markdown stringlengths 0 1.02M | code stringlengths 0 832k | output stringlengths 0 1.02M | license stringlengths 3 36 | path stringlengths 6 265 | repo_name stringlengths 6 127 |
|---|---|---|---|---|---|
Data cleaning* Filtra horários de aula* remover linhas incompletas (sistema fora do ar)* remover oulier (falhas na coleta de dados).* remover dias não-letivos* remover dias com falhas na medição (sistema fora do ar) | processed = raw.dropna()
processed = processed.set_index(pd.to_datetime (processed['momento'])).drop('momento', axis=1)
(ax1, ax2, ax3) = processed['2019-05-20 00:00:00' : '2019-05-25 00:00:00'].plot(subplots=True, sharex=True)
ax1.legend(loc='upper left')
ax2.legend(loc='upper left')
ax3.legend(loc='upper left')
#ax1... | _____no_output_____ | MIT | artificial_intelligence/01 - ConsumptionRegression/All campus/Fpolis.ipynb | LeonardoSanBenitez/LorisWeb |
Linear Regression | model1 = LinearRegression()
model1.fit (X_train, y_train)
pd.DataFrame(model1.coef_,X.columns,columns=['Coefficient'])
from sklearn import metrics
y_hat1 = model1.predict(X_test)
print ("MAE: ", metrics.mean_absolute_error(y_test, y_hat1))
print ("RMSE: ", np.sqrt(metrics.mean_squared_erro... | _____no_output_____ | MIT | artificial_intelligence/01 - ConsumptionRegression/All campus/Fpolis.ipynb | LeonardoSanBenitez/LorisWeb |
Random Forest | import sklearn.metrics as metrics
import math
from sklearn.ensemble import RandomForestRegressor
mae1 = {}
mae2 = {}
for k in range(1,15, 1):
model2 = RandomForestRegressor(max_depth=k, n_estimators=100, criterion='mae').fit(X_train,y_train)
y_hat = model2.predict(X_train)
mae1[k] = metrics.mean_absolute_er... | _____no_output_____ | MIT | artificial_intelligence/01 - ConsumptionRegression/All campus/Fpolis.ipynb | LeonardoSanBenitez/LorisWeb |
IntroductionLinear Regression is one of the most famous and widely used machine learning algorithms out there. It assumes that the target variable can be explained as a linear combination of the input features. What does this mean? It means that the target can be viewed as a weighted sum of each feature. Let’s use a p... | fixed_price = 5
ingredient_costs = {"meat": 10,
"fish": 13,
"vegetables": 2,
"fries": 3}
def price(**ingredients):
""" returns the price of a dish """
cost = 0
for name, quantity in ingredients.items():
cost += ingredient_costs[name... | _____no_output_____ | MIT | notebooks/Learning Units/Linear Regression/Linear Regression - Chapter 1 - Introduction.ipynb | ValentinCalomme/skratch |
The Iris flower data set is a multivariate data set introduced by the British statistician and biologist Ronald Fisher in his 1936 paper The use of multiple measurements in taxonomic problems. The dataset consists of 50 samples from each of three species of Iris (Iris Setosa, Iris virginica, and Iris versicolor). Four ... | import numpy as np
import pandas as pd
from pandas import Series, DataFrame
import seaborn as sns
import matplotlib.pyplot as plt
iris = pd.read_csv("iris.csv")
iris.head() | _____no_output_____ | MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
*We can see that we have a column named ID that we donot need , so let's drop it !* | iris.drop("Id", axis=1, inplace = True)
iris.info()
figure = iris[iris.Species == 'Iris-setosa'].plot(kind='scatter', x='SepalLengthCm', y='SepalWidthCm', color='red', label='Setosa')
iris[iris.Species == 'Iris-versicolor'].plot(kind='scatter', x='SepalLengthCm', y='SepalWidthCm', color='blue', label='Versicolor', ax=f... | _____no_output_____ | MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
Splitting The Data into Training And Testing Dataset | train, test = train_test_split(iris, test_size=0.2)
print(train.shape)
print(test.shape)
train_X = train[['SepalLengthCm','SepalWidthCm','PetalLengthCm','PetalWidthCm']]
train_y = train.Species
test_X = test[['SepalLengthCm','SepalWidthCm','PetalLengthCm','PetalWidthCm']]
test_y = test.Species | _____no_output_____ | MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
1. Logistic Regression | model1 = LogisticRegression()
model1.fit(train_X, train_y)
prediction1 = model1.predict(test_X)
print('Accuracy of Logistic Regression is: ', metrics.accuracy_score(prediction1, test_y)) | Accuracy of Logistic Regression is: 0.9333333333333333
| MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
2. SVM Classifier | model2 = svm.SVC()
model2.fit(train_X, train_y)
prediction2 = model2.predict(test_X)
print('Accuracy of SVM is: ', metrics.accuracy_score(prediction2, test_y)) | Accuracy of SVM is: 0.9666666666666667
| MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
3. K-Nearest Neighbors | model3 = KNeighborsClassifier(n_neighbors=3) # this examines 3 neighbors
model3.fit(train_X, train_y)
prediction3 = model3.predict(test_X)
print('Accuracy of KNN is: ', metrics.accuracy_score(prediction3, test_y)) | Accuracy of KNN is: 0.9666666666666667
| MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
4. Decision Tree | model4 = DecisionTreeClassifier()
model4.fit(train_X, train_y)
prediction4 = model4.predict(test_X)
print('Accuracy of Decision Tree is: ', metrics.accuracy_score(prediction4, test_y)) | Accuracy of Decision Tree is: 0.9
| MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
5. XGBoost | model5 = xgb.XGBClassifier()
model5.fit(train_X, train_y)
prediction5 = model5.predict(test_X)
print('Accuracy of xgb classifier is: ', metrics.accuracy_score(prediction5, test_y)) | Accuracy of xgb classifier is: 0.9333333333333333
| MIT | Machine Learning Problem-Statements/Iris/Iris_Dataset_Machine_Learning.ipynb | JukMR/Hacktoberfest2020 |
Lagged Price Machine Learning Testing | df1_50 = pd.read_csv(
Path("./Data/QM_50_6month.csv")
)
tickers = list(df1_50["Tickers"])
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
from skle... | _____no_output_____ | Unlicense | Back Test 2021_02-2021_04.ipynb | tonghuang-uw/Project_2 |
SVC | poly_kernel_svm_clf = Pipeline([
("scaler", StandardScaler()),
("svm_clf", SVC())
])
poly_kernel_svm_clf.fit(historical_train[cols],np.sign(historical_train["return"]))
price_3month["prediction"] = model.predict(price_3month[cols])
price_3month["prediction"].value_counts()
print(classification_report(pri... | _____no_output_____ | Unlicense | Back Test 2021_02-2021_04.ipynb | tonghuang-uw/Project_2 |
SMA | %%time
short_win = 5
long_win = 15
weighting = 1/50
strat = np.zeros(63)
actual = np.zeros(63)
for ticker in tickers:
historical = yf.Ticker(ticker).history(period="max")
historical["return"] = historical["Close"].pct_change()
historical["SMA_short"] = historical["Close"].rolling(window=short_win).... | _____no_output_____ | Unlicense | Back Test 2021_02-2021_04.ipynb | tonghuang-uw/Project_2 |
EMA | short_win = 12
long_win = 26
strat = np.zeros(63)
actual = np.zeros(63)
for ticker in tickers:
historical = yf.Ticker(ticker).history(period="2y")
historical["return"] = historical["Close"].pct_change()
historical["exp1"] = historical["Close"].ewm(span=short_win, adjust=False).mean().shift()
historic... | _____no_output_____ | Unlicense | Back Test 2021_02-2021_04.ipynb | tonghuang-uw/Project_2 |
MACD | short_win = 12
long_win = 26
signal_line = 9
strat = np.zeros(63)
actual = np.zeros(63)
for ticker in tickers:
historical = yf.Ticker(ticker).history(period="2y")
historical["return"] = historical["Close"].pct_change()
historical["exp1"] = historical["Close"].ewm(span=short_win, adjust=False).mean().shift... | _____no_output_____ | Unlicense | Back Test 2021_02-2021_04.ipynb | tonghuang-uw/Project_2 |
Tutorial 5: Trace - training control and debuggingIn this tutorial, we will talk about another important concept in FastEstimator - Trace.`Trace` is a class contains has 6 event functions below, each event function will be executed on different events of training loop when putting `Trace` inside `Estimator`. If you ar... | import tempfile
import numpy as np
import tensorflow as tf
import fastestimator as fe
from fastestimator.architecture import LeNet
from fastestimator.estimator.trace import Accuracy, ModelSaver
from fastestimator.network.loss import SparseCategoricalCrossentropy
from fastestimator.network.model import FEModel, ModelO... | _____no_output_____ | Apache-2.0 | tutorial/t05_trace_debug_training.ipynb | AriChow/fastestimator |
define trace | from fastestimator.estimator.trace import Trace
class ShowPred(Trace):
def on_batch_end(self, state):
if state["mode"] == "train":
batch_data = state["batch"]
print("step: {}".format(state["batch_idx"]))
print("batch data has following keys: {}".format(list(batch_data.ke... | ______ __ ______ __ _ __
/ ____/___ ______/ /_/ ____/____/ /_(_)___ ___ ____ _/ /_____ _____
/ /_ / __ `/ ___/ __/ __/ / ___/ __/ / __ `__ \/ __ `/ __/ __ \/ ___/
/ __/ / /_/ (__ ) /_/ /___(__ ) /_/ / / / / / / /_/ / /_/ /_/ / /
/_/ \__,_/____/\__/_____/... | Apache-2.0 | tutorial/t05_trace_debug_training.ipynb | AriChow/fastestimator |
Flopy MODFLOW Boundary ConditionsFlopy has a new way to enter boundary conditions for some MODFLOW packages. These changes are substantial. Boundary conditions can now be entered as a list of boundaries, as a numpy recarray, or as a dictionary. These different styles are described in this notebook.Flopy also now re... | #begin by importing flopy
import os
import sys
import numpy as np
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
workspace = os.path.join('data')
#make sure workspace directory exists
if n... | flopy is installed in /Users/jdhughes/Documents/Development/flopy_git/flopy_us/flopy
3.7.3 (default, Mar 27 2019, 16:54:48)
[Clang 4.0.1 (tags/RELEASE_401/final)]
numpy version: 1.16.2
flopy version: 3.2.12
| CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
List of Boundaries Boundary condition information is passed to a package constructor as stress_period_data. In its simplest form, stress_period_data can be a list of individual boundaries, which themselves are lists. The following shows a simple example for a MODFLOW River Package boundary: | stress_period_data = [
[2, 3, 4, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 5, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 6, 10.7, 5000., -5.7], #layer, row, column, stage,... | _____no_output_____ | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
If we look at the River Package created here, you see that the layer, row, and column numbers have been increased by one. | !head -n 10 'data/test.riv' | # RIV package for MODFLOW-2005, generated by Flopy.
3 0
3 0 # stress period 1
3 4 5 10.7 5000.0 -5.7
3 4 6 10.7 5000.0 -5.7
3 4 7 10... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
If this model had more than one stress period, then Flopy will assume that this boundary condition information applies until the end of the simulation | m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
dis = flopy.modflow.ModflowDis(m, nper=3)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data)
m.write_input()
!head -n 10 'data/test.riv' | # RIV package for MODFLOW-2005, generated by Flopy.
3 0
3 0 # stress period 1
3 4 5 10.7 5000.0 -5.7
3 4 6 10.7 5000.0 -5.7
3 4 7 10... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
Recarray of BoundariesNumpy allows the use of recarrays, which are numpy arrays in which each column of the array may be given a different type. Boundary conditions can be entered as recarrays. Information on the structure of the recarray for a boundary condition package can be obtained from that particular package.... | riv_dtype = flopy.modflow.ModflowRiv.get_default_dtype()
print(riv_dtype) | [('k', '<i8'), ('i', '<i8'), ('j', '<i8'), ('stage', '<f4'), ('cond', '<f4'), ('rbot', '<f4')]
| CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
Now that we know the structure of the recarray that we want to create, we can create a new one as follows. | stress_period_data = np.zeros((3), dtype=riv_dtype)
stress_period_data = stress_period_data.view(np.recarray)
print('stress_period_data: ', stress_period_data)
print('type is: ', type(stress_period_data)) | stress_period_data: [(0, 0, 0, 0., 0., 0.) (0, 0, 0, 0., 0., 0.) (0, 0, 0, 0., 0., 0.)]
type is: <class 'numpy.recarray'>
| CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
We can then fill the recarray with our boundary conditions. | stress_period_data[0] = (2, 3, 4, 10.7, 5000., -5.7)
stress_period_data[1] = (2, 3, 5, 10.7, 5000., -5.7)
stress_period_data[2] = (2, 3, 6, 10.7, 5000., -5.7)
print(stress_period_data)
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data... | # RIV package for MODFLOW-2005, generated by Flopy.
3 0
3 0 # stress period 1
3 4 5 10.7 5000.0 -5.7
3 4 6 10.7 5000.0 -5.7
3 4 7 10... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
As before, if we have multiple stress periods, then this recarray will apply to all of them. | m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
dis = flopy.modflow.ModflowDis(m, nper=3)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data)
m.write_input()
!head -n 10 'data/test.riv' | # RIV package for MODFLOW-2005, generated by Flopy.
3 0
3 0 # stress period 1
3 4 5 10.7 5000.0 -5.7
3 4 6 10.7 5000.0 -5.7
3 4 7 10... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
Dictionary of BoundariesThe power of the new functionality in Flopy3 is the ability to specify a dictionary for stress_period_data. If specified as a dictionary, the key is the stress period number (**as a zero-based number**), and the value is either a nested list, an integer value of 0 or -1, or a recarray for that... | sp1 = [
[2, 3, 4, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 5, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 6, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
]
print(sp1)
riv_dtype = fl... | # RIV package for MODFLOW-2005, generated by Flopy.
3 0
0 0 # stress period 1
3 0 # stress period 2
3 4 5 10.7 5000.0 -5.7
3 4 6 10.7 5000.0 -5.7
... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
MODFLOW Auxiliary VariablesFlopy works with MODFLOW auxiliary variables by allowing the recarray to contain additional columns of information. The auxiliary variables must be specified as package options as shown in the example below.In this example, we also add a string in the last column of the list in order to nam... | #create an empty array with an iface auxiliary variable at the end
riva_dtype = [('k', '<i8'), ('i', '<i8'), ('j', '<i8'),
('stage', '<f4'), ('cond', '<f4'), ('rbot', '<f4'),
('iface', '<i4'), ('boundname', object)]
riva_dtype = np.dtype(riva_dtype)
stress_period_data = np.zeros((3), dtype... | # RIV package for MODFLOW-2005, generated by Flopy.
3 0 aux iface
3 0 # stress period 1
3 4 5 10.7 5000.0 -5.7 1 riv1
3 4 6 10.7 5000.0 -5.7 2 ... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
Working with Unstructured GridsFlopy can create an unstructured grid boundary condition package for MODFLOW-USG. This can be done by specifying a custom dtype for the recarray. The following shows an example of how that can be done. | #create an empty array based on nodenumber instead of layer, row, and column
rivu_dtype = [('nodenumber', '<i8'), ('stage', '<f4'), ('cond', '<f4'), ('rbot', '<f4')]
rivu_dtype = np.dtype(rivu_dtype)
stress_period_data = np.zeros((3), dtype=rivu_dtype)
stress_period_data = stress_period_data.view(np.recarray)
print('st... | data
# RIV package for MODFLOW-2005, generated by Flopy.
3 0
3 0 # stress period 1
77 10.7 5000.0 -5.7
245 10.7 5000.0 -5.7
450034 10.7 5000.0 -5.7
| CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
Combining two boundary condition packages | ml = flopy.modflow.Modflow(modelname="test",model_ws=workspace)
dis = flopy.modflow.ModflowDis(ml,10,10,10,10)
sp_data1 = {3: [1, 1, 1, 1.0],5:[1,2,4,4.0]}
wel1 = flopy.modflow.ModflowWel(ml, stress_period_data=sp_data1)
ml.write_input()
!head -n 10 'data/test.wel'
sp_data2 = {0: [1, 1, 3, 3.0],8:[9,2,4,4.0]}
wel2 = fl... | WARNING: unit 20 of package WEL already in use
****Warning -- two packages of the same type: <class 'flopy.modflow.mfwel.ModflowWel'> <class 'flopy.modflow.mfwel.ModflowWel'>
replacing existing Package...
# WEL package for MODFLOW-2005, generated by Flopy.
1 0
1 0 # stress period ... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
Now we create a third wel package, using the ```MfList.append()``` method: | wel3 = flopy.modflow.ModflowWel(ml,stress_period_data=\
wel2.stress_period_data.append(
wel1.stress_period_data))
ml.write_input()
!head -n 10 'data/test.wel' | WARNING: unit 20 of package WEL already in use
****Warning -- two packages of the same type: <class 'flopy.modflow.mfwel.ModflowWel'> <class 'flopy.modflow.mfwel.ModflowWel'>
replacing existing Package...
# WEL package for MODFLOW-2005, generated by Flopy.
2 0
1 0 # stress period ... | CC0-1.0 | examples/Notebooks/flopy3_modflow_boundaries.ipynb | briochh/flopy |
!pip install --quiet transformers sentence-transformers nltk pyter3
import json
from pathlib import Path
def read_squad(path):
path = Path(path)
with open(path, 'rb') as f:
squad_dict = json.load(f)
contexts = []
questions = []
answers = []
for group in squad_dict['data']:
for ... | electra-base-squad2.txt
| MIT | src/test/resources/Baseline_QA/Baseline_QA_ELECTRA.ipynb | jenka2014/aigents-java-nlp | |
Machine Translation Inference Pipeline Packages | import os
import shutil
from typing import Dict
from transformers import T5Tokenizer, T5ForConditionalGeneration
from forte import Pipeline
from forte.data import DataPack
from forte.common import Resources, Config
from forte.processors.base import PackProcessor
from forte.data.readers import PlainTextReader | _____no_output_____ | Apache-2.0 | docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb | Xuezhi-Liang/forte |
BackgroundAfter a Data Scientist is satisfied with the results of a training model, they will have their notebook over to an MLE who has to convert their model into an inference model. Inference Workflow PipelineWe consider `t5-small` as a trained MT model to simplify the example. We should always consider pipeline f... | pipeline: Pipeline = Pipeline[DataPack]() | _____no_output_____ | Apache-2.0 | docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb | Xuezhi-Liang/forte |
ReaderAfter observing the dataset, it's a plain `txt` file. Therefore, we can use `PlainTextReader` directly. | pipeline.set_reader(PlainTextReader()) | _____no_output_____ | Apache-2.0 | docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb | Xuezhi-Liang/forte |
However, it's still beneficial to take a deeper look at how to design this class so that users can customize a reader when needed. ProcessorWe already have an inference model, `t5-small`, and we need a component to make an inference. Therefore, besides the model itself, there are several behaviors needed.1. tokenizati... |
class MachineTranslationProcessor(PackProcessor):
"""
Translate the input text and output to a file.
"""
def initialize(self, resources: Resources, configs: Config):
super().initialize(resources, configs)
# Initialize the tokenizer and model
model_name: str = self.configs.pretr... | _____no_output_____ | Apache-2.0 | docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb | Xuezhi-Liang/forte |
ExamplesWe have a working [MT translation pipeline example](https://github.com/asyml/forte/blob/master/docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb).There are several basic functions of the processor and internal functions defined in this example.* ``initialize()``: Pipeline will call it at the start of pro... | dir_path = os.path.abspath(
os.path.join("data_samples", "machine_translation")
) # notebook should be running from project root folder
pipeline.run(dir_path)
print("Done successfully") | _____no_output_____ | Apache-2.0 | docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb | Xuezhi-Liang/forte |
One can investigate the machine translation output in folder `mt_test_output` located under the script's directory.Then we remove the output folder below. | shutil.rmtree(MachineTranslationProcessor.default_configs()["output_folder"]) | _____no_output_____ | Apache-2.0 | docs/notebook_tutorial/wrap_MT_inference_pipeline.ipynb | Xuezhi-Liang/forte |
T81-558: Applications of Deep Neural Networks* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), School of Engineering and Applied Science, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)* For more information visit the [class website](https://sites.wustl.e... | from sklearn import preprocessing
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import shutil
import os
import requests
import base64
# Encode text values to dummy variables(i.e. [1,0,0],[0,1,0],[0,0,1] for red,green,blue)
def encode_text_dummy(df, name):
dummies = pd.get_dummies(df[name]... | _____no_output_____ | Apache-2.0 | assignments/assignment_yourname_class3.ipynb | Chuyi1202/T81-558-Application-of-Deep-Neural-Networks |
Assignment 3 Sample CodeThe following code provides a starting point for this assignment. | import os
import pandas as pd
from scipy.stats import zscore
# This is your student key that I emailed to you at the beginnning of the semester.
key = "qgABjW9GKV1vvFSQNxZW9akByENTpTAo2T9qOjmh" # This is an example key and will not work.
# You must also identify your source file. (modify for your local setup)
# fil... | _____no_output_____ | Apache-2.0 | assignments/assignment_yourname_class3.ipynb | Chuyi1202/T81-558-Application-of-Deep-Neural-Networks |
Checking Your SubmissionYou can always double check to make sure your submission actually happened. The following utility code will help with that. | import requests
import pandas as pd
import base64
import os
def list_submits(key):
r = requests.post("https://api.heatonresearch.com/assignment-submit",
headers={'x-api-key': key},
json={})
if r.status_code == 200:
print("Success: \n{}".format(r.text))
el... | _____no_output_____ | Apache-2.0 | assignments/assignment_yourname_class3.ipynb | Chuyi1202/T81-558-Application-of-Deep-Neural-Networks |
Variational Autoencoder From book - "Hands-On Machine Learning with Scikit-Learn and TensorFlow" $\bullet$ Perform PCA with an undercomplete linear autoencoder(Undercomplete Autoencode: The internal representation has a lower dimensionality than the input data) | import tensorflow as tf
from tensorflow.contrib.layers import fully_connected
n_inputs = 3 # 3 D input dimension
n_hidden = 2 # 2 D internal representation
n_outputs = n_inputs
learning_rate = 0.01
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
hidden = fully_connected(X, n_hidden, activation_fn = None)
ou... | _____no_output_____ | MIT | VAE/VAE.ipynb | DarrenZhang01/Machine-Learning |
Reading and writing fieldsThere are two main file formats to which a `discretisedfield.Field` object can be saved:- [VTK](https://vtk.org/) for visualisation using e.g., [ParaView](https://www.paraview.org/) or [Mayavi](https://docs.enthought.com/mayavi/mayavi/)- OOMMF [Vector Field File Format (OVF)](https://math.nis... | import discretisedfield as df
r = 5e-9
cell = (0.5e-9, 0.5e-9, 0.5e-9)
mesh = df.Mesh(p1=(-r, -r, -r), p2=(r, r, r), cell=cell)
def norm_fun(pos):
x, y, z = pos
if x**2 + y**2 + z**2 <= r**2:
return 1e6
else:
return 0
def value_fun(pos):
x, y, z = pos
c = 1e9
return (-c*... | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Let us have a quick view of the field we created | # NBVAL_IGNORE_OUTPUT
field.plane('z').k3d.vector(color_field=field.z) | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Writing the field to a fileThe main method used for saving field in different files is `discretisedfield.Field.write()`. It takes `filename` as an argument, which is a string with one of the following extensions:- `'.vtk'` for saving in the VTK format- `'.ovf'`, `'.omf'`, `'.ohf'` for saving in the OVF formatLet us fi... | vtkfilename = 'my_vtk_file.vtk'
field.write(vtkfilename) | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
We can check if the file was saved in the current directory. | import os
os.path.isfile(f'./{vtkfilename}') | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Now, we can delete the file: | os.remove(f'./{vtkfilename}') | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Next, we can save the field in the OVF format and check whether it was created in the current directory. | omffilename = 'my_omf_file.omf'
field.write(omffilename)
os.path.isfile(f'./{omffilename}') | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
There are three different possible representations of an OVF file: one ASCII (`txt`) and two binary (`bin4` or `bin8`). ASCII `txt` representation is a default representation when `discretisedfield.Field.write()` is called. If any different representation is required, it can be passed via `representation` argument. | field.write(omffilename, representation='bin8')
os.path.isfile(f'./{omffilename}') | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Reading the OVF fileThe method for reading OVF files is a class method `discretisedfield.Field.fromfile()`. By passing a `filename` argument, it reads the file and creates a `discretisedfield.Field` object. It is not required to pass the representation of the OVF file to the `discretisedfield.Field.fromfile()` method,... | read_field = df.Field.fromfile(omffilename) | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Like previouly, we can quickly visualise the field | # NBVAL_IGNORE_OUTPUT
read_field.plane('z').k3d.vector(color_field=read_field.z) | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Finally, we can delete the OVF file we created. | os.remove(f'./{omffilename}') | _____no_output_____ | BSD-3-Clause | docs/field-read-write.ipynb | ubermag/discretisedfield |
Now we get the theoretical earth orbital speed: | # Now let's compute the theoretical expectation. First, we load a pck file
# that contain miscellanoeus information, like the G*M values for different
# objects
# First, load the kernel
spiceypy.furnsh('../kernels/pck/gm_de431.tpc')
_, GM_SUN = spiceypy.bodvcd(bodyid=10, item='GM', maxn=1)
# Now compute the orbital s... | Theoretical orbital speed of the Earth around the Sun in km/s: 29.87838444261713
| MIT | Jorges Notes/Tutorial_1.ipynb | Chuly90/Astroniz-YT-Tutorials |
Lab 05 - "Convolutional Neural Networks (CNNs)" AssignmentsGSERM'21 course "Deep Learning: Fundamentals and Applications", University of St. Gallen In the last lab we learned how to enhance vanilla Artificial Neural Networks (ANNs) using `PyTorch` to classify even more complex images. Therefore, we used a special ty... | from IPython.display import YouTubeVideo
# NVIDIA: "Official Intro | GTC 2020 | I AM AI"
YouTubeVideo('e2_hsjpTi4w', width=1000, height=500) | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
As always, pls. don't hesitate to ask all your questions either during the lab, post them in our CANVAS (StudyNet) forum (https://learning.unisg.ch), or send us an email (using the course email). 1. Assignment Objectives: Similar today's lab session, after today's self-coding assignments you should be able to:> 1. Und... | # import standard python libraries
import os, urllib, io
from datetime import datetime
import numpy as np | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Import Python machine / deep learning libraries: | # import the PyTorch deep learning library
import torch, torchvision
import torch.nn.functional as F
from torch import nn, optim
from torch.autograd import Variable | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Import the sklearn classification metrics: | # import sklearn classification evaluation library
from sklearn import metrics
from sklearn.metrics import classification_report, confusion_matrix | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Import Python plotting libraries: | # import matplotlib, seaborn, and PIL data visualization libary
import matplotlib.pyplot as plt
import seaborn as sns
from PIL import Image | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Enable notebook matplotlib inline plotting: | %matplotlib inline | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Import Google's GDrive connector and mount your GDrive directories: | # import the Google Colab GDrive connector
from google.colab import drive
# mount GDrive inside the Colab notebook
drive.mount('/content/drive') | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Create a structure of Colab Notebook sub-directories inside of GDrive to store (1) the data as well as (2) the trained neural network models: | # create Colab Notebooks directory
notebook_directory = '/content/drive/MyDrive/Colab Notebooks'
if not os.path.exists(notebook_directory): os.makedirs(notebook_directory)
# create data sub-directory inside the Colab Notebooks directory
data_directory = '/content/drive/MyDrive/Colab Notebooks/data'
if not os.path.exi... | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Set a random `seed` value to obtain reproducable results: | # init deterministic seed
seed_value = 1234
np.random.seed(seed_value) # set numpy seed
torch.manual_seed(seed_value) # set pytorch seed CPU | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Google Colab provides the use of free GPUs for running notebooks. However, if you just execute this notebook as is, it will use your device's CPU. To run the lab on a GPU, got to `Runtime` > `Change runtime type` and set the Runtime type to `GPU` in the drop-down. Running this lab on a CPU is fine, but you will find th... | # set cpu or gpu enabled device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu').type
# init deterministic GPU seed
torch.cuda.manual_seed(seed_value)
# log type of device enabled
print('[LOG] notebook with {} computation enabled'.format(str(device))) | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Let's determine if we have access to a GPU provided by e.g. Google's COLab environment: | !nvidia-smi | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
3. Convolutional Neural Networks (CNNs) Assignments 3.1 CIFAR-10 Dataset Download and Data Assessment The **CIFAR-10 database** (**C**anadian **I**nstitute **F**or **A**dvanced **R**esearch) is a collection of images that are commonly used to train machine learning and computer vision algorithms. The database is wide... | cifar10_classes = ['plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'] | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Thereby the dataset contains 6,000 images for each of the ten classes. The CIFAR-10 is a straightforward dataset that can be used to teach a computer how to recognize objects in images.Let's download, transform and inspect the training images of the dataset. Therefore, we first will define the directory we aim to store... | train_path = data_directory + '/train_cifar10' | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Now, let's download the training data accordingly: | # define pytorch transformation into tensor format
transf = torchvision.transforms.Compose([torchvision.transforms.ToTensor(), torchvision.transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# download and transform training images
cifar10_train_data = torchvision.datasets.CIFAR10(root=train_path, train=True, tra... | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Verify the volume of training images downloaded: | # get the length of the training data
len(cifar10_train_data) | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Let's now decide on where we want to store the evaluation data: | eval_path = data_directory + '/eval_cifar10' | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
And download the evaluation data accordingly: | # define pytorch transformation into tensor format
transf = torchvision.transforms.Compose([torchvision.transforms.ToTensor(), torchvision.transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# download and transform validation images
cifar10_eval_data = torchvision.datasets.CIFAR10(root=eval_path, train=False, tr... | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Let's also verfify the volume of validation images downloaded: | # get the length of the training data
len(cifar10_eval_data) | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
3.2 Convolutional Neural Network (CNN) Model Training and Evaluation We recommend you to try the following exercises as part of the self-coding session:**Exercise 1: Train the neural network architecture of the lab with increased learning rate.** > Increase the learning rate of the network training to a value of **0.... | #### Step 1. define and init neural network architecture #############################################################
# ***************************************************
# INSERT YOUR SOLUTION/CODE HERE
# ***************************************************
#### Step 2. define loss, training hyperparameters and dat... | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
**2. Evaluation of "shallow" vs. "deep" neural network architectures.** > In addition to the architecture of the lab notebook, evaluate further (more **shallow** as well as more **deep**) neural network architectures by either **removing or adding convolutional layers** to the network. Train a model (using the architec... | #### Step 1. define and init neural network architecture #############################################################
# ***************************************************
# INSERT YOUR SOLUTION/CODE HERE
# ***************************************************
#### Step 2. define loss, training hyperparameters and dat... | _____no_output_____ | BSD-3-Clause | lab_05/lab_05_exercises.ipynb | HSG-AIML/LabGSERM |
Data Science Academy - Python Fundamentos - Capítulo 9 Download: http://github.com/dsacademybr Mini-Projeto 2 - Análise Exploratória em Conjunto de Dados do Kaggle Análise 3 | # Imports
import os
import subprocess
import stat
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import datetime
sns.set(style="white")
%matplotlib inline
# Dataset
clean_data_path = "dataset/autos.csv"
df = pd.read_csv(clean_data_path,encoding="latin-1") | _____no_output_____ | Apache-2.0 | Cap10/Mini-Projeto-Solucao/Mini-Projeto2 - Analise3.ipynb | CezarPoeta/Python-Fundamentos |
Preço médio do veículo por tipo de combustível e tipo de caixa de câmbio | # Crie um Barplot com o Preço médio do veículo por tipo de combustível e tipo de caixa de câmbio
fig, ax = plt.subplots(figsize=(8,5))
colors = ["#00e600", "#ff8c1a","#a180cc"]
sns.barplot(x="fuelType", y="price",hue="gearbox", palette="husl",data=df)
ax.set_title("Preço médio do veículo por tipo de combustível e tipo ... | _____no_output_____ | Apache-2.0 | Cap10/Mini-Projeto-Solucao/Mini-Projeto2 - Analise3.ipynb | CezarPoeta/Python-Fundamentos |
Potência média de um veículo por tipo de veículo e tipo de caixa de câmbio | # Crie um Barplot com a Potência média de um veículo por tipo de veículo e tipo de caixa de câmbio
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple"]
fig, ax = plt.subplots(figsize=(8,5))
sns.set_palette(sns.xkcd_palette(colors))
sns.barplot(x="vehicleType", y="powerPS",hue="gearbox",data=df)
... | _____no_output_____ | Apache-2.0 | Cap10/Mini-Projeto-Solucao/Mini-Projeto2 - Analise3.ipynb | CezarPoeta/Python-Fundamentos |
Calibrate mean and integrated intensity of a fluorescence marker versus concentration Requirements- Images with different concentrations of the fluorescent tag with the concentration clearly specified in the image namePrepare pure solutions of various concentrations of fluorescent tag in imaging media and collect imag... | #################################
# Don't modify the code below #
#################################
import intake_io
import os
import re
import numpy as np
import pylab as plt
import seaborn as sns
from skimage import io
import pandas as pd
from tqdm import tqdm
from skimage.measure import regionprops_table
from am... | _____no_output_____ | Apache-2.0 | notebooks/misc/calibrate_intensities.ipynb | stjude/punctatools |
Data & parameters`input_dir`: folder with images to be analyzed`output_dir`: folder to save results`channel_name`: name of the fluorecent tag (e.g. "GFP") Specify data paths and parameters | input_dir = "../../example_data/calibration"
output_dir = "../../test_output/calibration"
channel_name = 'GFP' | _____no_output_____ | Apache-2.0 | notebooks/misc/calibrate_intensities.ipynb | stjude/punctatools |
The following code lists all images in the input directory: | #################################
# Don't modify the code below #
#################################
samples = walk_dir(input_dir)
print(f'{len(samples)} images were found:')
print(np.array(samples)) | 4 images were found:
['../../example_data/calibration/05192021_GFPcalibration_1nM_-_Position_4_XY1621491830.tif'
'../../example_data/calibration/05192021_GFPcalibration_5.62uM_-_Position_5_XY1621485379.tif'
'../../example_data/calibration/05192021_GFPcalibration_31.6nM_-_Position_2_XY1621488646.tif'
'../../example_d... | Apache-2.0 | notebooks/misc/calibrate_intensities.ipynb | stjude/punctatools |
The following code loads a random image: | #################################
# Don't modify the code below #
#################################
sample = samples[np.random.randint(len(samples))]
dataset = intake_io.imload(sample)
if 'z' in dataset.dims:
dataset = dataset.max('z')
plt.figure(figsize=(7, 7))
io.imshow(dataset['image'].data) | /research/sharedresources/cbi/public/conda_envs/punctatools/lib/python3.9/site-packages/scikit_image-0.19.0-py3.9-linux-x86_64.egg/skimage/io/_plugins/matplotlib_plugin.py:150: UserWarning: Low image data range; displaying image with stretched contrast.
lo, hi, cmap = _get_display_range(image)
| Apache-2.0 | notebooks/misc/calibrate_intensities.ipynb | stjude/punctatools |
The following code quantifies all input images: | %%time
#################################
# Don't modify the code below #
#################################
def quantify(sample, input_dir, output_dir, channel_name):
dataset = intake_io.imload(sample)
img = np.array(dataset['image'].data)
df = pd.DataFrame(regionprops_table(label_image=np.ones_like(img... | _____no_output_____ | Apache-2.0 | notebooks/misc/calibrate_intensities.ipynb | stjude/punctatools |
The following code plots intensity versus concentration for sanity check | #################################
# Don't modify the code below #
#################################
for col in [rf'{channel_name} concentration nM', rf'{channel_name} mean intensity per image', rf'{channel_name} integrated intensity per image']:
df['Log ' + col] = np.log10(df[col])
for col in [rf'{channel_n... | _____no_output_____ | Apache-2.0 | notebooks/misc/calibrate_intensities.ipynb | stjude/punctatools |
Uncomment the following line to install [geemap](https://geemap.org) if needed. | # !pip install geemap
import ee
import geemap
geemap.show_youtube('N7rK2aV1R4c')
Map = geemap.Map()
Map
# Add Earth Engine dataset
image = ee.Image('USGS/SRTMGL1_003')
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5'],
}
# A... | _____no_output_____ | MIT | examples/notebooks/05_drawing_tools.ipynb | ppoon23/geemap |
Circuit Quantum Electrodynamics Contents1. [Introduction](intro)2. [The Schrieffer-Wolff Transformation](tswt)3. [Block-diagonalization of the Jaynes-Cummings Hamiltonian](bdotjch)4. [Full Transmon](full-transmon)5. [Qubit Drive with cQED](qdwcqed)6. [The Cross Resonance Entangling Gate](tcreg) 1. Introduction By an... | # import SymPy and define symbols
import sympy as sp
sp.init_printing(use_unicode=True)
wr = sp.Symbol('\omega_r') # resonator frequency
wq = sp.Symbol('\omega_q') # qubit frequency
g = sp.Symbol('g', real=True) # vacuum Rabi coupling
Delta = sp.Symbol('Delta', real=True) # wr - wq; defined later
# import operator rela... | _____no_output_____ | Apache-2.0 | content/ch-quantum-hardware/cQED-JC-SW.ipynb | muneerqu/qiskit-textbook |
As a note about `sympy`, we will need to used the methods `doit()`, `expand`, `normal_ordered_form`, and `qsimplify_pauli` to proceed with actually taking the commutator, expanding it into terms, normal ordering the bosonic modes (creation before annihilation), and simplify the Pauli algebra. Trying this with $\eta$ yi... | pauli.qsimplify_pauli(normal_ordered_form(eta.doit().expand())) | _____no_output_____ | Apache-2.0 | content/ch-quantum-hardware/cQED-JC-SW.ipynb | muneerqu/qiskit-textbook |
Now take $A$ and $B$ as the coefficients of $a^\dagger \sigma_-$ and $a\sigma_+$, respectively. Then the commutator | A = sp.Symbol('A')
B = sp.Symbol('B')
eta = A * Dagger(a) * sminus - B * a * splus;
pauli.qsimplify_pauli(normal_ordered_form(Commutator(H0, eta).doit().expand())) | _____no_output_____ | Apache-2.0 | content/ch-quantum-hardware/cQED-JC-SW.ipynb | muneerqu/qiskit-textbook |
This expression should be equal to $H_2$ | H2 | _____no_output_____ | Apache-2.0 | content/ch-quantum-hardware/cQED-JC-SW.ipynb | muneerqu/qiskit-textbook |
which implies $A = B = g/\Delta$ where $\Delta = \omega_r - \omega_q$ is the frequency detuning between the resonator and qubit. Therefore our $S^{(1)}$ is determined to be | S1 = eta.subs(A, g/Delta)
S1 = S1.subs(B, g/Delta); S1.factor() | _____no_output_____ | Apache-2.0 | content/ch-quantum-hardware/cQED-JC-SW.ipynb | muneerqu/qiskit-textbook |
Then we can calculate the effective second order correction to $H_0$ | Heff = H0 + 0.5*pauli.qsimplify_pauli(normal_ordered_form(Commutator(H2, S1).doit().expand())).simplify(); Heff | _____no_output_____ | Apache-2.0 | content/ch-quantum-hardware/cQED-JC-SW.ipynb | muneerqu/qiskit-textbook |
Tutorial sobre Scala Declaraciones Declaración de variables Existen dos categorias de variables: inmutables y mutables. Las variables mutables son aquellas en las que es posible modificar el contenido de la variable. Las variables inmutables son aquellas en las que no es posible alterar el contenido de las variables... | //Variable inmutable
val a:Int=1
//variable mutable
var b:Int=2 | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Tipos de datos 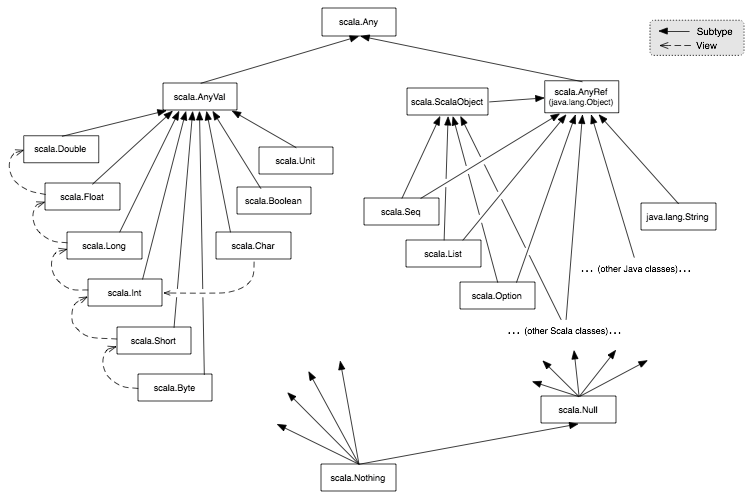 Siempre que se infiere un tipo en Scala, el tipo escogido será siempre el mas bajo posible en la jerarquía.Algunos tipos especiales:- **Any**: Es la clase de la que heredan todas las clases en Sc... | def funcion1(a:Int,b:Int):Int={
return a+b
}
def funcion2(a:Int,b:Int)={
a+b
}
def funcion3(a:Int,b:Int)=a+b | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Al igual que con la declaración de variables no es obligatorio declarar el tipo devuelto por la función. Si no se declara una sentencia `return`, el valor de la ultima instrucción es el devuelto por la función. Interpolación de cadenasLa interpolación de cadenas consiste insertar el valor de una variable dentro de una... | val valor=1
val expresion=2
println(s"El valor de la variable ${valor} y la expresion vale ${expresion+1}") | El valor de la variable 1 y la expresion vale 3
| MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Estructuras de selección If/Else | //Funciona igual que en Java
val verdad:Boolean=true;
if (verdad){
println("Hola")
}else{
println("Adios")
}
| _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
En Scala no existe la estructura `switch`, en su lugar existe lo conocido como *pattern matching* Match | val numero:Int=3
val nombre=numero match{ //Puede ir dentro de la llamada a una funcion
case 1=> "Uno"
case 2=> "Dos"
case 3=> "Tres"
case _=> "Ninguno" //Es obligatorio incluir una clausula con _ que se ejecuta cuando no hay coincidencia
}
println(nombre) | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Estructuras de repetición Bucle *While* | //Igual que en Java
var x=0
while(x<5){
print(x)
x+=1
} | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Bucle *Do While* | //Igual que en Java
var x=0
do{
print(x)
x+=1
}while(x<5) | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Bucle *For* | println("For to")
for(i<- 1 to 5){ //Hasta el limite inclusive
print(i)
}
println("\nFor until")
for(i<- 1 until 5){ //Hasta el limite exclusive
print(i)
}
println("\nFor para colecciones")
for(i <- List(1,2,3,4)){ //For para recorrer colecciones
print(i)
} | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
*foreach* | val lista=List(1,2,3,4)
lista.foreach(x=> print(x)) //La funcion no devuelve nada y no modifica el conjunto | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Clases Indicaciones previasSe deben declarar entre parentesis todos los atributos que vaya a usar la clase. Se pueden declarar otros constructores mediante la definición de this, pero siempre se debe llamar al constructor por defecto que es el que contiene todos los atributos.Los parametros de un constructor constitu... | //Declaracion de clases
class Saludo(mensaje: String) { //Estos son los atributos y son accesibles desde cualquier metodo de la clase
def diHola(nombre:String):Unit ={
println(mensaje+" "+nombre);
}
}
val saludo = new Saludo("Hola")
saludo.diHola("Pepe") | _____no_output_____ | MIT | Scala-basics.ipynb | FranciscoJavierMartin/Notebooks |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.