kfp_trigger.ipynb)). Setup Create a Cloud AI Platform Pipelines installationFollow the instructions in the [documentation](https://cloud.google.com/ai-platform/pipelines/docs) to create a Cloud AI Platform Pipelines installation. Identify (or create) a Cloud Storage bucket to use for the example **Before executing the next cell**, edit it to **set the `TRIGGER_BUCKET` environment variable** to a Google Cloud Storage bucket ([create a bucket first](https://console.cloud.google.com/storage/browser) if necessary). Do *not* include the `gs://` prefix in the bucket name.We'll deploy the GCF function so that it will trigger on new and updated files (blobs) in this bucket. | %env TRIGGER_BUCKET=REPLACE_WITH_YOUR_GCS_BUCKET_NAME | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
Give Cloud Function's service account the necessary accessFirst, make sure the Cloud Function API [is enabled](https://console.cloud.google.com/apis/library/cloudfunctions.googleapis.com?q=functions).Cloud Functions uses the project's 'appspot' acccount for its service account. It will have the form: `PROJECT_ID@appspot.gserviceaccount.com`. (This is also the project's App Engine service account).- Go to your project's [IAM - Service Account page](https://console.cloud.google.com/iam-admin/serviceaccounts).- Find the ` PROJECT_ID@appspot.gserviceaccount.com` account and copy its email address.- Find the project's Compute Engine (GCE) default service account (this is the default account used for the Pipelines installation). It will have a form like this: `PROJECT_NUMBER@developer.gserviceaccount.com`. Click the checkbox next to the GCE service account, and in the 'INFO PANEL' to the right, click **ADD MEMBER**. Add the Functions service account (`PROJECT_ID@appspot.gserviceaccount.com`) as a **Project Viewer** of the GCE service account. 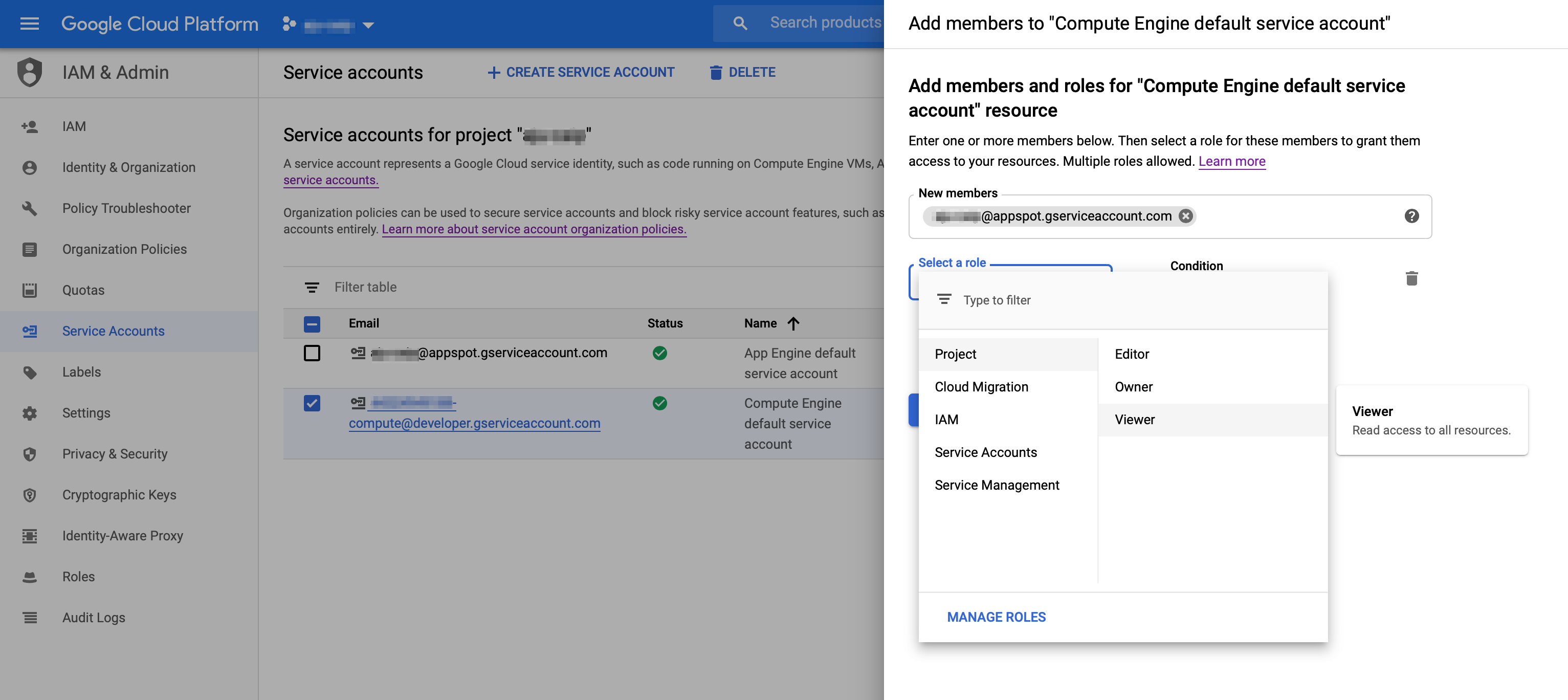 Next, configure your `TRIGGER_BUCKET` to allow the Functions service account access to that bucket. - Navigate in the console to your list of buckets in the [Storage Browser](https://console.cloud.google.com/storage/browser).- Click the checkbox next to the `TRIGGER_BUCKET`. In the 'INFO PANEL' to the right, click **ADD MEMBER**. Add the service account (`PROJECT_ID@appspot.gserviceaccount.com`) with `Storage Object Admin` permissions. (While not tested, giving both Object view and create permissions should also suffice).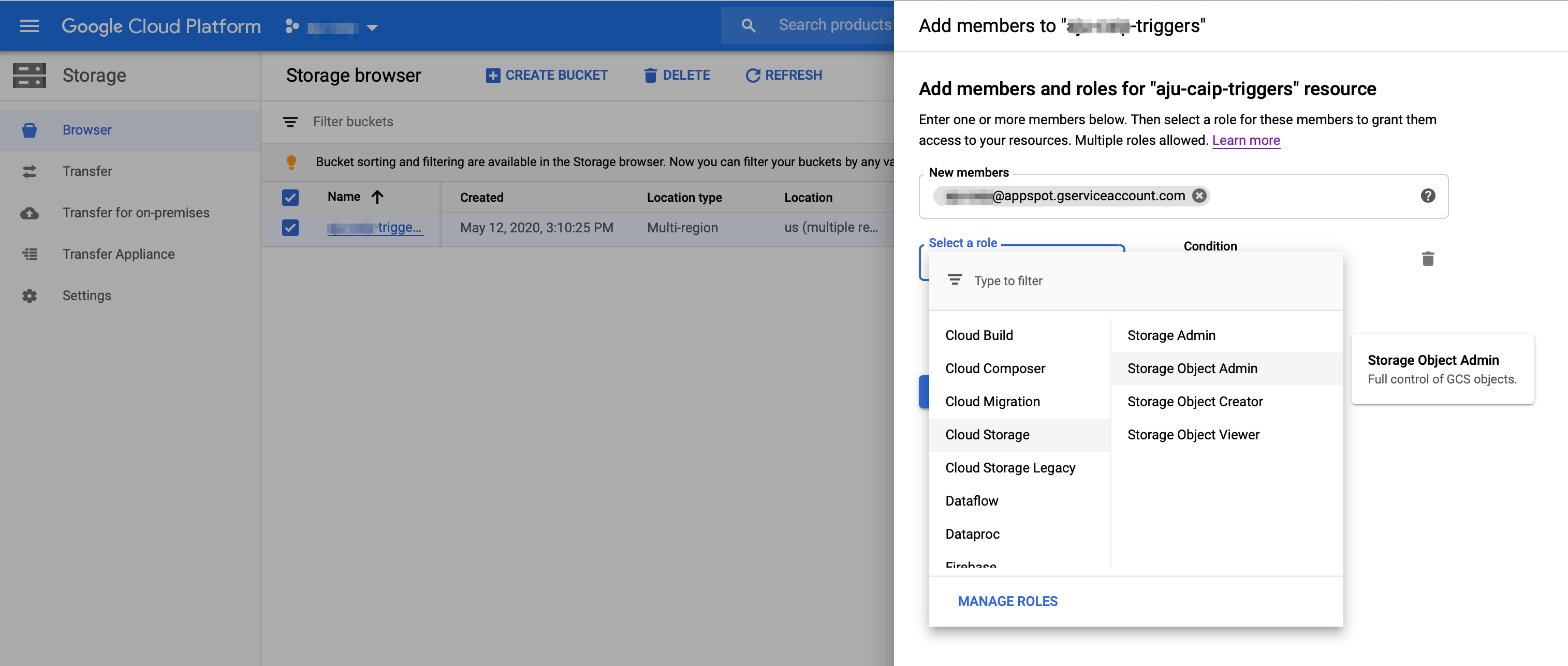 Create a simple GCF function to test your configurationFirst we'll generate and deploy a simple GCF function, to test that the basics are properly configured. | %%bash
mkdir -p functions | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
We'll first create a `requirements.txt` file, to indicate what packages the GCF code requires to be installed. (We won't actually need `kfp` for this first 'sanity check' version of a GCF function, but we'll need it below for the second function we'll create, that deploys a pipeline). | %%writefile functions/requirements.txt
kfp | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
Next, we'll create a simple GCF function in the `functions/main.py` file: | %%writefile functions/main.py
import logging
def gcs_test(data, context):
"""Background Cloud Function to be triggered by Cloud Storage.
This generic function logs relevant data when a file is changed.
Args:
data (dict): The Cloud Functions event payload.
context (google.cloud.functions.Context): Metadata of triggering event.
Returns:
None; the output is written to Stackdriver Logging
"""
logging.info('Event ID: {}'.format(context.event_id))
logging.info('Event type: {}'.format(context.event_type))
logging.info('Data: {}'.format(data))
logging.info('Bucket: {}'.format(data['bucket']))
logging.info('File: {}'.format(data['name']))
file_uri = 'gs://%s/%s' % (data['bucket'], data['name'])
logging.info('Using file uri: %s', file_uri)
logging.info('Metageneration: {}'.format(data['metageneration']))
logging.info('Created: {}'.format(data['timeCreated']))
logging.info('Updated: {}'.format(data['updated'])) | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
Deploy the GCF function as follows. (You'll need to **wait a moment or two for output of the deployment to display in the notebook**). You can also run this command from a notebook terminal window in the `functions` subdirectory. | %%bash
cd functions
gcloud functions deploy gcs_test --runtime python37 --trigger-resource ${TRIGGER_BUCKET} --trigger-event google.storage.object.finalize | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
After you've deployed, test your deployment by adding a file to the specified `TRIGGER_BUCKET`. You can do this easily by visiting the **Storage** panel in the Cloud Console, clicking on the bucket in the list, and then clicking on **Upload files** in the bucket details view.Then, check in the logs viewer panel (https://console.cloud.google.com/logs/viewer) to confirm that the GCF function was triggered and ran correctly. You can select 'Cloud Function' in the first pulldown menu to filter on just those log entries. Deploy a Pipeline from a GCF functionNext, we'll create a GCF function that deploys an AI Platform Pipeline when triggered. First, preserve your existing main.py in a backup file: | %%bash
cd functions
mv main.py main.py.bak | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
Then, **before executing the next cell**, **edit the `HOST` variable** in the code below. You'll replace `` with the correct value for your installation.To find this URL, visit the [Pipelines panel](https://console.cloud.google.com/ai-platform/pipelines/) in the Cloud Console. From here, you can find the URL by clicking on the **SETTINGS** link for the Pipelines installation you want to use, and copying the 'host' string displayed in the client example code (prepend `https://` to that string in the code below). You can alternately click on **OPEN PIPELINES DASHBOARD** for the Pipelines installation, and copy that URL, removing the `//pipelines` suffix. | %%writefile functions/main.py
import logging
import datetime
import logging
import time
import kfp
import kfp.compiler as compiler
import kfp.dsl as dsl
import requests
# TODO: replace with your Pipelines endpoint URL
HOST = 'https://<your_endpoint>.pipelines.googleusercontent.com'
@dsl.pipeline(
name='Sequential',
description='A pipeline with two sequential steps.'
)
def sequential_pipeline(filename='gs://ml-pipeline-playground/shakespeare1.txt'):
"""A pipeline with two sequential steps."""
op1 = dsl.ContainerOp(
name='filechange',
image='library/bash:4.4.23',
command=['sh', '-c'],
arguments=['echo "%s" > /tmp/results.txt' % filename],
file_outputs={'newfile': '/tmp/results.txt'})
op2 = dsl.ContainerOp(
name='echo',
image='library/bash:4.4.23',
command=['sh', '-c'],
arguments=['echo "%s"' % op1.outputs['newfile']]
)
def get_access_token():
url = 'http://metadata.google.internal/computeMetadata/v1/instance/service-accounts/default/token'
r = requests.get(url, headers={'Metadata-Flavor': 'Google'})
r.raise_for_status()
access_token = r.json()['access_token']
return access_token
def hosted_kfp_test(data, context):
logging.info('Event ID: {}'.format(context.event_id))
logging.info('Event type: {}'.format(context.event_type))
logging.info('Data: {}'.format(data))
logging.info('Bucket: {}'.format(data['bucket']))
logging.info('File: {}'.format(data['name']))
file_uri = 'gs://%s/%s' % (data['bucket'], data['name'])
logging.info('Using file uri: %s', file_uri)
logging.info('Metageneration: {}'.format(data['metageneration']))
logging.info('Created: {}'.format(data['timeCreated']))
logging.info('Updated: {}'.format(data['updated']))
token = get_access_token()
logging.info('attempting to launch pipeline run.')
ts = int(datetime.datetime.utcnow().timestamp() * 100000)
client = kfp.Client(host=HOST, existing_token=token)
compiler.Compiler().compile(sequential_pipeline, '/tmp/sequential.tar.gz')
exp = client.create_experiment(name='gcstriggered') # this is a 'get or create' op
res = client.run_pipeline(exp.id, 'sequential_' + str(ts), '/tmp/sequential.tar.gz',
params={'filename': file_uri})
logging.info(res)
| _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
Next, deploy the new GCF function. As before, **it will take a moment or two for the results of the deployment to display in the notebook**. | %%bash
cd functions
gcloud functions deploy hosted_kfp_test --runtime python37 --trigger-resource ${TRIGGER_BUCKET} --trigger-event google.storage.object.finalize | _____no_output_____ | Apache-2.0 | _notebooks/2020-05-12-hosted_kfp_gcf.ipynb | amygdala/fastpages |
Herramientas Estadisticas Contenido:1.Estadistica: - Valor medio. - Mediana. - Desviacion estandar. 2.Histogramas: - Histrogramas con python. - Histogramas con numpy. - Como normalizar un histograma. 3.Distribuciones: - Como obtener una distribucion a partir de un histograma. - Distribucion Normal - Distribucion de Poisson - Distribucion Binomial 1. Estadistica Promedio El promedio de una variable $x$ esta definado como:$\bar{x} = \dfrac{\sum{x_i}}{N} $ Mediana La mediana de un conjunto de datos, es el valor al cual el conjunto de datosse divide en dos: Ejemplo: sea $x$ = [1, 4, 7, 7, 3, 3, 1] la mediana de $median(x) = 3$Formalmente la mediana se define como el valor $x_m$ que divide la funcion de probabilidad $F(x)$ en partes iguales.$F(x_m) = \dfrac{1}{2}$ El valor mas probableEs el valor con mayor probabilidad $x_p$.Ejemplo: sea $x$ = [1, 4, 7, 7, 3, 2, 1] el valor mas probable es $x_p = 7$ | import matplotlib.pyplot as plt
import numpy as np
# %pylab inline
def mi_mediana(lista):
x = sorted(lista)
d = int(len(x)/2)
if(len(x)%2==0):
return (x[d-1] + x[d])*0.5
else:
return x[d-1]
x_input = [1,3,4,5,5,7,7,6,8,6]
mi_mediana(x_input)
print(mi_mediana(x_input) == np.median(x_input)) | True
| MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Problemas de no saber estadísticaEste tipo de conceptos parecen sencillos. Pero no siempre son claros para todo el mundo. | x = np.arange(1, 12)
y = np.random.random(11)*10
plt.figure(figsize=(12, 5))
fig = plt.subplot(1, 2, 1)
plt.scatter(x, y, c='purple', alpha=0.8, s=60)
y_mean = np.mean(y)
y_median = np.median(y)
plt.axhline(y_mean, c='g', lw=3, label=r"$\rm{Mean}$")
plt.axhline(y_median, c='r', lw=3, label=r"$\rm{Median}$")
plt.legend(fontsize=20)
fig = plt.subplot(1, 2, 2)
h = plt.hist(x, alpha=0.6, histtype='bar', ec='black')
print(y) | [9.33745032 0.46206052 3.07349261 8.65709198 6.44733954 2.5552359
8.93987727 8.24695437 5.62111292 4.64621772 0.05366015]
| MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Desviacion estandarEs el promedio de las incertidumbres de las mediciones $x_i$$\sigma = \sqrt{\dfrac{1}{n-1} \sum(x_{i} - \bar{x})^2}$Donde $n$ es el número de la muestraAdicionalmente la ${\bf{varianza}}$ se define como:$\bar{x^2} - \bar{x}^{2}$$\sigma^2 = \dfrac{1}{N} \sum(x_{i} - \bar{x})^2$Y es una medida similar a la desviacion estandar que da cuenta de la dispersion de los datos alrededor del promedio.Donde $N$ es la población total. Función de Correlación$cor(x, y) = \dfrac{}{\sigma_x \sigma_{y}} $ Ejercicio:Compruebe si se cumplen las siguientes propiedades:1. Cor(X,Y) = Cor(Y, X)2. Cor(X,X) = 13. Cor(X,-X) = -14. Cor(aX+b, cY + d) = Cor(X, Y), si a y c != 0 | x = np.arange(1, 12)
y = np.random.random(11)*10
plt.figure(figsize=(9, 5))
y_mean = np.mean(y)
y_median = np.median(y)
plt.axhline(y_mean, c='g', lw=3, label=r"$\rm{Mean}$")
plt.axhline(y_median, c='r', lw=3, label=r"$\rm{Median}$")
sigma_y = np.std(y)
plt.axhspan(y_mean-sigma_y, y_mean + sigma_y, facecolor='g', alpha=0.5, label=r"$\rm{\sigma}$")
plt.legend(fontsize=20)
plt.scatter(x, y, c='purple', alpha=0.8, s=60)
plt.ylim(-2, 14)
print ("Variancia = ", np.var(y))
print ("Desviacion estandar = ", np.std(y)) | Variancia = 7.888849132964844
Desviacion estandar = 2.8087095138096507
| MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Referencias: Para mas funciones estadisticas que se pueden usar en python ver: - NumPy: http://docs.scipy.org/doc/numpy/reference/routines.statistics.html- SciPy: http://docs.scipy.org/doc/scipy/reference/stats.html Histogramas 1. histhist es una funcion de python que genera un histograma a partir de un array de datos. | x = np.random.random(200)
plt.subplot(2,2,1)
plt.title("A simple hist")
h = plt.hist(x)
plt.subplot(2,2,2)
plt.title("bins")
h = plt.hist(x, bins=20)
plt.subplot(2,2,3)
plt.title("alpha")
h = plt.hist(x, bins=20, alpha=0.6)
plt.subplot(2,2,4)
plt.title("histtype")
h = plt.hist(x, bins=20, alpha=0.6, histtype='stepfilled') | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
2. Numpy-histogram | N, bins = np.histogram(caras, bins=15)
plt.plot(bins[0:-1], N) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Histogramas 2D | x = np.random.random(500)
y = np.random.random(500)
plt.subplot(4, 2, 1)
plt.hexbin(x, y, gridsize=15, cmap="gray")
plt.colorbar()
plt.subplot(4, 2, 2)
data = plt.hist2d(x, y, bins=15, cmap="binary")
plt.colorbar()
plt.subplot(4, 2, 3)
plt.hexbin(x, y, gridsize=15)
plt.colorbar()
plt.subplot(4, 2, 4)
data = plt.hist2d(x, y, bins=15)
plt.colorbar() | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Como normalizar un histograma. Normalizar un histograma significa que la integral del histograma sea 1. | x = np.random.random(10)*4
plt.title("Como no normalizar un histograma", fontsize=25)
h = plt.hist(x, normed="True")
print ("El numero tamaño del bin debe de ser de la unidad")
plt.title("Como normalizar un histograma", fontsize=25)
h = hist(x, normed="True", bins=4)
| _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Cual es la probabilidad de sacar 9 veces cara en 10 lanzamientos? Distribución de Probabilidad:Las distribuciones de probabilidad dan información de cual es la probabilidad de que una variable aleatoria $x$ aprezca en un intervalo dado. ¿Si tenemos un conjunto de datos como podemos conocer la distribucion de probabilidad? | x = np.random.random(100)*10
plt.subplot(1, 2, 1)
h = plt.hist(x)
plt.subplot(1, 2, 2)
histo, bin_edges = np.histogram(x, density=True)
plt.bar(bin_edges[:-1], histo, width=1)
plt.xlim(min(bin_edges), max(bin_edges)) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Distribución Normal: Descripcion Matemática.$f(x, \mu, \sigma) = \dfrac{1}{\sigma \sqrt(2\pi)} e^{-\dfrac{(x-\mu)^2}{2\sigma^2}} $donde $\sigma$ es la desviacion estandar y $\mu$ la media de los datos $x$Es una función de distribucion de probabilidad que esta totalmente determinada por los parametros $\mu$ y $\sigma$. La funcion es simetrica alrededor de $\mu$.En python podemos usar scipy para hacer uso de la función normal. | import scipy.stats
x = np.linspace(0, 1, 100)
n_dist = scipy.stats.norm(0.5, 0.1)
plt.plot(x, n_dist.pdf(x)) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Podemos generar numeros aleatorios con una distribucion normal: | x = np.random.normal(0.0, 1.0, 1000)
y = np.random.normal(0.0, 2.0, 1000)
w = np.random.normal(0.0, 3.0, 1000)
z = np.random.normal(0.0, 4.0, 1000)
histo = plt.hist(z, alpha=0.2, histtype="stepfilled", color='r')
histo = plt.hist(w, alpha=0.4, histtype="stepfilled", color='b')
histo = plt.hist(y, alpha=0.6, histtype="stepfilled", color='k')
histo = plt.hist(x, alpha=0.8, histtype="stepfilled", color='g')
plt.title(r"$\rm{Distribuciones\ normales\ con\ diferente\ \sigma}$", fontsize=20) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
**Intervalo de confianza**$\sigma_1$ = 68% de los datos van a estar dentro de 1$\sigma$$\sigma_2$ = 95% de los datos van a estar dentro de 2$\sigma$$\sigma_3$ = 99.7% de los datos van a estar dentro de 3$\sigma$ Ejercicio: Generen distribuciones normales con:- $\mu = 5$ y $\sigma = 2$ - $\mu = -3$ y $\sigma = -2$- $\mu = 4$ y $\sigma = 5$ Grafiquen las PDF,CDF sobre los mismos ejes, con distintos colores y leyendas. Qué observan? (Una gráfica con PDF y otra con CDF). Ejercicio:1. Realize graficas de: 1. Diferencia de Caras - Sellos para 40 y 20 mediciones cada una con mayor numero de lanzamientos que la anterior. (abs(cara-sello)vs Numero de lanzamientos) 2. La razon (sara/sello) en funcion del Numero de lanzamientos. Comente los resultados.2. Repita los graficos anteriores pero ahora hagalos en escala logaritmica. Comente los resultados.3. Haga graficos de el promedio de abs(cara - sello) en funcion del numero de lanzamientos en escala logaritmica.y otro con el promedio de (cara/sello). Comente los reultados.4. Repita el punto anterior pero esta vez con la desviación estandar. comente los resultados. Imaginemos por un momento el siguiente experimento: Queremos estudiar la probabilidad de que al lanzar una moneda obtengamos cara o sello, de antamento sabemos que esta es del 50%.Pero analizemos un poco mas a fondo, ¿Cual será la probabilidad de sacar 10 caras consecutivas?Para responder proponemos el siguiente método:1. Lanzamos una moneda 10 veces y miramos si sale cara o sello y guardamos estos datos. 2. Repetimos este procedimiento y 1000 veces. Funcion que lanza la moneda N veces. | def coinflip(N):
cara = 0
sello = 0
i=0
while i < N:
x = np.random.randint(0, 10)/5.0
if x >= 1.0:
cara+=1
elif x<1.0:
sello+=1
i+=1
return cara/N, sello/N | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
Función que hace M veces N lanzamientos. | def realizaciones(M, N):
caras=[]
for i in range(M):
x, y = coinflip(N)
caras.append(x)
return caras
hist(caras, normed=True, bins=20)
caras = realizaciones(100000, 30.) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
PDF | N, bins = np.histogram(x, density=True)
plt.plot(bins[0:-1], N) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
CDF | h = plt.hist(x, cumulative=True, bins=20) | _____no_output_____ | MIT | Notebooks/python 9 - Herramientas_Estadisticas.ipynb | diegour1/HerramientasComputacionales |
01 - Sentence Classification Model Building Parse & clearn labeled training data | import xml.etree.ElementTree as ET
tree = ET.parse('../data/Restaurants_Train.xml')
root = tree.getroot()
root
# Use this dataframe for multilabel classification
# Must use scikitlearn's multilabel binarizer
labeled_reviews = []
for sentence in root.findall("sentence"):
entry = {}
aterms = []
aspects = []
if sentence.find("aspectTerms"):
for aterm in sentence.find("aspectTerms").findall("aspectTerm"):
aterms.append(aterm.get("term"))
if sentence.find("aspectCategories"):
for aspect in sentence.find("aspectCategories").findall("aspectCategory"):
aspects.append(aspect.get("category"))
entry["text"], entry["terms"], entry["aspects"]= sentence[0].text, aterms, aspects
labeled_reviews.append(entry)
labeled_df = pd.DataFrame(labeled_reviews)
print("there are",len(labeled_reviews),"reviews in this training set")
# print(sentence.find("aspectCategories").findall("aspectCategory").get("category"))
# Save annotated reviews
labeled_df.to_pickle("annotated_reviews_df.pkl")
labeled_df.head() | _____no_output_____ | Apache-2.0 | 04-Aspect_Based_Opinion_Mining/code/01-Build_Model.ipynb | ayan1995/DS_projects |
Training the model with Naive Bayes1. replace pronouns with neural coref2. train the model with naive bayes | from neuralcoref import Coref
import en_core_web_lg
spacy = en_core_web_lg.load()
coref = Coref(nlp=spacy)
# Define function for replacing pronouns using neuralcoref
def replace_pronouns(text):
coref.one_shot_coref(text)
return coref.get_resolved_utterances()[0]
# Read annotated reviews df, which is the labeled dataset for training
# This is located in the pickled files folder
annotated_reviews_df = pd.read_pickle("../pickled_files/annotated_reviews_df.pkl")
annotated_reviews_df.head(3)
# Create a new column for text whose pronouns have been replaced
annotated_reviews_df["text_pro"] = annotated_reviews_df.text.map(lambda x: replace_pronouns(x))
# uncomment below to pickle the new df
# annotated_reviews_df.to_pickle("annotated_reviews_df2.pkl")
# Read pickled file with replaced pronouns if it exists already
annotated_reviews_df = pd.read_pickle("annotated_reviews_df2.pkl")
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MultiLabelBinarizer
# Convert the multi-labels into arrays
mlb = MultiLabelBinarizer()
y = mlb.fit_transform(annotated_reviews_df.aspects)
X = annotated_reviews_df.text_pro
# Split data into train and test set
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=0)
# save the the fitted binarizer labels
# This is important: it contains the how the multi-label was binarized, so you need to
# load this in the next folder in order to undo the transformation for the correct labels.
filename = 'mlb.pkl'
pickle.dump(mlb, open(filename, 'wb'))
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.pipeline import Pipeline
from sklearn.naive_bayes import MultinomialNB
from skmultilearn.problem_transform import LabelPowerset
import numpy as np
# LabelPowerset allows for multi-label classification
# Build a pipeline for multinomial naive bayes classification
text_clf = Pipeline([('vect', CountVectorizer(stop_words = "english",ngram_range=(1, 1))),
('tfidf', TfidfTransformer(use_idf=False)),
('clf', LabelPowerset(MultinomialNB(alpha=1e-1))),])
text_clf = text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
# Calculate accuracy
np.mean(predicted == y_test)
# Test if SVM performs better
from sklearn.linear_model import SGDClassifier
text_clf_svm = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf-svm', LabelPowerset(
SGDClassifier(loss='hinge', penalty='l2',
alpha=1e-3, max_iter=6, random_state=42)))])
_ = text_clf_svm.fit(X_train, y_train)
predicted_svm = text_clf_svm.predict(X_test)
#Calculate accuracy
np.mean(predicted_svm == y_test)
import pickle
# Train naive bayes on full dataset and save model
text_clf = Pipeline([('vect', CountVectorizer(stop_words = "english",ngram_range=(1, 1))),
('tfidf', TfidfTransformer(use_idf=False)),
('clf', LabelPowerset(MultinomialNB(alpha=1e-1))),])
text_clf = text_clf.fit(X, y)
# save the model to disk
filename = 'naive_model1.pkl'
pickle.dump(text_clf, open(filename, 'wb')) | _____no_output_____ | Apache-2.0 | 04-Aspect_Based_Opinion_Mining/code/01-Build_Model.ipynb | ayan1995/DS_projects |
At this point, we can move on to 02-Sentiment analysis notebook, which will load the fitted Naive bayes model. | #mlb.inverse_transform(predicted)
pred_df = pd.DataFrame(
{'text_pro': X_test,
'pred_category': mlb.inverse_transform(predicted)
})
pd.set_option('display.max_colwidth', -1)
pred_df.head() | _____no_output_____ | Apache-2.0 | 04-Aspect_Based_Opinion_Mining/code/01-Build_Model.ipynb | ayan1995/DS_projects |
Some scrap code below which wasn't used | # Save annotated reviews
labeled_df.to_pickle("annotated_reviews_df.pkl")
labeled_df.head()
# This code was for parsing out terms & their relations to aspects
# However, the terms were not always hyponyms of the aspects, so they were unusable
aspects = {"food":[],"service":[],"anecdotes/miscellaneous":[], "ambience":[], "price":[]}
for i in range(len(labeled_df)):
if len(labeled_df.aspects[i]) == 1:
if labeled_df.terms[i] != []:
for terms in labeled_df.terms[i]:
aspects[labeled_df.aspects[i][0]].append(terms.lower())
for key in aspects:
aspects[key] = list(set(aspects[key]))
terms = []
for i in labeled_df.terms:
for j in i:
if j not in terms:
terms.append(j)
print("there are", len(terms),"unique terms")
# Use this dataframe if doing the classifications separately as binary classifications
labeled_reviews2 = []
for sentence in root.findall("sentence"):
entry = {"food":0,"service":0,"anecdotes/miscellaneous":0, "ambience":0, "price":0}
aterms = []
aspects = []
if sentence.find("aspectTerms"):
for aterm in sentence.find("aspectTerms").findall("aspectTerm"):
aterms.append(aterm.get("term"))
if sentence.find("aspectCategories"):
for aspect in sentence.find("aspectCategories").findall("aspectCategory"):
if aspect.get("category") in entry.keys():
entry[aspect.get("category")] = 1
entry["text"], entry["terms"] = sentence[0].text, aterms
labeled_reviews2.append(entry)
labeled_df2 = pd.DataFrame(labeled_reviews2)
# print(sentence.find("aspectCategories").findall("aspectCategory").get("category"))
labeled_df2.iloc[:,:5].sum() | _____no_output_____ | Apache-2.0 | 04-Aspect_Based_Opinion_Mining/code/01-Build_Model.ipynb | ayan1995/DS_projects |
Network inference of categorical variables: non-sequential data | import sys
import numpy as np
from scipy import linalg
from sklearn.preprocessing import OneHotEncoder
import matplotlib.pyplot as plt
%matplotlib inline
import inference
import fem
# setting parameter:
np.random.seed(1)
n = 20 # number of positions
m = 5 # number of values at each position
l = int(((n*m)**2)) # number of samples
g = 2.
nm = n*m
def itab(n,m):
i1 = np.zeros(n)
i2 = np.zeros(n)
for i in range(n):
i1[i] = i*m
i2[i] = (i+1)*m
return i1.astype(int),i2.astype(int)
# generate coupling matrix w0:
def generate_interactions(n,m,g):
nm = n*m
w = np.random.normal(0.0,g/np.sqrt(nm),size=(nm,nm))
i1tab,i2tab = itab(n,m)
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
w[i1:i2,:] -= w[i1:i2,:].mean(axis=0)
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
w[i1:i2,i1:i2] = 0. # no self-interactions
for i in range(nm):
for j in range(nm):
if j > i: w[i,j] = w[j,i]
return w
i1tab,i2tab = itab(n,m)
w0 = inference.generate_interactions(n,m,g)
#plt.imshow(w0,cmap='rainbow',origin='lower')
#plt.clim(-0.5,0.5)
#plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
#plt.show()
#print(w0)
def generate_sequences2(w,n,m,l):
i1tab,i2tab = itab(n,m)
# initial s (categorical variables)
s_ini = np.random.randint(0,m,size=(l,n)) # integer values
#print(s_ini)
# onehot encoder
enc = OneHotEncoder(n_values=m)
s = enc.fit_transform(s_ini).toarray()
print(s)
nrepeat = 500
for irepeat in range(nrepeat):
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
h = s.dot(w[i1:i2,:].T) # h[t,i1:i2]
h_old = (s[:,i1:i2]*h).sum(axis=1) # h[t,i0]
k = np.random.randint(0,m,size=l)
for t in range(l):
if np.exp(h[t,k[t]] - h_old[t]) > np.random.rand():
s[t,i1:i2] = 0.
s[t,i1+k[t]] = 1.
return s
# 2018.11.07: Tai
def nrgy(s,w):
l = s.shape[0]
n,m = 20,3
i1tab,i2tab = itab(n,m)
p = np.zeros((l,n))
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
h = s.dot(w[i1:i2,:].T)
#e = (s[:,i1:i2]*h).sum(axis=1)
#p[:,i] = np.exp(e)
#p_sum = np.sum(np.exp(h),axis=1)
#p[:,i] /= p_sum
p[:,i] = np.exp((s[:,i1:i2]*h).sum(axis=1))/(np.exp(h).sum(axis=1))
#like = p.sum(axis=1)
return np.sum(np.log(p),axis=1)
# Vipul:
def nrgy_vp(onehot,w):
nrgy = onehot*(onehot.dot(w.T))
# print(nrgy - np.log(2*np.cosh(nrgy)))
return np.sum(nrgy - np.log(2*np.cosh(nrgy)),axis=1) #ln prob
def generate_sequences_vp(w,n_positions,n_residues,n_seq):
n_size = n_residues*n_positions
n_trial = 100*(n_size) #monte carlo steps to find the right sequences
b = np.zeros((n_size))
trial_seq = np.tile(np.random.randint(0,n_residues,size=(n_positions)),(n_seq,1))
print(trial_seq[0])
enc = OneHotEncoder(n_values=n_residues)
onehot = enc.fit_transform(trial_seq).toarray()
old_nrgy = nrgy(onehot,w) #+ n_positions*(n_residues-1)*np.log(2)
for trial in range(n_trial):
# print('before',np.mean(old_nrgy))
index_array = np.random.choice(range(n_positions),size=2,replace=False)
index,index1 = index_array[0],index_array[1]
r_trial = np.random.randint(0,n_residues,size=(n_seq))
r_trial1 = np.random.randint(0,n_residues,size=(n_seq))
mod_seq = np.copy(trial_seq)
mod_seq[:,index] = r_trial
mod_seq[:,index1] = r_trial1
mod_nrgy = nrgy(enc.fit_transform(mod_seq).toarray(),w) #+ n_positions*(n_residues-1)*np.log(2)
seq_change = mod_nrgy-old_nrgy > np.log(np.random.rand(n_seq))
#seq_change = mod_nrgy/(old_nrgy+mod_nrgy) > np.random.rand(n_seq)
if trial>n_size:
trial_seq[seq_change,index] = r_trial[seq_change]
trial_seq[seq_change,index1] = r_trial1[seq_change]
old_nrgy[seq_change] = mod_nrgy[seq_change]
else:
best_seq = np.argmax(mod_nrgy-old_nrgy)
trial_seq = np.tile(mod_seq[best_seq],(n_seq,1))
old_nrgy = np.tile(mod_nrgy[best_seq],(n_seq))
if trial%(10*n_size) == 0: print('after',np.mean(old_nrgy))#,trial_seq[0:5])
print(trial_seq[:10,:10])
#return trial_seq
return enc.fit_transform(trial_seq).toarray()
s = generate_sequences_vp(w0,n,m,l)
def generate_sequences_time_series(s_ini,w,n,m):
i1tab,i2tab = itab(n,m)
l = s_ini.shape[0]
# initial s (categorical variables)
#s_ini = np.random.randint(0,m,size=(l,n)) # integer values
#print(s_ini)
# onehot encoder
enc = OneHotEncoder(n_values=m)
s = enc.fit_transform(s_ini).toarray()
#print(s)
ntrial = 20*m
for t in range(l-1):
h = np.sum(s[t,:]*w[:,:],axis=1)
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
k = np.random.randint(0,m)
for itrial in range(ntrial):
k2 = np.random.randint(0,m)
while k2 == k:
k2 = np.random.randint(0,m)
if np.exp(h[i1+k2]- h[i1+k]) > np.random.rand():
k = k2
s[t+1,i1:i2] = 0.
s[t+1,i1+k] = 1.
return s
# generate non-sequences from time series
#l1 = 100
#s_ini = np.random.randint(0,m,size=(l1,n)) # integer values
#s = np.zeros((l,nm))
#for t in range(l):
# np.random.seed(t+10)
# s[t,:] = generate_sequences_time_series(s_ini,w0,n,m)[-1,:]
print(s.shape)
print(s[:10,:10])
## 2018.11.07: for non sequencial data
def fit_additive(s,n,m):
nloop = 10
i1tab,i2tab = itab(n,m)
nm = n*m
nm1 = nm - m
w_infer = np.zeros((nm,nm))
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
# remove column i
x = np.hstack([s[:,:i1],s[:,i2:]])
x_av = np.mean(x,axis=0)
dx = x - x_av
c = np.cov(dx,rowvar=False,bias=True)
c_inv = linalg.pinv(c,rcond=1e-15)
#print(c_inv.shape)
h = s[:,i1:i2].copy()
for iloop in range(nloop):
h_av = h.mean(axis=0)
dh = h - h_av
dhdx = dh[:,:,np.newaxis]*dx[:,np.newaxis,:]
dhdx_av = dhdx.mean(axis=0)
w = np.dot(dhdx_av,c_inv)
#w = w - w.mean(axis=0)
h = np.dot(x,w.T)
p = np.exp(h)
p_sum = p.sum(axis=1)
#p /= p_sum[:,np.newaxis]
for k in range(m):
p[:,k] = p[:,k]/p_sum[:]
h += s[:,i1:i2] - p
w_infer[i1:i2,:i1] = w[:,:i1]
w_infer[i1:i2,i2:] = w[:,i1:]
return w_infer
w2 = fit_additive(s,n,m)
plt.plot([-1,1],[-1,1],'r--')
plt.scatter(w0,w2)
i1tab,i2tab = itab(n,m)
nloop = 5
nm1 = nm - m
w_infer = np.zeros((nm,nm))
wini = np.random.normal(0.0,1./np.sqrt(nm),size=(nm,nm1))
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
x = np.hstack([s[:,:i1],s[:,i2:]])
y = s.copy()
# covariance[ia,ib]
cab_inv = np.empty((m,m,nm1,nm1))
eps = np.empty((m,m,l))
for ia in range(m):
for ib in range(m):
if ib != ia:
eps[ia,ib,:] = y[:,i1+ia] - y[:,i1+ib]
which_ab = eps[ia,ib,:] !=0.
xab = x[which_ab]
# ----------------------------
xab_av = np.mean(xab,axis=0)
dxab = xab - xab_av
cab = np.cov(dxab,rowvar=False,bias=True)
cab_inv[ia,ib,:,:] = linalg.pinv(cab,rcond=1e-15)
w = wini[i1:i2,:].copy()
for iloop in range(nloop):
h = np.dot(x,w.T)
for ia in range(m):
wa = np.zeros(nm1)
for ib in range(m):
if ib != ia:
which_ab = eps[ia,ib,:] !=0.
eps_ab = eps[ia,ib,which_ab]
xab = x[which_ab]
# ----------------------------
xab_av = np.mean(xab,axis=0)
dxab = xab - xab_av
h_ab = h[which_ab,ia] - h[which_ab,ib]
ha = np.divide(eps_ab*h_ab,np.tanh(h_ab/2.), out=np.zeros_like(h_ab), where=h_ab!=0)
dhdx = (ha - ha.mean())[:,np.newaxis]*dxab
dhdx_av = dhdx.mean(axis=0)
wab = cab_inv[ia,ib,:,:].dot(dhdx_av) # wa - wb
wa += wab
w[ia,:] = wa/m
w_infer[i1:i2,:i1] = w[:,:i1]
w_infer[i1:i2,i2:] = w[:,i1:]
#return w_infer
plt.plot([-1,1],[-1,1],'r--')
plt.scatter(w0,w_infer)
#plt.scatter(w0[0:3,3:],w[0:3,:]) | _____no_output_____ | MIT | old_versions/1main-v4-MCMC-symmetry.ipynb | danhtaihoang/categorical-variables |
1. You are provided the titanic dataset. Load the dataset and perform splitting into training and test sets with 70:30 ratio randomly using test train split.2. Use the Logistic regression created from scratch (from the prev question) in this question as well.3. Data cleaning plays a major role in this question. Report all the methods used by you in the ipynb.--> i. Check for missing valuesii. Drop Columns & Handle missing valuesiii. Create dummies for categorical featuresyou are free to perform other data cleaning to improve your results.4. Report accuracy score, Confusion matrix, heat map, classifiaction report and any other metrics you feel useful. dataset link : https://iiitaphyd-my.sharepoint.com/:f:/g/personal/apurva_jadhav_students_iiit_ac_in/Eictt5_qmoxNqezgQQiMWeIBph4sxlfA6jWAJNPnV2SF9Q?e=mQmYN0 (titanic.csv) | import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, f1_score,confusion_matrix,r2_score
sns.set(style="darkgrid")
df = pd.read_csv('titanic.csv')
df.head()
print('Missing Values in the columns : \n')
print(df.isnull().sum())
df.describe(include='all') | _____no_output_____ | MIT | question3.ipynb | kanishk779/SMAI-2 |
Data cleaning1. **Removal** :-- Remove *Name* column as this attribute does not affect the *Survived* status of the passenger. And moreover we can see that each person has a unique name hence there is no point considering this column.- Remove *Ticket* because there are 681 unique values of ticket and moreover if there is some correlation between the ticket and *Survived* status that can be captured by *Fare*.- Remove *Cabin* as there are lot of missing values | df = df.drop(columns=['Name', 'Ticket', 'Cabin', 'PassengerId'])
s1 = sns.barplot(data = df, y='Survived' , hue='Sex' , x='Sex')
s1.set_title('Male-Female Survival')
plt.show() | _____no_output_____ | MIT | question3.ipynb | kanishk779/SMAI-2 |
Females had a better survival rate than male. | sns.pairplot(df, hue='Survived') | _____no_output_____ | MIT | question3.ipynb | kanishk779/SMAI-2 |
Categorical dataFor categorical variables where no ordinal relationship exists, the integer encoding may not be enough, at best, or misleading to the model at worst.Forcing an ordinal relationship via an ordinal encoding and allowing the model to assume a natural ordering between categories may result in poor performance or unexpected results (predictions halfway between categories).In this case, a one-hot encoding can be applied to the ordinal representation. This is where the integer encoded variable is removed and one new binary variable is added for each unique integer value in the variable. Dummy VariablesThe one-hot encoding creates one binary variable for each category.The problem is that this representation includes redundancy. For example, if we know that [1, 0, 0] represents “blue” and [0, 1, 0] represents “green” we don’t need another binary variable to represent “red“, instead we could use 0 values for both “blue” and “green” alone, e.g. [0, 0].This is called a dummy variable encoding, and always represents C categories with C-1 binary variables. | from numpy import mean
s1 = sns.barplot(data = df, y='Survived' , hue='Embarked' , x='Embarked', estimator=mean)
s1.set_title('Survival vs Boarding place')
plt.show()
carrier_count = df['Embarked'].value_counts()
sns.barplot(x=carrier_count.index, y=carrier_count.values, alpha=0.9)
plt.title('Frequency Distribution of Boarding place')
plt.ylabel('Number of Occurrences', fontsize=12)
plt.xlabel('Places', fontsize=12)
plt.show()
df = pd.get_dummies(df, columns=['Sex', 'Embarked'], prefix=['Sex', 'Embarked'])
df.head()
print('Missing Values in the columns : \n')
print(df.isnull().sum())
df = df.fillna(df['Age'].mean())
print('Missing Values in the columns : \n')
print(df.isnull().sum())
df = df.astype(np.float64)
Y = df['Survived']
Y = np.array(Y)
df.drop(columns=['Survived'], inplace=True)
def standardise(df, col):
df[col] = (df[col] - df[col].mean())/df[col].std()
return df
for col in df.columns:
df = standardise(df, col)
import copy
X = copy.deepcopy(df.to_numpy())
X.shape
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=0.3, shuffle=True)
x_train.shape
class MyLogisticRegression:
def __init__(self, train_data, Y):
self.data = train_data # It is assumed that data is normalized and shuffled (rows, cols)
self.Y = Y[:, np.newaxis]
self.b = np.random.randn()
self.cols = self.data.shape[1]
self.rows = self.data.shape[0]
self.weights = np.random.randn(self.cols, 1) # Initialising weights to 1, shape (cols, 1)
self.num_iterations = 600
self.learning_rate = 0.0001
self.batch_size = 20
self.errors = []
@staticmethod
def sigmoid(x):
return 1/(1 + np.exp(-x))
def calc_mini_batches(self):
new_data = np.hstack((self.data, self.Y))
np.random.shuffle(new_data)
rem = self.rows % self.batch_size
num = self.rows // self.batch_size
till = self.batch_size * num
if num > 0:
dd = np.array(np.vsplit(new_data[ :till, :], num))
X_batch = dd[:, :, :-1]
Y_batch = dd[:, :, -1]
return X_batch, Y_batch
def update_weights(self, X, Y):
Y_predicted = self.predict(X) # Remember that X has data stored along the row for one sample
gradient = np.dot(np.transpose(X), Y_predicted - Y)
self.b = self.b - np.sum(Y_predicted - Y)
self.weights = self.weights - (self.learning_rate * gradient) # vector subtraction
def print_error(self):
Y_Predicted = self.predict(self.data)
class_one = self.Y == 1
class_two = np.invert(class_one)
val = np.sum(np.log(Y_Predicted[class_one]))
val += np.sum(np.log(1 - Y_Predicted[class_two]))
self.errors.append(-val)
print(-val)
def gradient_descent(self):
for j in range(self.num_iterations):
X, Y = self.calc_mini_batches()
num_batches = X.shape[0]
for i in range(num_batches):
self.update_weights(X[i, :, :], Y[i, :][:, np.newaxis]) # update the weights
if (j+1)%100 == 0:
self.print_error()
plt.plot(self.errors)
plt.style.use('ggplot')
plt.xlabel('iteration')
plt.ylabel('')
plt.title('Error Vs iteration')
plt.show()
def predict(self, X):
# X is 2 dimensional array, samples along the rows
return self.sigmoid(np.dot(X, self.weights) + self.b)
reg = MyLogisticRegression(x_train, y_train)
reg.gradient_descent()
y_pred = reg.predict(x_test)
pred = y_pred >= 0.5
pred = pred.astype(int)
print('accuracy : {a}'.format(a=accuracy_score(y_test, pred)))
print('f1 score : {a}'.format(a = f1_score(y_test, pred)))
confusion_matrix(y_test, pred)
sns.heatmap(confusion_matrix(y_test, pred))
from sklearn.metrics import classification_report
print(classification_report(y_test, pred)) | precision recall f1-score support
0.0 0.82 0.86 0.84 153
1.0 0.80 0.75 0.77 115
accuracy 0.81 268
macro avg 0.81 0.80 0.80 268
weighted avg 0.81 0.81 0.81 268
| MIT | question3.ipynb | kanishk779/SMAI-2 |
起手式,導入 numpy, matplotlib | from PIL import Image
import numpy as np
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
matplotlib.style.use('bmh')
matplotlib.rcParams['figure.figsize']=(8,5) | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
使用之前下載的 mnist 資料,載入訓練資料 `train_set` 和測試資料 `test_set` | import gzip
import pickle
with gzip.open('../Week02/mnist.pkl.gz', 'rb') as f:
train_set, validation_set, test_set = pickle.load(f, encoding='latin1')
train_X, train_y = train_set
validation_X, validation_y = validation_set
test_X, test_y = test_set | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
之前的看圖片函數 | from IPython.display import display
def showX(X):
int_X = (X*255).clip(0,255).astype('uint8')
# N*784 -> N*28*28 -> 28*N*28 -> 28 * 28N
int_X_reshape = int_X.reshape(-1,28,28).swapaxes(0,1).reshape(28,-1)
display(Image.fromarray(int_X_reshape))
# 訓練資料, X 的前 20 筆
showX(train_X[:20]) | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
train_set 是用來訓練我們的模型用的我們的模型是很簡單的 logistic regression 模型,用到的參數只有一個 784x10 的矩陣 W 和一個長度 10 的向量 b。我們先用均勻隨機亂數來設定 W 和 b 。 | W = np.random.uniform(low=-1, high=1, size=(28*28,10))
b = np.random.uniform(low=-1, high=1, size=10)
| _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
完整的模型如下將圖片看成是長度 784 的向量 x計算 $Wx+b$, 然後再取 $exp$。 最後得到的十個數值。將這些數值除以他們的總和。我們希望出來的數字會符合這張圖片是這個數字的機率。 $ \Pr(Y=i|x, W, b) = \frac {e^{W_i x + b_i}} {\sum_j e^{W_j x + b_j}}$ 先拿第一筆資料試試看, x 是輸入。 y 是這張圖片對應到的數字(以這個例子來說 y=5)。 | x = train_X[0]
y = train_y[0]
showX(x)
y | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
先計算 $e^{Wx+b} $ | Pr = np.exp(x @ W + b)
Pr.shape | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
然後 normalize,讓總和變成 1 (符合機率的意義) | Pr = Pr/Pr.sum()
Pr | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
由於 $W$ 和 $b$ 都是隨機設定的,所以上面我們算出的機率也是隨機的。正確解是 $y=5$, 運氣好有可能猜中為了要評斷我們的預測的品質,要設計一個評斷誤差的方式,我們用的方法如下(不是常見的方差,而是用熵的方式來算,好處是容易微分,效果好) $ loss = - \log(\Pr(Y=y|x, W,b)) $ 上述的誤差評分方式,常常稱作 error 或者 loss,數學式可能有點費解。實際計算其實很簡單,就是下面的式子 | loss = -np.log(Pr[y])
loss | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
想辦法改進。 我們用一種被稱作是 gradient descent 的方式來改善我們的誤差。因為我們知道 gradient 是讓函數上升最快的方向。所以我們如果朝 gradient 的反方向走一點點(也就是下降最快的方向),那麼得到的函數值應該會小一點。記得我們的變數是 $W$ 和 $b$ (裡面總共有 28*20+10 個變數),所以我們要把 $loss$ 對 $W$ 和 $b$ 裡面的每一個參數來偏微分。還好這個偏微分是可以用手算出他的形式,而最後偏微分的式子也不會很複雜。 $loss$ 展開後可以寫成$loss = \log(\sum_j e^{W_j x + b_j}) - W_i x - b_i$ 對 $k \neq i$ 時, $loss$ 對 $b_k$ 的偏微分是 $$ \frac{e^{W_k x + b_k}}{\sum_j e^{W_j x + b_j}} = \Pr(Y=k | x, W, b)$$對 $k = i$ 時, $loss$ 對 $b_k$ 的偏微分是 $$ \Pr(Y=k | x, W, b) - 1$$ | gradb = Pr.copy()
gradb[y] -= 1
print(gradb) | [ 1.11201478e-03 2.32129668e-06 3.47186834e-03 3.64416088e-03
9.89922844e-01 -9.99616538e-01 4.67890738e-09 3.02581069e-04
1.11720864e-07 1.16063080e-03]
| MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
對 $W$ 的偏微分也不難 對 $k \neq i$ 時, $loss$ 對 $W_{k,t}$ 的偏微分是 $$ \frac{e^{W_k x + b_k} W_{k,t} x_t}{\sum_j e^{W_j x + b_j}} = \Pr(Y=k | x, W, b) x_t$$對 $k = i$ 時, $loss$ 對 $W_{k,t}$ 的偏微分是 $$ \Pr(Y=k | x, W, b) x_t - x_t$$ | print(Pr.shape, x.shape, W.shape)
gradW = x.reshape(784,1) @ Pr.reshape(1,10)
gradW[:, y] -= x | (10,) (784,) (784, 10)
| MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
算好 gradient 後,讓 W 和 b 分別往 gradient 反方向走一點點,得到新的 W 和 b | W -= 0.1 * gradW
b -= 0.1 * gradb | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
再一次計算 $\Pr$ 以及 $loss$ | Pr = np.exp(x @ W + b)
Pr = Pr/Pr.sum()
loss = -np.log(Pr[y])
loss | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
Q* 看看 Pr , 然後找出機率最大者, predict y 值* 再跑一遍上面程序,看看誤差是否變小?* 拿其他的測試資料來看看,我們的 W, b 學到了什麼? 我們將同樣的方式輪流對五萬筆訓練資料來做,看看情形會如何 | W = np.random.uniform(low=-1, high=1, size=(28*28,10))
b = np.random.uniform(low=-1, high=1, size=10)
score = 0
N=50000*20
d = 0.001
learning_rate = 1e-2
for i in range(N):
if i%50000==0:
print(i, "%5.3f%%"%(score*100))
x = train_X[i%50000]
y = train_y[i%50000]
Pr = np.exp( x @ W +b)
Pr = Pr/Pr.sum()
loss = -np.log(Pr[y])
score *=(1-d)
if Pr.argmax() == y:
score += d
gradb = Pr.copy()
gradb[y] -= 1
gradW = x.reshape(784,1) @ Pr.reshape(1,10)
gradW[:, y] -= x
W -= learning_rate * gradW
b -= learning_rate * gradb
| 0 0.000%
50000 87.490%
100000 89.497%
150000 90.022%
200000 90.377%
250000 90.599%
300000 91.002%
350000 91.298%
400000 91.551%
450000 91.613%
500000 91.678%
550000 91.785%
600000 91.792%
650000 91.889%
700000 91.918%
750000 91.946%
800000 91.885%
850000 91.955%
900000 91.954%
950000 92.044%
| MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
結果發現正確率大約是 92%, 但這是對訓練資料而不是對測試資料而且,一筆一筆的訓練資也有點慢,線性代數的特點就是能夠向量運算。如果把很多筆 $x$ 當成列向量組合成一個矩陣(然後叫做 $X$),由於矩陣乘法的原理,我們還是一樣計算 $WX+b$ , 就可以同時得到多筆結果。下面的函數,可以一次輸入多筆 $x$, 同時一次計算多筆 $x$ 的結果和準確率。 | def compute_Pr(X):
Pr = np.exp(X @ W + b)
return Pr/Pr.sum(axis=1, keepdims=True)
def compute_accuracy(Pr, y):
return (Pr.argmax(axis=1)==y).mean() | _____no_output_____ | MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
下面是更新過得訓練過程, 當 i%100000 時,順便計算一下 test accuracy 和 valid accuracy。 | %%timeit -r 1 -n 1
def compute_Pr(X):
Pr = np.exp(X @ W + b)
return Pr/Pr.sum(axis=1, keepdims=True)
def compute_accuracy(Pr, y):
return (Pr.argmax(axis=1)==y).mean()
W = np.random.uniform(low=-1, high=1, size=(28*28,10))
b = np.random.uniform(low=-1, high=1, size=10)
score = 0
N=20000
batch_size = 128
learning_rate = 0.5
for i in range(0, N):
if (i+1)%2000==0:
test_score = compute_accuracy(compute_Pr(test_X), test_y)*100
train_score = compute_accuracy(compute_Pr(train_X), train_y)*100
print(i+1, "%5.2f%%"%test_score, "%5.2f%%"%train_score)
# 隨機選出一些訓練資料出來
rndidx = np.random.choice(train_X.shape[0], batch_size, replace=False)
X, y = train_X[rndidx], train_y[rndidx]
# 一次計算所有的 Pr
Pr = compute_Pr(X)
# 計算平均 gradient
Pr_one_y = Pr-np.eye(10)[y]
gradb = Pr_one_y.mean(axis=0)
gradW = X.T @ (Pr_one_y) / batch_size
# 更新 W 和 ba
W -= learning_rate * gradW
b -= learning_rate * gradb | 2000 90.50% 90.47%
4000 91.17% 91.56%
6000 91.72% 92.03%
8000 91.86% 92.25%
10000 92.03% 92.52%
12000 92.14% 92.88%
14000 92.34% 92.81%
16000 92.29% 92.99%
18000 92.18% 93.13%
20000 92.06% 93.12%
1 loop, best of 1: 1min 8s per loop
| MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
最後得到的準確率是 92%-93%不算完美,不過畢竟這只有一個矩陣而已。 光看數據沒感覺,我們來看看前十筆測試資料跑起來的情形可以看到前十筆只有錯一個 | Pr = compute_Pr(test_X[:10])
pred_y =Pr.argmax(axis=1)
for i in range(10):
print(pred_y[i], test_y[i])
showX(test_X[i]) | 7 7
| MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
看看前一百筆資料中,是哪些情況算錯 | Pr = compute_Pr(test_X[:100])
pred_y = Pr.argmax(axis=1)
for i in range(100):
if pred_y[i] != test_y[i]:
print(pred_y[i], test_y[i])
showX(test_X[i]) | 6 5
| MIT | Week05/From NumPy to Logistic Regression.ipynb | HowardNTUST/HackNTU_Data_2017 |
Скородумов Александр БВТ1904 Лабораторная работа №2 Методы поиска №1 | #Импорты
from IPython.display import HTML, display
from tabulate import tabulate
import random
import time
#Рандомная генерация
def random_matrix(m = 50, n = 50, min_limit = -250, max_limit = 1016):
return [[random.randint(min_limit, max_limit) for _ in range(n)] for _ in range(m)]
#Бинарный поиск
class BinarySearchMap:
def __init__(self):
self.data = [] # хранилище (key, value) значений
def search(self, key):
""" Поиск индекса (во всех случаях лучше левосторонний,
чтоб insert вставлял по убыванию) """
l = 0
r = len(self.data)
while l < r:
m = (l + r) // 2
if self.data[m][0] < key:
l = m + 1
else:
r = m
return l
def __setitem__(self, key, value):
""" Добавить элемент """
index = self.search(key)
# если ключ уже есть в таблице, то надо заменить значение
if index < len(self.data) and self.data[index][0] == key:
self.data[index] = (key, value)
else:
# иначе добавляем новую запись
self.data.insert(index, (key, value))
def __delitem__(self, key):
""" Удалить элемент """
index = self.search(key)
self.data.pop(index)
def __getitem__(self, key):
""" Получить элемент """
index = self.search(key)
found_key, val = self.data[index]
# если найденный индекс выдает запрашиваемый ключ
if found_key == key:
return val
raise KeyError()
#Фибоначчиев поиск
fib_c = [0, 1]
def fib(n):
if len(fib_c) - 1 < n:
fib_c.append(fib(n - 1) + fib(n - 2))
return fib_c[n]
class FibonacciMap(BinarySearchMap):
def search(self, key):
m = 0
while fib(m) < len(self.data):
m += 1
offset = 0
while fib(m) > 1:
i = min(offset + fib(m - 1), len(self.data) - 1)
if key > self.data[i][0]:
offset = i
elif key == self.data[i][0]:
return i
m -= 1
if len(self.data) and self.data[offset][0] < key:
return offset + 1
return 0
#Интерполяционный поиск
def nearest_mid(input_list, lower_bound_index, upper_bound_index, search_value):
return lower_bound_index + \
(upper_bound_index - lower_bound_index) * \
(search_value - input_list[lower_bound_index]) // \
(input_list[upper_bound_index][0] - input_list[lower_bound_index][0])
class InterpolateMap(BinarySearchMap):
def interpolation_search(self, term):
size_of_list = len(self.data) - 1
index_of_first_element = 0
index_of_last_element = size_of_list
while index_of_first_element <= index_of_last_element:
mid_point = nearest_mid(self.data, index_of_first_element, index_of_last_element, term)
if mid_point > index_of_last_element or mid_point < index_of_first_element:
return None
if self.data[mid_point][0] == term:
return mid_point
if term > self.data[mid_point][0]:
index_of_first_element = mid_point + 1
else:
index_of_last_element = mid_point - 1
if index_of_first_element > index_of_last_element:
return None
#Бинарное дерево
class Tree:
def __init__(self, key, value):
self.key = key
self.value = value
self.left = self.right = None
class BinaryTreeMap:
root = None
def insert(self, tree, key, value):
if tree is None:
return Tree(key, value)
if tree.key > key:
tree.left = self.insert(tree.left, key, value)
elif tree.key < key:
tree.right = self.insert(tree.right, key, value)
else:
tree.value = value
return tree
def search(self, tree, key):
if tree is None or tree.key == key:
return tree
if tree.key > key:
return self.search(tree.left, key)
return self.search(tree.right, key)
def __getitem__(self, key):
tree = self.search(self.root, key)
if tree is not None:
return tree.value
raise KeyError()
def __setitem__(self, key, value):
if self.root is None:
self.root = self.insert(self.root, key, value)
else: self.insert(self.root, key, value) | _____no_output_____ | MIT | .ipynb_checkpoints/Skorodumov.Lab2-checkpoint.ipynb | SkorodumovAlex/SIAODLabs |
№2 | #Простое рехеширование
class HashMap:
def __init__(self):
self.size = 0
self.data = []
self._resize()
def _hash(self, key, i):
return (hash(key) + i) % len(self.data)
def _find(self, key):
i = 0;
index = self._hash(key, i);
while self.data[index] is not None and self.data[index][0] != key:
i += 1
index = self._hash(key, i);
return index;
def _resize(self):
temp = self.data
self.data = [None] * (2*len(self.data) + 1)
for item in temp:
if item is not None:
self.data[self._find(item[0])] = item
def __setitem__(self, key, value):
if self.size + 1 > len(self.data) // 2:
self._resize()
index = self._find(key)
if self.data[index] is None:
self.size += 1
self.data[index] = (key, value)
def __getitem__(self, key):
index = self._find(key)
if self.data[index] is not None:
return self.data[index][1]
raise KeyError()
#Рехеширование с помощью псевдослучайных чисел
class RandomHashMap(HashMap):
_rand_c = [5323]
def _rand(self, i):
if len(self._rand_c) - 1 < i:
self._rand_c.append(self._rand(i - 1))
return (123456789 * self._rand_c[i] + 987654321) % 65546
def _hash(self, key, i):
return (hash(key) + self._rand(i)) % len(self.data)
#Метод Цепочек
class ChainMap:
def __init__(self):
self.size = 0
self.data = []
self._resize()
def _hash(self, key):
return hash(key) % len(self.data)
def _insert(self, index, item):
if self.data[index] is None:
self.data[index] = [item]
return True
else:
for i, item_ in enumerate(self.data[index]):
if item_[0] == item[0]:
self.data[index][i] = item
return False
self.data[index].append(item)
return True
def _resize(self):
temp = self.data
self.data = [None] * (2*len(self.data) + 1)
for bucket in temp:
if bucket is not None:
for key, value in bucket:
self._insert(self._hash(key), (key, value))
def __setitem__(self, key, value):
if self.size + 1 > len(self.data) // 1.5:
self._resize()
if self._insert(self._hash(key), (key, value)):
self.size += 1
def __getitem__(self, key):
index = self._hash(key)
if self.data[index] is not None:
for key_, value in self.data[index]:
if key_ == key:
return value
raise KeyError() | _____no_output_____ | MIT | .ipynb_checkpoints/Skorodumov.Lab2-checkpoint.ipynb | SkorodumovAlex/SIAODLabs |
Сравнение алгоритмов | алгоритмы = {
'Бинарный поиск': BinarySearchMap,
'Фибоначчиева поиск': FibonacciMap,
'Интерполяционный поиск': InterpolateMap,
'Бинарное дерево': BinaryTreeMap,
'Простое рехэширование': HashMap,
'Рехэширование с помощью псевдослучайных чисел': RandomHashMap,
'Метод цепочек': ChainMap,
'Стандартная функция поиска': dict
}
затраченное_время = {}
тестовые_набор = random_matrix(50, 1000)
for имя_алгоритма, Таблица in алгоритмы.items():
копия_наборов = тестовые_набор.copy()
время_начало = time.perf_counter()
for набор in копия_наборов:
таблица = Таблица()
for значение, ключ in enumerate(набор):
таблица[ключ] = значение
assert таблица[ключ] == значение, f'Найденный элемент не соответствует записанному'
время_конца = time.perf_counter()
затраченное_время[имя_алгоритма] = (время_конца - время_начало) / len(тестовые_набор)
отсортированная_таблица_затраченного_времени = sorted(затраченное_время.items(), key=lambda kv: kv[1])
tabulate(отсортированная_таблица_затраченного_времени, headers=['Алгоритм','Время'], tablefmt='html', showindex="always") | _____no_output_____ | MIT | .ipynb_checkpoints/Skorodumov.Lab2-checkpoint.ipynb | SkorodumovAlex/SIAODLabs |
№3 | #Вывод результата
def tag(x, color='white'):
return f'<td style="width:24px;height:24px;text-align:center;" bgcolor="{color}">{x}</td>'
th = ''.join(map(tag, ' abcdefgh '))
def chessboard(data):
row = lambda i: ''.join([
tag('<span style="font-size:24px">*</span>' * v,
color='white' if (i+j+1)%2 else 'silver')
for j, v in enumerate(data[i])])
tb = ''.join([f'<tr>{tag(8-i)}{row(i)}{tag(8-i)}</tr>' for i in range(len(data))])
return HTML(f'<table>{th}{tb}{th}</table>')
#Создание доски
arr = [[0] * 8 for i in range(8)]
arr[1][2] = 1
chessboard(arr)
#Алгоритм
def check_place(rows, row, column):
""" Проверяет, если board[column][row] под атакой других ферзей """
for i in range(row):
if rows[i] == column or \
rows[i] - i == column - row or \
rows[i] + i == column + row:
return False
return True
total_shown = 0
def put_queen(rows=[0]*8, row=0):
""" Пытается подобрать место для ферзя, которое не находится под атакой других """
if row == 8: # мы уместили всех 8 ферзей и можем показать доску
arr = [[0] * 8 for i in range(8)]
for row, column in enumerate(rows):
arr[row][column] = 1
return chessboard(arr)
else:
for column in range(8):
if check_place(rows, row, column):
rows[row] = column
board = put_queen(rows, row + 1)
if board: return board
put_queen() | _____no_output_____ | MIT | .ipynb_checkpoints/Skorodumov.Lab2-checkpoint.ipynb | SkorodumovAlex/SIAODLabs |
```Pythonx_train = HDF5Matrix("data.h5", "x_train")x_valid = HDF5Matrix("data.h5", "x_valid")```shapes should be:* (1355578, 432, 560, 1)* (420552, 432, 560, 1) | def gen_data(shape=0, name="input"):
data = np.random.rand(512, 512, 4)
label = data[:,:,-1]
return tf.constant(data.reshape(1,512,512,4).astype(np.float32)), tf.constant(label.reshape(1,512,512,1).astype(np.float32))
## NOTE:
## Tensor 4D -> Batch,X,Y,Z
## Tesnor max. float32!
d, l = gen_data(0,0)
print(d.shape, l.shape)
def unet():
inputs, label = gen_data()
input_shape = inputs.shape
#down0a = Conv2D(16, (3, 3), padding='same')(inputs)
down0a = Conv2D(16, kernel_size=(3, 3), padding='same', input_shape=input_shape)(inputs)
down0a_pool = MaxPooling2D((2, 2), strides=(2, 2))(down0a)
print("down0a.shape:",down0a.shape,"\ndwnpool.shap:", down0a_pool.shape)#?!? letztes != Batch?
#dim0 = Batch
#dim1,dim2 = X,Y
#dim3 = Kanaele
up1 = UpSampling2D((3, 3))(down0a)
print("upsamp.shape:",up1.shape) #UpSampling ändert dim1, dim2... somit (?,X,Y,?) evtl. Batch auf dim0 ?
unet()
def unet2(input_shape, output_length):
inputs = Input(shape=input_shape, name="input")
# 512
down0a = Conv2D(16, (3, 3), padding='same')(inputs)
down0a = BatchNormalization()(down0a)
down0a = Activation('relu')(down0a)
down0a = Conv2D(16, (3, 3), padding='same')(down0a)
down0a = BatchNormalization()(down0a)
down0a = Activation('relu')(down0a)
down0a_pool = MaxPooling2D((2, 2), strides=(2, 2))(down0a)
# 256
down0 = Conv2D(32, (3, 3), padding='same')(down0a_pool)
down0 = BatchNormalization()(down0)
down0 = Activation('relu')(down0)
down0 = Conv2D(32, (3, 3), padding='same')(down0)
down0 = BatchNormalization()(down0)
down0 = Activation('relu')(down0)
down0_pool = MaxPooling2D((2, 2), strides=(2, 2))(down0)
# 128
down1 = Conv2D(64, (3, 3), padding='same')(down0_pool)
down1 = BatchNormalization()(down1)
down1 = Activation('relu')(down1)
down1 = Conv2D(64, (3, 3), padding='same')(down1)
down1 = BatchNormalization()(down1)
down1 = Activation('relu')(down1)
down1_pool = MaxPooling2D((2, 2), strides=(2, 2))(down1)
# 64
down2 = Conv2D(128, (3, 3), padding='same')(down1_pool)
down2 = BatchNormalization()(down2)
down2 = Activation('relu')(down2)
down2 = Conv2D(128, (3, 3), padding='same')(down2)
down2 = BatchNormalization()(down2)
down2 = Activation('relu')(down2)
down2_pool = MaxPooling2D((2, 2), strides=(2, 2))(down2)
# 8
center = Conv2D(1024, (3, 3), padding='same')(down2_pool)
center = BatchNormalization()(center)
center = Activation('relu')(center)
center = Conv2D(1024, (3, 3), padding='same')(center)
center = BatchNormalization()(center)
center = Activation('relu')(center)
# center
up2 = UpSampling2D((2, 2))(center)
up2 = concatenate([down2, up2], axis=3)
up2 = Conv2D(128, (3, 3), padding='same')(up2)
up2 = BatchNormalization()(up2)
up2 = Activation('relu')(up2)
up2 = Conv2D(128, (3, 3), padding='same')(up2)
up2 = BatchNormalization()(up2)
up2 = Activation('relu')(up2)
up2 = Conv2D(128, (3, 3), padding='same')(up2)
up2 = BatchNormalization()(up2)
up2 = Activation('relu')(up2)
# 64
up1 = UpSampling2D((2, 2))(up2)
up1 = concatenate([down1, up1], axis=3)
up1 = Conv2D(64, (3, 3), padding='same')(up1)
up1 = BatchNormalization()(up1)
up1 = Activation('relu')(up1)
up1 = Conv2D(64, (3, 3), padding='same')(up1)
up1 = BatchNormalization()(up1)
up1 = Activation('relu')(up1)
up1 = Conv2D(64, (3, 3), padding='same')(up1)
up1 = BatchNormalization()(up1)
up1 = Activation('relu')(up1)
# 128
up0 = UpSampling2D((2, 2))(up1)
up0 = concatenate([down0, up0], axis=3)
up0 = Conv2D(32, (3, 3), padding='same')(up0)
up0 = BatchNormalization()(up0)
up0 = Activation('relu')(up0)
up0 = Conv2D(32, (3, 3), padding='same')(up0)
up0 = BatchNormalization()(up0)
up0 = Activation('relu')(up0)
up0 = Conv2D(32, (3, 3), padding='same')(up0)
up0 = BatchNormalization()(up0)
up0 = Activation('relu')(up0)
# 256
up0a = UpSampling2D((2, 2))(up0)
up0a = concatenate([down0a, up0a], axis=3)
up0a = Conv2D(16, (3, 3), padding='same')(up0a)
up0a = BatchNormalization()(up0a)
up0a = Activation('relu')(up0a)
up0a = Conv2D(16, (3, 3), padding='same')(up0a)
up0a = BatchNormalization()(up0a)
up0a = Activation('relu')(up0a)
up0a = Conv2D(16, (3, 3), padding='same')(up0a)
up0a = BatchNormalization()(up0a)
up0a = Activation('relu')(up0a)
# 512
output = Conv2D(1, (1, 1), activation='relu')(up0a)
model = Model(inputs=inputs, outputs=output)
model.compile(loss="mean_squared_error", optimizer='adam')
return model
d = unet2((512,512,4),(512,512,1))
| _____no_output_____ | BSD-3-Clause | Examples/simple_U-Net.ipynb | thgnaedi/DeepRain |
Anschließend:```Pythonoutput_length = 1input_length = output_length + 1input_shape=(432, 560, input_length)model_1 = unet(input_shape, output_length)model_1.fit(x_train_1, y_train_1, batch_size = 16, epochs = 25, validation_data=(x_valid_1, y_valid_1))``` | d.summary()
#ToDo: now learn something! | _____no_output_____ | BSD-3-Clause | Examples/simple_U-Net.ipynb | thgnaedi/DeepRain |
Implementing a Neural NetworkIn this exercise we will develop a neural network with fully-connected layers to perform classification, and test it out on the CIFAR-10 dataset. | # A bit of setup
import numpy as np
import matplotlib.pyplot as plt
from cs231n.classifiers.neural_net import TwoLayerNet
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y)))) | /Users/ayush/anaconda2/lib/python2.7/site-packages/matplotlib/font_manager.py:273: UserWarning: Matplotlib is building the font cache using fc-list. This may take a moment.
warnings.warn('Matplotlib is building the font cache using fc-list. This may take a moment.')
| MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
We will use the class `TwoLayerNet` in the file `cs231n/classifiers/neural_net.py` to represent instances of our network. The network parameters are stored in the instance variable `self.params` where keys are string parameter names and values are numpy arrays. Below, we initialize toy data and a toy model that we will use to develop your implementation. | # Create a small net and some toy data to check your implementations.
# Note that we set the random seed for repeatable experiments.
input_size = 4
hidden_size = 10
num_classes = 3
num_inputs = 5
def init_toy_model():
np.random.seed(0)
return TwoLayerNet(input_size, hidden_size, num_classes, std=1e-1)
def init_toy_data():
np.random.seed(1)
X = 10 * np.random.randn(num_inputs, input_size)
y = np.array([0, 1, 2, 2, 1])
return X, y
net = init_toy_model()
X, y = init_toy_data() | _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Forward pass: compute scoresOpen the file `cs231n/classifiers/neural_net.py` and look at the method `TwoLayerNet.loss`. This function is very similar to the loss functions you have written for the SVM and Softmax exercises: It takes the data and weights and computes the class scores, the loss, and the gradients on the parameters. Implement the first part of the forward pass which uses the weights and biases to compute the scores for all inputs. | scores = net.loss(X)
print 'Your scores:'
print scores
print
print 'correct scores:'
correct_scores = np.asarray([
[-0.81233741, -1.27654624, -0.70335995],
[-0.17129677, -1.18803311, -0.47310444],
[-0.51590475, -1.01354314, -0.8504215 ],
[-0.15419291, -0.48629638, -0.52901952],
[-0.00618733, -0.12435261, -0.15226949]])
print correct_scores
print
# The difference should be very small. We get < 1e-7
print 'Difference between your scores and correct scores:'
print np.sum(np.abs(scores - correct_scores)) | Your scores:
[[-0.81233741 -1.27654624 -0.70335995]
[-0.17129677 -1.18803311 -0.47310444]
[-0.51590475 -1.01354314 -0.8504215 ]
[-0.15419291 -0.48629638 -0.52901952]
[-0.00618733 -0.12435261 -0.15226949]]
correct scores:
[[-0.81233741 -1.27654624 -0.70335995]
[-0.17129677 -1.18803311 -0.47310444]
[-0.51590475 -1.01354314 -0.8504215 ]
[-0.15419291 -0.48629638 -0.52901952]
[-0.00618733 -0.12435261 -0.15226949]]
Difference between your scores and correct scores:
3.68027209255e-08
| MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Forward pass: compute lossIn the same function, implement the second part that computes the data and regularizaion loss. | loss, _ = net.loss(X, y, reg=0.1)
correct_loss = 1.30378789133
# should be very small, we get < 1e-12
print 'Difference between your loss and correct loss:'
print np.sum(np.abs(loss - correct_loss)) | Difference between your loss and correct loss:
1.79856129989e-13
| MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Backward passImplement the rest of the function. This will compute the gradient of the loss with respect to the variables `W1`, `b1`, `W2`, and `b2`. Now that you (hopefully!) have a correctly implemented forward pass, you can debug your backward pass using a numeric gradient check: | from cs231n.gradient_check import eval_numerical_gradient
# Use numeric gradient checking to check your implementation of the backward pass.
# If your implementation is correct, the difference between the numeric and
# analytic gradients should be less than 1e-8 for each of W1, W2, b1, and b2.
loss, grads = net.loss(X, y, reg=0.1)
# these should all be less than 1e-8 or so
for param_name in grads:
f = lambda W: net.loss(X, y, reg=0.1)[0]
param_grad_num = eval_numerical_gradient(f, net.params[param_name], verbose=False)
print param_grad_num.shape
print '%s max relative error: %e' % (param_name, rel_error(param_grad_num, grads[param_name])) | (4, 10)
W1 max relative error: 1.000000e+00
| MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Train the networkTo train the network we will use stochastic gradient descent (SGD), similar to the SVM and Softmax classifiers. Look at the function `TwoLayerNet.train` and fill in the missing sections to implement the training procedure. This should be very similar to the training procedure you used for the SVM and Softmax classifiers. You will also have to implement `TwoLayerNet.predict`, as the training process periodically performs prediction to keep track of accuracy over time while the network trains.Once you have implemented the method, run the code below to train a two-layer network on toy data. You should achieve a training loss less than 0.2. | net = init_toy_model()
stats = net.train(X, y, X, y,
learning_rate=1e-1, reg=1e-5,
num_iters=100, verbose=False)
print 'Final training loss: ', stats['loss_history'][-1]
# plot the loss history
plt.plot(stats['loss_history'])
plt.xlabel('iteration')
plt.ylabel('training loss')
plt.title('Training Loss history')
plt.show() | _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Load the dataNow that you have implemented a two-layer network that passes gradient checks and works on toy data, it's time to load up our favorite CIFAR-10 data so we can use it to train a classifier on a real dataset. | from cs231n.data_utils import load_CIFAR10
def get_CIFAR10_data(num_training=49000, num_validation=1000, num_test=1000):
"""
Load the CIFAR-10 dataset from disk and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 data
cifar10_dir = 'cs231n/datasets/cifar-10-batches-py'
X_train, y_train, X_test, y_test = load_CIFAR10(cifar10_dir)
# Subsample the data
mask = range(num_training, num_training + num_validation)
X_val = X_train[mask]
y_val = y_train[mask]
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean image
mean_image = np.mean(X_train, axis=0)
X_train -= mean_image
X_val -= mean_image
X_test -= mean_image
# Reshape data to rows
X_train = X_train.reshape(num_training, -1)
X_val = X_val.reshape(num_validation, -1)
X_test = X_test.reshape(num_test, -1)
return X_train, y_train, X_val, y_val, X_test, y_test
# Invoke the above function to get our data.
X_train, y_train, X_val, y_val, X_test, y_test = get_CIFAR10_data()
print 'Train data shape: ', X_train.shape
print 'Train labels shape: ', y_train.shape
print 'Validation data shape: ', X_val.shape
print 'Validation labels shape: ', y_val.shape
print 'Test data shape: ', X_test.shape
print 'Test labels shape: ', y_test.shape | _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Train a networkTo train our network we will use SGD with momentum. In addition, we will adjust the learning rate with an exponential learning rate schedule as optimization proceeds; after each epoch, we will reduce the learning rate by multiplying it by a decay rate. | input_size = 32 * 32 * 3
hidden_size = 50
num_classes = 10
net = TwoLayerNet(input_size, hidden_size, num_classes)
# Train the network
stats = net.train(X_train, y_train, X_val, y_val,
num_iters=1000, batch_size=200,
learning_rate=1e-4, learning_rate_decay=0.95,
reg=0.5, verbose=True)
# Predict on the validation set
val_acc = (net.predict(X_val) == y_val).mean()
print 'Validation accuracy: ', val_acc
| _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Debug the trainingWith the default parameters we provided above, you should get a validation accuracy of about 0.29 on the validation set. This isn't very good.One strategy for getting insight into what's wrong is to plot the loss function and the accuracies on the training and validation sets during optimization.Another strategy is to visualize the weights that were learned in the first layer of the network. In most neural networks trained on visual data, the first layer weights typically show some visible structure when visualized. | # Plot the loss function and train / validation accuracies
plt.subplot(2, 1, 1)
plt.plot(stats['loss_history'])
plt.title('Loss history')
plt.xlabel('Iteration')
plt.ylabel('Loss')
plt.subplot(2, 1, 2)
plt.plot(stats['train_acc_history'], label='train')
plt.plot(stats['val_acc_history'], label='val')
plt.title('Classification accuracy history')
plt.xlabel('Epoch')
plt.ylabel('Clasification accuracy')
plt.show()
from cs231n.vis_utils import visualize_grid
# Visualize the weights of the network
def show_net_weights(net):
W1 = net.params['W1']
W1 = W1.reshape(32, 32, 3, -1).transpose(3, 0, 1, 2)
plt.imshow(visualize_grid(W1, padding=3).astype('uint8'))
plt.gca().axis('off')
plt.show()
show_net_weights(net) | _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Tune your hyperparameters**What's wrong?**. Looking at the visualizations above, we see that the loss is decreasing more or less linearly, which seems to suggest that the learning rate may be too low. Moreover, there is no gap between the training and validation accuracy, suggesting that the model we used has low capacity, and that we should increase its size. On the other hand, with a very large model we would expect to see more overfitting, which would manifest itself as a very large gap between the training and validation accuracy.**Tuning**. Tuning the hyperparameters and developing intuition for how they affect the final performance is a large part of using Neural Networks, so we want you to get a lot of practice. Below, you should experiment with different values of the various hyperparameters, including hidden layer size, learning rate, numer of training epochs, and regularization strength. You might also consider tuning the learning rate decay, but you should be able to get good performance using the default value.**Approximate results**. You should be aim to achieve a classification accuracy of greater than 48% on the validation set. Our best network gets over 52% on the validation set.**Experiment**: You goal in this exercise is to get as good of a result on CIFAR-10 as you can, with a fully-connected Neural Network. For every 1% above 52% on the Test set we will award you with one extra bonus point. Feel free implement your own techniques (e.g. PCA to reduce dimensionality, or adding dropout, or adding features to the solver, etc.). | best_net = None # store the best model into this
#################################################################################
# TODO: Tune hyperparameters using the validation set. Store your best trained #
# model in best_net. #
# #
# To help debug your network, it may help to use visualizations similar to the #
# ones we used above; these visualizations will have significant qualitative #
# differences from the ones we saw above for the poorly tuned network. #
# #
# Tweaking hyperparameters by hand can be fun, but you might find it useful to #
# write code to sweep through possible combinations of hyperparameters #
# automatically like we did on the previous exercises. #
#################################################################################
pass
#################################################################################
# END OF YOUR CODE #
#################################################################################
# visualize the weights of the best network
show_net_weights(best_net) | _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Run on the test setWhen you are done experimenting, you should evaluate your final trained network on the test set; you should get above 48%.**We will give you extra bonus point for every 1% of accuracy above 52%.** | test_acc = (best_net.predict(X_test) == y_test).mean()
print 'Test accuracy: ', test_acc | _____no_output_____ | MIT | assignment1/two_layer_net.ipynb | ayush29feb/cs231n |
Social Media Analysis EDA | df = pd.read_csv('./meme_cleaning.csv')
df_sentiment = pd.read_csv('563_df_sentiments.csv')
df_sentiment = df_sentiment.drop(columns=['Unnamed: 0', 'Unnamed: 0.1', 'Unnamed: 0.1.1'])
df_sentiment.head()
#Extract all words that begin with # and turn the results into a dataframe
temp = df_sentiment['Tweet'].str.lower().str.extractall(r"(#\w+)")
temp.columns = ['unnamed']
# Convert the multiple hashtag values into a list
temp = temp.groupby(level = 0)['unnamed'].apply(list)
# Save the result as a feature in the original dataset
df_sentiment['hashtags'] = temp
for i in range(len(df_sentiment)):
if df_sentiment.loc[i, 'No_of_Retweets'] >= 4:
df_sentiment.loc[i, 'No_of_Retweets'] = 4
for i in range(len(df_sentiment)):
if df_sentiment.loc[i, 'No_of_Likes'] >= 10:
df_sentiment.loc[i, 'No_of_Likes'] = 10
retweet_df = df_sentiment.groupby(['No_of_Retweets', 'vaderSentiment']).vaderSentimentScores.agg(count='count').reset_index()
like_df = df_sentiment.groupby(['No_of_Likes', 'vaderSentiment']).vaderSentimentScores.agg(count='count').reset_index()
classify_df = df_sentiment.vaderSentiment.value_counts().reset_index()
df_sentiment.Labels = df_sentiment.Labels.fillna('')
df_likes_dict = df_sentiment.groupby('No_of_Likes').vaderSentimentScores.agg(count='count').to_dict()['count']
df_retweet_dict = df_sentiment.groupby('No_of_Retweets').vaderSentimentScores.agg(count='count').to_dict()['count']
for i in range(len(like_df)):
like_df.loc[i, 'Normalized_count'] = like_df.loc[i, 'count'] / df_likes_dict[like_df.loc[i, 'No_of_Likes']]
for i in range(len(retweet_df)):
retweet_df.loc[i, 'Normalized_count'] = retweet_df.loc[i, 'count'] / df_retweet_dict[retweet_df.loc[i, 'No_of_Retweets']] | _____no_output_____ | MIT | Final_Colab.ipynb | jared-garalde/sme_deploy_heroku |
Sentiment | g = sns.catplot(x = "No_of_Likes", y = "Normalized_count", hue = "vaderSentiment", data = like_df, kind = "bar")
g = sns.catplot(x = "No_of_Retweets", y = "Normalized_count", hue = "vaderSentiment", data = retweet_df, kind = "bar")
plt.pie(classify_df['vaderSentiment'], labels=classify_df['index']);
l = []
for i in range(len(df_sentiment)):
for element in df_sentiment.loc[i, 'Labels'].split():
if element != 'Font':
l.append(element) | _____no_output_____ | MIT | Final_Colab.ipynb | jared-garalde/sme_deploy_heroku |
Word Cloud | wordcloud = WordCloud(width = 800, height = 800,
background_color ='white',
min_font_size = 10).generate(str(l))
plt.figure(figsize = (8, 8), facecolor = None)
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show() | _____no_output_____ | MIT | Final_Colab.ipynb | jared-garalde/sme_deploy_heroku |
Topic Modeling | cv = CountVectorizer(stop_words='english')
data_cv = cv.fit_transform(df.Tweet)
words = cv.get_feature_names()
data_dtm = pd.DataFrame(data_cv.toarray(), columns=cv.get_feature_names())
pickle.dump(cv, open("cv_stop.pkl", "wb"))
data_dtm_transpose = data_dtm.transpose()
sparse_counts = scipy.sparse.csr_matrix(data_dtm_transpose)
corpus = matutils.Sparse2Corpus(sparse_counts)
cv = pickle.load(open("cv_stop.pkl", "rb"))
id2word = dict((v, k) for k, v in cv.vocabulary_.items())
word2id = dict((k, v) for k, v in cv.vocabulary_.items())
d = corpora.Dictionary()
d.id2token = id2word
d.token2id = word2id
lda = gensim.models.ldamodel.LdaModel(corpus=corpus, id2word=id2word, num_topics=3, passes=10)
lda.print_topics()
pyLDAvis.enable_notebook()
vis = pyLDAvis.gensim.prepare(lda, corpus, d)
vis | _____no_output_____ | MIT | Final_Colab.ipynb | jared-garalde/sme_deploy_heroku |
1. We shall use the same dataset used in previous assignment - digits. Make a 80-20 train/test split.[Hint: Explore datasets module from scikit learn] 2. Using scikit learn perform a LDA on the dataset. Find out the number of components in the projected subspace.[Hint: Refer to discriminant analysis module of scikit learn] 3. Transform the dataset and fit a logistic regression and observe the accuracy. Compare it with the previous model based on PCA in terms of accuracy and model complexity.[Hint: Project both the train and test samples to the new subspace] | import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.datasets import load_digits
digits = load_digits()
digits.data
digits.data.shape
digits.target.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(digits.data,digits.target, test_size=0.2, random_state = 5)
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
lda = LinearDiscriminantAnalysis()
lda.fit(X_train,y_train)
predictedlda = lda.predict(X_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test,predicted)
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(random_state=5)
logreg.fit(X_train,y_train)
logregpredict = logreg.predict(X_test)
accuracy_score(y_test, logregpredict)
dfcompare = pd.DataFrame({"Actual":y_test,"Predicted":logregpredict})
dfcompare.sample(10) | _____no_output_____ | MIT | Cls5-Dimentionality Reduction/DimensionalityReduction-CaseStudy2-solution.ipynb | tuhinssam/MLResources |
Project: Create a neural network class---Based on previous code examples, develop a neural network class that is able to classify any dataset provided. The class should create objects based on the desired network architecture:1. Number of inputs2. Number of hidden layers3. Number of neurons per layer4. Number of outputs5. Learning rateThe class must have the train, and predict functions.Test the neural network class on the datasets provided below: Use the input data to train the network, and then pass new inputs to predict on. Print the expected label and the predicted label for the input you used. Print the accuracy of the training after predicting on different inputs.Use matplotlib to plot the error that the train method generates.**Don't forget to install Keras and tensorflow in your environment!**--- Import the needed Packages | import numpy as np
import matplotlib.pyplot as plt
# Needed for the mnist data
from keras.datasets import mnist
from keras.utils import to_categorical | _____no_output_____ | MIT | Neural Network Assignment.ipynb | DeepLearningVision-2019/a3-neural-network-class-munozgce |
Define the class | class NeuralNetwork:
def __init__(self, architecture, alpha):
'''
layers: List of integers which represents the architecture of the network.
alpha: Learning rate.
'''
# TODO: Initialize the list of weights matrices, then store
# the network architecture and learning rate
self.layers = architecture
self.alpha = alpha
np.random.seed(13)
self.FW = np.random.randn(architecture[0], architecture[2])
self.MW = np.empty((architecture[0] - 1, architecture[2], architecture[2]))
for x in range(architecture[0] - 2):
self.MW[x] = np.random.randn(architecture[2], architecture[2])
self.LW = np.random.randn(architecture[2], architecture[3])
self.FB = np.random.randn(architecture[2])
self.MB = np.random.randn(architecture[1] - 1, architecture[2])
self.LB = np.random.randn(architecture[3])
pass
def __repr__(self):
return "NeuralNetwork: {}".format( "-".join(str(l) for l in self.layers))
def softmax(self, X):
expX = np.exp(X)
return expX / expX.sum(axis=1, keepdims=True)
def sigmoid(self, x):
# the sigmoid for a given input value
return 1.0 / (1.0 + np.exp(-x))
def sigmoid_deriv(self, x):
# the derivative of the sigmoid
return x * (1 - x)
def predict(self, inputs):
# TODO: Define the predict function
self.newWeights = np.empty((self.layers[1], inputs.shape[0], self.layers[2]))
self.newWeights[0] = self.sigmoid(np.dot(inputs, self.FW) + self.FB)
for x in range(self.layers[0] - 2):
self.newWeights[x+1] = self.sigmoid(np.dot(self.newWeights[x], self.MW[x]) + self.MB[x])
finalLevel = self.softmax( np.dot(self.newWeights[len(self.newWeights)-1], self.LW) + self.LB)
return finalLevel
def train(self, inputs, labels, epochs = 1000, displayUpdate = 100):
fail = []
for i in range(epochs):
hop = self.predict(inputs)
error = labels - hope
error1 = np.dot(error * self.sigmoid_deriv(hope), self.LW.T)
delta = np.dot(error * self.sigmoid_deriv(hope), self.LW.T) * self.sigmoid_deriv(self.newWeights[len(self.newWeights) - 1])
self.LB += * self.alpha
self.LW += np.dot(self.newWeights[len(self.newWeights) - 1].T, error * self.sigmoid_deriv(hope)) * self.alpha
for x in range(self.layers[1] - 1):
self.MW[(len(self.MW) - 1) - x] += np.dot(self.newWeights[(len(self.newWeights) - 2) - x].T, delta) * self.alpha
delta = np.sum(delta)
self.MB[x] += delta * self.alpha
delta = np.dot(delta, self.MW[(len(self.MW) - 1) - x]) * self.sigmoid_deriv(self.newWeights[(len(self.newWeights) - 2) - x])
self.FW += np.dot(inputs.T, delta) * self.alpha
delta2 = np.sum(delta)
self.FB += delta2 * self.alpha
fail.append(np.mean(np.abs(level_error)))
return fail
| _____no_output_____ | MIT | Neural Network Assignment.ipynb | DeepLearningVision-2019/a3-neural-network-class-munozgce |
Test datasets XOR | # input dataset
XOR_inputs = np.array([
[0,0],
[0,1],
[1,0],
[1,1]
])
# labels dataset
XOR_labels = np.array([[0,1,1,0]]).T
#TODO: Test the class with the XOR data
| _____no_output_____ | MIT | Neural Network Assignment.ipynb | DeepLearningVision-2019/a3-neural-network-class-munozgce |
Multiple classes | # Creates the data points for each class
class_1 = np.random.randn(700, 2) + np.array([0, -3])
class_2 = np.random.randn(700, 2) + np.array([3, 3])
class_3 = np.random.randn(700, 2) + np.array([-3, 3])
feature_set = np.vstack([class_1, class_2, class_3])
labels = np.array([0]*700 + [1]*700 + [2]*700)
one_hot_labels = np.zeros((2100, 3))
for i in range(2100):
one_hot_labels[i, labels[i]] = 1
plt.figure(figsize=(10,10))
plt.scatter(feature_set[:,0], feature_set[:,1], c=labels, s=30, alpha=0.5)
plt.show()
#TODO: Test the class with the multiple classes data
r = NeuralNetwork([2,2,5,3], 0.01)
fails = r.train(feature_set, one_hot_labels, 10000, 1000)
fig, ax = plt.subplots(1,1)
ax.set_ylabel('Error')
ax.plot(fails)
test = np.array([[0,-4]])
print(str(test) + str(network.predict(test))) | Error: 0.47806009636665425
Error: 0.007121601211763323
Error: 0.005938405234795827
Error: 0.005920131593441376
Error: 0.00585185558757003
Error: 0.0049490985751735606
Error: 0.004301948969147726
Error: 0.0038221899933325782
Error: 0.003507891190406313
Error: 0.003280260509683804
| MIT | Neural Network Assignment.ipynb | DeepLearningVision-2019/a3-neural-network-class-munozgce |
On the mnist data set---Train the network to classify hand drawn digits.For this data set, if the training step is taking too long, you can try to adjust the architecture of the network to have fewer layers, or you could try to train it with fewer input. The data has already been loaded and preprocesed so that it can be used with the network.--- | # Load the train and test data from the mnist data set
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# Plot a sample data point
plt.title("Label: " + str(train_labels[0]))
plt.imshow(train_images[0], cmap="gray")
# Standardize the data
# Flatten the images
train_images = train_images.reshape((60000, 28 * 28))
# turn values from 0-255 to 0-1
train_images = train_images.astype('float32') / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype('float32') / 255
# Create one hot encoding for the labels
train_labels = to_categorical(train_labels)
test_labels = to_categorical(test_labels)
# TODO: Test the class with the mnist data. Test the training of the network with the test_images data, and
# record the accuracy of the classification.
r = NeuralNetwork([2,2,5,3], 0.01)
fails = r.train(feature_set, one_hot_labels, 10000, 1000)
fig, ax = plt.subplots(1,1)
ax.set_ylabel('Error')
ax.plot(fails)
test = np.array([[0,-3]])
prf = network.predict(test_images[0:1000])
one_hot_test_labels = to_categorical(test_labels[0:1000])
np.set_printoptions(precision=10, suppress= True, linewidth=75)
guess = np.copy(prf)
guess[guess > 0.5] = 1
guess[guess < 0.5] = 0
fails = []
for index, (guess, label) in enumerate(zip(predictions[0:10], one_hot_test_labels[0:10])):
if not np.array_equal(prediction,label):
fails.append((index, prediction, label))
for img, plot in zip(fails, plots):
plot.imshow(test_images[img[0]].reshape(28,28), cmap = "gray")
plot.set_title(str(img[1])) | Error: 0.47806009636665425
Error: 0.007121601211763323
Error: 0.005938405234795827
Error: 0.005920131593441376
Error: 0.00585185558757003
Error: 0.0049490985751735606
Error: 0.004301948969147726
Error: 0.0038221899933325782
Error: 0.003507891190406313
Error: 0.003280260509683804
| MIT | Neural Network Assignment.ipynb | DeepLearningVision-2019/a3-neural-network-class-munozgce |
Titanic 4 > `Pclass, Sex, Age` | import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('seaborn')
sns.set(font_scale=2.5)
import missingno as msno
import warnings
warnings.filterwarnings('ignore')
%matplotlib inline
df_train=pd.read_csv('C:/Users/ehfus/Downloads/titanic/train.csv')
df_test=pd.read_csv('C:/Users/ehfus/Downloads/titanic/test.csv')
# violin plot
fig,ax=plt.subplots(1,2,figsize=(18,8))
sns.violinplot('Pclass','Age',hue='Survived',data=df_train,sacle='count',split=True,ax=ax[0])
ax[0].set_title('Pclass and Age vs Survived')
ax[0].set_yticks(range(0,110,10))
sns.violinplot('Sex','Age',hue='Survived',data=df_train,scale='count',split=True, ax=ax[1])
ax[1].set_title('Sexx and Age vs Survived')
ax[1].set_yticks(range(0,110,10))
plt.show() | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
- scale에도 option이 여러가지 있음, google에서 확인해볼 것 > `Embarked : 탑승한 항구` | f,ax=plt.subplots(1,1,figsize=(7,7))
df_train[['Embarked','Survived']]\
.groupby(['Embarked'], as_index=True).mean()\
.sort_values(by='Survived', ascending=False)\
.plot.bar(ax=ax) | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
- `sort_values` 또는 `sort_index`도 사용 가능 | f,ax=plt.subplots(2,2,figsize=(20,15)) #2차원임/ 1,2는 1차원
sns.countplot('Embarked',data=df_train, ax=ax[0,0])
ax[0,0].set_title('(1) No. Of Passengers Boared')
sns.countplot('Embarked',hue='Sex',data=df_train, ax=ax[0,1])
ax[0,1].set_title('(2) Male-Female split for embarked')
sns.countplot('Embarked', hue='Survived', data=df_train, ax=ax[1,0])
ax[1,0].set_title('(3) Embarked vs Survived')
sns.countplot('Embarked', hue='Pclass', data=df_train, ax=ax[1,1])
ax[1,1].set_title('(4) Embarked vs Survived')
plt.subplots_adjust(wspace=0.2, hspace=0.5) # 상하좌우간격 맞춰줌
plt.show() | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
> `Family - SibSp + ParCh` | df_train['FamilySize']=df_train['SibSp'] + df_train['Parch'] + 1
print('Maximum size of Family : ',df_train['FamilySize'].max())
print('Minimum size of Family : ',df_train['FamilySize'].min()) | Maximum size of Family : 11
Minimum size of Family : 1
| Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
- Pandas series는 연산이 가능 | f,ax=plt.subplots(1,3,figsize=(40,10))
sns.countplot('FamilySize', data=df_train, ax=ax[0])
ax[0].set_title('(1) No. Of Passenger Boarded', y=1.02)
sns.countplot('FamilySize', hue='Survived',data=df_train, ax=ax[1])
ax[1].set_title('(2) Survived countplot depending on FamilySize', y=1.02)
df_train[['FamilySize','Survived']].groupby(['FamilySize'],as_index=True).mean().sort_values(by='Survived',ascending=False).plot.bar(ax=ax[2])
ax[2].set_title('(3) Survived rate depending on FamilySize',y=1.02)
plt.subplots_adjust(wspace=0.2,hspace=0.5)
plt.show() | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
> `Fare : 요금, 연속형 변수` - distplot ?? 시리즈에 히스토그램을 그려줌,Skewness? 왜도임 + 첨도도 있음- 왜도? 첨도?- python에서 나타내는 함수는? | fig,ax=plt.subplots(1,1,figsize=(8,8))
g=sns.distplot(df_train['Fare'], color='b',label='Skewness{:.2f}'.format(df_train['Fare'].skew()),ax=ax)
g=g.legend(loc='best') | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
- skewness가 5정도로 꽤 큼 -> 좌로 많이 치우쳐져 있음 -> 그대로 모델에 학습시키면 성능이 낮아질 수 있음 | df_train['Fare']=df_train['Fare'].map(lambda i: np.log(i) if i>0 else 0) | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
df_train['Fare']의 값을 적절하게 변형 중 | fig,ax=plt.subplots(1,1,figsize=(8,8))
g=sns.distplot(df_train['Fare'], color='b',label='Skewness{:.2f}'.format(df_train['Fare'].skew()),ax=ax)
g=g.legend(loc='best') | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
이런 작업(log로 변환)을 통해 skewness가 0으로 근접하게 해주었음 | df_train['Ticket'].value_counts() | _____no_output_____ | Apache-2.0 | _notebooks/2021-12-26-titanic.ipynb | rhkrehtjd/kaggle |
Section 1.2: Dimension reduction and principal component analysis (PCA)One of the iron laws of data science is know as the "curse of dimensionality": as the number of considered features (dimensions) of a feature space increases, the number of data configurations can grow exponentially and thus the number observations (data points) needed to account for these configurations must also increase. Because this fact of life has huge ramifications for the time, computational effort, and memory required it is often desirable to reduce the number of dimensions we have to work with.One way to accomplish this is by reducing the number of features considered in an analysis. After all, not all features are created equal, and some yield more insight for a given analysis than others. While this type of feature engineering is necessary in any data-science project, we can really only take it so far; up to a point, considering more features can often increase the accuracy of a classifier. (For example, consder how many features could increase the accuracy of classifying images as cats or dogs.) PCA in theoryAnother way to reduce the number of dimensions that we have to work with is by projecting our feature space into a lower dimensional space. The reason why we can do this is that in most real-world problems, data points are not spread uniformly across all dimensions. Some features might be near constant, while others are highly correlated, which means that those data points lie close to a lower-dimensional subspace.In the image below, the data points are not spread across the entire plane, but are nicely clumped, roughly in an oval. Because the cluster (or, indeed, any cluster) is roughly elliptical, it can be mathematically described by two values: its major (long) axis and its minor (short) axis. These axes form the *principal components* of the cluster. In fact, we can construct a whole new feature space around this cluster, defined by two *eigenvectors* (the vectors that define the linear transformation to this new feature space), $c_{1}$ and $c_{2}$. Better still, we don't have to consider all of the dimensions of this new space. Intuitively, we can see that most of the points lie on or close to the line that runs through $c_{1}$. So, if we project the cluster down from two dimensions to that single dimension, we capture most of the information about this data sense while simplifying our analysis. This ability to extract most of the information from a dataset by considering only a fraction of its definitive eigenvectors forms the heart of principal component analysis (PCA). Import modules and datasetYou will need to clean and prepare the data in order to conduct PCA on it, so pandas will be essential. You will also need NumPy, a bit of Scikit Learn, and pyplot. | import pandas as pd
import numpy as np
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
%matplotlib inline | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
The dataset we’ll use here is the same one drawn from the [U.S. Department of Agriculture National Nutrient Database for Standard Reference](https://www.ars.usda.gov/northeast-area/beltsville-md-bhnrc/beltsville-human-nutrition-research-center/nutrient-data-laboratory/docs/usda-national-nutrient-database-for-standard-reference/) that you prepared in Section 1.1. Remember to set the encoding to `latin_1` (for those darn µg). | df = pd.read_csv('Data/USDA-nndb-combined.csv', encoding='latin_1') | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
We can check the number of columns and rows using the `info()` method for the `DataFrame`. | df.info() | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
> **Exercise**>> Can you think of a more concise way to check the number of rows and columns in a `DataFrame`? (***Hint:*** Use one of the [attributes](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) of the `DataFrame`.) Handle `null` valuesBecause this is a real-world dataset, it is a safe bet that it has `null` values in it. We could first check to see if this is true. However, later on in this section, we will have to transform our data using a function that cannot use `NaN` values, so we might as well drop rows containing those values. > **Exercise**>> Drop rows from the `DataFrame` that contain `NaN` values. (If you need help remembering which method to use, see [this page](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.dropna.html).) > **Exercise solution**>> The correct code to use is `df = df.dropna()`. Now let’s see how many rows we have left. | df.shape | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
Dropping those rows eliminated 76 percent of our data (8989 entries to 2190). An imperfect state of affairs, but we still have enough for our purposes in this section.> **Key takeaway:** Another solution to removing `null` values is to impute values for them, but this can be tricky. Should we handle missing values as equal to 0? What about a fatty food with `NaN` for `Lipid_Tot_(g)`? We could try taking the averages of values surrounding a `NaN`, but what about foods that are right next to rows containing foods from radically different food groups? It is possible to make justifiable imputations for missing values, but it can be important to involve subject-matter experts (SMEs) in that process. Split off descriptive columnsOut descriptive columns (such as `FoodGroup` and `Shrt_Desc`) pose challenges for us when it comes time to perform PCA because they are categorical rather than numerical features, so we will split our `DataFrame` in to one containing the descriptive information and one containing the nutritional information. | desc_df = df.iloc[:, [0, 1, 2]+[i for i in range(50,54)]]
desc_df.set_index('NDB_No', inplace=True)
desc_df.head() | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
> **Question**>> Why was it necessary to structure the `iloc` method call the way we did in the code cell above? What did it accomplish? Why was it necessary set the `desc_df` index to `NDB_No`? | nutr_df = df.iloc[:, :-5]
nutr_df.head() | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
> **Question**>> What did the `iloc` syntax do in the code cell above? | nutr_df = nutr_df.drop(['FoodGroup', 'Shrt_Desc'], axis=1) | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
> **Exercise**>> Now set the index of `nutr_df` to use `NDB_No`. > **Exercise solution**>> The correct code for students to use here is `nutr_df.set_index('NDB_No', inplace=True)`. Now let’s take a look at `nutr_df`. | nutr_df.head() | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
Check for correlation among featuresOne thing that can skew our classification results is correlation among our features. Recall that the whole reason that PCA works is that it exploits the correlation among data points to project our feature-space into a lower-dimensional space. However, if some of our features are highly correleted to begin with, these relationships might create spurious clusters of data in our PCA.The code to check for correlations in our data isn't long, but it takes too long (up to 10 to 20 minutes) to run for a course like this. Instead, the table below shows the output from that code:| | column | row | corr ||--:|------------------:|------------------:|-----:|| 0 | Folate\_Tot\_(µg) | Folate\_DFE\_(µg) | 0.98 || 1 | Folic\_Acid\_(µg) | Folate\_DFE\_(µg) | 0.95 || 2 | Folate\_DFE\_(µg) | Folate\_Tot\_(µg) | 0.98 || 3 | Vit\_A\_RAE | Retinol\_(µg) | 0.99 || 4 | Retinol\_(µg) | Vit\_A\_RAE | 0.99 || 5 | Vit\_D\_µg | Vit\_D\_IU | 1 || 6 | Vit\_D\_IU | Vit\_D\_µg | 1 |As it turns out, dropping `Folate_DFE_(µg)`, `Vit_A_RAE`, and `Vit_D_IU` will eliminate the correlations enumerated in the table above. | nutr_df.drop(['Folate_DFE_(µg)', 'Vit_A_RAE', 'Vit_D_IU'],
inplace=True, axis=1)
nutr_df.head() | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
Normalize and center the dataOur numeric data comes in a variety of mass units (grams, milligrams, and micrograms) and one energy unit (kilocalories). In order to make an apples-to-apples comparison (pun intended) of the nutritional data, we need to first *normalize* the data and make it more normally distributed (that is, make the distribution of the data look more like a familiar bell curve).To help see why we need to normalize the data, let's look at a histogram of all of the columns. | ax = nutr_df.hist(bins=50, xlabelsize=-1, ylabelsize=-1, figsize=(11,11)) | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
Not a bell curve in sight. Worse, a lot of the data is clumped at or around 0. We will use the Box-Cox Transformation on the data, but it requires strictly positive input, so we will add 1 to every value in each column. | nutr_df = nutr_df + 1 | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
Now for the transformation. The [Box-Cox Transformation](https://www.statisticshowto.datasciencecentral.com/box-cox-transformation/) performs the transformation $y(\lambda) = \dfrac{y^{\lambda}-1}{\lambda}$ for $\lambda \neq 0$ and $y(\lambda) = log y$ for $\lambda = 0$ for all values $y$ in a given column. SciPy has a particularly useful `boxcox()` function that can automatically calculate the $\lambda$ for each column that best normalizes the data in that column. (However, it is does not support `NaN` values; scikit-learn has a comparable `boxcox()` function that is `NaN`-safe, but it is not available on the version of scikit-learn that comes with Azure notebooks.) | from scipy.stats import boxcox
nutr_df_TF = pd.DataFrame(index=nutr_df.index)
for col in nutr_df.columns.values:
nutr_df_TF['{}_TF'.format(col)] = boxcox(nutr_df.loc[:, col])[0] | _____no_output_____ | MIT | Machine Learning 2_Using Advanced Machine Learning Models/Reference Material/190053-Reactors-DS-Tr2-Sec1-2-PCA.ipynb | raspyweather/Reactors |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.