text stringlengths 957 885k |
|---|
<filename>src/tiden/apps/ignite/components/ignitestaticinitmixin.py
#!/usr/bin/env python3
#
# Copyright 2017-2020 GridGain Systems.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from ...nodestatus import NodeStatus
from .ignitemixin import IgniteMixin
from ....tidenexception import TidenException
from ....util import encode_enums, write_yaml_file, log_print, read_yaml_file, decode_enums
class IgniteStaticInitMixin(IgniteMixin):
"""
Encapsulates 'static' Ignite initialization from previously dumped configuration file.
"""
nodes_config_path = None
nodes_config_name = 'nodes-config.yaml'
nodes_config_store_host = None
is_static_inited = False
def __init__(self, *args, **kwargs):
# print('IgniteStaticInitMixin.__init__')
super(IgniteStaticInitMixin, self).__init__(*args, **kwargs)
# used by static dump_nodes_config/restore_nodes_config
self.nodes_config_path = None
self.nodes_config_name = 'nodes-config.yaml'
self.nodes_config_store_host = None
self.is_static_inited = False
self._parse_static_init_params(kwargs)
def on_setup(self):
# TODO:
pass
def _parse_static_init_params(self, kwargs):
"""
Parse kwargs with static nodes config params for dump or restore
"""
self.nodes_config_path = kwargs.get('nodes_config_path',
self.config.get('environment', {}).get("nodes_config_path", self.nodes_config_path))
self.nodes_config_name = kwargs.get('nodes_config_name',
self.config.get('environment', {}).get("nodes_config_name",
self.nodes_config_name))
self.nodes_config_store_host = kwargs.get('nodes_config_store_host',
self.config.get('environment', {}).get("nodes_config_store_host",
self.nodes_config_store_host))
self.is_static_inited = kwargs.get('static_init', False)
def dump_nodes_config(self, strict=True, **kwargs):
"""
Write nodes config in yaml file and upload on selected node for storing
"""
if kwargs:
self._parse_static_init_params(kwargs)
if self.nodes_config_path is None:
if strict:
raise TidenException("Can't backup nodes config without nodes_config_path")
else:
return
nodes_config = encode_enums(self.nodes)
config_local_path = "{}/{}".format(self.config["tmp_dir"], self.nodes_config_name)
write_yaml_file(config_local_path, nodes_config)
if self.nodes_config_store_host is None:
self.nodes_config_store_host = self.nodes[min(self.nodes.keys())]['host']
remote_path = '{}/{}'.format(self.nodes_config_path, self.nodes_config_name)
log_print("Dump nodes config on host '{}' to '{}'".format(remote_path, self.nodes_config_store_host))
self.ssh.upload_on_host(self.nodes_config_store_host, [config_local_path], self.nodes_config_path)
cmd = "chmod 777 {}".format(remote_path)
self.ssh.exec_on_host(self.nodes_config_store_host, [cmd])
def restore_nodes_config(self, **kwargs):
"""
Download nodes config yaml file from storing place and parse into self.nodes
"""
if kwargs:
self._parse_static_init_params(kwargs)
if self.nodes_config_path is None:
raise TidenException("Can't restore nodes config without nodes_config_path")
config_local_path = "{}/{}".format(self.config["tmp_dir"], self.nodes_config_name)
config_remote_path = "{}/{}".format(self.nodes_config_path, self.nodes_config_name)
log_print(
"Restore nodes config from host '{}' path '{}'".format(config_remote_path, self.nodes_config_store_host))
self.ssh.download_from_host(self.nodes_config_store_host, config_remote_path, config_local_path)
configs = read_yaml_file(config_local_path)
self.nodes = decode_enums(configs, available_enums=[NodeStatus])
|
<gh_stars>0
# -*- coding: utf-8 -*-
import sys
import time
import os
import json
import argparse
from torch.autograd import Variable
from utils.logger import setup_logger
sys.path.insert(0, '../')
sys.dont_write_bytecode = True
import dataset
import torch
import torch.nn.functional as F
import torch.nn as nn
from torch.utils.data import DataLoader
from tensorboardX import SummaryWriter
from model import CCLDNet
from utils.lr_scheduler import get_scheduler
def clip_gradient(optimizer, grad_clip):
"""
Clips gradients computed during backpropagation to avoid explosion of gradients.
:param optimizer: optimizer with the gradients to be clipped
:param grad_clip: clip value
"""
for group in optimizer.param_groups:
for param in group["params"]:
if param.grad is not None:
param.grad.data.clamp_(-grad_clip, grad_clip)
def parse_option():
parser = argparse.ArgumentParser()
# data
parser.add_argument('--batchsize', type=int, default=20, help='training batch size')
parser.add_argument('--epoch', type=int, default=100, help='epoch number')
parser.add_argument('--crossnum', type=str, default='endo', help='CA1, CA2, CA3, CA4, CA5, endo, CVC')
# training
parser.add_argument('--hflip', type=bool,default=True, help='hflip data')
parser.add_argument('--vflip', type=bool,default=True, help='vflip data')
parser.add_argument('--checkpoint', type=int, default=1, help='use_checkpoint')
parser.add_argument('--ratio', type=float, default=1, help='hidden_number/input_number in the linear_layer')
parser.add_argument('--K', type=int, default=4, help='the number of weights')
parser.add_argument('--T', type=int, default=31, help='hyperparameter for sparsity')
parser.add_argument('--swin_type', type=str, default='base', help='base,large')
parser.add_argument('--dropout_rate', type=float, default=0.5, help='dropout_rate in swin_backbone')
parser.add_argument('--decoder', type=str, default='CCLD', help='CCLD')
parser.add_argument('--lr', type=float, default=0.8, help='learning rate')
parser.add_argument('--lr_rate', type=float, default=0.1, help='lr_backbone/lr_head')
parser.add_argument('--lr_scheduler', type=str, default='cosine',
choices=["step", "cosine"], help="learning rate scheduler")
parser.add_argument('--warmup_epoch', type=int, default=30, help='warmup epoch')
parser.add_argument('--warmup_multiplier', type=int, default=100, help='warmup multiplier')
parser.add_argument('--optim', type=str, default='SGD', help="SGD,AdamW")
parser.add_argument('--head_weight_decay', type=float, default=5e-4, help='weight decay')
parser.add_argument('--backbone_weight_decay', type=float, default=5e-4, help='weight decay')
parser.add_argument('--momentum', type=float, default=0.9, help='momentum for SGD')
parser.add_argument('--model_path', type=str, default=None, help='model path to pretrain')
parser.add_argument('--output_dir', type=str, default='./model_save', help='output director')
parser.add_argument('--data_dir', type=str, default='./data/TrainDataset_Endo', help='./data/TrainDataset_Endo, ./data/TrainDataset_CVC_Kva, ./data/TrainDataset_2018')
opt, unparsed = parser.parse_known_args()
opt.output_dir = os.path.join(opt.output_dir, str(int(time.time())) + '_' + opt.decoder + '_' + opt.swin_type + '_' + opt.crossnum)
return opt
def get_DC(SR_real, GT):
# DC : Dice Coefficient
GT = GT > 0.5
dice_sum = 0.0
for i in range(0,1):
SR = SR_real >= 0.5
inter = ((SR == GT) & (GT == 1)).sum(dim=(1,2,3))
union = SR.sum(dim=(1,2,3))+GT.sum(dim=(1,2,3))
dice_sum += 2.0*inter/union
return dice_sum
def iou_loss(pred, mask):
pred = torch.sigmoid(pred)
inter = (pred*mask).sum(dim=(2,3))
union = (pred+mask).sum(dim=(2,3))
iou = 1-(inter+1)/((union-inter+1)+1e-6)
return iou.mean()
def build_loader(opt):
num_gpus = torch.cuda.device_count()
print("========>num_gpus:{}==========".format(num_gpus))
train_data = dataset.Endo_ISIC_Dataset(opt.data_dir, mode='train'+opt.crossnum, trainsize=384, hflip=opt.hflip, vflip=opt.vflip)
train_loader = DataLoader(train_data,collate_fn=train_data.collate, batch_size=opt.batchsize*num_gpus, shuffle=True, pin_memory=False, num_workers=8)
val_data = dataset.Endo_ISIC_Dataset(opt.data_dir, trainsize=384,mode='val'+opt.crossnum)
val_loader = DataLoader(val_data, batch_size=1, shuffle=False, num_workers=8)
return train_loader, val_loader
def build_model(opt):
# build model
Network = CCLDNet(opt)
if torch.cuda.device_count() > 1:
print("Let's use", torch.cuda.device_count(), "GPUs!")
Network = nn.DataParallel(Network)
Network = Network.cuda()
return Network
def main(opt):
train_loader, val_loader = build_loader(opt)
Network = build_model(opt)
base, head = [], []
# build optimizer
for name, param in Network.named_parameters():
if 'bkbone' in name:
base.append(param)
else:
head.append(param)
if opt.optim == 'SGD':
optimizer = torch.optim.SGD([{'params':base, 'weight_decay': opt.backbone_weight_decay, 'lr': opt.lr * opt.lr_rate * opt.batchsize/128}, {'params':head}], lr=opt.lr*opt.batchsize/128, momentum=opt.momentum, weight_decay=opt.head_weight_decay, nesterov=True)
elif opt.optim == 'AdamW':
optimizer = torch.optim.AdamW([{'params':base, 'weight_decay': opt.head_weight_decay, 'lr': opt.lr * opt.lr_rate * opt.batchsize/128}, {'params':head}], lr=opt.lr*opt.batchsize/128, weight_decay=opt.head_weight_decay)
scheduler = get_scheduler(optimizer, len(train_loader), opt)
if opt.decoder == 'CCLD':
for epoch in range(opt.epoch):
train(train_loader, Network, opt, epoch, optimizer, scheduler)
validate(val_loader, Network, opt, epoch)
return os.path.join(opt.output_dir, "last_epoch.pth")
def train(data_loader, Network, opt, epoch, optimizer, scheduler):
net = Network
net.train(True)
global_step = 0
num = len(data_loader)
loss_finalmask_all = 0.0
loss_center_all = 0.0
loss_boundary_all = 0.0
loss_premask_all = 0.0
# routine
for step, (image, mask, center, boundary) in enumerate(data_loader):
optimizer.zero_grad()
image, mask, center, boundary = Variable(image).cuda(), Variable(mask).cuda(), Variable(center).cuda(), Variable(boundary).cuda()
out_premask, out_center, out_boundary, out_finalmask = net(image)
loss_premask = F.binary_cross_entropy_with_logits(out_premask, mask) + iou_loss(out_premask, mask)
loss_center = F.binary_cross_entropy_with_logits(out_center, center)
loss_boundary = F.binary_cross_entropy_with_logits(out_boundary, boundary)
loss_finalmask = F.binary_cross_entropy_with_logits(out_finalmask, mask) + iou_loss(out_finalmask, mask)
loss = (loss_premask + loss_center + loss_boundary + loss_finalmask)/4
loss.backward()
clip_gradient(optimizer, 0.5)
optimizer.step()
scheduler.step()
loss_premask_all += loss_premask.item()
loss_center_all += loss_center.item()
loss_boundary_all += loss_boundary.item()
loss_finalmask_all += loss_finalmask.item()
global_step += 1
sw.add_scalars('lr' , {'lr_backbone':optimizer.param_groups[0]['lr'], 'lr_head':optimizer.param_groups[0]['lr']}, global_step=epoch+1)
sw.add_scalars('trainloss', {'premask':loss_premask_all/num, 'center':loss_center_all/num, 'boundary':loss_boundary_all/num, 'mask':loss_finalmask_all/num}, global_step=epoch+1)
logger.info('step:%d/%d/%d | lr_backbone=%.6f | lr_head=%.6f | loss_premask_ave=%.6f | loss_center_ave=%.6f | loss_boundary_ave=%.6f | loss_finalmask_ave=%.6f'
%(global_step, epoch+1, opt.epoch, optimizer.param_groups[0]['lr'], optimizer.param_groups[0]['lr'], loss_premask_all/num, loss_center_all/num, loss_boundary_all/num, loss_finalmask_all/num))
if (epoch+1) % 10 == 0:
torch.save(net.state_dict(), os.path.join(opt.output_dir, "epoch_{}_ckpt.pth".format(epoch + 1)))
logger.info("model saved {}!, learning_rate {}".format(os.path.join(opt.output_dir, "epoch_{}_ckpt.pth".format(epoch + 1)), optimizer.param_groups[0]['lr']))
if (epoch+1) == opt.epoch:
torch.save(net.state_dict(), os.path.join(opt.output_dir, "last_epoch.pth"))
logger.info("last_model saved !")
def validate(data_loader, Network, opt, epoch):
net = Network
net.eval()
num = len(data_loader)
sum_loss = 0.0
sum_loss_pre = 0.0
dice_score = 0.0
dice_score_pre = 0.0
with torch.no_grad():
for image, mask, (H, W), name in data_loader:
image, shape = image.cuda().float(), (H, W)
mask = mask.cuda().float()
out_premask, out_center, out_boundary, out_finalmask = net(image, shape)
loss = F.binary_cross_entropy_with_logits(out_finalmask, mask) + iou_loss(out_finalmask, mask)
sum_loss += loss.item()
loss_pre = F.binary_cross_entropy_with_logits(out_premask, mask) + iou_loss(out_premask, mask)
sum_loss_pre += loss_pre.item()
out_finalmask_score = torch.nn.Sigmoid()(out_finalmask)
dice_score += get_DC(out_finalmask_score, mask)
out_premask_score = torch.nn.Sigmoid()(out_premask)
dice_score_pre += get_DC(out_premask_score, mask)
sw.add_scalars('testloss', {'test_final':sum_loss/num, 'test_pre':sum_loss_pre/num}, global_step=epoch+1)
logger.info('epoch:%d | testloss=%.6f | testloss_pre=%.6f | dice=%.4f | dice_pre=%.4f'
%(epoch+1, sum_loss/num, sum_loss_pre/num, dice_score/num, dice_score_pre/num))
if __name__=='__main__':
best_dice = 0.0
opt = parse_option()
print(opt)
os.makedirs(opt.output_dir, exist_ok=True)
sw = SummaryWriter(opt.output_dir)
logger = setup_logger(output=opt.output_dir, name="CCLDNet")
path = os.path.join(opt.output_dir, "config.json")
with open(path, 'w') as f:
json.dump(vars(opt), f, indent=2)
logger.info("Full config saved to {}".format(path))
ckpt_path = main(opt)
|
<reponame>YosefLab/SingleCellLineageTracing<gh_stars>10-100
import unittest
import networkx as nx
import numpy as np
from cassiopeia.data.CassiopeiaTree import CassiopeiaTree
from cassiopeia.simulator.LeafSubsampler import LeafSubsamplerError
from cassiopeia.simulator.UniformLeafSubsampler import UniformLeafSubsampler
class UniformLeafSubsamplerTest(unittest.TestCase):
def test_bad_parameters(self):
with self.assertRaises(LeafSubsamplerError):
uniform_sampler = UniformLeafSubsampler(
ratio=0.5, number_of_leaves=400
)
with self.assertRaises(LeafSubsamplerError):
uniform_sampler = UniformLeafSubsampler()
def test_bad_number_of_samples(self):
balanced_tree = nx.balanced_tree(2, 3, create_using=nx.DiGraph)
tree = CassiopeiaTree(tree=balanced_tree)
with self.assertRaises(LeafSubsamplerError):
uniform_sampler = UniformLeafSubsampler(number_of_leaves=0)
uniform_sampler.subsample_leaves(tree)
with self.assertRaises(LeafSubsamplerError):
uniform_sampler = UniformLeafSubsampler(ratio=0.0001)
uniform_sampler.subsample_leaves(tree)
def test_subsample_balanced_tree(self):
balanced_tree = nx.balanced_tree(2, 3, create_using=nx.DiGraph)
balanced_tree = nx.relabel_nodes(
balanced_tree,
dict([(i, "node" + str(i)) for i in balanced_tree.nodes]),
)
balanced_tree.add_node("node15")
balanced_tree.add_edge("node15", "node0")
tree = CassiopeiaTree(tree=balanced_tree)
np.random.seed(10)
uni = UniformLeafSubsampler(number_of_leaves=3)
res = uni.subsample_leaves(tree=tree, keep_singular_root_edge=False)
expected_edges = [
("node15", "node8"),
("node15", "node5"),
("node5", "node11"),
("node5", "node12"),
]
self.assertEqual(set(res.edges), set(expected_edges))
np.random.seed(10)
uni = UniformLeafSubsampler(ratio=0.65)
res = uni.subsample_leaves(tree=tree, keep_singular_root_edge=False)
expected_edges = [
("node15", "node2"),
("node15", "node3"),
("node2", "node14"),
("node2", "node5"),
("node5", "node11"),
("node5", "node12"),
("node3", "node7"),
("node3", "node8"),
]
self.assertEqual(set(res.edges), set(expected_edges))
def test_subsample_custom_tree(self):
custom_tree = nx.DiGraph()
custom_tree.add_nodes_from(["node" + str(i) for i in range(17)])
custom_tree.add_edges_from(
[
("node16", "node0"),
("node0", "node1"),
("node0", "node2"),
("node1", "node3"),
("node1", "node4"),
("node2", "node5"),
("node2", "node6"),
("node4", "node7"),
("node4", "node8"),
("node6", "node9"),
("node6", "node10"),
("node7", "node11"),
("node11", "node12"),
("node11", "node13"),
("node9", "node14"),
("node9", "node15"),
]
)
tree = CassiopeiaTree(tree=custom_tree)
for u, v in tree.edges:
tree.set_branch_length(u, v, 1.5)
np.random.seed(10)
uni = UniformLeafSubsampler(ratio=0.5)
res = uni.subsample_leaves(tree=tree)
expected_edges = {
("node16", "node0"): 1.5,
("node0", "node1"): 1.5,
("node0", "node5"): 3.0,
("node1", "node3"): 1.5,
("node1", "node11"): 4.5,
("node11", "node12"): 1.5,
("node11", "node13"): 1.5,
}
self.assertEqual(set(res.edges), set(expected_edges.keys()))
for u, v in res.edges:
self.assertEqual(
res.get_branch_length(u, v), expected_edges[(u, v)]
)
expected_times = {
"node16": 0.0,
"node0": 1.5,
"node1": 3.0,
"node5": 4.5,
"node3": 4.5,
"node11": 7.5,
"node12": 9.0,
"node13": 9.0,
}
for u in res.nodes:
self.assertEqual(res.get_time(u), expected_times[u])
np.random.seed(11)
uni = UniformLeafSubsampler(number_of_leaves=6)
res = uni.subsample_leaves(tree=tree, keep_singular_root_edge=True)
expected_edges = [
("node16", "node0"),
("node0", "node1"),
("node0", "node2"),
("node1", "node3"),
("node1", "node11"),
("node11", "node12"),
("node11", "node13"),
("node2", "node5"),
("node2", "node6"),
("node6", "node10"),
("node6", "node15"),
]
self.assertEqual(set(res.edges), set(expected_edges))
if __name__ == "__main__":
unittest.main()
|
<gh_stars>10-100
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import collections
import math
from tensorflow.python.framework import constant_op
from tensorflow.python.framework import dtypes
from tensorflow.python.ops import gradient_checker
from tensorflow.python.ops import nn_ops
import tensorflow.python.ops.nn_grad # pylint: disable=unused-import
from tensorflow.python.platform import test
import tensorflow as tf
import random
import numpy as np
import time
import sparse_tools as sp
import os
import direct_sparse_grad_ops
from tensorflow.python import debug as tf_debug
import direct_sparse_layer_definition as ld
def placeholder_inputs(batch_size, num_classes, tensor_in_sizes_):
tensor_in_sizes = np.array(tensor_in_sizes_, dtype=np.int64)
batch_label_sizes = [batch_size, num_classes]
pointclouds_pl = tf.sparse_placeholder(tf.float32, shape=tensor_in_sizes, name="sparse_placeholder")
labels_pl = tf.placeholder(tf.float32, shape=batch_label_sizes, name="labels_placeholder")
return pointclouds_pl, labels_pl
def get_model(sparse_data, train_labels, is_training, tensor_in_sizes, num_classes = 10, scope = "mn16-", initializer = None, regularizer = None):
strides = [1,1,1,1,1]
padding = "SAME"
dim = 5
pooling_sizes = [1,2,2,2,1]
batch_size = tensor_in_sizes[0]
total_size = 1
for i in range(1, len(tensor_in_sizes)): #skip batch size
total_size = total_size * tensor_in_sizes[i]
sd_converted = ld.create_sparse_data_to_direct_sparse(sparse_data, dim)
ops = [None]*6
d1 = 0.06
net, tensor_in_sizes = ld.create_sparse_conv_layer(sd_converted, [3,3,3,1,8], tensor_in_sizes, strides, padding, dim, d1, "K-RELU", name = scope + "sc1", initializer=initializer)
net, tensor_in_sizes = ld.create_sparse_conv_layer(net, [3,3,3,8,8], tensor_in_sizes, strides, padding, dim, d1, "K-RELU", name = scope + "sc2", initializer=initializer)
net, tensor_in_sizes = ld.create_sparse_conv_layer(net, [3,3,3,8,8], tensor_in_sizes, strides, padding, dim, d1, "K-RELU", name = scope + "sc3", initializer=initializer)
net, tensor_in_sizes = ld.create_sparse_pooling_layer(net, pooling_sizes, tensor_in_sizes, dim, 6 * d1)
d2 = 0.12
net, tensor_in_sizes = ld.create_sparse_conv_layer(net, [3,3,3,8,16], tensor_in_sizes, strides, padding, dim, d2, "K-RELU", name = scope + "sc4", initializer=initializer)
net, tensor_in_sizes = ld.create_sparse_conv_layer(net, [3,3,3,16,16], tensor_in_sizes, strides, padding, dim, d2, "K-RELU", name = scope + "sc5", initializer=initializer)
net, tensor_in_sizes = ld.create_sparse_conv_layer(net, [3,3,3,16,16], tensor_in_sizes, strides, padding, dim, d2, "K-ABS", name = scope + "sc6", initializer=initializer)
sd = ld.create_direct_sparse_to_dense(net, dim)
sd_flat = tf.reshape(sd, [batch_size, total_size * 2])
conv_out = tf.layers.dropout(sd_flat, 0.5, name="dropout", training=is_training)
fc512 = tf.layers.dense(conv_out, 1024, name="dense2")
fc10 = tf.layers.dense(fc512, num_classes, name="dense1")
#if train:
sd_out = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=fc10, labels=train_labels, name = "softmax_loss"))
p_sd_out = tf.nn.softmax(logits=fc10)
return [sd_out, p_sd_out, ops]
|
#!/usr/bin/env python
# coding=utf-8
# Copyright 2021 The HuggingFace Team All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Fine-tuning the library models for seq2seq, text to image.
Script adapted from run_summarization_flax.py
"""
# You can also adapt this script on your own sequence to sequence task. Pointers for this are left as comments.
import os
import copy
# set a common huggingface cache folder (used with datasets and transformers) and wandb cache folder (used with artifacts) # required before importing transformers & datasets
os.environ['WANDB_CACHE_DIR'] = '/media/storage/wandb/' # required before importing wandb
os.environ['HF_HOME'] = '/media/storage/huggingface/'
import logging as pylogging # To avoid collision with transformers.utils.logging
import sys
import time
from dataclasses import dataclass, field
from functools import partial
from pathlib import Path
from typing import Callable, Optional
import json
import datasets
import nltk # Here to have a nice missing dependency error message early on
import numpy as np
from datasets import Dataset, load_dataset, load_metric
from tqdm import tqdm
import jax
import jax.numpy as jnp
import optax
import transformers
from filelock import FileLock
from flax import jax_utils, traverse_util
from flax.serialization import from_bytes, to_bytes
import flax.linen as nn
from flax.jax_utils import unreplicate
from flax.training import train_state
from flax.training.common_utils import get_metrics, onehot, shard, shard_prng_key
from transformers import (
CONFIG_MAPPING,
FLAX_MODEL_FOR_SEQ_TO_SEQ_CAUSAL_LM_MAPPING,
AutoConfig,
AutoTokenizer,
FlaxAutoModelForSeq2SeqLM,
FlaxBartForConditionalGeneration,
HfArgumentParser,
TrainingArguments,
)
from transformers.file_utils import is_offline_mode
from transformers.models.t5.modeling_flax_t5 import *
import wandb
logger = pylogging.getLogger(__name__)
try:
nltk.data.find("tokenizers/punkt")
except (LookupError, OSError):
if is_offline_mode():
raise LookupError(
"Offline mode: run this script without TRANSFORMERS_OFFLINE first to download nltk data files"
)
with FileLock(".lock") as lock:
nltk.download("punkt", quiet=True)
MODEL_CONFIG_CLASSES = list(FLAX_MODEL_FOR_SEQ_TO_SEQ_CAUSAL_LM_MAPPING.keys())
MODEL_TYPES = tuple(conf.model_type for conf in MODEL_CONFIG_CLASSES)
# Model hyperparameters, for convenience
# TODO: the model has now it's own definition file and should be imported
OUTPUT_VOCAB_SIZE = 16384 + 1 # encoded image token space + 1 for bos
OUTPUT_LENGTH = 256 + 1 # number of encoded tokens + 1 for bos
BOS_TOKEN_ID = 16384
BASE_MODEL = 'Wikidepia/IndoT5-large' # we currently have issues with bart-large
@dataclass
class ModelArguments:
"""
Arguments pertaining to which model/config/tokenizer we are going to fine-tune, or train from scratch.
"""
model_name_or_path: Optional[str] = field(
default=BASE_MODEL,
metadata={
"help": "The model checkpoint for weights initialization."
"Don't set if you want to train a model from scratch."
},
)
model_type: Optional[str] = field(
default=None,
metadata={"help": "If training from scratch, pass a model type from the list: " + ", ".join(MODEL_TYPES)},
)
config_name: Optional[str] = field(
default=None, metadata={"help": "Pretrained config name or path if not the same as model_name"}
)
tokenizer_name: Optional[str] = field(
default=None, metadata={"help": "Pretrained tokenizer name or path if not the same as model_name"}
)
cache_dir: Optional[str] = field(
default=None, metadata={"help": "Where do you want to store the pretrained models downloaded from s3"}
)
use_fast_tokenizer: bool = field(
default=True,
metadata={"help": "Whether to use one of the fast tokenizer (backed by the tokenizers library) or not."},
)
dtype: Optional[str] = field(
default="float32",
metadata={
"help": "Floating-point format in which the model weights should be initialized and trained. Choose one of `[float32, float16, bfloat16]`."
},
)
from_checkpoint: Optional[str] = field(
default=None,
metadata={
"help": "Loads a pretrained wandb checkpoint. Use artifact reference."
},
)
@dataclass
class DataTrainingArguments:
"""
Arguments pertaining to what data we are going to input our model for training and eval.
"""
dataset_name: Optional[str] = field(
default=None, metadata={"help": "The name of the dataset to use (via the datasets library)."}
)
dataset_config_name: Optional[str] = field(
default=None, metadata={"help": "The configuration name of the dataset to use (via the datasets library)."}
)
text_column: Optional[str] = field(
default='caption',
metadata={"help": "The name of the column in the datasets containing the full texts (for summarization)."},
)
encoding_column: Optional[str] = field(
default='encoding',
metadata={"help": "The name of the column in the datasets containing the image encodings."},
)
train_file: Optional[str] = field(default=None, metadata={"help": "The input training data file (a text file)."})
validation_file: Optional[str] = field(
default=None,
metadata={"help": "An optional input evaluation data file to evaluate the perplexity on (a text file)."},
)
test_file: Optional[str] = field(
default=None,
metadata={"help": "An optional input predict data file to do prediction on (a text file)."},
)
max_source_length: Optional[int] = field(
default=128,
metadata={
"help": "The maximum total input sequence length after tokenization. Sequences longer "
"than this will be truncated, sequences shorter will be padded."
},
)
no_decay: bool = field(
default=False, metadata={"help": "Whether to use decay in the learning rate scheduler."}
)
max_target_length: Optional[int] = field(
default=OUTPUT_LENGTH,
metadata={
"help": "The maximum total sequence length for target text after tokenization. Sequences longer "
"than this will be truncated, sequences shorter will be padded."
},
)
val_max_target_length: Optional[int] = field(
default=OUTPUT_LENGTH,
metadata={
"help": "The maximum total sequence length for validation target text after tokenization. Sequences longer "
"than this will be truncated, sequences shorter will be padded. Will default to `max_target_length`."
"This argument is also used to override the `max_length` param of `model.generate`, which is used "
"during evaluation."
},
)
max_train_samples: Optional[int] = field(
default=None,
metadata={
"help": "For debugging purposes or quicker training, truncate the number of training examples to this "
"value if set."
},
)
max_eval_samples: Optional[int] = field(
default=None,
metadata={
"help": "For debugging purposes or quicker training, truncate the number of evaluation examples to this "
"value if set."
},
)
max_predict_samples: Optional[int] = field(
default=None,
metadata={
"help": "For debugging purposes or quicker training, truncate the number of prediction examples to this "
"value if set."
},
)
preprocessing_num_workers: Optional[int] = field(
default=80, # ensure we have the same datasets cached data and avoid using too much space
metadata={"help": "The number of processes to use for the preprocessing."},
)
source_prefix: Optional[str] = field(
default=None, metadata={"help": "A prefix to add before every source text (useful for T5 models)."}
)
predict_with_generate: bool = field(
default=False, metadata={"help": "Whether to use generate to calculate generative metrics."}
)
num_beams: Optional[int] = field(
default=None,
metadata={
"help": "Number of beams to use for evaluation. This argument will be passed to `model.generate`, "
"which is used during evaluation."
},
)
overwrite_cache: bool = field(
default=False, metadata={"help": "Overwrite the cached training and evaluation sets"}
)
log_interval: Optional[int] = field(
default=40,
metadata={
"help": "For debugging purposes or quicker training, truncate the number of training examples to this "
"value if set."
},
)
log_model: bool = field(
default=False, metadata={"help": "Overwrite the cached training and evaluation sets"}
)
save_model_steps: Optional[int] = field(
default=3000, # about once every hour in our experiments
metadata={
"help": "For logging the model more frequently. Used only when `log_model` is set."
},
)
def __post_init__(self):
if self.dataset_name is None and self.train_file is None and self.validation_file is None:
raise ValueError("Need either a dataset name or a training/validation file.")
else:
if self.train_file is not None:
extension = self.train_file.split(".")[-1]
assert extension in ["tsv", "csv", "json"], "`train_file` should be a tsv, csv or json file."
if self.validation_file is not None:
extension = self.validation_file.split(".")[-1]
assert extension in ["tsv", "csv", "json"], "`validation_file` should be a tsv, csv or json file."
if self.val_max_target_length is None:
self.val_max_target_length = self.max_target_length
class TrainState(train_state.TrainState):
dropout_rng: jnp.ndarray
grad_accum: jnp.ndarray
optimizer_step: int
def replicate(self):
return jax_utils.replicate(self).replace(dropout_rng=shard_prng_key(self.dropout_rng))
class CustomT5Module(FlaxT5Module):
def setup(self):
self.config.vocab_size_output = getattr(self.config, 'vocab_size_output', OUTPUT_VOCAB_SIZE)
self.config.max_position_embeddings_decoder = getattr(self.config, 'max_position_embeddings_decoder', OUTPUT_LENGTH)
self.shared = nn.Embed(
self.config.vocab_size,
self.config.d_model,
embedding_init=jax.nn.initializers.normal(self.config.initializer_factor * 1.0, self.dtype),
dtype=self.dtype,
)
self.decoder_embed = nn.Embed(
self.config.vocab_size_output,
self.config.d_model,
embedding_init=jax.nn.initializers.normal(self.config.initializer_factor * 1.0, self.dtype),
dtype=self.dtype,
)
encoder_config = copy.deepcopy(self.config)
encoder_config.causal = False
self.encoder = FlaxT5Stack(encoder_config, embed_tokens=self.shared, dtype=self.dtype)
decoder_config = copy.deepcopy(self.config)
decoder_config.causal = True
decoder_config.num_layers = self.config.num_decoder_layers
decoder_config.max_position_embeddings = self.config.max_position_embeddings_decoder
decoder_config.vocab_size = self.config.vocab_size_output
self.decoder = FlaxT5Stack(decoder_config, embed_tokens=self.decoder_embed, dtype=self.dtype)
class CustomFlaxT5ForConditionalGenerationModule(FlaxT5ForConditionalGenerationModule):
def setup(self):
# check config is valid, otherwise set default values
self.config.vocab_size_output = getattr(self.config, 'vocab_size_output', OUTPUT_VOCAB_SIZE)
self.config.max_position_embeddings_decoder = getattr(self.config, 'max_position_embeddings_decoder', OUTPUT_LENGTH)
self.shared = nn.Embed(
self.config.vocab_size,
self.config.d_model,
embedding_init=jax.nn.initializers.normal(self.config.initializer_factor * 1.0, self.dtype),
dtype=self.dtype,
)
self.decoder_embed = nn.Embed(
self.config.vocab_size_output,
self.config.d_model,
embedding_init=jax.nn.initializers.normal(self.config.initializer_factor * 1.0, self.dtype),
dtype=self.dtype,
)
encoder_config = copy.deepcopy(self.config)
encoder_config.causal = False
self.encoder = FlaxT5Stack(encoder_config, embed_tokens=self.shared, dtype=self.dtype)
decoder_config = copy.deepcopy(self.config)
decoder_config.causal = True
decoder_config.num_layers = self.config.num_decoder_layers
decoder_config.max_position_embeddings = self.config.max_position_embeddings_decoder
decoder_config.vocab_size = self.config.vocab_size_output
self.decoder = FlaxT5Stack(decoder_config, embed_tokens=self.decoder_embed, dtype=self.dtype)
self.model = CustomT5Module(config=self.config, dtype=self.dtype)
self.lm_head = nn.Dense(
self.config.vocab_size_output,
use_bias=False,
dtype=self.dtype,
kernel_init=jax.nn.initializers.normal(self.config.initializer_factor, self.dtype),
)
#self.final_logits_bias = self.param("final_logits_bias", self.bias_init, (1, self.config.vocab_size_output))
class CustomFlaxT5ForConditionalGeneration(FlaxT5ForConditionalGeneration):
module_class = CustomFlaxT5ForConditionalGenerationModule
def data_loader(rng: jax.random.PRNGKey, dataset: Dataset, batch_size: int, shuffle: bool = False):
"""
Returns batches of size `batch_size` from truncated `dataset`, sharded over all local devices.
Shuffle batches if `shuffle` is `True`.
"""
steps_per_epoch = len(dataset) // batch_size
if shuffle:
batch_idx = jax.random.permutation(rng, len(dataset))
else:
batch_idx = jnp.arange(len(dataset))
batch_idx = batch_idx[: steps_per_epoch * batch_size] # Skip incomplete batch.
batch_idx = batch_idx.reshape((steps_per_epoch, batch_size))
for idx in batch_idx:
batch = dataset[idx]
batch = {k: jnp.array(v) for k, v in batch.items()}
batch = shard(batch)
yield batch
def create_learning_rate_fn(
train_ds_size: int, train_batch_size: int, num_train_epochs: int, num_warmup_steps: int, learning_rate: float, no_decay: bool
) -> Callable[[int], jnp.array]:
"""Returns a linear warmup, linear_decay learning rate function."""
steps_per_epoch = train_ds_size // train_batch_size
num_train_steps = steps_per_epoch * num_train_epochs
warmup_fn = optax.linear_schedule(init_value=0.0, end_value=learning_rate, transition_steps=num_warmup_steps)
if no_decay:
return warmup_fn
decay_fn = optax.linear_schedule(
init_value=learning_rate, end_value=0, transition_steps=num_train_steps - num_warmup_steps
)
schedule_fn = optax.join_schedules(schedules=[warmup_fn, decay_fn], boundaries=[num_warmup_steps])
return schedule_fn
def wandb_log(metrics, step=None, prefix=None):
if jax.process_index() == 0:
log_metrics = {f'{prefix}/{k}' if prefix is not None else k: jax.device_get(v) for k,v in metrics.items()}
if step is not None:
log_metrics['train/step'] = step
wandb.log(log_metrics)
def main():
# See all possible arguments in src/transformers/training_args.py
# or by passing the --help flag to this script.
# We now keep distinct sets of args, for a cleaner separation of concerns.
parser = HfArgumentParser((ModelArguments, DataTrainingArguments, TrainingArguments))
if len(sys.argv) == 2 and sys.argv[1].endswith(".json"):
# If we pass only one argument to the script and it's the path to a json file,
# let's parse it to get our arguments.
model_args, data_args, training_args = parser.parse_json_file(json_file=os.path.abspath(sys.argv[1]))
else:
model_args, data_args, training_args = parser.parse_args_into_dataclasses()
if (
os.path.exists(training_args.output_dir)
and os.listdir(training_args.output_dir)
and training_args.do_train
and not training_args.overwrite_output_dir
):
raise ValueError(
f"Output directory ({training_args.output_dir}) already exists and is not empty."
"Use --overwrite_output_dir to overcome."
)
# Set up wandb run
wandb.init(
entity='munggok',
project='dalle-indo',
job_type='Seq2SeqVQGAN',
config=parser.parse_args()
)
# set default x-axis as 'train/step'
wandb.define_metric('train/step')
wandb.define_metric('*', step_metric='train/step')
# Make one log on every process with the configuration for debugging.
pylogging.basicConfig(
format="%(asctime)s - %(levelname)s - %(name)s - %(message)s",
datefmt="%m/%d/%Y %H:%M:%S",
level=pylogging.INFO,
)
# Setup logging, we only want one process per machine to log things on the screen.
logger.setLevel(pylogging.INFO if jax.process_index() == 0 else pylogging.ERROR)
if jax.process_index() == 0:
datasets.utils.logging.set_verbosity_warning()
transformers.utils.logging.set_verbosity_info()
else:
datasets.utils.logging.set_verbosity_error()
transformers.utils.logging.set_verbosity_error()
# Set the verbosity to info of the Transformers logger (on main process only):
logger.info(f"Training/evaluation parameters {training_args}")
# Get the datasets: you can either provide your own CSV/JSON training and evaluation files (see below)
# or just provide the name of one of the public datasets available on the hub at https://huggingface.co/datasets/
# (the dataset will be downloaded automatically from the datasets Hub).
#
data_files = {}
if data_args.train_file is not None:
data_files["train"] = data_args.train_file
if data_args.validation_file is not None:
data_files["validation"] = data_args.validation_file
if data_args.test_file is not None:
data_files["test"] = data_args.test_file
dataset = load_dataset("csv", data_files=data_files, cache_dir=model_args.cache_dir, delimiter="\t")
# See more about loading any type of standard or custom dataset (from files, python dict, pandas DataFrame, etc) at
# https://huggingface.co/docs/datasets/loading_datasets.html.
# Set up items to load or create
tokenizer = None
artifact_dir = None
def restore_state(state, artifact_dir):
# restore optimizer state
if (Path(artifact_dir) / 'opt_state.msgpack').exists():
with (Path(artifact_dir) / 'opt_state.msgpack').open('rb') as f:
opt_state = from_bytes(state.opt_state, f.read())
# restore steps
if (Path(artifact_dir) / 'training_state.json').exists():
with (Path(artifact_dir) / 'training_state.json').open('r') as f:
training_state = json.load(f)
step = training_state['step']
optimizer_step = step // training_args.gradient_accumulation_steps
state.replace(step=step, optimizer_step=optimizer_step)
if model_args.from_checkpoint is not None:
artifact = wandb.run.use_artifact(model_args.from_checkpoint)
artifact_dir = artifact.download()
model = CustomFlaxT5ForConditionalGeneration.from_pretrained(artifact_dir)
# some models will try to change bos (because of force_bos_token_to_be_generated)
# we ensure bos and eos are not forced
model.config.force_bos_token_to_be_generated = False
model.config.forced_bos_token_id = None
model.config.forced_eos_token_id = None
# used in the preprocessing function
config = model.config
# load tokenizer if present
if (Path(artifact_dir) / 'tokenizer_config.json').exists():
tokenizer = AutoTokenizer.from_pretrained(
model_args.model_name_or_path, cache_dir=model_args.cache_dir, use_fast=model_args.use_fast_tokenizer
)
else:
base_model = FlaxAutoModelForSeq2SeqLM.from_pretrained(
model_args.model_name_or_path, seed=training_args.seed, dtype=getattr(jnp, model_args.dtype), from_pt=True
)
# Set up our new model config
config = T5Config.from_pretrained(model_args.model_name_or_path)
config.tie_word_embeddings = False
config.decoder_start_token_id = BOS_TOKEN_ID # for first token
config.bos_token_id = BOS_TOKEN_ID # should not be used (due to forced_bos_token_id)
config.pos_token_id = BOS_TOKEN_ID # should not be needed (as we generate until max_length)
config.eos_token_id = BOS_TOKEN_ID + 1 # unreachable
config.forced_bos_token_id = None # we don't need this token
config.forced_eos_token_id = None # we don't need this token
config.force_bos_token_to_be_generated = False # otherwise it sets bos_token_id at loading
config.min_length = data_args.max_target_length
config.max_length = data_args.max_target_length
# Create a custom model and initialize it randomly
model = CustomFlaxT5ForConditionalGeneration(config, seed=training_args.seed, dtype=getattr(jnp, model_args.dtype))
# Use pre-trained weights for encoder
model.params['encoder'] = base_model.params['encoder']
model.params['shared'] = base_model.params['shared']
del base_model
# Load tokenizer if it has not been set
if tokenizer is None:
tokenizer = AutoTokenizer.from_pretrained(
model_args.model_name_or_path, cache_dir=model_args.cache_dir, use_fast=model_args.use_fast_tokenizer
)
print(f"TPUs: {jax.device_count()}")
assert jax.device_count() == 8, "TPUs in use, please check running processes"
prefix = data_args.source_prefix if data_args.source_prefix is not None else ""
# Preprocessing the datasets.
# We need to tokenize inputs and targets.
if training_args.do_train:
column_names = dataset["train"].column_names
elif training_args.do_eval:
column_names = dataset["validation"].column_names
elif training_args.do_predict:
column_names = dataset["test"].column_names
else:
logger.info("There is nothing to do. Please pass `do_train`, `do_eval` and/or `do_predict`.")
return
# Get the column names for input/target.
text_column = data_args.text_column
encoding_column = data_args.encoding_column
# Temporarily set max_target_length for training.
max_target_length = data_args.max_target_length
def shift_tokens_right(input_ids: np.array, decoder_start_token_id: int):
"""
Shift input ids one token to the right.
"""
shifted_input_ids = np.zeros(input_ids.shape)
shifted_input_ids[:, 1:] = input_ids[:, :-1]
shifted_input_ids[:, 0] = decoder_start_token_id
return shifted_input_ids
def preprocess_function(examples):
inputss = examples[text_column]
inputs = []
for inp in inputss:
if inp is not None:
inputs.append(prefix + inp)
else:
inputs.append('' + "sebuah gambar")
# Setting padding="max_length" as we need fixed length inputs for jitted functions
model_inputs = tokenizer(
inputs, max_length=data_args.max_source_length, padding="max_length", truncation=True, return_tensors="np"
)
# set up targets
# Note: labels correspond to our target indices
# decoder input ids are the same but shifted to the right with bos at the beginning (and without last token)
labels = [eval(indices) for indices in examples['encoding']]
labels = np.asarray(labels)
# We need the labels, in addition to the decoder_input_ids, for the compute_loss function
model_inputs["labels"] = labels
# In our case, this prepends the bos token and removes the last one
decoder_input_ids = shift_tokens_right(labels, config.decoder_start_token_id)
model_inputs["decoder_input_ids"] = decoder_input_ids
return model_inputs
if training_args.do_train:
if "train" not in dataset:
raise ValueError("--do_train requires a train dataset")
train_dataset = dataset["train"]
if data_args.max_train_samples is not None:
train_dataset = train_dataset.select(range(data_args.max_train_samples))
train_dataset = train_dataset.map(
preprocess_function,
batched=True,
num_proc=data_args.preprocessing_num_workers,
remove_columns=column_names,
load_from_cache_file=not data_args.overwrite_cache,
desc="Running tokenizer on train dataset",
)
if training_args.do_eval:
max_target_length = data_args.val_max_target_length
if "validation" not in dataset:
raise ValueError("--do_eval requires a validation dataset")
eval_dataset = dataset["validation"]
if data_args.max_eval_samples is not None:
eval_dataset = eval_dataset.select(range(data_args.max_eval_samples))
eval_dataset = eval_dataset.map(
preprocess_function,
batched=True,
num_proc=data_args.preprocessing_num_workers,
remove_columns=column_names,
load_from_cache_file=not data_args.overwrite_cache,
desc="Running tokenizer on validation dataset",
)
if training_args.do_predict:
max_target_length = data_args.val_max_target_length
if "test" not in dataset:
raise ValueError("--do_predict requires a test dataset")
predict_dataset = dataset["test"]
if data_args.max_predict_samples is not None:
predict_dataset = predict_dataset.select(range(data_args.max_predict_samples))
predict_dataset = predict_dataset.map(
preprocess_function,
batched=True,
num_proc=data_args.preprocessing_num_workers,
remove_columns=column_names,
load_from_cache_file=not data_args.overwrite_cache,
desc="Running tokenizer on prediction dataset",
)
# Initialize our training
rng = jax.random.PRNGKey(training_args.seed)
rng, dropout_rng = jax.random.split(rng)
# Store some constant
num_epochs = int(training_args.num_train_epochs)
train_batch_size = int(training_args.per_device_train_batch_size) * jax.device_count()
total_batch_size = int(train_batch_size) * training_args.gradient_accumulation_steps
eval_batch_size = int(training_args.per_device_eval_batch_size) * jax.device_count()
steps_per_epoch = len(train_dataset) // train_batch_size
total_steps = steps_per_epoch * num_epochs
total_optimization_steps = (len(train_dataset) // total_batch_size) * num_epochs
# Create learning rate schedule
linear_decay_lr_schedule_fn = create_learning_rate_fn(
len(train_dataset),
total_batch_size,
training_args.num_train_epochs,
training_args.warmup_steps,
training_args.learning_rate,
data_args.no_decay
)
# We use Optax's "masking" functionality to not apply weight decay
# to bias and LayerNorm scale parameters. decay_mask_fn returns a
# mask boolean with the same structure as the parameters.
# The mask is True for parameters that should be decayed.
# Note that this mask is specifically adapted for FlaxBart.
# For FlaxT5, one should correct the layer norm parameter naming
# accordingly - see `run_t5_mlm_flax.py` e.g.
def decay_mask_fn(params):
flat_params = traverse_util.flatten_dict(params)
layer_norm_params = [
(name, "scale") for name in ["self_attn_layer_norm", "layernorm_embedding", "final_layer_norm"]
]
flat_mask = {path: (path[-1] != "bias" and path[-2:] not in layer_norm_params) for path in flat_params}
return traverse_util.unflatten_dict(flat_mask)
# create adam optimizer
if training_args.adafactor:
# We use the default parameters here to initialize adafactor,
# For more details about the parameters please check https://github.com/deepmind/optax/blob/ed02befef9bf81cbbf236be3d2b0e032e9ed4a40/optax/_src/alias.py#L74
optimizer = optax.adafactor(
learning_rate=linear_decay_lr_schedule_fn,
)
else:
optimizer = optax.adamw(
learning_rate=linear_decay_lr_schedule_fn,
b1=training_args.adam_beta1,
b2=training_args.adam_beta2,
eps=training_args.adam_epsilon,
weight_decay=training_args.weight_decay,
mask=decay_mask_fn,
)
# Setup train state
state = TrainState.create(
apply_fn=model.__call__,
params=model.params,
tx=optimizer,
dropout_rng=dropout_rng,
grad_accum=jax.tree_map(jnp.zeros_like, model.params),
optimizer_step=0,
)
if model_args.from_checkpoint is not None:
# restore optimizer state, step and optimizer_step
restore_state(state, artifact_dir)
# label smoothed cross entropy
def loss_fn(logits, labels):
loss = optax.softmax_cross_entropy(logits, onehot(labels, logits.shape[-1]))
loss = loss.mean()
return loss
# Define gradient update step fn
def train_step(state, batch):
dropout_rng, new_dropout_rng = jax.random.split(state.dropout_rng)
def compute_loss(params):
labels = batch.pop("labels")
logits = state.apply_fn(**batch, params=params, dropout_rng=dropout_rng, train=True)[0]
loss = loss_fn(logits, labels)
return loss
grad_fn = jax.value_and_grad(compute_loss)
loss, grads = grad_fn(state.params)
grad_accum = jax.tree_multimap(lambda x, y: x + y, grads, state.grad_accum)
def update_fn():
grads = jax.tree_map(lambda x: x / training_args.gradient_accumulation_steps, grad_accum)
grads = jax.lax.pmean(grads, "batch")
new_state = state.apply_gradients(
grads=grads, grad_accum=jax.tree_map(jnp.zeros_like, grads), optimizer_step=state.optimizer_step + 1
)
return new_state
new_state = jax.lax.cond(
(state.step + 1) % training_args.gradient_accumulation_steps == 0,
lambda _: update_fn(),
lambda _: state.replace(grad_accum=grad_accum, step=state.step + 1),
None,
)
metrics = {"loss": loss, "learning_rate": linear_decay_lr_schedule_fn(state.optimizer_step)}
metrics = jax.lax.pmean(metrics, axis_name="batch")
return new_state.replace(dropout_rng=new_dropout_rng), metrics
# Define eval fn
def eval_step(params, batch):
labels = batch.pop("labels")
logits = model(**batch, params=params, train=False)[0]
loss = loss_fn(logits, labels)
# summarize metrics
metrics = {"loss": loss}
metrics = jax.lax.pmean(metrics, axis_name="batch")
return metrics
# Define generation function
max_length = (
data_args.val_max_target_length if data_args.val_max_target_length is not None else model.config.max_length
)
num_beams = data_args.num_beams if data_args.num_beams is not None else model.config.num_beams
gen_kwargs = {"max_length": max_length, "num_beams": num_beams}
def generate_step(params, batch):
model.params = params
output_ids = model.generate(batch["input_ids"], attention_mask=batch["attention_mask"], **gen_kwargs)
return output_ids.sequences
# Create parallel version of the train and eval step
p_train_step = jax.pmap(
train_step, "batch", donate_argnums=(0,)
)
p_eval_step = jax.pmap(eval_step, "batch")
p_generate_step = jax.pmap(generate_step, "batch")
# Replicate the train state on each device
state = state.replicate()
logger.info("***** Running training *****")
logger.info(f" Num examples = {len(train_dataset)}")
logger.info(f" Num Epochs = {num_epochs}")
logger.info(f" Instantaneous batch size per device = {training_args.per_device_train_batch_size}")
logger.info(
f" Total train batch size (w. parallel & distributed) = {train_batch_size * training_args.gradient_accumulation_steps}"
)
logger.info(f" Total global steps = {total_steps}")
logger.info(f" Total optimization steps = {total_optimization_steps}")
train_time = 0
epochs = tqdm(range(num_epochs), desc=f"Epoch ... (1/{num_epochs})", position=0)
global_step = 0
def run_evaluation():
# ======================== Evaluating ==============================
eval_metrics = []
if training_args.do_eval:
eval_preds = []
eval_labels = []
eval_loader = data_loader(input_rng, eval_dataset, eval_batch_size)
eval_steps = len(eval_dataset) // eval_batch_size
for _ in tqdm(range(eval_steps), desc="Evaluating...", position=2, leave=False):
# Model forward
batch = next(eval_loader)
labels = batch["labels"]
metrics = p_eval_step(state.params, batch)
eval_metrics.append(metrics)
# generation
if data_args.predict_with_generate:
generated_ids = p_generate_step(state.params, batch)
eval_preds.extend(jax.device_get(generated_ids.reshape(-1, gen_kwargs["max_length"])))
eval_labels.extend(jax.device_get(labels.reshape(-1, labels.shape[-1])))
# normalize eval metrics
eval_metrics = get_metrics(eval_metrics)
eval_metrics = jax.tree_map(jnp.mean, eval_metrics)
# log metrics
wandb_log(eval_metrics, step=global_step, prefix='eval')
# Print metrics and update progress bar
desc = f"Epoch... ({epoch + 1}/{num_epochs} | Eval Loss: {eval_metrics['loss']})"
epochs.write(desc)
epochs.desc = desc
return eval_metrics
def run_save_model(state, step, epoch, eval_metrics=None):
if jax.process_index() == 0:
params = jax.device_get(jax.tree_map(lambda x: x[0], state.params))
# save model locally
model.save_pretrained(
training_args.output_dir,
params=params,
)
# save tokenizer
tokenizer.save_pretrained(training_args.output_dir)
# save state
state = unreplicate(state)
with (Path(training_args.output_dir) / 'opt_state.msgpack').open('wb') as f:
f.write(to_bytes(state.opt_state))
with (Path(training_args.output_dir) / 'training_state.json').open('w') as f:
json.dump({'step': state.step.item()}, f)
# save to W&B
if data_args.log_model:
metadata = {'step': step, 'epoch': epoch}
if eval_metrics is not None:
metadata['eval/loss'] = eval_metrics['loss']
artifact = wandb.Artifact(
name=f"model-{wandb.run.id}", type="bart_model", metadata=metadata
)
artifact.add_file(str(Path(training_args.output_dir) / 'flax_model.msgpack'))
artifact.add_file(str(Path(training_args.output_dir) / 'config.json'))
artifact.add_file(str(Path(training_args.output_dir) / 'tokenizer.json'))
artifact.add_file(str(Path(training_args.output_dir) / 'tokenizer_config.json'))
artifact.add_file(str(Path(training_args.output_dir) / 'vocab.json'))
artifact.add_file(str(Path(training_args.output_dir) / 'merges.txt'))
artifact.add_file(str(Path(training_args.output_dir) / 'special_tokens_map.json'))
artifact.add_file(str(Path(training_args.output_dir) / 'opt_state.msgpack'))
artifact.add_file(str(Path(training_args.output_dir) / 'training_state.json'))
wandb.run.log_artifact(artifact)
# save to the hub
if training_args.push_to_hub:
model.save_pretrained(
training_args.output_dir,
params=params,
push_to_hub=training_args.push_to_hub,
commit_message=f"Saving weights and logs of epoch {epoch+1}",
temp_dir=True # avoid issues with being in a repository
)
for epoch in epochs:
# ======================== Training ================================
train_start = time.time()
# Create sampling rng
rng, input_rng = jax.random.split(rng)
# Generate an epoch by shuffling sampling indices from the train dataset
train_loader = data_loader(input_rng, train_dataset, train_batch_size, shuffle=True)
steps_per_epoch = len(train_dataset) // train_batch_size
# train
for step in tqdm(range(steps_per_epoch), desc="Training...", position=1, leave=False):
global_step +=1
batch = next(train_loader)
state, train_metric = p_train_step(state, batch)
if global_step % data_args.log_interval == 0 and jax.process_index() == 0:
# log metrics
wandb_log(unreplicate(train_metric), step=global_step, prefix='train')
if global_step % training_args.eval_steps == 0:
run_evaluation()
if global_step % data_args.save_model_steps == 0:
run_save_model(state, global_step, epoch)
# log final train metrics
wandb_log(unreplicate(train_metric), step=global_step, prefix='train')
train_time += time.time() - train_start
train_metric = unreplicate(train_metric)
epochs.write(
f"Epoch... ({epoch + 1}/{num_epochs} | Loss: {train_metric['loss']}, Learning Rate: {train_metric['learning_rate']})"
)
# Final evaluation
eval_metrics = run_evaluation()
# save checkpoint after each epoch and push checkpoint to the hub
run_save_model(state, global_step, epoch, eval_metrics)
# ======================== Prediction loop ==============================
if training_args.do_predict:
logger.info("*** Predict ***")
pred_metrics = []
pred_generations = []
pred_labels = []
pred_loader = data_loader(input_rng, predict_dataset, eval_batch_size)
pred_steps = len(predict_dataset) // eval_batch_size
for _ in tqdm(range(pred_steps), desc="Predicting...", position=2, leave=False):
# Model forward
batch = next(pred_loader)
labels = batch["labels"]
metrics = p_eval_step(state.params, batch)
pred_metrics.append(metrics)
# generation
if data_args.predict_with_generate:
generated_ids = p_generate_step(state.params, batch)
pred_generations.extend(jax.device_get(generated_ids.reshape(-1, gen_kwargs["max_length"])))
pred_labels.extend(jax.device_get(labels.reshape(-1, labels.shape[-1])))
# normalize prediction metrics
pred_metrics = get_metrics(pred_metrics)
pred_metrics = jax.tree_map(jnp.mean, pred_metrics)
# Print metrics
desc = f"Predict Loss: {pred_metrics['loss']})"
logger.info(desc)
if __name__ == "__main__":
main()
|
from __future__ import print_function, division
import numpy
from west.propagators import WESTPropagator
from west.systems import WESTSystem
from westpa.binning import RectilinearBinMapper
PI = numpy.pi
from numpy import sin, cos, exp
from numpy.random import normal as random_normal
pcoord_len = 21
pcoord_dtype = numpy.float32
class ODLDPropagator(WESTPropagator):
def __init__(self, rc=None):
super(ODLDPropagator,self).__init__(rc)
self.coord_len = pcoord_len
self.coord_dtype = pcoord_dtype
self.coord_ndim = 1
self.initial_pcoord = numpy.array([8.0], dtype=self.coord_dtype)
self.sigma = 0.001**(0.5)
self.A = 2
self.B = 10
self.C = 0.5
self.x0 = 1
# Implement a reflecting boundary at this x value
# (or None, for no reflection)
self.reflect_at = 10.0
#self.reflect_at = None
def get_pcoord(self, state):
'''Get the progress coordinate of the given basis or initial state.'''
state.pcoord = self.initial_pcoord.copy()
def gen_istate(self, basis_state, initial_state):
initial_state.pcoord = self.initial_pcoord.copy()
initial_state.istate_status = initial_state.ISTATE_STATUS_PREPARED
return initial_state
def propagate(self, segments):
A, B, C, x0 = self.A, self.B, self.C, self.x0
n_segs = len(segments)
coords = numpy.empty((n_segs, self.coord_len, self.coord_ndim), dtype=self.coord_dtype)
for iseg, segment in enumerate(segments):
coords[iseg,0] = segment.pcoord[0]
twopi_by_A = 2*PI/A
half_B = B/2
sigma = self.sigma
gradfactor = self.sigma*self.sigma/2
coord_len = self.coord_len
reflect_at = self.reflect_at
all_displacements = numpy.zeros((n_segs, self.coord_len, self.coord_ndim), dtype=self.coord_dtype)
for istep in xrange(1,coord_len):
x = coords[:,istep-1,0]
xarg = twopi_by_A*(x - x0)
eCx = numpy.exp(C*x)
eCx_less_one = eCx - 1.0
all_displacements[:,istep,0] = displacements = random_normal(scale=sigma, size=(n_segs,))
grad = half_B / (eCx_less_one*eCx_less_one)*(twopi_by_A*eCx_less_one*sin(xarg)+C*eCx*cos(xarg))
newx = x - gradfactor*grad + displacements
if reflect_at is not None:
# Anything that has moved beyond reflect_at must move back that much
# boolean array of what to reflect
to_reflect = newx > reflect_at
# how far the things to reflect are beyond our boundary
reflect_by = newx[to_reflect] - reflect_at
# subtract twice how far they exceed the boundary by
# puts them the same distance from the boundary, on the other side
newx[to_reflect] -= 2*reflect_by
coords[:,istep,0] = newx
for iseg, segment in enumerate(segments):
segment.pcoord[...] = coords[iseg,:]
segment.data['displacement'] = all_displacements[iseg]
segment.status = segment.SEG_STATUS_COMPLETE
return segments
class ODLDSystem(WESTSystem):
def initialize(self):
self.pcoord_ndim = 1
self.pcoord_dtype = pcoord_dtype
self.pcoord_len = pcoord_len
#self.bin_mapper = RectilinearBinMapper([[0,1.3] + list(numpy.arange(1.4, 10.1, 0.1)) + [float('inf')]])
self.bin_mapper = RectilinearBinMapper([ list(numpy.arange(0.0, 10.1, 0.1)) ])
self.bin_target_counts = numpy.empty((self.bin_mapper.nbins,), numpy.int_)
self.bin_target_counts[...] = 10
|
<filename>visualize_json.py
import sys
import cv2
import json
if len(sys.argv) < 4:
print("Usage: python visualize_json.py json_annotation input_im output_im")
exit()
def find_by_id(_id, objs):
for obj in objs:
if obj['id'] == _id:
return obj
return None
in_json_file = sys.argv[1]
in_im_file = sys.argv[2]
out_file = sys.argv[3]
in_obj = json.load(open(in_json_file))
im = cv2.imread(in_im_file, 1)
h,w,_ = im.shape
# role_colors is also used to color lineplot lines
role_colors = [ (255, 0, 0), (0, 255, 0), (0, 0, 255), (255, 255, 0), (255, 0, 255), (0, 255, 255),
(255, 128, 0)]
patch_color = (255, 0, 128)
patch_connect = (255, 128, 0)
element_colors = [(0, 255, 128), (128, 255, 0), (0, 128, 255), (128, 0, 255), (128, 128, 0), (128, 0, 128), (0, 128, 128) ]
text_role_mapping = {'chart_title': 0, 'axis_title': 1, 'tick_label': 2, 'legend_title': 3, 'legend_label': 4}
boxplot_mapping = {'median': 0, 'min': 1, 'max': 2, 'first_quartile': 3, 'third_quartile': 4}
# task 1
chart_type = in_obj['task1']['output']['chart_type']
cv2.putText(im, chart_type, (10, 10), cv2.FONT_HERSHEY_SIMPLEX, 0.5, (0, 0, 255), thickness=2, bottomLeftOrigin=False)
if chart_type in ['Pie', 'Donut']:
cv2.imwrite(out_file, im)
exit()
# task 2 & 3
for text_block_2_op, text_block_3_ip in zip(in_obj['task2']['output']['text_blocks'], in_obj['task3']['input']['task2_output']['text_blocks']):
bb = text_block_2_op['bb']
_id = text_block_3_ip['id']
obj = find_by_id(_id, in_obj['task3']['output']['text_roles'])
assert obj is not None
role = obj['role']
color = role_colors[text_role_mapping[role]]
p1 = (int(bb['x0']), int(bb['y0']))
p2 = (int(bb['x0'] + bb['width']), int(bb['y0'] + bb['height']))
cv2.rectangle(im, p1, p2, color, thickness=2)
#if role == 'axis_title':
cv2.putText(im, text_block_2_op['text'], p2, cv2.FONT_HERSHEY_SIMPLEX, 0.4, (0, 0, 255), thickness=2, bottomLeftOrigin=False)
# task 4
bb = in_obj['task4']['output']['_plot_bb']
p1 = (int(bb['x0']), int(bb['y0']))
p2 = (int(bb['x0'] + bb['width']), int(bb['y0'] + bb['height']))
cv2.rectangle(im, p1, p2, (0, 255, 0), thickness=2)
for axis, color in [('x', (255, 0, 0)), ('y', (255, 0, 255))]:
for tick_obj in in_obj['task4']['output']['axes']['%s-axis' % axis]:
_id = tick_obj['id']
pt = tick_obj['tick_pt']
x = pt['x']
y = pt['y']
cv2.circle(im, (x, y), 2, color, thickness=-1)
label_bb = find_by_id(_id, in_obj['task5']['input']['task2_output']['text_blocks'])['bb']
p = (int(label_bb['x0']), int(label_bb['y0']))
cv2.line(im, (x,y), p, color, thickness=1)
# task 5
for obj in in_obj['task5']['output']['legend_pairs']:
patch_bb = obj['bb']
p1 = (int(patch_bb['x0']), int(patch_bb['y0']))
p2 = (int(patch_bb['x0'] + patch_bb['width']), int(patch_bb['y0'] + patch_bb['height']))
cv2.rectangle(im, p1, p2, patch_color, thickness=2)
_id = obj['id']
label_bb = find_by_id(_id, in_obj['task5']['input']['task2_output']['text_blocks'])['bb']
p = (int(label_bb['x0']), int(label_bb['y0']))
cv2.line(im, p1, p, patch_connect, thickness=1)
# task 6
idx = 0
for bb in in_obj['task6']['output']['visual elements']['bars']:
p1 = (int(bb['x0']), int(bb['y0']))
p2 = (int(bb['x0'] + bb['width']), int(bb['y0'] + bb['height']))
color = element_colors[idx % len(element_colors)]
cv2.rectangle(im, p1, p2, color, thickness=2)
idx += 1
for boxplot in in_obj['task6']['output']['visual elements']['boxplots']:
for name, obj in boxplot.items():
color = element_colors[boxplot_mapping[name]]
x = obj['x']
y = obj['y']
bb = obj['_bb']
p1 = (int(bb['x0']), int(bb['y0']))
p2 = (int(bb['x0'] + bb['width']), int(bb['y0'] + bb['height']))
cv2.rectangle(im, p1, p2, color, thickness=1)
cv2.circle(im, (x, y), 3, color, thickness=-1)
idx = 0
for line in in_obj['task6']['output']['visual elements']['lines']:
for pt in line:
x = pt['x']
y = pt['y']
color = role_colors[idx % len(role_colors)]
cv2.circle(im, (x, y), 2, color, thickness=-1)
idx += 1
idx = 0
for pt in in_obj['task6']['output']['visual elements']['scatter points']:
x = pt['x']
y = pt['y']
color = element_colors[idx % len(element_colors)]
cv2.circle(im, (x, y), 2, color, thickness=-1)
idx += 1
cv2.imwrite(out_file, im)
|
<gh_stars>10-100
import torch
import argparse
import os
import os.path as osp
import numpy as np
import torch.nn as nn
from datetime import datetime
from torch.autograd import Variable
import torch.optim as optim
import warnings
from torch.utils import model_zoo
warnings.filterwarnings("ignore")
#===========
import network
import Utilizes
import Data
#==============================================================================
# Train Function
#==============================================================================
def train(config):
# ================== Pass the Config
dset_name = config['dset']
source_domain_set, target_domain_set, save_name, n_classes = \
Data.Get_Domain_Meta(dset_name)
Exptimes = int(config['exp_time'])
num_tasks = len(source_domain_set)
batch_size = int(config['bs'])
epochs = int(config['maxEpo'])
# ========== Hyperparameter of DRMEA ========
Manifold_dim = [1024,512]
Aligned_step = 0 # Apply Grass alignment at x-th epoch
PLGE_step = 10 # Apply target intra-class loss at x-th epoch
PLGE_Inter_step = 1 # Apply source inter-class loss at x-th epoch
PLGE_lambda_L2, PLGE_lambda_L1 = 1e1, 1e0 # \lambda_1
Grass_lambda = 5e3 # \lambda_2
Top_n = 1 # Top-k preserving scheme
# ===========================================
# ========= Downlaod Pretrained Model
url = 'https://download.pytorch.org/models/resnet50-19c8e357.pth'
pretrained_dict = model_zoo.load_url(url,model_dir='./pretrained_dict')
del pretrained_dict['fc.bias']
del pretrained_dict['fc.weight']
# ================== Training
ACC_Recorder = np.zeros((Exptimes,num_tasks)) # Result recorder
Total_Result = np.zeros((3,epochs,Exptimes))
for Domain_iter in range(num_tasks):
source_domain = source_domain_set[Domain_iter]
target_domain = target_domain_set[Domain_iter]
source_loader = Data.data_loader(dset_name, source_domain, batch_size)
target_loader = Data.data_loader(dset_name, target_domain, batch_size)
# Random Experiment
Exp_iter = 0
best_acc = 0
while Exp_iter < Exptimes:
# ============ Define Network Architecture
if config['Network'] == 'ResNet50':
e = network.ResNet50_Feature(network.Bottleneck, [3, 4, 6, 3])
fea_dim = 2048
c = network.C(fea_dim, Manifold_dim[0], Manifold_dim[1], n_classes)
e.cuda()
c.cuda()
e.load_state_dict(pretrained_dict)
c.apply(Utilizes.weights_init)
# ============= Define Optimizer
lr = 2e-4
beta1=0.9
beta2=0.999
Optimizer = optim.Adam([{'params':e.parameters(), 'lr': lr*0.1},
{'params':c.parameters()}],
lr*1.5, [beta1, beta2], weight_decay=0.01)
criterionQ_dis = nn.NLLLoss().cuda()
# ============= Training Epoches
result = np.zeros((3,epochs))
Exp_start_time = datetime.now()
print("************ %1s→%1s: %1s Start Experiment %1s training ************"%(source_domain,target_domain,Exp_start_time,Exp_iter+1))
for step in range(epochs):
epoch_time_start = datetime.now()
#=================== Initialize the mean vector ========
H_mean_update_L2 = torch.zeros(n_classes,512).cuda()
C_num_count = torch.zeros(n_classes,1).cuda()
H_mean_update_L1 = torch.zeros(n_classes,1024).cuda()
#=======================================================
Current_loss = np.array([0])
Current_Coral_Grass_loss_L1 = np.array([0])
Current_Coral_Grass_loss_L2 = np.array([0])
Current_PLGE_loss_L2 = np.array([0])
Current_PLGE_loss_L1 = np.array([0])
Current_PLGE_inter_loss_L2 = np.array([0])
Current_PLGE_inter_loss_L1 = np.array([0])
for (X, target), (X_t, target_test) in zip(source_loader,target_loader):
e.train()
c.train()
X, target = Variable(X), Variable(target)
X, target = X.cuda(), target.cuda()
X_t = Variable(X_t)
X_t = X_t.cuda()
# Init gradients
e.zero_grad()
c.zero_grad()
s_fe = e.forward(X)
s_fe_t = e.forward(X_t)
pred_s, h_s, h_s2 = c(s_fe)
pred_t, h_t, h_t2 = c(s_fe_t)
if step >= (PLGE_step - 1):
# =============== compute the class mean vector ==========
sam_count = 0
Tensor_size = target.shape
if Tensor_size:
target = target
else:
target = target.unsqueeze(0)
for i in target:
C_num_count[i] += 1
H_mean_update_L2[i,:] += h_s2[sam_count,:].data
H_mean_update_L1[i,:] += h_s[sam_count,:].data
sam_count += 1
# =========================================================
#==========================================================
# Loss Part
#==========================================================
CE_loss = criterionQ_dis(torch.log(pred_s+1e-4), target)
#===================== Align Loss =========================
if step <= (Aligned_step - 1):
Coral_Grass_loss_L1 = torch.zeros(1).squeeze(0).cuda()
Coral_Grass_loss_L2 = torch.zeros(1).squeeze(0).cuda()
else:
Coral_Grass_loss_L1, _, _ = Utilizes.grassmann_dist_Fast(h_s, h_t)
Coral_Grass_loss_L2, _, _ = Utilizes.grassmann_dist_Fast(h_s2, h_t2)
Align_loss = Grass_lambda*Coral_Grass_loss_L1 + Grass_lambda*Coral_Grass_loss_L2
#===================== Align Loss =========================
#================ Source Discriminative Loss ==============
if step <= (PLGE_Inter_step - 1):
PLGE_inter_loss_L2 = torch.zeros(1).squeeze(0).cuda()
PLGE_inter_loss_L1 = torch.zeros(1).squeeze(0).cuda()
else:
PLGE_inter_loss_L2 = PLGE_lambda_L2*Utilizes.Source_InterClass_sim_loss(h_s2, target, H_Tmean_use_L2, 'adj')
PLGE_inter_loss_L1 = PLGE_lambda_L1*Utilizes.Source_InterClass_sim_loss(h_s, target, H_Tmean_use_L1, 'adj')
Source_Discri_loss = PLGE_inter_loss_L2 + PLGE_inter_loss_L1
#================ Source Discriminative Loss ==============
#================ Target Discriminative Loss ==============
if step <= (PLGE_step - 1):
c_loss = CE_loss + Align_loss + Source_Discri_loss
else:
PLGE_loss_L2 = PLGE_lambda_L2*Utilizes.Target_IntraClass_sim_loss(h_t2, pred_t, H_mean_use_L2, Top_n)
PLGE_loss_L1 = PLGE_lambda_L1*Utilizes.Target_IntraClass_sim_loss(h_t, pred_t, H_mean_use_L1, Top_n)
Target_Discri_loss = PLGE_loss_L2 + PLGE_loss_L1
c_loss = CE_loss + Align_loss + Target_Discri_loss + Source_Discri_loss
#================ Target Discriminative Loss ==============
Current_loss = np.concatenate((Current_loss,c_loss.cpu().detach().numpy()[np.newaxis]),axis = 0)
Current_Coral_Grass_loss_L1 = np.concatenate((Current_Coral_Grass_loss_L1,Coral_Grass_loss_L1.cpu().detach().numpy()[np.newaxis]),axis = 0)
Current_Coral_Grass_loss_L2 = np.concatenate((Current_Coral_Grass_loss_L2,Coral_Grass_loss_L2.cpu().detach().numpy()[np.newaxis]),axis = 0)
Current_PLGE_inter_loss_L2 = np.concatenate((Current_PLGE_inter_loss_L2,PLGE_inter_loss_L2.cpu().detach().numpy()[np.newaxis]),axis = 0)
Current_PLGE_inter_loss_L1 = np.concatenate((Current_PLGE_inter_loss_L1,PLGE_inter_loss_L1.cpu().detach().numpy()[np.newaxis]),axis = 0)
if step > (PLGE_step - 1):
Current_PLGE_loss_L2 = np.concatenate((Current_PLGE_loss_L2,PLGE_loss_L2.cpu().detach().numpy()[np.newaxis]),axis = 0)
Current_PLGE_loss_L1 = np.concatenate((Current_PLGE_loss_L1,PLGE_loss_L1.cpu().detach().numpy()[np.newaxis]),axis = 0)
c_loss.backward()
Optimizer.step()
H_mean_use_L2 = H_mean_update_L2.mul(1/C_num_count) # Class mean matrix
H_Tmean_use_L2 = H_mean_update_L2.mean(0) # Total mean vector
H_mean_use_L1 = H_mean_update_L1.mul(1/C_num_count)
H_Tmean_use_L1 = H_mean_update_L1.mean(0)
del H_mean_update_L2, H_mean_update_L1
e.eval()
c.eval()
Test_start_time = datetime.now()
print('========================== %1s | Testing start! ==========================='%(Test_start_time))
source_acc = Utilizes.classification_accuracy(e,c,source_loader)
target_acc = Utilizes.classification_accuracy(e,c,target_loader)
Current_Coral_Grass_loss_L1 = np.sum(Current_Coral_Grass_loss_L1)/(Current_Coral_Grass_loss_L1.size - 1)
Current_Coral_Grass_loss_L2 = np.sum(Current_Coral_Grass_loss_L2)/(Current_Coral_Grass_loss_L2.size - 1)
Current_PLGE_inter_loss_L2 = np.sum(Current_PLGE_inter_loss_L2)/(Current_PLGE_inter_loss_L2.size - 1)
Current_PLGE_inter_loss_L1 = np.sum(Current_PLGE_inter_loss_L1)/(Current_PLGE_inter_loss_L1.size - 1)
if step > (PLGE_step - 1):
Current_PLGE_loss_L2 = np.sum(Current_PLGE_loss_L2)/(Current_PLGE_loss_L2.size - 1)
Current_PLGE_loss_L1 = np.sum(Current_PLGE_loss_L1)/(Current_PLGE_loss_L1.size - 1)
Current_loss = np.sum(Current_loss)/(Current_loss.size - 1)
result[:,step] = [target_acc,source_acc,Current_loss]
#====================== Time =====================
epoch_time_end = datetime.now()
seconds = (epoch_time_end - epoch_time_start).seconds
minutes = seconds//60
second = seconds%60
hours = minutes//60
minute = minutes%60
print('Source accuracy: {}'.format(source_acc))
print('Target accuracy: {}'.format(target_acc))
print('Max Target Accuracy: {}'.format(max(result[0,:])))
print('Total_Loss: {}'.format(Current_loss))
print('Current epoch time cost (including test): %1s Hour %1s'\
' Minutes %1s Seconds'%(hours,minute,second))
#========= Save the best model and write log
if target_acc > best_acc:
best_acc = target_acc
torch.save({
'Epoch': (step+1),
'state_dict_Backbone': e.state_dict(),
'state_dict_Manifold': c.state_dict(),
'Manifold_Dim': [1024,512],
'best_prec1': best_acc,
}, config["output_path"]+'/BestModel_'+save_name[Domain_iter]+'.tar')
log_str = 'Experiment: {:05d}, Epoch: {:05d}, test precision:'\
' {:.5f}'.format(Exp_iter+1, step+1, target_acc)
config["out_file"].write(save_name[Domain_iter]+'||'+log_str+'\n')
config["out_file"].flush()
#=========== If target accuracy reach 1, start new experiment
if max(result[0,:]) == 1:
print('Reach accuracy {1} at Epoch %1s !'%(step+1))
break
#============== End this Experiment ==============
seconds = (epoch_time_end - Exp_start_time).seconds
minutes = seconds//60
second = seconds%60
hours = minutes//60
minute = minutes%60
Total_Result[:,:,Exp_iter] = result
ACC_Recorder[Exp_iter,Domain_iter] = max(result[0,:])
print('Starting TIme: {}'.format(Exp_start_time))
print("Finishing TIme: {}".format(epoch_time_end))
print('Total TIme Cost: %1s Hour %1s Minutes %1s Seconds'%(hours,minute,second))
print("************ %1s→%1s: %1s End Experiment %1s training ************"%(source_domain,target_domain,epoch_time_end,Exp_iter+1))
Exp_iter += 1
#================= End this domain transfer task
if __name__ == "__main__":
parser = argparse.ArgumentParser(description='Discriminative Manifold Embedding and Alignment AAAI-2020')
parser.add_argument('--gpu_id', type=str, nargs='?', default='0', help="device id to run")
parser.add_argument('--net', type=str, default='ResNet50', choices=['ResNet50'])
parser.add_argument('--dset', type=str, default='ImageCLEF', choices=['ImageCLEF'])
parser.add_argument('--mEpo', type=str, nargs='?', default='50', help='Max epoches')
parser.add_argument('--ExpTime', type=str, nargs='?', default='10', help='Numbers of random experiments')
parser.add_argument('--BatchSize', type=str, nargs='?', default='32', help='Mini-Batch size')
args = parser.parse_args()
os.environ["CUDA_VISIBLE_DEVICES"] = args.gpu_id
config = {}
config['gpu'] = args.gpu_id
config["output_path"] = "Model_Log/" + args.dset
config['exp_time'] = args.ExpTime
config['bs'] = args.BatchSize
config['maxEpo'] = args.mEpo
config['Network'] = args.net
config['dset'] = args.dset
if not osp.exists(config["output_path"]):
os.system('mkdir -p '+config["output_path"])
config["out_file"] = open(osp.join(config["output_path"], "log.txt"), "w")
if not osp.exists(config["output_path"]):
os.mkdir(config["output_path"])
train(config)
|
document_class_array = ['article','ieeetran','proc','minimal','report','book','slides','memoir','letter','beamer']
color = ['apricot','aquamarine','bittersweet','black','blue','bluegreen','blueviolet','brickred','brown','burntorange','cadetblue','carnationpink','cerulean','cornflowerblue','cyan','dandelion','darkorchid','emerald','forestgreen','fuchsia','goldenrod','gray','green','greenyellow','junglegreen','lavender','limegreen','magenta','mahogany','maroon','melon','midnightblue','mulberry','navyblue','olivegreen','orange','orangered','orchid','peach','periwinkle','pinegreen','plum','processblue','purple','rawsienna','red','redorange','redviolet','rhodamine','royalblue','royalpurple','rubinered','salmon','seagreen','sepia','skyblue','springgreen','tan','tealblue','thistle','turquoise','violet','violetred','white','wildstrawberry','yellow','yellowgreen','yelloworange']
text_effect_array = ['emphasis','roman',' sans serif','teletype','italic','small capitals','uppercase','bold']
text_effect_code = ['\\emph','\\textrm','\\textsf','\\texttt','\\textit','\\textsc','\\uppercase','\\textbf']
text_size_array = ['tiny','small','normalsize','large','huge']
text_size_code = ['\\tiny','\small','\\normalsize','\large','\huge']
code_extension_array = ['abap','ada','alg','ass','awk','bash','bas','c','cpp','cob','csh','dil','e','ex','for','gplt','hk','html','java','l','obn','p','perl','php','pov','pro','py','rexx','ruby','sim','sql','tcl','tex','vbs','vrml','xml','xslt']
code_language_array = ['abap','ada','algol','assembler','awk','bash','basic','c','c++','cobol','csh','delphi','eiffel','euphoria','fortran','gnuplot','haskell','html','java','logo','oberon-2','pascal','perl','php','pov','prolog','python','rexx','ruby','simula','sql','tcl','tex','vbsscript','vrml','xml','xslt']
letter_component_array = ['name','signature','address','location','opening','closing','cc','encl']
letter_component_code = ['\\name','\signature','\\address','\location','\opening','\closing','\cc','\encl']
effects_1 = [
['superscript','\\textsuperscript{','fixltx2e'] ,
['subscript','\\textsubscript{','fixltx2e'] ,
['footnote','\\footnote{',''] ,
['marginnote','\marginnote{','marginnote'] ,
['mbox','\mbox{',''] ,
['fbox','\\fbox{',''] ,
['vbox','\\vbox{',''] ,
]
effects_2 = [
['abstract',''] ,
['equation',''] ,
['flushleft',''] ,
['flushright',''] ,
['center',''] ,
['frame',''] ,
['verbatim',''] ,
['sideways','rotating'] ,
['sidewaysfigure','rotating'] ,
['sidewaystable','rotating'] ,
]
maths = [
['arcsin','$\\arcsin{'] ,
['arccos','$\\arccos{'] ,
['arctan','$\\arctan{'] ,
['arg','$\\arg{'] ,
['cos','$\cos{'] ,
['cosh','$\cosh{'] ,
['cot','$\cot{'] ,
['coth','$\coth{'] ,
['csc','$\csc{'] ,
['deg','$\deg{'] ,
['det','$\det{'] ,
['dim','$\dim{'] ,
['exp','$\exp{'] ,
['gcd','$\gcd{'] ,
['hom','$\hom{'] ,
['inf','$\inf{'] ,
['ker','$\ker{'] ,
['lg','$\lg{'] ,
['lim','$\lim{'] ,
['liminf','$\liminf{'] ,
['limsup','$\limsup{'] ,
['ln','$\ln{'] ,
['log','$\log{'] ,
['max','$\max{'] ,
['min','$\min{'] ,
['Pr','$\Pr{'] ,
['sec','$\sec{'] ,
['sin','$\sin{'] ,
['sinh','$\sinh{'] ,
['sup','$\sup{'] ,
['tan','$\\tan{'] ,
['tanh','$\\tanh{'] ,
['sqrt','$\sqrt{'] ,
]
symbols = ['\\alpha']
frame_component_array = ['frametitle','framesubtitle']
format_array = ['usetheme','usecolortheme','useoutertheme','useinnertheme']
format_options = [
['Antibes','Bergen','Berkeley','Berlin','Copenhagen','Darmstadt','Dresden','Frankfurt','Goettingen','Hannover','Ilmenau','JuanLesPins','Luebeck','Madrid','Malmoe','Marburg','Montpellier','PaloAlto','Pittsburgh','Rochester','Singapore','Szeged','Warsaw','boxes','default'],['default','albatross','beaver','beetle','crane','dolphin','dove','fly','lily','orchid','rose','seagull','seahorse','whale','wolverine'],['infolines','miniframes','shadow','sidebar','smoothbars','smoothtree','split','tree'],['rectangles','circles','inmargin','rounded']
]
|
<reponame>Lucas-Nieto/Laboratorio_Fisica_Moderna
# -*- coding: utf-8 -*-
"""
Created on Tue Feb 15 21:56:02 2022
Experimento 1: Espectrometría
Objetivo: Determinar la constante de Rydberg con datos de las líneas espectrales de la serie
de Balmer exportados por Astrosurf IRIS
Updated on Sat Feb 26 07:23:59 2022:
Correciones:
- Corregido reporte de barras de error
- Corregido cálculo de incertidumbre de pendiente usando stats.linregress
@author: <NAME>
"""
# %% Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
# Al abrir en un notebook de Jupyter, sacar la línea siguiente del comentario
#%matplotlib inline
# %% Plots pixel vs grayscale value para hidrógeno y mercurio
"""Hidrógeno"""
HFile = pd.read_csv("Corrected_Hydrogen.csv",sep = ";")
DataH = HFile.to_numpy().transpose().tolist()
x_valuesH = np.array(DataH[0])
y_valuesH = np.array(DataH[1])
plt.scatter(x_valuesH, y_valuesH, c="crimson", s = 10)
plt.title("Espectro experimental del hidrógeno")
plt.xlabel("Pixels")
plt.ylabel("Grayscale intensity")
plt.grid()
plt.show()
"""Mercurio"""
HgFile = pd.read_csv("Corrected_Mercury.csv",sep = ";")
DataHg = HgFile.to_numpy().transpose().tolist()
x_valuesHg = np.array(DataHg[0])
y_valuesHg = np.array(DataHg[1])
plt.scatter(x_valuesHg, y_valuesHg, c="springgreen", s = 10)
plt.title("Espectro experimental del mercurio")
plt.xlabel("Pixels")
plt.ylabel("Grayscale intensity")
plt.grid()
plt.show()
# %% Conversión pixeles a nanómetros
def pix_to_nm(pix)->int:
Lambda_H_Alpha = 656.2797 # Valor teórico de longitud de onda de la línea H_alpha
wavelenght = (Lambda_H_Alpha*pix)/562 # Regla de 3 simple con el pixel en el que se encontró
# H_alpha en IRIS astronomy
return wavelenght
C = True
while C == True:
print()
print("Presione 1 para hacer conversión de pixeles a nanómetros.")
Prompt = input("Presione cualquier otra tecla para continuar: ")
if Prompt == "1":
conv = int(input("Digite la cantidad de pixeles que desea convertir a nanómetros: "))
print(conv,"pixeles es equivalente a:",pix_to_nm(conv),"nanómetros")
else:
C = False
print("\n---------------------------------------------------------------------------")
# %% Cálculo de la constante
Experimental_wavelenghts = [656.2797,476.4062,417.8894,393.5343] # Manualmente encontradas con el
# slice de IRIS Astronomy
ToMeters = Experimental_wavelenghts.copy()
i = 0
while i in range(len(ToMeters)):
ToMeters[i] = (ToMeters[i])*10e-10 # Conversión de nanómetros a metros (el factor de conversión
# es 10e-9 pero por error de truncamiento del intérprete fue
# necesario sumar 1 para tener orden de magnitud correcto)
i+=1
y = ToMeters.copy()
i = 0
while i in range(len(y)):
y[i] = 1/(y[i])
i+=1
x = [0.1388,0.1875,0.2100,0.2222] # De los niveles de energía de la fórmula de Balmer
x = np.array(x)
y = np.array(y)
regr = stats.linregress(x, y)
slope = regr[0]
intercept = regr[1]
def fit(x):
return slope*x + intercept
# Barras de error
S_y = np.std(y)
S_x = np.std(x)
ybar = (S_y/np.sqrt(len(y))) # Error típico o estándar de un estadístico
xbar = (S_x/np.sqrt(len(x)))
# Plot de regresión
rydberg = list(x)
i = 0
while i in range(len(rydberg)):
rydberg[i] = rydberg[i]*1.0973e+07 # Valor reportado en la literaturya de la constante de Rydberg
i += 1
plt.scatter(x,y, c = "midnightblue")
plt.errorbar(x, y, fmt = " ", yerr = ybar,ecolor = "k")
plt.plot(x,fit(x), c = "green", label = "R Experimental")
plt.plot(x,rydberg, linestyle='dashdot',c = "slateblue",label = "R Reportado por Beyer et al.")
plt.legend()
plt.title(r"Regresión lineal de $ \frac{1}{\lambda} = \left ( \frac{1}{n^{2}}- \frac{1}{m^{2}}\right )$")
plt.grid()
plt.show()
# %% Reporte de parámetros
# Pendiente de regresión
print("\nConstante experimental de Rydberg para el hidrógeno =","{:e}".format(slope))
# Incertidumbre de la pendiente
Sigma = regr[4]
print("\nIncertidumbre de la pendiente =","{:e}".format(Sigma))
# Coeficiente de correlación lineal
print(u"\nCoeficiente de correlación lineal R\u00b2 =",regr[2])
# Tamaño de barras de error
print("\nTamaño de barras de error:")
print("\n - Distancia de dato a límite =","{:e}".format(ybar))
print("\n - Distancia de límite a límite =","{:e}".format(2*ybar))
|
<filename>Train_cifar.py
from __future__ import print_function
import sys
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
import random
import os
import argparse
import numpy as np
from PreResNet import *
from sklearn.mixture import GaussianMixture
import dataloader_cifar as dataloader
from torch.utils.tensorboard import SummaryWriter
import pdb
import io
from PIL import Image
import matplotlib.pyplot as plt
import torchvision
from edl_losses import *
import warnings
from pathlib import Path
from sklearn.preprocessing import normalize
warnings.filterwarnings("ignore")
parser = argparse.ArgumentParser(description='PyTorch CIFAR Training')
parser.add_argument('--batch_size', default=64, type=int, help='train batchsize')
parser.add_argument('--slr', '--s_learning_rate', default=0.02, type=float, help='initial learning rate for netS')
parser.add_argument('--dlr', '--d_learning_rate', default=0.02, type=float, help='initial learning rate for netD')
parser.add_argument('--noise_mode', default='sym')
parser.add_argument('--alpha', default=4, type=float, help='parameter for Beta')
parser.add_argument('--lambda_u', default=25, type=float, help='weight for unsupervised loss')
parser.add_argument('--p_threshold', default=1/3, type=float, help='clean probability threshold')
parser.add_argument('--T', default=0.5, type=float, help='sharpening temperature')
parser.add_argument('--num_epochs', default=200, type=int)
parser.add_argument('--r', default=0.5, type=float, help='noise ratio')
parser.add_argument('--on', default=0, type=float, help='open noise ratio')
parser.add_argument('--id', default='')
parser.add_argument('--seed', default=123)
parser.add_argument('--gpuid', default=0, type=int)
parser.add_argument('--num_class', default=10, type=int)
parser.add_argument('--data_path', default='../../data/cifar10/cifar-10-batches-py', type=str, help='path to dataset')
parser.add_argument('--dataset', default='cifar10', type=str)
parser.add_argument('--noisy_dataset', default='cifar100', type=str)
parser.add_argument('--noise_dir', default='./noise', type=str)
parser.add_argument('--noise_data_dir', default='../../data/cifar100/cifar-100-python', type=str)
parser.add_argument('--inference', action='store_true')
parser.add_argument('--skip_warmup', action='store_true')
parser.add_argument('--start_epoch', default=0, type=int)
parser.add_argument('--load_state_dict', default=None, type=str)
parser.add_argument('--warmup_epochs_S', default=30, type=int)
parser.add_argument('--warmup_epochs_D', default=10, type=int)
parser.add_argument('--plots_dir', default='plots/', type=str)
parser.add_argument('--gmmc', default=3, type=int)
parser.add_argument('--cudaid', default=0)
args = parser.parse_args()
torch.cuda.set_device(args.gpuid)
random.seed(args.seed)
torch.manual_seed(args.seed)
torch.cuda.manual_seed_all(args.seed)
device = device = torch.device('cuda:{}'.format(args.cudaid))
writer = SummaryWriter('runs/r={}_on={}'.format(args.r, args.on))
args.uid = '{}_{}'.format(args.dataset, args.noisy_dataset)
args.plots_dir = os.path.join(args.plots_dir, args.uid)
args.plots_dir = os.path.join(args.plots_dir, 'r={}_on={}'.format(args.r, args.on))
args.save_dicts_dir = os.path.join('saveDicts', args.uid)
args.checkpoint_dir = os.path.join('./checkpoint',args.uid)
Path(os.path.join(args.plots_dir, 'netS')).mkdir(parents=True, exist_ok=True)
Path(os.path.join(args.plots_dir, 'netD')).mkdir(parents=True, exist_ok=True)
Path(os.path.join(args.save_dicts_dir,)).mkdir(parents=True, exist_ok=True)
Path(os.path.join(args.checkpoint_dir,)).mkdir(parents=True, exist_ok=True)
# Training
def train_D(epoch,netD,optimizer,labeled_trainloader,unlabeled_trainloader):
netD.train()
global iter_net
iter_idx = 1
unlabeled_train_iter = iter(unlabeled_trainloader)
num_iter = (len(labeled_trainloader.dataset)//args.batch_size)+1
for batch_idx, (inputs_x, inputs_x2, labels_x, w_x) in enumerate(labeled_trainloader):
try:
inputs_u, inputs_u2 = unlabeled_train_iter.next()
except:
unlabeled_train_iter = iter(unlabeled_trainloader)
inputs_u, inputs_u2 = unlabeled_train_iter.next()
batch_size = inputs_x.size(0)
# Transform label to one-hot
labels_x = torch.zeros(batch_size, args.num_class).scatter_(1, labels_x.view(-1,1), 1)
w_x = w_x.view(-1,1).type(torch.FloatTensor)
inputs_x, inputs_x2, labels_x, w_x = inputs_x.to(device), inputs_x2.to(device), labels_x.to(device), w_x.to(device)
inputs_u, inputs_u2 = inputs_u.to(device), inputs_u2.to(device)
with torch.no_grad():
# label guessing of unlabeled samples
outputs_u1 = netD(inputs_u)
outputs_u2 = netD(inputs_u2)
pu = (torch.softmax(outputs_u1, dim=1) + torch.softmax(outputs_u2, dim=1)) / 2
ptu = pu**(1/args.T) # temparature sharpening
targets_u = ptu / ptu.sum(dim=1, keepdim=True) # normalize
targets_u = targets_u.detach()
# label refinement of labeled samples
outputs_x = netD(inputs_x)
outputs_x2 = netD(inputs_x2)
px = (torch.softmax(outputs_x, dim=1) + torch.softmax(outputs_x2, dim=1)) / 2
px = w_x*labels_x + (1-w_x)*px
ptx = px**(1/args.T) # temparature sharpening
targets_x = ptx / ptx.sum(dim=1, keepdim=True) # normalize
targets_x = targets_x.detach()
# mixmatch
l = np.random.beta(args.alpha, args.alpha)
l = max(l, 1-l)
all_inputs = torch.cat([inputs_x, inputs_x2, inputs_u, inputs_u2], dim=0)
all_targets = torch.cat([targets_x, targets_x, targets_u, targets_u], dim=0)
idx = torch.randperm(all_inputs.size(0))
input_a, input_b = all_inputs, all_inputs[idx]
target_a, target_b = all_targets, all_targets[idx]
mixed_input = l * input_a + (1 - l) * input_b
mixed_target = l * target_a + (1 - l) * target_b
logits = netD(mixed_input)
logits_x = logits[:batch_size*2]
logits_u = logits[batch_size*2:]
Lx, Lu, lamb = criterion(logits_x, mixed_target[:batch_size*2], logits_u, mixed_target[batch_size*2:], epoch+batch_idx/num_iter)
# regularization
prior = torch.ones(args.num_class)/args.num_class
prior = prior.to(device)
pred_mean = torch.softmax(logits, dim=1).mean(0)
penalty = torch.sum(prior*torch.log(prior/pred_mean))
loss = Lx + lamb * Lu + penalty
# compute gradient and do SGD step
optimizer.zero_grad()
loss.backward()
optimizer.step()
iter_net[iter_idx] += 1
sys.stdout.write('\r')
sys.stdout.write('%s:%.1f-%s | Epoch [%3d/%3d] Iter[%3d/%3d]\t Labeled loss: %.2f Unlabeled loss: %.2f'
%(args.dataset, args.r, args.noise_mode, epoch, args.num_epochs, batch_idx+1, num_iter, Lx.item(), Lu.item()))
writer.add_scalar('Train/Loss/{}/Labelled'.format('netD'), Lx.item(), iter_net[iter_idx])
writer.add_scalar('Train/Loss/{}/Unlabelled'.format('netD'), Lu.item(), iter_net[iter_idx])
sys.stdout.flush()
def train_S(epoch,netS,optimizer,dataloader):
netS.train()
global iter_net
iter_idx = 0
num_iter = (len(dataloader.dataset)//dataloader.batch_size)+1
for batch_idx, (inputs, labels, path) in enumerate(dataloader):
inputs, labels = inputs.to(device), labels.to(device)
optimizer.zero_grad()
outputs = netS(inputs)
y = one_hot_embedding(labels.type(torch.LongTensor))
y = y.to(device)
loss = subjective_loss(outputs, y.float()).mean()
loss.backward()
optimizer.step()
iter_net[iter_idx] += 1
sys.stdout.write('\r')
sys.stdout.write('%s:%.1f-%s | Epoch [%3d/%3d] Iter[%3d/%3d]\t loss: %.4f'
%(args.dataset, args.r, args.noise_mode, epoch, args.num_epochs, batch_idx+1, num_iter, loss.item()))
writer.add_scalar('Train/Loss/All{}'.format('netS'), loss.item(), iter_net[iter_idx])
sys.stdout.flush()
def warmup(epoch,net,optimizer,dataloader,model_name):
net.train()
num_iter = (len(dataloader.dataset)//dataloader.batch_size)+1
for batch_idx, (inputs, labels, path) in enumerate(dataloader):
inputs, labels = inputs.to(device), labels.to(device)
optimizer.zero_grad()
outputs = net(inputs)
if model_name == 'netS':
warm_up_epochs = args.warmup_epochs_S
y = one_hot_embedding(labels)
y = y.to(device)
loss = subjective_loss(outputs, y.float()).mean()
else:
warm_up_epochs = args.warmup_epochs_D
loss = CEloss(outputs, labels)
if args.noise_mode=='asym': # penalize confident prediction for asymmetric noise
penalty = conf_penalty(outputs)
L = loss + penalty
elif args.noise_mode=='sym':
L = loss
L.backward()
optimizer.step()
sys.stdout.write('\r')
sys.stdout.write('%s:%.1f-%s | Epoch [%3d/%3d] Iter[%3d/%3d]\t loss: %.4f'
%(args.dataset, args.r, args.noise_mode, epoch, warm_up_epochs, batch_idx+1, num_iter, loss.item()))
writer.add_scalar('Warmup/Loss/{}'.format(model_name), loss.item(), epoch * num_iter + batch_idx)
sys.stdout.flush()
losses = get_losses_on_all(net, model_name)
def test(epoch,net,test_loader,model_name):
net.eval()
correct = 0
total = 0
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(test_loader):
inputs, targets = inputs.to(device), targets.to(device)
outputs = net(inputs)
_, predicted = torch.max(outputs, 1)
total += targets.size(0)
correct += predicted.eq(targets).cpu().sum().item()
acc = 100.*correct/total
print("\n| Test Epoch #%d\t Accuracy: %.2f%%\n" %(epoch,acc))
test_log.write('Epoch:%d Accuracy:%.2f\n'%(epoch,acc))
test_log.flush()
writer.add_scalar('Test/Accuracy/{}'.format(model_name), acc, epoch)
def get_losses_on_all(model, model_name):
eval_loader = loader.run('eval_train')
model.eval()
losses = torch.zeros(50000)
with torch.no_grad():
for batch_idx, (inputs, targets, index) in enumerate(eval_loader):
inputs, targets = inputs.to(device), targets.to(device)
outputs = model(inputs)
if model_name == 'netS':
y = one_hot_embedding(targets)
y = y.to(device)
loss = subjective_loss(outputs, y.float())
else:
loss = CE(outputs, targets)
for b in range(inputs.size(0)):
losses[index[b]]=loss[b]
losses = (losses-losses.min())/(losses.max()-losses.min()) # normalised losses for each image
return losses
def refine_labels(model, probs, zero_open=False):
probClean, probOpen, probClosed = probs[0], probs[1], probs[2]
predClean = (probClean > probOpen) & (probClean > probClosed)
predClosed = (probClosed > probClean) & (probClosed > probOpen)
predOpen = (probOpen > probClean) & (probOpen > probClosed)
model.eval()
targets = torch.zeros(50000)
w_x = probClosed
w_x = torch.from_numpy(np.expand_dims(w_x, axis=1)).to(device)
eval_loader = loader.run('eval_train')
with torch.no_grad():
for batch_idx, (inputs, labels, index) in enumerate(eval_loader):
inputs, labels = inputs.to(device), one_hot_embedding(labels).to(device)
outputs = model(inputs)
px = torch.softmax(outputs, dim=1).to(device)
px = (1-w_x[index])*labels + (w_x[index])*px
ptx = px**(1/args.T) # temparature sharpening
refined = ptx / ptx.sum(dim=1, keepdim=True) # normalize
refined = refined.detach()
for b in range(inputs.size(0)):
if zero_open:
if predOpen[index[b]]:
targets[index[b]] = -100
else:
targets[index[b]] = refined[b].argmax()
else:
targets[index[b]] = refined[b].argmax()
return targets
def fit_gmm_multiple_components(input_loss):
gmm = GaussianMixture(n_components=20,max_iter=10,tol=1e-2,reg_covar=5e-4)
gmm.fit(input_loss)
components_open = []
components_clean = []
components_closed = []
for n in range(gmm.n_components):
if (gmm.means_[n] > .3) & (gmm.means_[n] < .7):
components_open.extend([n])
elif (gmm.means_[n] < .3):
components_clean.extend([n])
else:
components_closed.extend([n])
prob = gmm.predict_proba(input_loss)
# transform this probability into a 3-component probability
prob_clean = np.sum(prob[:,components_clean],axis=1)
prob_closed = np.sum(prob[:,components_closed],axis=1)
prob_open = np.sum(prob[:,components_open], axis=1)
return prob_clean, prob_open, prob_closed
def eval_train(model,all_loss,epoch,model_name):
losses = get_losses_on_all(model, model_name)
all_loss.append(losses)
if args.r==0.9: # average loss over last 5 epochs to improve convergence stability
history = torch.stack(all_loss)
input_loss = history[-5:].mean(0)
input_loss = input_loss.reshape(-1,1)
else:
input_loss = losses.reshape(-1,1)
strongClean = input_loss.argmin()
strongClosed = input_loss.argmax()
probClean, probOpen, probClosed = fit_gmm_multiple_components(input_loss)
predClean = (probClean > probOpen) & (probClean > probClosed)
predClosed = (probClosed > probClean) & (probClosed > probOpen)
predOpen = (probOpen > probClean) & (probOpen > probClosed)
# guarantee that there is at least one sample in clean and closed
predClean[strongClean] = True
predClosed[strongClosed] = True
return [probClean, probOpen, probClosed], [predClean, predOpen, predClosed], all_loss
def linear_rampup(current, rampup_length=16):
current = np.clip((current) / rampup_length, 0.0, 1.0)
return args.lambda_u*float(current)
def printDataSplit(predClean, predOpen, predClosed):
cleanIdx = np.where(predClean)[0]
closedIdx = np.where(predClosed)[0]
openIdx = np.where(predOpen)[0]
clean, open_noise, closed_noise = loader.get_noise()
stats_log.write('Dividing dataset...\n')
if len(clean) != 0:
stats_log.write('Clean - clean:{:.2f}, closed:{:.2f}, open;{:.2f}\n'.format(len(set(cleanIdx).intersection(clean))/len(clean), len(set(closedIdx).intersection(clean))/len(clean), len(set(openIdx).intersection(clean))/len(clean)))
if len(closed_noise) != 0:
stats_log.write('Closed - clean:{:.2f}, closed:{:.2f}, open;{:.2f}\n'.format(len(set(cleanIdx).intersection(closed_noise))/len(closed_noise), len(set(closedIdx).intersection(closed_noise))/len(closed_noise), len(set(openIdx).intersection(closed_noise))/len(closed_noise)))
if len(open_noise) != 0:
stats_log.write('Open - clean:{:.2f}, closed:{:.2f}, open;{:.2f}\n\n'.format(len(set(cleanIdx).intersection(open_noise))/len(open_noise), len(set(closedIdx).intersection(open_noise))/len(open_noise), len(set(openIdx).intersection(open_noise))/len(open_noise)))
stats_log.flush()
class SemiLoss(object):
def __call__(self, outputs_x, targets_x, outputs_u, targets_u, epoch):
probs_u = torch.softmax(outputs_u, dim=1)
Lx = -torch.mean(torch.sum(F.log_softmax(outputs_x, dim=1) * targets_x, dim=1))
Lu = torch.mean((probs_u - targets_u)**2)
return Lx, Lu, linear_rampup(epoch)
class NegEntropy(object):
def __call__(self,outputs):
probs = torch.softmax(outputs, dim=1)
return torch.mean(torch.sum(probs.log()*probs, dim=1))
def create_model():
model = ResNet18(num_classes=args.num_class)
model = model.to(device)
return model
def plotHistogram(data, predictions=None, log=False, model_name='', phase='', title=''):
clean, open_noise, closed_noise = loader.get_noise()
fig = plt.figure()
if predictions is not None:
plt.subplot(121)
plt.hist(data[clean], bins=300, alpha=0.5, color='green')
if len(closed_noise) > 0:
plt.hist(data[closed_noise], bins=300, alpha=0.5, color='blue')
if len(open_noise) > 0:
plt.hist(data[open_noise], bins=300, alpha=0.5, color='red')
# plt.legend(loc='upper right')
if predictions is not None:
plt.subplot(122)
plt.hist(data[predictions[0]], bins=300, alpha=0.5, color='green', label='Predicted clean set')
plt.hist(data[predictions[2]], bins=300, alpha=0.5, color='blue', label='Predicted closed set')
plt.hist(data[predictions[1]], bins=300, alpha=0.5, color='red', label='Predicted open set')
plt.legend(loc='upper right')
if log:
print('\nlogging histogram...')
plt.savefig(os.path.join(args.plots_dir, '{}/{}_{}'.format(model_name, phase, title)), format='png')
else:
plt.show()
plt.close()
def get_logits(model):
eval_loader = loader.run('eval_train')
model.eval()
logits = np.zeros((50000, 10))
with torch.no_grad():
for batch_idx, (inputs, targets, index) in enumerate(eval_loader):
inputs, targets = inputs.to(device), targets.to(device)
outputs = model(inputs)
logits[index] = outputs.cpu()
return logits
def runExperiment():
print('| Building net')
netS = create_model()
netD = create_model()
if args.load_state_dict is not None:
print('Loading saved state dict from {}'.format(args.load_state_dict))
checkpoint = torch.load(args.load_state_dict)
netS.load_state_dict(checkpoint['netS_state_dict'])
netD.load_state_dict(checkpoint['netD_state_dict'])
cudnn.benchmark = True
optimizer1 = optim.SGD(netS.parameters(), lr=args.slr, momentum=0.9, weight_decay=5e-4)
optimizer2 = optim.SGD(netD.parameters(), lr=args.dlr, momentum=0.9, weight_decay=5e-4)
if args.noise_mode=='asym':
conf_penalty = NegEntropy()
all_loss = [[],[]] # save the history of losses from two networks
test_loader = loader.run('test')
if not args.skip_warmup:
warmup_trainloader = loader.run('warmup')
print('Warmup netS')
for epoch in range(args.warmup_epochs_S):
warmup(epoch,netS,optimizer1,warmup_trainloader,'netS')
if epoch % 3 == 0:
plotHistogram(get_losses_on_all(netS, 'netS'), log=True, model_name='netS', phase='warmup', title='epoch={}'.format(epoch))
test(epoch,netS,test_loader,'netS')
print('\nWarmup netD')
for epoch in range(args.warmup_epochs_D):
warmup(epoch,netD,optimizer2,warmup_trainloader,'netD')
if epoch % 3 == 0:
plotHistogram(get_losses_on_all(netD, 'netD'), log=True, model_name='netD', phase='warmup', title='epoch={}'.format(epoch))
test(epoch,netD,test_loader,'netD')
print('\nSaving warmup state dict...')
torch.save({
'netS_state_dict': netS.state_dict(),
'netD_state_dict': netD.state_dict(),
}, os.path.join(args.save_dicts_dir, 'warmup_%.1f_%0.2f.json'%(args.r,args.on)))
else:
test(args.start_epoch-1,netS,test_loader,'netS')
test(args.start_epoch-1,netD,test_loader,'netD')
for epoch in range(args.start_epoch, args.num_epochs+1):
slr=args.slr
dlr=args.dlr
if epoch > 100:
dlr /= 10
for param_group in optimizer1.param_groups:
param_group['lr'] = slr
for param_group in optimizer2.param_groups:
param_group['lr'] = dlr
clean_loader = loader.run('clean')
eval_loader = loader.run('eval_train')
#train_clean_accuracy(epoch, netS, netD, clean_loader)
probs, preds, all_loss[0] = eval_train(netS,all_loss[0], epoch, 'netS')
probClean, probOpen, probClosed = probs[0], probs[1], probs[2]
predClean, predOpen, predClosed = preds[0], preds[1], preds[2]
printDataSplit(predClean, predOpen, predClosed)
if epoch % 3 == 0:
plotHistogram(get_losses_on_all(netS, 'netS'), [predClean, predOpen, predClosed], log=True, model_name='netS', phase='train', title='epoch={}'.format(epoch))
plotHistogram(get_losses_on_all(netD, 'netD'), [predClean, predOpen, predClosed], log=True, model_name='netD', phase='train', title='epoch={}'.format(epoch))
print('\nTrain netD')
labeled_trainloader, unlabeled_trainloader = loader.run('trainD', predClean, predClosed, probClean) # divide
train_D(epoch,netD,optimizer2,labeled_trainloader, unlabeled_trainloader) # train netD
#refine labels using netD
targets = refine_labels(netD, [probClean, probOpen, probClosed])
print('\nTrain netS')
trainloader = loader.run('trainS', targets=targets)
train_S(epoch,netS,optimizer1,trainloader)
test(epoch,netS,test_loader,'netS')
test(epoch,netD,test_loader, 'netD')
if epoch % 10 == 0:
torch.save({
'netS_state_dict': netS.state_dict(),
'netD_state_dict': netD.state_dict(),
}, os.path.join(args.save_dicts_dir, 'train_%.1f_%0.2f_%d.json'%(args.r,args.on,epoch)))
def train_clean_accuracy(epoch,netS,netD,clean_loader):
netS.eval()
netD.eval()
correct = 0
total = 0
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(clean_loader):
inputs, targets = inputs.to(device), targets.to(device)
outputs1 = netS(inputs)
outputs2 = netD(inputs)
outputs = outputs1+outputs2
_, predicted = torch.max(outputs, 1)
total += targets.size(0)
correct += predicted.eq(targets).cpu().sum().item()
acc = 100.*correct/total
print("\n| Clean Set Epoch #%d\t Accuracy: %.2f%%\n" %(epoch,acc))
test_log.write('Epoch:%d Accuracy:%.2f\n'%(epoch,acc))
test_log.flush()
writer.add_scalar('Clean/Accuracy', acc, epoch)
def infer():
print('Loading saved state dict from {}'.format(args.load_state_dict))
checkpoint = torch.load(args.load_state_dict)
netS = create_model()
netD = create_model()
netS.load_state_dict(checkpoint['netS_state_dict'])
netD.load_state_dict(checkpoint['netD_state_dict'])
losses = get_losses_on_all(netS, 'netS')
plotHistogram(losses)
stats_log=open('./%s/%.1f_%.2f'%(args.checkpoint_dir,args.r,args.on)+'_stats.txt','w')
test_log=open('./%s/%.1f_%.2f'%(args.checkpoint_dir,args.r,args.on)+'_acc.txt','w')
loader = dataloader.cifar_dataloader(args.dataset, args.noisy_dataset, r=args.r, on=args.on, noise_mode=args.noise_mode,batch_size=args.batch_size,num_workers=5,\
root_dir=args.data_path, noise_data_dir=args.noise_data_dir, log=stats_log,noise_file='%s/%.1f_%0.2f_%s.json'%(args.noise_dir,args.r,args.on,args.noisy_dataset))
CE = nn.CrossEntropyLoss(reduction='none')
CEloss = nn.CrossEntropyLoss()
subjective_loss = edl_mse_loss
criterion = SemiLoss()
iter_net = [0, 0]
if args.inference is False:
runExperiment()
else:
infer() |
<gh_stars>1-10
#!/usr/bin/python3
"""
This module test the maximal number of TXs in one block.
"""
import math
import logging
import sys
from collections import deque
from bitcoinrpc.authproxy import AuthServiceProxy
import socket
from eval import get_txnum_eval_path
from protocol import State
from generate_keys import get_keys
from helpers import generate_peers
PORT = 18445
filepath = get_txnum_eval_path() + '/' + 'max_tx_num.txt'
def write(text: str) -> None:
"""Both log to console and write into output file"""
logger.info(text)
with open(filepath, 'a') as fd:
fd.write(text)
fd.write('\n')
def connect() -> AuthServiceProxy:
"""Establish a fresh connection to the deamon"""
return AuthServiceProxy(
"http://%s:%s@127.0.0.1:%s" % (
State.RPC_USER, State.RPC_PASSWORD, PORT))
if __name__ == '__main__':
# Logging
LOG_LEVEL = logging.DEBUG
logger = logging.getLogger("Blocksize Test")
logger.setLevel(LOG_LEVEL)
handler = logging.StreamHandler()
handler.setLevel(LOG_LEVEL)
logger.addHandler(handler)
try:
keys = get_keys(5000)
except FileNotFoundError:
logger.error(
"There is no key-file bin/keys_10000.pyc. Call "
"'anonboot/generate_keys.py "
"10000' to create the necessary file.")
sys.exit()
# Generate Peers
peers = generate_peers(keys, 5000)
# Generate addresses
NUM_ADDRESSES = int(len(peers) / 20)
logger.info(
"Generate {} addresses!".format(math.ceil(NUM_ADDRESSES / 50) * 50))
# Bitcoin allows each address to be used at most 25 times per block.
# We have to do this in several rounds because preserver has problems with
# socket timeouts.
a_counter = {}
inputs = deque()
for _ in range(0, NUM_ADDRESSES, 50):
complete = False
while not complete:
try:
addresses = connect().batch_(
['getnewaddress'] for i in range(50))
blocks = [b[0] for b in connect().batch_(
[['generatetoaddress', 1, addr] for addr in addresses])]
blocks = [block['tx'][0] for block in
connect().batch_(['getblock', b] for b in blocks)]
raws = connect().batch_(
['getrawtransaction', txid, 1] for txid in blocks)
for r in raws:
for v in r['vout']:
if 'addresses' in v['scriptPubKey']:
vout = v['n']
addr = v['scriptPubKey']['addresses'][0]
val = v['value']
inputs.append(
{'in': {'txid': r['txid'], 'vout': vout},
'addr': addr, 'amount': val})
a_counter[addr] = 0
break
complete = True
except socket.timeout as e:
logger.warning("Socket timeout occured.")
# Generate 101 blocks to make addresses usable
connect().generate(101)
logger.info("Address Generation Done!")
errors = 0
last_success = 0
for i in range(5100, 5300, 1):
if errors >= 2:
# 2 consecutive fails occurred
write("**********************************")
write(
"The largest block that could be mined contained {} "
"TXs.".format(
i - 2))
write("**********************************")
break
try:
ads = []
tmp_state = State(1, 1, 0)
for p in peers[:i]:
ads.append(p.advertise(tmp_state))
print("Attempt to write {} advertisements".format(len(ads)))
con = connect()
for ad in ads:
input = inputs.popleft()
raw = con.createrawtransaction([input['in']], [{'data': ad}])
funded = con.fundrawtransaction(raw, {
"changeAddress": input['addr']})
signed = con.signrawtransactionwithwallet(funded['hex'])
sent = con.sendrawtransaction(signed['hex'])
raw = con.getrawtransaction(sent, 1)
vout = -1
new_value = input['amount'] - 1
for v in raw['vout']:
if v['value'] >= new_value:
vout = v['n']
new_value = v['value']
break
if vout == -1:
logging.warning(
"WARNING: No suitable output found, will lead to "
"problems!")
else:
inputs.append(
{'in': {'txid': raw['txid'], 'vout': vout},
'addr': input['addr'], 'amount': new_value})
# For this eval we do not fill the remainder of the block
# because it does not have an influence.
connect().generate(1)
# No error
write("Successfully mined a block with {} TXs.".format(i))
errors = 0
except Exception as e:
write("The following error occured: {}".format(str(e)))
write("Did not manage to write block with {} TXs.".format(i))
errors += 1
|
<reponame>jamesbowman/py-eve<filename>loadable/grave.py
import sys
import datetime
from datetime import timezone
import time
import math
import struct
import numpy as np
from PIL import Image
from gameduino_spidriver import GameduinoSPIDriver
import registers as gd3
import common
import gameduino2.prep
import gameduino2.convert
import tmxreader
TD = 86
class Renderer:
def __init__(self, gd):
self.gd = gd
self.t = 0
version = 102
def load(self):
gd = self.gd
Mloc = 0
if gd.rd32(Mloc) != self.version:
ld = common.Loader(gd)
ld.add(struct.pack("4I", self.version, 0, 0, 0))
self.subload(ld)
def fetchtile(self, l, i, j):
world_map = self.world_map
used = self.used
def reindex(i):
if i == 0:
return None
else:
return used.index(i)
if (i < world_map.width) and (j < world_map.height):
return reindex(l.decoded_content[i + (j * world_map.width)])
else:
return None
def subload(self, ld):
gd = self.gd
world_map = tmxreader.TileMapParser().parse_decode("../tiled-maps/grave.tmx")
print(world_map.tile_sets[0].images[0].source)
used = list(sorted(set(sum([list(l.decoded_content) for l in world_map.layers], [])) - {0}))
print('used', used)
self.world_map = world_map
self.used = used
ts = world_map.tile_sets[0]
tw = int(ts.tilewidth)
th = int(ts.tileheight)
im = (Image.open(world_map.tile_sets[0].images[0].source))
def extract(i):
if hasattr(ts, 'columns'):
w = int(ts.columns)
elif not hasattr(ts, 'spacing'):
w = im.size[0] // tw
else:
w = (im.size[0] + ts.spacing) // (tw + ts.spacing)
x = ts.margin + (tw + ts.spacing) * (i % w)
y = ts.margin + (th + ts.spacing) * (i // w)
print(i, 'is at', (x, y))
r = im.crop((x + 0, y + 0, x + tw, y + th))
r.save("x%d.png" % i)
if 0 and scale:
r = r.resize((stw, sth), Image.ANTIALIAS)
return r
if 0:
for ti in used:
t = extract(ti).resize((90, 90), Image.BICUBIC)
print(ti, t)
gd.BitmapHandle(0)
ld.Lastc("grave-moon.astc")
gd.BitmapHandle(1)
ld.L4(Image.open("grave-bg0.png").convert("L"))
gd.BitmapSize(gd3.BILINEAR, gd3.REPEAT, gd3.BORDER, 1280, 95)
tilebase = ld.a
gd.BitmapHandle(2)
if 1:
for ti in used:
ti -= 1
print('loading', ti)
t = extract(ti).resize((TD, TD), Image.BICUBIC)
(_, d) = gameduino2.convert.convert(t, False, gd3.ARGB4)
ld.add(d)
gd.cmd_setbitmap(tilebase, gd3.ARGB4, TD, TD)
h = 120
walks = []
for i in range(1, 11):
walk = Image.open("zombie/Walk (%d).png" % i).resize((430 * h // 519, h), Image.BILINEAR)
walks.append(walk)
gd.BitmapHandle(3)
ld.ARGB4s(walks)
print('end', hex(ld.a))
def draw(self):
gd = self.gd
gd.Clear()
gd.SaveContext()
gd.ScissorSize(1280, 460)
gd.cmd_gradient(0, 0, 0x1a1a1a, 0, 400, 0x193439)
gd.RestoreContext()
gd.VertexFormat(3)
gd.Begin(gd3.BITMAPS)
gd.BitmapHandle(0)
gd.Vertex2f((1280 - 756) / 2, 40)
sx = 2 * self.t
def scale(n, rgb, y):
gd.SaveContext()
gd.cmd_scale(n, n)
gd.cmd_translate((947 - sx / 2) * n, 0)
gd.cmd_setmatrix()
gd.cmd_loadidentity()
gd.ColorRGB(*common.hex3(rgb))
gd.Vertex2f(0, y)
gd.ClearColorRGB(*common.hex3(rgb))
gd.ScissorXY(0, y + 95)
gd.Clear()
gd.RestoreContext()
if 1:
gd.BitmapHandle(1)
scale(0.6, 0x193439, 400)
scale(0.8, 0x162a2d, 440)
scale(1.0, 0x0d1516, 480)
if 1:
gd.BitmapHandle(2)
tb = (sx // TD)
y0 = 720 - 8 * TD
for j in range(8):
for i in range(16):
for l in self.world_map.layers:
ti = self.fetchtile(l, tb + i, 0 + j)
if ti is not None:
# print('draw', (i, j), ti)
gd.Cell(ti)
gd.Vertex2f(TD * i - sx % TD, y0 + TD * j)
if 0:
frame = (self.t // 4) % 10
gd.BitmapHandle(3)
gd.Cell(frame)
for i in range(8):
gd.Vertex2f(0, y0 + TD * i - 30)
if 0:
gd.Begin(gd3.LINES)
gd.Vertex2f(0, 0)
gd.Vertex2f(1280, 720)
# gd.swap(); gd.screenshot_im().save("out.png") ; sys.exit(0)
self.t += 2
if self.t > 600:
sys.exit(0)
print(self.t)
|
# coding=utf-8
import os
import shutil
import unittest
import pyid3tagger
class ID3v1Test(unittest.TestCase):
def compare_files(self, file_path_1, file_path_2):
file_1_content = open(file_path_1).read()
file_2_content = open(file_path_2).read()
self.assertEqual(file_1_content, file_2_content, 'Files are not equal')
def test_write_id3v1_tag(self):
source_path = 'TestData\\no_tag.mp3'
target_path = 'test.mp3'
compare_file_path = 'TestData\\id3v1\\id3v1_001_basic.mp3'
if os.path.exists(target_path):
os.remove(target_path)
shutil.copyfile(source_path, target_path)
tag = pyid3tagger.ID3v1Tag()
tag.title = 'Title'
tag.artist = 'Artist'
tag.album = 'Album'
tag.year = 2003
tag.comment = 'Comment'
tag.genre = pyid3tagger.ID3v1_GENERES.HIP_HOP
tag.write(target_path)
test_file_content = open(compare_file_path).read()
should_have_no_tag_file_content = open(target_path).read()
self.assertEqual(test_file_content, should_have_no_tag_file_content)
os.remove(target_path)
def test_overwrite_id3v1_tag(self):
source_path = 'TestData\\id3v1\\id3v1_274_extra.mp3'
target_path = 'test.mp3'
compare_file_path = 'TestData\\id3v1\\id3v1_001_basic.mp3'
if os.path.exists(target_path):
os.remove(target_path)
shutil.copyfile(source_path, target_path)
tag = pyid3tagger.ID3v1Tag()
tag.title = 'Title'
tag.artist = 'Artist'
tag.album = 'Album'
tag.year = 2003
tag.comment = 'Comment'
tag.genre = pyid3tagger.ID3v1_GENERES.HIP_HOP
tag.write(target_path)
test_file_content = open(compare_file_path).read()
should_have_no_tag_file_content = open(target_path).read()
self.assertEqual(test_file_content, should_have_no_tag_file_content)
os.remove(target_path)
# todo test exceptions
# test cases for year
def test_read_id3v1_test_case_1(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_001_basic.mp3')
self.assertEqual('Title', tag.title)
self.assertEqual('Artist', tag.artist)
self.assertEqual('Album', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('Comment', tag.comment)
self.assertEqual(7, tag.genre)
def test_read_id3v1_1_test_case_2(self):
tag = pyid3tagger.ID3v1_1Tag()
tag.read('TestData\\id3v1\\id3v1_002_basic.mp3')
self.assertEqual('Title', tag.title)
self.assertEqual('Artist', tag.artist)
self.assertEqual('Album', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('Comment', tag.comment)
self.assertEqual(12, tag.track)
self.assertEqual(7, tag.genre)
def test_read_id3v1_1_test_case_3(self):
tag = pyid3tagger.ID3v1Tag()
try:
tag.read('TestData\\id3v1\\id3v1_003_basic_F.mp3')
except pyid3tagger.PyID3TaggerInvalidData as e:
self.assertEqual('File does not contain a ID3v1 Tag', e.message)
else:
self.fail('No exception where PyID3TaggerInvalidData should be raised')
def test_read_id3v1_1_test_case_4(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_004_basic.mp3')
self.assertEqual('', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_5(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_005_basic.mp3')
self.assertEqual('aaaaaaaaaaaaaaaaaaaaaaaaaaaaaA', tag.title)
self.assertEqual('bbbbbbbbbbbbbbbbbbbbbbbbbbbbbB', tag.artist)
self.assertEqual('cccccccccccccccccccccccccccccC', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('dddddddddddddddddddddddddddddD', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_6(self):
tag = pyid3tagger.ID3v1_1Tag()
tag.read('TestData\\id3v1\\id3v1_006_basic.mp3')
self.assertEqual('aaaaaaaaaaaaaaaaaaaaaaaaaaaaaA', tag.title)
self.assertEqual('bbbbbbbbbbbbbbbbbbbbbbbbbbbbbB', tag.artist)
self.assertEqual('cccccccccccccccccccccccccccccC', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('dddddddddddddddddddddddddddD', tag.comment)
self.assertEqual(1, tag.track)
self.assertEqual(0, tag.genre)
def test_read_id3v1_test_case_7(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_007_basic_W.mp3')
self.assertEqual('12345', tag.title)
self.assertEqual('12345', tag.artist)
self.assertEqual('12345', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('12345', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_8(self):
tag = pyid3tagger.ID3v1_1Tag()
tag.read('TestData\\id3v1\\id3v1_008_basic_W.mp3')
self.assertEqual('12345', tag.title)
self.assertEqual('12345', tag.artist)
self.assertEqual('12345', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('12345', tag.comment)
self.assertEqual(1, tag.track)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_9(self):
tag = pyid3tagger.ID3v1_1Tag()
tag.read('TestData\\id3v1\\id3v1_009_basic.mp3')
self.assertEqual('', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('', tag.comment)
self.assertEqual(255, tag.track)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_10(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_010_year.mp3')
self.assertEqual('', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(0, tag.year)
self.assertEqual('', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_11(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_011_year.mp3')
self.assertEqual('', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(9999, tag.year)
self.assertEqual('', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_1_test_case_12(self):
tag = pyid3tagger.ID3v1Tag()
try:
tag.read('TestData\\id3v1\\id3v1_012_year_F.mp3')
except pyid3tagger.PyID3TaggerInvalidData as e:
self.assertEqual(e.message, 'Invalid year')
else:
self.fail('This should raise an PyID3TaggerInvalidData exeption')
def test_read_id3v1_1_test_case_13(self):
tag = pyid3tagger.ID3v1Tag()
try:
tag.read('TestData\\id3v1\\id3v1_013_year_F.mp3')
except pyid3tagger.PyID3TaggerInvalidData as e:
self.assertEqual(e.message, 'Invalid year')
else:
self.fail('This should raise an PyID3TaggerInvalidData exeption')
def test_read_id3v1_1_test_case_14(self):
tag = pyid3tagger.ID3v1Tag()
try:
tag.read('TestData\\id3v1\\id3v1_014_year_F.mp3')
except pyid3tagger.PyID3TaggerInvalidData as e:
self.assertEqual(e.message, 'Invalid year')
else:
self.fail('This should raise an PyID3TaggerInvalidData exeption')
def test_read_id3v1_test_case_genres(self):
for i in range(80):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_%03i_genre.mp3' % (i + 15,))
self.assertEqual(i, tag.genre)
self.assertEqual(pyid3tagger.ID3v1_GENERES.GENRE_2_NAME[i], tag.title)
def test_read_id3v1_test_case_genres_w(self):
for i in range(80, 148):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_%03i_genre_W.mp3' % (i + 15,))
self.assertEqual(i, tag.genre)
self.assertEqual(pyid3tagger.ID3v1_GENERES.GENRE_2_NAME[i], tag.title)
def test_read_id3v1_test_case_genres_f(self):
for i in range(148, 256):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_%03i_genre_f.mp3' % (i + 15,))
self.assertEqual(i, tag.genre)
self.assertEqual('Unknown/%i' % i, tag.title)
def test_read_id3v1_test_case_271(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_271_extra.mp3')
self.assertEqual(u'räksmörgås', tag.title.decode('latin-1'))
self.assertEqual(u'räksmörgås', tag.artist.decode('latin-1'))
self.assertEqual(u'räksmörgås', tag.album.decode('latin-1'))
self.assertEqual(2003, tag.year)
self.assertEqual(u'räksmörgås', tag.comment.decode('latin-1'))
self.assertEqual(0, tag.genre)
def test_read_id3v1_test_case_272(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_272_extra.mp3')
self.assertEqual(u'räksmörgås', tag.title.decode('utf-8'))
self.assertEqual(u'räksmörgås', tag.artist.decode('utf-8'))
self.assertEqual(u'räksmörgås', tag.album.decode('utf-8'))
self.assertEqual(2003, tag.year)
self.assertEqual(u'räksmörgås', tag.comment.decode('utf-8'))
self.assertEqual(0, tag.genre)
def test_read_id3v1_test_case_273(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_273_extra.mp3')
self.assertEqual('', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('http://www.id3.org/', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_test_case_274(self):
tag = pyid3tagger.ID3v1Tag()
tag.read('TestData\\id3v1\\id3v1_274_extra.mp3')
self.assertEqual('', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('www.id3.org/', tag.comment)
self.assertEqual(0, tag.genre)
def test_read_id3v1_as_id3v1_1(self):
tag = pyid3tagger.ID3v1_1Tag()
tag.read('TestData\\id3v1\\id3v1_039_genre.mp3')
self.assertEqual('Soundtrack', tag.title)
self.assertEqual('', tag.artist)
self.assertEqual('', tag.album)
self.assertEqual(2003, tag.year)
self.assertEqual('', tag.comment)
self.assertEqual(0, tag.track)
self.assertEqual(24, tag.genre)
def test_write_id3v1_test_case_1(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = 'Title'
tag.artist = 'Artist'
tag.album = 'Album'
tag.year = 2003
tag.comment = 'Comment'
tag.genre = pyid3tagger.ID3v1_GENERES.HIP_HOP
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_001_basic.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_2(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1_1Tag()
tag.title = 'Title'
tag.artist = 'Artist'
tag.album = 'Album'
tag.year = 2003
tag.comment = 'Comment'
tag.track = 12
tag.genre = pyid3tagger.ID3v1_GENERES.HIP_HOP
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_002_basic.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_4(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = ''
tag.artist = ''
tag.album = ''
tag.year = 2003
tag.comment = ''
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_004_basic.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_5(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = 'aaaaaaaaaaaaaaaaaaaaaaaaaaaaaA'
tag.artist = 'bbbbbbbbbbbbbbbbbbbbbbbbbbbbbB'
tag.album = 'cccccccccccccccccccccccccccccC'
tag.year = 2003
tag.comment = 'dddddddddddddddddddddddddddddD'
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_005_basic.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_6(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1_1Tag()
tag.title = 'aaaaaaaaaaaaaaaaaaaaaaaaaaaaaA'
tag.artist = 'bbbbbbbbbbbbbbbbbbbbbbbbbbbbbB'
tag.album = 'cccccccccccccccccccccccccccccC'
tag.year = 2003
tag.track = 1
tag.comment = 'dddddddddddddddddddddddddddD'
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_006_basic.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_9(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1_1Tag()
tag.title = ''
tag.artist = ''
tag.album = ''
tag.year = 2003
tag.track = 255
tag.comment = ''
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_009_basic.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_10(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = ''
tag.artist = ''
tag.album = ''
tag.year = 0
tag.comment = ''
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_010_year.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_1_test_case_11(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = ''
tag.artist = ''
tag.album = ''
tag.year = 9999
tag.comment = ''
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_011_year.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_genres(self):
for i in range(80):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = pyid3tagger.ID3v1_GENERES.GENRE_2_NAME[i]
tag.year = 2003
tag.genre = i
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_%03i_genre.mp3' % (i + 15,), 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_genres_w(self):
for i in range(80, 148):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = pyid3tagger.ID3v1_GENERES.GENRE_2_NAME[i]
tag.year = 2003
tag.genre = i
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_%03i_genre_W.mp3' % (i + 15,), 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_genres_f(self):
for i in range(148, 256):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = 'Unknown/%i' % i
tag.year = 2003
tag.genre = i
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_%03i_genre_F.mp3' % (i + 15,), 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_271(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = u'räksmörgås'.encode('latin-1')
tag.artist = u'räksmörgås'.encode('latin-1')
tag.album = u'räksmörgås'.encode('latin-1')
tag.year = 2003
tag.comment = u'räksmörgås'.encode('latin-1')
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_271_extra.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_272(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.title = u'räksmörgås'.encode('utf-8')
tag.artist = u'räksmörgås'.encode('utf-8')
tag.album = u'räksmörgås'.encode('utf-8')
tag.year = 2003
tag.comment = u'räksmörgås'.encode('utf-8')
tag.genre = pyid3tagger.ID3v1_GENERES.BLUES
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_272_extra.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_273(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.year = 2003
tag.comment = 'http://www.id3.org/'
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_273_extra.mp3', 'test.mp3')
os.remove('test.mp3')
def test_write_id3v1_test_case_274(self):
if os.path.exists('test.mp3'):
os.remove('test.mp3')
shutil.copyfile('TestData\\no_tag.mp3', 'test.mp3')
tag = pyid3tagger.ID3v1Tag()
tag.year = 2003
tag.comment = 'www.id3.org/'
tag.write('test.mp3')
self.compare_files('TestData\\id3v1\\id3v1_274_extra.mp3', 'test.mp3')
os.remove('test.mp3')
if __name__ == '__main__':
unittest.main()
|
<reponame>wrobstory/reconciler
# -*- coding: utf-8 -*-
"""
Reconciler: reconcile messages in S3 to those that have been loaded in Redshift
---------------------------
Given a list of S3 buckets, determine if any of the data has already
been loaded into redshift (via a successful COMMIT in the stl_load_commits tbl)
and diff the two. Can optionally delete the S3 objects that have been
loaded
"""
from __future__ import print_function
import os
from boto.s3.connection import S3Connection
from boto.s3.key import Key
import psycopg2
class Reconciler(object):
def __init__(self, aws_access_key_id=None, aws_secret_access_key=None,
database=None, user=None, password=None, host=None,
port=None):
"""
The Reconciler reconciles objects in S3 with those that have already
been loaded in Redshift.
The aws keys are not required if you have environmental params set
for boto to pick up:
http://boto.readthedocs.org/en/latest/s3_tut.html#creating-a-connection
Parameters
----------
aws_access_key_id: str
aws_secret_access_key: str
database: str
user: str
password: str
host: str
port: int
"""
if aws_access_key_id and aws_secret_access_key:
self.s3conn = S3Connection(aws_access_key_id, aws_secret_access_key)
else:
self.s3conn = S3Connection()
database = database or os.environ.get('PGDATABASE')
user = user or os.environ.get('PGUSER')
password = password or os.<PASSWORD>('PGPASSWORD')
host = host or os.environ.get('PGHOST')
port = port or os.environ.get('PGPORT') or 5439
print('Connecting to Redshift...')
self.conn = psycopg2.connect(database=database, user=user,
password=password, host=host,
port=port)
self.cur = self.conn.cursor()
self.bucket_cache = {}
def _get_bucket_from_cache(self, buckpath):
"""Get bucket from cache, or add to cache if does not exist"""
if buckpath not in self.bucket_cache:
self.bucket_cache[buckpath] = self.s3conn.get_bucket(buckpath)
return self.bucket_cache[buckpath]
def _get_bucket_and_key(self, path):
"""Get top-level bucket and nested key path"""
if '/' in path:
parts = path.split('/')
buckpath = parts[0]
keypath = os.path.join(*parts[1:])
else:
buckpath, keypath = path, ""
return buckpath, keypath
def get_committed_keys(self, start_date, end_date):
"""
Get all S3 LOADs that have been commited to Redshift in a given
time window.
`start_date` and `end_date` must be Redshift compatible dates:
Parameters
----------
start_date: str
end_date: str
Returns
-------
Set of object names
"""
query = """
select rtrim(l.filename)
from stl_load_commits l, stl_query q
where l.query=q.query
and exists
(select xid from stl_utilitytext
where xid=q.xid and rtrim("text")='COMMIT')
and q.starttime between %s and %s
and l.filename like 's3://%%'
and q.querytxt not like '%%CopyManifestJsonAutoNoload%%'
order by q.starttime desc;
"""
print('Getting keys already committed to Redshift...')
self.cur.execute(query, (start_date, end_date))
return {x[0] for x in self.cur}
def get_all_keys(self, bucket_path):
"""
Get all keys in a given keypath. Given a folder or bucket, will
get a set of all keys in that bucket/folder.
Parameters
----------
bucket_path: str
Ex: my.bucket/folder1/
"""
buckpath, keypath = self._get_bucket_and_key(bucket_path)
print('Getting bucket...')
bucket = self._get_bucket_from_cache(buckpath)
print('Getting all keys in bucket...')
return {os.path.join('s3://', k.bucket.name, k.name)
for k in bucket.list(keypath)}
def diff_redshift_and_bucket(self, start_date, end_date, bucket_path):
"""
Given a start date and end date, get the S3 keys that have been
committed in a load, the S3 keys currently in the given bucket
path, and the difference of the two.
Parameters
----------
start_date: str
end_date: str
bucket_path: str
Returns
-------
Dict: {'comitted_keys': {"s3://foo", ...},
'keys_in_bucket': {"s3://bar", ...},
'bucket_keys_to_be_committed': {"s3://bar", ...},
'bucket_keys_already_committed': {"s3://foo", ...}}
"""
ck = self.get_committed_keys(start_date, end_date)
keys = self.get_all_keys(bucket_path)
return {'committed_keys': ck,
'keys_in_bucket': keys,
'bucket_keys_to_be_committed': keys.difference(ck),
'bucket_keys_already_committed': keys.intersection(ck)}
def _iter_keys(self, keys):
"""Iterate through buckets/keys"""
for key in keys:
splitter = key.split('/')
buckname, keyval = splitter[2], os.path.join(*splitter[3:])
yield self._get_bucket_from_cache(buckname), keyval
def copy_committed_keys(self, diff, new_folder):
"""
Given the diff from `diff_redshift_and_bucket`, copy the keys
that have already been committed to a new bucket folder for
later validation
"""
for buck, k in self._iter_keys(diff['bucket_keys_already_committed']):
new_key = os.path.join(new_folder, k.split('/')[-1])
print("Copying {} to {}...".format(k, new_key))
buck.copy_key(new_key, buck.name, k)
def delete_committed_keys(self, diff):
"""
Given the diff from `diff_redshift_and_bucket`, delete the keys
that have already been committed.
Parameters
----------
diff: Dict
"""
for b, k in self._iter_keys(diff['bucket_keys_already_committed']):
print("Deleting key {}...".format(k))
b.delete_key(k) |
<filename>arhuaco/analysis/generative/rnn_gen.py<gh_stars>1-10
# Copyright (c) 2019 <NAME>.
# All Rights Reserved.
from __future__ import print_function
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.layers import LSTM
from keras.optimizers import RMSprop
from keras.utils.data_utils import get_file
from arhuaco.analysis.features.data_helpers\
import DataHelpers
from tqdm import tqdm
import numpy as np
import random
import sys
import string
import os
# This is the main class for RNN based generative models,
# That creates synthetic data based on previous examples.
class RnnGen:
def __init__(self, data_helpers, maxlen,
step, num_epochs, num_chars,
samples_per_epoch, weights_file,
model_file, generated_file
, number_generated):
# Parameters
self.maxlen = maxlen
self.step = step
self.num_epochs = num_epochs
self.num_chars = 0
self.model = None
self.data_helpers = data_helpers
self.data_generator = None
self.char_indices = None
self.indices_char = None
self.samples_per_epoch = samples_per_epoch
self.weights_file = weights_file
self.model_file = model_file
self.generated_file = generated_file
self.number_generated = number_generated
def get_data(self):
self.data_generator = self.data_helpers.generator_from_file(
self.data_helpers.data_source[1],
self.data_helpers.samples_per_batch)
# Initialize character set
chars = sorted(list(set(string.printable+"\n")))
print('total chars:', len(chars))
self.num_chars = len(chars)
self.char_indices = dict((c, i) for i, c in enumerate(chars))
self.indices_char = dict((i, c) for i, c in enumerate(chars))
def format_text(self, text):
# cut the text in semi-redundant
# sequences of maxlen characters
sentences = []
next_chars = []
for i in range(0, len(text) - self.maxlen, self.step):
sentences.append(text[i: i + self.maxlen])
next_chars.append(text[i + self.maxlen])
print('nb sequences:', len(sentences))
print('Vectorization...')
X = np.zeros((len(sentences), self.maxlen,
self.num_chars), dtype=np.bool)
y = np.zeros((len(sentences), self.num_chars),
dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
X[i, t, self.char_indices[char]] = 1
y[i, self.char_indices[next_chars[i]]] = 1
return (X,y)
def build_model(self):
# build the model: a single LSTM
print('Build model...')
self.model = Sequential()
self.model.add(LSTM(128,
input_shape=(self.maxlen,
self.num_chars)))
self.model.add(Dense(self.num_chars))
self.model.add(Activation('softmax'))
optimizer = RMSprop(lr=0.01)
self.model.compile(loss='categorical_crossentropy',
optimizer=optimizer)
def sample(self, preds, temperature=1.0):
# helper function to sample an index
# from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def train_model(self):
# train the model, output generated text
# after each iteration
if os.path.exists(self.weights_file):
self.model.load_weights(self.weights_file)
print("Model loaded from disk.")
x_train = next(self.data_generator)
text = self.data_helpers.get_text_from_list(
x_train)
else:
for iteration in range(1):
x_train = next(self.data_generator)
text = self.data_helpers.get_text_from_list(
x_train)
print('total chars in text:', len(text))
X, y = self.format_text(text)
print('-' * 50)
print('Iteration', iteration)
self.model.fit(X, y,
batch_size=self.samples_per_epoch,
nb_epoch=self.num_epochs)
# Save model
print("dumping weights to file...")
# serialize model to JSON
model_json = self.model.to_json()
with open(self.model_file, "w") as json_file:
json_file.write(model_json)
self.model.save_weights(self.weights_file,
overwrite=True)
self.test_model(text)
def test_model(self, text):
# Generate new data
print("Size of text:"+str(len(text)))
for diversity in [0.2, 0.5, 1.0, 1.2]:
start_index = random.randint(0, len(text)\
- self.maxlen - 1)
with open(self.generated_file+"-"+str(diversity),
"a") as gen_file:
print()
print('----- diversity:', diversity)
# Create a seed for generating data
generated = ''
sentence = text[start_index: start_index + self.maxlen]
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
for i in tqdm(range(self.number_generated)):
x = np.zeros((1, self.maxlen, self.num_chars))
for t, char in enumerate(sentence):
x[0, t, self.char_indices[char]] = 1.
preds = self.model.predict(x, verbose=0)[0]
next_index = self.sample(preds, diversity)
next_char = self.indices_char[next_index]
generated += next_char
sentence = sentence[1:] + next_char
gen_file.write(generated)
|
import pytest
from django.test import RequestFactory
from va_explorer.users.forms import (
ExtendedUserCreationForm,
UserSetPasswordForm,
UserUpdateForm,
)
from va_explorer.users.tests.factories import (
GroupFactory,
LocationFactory,
NewUserFactory,
)
pytestmark = pytest.mark.django_db
class TestUserCreationForm:
def test_valid_form_with_national_access(self, rf: RequestFactory):
# A user with proto_user params does not exist yet.
proto_user = NewUserFactory.build()
group = GroupFactory.create()
form = ExtendedUserCreationForm(
{
"name": proto_user.name,
"email": proto_user.email,
"group": group,
"geographic_access": "national",
"locations": [],
}
)
# Note: The form expects a request object to be set in order to save it
request = rf.get("/fake-url/")
form.request = request
assert form.is_valid()
def test_valid_form_with_location_specific_access(self, rf: RequestFactory):
# A user with proto_user params does not exist yet.
proto_user = NewUserFactory.build()
group = GroupFactory.create()
location = LocationFactory.create()
form = ExtendedUserCreationForm(
{
"name": proto_user.name,
"email": proto_user.email,
"group": group,
"geographic_access": "location-specific",
"locations": [location],
}
)
# Note: The form expects a request object to be set in order to save it
request = rf.get("/fake-url/")
form.request = request
assert form.is_valid()
def test_email_uniqueness(self):
# A user with existing_user params exists already.
existing_user = NewUserFactory.create()
group = GroupFactory.create()
location = LocationFactory.create()
form = ExtendedUserCreationForm(
{
"name": existing_user.name,
"email": existing_user.email,
"group": group,
"geographic_access": "location-specific",
"locations": [location],
}
)
assert not form.is_valid()
assert len(form.errors) == 1
assert "email" in form.errors
def test_basic_form_field_requirements(self):
form = ExtendedUserCreationForm(
{
"name": "",
"email": "",
"group": "",
"geographic_access": "",
"locations": "",
}
)
assert not form.is_valid()
assert len(form.errors) == 4
assert "email" in form.errors
assert "name" in form.errors
assert "group" in form.errors
assert "geographic_access" in form.errors
def test_location_required(self):
# A user with proto_user params does not exist yet.
proto_user = NewUserFactory.build()
group = GroupFactory.create()
form = ExtendedUserCreationForm(
{
"name": proto_user.name,
"email": proto_user.email,
"group": group,
"geographic_access": "location-specific",
"locations": [],
}
)
assert not form.is_valid()
assert len(form.errors) == 1
assert "locations" in form.errors
def test_location_not_required(self):
# A user with proto_user params does not exist yet.
proto_user = NewUserFactory.build()
group = GroupFactory.create()
location = LocationFactory.create()
form = ExtendedUserCreationForm(
{
"name": proto_user.name,
"email": proto_user.email,
"group": group,
"geographic_access": "national",
"locations": [location],
}
)
assert not form.is_valid()
assert len(form.errors) == 1
assert "locations" in form.errors
class TestUserUpdateForm:
def test_valid_form(self, rf: RequestFactory):
new_group = GroupFactory.create()
location = LocationFactory.create()
form = UserUpdateForm(
{
"name": "<NAME>",
"email": "<EMAIL>",
"group": new_group,
"is_active": False,
"geographic_access": "location-specific",
"locations": [location],
}
)
assert form.is_valid()
def test_group_required(self):
# A user with proto_user params does not exist yet.
proto_user = NewUserFactory.build()
location = LocationFactory.create()
form = UserUpdateForm(
{
"name": proto_user.name,
"email": proto_user.email,
"group": "",
"geographic_access": "location-specific",
"locations": [location],
}
)
assert not form.is_valid()
assert len(form.errors) == 1
assert "group" in form.errors
class TestUserSetPasswordForm:
def test_valid_form(self, rf: RequestFactory):
form = UserSetPasswordForm(
{
"password1": "<PASSWORD>!",
"password2": "<PASSWORD>!",
}
)
assert form.is_valid()
def test_invalid_form(self, rf: RequestFactory):
form = UserSetPasswordForm(
{
"password1": "<PASSWORD>!",
"password2": "<PASSWORD>!",
}
)
assert not form.is_valid()
assert "You must type the same password each time." in form.errors["password2"]
assert len(form.errors) == 1
|
<reponame>TheWardoctor/wardoctors-repo
# -*- coding: utf-8 -*-
'''
Bubbles Add-on
Copyright (C) 2016 Bubbles
This program is free software: you can redistribute it and/or modify
it under the terms of the GNU General Public License as published by
the Free Software Foundation, either version 3 of the License, or
(at your option) any later version.
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
GNU General Public License for more details.
You should have received a copy of the GNU General Public License
along with this program. If not, see <http://www.gnu.org/licenses/>.
'''
import re,urllib,urlparse,json,xbmc
from resources.lib.modules import cleantitle
from resources.lib.modules import client
from resources.lib.extensions import metadata
from resources.lib.extensions import tools
from resources.lib.extensions import network
class source:
def __init__(self):
self.pack = False # Checked by provider.py
self.priority = 0
self.language = ['un']
self.domains = ['yts.ag'] # Other YIFI domains do not have an API.
self.base_link = 'https://yts.ag'
self.search_link = '/api/v2/list_movies.json?query_term=%s&limit=50&sort_by=seeds&order_by=desc&with_rt_ratings=false'
def movie(self, imdb, title, localtitle, year):
try:
url = {'imdb': imdb, 'title': title, 'year': year}
url = urllib.urlencode(url)
return url
except:
return
def sources(self, url, hostDict, hostprDict):
sources = []
try:
if url == None:
raise Exception()
data = urlparse.parse_qs(url)
data = dict([(i, data[i][0]) if data[i] else (i, '') for i in data])
if 'exact' in data and data['exact']:
titleYear = title = data['tvshowtitle'] if 'tvshowtitle' in data else data['title']
year = None
season = None
episode = None
pack = False
packCount = None
else:
title = data['tvshowtitle'] if 'tvshowtitle' in data else data['title']
titleYear = '%s %s' % (title, str(data['year']))
year = int(data['year']) if 'year' in data and not data['year'] == None else None
season = int(data['season']) if 'season' in data and not data['season'] == None else None
episode = int(data['episode']) if 'episode' in data and not data['episode'] == None else None
query = data['imdb'] if 'imdb' in data and not data['imdb'] == None else title
url = urlparse.urljoin(self.base_link, self.search_link) % query
result = json.loads(client.request(url))
movie = result['data']['movies'][0]
name = movie['title_long'] + ' '
torrents = movie['torrents']
for torrent in torrents:
quality = torrent['quality']
if quality.lower() == '3d':
quality += ' HD1080'
jsonName = name + quality
jsonSize = torrent['size_bytes']
jsonSeeds = torrent['seeds']
jsonHash = torrent['hash']
jsonLink = network.Container(jsonHash).torrentMagnet(title = titleYear)
# Metadata
meta = metadata.Metadata(name = jsonName, title = title, year = year, season = season, episode = episode, link = jsonLink, size = jsonSize, seeds = jsonSeeds)
jsonLink = network.Container(jsonHash).torrentMagnet(title = meta.title(extended = True))
# Ignore
if meta.ignore(False):
continue
# Add
sources.append({'url' : jsonLink, 'debridonly' : False, 'direct' : False, 'source' : 'torrent', 'language' : self.language[0], 'quality': meta.videoQuality(), 'metadata' : meta, 'file' : jsonName})
return sources
except:
return sources
def resolve(self, url):
return url
|
<reponame>mpoiitis/iSpine<gh_stars>1-10
from tensorflow.keras.layers import Layer
import tensorflow as tf
class Sampling(Layer):
"""Uses (z_mean, z_log_var) to sample z, the vector encoding a digit."""
def call(self, inputs, **kwargs):
z_mean, z_log_var = inputs
batch = tf.shape(z_mean)[0]
dim = tf.shape(z_mean)[1]
epsilon = tf.keras.backend.random_normal(shape=(batch, dim))
return z_mean + tf.exp(0.5 * z_log_var) * epsilon
class VariationalEncoder(Layer):
def __init__(self, input_dim, output_dim, hidden_dim, num_features_nonzero, activation, dropout, is_sparse_inputs=True, name="variational_encoder", **kwargs):
super(VariationalEncoder, self).__init__(name=name, **kwargs)
self.proj = GraphConvolution(input_dim=input_dim, output_dim=hidden_dim, num_features_nonzero=num_features_nonzero,
activation=activation, dropout=dropout, is_sparse_inputs=True)
self.mean = GraphConvolution(input_dim=hidden_dim, output_dim=output_dim, num_features_nonzero=num_features_nonzero,
activation=lambda x: x, dropout=dropout)
self.log_var = GraphConvolution(input_dim=hidden_dim, output_dim=output_dim, num_features_nonzero=num_features_nonzero,
activation=lambda x: x, dropout=dropout)
self.sampling = Sampling()
def call(self, inputs, training):
x, support = inputs
h = self.proj((x, support), training)
z_mean = self.mean((h, support), training)
z_log_var = self.log_var((h, support), training)
z = self.sampling((z_mean, z_log_var))
return z_mean, z_log_var, z
class Encoder(Layer):
def __init__(self, input_dim, output_dim, num_features_nonzero, activation, dropout, is_sparse_inputs=True, name="encoder", **kwargs):
super(Encoder, self).__init__(name=name, **kwargs)
self.proj = GraphConvolution(input_dim=input_dim, output_dim=output_dim, num_features_nonzero=num_features_nonzero,
activation=activation, dropout=dropout, is_sparse_inputs=True)
def call(self, inputs, training):
z = self.proj(inputs, training)
return z
class Decoder(Layer):
"""Converts z, the encoded digit vector, back into a readable digit."""
def __init__(self, input_dim, dropout=0., act=tf.nn.sigmoid, name="decoder", **kwargs):
super(Decoder, self).__init__(name=name, **kwargs)
self.dropout = dropout
self.act = act
def call(self, inputs):
inputs = tf.nn.dropout(inputs, self.dropout)
x = tf.transpose(inputs)
x = tf.matmul(inputs, x)
x = tf.reshape(x, [-1])
outputs = self.act(x)
return outputs
def dot(x, y, sparse=False):
"""
Wrapper for tf.matmul (sparse vs dense).
"""
if sparse:
res = tf.sparse.sparse_dense_matmul(x, y)
else:
res = tf.matmul(x, y)
return res
def sparse_dropout(x, rate, noise_shape):
"""
Dropout for sparse tensors.
"""
random_tensor = 1 - rate
random_tensor += tf.random.uniform(noise_shape)
dropout_mask = tf.cast(tf.floor(random_tensor), dtype=tf.bool)
pre_out = tf.sparse.retain(x, dropout_mask)
return pre_out * (1./(1 - rate))
class GraphConvolution(Layer):
"""
Graph convolution layer.
"""
def __init__(self, input_dim, output_dim, num_features_nonzero,
dropout=0.,
is_sparse_inputs=False,
activation=tf.nn.relu,
bias=False,
featureless=False, **kwargs):
super(GraphConvolution, self).__init__(**kwargs)
self.dropout = dropout
self.activation = activation
self.is_sparse_inputs = is_sparse_inputs
self.featureless = featureless
self.bias = bias
self.num_features_nonzero = num_features_nonzero
self.weights_ = []
for i in range(1):
w = self.add_variable('weight' + str(i), [input_dim, output_dim])
self.weights_.append(w)
if self.bias:
self.bias = self.add_variable('bias', [output_dim])
def call(self, inputs, training=None):
x, support_ = inputs
# dropout
if training is not False and self.is_sparse_inputs:
x = sparse_dropout(x, self.dropout, self.num_features_nonzero)
elif training is not False:
x = tf.nn.dropout(x, self.dropout)
# convolve
supports = list()
for i in range(len(support_)):
if not self.featureless: # if it has features x
pre_sup = dot(x, self.weights_[i], sparse=self.is_sparse_inputs)
else:
pre_sup = self.weights_[i]
support = dot(support_[i], pre_sup, sparse=True)
supports.append(support)
output = tf.add_n(supports)
# bias
if self.bias:
output += self.bias
return self.activation(output)
|
# Licensed under a 3-clause BSD style license - see LICENSE.rst
"""
MAST Portal
===========
This module contains various methods for querying the MAST Portal.
"""
from __future__ import print_function, division
import warnings
import json
import time
import os
import keyring
import threading
import uuid
import numpy as np
from requests import HTTPError
from getpass import getpass
import astropy.units as u
import astropy.coordinates as coord
from astropy.table import Table, Row, vstack, MaskedColumn
from astropy.logger import log
from astropy.utils import deprecated
from astropy.utils.console import ProgressBarOrSpinner
from astropy.utils.exceptions import AstropyDeprecationWarning
from six.moves.urllib.parse import quote as urlencode
from ..query import QueryWithLogin
from ..utils import commons, async_to_sync
from ..utils.class_or_instance import class_or_instance
from ..exceptions import (TimeoutError, InvalidQueryError, RemoteServiceError,
ResolverError, MaxResultsWarning,
NoResultsWarning, InputWarning, AuthenticationWarning)
from . import conf
from . import fpl
__all__ = ['Observations', 'ObservationsClass',
'Mast', 'MastClass']
def _prepare_service_request_string(json_obj):
"""
Takes a mashup JSON request object and turns it into a url-safe string.
Parameters
----------
json_obj : dict
A Mashup request JSON object (python dictionary).
Returns
-------
response : str
URL encoded Mashup Request string.
"""
# Append cache breaker
if not 'cacheBreaker' in json_obj:
json_obj['cacheBreaker'] = _generate_uuid_string()
request_string = json.dumps(json_obj)
return 'request={}'.format(urlencode(request_string))
def _parse_type(dbtype):
"""
Takes a data type as returned by a database call and regularizes it into a
triplet of the form (human readable datatype, python datatype, default value).
Parameters
----------
dbtype : str
A data type, as returned by a database call (ex. 'char').
Returns
-------
response : tuple
Regularized type tuple of the form (human readable datatype, python datatype, default value).
For example:
_parse_type("short")
('integer', np.int64, -999)
"""
dbtype = dbtype.lower()
return {
'char': ('string', str, ""),
'string': ('string', str, ""),
'datetime': ('string', str, ""), # TODO: handle datetimes correctly
'date': ('string', str, ""), # TODO: handle datetimes correctly
'double': ('float', np.float64, np.nan),
'float': ('float', np.float64, np.nan),
'decimal': ('float', np.float64, np.nan),
'int': ('integer', np.int64, -999),
'short': ('integer', np.int64, -999),
'long': ('integer', np.int64, -999),
'number': ('integer', np.int64, -999),
'boolean': ('boolean', bool, None),
'binary': ('boolean', bool, None),
'unsignedbyte': ('byte', np.ubyte, -999)
}.get(dbtype, (dbtype, dbtype, dbtype))
def _generate_uuid_string():
"""
Generates a UUID using Python's UUID module
Parameters
----------
None
Returns
-------
response: str
Generated UUID string
"""
return str(uuid.uuid4())
def _mashup_json_to_table(json_obj, col_config=None):
"""
Takes a JSON object as returned from a Mashup request and turns it into an `~astropy.table.Table`.
Parameters
----------
json_obj : dict
A Mashup response JSON object (python dictionary)
col_config : dict, optional
Dictionary that defines column properties, e.g. default value.
Returns
-------
response : `~astropy.table.Table`
"""
data_table = Table(masked=True)
if not all(x in json_obj.keys() for x in ['fields', 'data']):
raise KeyError("Missing required key(s) 'data' and/or 'fields.'")
for col, atype in [(x['name'], x['type']) for x in json_obj['fields']]:
# Removing "_selected_" column
if col == "_selected_":
continue
# reading the colum config if given
ignore_value = None
if col_config:
col_props = col_config.get(col, {})
ignore_value = col_props.get("ignoreValue", None)
# regularlizing the type
reg_type = _parse_type(atype)
atype = reg_type[1]
ignore_value = reg_type[2] if (ignore_value is None) else ignore_value
# Make the column list (don't assign final type yet or there will be errors)
col_data = np.array([x.get(col, ignore_value) for x in json_obj['data']], dtype=object)
if ignore_value is not None:
col_data[np.where(np.equal(col_data, None))] = ignore_value
# no consistant way to make the mask because np.equal fails on ''
# and array == value fails with None
if atype == 'str':
col_mask = (col_data == ignore_value)
else:
col_mask = np.equal(col_data, ignore_value)
# add the column
data_table.add_column(MaskedColumn(col_data.astype(atype), name=col, mask=col_mask))
return data_table
def _fabric_json_to_table(json_obj):
"""
Takes a JSON object as returned from a MAST microservice request and turns it into an `~astropy.table.Table`.
Parameters
----------
json_obj : dict
A MAST microservice response JSON object (python dictionary)
Returns
-------
response : `~astropy.table.Table`
"""
data_table = Table(masked=True)
if not all(x in json_obj.keys() for x in ['info', 'data']):
raise KeyError("Missing required key(s) 'data' and/or 'info.'")
# determine database type key in case missing
type_key = 'type' if json_obj['info'][0].get('type') else 'db_type'
# for each item in info, store the type and column name
for idx, col, col_type, ignore_value in \
[(idx, x['name'], x[type_key], "NULL") for idx, x in enumerate(json_obj['info'])]:
# [(idx, x['name'], x[type_key], x['default_value']) for idx, x in enumerate(json_obj['info'])]:
# if default value is NULL, set ignore value to None
if ignore_value == "NULL":
ignore_value = None
# making type adjustments
if col_type == "char" or col_type == "STRING":
col_type = "str"
ignore_value = "" if (ignore_value is None) else ignore_value
elif col_type == "boolean" or col_type == "BINARY":
col_type = "bool"
elif col_type == "unsignedByte":
col_type = np.ubyte
elif col_type == "int" or col_type == "short" or col_type == "long" or col_type == "NUMBER":
# int arrays do not admit Non/nan vals
col_type = np.int64
ignore_value = -999 if (ignore_value is None) else ignore_value
elif col_type == "double" or col_type == "float" or col_type == "DECIMAL":
# int arrays do not admit Non/nan vals
col_type = np.float64
ignore_value = -999 if (ignore_value is None) else ignore_value
elif col_type == "DATETIME":
col_type = "str"
ignore_value = "" if (ignore_value is None) else ignore_value
# Make the column list (don't assign final type yet or there will be errors)
# Step through data array of values
col_data = np.array([x[idx] for x in json_obj['data']], dtype=object)
if ignore_value is not None:
col_data[np.where(np.equal(col_data, None))] = ignore_value
# no consistant way to make the mask because np.equal fails on ''
# and array == value fails with None
if col_type == 'str':
col_mask = (col_data == ignore_value)
else:
col_mask = np.equal(col_data, ignore_value)
# add the column
data_table.add_column(MaskedColumn(col_data.astype(col_type), name=col, mask=col_mask))
return data_table
@async_to_sync
class MastClass(QueryWithLogin):
"""
MAST query class.
Class that allows direct programatic access to the MAST Portal,
more flexible but less user friendly than `ObservationsClass`.
"""
def __init__(self, mast_token=None):
super(MastClass, self).__init__()
self._MAST_REQUEST_URL = conf.server + "/api/v0/invoke"
self._COLUMNS_CONFIG_URL = conf.server + "/portal/Mashup/Mashup.asmx/columnsconfig"
self._MAST_CATALOGS_REQUEST_URL = conf.catalogsserver + "/api/v0.1/"
self._MAST_CATALOGS_SERVICES = {
"panstarrs": {
"path": "panstarrs/{data_release}/{table}.json",
"args": {"data_release": "dr2", "table": "mean"}
}
}
self.TIMEOUT = conf.timeout
self.PAGESIZE = conf.pagesize
self._column_configs = dict()
self._current_service = None
self._SESSION_INFO_URL = conf.server + "/whoami"
self._MAST_DOWNLOAD_URL = conf.server + "/api/v0.1/Download/file"
self._MAST_BUNDLE_URL = conf.server + "/api/v0.1/Download/bundle"
if mast_token:
self.login(token=mast_token)
def _login(self, token=None, store_token=False, reenter_token=False): # pragma: no cover
"""
Log into the MAST portal.
Parameters
----------
token : string, optional
Default is None.
The token to authenticate the user.
This can be generated at
https://auth.mast.stsci.edu/token?suggested_name=Astroquery&suggested_scope=mast:exclusive_access.
If not supplied, it will be prompted for if not in the keyring or set via $MAST_API_TOKEN
store_token : bool, optional
Default False.
If true, MAST token will be stored securely in your keyring.
reenter_token : bool, optional
Default False.
Asks for the token even if it is already stored in the keyring or $MAST_API_TOKEN environment variable.
This is the way to overwrite an already stored password on the keyring.
"""
auth_link = (conf.server.replace("mast", "auth.mast") +
"/token?suggested_name=Astroquery&suggested_scope=mast:exclusive_access")
if token is None and "MAST_API_TOKEN" in os.environ:
token = os.environ["MAST_API_TOKEN"]
if token is None:
token = keyring.get_password("astroquery:mast.stsci.edu.token", "masttoken")
if token is None or reenter_token:
info_msg = "If you do not have an API token already, visit the following link to create one: "
log.info(info_msg + auth_link)
token = getpass("Enter MAST API Token: ")
# store token if desired
if store_token:
keyring.set_password("astroquery:mast.stsci.edu.token", "masttoken", token)
self._session.headers["Accept"] = "application/json"
self._session.cookies["mast_token"] = token
info = self.session_info(verbose=False)
if not info["anon"]:
log.info("MAST API token accepted, welcome {}".format(info["attrib"].get("display_name")))
else:
warn_msg = ("MAST API token invalid!\n"
"To make create a new API token visit to following link: " +
auth_link)
warnings.warn(warn_msg, AuthenticationWarning)
return not info["anon"]
@deprecated(since="v0.3.9", message=("The get_token function is deprecated, "
"session token is now the token used for login."))
def get_token(self):
return None
def session_info(self, silent=None, verbose=None):
"""
Displays information about current MAST user, and returns user info dictionary.
Parameters
----------
silent :
Deprecated. Use verbose instead.
verbose : bool, optional
Default True. Set to False to suppress output to stdout.
Returns
-------
response : dict
"""
# Dealing with deprecated argument
if (silent is not None) and (verbose is not None):
warnings.warn(("Argument 'silent' has been deprecated, "
"will be ignored in favor of 'verbose'"), AstropyDeprecationWarning)
elif silent is not None:
warnings.warn(("Argument 'silent' has been deprecated, "
"and will be removed in the future. "
" Use 'verbose' instead."), AstropyDeprecationWarning)
verbose = not silent
elif (silent is None) and (verbose is None):
verbose = True
# get user information
self._session.headers["Accept"] = "application/json"
response = self._session.request("GET", self._SESSION_INFO_URL)
info_dict = response.json()
if verbose:
for key, value in info_dict.items():
if isinstance(value, dict):
for subkey, subval in value.items():
print("{}.{}: {}".format(key, subkey, subval))
else:
print("{}: {}".format((key, value)))
return info_dict
def _fabric_request(self, method, url, params=None, data=None, headers=None,
files=None, stream=False, auth=None, cache=False):
"""
Override of the parent method:
A generic HTTP request method, similar to `~requests.Session.request`
This is a low-level method not generally intended for use by astroquery
end-users.
This method wraps the _request functionality to include raise_for_status
Caching is defaulted to False but may be modified as needed
Also parameters that allow for file download through this method are removed
Parameters
----------
method : 'GET' or 'POST'
url : str
params : None or dict
data : None or dict
headers : None or dict
auth : None or dict
files : None or dict
stream : bool
See `~requests.request`
cache : bool
Default False. Use of bulit in _request caching
Returns
-------
response : `~requests.Response`
The response from the server.
"""
start_time = time.time()
response = super(MastClass, self)._request(method, url, params=params, data=data, headers=headers,
files=files, cache=cache, stream=stream, auth=auth)
if (time.time() - start_time) >= self.TIMEOUT:
raise TimeoutError("Timeout limit of {} exceeded.".format(self.TIMEOUT))
response.raise_for_status()
return [response]
def _request(self, method, url, params=None, data=None, headers=None,
files=None, stream=False, auth=None, retrieve_all=True):
"""
Override of the parent method:
A generic HTTP request method, similar to `~requests.Session.request`
This is a low-level method not generally intended for use by astroquery
end-users.
The main difference in this function is that it takes care of the long
polling requirements of the mashup server.
Thus the cache parameter of the parent method is hard coded to false
(the MAST server does it's own caching, no need to cache locally and it
interferes with follow requests after an 'Executing' response was returned.)
Also parameters that allow for file download through this method are removed
Parameters
----------
method : 'GET' or 'POST'
url : str
params : None or dict
data : None or dict
headers : None or dict
auth : None or dict
files : None or dict
stream : bool
See `~requests.request`
retrieve_all : bool
Default True. Retrieve all pages of data or just the one indicated in the params value.
Returns
-------
response : `~requests.Response`
The response from the server.
"""
start_time = time.time()
all_responses = []
total_pages = 1
cur_page = 0
while cur_page < total_pages:
status = "EXECUTING"
while status == "EXECUTING":
response = super(MastClass, self)._request(method, url, params=params, data=data,
headers=headers, files=files, cache=False,
stream=stream, auth=auth)
if (time.time() - start_time) >= self.TIMEOUT:
raise TimeoutError("Timeout limit of {} exceeded.".format(self.TIMEOUT))
# Raising error based on HTTP status if necessary
response.raise_for_status()
result = response.json()
if not result: # kind of hacky, but col_config service returns nothing if there is an error
status = "ERROR"
else:
status = result.get("status")
all_responses.append(response)
if (status != "COMPLETE") or (not retrieve_all):
break
paging = result.get("paging")
if paging is None:
break
total_pages = paging['pagesFiltered']
cur_page = paging['page']
data = data.replace("page%22%3A%20"+str(cur_page)+"%2C", "page%22%3A%20"+str(cur_page+1)+"%2C")
return all_responses
def _get_col_config(self, service, fetch_name=None):
"""
Gets the columnsConfig entry for given service and stores it in `self._column_configs`.
Parameters
----------
service : string
The service for which the columns config will be fetched.
fetch_name : string, optional
If the columns-config associated with the service has a different name,
use this argument. The default sets it to the same as service.
"""
if not fetch_name:
fetch_name = service
headers = {"User-Agent": self._session.headers["User-Agent"],
"Content-type": "application/x-www-form-urlencoded",
"Accept": "text/plain"}
response = self._request("POST", self._COLUMNS_CONFIG_URL,
data=("colConfigId="+fetch_name), headers=headers)
self._column_configs[service] = response[0].json()
more = False # for some catalogs this is not enough information
if "tess" in fetch_name.lower():
all_name = "Mast.Catalogs.All.Tic"
more = True
elif "dd." in fetch_name.lower():
all_name = "Mast.Catalogs.All.DiskDetective"
more = True
if more:
mashup_request = {'service': all_name, 'params': {}, 'format': 'extjs'}
req_string = _prepare_service_request_string(mashup_request)
response = self._request("POST", self._MAST_REQUEST_URL, data=req_string, headers=headers)
json_response = response[0].json()
self._column_configs[service].update(json_response['data']['Tables'][0]
['ExtendedProperties']['discreteHistogram'])
self._column_configs[service].update(json_response['data']['Tables'][0]
['ExtendedProperties']['continuousHistogram'])
for col, val in self._column_configs[service].items():
val.pop('hist', None) # don't want to save all this unecessary data
def _parse_result(self, responses, verbose=False):
"""
Parse the results of a list of `~requests.Response` objects and returns an `~astropy.table.Table` of results.
Parameters
----------
responses : list of `~requests.Response`
List of `~requests.Response` objects.
verbose : bool
(presently does nothing - there is no output with verbose set to
True or False)
Default False. Setting to True provides more extensive output.
Returns
-------
response : `~astropy.table.Table`
"""
result_list = []
catalogs_service = True if self._current_service in self._MAST_CATALOGS_SERVICES else False
if catalogs_service:
self._current_service = None # clearing current service
if not isinstance(responses, list):
responses = [responses]
for resp in responses:
result = resp.json()
result_table = _fabric_json_to_table(result)
result_list.append(result_table)
else:
# loading the columns config
col_config = None
if self._current_service:
col_config = self._column_configs.get(self._current_service)
self._current_service = None # clearing current service
for resp in responses:
result = resp.json()
# check for error message
if result['status'] == "ERROR":
raise RemoteServiceError(result.get('msg', "There was an error with your request."))
result_table = _mashup_json_to_table(result, col_config)
result_list.append(result_table)
all_results = vstack(result_list)
# Check for no results
if not all_results:
warnings.warn("Query returned no results.", NoResultsWarning)
return all_results
def logout(self): # pragma: no cover
"""
Log out of current MAST session.
"""
self._session.cookies.clear_session_cookies()
self._authenticated = False
@class_or_instance
def service_request_async(self, service, params, pagesize=None, page=None, rdm=42, **kwargs):
"""
Given a Mashup service and parameters, builds and excecutes a Mashup query.
See documentation `here <https://mast.stsci.edu/api/v0/class_mashup_1_1_mashup_request.html>`__
for information about how to build a Mashup request.
Parameters
----------
service : str
The Mashup service to query.
params : dict
JSON object containing service parameters.
pagesize : int, optional
Default None.
Can be used to override the default pagesize (set in configs) for this query only.
E.g. when using a slow internet connection.
page : int, optional
Default None.
Can be used to override the default behavior of all results being returned to obtain
a specific page of results.
rdm : int, optional
Default 42.
Can be used to specify random seed number.
**kwargs :
See MashupRequest properties
`here <https://mast.stsci.edu/api/v0/class_mashup_1_1_mashup_request.html>`__
for additional keyword arguments.
Returns
-------
response : list of `~requests.Response`
"""
# setting random seed to obtain the same results next time
np.random_seed(rdm)
# setting self._current_service
if service not in self._column_configs.keys():
fetch_name = kwargs.pop('fetch_name', None)
self._get_col_config(service, fetch_name)
self._current_service = service
# setting up pagination
if not pagesize:
pagesize = self.PAGESIZE
if not page:
page = 1
retrieve_all = True
else:
retrieve_all = False
headers = {"User-Agent": self._session.headers["User-Agent"],
"Content-type": "application/x-www-form-urlencoded",
"Accept": "text/plain"}
mashup_request = {'service': service,
'params': params,
'format': 'json',
'pagesize': pagesize,
'page': page}
for prop, value in kwargs.items():
mashup_request[prop] = value
req_string = _prepare_service_request_string(mashup_request)
response = self._request("POST", self._MAST_REQUEST_URL, data=req_string, headers=headers,
retrieve_all=retrieve_all)
return response
@class_or_instance
def catalogs_service_request_async(self, service, params, page_size=None, page=None, **kwargs):
"""
Given a MAST fabric service and parameters, builds and excecutes a fabric microservice catalog query.
See documentation `here <https://catalogs.mast.stsci.edu/docs/index.html>`__
for information about how to build a MAST catalogs microservice request.
Parameters
----------
service : str
The MAST catalogs service to query. Should be present in self._MAST_CATALOGS_SERVICES
params : dict
JSON object containing service parameters.
page_size : int, optional
Default None.
Can be used to override the default pagesize (set in configs) for this query only.
E.g. when using a slow internet connection.
page : int, optional
Default None.
Can be used to override the default behavior of all results being returned to obtain
a specific page of results.
**kwargs :
See Catalogs.MAST properties in documentation referenced above
Returns
-------
response : list of `~requests.Response`
"""
self._current_service = service.lower()
service_config = self._MAST_CATALOGS_SERVICES.get(service.lower())
service_url = service_config.get('path')
compiled_service_args = {}
# Gather URL specific parameters
for service_argument, default_value in service_config.get('args').items():
found_argument = params.pop(service_argument, None)
if found_argument is None:
found_argument = kwargs.pop(service_argument, default_value)
compiled_service_args[service_argument] = found_argument.lower()
request_url = self._MAST_CATALOGS_REQUEST_URL + service_url.format(**compiled_service_args)
headers = {
'User-Agent': self._session.headers['User-Agent'],
'Content-type': 'application/x-www-form-urlencoded',
'Accept': 'application/json'
}
# Params as a list of tuples to allow for multiple parameters added
catalogs_request = []
if not page:
page = params.pop('page', None)
if not page_size:
page_size = params.pop('page_size', None)
if page is not None:
catalogs_request.append(('page', page))
if page_size is not None:
catalogs_request.append(('pagesize', page_size))
# Decompose filters, sort
for prop, value in kwargs.items():
params[prop] = value
catalogs_request.extend(self._build_catalogs_params(params))
response = self._fabric_request('POST', request_url, data=catalogs_request, headers=headers)
return response
def _build_catalogs_params(self, params):
"""
Gathers parameters for Catalogs.MAST usage and translates to valid API syntax tuples
Parameters
----------
params: dict
A dict of parameters to convert into valid API syntax. Will omit the "format" parameter
Returns
-------
response : list(tuple)
List of tuples representing API syntax parameters
"""
catalog_params = []
for prop, value in params.items():
if prop == 'format':
# Ignore format changes
continue
elif prop == 'page_size':
catalog_params.extend(('pagesize', value))
elif prop == 'sort_by':
# Loop through each value if list
if isinstance(value, list):
for sort_item in value:
# Determine if tuple with sort direction
if isinstance(sort_item, tuple):
catalog_params.append(('sort_by', sort_item[1] + '.' + sort_item[0]))
else:
catalog_params.append(('sort_by', sort_item))
else:
# A single sort
# Determine if tuple with sort direction
if isinstance(value, tuple):
catalog_params.append(('sort_by', value[0] + '.' + value[1]))
else:
catalog_params.append(('sort_by', value))
elif prop == 'columns':
catalog_params.extend(tuple(('columns', col) for col in value))
else:
if isinstance(value, list):
# A composed list of multiple filters for a single column
# Extract each filter value in list
for filter_value in value:
# Determine if tuple with filter decorator
if isinstance(filter_value, tuple):
catalog_params.append((prop + '.' + filter_value[0], filter_value[1]))
else:
# Otherwise just append the value without a decorator
catalog_params.append((prop, filter_value))
else:
catalog_params.append((prop, value))
return catalog_params
def _check_catalogs_criteria_params(self, criteria):
"""
Tests a dict of passed criteria for Catalogs.MAST to ensure that at least one parameter is for a given criteria
Parameters
----------
criteria: dict
A dict of parameters to test for at least one criteria parameter
Returns
-------
response : boolean
Whether the passed dict has at least one criteria parameter
"""
criteria_check = False
non_criteria_params = ["columns", "sort_by", "page_size", "pagesize", "page"]
criteria_keys = criteria.keys()
for key in criteria_keys:
if key not in non_criteria_params:
criteria_check = True
break
return criteria_check
def resolve_object(self, objectname):
"""
Resolves an object name to a position on the sky.
Parameters
----------
objectname : str
Name of astronomical object to resolve.
Returns
-------
response : `~astropy.coordinates.SkyCoord`
The sky position of the given object.
"""
service = 'Mast.Name.Lookup'
params = {'input': objectname,
'format': 'json'}
response = self.service_request_async(service, params)
result = response[0].json()
if len(result['resolvedCoordinate']) == 0:
raise ResolverError("Could not resolve {} to a sky position.".format(objectname))
ra = result['resolvedCoordinate'][0]['ra']
dec = result['resolvedCoordinate'][0]['decl']
coordinates = coord.SkyCoord(ra, dec, unit="deg")
return coordinates
def _build_filter_set(self, column_config_name, service_name=None, **filters):
"""
Takes user input dictionary of filters and returns a filterlist that the Mashup can understand.
Parameters
----------
column_config_name : string
The service for which the columns config will be fetched.
service_name : string, optional
The service that will use the columns config, default is to be the same as column_config_name.
**filters :
Filters to apply. At least one filter must be supplied.
Valid criteria are coordinates, objectname, radius (as in `query_region` and `query_object`),
and all observation fields listed `here <https://mast.stsci.edu/api/v0/_c_a_o_mfields.html>`__.
The Column Name is the keyword, with the argument being one or more acceptable values for that parameter,
except for fields with a float datatype where the argument should be in the form [minVal, maxVal].
For example: filters=["FUV","NUV"],proposal_pi="Osten",t_max=[52264.4586,54452.8914]
Returns
-------
response : list(dict)
The mashup json filter object.
"""
if not service_name:
service_name = column_config_name
if not self._column_configs.get(service_name):
self._get_col_config(service_name, fetch_name=column_config_name)
caom_col_config = self._column_configs[service_name]
mashup_filters = []
for colname, value in filters.items():
# make sure value is a list-like thing
if np.isscalar(value,):
value = [value]
# Get the column type and separator
col_info = caom_col_config.get(colname)
if not col_info:
warnings.warn("Filter {} does not exist. This filter will be skipped.".format(colname), InputWarning)
continue
colType = "discrete"
if (col_info.get("vot.datatype", col_info.get("type")) in ("double", "float", "numeric")) \
or col_info.get("treatNumeric"):
colType = "continuous"
separator = col_info.get("separator")
free_text = None
# validate user input
if colType == "continuous":
if len(value) < 2:
warning_string = "{} is continuous, ".format(colname) + \
"and filters based on min and max values.\n" + \
"Not enough values provided, skipping..."
warnings.warn(warning_string, InputWarning)
continue
elif len(value) > 2:
warning_string = "{} is continuous, ".format(colname) + \
"and filters based on min and max values.\n" + \
"Too many values provided, the first two will be " + \
"assumed to be the min and max values."
warnings.warn(warning_string, InputWarning)
else: # coltype is discrete, all values should be represented as strings, even if numerical
value = [str(x) for x in value]
# check for wildcards
for i, val in enumerate(value):
if ('*' in val) or ('%' in val):
if free_text: # free_text is already set cannot set again
warning_string = ("Only one wildcarded value may be used per filter, "
"all others must be exact.\n"
"Skipping {}...".format(val))
warnings.warn(warning_string, InputWarning)
else:
free_text = val.replace('*', '%')
value.pop(i)
# craft mashup filter entry
entry = {}
entry["paramName"] = colname
if separator:
entry["separator"] = separator
if colType == "continuous":
entry["values"] = [{"min": value[0], "max":value[1]}]
else:
entry["values"] = value
if free_text:
entry["freeText"] = free_text
mashup_filters.append(entry)
return mashup_filters
def _get_columnsconfig_metadata(self, colconf_name):
"""
Given a columns config id make a table of the associated metadata properties.
Parameters
----------
colconf_name : str
The columns config idea to find metadata for (ex. Mast.Caom.Cone).
Returns
-------
response : `~astropy.table.Table`
The metadata table.
"""
headers = {"User-Agent": self._session.headers["User-Agent"],
"Content-type": "application/x-www-form-urlencoded",
"Accept": "text/plain"}
response = self._request("POST", self._COLUMNS_CONFIG_URL,
data=("colConfigId={}".format(colconf_name)), headers=headers)
column_dict = response[0].json()
meta_fields = ["Column Name", "Column Label", "Data Type", "Units", "Description", "Examples/Valid Values"]
names = []
labels = []
data_types = []
field_units = []
descriptions = []
examples = []
for colname in column_dict:
# skipping the _selected column (gets rmoved in return table)
if colname == "_selected_":
continue
field = column_dict[colname]
# skipping any columns that are removed
if field.get("remove", False):
continue
names.append(colname)
labels.append(field.get("text", colname))
# datatype is a little more complicated
d_type = _parse_type(field.get("type", ""))[0]
if not d_type:
d_type = _parse_type(field.get("vot.datatype", ""))[0]
data_types.append(d_type)
# units
units = field.get("unit", "")
if not units:
units = field.get("vot.unit", "")
field_units.append(units)
descriptions.append(field.get("vot.description", ""))
examples.append(field.get("example", ""))
meta_table = Table(names=meta_fields, data=[names, labels, data_types, field_units, descriptions, examples])
# Removing any empty columns
for colname in meta_table.colnames:
if (meta_table[colname] == "").all():
meta_table.remove_column(colname)
return meta_table
@async_to_sync
class ObservationsClass(MastClass):
"""
MAST Observations query class.
Class for querying MAST observational data.
"""
def __init__(self, *args, **kwargs):
super(ObservationsClass, self).__init__(*args, **kwargs)
self._boto3 = None
self._botocore = None
self._pubdata_bucket = "stpubdata"
def list_missions(self):
"""
Lists data missions archived by MAST and avaiable through `astroquery.mast`.
Returns
--------
response : list
List of available missions.
"""
# getting all the histogram information
service = "Mast.Caom.All"
params = {}
response = self.service_request_async(service, params, format='extjs')
json_response = response[0].json()
# getting the list of missions
hist_data = json_response['data']['Tables'][0]['Columns']
for facet in hist_data:
if facet['text'] == "obs_collection":
mission_info = facet['ExtendedProperties']['histObj']
missions = list(mission_info.keys())
missions.remove('hist')
return missions
def get_metadata(self, query_type):
"""
Returns metadata about the requested query type.
Parameters
----------
query_type : str
The query to get metadata for. Options are observations, and products.
Returns
--------
response : `~astropy.table.Table`
The metadata table.
"""
if query_type.lower() == "observations":
colconf_name = "Mast.Caom.Cone"
elif query_type.lower() == "products":
colconf_name = "Mast.Caom.Products"
else:
raise InvalidQueryError("Unknown query type.")
return self._get_columnsconfig_metadata(colconf_name)
@class_or_instance
def query_region_async(self, coordinates, radius=0.2*u.deg, pagesize=None, page=None):
"""
Given a sky position and radius, returns a list of MAST observations.
See column documentation `here <https://mast.stsci.edu/api/v0/_c_a_o_mfields.html>`__.
Parameters
----------
coordinates : str or `~astropy.coordinates` object
The target around which to search. It may be specified as a
string or as the appropriate `~astropy.coordinates` object.
radius : str or `~astropy.units.Quantity` object, optional
Default 0.2 degrees.
The string must be parsable by `~astropy.coordinates.Angle`. The
appropriate `~astropy.units.Quantity` object from
`~astropy.units` may also be used. Defaults to 0.2 deg.
pagesize : int, optional
Default None.
Can be used to override the default pagesize for (set in configs) this query only.
E.g. when using a slow internet connection.
page : int, optional
Default None.
Can be used to override the default behavior of all results being returned to
obtain a specific page of results.
Returns
-------
response : list of `~requests.Response`
"""
# Put coordinates and radius into consistant format
coordinates = commons.parse_coordinates(coordinates)
# if radius is just a number we assume degrees
if isinstance(radius, (int, float)):
radius = radius * u.deg
radius = coord.Angle(radius)
service = 'Mast.Caom.Cone'
params = {'ra': coordinates.ra.deg,
'dec': coordinates.dec.deg,
'radius': radius.deg}
return self.service_request_async(service, params, pagesize, page)
@class_or_instance
def query_object_async(self, objectname, radius=0.2*u.deg, pagesize=None, page=None):
"""
Given an object name, returns a list of MAST observations.
See column documentation `here <https://mast.stsci.edu/api/v0/_c_a_o_mfields.html>`__.
Parameters
----------
objectname : str
The name of the target around which to search.
radius : str or `~astropy.units.Quantity` object, optional
Default 0.2 degrees.
The string must be parsable by `~astropy.coordinates.Angle`.
The appropriate `~astropy.units.Quantity` object from
`~astropy.units` may also be used. Defaults to 0.2 deg.
pagesize : int, optional
Default None.
Can be used to override the default pagesize for (set in configs) this query only.
E.g. when using a slow internet connection.
page : int, optional
Defaulte None.
Can be used to override the default behavior of all results being returned
to obtain a specific page of results.
Returns
-------
response : list of `~requests.Response`
"""
coordinates = self.resolve_object(objectname)
return self.query_region_async(coordinates, radius, pagesize, page)
@class_or_instance
def query_criteria_async(self, pagesize=None, page=None, **criteria):
"""
Given an set of criteria, returns a list of MAST observations.
Valid criteria are returned by ``get_metadata("observations")``
Parameters
----------
pagesize : int, optional
Can be used to override the default pagesize.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to obtain
one sepcific page of results.
**criteria
Criteria to apply. At least one non-positional criteria must be supplied.
Valid criteria are coordinates, objectname, radius (as in `query_region` and `query_object`),
and all observation fields returned by the ``get_metadata("observations")``.
The Column Name is the keyword, with the argument being one or more acceptable values for that parameter,
except for fields with a float datatype where the argument should be in the form [minVal, maxVal].
For non-float type criteria wildcards maybe used (both * and % are considered wildcards), however
only one wildcarded value can be processed per criterion.
RA and Dec must be given in decimal degrees, and datetimes in MJD.
For example: filters=["FUV","NUV"],proposal_pi="Ost*",t_max=[52264.4586,54452.8914]
Returns
-------
response : list of `~requests.Response`
"""
# Seperating any position info from the rest of the filters
coordinates = criteria.pop('coordinates', None)
objectname = criteria.pop('objectname', None)
radius = criteria.pop('radius', 0.2*u.deg)
if ('obstype' in criteria) and ('intentType' in criteria):
warn_string = ("Cannot specify both obstype and intentType, "
"obstype is the deprecated version of intentType and will be ignored.")
warnings.warn(warn_string, InputWarning)
criteria.pop('obstype', None)
# Temporarily issuing warning about change in behavior
# continuing old behavior
# grabbing the observation type (science vs calibration)
obstype = criteria.pop('obstype', None)
if obstype:
warn_string = ("Criteria obstype argument will disappear in May 2019. "
"Criteria 'obstype' is now 'intentType', options are 'science' or 'calibration', "
"if intentType is not supplied all observations (science and calibration) are returned.")
warnings.warn(warn_string, AstropyDeprecationWarning)
if obstype == "science":
criteria["intentType"] = "science"
elif obstype == "cal":
criteria["intentType"] = "calibration"
# Build the mashup filter object and store it in the correct service_name entry
if coordinates or objectname:
mashup_filters = self._build_filter_set("Mast.Caom.Cone", "Mast.Caom.Filtered.Position", **criteria)
else:
mashup_filters = self._build_filter_set("Mast.Caom.Cone", "Mast.Caom.Filtered", **criteria)
if not mashup_filters:
raise InvalidQueryError("At least one non-positional criterion must be supplied.")
# handle position info (if any)
position = None
if objectname and coordinates:
raise InvalidQueryError("Only one of objectname and coordinates may be specified.")
if objectname:
coordinates = self.resolve_object(objectname)
if coordinates:
# Put coordinates and radius into consitant format
coordinates = commons.parse_coordinates(coordinates)
# if radius is just a number we assume degrees
if isinstance(radius, (int, float)):
radius = radius * u.deg
radius = coord.Angle(radius)
# build the coordinates string needed by Mast.Caom.Filtered.Position
position = ', '.join([str(x) for x in (coordinates.ra.deg, coordinates.dec.deg, radius.deg)])
# send query
if position:
service = "Mast.Caom.Filtered.Position"
params = {"columns": "*",
"filters": mashup_filters,
"position": position}
else:
service = "Mast.Caom.Filtered"
params = {"columns": "*",
"filters": mashup_filters}
return self.service_request_async(service, params)
def query_region_count(self, coordinates, radius=0.2*u.deg, pagesize=None, page=None):
"""
Given a sky position and radius, returns the number of MAST observations in that region.
Parameters
----------
coordinates : str or `~astropy.coordinates` object
The target around which to search. It may be specified as a
string or as the appropriate `~astropy.coordinates` object.
radius : str or `~astropy.units.Quantity` object, optional
The string must be parsable by `~astropy.coordinates.Angle`. The
appropriate `~astropy.units.Quantity` object from
`~astropy.units` may also be used. Defaults to 0.2 deg.
pagesize : int, optional
Can be used to override the default pagesize for.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to
obtain a specific page of results.
Returns
-------
response : int
"""
# build the coordinates string needed by Mast.Caom.Filtered.Position
coordinates = commons.parse_coordinates(coordinates)
# if radius is just a number we assume degrees
if isinstance(radius, (int, float)):
radius = radius * u.deg
radius = coord.Angle(radius)
# turn coordinates into the format
position = ', '.join([str(x) for x in (coordinates.ra.deg, coordinates.dec.deg, radius.deg)])
service = "Mast.Caom.Filtered.Position"
params = {"columns": "COUNT_BIG(*)",
"filters": [],
"position": position}
return int(self.service_request(service, params, pagesize, page)[0][0])
def query_object_count(self, objectname, radius=0.2*u.deg, pagesize=None, page=None):
"""
Given an object name, returns the number of MAST observations.
Parameters
----------
objectname : str
The name of the target around which to search.
radius : str or `~astropy.units.Quantity` object, optional
The string must be parsable by `~astropy.coordinates.Angle`. The
appropriate `~astropy.units.Quantity` object from
`~astropy.units` may also be used. Defaults to 0.2 deg.
pagesize : int, optional
Can be used to override the default pagesize.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to obtain
one sepcific page of results.
Returns
-------
response : int
"""
coordinates = self.resolve_object(objectname)
return self.query_region_count(coordinates, radius, pagesize, page)
def query_criteria_count(self, pagesize=None, page=None, **criteria):
"""
Given an set of filters, returns the number of MAST observations meeting those criteria.
Parameters
----------
pagesize : int, optional
Can be used to override the default pagesize.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to obtain
one sepcific page of results.
**criteria
Criteria to apply. At least one non-positional criterion must be supplied.
Valid criteria are coordinates, objectname, radius (as in `query_region` and `query_object`),
and all observation fields listed `here <https://mast.stsci.edu/api/v0/_c_a_o_mfields.html>`__.
The Column Name is the keyword, with the argument being one or more acceptable values for that parameter,
except for fields with a float datatype where the argument should be in the form [minVal, maxVal].
For non-float type criteria wildcards maybe used (both * and % are considered wildcards), however
only one wildcarded value can be processed per criterion.
RA and Dec must be given in decimal degrees, and datetimes in MJD.
For example: filters=["FUV","NUV"],proposal_pi="Ost*",t_max=[52264.4586,54452.8914]
Returns
-------
response : int
"""
# Seperating any position info from the rest of the filters
coordinates = criteria.pop('coordinates', None)
objectname = criteria.pop('objectname', None)
radius = criteria.pop('radius', 0.2*u.deg)
# grabbing the observation type (science vs calibration)
obstype = criteria.pop('obstype', 'science')
# Build the mashup filter object and store it in the correct service_name entry
if coordinates or objectname:
mashup_filters = self._build_filter_set("Mast.Caom.Cone", "Mast.Caom.Filtered.Position", **criteria)
else:
mashup_filters = self._build_filter_set("Mast.Caom.Cone", "Mast.Caom.Filtered", **criteria)
# handle position info (if any)
position = None
if objectname and coordinates:
raise InvalidQueryError("Only one of objectname and coordinates may be specified.")
if objectname:
coordinates = self.resolve_object(objectname)
if coordinates:
# Put coordinates and radius into consitant format
coordinates = commons.parse_coordinates(coordinates)
# if radius is just a number we assume degrees
if isinstance(radius, (int, float)):
radius = radius * u.deg
radius = coord.Angle(radius)
# build the coordinates string needed by Mast.Caom.Filtered.Position
position = ', '.join([str(x) for x in (coordinates.ra.deg, coordinates.dec.deg, radius.deg)])
# send query
if position:
service = "Mast.Caom.Filtered.Position"
params = {"columns": "COUNT_BIG(*)",
"filters": mashup_filters,
"obstype": obstype,
"position": position}
else:
service = "Mast.Caom.Filtered"
params = {"columns": "COUNT_BIG(*)",
"filters": mashup_filters,
"obstype": obstype}
return self.service_request(service, params)[0][0].astype(int)
@class_or_instance
def get_product_list_async(self, observations):
"""
Given a "Product Group Id" (column name obsid) returns a list of associated data products.
See column documentation `here <https://masttest.stsci.edu/api/v0/_productsfields.html>`__.
Parameters
----------
observations : str or `~astropy.table.Row` or list/Table of same
Row/Table of MAST query results (e.g. output from `query_object`)
or single/list of MAST Product Group Id(s) (obsid).
See description `here <https://masttest.stsci.edu/api/v0/_c_a_o_mfields.html>`__.
Returns
-------
response : list of `~requests.Response`
"""
# getting the obsid list
if type(observations) == Row:
observations = observations["obsid"]
if np.isscalar(observations):
observations = [observations]
if type(observations) == Table:
observations = observations['obsid']
service = 'Mast.Caom.Products'
params = {'obsid': ','.join(observations)}
return self.service_request_async(service, params)
def filter_products(self, products, mrp_only=False, extension=None, **filters):
"""
Takes an `~astropy.table.Table` of MAST observation data products and filters it based on given filters.
Parameters
----------
products : `~astropy.table.Table`
Table containing data products to be filtered.
mrp_only : bool, optional
Default False. When set to true only "Minimum Recommended Products" will be returned.
extension : string or array, optional
Default None. Option to filter by file extension.
**filters :
Filters to be applied. Valid filters are all products fields listed
`here <https://masttest.stsci.edu/api/v0/_productsfields.html>`__.
The column name is the keyword, with the argument being one or more acceptable values
for that parameter.
Filter behavior is AND between the filters and OR within a filter set.
For example: productType="SCIENCE",extension=["fits","jpg"]
Returns
-------
response : `~astropy.table.Table`
"""
filter_mask = np.full(len(products), True, dtype=bool)
# Applying the special filters (mrp_only and extension)
if mrp_only:
filter_mask &= (products['productGroupDescription'] == "Minimum Recommended Products")
if extension:
if type(extension) == str:
extension = [extension]
mask = np.full(len(products), False, dtype=bool)
for elt in extension:
mask |= [False if isinstance(x, np.ma.core.MaskedConstant) else x.endswith(elt)
for x in products["productFilename"]]
filter_mask &= mask
# Applying the rest of the filters
for colname, vals in filters.items():
if type(vals) == str:
vals = [vals]
mask = np.full(len(products), False, dtype=bool)
for elt in vals:
mask |= (products[colname] == elt)
filter_mask &= mask
return products[np.where(filter_mask)]
def _download_curl_script(self, products, out_dir):
"""
Takes an `~astropy.table.Table` of data products and downloads a curl script to pull the datafiles.
Parameters
----------
products : `~astropy.table.Table`
Table containing products to be included in the curl script.
out_dir : str
Directory in which the curl script will be saved.
Returns
-------
response : `~astropy.table.Table`
"""
url_list = [("uri", url) for url in products['dataURI']]
download_file = "mastDownload_" + time.strftime("%Y%m%d%H%M%S")
local_path = os.path.join(out_dir.rstrip('/'), download_file + ".sh")
response = self._download_file(self._MAST_BUNDLE_URL + ".sh", local_path, data=url_list, method="POST")
status = "COMPLETE"
msg = None
if not os.path.isfile(local_path):
status = "ERROR"
msg = "Curl could not be downloaded"
manifest = Table({'Local Path': [local_path],
'Status': [status],
'Message': [msg]})
return manifest
@deprecated(since="v0.3.9", alternative="enable_cloud_dataset")
def enable_s3_hst_dataset(self):
return self.enable_cloud_dataset()
def enable_cloud_dataset(self, provider="AWS", profile=None, verbose=True):
"""
Enable downloading public files from S3 instead of MAST.
Requires the boto3 library to function.
Parameters
----------
provider : str
Which cloud data provider to use. We may in the future support multiple providers,
though at the moment this argument is ignored.
profile : str
Profile to use to identify yourself to the cloud provider (usually in ~/.aws/config).
verbose : bool
Default True.
Logger to display extra info and warning.
"""
import boto3
import botocore
if profile is not None:
self._boto3 = boto3.Session(profile_name=profile)
else:
self._boto3 = boto3
self._botocore = botocore
if verbose:
log.info("Using the S3 STScI public dataset")
log.warning("Your AWS account will be charged for access to the S3 bucket")
log.info("See Request Pricing in https://aws.amazon.com/s3/pricing/ for details")
log.info("If you have not configured boto3, follow the instructions here: "
"https://boto3.readthedocs.io/en/latest/guide/configuration.html")
@deprecated(since="v0.3.9", alternative="disable_cloud_dataset")
def disable_s3_hst_dataset(self):
return self.disable_cloud_dataset()
def disable_cloud_dataset(self):
"""
Disables downloading public files from S3 instead of MAST
"""
self._boto3 = None
self._botocore = None
@deprecated(since="v0.3.9", alternative="get_cloud_uris")
def get_hst_s3_uris(self, data_products, include_bucket=True, full_url=False):
return self.get_cloud_uris(data_products, include_bucket, full_url)
def get_cloud_uris(self, data_products, include_bucket=True, full_url=False):
"""
Takes an `~astropy.table.Table` of data products and returns the associated cloud data uris.
Parameters
----------
data_products : `~astropy.table.Table`
Table containing products to be converted into cloud data uris.
include_bucket : bool
When either to include the cloud bucket prefix in the result or not.
full_url : bool
Return a HTTP fetchable url instead of a uri.
Returns
-------
response : list
List of URIs generated from the data products, list way contain entries that are None
if data_products includes products not found in the cloud.
"""
return [self.get_cloud_uri(data_product, include_bucket, full_url) for data_product in data_products]
@deprecated(since="v0.3.9", alternative="get_cloud_uri")
def get_hst_s3_uri(self, data_product, include_bucket=True, full_url=False):
return self.get_cloud_uri(data_product, include_bucket, full_url)
def get_cloud_uri(self, data_product, include_bucket=True, full_url=False):
"""
For a given data product, returns the associated cloud URI.
If the product is from a mission that does not support cloud access an
exception is raised. If the mission is supported but the product
cannot be found in the cloud, the returned path is None.
Parameters
----------
data_product : `~astropy.table.Row`
Product to be converted into cloud data uri.
include_bucket : bool
When either to include the cloud bucket prefix in the result or not.
full_url : bool
Return a HTTP fetchable url instead of a uri.
Returns
-------
response : str or None
Cloud URI generated from the data product. If the product cannot be
found in the cloud, None is returned.
"""
if self._boto3 is None:
raise AtrributeError("Must enable s3 dataset before attempting to query the s3 information")
# This is a cheap operation and does not perform any actual work yet
s3_client = self._boto3.client('s3')
paths = fpl.paths(data_product)
if paths is None:
raise Exception("Unsupported mission {}".format(data_product['obs_collection']))
for path in paths:
try:
s3_client.head_object(Bucket=self._pubdata_bucket, Key=path, RequestPayer='requester')
if include_bucket:
path = "s3://{}/{}".format(self._pubdata_bucket, path)
elif full_url:
path = "http://s3.amazonaws.com/{}/{}".format(self._pubdata_bucket, path)
return path
except self._botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] != "404":
raise
warnings.warn("Unable to locate file {}.".format(data_product['productFilename']), NoResultsWarning)
return None
def _download_from_cloud(self, data_product, local_path, cache=True):
"""
Takes a data product in the form of an `~astropy.table.Row` and downloads it from the cloud into
the given directory.
Parameters
----------
data_product : `~astropy.table.Row`
Product to download.
local_path : str
Directory in which files will be downloaded.
cache : bool
Default is True. If file is found on disc it will not be downloaded again.
"""
# The following is a mishmash of BaseQuery._download_file and s3 access through boto
self._pubdata_bucket = 'stpubdata'
# This is a cheap operation and does not perform any actual work yet
s3 = self._boto3.resource('s3')
s3_client = self._boto3.client('s3')
bkt = s3.Bucket(self._pubdata_bucket)
bucket_path = self.get_cloud_uri(data_product, False)
info_lookup = s3_client.head_object(Bucket=self._pubdata_bucket, Key=bucket_path, RequestPayer='requester')
# Unfortunately, we can't use the reported file size in the reported product. STScI's backing
# archive database (CAOM) is frequently out of date and in many cases omits the required information.
# length = data_product["size"]
# Instead we ask the webserver (in this case S3) what the expected content length is and use that.
length = info_lookup["ContentLength"]
if cache and os.path.exists(local_path):
if length is not None:
statinfo = os.stat(local_path)
if statinfo.st_size != length:
log.warning("Found cached file {0} with size {1} that is "
"different from expected size {2}"
.format(local_path,
statinfo.st_size,
length))
else:
log.info("Found cached file {0} with expected size {1}."
.format(local_path, statinfo.st_size))
return
with ProgressBarOrSpinner(length, ('Downloading URL s3://{0}/{1} to {2} ...'.format(
self._pubdata_bucket, bucket_path, local_path))) as pb:
# Bytes read tracks how much data has been received so far
# This variable will be updated in multiple threads below
global bytes_read
bytes_read = 0
progress_lock = threading.Lock()
def progress_callback(numbytes):
# Boto3 calls this from multiple threads pulling the data from S3
global bytes_read
# This callback can be called in multiple threads
# Access to updating the console needs to be locked
with progress_lock:
bytes_read += numbytes
pb.update(bytes_read)
bkt.download_file(bucket_path, local_path, ExtraArgs={"RequestPayer": "requester"},
Callback=progress_callback)
def _download_files(self, products, base_dir, cache=True, cloud_only=False,):
"""
Takes an `~astropy.table.Table` of data products and downloads them into the directory given by base_dir.
Parameters
----------
products : `~astropy.table.Table`
Table containing products to be downloaded.
base_dir : str
Directory in which files will be downloaded.
cache : bool
Default is True. If file is found on disk it will not be downloaded again.
cloud_only : bool, optional
Default False. If set to True and cloud data access is enabled (see `enable_cloud_dataset`)
files that are not found in the cloud will be skipped rather than downloaded from MAST
as is the default behavior. If cloud access is not enables this argument as no affect.
Returns
-------
response : `~astropy.table.Table`
"""
manifest_array = []
for data_product in products:
local_path = os.path.join(base_dir, data_product['obs_collection'], data_product['obs_id'])
data_url = self._MAST_DOWNLOAD_URL + "?uri=" + data_product["dataURI"]
if not os.path.exists(local_path):
os.makedirs(local_path)
local_path = os.path.join(local_path, data_product['productFilename'])
status = "COMPLETE"
msg = None
url = None
try:
if self._boto3 is not None and fpl.has_path(data_product):
try:
self._download_from_cloud(data_product, local_path, cache)
except Exception as ex:
log.exception("Error pulling from S3 bucket: {}".format(ex))
if cloud_only:
log.warn("Skipping file...")
local_path = ""
status = "SKIPPED"
else:
log.warn("Falling back to mast download...")
self._download_file(data_url, local_path, cache=cache, head_safe=True, continuation=False)
else:
self._download_file(data_url, local_path, cache=cache, head_safe=True, continuation=False)
# check if file exists also this is where would perform md5,
# and also check the filesize if the database reliably reported file sizes
if (not os.path.isfile(local_path)) and (status != "SKIPPED"):
status = "ERROR"
msg = "File was not downloaded"
url = data_url
except HTTPError as err:
status = "ERROR"
msg = "HTTPError: {0}".format(err)
url = data_url
manifest_array.append([local_path, status, msg, url])
manifest = Table(rows=manifest_array, names=('Local Path', 'Status', 'Message', "URL"))
return manifest
def download_products(self, products, download_dir=None,
cache=True, curl_flag=False, mrp_only=False, cloud_only=False, **filters):
"""
Download data products.
If cloud access is enabled, files will be downloaded from the cloud if possible.
Parameters
----------
products : str, list, `~astropy.table.Table`
Either a single or list of obsids (as can be given to `get_product_list`),
or a Table of products (as is returned by `get_product_list`)
download_dir : str, optional
Optional. Directory to download files to. Defaults to current directory.
cache : bool, optional
Default is True. If file is found on disc it will not be downloaded again.
Note: has no affect when downloading curl script.
curl_flag : bool, optional
Default is False. If true instead of downloading files directly, a curl script
will be downloaded that can be used to download the data files at a later time.
mrp_only : bool, optional
Default False. When set to true only "Minimum Recommended Products" will be returned.
cloud_only : bool, optional
Default False. If set to True and cloud data access is enabled (see `enable_cloud_dataset`)
files that are not found in the cloud will be skipped rather than downloaded from MAST
as is the default behavior. If cloud access is not enables this argument as no affect.
**filters :
Filters to be applied. Valid filters are all products fields returned by
``get_metadata("products")`` and 'extension' which is the desired file extension.
The Column Name (or 'extension') is the keyword, with the argument being one or
more acceptable values for that parameter.
Filter behavior is AND between the filters and OR within a filter set.
For example: productType="SCIENCE",extension=["fits","jpg"]
Returns
-------
response : `~astropy.table.Table`
The manifest of files downloaded, or status of files on disk if curl option chosen.
"""
# If the products list is not already a table of products we need to
# get the products and filter them appropriately
if type(products) != Table:
if type(products) == str:
products = [products]
# collect list of products
product_lists = []
for oid in products:
product_lists.append(self.get_product_list(oid))
products = vstack(product_lists)
# apply filters
products = self.filter_products(products, mrp_only, **filters)
if not len(products):
warnings.warn("No products to download.", NoResultsWarning)
return
# set up the download directory and paths
if not download_dir:
download_dir = '.'
if curl_flag: # don't want to download the files now, just the curl script
manifest = self._download_curl_script(products, download_dir)
else:
base_dir = download_dir.rstrip('/') + "/mastDownload"
manifest = self._download_files(products, base_dir, cache, cloud_only)
return manifest
@async_to_sync
class CatalogsClass(MastClass):
"""
MAST catalog query class.
Class for querying MAST catalog data.
"""
def __init__(self):
super(CatalogsClass, self).__init__()
self.catalog_limit = None
def _parse_result(self, response, verbose=False):
results_table = super(CatalogsClass, self)._parse_result(response, verbose)
if len(results_table) == self.catalog_limit:
warnings.warn("Maximum catalog results returned, may not include all sources within radius.",
MaxResultsWarning)
return results_table
@class_or_instance
def query_region_async(self, coordinates, radius=0.2*u.deg, catalog="Hsc",
version=None, pagesize=None, page=None, **kwargs):
"""
Given a sky position and radius, returns a list of catalog entries.
See column documentation for specific catalogs `here <https://mast.stsci.edu/api/v0/pages.htmll>`__.
Parameters
----------
coordinates : str or `~astropy.coordinates` object
The target around which to search. It may be specified as a
string or as the appropriate `~astropy.coordinates` object.
radius : str or `~astropy.units.Quantity` object, optional
Default 0.2 degrees.
The string must be parsable by `~astropy.coordinates.Angle`. The
appropriate `~astropy.units.Quantity` object from
`~astropy.units` may also be used. Defaults to 0.2 deg.
catalog : str, optional
Default HSC.
The catalog to be queried.
version : int, optional
Version number for catalogs that have versions. Default is highest version.
pagesize : int, optional
Default None.
Can be used to override the default pagesize for (set in configs) this query only.
E.g. when using a slow internet connection.
page : int, optional
Default None.
Can be used to override the default behavior of all results being returned to
obtain a specific page of results.
**kwargs
Other catalog-specific keyword args.
These can be found in the (service documentation)[https://mast.stsci.edu/api/v0/_services.html]
for specific catalogs. For example one can specify the magtype for an HSC search.
Returns
-------
response : list of `~requests.Response`
"""
catalogs_service = False
# Put coordinates and radius into consistant format
coordinates = commons.parse_coordinates(coordinates)
# if radius is just a number we assume degrees
if isinstance(radius, (int, float)):
radius = radius * u.deg
radius = coord.Angle(radius)
# Figuring out the service
if catalog.lower() in self._MAST_CATALOGS_SERVICES:
catalogs_service = True
service = catalog
elif catalog.lower() == "hsc":
if version == 2:
service = "Mast.Hsc.Db.v2"
else:
if version not in (3, None):
warnings.warn("Invalid HSC version number, defaulting to v3.", InputWarning)
service = "Mast.Hsc.Db.v3"
self.catalog_limit = kwargs.get('nr', 50000)
elif catalog.lower() == "galex":
service = "Mast.Galex.Catalog"
self.catalog_limit = kwargs.get('maxrecords', 50000)
elif catalog.lower() == "gaia":
if version == 1:
service = "Mast.Catalogs.GaiaDR1.Cone"
else:
if version not in (2, None):
warnings.warn("Invalid Gaia version number, defaulting to DR2.", InputWarning)
service = "Mast.Catalogs.GaiaDR2.Cone"
else:
service = "Mast.Catalogs." + catalog + ".Cone"
self.catalog_limit = None
# basic params
params = {'ra': coordinates.ra.deg,
'dec': coordinates.dec.deg,
'radius': radius.deg}
if not catalogs_service:
# Hsc specific parameters (can be overridden by user)
params['nr'] = 50000
params['ni'] = 1
params['magtype'] = 1
# galex specific parameters (can be overridden by user)
params['maxrecords'] = 50000
# adding additional parameters
for prop, value in kwargs.items():
params[prop] = value
if catalogs_service:
return self.catalogs_service_request_async(service, params, pagesize, page)
return self.service_request_async(service, params, pagesize, page)
@class_or_instance
def query_object_async(self, objectname, radius=0.2*u.deg, catalog="Hsc",
pagesize=None, page=None, version=None, **kwargs):
"""
Given an object name, returns a list of catalog entries.
See column documentation for specific catalogs `here <https://mast.stsci.edu/api/v0/pages.html>`__.
Parameters
----------
objectname : str
The name of the target around which to search.
radius : str or `~astropy.units.Quantity` object, optional
Default 0.2 degrees.
The string must be parsable by `~astropy.coordinates.Angle`.
The appropriate `~astropy.units.Quantity` object from
`~astropy.units` may also be used. Defaults to 0.2 deg.
catalog : str, optional
Default HSC.
The catalog to be queried.
pagesize : int, optional
Default None.
Can be used to override the default pagesize for (set in configs) this query only.
E.g. when using a slow internet connection.
page : int, optional
Defaulte None.
Can be used to override the default behavior of all results being returned
to obtain a specific page of results.
version : int, optional
Version number for catalogs that have versions. Default is highest version.
**kwargs
Catalog-specific keyword args.
These can be found in the `service documentation <https://mast.stsci.edu/api/v0/_services.html>`__.
for specific catalogs. For example one can specify the magtype for an HSC search.
Returns
-------
response : list of `~requests.Response`
"""
coordinates = self.resolve_object(objectname)
return self.query_region_async(coordinates, radius, catalog,
version=version, pagesize=pagesize, page=page, **kwargs)
@class_or_instance
def query_criteria_async(self, catalog, pagesize=None, page=None, **criteria):
"""
Given an set of filters, returns a list of catalog entries.
See column documentation for specific catalogs `here <https://mast.stsci.edu/api/v0/pages.htmll>`__.
Parameters
----------
pagesize : int, optional
Can be used to override the default pagesize.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to obtain
one specific page of results.
**criteria
Criteria to apply. At least one non-positional criteria must be supplied.
Valid criteria are coordinates, objectname, radius (as in `query_region` and `query_object`),
and all fields listed in the column documentation for the catalog being queried.
The Column Name is the keyword, with the argument being one or more acceptable values for that parameter,
except for fields with a float datatype where the argument should be in the form [minVal, maxVal].
For non-float type criteria wildcards maybe used (both * and % are considered wildcards), however
only one wildcarded value can be processed per criterion.
RA and Dec must be given in decimal degrees, and datetimes in MJD.
For example: filters=["FUV","NUV"],proposal_pi="Ost*",t_max=[52264.4586,54452.8914]
For catalogs available through Catalogs.MAST (PanSTARRS), the Column Name is the keyword, and the argument
should be either an acceptable value for that parameter, or a list consisting values, or tuples of
decorator, value pairs (decorator, value). In addition, columns may be used to select the return columns,
consisting of a list of column names. Results may also be sorted through the query with the parameter
sort_by composed of either a single Column Name to sort ASC, or a list of Column Nmaes to sort ASC or
tuples of Column Name and Direction (ASC, DESC) to indicate sort order (Column Name, DESC).
Detailed information of Catalogs.MAST criteria usage can
be found `here <https://catalogs.mast.stsci.edu/docs/index.html>`__.
Returns
-------
response : list of `~requests.Response`
"""
catalogs_service = False
# Seperating any position info from the rest of the filters
coordinates = criteria.pop('coordinates', None)
objectname = criteria.pop('objectname', None)
radius = criteria.pop('radius', 0.2*u.deg)
mashup_filters = None
# Build the mashup filter object
if catalog.lower() in self._MAST_CATALOGS_SERVICES:
catalogs_service = True
service = catalog
mashup_filters = self._check_catalogs_criteria_params(criteria)
elif catalog.lower() == "tic":
service = "Mast.Catalogs.Filtered.Tic"
if coordinates or objectname:
service += ".Position"
service += ".Rows" # Using the rowstore version of the query for speed
mashup_filters = self._build_filter_set("Mast.Catalogs.Tess.Cone", service, **criteria)
elif catalog.lower() == "ctl":
service = "Mast.Catalogs.Filtered.Ctl"
if coordinates or objectname:
service += ".Position"
service += ".Rows" # Using the rowstore version of the query for speed
mashup_filters = self._build_filter_set("Mast.Catalogs.Tess.Cone", service, **criteria)
elif catalog.lower() == "diskdetective":
service = "Mast.Catalogs.Filtered.DiskDetective"
if coordinates or objectname:
service += ".Position"
mashup_filters = self._build_filter_set("Mast.Catalogs.Dd.Cone", service, **criteria)
else:
raise InvalidQueryError("Criteria query not available for {}".format(catalog))
if not mashup_filters:
raise InvalidQueryError("At least one non-positional criterion must be supplied.")
if objectname and coordinates:
raise InvalidQueryError("Only one of objectname and coordinates may be specified.")
if objectname:
coordinates = self.resolve_object(objectname)
if coordinates:
# Put coordinates and radius into consitant format
coordinates = commons.parse_coordinates(coordinates)
# if radius is just a number we assume degrees
if isinstance(radius, (int, float)):
radius = radius * u.deg
radius = coord.Angle(radius)
# build query
params = {}
if coordinates:
params["ra"] = coordinates.ra.deg
params["dec"] = coordinates.dec.deg
params["radius"] = radius.deg
if not catalogs_service:
params["filters"] = mashup_filters
# TIC and CTL need columns specified
if catalog.lower() in ("tic", "ctl"):
params["columns"] = "*"
if catalogs_service:
# For catalogs service append criteria to main parameters
for prop, value in criteria.items():
params[prop] = value
if catalogs_service:
return self.catalogs_service_request_async(service, params, page_size=pagesize, page=page)
return self.service_request_async(service, params, pagesize=pagesize, page=page)
@class_or_instance
def query_hsc_matchid_async(self, match, version=3, pagesize=None, page=None):
"""
Returns all the matches for a given Hubble Source Catalog MatchID.
Parameters
----------
match : int or `~astropy.table.Row`
The matchID or HSC entry to return matches for.
version : int, optional
The HSC version to match against. Default is v3.
pagesize : int, optional
Can be used to override the default pagesize.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to obtain
one sepcific page of results.
Returns
--------
response : list of `~requests.Response`
"""
if isinstance(match, Row):
match = match["MatchID"]
match = str(match) # np.int64 gives json serializer problems, so strigify right here
if version == 2:
service = "Mast.HscMatches.Db.v2"
else:
if version not in (3, None):
warnings.warn("Invalid HSC version number, defaulting to v3.", InputWarning)
service = "Mast.HscMatches.Db.v3"
params = {"input": match}
return self.service_request_async(service, params, pagesize, page)
@class_or_instance
def get_hsc_spectra_async(self, pagesize=None, page=None):
"""
Returns all Hubble Source Catalog spectra.
Parameters
----------
pagesize : int, optional
Can be used to override the default pagesize.
E.g. when using a slow internet connection.
page : int, optional
Can be used to override the default behavior of all results being returned to obtain
one sepcific page of results.
Returns
--------
response : list of `~requests.Response`
"""
service = "Mast.HscSpectra.Db.All"
params = {}
return self.service_request_async(service, params, pagesize, page)
def download_hsc_spectra(self, spectra, download_dir=None, cache=True, curl_flag=False):
"""
Download one or more Hubble Source Catalog spectra.
Parameters
----------
spectra : `~astropy.table.Table` or `~astropy.table.Row`
One or more HSC spectra to be downloaded.
download_dir : str, optional
Specify the base directory to download spectra into.
Spectra will be saved in the subdirectory download_dir/mastDownload/HSC.
If download_dir is not specified the base directory will be '.'.
cache : bool, optional
Default is True. If file is found on disc it will not be downloaded again.
Note: has no affect when downloading curl script.
curl_flag : bool, optional
Default is False. If true instead of downloading files directly, a curl script
will be downloaded that can be used to download the data files at a later time.
Returns
--------
response : list of `~requests.Response`
"""
# if spectra is not a Table, put it in a list
if isinstance(spectra, Row):
spectra = [spectra]
# set up the download directory and paths
if not download_dir:
download_dir = '.'
if curl_flag: # don't want to download the files now, just the curl script
download_file = "mastDownload_" + time.strftime("%Y%m%d%H%M%S")
url_list = []
path_list = []
for spec in spectra:
if spec['SpectrumType'] < 2:
url_list.append('https://hla.stsci.edu/cgi-bin/getdata.cgi?config=ops&dataset={0}'
.format(spec['DatasetName']))
else:
url_list.append('https://hla.stsci.edu/cgi-bin/ecfproxy?file_id={0}'
.format(spec['DatasetName']) + '.fits')
path_list.append(download_file + "/HSC/" + spec['DatasetName'] + '.fits')
description_list = [""]*len(spectra)
producttype_list = ['spectrum']*len(spectra)
service = "Mast.Bundle.Request"
params = {"urlList": ",".join(url_list),
"filename": download_file,
"pathList": ",".join(path_list),
"descriptionList": list(description_list),
"productTypeList": list(producttype_list),
"extension": 'curl'}
response = self.service_request_async(service, params)
bundler_response = response[0].json()
local_path = os.path.join(download_dir, "{}.sh".format(download_file))
self._download_file(bundler_response['url'], local_path, head_safe=True, continuation=False)
status = "COMPLETE"
msg = None
url = None
if not os.path.isfile(local_path):
status = "ERROR"
msg = "Curl could not be downloaded"
url = bundler_response['url']
else:
missing_files = [x for x in bundler_response['statusList'].keys()
if bundler_response['statusList'][x] != 'COMPLETE']
if len(missing_files):
msg = "{} files could not be added to the curl script".format(len(missing_files))
url = ",".join(missing_files)
manifest = Table({'Local Path': [local_path],
'Status': [status],
'Message': [msg],
"URL": [url]})
else:
base_dir = download_dir.rstrip('/') + "/mastDownload/HSC"
if not os.path.exists(base_dir):
os.makedirs(base_dir)
manifest_array = []
for spec in spectra:
if spec['SpectrumType'] < 2:
data_url = 'https://hla.stsci.edu/cgi-bin/getdata.cgi?config=ops&dataset=' \
+ spec['DatasetName']
else:
data_url = 'https://hla.stsci.edu/cgi-bin/ecfproxy?file_id=' \
+ spec['DatasetName'] + '.fits'
local_path = os.path.join(base_dir, "{}.fits".format(spec['DatasetName']))
status = "COMPLETE"
msg = None
url = None
try:
self._download_file(data_url, local_path, cache=cache, head_safe=True)
# check file size also this is where would perform md5
if not os.path.isfile(local_path):
status = "ERROR"
msg = "File was not downloaded"
url = data_url
except HTTPError as err:
status = "ERROR"
msg = "HTTPError: {0}".format(err)
url = data_url
manifest_array.append([local_path, status, msg, url])
manifest = Table(rows=manifest_array, names=('Local Path', 'Status', 'Message', "URL"))
return manifest
Observations = ObservationsClass()
Catalogs = CatalogsClass()
Mast = MastClass()
|
import os
import time
from urllib.parse import urlparse
import requests
from auth import get_auth
def get_resource_list(url):
"""
Returns a list of HC resources specified by the url basename (such as .../articles.json)
:param url: A full endpoint url, such as 'https://support.zendesk.com/api/v2/help_center/articles.json'
:return: List of resources, or False if the request failed.
"""
session = requests.Session()
session.auth = get_auth()
o = urlparse(url)
resource = os.path.splitext(os.path.basename(o.path))[0] # e.g., 'articles'
record_list = {resource: []}
while url:
response = session.get(url)
if response.status_code == 429:
print('Rate limited! Please wait.')
time.sleep(int(response.headers['retry-after']))
response = session.get(url)
if response.status_code != 200:
print('Error with status code {}'.format(response.status_code))
exit()
data = response.json()
if data[resource]: # guard against empty record list
record_list[resource].extend(data[resource])
url = data['next_page']
return record_list[resource]
def get_resource(url):
"""
Returns a single HC resource
:param url: A full endpoint url, such as 'https://support.zendesk.com/api/v2/help_center/articles/2342572.json'
:return: Dict of a resource, or False if the request failed.
"""
resource = None
response = requests.get(url, auth=get_auth())
if response.status_code == 429:
print('Rate limited! Please wait.')
time.sleep(int(response.headers['retry-after']))
response = requests.get(url, auth=get_auth())
if response.status_code != 200:
print('Failed to get record with error {}:'.format(response.status_code))
print(response.text)
return False
for k, v in response.json().items():
resource = v
if type(resource) is dict:
return resource
return None
def post_resource(url, data, status=201):
"""
:param url:
:param data:
:param status: HTTP status. Normally 201 but some POST requests return 200
:return: Python data, or False if the request failed.
"""
resource = None
headers = {'Content-Type': 'application/json'}
response = requests.post(url, json=data, auth=get_auth(), headers=headers)
if response.status_code == 429:
print('Rate limited! Please wait.')
time.sleep(int(response.headers['retry-after']))
response = requests.post(url, json=data, auth=get_auth(), headers=headers)
if response.status_code != status:
print('Failed to create record with error {}:'.format(response.status_code))
print(response.text)
return False
for k, v in response.json().items():
resource = v
if type(resource) is dict:
return resource
return None
def put_resource(url, data):
"""
:param url:
:param data:
:return: Python data, or False if the request failed.
"""
resource = None
headers = {'Content-Type': 'application/json'}
response = requests.put(url, json=data, auth=get_auth(), headers=headers)
if response.status_code == 429:
print('Rate limited! Please wait.')
time.sleep(int(response.headers['retry-after']))
response = requests.post(url, json=data, auth=get_auth(), headers=headers)
if response.status_code != 200:
print('Failed to update record with error {}:'.format(response.status_code))
print(response.text)
return False
for k, v in response.json().items():
resource = v
if type(resource) is dict:
return resource
return None
def delete_resource(url):
"""
Runs a DELETE request on any Delete endpoint in the Zendesk API
:param url: A full endpoint url, such as 'https://support.zendesk.com/api/v2/help_center/articles/2342572.json'
:return: If successful, a 204 status code. If not, None
"""
response = requests.delete(url, auth=get_auth())
if response.status_code == 429:
print('Rate limited! Please wait.')
time.sleep(int(response.headers['retry-after']))
response = requests.delete(url, auth=get_auth())
if response.status_code != 204:
print('Failed to delete record with error {}'.format(response.status_code))
print(response.text)
return False
return None
|
<gh_stars>0
from os.path import exists, dirname, join, abspath
from datetime import datetime
import pandas as pd
from two_thinning.environment import run_strategy_multiple_times
from two_thinning.strategies.always_accept_strategy import AlwaysAcceptStrategy
from two_thinning.strategies.local_reward_optimiser_strategy import LocalRewardOptimiserStrategy
from two_thinning.strategies.full_knowledge_DQN_strategy import FullKnowledgeDQNStrategy
from two_thinning.strategies.dp_strategy import DPStrategy
from two_thinning.strategies.random_strategy import RandomStrategy
from two_thinning.strategies.the_threshold_strategy import TheThresholdStrategy
from two_thinning.strategies.mean_thinning_strategy import MeanThinningStrategy
from two_thinning.full_knowledge.RL.DQN.constants import MAX_LOAD_POTENTIAL
from evaluation.two_thinning.hyperparameters import get_dqn_hyperparameters, get_threshold_hyperparameters
NMS = ((5, 5), (5, 10), (5, 25), (20, 20), (20, 60), (20, 400), (50, 50), (50, 200), (50, 2500))
STRATEGIES = ("always_accept", "local_reward_optimiser", "mean_thinning", "dp", "dqn", "threshold")
RUNS = 500
RE_TRAIN_DQN = 1
PRINT_BEHAVIOUR = False
def REWARD_FUN(loads):
return -max(loads)
def compare_strategies(nms=NMS, runs=RUNS, strategies=STRATEGIES, reward_fun=REWARD_FUN,
print_behaviour=PRINT_BEHAVIOUR, re_train_dqn=RE_TRAIN_DQN):
for n, m in nms:
for strategy_name in strategies:
print(f"n={n}, m={m}, strategy={strategy_name} started.")
if strategy_name == "dqn":
hyperparameters = get_dqn_hyperparameters(n=n, m=m)
scores = []
for _ in range(re_train_dqn):
save_path = join((dirname(dirname(abspath(__file__)))), "training_progression",
f'{str(datetime.now().strftime("%Y_%m_%d %H_%M_%S_%f"))}_{n}_{m}')
strategy = FullKnowledgeDQNStrategy(n=n, m=m, use_pre_trained=False, save_path=save_path,
**hyperparameters)
curr_scores = run_strategy_multiple_times(n=n, m=m, runs=runs // re_train_dqn, strategy=strategy,
reward=reward_fun,
print_behaviour=print_behaviour)
scores.extend(curr_scores)
else:
if strategy_name == "always_accept":
strategy = AlwaysAcceptStrategy(n=n, m=m)
elif strategy_name == "random":
strategy = RandomStrategy(n=n, m=m)
elif strategy_name == "mean_thinning":
strategy = MeanThinningStrategy(n=n, m=m)
elif strategy_name == "local_reward_optimiser":
strategy = LocalRewardOptimiserStrategy(n=n, m=m, reward_fun=reward_fun,
potential_fun=MAX_LOAD_POTENTIAL)
elif strategy_name == "threshold":
hyperparameters = get_threshold_hyperparameters(n=n, m=m)
strategy = TheThresholdStrategy(n=n, m=m, reward_fun=reward_fun, **hyperparameters)
elif strategy_name == "dp":
if n > 50 or m > 70: # these are out of the feasible range
continue
strategy = DPStrategy(n=n, m=m, reward_fun=reward_fun)
else:
raise Exception("No such strategy is known, check spelling!")
scores = run_strategy_multiple_times(n=n, m=m, runs=runs, strategy=strategy, reward=reward_fun,
print_behaviour=print_behaviour)
df = pd.DataFrame(data=scores, columns=["score"])
output_path = f'data/{n}_{m}_{strategy_name}.csv'
df.to_csv(output_path, mode='a', index=False, header=not exists(output_path))
if __name__ == "__main__":
compare_strategies()
|
import numpy as np
arr = np.arange(0, 11)
print(arr)
# [ 0 1 2 3 4 5 6 7 8 9 10]
print(arr[8])
# 8
print(arr[1:5])
# [1 2 3 4]
# change elements
arr[0:5] = 100
print(arr)
# [100 100 100 100 100 5 6 7 8 9 10]
# reset original array
arr = np.arange(0, 11)
# slicing
slice1 = arr[0:6]
print(slice1)
# [0 1 2 3 4 5]
# change all elements
slice1[:] = 99
print(slice1)
# [99 99 99 99 99 99]
# changes occured in the original array
print(arr)
# [99 99 99 99 99 99 6 7 8 9 10]
# to prevent this use .copy() function
arr_copy = arr.copy()
arr_copy[:3] = 11
print(arr)
# [99 99 99 99 99 99 6 7 8 9 10]
print(arr_copy)
# [11 11 11 99 99 99 6 7 8 9 10]
# indexing 2d array
arr_2d = np.array(([5,10,15],[20,25,30], [35,40,45]))
print(arr_2d)
'''
[[ 5 10 15]
[20 25 30]
[35 40 45]]
'''
print(arr_2d[1])
# [20 25 30]
print(arr_2d[1][0])
# 20
print(arr_2d[1,0])
# 20
# 2d array slicing
print(arr_2d[:2,1:])
'''
[[10 15]
[25 30]]
'''
new_arr = np.zeros((10,10))
print(new_arr)
'''
[[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]]
'''
new_arr_len = new_arr.shape[0]
print(new_arr_len)
# 10
for i in range(new_arr_len):
new_arr[i] = 1
print(new_arr)
'''
[[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]]
'''
for i in range(new_arr_len):
new_arr[i] = i
print(new_arr)
'''
[[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]
[ 3. 3. 3. 3. 3. 3. 3. 3. 3. 3.]
[ 4. 4. 4. 4. 4. 4. 4. 4. 4. 4.]
[ 5. 5. 5. 5. 5. 5. 5. 5. 5. 5.]
[ 6. 6. 6. 6. 6. 6. 6. 6. 6. 6.]
[ 7. 7. 7. 7. 7. 7. 7. 7. 7. 7.]
[ 8. 8. 8. 8. 8. 8. 8. 8. 8. 8.]
[ 9. 9. 9. 9. 9. 9. 9. 9. 9. 9.]]
'''
# fancy indexing
print(new_arr[[2,5]])
'''
[[ 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]
[ 5. 5. 5. 5. 5. 5. 5. 5. 5. 5.]]
'''
print(new_arr[[8,1,4]])
'''
[[ 8. 8. 8. 8. 8. 8. 8. 8. 8. 8.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 4. 4. 4. 4. 4. 4. 4. 4. 4. 4.]]
'''
|
<gh_stars>10-100
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import argparse
import datetime
from functools import partial
import json
import traceback
import imlib as im
import numpy as np
import pylib
import tensorflow as tf
import tensorflow.contrib.distributions as tfd
# from tensorflow_probability import distributions as tfd
import tflib as tl
import utils
import fid
# ==============================================================================
# = param =
# ==============================================================================
parser = argparse.ArgumentParser()
parser.add_argument('--dataset', dest='dataset_name', default='mnist', choices=['mnist', 'cifar10', 'celeba'])
parser.add_argument('--model', dest='model_name', default='conv_mnist', choices=['conv_mnist', 'conv_32', 'conv_64'])
parser.add_argument('--epoch', dest='epoch', type=int, default=600)
parser.add_argument('--batch_size', dest='batch_size', type=int, default=64)
parser.add_argument('--lr', dest='lr', type=float, default=0.1, help='learning rate')
parser.add_argument('--bn', dest='use_bn', type=lambda v: v.lower() in ('true', 'yes'), default=False,
help='use batchnorm or not')
parser.add_argument('--z_dim', dest='z_dim', type=int, default=32, help='dimension of latent space')
parser.add_argument('--init_steps', dest='init_steps', type=int, default=3000, help='initialization steps')
parser.add_argument('--zn_rec', dest='zn_rec_coeff', type=float, default=6e-2,
help='coefficient of latent reconstruction loss (z~N)')
parser.add_argument('--zh_rec', dest='zh_rec_coeff', type=float, default=0,
help='coefficient of latent reconstruction loss (z~H)')
parser.add_argument('--vrec', dest='vrec_coeff', type=float, default=2e-2,
help='coefficient of VAE reconstruction loss')
parser.add_argument('--vkld', dest='vkld_coeff', type=float, default=2e-2, help='coefficient of VAE KLD loss')
parser.add_argument('--nll', dest='nll_coeff', type=float, default=0, help='coefficient of NLL loss')
parser.add_argument('--experiment_name', dest='experiment_name',
default=datetime.datetime.now().strftime("%I:%M%p on %B %d, %Y"))
args = parser.parse_args()
dataset_name = args.dataset_name
model_name = args.model_name
epoch = args.epoch
batch_size = args.batch_size
lr = args.lr
use_bn = args.use_bn
z_dim = args.z_dim
init_steps = args.init_steps
zn_rec_coeff = args.zn_rec_coeff
zh_rec_coeff = args.zh_rec_coeff
vrec_coeff = args.vrec_coeff
vkld_coeff = args.vkld_coeff
nll_coeff = args.nll_coeff
experiment_name = args.experiment_name
pylib.mkdir('./output/%s' % experiment_name)
with open('./output/%s/setting.txt' % experiment_name, 'w') as f:
f.write(json.dumps(vars(args), indent=4, separators=(',', ':')))
inception_path = fid.check_or_download_inception('../data/inception_model/')
fid_stats_dict = {'mnist': '../data/fid/fid_stats_mnist_train.npz',
'cifar10': '../data/fid/fid_stats_cifar10_train.npz',
'celeba': '../data/fid/fid_stats_celeba.npz'}
fid_stats_path = fid_stats_dict[dataset_name] if dataset_name in fid_stats_dict else None
# dataset and models
Dataset, img_shape, get_imgs = utils.get_dataset(dataset_name)
dataset = Dataset(batch_size=batch_size)
# TODO: use a separate validation set
dataset_val = Dataset(batch_size=100)
Enc, Dec = utils.get_models(model_name)
Enc = partial(Enc, z_dim=z_dim, use_bn=use_bn, sigma=True)
Dec = partial(Dec, channels=img_shape[2], use_bn=use_bn)
# ==============================================================================
# = graph =
# ==============================================================================
def enc_dec(img, is_training=True):
# encode
z_mu, z_log_sigma_sq = Enc(img, is_training=is_training)
# decode
img_rec = Dec(z_mu, is_training=is_training)
return z_mu, z_log_sigma_sq, img_rec
def dec_enc(z, is_training=True, no_enc_grad=False):
# decode
img = Dec(z, is_training=is_training)
# encode
z_rec, _ = Enc(img, is_training=is_training)
if no_enc_grad:
z_rec -= Enc(tf.stop_gradient(img), is_training=is_training)[0] - tf.stop_gradient(z_rec)
return z_rec
# input
img = tf.placeholder(tf.float32, [None] + img_shape)
normal_dist = tfd.MultivariateNormalDiag(scale_diag=np.ones([z_dim], dtype=np.float32))
# encode & decode
z_mu, z_log_sigma_sq, img_rec = enc_dec(img)
zn_targ, zh_targ = normal_dist.sample(batch_size), tf.stop_gradient(z_mu)
zn_rec, zh_rec = dec_enc(zn_targ), dec_enc(zh_targ)
# loss
def log_det_jacobian(z):
z = tf.stop_gradient(z)
delta = tf.random_normal([batch_size, z_dim]) * (tf.exp(0.5 * z_log_sigma_sq) + 1e-2)
epsilon = tf.stop_gradient(tf.sqrt(tf.reduce_sum(tf.square(delta), 1, keepdims=True)))
return z_dim / 2 * tf.log(tf.reduce_sum(tf.square(1 / epsilon * (dec_enc(z + delta, no_enc_grad=True) -
dec_enc(z, no_enc_grad=True))), 1))
img_rec_loss = tf.losses.mean_squared_error(img, img_rec)
zn_rec_loss, zh_rec_loss = tf.losses.mean_squared_error(zn_targ, zn_rec), tf.losses.mean_squared_error(zh_targ, zh_rec)
vrec_loss = tf.reduce_mean(log_det_jacobian(z_mu)) / z_dim
vkld_loss = -tf.reduce_mean(0.5 * (1 + z_log_sigma_sq - z_mu ** 2 - tf.exp(z_log_sigma_sq)))
nll_enc_loss = -tf.reduce_mean(normal_dist.log_prob(z_mu)) / z_dim
global_step = tf.get_variable('global_steps', [], initializer=tf.zeros_initializer, trainable=False)
zn_rec_coeff = tf.cond(global_step > init_steps, lambda: zn_rec_coeff, lambda: 0.)
vrec_coeff = tf.cond(global_step > init_steps, lambda: vrec_coeff, lambda: 0.)
vkld_coeff = tf.cond(global_step > init_steps, lambda: vkld_coeff, lambda: 0.)
nll_coeff = tf.cond(global_step > init_steps, lambda: nll_coeff, lambda: 0.)
enc_loss = img_rec_loss + zn_rec_coeff * zn_rec_loss + \
vrec_coeff * vrec_loss + vkld_coeff * vkld_loss + nll_coeff * nll_enc_loss
if zh_rec_coeff > 0:
zh_rec_coeff = tf.cond(global_step > init_steps, lambda: zh_rec_coeff, lambda: 0.)
enc_loss += zh_rec_coeff * zh_rec_loss
dec_loss = img_rec_loss + vrec_coeff * vrec_loss
# otpim
enc_vars = []
dec_vars = []
for var in tf.trainable_variables():
if var.name.startswith('Enc'):
enc_vars.append(var)
elif var.name.startswith('Dec'):
dec_vars.append(var)
optimizer = tf.train.MomentumOptimizer(learning_rate=lr, momentum=0.9)
enc_gvs = optimizer.compute_gradients(enc_loss, enc_vars)
dec_gvs = optimizer.compute_gradients(dec_loss, dec_vars)
train_op = optimizer.apply_gradients(enc_gvs + dec_gvs, global_step=global_step)
# summary
summary = tl.summary({img_rec_loss: 'img_rec_loss',
zn_rec_loss: 'zn_rec_loss', zh_rec_loss: 'zh_rec_loss',
vrec_loss: 'vrec_loss', vkld_loss: 'vkld_loss', nll_enc_loss: 'nll_enc_loss'})
# sample
# TODO: compute running averages for different input batches respectively
z_intp_sample, _, img_rec_sample = enc_dec(img, is_training=True)
img_sample = Dec(normal_dist.sample([100]), is_training=True)
fid_sample = Dec(normal_dist.sample([1000]), is_training=True)
if dataset_name == 'mnist':
fid_sample = tf.image.grayscale_to_rgb(fid_sample)
z_intp_split, img_split = tf.split(z_intp_sample, 2), tf.split(img, 2)
img_intp_sample = [Dec((1 - i) * z_intp_split[0] + i * z_intp_split[1], is_training=True) for i in np.linspace(0, 1, 9)]
img_intp_sample = [img_split[0]] + img_intp_sample + [img_split[1]]
img_intp_sample = tf.concat(img_intp_sample, 2)
# ==============================================================================
# = train =
# ==============================================================================
# session
sess = tl.session()
# saver
saver = tf.train.Saver(max_to_keep=1)
# summary writer
summary_writer = tf.summary.FileWriter('./output/%s/summaries' % experiment_name, sess.graph)
# initialization
ckpt_dir = './output/%s/checkpoints' % experiment_name
pylib.mkdir(ckpt_dir)
try:
tl.load_checkpoint(ckpt_dir, sess)
except:
sess.run(tf.global_variables_initializer())
if fid_stats_path:
with np.load(fid_stats_path) as stats:
mu_real, sigma_real = stats['mu'][:], stats['sigma'][:]
fid.create_inception_graph(inception_path)
# train
try:
img_ipt_sample = get_imgs(dataset_val.get_next())
z_ipt_sample = np.random.normal(size=[100, z_dim])
it = -1
for ep in range(epoch):
dataset.reset()
it_per_epoch = it_in_epoch if it != -1 else -1
it_in_epoch = 0
for batch in dataset:
it += 1
it_in_epoch += 1
# batch data
img_ipt = get_imgs(batch)
# train & add summary
if (it + 1) % 100 == 0:
summary_opt, _ = sess.run([summary, train_op], feed_dict={img: img_ipt})
summary_writer.add_summary(summary_opt, it)
else:
sess.run([train_op], feed_dict={img: img_ipt})
# display
if (it + 1) % 100 == 0:
print("Epoch: (%3d) (%5d/%5d)" % (ep, it_in_epoch, it_per_epoch))
# sample
if (it + 1) % 2000 == 0:
save_dir = './output/%s/sample_training' % experiment_name
pylib.mkdir(save_dir)
img_rec_opt_sample, img_intp_opt_sample = sess.run([img_rec_sample, img_intp_sample],
feed_dict={img: img_ipt_sample})
img_rec_opt_sample, img_intp_opt_sample = img_rec_opt_sample.squeeze(), img_intp_opt_sample.squeeze()
# ipt_rec = np.concatenate((img_ipt_sample, img_rec_opt_sample), axis=2).squeeze()
img_opt_sample = sess.run(img_sample).squeeze()
# im.imwrite(im.immerge(ipt_rec, padding=img_shape[0] // 8),
# '%s/Epoch_(%d)_(%dof%d)_img_rec.png' % (save_dir, ep, it_in_epoch, it_per_epoch))
im.imwrite(im.immerge(img_intp_opt_sample, n_col=1, padding=0),
'%s/Epoch_(%d)_(%dof%d)_img_intp.png' % (save_dir, ep, it_in_epoch, it_per_epoch))
im.imwrite(im.immerge(img_opt_sample),
'%s/Epoch_(%d)_(%dof%d)_img_sample.png' % (save_dir, ep, it_in_epoch, it_per_epoch))
if fid_stats_path:
try:
mu_gen, sigma_gen = fid.calculate_activation_statistics(im.im2uint(
np.concatenate([sess.run(fid_sample).squeeze() for _ in range(5)], 0)), sess,
batch_size=100)
fid_value = fid.calculate_frechet_distance(mu_gen, sigma_gen, mu_real, sigma_real)
except:
fid_value = -1.
fid_summary = tf.Summary()
fid_summary.value.add(tag='FID', simple_value=fid_value)
summary_writer.add_summary(fid_summary, it)
print("FID: %s" % fid_value)
save_path = saver.save(sess, '%s/Epoch_%d.ckpt' % (ckpt_dir, ep))
print('Model is saved in file: %s' % save_path)
except:
traceback.print_exc()
finally:
sess.close()
|
# -*- coding:utf-8 -*-
# ###########################
# File Name: hdataset.py
# Author: geekinglcq
# Mail: <EMAIL>
# Created Time: 2020-12-28 20:17:47
# ###########################
import pandas as pd
import os
import logging
from collections import defaultdict
from torch.utils.data import DataLoader, Dataset
from .enum_type import FeatureSource as FS
from .enum_type import item_type_dict
from .dataset import DataSet, SubSet
class HDataSet(DataSet):
"""
Dataset used for heterogenous items
"""
def __init__(self, config, restore_path=None):
self.config = config
self._init_setting()
if restore_path is None:
self._load_feats()
else:
# TODO
pass
self._preprocessing()
def _load_feats(self):
self.user_feat = self._load_meta_feats(self.config["user_feat_path"],
FS.USER, "user_id")
self.item_feat = self._load_item_feats(self.config["item_feat_path"],
FS.ITEM)
self.inter_feat = pd.read_csv(self.config["inter_feat_path"]).sample(
frac=1, random_state=28)
mask = None
if len(self.types) < 3:
for item_type, item_feat in self.item_feat.items():
new_mask = self.inter_feat[self.iid_field].isin(
item_feat[self.iid_field])
if mask is not None:
mask = mask | new_mask
else:
mask = new_mask
self.inter_feat = self.inter_feat[mask]
self.h_inter_feat = {}
self.user_num = len(self.user_feat)
self.item_num = sum([len(i) for i in self.item_feat.values()])
self.item_nums = {k: len(v) for k, v in self.item_feat.items()}
print(f'user num: {self.user_num}')
print(f'item num: {self.item_num}')
print(f'item nums: {self.item_nums}')
def _preprocessing(self):
self._normalize()
if len(self.types) < 3:
self._reID(self.iid_field)
self._reID(self.uid_field)
def _load_item_feats(self, paths, source):
item_feat = {}
for item_type, item_path in paths.items():
if item_type not in self.types:
continue
if os.path.isfile(item_path):
feat = pd.read_csv(item_path)
item_feat[item_type] = feat
else:
raise ValueError("Dataset file not fountd.")
return item_feat
def _init_setting(self):
self.logger = logging.getLogger()
self.name = self.config['name']
print(self.config)
self.uid_field = self.config["USER_ID_FIELD"]
self.iid_field = self.config["ITEM_ID_FIELD"]
self.label_field = self.config["LABEL_FIELD"]
self.itype_field = self.config["TYPE_FIELD"]
self.types = self.config["type"]
self.field2type = {}
self.field2source = {}
self.field2id_token = defaultdict(dict)
self.field2token_id = defaultdict(dict)
self.user_feat_fields = []
self.item_feat_fields = defaultdict(list)
for feat_name, feat_value in self.config['feat'].items():
source = feat_value['source']
self.field2type[feat_name] = feat_value['type']
self.field2source[feat_name] = feat_value['source']
if source == 'user' and feat_name != self.uid_field:
self.user_feat_fields.append(feat_name)
if source.startswith('item') and feat_name != self.iid_field:
item_type = source.split("_")[1]
if item_type in self.types:
self.item_feat_fields[item_type].append(feat_name)
def num(self, field):
if field == self.uid_field:
return self.user_num
if field == self.iid_field:
return self.item_num
if field not in self.field2type:
raise ValueError('field {} not in dataset'.format(field))
# if field not in self.field2token_id:
# raise ValueError('field {} is not token type'.format(field))
if len(self.field2token_id[field]) == 0:
if field in self.user_feat_fields:
return len(self.user_feat[field].unique())
else:
for item_type, item_feat_fields in self.item_feat_fields.items(
):
if field in item_feat_fields:
return len(self.item_feat[item_type][field].unique())
return len(self.field2token_id[field])
def _reID(self, field):
"""
Re-ID the token-type feature, save the id map in self.field2token_id
"""
self.logger.info(f'ReID field {field}.')
ftype = self.field2type.get(field)
assert ftype == 'token'
source = self.field2source.get(field)
if type(source) is str and source.startswith("item_"):
item_type = source.split("_")[1]
dataframe = self.item_feat[item_type]
elif source is FS.ITEM_ID or source == "item":
dataframe = pd.concat(list(self.item_feat.values()), join='inner')
elif source == 'user' or source is FS.USER_ID:
dataframe = self.user_feat
else:
dataframe = self.inter_feat
id_map = {v: k for k, v in enumerate(dataframe[field].unique())}
self.field2token_id[field].update(id_map)
dataframe[field] = dataframe[field].map(id_map)
if source in ['item', 'user', FS.ITEM_ID, FS.USER_ID]:
if field in self.inter_feat:
self.inter_feat[field] = self.inter_feat[field].map(id_map)
for item_type, item_feat in self.item_feat.items():
if field in item_feat:
item_feat[field] = item_feat[field].map(id_map)
def join(self, df):
"""
Join user/item features to interactions.
"""
if self.user_feat is not None and self.uid_field in df:
df = pd.merge(df,
self.user_feat,
on=self.uid_field,
how='left',
suffixes=('_inter', '_user'))
if self.item_feat is not None and self.iid_field in df:
for item_type, item_feat in self.item_feat.items():
df = pd.merge(df,
item_feat,
on=self.iid_field,
how='left',
suffixes=(f'_{item_type}', '_inter'))
type_c = [i for i in df.columns if i.startswith(self.itype_field)]
df[self.itype_field] = df[type_c].agg(sum, axis=1)
return df
def join_interaction(self):
self.inter_feat = self.join(self.inter_feat)
if 'sample' in self.config:
sample_ratio = self.config['sample']
sampled = []
for kind in self.types:
ratio = sample_ratio.get(kind, 1.0)
kind_id = item_type_dict[kind]
# preverse the data for val & test
new_df = self.inter_feat[self.inter_feat['type'] ==
kind_id].sample(frac=ratio * 0.7 +
0.3,
random_state=16)
print(kind, kind_id, ratio, new_df.shape)
sampled.append(new_df)
self.inter_feat = pd.concat(sampled, ignore_index=True)
self.inter_feat = self.inter_feat.sample(frac=1.).reset_index(
drop=True)
def train_val_test_split(self,
ratios=[0.7, 0.2, 0.1],
group_by=None,
**kwargs):
assert len(ratios) == 3
if 'sample' in self.config:
train, val, test = self.split_by_ratio_sampled(
ratios, create_new_dataset=False)
else:
train, val, test = self.split_by_ratio(ratios,
group_by=group_by,
create_new_dataset=False)
user_fs = self.user_feat_fields
item_fs = self.item_feat_fields
type_field = self.itype_field
self.train_inter_subset = {}
self.val_inter_subset = {}
self.test_inter_subset = {}
for item_type in self.types:
item_type_id = item_type_dict[item_type]
self.train_inter_subset[item_type] = SubSet(
train[train[type_field] == item_type_id], self.uid_field,
self.iid_field, self.itype_field, self.label_field, user_fs,
item_fs[item_type])
self.val_inter_subset[item_type] = SubSet(
val[val[type_field] == item_type_id], self.uid_field,
self.iid_field, self.itype_field, self.label_field, user_fs,
item_fs[item_type])
self.test_inter_subset[item_type] = SubSet(
test[test[type_field] == item_type_id], self.uid_field,
self.iid_field, self.itype_field, self.label_field, user_fs,
item_fs[item_type])
self.all_inter_feat = self.inter_feat
self.logger.info(
"Replace interaction features with train interaction fatures.")
self.logger.info(
"Interaction features are stored in self.all_inter_feat")
self.inter_feat = train
def init_data_loader(self, batch_size=256, num_workers=1):
self.train_data_loader = {}
self.val_data_loader = {}
self.test_data_loader = {}
for item_type in self.types:
self.train_data_loader[item_type] = DataLoader(
self.train_inter_subset[item_type],
batch_size=batch_size,
# pin_memory=True,
num_workers=num_workers)
self.val_data_loader[item_type] = DataLoader(
self.val_inter_subset[item_type],
batch_size=batch_size,
num_workers=num_workers)
self.test_data_loader[item_type] = DataLoader(
self.test_inter_subset[item_type],
batch_size=batch_size,
num_workers=num_workers)
class HSubSet(Dataset):
def __init__(self, dataframes, uid_field, iid_field, label_field,
u_feat_fields, i_feat_fields):
self.types = dataframes.keys()
self.dfs = dataframes
self.uid = uid_field
self.iid = iid_field
self.label = label_field
def __len__(self):
return min([len(df.index) for df in self.dfs])
|
import copy
import os
import yaml
from utils import write_conf
def get_expelled_srv_conf(uuid):
return {uuid: "expelled"}
def get_srv_conf(uuid, rpl_uuid, uri=None, disabled=False):
return {
uuid: {
'disabled': disabled,
'replicaset_uuid': rpl_uuid,
'uri': uri if uri is not None else '%s-uri' % uuid,
}
}
def get_rpl_conf(uuid, leaders, alias=None):
return {
uuid: {
'alias': alias if alias is not None else 'unnamed',
'master': leaders,
'roles': {'vshard-storage': True},
'vshard_group': 'default',
'weight': 1
},
}
def get_topology_conf(instances, replicasets):
conf = {
'failover': False,
'replicasets': {},
'servers': {},
}
for instance in instances:
conf['servers'].update(instance)
for replicaset in replicasets:
conf['replicasets'].update(replicaset)
return conf
def get_one_file_conf(instances, replicasets):
return {
'topology': get_topology_conf(instances, replicasets)
}
class ClusterwideConfig:
def __init__(self, conf, instance_uuid=None, replicaset_uuid=None, instance_uri=None, one_file=False):
self.conf = conf
self.instance_uuid = instance_uuid
self.replicaset_uuid = replicaset_uuid
self.instance_uri = instance_uri
self.one_file = one_file
def write_instances_topology_conf(data_dir, app_name, conf, instances, one_file=False):
conf_paths = []
for instance in instances:
work_dir = os.path.join(data_dir, '%s.%s' % (app_name, instance))
os.makedirs(work_dir, exist_ok=True)
if one_file:
conf_path = os.path.join(work_dir, 'config.yml')
else:
conf_dir = os.path.join(work_dir, 'config')
os.makedirs(conf_dir, exist_ok=True)
conf_path = os.path.join(conf_dir, 'topology.yml')
conf_paths.append(conf_path)
write_conf(conf_path, conf)
return conf_paths
def assert_conf_changed(conf_paths, other_app_conf_paths, old_conf, new_conf):
for conf_path in conf_paths:
assert os.path.exists(conf_path)
with open(conf_path, 'r') as f:
conf = yaml.safe_load(f.read())
assert conf == new_conf
# check backup
backup_conf_path = '%s.bak' % conf_path
assert os.path.exists(backup_conf_path)
with open(backup_conf_path, 'r') as f:
conf = yaml.safe_load(f.read())
assert conf == old_conf
if other_app_conf_paths is None:
return
# check that other app config wasn't changed
for conf_path in other_app_conf_paths:
assert os.path.exists(conf_path)
with open(conf_path, 'r') as f:
conf = yaml.safe_load(f.read())
assert conf == old_conf
# check backup
backup_conf_path = '%s.bak' % conf_path
assert not os.path.exists(backup_conf_path)
def assert_conf_not_changed(conf_paths, old_conf):
for conf_path in conf_paths:
assert os.path.exists(conf_path)
with open(conf_path, 'r') as f:
conf = yaml.safe_load(f.read())
assert conf == old_conf
# check backup
backup_conf_path = '%s.bak' % conf_path
assert not os.path.exists(backup_conf_path)
def get_conf_with_new_uri(conf, instance_uuid, new_uri):
new_conf = copy.deepcopy(conf)
if new_conf.get('servers') is not None:
new_conf['servers'][instance_uuid]['uri'] = new_uri
else:
new_conf['topology']['servers'][instance_uuid]['uri'] = new_uri
return new_conf
def get_conf_with_removed_instance(conf, instance_uuid):
new_conf = copy.deepcopy(conf)
if new_conf.get('topology') is None:
topology_conf = new_conf
else:
topology_conf = new_conf['topology']
while True:
if topology_conf['servers'][instance_uuid] == 'expelled':
break
replicaset_uuid = topology_conf['servers'][instance_uuid]['replicaset_uuid']
# if there is no replicaset instance belong to - break
if replicaset_uuid not in topology_conf['replicasets']:
break
# if instance not in replicaset leaders - break
new_leaders = topology_conf['replicasets'][replicaset_uuid]['master']
if isinstance(new_leaders, list) and instance_uuid not in new_leaders:
break
if isinstance(new_leaders, str) and instance_uuid != new_leaders:
break
rpl_other_instances = [
uuid for uuid, instance_conf
in topology_conf['servers'].items()
if instance_conf.get('replicaset_uuid') == replicaset_uuid
and uuid != instance_uuid
]
rpl_other_instances.sort()
# if instance was the last leader in replicaset, check if there are
# other instances of this replicaset that aren't in the leaders list
if isinstance(new_leaders, str):
if len(rpl_other_instances) > 0:
topology_conf['replicasets'][replicaset_uuid]['master'] = rpl_other_instances[0]
else:
del topology_conf['replicasets'][replicaset_uuid]
if isinstance(new_leaders, list):
new_leaders.remove(instance_uuid)
if len(new_leaders) == 0:
if len(rpl_other_instances) > 0:
new_leaders.append(rpl_other_instances[0])
# leaders list is still empty - remove this replicaset
if len(new_leaders) == 0:
del topology_conf['replicasets'][replicaset_uuid]
break
del topology_conf['servers'][instance_uuid]
return new_conf
def get_conf_with_new_leader(conf, replicaset_uuid, instance_uuid):
new_conf = copy.deepcopy(conf)
new_leaders = new_conf['replicasets'][replicaset_uuid]['master']
if instance_uuid in new_leaders:
new_leaders.remove(instance_uuid)
new_leaders.insert(0, instance_uuid)
return conf
|
import time
from copy import deepcopy
import torch
import torch.optim as optim
from torch.utils.data.sampler import BatchSampler, SubsetRandomSampler
from torch.distributions import kl_divergence
import numpy as np
from rl.algos import PPO
from rl.policies.actor import GaussianMLP_Actor
from rl.policies.critic import GaussianMLP_Critic
from rl.envs.normalize import get_normalization_params, PreNormalizer
from rl.envs.wrappers import SymmetricEnv
import functools
# TODO:
# env.mirror() vs env.matrix?
# TODO: use magic to make this reuse more code (callbacks etc?)
class MirrorPPO(PPO):
def update(self, policy, old_policy, optimizer,
critic, critic_optimizer,
observations, actions, returns, advantages,
env_fn
):
env = env_fn()
mirror_observation = env.mirror_observation
if env.clock_based:
mirror_observation = env.mirror_clock_observation
mirror_action = env.mirror_action
minibatch_size = self.minibatch_size or advantages.numel()
for _ in range(self.epochs):
losses = []
sampler = BatchSampler(
SubsetRandomSampler(range(advantages.numel())),
minibatch_size,
drop_last=True
)
for indices in sampler:
indices = torch.LongTensor(indices)
obs_batch = observations[indices]
# obs_batch = torch.cat(
# [obs_batch,
# obs_batch @ torch.Tensor(env.obs_symmetry_matrix)]
# ).detach()
action_batch = actions[indices]
# action_batch = torch.cat(
# [action_batch,
# action_batch @ torch.Tensor(env.action_symmetry_matrix)]
# ).detach()
return_batch = returns[indices]
# return_batch = torch.cat(
# [return_batch,
# return_batch]
# ).detach()
advantage_batch = advantages[indices]
# advantage_batch = torch.cat(
# [advantage_batch,
# advantage_batch]
# ).detach()
values, pdf = policy.evaluate(obs_batch)
# TODO, move this outside loop?
with torch.no_grad():
_, old_pdf = old_policy.evaluate(obs_batch)
old_log_probs = old_pdf.log_prob(action_batch).sum(-1, keepdim=True)
log_probs = pdf.log_prob(action_batch).sum(-1, keepdim=True)
ratio = (log_probs - old_log_probs).exp()
cpi_loss = ratio * advantage_batch
clip_loss = ratio.clamp(1.0 - self.clip, 1.0 + self.clip) * advantage_batch
actor_loss = -torch.min(cpi_loss, clip_loss).mean()
critic_loss = 0.5 * (return_batch - values).pow(2).mean()
# Mirror Symmetry Loss
_, deterministic_actions = policy(obs_batch)
if env.clock_based:
mir_obs = mirror_observation(obs_batch, env.clock_inds)
_, mirror_actions = policy(mir_obs)
else:
_, mirror_actions = policy(mirror_observation(obs_batch))
mirror_actions = mirror_action(mirror_actions)
mirror_loss = 4 * (deterministic_actions - mirror_actions).pow(2).mean()
entropy_penalty = -self.entropy_coeff * pdf.entropy().mean()
optimizer.zero_grad()
(actor_loss + mirror_loss + entropy_penalty).backward()
# Clip the gradient norm to prevent "unlucky" minibatches from
# causing pathalogical updates
torch.nn.utils.clip_grad_norm_(policy.parameters(), self.grad_clip)
optimizer.step()
critic_optimizer.zero_grad()
critic_loss.backward()
# Clip the gradient norm to prevent "unlucky" minibatches from
# causing pathalogical updates
torch.nn.utils.clip_grad_norm_(critic.parameters(), self.grad_clip)
critic_optimizer.step()
losses.append([actor_loss.item(),
pdf.entropy().mean().item(),
critic_loss.item(),
ratio.mean().item(),
mirror_loss.item()])
# TODO: add verbosity arguments to suppress this
print(' '.join(["%g"%x for x in np.mean(losses, axis=0)]))
# Early stopping
if kl_divergence(pdf, old_pdf).mean() > 0.02:
print("Max kl reached, stopping optimization early.")
break
def train(self,
env_fn,
policy,
policy_copy,
critic,
n_itr,
logger=None):
# old_policy = deepcopy(policy)
old_policy = policy_copy
optimizer = optim.Adam(policy.parameters(), lr=self.lr, eps=self.eps)
critic_optimizer = optim.Adam(critic.parameters(), lr=self.lr, eps=self.eps)
start_time = time.time()
for itr in range(n_itr):
print("********** Iteration {} ************".format(itr))
sample_start = time.time()
batch = self.sample_parallel(env_fn, policy, critic, self.num_steps, self.max_traj_len)
print("time elapsed: {:.2f} s".format(time.time() - start_time))
print("sample time elapsed: {:.2f} s".format(time.time() - sample_start))
observations, actions, returns, values = map(torch.Tensor, batch.get())
advantages = returns - values
advantages = (advantages - advantages.mean()) / (advantages.std() + self.eps)
minibatch_size = self.minibatch_size or advantages.numel()
print("timesteps in batch: %i" % advantages.numel())
self.total_steps += advantages.numel()
old_policy.load_state_dict(policy.state_dict()) # WAY faster than deepcopy
optimizer_start = time.time()
self.update(policy, old_policy, optimizer, critic, critic_optimizer, observations, actions, returns, advantages, env_fn)
print("optimizer time elapsed: {:.2f} s".format(time.time() - optimizer_start))
if logger is not None:
evaluate_start = time.time()
test = self.sample_parallel(env_fn, policy, critic, 800 // self.n_proc, self.max_traj_len, deterministic=True)
print("evaluate time elapsed: {:.2f} s".format(time.time() - evaluate_start))
pdf = policy.evaluate(observations)
old_pdf = old_policy.evaluate(observations)
entropy = pdf.entropy().mean().item()
kl = kl_divergence(pdf, old_pdf).mean().item()
logger.add_scalar("Test/Return", avg_eval_reward, itr)
logger.add_scalar("Train/Return", np.mean(batch.ep_returns), itr)
logger.add_scalar("Train/Mean Eplen", np.mean(batch.ep_lens), itr)
logger.add_scalar("Train/Mean KL Div", kl, itr)
logger.add_scalar("Train/Mean Entropy", entropy, itr)
logger.add_scalar("Misc/Timesteps", self.total_steps, itr)
logger.dump()
# TODO: add option for how often to save model
if np.mean(test.ep_returns) > self.max_return:
self.max_return = np.mean(test.ep_returns)
self.save(policy)
print("Total time: {:.2f} s".format(time.time() - start_time))
def run_experiment(args):
torch.set_num_threads(1) # see: https://github.com/pytorch/pytorch/issues/13757
from apex import env_factory, create_logger
# # Environment
# if(args.env in ["Cassie-v0", "Cassie-mimic-v0", "Cassie-mimic-walking-v0"]):
# # NOTE: importing cassie for some reason breaks openai gym, BUG ?
# from cassie import CassieEnv, CassieTSEnv, CassieIKEnv
# from cassie.no_delta_env import CassieEnv_nodelta
# from cassie.speed_env import CassieEnv_speed
# from cassie.speed_double_freq_env import CassieEnv_speed_dfreq
# from cassie.speed_no_delta_env import CassieEnv_speed_no_delta
# # set up cassie environment
# # import gym_cassie
# # env_fn = gym_factory(args.env_name)
# #env_fn = make_env_fn(state_est=args.state_est)
# #env_fn = functools.partial(CassieEnv_speed_dfreq, "walking", clock_based = True, state_est=args.state_est)
# env_fn = functools.partial(CassieIKEnv, clock_based=True, state_est=args.state_est)
# print(env_fn().clock_inds)
# obs_dim = env_fn().observation_space.shape[0]
# action_dim = env_fn().action_space.shape[0]
# # Mirror Loss
# if args.mirror:
# if args.state_est:
# # with state estimator
# env_fn = functools.partial(SymmetricEnv, env_fn, mirrored_obs=[0.1, 1, 2, 3, 4, -10, -11, 12, 13, 14, -5, -6, 7, 8, 9, 15, 16, 17, 18, 19, 20, -26, -27, 28, 29, 30, -21, -22, 23, 24, 25, 31, 32, 33, 37, 38, 39, 34, 35, 36, 43, 44, 45, 40, 41, 42, 46, 47, 48], mirrored_act=[-5, -6, 7, 8, 9, -0.1, -1, 2, 3, 4])
# else:
# # without state estimator
# env_fn = functools.partial(SymmetricEnv, env_fn, mirrored_obs=[0.1, 1, 2, 3, 4, 5, -13, -14, 15, 16, 17,
# 18, 19, -6, -7, 8, 9, 10, 11, 12, 20, 21, 22, 23, 24, 25, -33,
# -34, 35, 36, 37, 38, 39, -26, -27, 28, 29, 30, 31, 32, 40, 41, 42],
# mirrored_act = [-5, -6, 7, 8, 9, -0.1, -1, 2, 3, 4])
# else:
# import gym
# env_fn = gym_factory(args.env_name)
# #max_episode_steps = env_fn()._max_episode_steps
# obs_dim = env_fn().observation_space.shape[0]
# action_dim = env_fn().action_space.shape[0]
# max_episode_steps = 1000
# wrapper function for creating parallelized envs
env_fn = env_factory(args.env_name, state_est=args.state_est, mirror=args.mirror, speed=args.speed)
obs_dim = env_fn().observation_space.shape[0]
action_dim = env_fn().action_space.shape[0]
# Set seeds
torch.manual_seed(args.seed)
np.random.seed(args.seed)
if args.previous is not None:
policy = torch.load(args.previous)
print("loaded model from {}".format(args.previous))
else:
policy = GaussianMLP_Actor(
obs_dim, action_dim,
env_name=args.env_name,
nonlinearity=torch.nn.functional.relu,
bounded=True,
init_std=np.exp(-2),
learn_std=False,
normc_init=False
)
policy_copy = GaussianMLP_Actor(
obs_dim, action_dim,
env_name=args.env_name,
nonlinearity=torch.nn.functional.relu,
bounded=True,
init_std=np.exp(-2),
learn_std=False,
normc_init=False
)
critic = GaussianMLP_Critic(
obs_dim,
env_name=args.env_name,
nonlinearity=torch.nn.functional.relu,
bounded=True,
init_std=np.exp(-2),
learn_std=False,
normc_init=False
)
policy.obs_mean, policy.obs_std = map(torch.Tensor, get_normalization_params(iter=args.input_norm_steps, noise_std=1, policy=policy, env_fn=env_fn))
critic.obs_mean = policy.obs_mean
policy_copy.obs_mean = policy.obs_mean
critic.obs_std = policy.obs_std
policy_copy.obs_std = policy.obs_std
policy_copy.train(0)
policy.train(0)
critic.train(0)
print("obs_dim: {}, action_dim: {}".format(obs_dim, action_dim))
if args.mirror:
algo = MirrorPPO(args=vars(args))
else:
algo = PPO(args=vars(args))
# create a tensorboard logging object
logger = create_logger(args)
print()
print("Synchronous Distributed Proximal Policy Optimization:")
print("\tenv: {}".format(args.env_name))
print("\tmax traj len: {}".format(args.max_traj_len))
print("\tseed: {}".format(args.seed))
print("\tmirror: {}".format(args.mirror))
print("\tnum procs: {}".format(args.num_procs))
print("\tlr: {}".format(args.lr))
print("\teps: {}".format(args.eps))
print("\tlam: {}".format(args.lam))
print("\tgamma: {}".format(args.gamma))
print("\tentropy coeff: {}".format(args.entropy_coeff))
print("\tclip: {}".format(args.clip))
print("\tminibatch size: {}".format(args.minibatch_size))
print("\tepochs: {}".format(args.epochs))
print("\tnum steps: {}".format(args.num_steps))
print("\tuse gae: {}".format(args.use_gae))
print("\tmax grad norm: {}".format(args.max_grad_norm))
print("\tmax traj len: {}".format(args.max_traj_len))
print()
algo.train(env_fn, policy, policy_copy, critic, args.n_itr, logger=logger) |
# -*- coding: utf-8 -*-
"""
Created on Tue Feb 12 23:01:53 2019
@author: <NAME>
<EMAIL>
"""
from sklearn.datasets import load_digits, load_breast_cancer, load_diabetes
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.cluster import KMeans
from sklearn.decomposition import KernelPCA
# =============================================================================
# DATASETS
# =============================================================================
diabetes = load_diabetes()
bc = load_breast_cancer()
digits = load_digits()
images_and_labels = list(zip(digits.images, digits.target))
for index, (image, label) in enumerate(images_and_labels[10:20]):
plt.subplot(2, 5, index + 1)
plt.axis('off')
plt.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
plt.title('Target: %i' % label)
# =============================================================================
# CLASSIFICATION
# =============================================================================
f = lambda x: 2 * x - 5
pos = []
neg = []
for i in range(30):
x = np.random.randint(15)
y = np.random.randint(15)
if f(x) < y:
pos.append([x,y])
else:
neg.append([x,y])
plt.figure()
plt.xticks([])
plt.yticks([])
plt.scatter(*zip(*pos))
plt.scatter(*zip(*neg))
plt.plot([0,10],[f(0),f(10)], linestyle='--', color='m')
plt.xlabel('x')
plt.ylabel('y')
plt.title('Classification')
# =============================================================================
# REGRESSION
# =============================================================================
dat = []
for i in range(30):
x = np.random.uniform(10)
y = f(x) + np.random.uniform(-2.0,2.0)
dat.append([x,y])
plt.figure()
plt.xticks([])
plt.yticks([])
plt.scatter(*zip(*dat))
plt.plot([0,10],[f(0),f(10)], linestyle='--', color='m')
plt.xlabel('x')
plt.ylabel('y')
plt.title('Regression')
# =============================================================================
# CLUSTERING
# =============================================================================
km = KMeans(n_clusters=3)
dat = []
t = 0.5
for i in range(300):
c = np.random.randint(3)
a = np.random.uniform() * 2 * 3.14
r = t * np.sqrt(np.random.uniform())
x = r * np.cos(a)
y = r * np.sin(a)
dat.append([c+x, c+y])
c = km.fit_predict(dat)
plt.figure()
plt.xticks([])
plt.yticks([])
plt.scatter(*zip(*dat),c=c)
plt.xlabel('x')
plt.ylabel('y')
plt.title('Clustering')
# =============================================================================
# PCA
# =============================================================================
from sklearn.datasets import make_circles
pca = KernelPCA(kernel="rbf", fit_inverse_transform=True, gamma=10)
x, y = make_circles(n_samples=400, factor=.3, noise=.05)
pp = pca.fit_transform(x)
plt.figure()
plt.xticks([])
plt.yticks([])
plt.scatter(pp[:,0], pp[:,1], c=y)
plt.xlabel('x')
plt.ylabel('y')
plt.title('Clustering')
# =============================================================================
# TSNE
# =============================================================================
from sklearn.manifold import TSNE
tsne = TSNE()
dat = tsne.fit_transform(bc.data)
reds = bc.target == 0
blues = bc.target == 1
plt.scatter(dat[reds,0], dat[reds,1], label='malignant')
plt.scatter(dat[blues,0], dat[blues,1], label='benign')
plt.xlabel('1st Component')
plt.ylabel('2nd Component')
plt.title('Breast Cancer Data')
plt.legend()
# =============================================================================
# ROC
# =============================================================================
import numpy as np
from sklearn import metrics
ax1 = plt.subplot()
ax1.margins(0)
np.random.seed(856522)
y = np.random.choice([1,2], 30)
scores = np.random.choice([i/100 for i in range(0,100)], 30)
fpr, tpr, thresholds = metrics.roc_curve(y, scores, pos_label=2)
x = [i/100 for i in range(0,100)]
y = [i/100 for i in range(0,100)]
plt.plot(x, y, linestyle='-.')
plt.plot(fpr, tpr, label='ROC curve')
plt.xlabel('Specificity')
plt.ylabel('Sensitivity')
plt.title('ROC')
plt.legend()
|
#-------------------------------------------------------------------------------
#
# Project: EOxServer <http://eoxserver.org>
# Authors: <NAME> <<EMAIL>>
#
#-------------------------------------------------------------------------------
# Copyright (C) 2011 EOX IT Services GmbH
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all
# copies of this Software or works derived from this Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN
# THE SOFTWARE.
#-------------------------------------------------------------------------------
from eoxserver.contrib.mapserver import Layer
from eoxserver.services.mapserver.wms.layerfactories.base import (
AbstractLayerFactory, OffsiteColorMixIn
)
class CoverageBandsLayerFactory(OffsiteColorMixIn, AbstractLayerFactory):
suffixes = ("_bands",)
requires_connection = True
def generate(self, eo_object, group_layer, suffix, options):
name = eo_object.identifier + "_bands"
layer = Layer(name)
layer.setMetaData("ows_title", name)
layer.setMetaData("wms_label", name)
layer.addProcessing("CLOSE_CONNECTION=CLOSE")
coverage = eo_object.cast()
range_type = coverage.range_type
req_bands = options["bands"]
band_indices = []
bands = []
for req_band in req_bands:
if isinstance(req_band, int):
band_indices.append(req_band + 1)
bands.append(range_type[req_band])
else:
for i, band in enumerate(range_type):
if band.name == req_band:
band_indices.append(i + 1)
bands.append(band)
break
else:
raise Exception(
"Coverage '%s' does not have a band with name '%s'."
% (coverage.identifier, req_band)
)
if len(req_bands) in (3, 4):
indices_str = ",".join(map(str, band_indices))
offsite_indices = list(map(lambda v: v-1, band_indices[:3]))
elif len(req_bands) == 1:
indices_str = ",".join(map(str, band_indices * 3))
v = band_indices[0] - 1
offsite_indices = [v, v, v]
else:
raise Exception("Invalid number of bands requested.")
offsite = self.offsite_color_from_range_type(
range_type, offsite_indices
)
options = self.get_render_options(coverage)
self.set_render_options(layer, offsite, options)
layer.setProcessingKey("BANDS", indices_str)
if options.bands_scale_min and options.bands_scale_max:
bands_scale_min = str(options.bands_scale_min).split(',')
bands_scale_max = str(options.bands_scale_max).split(',')
idx1, idx2, idx3 = offsite_indices
layer.setProcessingKey("SCALE_1", "%s,%s" % (
bands_scale_min[idx1], bands_scale_max[idx1]
))
layer.setProcessingKey("SCALE_2", "%s,%s" % (
bands_scale_min[idx2], bands_scale_max[idx2]
))
layer.setProcessingKey("SCALE_3", "%s,%s" % (
bands_scale_min[idx3], bands_scale_max[idx3]
))
yield (layer, coverage.data_items.all())
def generate_group(self, name):
return Layer(name)
|
import logging
import time
import os
import importlib
import WorkManager
import FileManager
from pandayoda.common import MessageTypes
from pandayoda.common.yoda_multiprocessing import Process, Event
logger = logging.getLogger(__name__)
config_section = os.path.basename(__file__)[:os.path.basename(__file__).rfind('.')]
class Yoda(Process):
def __init__(self, queues, config, rank, worldsize):
''' config: configuration of Yoda
'''
# call Thread constructor
super(Yoda, self).__init__()
# message queues
self.queues = queues
# rank number
self.rank = rank
# world size
self.worldsize = worldsize
# config settings
self.config = config
# keep track of if the wallclock has expired
self.wallclock_expired = Event()
# this is used to trigger the thread exit
self.exit = Event()
def stop(self):
''' this function can be called by outside threads to cause the Yoda thread to exit'''
self.exit.set()
# this runs when 'yoda_instance.start()' is called
def run(self):
''' this is the function called when the user runs yoda_instance.start() '''
try:
self.subrun()
except Exception:
logger.exception('Yoda failed with uncaught exception')
raise
def subrun(self):
''' this function is the business logic, but wrapped in exception '''
self.read_config()
# set logging level
logger.info('Yoda Thread starting')
logger.info('loglevel: %s', self.loglevel)
logger.info('loop_timeout: %s', self.loop_timeout)
top_working_path = os.getcwd()
logger.debug('cwd: %s', top_working_path)
# setup harvester messenger to share with FileManager and WorkManager
logger.debug('setup harvester messenger')
harvester_messenger = self.get_harvester_messenger()
harvester_messenger.setup(self.config)
# wait for setup to complete
harvester_messenger.sfm_har_config_done.wait()
# a list of ranks that have exited
self.exited_droids = []
# a dictionary of subthreads
subthreads = {}
# create WorkManager thread
subthreads['WorkManager'] = WorkManager.WorkManager(self.config, self.queues, harvester_messenger)
subthreads['WorkManager'].start()
# create FileManager thread
subthreads['FileManager'] = FileManager.FileManager(self.config, self.queues, top_working_path,
harvester_messenger)
subthreads['FileManager'].start()
# start message loop
while not self.exit.is_set():
logger.debug('start loop')
# process incoming messages from other threads or ranks
self.process_incoming_messages()
# check if all droids have exited
if len(self.exited_droids) >= (self.worldsize - 1):
logger.info('all droids have exited, exiting yoda')
self.stop()
break
# check the status of each subthread
logger.debug('checking all threads still alive')
keys = subthreads.keys()
for name in keys:
thread = subthreads[name]
# if the thread is not alive, throw an error
if not thread.is_alive():
logger.warning('%s is no longer running.', name)
del subthreads[name]
if name == 'WorkManager':
self.stop()
continue
# else:
# logger.debug('%s %s is running.',self.prelog,name)
if len(subthreads) == 0:
logger.info('no subthreads remaining, exiting')
self.stop()
break
if self.queues['Yoda'].empty():
logger.debug('sleeping %s', self.loop_timeout)
self.exit.wait(timeout=self.loop_timeout)
# send the exit signal to all droid ranks
logger.info('sending exit signal to droid ranks')
for ranknum in range(1, self.worldsize):
if ranknum not in self.exited_droids:
if self.wallclock_expired.is_set():
self.queues['MPIService'].put(
{'type': MessageTypes.WALLCLOCK_EXPIRING, 'destination_rank': ranknum})
else:
self.queues['MPIService'].put({'type': MessageTypes.DROID_EXIT, 'destination_rank': ranknum})
# send the exit signal to all subthreads
for name, thread in subthreads.iteritems():
logger.info('sending exit signal to %s', name)
thread.stop()
# wait for sub threads to exit
for name, thread in subthreads.iteritems():
logger.info('waiting for %s to join', name)
thread.join()
logger.info('%s has joined', name)
while not self.queues['MPIService'].empty():
logger.info('waiting for MPIService to send exit messages to Droid, sleep for %s', self.loop_timeout)
time.sleep(self.loop_timeout)
logger.info('Yoda is exiting')
def read_config(self):
if config_section in self.config:
# read log level:
if 'loglevel' in self.config[config_section]:
self.loglevel = self.config[config_section]['loglevel']
logger.info('%s loglevel: %s', config_section, self.loglevel)
logger.setLevel(logging.getLevelName(self.loglevel))
else:
logger.warning('no "loglevel" in "%s" section of config file, keeping default', config_section)
# read loop timeout:
if 'loop_timeout' in self.config[config_section]:
self.loop_timeout = int(self.config[config_section]['loop_timeout'])
logger.info('%s loop_timeout: %s', config_section, self.loop_timeout)
else:
logger.warning('no "loop_timeout" in "%s" section of config file, keeping default %s', config_section,
self.loop_timeout)
# messenger_plugin_module
if 'messenger_plugin_module' in self.config[config_section]:
self.messenger_plugin_module = self.config[config_section]['messenger_plugin_module']
else:
raise Exception(
'Failed to retrieve "messenger_plugin_module" from config file section %s' % config_section)
else:
raise Exception('no %s section in the configuration' % config_section)
def process_incoming_messages(self):
while not self.queues['Yoda'].empty():
qmsg = self.queues['Yoda'].get(block=False)
# process message
logger.debug('received message: %s', qmsg)
if qmsg['type'] == MessageTypes.DROID_HAS_EXITED:
logger.debug(' droid rank %d has exited', qmsg['source_rank'])
self.exited_droids.append(qmsg['source_rank'])
logger.debug('%s droid ranks have exited', len(self.exited_droids))
else:
logger.error(' could not interpret message: %s', qmsg)
def get_harvester_messenger(self):
# try to import the module specified in the config
# if it is not in the PYTHONPATH this will fail
try:
return importlib.import_module(self.messenger_plugin_module)
except ImportError:
logger.exception('Failed to import messenger_plugin: %s', self.messenger_plugin_module)
raise
|
<reponame>seblee97/student_teacher_catastrophic
import abc
import math
from typing import List
from typing import Union
import torch
from cata import constants
from cata.teachers import classification_teacher
from cata.teachers import regression_teacher
class BaseTeacherEnsemble(abc.ABC):
"""Base class for sets/ensembles of teachers
(as opposed to single teacher network)."""
def __init__(
self,
input_dimension: int,
hidden_dimensions: List[int],
output_dimension: int,
bias: bool,
loss_type: str,
nonlinearities: List[str],
scale_hidden_lr: bool,
forward_scaling: float,
unit_norm_teacher_head: bool,
weight_normalisation: bool,
noise_stds: List[Union[int, float]],
num_teachers: int,
initialisation_std: float,
) -> None:
self._input_dimension = input_dimension
self._hidden_dimensions = hidden_dimensions
self._output_dimension = output_dimension
self._bias = bias
self._loss_type = loss_type
self._nonlinearities = nonlinearities
self._forward_scaling = forward_scaling
self._unit_norm_teacher_head = unit_norm_teacher_head
self._weight_normalisation = weight_normalisation
self._noise_stds = noise_stds
self._num_teachers = num_teachers
self._initialisation_std = initialisation_std
if scale_hidden_lr:
self._forward_hidden_scaling = 1 / math.sqrt(input_dimension)
else:
self._forward_hidden_scaling = 1.0
self._teachers = self._setup_teachers()
@property
def teachers(self) -> List:
"""Getter method for teacher networks."""
return self._teachers
@property
def cross_overlaps(self):
overlaps = []
with torch.no_grad():
for i in range(len(self._teachers)):
for j in range(i, len(self._teachers)):
if i != j:
overlap = (
torch.mm(
self._teachers[i].layers[0].weight.data,
self._teachers[j].layers[0].weight.data.T,
)
/ self._input_dimension
)
overlaps.append(overlap)
return overlaps
@abc.abstractmethod
def _setup_teachers(self) -> None:
"""instantiate teacher network(s)"""
pass
def forward(self, teacher_index: int, batch: torch.Tensor) -> torch.Tensor:
"""Call to current teacher forward."""
output = self._teachers[teacher_index](batch)
return output
def _init_teacher(
self, nonlinearity: str, noise_std: Union[float, int], zero_head: bool = False
):
if self._loss_type == constants.CLASSIFICATION:
teacher = classification_teacher.ClassificationTeacher
elif self._loss_type == constants.REGRESSION:
teacher = regression_teacher.RegressionTeacher
else:
raise ValueError(f"Loss type {self._loss_type} is not recognised.")
return teacher(
input_dimension=self._input_dimension,
hidden_dimensions=self._hidden_dimensions,
output_dimension=self._output_dimension,
bias=self._bias,
nonlinearity=nonlinearity,
forward_hidden_scaling=self._forward_hidden_scaling,
forward_scaling=self._forward_scaling,
unit_norm_teacher_head=self._unit_norm_teacher_head,
weight_normalisation=self._weight_normalisation,
noise_std=noise_std,
initialisation_std=self._initialisation_std,
zero_head=zero_head,
)
def save_all_teacher_weights(self, save_path: str) -> None:
"""Save weights associated with each teacher
Args:
save_path: path to save weights, will be concatenated with
_i where i is the index of the teacher.
"""
for t, teacher in enumerate(self._teachers):
torch.save(teacher.state_dict(), f"{save_path}_{t}")
def save_weights(self, teacher_index: int, save_path: str) -> None:
"""Save weights associated with given teacher index"""
torch.save(self._teachers[teacher_index].state_dict(), save_path)
def forward_all(self, batch: torch.Tensor) -> List[torch.Tensor]:
"""Call to forward of all teachers (used primarily for evaluation)"""
outputs = [self.forward(t, batch) for t in range(self._num_teachers)]
return outputs
|
<reponame>JBurkinshaw/ogc-api-fast-features
import os
from asyncio import get_event_loop
from typing import Type
from unittest.mock import patch
from uuid import uuid4
from oaff.app.configuration.data import get_layer, get_layers
from oaff.app.configuration.frontend_configuration import FrontendConfiguration
from oaff.app.data.retrieval.feature_provider import FeatureProvider
from oaff.app.data.retrieval.feature_set_provider import FeatureSetProvider
from oaff.app.data.sources.common.data_source import DataSource
from oaff.app.data.sources.common.layer import Layer
from oaff.app.gateway import cleanup, configure
from oaff.app.responses.response_format import ResponseFormat
from oaff.app.responses.response_type import ResponseType
from oaff.app.settings import ENV_VAR_PREFIX
def setup_module():
os.environ[f"{ENV_VAR_PREFIX}DATA_SOURCE_TYPES"] = "postgresql"
def teardown_module():
del os.environ[f"{ENV_VAR_PREFIX}DATA_SOURCE_TYPES"]
def teardown_function():
get_event_loop().run_until_complete(cleanup())
@patch("oaff.app.data.sources.postgresql.postgresql_manager.PostgresqlManager")
def test_empty(PostgresqlManagerMock):
PostgresqlManagerMock.return_value.get_data_sources.return_value = []
get_event_loop().run_until_complete(
configure(
FrontendConfiguration(
asset_url_base="",
api_url_base="",
endpoint_format_switcher=_endpoint_format_switcher,
next_page_link_generator=_next_page_link_generator,
prev_page_link_generator=_prev_page_link_generator,
openapi_path_html="/html",
openapi_path_json="/json",
)
)
)
assert len(get_layers()) == 0
@patch("oaff.app.data.sources.postgresql.postgresql_manager.PostgresqlManager")
def test_with_layers(PostgresqlManagerMock):
PostgresqlManagerMock.return_value.get_data_sources.return_value = [
_TestDataSource1(str(uuid4())),
_TestDataSource2(str(uuid4())),
]
get_event_loop().run_until_complete(
configure(
FrontendConfiguration(
asset_url_base="",
api_url_base="",
endpoint_format_switcher=_endpoint_format_switcher,
next_page_link_generator=_next_page_link_generator,
prev_page_link_generator=_prev_page_link_generator,
openapi_path_html="/html",
openapi_path_json="/json",
)
)
)
assert len(get_layers()) == 3
for lyrnum in [1, 2, 3]:
assert get_layer(f"layer{lyrnum}").title == f"title{lyrnum}"
assert get_layer(f"layer{lyrnum}").description == f"description{lyrnum}"
assert len(get_layer(f"layer{lyrnum}").bboxes) == 1
assert get_layer(f"layer{lyrnum}").bboxes[0] == [
int(f"{lyrnum}1"),
int(f"{lyrnum}2"),
int(f"{lyrnum}3"),
int(f"{lyrnum}4"),
]
def _endpoint_format_switcher(
url: str, format: ResponseFormat, type: ResponseType
) -> str:
return url
def _next_page_link_generator(url: str) -> str:
return url
def _prev_page_link_generator(url: str) -> str:
return url
class _TestDataSource1(DataSource):
async def get_layers(self):
return [
Layer(
id="layer1",
title="title1",
description="description1",
bboxes=[[11, 12, 13, 14]],
intervals=[[None, None]],
data_source_id=self.id,
geometry_crs_auth_name="EPSG",
geometry_crs_auth_code=3857,
temporal_attributes=[],
),
Layer(
id="layer2",
title="title2",
description="description2",
bboxes=[[21, 22, 23, 24]],
intervals=[[None, None]],
data_source_id=self.id,
geometry_crs_auth_name="EPSG",
geometry_crs_auth_code=4326,
temporal_attributes=[],
),
]
async def disconnect(self):
pass
async def get_feature_set_provider(self) -> Type[FeatureSetProvider]:
pass
async def get_feature_provider(self) -> Type[FeatureProvider]:
pass
async def initialize(self):
pass
class _TestDataSource2(DataSource):
async def get_layers(self):
return [
Layer(
id="layer3",
title="title3",
description="description3",
bboxes=[[31, 32, 33, 34]],
intervals=[[None, None]],
data_source_id=self.id,
geometry_crs_auth_name="EPSG",
geometry_crs_auth_code=3857,
temporal_attributes=[],
)
]
async def disconnect(self):
pass
async def get_feature_set_provider(self) -> Type[FeatureSetProvider]:
pass
async def get_feature_provider(self) -> Type[FeatureProvider]:
pass
async def initialize(self):
pass
|
#!/usr/bin/env python
"""Simple script to package cache folder into GeoPackage.
Includes GlobalGeodetic class from gdal2tiles.py by <NAME>, klokan at klokan dot cz
licensed under MIT.
"""
__author__ = '<NAME>'
__copyright__ = "Copyright 2015, Esri"
__license__ = "ASL 2.0"
__version__ = "1.1"
__credits__ = ["<NAME>"]
# $Id$
import os
import sys
import re
import math
import sqlite3
import fnmatch
import collections
try:
import xml.etree.cElementTree as ET
except ImportError:
import xml.etree.ElementTree as ET
TILES_TABLE_SUFFIX = '_tiles' # Added to basename to create table_name
TILES_TABLE_PREFIX = 'table_' # Used if basename starts with a non alphabet character
class GlobalGeodetic(object):
"""
GlobalGeodetic class is licensed under MIT:
Copyright (c) 2008, <NAME>
Permission is hereby granted, free of charge, to any person obtaining a
copy of this software and associated documentation files (the "Software"),
to deal in the Software without restriction, including without limitation
the rights to use, copy, modify, merge, publish, distribute, sublicense,
and/or sell copies of the Software, and to permit persons to whom the
Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included
in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS
OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
DEALINGS IN THE SOFTWARE.
TMS Global Geodetic Profile
---------------------------
Functions necessary for generation of global tiles in Plate Carre projection,
EPSG:4326, "unprojected profile".
Such tiles are compatible with Google Earth (as any other EPSG:4326 rasters)
and you can overlay the tiles on top of OpenLayers base map.
Pixel and tile coordinates are in TMS notation (origin [0,0] in bottom-left).
What coordinate conversions do we need for TMS Global Geodetic tiles?
Global Geodetic tiles are using geodetic coordinates (latitude,longitude)
directly as planar coordinates XY (it is also called Unprojected or Plate
Carre). We need only scaling to pixel pyramid and cutting to tiles.
Pyramid has on top level two tiles, so it is not square but rectangle.
Area [-180,-90,180,90] is scaled to 512x256 pixels.
TMS has coordinate origin (for pixels and tiles) in bottom-left corner.
Rasters are in EPSG:4326 and therefore are compatible with Google Earth.
LatLon <-> Pixels <-> Tiles
WGS84 coordinates Pixels in pyramid Tiles in pyramid
lat/lon XY pixels Z zoom XYZ from TMS
EPSG:4326
.----. ----
/ \ <-> /--------/ <-> TMS
\ / /--------------/
----- /--------------------/
WMS, KML Web Clients, Google Earth TileMapService
"""
def __init__(self, tileSize=256):
self.tileSize = tileSize
self.resFact = 360.0 / self.tileSize
self.origin = [-180, -90, 0, 90]
def LatLonToPixels(self, lat, lon, zoom):
"Converts lat/lon to pixel coordinates in given zoom of the EPSG:4326 pyramid"
res = self.resFact / 2**zoom
px = (180 + lat) / res
py = (90 + lon) / res
return px, py
def PixelsToTile(self, px, py):
"Returns coordinates of the tile covering region in pixel coordinates"
tx = int(math.ceil(px / float(self.tileSize)) - 1)
ty = int(math.ceil(py / float(self.tileSize)) - 1)
return tx, ty
def LatLonToTile(self, lat, lon, zoom):
"Returns the tile for zoom which covers given lat/lon coordinates"
px, py = self.LatLonToPixels(lat, lon, zoom)
return self.PixelsToTile(px, py)
def MatrixDim(self, zoom):
"Matrix Wdith and Height for given zoom level"
return 2**zoom
def Resolution(self, zoom):
"Resolution (arc/pixel) for given zoom level (measured at Equator)"
return self.resFact / 2**zoom
#return 180 / float( 1 << (8+zoom) )
def ZoomForPixelSize(self, pixelSize):
"Maximal scaledown zoom of the pyramid closest to the pixelSize."
for i in range(MAXZOOMLEVEL):
if pixelSize > self.Resolution(i):
if i != 0:
return i - 1
else:
return 0 # We don't want to scale up
def TileBounds(self, tx, ty, zoom):
"Returns bounds of the given tile"
res = self.resFact / 2**zoom
return (tx * self.tileSize * res - 180, ty * self.tileSize * res - 90,
(tx + 1) * self.tileSize * res - 180,
(ty + 1) * self.tileSize * res - 90)
def TileLatLonBounds(self, tx, ty, zoom):
"Returns bounds of the given tile in the SWNE form"
b = self.TileBounds(tx, ty, zoom)
return (b[1], b[0], b[3], b[2])
class Point:
def __init__(self):
self.x = 0.0
self.y = 0.0
def __init__(self, x, y):
self.x = x
self.y = y
class Cache:
def __init__(self):
self.path = None
self.xmlTree = None
self.wkt = None
self.min_x = None
self.min_y = None
self.max_x = None
self.max_y = None
self.matrix_min_x = None
self.matrix_min_y = None
self.matrix_max_x = None
self.matrix_max_y = None
self.tileStart = None
self.tileStop = None
self.levels = []
self.geodetic = GlobalGeodetic()
self.verbose = False
self.srs_id = 0
self.srs_org_id = 3857
self.srs_org_name = 'EPSG'
self.level_info = collections.namedtuple('level_info', ['startX', 'startY',
'stopX', 'stopY',
'matrix_width', 'matrix_height',
'zoom_level',
'pixel_x_size', 'pixel_y_size',
'offset_x', 'offset_y'])
self.level_infos = []
def deg2num(self, lat_deg, lon_deg, zoom):
lat_rad = math.radians(lat_deg)
n = 2.0 ** zoom
xtile = int((lon_deg + 180.0) / 360.0 * n)
ytile = int((1.0 - math.log(math.tan(lat_rad) + (1 / math.cos(lat_rad))) / math.pi) / 2.0 * n)
return (xtile, ytile)
def num2deg(self, xtile, ytile, zoom):
n = 2.0 ** zoom
lon_deg = xtile / n * 360.0 - 180.0
lat_rad = math.atan(math.sinh(math.pi * (1 - 2 * ytile / n)))
lat_deg = math.degrees(lat_rad)
return (lat_deg, lon_deg)
def parseXML(self, path):
self.xmlTree = ET.ElementTree(file=path)
wktElement = self.xmlTree.iterfind('SpatialReference/WKT')
if wktElement is not None:
self.wkt = next(wktElement).text
originElement = self.xmlTree.iterfind('XMin')
if originElement is not None:
self.min_x = float(next(originElement).text)
originElement = self.xmlTree.iterfind('YMin')
if originElement is not None:
self.min_y = float(next(originElement).text)
originElement = self.xmlTree.iterfind('XMax')
if originElement is not None:
self.max_x = float(next(originElement).text)
originElement = self.xmlTree.iterfind('YMax')
if originElement is not None:
self.max_y = float(next(originElement).text)
print("self.max_x = {0}".format(self.max_x))
print("self.max_y = {0}".format(self.max_y))
if self.wkt is None or self.min_x is None or self.min_y is None or self.max_x is None or self.max_y is None:
return False
latestWKIDElement = self.xmlTree.iterfind('SpatialReference/LatestWKID')
if latestWKIDElement is not None:
self.srs_id = int(next(latestWKIDElement).text)
ulLatLon = self.min_x, self.max_y
lrLatLon = self.max_x, self.min_y
print("Lat/Lon: {0}, {1}, {2}, {3}".format(ulLatLon[0], ulLatLon[1], lrLatLon[0], lrLatLon[1]))
#startX, startY, stopX, stopY = self.getTileStartStopLL(ulLatLon[0], ulLatLon[1], lrLatLon[0], lrLatLon[1], self.levels[0])
#ulLatLon = self.num2deg(startX, startY, self.levels[0])
#lrLatLon = self.num2deg(stopX, stopY, self.levels[0])
self.matrix_min_x, self.matrix_max_y = ulLatLon[0], ulLatLon[1]
self.matrix_max_x, self.matrix_min_y = lrLatLon[0], lrLatLon[1]
for index, level in enumerate(self.levels):
if index == 0:
startX, startY, stopX, stopY = self.getTileStartStopL0(self.levels[0])
else:
prev = self.level_infos[index - 1]
startX = prev.startX * 2
startY = prev.startY * 2
stopX = prev.stopX * 2
stopY = prev.stopY * 2
self.level_infos.append(self.level_info(startX=startX, startY=startY, stopX=stopX, stopY=stopY,
matrix_width=self.geodetic.MatrixDim(level+1),
matrix_height=self.geodetic.MatrixDim(level+1),
zoom_level=level,
pixel_x_size=self.geodetic.Resolution(level+1),
pixel_y_size=self.geodetic.Resolution(level+1),
offset_x=0, offset_y=0))
print("Tile(s)[{0}]: {1}, {2}, {3}, {4}".format(level, startX, startY, stopX-1, stopY-1))
return True
def findZ(self, path):
for entry in os.listdir(path):
entry_path = os.path.join(path, entry)
entry_lower = entry.lower()
if os.path.isdir(entry_path) and fnmatch.fnmatch(entry_lower, 'l??'):
level_str = entry_lower.split('l')
if len(level_str) == 2:
self.levels.append(int(level_str[1]))
if not self.levels:
return False
self.levels = sorted(self.levels)
print("Found level(s): {0}".format(str(self.levels).strip('[]')))
return True
def getTileStartStopL0(self, level):
level0Path = os.path.join(self.path, "_alllayers", "L{0:02d}".format(level))
rows = sorted(os.listdir(level0Path))
cols = sorted(os.listdir(os.path.join(level0Path, rows[0])))
startX = int(cols[0][1:-4], 16)
stopX = int(cols[-1][1:-4], 16)
startY = int(rows[0][1:], 16)
stopY = int(rows[-1][1:], 16)
return startX, startY, stopX + 1, stopY + 1
def getTileStartStopLL(self, min_x, min_y, max_x, max_y, level):
if level not in self.levels:
return (0, 0), (0, 0)
startX, startY = self.deg2num(min_x, min_y, level)
stopX, stopY = self.deg2num(max_x, max_y, level)
return startX, startY, stopX+1, stopY+1
def getTileStartStop(self, level):
if level not in self.levels:
return (0, 0), (0, 0)
tileStart = self.geodetic.LatLonToTile(self.min_x, self.min_y, level)
tileStop = self.geodetic.LatLonToTile(self.max_x, self.max_y, level)
startX = tileStart[0]
startY = tileStart[1]
stopX = tileStop[0]
stopY = tileStop[1]
return startX, startY, stopX+1, stopY+1
def getTilePath(self, x, y, level):
levelPath = "L{0:02d}".format(level)
rowPath = "R{0:08x}".format(y)
columnPath = "C{0:08x}".format(x)
return os.path.join(self.path, "_alllayers", levelPath, rowPath, columnPath)
def findTile(self, path):
jpgPath = path + '.jpg'
if os.path.exists(jpgPath):
return jpgPath
jpegPath = path + '.jpeg'
if os.path.exists(jpegPath):
return jpegPath
pngPath = path + '.png'
if os.path.exists(pngPath):
return pngPath
return None
def checkTiles(self):
for level in self.levels:
startX, startY, stopX, stopY = self.getTileStartStop(level)
for y in range(startY, stopY):
for x in range(startX, stopX):
tilePath = self.getTilePath(x, y, level)
foundTilePath = self.findTile(tilePath)
if foundTilePath is None:
print("Missing tile: {0}.png/.jpg".format(tilePath))
elif self.verbose:
print("Found tile: {0}".format(tilePath))
print("Required tiles found at expected locations.")
return True
def open(self, path):
if not os.path.isdir(path):
return False
self.path = path
levelsDir = os.path.join(self.path, '_alllayers')
if not self.findZ(levelsDir):
return False
xmlFile = os.path.join(self.path, 'conf.cdi')
if not self.parseXML(xmlFile):
return False
return True
class GeoPackage:
"""
Simple class to add tiles to an existing or new GeoPackage using GDAL.
"""
def __init__(self):
self.connection = None
self.filename = None
self.tile_width = 256
self.tile_height = 256
self.sr_organization = "NONE"
self.sr_organization_coordsys_id = 0
self.sr_description = None
self.description = None
self.cache = Cache()
self.verbose = False
def __del__(self):
if self.connection is not None:
self.connection.close()
def write_srs(self, srs_name):
"""
Write SRS to gpkg_spatial_ref_sys table and return srs_id.
@param wkt: WKT string.
@param srs_name: Value for srs_name field.
@return: srs_id for new entry or -1 (undefined cartesian)
"""
if self.cache.wkt is None:
return -1
result = self.connection.execute("""SELECT * FROM gpkg_spatial_ref_sys WHERE srs_id=?;""",
(self.cache.srs_id,)).fetchone()
if result is None:
self.connection.execute(
"""
INSERT INTO gpkg_spatial_ref_sys(srs_name, srs_id, organization, organization_coordsys_id, definition)
VALUES(?, ?, ?, ?, ?)
""", (srs_name, self.cache.srs_id, self.cache.srs_org_name, self.cache.srs_org_id, self.cache.wkt))
self.connection.commit()
return self.cache.srs_id
else:
return result['srs_id']
def add_cache(self, path):
if not self.cache.open(path):
return False
identifier = os.path.basename(path)
table_name = re.sub('[.~,;-]', '', identifier + TILES_TABLE_SUFFIX)
if not table_name[0].isalpha():
table_name = TILES_TABLE_PREFIX + table_name
#table_name = table_name.lower()
if self.connection.execute("""SELECT * FROM gpkg_contents WHERE identifier=? OR table_name=?""",
(identifier, table_name)).fetchone() is not None:
print("An entry with identifier {0} and/or table_name {1} already exists in gpkg_contents.".format(identifier, table_name))
return False
if self.cache.srs_id == 3857:
srs_id = self.write_srs('Web Mercator')
if self.description is None:
self.description = path
try:
self.connection.execute(
"""
INSERT INTO gpkg_contents(table_name, data_type, identifier, description, min_x, min_y, max_x, max_y, srs_id)
VALUES(?, 'tiles', ?, ?, ?, ?, ?, ?, ?);
""",
(table_name, identifier, self.description, self.cache.matrix_min_x, self.cache.matrix_min_y, self.cache.matrix_max_x, self.cache.matrix_max_y, self.cache.srs_id)
)
self.connection.execute(
"""
INSERT INTO gpkg_tile_matrix_set(table_name, srs_id, min_x, min_y, max_x, max_y)
VALUES(?, ?, ?, ?, ?, ?);
""",
(table_name, self.cache.srs_id, self.cache.geodetic.origin[0], self.cache.geodetic.origin[1],
self.cache.geodetic.origin[2], self.cache.geodetic.origin[3])
)
sql_string = """
CREATE TABLE """ + table_name + """ (
id INTEGER PRIMARY KEY AUTOINCREMENT,
zoom_level INTEGER NOT NULL,
tile_column INTEGER NOT NULL,
tile_row INTEGER NOT NULL,
tile_data BLOB NOT NULL,
UNIQUE (zoom_level, tile_column, tile_row) );
"""
self.connection.execute(sql_string)
sql_string = """
CREATE TRIGGER '""" + table_name + """_zoom_insert'
BEFORE INSERT ON '""" + table_name + """'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table """ + table_name + """ violates constraint: zoom_level not specified for table in gpkg_tile_matrix')
WHERE NOT (NEW.zoom_level IN (SELECT zoom_level FROM gpkg_tile_matrix WHERE table_name = '""" + table_name + """')) ;
END
"""
self.connection.execute(sql_string)
sql_string = """
CREATE TRIGGER '""" + table_name + """_zoom_update'
BEFORE UPDATE OF zoom_level ON '""" + table_name + """'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table """ + table_name + """ violates constraint: zoom_level not specified for table in gpkg_tile_matrix')
WHERE NOT (NEW.zoom_level IN (SELECT zoom_level FROM gpkg_tile_matrix WHERE table_name = '""" + table_name + """')) ;
END
"""
self.connection.execute(sql_string)
sql_string = """
CREATE TRIGGER '""" + table_name + """_tile_column_insert'
BEFORE INSERT ON '""" + table_name + """'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table """ + table_name + """ violates constraint: tile_column cannot be < 0')
WHERE (NEW.tile_column < 0) ;
SELECT RAISE(ABORT, 'insert on table """ + table_name + """ violates constraint: tile_column must by < matrix_width specified for table and zoom level in gpkg_tile_matrix')
WHERE NOT (NEW.tile_column < (SELECT matrix_width FROM gpkg_tile_matrix WHERE table_name = '""" + table_name + """' AND zoom_level = NEW.zoom_level));
END
"""
self.connection.execute(sql_string)
sql_string = """
CREATE TRIGGER '""" + table_name + """_tile_column_update'
BEFORE UPDATE OF tile_column ON '""" + table_name + """'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table """ + table_name + """ violates constraint: tile_column cannot be < 0')
WHERE (NEW.tile_column < 0) ;
SELECT RAISE(ABORT, 'update on table """ + table_name + """ violates constraint: tile_column must by < matrix_width specified for table and zoom level in gpkg_tile_matrix')
WHERE NOT (NEW.tile_column < (SELECT matrix_width FROM gpkg_tile_matrix WHERE table_name = '""" + table_name + """' AND zoom_level = NEW.zoom_level));
END
"""
self.connection.execute(sql_string)
sql_string = """
CREATE TRIGGER '""" + table_name + """_tile_row_insert'
BEFORE INSERT ON '""" + table_name + """'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table """ + table_name + """ violates constraint: tile_row cannot be < 0')
WHERE (NEW.tile_row < 0) ;
SELECT RAISE(ABORT, 'insert on table """ + table_name + """ violates constraint: tile_row must by < matrix_height specified for table and zoom level in gpkg_tile_matrix')
WHERE NOT (NEW.tile_row < (SELECT matrix_height FROM gpkg_tile_matrix WHERE table_name = '""" + table_name + """' AND zoom_level = NEW.zoom_level));
END
"""
self.connection.execute(sql_string)
sql_string = """
CREATE TRIGGER '""" + table_name + """_tile_row_update'
BEFORE UPDATE OF tile_row ON '""" + table_name + """'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table """ + table_name + """ violates constraint: tile_row cannot be < 0')
WHERE (NEW.tile_row < 0) ;
SELECT RAISE(ABORT, 'update on table """ + table_name + """ violates constraint: tile_row must by < matrix_height specified for table and zoom level in gpkg_tile_matrix')
WHERE NOT (NEW.tile_row < (SELECT matrix_height FROM gpkg_tile_matrix WHERE table_name = '""" + table_name + """' AND zoom_level = NEW.zoom_level));
END
"""
self.connection.execute(sql_string)
except sqlite3.Error as e:
print("Error inserting entries into gpkg_contents and/or other tables: {0}".format(e.args[0]))
return False
self.connection.commit()
for level in self.cache.level_infos:
try:
self.connection.execute(
"""
INSERT INTO gpkg_tile_matrix(table_name, zoom_level, matrix_width, matrix_height, tile_width,
tile_height, pixel_x_size, pixel_y_size)
VALUES(?, ?, ?, ?, ?, ?, ?, ?);
""",
(table_name, level.zoom_level, level.matrix_width, level.matrix_height,
self.tile_width, self.tile_height, level.pixel_x_size, level.pixel_y_size)
)
except sqlite3.Error as e:
print("Error inserting entry into gpkg_tile_matrix for overview {0}: {1}".format(level.zoom_level, e.args[0]))
return False
if not self.write_level(table_name, level):
print("Error writing full resolution tiles.")
return False
self.connection.commit()
return True
def write_level(self, table_name, level):
"""
Write one zoom/resolution level into pyramid data table.
@param table_name: Name of table to write pyramid data into.
@param zoom_level: Zoom/Resolution level to write.
@return: True on success, False on failure.
"""
for tile_row in range(level.startY, level.stopY):
for tile_column in range(level.startX, level.stopX):
tilePath = self.cache.getTilePath(tile_column, tile_row, level.zoom_level)
foundTilePath = self.cache.findTile(tilePath)
if foundTilePath is None:
if self.verbose:
print("{0}[.jpg/.png] not found, skipping.".format(tilePath))
else:
if not self.write_tile(foundTilePath, table_name, level.zoom_level,
(tile_row ) + level.offset_y,
(tile_column ) + level.offset_x):
print("Error writing image tiles for level {0} to database.".format(level.zoom_level))
return False
return True
def write_tile(self, filename, table_name, zoom_level, tile_row, tile_column):
"""
Extract specified tile from source dataset and write as a blob into GeoPackage, expanding colormap if required.
@param table_name: Name of table to write pyramid data into.
@param zoom_level: Zoom/Resolution level to write.
@param tile_row: Tile index (Y).
@param tile_column: Tile index (X).
@return: True on success, False on failure.
"""
size = os.stat(filename).st_size
if size == 0:
print("Tile {0} is 0 bytes, ignoring.".format(filename))
return True
try:
in_file = open(filename, 'rb')
tile_data = in_file.read(size)
in_file.close()
except IOError as e:
print("Error reading tile {0} : {1}".format(filename, e.args[0]))
return False
try:
self.connection.execute(
"""
INSERT INTO """ + table_name + """(zoom_level, tile_column, tile_row, tile_data)
VALUES (?, ?, ?, ?);
""",
(zoom_level, tile_column, tile_row, buffer(tile_data))
)
except sqlite3.Error as e:
print("Error inserting blob for tile {0}, {1}: {2}".format(tile_column, tile_row, e.args[0]))
return False
return True
def open(self, filename):
"""
Create or open a GeoPackage and create necessary tables and triggers.
@param filename: Name of sqlite3 database.
@return: True on success, False on failure.
"""
self.filename = filename
try:
self.connection = sqlite3.connect(filename)
except sqlite3.Error as e:
print("Error opening ", filename, ": ", e.args[0])
return False
self.connection.row_factory = sqlite3.Row
try:
self.connection.execute(
"""
PRAGMA application_id = 1196437808;
"""
)
self.connection.execute(
"""
CREATE TABLE IF NOT EXISTS gpkg_spatial_ref_sys ( \
srs_name TEXT NOT NULL, \
srs_id INTEGER NOT NULL PRIMARY KEY, \
organization TEXT NOT NULL, \
organization_coordsys_id INTEGER NOT NULL, \
definition TEXT NOT NULL, \
description TEXT );
"""
)
self.connection.execute(
"""
INSERT INTO gpkg_spatial_ref_sys(srs_name,srs_id,organization,organization_coordsys_id,definition)
SELECT 'Undefined Cartesian', -1, 'NONE', -1, 'undefined'
WHERE NOT EXISTS(SELECT 1 FROM gpkg_spatial_ref_sys WHERE srs_id=-1);
"""
)
self.connection.execute(
"""
INSERT INTO gpkg_spatial_ref_sys(srs_name,srs_id,organization,organization_coordsys_id,definition)
SELECT 'Undefined Geographic', 0, 'NONE', 0, 'undefined'
WHERE NOT EXISTS(SELECT 1 FROM gpkg_spatial_ref_sys WHERE srs_id=0);
"""
)
self.connection.execute(
"""
INSERT INTO gpkg_spatial_ref_sys(srs_name,srs_id,organization,organization_coordsys_id,definition)
SELECT 'WGS84', 4326, 'EPSG', 4326, 'GEOGCS["WGS 84",
DATUM["WGS_1984",
SPHEROID["WGS 84",6378137,298.257223563,
AUTHORITY["EPSG","7030"]],
AUTHORITY["EPSG","6326"]],
PRIMEM["Greenwich",0,
AUTHORITY["EPSG","8901"]],
UNIT["degree",0.01745329251994328,
AUTHORITY["EPSG","9122"]],
AUTHORITY["EPSG","4326"]]'
WHERE NOT EXISTS(SELECT 1 FROM gpkg_spatial_ref_sys WHERE srs_id=4326);
"""
)
self.connection.execute(
"""
CREATE TABLE IF NOT EXISTS gpkg_contents (
table_name TEXT NOT NULL PRIMARY KEY, \
data_type TEXT NOT NULL, \
identifier TEXT UNIQUE, \
description TEXT DEFAULT '', \
last_change DATETIME NOT NULL DEFAULT (strftime('%Y-%m-%dT%H:%M:%fZ',CURRENT_TIMESTAMP)), \
min_x DOUBLE, \
min_y DOUBLE, \
max_x DOUBLE, \
max_y DOUBLE, \
srs_id INTEGER, \
CONSTRAINT fk_gc_r_srs_id FOREIGN KEY (srs_id) REFERENCES gpkg_spatial_ref_sys(srs_id) );
"""
)
self.connection.execute(
"""
CREATE TABLE IF NOT EXISTS gpkg_tile_matrix (
table_name TEXT NOT NULL,
zoom_level INTEGER NOT NULL,
matrix_width INTEGER NOT NULL,
matrix_height INTEGER NOT NULL,
tile_width INTEGER NOT NULL,
tile_height INTEGER NOT NULL,
pixel_x_size DOUBLE NOT NULL,
pixel_y_size DOUBLE NOT NULL,
CONSTRAINT pk_ttm PRIMARY KEY (table_name, zoom_level),
CONSTRAINT fk_tmm_table_name FOREIGN KEY (table_name) REFERENCES gpkg_contents(table_name) );
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_zoom_level_insert'
BEFORE INSERT ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table ''gpkg_tile_matrix'' violates constraint: zoom_level cannot be less than 0')
WHERE (NEW.zoom_level < 0);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_zoom_level_update'
BEFORE UPDATE OF zoom_level ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table ''gpkg_tile_matrix'' violates constraint: zoom_level cannot be less than 0')
WHERE (NEW.zoom_level < 0);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_matrix_width_insert'
BEFORE INSERT ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table ''gpkg_tile_matrix'' violates constraint: matrix_width cannot be less than 1')
WHERE (NEW.matrix_width < 1);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_matrix_width_update'
BEFORE UPDATE OF matrix_width ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table ''gpkg_tile_matrix'' violates constraint: matrix_width cannot be less than 1')
WHERE (NEW.matrix_width < 1);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_matrix_height_insert'
BEFORE INSERT ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table ''gpkg_tile_matrix'' violates constraint: matrix_height cannot be less than 1')
WHERE (NEW.matrix_height < 1);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_matrix_height_update'
BEFORE UPDATE OF matrix_height ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table ''gpkg_tile_matrix'' violates constraint: matrix_height cannot be less than 1')
WHERE (NEW.matrix_height < 1);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_pixel_x_size_insert'
BEFORE INSERT ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table ''gpkg_tile_matrix'' violates constraint: pixel_x_size must be greater than 0')
WHERE NOT (NEW.pixel_x_size > 0);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_pixel_x_size_update'
BEFORE UPDATE OF pixel_x_size ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table ''gpkg_tile_matrix'' violates constraint: pixel_x_size must be greater than 0')
WHERE NOT (NEW.pixel_x_size > 0);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_pixel_y_size_insert'
BEFORE INSERT ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'insert on table ''gpkg_tile_matrix'' violates constraint: pixel_y_size must be greater than 0')
WHERE NOT (NEW.pixel_y_size > 0);
END
"""
)
self.connection.execute(
"""
CREATE TRIGGER IF NOT EXISTS 'gpkg_tile_matrix_pixel_y_size_update'
BEFORE UPDATE OF pixel_y_size ON 'gpkg_tile_matrix'
FOR EACH ROW BEGIN
SELECT RAISE(ABORT, 'update on table ''gpkg_tile_matrix'' violates constraint: pixel_y_size must be greater than 0')
WHERE NOT (NEW.pixel_y_size > 0);
END
"""
)
self.connection.execute(
"""
CREATE TABLE IF NOT EXISTS gpkg_tile_matrix_set (
table_name TEXT NOT NULL PRIMARY KEY,
srs_id INTEGER NOT NULL,
min_x DOUBLE NOT NULL,
min_y DOUBLE NOT NULL,
max_x DOUBLE NOT NULL,
max_y DOUBLE NOT NULL,
CONSTRAINT fk_gtms_table_name FOREIGN KEY (table_name) REFERENCES gpkg_contents(table_name),
CONSTRAINT fk_gtms_srs FOREIGN KEY (srs_id) REFERENCES gpkg_spatial_ref_sys (srs_id) );
"""
)
self.connection.commit()
except sqlite3.Error as e:
print("ERROR: SQLite error while creating core tables and triggers: {0}".format(e.args[0]))
return False
return True
def usage():
print("Usage: gdal2gpkg [-sr_org organization] [-sr_sysid identifier] [-sr_desc description]\n"
" [-v] path gpkgname")
return 2
def equal(a, b):
"""
Case insensitive string compare.
@param a: String to compare.
@param b: String to compare.
@return: True if equal, False if not.
"""
return a.lower() == b.lower()
def cache2gpkg(cache_path, gpkg_filename, verbose=False, **kwargs):
gpkg = GeoPackage()
gpkg.verbose = gpkg.cache.verbose = verbose
if kwargs is not None:
for key, value in kwargs.iteritems():
if equal(key, "sr_org"):
gpkg.sr_organization = value
elif equal(key, "sr_sysid"):
gpkg.sr_organization_coordsys_id = value
elif equal(key, "-sr_desc"):
gpkg.sr_description = value
if cache_path is None or gpkg_filename is None:
print("ERROR: Failed to open or create {0}".format(gpkg_filename))
return False
if not gpkg.open(gpkg_filename):
print("ERROR: Failed to open or create {0}".format(gpkg_filename))
return False
if not gpkg.add_cache(cache_path):
print("ERROR: Adding {0} to {1} failed".format(cache_path, gpkg_filename))
return False
gpkg = None
return True
|
<reponame>miquelramirez/tulip-control
"""
Tests for the abstraction from continuous dynamics to logic
"""
import logging
logging.basicConfig(level=logging.INFO)
logger = logging.getLogger(__name__)
# logging.getLogger('tulip').setLevel(logging.ERROR)
logger.setLevel(logging.DEBUG)
from nose.tools import assert_raises
import matplotlib
# to avoid the need for using: ssh -X when running tests remotely
matplotlib.use('Agg')
import networkx as nx
import numpy as np
from tulip import abstract
from tulip.abstract import feasible
from tulip import hybrid
import polytope as pc
input_bound = 0.4
def subsys0():
dom = pc.box2poly([[0., 3.], [0., 2.]])
A = np.eye(2)
B = np.eye(2)
U = pc.box2poly([[0., 1.],
[0., 1.]])
U.scale(input_bound)
sys_dyn = hybrid.LtiSysDyn(A, B, Uset=U, domain=dom)
return sys_dyn
def subsys1():
dom = pc.box2poly([[0., 3.], [0., 2.]])
A = np.eye(2)
B = np.eye(2)
U = pc.box2poly([[0., 0.],
[-1., 0.]])
U.scale(input_bound)
sys_dyn = hybrid.LtiSysDyn(A, B, Uset=U, domain=dom)
return sys_dyn
def transition_directions_test():
"""Unit test for correctness of abstracted transition directions, with:
- uni-directional control authority
- no disturbance
"""
modes = list()
modes.append(('normal', 'fly'))
modes.append(('refuel', 'fly'))
env_modes, sys_modes = zip(*modes)
# dynamics
cont_state_space = pc.box2poly([[0.0, 3.0], [0.0, 2.0]])
pwa_sys = dict()
pwa_sys[('normal', 'fly')] = hybrid.PwaSysDyn(
[subsys0()], cont_state_space)
pwa_sys[('refuel', 'fly')] = hybrid.PwaSysDyn(
[subsys1()], cont_state_space)
switched_dynamics = hybrid.SwitchedSysDyn(
disc_domain_size=(len(env_modes), len(sys_modes)),
dynamics=pwa_sys,
env_labels=env_modes,
disc_sys_labels=sys_modes,
cts_ss=cont_state_space)
# propositions
cont_props = dict()
cont_props['home'] = pc.box2poly([[0.0, 1.0], [0.0, 1.0]])
cont_props['lot'] = pc.box2poly([[2.0, 3.0], [1.0, 2.0]])
# partition
ppp = abstract.prop2part(cont_state_space, cont_props)
ppp, new2old = abstract.part2convex(ppp)
# configure discretization
N = 8
trans_len = 1
disc_params = dict()
for mode in modes:
disc_params[mode] = dict(N=N, trans_length=trans_len)
# discretize
swab = abstract.discretize_switched(
ppp, switched_dynamics, disc_params,
plot=True, show_ts=True, only_adjacent=False)
# assertions
ts = swab.modes[('normal', 'fly')].ts
edges = {(0, 0), (1, 1), (2, 2), (3, 3),
(4, 4), (5, 5),
(1, 2), (1, 4), (1, 5),
(2, 3), (2, 5), (2, 0),
(3, 0),
(4, 5),
(5, 0)}
h = nx.MultiDiGraph()
h.add_edges_from(edges)
assert nx.is_isomorphic(ts, h)
ts = swab.ts
assert nx.is_isomorphic(ts, h)
for _, _, d in ts.edges(data=True):
assert d['env_actions'] == 'normal'
assert d['sys_actions'] == 'fly'
transition_directions_test.slow = True
def test_transient_regions():
"""drift is too strong, so no self-loop must exist
This bug caused when running union is taken between Presets
during solve_feasible, as happened with old use_all_horizon,
cf:
- 5b1e9681918739b276a221fcc1fd6eebfd058ce3
- f5f4934ab9d21062f633eef3861ad935c3d3b54b
"""
dom = pc.box2poly([[0.0, 4.0], [0.0, 3.0]])
def cont_predicates():
p = dict()
p['safe'] = pc.box2poly([[0.5, 3.5], [0.5, 2.5]])
ppp = abstract.prop2part(dom, p)
ppp, new2old_reg = abstract.part2convex(ppp)
ppp.plot()
return ppp
def drifting_dynamics():
A = np.array([[1.0, 0.0],
[0.0, 1.0]])
B = np.array([[1.0],
[0.0]])
U = pc.box2poly([[0.0, 1.0]])
K = np.array([[-100.0],
[0.0]])
sys = hybrid.LtiSysDyn(A, B, None, K, U, None, dom)
return sys
ppp = cont_predicates()
sys = drifting_dynamics()
logger.info(sys)
ab = abstract.discretize(ppp, sys, N=1, use_all_horizon=True,
trans_length=1)
logger.debug(ab.ts)
self_loops = {i for i, j in ab.ts.transitions() if i == j}
logger.debug('self loops at states: ' + str(self_loops))
assert(not self_loops)
# ax = ab.plot(show_ts=True)
# ax.figure.savefig('./very_simple.pdf')
def define_partition(dom):
p = dict()
p['a'] = pc.box2poly([[0.0, 10.0], [15.0, 18.0]])
p['b'] = pc.box2poly([[0.0, 1.0], [0.0, 20.0]])
ppp = abstract.prop2part(dom, p)
ppp, new2old_reg = abstract.part2convex(ppp)
return ppp
def define_dynamics(dom):
A = np.eye(2)
B = np.array([[1.0, -1.0],
[0.0, +1.0]])
U = pc.box2poly([[0.0, 3.0],
[-3.0, 3.0]])
E = np.array([[0.0],
[-1.0]])
W = pc.box2poly([[-1.0, 1.0]])
W.scale(0.4)
K = np.array([[0.0],
[-0.4]])
sys = hybrid.LtiSysDyn(A, B, E, K, U, W, dom)
return sys
def define_dynamics_dual():
# Continuous state space
cont_state_space = pc.box2poly([[-1.5, 1.5]])
# Continuous dynamics
# (continuous-state, discrete-time)
A = np.array([[2]])
B = np.array([[1]])
# Available control, possible disturbances
U = np.array([[-2.0, 2.0]])
# Convert to polyhedral representation
U = pc.box2poly(U)
# Construct the LTI system describing the dynamics
sys_dyn = hybrid.LtiSysDyn(A, B, None, None, U, None, cont_state_space)
# @dynamics_section_end@
# @partition_section@
# Define atomic propositions for relevant regions of state space
cont_props = {}
cont_props['a'] = pc.box2poly([[-1.5, -1]])
cont_props['b'] = pc.box2poly([[-1, 1]])
cont_props['c'] = pc.box2poly([[1, 1.5]])
part = []
part.append(pc.box2poly([[-1.5, -1]]))
part.append(pc.box2poly([[-1, 1]]))
part.append(pc.box2poly([[1, 1.5]]))
part.append(pc.box2poly([[-1, 0.5]]))
part.append(pc.box2poly([[-0.5, 1]]))
part.append(pc.box2poly([[-0.5, 0.5]]))
part.append(pc.box2poly([[-1.25, -1]]))
part.append(pc.box2poly([[1, 1.25]]))
# Compute the proposition preserving partition of the continuous state
# space
cont_partition = abstract.prop2part(cont_state_space, cont_props)
return sys_dyn, cont_partition, part
def test_abstract_the_dynamics():
"""test_abstract_the_dynamics (known to fail without GLPK)"""
dom = pc.box2poly([[0.0, 10.0], [0.0, 20.0]])
ppp = define_partition(dom)
sys = define_dynamics(dom)
logger.info(sys)
disc_options = {'N': 3, 'trans_length': 2, 'min_cell_volume': 1.5}
ab = abstract.discretize(ppp, sys, plotit=False,
save_img=False, **disc_options)
assert ab.ppp.compute_adj()
assert ab.ppp.is_partition()
# ax = ab.plot(show_ts=True, color_seed=0)
# sys.plot(ax, show_domain=False)
# print(ab.ts)
# self_loops = {i for i,j in ab.ts.transitions() if i==j}
# print('self loops at states: ' + str(self_loops))
test_abstract_the_dynamics.slow = True
def test_abstract_the_dynamics_dual():
"""test_abstract_the_dynamics using dual-simulation algorithm"""
dom = pc.box2poly([[0.0, 10.0], [0.0, 20.0]])
ppp = define_partition(dom)
sys = define_dynamics(dom)
logger.info(sys)
disc_options = {'N': 3, 'trans_length': 2, 'min_cell_volume': 1.5}
ab = abstract.discretize(ppp, sys, plotit=False,
save_img=False, simu_type='dual', **disc_options)
assert ab.ppp.compute_adj()
[sys_dyn, cont_partition, part] = define_dynamics_dual()
disc_options = {'N': 1, 'trans_length': 1000, 'min_cell_volume': 0.0}
ab_2 = abstract.discretize(cont_partition, sys_dyn,
simu_type='dual', **disc_options)
table = np.zeros([len(part), 1])
for i in ab_2.ppp.regions:
for j in range(len(part)):
if i == part[j]:
table[j] = 1
assert np.sum(table) == len(part)
test_abstract_the_dynamics_dual.slow = True
def test_is_feasible():
"""Difference between attractor and fixed horizon."""
dom = pc.box2poly([[0.0, 4.0], [0.0, 3.0]])
sys = drifting_dynamics(dom)
p1 = pc.box2poly([[0.0, 1.0], [0.0, 1.0]])
p2 = pc.box2poly([[2.0, 3.0], [0.0, 1.0]])
n = 10
r = feasible.is_feasible(p1, p2, sys, n, use_all_horizon=False)
assert r is False, r
r = feasible.is_feasible(p1, p2, sys, n, use_all_horizon=True)
assert r is True, r
def drifting_dynamics(dom):
A = np.array([[1.0, 0.0],
[0.0, 1.0]])
B = np.array([[1.0],
[0.0]])
U = pc.box2poly([[0.0, 1.0]])
K = np.array([[1.0],
[0.0]])
sys = hybrid.LtiSysDyn(A, B, None, K, U, None, dom)
return sys
if __name__ == '__main__':
test_abstract_the_dynamics()
test_abstract_the_dynamics_dual()
|
'''
Created on Nov 15, 2010
@author: octi
'''
import dbbgm_batch
import multiprocessing
import Queue
from bbgm_utils import saveImg,dbvalue,openImage,frameSuffix,frameSuffixWrite
class GUIInvoker(multiprocessing.Process):
def __init__(self,model,inv_model,fwd_model,pMap):
self.mdl=model
self.invmdl=inv_model
self.fwd=fwd_model
self.pmap=pMap
multiprocessing.Process.__init__(self)
def run(self):
dbbgm_batch.init_process("bbgmInvokeGUIProcess");
dbbgm_batch.set_input_from_db(0,self.mdl);
dbbgm_batch.set_input_from_db(1,self.invmdl);
dbbgm_batch.set_input_from_db(2,self.fwd);
dbbgm_batch.set_input_from_db(3,self.pmap);
print dbbgm_batch.run_process();
return []
def executeJobs(jobs, num_procs=5):
work_queue=multiprocessing.Queue();
result_queue=multiprocessing.Queue();
for job in jobs:
work_queue.put(job)
for i in range(num_procs):
worker= measureProbWorker(work_queue,result_queue)
worker.start();
print("worker with name ",worker.name," started!")
class bbgmJob():
def __init__(self,wavelet,attribute,tolerance,interp_functor,data_path,input_path,output_path):
self.wavelet=wavelet;
self.test_image_path=input_path;
self.attribute=attribute;
self.tolerance=tolerance;
self.interp_functor=interp_functor;
self.data_path=data_path;
self.output_path=output_path;
class measureProbWorker(multiprocessing.Process):
def __init__(self,work_queue,result_queue):
# base class initialization
multiprocessing.Process.__init__(self)
# job management stuff
self.work_queue = work_queue
self.result_queue = result_queue
self.kill_received = False
def run(self):
while not self.kill_received:
# get a task
try:
job = self.work_queue.get_nowait()
except Queue.Empty:
break
# store the result
[test_image,ni,nj]=openImage(job.test_image_path);
dbbgm_batch.init_process("bbgmMeasureWvLookupProcess");
dbbgm_batch.set_input_from_db(0,job.wavelet);
dbbgm_batch.set_input_from_db(1,test_image);
dbbgm_batch.set_input_string(2,job.attribute);
dbbgm_batch.set_input_float(3,job.tolerance);
dbbgm_batch.set_input_string(4,job.interp_functor);
dbbgm_batch.set_input_string(5,job.data_path);
print dbbgm_batch.run_process();
out_image=dbvalue(0,"")
(out_image.id,out_image.type)=dbbgm_batch.commit_output(0);
saveImg(out_image,job.output_path)
class measureProbQueue(multiprocessing.Process):
def __init__(self,wavelet,attribute,tolerance,interp_functor,data_path,input_path,output_path,input_queue):
multiprocessing.Process.__init__(self)
self.wavelet=wavelet;
self.input_path=input_path
self.attribute=attribute;
self.tolerance=tolerance;
self.interp_functor=interp_functor;
self.data_path=data_path;
self.input_queue=input_queue;
self.output_path=output_path;
self.kill_received=False
def run(self):
while not self.kill_received:
try:
index = self.input_queue.get_nowait()
except Queue.Empty:
break
# store the result
[test_image,ni,nj]=openImage(self.input_path+frameSuffix(index,5)+".tiff");
dbbgm_batch.init_process("bbgmMeasureWvLookupProcess");
dbbgm_batch.set_input_from_db(0,self.wavelet);
dbbgm_batch.set_input_from_db(1,test_image);
dbbgm_batch.set_input_string(2,self.attribute);
dbbgm_batch.set_input_float(3,self.tolerance);
dbbgm_batch.set_input_string(4,self.interp_functor);
dbbgm_batch.set_input_string(5,self.data_path);
print dbbgm_batch.run_process();
out_image=dbvalue(0,"")
(out_image.id,out_image.type)=dbbgm_batch.commit_output(0)
saveImg(out_image,self.output_path+frameSuffix(index,3)+".tiff")
class measureProb(multiprocessing.Process):
def __init__(self,wavelet,test_image,attribute,tolerance,interp_functor,data_path,output_path,index):
multiprocessing.Process.__init__(self)
self.wavelet=wavelet;
self.test_image=test_image;
self.attribute=attribute;
self.tolerance=tolerance;
self.interp_functor=interp_functor;
self.data_path=data_path;
self.output_path=output_path;
self.index=index
self.done=0;
def run(self):
dbbgm_batch.init_process("bbgmMeasureWvLookupProcess");
dbbgm_batch.set_input_from_db(0,self.wavelet);
dbbgm_batch.set_input_from_db(1,self.test_image);
dbbgm_batch.set_input_string(2,self.attribute);
dbbgm_batch.set_input_float(3,self.tolerance);
dbbgm_batch.set_input_string(4,self.interp_functor);
dbbgm_batch.set_input_string(5,self.data_path);
print dbbgm_batch.run_process();
out_image=dbvalue(0,"")
(out_image.id,out_image.type)=dbbgm_batch.commit_output(0)
saveImg(out_image,self.output_path+frameSuffixWrite(self.index,300)+".tiff")
|
<filename>vpc.py
import boto3
import pprint
import sys
ec2_client = boto3.client('ec2')
ec2_res = boto3.resource('ec2')
def createVpc(offset):
vpc = ec2_res.create_vpc(CidrBlock = '10.' + str(offset) + '.0.0/16')
vpc.create_tags(
Tags = [ { 'Key': 'Name', 'Value': 'VPC-' + str(offset) }, ]
)
vpc.wait_until_available()
return(vpc)
def destroyVpc(vpcid):
print('Removing VPC ({}) from AWS'.format(vpcid))
ec2 = boto3.resource('ec2')
ec2client = ec2.meta.client
vpc = ec2.Vpc(vpcid)
# detach and delete all gateways associated with the vpc
for gw in vpc.internet_gateways.all():
print('Removing igw {}'.format(gw.id))
vpc.detach_internet_gateway(InternetGatewayId=gw.id)
gw.delete()
# delete all route table associations
for rt in vpc.route_tables.all():
for rta in rt.associations:
if not rta.main:
rta.delete()
# delete any instances
for subnet in vpc.subnets.all():
for instance in subnet.instances.all():
instance.terminate()
# delete our endpoints
for ep in ec2client.describe_vpc_endpoints(
Filters=[{
'Name': 'vpc-id',
'Values': [vpcid]
}])['VpcEndpoints']:
ec2client.delete_vpc_endpoints(VpcEndpointIds=[ep['VpcEndpointId']])
# delete our security groups
for sg in vpc.security_groups.all():
if sg.group_name != 'default':
sg.delete()
# delete any vpc peering connections
for vpcpeer in ec2client.describe_vpc_peering_connections(
Filters=[{
'Name': 'requester-vpc-info.vpc-id',
'Values': [vpcid]
}])['VpcPeeringConnections']:
ec2.VpcPeeringConnection(vpcpeer['VpcPeeringConnectionId']).delete()
# delete non-default network acls
for netacl in vpc.network_acls.all():
if not netacl.is_default:
netacl.delete()
# delete network interfaces
for subnet in vpc.subnets.all():
for interface in subnet.network_interfaces.all():
interface.delete()
subnet.delete()
# finally, delete the vpc
ec2client.delete_vpc(VpcId=vpcid)
print(" VPC deleted")
def bulkCreate(qty):
vpc_id_list = []
for i in range(qty):
vpc = createVpc(i)
print("Created VPC {}".format(vpc.id))
igw = ec2_res.create_internet_gateway()
vpc.attach_internet_gateway(InternetGatewayId=igw.id)
default_rt = ec2_client.describe_route_tables(Filters=[{'Name': 'vpc-id','Values': [vpc.id,]},])['RouteTables'][0]['RouteTableId']
print(" default route table is is {}".format(default_rt))
rt = ec2_res.RouteTable(default_rt)
route = rt.create_route(DestinationCidrBlock='0.0.0.0/0',
GatewayId = igw.id)
subnetString = "10.{}.{}.0/24".format(i,i)
subnet = ec2_res.create_subnet(CidrBlock=subnetString, VpcId=vpc.id)
rt.associate_with_subnet(SubnetId=subnet.id)
sec_group = ec2_res.create_security_group(GroupName='slice_0',
Description='slice_0 sec group',
VpcId=vpc.id)
sec_group.authorize_ingress(CidrIp='0.0.0.0/0',
IpProtocol='icmp',
FromPort=-1,
ToPort=-1)
vpc_id_list.append(vpc.id)
return(vpc_id_list)
allVpcs = ec2_client.describe_vpcs()['Vpcs']
print("Found {} VPCs".format(len(allVpcs)))
for vpc in allVpcs:
print("VPC id {}".format(vpc['VpcId']))
vpcList = bulkCreate(4)
user = input("Delete VPCs now [Y/y]")
for vpc in vpcList:
destroyVpc(vpc)
|
"""
This is a script to test a particle simulation.
Please change any ..._path to your corresponding file path.
"""
import sys
sys.path.append('..')
import numpy as np
import matplotlib.pyplot as pp
from flow import Flow
from animationparticles import AnimationParticles
from text.text_particles import read_particles, write_particles
from test_multiple_particles_scripts import prepare_initial_positions, spread_particles
from axes_world import one_by_two
#==============================================================================
# Prepare flow
parent_folder = r'../'
case = 'uniform'
flow_path = parent_folder + 'flows/%s' % case
elements_path = parent_folder + 'elements/%s' % case
geometries_path = parent_folder + 'geometries/%s' % case
#flow = Flow()
#flow.load_flow(flow_path, elements_path, geometries_path)
# Prepare particle
diameter = 0.1
density = 2.0
births = range(0,1)
lifetime = 20
# Initial values
x0 = -0.75
list_y = np.array([0.])
#list_y = np.array([0.2, 0.423, 0.6])
#list_y = np.array([0.415, 0.6])
initial_positions = prepare_initial_positions(x0, list_y)
#u0 = None
u0 = np.array([0,0.])
# Arguments to create a list of particles
fill_args = diameter, density, births, lifetime, initial_positions, u0
# Ready to compute
list_factors = np.array([1,2,4,8,16,32,64,128])
#list_trajectories = []
#for factor in list_factors:
# particles, captured_ones = spread_particles(fill_args, flow, factor,
# printIt=False, too_far_stop=False)
#
# traj = particles[0].trajectory
# list_trajectories.append(traj)
#
#delta_t = 0.1
#def one_each_two(l):
# return np.array([l[i] for i in range(0,len(l),2)])
#
#list_epsilon = []
#for i in range(len(list_trajectories)-1):
# traj_coarse, traj_fine = list_trajectories[i], list_trajectories[i+1]
#
# epsilon = np.sqrt( delta_t/list_factors[i] \
# * np.sum( (traj_coarse - one_each_two(traj_fine))**2 ) )
# list_epsilon.append(epsilon)
#list_epsilon_big = np.array(list_epsilon)
# =============================================================================
def plot_discretisation_error_L2(ax, list_dt, errorsL2, color, marker):
ax.plot(list_dt[1:], errorsL2, linestyle='-',
linewidth=1, color=color, marker=marker, markeredgecolor='black',
markeredgewidth=0.5, alpha=0.75)
ax.set(xscale='log', yscale='log')
ylabel = r'$\varepsilon_{\Delta t}$'
ax.set_ylabel(ylabel, fontsize=12)
ax.set_xlabel(r'$\Delta t$', fontsize=12)
def plot_estimated_order_L2(ax, list_dt, errorsL2, color, marker):
obs_p = np.log(errorsL2[:-1]/errorsL2[1:])/np.log(2)
print(obs_p)
ax.plot(list_dt[2:], obs_p, linestyle='', linewidth=1,
color=color, marker=marker, markeredgecolor='black',
markeredgewidth=0.5, alpha=0.75)
ax.set_xscale('log')
ax.set_ylim([0.89, 1.11])
# ax.set_ylim([1.89, 2.11])
ax.set_yticks([0.9, 0.95, 1, 1.05, 1.1])
ax.set_xlabel(r'$\Delta t$', fontsize=12)
ax.set_ylabel(r'$\hat{p}_{\Delta t}$', fontsize=12)
# =============================================================================
list_dt = 0.1/list_factors
ax_a, ax_b = one_by_two()
ax_a.plot(list_dt[1:5], 2e-4*list_dt[:4], linestyle='--',
color='black')
ax_b.plot(list_dt[2:], 1 + 0*list_dt[2:], linestyle='--',
color='black')
plot_discretisation_error_L2(ax_a, list_dt, list_epsilon_small, 'blue', '.')
plot_estimated_order_L2(ax_b, list_dt, list_epsilon_small, 'blue', '.')
plot_discretisation_error_L2(ax_a, list_dt, list_epsilon_big, 'gray', 'o')
plot_estimated_order_L2(ax_b, list_dt, list_epsilon_big, 'gray', 'o')
|
import datetime
import logging
import Mollie
from django.conf import settings
from django.contrib.auth.decorators import login_required
from django.template.context_processors import csrf
from django.core.urlresolvers import reverse
from django.http import HttpResponse, HttpResponseRedirect
from django.shortcuts import render_to_response, get_object_or_404
from django.utils.html import escape
from django.utils.safestring import mark_safe
from django.utils.translation import ugettext as _
from django.views.decorators.csrf import csrf_exempt
from django.views.generic.list import ListView
from subscribe.models import Event, EventQuestion
from subscribe.forms import Registration, SubscribeForm, fill_subscription
def event_message(request, event, message):
c = {"event": event, "request": request, "message": message}
return render_to_response("subscribe/event_message.html", c)
def register(request, slug):
logger = logging.getLogger(__name__)
# Get the event
event = get_object_or_404(Event, slug=slug)
# If not staff, check if allowed
if not request.user.is_staff:
now = datetime.datetime.now()
if event.start_registration > now or event.end_registration < now:
return event_message(request, event, _("Inschrijving gesloten."))
if event.is_full():
return event_message(request, event, _("Helaas is het maximum aantal inschrijvingen bereikt."))
# If this is not a POST request
if request.method != "POST":
# then just create the form...
form = SubscribeForm(event)
c = {"event": event, "request": request, "form": form, "user_is_staff": request.user.is_staff}
c.update(csrf(request))
return render_to_response("subscribe/form.html", c)
# It is a POST request, check if the form is valid...
form = SubscribeForm(event, request.POST)
if not form.is_valid():
c = {"event": event, "request": request, "form": form, "user_is_staff": request.user.is_staff}
c.update(csrf(request))
return render_to_response("subscribe/form.html", c)
# It is a POST request, and the form is valid, check if this is the confirmation page
if 'registration_preview' not in request.POST and not form.is_free_event:
form.confirm_page()
c = {"event": event, "request": request, "form": form, "user_is_staff": request.user.is_staff}
c.update(csrf(request))
return render_to_response("subscribe/form.html", c)
# Maybe this is just a test of the confirmation email?
if 'testsubmit' in request.POST:
subscription = fill_subscription(form, event)
msg = subscription.send_confirmation_email()
subscription.delete()
msg = '<br/>'.join(escape(msg).split('\n'))
return event_message(request, event, mark_safe("De volgende email is verstuurd:<br/><br/>{}".format(msg)))
# We confirmed. Create the subscription in the database...
subscription = fill_subscription(form, event)
# Check if the subscription form could be saved
if not subscription:
# Error Filling subscription
error_str = "Error in saving form."
return HttpResponse(_(error_str))
# Check (again) if maybe the number of registrations is over the limit...
if subscription in event.get_registrations_over_limit():
subscription.delete()
if event.is_full():
error_str = "De inschrijving kan niet worden voltooid, omdat het maximum aantal inschrijvingen is bereikt."
else:
error_str = "De inschrijving kan niet worden voltooid, omdat een van de gekozen opties het maximum aantal inschrijvingen heeft bereikt."
return event_message(request, event, _(error_str))
# Check if we need to pay or not...
if subscription.price <= 0:
subscription.paid = True
subscription.send_confirmation_email()
subscription.save()
return event_message(request, event, _("Inschrijving geslaagd. Ter bevestiging is een e-mail verstuurd."))
# Payment required...
try:
mollie = Mollie.API.Client()
mollie.setApiKey(settings.MOLLIE_KEY)
# METADATA TOEVOEGEN
webhookUrl = request.build_absolute_uri(reverse("webhook", args=[subscription.id]))
redirectUrl = request.build_absolute_uri(reverse("return_page", args=[subscription.id]))
payment = mollie.payments.create({
'amount': float(subscription.price) / 100.0,
'description': subscription.event.name,
'webhookUrl': webhookUrl,
'redirectUrl': redirectUrl,
})
subscription.trxid = payment["id"]
subscription.save()
return HttpResponseRedirect(payment.getPaymentUrl())
except Mollie.API.Error as e:
error_str = "register: Technische fout, probeer later opnieuw.\n\n" + str(e)
logger.error(error_str)
return event_message(request, event, _(error_str))
def check_transaction(subscription):
logger = logging.getLogger(__name__)
logger.info('check_transaction: Checking transaction %d with id %s' % (subscription.id, subscription.trxid))
mollie = Mollie.API.Client()
mollie.setApiKey(settings.MOLLIE_KEY)
payment = mollie.payments.get(subscription.trxid)
logger.info("check_transaction: Transaction %s has status %s" % (subscription.id, payment['status']))
subscription.status = payment['status']
subscription.paid = payment.isPaid()
subscription.save()
if subscription.paid:
subscription.send_confirmation_email()
# called when the user returns from Mollie
def return_page(request, id):
logger = logging.getLogger(__name__)
logger.info('views::return_page() - registration id: ' + str(id))
# Retrieve the registration
try:
subscription = Registration.objects.get(id=id)
except:
return HttpResponse(_("iDEAL error (onbekende inschrijving): Neem contact op met <EMAIL>. Controleer of uw betaling is afgeschreven alvorens de betaling opnieuw uit te voeren."))
# If status unknown, then check it...
if subscription.status == "":
try:
check_transaction(subscription)
except Mollie.API.Error as e:
error_str = "return_page: Technische fout, probeer later opnieuw." + "\n\n%s" % (str(e),)
logger.error(error_str)
return event_message(request, subscription.event, _(error_str))
if subscription.status == "paid":
return event_message(request, subscription.event, _("Betaling geslaagd. Ter bevestiging is een e-mail verstuurd."))
elif subscription.status == "cancelled" or subscription.status == "expired":
return event_message(request, subscription.event, _("Je betaling is geannuleerd."))
elif subscription.status == "open" or subscription.status == "pending":
return event_message(request, subscription.event, _("Je betaling staat geregistreerd in ons systeem, maar wordt nog verwerkt door onze bank. Als je binnen een uur geen bevestigingsmail ontvangt, is er mogelijk iets fout gegaan met de betaling. Neem in dat geval contact op met <EMAIL>."))
else:
return event_message(request, subscription.event, _("Er is een fout opgetreden bij het verwerken van je iDEAL transactie. Neem contact op met <EMAIL> of probeer het later nogmaals. Controleer of je betaling is afgeschreven alvorens de betaling opnieuw uit te voeren."))
@csrf_exempt
def webhook(request, id):
# trigger checking
if request.method == "POST":
transaction_id = request.POST['id']
else:
transaction_id = request.GET['id']
logger = logging.getLogger(__name__)
logger.info('views::check() - id: %s, transaction id: %s' % (id, transaction_id))
try:
subscription = Registration.objects.get(id=id, trxid=transaction_id)
except:
logger.error("views::check() - cannot find matching subscription")
return HttpResponse(_("NOT OK"))
try:
check_transaction(subscription)
except Mollie.API.Error as e:
logger.error("webhook: error %s" % (str(e),))
return HttpResponse(_("OK"))
@login_required
def delete_event_question(request):
questionId = request.GET['questionId']
warning = int(request.GET['warning'])
if warning == 0:
eventQuestion = EventQuestion.objects.get(pk=questionId)
eventQuestion.delete()
return HttpResponse(_('Vraag verwijderd. <br /> <a href="/admin/">Terug naar admin.</a>'))
else:
return HttpResponse(_("""Weet je zeker dat je deze vraag wilt verwijderen? <br />
<a href="/deleteEventQuestion/?questionId=%d&warning=%d">Ja</a>
<a href="/admin/">Nee</a>""" % (int(questionId), 0)))
class HomeView(ListView):
model = Event
queryset = Event.objects.order_by('-end_registration', '-start_registration')
template_name = "subscribe/index.html"
context_object_name = "events"
def get(self, request, *args, **kwargs):
if not request.user.is_staff:
now = datetime.datetime.now()
self.queryset = self.queryset.filter(start_registration__lte=now, end_registration__gte=now)
return super().get(self, request, *args, **kwargs)
|
# copyright (c) 2021 PaddlePaddle Authors. All Rights Reserve.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Functions for computing metrics like precision, recall, CorLoc and etc."""
import numpy as np
def compute_precision_recall(scores, labels, num_gt):
"""Compute precision and recall.
Args:
scores: A float numpy array representing detection score
labels: A boolean numpy array representing true/false positive labels
num_gt: Number of ground truth instances
Raises:
ValueError: if the input is not of the correct format
Returns:
precision: Fraction of positive instances over detected ones. This
value is None if no ground truth labels are present.
recall: Fraction of detected positive instance over all positive
instances. This value is None if no ground truth labels are
present.
"""
if (not isinstance(labels, np.ndarray) or labels.dtype != np.bool
or len(labels.shape) != 1):
raise ValueError('labels must be single dimension bool numpy array')
if not isinstance(scores, np.ndarray) or len(scores.shape) != 1:
raise ValueError('scores must be single dimension numpy array')
if num_gt < np.sum(labels):
raise ValueError(
'Number of true positives must be smaller than num_gt.')
if len(scores) != len(labels):
raise ValueError('scores and labels must be of the same size.')
if num_gt == 0:
return None, None
sorted_indices = np.argsort(scores)
sorted_indices = sorted_indices[::-1]
labels = labels.astype(int)
true_positive_labels = labels[sorted_indices]
false_positive_labels = 1 - true_positive_labels
cum_true_positives = np.cumsum(true_positive_labels)
cum_false_positives = np.cumsum(false_positive_labels)
precision = cum_true_positives.astype(float) / (
cum_true_positives + cum_false_positives)
recall = cum_true_positives.astype(float) / num_gt
return precision, recall
def compute_average_precision(precision, recall):
"""Compute Average Precision according to the definition in VOCdevkit.
Precision is modified to ensure that it does not decrease as recall
decrease.
Args:
precision: A float [N, 1] numpy array of precisions
recall: A float [N, 1] numpy array of recalls
Raises:
ValueError: if the input is not of the correct format
Returns:
average_precison: The area under the precision recall curve. NaN if
precision and recall are None.
"""
if precision is None:
if recall is not None:
raise ValueError('If precision is None, recall must also be None')
return np.NAN
if not isinstance(precision, np.ndarray) or not isinstance(
recall, np.ndarray):
raise ValueError('precision and recall must be numpy array')
if precision.dtype != np.float or recall.dtype != np.float:
raise ValueError('input must be float numpy array.')
if len(precision) != len(recall):
raise ValueError('precision and recall must be of the same size.')
if not precision.size:
return 0.0
if np.amin(precision) < 0 or np.amax(precision) > 1:
raise ValueError('Precision must be in the range of [0, 1].')
if np.amin(recall) < 0 or np.amax(recall) > 1:
raise ValueError('recall must be in the range of [0, 1].')
if not all(recall[i] <= recall[i + 1] for i in range(len(recall) - 1)):
raise ValueError('recall must be a non-decreasing array')
recall = np.concatenate([[0], recall, [1]])
precision = np.concatenate([[0], precision, [0]])
# Preprocess precision to be a non-decreasing array
for i in range(len(precision) - 2, -1, -1):
precision[i] = np.maximum(precision[i], precision[i + 1])
indices = np.where(recall[1:] != recall[:-1])[0] + 1
average_precision = np.sum(
(recall[indices] - recall[indices - 1]) * precision[indices])
return average_precision
def compute_cor_loc(num_gt_imgs_per_class,
num_images_correctly_detected_per_class):
"""Compute CorLoc according to the definition in the following paper.
https://www.robots.ox.ac.uk/~vgg/rg/papers/deselaers-eccv10.pdf
Returns nans if there are no ground truth images for a class.
Args:
num_gt_imgs_per_class: 1D array, representing number of images
containing at least one object instance of a particular class
num_images_correctly_detected_per_class: 1D array, representing number
of images that are correctly detected at least one object instance
of a particular class
Returns:
corloc_per_class: A float numpy array represents the corloc score of
each class
"""
# Divide by zero expected for classes with no gt examples.
with np.errstate(divide='ignore', invalid='ignore'):
return np.where(
num_gt_imgs_per_class == 0, np.nan,
num_images_correctly_detected_per_class / num_gt_imgs_per_class)
|
# Copyright 2013 Cisco Systems, Inc.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
This module contains APIs to facilitate UcsCentral backup and import
"""
import os
import platform
import time
import datetime
import logging
from ..ucscexception import UcscValidationException, \
UcscWarning, \
UcscOperationError
log = logging.getLogger('ucscentral')
ucsc_base_dn = "org-root/deviceprofile-default"
def _validate_remote_host_args(protocol, hostname, username, password):
if not protocol:
raise UcscValidationException("Missing protocol argument")
if not hostname:
raise UcscValidationException("Missing hostname argument")
if protocol == 'tftp':
return
if not username:
raise UcscValidationException("Missing username argument")
if not password:
raise UcscValidationException("Missing password argument")
def _backup(handle, file_dir, file_name, timeout=600,
remote_enabled=False, protocol=None,
hostname="localhost", username=None, password="",
remove_from_ucsc=False,
preserve_pooled_values=False):
"""
_backup internal method helps create UcsCentral full-state backup and
download it locally or to remote location.
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download backup file to
file_name (str): name for the backup file
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
remote_enabled (boolean): True if Remote backup is enabled
False - by default
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
username (str): Username for remote backup
password (str): <PASSWORD> remote <PASSWORD>
remove_from_ucsc (boolean): True/False, False - by default
preserve_pooled_values (boolean): True/False, False - by default
Example:
file_dir = "/home/user/backup"\n
file_name = "config_backup.tgz"\n
_backup(handle, file_dir=file_dir, file_name=file_name)\n
_backup(handle, file_dir=file_dir, file_name=file_name,
remote_enabled=True, protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
from ..mometa.mgmt.MgmtBackup import MgmtBackup, MgmtBackupConsts
from ..mometa.top.TopSystem import TopSystem
backup_type = "full-state"
if not file_dir:
raise UcscValidationException("Missing file_dir argument")
if not file_name:
raise UcscValidationException("Missing file_name argument")
top_system = TopSystem()
if remote_enabled:
_validate_remote_host_args(protocol, hostname, username, password)
if (not file_name.endswith('.tgz')):
raise UcscValidationException(
"file_name must be .tgz format")
file_path = os.path.join(file_dir, file_name)
mgmt_backup = MgmtBackup(
parent_mo_or_dn=top_system,
hostname=hostname,
admin_state=MgmtBackupConsts.ADMIN_STATE_ENABLED,
proto=protocol,
type=backup_type,
remote_file=file_path,
user=username,
pwd=password)
else:
if not os.path.exists(file_dir):
os.makedirs(file_dir)
file_string = platform.node().lower() \
+ datetime.datetime.now().strftime("%Y%m%d%H%M%S")
file_path = "/" + file_string + "_" + backup_type + "_backup.tgz"
mgmt_backup = MgmtBackup(
parent_mo_or_dn=top_system,
hostname=hostname,
admin_state=MgmtBackupConsts.ADMIN_STATE_ENABLED,
proto=MgmtBackupConsts.PROTO_HTTP,
type=backup_type,
remote_file=file_path)
if preserve_pooled_values:
mgmt_backup.preserve_pooled_values = \
MgmtBackupConsts.PRESERVE_POOLED_VALUES_YES
else:
mgmt_backup.preserve_pooled_values = \
MgmtBackupConsts.PRESERVE_POOLED_VALUES_NO
handle.add_mo(mgmt_backup, modify_present=True)
handle.commit()
mgmt_backup = handle.query_dn(dn=mgmt_backup.dn)
admin_state_temp = mgmt_backup.admin_state
# Checking for the backup to compete.
duration = timeout
poll_interval = 2
log.debug("Starting Backup ")
while True:
mgmt_backup = handle.query_dn(dn=mgmt_backup.dn)
admin_state_temp = mgmt_backup.admin_state
# Break condition:- if state id disabled then break
if admin_state_temp == MgmtBackupConsts.ADMIN_STATE_DISABLED:
break
time.sleep(min(duration, poll_interval))
duration = max(0, (duration - poll_interval))
if duration == 0:
handle.remove_mo(mgmt_backup)
handle.commit()
raise UcscOperationError(
"Backup UcsCentral", " operation timed out")
if remote_enabled:
if mgmt_backup.over_all_status == \
MgmtBackupConsts.OVER_ALL_STATUS_FAILED:
log.debug("Backup failed")
handle.remove_mo(mgmt_backup)
handle.commit()
raise UcscOperationError(
"Backup UcsCentral", "%s" % mgmt_backup.fsm_rmt_inv_err_descr)
if not remote_enabled:
file_source = "backupfile" + file_path
if handle.is_local_download_supported():
try:
log.debug("Starting Download ")
handle.file_download(url_suffix=file_source,
file_dir=file_dir,
file_name=file_name)
except Exception as err:
log.debug("Download backup Failed")
UcscWarning(str(err))
else:
log.error("Local download not supported from sdk for this "
"version of UcsCentral, skipping it")
if remove_from_ucsc:
# remove backup from UcsCentral
log.debug("Removing backup from UcsCentral")
handle.remove_mo(mgmt_backup)
handle.commit()
def backup_local(handle, file_dir, file_name, preserve_pooled_values=False,
remove_from_ucsc=False, timeout=600):
"""
backup_local helps create UcsCentral full-state backup and dowload it
locally.
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
preserve_pooled_values (boolean): True/False, False - by default
remove_from_ucsc (boolean): True/False, False - by default
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
Example:
file_dir = "/home/user/backup"\n
file_name = "full-state_backup.tgz"\n
backup_local(handle, file_dir=file_dir, file_name=file_name)\n
"""
_backup(handle, file_dir=file_dir, file_name=file_name,
remote_enabled=False, hostname="localhost",
preserve_pooled_values=preserve_pooled_values,
remove_from_ucsc=remove_from_ucsc,
timeout=timeout)
def backup_remote(handle, file_dir, file_name, hostname,
protocol="scp", username=None, password="",
preserve_pooled_values=False,
remove_from_ucsc=False,
timeout=600):
"""
backup_remote helps create and download UcsCentral full-state backup to
remote location.
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
(supported file extension '.tgz')
hostname (str): Hostname/IP for the remote backup
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
username (str): Username for remote backup
password (str): Password for remote backup
preserve_pooled_values (boolean): True/False, False - by default
remove_from_ucsc (boolean): True/False, False - by default
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
Example:
file_dir = "/home/user/backup"\n
file_name = "full-state_backup.tgz"\n
backup_remote(handle, file_dir=file_dir, file_name=file_name,
protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
_backup(handle, file_dir=file_dir, file_name=file_name,
remote_enabled=True,
hostname=hostname, protocol=protocol,
username=username, password=password,
preserve_pooled_values=preserve_pooled_values,
remove_from_ucsc=remove_from_ucsc,
timeout=timeout)
def _export_config(handle, file_dir, file_name, timeout=600,
remote_enabled=False, protocol=None,
hostname="localhost", username=None, password="",
remove_from_ucsc=False,
preserve_pooled_values=False):
"""
_export_config internal method helps export UcsCentral config-all backup
and download it locally or to remote location.
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
remote_enabled (boolean): True if Remote backup is enabled
False - by default
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
username (str): Username for remote backup
password (str): <PASSWORD>
remove_from_ucsc (boolean): True/False, False - by default
preserve_pooled_values (boolean): True/False, False - by default
Example:
file_dir = "/home/user/backup"\n
file_name = "export_config.tgz"\n
_export_config(handle, file_dir=test_support,
file_name=file_name)\n
_export_config(handle, file_dir=test_support,
file_name=file_name,remote_enabled=True,
protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
from ..mometa.mgmt.MgmtDataExporter import MgmtDataExporter, \
MgmtDataExporterConsts
from ..mometa.top.TopSystem import TopSystem
backup_type = "config-all"
if not file_dir:
raise UcscValidationException("Missing file_dir argument")
if not file_name:
raise UcscValidationException("Missing file_name argument")
top_system = TopSystem()
if remote_enabled:
_validate_remote_host_args(protocol, hostname, username, password)
if (not file_name.endswith('.tgz')):
raise UcscValidationException(
"file_name must be .tgz format")
file_path = os.path.join(file_dir, file_name)
mgmt_export = MgmtDataExporter(
parent_mo_or_dn=top_system,
hostname=hostname,
admin_state=MgmtDataExporterConsts.ADMIN_STATE_ENABLED,
proto=protocol,
type=backup_type,
remote_file=file_path,
user=username,
pwd=password)
else:
if not os.path.exists(file_dir):
os.makedirs(file_dir)
file_string = platform.node().lower() \
+ datetime.datetime.now().strftime("%Y%m%d%H%M%S")
file_path = "/" + file_string + "_" + backup_type + "_backup.tgz"
mgmt_export = MgmtDataExporter(
parent_mo_or_dn=top_system,
hostname=hostname,
admin_state=MgmtDataExporterConsts.ADMIN_STATE_ENABLED,
proto=MgmtDataExporterConsts.PROTO_HTTP,
type=backup_type,
remote_file=file_path)
if preserve_pooled_values:
mgmt_export.preserve_pooled_values = \
MgmtDataExporterConsts.PRESERVE_POOLED_VALUES_YES
else:
mgmt_export.preserve_pooled_values = \
MgmtDataExporterConsts.PRESERVE_POOLED_VALUES_NO
handle.add_mo(mgmt_export, modify_present=True)
handle.commit()
mgmt_export = handle.query_dn(dn=mgmt_export.dn)
admin_state_temp = mgmt_export.admin_state
log.debug("Starting export config")
# Checking for the backup to compete.
duration = timeout
poll_interval = 2
while True:
mgmt_export = handle.query_dn(dn=mgmt_export.dn)
admin_state_temp = mgmt_export.admin_state
# Break condition:- if state id disabled then break
if admin_state_temp == MgmtDataExporterConsts.ADMIN_STATE_DISABLED:
break
time.sleep(min(duration, poll_interval))
duration = max(0, (duration - poll_interval))
if duration == 0:
handle.remove_mo(mgmt_export)
handle.commit()
raise UcscOperationError(
"Config Export UcsCentral", "operation timed out")
if remote_enabled:
if mgmt_export.over_all_status == \
MgmtDataExporterConsts.OVER_ALL_STATUS_FAILED:
log.debug("Config exported failed")
handle.remove_mo(mgmt_export)
handle.commit()
raise UcscOperationError(
"Config Export UcsCentral", "%s" % mgmt_export.fsm_rmt_inv_err_descr)
if not remote_enabled:
file_source = "backupfile" + file_path
if handle.is_local_download_supported():
try:
log.debug("Starting download")
handle.file_download(url_suffix=file_source,
file_dir=file_dir,
file_name=file_name)
except Exception as err:
log.debug("Download export config Failed")
UcscWarning(str(err))
else:
log.error("Local download not supported from sdk for this "
"version of UcsCentral, skipping it")
if remove_from_ucsc:
# remove backup from UcsCentral
log.debug("Removing export config from UcsCentral")
handle.remove_mo(mgmt_export)
handle.commit()
def export_config_local(handle, file_dir, file_name,
preserve_pooled_values=False,
remove_from_ucsc=False,
timeout=600):
"""
export_config_local helps export UcsCentral config-all backup and download
it locally.
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
preserve_pooled_values (boolean): True/False, False - by default
remove_from_ucsc (boolean): True/False, False - by default
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
Example:
file_dir = "/home/user/backup"\n
file_name = "export_config.tgz"\n
export_config_local(handle, file_dir=file_dir,
file_name=file_name)\n
"""
_export_config(handle, file_dir=file_dir, file_name=file_name,
remote_enabled=False, hostname="localhost",
preserve_pooled_values=preserve_pooled_values,
remove_from_ucsc=remove_from_ucsc,
timeout=timeout)
def export_config_remote(handle, file_dir, file_name, hostname,
protocol="scp", username=None, password="",
preserve_pooled_values=False,
remove_from_ucsc=False,
timeout=600):
"""
export_config_remote helps export UcsCentral config-all backup and
download it to remote location.
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
(supported file extension '.tgz')
hostname (str): Hostname/IP for the remote backup
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
username (str): Username for remote backup
password (str): Password for remote backup
preserve_pooled_values (boolean): True/False, False - by default
remove_from_ucsc (boolean): True/False, False - by default
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
Example:
file_dir = "/home/user/backup"\n
file_name = "export_config.tgz"\n
export_config_remote(handle, file_dir=file_dir,
file_name=file_name,
protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
_export_config(handle, file_dir=file_dir, file_name=file_name,
remote_enabled=True,
hostname=hostname, protocol=protocol,
username=username, password=password,
preserve_pooled_values=preserve_pooled_values,
remove_from_ucsc=remove_from_ucsc,
timeout=timeout)
def _fail_and_remove_domain_backup(handle, backup_status_mo, err):
if backup_status_mo:
handle.remove_mo(backup_status_mo)
handle.commit(dme="resource-mgr")
raise UcscOperationError("Domain backup/export_config", err)
def _backup_or_exportconfig_domain(handle, backup_type, file_dir, file_name,
domain_ip, domain_name, hostname,
preserve_pooled_values, protocol,
username, password, timeout):
"""
This internal function helps create domain full_state backup or export
config to remote location
Note: This is internal function, should use either backup_domain or
backup_export_config
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
(supported file extension is '.tgz')
backup_type (str): Either full-state or config-all.
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
domain_ip(str): IP of domain, set 'None' if domain_name is valid
domain_name(str): Domain name, valid only if domain_ip is None
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
username (str): Username for remote backup
password (str): Password for remote backup
preserve_pooled_values (boolean): True/False,
False - by default
"""
from ..mometa.mgmt.MgmtBackupOperation import MgmtBackupOperation, \
MgmtBackupOperationConsts
from ..mometa.mgmt.MgmtBackup import MgmtBackupConsts
from .ucscdomain import get_domain, _is_domain_available
preserve_pooled_values = False
if not file_dir:
raise UcscValidationException("Missing file_dir argument")
if not file_name:
raise UcscValidationException("Missing file_name argument")
if not domain_ip:
raise UcscValidationException("Missing domain_ip argument")
if backup_type == 'full-state':
if (not file_name.endswith('.tgz')):
raise UcscValidationException(
"file_name must be .tgz format")
elif backup_type == 'config-all':
if (not file_name.endswith('.xml')):
raise UcscValidationException(
"file_name must be .xml format")
_validate_remote_host_args(protocol, hostname, username, password)
domain = get_domain(handle, domain_ip, domain_name)
if _is_domain_available(handle, domain.id):
domain_dn = domain.dn
else:
raise UcscOperationError("Backup or Export_config",
"Domain with IP %s or name %s not "
"registered or lost visibility "
"with UcsCentral" %
(domain_ip, domain_name))
file_path = os.path.join(file_dir, file_name)
mgmt_backup = MgmtBackupOperation(
parent_mo_or_dn=domain_dn,
hostname=hostname,
admin_state=MgmtBackupOperationConsts.ADMIN_STATE_ENABLED,
proto=protocol,
type=backup_type,
remote_file=file_path,
user=username,
pwd=password)
if preserve_pooled_values:
mgmt_backup.preserve_pooled_values = \
MgmtBackupOperationConsts.PRESERVE_POOLED_VALUES_YES
else:
mgmt_backup.preserve_pooled_values = \
MgmtBackupOperationConsts.PRESERVE_POOLED_VALUES_NO
handle.set_mo(mgmt_backup)
handle.commit()
log.debug("Triggering Domain Backup ")
duration = 30
poll_interval = 2
backup_status_dn = "extpol/reg/clients/client-" + \
domain.id + "/backup-" + hostname
while True:
backup_status = handle.query_dn(
dn=backup_status_dn, dme="resource-mgr")
if backup_status is not None:
break
time.sleep(min(duration, poll_interval))
duration = max(0, (duration - poll_interval))
if duration == 0:
raise UcscOperationError(
"Backup or export config of domain", "not triggered")
log.debug("Domain Backup Triggered")
# Checking for the backup to become available.
log.debug("Waiting for Domain Backup to become available")
duration = timeout
poll_interval = 5
while True:
backup_status = handle.query_dn(backup_status_dn, dme="resource-mgr")
if backup_status.over_all_status == \
MgmtBackupConsts.OVER_ALL_STATUS_ALL_SUCCESS:
break
if backup_status.over_all_status != \
MgmtBackupConsts.OVER_ALL_STATUS_WORK_IN_PROGRESS:
_fail_and_remove_domain_backup(
handle, backup_status, 'operation failed')
time.sleep(min(duration, poll_interval))
duration = max(0, (duration - poll_interval))
if duration == 0:
_fail_and_remove_domain_backup(
handle, backup_status, 'operation timed out')
log.debug("Domain backup is available")
def backup_domain_remote(handle, file_dir, file_name,
domain_ip, protocol, hostname,
username=None, password="",
domain_name=None, preserve_pooled_values=False,
timeout=600):
"""
backup_domain_remote helps create domain full_state backup and download
it to remote location.
Note: Domain backup will always be remote backup
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
(supported file extension is '.tgz')
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
domain_ip(str): IP of domain, set 'None' if domain_name is valid
domain_name(str): Domain name, valid only if domain_ip is None
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
username (str): Username for remote backup
password (str): Password for remote backup
preserve_pooled_values (boolean): True/False,
False - by default
Example:
file_dir = "/home/user/backup"\n
file_name = "domain_backup.tgz"\n
backup_domain_remote(handle, file_dir=file_dir, file_name=file_name,
domain_ip='10.10.10.1', protocol='scp',
hostname='192.168.1.1',
username='admin',password='password')\n
"""
backup_type = "full-state"
return _backup_or_exportconfig_domain(handle, backup_type, file_dir,
file_name, domain_ip, domain_name,
hostname, preserve_pooled_values,
protocol, username, password,
timeout)
def export_config_domain_remote(handle, file_dir, file_name,
domain_ip, hostname, protocol,
username=None, password="",
domain_name=None, preserve_pooled_values=False,
timeout=600):
"""
export_config_domain_remote helps create domain export config and download
it to remote location.
Note: Domain export export config will always be remote export
Args:
handle (UcscHandle): UcsCentral Connection handle
file_dir (str): directory to download ucs backup file to
file_name (str): name for the backup file
(supported file extension is '.xml')
timeout (number) : time in seconds for which method waits
for the backUp file to generate before it exits.
domain_ip(str): IP of domain, set 'None' if domain_name is valid
domain_name(str): Domain name, valid only if domain_ip is None
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
username (str): Username for remote backup
password (str): Password for remote backup
preserve_pooled_values (boolean): True/False,
False - by default
Example:
file_dir = "/home/user/backup"\n
file_name = "config_backup.xml"\n
export_config_domain(handle, file_dir=test_support,
file_name=file_name,
username='admin', password='password',
domain_ip='10.10.10.1', protocol='scp',
hostname='192.168.1.1')
"""
backup_type = "config-all"
return _backup_or_exportconfig_domain(handle, backup_type, file_dir,
file_name, domain_ip, domain_name,
hostname, preserve_pooled_values,
protocol, username, password,
timeout)
def _is_backup_file_on_server(handle, hostname_str, backup_file):
backup_path_str = "/" + hostname_str + "/cfg-backups"
filter_str = '(file_path, %s, type="eq")' % backup_path_str
backups = handle.query_classid(
class_id='ConfigBackup', filter_str=filter_str)
for backup in backups:
if backup.file_name == backup_file:
log.debug("Backup file '%s' exist on UcsCentral" % backup_file)
return True
return False
def _import_config(handle, file_name, file_location="ucscentral",
file_dir=None, merge=True, protocol=None,
hostname="localhost",
username=None, password="", timeout=120):
"""
This internal method imports a UcsCentral config from local, remote or
ucscentral
Args:
handle (UcscHandle): connection handle
file_name (str): backup file name to be imported
file_location (str): file location where the config file is, it can be:
['ucscentral','local','remote']
file_dir (str): directory containing ucscentral backup file,
used only for import from local or remote
merge (boolean): True/False, specifies whether to merge the backup
config with the existing UCS Central configuration
protocol (str): Transfer protocol for remote import
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote import
username (str): Username for remote import
password (str): <PASSWORD>
timeout (number) : time in seconds for which method waits for
import operation success before timing out
Example:
file_dir = "/home/user/backup"\n
file_name = "config_export.tgz"\n
import from ucscentral:
_import_config(handle, file_name=file_name,
file_location="ucscentral")
import from local:
_import_config(handle, file_name=file_name,
file_location="local",
file_dir=file_dir,
merge=True)\n
import from remote:
_import_config(handle, file_name=file_name,
file_location="remote",
file_dir=file_dir
protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
from ..mometa.top.TopSystem import TopSystem
from ..mometa.mgmt.MgmtDataImporter import MgmtDataImporter, \
MgmtDataImporterConsts
if not file_name:
raise UcscValidationException("Missing file_name argument")
if file_location != "ucscentral":
if not file_dir:
raise UcscValidationException("Missing file_dir argument")
if (not file_name.endswith('.tgz')):
raise UcscValidationException("file_name must be .tgz format")
top_system = TopSystem()
if file_location == "remote":
file_path = os.path.join(file_dir, file_name)
_validate_remote_host_args(protocol, hostname, username, password)
mgmt_importer = MgmtDataImporter(
parent_mo_or_dn=top_system,
hostname=hostname,
remote_file=file_path,
proto=protocol,
user=username,
pwd=password,
admin_state=MgmtDataImporterConsts.ADMIN_STATE_ENABLED)
elif file_location == "local":
file_path = os.path.join(file_dir, file_name)
if not os.path.exists(file_path):
raise UcscOperationError("Import config",
"Backup File '%s' not found" %
file_path)
mgmt_importer = MgmtDataImporter(
parent_mo_or_dn=top_system,
hostname="localhost",
remote_file='/' + file_name,
proto=MgmtDataImporterConsts.PROTO_HTTP,
admin_state=MgmtDataImporterConsts.ADMIN_STATE_ENABLED)
elif file_location == "ucscentral":
if not _is_backup_file_on_server(handle, "ucs-central", file_name):
raise UcscOperationError("Import config",
"Backup File '%s' not found "
"on UcsCentral" % file_name)
mgmt_importer = MgmtDataImporter(
parent_mo_or_dn=top_system,
hostname="localhost",
remote_file='/ucs-central/cfg-backups/' + file_name,
proto=MgmtDataImporterConsts.PROTO_TFTP,
admin_state=MgmtDataImporterConsts.ADMIN_STATE_ENABLED)
else:
raise UcscOperationError(
"Import config",
"Invalid file_location argument."
"It must be either ucscentral,local or remote")
if merge:
mgmt_importer.action = MgmtDataImporterConsts.ACTION_MERGE
else:
mgmt_importer.action = MgmtDataImporterConsts.ACTION_REPLACE
if file_location == "local":
try:
log.debug("Start uploading config")
uri_suffix = "operations/file-%s/importconfig.txt?Cookie=%s" % (
file_name, handle.cookie)
handle.file_upload(url_suffix=uri_suffix,
file_dir=file_dir,
file_name=file_name)
except Exception as err:
UcscWarning(str(err))
raise UcscOperationError("Upload config", "upload failed")
handle.add_mo(mgmt_importer, modify_present=True)
handle.commit()
duration = timeout
poll_interval = 2
log.debug("Importing UcsCentral config")
while True:
mgmt_importer = handle.query_dn(dn=mgmt_importer.dn)
admin_state = mgmt_importer.admin_state
# Break condition:- if state id disabled then break
if admin_state == MgmtDataImporterConsts.ADMIN_STATE_DISABLED:
break
time.sleep(min(duration, poll_interval))
duration = max(0, (duration - poll_interval))
if duration == 0:
raise UcscOperationError(
"Import config", "operation timed out")
if mgmt_importer.over_all_status != \
MgmtDataImporterConsts.OVER_ALL_STATUS_ALL_SUCCESS:
raise UcscOperationError(
"Import config",
("operational status %s " % mgmt_importer.over_all_status))
log.debug("Import config to UcsCentral was successfull")
return mgmt_importer
def import_config_ucscentral(handle, file_name, merge=True,
timeout=120):
"""
import_config_ucscentral imports existing UcsCentral config ie.available
on UcsCentral(taken via schedule_export_config) to the UcsCentral
Args:
handle (UcscHandle): connection handle
file_name (str): backup file name to be imported
merge (boolean): True/False, specifies whether to merge the backup
config with the existing UCS Central configuration
timeout (number) : time in seconds for which method waits for
import operation success before timing out
Example:
file_dir = "/home/user/backup"\n
file_name = "config_export.tgz"\n
import from ucscentral:
import_config_ucscentral(handle, file_name=file_name)
"""
_import_config(handle, file_name=file_name, merge=merge,
file_location="ucscentral",
hostname="localhost",
timeout=timeout)
def import_config_local(handle, file_dir, file_name, merge=True,
timeout=120):
"""
import_config_ucscentral imports UcsCentral config available locally to
UcsCentral
Args:
handle (UcscHandle): connection handle
file_dir (str): directory containing ucscentral backup file,
file_name (str): backup file name to be imported
merge (boolean): True/False, specifies whether to merge the backup
config with the existing UCS Central configuration
timeout (number) : time in seconds for which method waits for
import operation success before timing out
Example:
file_dir = "/home/user/backup"\n
file_name = "config_export.tgz"\n
import from local:
import_config_local(handle, file_name=file_name,
file_dir=file_dir,
merge=True)\n
"""
_import_config(handle, file_name=file_name, file_dir=file_dir, merge=merge,
file_location="local",
hostname="localhost",
timeout=timeout)
def import_config_remote(handle, file_dir, file_name, hostname,
merge=True,
protocol="scp",
username=None, password="",
timeout=120):
"""
import_config_remote imports UcsCentral config from remote location to
UcsCentral
Args:
handle (UcscHandle): connection handle
file_dir (str): directory containing ucscentral backup file,
file_name (str): backup file name to be imported
hostname (str): Hostname/IP for the remote import
merge (boolean): True/False, specifies whether to merge the backup
config with the existing UCS Central configuration
protocol (str): Transfer protocol for remote import
['ftp','sftp','tftp','scp']
username (str): Username for remote import
password (str): Password for remote import
timeout (number) : time in seconds for which method waits for
import operation success before timing out
Example:
file_dir = "/home/user/backup"\n
file_name = "config_export.tgz"\n
import from remote:
import_config_remote(handle, file_name=file_name,
file_dir=file_dir
protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
_import_config(handle, file_name=file_name, file_dir=file_dir, merge=merge,
file_location="remote",
protocol=protocol, hostname=hostname,
username=username, password=password,
timeout=timeout)
def import_config_domain(handle, to_domain_ip, from_domain_ip, config_file,
merge=True,
to_domain_name=None, from_domain_name=None,
timeout=120):
"""
This operation imports UcsCentral Domain's config created earlier via
schedule backup of the same or other Domains.
Note: Only config-all backup on UcsCentral are available for import, local
or remote import is not supported for domain
Args:
handle (UcscHandle): connection handle
to_domain_ip(str): IP of domain To which you want to import
set 'None' if domain_name is valid
from_domain_ip(str): IP of domain From which you want to import
set 'None' if domain_name is valid
config_file(str): From domain's config file which you want to import
merge (boolean): True/False, specifies whether to merge the backup
config with the existing domain configuration
to_domain_name(str): to domain name, valid only if to_domain_ip is None
from_domain_name(str): from domain name, valid only if
from_domain_ip is None
timeout (number) : time in seconds for which method waits for
import operation success before timing out
Example:
import_config_domain(handle, to_domain_ip="10.10.10.100",
from_domain_ip="192.168.1.1",
config_file="all-cfg.1.tgz")\n
"""
from ..mometa.mgmt.MgmtImporter import MgmtImporter, MgmtImporterConsts
from .ucscdomain import get_domain, _is_domain_available
to_domain = get_domain(handle, to_domain_ip, to_domain_name)
if not _is_domain_available(handle, to_domain.id):
raise UcscOperationError("Import config domain",
"Domain with IP %s or name %s not "
"registered or lost visibility "
"with UcsCentral" %
(to_domain_ip, to_domain_name))
from_domain = get_domain(handle, from_domain_ip, from_domain_name)
if not _is_domain_available(handle, from_domain.id):
raise UcscOperationError("Import config domain",
"Domain with IP %s or name %s not "
"registered or lost visibility "
"with UcsCentral" %
(from_domain_ip, from_domain_name))
if not _is_backup_file_on_server(handle, from_domain.address, config_file):
raise UcscOperationError("Import config domain",
"Backup File '%s' not found "
"on UcsCentral" % config_file)
filter_str = '(connector_id, %s, type="eq")' % to_domain.id
consumer_inst = handle.query_classid(
class_id='ConsumerInst', filter_str=filter_str)
if (len(consumer_inst) <= 0):
raise UcscOperationError("Import config domain",
"Unable to get Domain instance"
"with IP %s for import " %
to_domain.address)
consumer_dn = consumer_inst[0].dn
top_system = handle.query_dn("sys")
mgmt_importer = MgmtImporter(
parent_mo_or_dn=consumer_dn,
hostname=top_system.address,
remote_file='/' + from_domain.address + '/cfg-backups/' + config_file,
proto=MgmtImporterConsts.PROTO_TFTP,
admin_state=MgmtImporterConsts.ADMIN_STATE_ENABLED)
if merge:
mgmt_importer.action = MgmtImporterConsts.ACTION_MERGE
else:
mgmt_importer.action = MgmtImporterConsts.ACTION_REPLACE
handle.add_mo(mgmt_importer, modify_present=True)
handle.commit(dme="operation-mgr")
duration = timeout
poll_interval = 2
log.debug("Importing domain config")
while True:
mgmt_importer = handle.query_dn(
dn=mgmt_importer.dn,
dme="operation-mgr")
admin_state = mgmt_importer.admin_state
# Break condition:- if state id disabled then break
if admin_state == MgmtImporterConsts.ADMIN_STATE_DISABLED:
break
time.sleep(min(duration, poll_interval))
duration = max(0, (duration - poll_interval))
if duration == 0:
raise UcscOperationError(
"Import config", "operation timed out")
if mgmt_importer.op_status != \
MgmtImporterConsts.OP_STATUS_ALL_SUCCESS:
raise UcscOperationError(
"Import config",
("operational status %s " % mgmt_importer.op_status))
log.debug("Import config to domain was successfull")
return mgmt_importer
def _schedule_backup(handle, descr, sched_name, max_bkup_files, remote_enabled,
protocol, hostname, file_path,
username, password, parent_mo_or_dn):
"""
Internal method to schedule and take remote backup of UcsCentral and Domain
full-state backup
"""
from ..mometa.mgmt.MgmtBackupPolicy import MgmtBackupPolicy, \
MgmtBackupPolicyConsts
if remote_enabled:
if not file_path:
raise UcscValidationException("Missing file_path argument")
_validate_remote_host_args(protocol, hostname, username, password)
proto = protocol
host = hostname
remote_file = file_path
user = username
pwd = password
else:
proto = MgmtBackupPolicyConsts.PROTO_NFS_COPY
host = ""
remote_file = " "
user = ""
pwd = ""
backup_pol = MgmtBackupPolicy(
parent_mo_or_dn=parent_mo_or_dn,
descr=descr,
admin_state=MgmtBackupPolicyConsts.ADMIN_STATE_ENABLE,
sched_name=sched_name,
max_files=str(max_bkup_files),
proto=proto,
host=host,
remote_file=remote_file,
user=user,
pwd=pwd,
name="default")
handle.add_mo(backup_pol, modify_present=True)
handle.commit()
return backup_pol
def schedule_backup(handle, descr="Database Backup Policy",
sched_name="global-default", max_bkup_files="2",
remote_enabled=False, protocol=None,
hostname=None, file_path=None,
username=None, password=""):
"""
schedule_backup helps schedule and optionally take remote backup of
UcsCentral full-state backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
descr (str): Description of the policy
sched_name (str): Name of the schedule
max_bkup_files (str): Number of files to keep as backup
remote_enabled (boolean): True if Remote backup is enabled
False - by default
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
file_path (str): Absolute file path on remote host
username (str): Username for remote backup
password (str): Password for remote backup
Example:
file_path = "/ws/usr-admin/backup.tgz"\n
schedule_backup(handle, max_bkup_files=3)
schedule_backup(handle, file_path=file_path,
remote_enabled=True,
protocol='scp',hostname='192.168.1.1',
username='admin',password='password')\n
"""
return _schedule_backup(
handle, descr=descr, sched_name=sched_name,
max_bkup_files=max_bkup_files,
remote_enabled=remote_enabled, protocol=protocol,
hostname=hostname, file_path=file_path,
username=username, password=password,
parent_mo_or_dn=ucsc_base_dn)
def schedule_backup_domain(handle, descr="Database Backup Policy",
sched_name="global-default", max_bkup_files="2",
remote_enabled=False, protocol=None,
hostname=None, file_path=None,
username=None, password="",
domain_group="root"):
"""
schedule_backup_domain helps schedule and optionally take remote backup of
domain's full-state backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
descr (str): Description of the policy
sched_name (str): Name of the schedule
max_bkup_files (str): Number of files to keep as backup
remote_enabled (boolean): True if Remote backup is enabled
False - by default
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
file_path (str): Absolute file path on remote host
username (str): Username for remote backup
password (<PASSWORD>): <PASSWORD>
domain_group (str): Fully qualified domain group name
Example:
file_path = "/ws/usr-admin/backup.tgz"\n
schedule_backup_domain(handle, max_bkup_files=3)
schedule_backup_domain(handle, file_path=file_path,
remote_enabled=True,
protocol='scp',hostname='192.168.1.1',
username='admin',password='password',
domain_group="root/demo_domgrp")\n
"""
from ucscsdk.utils.ucscdomain import get_domain_group_dn
domain_group_dn = get_domain_group_dn(handle, domain_group)
return _schedule_backup(
handle, descr=descr, sched_name=sched_name,
max_bkup_files=max_bkup_files,
remote_enabled=remote_enabled, protocol=protocol,
hostname=hostname, file_path=file_path,
username=username, password=password,
parent_mo_or_dn=domain_group_dn)
def _schedule_export_config(handle, descr, sched_name,
max_bkup_files, remote_enabled,
protocol, hostname, file_path,
username, password,
parent_mo_or_dn):
"""
Internal method to schedule and take remote backup of UcsCentral and Domain
config-all backup
"""
from ..mometa.mgmt.MgmtCfgExportPolicy import MgmtCfgExportPolicy, \
MgmtCfgExportPolicyConsts
if remote_enabled:
if not file_path:
raise UcscValidationException("Missing file_path argument")
_validate_remote_host_args(protocol, hostname, username, password)
proto = protocol
host = hostname
remote_file = file_path
user = username
pwd = password
else:
proto = MgmtCfgExportPolicyConsts.PROTO_NFS_COPY
host = ""
remote_file = " "
user = ""
pwd = ""
cfg_export_pol = MgmtCfgExportPolicy(
parent_mo_or_dn=parent_mo_or_dn,
descr=descr,
admin_state=MgmtCfgExportPolicyConsts.ADMIN_STATE_ENABLE,
sched_name=sched_name,
max_files=str(max_bkup_files),
proto=proto,
host=host,
remote_file=remote_file,
user=user,
pwd=<PASSWORD>,
name="default")
handle.add_mo(cfg_export_pol, modify_present=True)
handle.commit()
return cfg_export_pol
def schedule_export_config(handle, descr="Configuration Export "
"Policy", sched_name="global-default",
max_bkup_files="2",
remote_enabled=False, protocol=None,
hostname="", file_path="",
username=None, password=""):
"""
schedule_export_config helps schedule and optionally take remote backup of
UcsCentral's config-all backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
descr (str): Description of the policy
sched_name (str): Name of the schedule
max_bkup_files (str): Number of files to keep as backup
remote_enabled (boolean): True if Remote backup is enabled
False - by default
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
file_path (str): Absolute file path on remote host
username (str): Username for remote backup
password (str): Password for remote backup
Example:
file_path = "/ws/usr-admin/config.tgz"\n
schedule_export_config(handle, max_bkup_files=3)
schedule_export_config(handle, file_path=file_path,
remote_enabled=True,
protocol='scp',hostname='172.16.58.3',
username='admin',password='<PASSWORD>')\n
"""
return _schedule_export_config(
handle, descr=descr, sched_name=sched_name,
max_bkup_files=max_bkup_files,
remote_enabled=remote_enabled, protocol=protocol,
hostname=hostname, file_path=file_path,
username=username, password=password,
parent_mo_or_dn=ucsc_base_dn)
def schedule_export_config_domain(handle, descr="Configuration Export "
"Policy", sched_name="global-default",
max_bkup_files="2",
remote_enabled=False, protocol=None,
hostname="", file_path="",
username=None, password=None,
domain_group="root"):
"""
schedule_export_config_domain helps schedule and optionally take remote
backup of domain's config-all backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
descr (str): Description of the policy
sched_name (str): Name of the schedule
max_bkup_files (str): Number of files to keep as backup
remote_enabled (boolean): True if Remote backup is enabled
False - by default
protocol (str): Transfer protocol for remote backup
['ftp','sftp','tftp','scp']
hostname (str): Hostname/IP for the remote backup
file_path (str): Absolute file path on remote host
username (str): Username for remote backup
password (str): Password for remote backup
domain_group (str): Fully qualified domain group name
Example:
file_path = "/ws/usr-admin/config.tgz"\n
schedule_export_config_domain(handle, max_bkup_files=3)
schedule_export_config_domain(handle, file_path=file_path,
remote_enabled=True,
protocol='scp',hostname='172.16.58.3',
username='admin',password='<PASSWORD>',
domain_group="root/demo_domgrp")\n
"""
from ucscsdk.utils.ucscdomain import get_domain_group_dn
domain_group_dn = get_domain_group_dn(handle, domain_group)
return _schedule_export_config(
handle, descr=descr, sched_name=sched_name,
max_bkup_files=max_bkup_files,
remote_enabled=remote_enabled, protocol=protocol,
hostname=hostname, file_path=file_path,
username=username, password=password,
parent_mo_or_dn=domain_group_dn)
def remove_schedule_backup(handle):
"""
remvove_schedule_backup removes the schedule of UcsCentral's
full backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
Example:
remove_schedule_backup(handle)
"""
from ..mometa.mgmt.MgmtBackupPolicy import MgmtBackupPolicyConsts
dn = ucsc_base_dn + "/db-backup-policy-default"
mo = handle.query_dn(dn=dn)
if not mo:
raise UcscOperationError("Remove backup schedule",
"Backup Schedule doesn't exist")
mo.admin_state = MgmtBackupPolicyConsts.ADMIN_STATE_DISABLE
handle.set_mo(mo)
handle.commit()
def remove_schedule_export_config(handle):
"""
remvove_schedule_export_config removes the schedule of
UcsCentral config-all backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
Example:
remove_schedule_export_config(handle)
"""
from ..mometa.mgmt.MgmtCfgExportPolicy import MgmtCfgExportPolicyConsts
dn = ucsc_base_dn + "/cfg-exp-policy-default"
mo = handle.query_dn(dn=dn)
if not mo:
raise UcscOperationError("Remove export config",
"Export config schedule doesn't exist")
mo.admin_state = MgmtCfgExportPolicyConsts.ADMIN_STATE_DISABLE
handle.set_mo(mo)
handle.commit()
def remove_schedule_backup_domain(handle, domain_group="root"):
"""
remvove_schedule_backup_domain removes the schedule policy of Domain's
full backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
domain_group (str): Fully qualified domain group name
Example:
remove_schedule_backup_domain(handle, domain_group="root/demo_domgrp")
"""
from ..mometa.mgmt.MgmtBackupPolicy import MgmtBackupPolicyConsts
from ucscsdk.utils.ucscdomain import get_domain_group_dn
domain_group_dn = get_domain_group_dn(handle, domain_group)
dn = domain_group_dn + "/db-backup-policy-default"
mo = handle.query_dn(dn=dn)
if not mo:
raise UcscOperationError("Remove backup schedule",
"Backup Schedule doesn't exist")
mo.admin_state = MgmtBackupPolicyConsts.ADMIN_STATE_DISABLE
handle.set_mo(mo)
if domain_group != "root":
handle.remove_mo(mo)
handle.commit()
def remove_schedule_export_config_domain(handle, domain_group="root"):
"""
remvove_schedule_export_config_domain removes the schedule policy of
Domain's config-all backup.
Args:
handle (UcscHandle): UcsCentral Connection handle
domain_group (str): Fully qualified domain group name
Example:
remove_schedule_export_config_domain(handle,
domain_group="root/demo_domgrp")\n
"""
from ..mometa.mgmt.MgmtCfgExportPolicy import MgmtCfgExportPolicyConsts
from ucscsdk.utils.ucscdomain import get_domain_group_dn
domain_group_dn = get_domain_group_dn(handle, domain_group)
dn = domain_group_dn + "/cfg-exp-policy-default"
mo = handle.query_dn(dn=dn)
if not mo:
raise UcscOperationError("Remove export config",
"Export config schedule doesn't exist")
mo.admin_state = MgmtCfgExportPolicyConsts.ADMIN_STATE_DISABLE
handle.set_mo(mo)
if domain_group != "root":
handle.remove_mo(mo)
handle.commit()
|
<reponame>ska-sa/tango-simlib
#########################################################################################
# Author: <EMAIL> #
# Copyright 2018 SKA South Africa (http://ska.ac.za/) #
# #
# BSD license - see LICENSE.txt for details #
#########################################################################################
from __future__ import absolute_import, division, print_function
from future import standard_library
standard_library.install_aliases() # noqa: E402
import logging
import pkg_resources
import unittest
from mock import patch
import tango
from katcp.testutils import start_thread_with_cleanup
from tango.test_context import DeviceTestContext
from tango_simlib import model, tango_sim_generator
from tango_simlib.examples import override_class
from tango_simlib.utilities import simdd_json_parser, helper_module, sim_xmi_parser
from tango_simlib.utilities.testutils import cleanup_tempfile, ClassCleanupUnittestMixin
MODULE_LOGGER = logging.getLogger(__name__)
TANGO_CMD_PARAMS_NAME_MAP = {
"name": "cmd_name",
"doc_in": "in_type_desc",
"dtype_in": "in_type",
"doc_out": "out_type_desc",
"dtype_out": "out_type",
}
# Mandatory parameters required to create a well configure Tango command.
EXPECTED_MANDATORY_CMD_PARAMETERS = frozenset(
["dformat_in", "dformat_out", "doc_in", "doc_out", "dtype_in", "dtype_out", "name"]
)
# Mandatory parameters required by each override_class.
EXPECTED_MANDATORY_OVERRIDE_CLASS_PARAMETERS = frozenset(
["class_name", "module_directory", "module_name", "name"]
)
# The desired information for the attribute temperature when the Weather_SimDD
# json file is parsed by the SimddParser.
EXPECTED_TEMPERATURE_ATTR_INFO = {
"abs_change": "0.5",
"archive_abs_change": "0.5",
"archive_period": "1000",
"archive_rel_change": "10",
"data_format": "Scalar",
"data_type": tango._tango.CmdArgType.DevDouble,
"format": "6.2f",
"delta_t": "1000",
"delta_val": "0.5",
"description": "Current actual temperature outside near the telescope.",
"display_level": "OPERATOR",
"event_period": "1000",
"label": "Outside Temperature",
"max_alarm": "50",
"max_bound": "50",
"max_dim_x": "1",
"max_dim_y": "0",
"max_slew_rate": "1",
"max_value": "51",
"mean": "25",
"min_alarm": "-9",
"min_bound": "-10",
"min_value": "-10",
"min_warning": "-8",
"max_warning": "49",
"name": "temperature",
"quantity_simulation_type": "GaussianSlewLimited",
"period": "1000",
"rel_change": "10",
"std_dev": "5",
"unit": "Degrees Centrigrade",
"update_period": "1000",
"writable": "READ",
}
class GenericSetup(unittest.TestCase):
"""A class providing the setUp method definition for the other test classes."""
longMessage = True
def setUp(self):
super(GenericSetup, self).setUp()
self.simdd_json_file = [
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "Weather_SimDD.json"
)
]
self.simdd_parser = simdd_json_parser.SimddParser()
self.simdd_parser.parse(self.simdd_json_file[0])
class test_SimddJsonParser(GenericSetup):
"""A test class that tests that the SimddJsonParser works correctly."""
def test_parsed_attributes(self):
"""Testing that the attribute information parsed matches with the one captured
in the SimDD json file.
"""
actual_parsed_attrs = self.simdd_parser.get_device_attribute_metadata()
expected_attr_list = [
"input-comms-ok",
"insolation",
"pressure",
"rainfall",
"relative-humidity",
"temperature",
"wind-direction",
"wind-speed",
]
actual_parsed_attr_list = sorted(actual_parsed_attrs.keys())
self.assertGreater(
len(actual_parsed_attr_list), 0, "There is no attribute information parsed"
)
self.assertEquals(
set(expected_attr_list),
set(actual_parsed_attr_list),
"There are missing attributes",
)
# Test if all the parsed attributes have the mandatory properties
for attr_name, attribute_metadata in actual_parsed_attrs.items():
for param in helper_module.DEFAULT_TANGO_ATTRIBUTE_PARAMETER_TEMPLATE.keys():
self.assertIn(
param,
attribute_metadata.keys(),
"The parsed attribute '%s' does not have the mandotory parameter "
"'%s' " % (attr_name, param),
)
# Using the made up temperature attribute expected results as we
# haven't generated the full test data for the other attributes.
self.assertIn(
"temperature",
actual_parsed_attrs.keys(),
"The attribute temperature is not in the parsed attribute list",
)
actual_parsed_temperature_attr_info = actual_parsed_attrs["temperature"]
# Compare the values of the attribute properties captured in the POGO
# generated xmi file and the ones in the parsed attribute data structure.
for prop in EXPECTED_TEMPERATURE_ATTR_INFO:
self.assertEquals(
actual_parsed_temperature_attr_info[prop],
EXPECTED_TEMPERATURE_ATTR_INFO[prop],
"The expected value for the parameter '%s' does "
"not match with the actual value" % (prop),
)
def test_parsed_override_info(self):
"""Testing that the class override information parsed matches with the one
captured in the SimDD json file.
"""
actual_override_info = self.simdd_parser.get_device_cmd_override_metadata()
for klass_info in actual_override_info.values():
for param in EXPECTED_MANDATORY_OVERRIDE_CLASS_PARAMETERS:
self.assertIn(
param,
klass_info.keys(),
"Class override info missing" " some important parameter.",
)
class test_PopulateModelQuantities(GenericSetup):
def test_model_quantities(self):
"""Testing that the model quantities that are added to the model match with
the attributes specified in the XMI file.
"""
device_name = "tango/device/instance"
pmq = model.PopulateModelQuantities(self.simdd_parser, device_name)
self.assertEqual(
device_name,
pmq.sim_model.name,
"The device name and the model name do not match.",
)
expected_quantities_list = [
"insolation",
"temperature",
"pressure",
"input-comms-ok",
"rainfall",
"relative-humidity",
"wind-direction",
"wind-speed",
]
actual_quantities_list = pmq.sim_model.sim_quantities.keys()
self.assertEqual(
set(expected_quantities_list),
set(actual_quantities_list),
"The are quantities missing in the model",
)
def test_model_quantities_metadata(self):
"""Testing that the metadata of the quantities matches with the metadata
data of the parsed attribute data captured in the SDD xml file.
"""
device_name = "tango/device/instance"
pmq = model.PopulateModelQuantities(self.simdd_parser, device_name)
self.assertEqual(
device_name,
pmq.sim_model.name,
"The device name and the model name do not match.",
)
attribute_metadata = self.simdd_parser.get_device_attribute_metadata()
for sim_quantity_name, sim_quantity in pmq.sim_model.sim_quantities.items():
sim_quantity_metadata = getattr(sim_quantity, "meta")
attr_meta = attribute_metadata[sim_quantity_name]
for attr_param_name, attr_param_val in attr_meta.items():
self.assertTrue(
attr_param_name in sim_quantity_metadata,
"The param '%s' was not added to the model quantity"
" '%s'" % (attr_param_name, sim_quantity_name),
)
self.assertEqual(
sim_quantity_metadata[attr_param_name],
attr_param_val,
"The value of the param '%s' in the model quantity '%s' is "
"not the same with the one captured in the SDD xml file "
"for the monitoring point '%s'."
% (attr_param_name, sim_quantity_name, attr_param_name),
)
EXPECTED_ACTION_SET_TEMPERATURE_METADATA = {
"name": "SetTemperature",
"description": "Sets the temperature value",
"dtype_in": "Double",
"doc_in": "Value to set quantity",
"dformat_in": "",
"dtype_out": "String",
"doc_out": "Command responds",
"dformat_out": "",
"actions": [
{"behaviour": "input_transform", "destination_variable": "temporary_variable"},
{
"behaviour": "side_effect",
"source_variable": "temporary_variable",
"destination_quantity": "temperature",
},
{"behaviour": "output_return", "source_variable": "temporary_variable"},
],
}
class test_PopulateModelActions(GenericSetup):
def test_model_actions(self):
"""Testing that the model actions that are added to the model match with
the commands specified in the XMI file.
"""
device_name = "tango/device/instance"
pmq = model.PopulateModelQuantities(self.simdd_parser, device_name)
sim_model = pmq.sim_model
cmd_info = self.simdd_parser.get_device_command_metadata()
override_info = self.simdd_parser.get_device_cmd_override_metadata()
model.PopulateModelActions(cmd_info, override_info, device_name, sim_model)
actual_actions_list = sim_model.sim_actions.keys()
expected_actions_list = [
"On",
"Off",
"StopRainfall",
"SetTemperature",
"Add",
"StopQuantitySimulation",
"MultiplyStringBy3",
]
self.assertEqual(
set(actual_actions_list),
set(expected_actions_list),
"There are actions missing in the model",
)
def test_model_actions_metadata(self):
"""Testing that the model action metadata has been added correctly to the model.
"""
device_name = "tango/device/instance"
pmq = model.PopulateModelQuantities(self.simdd_parser, device_name)
sim_model = pmq.sim_model
cmd_info = self.simdd_parser.get_device_command_metadata()
override_info = self.simdd_parser.get_device_cmd_override_metadata()
model.PopulateModelActions(cmd_info, override_info, device_name, sim_model)
sim_model_actions_meta = sim_model.sim_actions_meta
for cmd_name, cmd_metadata in cmd_info.items():
model_act_meta = sim_model_actions_meta[cmd_name]
for action_parameter in EXPECTED_MANDATORY_CMD_PARAMETERS:
self.assertIn(
action_parameter,
model_act_meta,
"The parameter is not in the action's metadata",
)
self.assertEqual(
cmd_metadata,
model_act_meta,
"The action's %s metadata was not processed correctly" % cmd_name,
)
def test_model_actions_overrides(self):
"""Testing that the On command defined in the SimDD file is mapped to the
correct user-defined action handler provided in the override class.
"""
device_name = "tango/device/instance"
pmq = model.PopulateModelQuantities(self.simdd_parser, device_name)
sim_model = pmq.sim_model
cmd_info = self.simdd_parser.get_device_command_metadata()
override_info = self.simdd_parser.get_device_cmd_override_metadata()
model.PopulateModelActions(cmd_info, override_info, device_name, sim_model)
action_on = sim_model.sim_actions["On"]
self.assertEqual(
action_on.func.__self__.__class__, override_class.OverrideWeather
)
def test_model_action_behaviour(self):
device_name = "tango/device/instance"
pmq = model.PopulateModelQuantities(self.simdd_parser, device_name)
sim_model = pmq.sim_model
cmd_info = self.simdd_parser.get_device_command_metadata()
override_info = self.simdd_parser.get_device_cmd_override_metadata()
model.PopulateModelActions(cmd_info, override_info, device_name, sim_model)
action_set_temperature = sim_model.sim_actions["SetTemperature"]
data_in = 25.00
self.assertEqual(action_set_temperature(data_in), data_in)
class test_SimddDeviceIntegration(ClassCleanupUnittestMixin, unittest.TestCase):
longMessage = True
@classmethod
def setUpClassWithCleanup(cls):
cls.tango_db = cleanup_tempfile(cls, prefix="tango", suffix=".db")
cls.data_descr_file = [
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "Weather_SimDD.json"
)
]
cls.device_name = "test/nodb/tangodeviceserver"
model = tango_sim_generator.configure_device_models(
cls.data_descr_file, cls.device_name
)
cls.TangoDeviceServer = tango_sim_generator.get_tango_device_server(
model, cls.data_descr_file
)[0]
cls.tango_context = DeviceTestContext(
cls.TangoDeviceServer, device_name=cls.device_name, db=cls.tango_db
)
with patch("tango_simlib.utilities.helper_module.get_database"):
start_thread_with_cleanup(cls, cls.tango_context)
def setUp(self):
super(test_SimddDeviceIntegration, self).setUp()
self.device = self.tango_context.device
self.instance = self.TangoDeviceServer.instances[self.device.name()]
self.instance.model.paused = True
with patch("tango_simlib.utilities.helper_module.get_database"):
self.device.Init()
self.simdd_json_parser = simdd_json_parser.SimddParser()
self.simdd_json_parser.parse(self.data_descr_file[0])
default_metadata_values = {}
for quantity in self.instance.model.sim_quantities.keys():
if hasattr(self.instance.model.sim_quantities[quantity], "max_bound"):
default_metadata_values[quantity] = self.instance.model.sim_quantities[
quantity
].max_bound
self.addCleanup(self._restore_model, default_metadata_values)
def _restore_model(self, default_metadata_values):
for quantity in self.instance.model.sim_quantities.keys():
if hasattr(self.instance.model.sim_quantities[quantity], "max_bound"):
self.instance.model.sim_quantities[
quantity
].max_bound = default_metadata_values[quantity]
def test_attribute_list(self):
"""Testing whether the attributes specified in the POGO generated xmi file
are added to the TANGO device
"""
attributes = set(self.device.get_attribute_list())
expected_attributes = []
default_attributes = helper_module.DEFAULT_TANGO_DEVICE_ATTRIBUTES
expected_attributes = (
self.simdd_json_parser.get_device_attribute_metadata().keys()
)
self.assertEqual(
set(expected_attributes),
attributes - default_attributes,
"Actual tango device attribute list differs from expected " "list!",
)
def test_command_list(self):
"""Testing that the command list in the Tango device matches with the one
specified in the SimDD data description file.
"""
actual_device_commands = set(self.device.get_command_list())
expected_command_list = (
self.simdd_json_parser.get_device_command_metadata().keys()
)
expected_command_list = list(expected_command_list)
expected_command_list.extend(helper_module.DEFAULT_TANGO_DEVICE_COMMANDS)
self.assertEquals(
actual_device_commands,
set(expected_command_list),
"The commands specified in the SimDD file are not present in" " the device",
)
def test_command_properties(self):
"""Testing that the command parameter information matches with the information
captured in the SimDD data description file.
"""
command_data = self.simdd_json_parser.get_device_command_metadata()
extra_command_parameters = ["dformat_in", "dformat_out", "description", "actions"]
for cmd_name, cmd_metadata in command_data.items():
cmd_config_info = self.device.get_command_config(cmd_name)
for cmd_prop, cmd_prop_value in cmd_metadata.items():
# Exclude parameters that are not part of the TANGO command configuration
# information.
if cmd_prop in extra_command_parameters:
continue
self.assertTrue(
hasattr(cmd_config_info, TANGO_CMD_PARAMS_NAME_MAP[cmd_prop]),
"The cmd parameter '%s' for the cmd '%s' was not translated"
% (cmd_prop, cmd_name),
)
if cmd_prop_value == "none" or cmd_prop_value == "":
cmd_prop_value = "Uninitialised"
self.assertEqual(
getattr(cmd_config_info, TANGO_CMD_PARAMS_NAME_MAP[cmd_prop]),
cmd_prop_value,
"The cmd %s parameter '%s/%s' values do not match"
% (cmd_name, cmd_prop, TANGO_CMD_PARAMS_NAME_MAP[cmd_prop]),
)
def test_On_command(self):
"""Testing that the On command changes the value of the State attribute of the
Tango device to ON.
"""
command_name = "On"
expected_result = None
self.device.command_inout(command_name)
self.assertEqual(self.device.command_inout(command_name), expected_result)
self.assertEqual(
getattr(self.device.read_attribute("State"), "value"), tango.DevState.ON
)
def test_Add_command(self):
"""Testing that the Tango device command can take input of an array type and
return a output value of type double.
"""
command_name = "Add"
command_args = [12, 45, 53, 32, 2.1, 0.452]
expected_return_value = 144.552
actual_return_value = self.device.command_inout(command_name, command_args)
self.assertEqual(
expected_return_value,
actual_return_value,
"The actual return" "value does not match with the expected return value.",
)
def test_MultiplyStringBy3_command(self):
"""Testing that the Tango device command can take input of type string and
return an output value of type string.
"""
command_name = "MultiplyStringBy3"
command_args = "LMC"
expected_return_value = "LMCLMCLMC"
actual_return_value = self.device.command_inout(command_name, command_args)
self.assertEqual(
expected_return_value,
actual_return_value,
"The actual return" "value does not match with the expected return value.",
)
def test_Off_command(self):
"""Testing that the Off command changes the State attributes value of the Tango
device to OFF.
"""
command_name = "Off"
expected_result = None
self.assertEqual(self.device.command_inout(command_name), expected_result)
self.assertEqual(
getattr(self.device.read_attribute("State"), "value"), tango.DevState.OFF
)
def test_set_temperature_command(self):
"""Testing that the SetTemperature command changes the temperature
attributes value of the Tango device to the specified input parameter.
"""
command_name = "SetTemperature"
data_in = 25.0
expected_result = data_in
self.assertEqual(
self.device.command_inout(command_name, data_in), expected_result
)
self.instance.model.last_update_time = 0
# The tango device temperature attribute value return a floating number
# thus it is rounded to two decimal places before checking if it's the
# same as the `data_in` value
self.assertEqual(
round(getattr(self.device.read_attribute("Temperature"), "value"), 2), data_in
)
MKAT_VDS_ATTRIBUTE_LIST = frozenset(
[
"camera_power_on",
"flood_lights_on",
"focus_position",
"pan_position",
"pdu_connected",
"ptz_controller_connected",
"snmpd_trap_running",
"tilt_position",
"zoom_position",
]
)
MKAT_VDS_COMMAND_LIST = frozenset(
[
"CameraPowerOn",
"FloodLightOn",
"Focus",
"Pan",
"PresetClear",
"PresetGoto",
"PresetSet",
"Stop",
"Tilt",
"Zoom",
]
)
class test_XmiSimddDeviceIntegration(ClassCleanupUnittestMixin, unittest.TestCase):
longMessage = True
@classmethod
def setUpClassWithCleanup(cls):
cls.tango_db = cleanup_tempfile(cls, prefix="tango", suffix=".db")
cls.data_descr_files = []
cls.data_descr_files.append(
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "MkatVds.xmi"
)
)
cls.data_descr_files.append(
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "MkatVds_SimDD.json"
)
)
cls.device_name = "test/nodb/tangodeviceserver"
model = tango_sim_generator.configure_device_models(
cls.data_descr_files, cls.device_name
)
cls.TangoDeviceServer = tango_sim_generator.get_tango_device_server(
model, cls.data_descr_files
)[0]
cls.tango_context = DeviceTestContext(
cls.TangoDeviceServer, device_name=cls.device_name, db=cls.tango_db
)
with patch("tango_simlib.utilities.helper_module.get_database"):
start_thread_with_cleanup(cls, cls.tango_context)
def setUp(self):
super(test_XmiSimddDeviceIntegration, self).setUp()
self.device = self.tango_context.device
self.instance = self.TangoDeviceServer.instances[self.device.name()]
def test_attribute_list(self):
"""Test device attribute list.
Check whether the attributes specified in the POGO generated xmi file
are added to the TANGO device
"""
attributes = set(self.device.get_attribute_list())
default_attributes = helper_module.DEFAULT_TANGO_DEVICE_ATTRIBUTES
self.assertEqual(
MKAT_VDS_ATTRIBUTE_LIST,
attributes - default_attributes,
"Actual tango device attribute list differs from expected "
"list! \n\n Missing attributes: \n {}".format(
MKAT_VDS_ATTRIBUTE_LIST - attributes
),
)
def test_command_list(self):
"""Testing device command list.
Check that the command list in the Tango device matches with the one
specified in the SimDD data description file.
"""
actual_device_commands = set(self.device.get_command_list())
self.assertEquals(
actual_device_commands - helper_module.DEFAULT_TANGO_DEVICE_COMMANDS,
MKAT_VDS_COMMAND_LIST,
"The commands specified in the SimDD file are not present in" " the device",
)
class test_SourceSimulatorInfo(unittest.TestCase):
"""This class is not testing the code, but only testing that the test XMI and SimDD
files are consistent with each other.
"""
longMessage = True
def setUp(self):
super(test_SourceSimulatorInfo, self).setUp()
self.sim_xmi_file = [
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "MkatVds.xmi"
)
]
self.simdd_json_file = [
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "MkatVds_SimDD.json"
)
]
self.simdd_parser = simdd_json_parser.SimddParser()
self.xmi_parser = sim_xmi_parser.XmiParser()
self.xmi_parser.parse(self.sim_xmi_file[0])
self.simdd_parser.parse(self.simdd_json_file[0])
def test_source_data_attributes(self):
"""Testing attribute information from data files.
Check if the attribute information in the SimDD is consistent with the"
information captured in the XMI file generated using POGO.
"""
xmi_parser_attributes = self.xmi_parser.get_device_attribute_metadata()
simdd_parser_attributes = self.simdd_parser.get_device_attribute_metadata()
for attribute_name in MKAT_VDS_ATTRIBUTE_LIST:
self.assertIn(
attribute_name,
xmi_parser_attributes,
"The attribute '{}' is missing from the file: '{}'.".format(
attribute_name, self.sim_xmi_file[0]
),
)
for attribute_name in simdd_parser_attributes:
self.assertIn(
attribute_name,
xmi_parser_attributes,
"The attribute '{}' specified in the file: '{}' is not"
" captured in the main config file: '{}'.".format(
attribute_name, self.simdd_json_file[0], self.sim_xmi_file[0]
),
)
def test_source_data_commands(self):
"""Testing command information from data files.
Check if the commands information in the SimDD is consistent with the"
information captured in the XMI file generated using POGO.
"""
xmi_parser_commands = self.xmi_parser.get_device_command_metadata()
simdd_parser_commands = self.simdd_parser.get_device_command_metadata()
for command_name in MKAT_VDS_COMMAND_LIST:
self.assertIn(
command_name,
xmi_parser_commands,
"The command '{}' is missing from the file: '{}'.".format(
command_name, self.sim_xmi_file[0]
),
)
for command_name in simdd_parser_commands:
self.assertIn(
command_name,
xmi_parser_commands,
"The command '{}' specified in the file: '{}' is not captured"
"in the main config file: '{}'.".format(
command_name, self.simdd_json_file[0], self.sim_xmi_file[0]
),
)
def test_source_data_device_properties(self):
"""Testing device properties information from data files.
Check if the device properties information in the SimDD is consistent with the
information captured in the XMI file generated using POGO.
"""
xmi_parser_properties = self.xmi_parser.get_device_properties_metadata(
"deviceProperties"
)
simdd_parser_properties = self.simdd_parser.get_device_properties_metadata(
"deviceProperties"
)
for property_name in simdd_parser_properties:
self.assertIn(
property_name,
xmi_parser_properties,
"The property '{}' specified in the file: '{}' is not captured"
" in the main config file: '{}'.".format(
property_name, self.simdd_json_file[0], self.sim_xmi_file[0]
),
)
def test_source_data_class_properties(self):
"""Testing if the class properties information in the SimDD is consistent with the
information captured in the XMI file generated using POGO.
"""
xmi_parser_properties = self.xmi_parser.get_device_properties_metadata(
"classProperties"
)
simdd_parser_properties = self.simdd_parser.get_device_properties_metadata(
"classProperties"
)
for property_name in simdd_parser_properties:
self.assertIn(
property_name,
xmi_parser_properties,
"The property '{}' specified in the file: '{}' is not captured"
" in the main config file: '{}'.".format(
property_name, self.simdd_json_file[0], self.sim_xmi_file[0]
),
)
class test_XmiSimddSupplementaryDeviceIntegration(
ClassCleanupUnittestMixin, unittest.TestCase
):
"""A test class that tests the use of both the xmi and simdd.
This ensures that the specified parameters in the simdd override that of
the xmi when a simulator is generated.
"""
longMessage = True
@classmethod
def setUpClassWithCleanup(cls):
cls.tango_db = cleanup_tempfile(cls, prefix="tango", suffix=".db")
cls.data_descr_files = []
cls.data_descr_files.append(
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "Weather.xmi"
)
)
cls.data_descr_files.append(
pkg_resources.resource_filename(
"tango_simlib.tests.config_files", "Weather_SimDD_2.json"
)
)
cls.device_name = "test/nodb/tangodeviceserver"
model = tango_sim_generator.configure_device_models(
cls.data_descr_files, cls.device_name
)
cls.TangoDeviceServer = tango_sim_generator.get_tango_device_server(
model, cls.data_descr_files
)[0]
cls.tango_context = DeviceTestContext(
cls.TangoDeviceServer, device_name=cls.device_name, db=cls.tango_db
)
with patch("tango_simlib.utilities.helper_module.get_database"):
start_thread_with_cleanup(cls, cls.tango_context)
def setUp(self):
super(test_XmiSimddSupplementaryDeviceIntegration, self).setUp()
self.device = self.tango_context.device
self.instance = self.TangoDeviceServer.instances[self.device.name()]
def test_xmi_simdd_attribute_parameters_when_both_specified(self):
"""Testing attribute parameters when both xmi and simdd are specified.
Check whether the attribute parameters specified in the xmi and
simdd files are properly parsed to the device and also ensuring that
those of the simdd override the ones in xmi in the configured model
"""
attr_with_overrriden_info = "temperature"
simdd_specified_temperature_attr_params = {
"description": "Current actual " "temperature outside near the telescope.",
"min_value": "-15",
"max_value": "55",
}
for data_file in self.data_descr_files:
if ".xmi" in data_file.lower():
xmi_parser = sim_xmi_parser.XmiParser()
xmi_parser.parse(data_file)
expected_device_attr_xmi_info = xmi_parser.get_device_attribute_metadata()
expected_device_temperature_attr_overridden_info = dict(
expected_device_attr_xmi_info[attr_with_overrriden_info],
**simdd_specified_temperature_attr_params
)
# Creating a copy of the attribute info as specified in the xmi and
# overriding it with that specified in the simdd then create a
# structure of what is expected as a result of the combination of the two.
expected_device_attr_xmi_info_copy = expected_device_attr_xmi_info.copy()
expected_device_attr_xmi_info_copy[
attr_with_overrriden_info
] = expected_device_temperature_attr_overridden_info
expected_device_attr_xmi_overridden = expected_device_attr_xmi_info_copy
sim_quantities = self.instance.model.sim_quantities
for expected_quantity in expected_device_attr_xmi_info.keys():
self.assertIn(
expected_quantity,
sim_quantities,
"The attribute {} is not in the parsed "
"attribute list".format(expected_quantity),
)
actual_device_attr_info = sim_quantities[expected_quantity].meta
for prop in expected_device_attr_xmi_info[expected_quantity]:
# The 'inherited' parameter is not part of the Tango device attribute
# properties.
if prop == "inherited":
continue
if prop not in simdd_specified_temperature_attr_params.keys():
self.assertEquals(
expected_device_attr_xmi_info[expected_quantity][prop],
actual_device_attr_info[prop],
"The {} quantity expected value for the parameter "
"'{}' does not match with the actual value in the "
"device model".format(expected_quantity, prop),
)
self.assertEquals(
expected_device_attr_xmi_overridden[expected_quantity][prop],
actual_device_attr_info[prop],
"The {} quantity expected value for the overridden "
"parameter '{}' does not match with the actual value "
"in the device model".format(expected_quantity, prop),
)
def test_xmi_simdd_command_parameters_when_both_specified(self):
"""Testing command parameters when both xmi and simdd are specified.
Check whether the command parameters specified in the xmi and
simdd files are properly parsed to the device and also ensuring that
those of the simdd override the ones in xmi in the configured model
"""
cmd_with_overrriden_info = "On"
simdd_specified_on_cmd_params = {
"doc_in": "No input parameter required",
"doc_out": "Command responds only",
}
for data_file in self.data_descr_files:
if ".xmi" in data_file.lower():
xmi_parser = sim_xmi_parser.XmiParser()
xmi_parser.parse(data_file)
expected_device_cmd_xmi_info = xmi_parser.get_device_command_metadata()
expected_device_on_cmd_overridden_info = dict(
expected_device_cmd_xmi_info[cmd_with_overrriden_info],
**simdd_specified_on_cmd_params
)
# Creating a copy of the command info as specified in the xmi and
# overriding it with that specified in the simdd then create a
# structure of what is expected as a result of the combination of the two.
expected_device_cmd_xmi_info_copy = expected_device_cmd_xmi_info.copy()
expected_device_cmd_xmi_info_copy[
cmd_with_overrriden_info
] = expected_device_on_cmd_overridden_info
expected_device_cmd_xmi_overridden = expected_device_cmd_xmi_info_copy
sim_actions = self.instance.model.sim_actions_meta
for expected_action in expected_device_cmd_xmi_info.keys():
if expected_action not in helper_module.DEFAULT_TANGO_DEVICE_COMMANDS:
self.assertIn(
expected_action,
sim_actions.keys(),
"The command {} is not in the parsed "
"command list".format(expected_action),
)
actual_device_attr_info = sim_actions[expected_action]
for prop in expected_device_cmd_xmi_info[expected_action]:
# The 'inherited' parameter is not part of the Tango device attribute
# properties.
if prop == "inherited":
continue
if prop not in simdd_specified_on_cmd_params.keys():
self.assertEquals(
expected_device_cmd_xmi_info[expected_action][prop],
actual_device_attr_info[prop],
"The {} action expected value for the parameter "
"'{}' does not match with the actual value in the "
"device model".format(expected_action, prop),
)
self.assertEquals(
expected_device_cmd_xmi_overridden[expected_action][prop],
actual_device_attr_info[prop],
"The {} action expected value for the overridden "
"parameter '{}' does not match with the actual value "
"in the device model".format(expected_action, prop),
)
|
<filename>matterapi/endpoints/sync_api/webhooks.py
""" Module to access the Webhooks endpoints """
# pylint: disable=too-many-lines,too-many-locals,too-many-public-methods,too-few-public-methods
from typing import Any, Dict, List, Optional, Union
from pydantic import BaseModel
from ...models import (
CreateIncomingWebhookJsonBody,
CreateOutgoingWebhookJsonBody,
IncomingWebhook,
OutgoingWebhook,
StatusOK,
UpdateIncomingWebhookJsonBody,
UpdateOutgoingWebhookJsonBody,
)
from ..base import ApiBaseClass
class WebhooksApi(ApiBaseClass):
"""Endpoints for creating, getting and updating webhooks."""
def get_incoming_webhooks(
self,
*,
page: Optional[int] = 0,
per_page: Optional[int] = 60,
team_id: Optional[str] = None,
) -> List[IncomingWebhook]:
"""List incoming webhooks
Get a page of a list of incoming webhooks. Optionally filter for a
specific team using query parameters.
Permissions:
`manage_webhooks` for the system or `manage_webhooks` for
the specific team.
Api Reference:
`GetIncomingWebhooks <https://api.mattermost.com/#operation/GetIncomingWebhooks>`_
"""
url = "/hooks/incoming"
params: Dict[str, Any] = {
"page": page,
"per_page": per_page,
"team_id": team_id,
}
params = {k: v for k, v in params.items() if v is not None}
request_kwargs = {
"url": url,
"params": params,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.get(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = []
_response200 = response.json()
for response200_item_data in _response200:
response200_item = IncomingWebhook.parse_obj(response200_item_data)
response200.append(response200_item)
return response200
return response
def create_incoming_webhook(
self,
*,
json_body: Union[CreateIncomingWebhookJsonBody, Dict],
) -> IncomingWebhook:
"""Create an incoming webhook
Create an incoming webhook for a channel.
`manage_others_incoming_webhooks` for the team the webhook is in if the
user is different than the requester.
Permissions:
`manage_webhooks` for the team the webhook is in.
Api Reference:
`CreateIncomingWebhook <https://api.mattermost.com/#operation/CreateIncomingWebhook>`_
"""
url = "/hooks/incoming"
if isinstance(json_body, BaseModel):
json_json_body = json_body.dict(exclude_unset=True)
else:
json_json_body = json_body
request_kwargs = {
"url": url,
"json": json_json_body,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.post(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 201:
response201 = IncomingWebhook.parse_obj(response.json())
return response201
return response
def get_incoming_webhook(
self,
hook_id: str,
) -> IncomingWebhook:
"""Get an incoming webhook
Get an incoming webhook given the hook id.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`GetIncomingWebhook <https://api.mattermost.com/#operation/GetIncomingWebhook>`_
"""
url = f"/hooks/incoming/{hook_id}"
request_kwargs = {
"url": url,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.get(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = IncomingWebhook.parse_obj(response.json())
return response200
return response
def update_incoming_webhook(
self,
hook_id: str,
*,
json_body: Union[UpdateIncomingWebhookJsonBody, Dict],
) -> IncomingWebhook:
"""Update an incoming webhook
Update an incoming webhook given the hook id.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`UpdateIncomingWebhook <https://api.mattermost.com/#operation/UpdateIncomingWebhook>`_
"""
url = f"/hooks/incoming/{hook_id}"
if isinstance(json_body, BaseModel):
json_json_body = json_body.dict(exclude_unset=True)
else:
json_json_body = json_body
request_kwargs = {
"url": url,
"json": json_json_body,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.put(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = IncomingWebhook.parse_obj(response.json())
return response200
return response
def delete_incoming_webhook(
self,
hook_id: str,
) -> StatusOK:
"""Delete an incoming webhook
Delete an incoming webhook given the hook id.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`DeleteIncomingWebhook <https://api.mattermost.com/#operation/DeleteIncomingWebhook>`_
"""
url = f"/hooks/incoming/{hook_id}"
request_kwargs = {
"url": url,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.delete(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = StatusOK.parse_obj(response.json())
return response200
return response
def get_outgoing_webhooks(
self,
*,
page: Optional[int] = 0,
per_page: Optional[int] = 60,
team_id: Optional[str] = None,
channel_id: Optional[str] = None,
) -> List[OutgoingWebhook]:
"""List outgoing webhooks
Get a page of a list of outgoing webhooks. Optionally filter for a
specific team or channel using query parameters.
Permissions:
`manage_webhooks` for the system or `manage_webhooks` for
the specific team/channel.
Api Reference:
`GetOutgoingWebhooks <https://api.mattermost.com/#operation/GetOutgoingWebhooks>`_
"""
url = "/hooks/outgoing"
params: Dict[str, Any] = {
"page": page,
"per_page": per_page,
"team_id": team_id,
"channel_id": channel_id,
}
params = {k: v for k, v in params.items() if v is not None}
request_kwargs = {
"url": url,
"params": params,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.get(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = []
_response200 = response.json()
for response200_item_data in _response200:
response200_item = OutgoingWebhook.parse_obj(response200_item_data)
response200.append(response200_item)
return response200
return response
def create_outgoing_webhook(
self,
*,
json_body: Union[CreateOutgoingWebhookJsonBody, Dict],
) -> OutgoingWebhook:
"""Create an outgoing webhook
Create an outgoing webhook for a team.
`manage_others_outgoing_webhooks` for the team the webhook is in if the
user is different than the requester.
Permissions:
`manage_webhooks` for the team the webhook is in.
Api Reference:
`CreateOutgoingWebhook <https://api.mattermost.com/#operation/CreateOutgoingWebhook>`_
"""
url = "/hooks/outgoing"
if isinstance(json_body, BaseModel):
json_json_body = json_body.dict(exclude_unset=True)
else:
json_json_body = json_body
request_kwargs = {
"url": url,
"json": json_json_body,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.post(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 201:
response201 = OutgoingWebhook.parse_obj(response.json())
return response201
return response
def get_outgoing_webhook(
self,
hook_id: str,
) -> OutgoingWebhook:
"""Get an outgoing webhook
Get an outgoing webhook given the hook id.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`GetOutgoingWebhook <https://api.mattermost.com/#operation/GetOutgoingWebhook>`_
"""
url = f"/hooks/outgoing/{hook_id}"
request_kwargs = {
"url": url,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.get(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = OutgoingWebhook.parse_obj(response.json())
return response200
return response
def update_outgoing_webhook(
self,
hook_id: str,
*,
json_body: Union[UpdateOutgoingWebhookJsonBody, Dict],
) -> OutgoingWebhook:
"""Update an outgoing webhook
Update an outgoing webhook given the hook id.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`UpdateOutgoingWebhook <https://api.mattermost.com/#operation/UpdateOutgoingWebhook>`_
"""
url = f"/hooks/outgoing/{hook_id}"
if isinstance(json_body, BaseModel):
json_json_body = json_body.dict(exclude_unset=True)
else:
json_json_body = json_body
request_kwargs = {
"url": url,
"json": json_json_body,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.put(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = OutgoingWebhook.parse_obj(response.json())
return response200
return response
def delete_outgoing_webhook(
self,
hook_id: str,
) -> StatusOK:
"""Delete an outgoing webhook
Delete an outgoing webhook given the hook id.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`DeleteOutgoingWebhook <https://api.mattermost.com/#operation/DeleteOutgoingWebhook>`_
"""
url = f"/hooks/outgoing/{hook_id}"
request_kwargs = {
"url": url,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.delete(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = StatusOK.parse_obj(response.json())
return response200
return response
def regen_outgoing_hook_token(
self,
hook_id: str,
) -> StatusOK:
"""Regenerate the token for the outgoing webhook.
Regenerate the token for the outgoing webhook.
Permissions:
`manage_webhooks` for system or `manage_webhooks` for the
specific team or `manage_webhooks` for the channel.
Api Reference:
`RegenOutgoingHookToken <https://api.mattermost.com/#operation/RegenOutgoingHookToken>`_
"""
url = f"/hooks/outgoing/{hook_id}/regen_token"
request_kwargs = {
"url": url,
}
# pylint: disable-next=protected-access
with self.client._get_httpx_client() as httpx_client:
response = httpx_client.post(
**request_kwargs,
)
if self.skip_response_parsing:
return response
if response.status_code == 200:
response200 = StatusOK.parse_obj(response.json())
return response200
return response
|
# Copyright (c) 2019 Juniper Networks, Inc. All rights reserved.
import json
import uuid
from cfgm_common.tests import test_utils
from keystonemiddleware import auth_token
import mock
from vnc_api.vnc_api import Project
from vnc_api.vnc_api import VncApi
from vnc_cfg_api_server.tests import test_case
def get_token(user_name, project_name, domain_name, role_name, project_id=None,
domain_id=None):
token_dict = {
'X-User': user_name,
'X-User-Name': user_name,
'X-Project-Name': project_name,
'X-Project-Id': project_id or '',
'X-Domain-Id': domain_id or '',
'X-Domain-Name': domain_name,
'X-Role': role_name,
}
rval = json.dumps(token_dict)
return rval
def ks_admin_authenticate(self, response=None, headers=None):
rval = get_token('admin', 'admin', 'default-domain', 'cloud-admin')
new_headers = {}
new_headers['X-AUTH-TOKEN'] = rval
return new_headers
class TestPostAuthKeystone(test_case.ApiServerTestCase):
@classmethod
def setUpClass(cls):
extra_mocks = [
(auth_token, 'AuthProtocol', test_utils.FakeAuthProtocol),
(VncApi, '_authenticate', ks_admin_authenticate),
]
extra_config_knobs = [
('DEFAULTS', 'aaa_mode', 'rbac'),
('DEFAULTS', 'auth', 'keystone'),
('DEFAULTS', 'cloud_admin_role', 'cloud-admin'),
]
super(TestPostAuthKeystone, cls).setUpClass(
extra_mocks=extra_mocks,
extra_config_knobs=extra_config_knobs)
def test_default_kestone_domain_id_replaced(self):
admin_token = get_token(
'admin',
'admin',
'Default',
'cloud-admin',
uuid.uuid4().hex,
'default')
self._vnc_lib.set_auth_token(admin_token)
project = Project('project-%s' % self.id())
default_domain = self._vnc_lib.domain_read(['default-domain'])
with mock.patch('vnc_cfg_api_server.vnc_cfg_api_server.VncApiServer.'
'default_domain', new_callable=mock.PropertyMock) as\
dd_prop_mock:
dd_prop_mock.return_value = default_domain.serialize_to_json()
self._vnc_lib.project_create(project)
dd_prop_mock.assert_called()
class TestPostAuthKeystone2(test_case.ApiServerTestCase):
changed_default_doamin_id = 'changed-default-domain-id'
@classmethod
def setUpClass(cls):
extra_mocks = [
(auth_token, 'AuthProtocol', test_utils.FakeAuthProtocol),
(VncApi, '_authenticate', ks_admin_authenticate),
]
extra_config_knobs = [
('DEFAULTS', 'aaa_mode', 'rbac'),
('DEFAULTS', 'auth', 'keystone'),
('DEFAULTS', 'cloud_admin_role', 'cloud-admin'),
('KEYSTONE', 'default_domain_id', cls.changed_default_doamin_id),
]
super(TestPostAuthKeystone2, cls).setUpClass(
extra_mocks=extra_mocks,
extra_config_knobs=extra_config_knobs)
def test_custom_default_kestone_domain_id_replaced(self):
admin_token = get_token(
'admin',
'admin',
'Default',
'cloud-admin',
uuid.uuid4().hex,
self.changed_default_doamin_id)
self._vnc_lib.set_auth_token(admin_token)
project = Project('project-%s' % self.id())
default_domain = self._vnc_lib.domain_read(['default-domain'])
with mock.patch('vnc_cfg_api_server.vnc_cfg_api_server.VncApiServer.'
'default_domain', new_callable=mock.PropertyMock) as\
dd_prop_mock:
dd_prop_mock.return_value = default_domain.serialize_to_json()
self._vnc_lib.project_create(project)
dd_prop_mock.assert_called()
|
<filename>cellular/cellular.py
import collections
import itertools
import random
from .util import util
from fractions import Fraction
from PIL import Image, ImageDraw
from math import log2
class TotalisticCellularAutomaton:
def __init__(self, width, states=5, radius=1, colors=None, rules=None):
self.n_cells = width
self.n_states = states
self.radius = radius
if colors is None:
self.colors = ['black'] + [util.randcolor() for _ in range(self.n_states-1)]
else:
if len(colors) != self.n_states:
raise ValueError("Invalid number of colors. Expected {}.".format(self.n_states))
self.colors = colors
self.reseed()
n_rules = (2*self.radius + 1) * (self.n_states - 1) + 1
if rules is None:
self.rules = [0] + [random.randrange(1, self.n_states) for _ in range(n_rules - 1)]
else:
if len(rules) != n_rules:
raise ValueError("Invalid number of rules. Expected {}.".format(n_rules))
self.rules = rules
def run(self, ngens):
for n in range(ngens):
c = self.next_gen()
def resume(self, ngens):
self.history = [self.history[-1]]
self.history_set = {tuple(self.history[0])}
self.run(ngens)
def draw(self):
n = len(self.history)
m = len(self.history[0])
image = Image.new('RGB', (m, n))
draw = ImageDraw.Draw(image)
for i in range(n):
for j in range(m):
state = self.history[i][j]
draw.point((j, i), fill=self.colors[state])
return image
def print_stats(self):
print('[' + ' '.join(str(r) for r in self.rules) + ']')
print(("{:10s} " * 6).format('lambda', 'lambda_t', 'entropy', 'entropy_t', 'entropy_p', 'entropy_a'))
print(("{:10.8f} " * 6).format(self.lam, self.lam_t, self.entropy, self.entropy_t, self.entropy_p, self.entropy_a))
def neighbor_sum(self, pos):
return sum(self.cells[(pos+i)%self.n_cells] for i in range(-self.radius, self.radius+1))
def next_gen(self):
self.cells = [self.rules[self.neighbor_sum(i)] for i in range(self.n_cells)]
'''
if self.cells == self.history[-1]:
return 1
elif tuple(self.cells) in self.history_set:
return 2
'''
self.history.append(self.cells)
self.history_set.add(tuple(self.cells))
def decimate(self):
nonzeroes = [i for i in range(len(self.rules)) if self.rules[i] != 0]
if len(nonzeroes) != 0:
self.rules[random.choice(nonzeroes)] = 0
def reseed(self):
self.cells = [random.randrange(0, self.n_states) for _ in range(self.n_cells)]
self.history = [self.cells]
self.history_set = {tuple(self.cells)}
@property
def lam(self):
"""Currently only works with machines of radius 1 and 5 states"""
N = 2*self.radius + 1
T = pow(self.n_states, 2*self.radius +1)
def n(s):
tot = 0
for i in range(0, len(self.rules)):
if self.rules[i] == s:
tot += util.C(N, i, self.n_states-1)
return tot
return 1.0 - n(0) / T
@property
def lam_t(self):
return 1.0 - self.rules.count(0) / len(self.rules)
@property
def entropy(self):
N = 2*self.radius + 1
def n(s):
tot = 0
for i in range(0, len(self.rules)):
if self.rules[i] == s:
tot += util.C(N, i, self.n_states-1)
return tot
ent = 0
for i in range(0, self.n_states):
p_s = n(i) / pow(self.n_states, N)
if p_s == 0:
continue
else:
ent += p_s * log2(p_s)
return -1 * ent
@property
def entropy_t(self):
probs = [self.rules.count(state) / len(self.rules) for state in range(self.n_states)]
return -sum(p*log2(p) for p in probs if p != 0)
def get_probs(self, iters=5):
N = self.radius*2 + 1
probs = [Fraction(1, self.n_states) for _ in range(self.n_states)]
for x in range(iters):
new_probs = [0 for _ in probs]
for neighborhood in itertools.product(*[range(self.n_states) for _ in range(N)]):
p_n = util.product(probs[state] for state in neighborhood)
new_state = self.rules[sum(neighborhood)]
new_probs[new_state] += p_n
probs = new_probs
return [float(p) for p in probs]
@property
def entropy_p(self):
return -sum(p*log2(p) for p in self.get_probs() if p != 0)
@property
def entropy_a(self):
return -sum(p*log2(p) for p in self.get_real_probs() if p != 0)
def get_real_probs(self):
total = len(self.history) * len(self.history[0])
c = collections.Counter()
for row in self.history:
c.update(row)
probs = [c[state]/total for state in range(self.n_states)]
return probs
def __str__(self):
return '-'.join(str(r) for r in self.rules)
|
#!/usr/bin/env python
import os
import numpy as np
import tables
import pandas
from opty.utils import parse_free
def compute_gain_error(filename):
# root mean square of gain error
df = load_results_table(filename)
rms = []
for run_id, sim_dur, sample_rate in zip(df['run_id'],
df['sim_duration'],
df['sample_rate']):
run_dict = load_run(filename, run_id)
num_states = len(run_dict['initial_conditions'])
num_time_steps = int(sim_dur * sample_rate)
__, __, known_gains = parse_free(run_dict['known_solution'],
num_states, 0, num_time_steps)
__, __, optimal_gains = parse_free(run_dict['optimal_solution'],
num_states, 0, num_time_steps)
rms.append(np.sqrt(np.sum((known_gains - optimal_gains)**2)))
df['RMS of Gains'] = np.asarray(rms)
return df
def load_results_table(filename):
handle = tables.openFile(filename, 'r')
df = pandas.DataFrame.from_records(handle.root.results[:])
handle.close()
return df
def load_run(filename, run_id):
handle = tables.openFile(filename, 'r')
group = getattr(handle.root.arrays, run_id)
d = {}
for array_name in group.__members__:
d[array_name] = getattr(group, array_name)[:]
handle.close()
return d
def parse_ipopt_output(file_name):
"""Returns a dictionary with the IPOPT summary results.
Notes
-----
This is an example of the summary at the end of the file:
Number of Iterations....: 1013
(scaled) (unscaled)
Objective...............: 2.8983286604029537e-04 2.8983286604029537e-04
Dual infeasibility......: 4.7997817057236348e-09 4.7997817057236348e-09
Constraint violation....: 9.4542809291867735e-09 9.8205754639479892e-09
Complementarity.........: 0.0000000000000000e+00 0.0000000000000000e+00
Overall NLP error.......: 9.4542809291867735e-09 9.8205754639479892e-09
Number of objective function evaluations = 6881
Number of objective gradient evaluations = 1014
Number of equality constraint evaluations = 6900
Number of inequality constraint evaluations = 0
Number of equality constraint Jacobian evaluations = 1014
Number of inequality constraint Jacobian evaluations = 0
Number of Lagrangian Hessian evaluations = 0
Total CPU secs in IPOPT (w/o function evaluations) = 89.023
Total CPU secs in NLP function evaluations = 457.114
"""
with open(file_name, 'r') as f:
output = f.readlines()
results = {}
for line in output:
if 'Number of Iterations....:' in line and 'Maximum' not in line:
results['num_iterations'] = int(line.split(':')[1].strip())
elif 'Number of objective function evaluations' in line:
results['num_obj_evals'] = int(line.split('=')[1].strip())
elif 'Number of objective gradient evaluations' in line:
results['num_obj_grad_evals'] = int(line.split('=')[1].strip())
elif 'Number of equality constraint evaluations' in line:
results['num_con_evals'] = int(line.split('=')[1].strip())
elif 'Number of equality constraint Jacobian evaluations' in line:
results['num_con_jac_evals'] = int(line.split('=')[1].strip())
elif 'Total CPU secs in IPOPT (w/o function evaluations)' in line:
results['time_ipopt'] = float(line.split('=')[1].strip())
elif 'Total CPU secs in NLP function evaluations' in line:
results['time_func_evals'] = float(line.split('=')[1].strip())
return results
def create_database(file_name):
"""Creates an empty optimization results database on disk if it doesn't
exist."""
class RunTable(tables.IsDescription):
run_id = tables.StringCol(40) # sha1 hashes are 40 char long
init_type = tables.StringCol(10)
datetime = tables.Time32Col()
num_links = tables.Int32Col()
sim_duration = tables.Float32Col()
sample_rate = tables.Float32Col()
sensor_noise = tables.BoolCol()
num_iterations = tables.Int32Col()
num_obj_evals = tables.Int32Col()
num_obj_grad_evals = tables.Int32Col()
num_con_evals = tables.Int32Col()
num_con_jac_evals = tables.Int32Col()
time_ipopt = tables.Float32Col()
time_func_evals = tables.Float32Col()
if not os.path.isfile(file_name):
title = 'Inverted Pendulum Direct Collocation Results'
h5file = tables.open_file(file_name,
mode='w',
title=title)
h5file.create_table('/', 'results', RunTable,
'Optimization Results Table')
h5file.create_group('/', 'arrays', 'Optimization Parameter Arrays')
h5file.close()
def add_results(file_name, results):
if not os.path.isfile(file_name):
create_database(file_name)
h5file = tables.open_file(file_name, mode='a')
print('Adding run {} to the database.'.format(results['run_id']))
group_name = 'Optimization Run #{}'.format(results['run_id'])
run_array_dir = h5file.create_group(h5file.root.arrays,
results['run_id'],
group_name)
arrays = ['initial_guess',
'known_solution',
'optimal_solution',
'initial_guess_constraints',
'known_solution_constraints',
'optimal_solution_constraints',
'initial_conditions',
'lateral_force']
for k in arrays:
v = results.pop(k)
h5file.create_array(run_array_dir, k, v)
table = h5file.root.results
opt_row = table.row
for k, v in results.items():
opt_row[k] = v
opt_row.append()
table.flush()
h5file.close()
|
'''
Module containing Univariate Function Noise Generators.
Classes embody Stochastic Noise Distributions,
combined additively or multiplicatively with function gradient.
'''
import numpy
from . import univariate
class Beta:
'''
Beta Probability Distribution Function
Mathematically, p(x) = x ^ (p1 - 1) * (1 - x) ^ (p2 - 1) / integral_(0, 1)((x ^ (p1 - 1) * (1 - x) ^ (p2 - 1)) dx)
'''
def __init__(self, alpha = None, beta = None, scale = None):
'''
Constructor
: param alpha : p1, as given in its mathematical expression
: param beta : p2, as given in its mathematical expression
: param scale : factor by which random sample is scaled
'''
self.alpha = alpha if alpha is not None else 1.0
self.beta = beta if beta is not None else 1.0
self.scale = scale if scale is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return self.scale * numpy.random.beta(self.alpha, self.beta)
class Bernoulli:
'''
Bernoulli Probability Distribution Function
Mathematically, p(x) = C(p1, x) * p2 ^ x * (1 - p2) ^ (p1 - x)
'''
def __init__(self, probability = None, trials = None, scale = None):
'''
Constructor
: param probability : p2, as given in its mathematical expression
: param trials : p1, as given in its mathematical expression
: param scale : factor by which random sample is scaled
'''
self.probability = probability if probability is not None else 0.5
self.trials = trials if trials is not None else 1.0
self.scale = scale if scale is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return self.scale * numpy.random.binomial(self.trials, self.probability)
class Gamma:
'''
Gamma Probability Distribution Function
Mathematically, p(x) = x ^ (p1 - 1) * exp(-x / p2) / ((p2 ^ p1) * Gamma(p1))
'''
def __init__(self, exponent = None, scale = None):
'''
Constructor
: param exponent : p1, as given in its mathematical expression
: param scale : p2, as given in its mathematical expression
'''
self.exponent = exponent if exponent is not None else 1.0
self.scale = scale if scale is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return numpy.random.gamma(self.exponent, self.scale)
class Geometric:
'''
Geometric Probability Distribution Function
Mathematically, p(x) = p1 * (1 - p1) ^ (x - 1)
'''
def __init__(self, probability = None, scale = None):
'''
Constructor
: param probability : p1, as given in its mathematical expression
: param scale : factor by which random sample is scaled
'''
self.probability = probability if probability is not None else 0.5
self.scale = scale if scale is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return self.scale * numpy.random.geometric(self.probability)
class Gaussian:
'''
Gaussian Probability Distribution Function
Mathematically, p(x) = exp(-(x - p1) ^ 2 / (2 * p2 ^ 2)) / (2 * pi * p2 ^ 2) ^ 0.5
'''
def __init__(self, mean = None, variance = None):
'''
Constructor
: param mean : p1, as given in its mathematical expression
: param variance : p2, as given in its mathematical expression
'''
self.mean = mean if mean is not None else 0.0
self.variance = variance if variance is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return numpy.random.normal(self.mean, self.variance)
class Poisson:
'''
Poisson Probability Distribution Function
Mathematically, p(x) = p1 ^ x * exp(-x) / x!
'''
def __init__(self, lamda = None, scale = None):
'''
Constructor
: param lamda : p1, as given in its mathematical expression
: param scale : factor by which random sample is scaled
'''
self.lamda = lamda if lamda is not None else 1.0
self.scale = scale if scale is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return self.scale * numpy.random.poisson(self.lamda)
class Uniform:
'''
Uniform Probability Distribution Function
Mathematically, p(x) = 1 / (p1 - p2)
'''
def __init__(self, lowerlimit = None, upperlimit = None):
'''
Constructor
: param upperlimit : p2, as given in its mathematical expression
: param lowerlimit : p1, as given in its mathematical expression
'''
self.lowerlimit = lowerlimit if lowerlimit is not None else 0.0
self.upperlimit = upperlimit if upperlimit is not None else 1.0
def sample(self):
'''
Method to return a random sample from the probability distribution
'''
return numpy.random.uniform(self.lowerlimit, self.upperlimit)
# apply noise before piecing together as Curve
class NoisyUnivariate(univariate.Univariate):
'''
Univariate Function Stochastic Noise Wrapper
'''
def __init__(self, pureunivariate, noise, additive = None):
'''
Constructor
: param pureunivariate : pure univariate function to wrap with noises
: param noise : noise probability distribution function
: param additive : additive or multiplicative wrapper
'''
univariate.Univariate.__init__(self, pureunivariate.begin, pureunivariate.end, pureunivariate.value, pureunivariate.point, pureunivariate.shift, pureunivariate.lowerlimit, pureunivariate.upperlimit, pureunivariate.stepsize, pureunivariate.functiondepictor, pureunivariate.derivativedepictor)
self.purefunction = pureunivariate.function
self.purederivative = pureunivariate.derivative
self.noise = noise
self.additive = additive if additive is not None else True
def function(self, point):
'''
Method to evaluate noisy univariate function
: param inputvector : point of evaluation in parameter space
: returns : evaluated function value at point in parameter space
'''
value = self.purefunction(point)
noise = self.noise.sample()
return value + noise * (point - self.point) if self.additive else value * noise
def derivative(self, point):
'''
Method to evaluate noisy univariate derivative
: param inputvector : point of evaluation in parameter space
: returns : evaluated derivative value at point in parameter space
'''
value = self.purederivative(point)
noise = self.noise.sample()
return value + noise if self.additive else value * noise
|
<gh_stars>10-100
# Copyright (c) 2016 Uber Technologies, Inc.
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN
# THE SOFTWARE.
from __future__ import absolute_import, unicode_literals, print_function
from textwrap import dedent
import pytest
from thriftrw.errors import ThriftCompilerError
from thriftrw.loader import Loader
def test_load_from_file(tmpdir):
tmpdir.join('my_service.thrift').write('''
struct Foo {
1: required string a
2: optional string b
}
''')
my_service = Loader().load(str(tmpdir.join('my_service.thrift')))
my_service.Foo(b='b', a='a')
def test_load_from_file_missing_requiredness(tmpdir):
tmpdir.join('my_service.thrift').write('''
struct Foo {
1: required string a
2: string b
}
''')
with pytest.raises(ThriftCompilerError) as exc_info:
Loader().load(str(tmpdir.join('my_service.thrift')))
assert (
'"b" of "Foo" on line 4 does not explicitly specify requiredness.'
in str(exc_info)
)
def test_load_from_file_non_strict_missing_requiredness(tmpdir):
tmpdir.join('my_service.thrift').write('''
struct Foo {
1: required string a
2: string b
}
exception Bar {
1: string message
}
''')
loader = Loader(strict=False)
m = loader.load(str(tmpdir.join('my_service.thrift')))
m.Foo(b='b', a='a')
m.Bar(message='foo')
m.Bar()
def test_caching(tmpdir, monkeypatch):
tmpdir.join('my_service.thrift').write('''
struct Foo {
1: required string a
2: optional string b
}
''')
path = str(tmpdir.join('my_service.thrift'))
loader = Loader()
mod1 = loader.load(path)
mod2 = loader.load(path)
assert mod1 is mod2
@pytest.mark.unimport('foo.bar.svc')
def test_install_absolute(tmpdir, monkeypatch):
module_root = tmpdir.mkdir('foo')
module_root.join('__init__.py').ensure()
thrift_file = module_root.join('service.thrift')
thrift_file.write(
'struct Foo { 1: required string a; 2: optional string b }'
)
py_file = module_root.join('bar.py')
py_file.write(
dedent('''
import thriftrw
thriftrw.install(%r, name='svc')
''' % str(thrift_file))
)
monkeypatch.syspath_prepend(str(tmpdir))
from foo.bar.svc import Foo
assert Foo(a='bar') == Foo(a='bar')
@pytest.mark.unimport('foo.service', 'foo.bar')
def test_install_relative(tmpdir, monkeypatch):
module_root = tmpdir.mkdir('foo')
module_root.join('service.thrift').write('struct Bar {}')
module_root.join('bar.py').write(dedent('''
from __future__ import absolute_import
from .service import Bar
'''))
module_root.join('__init__.py').write(dedent('''
import thriftrw
thriftrw.install('service.thrift')
'''))
monkeypatch.syspath_prepend(str(tmpdir))
from foo.bar import Bar
assert Bar() == Bar()
@pytest.mark.unimport('foo.service')
def test_install_twice(tmpdir, monkeypatch):
module_root = tmpdir.mkdir('foo')
module_root.join('__init__.py').write(dedent('''
import thriftrw
def go():
return thriftrw.install('service.thrift')
'''))
module_root.join('service.thrift').write(
'struct Foo { 1: required string a 2: optional string b }'
)
monkeypatch.syspath_prepend(str(tmpdir))
from foo import go
with pytest.raises(ImportError):
from foo.service import Foo
Foo()
assert go() is go()
from foo.service import Foo
assert go().Foo is Foo
|
# Copyright 2014 ETH Zurich
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
:mod:`scion` --- SCION packets
==============================
"""
# Stdlib
import copy
import struct
# SCION
from lib.defines import LINE_LEN, MAX_HOPBYHOP_EXT, SCION_PROTO_VERSION
from lib.errors import SCIONIndexError, SCIONParseError
from lib.packet.ext_hdr import ExtensionHeader
from lib.packet.ext_util import parse_extensions
from lib.packet.host_addr import HostAddrInvalidType, haddr_get_type
from lib.packet.ctrl_pld import SignedCtrlPayload
from lib.packet.opaque_field import OpaqueField
from lib.packet.packet_base import (
Serializable,
L4HeaderBase,
PacketBase,
PayloadRaw,
)
from lib.packet.path import SCIONPath, parse_path
from lib.packet.scion_addr import ISD_AS, SCIONAddr
from lib.packet.scion_l4 import parse_l4_hdr
from lib.packet.scmp.errors import (
SCMPBadDstType,
SCMPBadEnd2End,
SCMPBadHOFOffset,
SCMPBadHopByHop,
SCMPBadHost,
SCMPBadIOFOffset,
SCMPBadPktLen,
SCMPBadSrcType,
SCMPBadVersion,
)
from lib.packet.scmp.ext import SCMPExt
from lib.packet.scmp.hdr import SCMPHeader
from lib.packet.scmp.payload import SCMPPayload
from lib.packet.svc import SVCType
from lib.types import (
AddrType,
ExtHopByHopType,
ExtensionClass,
L4Proto,
)
from lib.util import Raw, calc_padding
class SCIONCommonHdr(Serializable):
"""
Encapsulates the common header for SCION packets.
"""
NAME = "SCIONCommonHdr"
LEN = 8
def __init__(self, raw=None): # pragma: no cover
self.version = 0 # Version of SCION packet.
self.dst_addr_type = None
self.src_addr_type = None
self.addrs_len = None # Length of the address block
self.total_len = None # Total length of the packet.
self.hdr_len = None # Header length including the path.
self._iof_idx = None # Index of the current Info Opaque Field
self._hof_idx = None # Index of the current Hop Opaque Field
self.next_hdr = None # Type of the next hdr field (IP protocol numbers)
super().__init__(raw)
def _parse(self, raw):
"""
Parses the raw data and populates the fields accordingly.
"""
data = Raw(raw, self.NAME, self.LEN)
(types, self.total_len, self.hdr_len, iof_off, hof_off,
self.next_hdr) = struct.unpack("!HHBBBB", data.pop())
self.version = types >> 12
if self.version != SCION_PROTO_VERSION:
raise SCMPBadVersion("Unsupported SCION version: %s" % self.version)
self.dst_addr_type = (types >> 6) & 0x3f
self.src_addr_type = types & 0x3f
self.addrs_len, _ = SCIONAddrHdr.calc_lens(
self.dst_addr_type, self.src_addr_type)
if self.hdr_len_bytes() < self.LEN + self.addrs_len:
# Can't send an SCMP error, as there isn't enough information to
# parse the path and the l4 header.
raise SCIONParseError(
"hdr_len (%sB) < common header len (%sB) + addrs len (%sB) " %
(self.hdr_len_bytes(), self.LEN, self.addrs_len))
if iof_off == hof_off == 0:
self._iof_idx = self._hof_idx = 0
return
if iof_off == 0 or hof_off <= iof_off:
raise SCIONParseError(
"invalid CurrINF, CurrHF combination: (%s, %s) " % (iof_off, hof_off))
first_of_offset = self.LEN + self.addrs_len
# FIXME(kormat): NB this assumes that all OFs have the same length.
self._iof_idx = (iof_off * LINE_LEN - first_of_offset) // OpaqueField.LEN
self._hof_idx = (hof_off * LINE_LEN - first_of_offset) // OpaqueField.LEN
@classmethod
def from_values(cls, dst_type, src_type, next_hdr):
"""
Returns a SCIONCommonHdr object with the values specified.
:param int dst_type: Destination address type.
:param int src_type: Source address type.
:param int next_hdr: Next header type.
"""
inst = cls()
inst.dst_addr_type = dst_type
inst.src_addr_type = src_type
inst.addrs_len, _ = SCIONAddrHdr.calc_lens(dst_type, src_type)
inst.next_hdr = next_hdr
inst.total_len = cls.LEN + inst.addrs_len
inst.hdr_len = cls.bytes_to_hdr_len(inst.total_len)
inst._iof_idx = inst._hof_idx = 0
return inst
def pack(self):
packed = []
types = ((self.version << 12) | (self.dst_addr_type << 6) |
self.src_addr_type)
packed.append(struct.pack("!HHB", types, self.total_len, self.hdr_len))
curr_iof_p = curr_hof_p = 0
# FIXME(kormat): NB this assumes that all OFs have the same length.
if self._iof_idx or self._hof_idx:
curr_iof_p = self.LEN + self.addrs_len + self._iof_idx * OpaqueField.LEN
if self._hof_idx:
curr_hof_p = self.LEN + self.addrs_len + self._hof_idx * OpaqueField.LEN
packed.append(struct.pack("!BBB", curr_iof_p//LINE_LEN,
curr_hof_p//LINE_LEN, self.next_hdr))
raw = b"".join(packed)
assert len(raw) == self.LEN
return raw
def validate(self, pkt_len, path_len):
if pkt_len != self.total_len:
raise SCMPBadPktLen(
"Packet length incorrect. Expected: %sB. Actual: %sB" %
(self.total_len, pkt_len), 0)
if path_len == 0:
# Empty path
if self._iof_idx != 0:
raise SCMPBadIOFOffset(
"Non-zero IOF index for empty path: %s" % self._iof_idx)
if self._hof_idx != 0:
raise SCMPBadHOFOffset(
"Non-zero HOF index for empty path: %s" % self._hof_idx)
elif self._hof_idx == 0:
raise SCMPBadHOFOffset("Zero HOF index for non-empty path")
def get_of_idxs(self): # pragma: no cover
return self._iof_idx, self._hof_idx
def set_of_idxs(self, iof_idx, hof_idx): # pragma: no cover
self._iof_idx = iof_idx
self._hof_idx = hof_idx
@classmethod
def bytes_to_hdr_len(cls, bytes_):
assert bytes_ % LINE_LEN == 0
return bytes_ // LINE_LEN
def hdr_len_bytes(self): # pragma: no cover
return self.hdr_len * LINE_LEN
def __len__(self): # pragma: no cover
return self.LEN
def __str__(self):
values = {
"dst_addr_type": haddr_get_type(self.dst_addr_type).name(),
"src_addr_type": haddr_get_type(self.src_addr_type).name(),
"hdr_len": self.hdr_len_bytes(),
}
for i in ("version", "total_len", "_iof_idx", "_hof_idx", "next_hdr"):
values[i] = getattr(self, i)
return (
"CH ver: %(version)s, dst type: %(dst_addr_type)s, src type: %(src_addr_type)s, "
"total len: %(total_len)sB, hdr len: %(hdr_len)sB, "
"IOF idx: %(_iof_idx)s, HOF idx: %(_hof_idx)s, "
"next hdr: %(next_hdr)s" % values)
class SCIONAddrHdr(Serializable):
"""SCION Address header."""
NAME = "SCIONAddrHdr"
BLK_SIZE = 8
def __init__(self, raw_values=()): # pragma: no cover
"""
:param tuple raw:
Tuple of dst addr type, src addr type, and raw addr bytes.
"""
super().__init__()
self.dst = None
self.src = None
self._pad_len = None
self._total_len = None
if raw_values:
self._parse(*raw_values)
def _parse(self, dst_type, src_type, raw):
data = Raw(raw, self.NAME, self.calc_lens(dst_type, src_type)[0])
dst_ia = ISD_AS(data.pop(ISD_AS.LEN))
src_ia = ISD_AS(data.pop(ISD_AS.LEN))
dst_addr_t = haddr_get_type(dst_type)
dst_addr = dst_addr_t(data.pop(dst_addr_t.LEN))
self.dst = SCIONAddr.from_values(dst_ia, dst_addr)
src_addr_t = haddr_get_type(src_type)
src_addr = src_addr_t(data.pop(src_addr_t.LEN))
self.src = SCIONAddr.from_values(src_ia, src_addr)
self.update()
if self.src.host.TYPE == AddrType.SVC:
raise SCMPBadSrcType("Invalid source type: SVC")
@classmethod
def from_values(cls, dst, src): # pragma: no cover
"""
dst/src must be a :any:`SCIONAddr`
"""
inst = cls()
inst.dst = dst
inst.src = src
inst.update()
return inst
def pack(self):
self.update()
packed = []
packed.append(self.dst.isd_as.pack())
packed.append(self.src.isd_as.pack())
packed.append(self.dst.host.pack())
packed.append(self.src.host.pack())
packed.append(bytes(self._pad_len))
raw = b"".join(packed)
assert len(raw) % self.BLK_SIZE == 0
assert len(raw) == self._total_len
return raw
def validate(self): # pragma: no cover
if self.dst.host.TYPE == AddrType.SVC:
if self.dst.host.anycast() not in [SVCType.BS_A, SVCType.PS_A,
SVCType.CS_A, SVCType.SB_A]:
raise SCMPBadHost("Invalid dest SVC: %s" % self.dst.host.addr)
if self.src.host.TYPE == AddrType.SVC:
raise SCMPBadSrcType("Invalid source type: SVC")
def update(self):
self._total_len, self._pad_len = self.calc_lens(
self.dst.host.TYPE, self.src.host.TYPE)
@classmethod
def calc_lens(cls, dst_type, src_type):
try:
data_len = SCIONAddr.calc_len(dst_type)
except HostAddrInvalidType:
raise SCMPBadDstType(
"Unsupported dst address type: %s" % dst_type) from None
try:
data_len += SCIONAddr.calc_len(src_type)
except HostAddrInvalidType:
raise SCMPBadSrcType(
"Unsupported src address type: %s" % src_type) from None
pad_len = calc_padding(data_len, cls.BLK_SIZE)
total_len = data_len + pad_len
assert total_len % cls.BLK_SIZE == 0
return total_len, pad_len
def reverse(self):
self.dst, self.src = self.src, self.dst
self.update()
def dst_type(self): # pragma: no cover
return self.dst.host.TYPE
def src_type(self): # pragma: no cover
return self.src.host.TYPE
def __len__(self): # pragma: no cover
assert self._total_len is not None
return self._total_len
def __str__(self):
return "%s(%sB): Dst:<%s> Src:<%s>" % (self.NAME, len(self), self.dst, self.src)
class SCIONBasePacket(PacketBase):
"""
Encasulates the basic headers (common header, address header, and path
header). Everything else is stored as payload.
"""
NAME = "SCIONBasePacket"
MIN_LEN = SCIONCommonHdr.LEN
def __init__(self, raw=None): # pragma: no cover
self.cmn_hdr = None
self.addrs = None
self.path = None
self._l4_proto = L4Proto.NONE
self._payload = b""
super().__init__(raw)
def _parse(self, raw):
data = Raw(raw, self.NAME, self.MIN_LEN, min_=True)
self._inner_parse(data)
self.set_payload(PayloadRaw(data.get()))
def _inner_parse(self, data): # pragma: no cover
self.cmn_hdr = SCIONCommonHdr(data.pop(SCIONCommonHdr.LEN))
self._parse_addrs(data)
self._parse_path(data)
def _parse_addrs(self, data):
self.addrs = SCIONAddrHdr((
self.cmn_hdr.dst_addr_type,
self.cmn_hdr.src_addr_type,
data.get(self.cmn_hdr.addrs_len),
))
data.pop(len(self.addrs))
def _parse_path(self, data):
count = self.cmn_hdr.hdr_len_bytes() - data.offset()
if count < 0:
raise SCIONParseError(
"Bad header len field (%sB), implies negative path length" %
self.cmn_hdr.hdr_len_bytes(),
)
if count > len(data):
raise SCIONParseError(
"Bad header len field (%sB), "
"implies path is longer than packet (%sB)"
% (self.cmn_hdr.hdr_len_bytes(), len(data) + data.offset())
)
self.path = parse_path(data.get(count))
data.pop(len(self.path))
iof_idx, hof_idx = self.cmn_hdr.get_of_idxs()
self.path.set_of_idxs(iof_idx, hof_idx)
@classmethod
def from_values(cls, cmn_hdr, addr_hdr, path_hdr, payload=None):
inst = cls()
inst._inner_from_values(cmn_hdr, addr_hdr, path_hdr)
if payload is None:
payload = PayloadRaw()
inst.set_payload(payload)
inst.update()
return inst
def _inner_from_values(self, cmn_hdr, addr_hdr, path_hdr):
assert isinstance(cmn_hdr, SCIONCommonHdr), type(cmn_hdr)
self.cmn_hdr = cmn_hdr
assert isinstance(addr_hdr, SCIONAddrHdr), type(addr_hdr)
self.addrs = addr_hdr
assert isinstance(path_hdr, SCIONPath), type(path_hdr)
self.path = path_hdr
def get_fwd_ifid(self):
"""
Returns the next forwarding interface ID of the path or 0 if the path is
empty.
"""
if self.path:
return self.path.get_fwd_if()
return 0
def pack(self):
self.update()
packed = []
inner = self._inner_pack()
self.cmn_hdr.total_len = self.cmn_hdr.hdr_len_bytes() + len(inner)
packed.append(self.cmn_hdr.pack())
packed.append(self.addrs.pack())
packed.append(self.path.pack())
packed.append(inner)
raw = b"".join(packed)
assert len(raw) == self.cmn_hdr.total_len
return raw
def _inner_pack(self): # pragma: no cover
return b""
def _pack_payload(self): # pragma: no cover
return self._payload.pack()
def validate(self, pkt_len):
"""Called after parsing, to check for errors that don't break parsing"""
path_len = len(self.path)
self.cmn_hdr.validate(pkt_len, path_len)
self.addrs.validate()
if path_len:
self._validate_of_idxes()
assert isinstance(self._payload, PayloadRaw), type(self._payload)
def _validate_of_idxes(self):
try:
self.path.get_iof()
except SCIONIndexError as e:
raise SCMPBadIOFOffset("%s" % e) from None
try:
self.path.get_hof()
except SCIONIndexError as e:
raise SCMPBadHOFOffset("%s" % e) from None
def update(self):
self.addrs.update()
self._update_cmn_hdr()
def _update_cmn_hdr(self):
hdr = self.cmn_hdr
hdr.dst_addr_type = self.addrs.dst_type()
hdr.src_addr_type = self.addrs.src_type()
hdr.addrs_len = len(self.addrs)
hdr.hdr_len = hdr.bytes_to_hdr_len(len(hdr) + len(self.addrs) + len(self.path))
hdr.total_len = hdr.hdr_len_bytes() + self._get_offset_len()
hdr.set_of_idxs(*self.path.get_of_idxs())
hdr.next_hdr = self._get_next_hdr()
def _get_offset_len(self): # pragma: no cover
return 0
def _get_next_hdr(self): # pragma: no cover
return self._l4_proto
def reverse(self):
self.addrs.reverse()
self.path.reverse()
def reversed_copy(self): # pragma: no cover
inst = copy.deepcopy(self)
inst.reverse()
return inst
def convert_to_scmp_error(self, addr, class_, type_, pkt, *args,
hopbyhop=False, **kwargs):
self.addrs.src = addr
if self.ext_hdrs:
if self.ext_hdrs[0].EXT_TYPE == ExtHopByHopType.SCMP:
# Remove any existing SCMP ext header
del self.ext_hdrs[0]
# Insert SCMP ext at start of headers
self.ext_hdrs.insert(0, SCMPExt.from_values(hopbyhop=hopbyhop))
# Trim any extra headers, in the case of SCMPTooManyHopByHop, max+1 as
# the SCMP ext header isn't counted.
self.ext_hdrs = self.ext_hdrs[:MAX_HOPBYHOP_EXT + 1]
# Create SCMP payload.
pld = SCMPPayload.from_pkt(class_, type_, pkt, *args, **kwargs)
self.l4_hdr = SCMPHeader.from_values(self.addrs.src, self.addrs.dst,
class_, type_)
self.set_payload(pld)
def short_desc(self):
s = []
s.append("%s(%dB):" % (self.NAME, len(self)))
s.append(" %s" % self.cmn_hdr)
s.append(" %s" % self.addrs)
s.extend(self._inner_str())
return "\n".join(s)
def __len__(self): # pragma: no cover
return self.cmn_hdr.total_len
def __str__(self):
s = []
s.append("%s(%dB):" % (self.NAME, len(self)))
s.append(" %s" % self.cmn_hdr)
s.append(" %s" % self.addrs)
for line in str(self.path).splitlines():
s.append(" %s" % line)
s.extend(self._inner_str())
s.append(" Payload:")
for line in str(self._payload).splitlines():
s.append(" %s" % line)
return "\n".join(s)
def _inner_str(self): # pragma: no cover
return []
class SCIONExtPacket(SCIONBasePacket):
"""
Extends :any:`SCIONBasePacket` to handle extension headers.
"""
NAME = "SCIONExtPacket"
def __init__(self, raw=None): # pragma: no cover
self.ext_hdrs = []
self._unknown_exts = {}
super().__init__(raw)
def _inner_parse(self, data): # pragma: no cover
super()._inner_parse(data)
# Parse extension headers
self.ext_hdrs, self._l4_proto, self._unknown_exts = parse_extensions(
data, self.cmn_hdr.next_hdr)
@classmethod
def from_values(cls, cmn_hdr, addr_hdr, path_hdr, ext_hdrs, payload=b""):
inst = cls()
inst._inner_from_values(cmn_hdr, addr_hdr, path_hdr, ext_hdrs)
inst.set_payload(payload)
return inst
def _inner_from_values(self, cmn_hdr, addr_hdr, path_hdr, ext_hdrs):
super()._inner_from_values(cmn_hdr, addr_hdr, path_hdr)
for hdr in ext_hdrs:
assert isinstance(hdr, ExtensionHeader), type(hdr)
self.ext_hdrs.append(hdr)
def get_fwd_ifid(self):
"""
Returns the next forwarding interface ID depending on the extension
headers and the path in the packet.
"""
for hdr in self.ext_hdrs:
if_id = hdr.get_next_ifid()
if if_id is not None:
return if_id
return super().get_fwd_ifid()
def pack_exts(self):
packed = []
max_idx = len(self.ext_hdrs) - 1
for i, hdr in enumerate(self.ext_hdrs):
ext_packed = []
next_hdr = self._l4_proto
if i < max_idx:
next_hdr = self.ext_hdrs[i+1].EXT_CLASS
ext_packed.append(struct.pack("!BBB", next_hdr, hdr.hdr_len(),
hdr.EXT_TYPE))
ext_packed.append(hdr.pack())
ext = b"".join(ext_packed)
assert len(ext) % ExtensionHeader.LINE_LEN == 0
packed.append(ext)
return b"".join(packed)
def _inner_pack(self):
return super()._inner_pack() + self.pack_exts()
def _get_offset_len(self):
l = super()._get_offset_len()
for hdr in self.ext_hdrs:
l += len(hdr)
return l
def _get_next_hdr(self):
if self.ext_hdrs:
return self.ext_hdrs[0].EXT_CLASS
else:
return self._l4_proto
def validate(self, pkt_len):
super().validate(pkt_len)
if not self._unknown_exts:
return True
# Use the first unknown extension, and use that for the SCMP error
# message.
hbh = self._unknown_exts.get(ExtensionClass.HOP_BY_HOP)
if hbh:
raise SCMPBadHopByHop(hbh[0])
e2e = self._unknown_exts.get(ExtensionClass.END_TO_END)
if e2e:
raise SCMPBadEnd2End(e2e[0])
def _inner_str(self): # pragma: no cover
s = super()._inner_str()
for hdr in self.ext_hdrs:
for line in str(hdr).splitlines():
s.append(" %s" % line)
return s
def reverse(self): # pragma: no cover
for hdr in self.ext_hdrs:
hdr.reverse()
super().reverse()
class SCIONL4Packet(SCIONExtPacket):
"""
Extends :any:`SCIONExtPacket` to handle L4 headers.
"""
NAME = "SCIONL4Packet"
def __init__(self, raw=None): # pragma: no cover
self.l4_hdr = None
super().__init__(raw)
def _inner_parse(self, data):
super()._inner_parse(data)
# Parse L4 header
self.l4_hdr = parse_l4_hdr(
self._l4_proto, data, dst=self.addrs.dst, src=self.addrs.src)
@classmethod
def from_values(cls, cmn_hdr, addr_hdr, path_hdr, ext_hdrs, l4_hdr,
payload=None):
inst = cls()
inst._inner_from_values(cmn_hdr, addr_hdr, path_hdr, ext_hdrs, l4_hdr)
if payload is None:
payload = PayloadRaw()
inst.set_payload(payload)
inst.update()
return inst
def _inner_from_values(self, cmn_hdr, addr_hdr, path_hdr, ext_hdrs, l4_hdr):
super()._inner_from_values(cmn_hdr, addr_hdr, path_hdr, ext_hdrs)
assert isinstance(l4_hdr, L4HeaderBase), type(l4_hdr)
self.l4_hdr = l4_hdr
self._l4_proto = l4_hdr.TYPE
def _inner_pack(self):
self.update()
packed = [super()._inner_pack()]
pld = super()._pack_payload()
if self.l4_hdr:
packed.append(self.l4_hdr.pack(pld))
packed.append(pld)
return b"".join(packed)
def _pack_payload(self): # pragma: no cover
# Payload is already packed and included as part of _inner_pack
return b""
def validate(self, pkt_len): # pragma: no cover
super().validate(pkt_len)
if self.l4_hdr:
self.l4_hdr.validate(self._payload.pack())
def update(self):
if self.l4_hdr:
self.l4_hdr.update(src=self.addrs.src, dst=self.addrs.dst)
self._l4_proto = self.l4_hdr.TYPE
super().update()
def reverse(self): # pragma: no cover
if self.l4_hdr:
self.l4_hdr.reverse()
super().reverse()
def parse_payload(self):
if not self.l4_hdr:
raise SCIONParseError("Cannot parse payload of non-L4 packet")
praw = self._payload.pack()
if self.l4_hdr.TYPE == L4Proto.UDP:
# Treat as SCION control message
pld = SignedCtrlPayload.from_raw(praw).pld()
elif self.l4_hdr.TYPE == L4Proto.SCMP:
pld = SCMPPayload((self.l4_hdr.class_, self.l4_hdr.type, praw))
self.set_payload(pld)
return pld
def _get_offset_len(self):
l = super()._get_offset_len()
if self.l4_hdr:
l += self.l4_hdr.total_len
return l
def _inner_str(self): # pragma: no cover
s = super()._inner_str()
s.append(" %s" % self.l4_hdr)
return s
def get_l4_proto(self): # pragma: no cover
return self._l4_proto
def build_base_hdrs(dst, src, l4=L4Proto.UDP):
cmn_hdr = SCIONCommonHdr.from_values(dst.host.TYPE, src.host.TYPE, l4)
addr_hdr = SCIONAddrHdr.from_values(dst, src)
return cmn_hdr, addr_hdr
|
import click, os, sys, tempfile
from sqlalchemy.orm import relationship
from lah.db import LahDb
from lah.models import *
from lah.haplotig_iters import HaplotigIterator
@click.command(short_help="generate haplotig seqfile")
@click.argument("hid", type=click.STRING)
@click.option("--output", required=False, type=click.STRING, help="Send output to this file instead of deafult location")
def haplotig_seqfile_cmd(hid, output):
"""
Generate Haplotig Seqfile
Fetch the reads from the known seqfiles amd write a fastq for a given haplotig id. Default output file loaction is in the "haplotigs" sub directory. Optionally, save the reads to a different file.
"""
print("Generate haplotig seqfile ...")
print("Haplotig ID: {}".format(hid))
session = LahDb.session()
haplotig = session.query(Haplotig).get(hid)
if not haplotig:
raise Exception("Failed to get haplotig {} from db!".format(hid))
dn = session.query(Metadata).filter_by(name="directory").one().value
haplotigs_bn = session.query(Metadata).filter_by(name="haplotigs_fn").one().value
haplotigs_headers = session.query(Metadata).filter_by(name="haplotig_headers").one().value
headers = haplotigs_headers.split(",")
h_i = HaplotigIterator(in_fn=os.path.join(dn, haplotigs_bn), headers=headers, pos=haplotig.file_pos)
h_i.load_haplotig_reads(haplotig)
source_seqfiles = session.query(Seqfile).all()
if not len(source_seqfiles):
raise Exception("No seqfiles fround in database!")
if output is None:
output = haplotig.seqfile_fn(dn)
print("Output: {}".format(output))
create_seqfile(haplotig, sources=source_seqfiles, output=output)
print("Generate haplotig seqfile ... SUCCESS")
#-- haplotig_seqfile_cmd
def create_seqfile(haplotig, sources, output):
if not hasattr(haplotig, "reads") or len(haplotig.reads) == 0:
raise Exception("No reads loaded for haplotig!")
if not len(sources):
raise Exception("No source seqfiles given!")
if os.path.exists(output):
os.remove(output)
rds = haplotig.reads
temp_f = tempfile.NamedTemporaryFile(mode="a")
output_f = open(output, "w")
for seqfile in sources:
if len(rds) == 0:
print("Found allreads, skipping remaining seqfiles.")
break
print("Seqfile: {}".format(seqfile.fn))
print("Reads remaining: {}".format(len(rds)))
idx_fn = seqfile.idx_fn()
with open(seqfile.fn, "r") as seqfile_f, open(idx_fn, "r") as idx_f:
for l in idx_f.readlines():
rd_fai = l.rstrip().split("\t")
if rd_fai[0] in rds:
seqfile_f.seek( int(rd_fai[2]) )
output_f.write("@" + rd_fai[0] + "\n")
output_f.write( seqfile_f.read(int(rd_fai[1])) + "\n" )
seqfile_f.seek( int(rd_fai[5]) )
output_f.write("+\n")
output_f.write( seqfile_f.read(int(rd_fai[1])) + "\n" )
rds.remove(rd_fai[0])
output_f.close()
if len(rds) != 0:
raise Exception("Failed to find haplotig {} {} reads: {}".format(haplotig.id, haplotig.name, " ".join(rds)))
#-- create_seqfile
|
<filename>awsbw/awsbw.py<gh_stars>0
#!/usr/bin/env python3
import boto3
import curses
from curses import wrapper
from curses import panel
import sys
import argparse
import time
from datetime import datetime
class AWSBW():
def __init__(self, stdscr, jobQueues):
self.__currentJobs__ = []
self.__max_age_days__ = 7
# screen stuff
try:
curses.curs_set(0)
except:
pass
self.__stdscr__ = stdscr
self.__stdscr__.nodelay(True)
(curH, curW) = stdscr.getmaxyx()
self.__stdscr__.clear()
# add a window for the job listing
self.__jobsWin__ = curses.newwin(
curH - 2,
curW - 2,
1, 1
)
self.__termHeight__ = None
self.__termWidth__ = None
# Job stuff
self.__jobStatuses__ = [
'RUNNING',
'RUNNABLE',
'SUCCEEDED',
'FAILED',
'STARTING',
]
self.__jobQueues__ = jobQueues
self.__curJobQueue__ = jobQueues[0]
self.__curJobId__ = None
self.getJobs()
# Now display!
self.screenRefresh()
def screenRefresh(self, forceRedraw=False):
(curH, curW) = self.__stdscr__.getmaxyx()
if self.__termHeight__ != curH or self.__termWidth__ != curW:
curses.resizeterm(curH, curW)
self.__termHeight__ = curH
self.__termWidth__ = curW
self.__jobsWin__.resize(
curH - 2,
curW - 2,
)
self.__stdscr__.border()
self.__stdscr__.refresh()
self.showJobs()
elif forceRedraw:
self.__stdscr__.border()
self.__stdscr__.refresh()
self.showJobs()
# Header: Use it to show the queues including which is current.
x = 1
for q in self.__jobQueues__:
if x + len(q) > curW:
break
if q == self.__curJobQueue__:
self.__stdscr__.addstr(
0, x,
q,
curses.A_UNDERLINE
)
x += len(q) + 1
else:
self.__stdscr__.addstr(
0, x,
q,
)
x += len(q) + 1
if x + 20 < curW:
# If we have space, add the timestamp of the last check
self.__stdscr__.addstr(
0, curW - 20,
datetime.fromtimestamp(
self.__lastJobCheck__).strftime('%Y-%m-%d %H:%M:%S')
)
# Footer
if curW > 71:
self.__stdscr__.addstr(
curH - 1,
max(
1,
int(curW / 2) - 34
),
" < > queues. D details. L logs. T terminate. Q quit. "
)
self.__stdscr__.refresh()
def showJobs(self, moveKey=None):
win = self.__jobsWin__
# Limit to the current queue and recency:
cutoff_ts = (time.time() - self.__max_age_days__ * 24 * 3600) * 1000
jobs = [
j for j in
self.__currentJobs__
if j['queue'] == self.__curJobQueue__ and j['createdAt'] >= cutoff_ts
]
(winH, winW) = win.getmaxyx()
if len(jobs) == 0:
win.addnstr(
1,
0,
"No Jobs",
winW
)
return
statuses = [s for s in self.__jobStatuses__ if s in {j['status'] for j in jobs}]
col_width = max([
max(len(s) + 1 for s in statuses),
max(len(j['jobName']) + 1 for j in jobs),
])
maxJobs = winH - 2
maxCols = int((winW - 2) / col_width)
if self.__curJobId__ not in [j['jobId'] for j in jobs if j['status'] in statuses[0:maxCols]]:
selected_status_i = 0
selected_job_i = 0
else:
# Job ID is in our list, search for it
curJob = [j for j in jobs if j['jobId'] == self.__curJobId__][0]
selected_status_i = statuses.index(curJob['status'])
selected_job_i = [j['jobId'] for j in jobs if j['status'] == curJob['status']].index(curJob['jobId'])
# Do a bit of screen geometry sanity here
if selected_status_i > maxCols:
selected_status_i = 0
selected_job_i = 0
if selected_job_i > maxJobs:
selected_status_i = 0
selected_job_i = 0
if moveKey is not None:
if moveKey == curses.KEY_UP:
selected_job_i = max([
0,
selected_job_i - 1
])
elif moveKey == curses.KEY_DOWN:
selected_job_i = min([
selected_job_i + 1,
maxJobs,
len([j for j in jobs if j['status'] == statuses[selected_status_i]]) - 1
])
elif moveKey == curses.KEY_RIGHT:
selected_status_i = min(
len(statuses) - 1,
maxCols,
selected_status_i + 1
)
selected_job_i = min([
selected_job_i,
len([j for j in jobs if j['status'] == statuses[selected_status_i]]) - 1
])
elif moveKey == curses.KEY_LEFT:
selected_status_i = max(
0,
selected_status_i - 1
)
selected_job_i = min([
selected_job_i,
len([j for j in jobs if j['status'] == statuses[selected_status_i]]) - 1
])
win.addnstr(
0,
0,
"".join([s.ljust(col_width) for s in statuses[:maxCols]]).ljust(winW),
winW,
curses.A_UNDERLINE
)
for status_i, status in enumerate(statuses):
if status_i >= maxCols:
break
status_jobs = [j for j in jobs if j['status'] == status]
for job_i, job in enumerate(status_jobs):
if job_i > maxJobs:
break
if (job_i == selected_job_i) and (status_i == selected_status_i):
self.__curJobId__ = job['jobId']
win.addnstr(
job_i + 1,
col_width * status_i,
job['jobName'].ljust(col_width),
winW,
curses.A_REVERSE
)
else:
win.addnstr(
job_i + 1,
col_width * status_i,
job['jobName'].ljust(col_width),
winW
)
# Clearing out the remainder of the column
for y in range(job_i + 2, winH):
win.addnstr(
y,
col_width * status_i,
"".ljust(col_width),
winW
)
# Clearing the right column
right_pad = winW - col_width * len(statuses[:maxCols]) - 1
if right_pad > 0:
for y in range(1, winH):
win.addnstr(
y,
col_width * len(statuses[:maxCols]),
"".ljust(right_pad),
winW
)
win.refresh()
def queueJobs(self, queue, status='RUNNING'):
batch_client = boto3.client('batch')
jobs_running = batch_client.list_jobs(
jobQueue=queue,
jobStatus=status,
)
try:
for j in jobs_running['jobSummaryList']:
j.update({'queue': queue})
return jobs_running['jobSummaryList']
except:
return []
def jobDetails(self, jobId):
batch_client = boto3.client('batch')
try:
job_info = batch_client.describe_jobs(
jobs=[
jobId,
]
)['jobs'][0]
except:
job_info = None
return job_info
def terminateJob(self, jobId):
batch_client = boto3.client('batch')
batch_client.terminate_job(
jobId=jobId,
reason='Terminated by user'
)
def terminateJobDialog(self):
try:
job = [j for j in self.__currentJobs__ if j['jobId'] == self.__curJobId__][0]
except:
return
p = panel.new_panel(self.__stdscr__)
p.top()
p.show()
p_win = p.window()
p_win.clear()
p_win.border()
p_win.nodelay(False)
winH, winW = p_win.getmaxyx()
question_str = "To terminate job {} type Y".format(job['jobName'])
p_win.addnstr(
int(winH / 2) - 1,
max(
1,
int(winW / 2) - int(len(question_str) / 2),
),
question_str,
winW - 2,
)
p_win.refresh()
c = p_win.getch()
if c == 121 or c == 89:
self.terminateJob(job['jobId'])
p_win.addnstr(
int(winH / 2) + 2,
max(
1,
int(winW / 2) - 5,
),
"Terminated",
winW - 2,
curses.A_REVERSE
)
p_win.refresh()
time.sleep(1)
p_win.nodelay(True)
p.hide()
self.screenRefresh(forceRedraw=True)
def getJobs(self):
self.__currentJobs__ = []
for queue in self.__jobQueues__:
queue_jobs = []
for status in self.__jobStatuses__:
queue_jobs += self.queueJobs(
queue,
status=status
)
for j in queue_jobs:
j['queue'] = queue
self.__currentJobs__ += sorted(
queue_jobs,
key=lambda j: -j['createdAt']
)
self.__lastJobCheck__ = time.time()
def refreshJobs(self, MIN_DELAY=30):
if time.time() - self.__lastJobCheck__ >= MIN_DELAY:
self.getJobs()
self.showJobs()
return True
else:
return False
def queueRight(self):
prior_queue = self.__curJobQueue__
self.__curJobQueue__ = self.__jobQueues__[
min(
self.__jobQueues__.index(self.__curJobQueue__) + 1,
len(self.__jobQueues__) - 1
)]
if prior_queue != self.__curJobQueue__:
self.showJobs()
def queueLeft(self):
prior_queue = self.__curJobQueue__
self.__curJobQueue__ = self.__jobQueues__[
max(
self.__jobQueues__.index(self.__curJobQueue__) - 1,
0
)]
if prior_queue != self.__curJobQueue__:
self.showJobs()
def displayList(self, L, win, Hoffset, Hmax, Woffset, Wmax):
L_i = 0
for line in L:
if Hmax <= (L_i + Hoffset):
break
line_chunks = [
line[i:i + int(Wmax)]
for i in range(
0,
len(line),
Wmax
)
]
for line_chunk in line_chunks:
if Hmax <= (L_i + Hoffset):
break
win.addstr(
L_i + Hoffset,
Woffset,
line_chunk.ljust(Wmax)
)
L_i += 1
win.refresh()
def detail_panel(self):
try:
job = [j for j in self.__currentJobs__ if j['jobId'] == self.__curJobId__][0]
except:
return
dp = panel.new_panel(self.__stdscr__)
dp.top()
dp.show()
dp_win = dp.window()
dp_win.clear()
dp_win.border()
dp_win.refresh()
winH, winW = dp_win.getmaxyx()
if winH < 5:
dp.hide()
self.__stdscr__.border()
return
dp_win.addstr(
winH - 1,
int(winW / 2) - 3,
"ESC to close"
)
# Title!
dp_win.addnstr(
1,
1,
"{} (id: {}) on {}".format(
job['jobName'],
job['jobId'],
job['queue']
),
winW - 2,
)
# Timing
timingStr = "Created: {}.".format(
datetime.fromtimestamp(job['createdAt'] / 1000).strftime('%Y-%m-%d %H:%M:%S')
)
if 'startedAt' in job:
timingStr += "\t Started: {}.".format(
datetime.fromtimestamp(
job['startedAt'] / 1000).strftime('%Y-%m-%d %H:%M:%S')
)
if 'stoppedAt' in job:
timingStr += "\t Stopped: {}.".format(
datetime.fromtimestamp(
job['stoppedAt'] / 1000).strftime('%Y-%m-%d %H:%M:%S')
)
dp_win.addnstr(
2,
1,
timingStr,
winW - 2,
)
jobDetails = self.jobDetails(job['jobId'])
if jobDetails is not None:
dp_win.addnstr(
3,
1,
"Job Description: {}".format(
jobDetails['jobDefinition'].split('/')[-1]
),
winW - 2
)
dp_win.addnstr(
4,
1,
"Image: {}".format(
jobDetails['container']['image'],
),
winW - 2
)
dp_win.addnstr(
5,
1,
"\tvcpu: {}\t\tmem: {:,} MB.".format(
jobDetails['container'].get('vcpus'),
jobDetails['container'].get('memory')
),
winW - 2
)
cmd_start = 0
commands = jobDetails['container']['command']
else:
cmd_start = 0
commands = []
self.displayList(
commands[cmd_start:],
win=dp_win,
Hoffset=7,
Hmax=winH - 7,
Woffset=1,
Wmax=winW - 2
)
# Detail window loop!
while True:
c = self.__stdscr__.getch()
if c == 27:
dp_win.clear()
dp.hide()
self.screenRefresh(forceRedraw=True)
break
elif c == curses.KEY_DOWN:
if cmd_start < len(commands):
cmd_start += 1
self.displayList(
commands[cmd_start:],
win=dp_win,
Hoffset=7,
Hmax=winH - 7,
Woffset=1,
Wmax=winW - 2
)
elif c == curses.KEY_UP:
if cmd_start > 0:
cmd_start -= 1
self.displayList(
commands[cmd_start:],
win=dp_win,
Hoffset=7,
Hmax=winH - 7,
Woffset=1,
Wmax=winW - 2
)
def getLog(self, jobStreamName, startFromHead=False):
logs_client = boto3.client('logs')
try:
jobLog = logs_client.get_log_events(
logGroupName='/aws/batch/job',
logStreamName=jobStreamName,
startFromHead=startFromHead,
)
if startFromHead:
events = sorted(
jobLog['events'],
key=lambda e: e['timestamp']
)
else:
events = sorted(
jobLog['events'],
key=lambda e: -e['timestamp']
)
except:
events = []
return events
def log_panel(self):
try:
job = [j for j in self.__currentJobs__ if j['jobId'] == self.__curJobId__][0]
except:
return
lp = panel.new_panel(self.__stdscr__)
lp.top()
lp.show()
lp_win = lp.window()
winH, winW = lp_win.getmaxyx()
if winH < 5:
lp.hide()
self.__stdscr__.border()
return
lp_win.clear()
lp_win.border()
lp_win.addstr(
winH - 1,
int(winW / 2) - 3,
"ESC to close"
)
# Title!
lp_win.addnstr(
1,
1,
"Logs for {} (id: {}) on {}".format(
job['jobName'],
job['jobId'],
job['queue']
),
winW - 2,
)
lp_win.addnstr(
3,
1,
"Loading logs......".ljust(winW - 2),
winW - 2,
)
lp_win.refresh()
jobDetails = self.jobDetails(job['jobId'])
if jobDetails is None:
return
try:
jobStreamName = jobDetails['container']['logStreamName']
except:
return
# Get the log
startFromHead = True
events = self.getLog(jobStreamName, startFromHead)
event_first = 0
self.displayList(
[
e['message'] for e
in events[event_first:]
],
win=lp_win,
Hoffset=3,
Hmax=winH - 2,
Woffset=1,
Wmax=winW - 2,
)
# Log window loop!
while True:
c = self.__stdscr__.getch()
if c == 27: # esc
lp_win.clear()
lp.hide()
self.screenRefresh(forceRedraw=True)
break
elif c == 79 or c == 111: # O or o
lp_win.addnstr(
3,
1,
"Loading reversed logs ......".ljust(winW - 2),
winW - 2,
)
lp_win.refresh()
startFromHead = not startFromHead
events = self.getLog(jobStreamName, startFromHead)
event_first = 0
self.displayList(
[
e['message'] for e
in events[event_first:]
],
win=lp_win,
Hoffset=3,
Hmax=winH - 2,
Woffset=1,
Wmax=winW - 2,
)
elif c == curses.KEY_NPAGE or c == 32: # or space
if (event_first + winH - 2) < len(events):
event_first += (winH - 2)
self.displayList(
[
e['message'] for e
in events[event_first:]
],
win=lp_win,
Hoffset=3,
Hmax=winH - 2,
Woffset=1,
Wmax=winW - 2,
)
elif c == curses.KEY_DOWN:
if event_first < len(events):
event_first += 1
self.displayList(
[
e['message'] for e
in events[event_first:]
],
win=lp_win,
Hoffset=3,
Hmax=winH - 2,
Woffset=1,
Wmax=winW - 2,
)
elif c == curses.KEY_UP:
if event_first > 0:
event_first -= 1
self.displayList(
[
e['message'] for e
in events[event_first:]
],
win=lp_win,
Hoffset=3,
Hmax=winH - 2,
Woffset=1,
Wmax=winW - 2,
)
def handleInput(self, c):
if c == curses.KEY_UP or c == curses.KEY_DOWN:
self.showJobs(c)
if c == curses.KEY_LEFT or c == curses.KEY_RIGHT:
self.showJobs(c)
if c == 62 or c == 46:
self.queueRight()
if c == 60 or c == 44:
self.queueLeft()
if c == 68 or c == 100:
self.detail_panel()
if c == 108 or c == 76:
self.log_panel()
if c == 84 or c == 116:
self.terminateJobDialog()
def actionLoop(self):
while True:
c = self.__stdscr__.getch()
if c == 113 or c == 81:
break
self.handleInput(c)
self.refreshJobs()
self.screenRefresh()
def start(stdscr, args):
awsbw = AWSBW(
stdscr,
args.queue,
)
awsbw.actionLoop()
def main():
parser = argparse.ArgumentParser(description="AWS Batch Watcher")
parser.add_argument(
'-Q', '--queue',
required=True,
help='AWS batch queue(s) to monitor',
nargs='+'
)
args = parser.parse_args()
wrapper(start, args)
if __name__ == '__main__':
main()
|
#!/usr/bin/env python
#
# Copyright 2007 Google Inc.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
"""Datastore backed Blobstore API stub.
Class:
BlobstoreServiceStub: BlobstoreService stub backed by datastore.
"""
import base64
import os
import time
import urlparse
from google.appengine.api import apiproxy_stub
from google.appengine.api import blobstore
from google.appengine.api import datastore
from google.appengine.api import datastore_errors
from google.appengine.api import datastore_types
from google.appengine.api import users
from google.appengine.api.blobstore import blobstore_service_pb
from google.appengine.runtime import apiproxy_errors
__all__ = ['BlobStorage',
'BlobstoreServiceStub',
'ConfigurationError',
'CreateUploadSession',
'Error',
]
class Error(Exception):
"""Base blobstore error type."""
class ConfigurationError(Error):
"""Raised when environment is not correctly configured."""
_UPLOAD_SESSION_KIND = '__BlobUploadSession__'
_GS_INFO_KIND = '__GsFileInfo__'
def CreateUploadSession(creation,
success_path,
user,
max_bytes_per_blob,
max_bytes_total,
bucket_name=None):
"""Create upload session in datastore.
Creates an upload session and puts it in Datastore to be referenced by
upload handler later.
Args:
creation: Creation timestamp.
success_path: Path in users application to call upon success.
user: User that initiated this upload, if any.
max_bytes_per_blob: Maximum number of bytes for any blob in the upload.
max_bytes_total: Maximum aggregate bytes for all blobs in the upload.
bucket_name: Name of the Google Storage bucket tio upload the files.
Returns:
String encoded key of new Datastore entity.
"""
entity = datastore.Entity(_UPLOAD_SESSION_KIND, namespace='')
entity_dict = {'creation': creation,
'success_path': success_path,
'user': user,
'state': 'init',
'max_bytes_per_blob': max_bytes_per_blob,
'max_bytes_total': max_bytes_total}
if bucket_name:
entity_dict['gs_bucket_name'] = bucket_name
entity.update(entity_dict)
datastore.Put(entity)
return str(entity.key())
class BlobStorage(object):
"""Base class for defining how blobs are stored.
This base class merely defines an interface that all stub blob-storage
mechanisms must implement.
"""
def StoreBlob(self, blob_key, blob_stream):
"""Store blob stream.
Implement this method to persist blob data.
Args:
blob_key: Blob key of blob to store.
blob_stream: Stream or stream-like object that will generate blob content.
"""
raise NotImplementedError('Storage class must override StoreBlob method.')
def OpenBlob(self, blob_key):
"""Open blob for streaming.
Args:
blob_key: Blob-key of existing blob to open for reading.
Returns:
Open file stream for reading blob. Caller is responsible for closing
file.
"""
raise NotImplementedError('Storage class must override OpenBlob method.')
def DeleteBlob(self, blob_key):
"""Delete blob data from storage.
Args:
blob_key: Blob-key of existing blob to delete.
"""
raise NotImplementedError('Storage class must override DeleteBlob method.')
class BlobstoreServiceStub(apiproxy_stub.APIProxyStub):
"""Datastore backed Blobstore service stub.
This stub stores manages upload sessions in the Datastore and must be
provided with a blob_storage object to know where the actual blob
records can be found after having been uploaded.
This stub does not handle the actual creation of blobs, neither the BlobInfo
in the Datastore nor creation of blob data in the blob_storage. It does,
however, assume that another part of the system has created these and
uses these objects for deletion.
An upload session is created when the CreateUploadURL request is handled and
put in the Datastore under the __BlobUploadSession__ kind. There is no
analog for this kind on a production server. Other than creation, this stub
not work with session objects. The URLs created by this service stub are:
http://<appserver-host>:<appserver-port>/<uploader-path>/<session-info>
This is very similar to what the URL is on a production server. The session
info is the string encoded version of the session entity
"""
_ACCEPTS_REQUEST_ID = True
GS_BLOBKEY_PREFIX = 'encoded_gs_file:'
def __init__(self,
blob_storage,
time_function=time.time,
service_name='blobstore',
uploader_path='_ah/upload/',
request_data=None):
"""Constructor.
Args:
blob_storage: BlobStorage class instance used for blob storage.
time_function: Used for dependency injection in tests.
service_name: Service name expected for all calls.
uploader_path: Path to upload handler pointed to by URLs generated
by this service stub.
request_data: A apiproxy_stub.RequestData instance used to look up state
associated with the request that generated an API call.
"""
super(BlobstoreServiceStub, self).__init__(service_name,
request_data=request_data)
self.__storage = blob_storage
self.__time_function = time_function
self.__next_session_id = 1
self.__uploader_path = uploader_path
@classmethod
def ToDatastoreBlobKey(cls, blobkey):
"""Given a string blobkey, return its db.Key."""
kind = blobstore.BLOB_INFO_KIND
if blobkey.startswith(cls.GS_BLOBKEY_PREFIX):
kind = _GS_INFO_KIND
return datastore_types.Key.from_path(kind,
blobkey,
namespace='')
@property
def storage(self):
"""Access BlobStorage used by service stub.
Returns:
BlobStorage instance used by blobstore service stub.
"""
return self.__storage
def _GetEnviron(self, name):
"""Helper method ensures environment configured as expected.
Args:
name: Name of environment variable to get.
Returns:
Environment variable associated with name.
Raises:
ConfigurationError if required environment variable is not found.
"""
try:
return os.environ[name]
except KeyError:
raise ConfigurationError('%s is not set in environment.' % name)
def _CreateSession(self,
success_path,
user,
max_bytes_per_blob=None,
max_bytes_total=None,
bucket_name=None):
"""Create new upload session.
Args:
success_path: Application path to call upon successful POST.
user: User that initiated the upload session.
max_bytes_per_blob: Maximum number of bytes for any blob in the upload.
max_bytes_total: Maximum aggregate bytes for all blobs in the upload.
bucket_name: The name of the Cloud Storage bucket where the files will be
uploaded.
Returns:
String encoded key of a new upload session created in the datastore.
"""
return CreateUploadSession(self.__time_function(),
success_path,
user,
max_bytes_per_blob,
max_bytes_total,
bucket_name)
def _Dynamic_CreateUploadURL(self, request, response, request_id):
"""Create upload URL implementation.
Create a new upload session. The upload session key is encoded in the
resulting POST URL. This URL is embedded in a POST form by the application
which contacts the uploader when the user posts.
Args:
request: A fully initialized CreateUploadURLRequest instance.
response: A CreateUploadURLResponse instance.
request_id: A unique string identifying the request associated with the
API call.
"""
max_bytes_per_blob = None
max_bytes_total = None
bucket_name = None
if request.has_max_upload_size_per_blob_bytes():
max_bytes_per_blob = request.max_upload_size_per_blob_bytes()
if request.has_max_upload_size_bytes():
max_bytes_total = request.max_upload_size_bytes()
if request.has_gs_bucket_name():
bucket_name = request.gs_bucket_name()
session = self._CreateSession(request.success_path(),
users.get_current_user(),
max_bytes_per_blob,
max_bytes_total,
bucket_name)
# AppScale: Keep scheme for upload URL consistent with current context.
scheme = self.request_data.get_scheme(request_id)
protocol, host, _, _, _, _ = urlparse.urlparse(
self.request_data.get_request_url(request_id, scheme=scheme))
# End AppScale.
response.set_url('%s://%s/%s%s' % (protocol, host, self.__uploader_path,
session))
@classmethod
def DeleteBlob(cls, blobkey, storage):
"""Delete a blob.
Args:
blobkey: blobkey in str.
storage: blobstore storage stub.
"""
datastore.Delete(cls.ToDatastoreBlobKey(blobkey))
blobinfo = datastore_types.Key.from_path(blobstore.BLOB_INFO_KIND,
blobkey,
namespace='')
datastore.Delete(blobinfo)
storage.DeleteBlob(blobkey)
def _Dynamic_DeleteBlob(self, request, response, unused_request_id):
"""Delete a blob by its blob-key.
Delete a blob from the blobstore using its blob-key. Deleting blobs that
do not exist is a no-op.
Args:
request: A fully initialized DeleteBlobRequest instance.
response: Not used but should be a VoidProto.
"""
for blobkey in request.blob_key_list():
self.DeleteBlob(blobkey, self.__storage)
def _Dynamic_FetchData(self, request, response, unused_request_id):
"""Fetch a blob fragment from a blob by its blob-key.
Fetches a blob fragment using its blob-key. Start index is inclusive,
end index is inclusive. Valid requests for information outside of
the range of the blob return a partial string or empty string if entirely
out of range.
Args:
request: A fully initialized FetchDataRequest instance.
response: A FetchDataResponse instance.
Raises:
ApplicationError when application has the following errors:
INDEX_OUT_OF_RANGE: Index is negative or end > start.
BLOB_FETCH_SIZE_TOO_LARGE: Request blob fragment is larger than
MAX_BLOB_FRAGMENT_SIZE.
BLOB_NOT_FOUND: If invalid blob-key is provided or is not found.
"""
start_index = request.start_index()
if start_index < 0:
raise apiproxy_errors.ApplicationError(
blobstore_service_pb.BlobstoreServiceError.DATA_INDEX_OUT_OF_RANGE)
end_index = request.end_index()
if end_index < start_index:
raise apiproxy_errors.ApplicationError(
blobstore_service_pb.BlobstoreServiceError.DATA_INDEX_OUT_OF_RANGE)
fetch_size = end_index - start_index + 1
if fetch_size > blobstore.MAX_BLOB_FETCH_SIZE:
raise apiproxy_errors.ApplicationError(
blobstore_service_pb.BlobstoreServiceError.BLOB_FETCH_SIZE_TOO_LARGE)
blobkey = request.blob_key()
info_key = self.ToDatastoreBlobKey(blobkey)
try:
datastore.Get(info_key)
except datastore_errors.EntityNotFoundError:
raise apiproxy_errors.ApplicationError(
blobstore_service_pb.BlobstoreServiceError.BLOB_NOT_FOUND)
blob_file = self.__storage.OpenBlob(blobkey)
blob_file.seek(start_index)
response.set_data(blob_file.read(fetch_size))
def _Dynamic_DecodeBlobKey(self, request, response, unused_request_id):
"""Decode a given blob key: data is simply base64-decoded.
Args:
request: A fully-initialized DecodeBlobKeyRequest instance
response: A DecodeBlobKeyResponse instance.
"""
for blob_key in request.blob_key_list():
response.add_decoded(blob_key.decode('base64'))
@classmethod
def CreateEncodedGoogleStorageKey(cls, filename):
"""Create an encoded blob key that represents a Google Storage file.
For now we'll just base64 encode the Google Storage filename, APIs that
accept encoded blob keys will need to be able to support Google Storage
files or blobstore files based on decoding this key.
Any stub that creates GS files should use this function to convert
a gs filename to a blobkey. The created blobkey should be used both
as its _GS_FILE_INFO entity's key name and as the storage key to
store its content in blobstore. This ensures the GS files created
can be operated by other APIs.
Note this encoding is easily reversible and is not encryption.
Args:
filename: gs filename of form 'bucket/filename'
Returns:
blobkey string of encoded filename.
"""
return cls.GS_BLOBKEY_PREFIX + base64.urlsafe_b64encode(filename)
def _Dynamic_CreateEncodedGoogleStorageKey(self, request, response,
unused_request_id):
"""Create an encoded blob key that represents a Google Storage file.
For now we'll just base64 encode the Google Storage filename, APIs that
accept encoded blob keys will need to be able to support Google Storage
files or blobstore files based on decoding this key.
Args:
request: A fully-initialized CreateEncodedGoogleStorageKeyRequest
instance.
response: A CreateEncodedGoogleStorageKeyResponse instance.
"""
filename = request.filename()[len(blobstore.GS_PREFIX):]
response.set_blob_key(
self.CreateEncodedGoogleStorageKey(filename))
def CreateBlob(self, blob_key, content):
"""Create new blob and put in storage and Datastore.
This is useful in testing where you have access to the stub.
Args:
blob_key: String blob-key of new blob.
content: Content of new blob as a string.
Returns:
New Datastore entity without blob meta-data fields.
"""
entity = datastore.Entity(blobstore.BLOB_INFO_KIND,
name=blob_key, namespace='')
entity['size'] = len(content)
datastore.Put(entity)
self.storage.CreateBlob(blob_key, content)
return entity
|
<gh_stars>0
import copy
import re
class Player:
"""
A class to represent the player.
- Constructor Parameters
:param token: :type str:
- The player's token in the board.
:param name: :type str:
- The player's name.
"""
def __init__(self, token, name, *args, **kwargs):
self.token = token
self.name = name
class Slot:
"""
A slot will contain a space from the field.
- Constructor Parameters
:param token: :type str:
- The initial token for the slot.
"""
def __init__(self, token, *args, **kwargs):
self.token = token
def get_token(self):
"""
Returns the token if available, else, a single space.
"""
return self.token or ' '
def update_token(self, token):
"""
Updates the token in the slot if the slot is empty. Returns a boolean
value stating if the update was a success or not.
- Parameters:
:param token: :type str:
- The new token.
"""
empty = self.is_empty()
if empty:
self.token = token
return empty
return empty
def is_empty(self):
"""
Returns False if the token is empty or only contains spaces.
"""
return [False, True][re.search('^\s*$', self.token) is not None]
def reset(self):
"""
Resets the slot as empty.
"""
self.token = ' '
class Board:
"""
The playing field.
- Constructor Parameters
:param height: :type int:
- The board's height.
:param width: :type int:
- The board's width.
"""
def __init__(self, height, width, *args, **kwargs):
field = []
for i in range(width):
field.append([])
for j in range(height):
field[i].append(Slot(' '))
self.field = field
self.height = height
self.width = width
@property
def full(self):
"""
Returns if the board's slots are full.
"""
filled = 0
for i in range(self.width):
for j in range(self.height):
if not self.field[i][j].is_empty():
filled += 1
return filled == self.width * self.height
def draw(self, use_tokens=True):
"""
Prints the playing field in the command line.
- Parameters
:param use_tokens: :type bool:
- Defaults as True. If set as false,
the board is drawn in guide mode.
"""
printed = ''
ctr = 1
for i in range(self.height):
for j in range(self.width):
if use_tokens:
token = self.field[i][j].token
else:
token = str(ctr)
ctr += 1
if j == 0:
printed += f' {token} | '
else:
printed += token
if 0 < j < self.width - 1:
printed += ' | '
printed += '\n'
if i < self.height - 1:
printed += "-------------"
printed += "\n"
return printed
def write_turn(self, player, move):
"""
Writes a players turn into the board then returns a boolean value
representing if the move is valid.
- Parameters
:param player: :type Player:
The instance of the player making the turn.
:param input: :type str:
The player's move.
"""
ctr = 1
for i in range(self.width):
for j in range(self.height):
if int(move) == ctr:
return self.field[i][j].update_token(player.token)
else:
ctr += 1
return False
def equal_all_slots(self, slots):
"""
Checks if all tokens from a collection of slots are equal.
Returns True if they are.
- Parameters
:param slots: :type list:
- The collection of slots.
"""
for slot in slots:
if slot.is_empty():
return False
return len(set([slot.token for slot in slots])) == 1
def check_win(self):
"""
Returns a boolean value representing the
if a win condition is already met.
"""
win = False
for i in range(self.width):
horizontal_slots = []
vertical_slots = []
diagonal_top_to_bottom_slots = []
diagonal_bottom_to_top_slots = []
# Check horizontal and vertical.
for j in range(self.height):
h_slot = self.field[i][j]
v_slot = self.field[j][i]
horizontal_slots.append(h_slot)
vertical_slots.append(v_slot)
for j in range(self.height):
d_slot = self.field[j][j]
diagonal_top_to_bottom_slots.append(d_slot)
_height = copy.deepcopy(self.height - 1)
for j in range(self.height):
d_slot_1 = self.field[_height][j]
d_slot_2 = self.field[j][_height]
diagonal_bottom_to_top_slots.extend([d_slot_1, d_slot_2])
_height -= 1
h_win = self.equal_all_slots(horizontal_slots)
v_win = self.equal_all_slots(vertical_slots)
d1_win = self.equal_all_slots(diagonal_top_to_bottom_slots)
d2_win = self.equal_all_slots(diagonal_bottom_to_top_slots)
if h_win or v_win or d1_win or d2_win:
win = True
break
return win
def reset(self):
"""
Resets the fields of the board.
"""
for i in range(self.width):
for j in range(self.height):
self.field[i][j].reset()
|
# Copyright 2018 Google LLC.
#
# Redistribution and use in source and binary forms, with or without
# modification, are permitted provided that the following conditions
# are met:
#
# 1. Redistributions of source code must retain the above copyright notice,
# this list of conditions and the following disclaimer.
#
# 2. Redistributions in binary form must reproduce the above copyright
# notice, this list of conditions and the following disclaimer in the
# documentation and/or other materials provided with the distribution.
#
# 3. Neither the name of the copyright holder nor the names of its
# contributors may be used to endorse or promote products derived from this
# software without specific prior written permission.
#
# THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
# AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
# IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE
# ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE
# LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR
# CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF
# SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS
# INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN
# CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE)
# ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE
# POSSIBILITY OF SUCH DAMAGE.
"""Tests for third_party.nucleus.io.bed."""
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import sys
if 'google' in sys.modules and 'google.protobuf' not in sys.modules:
del sys.modules['google']
from absl.testing import absltest
from absl.testing import parameterized
from third_party.nucleus.io import bed
from third_party.nucleus.protos import bed_pb2
from third_party.nucleus.testing import test_utils
_VALID_NUM_BED_FIELDS = [3, 4, 5, 6, 8, 9, 12]
class BedReaderTests(parameterized.TestCase):
@parameterized.parameters('test_regions.bed', 'test_regions.bed.gz',
'test_regions.bed.tfrecord',
'test_regions.bed.tfrecord.gz')
def test_iterate_bed_reader(self, bed_filename):
bed_path = test_utils.genomics_core_testdata(bed_filename)
expected = [('chr1', 10, 20), ('chr1', 100, 200)]
with bed.BedReader(bed_path) as reader:
records = list(reader.iterate())
self.assertLen(records, 2)
self.assertEqual([(r.reference_name, r.start, r.end) for r in records],
expected)
@parameterized.parameters('test_regions.bed', 'test_regions.bed.gz')
def test_native_bed_header(self, bed_filename):
bed_path = test_utils.genomics_core_testdata(bed_filename)
with bed.BedReader(bed_path) as reader:
self.assertEqual(reader.header.num_fields, 12)
with bed.NativeBedReader(bed_path) as native_reader:
self.assertEqual(native_reader.header.num_fields, 12)
@parameterized.parameters(1, 2, 7, 10, 11, 13)
def test_invalid_num_fields(self, invalid_num_fields):
bed_path = test_utils.genomics_core_testdata('test_regions.bed')
with self.assertRaisesRegexp(ValueError, 'Invalid requested number of fie'):
_ = bed.BedReader(bed_path, num_fields=invalid_num_fields)
class BedWriterTests(parameterized.TestCase):
"""Tests for BedWriter."""
def setUp(self):
self.records = [
bed_pb2.BedRecord(
reference_name='chr1', start=30, end=40, name='first', score=55.5),
bed_pb2.BedRecord(
reference_name='chr2', start=32, end=38, name='second', score=0),
bed_pb2.BedRecord(
reference_name='chr3', start=40, end=50, name='third', score=99),
]
self.tokens = [
[
'chr1', '30', '40', 'first', '55.5', '+', '35', '38', '128,242,16',
'2', '5,3', '30,37'
],
[
'chr2', '32', '38', 'second', '0', '.', '32', '38', '128,128,128',
'1', '6', '32'
],
[
'chr3', '40', '50', 'third', '99', '-', '40', '44', '0,0,0', '3',
'40,43,48', '3,2,2'
],
]
@parameterized.parameters('test_raw.bed', 'test_zipped.bed.gz',
'test_raw.tfrecord', 'test_zipped.tfrecord.gz')
def test_roundtrip_writer(self, filename):
output_path = test_utils.test_tmpfile(filename)
with bed.BedWriter(
output_path, header=bed_pb2.BedHeader(num_fields=5)) as writer:
for record in self.records:
writer.write(record)
with bed.BedReader(output_path) as reader:
v2_records = list(reader.iterate())
self.assertEqual(self.records, v2_records)
@parameterized.parameters(3, 4, 5, 6, 8, 9, 12)
def test_roundtrip_num_fields(self, num_fields):
all_num_fields_in_file = [
n for n in _VALID_NUM_BED_FIELDS if n >= num_fields
]
for num_fields_in_file in all_num_fields_in_file:
lines = ['\t'.join(line[:num_fields_in_file]) for line in self.tokens]
contents = '{}\n'.format('\n'.join(lines))
input_path = test_utils.test_tmpfile('test_field.bed', contents=contents)
with bed.BedReader(input_path, num_fields=num_fields) as reader:
records = list(reader.iterate())
output_path = test_utils.test_tmpfile('test_field2.bed')
with bed.BedWriter(output_path, header=reader.header) as writer:
for record in records:
writer.write(record)
with bed.BedReader(output_path) as reader2:
v2_records = list(reader2.iterate())
self.assertLen(records, 3)
self.assertEqual(records, v2_records)
if __name__ == '__main__':
absltest.main()
|
import pygame
from pygame.locals import *
import random
# Generate Food Dot
def GenerateFood():
global screen, add
global height, width
global startX, startY
global positionHistory
RED = (255, 0, 0)
while True:
w = random.randint(0, height - add)
h = random.randint(0, width - add)
w -= w % add
h -= h % add
flag = 1
for x in positionHistory:
if x == (w,h):
flag = 0
if flag:
break
pygame.draw.rect(screen, RED, pygame.Rect(w, h, add, add))
pygame.display.update()
return w, h
def PositionAppend(positionArray, direction):
global snakeLen
if direction == 0:
temp = (positionHistory[snakeLen - 1][0] + add)
if temp >= width:
temp -= width
positionHistory.append((temp, positionHistory[snakeLen - 1][1]))
elif direction == 1:
temp = (positionHistory[snakeLen - 1][1] - add)
if temp < 0:
temp += height
positionHistory.append((positionHistory[snakeLen - 1][0], temp))
elif direction == 2:
temp = (positionHistory[snakeLen - 1][0] - add)
if temp < 0:
temp += width
positionHistory.append((temp, positionHistory[snakeLen - 1][1]))
else:
temp = (positionHistory[snakeLen - 1][1] + add)
if temp >= height:
temp -= height
positionHistory.append((positionHistory[snakeLen - 1][0], temp))
def CheckGame():
global total, snakeLen
if total == snakeLen:
return True
return False
def CheckDead(positionArray):
arrLen = len(positionArray) - 1
for index, x in enumerate(positionArray):
if index == arrLen:
return False
if x == positionArray[arrLen]:
return True
return False
# Initialize Game
pygame.init()
pygame.display.set_caption("New Caption")
# Initialize Variable
GREEN = (127,255,0)
BLACK = (0, 0, 0)
HEAD = (32,178,170)
size = 1000, 1000
add = 40
direction = 0 #0 Right, 1 Up, 2 Left, 3 Bottom
width, height = size
startX = width // 2
startY = height // 2
startX -= startX % add
startY -= startY % add
# Total Length to win the game
total = (size[0] // add) * (size[1] // add)
# Food Array
food = []
# Pygame Screen
screen = pygame.display.set_mode(size)
screen.fill((0, 0, 0))
pygame.display.update()
# Position History For Snake
positionHistory = [(startX, startY), (startX, startY + add)]
snakeLen = 2
# Draw First 2 Dot
pygame.draw.rect(screen, GREEN, pygame.Rect(positionHistory[0][0], positionHistory[0][1], add, add))
pygame.draw.rect(screen, GREEN, pygame.Rect(positionHistory[1][0], positionHistory[1][1], add, add))
pygame.display.update()
running = True
win = False
# Unlimited Loop
while(running):
for event in pygame.event.get():
if event.type == pygame.QUIT:
running = False
if event.type == KEYDOWN:
if event.key == K_RIGHT and direction != 2:
direction = 0
elif event.key == K_UP and direction != 3:
direction = 1
elif event.key == K_LEFT and direction != 0:
direction = 2
elif event.key == K_DOWN and direction != 1:
direction = 3
if len(food) == 0:
food.append(GenerateFood())
PositionAppend(positionHistory, direction)
flag = 1
if positionHistory[snakeLen] == food[0]:
flag = 0
food.pop()
snakeLen+=1
for index, x in enumerate(positionHistory):
if index == 0 and flag == 1:
pygame.draw.rect(screen, BLACK, pygame.Rect(x[0], x[1], add, add))
elif index == snakeLen:
pygame.draw.rect(screen, HEAD, pygame.Rect(x[0], x[1], add, add))
else:
pygame.draw.rect(screen, GREEN, pygame.Rect(x[0], x[1], add, add))
if flag:
positionHistory.pop(0)
pygame.display.update()
if CheckGame():
win = True
break
if CheckDead(positionHistory, ):
break
pygame.time.wait(100)
if win:
print("You Win")
else:
print("Game Over")
pygame.quit()
|
<gh_stars>0
import glob
import os
import subprocess
import unittest
import netCDF4
import numpy as np
import bald
from bald.tests import BaldTestCase
from rdflib import Graph
# a module level graph, to share for memory and performance
thisGraph = [Graph()]
loaded_boolean = []
class Test(BaldTestCase):
def setUp(self):
self.cdl_path = os.path.join(os.path.dirname(__file__), 'CDL')
# Check to see if another test has already loaded the graph.
if not loaded_boolean:
# load bald graphs from cdl
for cdl_file in glob.glob(os.path.join(self.cdl_path, '*.cdl')):
with self.temp_filename('.nc') as tfile:
subprocess.check_call(['ncgen', '-o', tfile, cdl_file])
root_container = bald.load_netcdf(tfile, cache=self.acache)
curr_g = root_container.rdfgraph()
#merge into graph in test obj
thisGraph[0] = thisGraph[0] + curr_g
print('setting loaded_boolean')
loaded_boolean.append(True)
self.graph = thisGraph[0]
def test_sparql_count_standard_names(self):
#query standard_name values used and frequency
qres = self.graph.query(
""" PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX bald: <http://binary-array-ld.net/latest/>
SELECT ?name (COUNT(?name) as ?NELEMENTS)
WHERE {
?contained a bald:Array .
?contained ?pred ?name
FILTER(regex(str(?pred), "standard_name"))
}
GROUP BY ?name
ORDER BY DESC(?NELEMENTS)
""")
# for row in qres:
# print("%s :: %s" % row)
# print( len(qres))
expected_result_rows = 15
self.assertTrue(len(qres) >= expected_result_rows)
def test_sparql_demo_graph_viz_labels(self):
#query standard_name values used and frequency
qres = self.graph.query(
"""
PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX bald: <http://binary-array-ld.net/latest/>
PREFIX cf: <http://def.scitools.org.uk/CFTerms/>
SELECT ?container ?contained ?containerName ?containedlabel
WHERE {
?container a bald:Container .
?container bald:contains ?contained .
?contained a bald:Array .
{ ?contained cf:long_name ?containedlabel }
UNION
{ ?contained ?lnprop ?containedlabel
FILTER(regex(str(?lnprop), "long_name", "i"))
}
BIND( str(?container) as ?containerName) }
""")
# for row in qres:
# print("%s, %s, %s, %s" % row)
# print( len(qres))
expected_result_rows = 150
self.assertTrue(len(qres) >= expected_result_rows)
def test_sparql_demo_map_viz_labels(self):
#query standard_name values used and frequency
qres = self.graph.query(
"""
PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX bald: <http://binary-array-ld.net/latest/>
PREFIX geo: <http://www.opengis.net/ont/geosparql#>
SELECT ?contained ?geoWKT
WHERE
{
?container a bald:Container .
?container bald:contains ?contained .
?contained geo:asWKT ?geoWKT
}
""")
# for row in qres:
# print("%s, %s" % row)
# print( len(qres))
expected_result_rows = 2
self.assertTrue(len(qres) >= expected_result_rows)
|
<filename>talking_heads/network_ops.py<gh_stars>1-10
# -*- coding: utf-8 -*-
#/usr/bin/python3
import tensorflow as tf
##################################################################################
# Initialization
##################################################################################
# Xavier : tf_contrib.layers.xavier_initializer()
# He : tf_contrib.layers.variance_scaling_initializer()
# Normal : tf.random_normal_initializer(mean=0.0, stddev=0.02)
# Truncated_normal : tf.truncated_normal_initializer(mean=0.0, stddev=0.02)
# Orthogonal : tf.orthogonal_initializer(1.0) / relu = sqrt(2), the others = 1.0
##################################################################################
# Regularizers
##################################################################################
def orthogonal_regularizer(scale) :
""" Defining the Orthogonal regularizer and return the function at last to be used in Conv layer as kernel regularizer"""
def ortho_reg(w) :
""" Reshaping the matrxi in to 2D tensor for enforcing orthogonality"""
c = w.get_shape().as_list()[-1]
w = tf.reshape(w, [-1, c])
""" Declaring a Identity Tensor of appropriate size"""
identity = tf.eye(c)
""" Regularizer Wt*W - I """
w_transpose = tf.transpose(w)
w_mul = tf.matmul(w_transpose, w)
reg = tf.subtract(w_mul, identity)
"""Calculating the Loss Obtained"""
ortho_loss = tf.nn.l2_loss(reg)
return scale * ortho_loss
return ortho_reg
def orthogonal_regularizer_fully(scale) :
""" Defining the Orthogonal regularizer and return the function at last to be used in Fully Connected Layer """
def ortho_reg_fully(w) :
""" Reshaping the matrix in to 2D tensor for enforcing orthogonality"""
c = w.get_shape().as_list()[-1]
"""Declaring a Identity Tensor of appropriate size"""
identity = tf.eye(c)
w_transpose = tf.transpose(w)
w_mul = tf.matmul(w_transpose, w)
reg = tf.subtract(w_mul, identity)
""" Calculating the Loss """
ortho_loss = tf.nn.l2_loss(reg)
return scale * ortho_loss
return ortho_reg_fully
##################################################################################
# Regularization
##################################################################################
# l2_decay : tf_contrib.layers.l2_regularizer(0.0001)
# orthogonal_regularizer : orthogonal_regularizer(0.0001) / orthogonal_regularizer_fully(0.0001)
weight_init = tf.truncated_normal_initializer(mean=0.0, stddev=0.02)
weight_regularizer = orthogonal_regularizer(0.0001)
weight_regularizer_fully = orthogonal_regularizer_fully(0.0001)
# Regularization only G in BigGAN
##################################################################################
# Layer
##################################################################################
# pad = ceil[ (kernel - stride) / 2 ]
def conv(x, channels, kernel=4, stride=2, pad=0, pad_type='zero', use_bias=True, sn=False, scope='conv_0'):
with tf.variable_scope(scope):
if pad > 0:
h = x.get_shape().as_list()[1]
if h % stride == 0:
pad = pad * 2
else:
pad = max(kernel - (h % stride), 0)
pad_top = pad // 2
pad_bottom = pad - pad_top
pad_left = pad // 2
pad_right = pad - pad_left
padding = [[0, 0], [pad_top, pad_bottom], [pad_left, pad_right], [0, 0]]
if pad_type == 'zero' :
x = tf.pad(x, padding)
if pad_type == 'reflect' :
x = tf.pad(x, padding, mode='REFLECT')
if scope.__contains__('generator') :
w = tf.get_variable("kernel", shape=[kernel, kernel, x.get_shape()[-1], channels], initializer=weight_init,
regularizer=weight_regularizer)
else :
w = tf.get_variable("kernel", shape=[kernel, kernel, x.get_shape()[-1], channels], initializer=weight_init,
regularizer=None)
if sn:
x = tf.nn.conv2d(input=x, filter=spectral_norm(w),
strides=[1, stride, stride, 1], padding='VALID')
else:
x = tf.nn.conv2d(input=x, filter=w,
strides=[1, stride, stride, 1], padding='VALID')
if use_bias :
bias = tf.get_variable("bias", [channels], initializer=tf.constant_initializer(0.0))
x = tf.nn.bias_add(x, bias)
return x
def deconv(x, channels, kernel=4, stride=2, padding='SAME', use_bias=True, sn=False, scope='deconv_0'):
with tf.variable_scope(scope):
x_shape = x.get_shape().as_list()
if padding == 'SAME':
output_shape = tf.stack([tf.shape(x)[0], x_shape[1] * stride, x_shape[2] * stride, channels])
else:
output_shape = tf.stack([tf.shape(x)[0], x_shape[1] * stride + max(kernel - stride, 0), x_shape[2] * stride + max(kernel - stride, 0), channels])
w = tf.get_variable("kernel", shape=[kernel, kernel, channels, x_shape[-1]], initializer=weight_init, regularizer=weight_regularizer)
if sn:
x = tf.nn.conv2d_transpose(x, filter=spectral_norm(w), output_shape=output_shape, strides=[1, stride, stride, 1], padding=padding)
else:
x = tf.nn.conv2d_transpose(x, filter=w, output_shape=output_shape, strides=[1, stride, stride, 1], padding=padding)
if use_bias :
bias = tf.get_variable("bias", [channels], initializer=tf.constant_initializer(0.0))
x = tf.nn.bias_add(x, bias)
return x
def fully_connected(x, units, use_bias=True, is_training=True, sn=False, scope='fully_0'):
with tf.variable_scope(scope):
x = flatten(x)
shape = x.get_shape().as_list()
channels = shape[-1]
if scope.__contains__('generator'):
w = tf.get_variable("kernel", [channels, units], tf.float32, initializer=weight_init,
regularizer=weight_regularizer_fully)
else :
w = tf.get_variable("kernel", [channels, units], tf.float32, initializer=weight_init,
regularizer=None)
if sn:
x = tf.matmul(x, spectral_norm(w))
else:
x = tf.matmul(x, w)
if use_bias:
bias = tf.get_variable("bias", [units], initializer=tf.constant_initializer(0.0), trainable=is_training)
x = tf.nn.bias_add(x, bias)
return x
def flatten(x) :
return tf.keras.layers.Flatten()(x)
def hw_flatten(x) :
_,h,w,c = x.get_shape().as_list()
return tf.reshape(x, shape=[-1, h*w, c])
##################################################################################
# Residual-block, Self-Attention-block
##################################################################################
def resblock(x_init, channels, kernel=3, pads=[1,1], use_bias=True, is_training=True, sn=False, scope='resblock'):
with tf.variable_scope(scope):
with tf.variable_scope('res1'):
x = conv(x_init, channels, kernel=kernel, stride=1, pad=pads[0], use_bias=use_bias, sn=sn)
x = instance_norm(x, is_training)
x = relu(x)
with tf.variable_scope('res2'):
x = conv(x, channels, kernel=kernel, stride=1, pad=pads[1], use_bias=use_bias, sn=sn)
x = instance_norm(x, is_training)
return relu(x + x_init)
def resblock_condition(x_init, conds, channels, kernel=3, pads=[1,1], use_bias=True, is_training=True, sn=False, scope='resblock'):
with tf.variable_scope(scope):
with tf.variable_scope('res1'):
x = conv(x_init, channels, kernel=kernel, stride=1, pad=pads[0], use_bias=use_bias, sn=sn)
x = adaptive_instance_norm(x, conds[:2], is_training)
x = relu(x)
with tf.variable_scope('res2'):
x = conv(x, channels, kernel=kernel, stride=1, pad=pads[1], use_bias=use_bias, sn=sn)
x = adaptive_instance_norm(x, conds[2:], is_training)
return relu(x + x_init)
def resblock_up(x_init, channels, use_bias=True, is_training=True, sn=False, scope='resblock_up'):
with tf.variable_scope(scope):
with tf.variable_scope('res1'):
x = instance_norm(x_init, is_training)
x = relu(x)
x = deconv(x, channels, kernel=3, stride=2, use_bias=use_bias, sn=sn)
with tf.variable_scope('res2') :
x = instance_norm(x, is_training)
x = relu(x)
x = deconv(x, channels, kernel=3, stride=1, use_bias=use_bias, sn=sn)
with tf.variable_scope('skip') :
x_skip = deconv(x_init, channels, kernel=3, stride=2, use_bias=use_bias, sn=sn)
return x + x_skip
def resblock_up_condition(x_init, conds, channels, use_bias=True, is_training=True, sn=False, scope='resblock_up'):
with tf.variable_scope(scope):
with tf.variable_scope('res1'):
x = adaptive_instance_norm(x_init, conds[:2], is_training)
x = relu(x)
x = deconv(x, channels, kernel=3, stride=2, use_bias=use_bias, sn=sn)
with tf.variable_scope('res2') :
x = adaptive_instance_norm(x, conds[2:], is_training)
x = relu(x)
x = deconv(x, channels, kernel=3, stride=1, use_bias=use_bias, sn=sn)
with tf.variable_scope('skip') :
x_skip = deconv(x_init, channels, kernel=3, stride=2, use_bias=use_bias, sn=sn)
return x + x_skip
def resblock_down(x_init, channels, use_bias=True, is_training=True, sn=False, scope='resblock_down'):
with tf.variable_scope(scope):
with tf.variable_scope('res1'):
x = instance_norm(x_init, is_training)
x = relu(x)
x = conv(x, channels, kernel=3, stride=2, pad=1, use_bias=use_bias, sn=sn)
with tf.variable_scope('res2') :
x = instance_norm(x, is_training)
x = relu(x)
x = conv(x, channels, kernel=3, stride=1, pad=1, use_bias=use_bias, sn=sn)
with tf.variable_scope('skip') :
x_skip = conv(x_init, channels, kernel=3, stride=2, pad=1, use_bias=use_bias, sn=sn)
return x + x_skip
def resblock_down_no_instance_norm(x_init, channels, use_bias=True, is_training=True, sn=False, scope='resblock_down'):
with tf.variable_scope(scope):
with tf.variable_scope('res1'):
x = relu(x_init)
x = conv(x, channels, kernel=3, stride=2, pad=1, use_bias=use_bias, sn=sn)
with tf.variable_scope('res2') :
x = relu(x)
x = conv(x, channels, kernel=3, stride=1, pad=1, use_bias=use_bias, sn=sn)
with tf.variable_scope('skip') :
x_skip = conv(x_init, channels, kernel=3, stride=2, pad=1, use_bias=use_bias, sn=sn)
return x + x_skip
def self_attention(x, channels, sn=False, scope='self_attention'):
with tf.variable_scope(scope):
f = conv(x, channels // 8, kernel=1, stride=1, sn=sn, scope='f_conv') # [bs, h, w, c']
g = conv(x, channels // 8, kernel=1, stride=1, sn=sn, scope='g_conv') # [bs, h, w, c']
h = conv(x, channels, kernel=1, stride=1, sn=sn, scope='h_conv') # [bs, h, w, c]
# N = h * w
s = tf.matmul(hw_flatten(g), hw_flatten(f), transpose_b=True) # # [bs, N, N]
beta = tf.nn.softmax(s) # attention map
# TODO: check that softmax along the last dimension was the correct one
o = tf.matmul(beta, hw_flatten(h)) # [bs, N, C]
gamma = tf.get_variable("gamma", [1], initializer=tf.constant_initializer(0.0))
o = tf.reshape(o, shape=[-1, x.shape[1], x.shape[2], x.shape[3]]) # [bs, h, w, C]
x = gamma * o + x
return x
def self_attention_2(x, channels, sn=False, scope='self_attention'):
with tf.variable_scope(scope):
f = conv(x, channels // 8, kernel=1, stride=1, sn=sn, scope='f_conv') # [bs, h, w, c']
f = max_pooling(f)
g = conv(x, channels // 8, kernel=1, stride=1, sn=sn, scope='g_conv') # [bs, h, w, c']
h = conv(x, channels // 2, kernel=1, stride=1, sn=sn, scope='h_conv') # [bs, h, w, c]
h = max_pooling(h)
# N = h * w
s = tf.matmul(hw_flatten(g), hw_flatten(f), transpose_b=True) # # [bs, N, N]
beta = tf.nn.softmax(s) # attention map
# TODO: check that softmax along the last dimension was the correct one
o = tf.matmul(beta, hw_flatten(h)) # [bs, N, C]
gamma = tf.get_variable("gamma", [1], initializer=tf.constant_initializer(0.0))
o = tf.reshape(o, shape=[-1, x.shape[1], x.shape[2], channels // 2]) # [bs, h, w, C]
o = conv(o, channels, kernel=1, stride=1, sn=sn, scope='attn_conv')
x = gamma * o + x
return x
##################################################################################
# Sampling
##################################################################################
def global_avg_pooling(x):
gap = tf.reduce_mean(x, axis=[1, 2])
return gap
def global_sum_pooling(x) :
gsp = tf.reduce_sum(x, axis=[1, 2])
return gsp
def max_pooling(x) :
x = tf.nn.max_pool(x, ksize=[1,2,2,1], strides=[1,2,2,1], padding='SAME')
return x
def up_sample(x, scale_factor=2):
_, h, w, _ = x.get_shape().as_list()
new_size = [h * scale_factor, w * scale_factor]
return tf.image.resize_nearest_neighbor(x, size=new_size)
##################################################################################
# Activation function
##################################################################################
def lrelu(x, alpha=0.2):
return tf.nn.leaky_relu(x, alpha)
def relu(x):
return tf.nn.relu(x)
def tanh(x):
return tf.tanh(x)
def sigmoid(x):
return tf.nn.sigmoid(x)
##################################################################################
# Normalization function
##################################################################################
def instance_norm(x, is_training=True, scope='IN'):
with tf.variable_scope(scope) :
_, w, h, c = x.get_shape().as_list()
epsilon = 1e-05
beta = tf.get_variable("beta", shape=[c], dtype=tf.float32, initializer=tf.constant_initializer(0.0))
gamma = tf.get_variable("gamma", shape=[c], dtype=tf.float32, initializer=tf.constant_initializer(0.0))
batch_mean, batch_var = tf.nn.moments(x, [1, 2])
x = tf.reshape(tf.transpose(x, perm = [0,3,1,2]), shape = [-1,c,w*h])
x = tf.nn.batch_normalization(x,
tf.expand_dims(batch_mean, axis=-1), tf.expand_dims(batch_var, axis=-1),
tf.expand_dims(beta, axis=-1), tf.expand_dims(gamma, axis=-1), epsilon)
x = tf.transpose(tf.reshape(x, shape = [-1,c,w,h]), perm = [0,2,3,1])
return x
#def instance_norm(x, is_training=True, scope='IN'):
# epsilon = 1e-05
# return tf.contrib.layers.instance_norm(x, epsilon = epsilon, scope = scope, trainable = is_training)
def adaptive_instance_norm(x, z, is_training=True, scope='AdaIN'):
with tf.variable_scope(scope) :
_, w, h, c = x.get_shape().as_list()
epsilon = 1e-05
beta = z[0]
gamma = z[1]
batch_mean, batch_var = tf.nn.moments(x, [1, 2])
x = tf.reshape(tf.transpose(x, perm = [0,3,1,2]), shape = [-1,c,w*h])
x = tf.nn.batch_normalization(x,
tf.expand_dims(batch_mean, axis=-1), tf.expand_dims(batch_var, axis=-1),
tf.expand_dims(beta, axis=-1), tf.expand_dims(gamma, axis=-1), epsilon)
x = tf.transpose(tf.reshape(x, shape = [-1,c,w,h]), perm = [0,2,3,1])
return x
def spectral_norm(w, iteration=1):
w_shape = w.shape.as_list()
w = tf.reshape(w, [-1, w_shape[-1]])
u = tf.get_variable("u", [1, w_shape[-1]], initializer=tf.random_normal_initializer(), trainable=False)
u_hat = u
v_hat = None
for i in range(iteration):
"""
power iteration
Usually iteration = 1 will be enough
"""
v_ = tf.matmul(u_hat, tf.transpose(w))
v_hat = tf.nn.l2_normalize(v_)
u_ = tf.matmul(v_hat, w)
u_hat = tf.nn.l2_normalize(u_)
u_hat = tf.stop_gradient(u_hat)
v_hat = tf.stop_gradient(v_hat)
sigma = tf.matmul(tf.matmul(v_hat, w), tf.transpose(u_hat))
with tf.control_dependencies([u.assign(u_hat)]):
w_norm = w / sigma
w_norm = tf.reshape(w_norm, w_shape)
return w_norm |
########################################################################
# ___ _ _____________
# / | / | / /_ __/ ___/
# / /| | / |/ / / / \__ \
# / ___ |/ /| / / / ___/ /
# /_/ |_/_/ |_/ /_/ /____/
#
# Test Problems for constucting material cross sections for materials
# listed below. The cross sections are compared to the original one-
# dimenionsal 'discrete1' package.
#
# Materials: "uranium", "uranium-hydride", "plutonium"
# "stainless-steel-440", "high-density-polyethyene-618",
# "high-density-polyethyene-087", "carbon", "uranium-235",
# "uranium-238", "water-uranium-dioxide", "plutonium-239",
# "plutonium-240"
#
########################################################################
from ants.materials import Materials
from discrete1 import generate, fixed
import pytest
import numpy as np
import pkg_resources
ENR_PATH = pkg_resources.resource_filename('ants','data/energy/')
_ENERGY_BOUNDS = np.load(ENR_PATH + 'energy_bounds.npz')
@pytest.mark.enrichment
def test_uranium_enrichment(enrich):
legacy_xs = generate.XSGenerate087('U', \
enrich=int(enrich)*0.01).cross_section()
material_list = ["uranium-%{}%".format(enrich)]
ants_problem = Materials(material_list, g087r, _ENERGY_BOUNDS['87'])
ants_xs = ants_problem.data["material-" + material_list[0]]
assert [np.all(np.allclose(ants_xs[xs], legacy_xs[xs], atol=1e-12)) \
for xs in range(3)]
@pytest.mark.enrichment
@pytest.mark.energy087r
def test_uranium_energy_coarsen(enrich, g087r):
legacy_xs = fixed.UraniumStainless.steady(g087r, 2, \
enrich=int(enrich)*0.01)
legacy_xs = [legacy_xs[4][500], legacy_xs[5][500], legacy_xs[6][500]]
material_list = ["uranium-%{}%".format(enrich)]
ants_problem = Materials(material_list, g087r, _ENERGY_BOUNDS['87'])
ants_xs = ants_problem.data["material-" + material_list[0]]
assert [np.all(np.allclose(ants_xs[xs], legacy_xs[xs], atol=1e-12)) \
for xs in range(3)]
@pytest.mark.enrichment
def test_uranium_hydride_enrichment(enrich):
legacy_xs = generate.XSGenerate087('UH3', \
enrich=int(enrich)*0.01).cross_section()
material_list = ["uranium-hydride-%{}%".format(enrich)]
ants_problem = Materials(material_list, 87, _ENERGY_BOUNDS['87'])
ants_xs = ants_problem.data["material-" + material_list[0]]
assert [np.all(np.allclose(ants_xs[xs], legacy_xs[xs], atol=1e-12)) \
for xs in range(3)]
@pytest.mark.enrichment
@pytest.mark.energy087r
def test_uranium_hydride_energy_coarsen(enrich, g087r):
legacy_xs = []
temp_xs = generate.XSGenerate087('UH3', \
enrich=int(enrich)*0.01).cross_section()
idx = generate.ReduceTools.index_generator(87, g087r)
legacy_xs.append(generate.ReduceTools.vector_reduction(temp_xs[0], idx))
legacy_xs.append(generate.ReduceTools.matrix_reduction(temp_xs[1], idx))
legacy_xs.append(generate.ReduceTools.matrix_reduction(temp_xs[2], idx))
material_list = ["uranium-hydride-%{}%".format(enrich)]
ants_problem = Materials(material_list, g087r, _ENERGY_BOUNDS['87'])
ants_xs = ants_problem.data["material-" + material_list[0]]
assert [np.all(np.allclose(ants_xs[xs], legacy_xs[xs], atol=1e-12)) \
for xs in range(3)]
@pytest.mark.enrichment
def test_plutonium_enrichment(enrich):
legacy_xs = generate.XSGenerate618.cross_section(1 - int(enrich)*0.01)
material_list = ["plutonium-%{}%".format(enrich)]
ants_problem = Materials(material_list, 618, _ENERGY_BOUNDS['618'])
ants_xs = ants_problem.data["material-" + material_list[0]]
assert [np.all(np.allclose(ants_xs[xs], legacy_xs[xs], atol=1e-12)) \
for xs in range(3)]
@pytest.mark.enrichment
@pytest.mark.energy618r
def test_plutonium_energy_coarsen(enrich, g618r):
legacy_xs = generate.XSGenerate618.cross_section_reduce(g618r, \
1 - int(enrich)*0.01)
material_list = ["plutonium-%{}%".format(enrich)]
ants_problem = Materials(material_list, g618r, _ENERGY_BOUNDS['618'])
ants_xs = ants_problem.data["material-" + material_list[0]]
assert [np.all(np.allclose(ants_xs[xs], legacy_xs[xs], atol=1e-12)) \
for xs in range(3)]
# _, _, _, _, orig_total, orig_scatter, orig_fission, _, _, _, _,
# orig_total = orig_total[500].copy()
# orig_scatter = orig_scatter[500].copy()
# orig_fission = orig_fission[500].copy()
# from discrete1.generate import generate.XSGenerate087, XSGenerate618
# from ants.materials import Materials
# from discrete1.fixed import SHEM, UraniumStainless
# import numpy as np
# import matplotlib.pyplot as plt
# """
# [x] uranium
# [x] uranium-hydride
# [.] plutonium
# [x] stainless-steel-440
# [.] high-density-polyethyene-618
# [x] high-density-polyethyene-087
# [x] carbon
# [x] uranium-235
# [x] uranium-238
# [ ] water-uranium-dioxide
# [.] plutonium-239
# [.] plutonium-240
# """
# _ENERGY_BOUNDS = np.load('ants/ants/materials_sources/energy_bounds.npz')['87']
# new_problem = Materials([("uranium",0.20), ("high-density-polyethyene-087",)], 43, energy_bounds)
# new_problem.add_source('14.1-mev',0)
# new_source = new_problem.data['source-14.1-mev']
# _, _, _, _, _, _, _, _, _, _, orig_source, = UraniumStainless.steady(43, 8)
# print(np.array_equal(new_source[0], orig_source[1]))
# energy_bounds = np.load('ants/ants/materials_sources/energy_bounds.npz')['361']
# energy_idx = np.load('ants/ants/materials_sources/group_indices_361G.npz')['240']
# new_problem = Materials([("water-uranium-dioxide",0.20)], 240, energy_bounds, energy_idx)
# new_problem.add_source('ambe-point',0)
# new_source = new_problem.data['source-ambe-point']
# _, _, _, _, _, _, _, _, _, _, orig_source, = SHEM.steady(240, 8)
# print(np.array_equal(new_source[0], orig_source[1]))
# new_problem.compile_cross_section()
# print(new_problem.data.keys())
# stainless steel
# for ii in [87, 80, 60, 43, 21, 10]:
# # for ii in [87]:
# # orig_total, orig_scatter, orig_fission = XSGenerate087('SS440').cross_section()
# _, _, _, _, orig_total, orig_scatter, orig_fission, _, _, _, _, = UraniumStainless.steady(ii, 8)
# orig_total = orig_total[500].copy()
# orig_scatter = orig_scatter[500].copy()
# orig_fission = orig_fission[500].copy()
# # None is energy bounds
# energy_bounds = np.load('ants/ants/materials_sources/energy_bounds.npz')['87']
# new_problem = Materials([("uranium",0.20)], ii, energy_bounds)
# new_total, new_scatter, new_fission = new_problem.material_cross_section()
# print('Groups ',ii)
# print('Total:', np.array_equal(orig_total, new_total), np.all(np.isclose(orig_total, new_total, atol=1e-12)))
# print('Scatter:', np.array_equal(orig_scatter, new_scatter), np.all(np.isclose(orig_scatter, new_scatter, atol=1e-12)))
# print('Fission:', np.array_equal(orig_fission, new_fission), np.all(np.isclose(orig_fission, new_fission, atol=1e-12)))
# print(np.amax(orig_total - new_total))
# print(np.amax(orig_scatter - new_scatter))
# print(np.amax(orig_fission - new_fission))
# print()
# # SHEM
# _, _, _, _, orig_total, orig_scatter, orig_fission, _, _, _, _, = SHEM.steady(240, 8)
# orig_total = orig_total[0].copy()
# orig_scatter = orig_scatter[0].copy()
# orig_fission = orig_fission[0].copy()
# # None is energy bounds
# energy_bounds = np.load('ants/ants/materials_sources/energy_bounds.npz')['361']
# energy_idx = np.load('ants/ants/materials_sources/group_indices_361G.npz')['240']
# new_problem = Materials([("water-uranium-dioxide",)], 240, energy_bounds, energy_idx)
# new_total, new_scatter, new_fission = new_problem.material_cross_section()
# print('SHEM')
# print('Total:', np.array_equal(orig_total, new_total))
# print('Scatter:', np.array_equal(orig_scatter, new_scatter))
# print('Fission:', np.array_equal(orig_fission, new_fission))
# print(np.amax(orig_total - new_total))
# print(np.amax(orig_scatter - new_scatter))
# print(np.amax(orig_fission - new_fission))
# print()
# for enrichment in [0.0, 0.25, 0.5, 0.75, 1.0]:
# # for enrichment in [1.0]:
# # uranium
# orig_total, orig_scatter, orig_fission = XSGenerate087('UH3',enrich=enrichment).cross_section()
# # None is energy bounds
# new_problem = Materials([("uranium-hydride",enrichment)], 87, None)
# new_total, new_scatter, new_fission = new_problem.material_cross_section()
# for enrichment in [0.0, 0.25, 0.5, 0.75, 1.0]:
# # for enrichment in [0.25]:
# # uranium
# orig_total, orig_scatter, orig_fission = XSGenerate618.cross_section(enrichment)
# # None is energy bounds
# # new_problem = Materials([("high-density-polyethyene-618",1-enrichment)], 618, None)
# new_problem = Materials([("plutonium-240",1-enrichment)], 618, None)
# new_total, new_scatter, new_fission = new_problem.material_cross_section()
# print('Uranium Hydride - {}'.format(enrichment))
# print('Total:', np.array_equal(orig_total[2], new_total))
# print('Scatter:', np.array_equal(orig_scatter[2], new_scatter))
# print('Fission:', np.array_equal(orig_fission[2], new_fission))
# print(np.amax(orig_total[2] - new_total))
# print(np.amax(orig_scatter[2] - new_scatter))
# print(np.amax(orig_fission[2] - new_fission))
# print()
|
<gh_stars>1-10
import numpy as np
import tensorflow as tf
class ClockworkRNN(object):
'''
A Clockwork RNN - Koutnik et al. 2014 [arXiv, https://arxiv.org/abs/1402.3511]
The Clockwork RNN (CW-RNN), in which the hidden layer is partitioned into separate modules,
each processing inputs at its own temporal granularity, making computations only at its prescribed clock rate.
Rather than making the standard RNN models more complex, CW-RNN reduces the number of RNN parameters,
improves the performance significantly in the tasks tested, and speeds up the network evaluation
'''
def __init__(self, config):
self.config = config
# Check if the number of groups (periods) in the hidden layer
# is compatible with the total number of units in the layer. Note that
# this is not a requirement in the paper; there the extra neurons are
# divided over the higher frequency groups.
assert self.config.num_hidden % len(self.config.periods) == 0
# Global training step
self.global_step = tf.Variable(0, name='global_step', trainable=False)
# Initialize placeholders
self.inputs = tf.placeholder(
dtype=tf.float32,
shape=[None, self.config.num_steps, self.config.num_input],
name="inputs")
self.targets = tf.placeholder(
dtype=tf.float32,
shape=[None, self.config.num_output],
name="targets")
# Build the complete model
self._build_model()
# Initialize the optimizer with gradient clipping
self._init_optimizer()
# Operations for creating summaries
self._build_summary_ops()
def _build_model(self):
# Weight and bias initializers
initializer_weights = tf.contrib.layers.variance_scaling_initializer()
# initializer_weights = tf.contrib.layers.xavier_initializer(uniform=False, seed=None, dtype=tf.float32)
initializer_bias = tf.constant_initializer(0.0)
group_size = int(self.config.num_hidden / len(self.config.periods))
counter = 0
# Activation functions of the hidden and output state
activation_hidden = tf.tanh
# activation_output = tf.tanh
activation_output = None
# Split into list of tensors, one for each timestep
x_list = [tf.squeeze(x, axis=[1])
for x in tf.split(
axis=1, num_or_size_splits=self.config.num_steps,
value=self.inputs, name="inputs_list")]
# Periods of each group: 1,2,4, ..., 256 (in the case num_periods=9)
self.clockwork_periods = self.config.periods
# Mask for matrix W_I to make sure it's upper triangular
# self.clockwork_mask = tf.constant(np.triu(np.ones([self.config.num_hidden, self.config.num_hidden])), dtype=tf.float32, name="mask")
self.clockwork_mask = tf.constant(np.ones([self.config.num_hidden, self.config.num_hidden]), dtype=tf.float32, name="mask")
mask = np.ones([self.config.num_hidden, self.config.num_hidden])
for i in range(1, len(self.config.periods)):
k = int(i*group_size)
mask[k:, :max(k, 0)] = 0
self.clockwork_mask = mask
with tf.variable_scope("input"):
self.input_W = tf.get_variable("W", shape=[self.config.num_input, self.config.num_hidden], initializer=initializer_weights) # W_I
self.input_b = tf.get_variable("b", shape=[self.config.num_hidden], initializer=initializer_bias) # b_I
with tf.variable_scope("hidden"):
self.hidden_W = tf.get_variable("W", shape=[self.config.num_hidden, self.config.num_hidden], initializer=initializer_weights) # W_H
self.hidden_W = tf.multiply(self.hidden_W, self.clockwork_mask) # => upper triangular matrix # W_H
self.hidden_b = tf.get_variable("b", shape=[self.config.num_hidden], initializer=initializer_bias) # b_H
with tf.variable_scope("output"):
self.output_W = tf.get_variable("W", shape=[self.config.num_hidden, self.config.num_output], initializer=initializer_weights) # W_O
self.output_b = tf.get_variable("b", shape=[self.config.num_output], initializer=initializer_bias) # b_O
with tf.variable_scope("clockwork_cell") as scope:
# Initialize the hidden state of the cell to zero (this is y_{t_1})
self.state = tf.get_variable("state", shape=[self.config.batch_size, self.config.num_hidden], initializer=tf.zeros_initializer(), trainable=False)
pred = [] # initialize list of predictions
for time_step in range(1, self.config.num_steps + 1):
# print("new time step")
# print("#" * 50)
# Only initialize variables in the first step
if time_step > 1: scope.reuse_variables()
# Find the groups of the hidden layer that are active
group_index = 0
notmod = []
mod = []
for i in range(len(self.clockwork_periods)):
# Check if (t MOD T_i == 0)
if time_step % self.clockwork_periods[i] == 0:
# print("time step : " + str(time_step))
# counter = i+1
# print("current time step divided by period in index :" + str(i))
mod.append(i) # store period indcies to update
group_index = (i + 1) * group_size
# print("group size: " + str(group_size))
# print("group index " + str(group_index))
# print("not mod: " + str(notmod))
# print("mod: " + str(mod))
# Compute (W_I*x_t + b_I)
WI_x = tf.matmul(x_list[time_step-1], tf.slice(self.input_W, [0, 0], [-1, group_index]))
WI_x = tf.nn.bias_add(WI_x, tf.slice(self.input_b, [0], [group_index]), name="WI_x")
# Compute (W_H*y_{t-1} + b_H), note the multiplication of the clockwork mask (upper triangular matrix)
self.hidden_W = tf.multiply(self.hidden_W, self.clockwork_mask)
WH_y = tf.matmul(self.state, tf.slice(self.hidden_W, [0, 0], [-1, group_index]))
WH_y = tf.nn.bias_add(WH_y, tf.slice(self.hidden_b, [0], [group_index]), name="WH_y")
# Compute y_t = (...) and update the cell state
y_update = tf.add(WH_y, WI_x, name="state_update")
y_update = activation_hidden(y_update)
# print("y_update")
# print(y_update.shape)
temp = []
for i in range(len(self.config.periods)):
dx = i * group_size
if i in mod:
temp.append(tf.slice(y_update, [0, dx], [-1, group_size]))
else:
temp.append(tf.slice(self.state, [0, dx], [-1, group_size]))
# Copy the updates to the cell state (original)
# self.state = tf.concat(
# axis=1, values=[y_update, tf.slice(self.state, [0, group_index], [-1,-1])])
self.state = tf.concat(axis=1, values=temp) # for saving state
# Save the final hidden state
self.final_state = self.state
# Compute the output, y = f(W_O*y_t + b_O) (full prediction)
self.predictions = tf.matmul(self.final_state, self.output_W)
self.predictions = tf.nn.bias_add(self.predictions, self.output_b, name="predictions")
# Compute the loss (original)
# self.error = tf.reduce_sum(tf.square(self.targets - self.predictions), axis=1)
# self.loss = tf.reduce_mean(self.error, name="loss")
# other loss funcs
self.loss = tf.losses.mean_squared_error(self.targets, self.predictions, weights=1.0, scope=None)
def _init_optimizer(self):
# Learning rate decay, note that is self.learning_rate_decay == 1.0,
# the decay schedule is disabled, i.e. learning rate is constant.
self.learning_rate = tf.train.exponential_decay(
self.config.learning_rate,
self.global_step,
self.config.learning_rate_step,
self.config.learning_rate_decay,
staircase=True
)
self.learning_rate = tf.maximum(self.learning_rate, self.config.learning_rate_min)
tf.summary.scalar("learning_rate", self.learning_rate)
# Definition of the optimizer and computing gradients operation
if self.config.optimizer == 'adam':
# Adam optimizer
self.optimizer = tf.train.AdamOptimizer(learning_rate=self.learning_rate)
elif self.config.optimizer == 'rmsprop':
# RMSProper optimizer
self.optimizer = tf.train.RMSPropOptimizer(learning_rate=self.learning_rate)
elif self.config.optimizer == 'adagrad':
# AdaGrad optimizer
self.optimizer = tf.train.AdagradOptimizer(learning_rate=self.learning_rate)
else:
raise ValueError("Unknown optimizer specified")
# Compute the gradients for each variable
self.grads_and_vars = self.optimizer.compute_gradients(self.loss)
# Optionally perform gradient clipping by max-norm
if self.config.max_norm_gradient > 0:
# Perform gradient clipping by the global norm
grads, variables = zip(*self.grads_and_vars)
grads_clipped, _ = tf.clip_by_global_norm(
grads, clip_norm=self.config.max_norm_gradient)
# Apply the gradients after clipping them
self.train_op = self.optimizer.apply_gradients(
zip(grads_clipped, variables),
global_step=self.global_step
)
else:
# Unclipped gradients
self.train_op = self.optimizer.apply_gradients(
self.grads_and_vars,
global_step=self.global_step
)
# Keep track of gradient values and their sparsity
grad_summaries = []
for g, v in self.grads_and_vars:
if g is not None:
grad_hist_summary = tf.summary.histogram("gradients/{}/hist".format(v.name), g)
sparsity_summary = tf.summary.scalar("gradients/{}/sparsity".format(v.name), tf.nn.zero_fraction(g))
grad_summaries.append(grad_hist_summary)
grad_summaries.append(sparsity_summary)
self.gradient_summaries_merged = tf.summary.merge(grad_summaries)
def _build_summary_ops(self):
# Training summaries
training_summaries = [
tf.summary.scalar("train/loss", self.loss),
tf.summary.scalar("train/learning_rate", self.learning_rate),
]
# Combine the training summaries with the gradient summaries
self.train_summary_op = tf.summary.merge(
[training_summaries, self.gradient_summaries_merged]) |
#!/usr/bin/python
import csv
import sys
import json
import logging
from util import django_utils
from optparse import OptionParser
django_utils.SetupDjango()
from gibbs import models
from gibbs import constants
# Column names
KEGG_ID = '!MiriamID::urn:miriam:kegg.compound'
NAME = '!Name'
INCHI = '!InChI'
SOURCE = '!Source'
FORMATION = '!FormationEnergy'
ROW_ORDER = [NAME, KEGG_ID, INCHI, FORMATION, SOURCE]
def GenFormationEnergyData(pH=constants.DEFAULT_PH,
ionic_strength=constants.DEFAULT_IONIC_STRENGTH):
"""Returns a list of dictionaries of compound formation energies.
TODO(flamholz): return data from multiple sources per compound when possible.
Args:
pH: the pH.
ionic_strength: the ionic strength.
"""
dicts = []
for compound in models.Compound.objects.all():
dG = compound.DeltaG0Prime(pH=pH, ionic_strength=ionic_strength)
if dG:
dG = round(dG, 3)
if dG is not None and not compound.dg_source:
logging.error('%s has a dg but no source' % compound)
d = {KEGG_ID: compound.kegg_id,
NAME: compound.FirstName(),
FORMATION: dG,
INCHI: compound.inchi,
SOURCE: None}
if dG is not None:
d[SOURCE] = compound.dg_source.name
dicts.append(d)
return dicts
def MakeOpts():
"""Returns an OptionParser object with all the default options."""
opt_parser = OptionParser()
opt_parser.add_option("-p", "--ph", dest="pH", type="float",
default=constants.DEFAULT_PH, help="The pH")
opt_parser.add_option("-i", "--ionic_strength", dest="ionic_strength",
type="float", default=constants.DEFAULT_IONIC_STRENGTH,
help="The ionic strength, M")
opt_parser.add_option("-o", "--output_name", dest="output_name",
type="string", default="compound_formation_energies",
help="The name of the file to write csv output to.")
return opt_parser
def WriteHeader(dict_writer, row_order=ROW_ORDER):
"""writeheader() is new in Python 2.7"""
if hasattr(dict_writer, 'writeheader'):
dict_writer.writeheader()
else:
dict_writer.writer.writerow(ROW_ORDER)
def main():
options, _ = MakeOpts().parse_args(sys.argv)
print 'Using pH = %.2f and ionic strength = %.3f' % (options.pH,
options.ionic_strength)
output_name = '%s_pH%.2f_is%.3f' % (options.output_name, options.pH,
options.ionic_strength)
output_tsv_name = output_name + '.tsv'
output_json_name = output_name + '.json'
print 'Will write tsv output to %s' % output_tsv_name
print 'Will write json output to %s' % output_json_name
dicts = GenFormationEnergyData(pH=options.pH,
ionic_strength=options.ionic_strength)
sorted_data = sorted(dicts, key=lambda x: (x[KEGG_ID], x[SOURCE]))
csv_file = open(output_tsv_name, 'w')
writer = csv.DictWriter(csv_file, ROW_ORDER, dialect=csv.excel_tab)
WriteHeader(writer, ROW_ORDER)
writer.writerows(sorted_data)
csv_file.close()
json_file = open(output_json_name, 'w')
json.dump(sorted_data, json_file, sort_keys=True, indent=4)
json_file.close()
print 'Done.'
if __name__ == '__main__':
main()
|
import torch
import torch.nn.functional as F
import numpy as np
from utils.proxy import proxy_reward
def test(args, policy_net, env):
device = next(policy_net.parameters()).device
width, height = 84, 84
num_ales = args.evaluation_episodes
if args.use_openai_test_env:
observation = torch.from_numpy(env.reset()).squeeze(1)
else:
observation = env.reset(initial_steps=50).squeeze(-1)
lengths = torch.zeros(num_ales, dtype=torch.int32)
rewards = torch.zeros(num_ales, dtype=torch.float32)
true_rewards = torch.zeros(num_ales, dtype=torch.float32)
all_done = torch.zeros(num_ales, dtype=torch.bool)
not_done = torch.ones(num_ales, dtype=torch.bool)
fire_reset = torch.zeros(num_ales, dtype=torch.bool)
actions = torch.ones(num_ales, dtype=torch.uint8)
maybe_npy = lambda a: a.numpy() if args.use_openai_test_env else a
info = env.step(maybe_npy(actions))[-1]
if args.use_openai_test_env:
lives = torch.IntTensor([d['ale.lives'] for d in info])
else:
lives = info['ale.lives'].clone()
states = torch.zeros((num_ales, args.num_stack, width, height), device=device, dtype=torch.float32)
states[:, -1] = observation.to(device=device, dtype=torch.float32)
policy_net.eval()
while not all_done.all():
logit = policy_net(states)[1]
actions = F.softmax(logit, dim=1).multinomial(1).cpu()
actions[fire_reset] = 1
cached_ram = env.ram.to(device=device, dtype=torch.uint8)
observation, reward, done, info = env.step(maybe_npy(actions))
ram = env.ram.to(device=device, dtype=torch.uint8)
if args.use_openai_test_env:
# convert back to pytorch tensors
observation = torch.from_numpy(observation)
reward = torch.from_numpy(reward.astype(np.float32))
done = torch.from_numpy(done.astype(np.bool))
new_lives = torch.IntTensor([d['ale.lives'] for d in info])
else:
new_lives = info['ale.lives'].clone()
true_reward = reward.detach().clone()
reward = proxy_reward(
reward,
None,
ram,
cached_ram,
diver_bonus=args.diver_bonus,
prox_bonus=args.proximity_bonus,
o2_pen=args.o2_penalty,
lives_pen=args.lives_penalty,
bullet_pen=args.bullet_penalty,
space_reward=args.space_reward)
fire_reset = new_lives < lives
lives.copy_(new_lives)
observation = observation.to(device=device, dtype=torch.float32)
states[:, :-1].copy_(states[:, 1:].clone())
states *= (1.0 - done.to(device=device, dtype=torch.float32)).view(-1, *[1] * (observation.dim() - 1))
states[:, -1].copy_(observation.view(-1, *states.size()[-2:]))
# update episodic reward counters
lengths += not_done.int()
rewards += reward.cpu() * not_done.float().cpu()
true_rewards += true_reward.cpu() * not_done.float().cpu()
all_done |= done.cpu()
all_done |= (lengths >= args.max_episode_length)
not_done = (all_done == False).int()
policy_net.train()
return lengths, rewards, true_rewards
|
<filename>zbuilder.py<gh_stars>1-10
#!/usr/bin/python
import argparse
import docker
import json
import logging
import os
import shutil
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s: %(message)s', level=logging.INFO)
class zbuilder():
def __init__(self, config):
js = json.load(config)
self.docker_files = []
self.build_succeeded_file = "/tmp/build_succeeded"
packages = js.get("packages")
if not packages:
logging.error("core: there is no 'packages' object, nothing to build")
return
logging.info("Starting parse different build types")
for package_type, package in packages.items():
images = []
if package_type == "deb":
img = js.get("deb-images")
if img:
images += img
elif package_type == "rpm":
img = js.get("rpm-images")
if img:
images += img
else:
logging.error("%s: unsupported package type", package_type)
continue
logging.info("%s: starting to parse commands", package_type)
pre_build_commands = package.get("pre-build-commands")
build_commands = package.get("build-commands")
if build_commands:
build_commands.append("echo success > %s" % (self.build_succeeded_file))
post_build = package.get("post-build-commands")
final_commands = {}
if post_build:
pbs = post_build.get("success")
if pbs:
final_commands["success"] = pbs
pbf = post_build.get("fail")
if pbf:
final_commands["fail"] = pbf
pba = post_build.get("always")
if pba:
final_commands["always"] = pba
sources = package.get("sources")
if not sources:
logging.error("%s: there is no 'sources' object, nothing to build", package_type)
break
for name, source in sources.items():
logging.info("%s/%s: starting to parse source", package_type, name)
include_images = source.get("include-images")
if include_images:
images += include_images
exclude_images = source.get("exclude-images")
if exclude_images:
tmp = []
for x in images:
if x in exclude_images:
continue
tmp.append(x)
images = tmp
logging.info("%s/%s: images: %s", package_type, name, ', '.join(images))
fetch_commands = []
try:
stype = source["type"]
repo = source["repository"]
branch = source.get("branch", "master")
if stype == "git":
fetch_commands.append("rm -rf %s" % (name))
fetch_commands.append("git clone %s %s" % (repo, name))
fetch_commands.append("cd %s" % (name))
fetch_commands.append("git checkout %s" % (branch))
build_commands.append("cd %s" % (name))
else:
logging.error("%s/%s: unsupported source type '%s'", package_type, name, stype)
continue
except Exception as e:
logging.error("%s/%s: invalid source: %s", package_type, name, e)
continue
logging.info("%s/%s: fetch commands: %s", package_type, name, ', '.join(fetch_commands))
commands = []
try:
commands.append(pre_build_commands)
commands.append(fetch_commands)
commands.append(build_commands)
except Exception as e:
logging.notice("%s/%s: could not append command: %s", package_type, name, e)
for image in images:
df = self.generate_dockerfile(name, image, commands, final_commands)
self.docker_files.append(df)
def generate_dockerfile(self, name, image, commands, final_commands):
df = "Dockerfile.%s.%s" % (name, image)
with open(df, 'w+') as f:
f.write("FROM %s\n" % (image))
f.write("ENV ZBUILDER_IMAGE=%s ZBUILDER_NAME=%s DEBIAN_FRONTEND=noninteractive\n" % (image, name))
f.write("ADD conf.d conf.d\n")
for cmd_set in commands:
cs = "RUN %s\n" % (' && \\\n'.join(cmd_set))
f.write(cs)
success = final_commands.get("success")
if success:
cs = "RUN test -f %s && \\\n %s\n" % (self.build_succeeded_file, ' && \\\n'.join(success))
f.write(cs)
fail = final_commands.get("fail")
if fail:
cs = "RUN test -f %s || \\\n %s\n" % (self.build_succeeded_file, ' && \\\n'.join(fail))
f.write(cs)
always = final_commands.get("always")
if always:
cs = "RUN %s\n" % ' && \\\n'.join(always)
f.write(cs)
return df
def run(self, name = None, build_dir = '.'):
c = docker.Client(base_url='unix://var/run/docker.sock')
for path in self.docker_files:
if name and not name in path:
continue
try:
shutil.rmtree(path="%s/" % build_dir, ignore_errors=True)
os.mkdir("%s/" % build_dir)
shutil.copy(path, "%s/" % build_dir)
shutil.copytree("conf.d", "%s/conf.d" % build_dir)
except Exception as e:
logging.error("Could not copy local content to destination build dir %s: %s",
build_dir, e)
continue
with open("%s.build.log" % (path), "w+") as out:
response = c.build(path=build_dir, dockerfile=path, rm=False, pull=False, forcerm=False)
for r in response:
out.write(r)
logging.info("%s: %s", path, r)
if __name__ == '__main__':
bparser = argparse.ArgumentParser(description='Builder arguments.', add_help=True)
bparser.add_argument("--conf", dest='conf', action='store', type=argparse.FileType('r'),
required=True, help='Input config file.')
bparser.add_argument("--build-dir", dest='build_dir', action='store', default=".",
help='Local directory where build process will run.')
bparser.add_argument("--image", dest='image', action='store',
help='Build only images containing this substring.')
args = bparser.parse_args()
try:
zb = zbuilder(config=args.conf)
try:
zb.run(name=args.image, build_dir=args.build_dir)
except Exception as e:
logging.error("Could not run build, name: %s: %s", args.image, e)
except Exception as e:
logging.error("Could not create zbuilder object: %s", e)
|
<gh_stars>0
# (c) 2017, Red Hat, inc
#
# This file is part of Ansible
#
# Ansible is free software: you can redistribute it and/or modify
# it under the terms of the GNU General Public License as published by
# the Free Software Foundation, either version 3 of the License, or
# (at your option) any later version.
#
# Ansible is distributed in the hope that it will be useful,
# but WITHOUT ANY WARRANTY; without even the implied warranty of
# MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# GNU General Public License for more details.
#
# You should have received a copy of the GNU General Public License
# along with Ansible. If not, see <https://www.gnu.org/licenses/>.
# Make coding more python3-ish
from __future__ import (absolute_import, division, print_function)
__metaclass__ = type
import hashlib
import os
import re
import string
from collections import Mapping
from ansible.errors import AnsibleError, AnsibleParserError
from ansible.plugins import AnsiblePlugin
from ansible.plugins.cache import InventoryFileCacheModule
from ansible.module_utils._text import to_bytes, to_native
from ansible.module_utils.parsing.convert_bool import boolean
from ansible.module_utils.six import string_types
from ansible.template import Templar
try:
from __main__ import display
except ImportError:
from ansible.utils.display import Display
display = Display()
_SAFE_GROUP = re.compile("[^A-Za-z0-9_]")
# Helper methods
def to_safe_group_name(name):
''' Converts 'bad' characters in a string to underscores so they can be used as Ansible hosts or groups '''
return _SAFE_GROUP.sub("_", name)
def detect_range(line=None):
'''
A helper function that checks a given host line to see if it contains
a range pattern described in the docstring above.
Returns True if the given line contains a pattern, else False.
'''
return '[' in line
def expand_hostname_range(line=None):
'''
A helper function that expands a given line that contains a pattern
specified in top docstring, and returns a list that consists of the
expanded version.
The '[' and ']' characters are used to maintain the pseudo-code
appearance. They are replaced in this function with '|' to ease
string splitting.
References: https://docs.ansible.com/ansible/latest/user_guide/intro_inventory.html#hosts-and-groups
'''
all_hosts = []
if line:
# A hostname such as db[1:6]-node is considered to consists
# three parts:
# head: 'db'
# nrange: [1:6]; range() is a built-in. Can't use the name
# tail: '-node'
# Add support for multiple ranges in a host so:
# db[01:10:3]node-[01:10]
# - to do this we split off at the first [...] set, getting the list
# of hosts and then repeat until none left.
# - also add an optional third parameter which contains the step. (Default: 1)
# so range can be [01:10:2] -> 01 03 05 07 09
(head, nrange, tail) = line.replace('[', '|', 1).replace(']', '|', 1).split('|')
bounds = nrange.split(":")
if len(bounds) != 2 and len(bounds) != 3:
raise AnsibleError("host range must be begin:end or begin:end:step")
beg = bounds[0]
end = bounds[1]
if len(bounds) == 2:
step = 1
else:
step = bounds[2]
if not beg:
beg = "0"
if not end:
raise AnsibleError("host range must specify end value")
if beg[0] == '0' and len(beg) > 1:
rlen = len(beg) # range length formatting hint
if rlen != len(end):
raise AnsibleError("host range must specify equal-length begin and end formats")
def fill(x):
return str(x).zfill(rlen) # range sequence
else:
fill = str
try:
i_beg = string.ascii_letters.index(beg)
i_end = string.ascii_letters.index(end)
if i_beg > i_end:
raise AnsibleError("host range must have begin <= end")
seq = list(string.ascii_letters[i_beg:i_end + 1:int(step)])
except ValueError: # not an alpha range
seq = range(int(beg), int(end) + 1, int(step))
for rseq in seq:
hname = ''.join((head, fill(rseq), tail))
if detect_range(hname):
all_hosts.extend(expand_hostname_range(hname))
else:
all_hosts.append(hname)
return all_hosts
class BaseInventoryPlugin(AnsiblePlugin):
""" Parses an Inventory Source"""
TYPE = 'generator'
def __init__(self):
super(BaseInventoryPlugin, self).__init__()
self._options = {}
self.inventory = None
self.display = display
self.cache = None
def parse(self, inventory, loader, path, cache=True):
''' Populates inventory from the given data. Raises an error on any parse failure
:arg inventory: a copy of the previously accumulated inventory data,
to be updated with any new data this plugin provides.
The inventory can be empty if no other source/plugin ran successfully.
:arg loader: a reference to the DataLoader, which can read in YAML and JSON files,
it also has Vault support to automatically decrypt files.
:arg path: the string that represents the 'inventory source',
normally a path to a configuration file for this inventory,
but it can also be a raw string for this plugin to consume
:arg cache: a boolean that indicates if the plugin should use the cache or not
you can ignore if this plugin does not implement caching.
'''
self.loader = loader
self.inventory = inventory
self.templar = Templar(loader=loader)
def verify_file(self, path):
''' Verify if file is usable by this plugin, base does minimal accessibility check
:arg path: a string that was passed as an inventory source,
it normally is a path to a config file, but this is not a requirement,
it can also be parsed itself as the inventory data to process.
So only call this base class if you expect it to be a file.
'''
b_path = to_bytes(path, errors='surrogate_or_strict')
return (os.path.exists(b_path) and os.access(b_path, os.R_OK))
def _populate_host_vars(self, hosts, variables, group=None, port=None):
if not isinstance(variables, Mapping):
raise AnsibleParserError("Invalid data from file, expected dictionary and got:\n\n%s" % to_native(variables))
for host in hosts:
self.inventory.add_host(host, group=group, port=port)
for k in variables:
self.inventory.set_variable(host, k, variables[k])
def _read_config_data(self, path):
''' validate config and set options as appropriate
:arg path: path to common yaml format config file for this plugin
'''
config = {}
try:
# avoid loader cache so meta: refresh_inventory can pick up config changes
# if we read more than once, fs cache should be good enough
config = self.loader.load_from_file(path, cache=False)
except Exception as e:
raise AnsibleParserError(to_native(e))
if not config:
# no data
raise AnsibleParserError("%s is empty" % (to_native(path)))
elif config.get('plugin') != self.NAME:
# this is not my config file
raise AnsibleParserError("Incorrect plugin name in file: %s" % config.get('plugin', 'none found'))
elif not isinstance(config, Mapping):
# configs are dictionaries
raise AnsibleParserError('inventory source has invalid structure, it should be a dictionary, got: %s' % type(config))
self.set_options(direct=config)
if self._options.get('cache'):
self._set_cache_options(self._options)
return config
def _set_cache_options(self, options):
self.cache = InventoryFileCacheModule(plugin_name=options.get('cache_plugin'),
timeout=options.get('cache_timeout'),
cache_dir=options.get('cache_connection'))
def _consume_options(self, data):
''' update existing options from alternate configuration sources not normally used by Ansible.
Many API libraries already have existing configuration sources, this allows plugin author to leverage them.
:arg data: key/value pairs that correspond to configuration options for this plugin
'''
for k in self._options:
if k in data:
self._options[k] = data.pop(k)
def clear_cache(self):
pass
class BaseFileInventoryPlugin(BaseInventoryPlugin):
""" Parses a File based Inventory Source"""
TYPE = 'storage'
def __init__(self):
super(BaseFileInventoryPlugin, self).__init__()
class Cacheable(object):
_cache = {}
def get_cache_key(self, path):
return "{0}_{1}_{2}".format(self.NAME, self._get_cache_prefix(path), self._get_config_identifier(path))
def _get_cache_prefix(self, path):
''' create predictable unique prefix for plugin/inventory '''
m = hashlib.sha1()
m.update(to_bytes(self.NAME, errors='surrogate_or_strict'))
d1 = m.hexdigest()
n = hashlib.sha1()
n.update(to_bytes(path, errors='surrogate_or_strict'))
d2 = n.hexdigest()
return 's_'.join([d1[:5], d2[:5]])
def _get_config_identifier(self, path):
''' create predictable config-specific prefix for plugin/inventory '''
return hashlib.md5(path.encode()).hexdigest()
def clear_cache(self):
self._cache = {}
class Constructable(object):
def _compose(self, template, variables):
''' helper method for plugins to compose variables for Ansible based on jinja2 expression and inventory vars'''
t = self.templar
t.set_available_variables(variables)
return t.template('%s%s%s' % (t.environment.variable_start_string, template, t.environment.variable_end_string), disable_lookups=True)
def _set_composite_vars(self, compose, variables, host, strict=False):
''' loops over compose entries to create vars for hosts '''
if compose and isinstance(compose, dict):
for varname in compose:
try:
composite = self._compose(compose[varname], variables)
except Exception as e:
if strict:
raise AnsibleError("Could not set %s for host %s: %s" % (varname, host, to_native(e)))
continue
self.inventory.set_variable(host, varname, composite)
def _add_host_to_composed_groups(self, groups, variables, host, strict=False):
''' helper to create complex groups for plugins based on jinja2 conditionals, hosts that meet the conditional are added to group'''
# process each 'group entry'
if groups and isinstance(groups, dict):
self.templar.set_available_variables(variables)
for group_name in groups:
conditional = "{%% if %s %%} True {%% else %%} False {%% endif %%}" % groups[group_name]
try:
result = boolean(self.templar.template(conditional))
except Exception as e:
if strict:
raise AnsibleParserError("Could not add host %s to group %s: %s" % (host, group_name, to_native(e)))
continue
if result:
# ensure group exists
self.inventory.add_group(group_name)
# add host to group
self.inventory.add_child(group_name, host)
def _add_host_to_keyed_groups(self, keys, variables, host, strict=False):
''' helper to create groups for plugins based on variable values and add the corresponding hosts to it'''
if keys and isinstance(keys, list):
groups = []
for keyed in keys:
if keyed and isinstance(keyed, dict):
try:
key = self._compose(keyed.get('key'), variables)
except Exception as e:
if strict:
raise AnsibleParserError("Could not generate group from %s entry: %s" % (keyed.get('key'), to_native(e)))
continue
if key:
prefix = keyed.get('prefix', '')
sep = keyed.get('separator', '_')
if isinstance(key, string_types):
groups.append('%s%s%s' % (prefix, sep, key))
elif isinstance(key, list):
for name in key:
groups.append('%s%s%s' % (prefix, sep, name))
elif isinstance(key, Mapping):
for (gname, gval) in key.items():
name = '%s%s%s' % (gname, sep, gval)
groups.append('%s%s%s' % (prefix, sep, name))
else:
raise AnsibleParserError("Invalid group name format, expected a string or a list of them or dictionary, got: %s" % type(key))
else:
if strict:
raise AnsibleParserError("No key or key resulted empty, invalid entry")
else:
raise AnsibleParserError("Invalid keyed group entry, it must be a dictionary: %s " % keyed)
# now actually add any groups
for group_name in groups:
gname = to_safe_group_name(group_name)
self.inventory.add_group(gname)
self.inventory.add_child(gname, host)
|
# -*- coding: iso-8859-1 -*-
"""
MoinMoin - RenamePage action
This action allows you to rename a page.
@copyright: 2002-2004 <NAME> <<EMAIL>>,
2006-2007 MoinMoin:ThomasWaldmann,
2007 MoinMoin:ReimarBauer
@license: GNU GPL, see COPYING for details.
"""
import re
from MoinMoin import wikiutil
from MoinMoin.Page import Page
from MoinMoin.PageEditor import PageEditor
from MoinMoin.action import ActionBase
class RenamePage(ActionBase):
""" Rename page action
Note: the action name is the class name
"""
def __init__(self, pagename, request):
ActionBase.__init__(self, pagename, request)
self.use_ticket = True
_ = self._
self.form_trigger = 'rename'
self.form_trigger_label = _('Rename Page')
filterfn = re.compile(ur"^%s/.*$" % re.escape(pagename), re.U).match
subpagenames = request.rootpage.getPageList(user='', exists=1, filter=filterfn)
self.subpages = [pagename for pagename in subpagenames if self.request.user.may.delete(pagename)]
try:
self.show_redirect = request.cfg.show_rename_redirect
except AttributeError:
self.show_redirect = False
try:
self.rename_redirect = int(self.request.form.get('rename_redirect', '0'))
except ValueError:
self.rename_redirect = 0
def is_allowed(self):
may = self.request.user.may
return may.write(self.pagename) and may.delete(self.pagename)
def check_condition(self):
_ = self._
if not self.page.exists():
return _('This page is already deleted or was never created!')
else:
return None
def do_action(self):
""" Rename this page to "pagename" """
_ = self._
form = self.form
newpagename = form.get('newpagename', u'')
newpagename = wikiutil.normalize_pagename(newpagename, self.cfg)
comment = form.get('comment', u'')
comment = wikiutil.clean_input(comment)
try:
rename_subpages = int(self.request.form.get('rename_subpages', '0'))
except ValueError:
rename_subpages = 0
self.page = PageEditor(self.request, self.pagename)
success, msgs = self.page.renamePage(newpagename, comment)
if not success:
return success, msgs
msgs = [msgs]
if self.show_redirect and self.rename_redirect:
self.page = PageEditor(self.request, self.pagename)
self.page.saveText('#redirect %s' % newpagename, 0)
if rename_subpages and self.subpages:
for name in self.subpages:
self.page = PageEditor(self.request, name)
new_subpagename = name.replace(self.pagename, newpagename, 1)
success_i, msg = self.page.renamePage(new_subpagename, comment)
msgs.append(msg)
if self.show_redirect and self.rename_redirect and success_i:
self.page = PageEditor(self.request, name)
self.page.saveText('#redirect %s' % new_subpagename, 0)
msgs = ' '.join([msg for msg in msgs if msg])
self.newpagename = newpagename # keep there for finish
return success, msgs
def do_action_finish(self, success):
if success:
url = Page(self.request, self.newpagename).url(self.request)
self.request.http_redirect(url, code=301)
else:
self.render_msg(self.make_form(), "dialog")
def get_form_html(self, buttons_html):
_ = self._
if self.subpages:
redirect_label = _('Create redirect for renamed page(s)?')
subpages = ' '.join([wikiutil.escape(page) for page in self.subpages])
subpages_html = """
<tr>
<dd>
%(subpage_label)s<input type="checkbox" name="rename_subpages" value="1" %(subpages_checked)s>
</dd>
<dd>
<class="label"><subpage> %(subpage)s</subpage>
</dd>
</tr>
""" % {
'subpage': subpages,
'subpages_checked': ('', 'checked')[self.request.args.get('subpages_checked', '0') == '1'],
'subpage_label': _('Rename all /subpages too?'),
}
else:
redirect_label = _('Create redirect for renamed page?')
subpages_html = ""
if self.show_redirect:
redirect_html = '<tr><dd>%(redirect_label)s<input type="checkbox" name="rename_redirect" value="1" %(redirect)s></dd></tr>' % {
'redirect': self.rename_redirect,
'redirect_label': redirect_label,
}
else:
redirect_html = ''
if self.show_redirect or self.subpages:
options_html = """
<table>
%(subpages_html)s
%(redirect_html)s
</table>
""" % {
"subpages_html": subpages_html,
"redirect_html": redirect_html,
}
else:
options_html = ""
d = {
'querytext': _('Really rename this page?'),
'pagename': wikiutil.escape(self.pagename, True),
'newname_label': _("New name"),
'comment_label': _("Optional reason for the renaming"),
'buttons_html': buttons_html,
'options_html': options_html,
}
return '''
<strong>%(querytext)s</strong>
<br>
<br>
%(options_html)s
<table>
<tr>
<td class="label"><label>%(newname_label)s</label></td>
<td class="content">
<input type="text" name="newpagename" value="%(pagename)s" size="80">
</td>
</tr>
<tr>
<td class="label"><label>%(comment_label)s</label></td>
<td class="content">
<input type="text" name="comment" size="80" maxlength="200">
</td>
</tr>
<tr>
<td></td>
<td class="buttons">
%(buttons_html)s
</td>
</tr>
</table>
''' % d
def execute(pagename, request):
""" Glue code for actions """
RenamePage(pagename, request).render()
|
import sys
import os
import threading
import multiprocessing
import multiprocessing.pool
import traceback
from . import logger
class SafeProcess(
multiprocessing.Process,
):
def __init__(
self,
*args,
**kwargs
):
super().__init__(
*args,
**kwargs
)
pipe = multiprocessing.Pipe()
self._parent_pipe = pipe[0]
self._child_pipe = pipe[1]
self._exception = None
def run(
self,
):
try:
multiprocessing.Process.run(
self,
)
self._child_pipe.send(None)
except Exception as exception:
self._child_pipe.send(
{
'exception': exception,
'traceback': traceback.format_exc(),
}
)
@property
def exception(
self,
):
if self._parent_pipe.poll():
self._exception = self._parent_pipe.recv()
return self._exception
class Supervisor:
def __init__(
self,
worker_class,
concurrent_workers,
):
self.logger = logger.logger.Logger(
logger_name='Supervisor',
)
self.worker_class = worker_class
self.concurrent_workers = concurrent_workers
self.task = self.worker_class()
self.workers_processes = []
self.should_work_event = threading.Event()
self.should_work_event.set()
multiprocessing.set_start_method(
method='spawn',
force=True,
)
def worker_watchdog(
self,
function,
):
process = None
while self.should_work_event.is_set():
try:
process = SafeProcess(
target=function,
kwargs={},
)
process.start()
self.workers_processes.append(process)
process.join(
timeout=self.task.config['timeouts']['global_timeout'] or None,
)
if process.exception:
self.logger.critical(
msg='supervisor has thrown an exception',
extra={
'exception': {
'type': process.exception['exception'].__class__.__name__,
'message': str(process.exception['exception']),
},
'traceback': process.exception['traceback'],
'additional': dict(),
},
)
except Exception as exception:
self.logger.critical(
msg='supervisor has thrown an exception',
extra={
'exception': {
'type': exception.__class__.__name__,
'message': str(exception),
},
'traceback': traceback.format_exc(),
'additional': dict(),
},
)
finally:
if process:
process.terminate()
try:
os.waitpid(
process.pid,
0,
)
except ChildProcessError:
pass
self.workers_processes.remove(process)
def start(
self,
):
threads = []
for i in range(self.concurrent_workers):
thread = threading.Thread(
target=self.worker_watchdog,
kwargs={
'function': self.task.work_loop,
},
)
thread.daemon = True
thread.start()
threads.append(thread)
try:
for thread in threads:
thread.join()
except KeyboardInterrupt:
pass
finally:
self.should_work_event.clear()
for worker_process in self.workers_processes:
worker_process.terminate()
sys.exit(0)
def __getstate__(
self,
):
state = {
'worker_class': self.worker_class,
'concurrent_workers': self.concurrent_workers,
}
return state
def __setstate__(
self,
value,
):
self.__init__(
worker_class=value['worker_class'],
concurrent_workers=value['concurrent_workers'],
)
|
<reponame>MrHamdulay/rsa-chat
import socket
import threading
import SocketServer
from protocol import Protocol
from time import time
protocol = Protocol()
global_lock = threading.Lock()
public_keys = {}
sockets = {}
class ServerServer(SocketServer.ThreadingMixIn, SocketServer.TCPServer):
''' Server that doesn't close requests after we handle messages.
Logs all server messages to file'''
daemon = True
def __init__(self, *args, **kwargs):
SocketServer.TCPServer.__init__(self, *args, **kwargs)
self.log_file = open('server.log', 'a')
def close_request(self, request):
pass
def shutdown_request(self, request):
pass
def log_client(self, socket, line):
line = '%s [%s]: %s\n' % (time(), socket.getpeername(), str(line))
self.log_file.write(line)
self.log_file.flush()
class ChatRequestHandler(SocketServer.BaseRequestHandler):
''' Handle client incoming messages, all we do pretty much is
distribute messages between client sockets'''
def __init__(self, request, client_address, server):
self.name = None
self.buffer = bytes()
SocketServer.BaseRequestHandler.__init__(self, request, client_address, server)
def handle_helo(self, result):
''' hello handler from clients '''
global public_keys, sockets
self.name = name = result[1]
print self.name, 'has joined'
self.version = int(result[0])
# store this clients socket
sockets[name] = self.request
public_key = result[2:]
# ensure client is using latest vesrion
if self.version != Protocol.version:
self.request.sendall(protocol.gen_mesg(
'God',
self.name,
'Your client is out of date, please update',
public_key))
return
# let's tell this person all the public keys we know
for other_name, other_public_key in public_keys.iteritems():
# why would we send someone their own keys?
if name == other_name:
continue
self.request.sendall(protocol.gen_hello(
other_name, other_public_key))
sockets[other_name].sendall(self.buffer)
public_keys[name] = public_key
def handle_mesg(self, result):
''' distribute all messages to everyone (purposefully insecure '''
global sockets
# let's broadcast to everyone (purposefully insecure)
for name, sock in sockets.iteritems():
sock.sendall(self.buffer)
def handle_disconnection(self):
''' when a client disconnects free all the resources'''
self.server.log_client(self.request, self.name+' has disconnected')
global sockets
global_lock.acquire()
if self.name in sockets:
del sockets[self.name]
if self.name in public_keys:
del public_keys[self.name]
# tell all other connected clients that this user has disconnected
for name, sock in sockets.iteritems():
if name == self.name:
continue
sock.sendall(protocol.gen_bye(self.name))
global_lock.release()
def handle_ping(self, result):
self.request.sendall('pong pong')
def on_parsed_data(self, data):
self.server.log_client(self.request, data)
global_lock.acquire()
# find the handler method for this request
if hasattr(self, 'handle_'+data[0]):
getattr(self, 'handle_'+data[0])(data[1])
else:
# we couldn't find a handler message for this type of message
print 'i do not know what', data[0], 'is'
global_lock.release()
def handle(self):
try:
while True:
self.buffer += self.request.recv(4096)
# parse raw data into nice arrays
result = protocol.parse(self.buffer)
if not result:
return
# handle this message
self.on_parsed_data(result)
self.buffer = bytes()
finally:
print self.name, 'has disconnected'
self.handle_disconnection()
def finish(self):
''' overload to prevent socketserver from closing socket on disconnect '''
pass
print 'starting server'
server = ServerServer(('0.0.0.0', 9001), ChatRequestHandler)
try:
server.serve_forever()
finally:
server.shutdown()
|
import functools
import time
from typing import List, Dict, Any, Iterable, Set, Tuple, Optional
from dbt.logger import (
GLOBAL_LOGGER as logger,
TextOnly,
HookMetadata,
UniqueID,
TimestampNamed,
DbtModelState,
)
from dbt.exceptions import InternalException
from dbt.node_types import NodeType, RunHookType
from dbt.node_runners import ModelRunner
import dbt.exceptions
import dbt.flags
from dbt.hooks import get_hook
from dbt.ui.printer import \
print_hook_start_line, \
print_hook_end_line, \
print_timestamped_line, \
print_run_end_messages, \
get_counts
from dbt.compilation import compile_node
from dbt.contracts.graph.compiled import CompileResultNode
from dbt.contracts.graph.parsed import ParsedHookNode
from dbt.task.compile import CompileTask
class Timer:
def __init__(self):
self.start = None
self.end = None
@property
def elapsed(self):
if self.start is None or self.end is None:
return None
return self.end - self.start
def __enter__(self):
self.start = time.time()
return self
def __exit__(self, exc_type, exc_value, exc_tracebck):
self.end = time.time()
@functools.total_ordering
class BiggestName(str):
def __lt__(self, other):
return True
def __eq__(self, other):
return isinstance(other, self.__class__)
def _hook_list() -> List[ParsedHookNode]:
return []
def get_hooks_by_tags(
nodes: Iterable[CompileResultNode],
match_tags: Set[str],
) -> List[ParsedHookNode]:
matched_nodes = []
for node in nodes:
if not isinstance(node, ParsedHookNode):
continue
node_tags = node.tags
if len(set(node_tags) & match_tags):
matched_nodes.append(node)
return matched_nodes
class RunTask(CompileTask):
def __init__(self, args, config):
super().__init__(args, config)
self.ran_hooks = []
self._total_executed = 0
def index_offset(self, value: int) -> int:
return self._total_executed + value
def raise_on_first_error(self):
return False
def populate_adapter_cache(self, adapter):
adapter.set_relations_cache(self.manifest)
def get_hook_sql(self, adapter, hook, idx, num_hooks, extra_context):
compiled = compile_node(adapter, self.config, hook, self.manifest,
extra_context)
statement = compiled.injected_sql
hook_index = hook.index or num_hooks
hook_obj = get_hook(statement, index=hook_index)
return hook_obj.sql or ''
def _hook_keyfunc(self, hook: ParsedHookNode) -> Tuple[str, Optional[int]]:
package_name = hook.package_name
if package_name == self.config.project_name:
package_name = BiggestName('')
return package_name, hook.index
def get_hooks_by_type(
self, hook_type: RunHookType
) -> List[ParsedHookNode]:
if self.manifest is None:
raise InternalException(
'self.manifest was None in get_hooks_by_type'
)
nodes = self.manifest.nodes.values()
# find all hooks defined in the manifest (could be multiple projects)
hooks: List[ParsedHookNode] = get_hooks_by_tags(nodes, {hook_type})
hooks.sort(key=self._hook_keyfunc)
return hooks
def run_hooks(self, adapter, hook_type: RunHookType, extra_context):
ordered_hooks = self.get_hooks_by_type(hook_type)
# on-run-* hooks should run outside of a transaction. This happens
# b/c psycopg2 automatically begins a transaction when a connection
# is created.
adapter.clear_transaction()
if not ordered_hooks:
return
num_hooks = len(ordered_hooks)
plural = 'hook' if num_hooks == 1 else 'hooks'
with TextOnly():
print_timestamped_line("")
print_timestamped_line(
'Running {} {} {}'.format(num_hooks, hook_type, plural)
)
startctx = TimestampNamed('node_started_at')
finishctx = TimestampNamed('node_finished_at')
for idx, hook in enumerate(ordered_hooks, start=1):
sql = self.get_hook_sql(adapter, hook, idx, num_hooks,
extra_context)
hook_text = '{}.{}.{}'.format(hook.package_name, hook_type,
hook.index)
hook_meta_ctx = HookMetadata(hook, self.index_offset(idx))
with UniqueID(hook.unique_id):
with hook_meta_ctx, startctx:
print_hook_start_line(hook_text, idx, num_hooks)
status = 'OK'
with Timer() as timer:
if len(sql.strip()) > 0:
status, _ = adapter.execute(sql, auto_begin=False,
fetch=False)
self.ran_hooks.append(hook)
with finishctx, DbtModelState({'node_status': 'passed'}):
print_hook_end_line(
hook_text, status, idx, num_hooks, timer.elapsed
)
self._total_executed += len(ordered_hooks)
with TextOnly():
print_timestamped_line("")
def safe_run_hooks(
self, adapter, hook_type: RunHookType, extra_context: Dict[str, Any]
) -> None:
try:
self.run_hooks(adapter, hook_type, extra_context)
except dbt.exceptions.RuntimeException:
logger.info("Database error while running {}".format(hook_type))
raise
def print_results_line(self, results, execution_time):
nodes = [r.node for r in results] + self.ran_hooks
stat_line = get_counts(nodes)
execution = ""
if execution_time is not None:
execution = " in {execution_time:0.2f}s".format(
execution_time=execution_time)
with TextOnly():
print_timestamped_line("")
print_timestamped_line(
"Finished running {stat_line}{execution}."
.format(stat_line=stat_line, execution=execution))
def before_run(self, adapter, selected_uids):
with adapter.connection_named('master'):
self.create_schemas(adapter, selected_uids)
self.populate_adapter_cache(adapter)
self.safe_run_hooks(adapter, RunHookType.Start, {})
def after_run(self, adapter, results):
# in on-run-end hooks, provide the value 'database_schemas', which is a
# list of unique database, schema pairs that successfully executed
# models were in. for backwards compatibility, include the old
# 'schemas', which did not include database information.
database_schema_set: Set[Tuple[Optional[str], str]] = {
(r.node.database, r.node.schema) for r in results
if not any((r.error is not None, r.fail, r.skipped))
}
self._total_executed += len(results)
extras = {
'schemas': list({s for _, s in database_schema_set}),
'results': results,
'database_schemas': list(database_schema_set),
}
with adapter.connection_named('master'):
self.safe_run_hooks(adapter, RunHookType.End, extras)
def after_hooks(self, adapter, results, elapsed):
self.print_results_line(results, elapsed)
def build_query(self):
return {
"include": self.args.models,
"exclude": self.args.exclude,
"resource_types": [NodeType.Model],
"tags": []
}
def get_runner_type(self):
return ModelRunner
def task_end_messages(self, results):
if results:
print_run_end_messages(results)
|
<filename>line_analysis_BSNIP.py
'''
TODO: Write a function to calculate the initial flux errors (to be used in the spline
weighting) by heavily smoothing the spectrum and calculating the stddev of the points
around the smoothed flux
'''
import os
from collections import namedtuple
from astropy.io import fits
from astropy.io import ascii as asc
from astropy.table import Table
from astropy.modeling import models,fitting
from astropy.convolution import convolve, Box1DKernel
from astropy.time import Time
import numpy as np
from scipy import signal, interpolate
from matplotlib import pyplot as plt
import matplotlib.collections as collections
from matplotlib.backends.backend_pdf import PdfPages
from utilities_az import spectroscopy as spec
endpoint = namedtuple('endpoint', ['wave', 'flux', 'error'])
FIG_DIR = '../figures'
def read_iraf_spectrum(filename, redshift=0.0069):
ofile = fits.open(filename)
flux = ofile[0].data[0,0,:]
err = ofile[0].data[3,0,:]
wave = spec.calc_wavelength(ofile[0].header, np.arange(len(flux))+1)
rest_wave = spec.apply_redshift(wave, redshift)
return(spec.spectrum1d(rest_wave, flux, error=err))
def smooth_signal(flux, width, poly_deg):
smoothed_flux = signal.savgol_filter(flux, width, poly_deg)
return smoothed_flux
def find_blue_edge(wave, flux, wcenter, binsize, wmin=None):
'''
Calculate the slope in each bin starting at wmin, until the bin changes sign, use center for blue_edge
'''
wcenter_indx = np.argmin(np.abs(wave-wcenter))
ifit = np.polyfit(wave[wcenter_indx-binsize: wcenter_indx+1],flux[wcenter_indx-binsize: wcenter_indx+1], 1)
slope_product = 1
#plt.plot(wave, flux)
#plt.xlim(wmin, wcenter)
if wmin is None:
search_indx = np.arange(binsize,wcenter_indx+1)
else:
min_indx = np.argmin(np.abs(wave - wmin))
search_indx = np.arange(min_indx, wcenter_indx)
for indx in search_indx[::-1]:
last_slope = ifit[0]
if indx-binsize < 0:
break
ifit = np.polyfit(wave[indx-binsize:indx+1], flux[indx-binsize:indx+1], 1)
#plt.plot(wave[indx-binsize:indx+1], flux[indx-binsize:indx+1])
#plt.plot(wave[indx-binsize:indx+1], np.polyval(ifit, wave[indx-binsize:indx+1]))
slope_product = last_slope*ifit[0] #if this is negative then the slope has changed sign
if slope_product < 0:
break
edge_indx = indx - binsize//2
return edge_indx, wave[edge_indx]
def find_red_edge(wave, flux, wcenter, binsize, wmax = None):
'''
Calculate the slope in each bin starting at wmin, until the bin changes sign, use center for red_edge
binsize is in pixels
'''
wcenter_indx = np.argmin(np.abs(wave-wcenter))
#fig = plt.figure()
#ax1 = fig.add_subplot(2,1,1)
#ax2 = fig.add_subplot(2,1,2, sharex=ax1)
#ax1.plot(wave, flux)
#ax2.plot(wave, wave-wcenter)
ifit = np.polyfit(wave[wcenter_indx:wcenter_indx+binsize+1],flux[wcenter_indx:wcenter_indx+binsize+1], 1)
slope_product = 1
#plt.plot(wave, flux)
#plt.xlim(wmin, wcenter)
#plt.axvline(wave[wcenter_indx], color='y')
if wmax is None:
search_indx = np.arange(wcenter_indx, len(flux))
else:
max_indx = np.argmin(np.abs(wave - wmax))
search_indx = np.arange(wcenter_indx, max_indx+1)
#plt.plot(wave[search_indx], flux[search_indx])
for indx in search_indx:
last_slope = ifit[0]
ifit = np.polyfit(wave[indx:indx+binsize+1], flux[indx:indx+binsize+1], 1)
slope_product = last_slope*ifit[0] #if this is negative then the slope has changed sign
#plt.plot(wave[indx:indx+binsize+1], flux[indx:indx+binsize+1])
#plt.plot(wave[indx:indx+binsize+1], np.polyval(ifit, wave[indx:indx+binsize+1]))
slope_product = last_slope*ifit[0] #if this is negative then the slope has changed sign
if slope_product < 0:
break
edge_indx = indx + binsize//2
return edge_indx, wave[edge_indx]
def check_max(wave, flux, edge_indx, binsize, absorption=True):
'''
Fit a quadratic and verify that it is the correct direction
'''
wmin_indx = edge_indx - binsize//2
wmax_indx = edge_indx + binsize//2
fit = np.polyfit(wave[wmin_indx:wmax_indx+1], flux[wmin_indx:wmax_indx+1], 2)
if fit[0]>0:
concavity = 'up'
if fit[0]<0:
concavity = 'down'
if ((absorption is True) and (concavity is 'down')) or ((absorption is False) and (concavity is 'up')):
good_fit = True
else:
good_fit = False
return good_fit
def calc_rmse(data, model):
rmse = np.sqrt(np.sum((model-data)**2)/len(data))
print('rmse calculated over {} points'.format(len(data)))
return rmse
def find_boundary(wave, flux, wmin, wmax, binsize, visualize=False):
wmin_indx = np.argmin(np.abs(wave-wmin))
wmax_indx = np.argmin(np.abs(wave-wmax))
slope_product = []
ifit = np.polyfit(wave[wmin_indx-binsize//2:wmin_indx+binsize//2+1], flux[wmin_indx-binsize//2:wmin_indx+binsize//2+1], 1)
search_indx = np.arange(wmin_indx, wmax_indx+1)
if visualize:
fig = plt.figure()
ax1 = fig.add_subplot(2,1,1)
ax2 = fig.add_subplot(2,1,2, sharex=ax1)
ax1.set_xlim(wmin, wmax)
for indx in search_indx:
last_slope = ifit[0]
ifit = np.polyfit(wave[indx-int(binsize//2):indx+int(binsize//2)+1], flux[indx-int(binsize//2):indx+int(binsize//2)+1], 1)
slope_product.append(last_slope*ifit[0]) #if this is negative then the slope has changed sign
if visualize:
ax1.plot(wave[indx-int(binsize//2):indx+int(binsize//2)+1], np.polyval(ifit, wave[indx-int(binsize//2):indx+int(binsize//2)+1]))
slope_product = np.array(slope_product)
slope_change_indx = search_indx[slope_product<0]
if visualize:
ax1.set_title('Slope Plot binsize={}, fit_wmin={:4.2f}, fit_wmax={:4.2f}'.format(binsize, wave[wmin_indx-binsize//2], wave[indx+int(binsize//2)]))
ax1.plot(wave, flux)
ax2.plot(wave[search_indx], slope_product)
ax2.axhline(0, color='k', linestyle=':')
if len(slope_change_indx) == 3:
blue_edge_indx, wcenter_indx, red_edge_indx = slope_change_indx
return blue_edge_indx, wcenter_indx, red_edge_indx
else:
return None, None, None
def determine_error_binsize(wave, wave_bin=100):
'''
We should be calculating noise over the same wavelength range
rather than the same number of pixels as long as one wavelength
bin includes enough pixels. Set binsize to be 100A. If there are
fewer than 10 pixels in 100A (dispersion is greater than 10A/pix)
then issue a warning and make binsize 10 pixels regardless of how
many angstroms this represents
wave: array of wavelengths
wave_bin: size of bin in angstroms
outputs: binsize in pixels
Note: right now this calculates the dispersion for the full wavelength range.
For a grating/grism with a large variation in dispersion, it might make sense to
just calculate this over the feature wavelength range.
'''
dispersion = np.median(wave[1:]-wave[:-1])
binsize = np.ceil(wave_bin/dispersion)
if binsize < 10:
print('WARNING: dispersion = {}, \
leading to binsize < 10 for {}$\AA$ bins, \
setting binsize=10, making wave_bin={}'.format(dispersion, wave_bin, 10*dispersion))
binsize=10
return binsize
def define_continuum(wave, flux, edge_indx, binsize, err_binsize, absorption=True, visualize=False):
'''
Fit a quadratic and verify that it is the correct direction
'''
#Silverman says: "Once these two endpoints are determined, a quadratic function is
# fit to the data in wavelength bins centred on each endpoint."
#Let's start with fitting over 2 wavelength bins?
wmin_indx = edge_indx - int(np.floor(1*binsize))
wmax_indx = edge_indx + int(np.ceil(1*binsize))
quad_model = models.Polynomial1D(degree=2)
fitter = fitting.LinearLSQFitter()
fit = fitter(quad_model, wave[wmin_indx:wmax_indx+1], flux[wmin_indx:wmax_indx+1])
fit_extreme_wl = -fit.c1.value/(2*fit.c2.value)
#calc rmse over edge_indx +/- 20 pixels
wmin_rmse = edge_indx - int(err_binsize//2)
wmax_rmse = edge_indx + int(err_binsize//2)
rmse = calc_rmse(flux[wmin_rmse:wmax_rmse], fit(wave[wmin_rmse:wmax_rmse]))
if fit.c2.value>0:
concavity = 'up'
if fit.c2.value<0:
concavity = 'down'
if ((absorption is True) and (concavity is 'down')) or ((absorption is False) and (concavity is 'up')):
good_fit = True
else:
good_fit = False
if visualize:
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(wave, flux)
ax.set_xlim(wave[edge_indx-4*binsize], wave[edge_indx+4*binsize])
ax.plot(wave[wmin_rmse:wmax_rmse], fit(wave[wmin_rmse:wmax_rmse]), label='RMSE range')
ax.plot(wave[wmin_indx:wmax_indx+1], fit(wave[wmin_indx:wmax_indx+1]), label='fit range')
ax.axvline(wave[edge_indx], label='input continuum', color='k')
ax.errorbar(fit_extreme_wl, fit(fit_extreme_wl), yerr=rmse, fmt='.', label='Edge w/error', zorder=10 )
ax.set_ylim(0.9*np.min(flux[wmin_rmse:wmax_rmse]), 1.1*np.max(flux[wmin_rmse:wmax_rmse]))
ax.legend(loc='best')
ax.set_title('absorption={}, concavity={}, good_fit={}'.format(absorption, concavity, good_fit))
endpt = endpoint(fit_extreme_wl, fit(fit_extreme_wl), rmse)
return good_fit, endpt
def calc_pseudo_ew(wave, flux, continuum_l, continuum_r, absorption=True, visualize=False):
'''
wave: array
array of wavelength (can be whole spectrum)
flux: array
array of fluxes (can be whole spectrum)
* Create a fit to the continuum and define the continuum for each wavelength in wave
* Use continuum wavelengths to define index location of feature
* Calc pseudo equivalent width using flux, continuum, and delta wave as calculated from the
wave array
'''
fitter = fitting.LinearLSQFitter()
lin_mod = models.Linear1D()
continuum_fit = fitter(lin_mod,[continuum_l.wave, continuum_r.wave], [continuum_l.flux, continuum_r.flux])
line_indx = np.int_(np.arange(len(wave))[(wave>=continuum_l.wave)&(wave<=continuum_r.wave)])
continuum = continuum_fit(wave[line_indx])
delta_lambda = wave[line_indx]-wave[line_indx-1]
if absorption is True:
pew = np.sum(delta_lambda*(continuum - flux[line_indx])/continuum)
else:
pew = np.sum(delta_lambda*(flux[line_indx]-continuum)/continuum) #Check that this is true
if visualize is True:
fig = plt.figure()
ax1 = fig.add_subplot(1,2,1)
ax2 = fig.add_subplot(1,2,2)
ax2.axhline(1, color='k')
ax1.plot(wave, flux)
ax1.plot(wave[line_indx], flux[line_indx], label='data')
ax1.plot(wave[line_indx], continuum, label='continuum')
ax1.set_xlim(continuum_l.wave-10, continuum_r.wave+10)
if absorption is True:
ax2.plot(wave[line_indx], (continuum - flux[line_indx])/continuum, label='sum for pEW')
else:
ax2.plot(wave[line_indx], (flux[line_indx]-continuum)/continuum, label='sum for pEW')
return pew
def calc_continuum(wave, continuum_l, continuum_r):
fitter = fitting.LinearLSQFitter()
lin_mod = models.Linear1D()
continuum_fit = fitter(lin_mod,[continuum_l.wave, continuum_r.wave], [continuum_l.flux, continuum_r.flux])
continuum = continuum_fit(wave)
return continuum
def find_velocity(wave, flux, error, wcenter, continuum_l, continuum_r, binsize, visualize=False):
line_indx = np.int_(np.arange(len(wave))[(wave>=continuum_l.wave)&(wave<=continuum_r.wave)])
windx_min = int(line_indx[0]-binsize//2)
windx_max = int(line_indx[-1]+binsize//2)
fitter = fitting.LinearLSQFitter()
lin_mod = models.Linear1D()
continuum_fit = fitter(lin_mod,[continuum_l.wave, continuum_r.wave], [continuum_l.flux, continuum_r.flux])
continuum = continuum_fit(wave[windx_min:windx_max])
weight = 1./error[windx_min:windx_max]
fit = interpolate.UnivariateSpline(wave[windx_min:windx_max], flux[windx_min:windx_max]-continuum, w=weight)
if visualize is True:
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(wave[windx_min:windx_max], flux[windx_min:windx_max]-continuum)
s = np.sum((weight * ((flux[windx_min:windx_max]-continuum)-fit(wave[windx_min:windx_max])))**2)
ax.errorbar(wave[windx_min:windx_max], flux[windx_min:windx_max]-continuum, error[windx_min:windx_max], fmt='.', label='spectrum', zorder=1, color='b')
ax.plot(wave[windx_min:windx_max], flux[windx_min:windx_max]-continuum, zorder=2, color='r')
ax.plot(wave[windx_min:windx_max], fit(wave[windx_min:windx_max]), label='fit, s={:2.2f}, len(w)={:2.2f}, med(std)={:2.2e}'.format(s, len(weight), np.median(error[windx_min:windx_max])), color='gold', zorder=3)
min_wave = wave[line_indx][np.argmin(fit(wave[line_indx]))]
ax.axvline(min_wave)
knots = fit.get_knots()
ax.vlines(knots, ymin=ax.get_ylim()[0], ymax=ax.get_ylim()[1], linestyle=':')
ax.legend(loc='best')
return fit
def calc_flux_variance(data, model, err_binsize):
kernel = Box1DKernel(err_binsize)
errors = convolve((data-model)**2, kernel, boundary=None)
errors = errors
errors = np.trim_zeros(errors)
return errors
def calc_continuum_variance(wave, continuum_l, continuum_r):
var = (1./(continuum_l.wave - continuum_r.wave))**2 * \
((wave - continuum_r.wave)**2 * continuum_l.error**2 +
(wave - continuum_l.wave)**2 * continuum_r.error**2)
return var
def calc_pew_variance(flux, continuum, delta_wave, flux_var, continuum_var, visualize=False, wave=None):
'''
Calculate the variance of the equivalent width
Parameters:
-----------
flux: array
flux values over which equivalent width is calculated
continuum: array
continuum values over which equivalent width is calculated
delta_wave: int
the wavelength bin size (in angstroms) used in the equivalent width calculation
flux_var: array
variance in the flux
continuum_var: array
variance in the continuum
Output:
variance in the equivalent width
'''
pew_var_indiv = ((flux/(continuum**2)*delta_wave)**2 * continuum_var) + \
((delta_wave/continuum)**2*flux_var)
pew_var = np.sum(pew_var_indiv)
if visualize is True:
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
if wave is not None:
ax.errorbar(wave, (continuum-flux)/continuum, np.sqrt(pew_err))
else:
print('wavelength not defined')
wave = np.arange(len(flux))
ax.errorbar(wave,(continuum-flux)/continuum, np.sqrt(pew_err))
return pew_var
def calc_velocity_error(wave, flux, vel_fit, continuum = None, visualize=False):
'''
Calculate 1-sigma errors
'''
min_indx = np.argmin(vel_fit(wave))
indx = np.argsort(wave - wave[min_indx])
min_indx_indx = int(indx[indx==min_indx])
cum_sum_right = 0
if continuum is None:
flux_sub = flux
else:
flux_sub = flux - continuum
total = np.sum((flux_sub))
for i in indx[min_indx_indx:]:
cum_sum_right += (flux_sub)[i]
if cum_sum_right/total > .341:
break
right_err = wave[i]-wave[min_indx]
cum_sum_left = 0
j=0
for j in indx[:min_indx_indx][::-1]:
cum_sum_left += (flux_sub)[j]
if cum_sum_left/total > .341:
break
left_err = wave[min_indx]-wave[j]
if visualize is True:
from visualization import make_color_wheel
colors = make_color_wheel(wave)
plt.figure()
for c, ind in zip(colors, indx):
plt.plot(wave[ind], flux[ind], marker='o', ls='none', color=c)
for k in indx[min_indx_indx:i]:
plt.plot(wave[k], flux[k], marker='s', ls='none', color=colors[k])
for k in indx[j:min_indx_indx]:
plt.plot(wave[k], flux[k], marker='s', ls='none', color=colors[k])
plt.axvline(wave[i], label='1 $\sigma$ right error')
plt.axvline(wave[j], label='1 $\sigma$ left error')
plt.xlabel('Wavelength')
plt.ylabel('Flux')
plt.legend(loc='best')
return left_err, right_err
def find_edges(spectrum, feature_dict, smooth_flux, filename, vis=False, pp=None):
'''
Fit incremental slopes to find where spectrum turns over
Fit quadratic to the turnover points to get spectrum edges
'''
#Estimate the edges and center of the feature
blue_edge_indx = None
red_edge_indx = None
good_fit_blue = False
good_fit_red = False
adjust_binsize = feature_dict['edge_param']['binsize']
#increase the binsize until only 3 turning points are found (edges and center)
wmin, wmax = find_wavelength_range(feature_dict, filename)
npts_feature = len(spectrum.wave[(spectrum.wave>=wmin) & (spectrum.wave <= wmax)])
while ((blue_edge_indx is None) or (red_edge_indx is None) or
(good_fit_blue is False) or (good_fit_red is False)) and \
(adjust_binsize < 0.4*npts_feature) and \
(adjust_binsize < feature_dict['edge_param']['binmax']): #TODO figure out a cutoff for this
if plt.get_fignums() is not False:
for ifig in plt.get_fignums():
plt.close(ifig)
adjust_binsize += 2
blue_edge_indx, wcenter_indx, red_edge_indx = find_boundary(spectrum.wave,
smooth_flux,
wmin,
wmax,
adjust_binsize,
visualize=vis)
if (blue_edge_indx is not None) and (red_edge_indx is not None):
err_binsize = determine_error_binsize(spectrum.wave, wave_bin=100)
#Find the feature edges and errors
good_fit_blue, continuum_l = define_continuum(spectrum.wave, smooth_flux, blue_edge_indx, feature_dict['edge_param']['concavity_binsize'], err_binsize, absorption=True, visualize=vis)
good_fit_red, continuum_r = define_continuum(spectrum.wave, smooth_flux, red_edge_indx, feature_dict['edge_param']['concavity_binsize'], err_binsize, absorption=True, visualize=vis)
if continuum_l.wave > continuum_r.wave:
print('**** WARNING: {}, left edge {} is greater than right edge {}****'.format(os.path.basename(filename), continuum_l.wave, continuum_r.wave))
return None
if adjust_binsize > 0.4*npts_feature:
blue_edge_indx, wcenter_indx, red_edge_indx = find_boundary(spectrum.wave,
smooth_flux,
wmin,
wmax,
feature_dict['edge_param']['binsize'],
visualize=True)
print('Unable to find edges for {}, {}'.format(feature_dict['name'],os.path.basename(filename)))
import pdb; pdb.set_trace()
return None
else:
print('filename = ',os.path.basename(filename))
print('\tinput binsize = ', feature_dict['edge_param']['binsize'])
print('\tadjusted binsize = ', adjust_binsize)
print('good_fit_blue={}, good_fit_red={}, combine={}'.format(good_fit_blue, good_fit_red, ((good_fit_blue is False) or (good_fit_red is False))))
if vis is True:
fig1 = plt.figure(1)
pp.savefig(fig1)
plt.close(fig1)
fig2 = plt.figure(2)
pp.savefig(fig2)
plt.close(fig2)
fig3 = plt.figure(3)
pp.savefig(fig3)
plt.close(fig3)
return err_binsize, blue_edge_indx, red_edge_indx, wcenter_indx, continuum_l, continuum_r
def find_wavelength_range(feature_dict, filename):
date = Time(fits.getval(filename, 'date-obs', 0))
phase = date.jd - feature_dict['texpl']
delta_wave = feature_dict['slope']*phase
wmin = feature_dict['wmin']+delta_wave
wmax = feature_dict['wmax']+delta_wave
return wmin, wmax
def final_plot(wave, flux, flux_err, continuum_l, continuum_r, vel_fit, vel_min, vel_err, pew, pew_err):
continuum = calc_continuum(wave, continuum_l, continuum_r)
plt.ioff()
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
#ax.plot(wave, flux/continuum, label='Spectrum')
ax.errorbar(wave, flux/continuum, flux_err/continuum, fmt='.', label='Spectrum')
ax.errorbar(np.array([continuum_l.wave, continuum_r.wave]),
np.array([continuum_l.flux, continuum_r.flux])/calc_continuum(np.array([continuum_l.wave, continuum_r.wave]), continuum_l, continuum_r),
np.array([continuum_l.error, continuum_r.error])/calc_continuum(np.array([continuum_l.wave, continuum_r.wave]), continuum_l, continuum_r),
fmt='o', label='Feature Edges')
min_continuum = calc_continuum(np.array([vel_min]), continuum_l, continuum_r)[0]
ax.plot(wave, (vel_fit(wave)+continuum)/continuum, label='Velocity fit')
ax.errorbar(vel_min, np.min((vel_fit(vel_min)+min_continuum)/min_continuum), xerr=(vel_err,), fmt='s', label='Velocity')
pew_collection_err = collections.BrokenBarHCollection.span_where(wave, ymin=0, ymax=1,
where= (wave>=vel_min-pew/2-np.sqrt(pew_err))&(wave<=vel_min+pew/2+np.sqrt(pew_err)),
color='k', alpha=0.1)
pew_collection = collections.BrokenBarHCollection.span_where(wave, ymin=0, ymax=1,
where= (wave>=vel_min-pew/2)&(wave<=vel_min+pew/2),
color='k', alpha=0.1, label = 'pEW')
ax.add_collection(pew_collection_err)
ax.add_collection(pew_collection)
ax.legend(loc='best')
ax.set_xlabel('Wavelength')
ax.set_ylabel('Continuum subtracted flux')
#ax.set_ylim(-0.05, 1.1)
plt.ion()
return fig
def characterize_line(feature_dict, filename, visualization_level=0):
final_vis = False
intermediate_vis = False
pp = None
if (visualization_level == 1) or (visualization_level ==2):
final_vis = True
if visualization_level == 2:
intermediate_vis = True
pp = PdfPages(os.path.join(FIG_DIR,
'line_fit_intermed_{}_{}.pdf'.format(feature_dict['name'],
os.path.basename(filename).split('.pdf')[0])))
#Read in spectrum
spectrum = read_iraf_spectrum(filename)
#Remove CR and other large deviations
smooth_flux = smooth_signal(spectrum.flux,
feature_dict['smooth_param']['width'],
feature_dict['smooth_param']['deg'])
edge_results = find_edges(spectrum, feature_dict, smooth_flux, filename, vis=intermediate_vis, pp=pp)
if edge_results is not None:
err_binsize, blue_edge_indx, red_edge_indx, wcenter_indx, continuum_l, continuum_r = edge_results
#Calculate the pseudo equivalent widths
pew = calc_pseudo_ew(spectrum.wave, smooth_flux, continuum_l, continuum_r, visualize=intermediate_vis)
if intermediate_vis is True:
pp.savefig()
plt.close()
#Calculate the most common velocity
wcenter = spectrum.wave[wcenter_indx]
vel_fit = find_velocity(spectrum.wave, smooth_flux, spectrum.error, wcenter, continuum_l, continuum_r, err_binsize, visualize=intermediate_vis)
if intermediate_vis is True:
pp.savefig()
plt.close()
#Find the error in the pseudo equivalent width
line_indx = np.arange(len(spectrum.wave))[(spectrum.wave>=continuum_l.wave)&(spectrum.wave<=continuum_r.wave)]
min_indx = int(np.floor(line_indx[0]-err_binsize/2))
max_indx = int(np.ceil(line_indx[-1]+err_binsize/2+1))
continuum_extended = calc_continuum(spectrum.wave[min_indx:max_indx], continuum_l, continuum_r)
flux_var = calc_flux_variance(spectrum.flux[min_indx:max_indx]-continuum_extended,
vel_fit(spectrum.wave[min_indx:max_indx]), err_binsize) #These don't include the errors from the continuum subtraction yet; ok for EW calc
if len(flux_var) > len(spectrum.flux[line_indx]):
flux_var = flux_var[1:-1]
continuum = calc_continuum(spectrum.wave[line_indx], continuum_l, continuum_r)
continuum_var = calc_continuum_variance(spectrum.wave[line_indx], continuum_l, continuum_r)
delta_wave = np.median(spectrum.wave[1:]-spectrum.wave[:-1])
pew_var = calc_pew_variance(spectrum.flux[line_indx], continuum, delta_wave, flux_var, continuum_var, wave=spectrum.wave[line_indx])
#Find the velocity error
vel_err = calc_velocity_error(spectrum.wave[line_indx], spectrum.flux[line_indx], vel_fit, continuum=continuum)
vel_min = spectrum.wave[line_indx][np.argmin(vel_fit(spectrum.wave[line_indx]))]
if final_vis is True:
fig = final_plot(spectrum.wave[min_indx:max_indx], spectrum.flux[min_indx:max_indx],spectrum.error[min_indx:max_indx], continuum_l, continuum_r, vel_fit, vel_min, vel_err, pew, pew_var)
fig.suptitle(os.path.basename(filename))
if intermediate_vis:
pp.close()
return pew, pew_var, vel_min, vel_err, fig |
<reponame>d02d33pak/PyQt5-Apps<filename>calculator/ui.py
"""
UI Doc for Calculator App
"""
from PyQt5 import QtWidgets as qtw
from PyQt5 import QtGui as qtg
class MainUI:
def init_ui(self):
self.lcd_display = qtw.QLCDNumber()
self.lcd_display.setDigitCount(10)
self.lcd_display.setMinimumHeight(100)
self.main_v_layout = qtw.QVBoxLayout()
self.main_v_layout.addWidget(self.lcd_display)
grid_layout = qtw.QGridLayout()
grid_layout.setSpacing(8)
grid_layout.setContentsMargins(4, 12, 4, 4) # top margin from display
font = qtg.QFont()
font.setPointSize(20)
font.setBold(True)
font.setWeight(60)
self.n0 = qtw.QPushButton("0")
self.n1 = qtw.QPushButton("1")
self.n2 = qtw.QPushButton("2")
self.n3 = qtw.QPushButton("3")
self.n4 = qtw.QPushButton("4")
self.n5 = qtw.QPushButton("5")
self.n6 = qtw.QPushButton("6")
self.n7 = qtw.QPushButton("7")
self.n8 = qtw.QPushButton("8")
self.n9 = qtw.QPushButton("9")
self.mod = qtw.QPushButton("%")
self.add = qtw.QPushButton("+")
self.sub = qtw.QPushButton("-")
self.mul = qtw.QPushButton("x")
self.div = qtw.QPushButton("÷")
self.eq = qtw.QPushButton("=")
self.ac = qtw.QPushButton("AC")
self.bs = qtw.QPushButton("DEL")
self.n0.setFont(font)
self.n1.setFont(font)
self.n2.setFont(font)
self.n3.setFont(font)
self.n4.setFont(font)
self.n5.setFont(font)
self.n6.setFont(font)
self.n7.setFont(font)
self.n8.setFont(font)
self.n9.setFont(font)
self.mod.setFont(font)
self.add.setFont(font)
self.sub.setFont(font)
self.mul.setFont(font)
self.div.setFont(font)
self.eq.setFont(font)
self.ac.setFont(font)
self.bs.setFont(font)
# so the button can expand in Y axis
self.eq.setSizePolicy(qtw.QSizePolicy.Expanding, qtw.QSizePolicy.Expanding)
self.ac.setSizePolicy(qtw.QSizePolicy.Expanding, qtw.QSizePolicy.Expanding)
# some simple styling
self.mod.setStyleSheet("QPushButton{color:#1976D2}")
self.add.setStyleSheet("QPushButton{color:#1976D2}")
self.sub.setStyleSheet("QPushButton{color:#1976D2}")
self.mul.setStyleSheet("QPushButton{color:#1976D2}")
self.div.setStyleSheet("QPushButton{color:#1976D2}")
self.eq.setStyleSheet("QPushButton{color:#4CAF50}")
self.ac.setStyleSheet("QPushButton{color:#F44336}")
self.bs.setStyleSheet("QPushButton{color:#FFC107}")
# tooltip for operators only
self.mod.setToolTip("Mod")
self.add.setToolTip("Add")
self.sub.setToolTip("Subract")
self.mul.setToolTip("Multiply")
self.div.setToolTip("Divide")
self.eq.setToolTip("Equals")
self.bs.setToolTip("Backspace")
self.ac.setToolTip("All Clear")
# first row on calc
grid_layout.addWidget(self.div, 0, 0)
grid_layout.addWidget(self.mul, 0, 1)
grid_layout.addWidget(self.sub, 0, 2)
grid_layout.addWidget(self.bs, 0, 3)
# second row on calc
grid_layout.addWidget(self.n7, 1, 0)
grid_layout.addWidget(self.n8, 1, 1)
grid_layout.addWidget(self.n9, 1, 2)
grid_layout.addWidget(self.ac, 1, 3)
# third row on calc
grid_layout.addWidget(self.n4, 2, 0)
grid_layout.addWidget(self.n5, 2, 1)
grid_layout.addWidget(self.n6, 2, 2)
grid_layout.addWidget(self.eq, 2, 3, 3, 1)
# fourth row on calc
grid_layout.addWidget(self.n1, 3, 0)
grid_layout.addWidget(self.n2, 3, 1)
grid_layout.addWidget(self.n3, 3, 2)
# fifth row on calc
grid_layout.addWidget(self.mod, 4, 0)
grid_layout.addWidget(self.n0, 4, 1)
grid_layout.addWidget(self.add, 4, 2)
self.main_v_layout.addLayout(grid_layout)
|
<reponame>dpa-newslab/livebridge-liveblog
# -*- coding: utf-8 -*-
#
# Copyright 2016 dpa-infocom GmbH
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import asynctest
from datetime import datetime
from livebridge_liveblog import LiveblogPost
from tests import load_json
class LiveblogPostTest(asynctest.TestCase):
def setUp(self):
self.post = load_json('post_to_convert.json')
self.images = ["/tmp/one.jpg"]
self.content= "foobaz"
self.lp = LiveblogPost(self.post, content=self.content, images=self.images)
@asynctest.fail_on(unused_loop=False)
def test_init(self):
assert self.lp.data == self.post
assert hasattr(self.lp, "is_deleted") == True
assert hasattr(self.lp, "is_update") == True
assert hasattr(self.lp, "is_sticky") == True
assert self.lp.id == "urn:newsml:localhost:2016-04-28T11:24:22.973191:666890f6-9054-4f81-81ac-cc6d5f02b2c9"
assert self.lp.source_id == "56fceedda505e600f71959c8"
assert type(self.lp.updated) == datetime
assert type(self.lp.created) == datetime
assert self.lp.created.year == 2016
assert self.lp.created.minute == 24
assert self.lp.updated.year == 2016
assert self.lp.updated.second == 22
assert self.lp.images == self.images
assert self.lp.content == self.content
@asynctest.fail_on(unused_loop=False)
def test_get_action(self):
# ignore/submitted
self.lp._existing = None
self.post["post_status"] = "submitted"
assert self.lp.get_action() == "ignore"
# ignore/draft
self.post["post_status"] = "draft"
assert self.lp.get_action() == "ignore"
# no ignore, post is known
self.lp._existing = {"foo":"baz"}
assert self.lp.get_action() != "ignore"
# should be update
self.post["post_status"] = ""
assert self.lp.get_action() == "update"
# test delete
self.lp._deleted = True
assert self.lp.get_action() == "delete"
# test ignore for unknown
self.lp._deleted = None
self.lp._existing = None
assert self.lp.get_action() == "create"
# test ignore for deleted
self.lp._deleted = True
assert self.lp.get_action() == "ignore"
@asynctest.fail_on(unused_loop=False)
def test_is_not_delete(self):
assert self.lp.is_deleted == False
@asynctest.fail_on(unused_loop=False)
def test_is_deleted(self):
self.lp.data["deleted"] = True
assert self.lp.is_deleted == True
self.lp._deleted = False
assert self.lp.is_deleted == False
@asynctest.fail_on(unused_loop=False)
def test_is_deleted_unpublished(self):
self.lp.data["unpublished_date"] = "2016-05-06T15:00:59+00:00"
self.lp.data["published_date"] = "2016-05-06T15:00:39+00:00"
assert self.lp.is_deleted == True
@asynctest.fail_on(unused_loop=False)
def test_is_sticky(self):
assert self.lp.is_sticky == False
self.lp.data["sticky"] = True
assert self.lp.is_sticky == True
@asynctest.fail_on(unused_loop=False)
def test_is_highlighted(self):
assert self.lp.is_highlighted == False
self.lp.data["lb_highlight"] = True
assert self.lp.is_highlighted == True
@asynctest.fail_on(unused_loop=False)
def test_is_submitted(self):
assert self.lp.is_submitted == False
self.lp.data["post_status"] = "submitted"
assert self.lp.is_submitted == True
@asynctest.fail_on(unused_loop=False)
def test_is_draft(self):
assert self.lp.is_draft == False
self.lp.data["post_status"] = "draft"
assert self.lp.is_draft == True
@asynctest.fail_on(unused_loop=False)
def test_is_update(self):
self.lp.data["_created"] = "new"
self.lp.data["_updated"] = "new"
assert self.lp.is_update == False
self.lp.data["_updated"] = "new2"
assert self.lp.is_update == True
@asynctest.fail_on(unused_loop=False)
def test_existing(self):
assert self.lp.get_existing() == None
assert self.lp.is_known == False
self.lp.set_existing({"foo": "baz"})
assert self.lp.get_existing() == {"foo": "baz"}
assert self.lp.is_known == True
@asynctest.fail_on(unused_loop=False)
def test_target_doc(self):
assert self.lp.target_doc == None
self.lp._existing = {"target_doc": {"doc": "foo"}}
assert self.lp.target_doc == self.lp._existing["target_doc"]
@asynctest.fail_on(unused_loop=False)
def test_target_id(self):
assert self.lp._target_id == None
self.lp._target_id = "foobaz"
assert self.lp.target_id == "foobaz"
@asynctest.fail_on(unused_loop=False)
def test_target_id_from_existing(self):
self.lp.set_existing({"target_id": "foobaz"})
assert self.lp.target_id == "foobaz"
|
#!/usr/bin/env python3
import numpy as np
from scipy.io import netcdf
import sys, os
def main(file):
print()
print("Usage: "+file+" quasisymmetry_out.*.nc")
#if len(sys.argv) != 2:
# print("Error! You must specify 1 argument: the quasisymmetry_out.*.nc file")
# exit(1)
def toString(ncVar):
temp = [c.decode('UTF-8') for c in ncVar]
return (''.join(temp)).strip()
filename = file#sys.argv[1]
print("Reading filename "+filename)
f = netcdf.netcdf_file(filename,mode='r',mmap=False)
general_option = toString(f.variables['general_option'][()])
if general_option != "single":
print("Error! This script is designed for plotting single runs, but the quasisymmetry_out file you provided is a scan.")
f.close()
exit(1)
nfp = f.variables['nfp'][()]
B0 = f.variables['B0'][()]
r = f.variables['r'][()]
eta_bar = f.variables['eta_bar'][()]
mpol = f.variables['mpol'][()]
ntor = f.variables['ntor'][()]
RBC = f.variables['RBC'][()]
RBS = f.variables['RBS'][()]
ZBC = f.variables['ZBC'][()]
ZBS = f.variables['ZBS'][()]
R0c = f.variables['R0c'][()]
R0s = f.variables['R0s'][()]
Z0c = f.variables['Z0c'][()]
Z0s = f.variables['Z0s'][()]
print("RBC.shape:",RBC.shape)
phi = f.variables['phi'][()]
X1c = f.variables['X1c'][()]
Y1c = f.variables['Y1c'][()]
Y1s = f.variables['Y1s'][()]
sigma = f.variables['sigma'][()]
curvature = f.variables['curvature'][()]
torsion = f.variables['torsion'][()]
elongation = f.variables['elongation'][()]
elongation_in_Rz_plane = f.variables['elongation_in_Rz_plane'][()]
modBinv_sqrt_half_grad_B_colon_grad_B = f.variables['modBinv_sqrt_half_grad_B_colon_grad_B'][()]
#order_r_option = f.variables["order_r_option"][()]
#order_r_option = ''.join(str(f.variables["order_r_option"][()]))
order_r_option = toString(f.variables["order_r_option"][()])
print("order_r_option:",order_r_option)
order_r_squared = (order_r_option != 'r1' and order_r_option != 'r1_compute_B2')
print("order_r_squared:",order_r_squared)
order_r_cubed = (order_r_option != 'r1' and order_r_option != 'r1_compute_B2' and order_r_option != 'r2')
print("order_r_cubed:",order_r_cubed)
order_r1_compute_B2 = order_r_option == 'r1_compute_B2'
if order_r1_compute_B2:
B20 = f.variables['B20'][()]
B2s_array = f.variables['B2s_array'][()]
B2c_array = f.variables['B2c_array'][()]
B02 = f.variables['B02'][()]
if order_r_squared:
B2s = f.variables['B2s'][()]
B2c = f.variables['B2c'][()]
B20_mean = f.variables['B20_mean'][()]
B20 = f.variables['B20'][()]
X20 = f.variables['X20'][()]
X2s = f.variables['X2s'][()]
X2c = f.variables['X2c'][()]
Y20 = f.variables['Y20'][()]
Y2s = f.variables['Y2s'][()]
Y2c = f.variables['Y2c'][()]
Z20 = f.variables['Z20'][()]
Z2s = f.variables['Z2s'][()]
Z2c = f.variables['Z2c'][()]
r_singularity_vs_zeta = f.variables['r_singularity_vs_zeta'][()]
try:
r_singularity_basic_vs_zeta = f.variables['r_singularity_basic_vs_zeta'][()]
except:
r_singularity_basic_vs_zeta = r_singularity_vs_zeta # Old output files might not have this field
for j in range(len(phi)):
if r_singularity_vs_zeta[j] > 1.0e10:
r_singularity_vs_zeta[j] = np.nan
if r_singularity_basic_vs_zeta[j] > 1.0e10:
r_singularity_basic_vs_zeta[j] = np.nan
if order_r_cubed:
X3s1 = f.variables['X3s1'][()]
X3c1 = f.variables['X3c1'][()]
X3s3 = f.variables['X3s3'][()]
X3c3 = f.variables['X3c3'][()]
Y3s1 = f.variables['Y3s1'][()]
Y3c1 = f.variables['Y3c1'][()]
Y3s3 = f.variables['Y3s3'][()]
Y3c3 = f.variables['Y3c3'][()]
Z3s1 = f.variables['Z3s1'][()]
Z3c1 = f.variables['Z3c1'][()]
Z3s3 = f.variables['Z3s3'][()]
Z3c3 = f.variables['Z3c3'][()]
try:
B3s1 = f.variables['B3s1'][()]
B3c1 = f.variables['B3c1'][()]
B3s3 = f.variables['B3s3'][()]
B3c3 = f.variables['B3c3'][()]
except:
B3s1 = phi * 0
B3s3 = phi * 0
B3c1 = phi * 0
B3c3 = phi * 0
f.close()
my_xlim = [0,phi[-1]]
N_theta = 150
N_phi = 8
Nfig = N_phi
theta1D = np.linspace(0,2*np.pi,N_theta)
phi1D = np.linspace(0,2*np.pi/nfp,N_phi,endpoint=False)
phi2D,theta2D = np.meshgrid(phi1D,theta1D)
#print "theta2D:"
#print theta2D
R = np.zeros((N_theta,N_phi))
z = np.zeros((N_theta,N_phi))
for m in range(mpol+1):
for jn in range(ntor*2+1):
n = jn-ntor
angle = m * theta2D - nfp * n * phi2D
sinangle = np.sin(angle)
cosangle = np.cos(angle)
R += RBC[m,jn] * cosangle + RBS[m,jn] * sinangle
z += ZBC[m,jn] * cosangle + ZBS[m,jn] * sinangle
R0 = np.zeros(N_phi)
z0 = np.zeros(N_phi)
for n in range(len(R0c)):
angle = nfp * n * phi1D
sinangle = np.sin(angle)
cosangle = np.cos(angle)
R0 += R0c[n] * cosangle + R0s[n] * sinangle
z0 += Z0c[n] * cosangle + Z0s[n] * sinangle
#exit(0)
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(16,7))
fig.patch.set_facecolor('white')
if order_r_cubed:
numRows = 5
numCols = 7
elif order_r_squared:
numRows = 4
numCols = 5
elif order_r1_compute_B2:
numRows = 3
numCols = 4
else:
numRows = 2
numCols = 4
plotNum = 1
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,curvature)
plt.title('curvature')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,torsion)
plt.title('torsion')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X1c)
plt.title('X1c')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y1s)
plt.title('Y1s')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y1c)
plt.title('Y1c')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,sigma)
plt.title('sigma')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,elongation,label='XY plane')
plt.plot(phi,elongation_in_Rz_plane,label='Rz plane')
plt.title('elongation')
plt.legend(loc=0,fontsize=6)
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,modBinv_sqrt_half_grad_B_colon_grad_B)
#plt.title('modBinv_sqrt_half_grad_B_colon_grad_B')
plt.title('Frobenius')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
if order_r1_compute_B2:
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B20)
plt.title('B20')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B2s_array)
plt.title('B2s_array')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B2c_array)
plt.title('B2c_array')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B02)
plt.title('B_0^{(2)}')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
if order_r_squared:
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X20)
plt.title('X20')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X2s)
plt.title('X2s')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X2c)
plt.title('X2c')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y20)
plt.title('Y20')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y2s)
plt.title('Y2s')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y2c)
plt.title('Y2c')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z20)
plt.title('Z20')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z2s)
plt.title('Z2s')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z2c)
plt.title('Z2c')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B20)
plt.title('B20')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,r_singularity_vs_zeta,label='refined')
plt.plot(phi,r_singularity_basic_vs_zeta,':k',label='unrefined')
plt.legend(fontsize=6,loc=0)
plt.title('r_singularity_vs_zeta')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
if order_r_cubed:
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X3s1)
plt.title('X3s1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X3c1)
plt.title('X3c1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X3s3)
plt.title('X3s3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,X3c3)
plt.title('X3c3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y3s1)
plt.title('Y3s1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y3c1)
plt.title('Y3c1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y3s3)
plt.title('Y3s3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Y3c3)
plt.title('Y3c3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z3s1)
plt.title('Z3s1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z3c1)
plt.title('Z3c1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z3s3)
plt.title('Z3s3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,Z3c3)
plt.title('Z3c3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B3s1)
plt.title('B3s1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B3c1)
plt.title('B3c1')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B3s3)
plt.title('B3s3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.subplot(numRows,numCols,plotNum)
plotNum += 1
plt.plot(phi,B3c3)
plt.title('B3c3')
plt.xlabel('$\phi$')
plt.xlim(my_xlim)
plt.tight_layout()
titleString = "Plot generated by "+ os.path.abspath(file)
plt.figtext(0.5,0.004,titleString,horizontalalignment='center',verticalalignment='bottom',fontsize=8)
plt.figtext(0.5,0.997,'File = '+os.path.abspath(filename),horizontalalignment='center',verticalalignment='top',fontsize=8)
plt.savefig(file+'_params.pdf', bbox_inches = 'tight', pad_inches = 0)
###################################################################3
fig = plt.figure(figsize=(16,7))
fig.patch.set_facecolor('white')
numRows = 3
numCols = 3
plt.subplot(numRows,numCols,1)
for jphi in range(N_phi):
plt.plot(R[:,jphi],z[:,jphi])
plt.xlabel('R')
plt.ylabel('z')
plt.gca().set_aspect('equal',adjustable='box')
Rfig=R
Zfig=z
R0fig=R0
Z0fig=z0
for jphi in range(N_phi):
plt.subplot(numRows,numCols,jphi+2)
for k in range(N_theta):
plt.plot([R0[jphi],R[k,jphi]], [z0[jphi],z[k,jphi]],'-')
plt.plot(R[:,jphi],z[:,jphi])
plt.plot(R0[jphi],z0[jphi],'o')
plt.xlabel('R')
plt.ylabel('z')
plt.gca().set_aspect('equal',adjustable='box')
plt.tight_layout()
titleString = "Plot generated by "+ os.path.abspath(__file__)
plt.figtext(0.5,0.004,titleString,horizontalalignment='center',verticalalignment='bottom',fontsize=8)
plt.figtext(0.5,0.997,'File = '+os.path.abspath(filename),horizontalalignment='center',verticalalignment='top',fontsize=8)
########################################################
# Now make 3D surface plot
########################################################
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from numpy.matlib import repmat
fig = plt.figure(figsize=(16,7))
fig.patch.set_facecolor('white')
# Zoom in:
#factor = 1
factor = 0.2
fig.subplots_adjust(bottom=-factor+0.0,top=0.95+factor)
N_theta = 40
N_phi = 150
theta1D = np.linspace(0,2*np.pi,N_theta)
phi1D = np.linspace(0,2*np.pi,N_phi)
phi2D,theta2D = np.meshgrid(phi1D,theta1D)
#print "theta2D:"
#print theta2D
R = np.zeros((N_theta,N_phi))
z = np.zeros((N_theta,N_phi))
for m in range(mpol+1):
for jn in range(ntor*2+1):
n = jn-ntor
angle = m * theta2D - nfp * n * phi2D
sinangle = np.sin(angle)
cosangle = np.cos(angle)
R += RBC[m,jn] * cosangle + RBS[m,jn] * sinangle
z += ZBC[m,jn] * cosangle + ZBS[m,jn] * sinangle
x = R * np.cos(phi2D)
y = R * np.sin(phi2D)
B = B0 * (1 + r * eta_bar * np.cos(theta2D))
if order_r_squared:
#B20_2D = repmat(B20,N_theta,1)
B20_interpolated = np.interp(phi1D,phi,B20,period=2*np.pi/nfp)
#print "B20:",B20
#print "B20_interpolated:",B20_interpolated
B20_2D = repmat(B20_interpolated,N_theta,1)
B += r * r * (B2s * np.sin(2*theta2D) + B2c * np.cos(2*theta2D) + B20_2D)
# Rescale to lie in [0,1]:
B_rescaled = (B - B.min()) / (B.max() - B.min())
ax = fig.gca(projection='3d')
#ax.set_aspect('equal')
ax.plot_surface(x, y, z, facecolors = cm.viridis(B_rescaled), rstride=1, cstride=1, antialiased=False)
max_range = np.array([x.max()-x.min(), y.max()-y.min(), z.max()-z.min()]).max() / 2.0
mid_x = (x.max()+x.min()) * 0.5
mid_y = (y.max()+y.min()) * 0.5
mid_z = (z.max()+z.min()) * 0.5
ax.set_xlim(mid_x - max_range, mid_x + max_range)
ax.set_ylim(mid_y - max_range, mid_y + max_range)
ax.set_zlim(mid_z - max_range, mid_z + max_range)
plt.show()
print("Save PDF")
fig = plt.figure(figsize=(14,7))
fig.patch.set_facecolor('white')
for jphi in range(Nfig):
plt.plot(Rfig[:,jphi],Zfig[:,jphi])
plt.plot(R0fig[jphi],Z0fig[jphi])
plt.xlabel('R')
plt.ylabel('Z')
plt.gca().set_aspect('equal',adjustable='box')
plt.savefig(file+'_surfs.pdf', bbox_inches = 'tight', pad_inches = 0)
|
<reponame>ablancha/gppath<gh_stars>1-10
import numpy as np
import time
import GPy
from .augmented_inputs import AugmentedInputs
from gpsearch.core.kernels import *
from gpsearch.core.acquisitions.check_acquisition import check_acquisition
class OptimalPath(object):
"""A class for Bayesian path-planning algorithm. This class assumes
that the objective function *may* change with time.
Parameters
---------
X_pose : array_like
The robot's starting pose. Must be in the form
(x_pos, y_pos, angle)
Y_pose : array_like
Observation at starting position.
my_map: instance of `BlackBox`
The black-box objective function.
inputs : instance of `Inputs`
The input space.
static : boolean, optional
Whether or not the objective function changes in time. If `True`,
sets the lengthscale for the temporal variable to +Inf.
fix_noise : boolean, optional
Whether or not to fix the noise variance in the GP model.
noise_var : float, optional
Variance for additive Gaussian noise. Default is None, in
which case the noise variance from BlackBox is used. If fix_noise
is False, noise_var is merely used to initialize the GP model.
normalize_Y : boolean, optional
Whether or not to normalize the output in the GP model.
Attributes
----------
my_map, inputs : see Parameters
X : array
Array of input points.
Y : array
Array of record_time.
X_pose : array_like
The robot's current pose.
model : instance of `GPRegression`
Current GPy model.
"""
def __init__(self, X_pose, Y_pose, my_map, inputs, static=False,
fix_noise=False, noise_var=None, normalize_Y=True):
self.my_map = my_map
self.inputs = AugmentedInputs(inputs)
self.X = np.atleast_2d(np.append(X_pose[0:2], 0))
self.Y = np.atleast_2d(Y_pose)
self.X_pose = X_pose
if noise_var is None:
noise_var = my_map.noise_var
# Currently only the RBF kernel is supported.
ker = RBF(input_dim=self.inputs.input_dim, ARD=True)
if static:
ker.lengthscale[-1:].fix(np.inf)
self.model = GPy.models.GPRegression(X=self.X,
Y=self.Y,
kernel=ker,
normalizer=normalize_Y,
noise_var=noise_var)
if fix_noise:
self.model.Gaussian_noise.variance.fix(noise_var)
def optimize(self, record_time, acquisition, path_planner,
callback=True, save_iter=True, prefix=None, kwargs_GPy=None):
"""Runs the Bayesian path-planning algorithm.
Parameters
----------
record_time : array_like
Time vector for when measurements are made.
acquisition : str or instance of `Acquisition`
Acquisition function for determining the next best point.
If a string, must be one of
- "PI": Probability of Improvement
- "EI": Expected Improvement
- "US": Uncertainty Sampling
- "US-BO": US repurposed for Bayesian Optimization (BO)
- "US-LW": Likelihood-Weighted US
- "US-LWBO": US-LW repurposed for BO
- "US-LWraw" : US-LW with no GMM approximation
- "US-LWBOraw": US-LWBO with no GMM approximation
- "LCB" : Lower Confidence Bound
- "LCB-LW" : Likelihood-Weighted LCB
- "LCB-LWraw" : LCB-LW with no GMM approximation
- "IVR" : Integrated Variance Reduction
- "IVR-IW" : Input-Weighted IVR
- "IVR-LW" : Likelihood-Weighted IVR
- "IVR-BO" : IVR repurposed for BO
- "IVR-LWBO": IVR-LW repurposed for BO
path_planner : instance of `PathPlanner`
Parametrization of the path by <NAME>.
callback : boolean, optional
Whether or not to display log at each iteration.
save_iter : boolean, optional
Whether or not to save the GP model at each iteration.
prefix : string, optional
Prefix for file naming
kwargs_GPy : dict, optional
Dictionary of arguments to be passed to the GP model.
Returns
-------
m_list : list
A list of trained GP models, one for each iteration of
the algorithm.
p_list : list
A list of paths, one for each iteration of the algorithm.
s_list : list
A list of trained GP models, one for each sample point.
"""
if prefix is None:
prefix = "model"
if kwargs_GPy is None:
kwargs_GPy = dict(num_restarts=10, optimizer="bfgs",
max_iters=1000, verbose=False)
m_list, p_list = [], []
t_cur = 0
t_fin = record_time[-1]
ii = -1
while t_cur <= t_fin:
ii += 1
tic = time.time()
filename = (prefix+"%.4d")%(ii)
if ii == 0:
# Densify frontier for truly uniform sampling
tmp = path_planner.n_frontier
path_planner.n_frontier = 1000
paths = path_planner.make_paths(self.X_pose)
popt = paths[np.random.randint(len(paths))]
path_planner.n_frontier = tmp
else:
reward = []
paths = path_planner.make_paths(self.X_pose)
max_length = np.max([p.path_length() for p in paths])
self.inputs.set_bounds([t_cur, t_cur+max_length])
acq = check_acquisition(acquisition, self.model, self.inputs)
for p in paths:
qs, ts = path_planner.make_itinerary(p,200)
qs, ts = np.array(qs), np.array(ts)
x_eval = np.hstack(( qs[:,0:2], t_cur+ts[:,None] ))
ac = acq.evaluate(x_eval)
reward.append(np.trapz(ac.squeeze(),ts))
popt = paths[np.argmin(reward)]
t_int = t_cur + popt.path_length()
time_stamps = record_time[ (record_time > t_cur) & \
(record_time <= t_int) ]
samples = []
for tt in time_stamps:
samples.append(popt.sample(tt-t_cur))
coords = np.atleast_2d(samples)[:,0:2]
xopt = np.hstack((coords, time_stamps[:,None]))
yopt = np.empty((xopt.shape[0],1))
for jj in range(xopt.shape[0]):
self.my_map.kwargs["time"] = time_stamps[jj]
yopt[jj] = self.my_map.evaluate(coords[jj])
t_cur = t_int
self.X_pose = popt.path_endpoint()
self.X = np.vstack((self.X, xopt))
self.Y = np.vstack((self.Y, yopt))
self.model.set_XY(self.X, self.Y)
self.model.optimize_restarts(**kwargs_GPy)
m_list.append(self.model.copy())
p_list.append(popt)
if ii == 0:
s_list = [ self.model.copy() ]
s_list.extend([self.model.copy()]*len(yopt))
if callback:
self._callback(ii, time.time()-tic)
if save_iter:
self.model.save_model(filename)
return m_list, p_list, s_list
@staticmethod
def _callback(ii, time):
m, s = divmod(time, 60)
print("Iteration {:3d} \t Optimization completed in {:02d}:{:02d}"
.format(ii, int(m), int(s)))
class OptimalPathStatic(object):
"""A class for Bayesian path-planning algorithm. This class assumes
that the objective function does not change with time.
Parameters
---------
X_pose : array_like
The robot's starting pose. Must be in the form
(x_pos, y_pos, angle)
Y_pose : array_like
Observation at starting position.
my_map: instance of `BlackBox`
The black-box objective function.
inputs : instance of `Inputs`
The input space.
fix_noise : boolean, optional
Whether or not to fix the noise variance in the GP model.
noise_var : float, optional
Variance for additive Gaussian noise. Default is None, in
which case the noise variance from BlackBox is used. If fix_noise
is False, noise_var is merely used to initialize the GP model.
normalize_Y : boolean, optional
Whether or not to normalize the output in the GP model.
Attributes
----------
my_map, inputs : see Parameters
X : array
Array of input points.
Y : array
Array of observations.
X_pose : array_like
The robot's current pose.
input_dim : int
Dimensionality of the input space
model : instance of `GPRegression`
Current GPy model.
"""
def __init__(self, X_pose, Y_pose, my_map, inputs, fix_noise=False,
noise_var=None, normalize_Y=True):
self.my_map = my_map
self.inputs = inputs
self.input_dim = inputs.input_dim
self.X = np.atleast_2d(X_pose[0:2])
self.Y = np.atleast_2d(Y_pose)
self.X_pose = X_pose
if noise_var is None:
noise_var = my_map.noise_var
# Currently only the RBF kernel is supported.
ker = RBF(input_dim=self.input_dim, ARD=True)
self.model = GPy.models.GPRegression(X=self.X,
Y=self.Y,
kernel=ker,
normalizer=normalize_Y,
noise_var=noise_var)
if fix_noise:
self.model.Gaussian_noise.variance.fix(noise_var)
def optimize(self, record_time, acquisition, path_planner,
callback=True, save_iter=True, prefix=None, kwargs_GPy=None):
"""Runs the Bayesian path-planning algorithm.
Parameters
----------
record_time : array_like
Time vector for when measurements are made.
acquisition : str or instance of `Acquisition`
Acquisition function for determining the next best point.
If a string, must be one of
- "PI": Probability of Improvement
- "EI": Expected Improvement
- "US": Uncertainty Sampling
- "US-BO": US repurposed for Bayesian Optimization (BO)
- "US-LW": Likelihood-Weighted US
- "US-LWBO": US-LW repurposed for BO
- "US-LWraw" : US-LW with no GMM approximation
- "US-LWBOraw": US-LWBO with no GMM approximation
- "LCB" : Lower Confidence Bound
- "LCB-LW" : Likelihood-Weighted LCB
- "LCB-LWraw" : LCB-LW with no GMM approximation
- "IVR" : Integrated Variance Reduction
- "IVR-IW" : Input-Weighted IVR
- "IVR-LW" : Likelihood-Weighted IVR
- "IVR-BO" : IVR repurposed for BO
- "IVR-LWBO": IVR-LW repurposed for BO
path_planner : instance of `PathPlanner`
Parametrization of the path by <NAME>.
callback : boolean, optional
Whether or not to display log at each iteration.
save_iter : boolean, optional
Whether or not to save the GP model at each iteration.
prefix : string, optional
Prefix for file naming
kwargs_GPy : dict, optional
Dictionary of arguments to be passed to the GP model.
Returns
-------
m_list : list
A list of trained GP models, one for each iteration of
the algorithm.
p_list : list
A list of paths, one for each iteration of the algorithm.
s_list : list
A list of trained GP models, one for each sample point.
"""
if prefix is None:
prefix = "model"
if kwargs_GPy is None:
kwargs_GPy = dict(num_restarts=10, optimizer="bfgs",
max_iters=1000, verbose=False)
m_list, p_list = [], []
t_cur = 0
t_fin = record_time[-1]
ii = -1
while t_cur <= t_fin:
ii += 1
tic = time.time()
filename = (prefix+"%.4d")%(ii)
if ii == 0:
# Densify frontier for truly uniform sampling
tmp = path_planner.n_frontier
path_planner.n_frontier = 1000
paths = path_planner.make_paths(self.X_pose)
popt = paths[np.random.randint(len(paths))]
path_planner.n_frontier = tmp
else:
if ii == 1:
acq = check_acquisition(acquisition, self.model,
self.inputs)
acq.model = self.model.copy()
acq.update_parameters()
reward = []
paths = path_planner.make_paths(self.X_pose)
for p in paths:
qs, ts = path_planner.make_itinerary(p,200)
qs, ts = np.array(qs), np.array(ts)
ac = acq.evaluate(qs[:,0:2])
reward.append(np.trapz(ac.squeeze(),ts))
popt = paths[np.argmin(reward)]
t_int = t_cur + popt.path_length()
time_stamps = record_time[ (record_time > t_cur) & \
(record_time <= t_int) ]
samples = []
for tt in time_stamps:
samples.append(popt.sample(tt-t_cur))
xopt = np.atleast_2d(samples)[:,0:2]
yopt = self.my_map.evaluate(xopt)
t_cur = t_int
self.X_pose = popt.path_endpoint()
self.X = np.vstack((self.X, xopt))
self.Y = np.vstack((self.Y, yopt))
self.model.set_XY(self.X, self.Y)
self.model.optimize_restarts(**kwargs_GPy)
m_list.append(self.model.copy())
p_list.append(popt)
if ii == 0:
s_list = [ self.model.copy() ]
s_list.extend([self.model.copy()]*len(yopt))
if callback:
self._callback(ii, time.time()-tic)
if save_iter:
self.model.save_model(filename)
return m_list, p_list, s_list
@staticmethod
def _callback(ii, time):
m, s = divmod(time, 60)
print("Iteration {:3d} \t Optimization completed in {:02d}:{:02d}"
.format(ii, int(m), int(s)))
|
<reponame>stephen-w-bailey/fast-n-deep-faces
import logging
try:
import maya.api.OpenMaya as om
import pymel
import pymel.core
usingMaya = True
except:
logging.warning('PoseGenerator not running in maya')
usingMaya = False
import functools
import numpy as np
import random
import socket
import struct
import time
class PoseGenerator(object):
def __init__(self,controlFile,geoFile):
self.loadControls(controlFile)
self.loadGeoNodes(geoFile)
self.sampler = None
# Read a file specifying the rig controls and the range in which
# they should be randomly set
# Each line is as follows:
# ControlName ControlType Index MinVAlue MaxValue
# Control type is either r-rotation, t-translation, or s-scale
# Index is either x, y, or z
def loadControls(self,fileName):
self.nodes = {}
self.numControls = 0
self.nameOrder = []
self.defaultPose = []
self.poseRange = []
self._active = None
with open(fileName) as file:
for line in file:
args = line.split(' ')
nodeName = args[0]
if len(args) < 6:
self.defaultPose.append(0) # Assume that the default value is 0 if not provided
else:
self.defaultPose.append(float(args[5]))
if nodeName not in self.nodes:
if usingMaya:
node = pymel.core.general.PyNode(nodeName)
else:
node = None
self.nodes[nodeName] = [node]
self.nameOrder.append(nodeName)
controlType = args[1]
if args[2].isdigit():
index = int(args[2])
else:
coords = ['x','y','z']
if args[2] in coords:
index = args[2]
else:
if usingMaya:
controlNode = pymel.core.general.PyNode(nodeName+'.'+args[2])
index = controlNode
else:
controlNode = None
index = -1
minValue = float(args[3])
maxValue = float(args[4])
self.nodes[nodeName].append((controlType,index,minValue,maxValue))
self.poseRange.append((minValue,maxValue))
self.numControls += 1
self.defaultPose = np.asarray(self.defaultPose)
def loadGeoNodes(self,geoFile):
if not usingMaya:
return
self.geoNodes = []
self.omGeoNodes = []
self.geoNames = []
self.useFull = []
with open(geoFile) as file:
for line in file:
name = line.strip()
if name[-1] == '*':
self.useFull.append(True)
name = name[:-1]
else:
self.useFull.append(False)
self.geoNames.append(name)
self.geoNodes.append(pymel.core.general.PyNode(name))
self.geoNodes[-1].select()
selectionLs = om.MGlobal.getActiveSelectionList()
selObj = selectionLs.getDagPath(0) # index 0 in selection
self.omGeoNodes.append(om.MFnMesh(selObj))
def setSampler(self,sampler):
self.sampler = sampler
def createRandomPose(self):
if self.sampler is None:
pose = []
for name in self.nameOrder:
nodeList = self.nodes[name]
for control in nodeList[1:]:
controlType,index,minValue,maxValue = control
val = random.random()*(maxValue-minValue)+minValue
pose.append(val)
return pose
else:
pose = self.sampler.getRandomPose()
return pose
def setRandomPose(self):
pose = self.createRandomPose()
self.setPose(pose)
return pose
def getControls(self,attrs):
res = {}
for attr in attrs:
res[attr] = attr.get()
return res
def setControls(self,attrs):
for attr in attrs:
attr.set(attrs[attr])
def setPose(self,pose):
if not usingMaya:
return
indexMap = {'x':0,'y':1,'z':2}
pose = [float(p) for p in pose]
for i in range(len(pose)):
pose[i] = min(max(pose[i],self.poseRange[i][0]),self.poseRange[i][1])
pose = iter(pose)
for name in self.nameOrder:
nodeList = self.nodes[name]
node = nodeList[0]
attrs = [i[1] for i in nodeList[1:] if not isinstance(i[1],str)]
nodeSetFunctions = {'r':node.setRotation if hasattr(node,'setRotation') else None,
't':node.setTranslation if hasattr(node,'setTranslation') else None,
'c':self.setControls}
nodeGetFunctions = {'r':node.getRotation if hasattr(node,'getRotation') else None,
't':node.getTranslation if hasattr(node,'getTranslation') else None,
'c':functools.partial(self.getControls,attrs)}
controls = {}
for control in nodeList[1:]:
controlType,index,minValue,maxValue = control
val = next(pose)
if controlType not in controls:
controls[controlType] = nodeGetFunctions[controlType]()
controls[controlType][indexMap[index] if isinstance(index,str) else index] = val
for control in controls:
nodeSetFunctions[control](controls[control])
def getVertices(self,index=None):
verts = []
if not usingMaya:
return None
if index is None:
for mesh,node in zip(self.omGeoNodes,self.geoNodes):
points = mesh.getPoints()
v = np.array(points)[:,:3]
T = np.array(node.getMatrix())
R = T[:3,:3]
t = T[3,:3]
v = v.dot(R)+t
verts.append(v)
else:
mesh = self.omGeoNodes[index]
points = mesh.getPoints()
verts.append(np.array(points)[:,:3])
return np.concatenate(verts,0)
# Returns a boolean mask for each control indicating if the control moves
# each vertex in the mesh
# n is the number of samples to make per control to estimate the influence
def getControlInfluence(self,n=16):
if self._active is None:
self.computeActiveVertices()
mesh = self.getVertices()[self.active]
masks = np.zeros((self.numControls,len(mesh)),dtype='bool')
# Run through each control separately
eps = 1e-6
for i in range(self.numControls):
p = np.stack([self.createRandomPose() for _ in range(n+1)],0)
self.setPose(p[0])
base = self.getVertices()[self.active]
newP = np.repeat(p[[0]],n,0)
newP[:,i] = p[1:,i]
meshes = []
for pose in newP:
self.setPose(pose)
meshes.append(self.getVertices()[self.active])
meshes = np.stack(meshes,0)
diff = np.sum(np.square(base-meshes),-1)
diff = np.mean(diff,0)
masks[i] = diff>eps
return masks
def getEdges(self,useActive=True):
if len(self.geoNames) > 1:
raise NotImplementedError('Cannot get edges for multiple meshes')
verts = self.getVertices()
idx = np.arange(len(verts))
idxMap = idx.copy()
if useActive:
if not hasattr(self,'active'):
self.computeActiveVertices()
idx = idx[self.active]
idxMap[:] = -1
idxMap[self.active] = np.arange(len(idx))
logging.info('Finding edges from '+str(len(idx))+' vertices')
# Get the vertices
baseName = self.geoNames[0]+'.vtx'
vs = []
for i in idx:
name = baseName+'['+str(i)+']'
vs.append(pymel.core.general.PyNode(name))
# Get the edges
es = pymel.core.modeling.polyListComponentConversion(vs,fv=True,te=True)
es = [pymel.core.general.PyNode(edge) for edge in es]
# Get the connected verties
e = []
for edges in es:
itr = iter(edges)
for edge in itr:
a,b = edge.connectedVertices()
a,b = idxMap[a.index()],idxMap[b.index()]
if a == -1 or b == -1:
continue
e.append((min(a,b),max(a,b)))
return e
def getFacesOnMesh(self,meshIdx,useActive=True):
#if len(self.geoNames) > 1:
# raise NotImplementedError('Cannot get edges for multiple meshes')
verts = self.getVertices(meshIdx)
idx = np.arange(len(verts))
idxMap = idx.copy()
if useActive:
if self._active is None:
self.computeActiveVertices()
idx = idx[self._active[meshIdx]]
idxMap[:] = -1
idxMap[self._active[meshIdx]] = np.arange(len(idx))
logging.info('Finding faces from '+str(len(idx))+' vertices')
# Get the vertices
baseName = self.geoNames[meshIdx]+'.vtx'
vs = []
for i in idx:
name = baseName+'['+str(i)+']'
vs.append(pymel.core.general.PyNode(name))
# Get the faces
fs = pymel.core.modeling.polyListComponentConversion(vs,fv=True,tf=True)
fs = [pymel.core.general.PyNode(face) for face in fs]
# Get the vertices on the faces
f = []
for faces in fs:
itr = iter(faces)
for face in itr:
vs = face.getVertices()
vs = [idxMap[v] for v in vs]
if any([v==-1 for v in vs]):
continue # Found a face with a vertex not being used
if len(vs) == 3:
f.append(vs)
elif len(vs) == 4:
f.append([vs[0],vs[1],vs[2]])
f.append([vs[2],vs[3],vs[0]])
else:
logging.warning('Face with '+str(len(vs))+' vertices encountered, ignoring face')
return f
def getFaces(self,useActive=True):
fs = []
if self._active is None:
self.computeActiveVertices()
for i in range(len(self.geoNames)):
f = np.asarray(self.getFacesOnMesh(i,useActive))
fs.append(f)
fullF = []
count = 0
for f,a in zip(fs,self._active):
if len(f) == 0:
continue
fullF.append(f+count)
if useActive:
count += np.sum(a)
else:
count += len(a)
f = np.concatenate(fullF,0)
return f
def getUVIndexOnMesh(self,meshIndex,useActive=True):
#if len(self.geoNames) > 1:
# raise NotImplementedError('Cannot get edges for multiple meshes')
verts = self.getVertices(meshIndex)
idx = np.arange(len(verts))
idxMap = idx.copy()
if useActive:
if self._active is None:
self.computeActiveVertices()
idx = idx[self._active[meshIndex]]
idxMap[:] = -1
idxMap[self._active[meshIndex]] = np.arange(len(idx))
logging.info('Finding faces from '+str(len(idx))+' vertices')
# Get the vertices
baseName = self.geoNames[meshIndex]+'.vtx'
vs = []
for i in idx:
name = baseName+'['+str(i)+']'
vs.append(pymel.core.general.PyNode(name))
# Get the faces
fs = pymel.core.modeling.polyListComponentConversion(vs,fv=True,tf=True)
fs = [pymel.core.general.PyNode(face) for face in fs]
# Get the uv indices on the faces
uv = []
uv = -np.ones(np.sum(idxMap>-1),dtype='int32')
for faces in fs:
itr = iter(faces)
for face in itr:
uvs = len(face.getUVs()[0])
uvs = [face.getUVIndex(i) for i in range(uvs)]
vs = face.getVertices()
vs = [idxMap[v] for v in vs]
for v,u in zip(vs,uvs):
uv[v] = u
"""if any([v==-1 for v in vs]):
continue # Found a face with a vertex not being used
if len(uvs) == 3:
uv.append(uvs)
elif len(uvs) == 4:
uv.append([uvs[0],uvs[1],uvs[2]])
uv.append([uvs[2],uvs[3],uvs[0]])
else:
logging.warning('Face with '+str(len(uvs))+' vertices encountered, ignoring face')"""
return uv
def getUVIndex(self,useActive=True):
uvs = []
if self._active is None:
self.computeActiveVertices()
for i in range(len(self.geoNames)):
uv = np.asarray(self.getUVIndexOnMesh(i,useActive))
uvs.append(uv)
fullUV = []
count = 0
for u,mesh,a in zip(uvs,self.geoNodes,self._active):
uvMesh = mesh.getUVs()
uvMesh = [list(i) for i in uvMesh]
uvMesh = np.asarray(uvMesh).T
if len(uvMesh) == 0:
continue
fullUV.append(u+count)
count += len(uvMesh)
uv = np.concatenate(fullUV,0)
return uv
def getUVs(self):
#if len(self.geoNames) > 1:
# raise NotImplementedError('Cannot get edges for multiple meshes')
uvs = []
for mesh in self.geoNodes:
uv = mesh.getUVs()
uv = [list(u) for u in uv]
uv = np.asarray(uv).T
uvs.append(uv)
uvs = np.concatenate(uvs,0)
return uvs
def computeActiveVertices(self, reps=25):
v0 = self.getVertices()
eps = 1e-6
active = np.zeros(len(v0),dtype=bool)
for _ in range(reps):
self.setRandomPose()
v = self.getVertices()
diff = np.sum(np.square(v-v0),-1)
active = np.logical_or(active,diff>eps)
self.active = active
@property
def active(self):
return np.concatenate(self._active,0)
@active.setter
def active(self,value):
verts = []
for mesh in self.omGeoNodes:
points = mesh.getPoints()
verts.append(np.array(points)[:,:3])
self._active = []
for v,useFull in zip(verts,self.useFull):
a = value[:len(v)].copy()
if useFull:
a[:] = True
self._active.append(a)
value = value[len(v):]
def generateBatch(self, n=256):
if not hasattr(self,'active'):
self.computeActiveVertices()
startTime = time.time()
setTime = 0
meshTime = 0
mesh = np.zeros((n,np.sum(self.active),3))
pose = np.zeros((n,self.numControls))
for i in range(n):
startSet = time.time()
pose[i] = self.setRandomPose()
endSet = time.time()
setTime += endSet-startSet
mesh[i] = self.getVertices()[self.active]
endMesh = time.time()
meshTime += endMesh-endSet
endTime = time.time()
logging.info('Time to generate poses: '+str(endTime-startTime)+' seconds')
logging.info('Time to set poses: '+str(setTime)+' seconds')
logging.info('Time to get mesh: '+str(meshTime)+' seconds')
return pose,mesh
class PoseGeneratorRemote(PoseGenerator):
def __init__(self,controlFile,geoFile,host,port,isServer=False):
super(PoseGeneratorRemote,self).__init__(controlFile,geoFile)
self.host = host
self.port = port
self.isServer = isServer
self.client = None
@property
def active(self):
return self._active
def connect(self):
# Create the connection
client = socket.socket()
if self.isServer:
logging.info('Running PoseGeneratorRemote as server')
client.bind(('',self.port))
client.listen(1)
connection,address = client.accept()
self.server = client
client = connection
else:
try:
client.connect((self.host,self.port))
except socket.error:
logging.info('Connection to '+self.host+':'+str(self.port)+' failed')
return
self.client = client
def setPose(self,pose):
pose = np.asarray(pose).astype('float32')
command = b'p' + packMatrix(pose.reshape(-1))
sendMessage(command,self.client)
def getVertices(self):
command = b'v'
sendMessage(command,self.client)
msg = receiveMessage(self.client)
return unpackMatrix(msg)
def computeActiveVertices(self, reps=5):
command = b'a'
sendMessage(command,self.client)
msg = receiveMessage(self.client)
active = unpackMatrix(msg)
self._active = active.astype('bool')
def getEdges(self,useActive=True):
if useActive:
command = b'eA'
else:
command = b'e'
sendMessage(command,self.client)
msg = receiveMessage(self.client)
return unpackMatrix(msg)
def getFaces(self,useActive=True):
if useActive:
command = b'fA'
else:
command = b'f'
sendMessage(command,self.client)
msg = receiveMessage(self.client)
return unpackMatrix(msg)
def getUVs(self):
command = b'u'
sendMessage(command,self.client)
msg = receiveMessage(self.client)
return unpackMatrix(msg)
def getUVIndex(self,useActive=True):
if useActive:
command = b'UA'
else:
command = b'U'
sendMessage(command,self.client)
msg = receiveMessage(self.client)
return unpackMatrix(msg)
def setActiveVertices(self,active):
active = active.astype('int32')
command = b'A'+packMatrix(active)
sendMessage(command,self.client)
self._active = active.astype('bool')
def close(self):
command = b'c'
sendMessage(command,self.client)
self.client.close()
self.client = None
if self.isServer:
self.server.close()
def sendMessage(data,connection):
if connection is None:
raise RuntimeError('Connection does not exist. Is PoseGeneratorServer running?')
if not isinstance(data,bytes):
raise ValueError('Cannot send data of type '+str(type(data)))
length = struct.pack('<Q',len(data))
buff = length+data
connection.sendall(buff)
def receiveMessage(connection):
length = connection.recv(8)
length = struct.unpack('<Q',length)[0]
data = connection.recv(length)
while len(data) != length:
data += connection.recv(length-len(data))
return data
def packMatrix(matrix):
dtypes = {np.dtype('float32'):b'f',
np.dtype('float64'):b'd',
np.dtype('int32'):b'i'}
dtype = ''
for t in dtypes:
if matrix.dtype is t:
dtype = dtypes[t]
break
if dtype == '':
raise ValueError('Cannot handle matrix dtype '+str(matrix.dtype))
shape = matrix.shape
data = struct.pack('c',dtype)
data += struct.pack('B',len(shape))
data += struct.pack('<'+'I'*len(shape),*shape)
data += matrix.tostring()
return data
def unpackMatrix(data):
dtypes = {b'f':np.dtype('float32'),
b'd':np.dtype('float64'),
b'i':np.dtype('int32')}
dtype = struct.unpack('c',data[0:1])[0]
shapeLength = struct.unpack('<B',data[1:2])[0]
shape = struct.unpack('<'+'I'*shapeLength,data[2:2+shapeLength*4])
matrix = np.frombuffer(data[2+shapeLength*4:],dtype=dtypes[dtype]).reshape(shape)
return matrix
class PoseGeneratorServer:
def __init__(self,controlFile,geoFile,port=9001,isServer=True):
if usingMaya:
self.generator = PoseGenerator(controlFile,geoFile)
else:
self.generator = None
self.port = port
self.isServer = isServer
def startServer(self,hostname=None):
server = socket.socket()
if self.isServer:
server.bind(('',self.port))
server.listen(1)
connection,address = server.accept()
else:
logging.info('Running PoseGeneratorServer as client')
server.connect((hostname,self.port))
connection = server
while True:
buff = receiveMessage(connection)
if not buff:
break
if not self.processBuffer(buff,connection):
break
connection.close()
server.close()
def processBuffer(self,buff,connection):
commandNames = {'A':'setActiveVertices',
'a':'getActiveVertices',
'e':'getEdges',
'f':'getFaces',
'p':'setPose',
'u':'getUVs',
'U':'getUVIndex',
'v':'getVertices',
'c':'close'}
commands = {'A':self.setActiveVertices,
'a':self.getActiveVertices,
'e':self.getEdges,
'f':self.getFaces,
'p':self.setPose,
'u':self.getUVs,
'U':self.getUVIndex,
'v':self.getVertices,
'c':self.close}
command = buff[0:1].decode('utf-8')
if command not in commands:
logging.warning('Received unknown command '+str(command))
return False
logging.info('Received command '+commandNames[command])
return commands[command](buff[1:],connection)
def close(self,buff,connection):
return False
def setPose(self,buff,connection):
pose = list(unpackMatrix(buff).reshape(-1))
self.generator.setPose(pose)
return True
def getVertices(self,buff,connection):
v = self.generator.getVertices().astype('float32')
data = packMatrix(v)
sendMessage(data,connection)
return True
def getEdges(self,buff,connection):
if len(buff) > 0:
useActive = buff[0:1].decode('utf-8')
useActive = useActive=='A'
else:
useActive = False
e = self.generator.getEdges(useActive=useActive)
e = np.asarray(e).astype('int32')
data = packMatrix(e)
sendMessage(data,connection)
return True
def getFaces(self,buff,connection):
if len(buff) > 0:
useActive = buff[0:1].decode('utf-8')
useActive = useActive=='A'
else:
useActive = False
f = self.generator.getFaces(useActive=useActive)
f = np.asarray(f).astype('int32')
data = packMatrix(f)
sendMessage(data,connection)
return True
def getUVs(self,buff,connection):
uv = self.generator.getUVs()
uv = np.asarray(uv).astype('float32')
data = packMatrix(uv)
sendMessage(data,connection)
return True
def getUVIndex(self,buff,connection):
if len(buff) > 0:
useActive = buff[0:1].decode('utf-8')
useActive = useActive=='A'
else:
useActive = False
uv = self.generator.getUVIndex(useActive=useActive)
uv = np.asarray(uv).astype('int32')
data = packMatrix(uv)
sendMessage(data,connection)
return True
def getActiveVertices(self,buff,connection):
if not hasattr(self.generator,'active'):
self.generator.computeActiveVertices()
active = self.generator.active.astype('int32')
data = packMatrix(active)
sendMessage(data,connection)
return True
def setActiveVertices(self,buff,connection):
active = unpackMatrix(buff).astype('bool')
self.generator.active = active
return True
|
#!/usr/bin/env python
# coding: utf-8
# # QDA + Pseudo Labeling + Gaussian Mixture = LB 0.975
# The dataset for Kaggle competition "Instant Gratification" appears to be 512 datasets concatenated where each sub dataset is believed to be created by Sklearn's `make_classification`. EDA suggests the following parameters:
#
# X, y = make_classification(n_samples=1024, n_features=255, n_informative=33+x,
# n_redundant=0, n_repeated=0, n_classes=2, n_clusters_per_class=3,
# weights=None, flip_y=0.05, class_sep=1.0, hypercube=True, shift=0.0,
# scale=1.0, shuffle=True, random_state=None) # where 0<=x<=14
#
# The important parameters to note are `n_clusters_per_class=3` and `n_informative=33+x`. This means that the data resides in `33+x` dimensional space within 6 hyper-ellipsoids. Each hyper-ellipsoid is a multivariate Gaussian distribution therefore the best classifiers to use are QDA, Pseudo Labeling, and Gaussian Mixture. (See appendix for EDA showing 3 clusters per class).
#
# 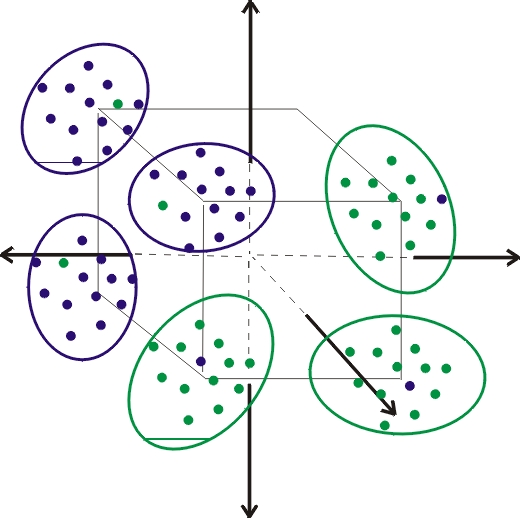
#
# # Load Libraries and Data
# In[1]:
# IMPORT LIBRARIES
import numpy as np, pandas as pd, os
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.mixture import GaussianMixture
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import VarianceThreshold
from sklearn.metrics import roc_auc_score
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
# In[2]:
# LOAD TRAIN AND TEST
df_train = pd.read_csv('../input/instant-gratification/train.csv')
df_test = pd.read_csv('../input/instant-gratification/test.csv')
sample = pd.read_csv('../input/instant-gratification/sample_submission.csv')
# In[3]:
df_train.info()
# In[4]:
df_test.info()
# In[5]:
df_train.head()
# In[6]:
df_test.head()
# In[7]:
print("Train shape: {}, Test shape: {}".format(df_train.shape, df_test.shape))
# Interactions
#
# Below shows the interactions between wheezy-copper-turtle-magic and the other variables. Each variable by itself cannot predict target well, but when wheezy-copper-turtle-magic equals a specific value then other variables can predict target well. For example, when wheezy-copper-turtle-magic = 0 then zippy-harlequin-otter-grandmaster is positively correlated with target. And when wheezy-copper-turtle-magic = 0 then hasty-puce-fowl-fepid is negatively correlated with target.
# In[8]:
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
plt.figure(figsize=(15,5))
# PLOT ALL ZIPPY
plt.subplot(1,2,1)
sns.distplot(df_train[ (df_train['target']==0) ]['zippy-harlequin-otter-grandmaster'], label = 't=0')
sns.distplot(df_train[ (df_train['target']==1) ]['zippy-harlequin-otter-grandmaster'], label = 't=1')
plt.title("Without interaction, zippy has no correlation \n (showing all rows)")
plt.xlim((-5,5))
plt.legend()
# PLOT ZIPPY WHERE WHEEZY-MAGIC=0
plt.subplot(1,2,2)
sns.distplot(df_train[ (df_train['wheezy-copper-turtle-magic']==0) & (df_train['target']==0) ]
['zippy-harlequin-otter-grandmaster'], label = 't=0')
sns.distplot(df_train[ (df_train['wheezy-copper-turtle-magic']==0) & (df_train['target']==1) ]
['zippy-harlequin-otter-grandmaster'], label = 't=1')
plt.title("With interaction, zippy has postive correlation \n (only showing rows where wheezy-copper-turtle-magic=0)")
plt.legend()
plt.show()
# In[9]:
plt.figure(figsize=(15,5))
# PLOT ALL HASTY
plt.subplot(1,2,1)
sns.distplot(df_train[ (df_train['target']==0) ]['hasty-puce-fowl-fepid'], label = 't=0')
sns.distplot(df_train[ (df_train['target']==1) ]['hasty-puce-fowl-fepid'], label = 't=1')
plt.title("Without interaction, hasty has no correlation \n (showing all rows)")
plt.xlim((-5,5))
plt.legend()
# PLOT HASTY WHERE WHEEZY-MAGIC=0
plt.subplot(1,2,2)
sns.distplot(df_train[ (df_train['wheezy-copper-turtle-magic']==0) & (df_train['target']==0) ]
['hasty-puce-fowl-fepid'], label = 't=0')
sns.distplot(df_train[ (df_train['wheezy-copper-turtle-magic']==0) & (df_train['target']==1) ]
['hasty-puce-fowl-fepid'], label = 't=1')
plt.title("With interaction, hasty has negative correlation \n (only showing rows where wheezy-copper-turtle-magic=0)")
plt.legend()
plt.show()
# In[14]:
df_train = pd.read_csv('../input/instant-gratification/train.csv')
df_test = pd.read_csv('../input/instant-gratification/test.csv')
# Structure of Entire Dataset
#
# The dataset for Kaggle's "Instant Gratification" competition appears to be 512 partial datasets combined as confirmed by experiments and EDA. Consequently each variable (column) appears to be the sum of 512 Gaussians. Below is a sketch of the dataset sorted by the column wheezy-copper-turtle-magic (using 1-based indexing).
#
# 
#
# In the dataset drawing above, column one is the summation of Gaussians DATA-1-1, DATA-2-1, and DATA-3-1. If you plot these histograms separately, you see that they are Gaussians.
#
# 
#
# **Structure of Partial Datasets**
#
# **There are 512 unique values of wheezy-copper-turtle-magic and there are 262144 rows of data. For each value of magic there are approximately 512 = 262144/512 associated rows that form a partial dataset.**
#
#
# **These partial datasets each have (approx) 512 rows and 255 features. We can model each partial dataset individually** and this can achieve high overall accuracy of at least LB 0.928 as shown here. (Predictions on test.csv are made using the appropriate model). The features for model one are DATA-1-1, DATA-1-2, DATA-1-3, etc. Each feature has a Gaussian distribution. When building a model with only 512 observations and 255 features, it is important to reduce the number of features to prevent overfitting.
# Determine structure of private dataset
#
# As described here, **both the train and public test dataset appear to be 512 datasets combined. Each partial dataset has 256 variables. Therefore there are 131072 = 512 * 216 blocks of data. Each block of data has either standard deviation approx 1.0 or approx 3.75. The blocks with standard deviation 1.0 seem to be useless while the blocks with standard deviation 3.75 seem to be useful.** Does the private test dataset have this same structure?
#
# (There are 512 partial datasets within the full dataset. **And each partial dataset has a different set of approximately 40 important features as identified by different standard deviations.**) We have observed that both the training dataset and public test dataset have this structure. Does the private dataset also have this structure? In this kernel, we probe the private dataset and confirm that it has the same structure.
# In[15]:
print("Train shape: {}, Test shape: {}".format(df_train.shape, df_test.shape))
# In[16]:
df_train.describe()
# In[17]:
df_test.head()
# In[18]:
sample.head()
# # Identify Useful Features
# the data appears to be 131072 = 512 * 256 blocks of data where some are useful and some are useless.
# In[19]:
# IDENTIFY USEFUL FEATURES PER MAGIC SUB-DATASET
useful = np.zeros((256,512))
for i in range(512):
partial = df_train[ df_train['wheezy-copper-turtle-magic']==i ]
useful[:,i] = np.std(partial.iloc[:,1:-1], axis=0)
useful = useful > 1.5
# In[20]:
import matplotlib.pyplot as plt
# PLOT BOOLEANS OF USEFUL BLOCKS
plt.figure(figsize=(10,20))
plt.matshow(useful.transpose(),fignum=1)
plt.title('The useful datablocks of dataset', fontsize=24)
plt.xlabel('256 Variable columns', fontsize=16)
plt.ylabel('512 Partial datasets', fontsize=16)
plt.show()
# In[21]:
useful = np.sum( useful, axis=0 )
# What is Pseudo Labeling?
#
# Pseudo labeling is the process of adding confident predicted test data to your training data. Pseudo labeling is a 5 step process. (1) Build a model using training data. (2) Predict labels for an unseen test dataset. (3) Add confident predicted test observations to our training data (4) Build a new model using combined data. And (5) use your new model to predict the test data and submit to Kaggle. Here is a pictorial explanation using sythetic 2D data.
# Why does Pseudo Labeling work?
#
# Pseudo labeling helps all types of models because all models can be visualized as finding shapes of target=1 and target=0 in p-dimensional space. See here for examples. More points allow for better estimation of shapes.
# # Model and Predict
# 1. First we use QDA plus Pseudo Labeling
# 2. Next we will use these predictions (pseudo labels) to find the 6 ellipses. We separately find the 3 ellipses of the target=1 data and 3 ellipses of the target=0 data using Sklearn GaussianMixture.
# 3. Then we label each point with 0, 1, 2, 3, 4, 5 representing which ellipse it belongs to.
# 4. Finally we train QDA on these 6 ellipses and use QDA to make our final predictions with `Pr(target=1) = Pr(in ellipse 3) + Pr(in ellipse 4) + Pr(in ellipse 5)`. (See appendix 2 for advanced techniques).
#
# For validation, we didn't use typical k-folds CV. Instead we created synthetic data and optimized our technique on synthetic data. This has proven to be more reliable than CV. Also it allows our model to use all 1024 rows of sub datasets when building models. Our model has demonstrated that on average it can perfectly classify Sklearn's `make_classification` data. However many other participants can too. So randomness is added which allows us to do better than perfect sometimes. Then two random versions' output were submitted to Kaggle and hopefully those are the high scoring ones! :P
#
# When the code below is run locally, it generates synthetic data and calculates validation AUC. When the code is submitted to Kaggle, it uses real data and predicts `test.csv`.
#
# [1]: https://www.kaggle.com/cdeotte/pseudo-labeling-qda-0-969
# In[22]:
# RUN LOCALLY AND VALIDATE
models = 512
RunLocally = True
# RUN SUBMITTED TO KAGGLE
if len(df_test)>512*300:
repeat = 1
models = 512 * repeat
RunLocally = False
# INITIALIZE
all_preds = np.zeros(len(df_test))
all_y_pu = np.array([])
all_y_pr = np.array([])
all_preds_pu = np.array([])
all_preds_pr = np.array([])
# MODEL AND PREDICT
for k in range(models):
# IF RUN LOCALLY AND VALIDATE
# THEN USE SYNTHETIC DATA
if RunLocally:
obs = 512
X, y = make_classification(n_samples=1024, n_features=useful[k%512],
n_informative=useful[k%512], n_redundant=0, n_repeated=0,
n_classes=2, n_clusters_per_class=3, weights=None, flip_y=0.05,
class_sep=1.0, hypercube=True, shift=0.0, scale=1.0, shuffle=True,
random_state=None)
# IF RUN SUBMITTED TO KAGGLE
# THEN USE REAL DATA
else:
df_train2 = df_train[df_train['wheezy-copper-turtle-magic']==k%512]
df_test2 = df_test[df_test['wheezy-copper-turtle-magic']==k%512]
sel = VarianceThreshold(1.5).fit(df_train2.iloc[:,1:-1])
df_train3 = sel.transform(df_train2.iloc[:,1:-1])
df_test3 = sel.transform(df_test2.iloc[:,1:])
obs = df_train3.shape[0]
X = np.concatenate((df_train3,df_test3),axis=0)
y = np.concatenate((df_train2['target'].values,np.zeros(len(df_test2))))
# TRAIN AND TEST DATA
train = X[:obs,:]
train_y = y[:obs]
test = X[obs:,:]
test_y = y[obs:]
comb = X
# FIRST MODEL : QDA
clf = QuadraticDiscriminantAnalysis(priors = [0.5,0.5])
clf.fit(train,train_y)
test_pred = clf.predict_proba(test)[:,1]
# SECOND MODEL : PSEUDO LABEL + QDA
test_pred = test_pred > np.random.uniform(0,1,len(test_pred)) #randomness
clf = QuadraticDiscriminantAnalysis(priors = [0.5, 0.5])
clf.fit(comb, np.concatenate((train_y,test_pred)) )
test_pred = clf.predict_proba(test)[:,1]
# THIRD MODEL : PSEUDO LABEL + GAUSSIAN MIXTURE
test_pred = test_pred > np.random.uniform(0,1,len(test_pred)) #randomness
all_y = np.concatenate((train_y,test_pred))
least = 0; ct = 1; thx=150
while least<thx:
# STOPPING CRITERIA
if ct>=10: thx -= 10
else: thx = 150
# FIND CLUSTERS
clusters = np.zeros((len(comb),6))
# FIND THREE TARGET=1 CLUSTERS
train4 = comb[ all_y==1, :]
clf = GaussianMixture(n_components=3).fit(train4) #randomness
clusters[ all_y==1, 3:] = clf.predict_proba(train4)
# FIND THREE TARGET=0 CLUSTERS
train4 = comb[ all_y==0, :]
clf = GaussianMixture(n_components=3).fit(train4) #randomness
clusters[ all_y==0, :3] = clf.predict_proba(train4)
# ADJUST CLUSTERS (EXPLAINED IN KERNEL COMMENTS)
for j in range(5): clusters[:,j+1] += clusters[:,j]
rand = np.random.uniform(0,1,clusters.shape[0])
for j in range(6): clusters[:,j] = clusters[:,j]>rand #randomness
clusters2 = 6 - np.sum(clusters,axis=1)
# IF IMBALANCED TRY AGAIN
least = pd.Series(clusters2).value_counts().min(); ct += 1
# FOURTH MODEL : GAUSSIAN MIXTURE + QDA
clf = QuadraticDiscriminantAnalysis(priors = [0.167, 0.167, 0.167, 0.167, 0.167, 0.167])
clf.fit(comb,clusters2)
pds = clf.predict_proba(test)
test_pred = pds[:,3]+pds[:,4]+pds[:,5]
# IF RUN LOCALLY, STORE TARGETS AND PREDS
if RunLocally:
all_y_pu = np.append(all_y_pu, test_y[:256])
all_y_pr = np.append(all_y_pr, test_y[256:])
all_preds_pu = np.append(all_preds_pu, test_pred[:256])
all_preds_pr = np.append(all_preds_pr, test_pred[256:])
# IF RUN SUBMIT TO KAGGLE, PREDICT TEST.CSV
else:
all_preds[df_test2.index] += test_pred / repeat
# PRINT PROGRESS
if ((k+1)%64==0)|(k==0): print('modeled and predicted',k+1,'magic sub datasets')
# IF RUN LOCALLY, COMPUTE AND PRINT VALIDATION AUCS
if RunLocally:
all_y_pu_pr = np.concatenate((all_y_pu,all_y_pr))
all_preds_pu_pr = np.concatenate((all_preds_pu,all_preds_pr))
auc1 = roc_auc_score(all_y_pu_pr, all_preds_pu_pr)
auc2 = roc_auc_score(all_y_pu, all_preds_pu)
auc3 = roc_auc_score(all_y_pr, all_preds_pr)
print()
print('Validation AUC =',np.round(auc1,5))
print('Approx Public LB =',np.round(auc2,5))
print('Approx Private LB =',np.round(auc3,5))
# # Submit to Kaggle
# Alright, let's cross our fingers and hope that our submission has a high private LB !!
# In[23]:
sub = pd.read_csv('../input/instant-gratification/sample_submission.csv')
sub['target'] = all_preds
sub.to_csv('submission.csv',index=False)
plt.hist( test_pred ,bins=100)
plt.title('Model 512 test predictions')
plt.show()
# # Appendix 1 - EDA revealing n_clusters_per_class = 3
# We believe the data was made from Sklearn's `make_classification`. An important question is what parameter for `n_clusters_per_class` did Kaggle use?
#
# According to Sklearn's documentation [here][1]:
#
# > This initially creates clusters of points normally distributed (std=1) about vertices of an n-informative-dimensional hypercube with sides of length 2 * class_sep and assigns an equal number of clusters to each class.
#
# In three dimensions that means, that the clusters will be centered at one of these 8 locations: (-1, -1, -1), (-1, -1, 1), (-1, 1, -1), (-1, 1, 1), (1, -1, -1), (1, -1, 1), (1, 1, -1), (1, 1, 1) where you replace all 1's by `class_sep`. If you create 1024 rows of data and have 2 clusters per class, then for `target=1`, you may have 256 points centered at (-1, 1, -1) and 256 points centered at (1, 1, -1). Then for `target=0`, you may have 256 points centered at (1, 1, 1) and 256 points centered at (-1, -1, 1).
#
# Using EDA, we can determine the number of clusters per class of the real data. Sklearn's `make_classification` generates data (ellipses) at hypercube corners. Therefore if there is only `n_clusters_per_class=1` then the center of each ellipse (target=1 and target=0) of data will have all coordinates 1's and -1's, for example (1,1,-1,1,-1,1,...). So if we plot a histogram of all the variables' means (center coordinates), we will see a bump at 1 and -1. (By variable we mean within each sub dataset. That's 768 rows of train and public test. We don't mean all 262144 rows of original train columns).
#
# If `n_clusters_per_class=2`, then within one sub dataset there will be 2 ellipses for target=1. For example, there may be one ellipse centered at (-1,1,...) and one at (1,1,...) and the first coordinates of 1 and -1 will average to 0 when we compute that variable's mean. Therefore, if `clusters=2`, we will see histogram bumps at -1, 0, and 1. If `n_clusters_per_class=3`, we will see 4 bumps. Etc, etc. We can confirm this with synthetic data. We will use `n_samples=768` because we only have training and public test data to compare with which only has 768 rows per `wheezy-magic` sub dataset.
#
# Afterward, we will plot a histogram of the real data's variable means (within sub datasets) and see which `n_clusters_per_class` it matches. Alternatively, we can build a model and assume that `n_clusters_per_class` equals 1, 2, 3, or 4. Then we check which `n_clusters_per_class` has the greatest CV. Both of these methods determine that `n_clusters_per_class=3`.
#
# [1]: https://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_classification.html
# In[27]:
for clusters in range(4):
centers = np.array([])
for k in range(512):
X, y = make_classification(n_samples=768, n_features=useful[k],
n_informative=useful[k], n_redundant=0, n_repeated=0,
n_classes=2, n_clusters_per_class=clusters+1, weights=None,
flip_y=0.05, class_sep=1.0, hypercube=True, shift=0.0, scale=1.0,
shuffle=True, random_state=None)
centers = np.append(centers,np.mean(X[ np.argwhere(y==0).flatten() ,:],axis=0))
centers = np.append(centers,np.mean(X[ np.argwhere(y==1).flatten() ,:],axis=0))
plt.hist(centers,bins=100)
plt.title('Variable means if clusters='+str(clusters+1))
plt.show()
# ## Now let's plot the real data
# First we will use QDA to create pseudo labels for the public test data. Then we will plot a histogram of the variable means (data centers' coordinates) of target=0 and target=1 using all training and pseudo labeled public test data combined (768 rows per sub dataset). The plot below shows that Kaggle used `n_clusters_per_class=3`.
# In[28]:
centers = np.array([])
for k in range(512):
# REAL DATA
df_train2 = df_train[df_train['wheezy-copper-turtle-magic']==k]
df_test2 = df_test[df_test['wheezy-copper-turtle-magic']==k]
sel = VarianceThreshold(1.5).fit(df_train2.iloc[:,1:-1])
df_train3 = sel.transform(df_train2.iloc[:,1:-1])
df_test3 = sel.transform(df_test2.iloc[:,1:])
obs = df_train3.shape[0]
X = np.concatenate((df_train3,df_test3),axis=0)
y = np.concatenate((df_train2['target'].values,np.zeros(len(df_test2))))
# TRAIN AND TEST DATA
train = X[:obs,:]
train_y = y[:obs]
test = X[obs:,:]
test_y = y[obs:]
comb = X
# FIRST MODEL : QDA
clf = QuadraticDiscriminantAnalysis(priors = [0.5,0.5])
clf.fit(train,train_y)
test_pred = clf.predict_proba(test)[:,1]
# SECOND MODEL : PSEUDO LABEL + QDA
test_pred = test_pred > np.random.uniform(0,1,len(test_pred))
clf = QuadraticDiscriminantAnalysis(priors = [0.5, 0.5])
clf.fit(comb, np.concatenate((train_y,test_pred)) )
test_pred = clf.predict_proba(test)[:,1]
# PSEUDO LABEL TEST DATA
test_pred = test_pred > np.random.uniform(0,1,len(test_pred))
y[obs:] = test_pred
# COLLECT CENTER COORDINATES
centers = np.append(centers,np.mean(X[ np.argwhere(y==0).flatten() ,:],axis=0))
centers = np.append(centers,np.mean(X[ np.argwhere(y==1).flatten() ,:],axis=0))
# PLOT CENTER COORDINATES
plt.hist(centers,bins=100)
plt.title('Real Data Variable Means (match clusters=3)')
plt.show()
# # Appendix 2 - Advanced Techniques
# Since the code above already classifies Sklearn's `make_classification` on average prefectly, there is no need to improve it. However, below are some ideas that could be used if improvement was possible and/or neccessary.
#
# 1. After building model 4's classifier, you could use it to classify the training data. Then all training data with `abs(oof - true)>0.9` are erroneous training data with their labels flippped. Next correct those training labels and run the entire kernel a second time.
#
# 2. Since each cluster is centered at a hypercube corner, you can modify Sklearn's Quadratic Discriminant Analysis code by adding `meang[ np.argwhere(meang>=0) ] = 1.0` and `meang[ np.argwhere(meang<0) ] = -1.0`. This moves the centers of all clusters to hypercube corners.
#
# 3. Computer accuracy cannot distinguish between predictions that are close to 1. Using an example with 6 digit accuracy, the numbers 1.000001 and 1.000002 are the same because both become 1.00000. To help improve AUC, you can add the following code to this kernel. `temp = np.log(pds[:,0]+pds[:,1]+pds[:,2])`; `temp[ np.isinf(temp) ] = -1e6`; `test_pred -= temp`. This improves AUC by differentiating between predictions close to 1. Note that this isn't a problem for predictions close to 0 because the numbers 0.0000001 and 0.0000002 are 1.0e-7 and 2.0e-7 and the computer can already differentiate them.
#
# 4. After making predictions for `test.csv`, you can use them as pseudo labels and run the entire kernel a second time. Then use those labels and run the kernel a third time. Each iteration can give a slight boost.
#
# 5. You can run this kernel multiple times and take an average. Or use k-folds. This removes this code's variance (randomness) and achieves close to prefection everytime but it also removes the possibility of scoring LB 0.00050 more or less than perfection.
#
# 6. We can also remove this code's variance (randomness) by modifying Sklearn's code for Quadratic Discriminant Analysis and Gaussian Mixture. Each of these models can only accept training labels that are either 0 or 1. By adding a few lines of code, we can allow these models to accept continuous probabilities and use them as weights. This would allow us to remove the randomization line `test_pred = test_pred > np.random.uniform(0,1,len(test_pred))`. Instead we can leave pseudo labels as probabilities between 0 and 1 and still call `QuadraticDiscriminantAnalysis.fit(test_data,test_pred)`.
#
# Using combinations of these additional advanced techniques, this kernel was able to score LB 0.97489 on this competition's public leaderboard. But validation showed that these techniques didn't make the basic perfect classifier any more perfect. Therefore for final submission, the basic classifier was used.
# # Appendix 3 - Final Submission Strategy
# By applying our model to synthetic data, we can learn how it performs on a simulated public and private leaderboard. We observe that this kernel achieves perfection on average (if Kaggle used `make_classification` with the parameters we suspect). Sklearn's code for `make_classification` includes
#
# # Randomly replace labels
# if flip_y >= 0.0:
# flip_mask = generator.rand(n_samples) < flip_y
# y[flip_mask] = generator.randint(n_classes, size=flip_mask.sum())
#
# Before variable `y` gets rewritten, you can store it by adding `y_orig = y.copy()`. Next update the shuffle line to `X, y, y_orig = util_shuffle(X, y, y_orig, random_state=generator)`. Then change the last line of `make_classification` to `return X, y, y_orig`. By doing this, we can compute the AUC of a perfect classifier with `prefect = roc_auc_score(y, y_orig)`.
#
# Now we can make hundreds of synthetic datasets that are the similar to this competition's data and apply this kernel to see how well it does compared with a perfect classifier. For each synthetic dataset, we will run this kernel 10 times. This will show us patterns and help us decide how to choose our two final submissions.
#
# We observe that sometimes this kenel does better than perfect and sometimes it does worse than perfect. It's interesting to note that there is no correlation between its performance on public LB versus private LB. In the example plots below, perfect classification is represented by the black dotted line.
#
# If we take the average of this kernel's performance over many sythnetic datasets, it achieves perfect classification. Therefore there is no reason to improve this kernel. We can never expect to perform better than perfect classification on average. The only change we could consider is altering this kernel's standard deviation from perfection. We could either try to achieve perfection every kernel run, or (leave it as is and) randomly try to exceed perfection by a desired amount on some kernel runs.
#
# ## Synthetic Dataset 1
# 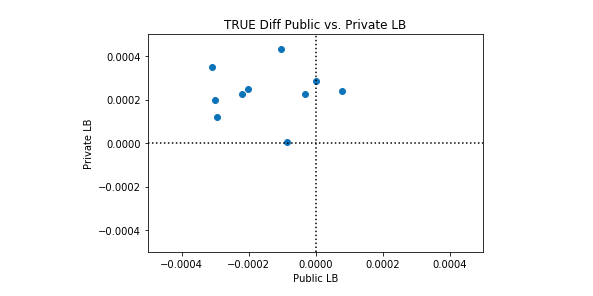
# ## Synthetic Dataset 2
# 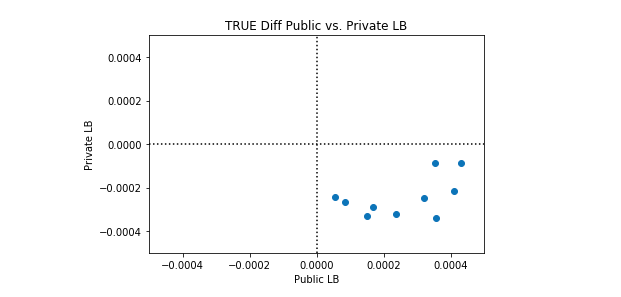
# ## Synthetic Dataset 3
# 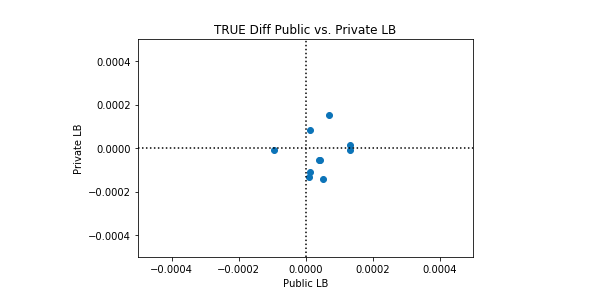
# ## Synthetic Dataset 4
# 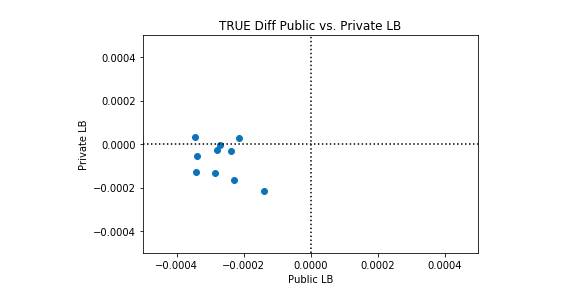
# ## Synthetic Dataset 5
# 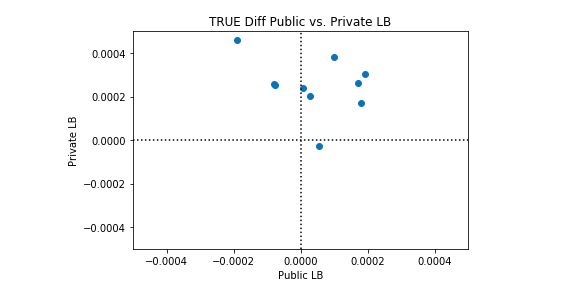
# ## Many Synthetic Datasets Together
# 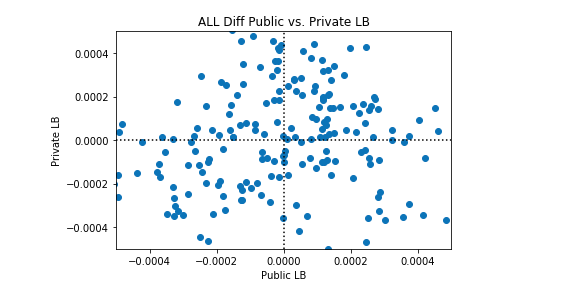
|
#!/usr/bin/env python2
# If you have virtualenv installed:
#
# $ python2 virtualenv.py venv
# $ venv/bin/pip install pygit2 dateparser
# To run:
# $ venv/bin/python tools/get_commits.py $HOME/ipfs_stuff/repos
# Pass this tool a folder that contains all of the IPFS repos you wish to scan
# It will output a list of authors
import pygit2
import dateparser
import glob
import argparse
import sys
import os.path
import time
from datetime import datetime,timedelta
from pprint import pprint
import urllib2
import json
name_map = {
('Jeromy', '<EMAIL>'): '<NAME>',
'<EMAIL>': '<NAME>',
'<EMAIL>': '<NAME>',
'<EMAIL>': '<NAME>',
}
def apply_name_map(name, email):
"Attempt to convert a name+email into a name"
if (name, email) in name_map:
return name_map[(name, email)]
if email in name_map:
return name_map[email]
return name
author_repo_map = {}
def main(repo_path, start, end):
if not start < end:
raise ValueError("`start` must be before `end`", start, end)
print "Getting all commits made between %s and %s" % (start.isoformat(), end.isoformat())
# get a full list of ipfs org repos
all_repos = json.load(urllib2.urlopen("https://api.github.com/orgs/ipfs/repos"))
for all_repo in all_repos:
if all_repo['fork']:
print "Skipping fork", all_repo['name']
continue
# does this repo exist locally?
local_repo_path = os.path.join(repo_path, all_repo['name'])
if not os.path.exists(local_repo_path):
print all_repo['name'], "does not exist locally! It will now be cloned..."
pygit2.clone_repository(all_repo['clone_url'], local_repo_path)
authors = set()
repos = glob.glob(os.path.join(repo_path, "*", ".git"))
print "Scanning for repos... Found %d repos" % len(repos)
print "If they have a remote named `origin`, it will be fetched from..."
for repo_path in repos:
repo = pygit2.Repository(repo_path)
for remote in repo.remotes:
if remote.name == "origin":
try:
transfer = remote.fetch()
while transfer.received_objects < transfer.total_objects:
time.sleep(0.1)
print "\r%d of %d " % (transfer.received_objects, transfer.total_objects) ,
print "Fetch complete for", remote.url
except Exception as ex:
print "Error while fetching for", repo_path, remote.url
print ex
master_oid = None
try:
master_oid = repo.lookup_reference("refs/remotes/origin/master").target
except KeyError:
print "Skipping %r, it doesn't seem to have the refs we want"
for commit in repo.walk(master_oid, pygit2.GIT_SORT_TIME):
commit_time = datetime.fromtimestamp(commit.commit_time)
if commit_time >= start and commit_time <= end:
n = apply_name_map(commit.author.name, commit.author.email)
#print "name:", repr(n), commit.oid
if type(n) == str:
n = n.decode("utf-8")
authors.add(n)
author_repo_map[n] = (repo_path.split(os.sep)[-2], commit.oid)
print "\nContributors for this period (%s to %s):" % (start.isoformat(), end.isoformat())
for a in sorted(authors):
try:
f = urllib2.urlopen("https://api.github.com/repos/ipfs/%s/commits/%s" % author_repo_map[a])
js = json.load(f)
# example:
# @eminence (<NAME>)
# [@githubname](githuburl) (author_name)
print "* [@%s](%s) (%s)" % (js['author']['login'], js['author']['html_url'], js['commit']['author']['name'])
except:
print "*", a, "https://github.com/ipfs/%s/commit/%s" % author_repo_map[a]
print ""
if __name__ == "__main__":
parser = argparse.ArgumentParser()
parser.add_argument('--start')
parser.add_argument('--end')
parser.add_argument('repo_dirs', help="Path to directories container all IPFS repos")
args = parser.parse_args()
start = None
end = None
if args.start:
start = dateparser.parse(args.start)
else:
print "Error: Failed to parse start date %r" % args.start
sys.exit(1)
if args.end:
end = dateparser.parse(args.end)
else:
print "Error: Failed to parse end date %r" % args.end
sys.exit(1)
main(args.repo_dirs, start, end)
|
#########################################
# Programmers: <NAME>, <NAME>, <NAME>
# File Name: PreviewTimetable.py
# Description: Contains UI code for the PreviewTimetableFrame
#########################################
from tkinter import *
from SQLWrapper import *
from Timetable import *
sqlWrapper = SQLWrapper()
class PreviewTimetableFrame(Frame):
def __init__(self, parent, studentId):
'''Initializes a PreviewTimetableFrame.'''
Frame.__init__(self, parent)
self.parent = parent
self.studentId = studentId
# Retrieve the student using the given studentId
self.student = sqlWrapper.getStudent(self.studentId)
# Convert final courses to a list (they are stored as a string in the database)
self.finalCourses = eval(self.student.finalCourses)
self.initUI()
def centerWindow(self):
'''Centers the window on the screen.'''
screenWidth = self.parent.winfo_screenwidth()
screenHeight = self.parent.winfo_screenheight()
# Calculate (x, y) position of window
x = (screenWidth/2) - (650/2)
y = (screenHeight/2) - (450/2)
self.parent.geometry('%dx%d+%d+%d' % (650, 450, x, y))
def initUI(self):
'''Initializes the user interface by configuring the table layout, adding buttons, etc.'''
self.parent.title("Timetable")
self.centerWindow()
self.pack(fill=BOTH, expand=1)
# Configure table layout
self.columnconfigure(0, weight=1)
self.columnconfigure(1, weight=1)
self.rowconfigure(0, weight=1)
self.rowconfigure(1, weight=1)
self.rowconfigure(2, weight=1)
self.rowconfigure(3, weight=1)
self.rowconfigure(4, weight=1)
self.rowconfigure(5, weight=1)
self.rowconfigure(6, weight=1)
self.rowconfigure(7, weight=1)
# Timetable title
idLabel = Label(self, text="Final Timetable", font=(26))
idLabel.grid(row=0, columnspan=20, column=0, sticky=S)
# Semester 1 title
sem1Label = Label(self, text="Semester 1:")
sem1Label.grid(row=1, column=0, sticky=S)
# Non-functioning buttons for each period slot on the timetable
self.sem1period1 = Button(self,text=self.finalCourses[0])
self.sem1period1.grid(row=2, column=0)
self.sem1period2 = Button(self,text=self.finalCourses[1])
self.sem1period2.grid(row=3, column=0)
self.sem1period3 = Button(self,text=self.finalCourses[2])
self.sem1period3.grid(row=4, column=0)
self.sem1period4 = Button(self,text=self.finalCourses[3])
self.sem1period4.grid(row=5, column=0)
self.sem1period5 = Button(self,text=self.finalCourses[4])
self.sem1period5.grid(row=6, column=0)
# Semester 2 title
sem2Label = Label(self, text="Semester 2:")
sem2Label.grid(row=1, column=1, sticky=S)
self.sem2period1 = Button(self,text=self.finalCourses[5])
self.sem2period1.grid(row=2, column=1)
self.sem2period2 = Button(self,text=self.finalCourses[6])
self.sem2period2.grid(row=3, column=1)
self.sem2period3 = Button(self,text=self.finalCourses[7])
self.sem2period3.grid(row=4, column=1)
self.sem2period4 = Button(self,text=self.finalCourses[8])
self.sem2period4.grid(row=5, column=1)
self.sem2period5 = Button(self,text=self.finalCourses[9])
self.sem2period5.grid(row=6, column=1)
|
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Mon Sep 7 20:55:52 2020
@author: zhouziyi
"""
real = []
imag = []
i = 0
with open("00_Data.txt") as Data:
readCSV = csv.reader(Data, delimiter=',')
next(readCSV)
for row in readCSV:
betar = float(row[1].split(",")[0])
#if (betar == beta) & (i == 0):
# n = row[0]
# s_raw = row[2].strip('[').strip(']').strip(' ').split(',')
# s = [int(i) for i in s_raw]
# print('********Input********')
# print('n:', n)
# print('beta:', beta)
# print('encoded string:', s)
# i = 1
#elif betar < beta:
it_raw = row[2].strip('[').strip(']').strip(' ').split(',')
it = [int(i) for i in it_raw]
n = int(row[0])
coe = [0 for i in range(n + 1)]
coe[0] = 1
coe[n] = -1
for j in it:
sumit = it.index(j)
coe[j + 1] = coe[j + 1] + (-2) * (sumit%2 * (-2) + 1)
roots = np.roots(coe)
for j in roots:
real.append(j.real)
imag.append(j.imag)
#for c in clist:
#print('c:', complex(c[0], c[1]))
#print('********Slice of Teapot when beta = ', beta,', c =',c, '********')
plt.show()
fig, ax = plt.subplots(figsize=(15, 15))
plt.scatter(real,imag,s=.0003,marker='.')
plt.xlim(-1.5, 1.5)
plt.ylim(-1.5, 1.5)
for i in Max:
R = i
n = 64
t = np.linspace(0, 2*np.pi, n+1)
x = R*np.cos(t)
y = R*np.sin(t)
plt.plot(x,y,'#4b8bbe')
for i in Min:
R = i
n = 64
t = np.linspace(0, 2*np.pi, n+1)
x = R*np.cos(t)
y = R*np.sin(t)
plt.plot(x,y,'orange')
nlist = [i+3 for i in range(len(Max))]
from matplotlib.pyplot import MultipleLocator
fig, ax = plt.subplots(figsize=(10, 5))
l1, = plt.plot(nlist, Max)
l2, = plt.plot(nlist, Min)
plt.legend(handles=[l1,l2],labels=['Maximum','Minimum'])
plt.xlabel('orbit length')
x_major_locator=MultipleLocator(1)
ax=plt.gca()
ax.xaxis.set_major_locator(x_major_locator)
plt.xlim(3, 24)
plt.show()
rate_Max = []
for i in range(len(Max)-1):
rate = Max[i + 1] / Max[i]
rate_Max.append(rate)
nlist2 = [i+3 for i in range(len(rate_Max))]
rate_Min = []
for i in range(len(Min)-1):
rate = Min[i + 1] / Min[i]
rate_Min.append(rate)
fig, ax = plt.subplots(figsize=(10, 5))
l1, = plt.plot(nlist2, rate_Max)
l2, = plt.plot(nlist2, rate_Min)
plt.legend(handles=[l1,l2],labels=['Maximum','Minimum'])
plt.xlabel('orbit length')
x_major_locator=MultipleLocator(1)
ax=plt.gca()
ax.xaxis.set_major_locator(x_major_locator)
plt.axhline(y=1, color='r', linestyle=':')
plt.show()
#plt.show() |
# -*- coding: utf-8 -*-
from collections import OrderedDict
import six
from fixtures_mongoengine import FixturesMongoengineException
from fixtures_mongoengine.fixture import Fixture, get_fixture_class, BaseFixture
"""
Metaclass idea and parts of code taken from https://github.com/croach/Flask-Fixtures
"""
TEST_SETUP_NAMES = ('setUp',)
TEST_TEARDOWN_NAMES = ('tearDown',)
def setup(obj):
"""
:type obj: FixturesMixin
"""
obj.unload_fixtures()
obj.load_fixtures()
def teardown(obj):
"""
:type obj: FixturesMixin
"""
obj.unload_fixtures()
class MetaFixturesMixin(type):
def __new__(mcs, name, bases, attrs):
fixtures_conf = attrs.get('fixtures_conf', [])
# We only need to do something if there's a set of fixtures,
# otherwise, do nothing. The main reason this is here is because this
# method is called when the FixturesMixin class is created and we
# don't want to do any test setup on that class.
if fixtures_conf:
child_setup_fn = mcs.get_child_fn(attrs, TEST_SETUP_NAMES, bases)
child_teardown_fn = mcs.get_child_fn(attrs, TEST_TEARDOWN_NAMES, bases)
attrs[child_setup_fn.__name__] = mcs.setup_handler(setup, child_setup_fn)
attrs[child_teardown_fn.__name__] = mcs.teardown_handler(teardown, child_teardown_fn)
return super(MetaFixturesMixin, mcs).__new__(mcs, name, bases, attrs)
@staticmethod
def setup_handler(setup_fixtures_fn, setup_fn):
"""Returns a function that adds fixtures handling to the setup method.
Makes sure that fixtures are setup before calling the given setup method.
"""
def handler(obj):
setup_fixtures_fn(obj)
setup_fn(obj)
return handler
@staticmethod
def teardown_handler(teardown_fixtures_fn, teardown_fn):
"""Returns a function that adds fixtures handling to the teardown method.
Calls the given teardown method first before calling the fixtures teardown.
"""
def handler(obj):
teardown_fn(obj)
teardown_fixtures_fn(obj)
return handler
@staticmethod
def get_child_fn(attrs, names, bases):
"""Returns a function from the child class that matches one of the names.
Searches the child class's set of methods (i.e., the attrs dict) for all
the functions matching the given list of names. If more than one is found,
an exception is raised, if one is found, it is returned, and if none are
found, a function that calls the default method on each parent class is
returned.
"""
def call_method(obj, method):
"""Calls a method as either a class method or an instance method.
"""
# The __get__ method takes an instance and an owner which changes
# depending on the calling object. If the calling object is a class,
# the instance is None and the owner will be the object itself. If the
# calling object is an instance, the instance will be the calling object
# and the owner will be its class. For more info on the __get__ method,
# see http://docs.python.org/2/reference/datamodel.html#object.__get__.
if isinstance(obj, type):
instance = None
owner = obj
else:
instance = obj
owner = obj.__class__
method.__get__(instance, owner)()
# Create a default function that calls the default method on each parent
default_name = names[0]
def default_fn(obj):
for cls in bases:
if hasattr(cls, default_name):
call_method(obj, getattr(cls, default_name))
default_fn.__name__ = default_name
# Get all of the functions in the child class that match the list of names
fns = [(name, attrs[name]) for name in names if name in attrs]
# Raise an error if more than one setup/teardown method is found
if len(fns) > 1:
raise RuntimeError("Cannot have more than one setup or teardown method per context (class or test).")
# If one setup/teardown function was found, return it
elif len(fns) == 1:
name, fn = fns[0]
def child_fn(obj):
call_method(obj, fn)
child_fn.__name__ = name
return child_fn
# Otherwise, return the default function
else:
return default_fn
class FixturesMixin(six.with_metaclass(MetaFixturesMixin, object)):
fixtures_conf = {}
"""
Declares the fixtures that are needed by the current test case.
The return value of this method must be an array of fixture configurations. For example,
```python
[
'users' => UserFixture,
'articles' => ArticleFixture
]
```
"""
__fixtures = None
def __getattr__(self, name):
if name in self.__dict__:
return self.__dict__[name]
if name in self.get_fixtures():
return self.get_fixtures()[name]
raise AttributeError('Attribute "{}" not found.'.format(name))
def load_fixtures(self):
for fixture in self.get_fixtures().values():
fixture.before_load()
for fixture in self.get_fixtures().values():
fixture.load()
fixtures = list(self.get_fixtures().values())
fixtures.reverse()
for fixture in fixtures:
fixture.after_load()
def unload_fixtures(self):
for fixture in self.get_fixtures().values():
fixture.before_unload()
fixtures = list(self.get_fixtures().values())
fixtures.reverse()
for fixture in fixtures:
fixture.unload()
for fixture in fixtures:
fixture.after_unload()
def get_fixtures(self):
"""
:rtype: OrderedDict[Fixture]
"""
if self.__fixtures is None:
self.__fixtures = self._create_fixtures()
return self.__fixtures
def _create_fixtures(self):
aliases = {}
for name, fixture_class in six.iteritems(self.fixtures_conf):
aliases[fixture_class] = name
instances = OrderedDict()
stack = [fixture_class for name, fixture_class in six.iteritems(self.fixtures_conf)]
stack.reverse()
while len(stack) > 0:
fixture = stack.pop()
if isinstance(fixture, BaseFixture):
fixture_class = fixture.__class__
if fixture_class in instances:
del instances[fixture_class]
instances[fixture_class] = fixture
else:
fixture_class = fixture
if fixture_class not in instances:
instances[fixture_class] = None
fixture = fixture_class()
stack.append(fixture)
for dep in fixture.depends.values():
if isinstance(dep, six.string_types):
dep = get_fixture_class(dep)
stack.append(dep)
elif instances[fixture_class] is None:
msg = 'A circular dependency is detected for fixture {}.'.format(fixture_class.__name__)
raise FixturesMongoengineException(msg)
fixtures = OrderedDict()
for fixture_class, fixture in six.iteritems(instances):
fixture.init_depended_fixtures(instances)
name = aliases[fixture_class] if fixture_class in aliases else fixture_class.__name__
fixtures[name] = fixture
return fixtures
|
<reponame>SepioSystems/demisto-sdk<filename>demisto_sdk/commands/update_release_notes/tests/update_rn_test.py
import os
import shutil
import unittest
from demisto_sdk.commands.common.git_tools import git_path
class TestRNUpdate(unittest.TestCase):
FILES_PATH = os.path.normpath(os.path.join(__file__, f'{git_path()}/demisto_sdk/tests', 'test_files'))
def test_build_rn_template_integration(self):
"""
Given:
- a dict of changed items
When:
- we want to produce a release notes template
Then:
- return a markdown string
"""
expected_result = "\n#### Integrations\n- __Hello World Integration__\n%%UPDATE_RN%%\n" \
"\n#### Playbooks\n- __Hello World Playbook__\n%%UPDATE_RN%%\n" \
"\n#### Scripts\n- __Hello World Script__\n%%UPDATE_RN%%\n" \
"\n#### IncidentFields\n- __Hello World IncidentField__\n%%UPDATE_RN%%\n" \
"\n#### Classifiers\n- __Hello World Classifier__\n%%UPDATE_RN%%\n" \
"\n#### Layouts\n- __Hello World Layout__\n%%UPDATE_RN%%\n" \
"\n#### IncidentTypes\n- __Hello World Incident Type__\n%%UPDATE_RN%%\n"
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='minor', pack_files={'HelloWorld'})
changed_items = {
"Hello World Integration": "Integration",
"Hello World Playbook": "Playbook",
"Hello World Script": "Script",
"Hello World IncidentField": "IncidentFields",
"Hello World Classifier": "Classifiers",
"N/A": "Integration",
"Hello World Layout": "Layouts",
"Hello World Incident Type": "IncidentTypes",
}
release_notes = update_rn.build_rn_template(changed_items)
assert expected_result == release_notes
def test_find_corresponding_yml(self):
"""
Given:
- a filepath containing a python file
When:
- determining the changed file
Then:
- return only the yml of the changed file
"""
expected_result = "Integration/HelloWorld.yml"
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='minor', pack_files={'HelloWorld'})
filepath = 'Integration/HelloWorld.py'
filename = update_rn.find_corresponding_yml(filepath)
assert expected_result == filename
def test_return_release_notes_path(self):
"""
Given:
- a pack name and version
When:
- building the release notes file within the ReleaseNotes directory
Then:
- the filepath of the correct release notes.
"""
expected_result = 'Packs/HelloWorld/ReleaseNotes/1_1_1.md'
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='minor', pack_files={'HelloWorld'})
input_version = '1.1.1'
result = update_rn.return_release_notes_path(input_version)
assert expected_result == result
def test_bump_version_number_minor(self):
"""
Given:
- a pack name and version
When:
- bumping the version number in the metadata.json
Then:
- return the correct bumped version number
"""
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/_pack_metadata.json'))
expected_version = '1.1.0'
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='minor', pack_files={'HelloWorld'})
update_rn.metadata_path = os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json')
version_number = update_rn.bump_version_number(pre_release=False)
assert version_number == expected_version
os.remove(os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'))
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/_pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'))
def test_bump_version_number_major(self):
"""
Given:
- a pack name and version
When:
- bumping the version number in the metadata.json
Then:
- return the correct bumped version number
"""
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/_pack_metadata.json'))
expected_version = '2.0.0'
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='major', pack_files={'HelloWorld'})
update_rn.metadata_path = os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json')
version_number = update_rn.bump_version_number(pre_release=False)
assert version_number == expected_version
os.remove(os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'))
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/_pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'))
def test_bump_version_number_revision(self):
"""
Given:
- a pack name and version
When:
- bumping the version number in the metadata.json
Then:
- return the correct bumped version number
"""
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/_pack_metadata.json'))
expected_version = '1.0.1'
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='revision', pack_files={'HelloWorld'})
update_rn.metadata_path = os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json')
version_number = update_rn.bump_version_number(pre_release=False)
assert version_number == expected_version
os.remove(os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'))
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/_pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack/pack_metadata.json'))
def test_bump_version_number_revision_overflow(self):
"""
Given:
- a pack name and a version before an overflow condition
When:
- bumping the version number in the metadata.json
Then:
- return ValueError
"""
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/_pack_metadata.json'))
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='revision', pack_files={'HelloWorld'})
update_rn.metadata_path = os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json')
self.assertRaises(ValueError, update_rn.bump_version_number)
os.remove(os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'))
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/_pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'))
def test_bump_version_number_minor_overflow(self):
"""
Given:
- a pack name and a version before an overflow condition
When:
- bumping the version number in the metadata.json
Then:
- return ValueError
"""
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/_pack_metadata.json'))
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='minor', pack_files={'HelloWorld'})
update_rn.metadata_path = os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json')
self.assertRaises(ValueError, update_rn.bump_version_number)
os.remove(os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'))
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/_pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'))
def test_bump_version_number_major_overflow(self):
"""
Given:
- a pack name and a version before an overflow condition
When:
- bumping the version number in the metadata.json
Then:
- return ValueError
"""
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/_pack_metadata.json'))
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="HelloWorld", update_type='major', pack_files={'HelloWorld'})
update_rn.metadata_path = os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json')
self.assertRaises(ValueError, update_rn.bump_version_number)
os.remove(os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'))
shutil.copy(src=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/_pack_metadata.json'),
dst=os.path.join(TestRNUpdate.FILES_PATH, 'fake_pack_invalid/pack_metadata.json'))
class TestRNUpdateUnit:
FILES_PATH = os.path.normpath(os.path.join(__file__, f'{git_path()}/demisto_sdk/tests', 'test_files'))
def test_ident_changed_file_type_integration(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'Integration')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'Integration/VulnDB/VulnDB.py')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_ident_changed_file_type_script(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'Script')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'Script/VulnDB/VulnDB.py')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_ident_changed_file_type_playbooks(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'Playbook')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'Playbooks/VulnDB/VulnDB_playbook.yml')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_ident_changed_file_type_incident_fields(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'IncidentFields')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'IncidentFields/VulnDB/VulnDB.json')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_ident_changed_file_type_incident_types(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'IncidentTypes')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'IncidentTypes/VulnDB/VulnDB.json')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_ident_changed_file_type_classifiers(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'Classifiers')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'Classifiers/VulnDB/VulnDB.json')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_ident_changed_file_type_layouts(self, mocker):
"""
Given:
- a filepath of a changed file
When:
- determining the type of item changed (e.g. Integration, Script, Layout, etc.)
Then:
- return tuple where first value is the pack name, and second is the item type
"""
expected_result = ('VulnDB', 'Layout')
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'Layouts/VulnDB/VulnDB.json')
mocker.patch.object(UpdateRN, 'find_corresponding_yml', return_value='Integrations/VulnDB/VulnDB.yml')
mocker.patch.object(UpdateRN, 'get_display_name', return_value='VulnDB')
result = update_rn.ident_changed_file_type(filepath)
assert expected_result == result
def test_check_rn_directory(self):
"""
Given:
- a filepath for a release notes directory
When:
- determining if the directory exists
Then:
- create the directory if it does not exist
"""
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'ReleaseNotes')
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
update_rn.check_rn_dir(filepath)
def test_create_markdown(self):
"""
Given:
- a filepath for a release notes file and a markdown string
When:
- creating a new markdown file
Then:
- create the file or skip if it exists.
"""
from demisto_sdk.commands.update_release_notes.update_rn import UpdateRN
update_rn = UpdateRN(pack="VulnDB", update_type='minor', pack_files={'HelloWorld'})
filepath = os.path.join(TestRNUpdate.FILES_PATH, 'ReleaseNotes/1_1_1.md')
md_string = '### Test'
update_rn.create_markdown(release_notes_path=filepath, rn_string=md_string)
|
<reponame>thesteau/Portfolio-Janggi
# All pieces that are used in the game of Janggi
class Pieces:
""" Represents the Janggi pieces.
Each piece will hold its individual information."""
def __init__(self, player):
""" Initializes the Janggi pieces.
Data member:
player: The player in which the piece will now belong to."""
self._player = player.lower()
def get_player(self):
""" Returns the owner of the piece."""
return self._player
class Chariot(Pieces):
""" Represents the Chariot piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits."""
super().__init__(player)
self._name = 'Chariot'
self._move = 10 # Within the game limits
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
class Elephant(Pieces):
""" Represents the Elephant piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits."""
super().__init__(player)
self._name = 'Elephant'
self._move = 5
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
def get_beast_num(self):
""" Returns the beast number (Colloquial on diagonal move difference)."""
return 3
class Horse(Pieces):
""" Represents the Horse piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits."""
super().__init__(player)
self._name = 'Horse'
self._move = 3
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
def get_beast_num(self):
""" Returns the beast number (Colloquial on diagonal move difference)."""
return 2
class Cannon(Pieces):
""" Represents the Cannon piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits."""
super().__init__(player)
self._name = 'Cannon'
self._move = 10
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
class Soldier(Pieces):
""" Represents the Soldier piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits."""
super().__init__(player)
self._name = 'Soldier'
self._move = 1
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
class Guard(Pieces):
""" Represents the Guard piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits."""
super().__init__(player)
self._name = 'Guard'
self._move = 1
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
class General(Pieces):
""" Represents the General piece."""
def __init__(self, player):
""" Initiates the piece's data accordingly.
Data members:
name: The name of the piece.
move: max movement limits.
check: The check status of the piece.
chk_count: The number of check have applied to the piece."""
super().__init__(player)
self._name = 'General'
self._move = 1
self._check = False
self._chk_count = 0
def __repr__(self):
""" Pretty print the name of the piece."""
return self.get_player()[0].upper()+'.'+self._name[:3]
def get_name(self):
""" Returns the name of the piece."""
return self._name
def get_move(self):
""" Returns the maximum movement square."""
return self._move
def get_check_status(self):
""" Returns whether the piece is in check or not."""
return self._check
def get_chk_count(self):
""" Returns the amount of checks imposed to the piece."""
return self._chk_count
def set_check_status(self, boolean=False):
""" Sets the check status of the piece."""
self._check = boolean
def add_chk_count(self):
""" Adds a check count to the piece."""
self._chk_count += 1
def sub_chk_count(self):
""" Subtracts a check count to the piece."""
self._chk_count -= 1
if self._chk_count < 0:
self._chk_count = 0 # Negative check count is not possible.
|
# -*- coding: UTF-8 -*-
import os
import sys
import json
import re
import sqlite3
import time
from selenium import webdriver
from utils.common import *
if PY3:
import urllib.request
import _thread
else:
import urllib2
import thread
driver = webdriver.Chrome()
REQ_HEADERS = {
"Accept": "*/*",
"Accept-Language": "en-US,en;q=0.9",
"Accept-Charset": "big5,ISO-8859-1,utf-8;q=0.7,*;q=0.3",
"Connection": "keep-alive",
"Origin": "https://webgis.sinica.edu.tw/",
"Referer": "https://webgis.sinica.edu.tw/",
"User-Agent": "Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/66.0.3359.139 Safari/537.36"
}
KEY_WORD = sys.argv[1]
SEARCH_URL = "http://webgis.sinica.edu.tw/place/query.asp?A1=%AC%D9%A5%F7&B1=containing&C1={}&Page_setup=50&D1=AND&A2=99&B2=containing&C2=&D2=AND&A3=99&B3=containing&C3=&page=".format(KEY_WORD)
INFO_URL_PREFIX = "http://webgis.sinica.edu.tw/place/"
DB_PATH = "data.db"
DATA_PATH = "data.json"
conn = sqlite3.connect(DB_PATH)
conn.text_factory = str
c = conn.cursor()
c.execute('''CREATE TABLE IF NOT EXISTS `INFO_1` (`ID` INTEGER NOT NULL, `省份` TEXT, `地區` TEXT, `卷數` TEXT, `編修者年代` TEXT, `人名` TEXT, `年代` TEXT, `西元` TEXT, `性質` TEXT,`館藏地` TEXT,`註記` TEXT, PRIMARY KEY(`ID`));''')
c.execute('''CREATE TABLE IF NOT EXISTS `INFO_2` (`ID` INTEGER NOT NULL, `省份` TEXT, `地區` TEXT, `地方志名` TEXT, `卷數` TEXT, `修纂時間` TEXT, `編纂單位` TEXT, `叢書名` TEXT, `出版地` TEXT, `出版時間` TEXT, `稽核項` TEXT, `館藏/藏書者` TEXT, `版本` TEXT, `備考/附註` TEXT, PRIMARY KEY(`ID`));''')
conn.commit()
def get_response_content(request_url):
if PY3:
# python 3.x
request = urllib.request.Request(request_url, headers=REQ_HEADERS)
response = urllib.request.urlopen(request)
else:
# python 2.x
request = urllib2.Request(request_url, headers=REQ_HEADERS)
response = urllib2.urlopen(request)
if response:
return response.read()
return None
def get_total_pages():
content = get_response_content(SEARCH_URL + "1")
content = content.decode("big5", errors="ignore")
total_regex = r"<font face=\"Arial, Helvetica, sans-serif\">(\d*)</font>"
return int(re.findall(total_regex, content)[0])
def get_info_urls(page_url):
content = get_response_content(page_url)
content = content.decode("big5", errors="ignore")
content = content.replace("\t", "").replace("\r", "").replace("\n", "")
url_regex = r"(detail\.asp\?ID=\d*\&Source=\d)"
return re.findall(url_regex, content)
def get_all_pages(start):
total = get_total_pages()
print("Totals: " + str(total))
for i in range(start, total):
info_urls = get_info_urls(SEARCH_URL + str(i+1))
print("Getting Pages from:" + str(i+1))
for info_url in info_urls:
get_info(info_url)
def get_info(info_url):
url = INFO_URL_PREFIX + info_url
print("Getting Data from URL:" + url)
driver.get(url)
content = driver.page_source
content = content.replace("\t", "").replace("\r", "").replace("\n", "")
id_regex = r"detail\.asp\?ID=(\d*)\&Source=\d"
source_regex = r"detail\.asp\?ID=\d*\&Source=(\d)"
info_regex = r"width=\"100\">([^<]*): </th><td class=\"calc\" align=\"left\" valign=\"top\">\xa0([^<]*)</td>"
id = re.findall(id_regex, url)[0]
source = re.findall(source_regex, url)[0]
info = {}
for key, value in re.findall(info_regex, content):
info[key] = value
write_to_db(id, info, source)
def write_to_db(id, info, source):
try:
if source == "1":
c.execute('''INSERT OR IGNORE INTO INFO_1 (`ID`, `省份`, `地區`, `卷數`, `編修者年代`, `人名`, `年代`, `西元`, `性質`,`館藏地`,`註記`) VALUES (?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?);''',
(id, info["省份"], info["地區"], info["卷數"], info["編修者年代"], info["人名"], info["年代"], info["西元"], info["性質"], info["館藏地"], info["註記"]))
conn.commit()
if source == "2":
c.execute('''INSERT OR IGNORE INTO INFO_2 (`ID`, `省份`, `地區`, `地方志名`, `卷數`, `修纂時間`, `編纂單位`, `叢書名`, `出版地`, `出版時間`, `稽核項`, `館藏/藏書者`, `版本`, `備考/附註`) VALUES (?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?);''',
(id, info["省份"], info["地區"], info["地方志名"], info["卷數"], info["修纂時間"], info["編纂單位"], info["叢書名"], info["出版地"], info["出版時間"], info["稽核項"], info["館藏/藏書者"], info["版本"], info["備考/附註"]))
conn.commit()
except Exception as e:
print(e)
get_all_pages(0)
conn.close()
|
import os #to use the OS structure commands
import gtts #to translate text to speech
import random #for the random integer generator
from playsound import playsound #to play the output sound
#converts text to speech
def text_speech(speech):
speech_file = gtts.gTTS(speech) #converting text to speech using google text to speech (gTTS)
speech_file.save("speech.mp3") #saving the audio output recieved from gTTS
playsound("speech.mp3") #used to play the saved audio file
os.remove("speech.mp3") #used in removing the audio file after use
#generates random number to determine computer choice of either rock, paper and scissors
def random_number():
r_int = random.randint(0, 2)
if r_int == 0:
return "rock"
elif r_int == 1:
return "paper"
elif r_int == 2:
return "scissors"
#checks who wins or weather the game ends on draw
def check_win(user_choice, comp_choice):
if user_choice == comp_choice:
return "-- Its a Draw!"
elif comp_choice == "paper" and user_choice == "rock":
return "-- Computer wins the game!"
elif comp_choice == "rock" and user_choice == "scissors":
return "-- Computer wins the game!"
elif comp_choice == "scissors" and user_choice == "paper":
return "-- Computer wins the game!"
else:
return "-- User won the game this time!"
#checks if the input entered by user is valid or not
def check_value(user_input):
valid_tpl = ("rock", "scissors", "paper")
if user_input.lower() in valid_tpl:
return True
else:
return False
#prints the output according the fulfilled conditions
def output_text(user_choice, comp_choice, result):
if result == "-- Its a Draw!":
text_speech(f"You entered {user_choice}. Computer chose {comp_choice}.")
print("-- Its a Draw!")
elif result == "-- Computer wins the game!":
text_speech(f"You entered {user_choice}. Computer chose {comp_choice}.")
print("-- Computer wins the game!")
else:
text_speech(f"You entered {user_choice}. Computer chose {comp_choice}.")
print("-- User won the game this time!")
#counts and stores scores of number of draws, computer wins and user wins
def scores(result, scores_list):
if result == "-- Its a Draw!":
scores_list[0] += 1
elif result == "-- User won the game this time!":
scores_list[1] += 1
else:
scores_list[2] += 1
return scores_list
#first = draw counts, second = user wins counts and third = computer wins count
scores_list = [0, 0, 0]
print("\n") #line break
flag = True
while flag:
#executed when the game is started for the first time
text_speech("Please enter either Rock, Paper or Scissors : ")
user_choice = input("Please enter either Rock, Paper or Scissors : ")
user_choice = user_choice.lower()
check = check_value(user_choice)
while check is False: #executed when the user input is incorrect
text_speech("Error! Please input Rock, Paper or Scissors.")
user_choice = input("Choose either Rock, Paper or Sciccors >>>")
check = check_value(user_choice)
comp_choice = random_number()
result = check_win(user_choice, comp_choice)
output_text(user_choice, comp_choice, result)
scores_list = scores(result, scores_list)
print("\n") #line break
count = input("Do you like to continue? (yes/no) : ") #executed to know if the user wants to continue the game.
if count.lower() != "yes":
flag = False
else:
text_speech("Thank you so much for playing this game!") #executed before final scoreboard is displayed
#final scoreboard displaying draws, user wins and computer wins
print(f"""\n\t\t----SCOREBOARD----\n
\t\tNumber of draws : {scores_list[0]}
\t\tUser Wins : {scores_list[1]}
\t\tComputer Wins : {scores_list[2]}""")
|
import skrf
import tkinter as tk
from matplotlib.figure import Figure
from matplotlib.backends.backend_tkagg import FigureCanvasTkAgg
import numpy as np
import CircuitFig
from PIL import ImageTk, Image, ImageDraw
import io
import MatchCal
l2z = lambda l: l[0] + 1j * l[1]
s4cmp = lambda sf: 'nH' if sf == 'l' else 'pF'
def ld4img2gui(label: tk.Label,
color: str, stage: int, sh_se: bool,
cmp_l: list, cmp_v: list, z_val: str = '50+0j',
valid: bool = True):
cr_cfg = CircuitFig.CircuitFig(color, stage, sh_se, cmp_l, cmp_v, z_val)
image = Image.open(io.BytesIO(cr_cfg.image_data)).resize((300, 180), Image.ANTIALIAS)
im = Image.new('RGBA', (300, 180), (255, 255, 255, 255))
draw = ImageDraw.Draw(im)
im.paste(image, (0, 0))
if not valid:
draw.line((0, 0, 300, 180), fill=(255, 0, 0, 255), width=5)
draw.line((0, 180, 300, 0), fill=(255, 0, 0, 255), width=5)
label.image = ImageTk.PhotoImage(im)
label.configure(image=label.image)
class TkGui:
def __init__(self, master):
self.master = master
self.top_frame = tk.Frame(self.master)
self.top_frame.pack(side=tk.LEFT)
self.right_frame = tk.Frame(self.master)
self.right_frame.pack(side=tk.LEFT, fill=tk.BOTH)
self.upper_sch_f = tk.Frame(self.right_frame)
self.upper_sch_f.grid(row=0, padx=(0, 5), pady=(5, 0), sticky="nsew")
self.lower_ety_f = tk.Frame(self.right_frame)
self.lower_ety_f.grid(row=1, padx=(0, 5), pady=(0, 5), sticky="nsew")
self.fig = Figure(figsize=(5, 6), dpi=100)
self.fig_cvs = FigureCanvasTkAgg(self.fig, master=self.top_frame)
self.ax: Figure = self.fig.gca()
self.fig_cvs.get_tk_widget().pack(side=tk.LEFT, padx=5, pady=5)
try:
with open('ring slot.s1p', 'r'):
pass
except IOError:
with open('ring slot.s1p', 'a+') as wf:
wf.write("""!Created with skrf (http://scikit-rf.org).
# GHz S RI R 50.0
!freq ReS11 ImS11
75.0 -0.503723180993 0.457844804761""")
self.my_slot = skrf.Network('ring slot.s1p')
self.to_match_z = [50, 0]
self.ser_match_z = [50, 0]
self.shu_match_z = [50, 0]
self.shu_ser_match_z_a = [50, 0]
self.shu_ser_match_z_b = [50, 0]
self.ser_shu_match_z_a = [50, 0]
self.ser_shu_match_z_b = [50, 0]
self.plt_z0 = 50 + 0j
self.plt_freq = 2.45e9
self.up2chart()
self.lb1 = tk.Label(self.upper_sch_f, relief="ridge")
self.lb1_tit = tk.Label(
self.upper_sch_f, text='Shunt Matching', relief="raised").grid(
row=0, column=0, sticky="nsew")
self.lb1.grid(row=1, column=0)
self.lb2 = tk.Label(self.upper_sch_f, relief="ridge")
self.lb2_tit = tk.Label(
self.upper_sch_f, text='Series Matching', relief="raised").grid(
row=0, column=1, sticky="nsew")
self.lb2.grid(row=1, column=1)
self.lb3 = tk.Label(self.upper_sch_f, relief="ridge")
self.lb3_tit = tk.Label(
self.upper_sch_f, text='Shunt-Series Matching', relief="raised").grid(
row=2, column=0, sticky="nsew")
self.lb3.grid(row=3, column=0)
self.lb4 = tk.Label(self.upper_sch_f, relief="ridge")
self.lb4_tit = tk.Label(
self.upper_sch_f, text='Shunt-Series Matching', relief="raised").grid(
row=2, column=1, sticky="nsew")
self.lb4.grid(row=3, column=1)
self.lb5 = tk.Label(self.upper_sch_f, relief="ridge")
self.lb5_tit = tk.Label(
self.upper_sch_f, text='Series-Shunt Matching', relief="raised").grid(
row=4, column=0, sticky="nsew")
self.lb5.grid(row=5, column=0)
self.lb6 = tk.Label(self.upper_sch_f, relief="ridge")
self.lb6_tit = tk.Label(
self.upper_sch_f, text='Series-Shunt Matching', relief="raised").grid(
row=4, column=1, sticky="nsew")
self.lb6.grid(row=5, column=1)
ld4img2gui(self.lb1, 'b', 1, False, ['c', 'l', 'c'], ['NC', 'SHORT', ''])
ld4img2gui(self.lb2, 'y', 1, True, ['c', 'l', 'c'], ['', 'SHORT', ''])
ld4img2gui(self.lb3, 'g', 2, False, ['c', 'l', 'c'], ['NC', 'SHORT', ''])
ld4img2gui(self.lb4, 'purple', 2, False, ['c', 'l', 'c'], ['NC', 'SHORT', ''])
ld4img2gui(self.lb5, 'orange', 2, True, ['c', 'l', 'c'], ['', 'SHORT', 'NC'])
ld4img2gui(self.lb6, 'brown', 2, True, ['c', 'l', 'c'], ['', 'SHORT', 'NC'])
###################################################################
self.to_match_r = tk.StringVar(value=str(self.to_match_z[0]))
self.to_match_i = tk.StringVar(value=str(self.to_match_z[1]))
self.ety_lb1 = tk.Label(self.lower_ety_f, text='To Match Complex Value')
self.ety_lb1.pack(side=tk.TOP)
self.ety_lb1b = tk.Label(self.lower_ety_f, text='Z = ')
self.ety_lb1b.pack(side=tk.LEFT)
self.ety1_r = tk.Entry(self.lower_ety_f, textvariable=self.to_match_r)
self.ety1_r.pack(side=tk.LEFT)
self.ety_lb1c = tk.Label(self.lower_ety_f, text=' + ')
self.ety_lb1c.pack(side=tk.LEFT)
self.ety1_i = tk.Entry(self.lower_ety_f, textvariable=self.to_match_i)
self.ety1_i.pack(side=tk.LEFT)
self.ety_lb1c = tk.Label(self.lower_ety_f, text='j')
self.ety_lb1c.pack(side=tk.LEFT)
self.enter = tk.Button(self.lower_ety_f, text="Start Auto Solver",
command=self.ld2chart)
self.enter.pack(side=tk.LEFT)
def ld2chart(self):
self.to_match_z = [float(self.ety1_r.get()), float(self.ety1_i.get())]
tmp_cal = MatchCal.MatchCal()
tmp_cal.tar_freq = self.plt_freq
to_mat = float(self.ety1_r.get()) + 1j * float(self.ety1_i.get())
tmp_cal.shu_0_sol(to_mat)
disp_str = f'{tmp_cal.shu:.2f} {s4cmp(tmp_cal.shu_t)}' if tmp_cal.shu else 'NC'
ld4img2gui(self.lb1, 'b', 1, False, [tmp_cal.shu_t, 'l', 'c'],
[disp_str, 'SHORT', ''],
f'{int(tmp_cal.tmp_z.real)}+{int(tmp_cal.tmp_z.imag)}j',
tmp_cal.sol_valid)
self.ser_match_z = [tmp_cal.tmp_z.real, tmp_cal.tmp_z.imag]
tmp_cal.ser_0_sol(to_mat)
disp_str = f'{tmp_cal.ser:.2f} {s4cmp(tmp_cal.ser_t)}' if tmp_cal.ser else 'SHORT'
ld4img2gui(self.lb2, 'y', 1, True, ['c', tmp_cal.ser_t, 'c'],
['', disp_str, ''],
f'{int(tmp_cal.tmp_z.real)}+{int(tmp_cal.tmp_z.imag)}j',
tmp_cal.sol_valid)
self.shu_match_z = [tmp_cal.tmp_z.real, tmp_cal.tmp_z.imag]
tmp_cal.sol_2stage(to_mat, True)
disp_str1 = f'{tmp_cal.ser:.2f} {s4cmp(tmp_cal.ser_t)}' if tmp_cal.ser else 'SHORT'
disp_str2 = f'{tmp_cal.shu:.2f} {s4cmp(tmp_cal.shu_t)}' if tmp_cal.shu else 'NC'
ld4img2gui(self.lb3, 'g', 2, False, [tmp_cal.shu_t, tmp_cal.ser_t, 'c'],
[disp_str2, disp_str1, ''],
f'{int(tmp_cal.tmp_z.real)}+{int(tmp_cal.tmp_z.imag)}j',
tmp_cal.sol_valid)
self.shu_ser_match_z_a = [tmp_cal.tmp_z.real, tmp_cal.tmp_z.imag]
tmp_cal.sol_2stage(to_mat, True, True)
disp_str1 = f'{tmp_cal.ser:.2f} {s4cmp(tmp_cal.ser_t)}' if tmp_cal.ser else 'SHORT'
disp_str2 = f'{tmp_cal.shu:.2f} {s4cmp(tmp_cal.shu_t)}' if tmp_cal.shu else 'NC'
ld4img2gui(self.lb4, 'purple', 2, False, [tmp_cal.shu_t, tmp_cal.ser_t, 'c'],
[disp_str2, disp_str1, ''],
f'{int(tmp_cal.tmp_z.real)}+{int(tmp_cal.tmp_z.imag)}j',
tmp_cal.sol_valid)
self.shu_ser_match_z_b = [tmp_cal.tmp_z.real, tmp_cal.tmp_z.imag]
tmp_cal.sol_2stage(to_mat)
disp_str1 = f'{tmp_cal.ser:.2f} {s4cmp(tmp_cal.ser_t)}' if tmp_cal.ser else 'SHORT'
disp_str2 = f'{tmp_cal.shu:.2f} {s4cmp(tmp_cal.shu_t)}' if tmp_cal.shu else 'NC'
ld4img2gui(self.lb5, 'orange', 2, True, ['c', tmp_cal.ser_t, tmp_cal.shu_t],
['', disp_str1, disp_str2],
f'{int(tmp_cal.tmp_z.real)}+{int(tmp_cal.tmp_z.imag)}j',
tmp_cal.sol_valid)
self.ser_shu_match_z_a = [tmp_cal.tmp_z.real, tmp_cal.tmp_z.imag]
tmp_cal.sol_2stage(to_mat, ans_sel=True)
disp_str1 = f'{tmp_cal.ser:.2f} {s4cmp(tmp_cal.ser_t)}' if tmp_cal.ser else 'SHORT'
disp_str2 = f'{tmp_cal.shu:.2f} {s4cmp(tmp_cal.shu_t)}' if tmp_cal.shu else 'NC'
ld4img2gui(self.lb6, 'brown', 2, True, ['c', tmp_cal.ser_t, tmp_cal.shu_t],
['', disp_str1, disp_str2],
f'{int(tmp_cal.tmp_z.real)}+{int(tmp_cal.tmp_z.imag)}j',
tmp_cal.sol_valid)
self.ser_shu_match_z_b = [tmp_cal.tmp_z.real, tmp_cal.tmp_z.imag]
self.up2chart()
def up2chart(self):
self.ax.clear()
self.fig2gui(np.array([[[l2z(self.to_match_z)]]]), 'To Match', 'r', 's')
self.fig2gui(np.array([[[l2z(self.ser_match_z)]]]), 'After Match', 'b', 'o')
self.fig2gui(np.array([[[l2z(self.shu_match_z)]]]), 'After Match', 'y', 'o')
self.fig2gui(np.array([[[l2z(self.shu_ser_match_z_a)]]]), 'After Match', 'g', 'o')
self.fig2gui(np.array([[[l2z(self.shu_ser_match_z_b)]]]), 'After Match', 'purple', 'o')
self.fig2gui(np.array([[[l2z(self.ser_shu_match_z_a)]]]), 'After Match', 'orange', 'o')
self.fig2gui(np.array([[[l2z(self.ser_shu_match_z_b)]]]), 'After Match', 'brown', 'o')
def fig2gui(self, plt_data: np.array,
label: str = '', color: str = 'r', mark: str = 's',
plt_sel: bool = False) -> None:
self.my_slot.frequency = self.plt_freq
self.my_slot.z0 = self.plt_z0
self.my_slot.z = plt_data
if plt_sel:
self.my_slot.plot_s_db(ax=self.ax)
else:
self.my_slot.plot_s_smith(ax=self.ax, draw_labels=True, show_legend=False,
label=label, color=color, chart_type='zy', marker=mark)
self.ax.legend(bbox_to_anchor=(0.5, 1.05), loc='lower center', ncol=3,
fancybox=True, shadow=True)
self.fig_cvs.draw()
|
<reponame>spmuppar/Adaptive-Tabulated-real-fluid-thermo<filename>quadtree_table_t.py
from quadtree_t import Node, QuadTree
#from quad_plot import draw_rectangle
import sys
import random
from pdb import set_trace as keyboard
from matplotlib.patches import Rectangle
import matplotlib.pyplot as plt
import NIST_reader as NIST
import pickle
import quad_utilities as utility
import numpy as np
from CoolProp.CoolProp import PropsSI
from skimage.transform import ProjectiveTransform
class CNode(Node):
#_______________________________________________________
# Overrides the base class method.
# Ensures Node.subdivide() uses instances of our custom
# class rather than instances of the base 'Node' class.
def getinstance(self, rect, rect_prop, index, n, accuracy, response, trans):
return CNode(self, rect, rect_prop, index, n, accuracy, response, trans)
# Test if the reconstructed values are within the error limits of the acutal values, if not subdivide
def spans_feature(self, rect, rect_prop, depth, accuracy, response, trans):
x0,z0,x1,z1 = rect
# dataNIST=NIST.readNIST(isoType = "isotherm", fluid = 'O2', T=x_mid, P=z_mid/1.0E6, tmin=x_mid, tmax=x_mid, pmin = z_mid/1.0E6, pmax = z_mid/1.0E6, N=1)
# dataNIST = np.ndarray.tolist(dataNIST)
# dataNIST = [item for sublist in dataNIST for item in sublist]
# del dataNIST[:2]
#dataNIST = utility.get_coolprop_TPS(x_mid, z_mid, response)
point_00 = [[rect[0],rect[1]], rect_prop[0]]
point_10 = [[rect[2],rect[1]], rect_prop[1]]
point_01 = [[rect[0],rect[3]], rect_prop[2]]
point_11 = [[rect[2],rect[3]], rect_prop[3]]
points = [point_00, point_10, point_01, point_11]
#reconstructing property values at num x num points inside each box
num = 7.0
del_x = (x1-x0)/(num-1.0)
del_z = (z1-z0)/(num-1.0)
x_l = np.linspace(x0+del_x,x1-del_x,num-2.0)
z_l = np.linspace(z0+del_z,z1-del_z,num-2.0)
xv = np.ndarray.flatten(np.meshgrid(x_l,z_l)[0])
zv = np.ndarray.flatten(np.meshgrid(x_l,z_l)[1])
int_points = zip(xv, zv)
mid_prop = [None]*25
dataNIST = [None]*25
for i,int_point in enumerate(int_points):
x_c = int_point[0]
z_c = int_point[1]
mid_prop[i] = self.bilinear_interpolation(x_c, z_c, points, trans)
#check if they're out of bounds
check = utility.check_out_of_bound(x_c, z_c, response, trans)
if check:
dataNIST[i] = mid_prop[i]
return False
dataNIST[i] = utility.get_coolprop_TPS(x_c, z_c, response, trans)
glob_error = [None]*25
for n in range(25):
lc_error = [None]*7
for e in range(7):
lc_error[e] = abs(mid_prop[n][e] - dataNIST[n][e])/dataNIST[n][e]
glob_error[n] = max(lc_error)
if (max(glob_error)<(accuracy/100.0)) or depth >= 13:
return False
# if depth >= 50:
# print error_rho, "This is the error in density"
# return False
# if response=='t':
# return True
# else:
# return False
# response = raw_input("Response")
# if response == 'Y':
return True
def bilinear_interpolation(self, x, y, points, trans):
q00=[None]*9
q01=[None]*9
q10=[None]*9
q11=[None]*9
prop_array=[]
##################------here points x0, y0 are of the order BL, TL, BR, TR
x, y = np.ndarray.tolist(trans.inverse([x, y])[0])
for p in range(9):
q00[p] = points[0][1][p]
x0, y0 = np.ndarray.tolist(trans.inverse(points[0][0])[0])
q01[p] = points[2][1][p]
_x0, y1 = np.ndarray.tolist(trans.inverse(points[2][0])[0])
q10[p] = points[1][1][p]
x1, _y0 = np.ndarray.tolist(trans.inverse(points[1][0])[0])
q11[p] = points[3][1][p]
_x1, _y1= np.ndarray.tolist(trans.inverse(points[3][0])[0])
#commented due to the roundoff error by the transforming function
# if x0 != _x0 or x1 != _x1 or y0 != _y0 or y1 != _y1:
# raise ValueError('points do not form a rectangle')
# sys.exit()
if not x0 <= x <= x1 or not y0 <= y <= y1:
raise ValueError('(x, y) not within the rectangle')
sys.exit()
bil = (q00[p] * (x1 - x) * (y1 - y) + q10[p] * (x - x0) * (y1 - y) + q01[p] * (x1 - x) * (y - y0) + q11[p] * (x - x0) * (y - y0)) / ((x1 - x0) * (y1 - y0) + 0.0)
prop_array.append(bil)
return prop_array
class CQuadTree(QuadTree):
#_______________________________________________________
def __init__(self, rootnode, minrect, accuracy, response, trans):
QuadTree.__init__(self, rootnode, minrect, accuracy, response, trans)
if __name__=="__main__":
print "#######################----ADAPTIVE TABULATION PROGRAM FOR THERMODYNAMIC EQUATION OF STATE------###########################"
print " "
response = raw_input("Enter the table index variables, T-P or rho-e: ")
if response == "T-P":
#T_min, T_max = [float(x) for x in raw_input("Enter the range of temperatures [T_min, T_max] in K (WITHOUT BRACES): ").split(',')]
#P_min, P_max = [float(x) for x in raw_input("Enter the range of temperatures [P_min, P_max] in Pa: (WITHOUT BRACES) ").split(',')]
#rootrect = [T_min, P_min, T_max, P_max]
rootrect = [200, 0.02E6, 600, 10.0E6]
b_left = [rootrect[0],rootrect[1]]
b_right = [rootrect[2],rootrect[1]]
t_left = [rootrect[0],rootrect[3]]
t_right = [rootrect[2],rootrect[3]]
### Transform to a square of dimensions [1024, 1024] ###
trans = ProjectiveTransform()
src = np.asarray([b_left, b_right, t_left, t_right])
dst = np.asarray([[0,0], [1023,0], [0,1023] ,[1024,1024]])
rootrect = [0, 0, 1024, 1024]
if not trans.estimate(src, dst):
raise Exception("estimate failed")
keyboard()
point_00 = [rootrect[0],rootrect[1]]
point_10 = [rootrect[2],rootrect[1]]
point_01 = [rootrect[0],rootrect[3]]
point_11 = [rootrect[2],rootrect[3]]
print("The initial rectangular domain of PT is", rootrect)
# points = [point_00, point_10, point_01, point_11]
elif response == "rho-e":
#rho_min, rho_max = [float(x) for x in raw_input("Enter the range of density [rho_min, rho_max] in Kg/m3 (WITHOUT BRACES): ").split(',')]
#e_min, e_max = [float(x) for x in raw_input("Enter the range of energies [e_min, e_max] in J/kg: (WITHOUT BRACES) ").split(',')]
#rootrect = [rho_min, e_min, rho_max, rho_max]
#rootrect = [0.14,-154110, 1208, 333470]
#rootrect = [1.4,-12, 717, 163000]
#rootrect = [1.4,-12000, 78, 163000]
rootrect = [0.14, -70793, 1305.20, 600000]
b_left = [rootrect[0],rootrect[1]]
b_right = [rootrect[2],rootrect[1]]
t_left = [rootrect[0],rootrect[3]]
t_right = [rootrect[2],rootrect[3]]
### Transform to a square of dimensions [1024, 1024] ###
trans = ProjectiveTransform()
src = np.asarray([b_left, b_right, t_left, t_right])
dst = np.asarray([[0,0], [1024,0], [0,1024] ,[1024,1024]])
rootrect = [0, 0, 1024, 1024]
if not trans.estimate(src, dst):
raise Exception("estimate failed")
keyboard()
point_00 = [rootrect[0],rootrect[1]]
point_10 = [rootrect[2],rootrect[1]]
point_01 = [rootrect[0],rootrect[3]]
point_11 = [rootrect[2],rootrect[3]]
print("The initial rectangular domain of rho-e is ", rootrect)
else:
print "The variables not recongnized !!"
sys.exit()
# points = [point_00, point_10, point_01, point_11]
rootrect_prop = [] #the properties at those given ranges(only on 4 boundary points)
rootrect_prop = utility.get_coolprop_TP(rootrect, response, trans)
resolution = 1
accuracy = float(raw_input("Enter the required accuracy in (%) "))
rootnode = CNode(None, rootrect, rootrect_prop, 0, 0, accuracy, response, trans)
tree = CQuadTree(rootnode, resolution, accuracy, response, trans)
#print "Done"
#pdb.set_trace()
#f=open("quadtree.pickle", "wb" )
#pickle.dump((tree), f )
#f.close()
print "End of program"
|
<gh_stars>1-10
"""
Contains all the machinery to register and load plugins.
"""
from __future__ import annotations
import importlib
import inspect
import sys
from abc import ABC
from dataclasses import dataclass
from pathlib import Path
from typing import Callable, List, Optional, Type
import wx
from .logging import logger
KNOWN_PLUGINS: List[Type[PluginBase]] = []
"""List of plugins registered as subclasses of PluginBase."""
@dataclass
class MenuTool:
"""Compiles the information required to add menu and toolbar items."""
menu: str = ""
id: int = wx.ID_ANY
text: str = ""
description: str = ""
short_help: str = ""
callback: Optional[Callable[[wx.Event], None]] = None
kind: wx.ItemKind = wx.ITEM_NORMAL
bitmap: Optional[wx.Bitmap] = None
@dataclass
class Tab:
"""Compiles the information required to define add tabs."""
page: wx.Window
text: str
select: bool = False
imageId: int = wx.NO_IMAGE
order: int = 0
class PluginBase(ABC):
"""
Base class that defines the required API that all the views of the different plugins
will need to provide.
"""
def __init_subclass__(cls: Type[PluginBase]):
if cls not in KNOWN_PLUGINS:
KNOWN_PLUGINS.append(cls)
def menu_entries(self) -> List[MenuTool]:
"""Return a list of menu entries provided by this plugin.
The items of the list must be MenuTool objects.
Returns:
A list of MenuTool objects provided by this plugin to be added to the
application menu bar.
"""
return []
def toolbar_items(self) -> List[MenuTool]:
"""Return a list of toolbar items provided by this plugin.
The items of the list must be MenuTool objects.
Returns:
A list of MenuTool objects provided by this plugin to be added to the
application toolbar.
"""
return []
def tabs(self, parent=None) -> List[Tab]:
"""Create and return the list of tabs provided by this plugin.
The elements of the list must be Tab objects.
Args:
parent: The application notebook, to be used as parent during the creation
of the tabs.
Returns:
A list of Tab objects provided by this plugin to be added to the application
notebook.
"""
return []
def central(self, parent=None) -> Optional[wx.Window]:
"""Central widget provided by this plugin.
Args:
parent: The main window, to be used as parent during the creation of the
widget.
Returns:
None (if no central widget provided) or any object derived from
wx.MainWindow (pretty much, any widget).
"""
return None
def collect_plugins(
path: Path, package: Optional[str] = None, add_to_path: bool = False
) -> List[str]:
"""Collects the plugins from the given location.
This function collects the modules and packages available in chosen location. If
they are not directly importable by Python (i.e. they are not in the path), either
the name of the containing package should be supplied or the add_to_path flag must
be set to True.
Eg. If path = "/some/location/with/tools/mytool", and "mytool" is an importable
package with "import mytool", plugins within "mytool" folder will be included with
no extra arguments.
If "mytool" is not importable but "tools" is, i.e. 'import tools.mytool' works,
then you should add "package='tools'".
If neither "mytool" nor anything else in its path is importable, then the path can
be added to the module's import path, sys.path, so the plugins within can be
imported as "import plugin".
Args:
path: Directory to explore.
package: Package in which the directory is contained.
add_to_path: If the directory should be added to the import path.
Returns:
List of modules and packages to be imported in the form
"package.subpackage.plugin", where "plugin" can be a module (a python file) or
a package itself (a folder containing a __init__.py file).
"""
if add_to_path:
sys.path.insert(1, str(path))
pkg = ""
else:
pkg = f"{path.stem}." if package is None else f"{package}.{path.stem}."
plugin_names = []
for p in path.glob("*.py"):
if not p.stem.startswith("__"):
plugin_names.append(f"{pkg}{p.stem}")
for p in path.glob("*/"):
if p.is_dir() and (p / "__init__.py").exists():
plugin_names.append(f"{pkg}{p.stem}")
return plugin_names
def collect_builtin_extensions():
"""Search for plugins to be loaded in the "extensions" subfolder.
These plugins can be single modules (simple_plugin.py) or more complex modules
defined as packages (i.e. they are folders with a __init__.py file).
Returns:
A list of plugins names to be loaded.
"""
frame = inspect.stack()[1]
caller_file = Path(frame[0].f_code.co_filename)
extensions = caller_file.parent / "extensions"
return collect_plugins(extensions, caller_file.parent.stem)
def load_plugins(plugin_list: List[str]):
"""Loads the plugins of the list.
Args:
plugin_list: A list of plugins to be loaded.
"""
for plugin in plugin_list:
try:
importlib.import_module(plugin)
except ModuleNotFoundError as err:
logger.warning(f"Plugin '{err.name}' could not be loaded. {err}")
|
<gh_stars>10-100
"""SQLite based metrics repositories."""
import logging
from typing import Optional, Iterable, List, Final
from sqlalchemy import insert, MetaData, Table, Column, Integer, Boolean, DateTime, String, Unicode, \
ForeignKey, Float, UnicodeText, JSON, update, select, delete
from sqlalchemy.engine import Connection, Result
from sqlalchemy.exc import IntegrityError
from jupiter.domain.adate import ADate
from jupiter.domain.difficulty import Difficulty
from jupiter.domain.eisen import Eisen
from jupiter.domain.entity_name import EntityName
from jupiter.domain.metrics.infra.metric_entry_repository import MetricEntryRepository, MetricEntryNotFoundError
from jupiter.domain.metrics.infra.metric_repository import MetricRepository, MetricAlreadyExistsError, \
MetricNotFoundError
from jupiter.domain.metrics.metric import Metric
from jupiter.domain.metrics.metric_entry import MetricEntry
from jupiter.domain.metrics.metric_key import MetricKey
from jupiter.domain.metrics.metric_unit import MetricUnit
from jupiter.domain.recurring_task_due_at_day import RecurringTaskDueAtDay
from jupiter.domain.recurring_task_due_at_month import RecurringTaskDueAtMonth
from jupiter.domain.recurring_task_due_at_time import RecurringTaskDueAtTime
from jupiter.domain.recurring_task_gen_params import RecurringTaskGenParams
from jupiter.domain.recurring_task_period import RecurringTaskPeriod
from jupiter.framework.base.entity_id import EntityId, BAD_REF_ID
from jupiter.framework.base.timestamp import Timestamp
from jupiter.repository.sqlite.infra.events import build_event_table, upsert_events
LOGGER = logging.getLogger(__name__)
class SqliteMetricRepository(MetricRepository):
"""A repository for metrics."""
_connection: Final[Connection]
_metric_table: Final[Table]
_metric_event_table: Final[Table]
def __init__(self, connection: Connection, metadata: MetaData) -> None:
"""Constructor."""
self._connection = connection
self._metric_table = Table(
'metric',
metadata,
Column('ref_id', Integer, primary_key=True, autoincrement=True),
Column('archived', Boolean, nullable=False),
Column('created_time', DateTime, nullable=False),
Column('last_modified_time', DateTime, nullable=False),
Column('archived_time', DateTime, nullable=True),
Column('the_key', String(64), nullable=False, index=True, unique=True),
Column('name', Unicode(), nullable=False),
Column('collection_project_ref_id', Integer, nullable=True),
Column('collection_period', String(), nullable=True),
Column('collection_eisen', JSON, nullable=True),
Column('collection_difficulty', String, nullable=True),
Column('collection_actionable_from_day', Integer, nullable=True),
Column('collection_actionable_from_month', Integer, nullable=True),
Column('collection_due_at_time', String, nullable=True),
Column('collection_due_at_day', Integer, nullable=True),
Column('collection_due_at_month', Integer, nullable=True),
Column('metric_unit', String(), nullable=True),
keep_existing=True)
self._metric_event_table = build_event_table(self._metric_table, metadata)
def create(self, metric: Metric) -> Metric:
"""Create a metric."""
try:
result = self._connection.execute(insert(self._metric_table).values(
ref_id=metric.ref_id.as_int() if metric.ref_id != BAD_REF_ID else None,
archived=metric.archived,
created_time=metric.created_time.to_db(),
last_modified_time=metric.last_modified_time.to_db(),
archived_time=metric.archived_time.to_db()
if metric.archived_time else None,
the_key=str(metric.key),
name=str(metric.name),
collection_project_ref_id=
metric.collection_params.project_ref_id.as_int() if metric.collection_params else None,
collection_period=metric.collection_params.period.value if metric.collection_params else None,
collection_eisen=[e.value for e in metric.collection_params.eisen] if metric.collection_params else [],
collection_difficulty=metric.collection_params.difficulty.value
if metric.collection_params and metric.collection_params.difficulty else None,
collection_actionable_from_day=metric.collection_params.actionable_from_day.as_int()
if metric.collection_params and metric.collection_params.actionable_from_day else None,
collection_actionable_from_month=metric.collection_params.actionable_from_month.as_int()
if metric.collection_params and metric.collection_params.actionable_from_month else None,
collection_due_at_time=str(metric.collection_params.due_at_time)
if metric.collection_params and metric.collection_params.due_at_time else None,
collection_due_at_day=metric.collection_params.due_at_day.as_int()
if metric.collection_params and metric.collection_params.due_at_day else None,
collection_due_at_month=metric.collection_params.due_at_month.as_int()
if metric.collection_params and metric.collection_params.due_at_month else None,
metric_unit=metric.metric_unit.value if metric.metric_unit else None))
except IntegrityError as err:
raise MetricAlreadyExistsError(f"Metric with key {metric.key} already exists") from err
metric.assign_ref_id(EntityId(str(result.inserted_primary_key[0])))
upsert_events(self._connection, self._metric_event_table, metric)
return metric
def save(self, metric: Metric) -> Metric:
"""Save a metric - it should already exist."""
result = self._connection.execute(
update(self._metric_table)
.where(self._metric_table.c.ref_id == metric.ref_id.as_int())
.values(
archived=metric.archived,
created_time=metric.created_time.to_db(),
last_modified_time=metric.last_modified_time.to_db(),
archived_time=metric.archived_time.to_db() if metric.archived_time else None,
the_key=str(metric.key),
name=str(metric.name),
collection_project_ref_id=metric.collection_params.project_ref_id.as_int()
if metric.collection_params else None,
collection_period=metric.collection_params.period.value if metric.collection_params else None,
collection_eisen=[e.value for e in metric.collection_params.eisen] if metric.collection_params else [],
collection_difficulty=metric.collection_params.difficulty.value
if metric.collection_params and metric.collection_params.difficulty else None,
collection_actionable_from_day=metric.collection_params.actionable_from_day.as_int()
if metric.collection_params and metric.collection_params.actionable_from_day else None,
collection_actionable_from_month=metric.collection_params.actionable_from_month.as_int()
if metric.collection_params and metric.collection_params.actionable_from_month else None,
collection_due_at_time=str(metric.collection_params.due_at_time)
if metric.collection_params and metric.collection_params.due_at_time else None,
collection_due_at_day=metric.collection_params.due_at_day.as_int()
if metric.collection_params and metric.collection_params.due_at_day else None,
collection_due_at_month=metric.collection_params.due_at_month.as_int()
if metric.collection_params and metric.collection_params.due_at_month else None,
metric_unit=metric.metric_unit.value if metric.metric_unit else None))
if result.rowcount == 0:
raise MetricNotFoundError(f"Metric with key {metric.key} does not exist")
upsert_events(self._connection, self._metric_event_table, metric)
return metric
def load_by_key(self, key: MetricKey) -> Metric:
"""Find a metric by key."""
query_stmt = select(self._metric_table).where(self._metric_table.c.the_key == str(key))
result = self._connection.execute(query_stmt).first()
if result is None:
raise MetricNotFoundError(f"Metric with key {key} does not exist")
return self._row_to_entity(result)
def load_by_id(self, ref_id: EntityId, allow_archived: bool = False) -> Metric:
"""Find a metric by id."""
query_stmt = select(self._metric_table).where(self._metric_table.c.ref_id == ref_id.as_int())
if not allow_archived:
query_stmt = query_stmt.where(self._metric_table.c.archived.is_(False))
result = self._connection.execute(query_stmt).first()
if result is None:
raise MetricNotFoundError(f"Metric with id {ref_id} does not exist")
return self._row_to_entity(result)
def find_all(
self,
allow_archived: bool = False,
filter_ref_ids: Optional[Iterable[EntityId]] = None,
filter_keys: Optional[Iterable[MetricKey]] = None) -> List[Metric]:
"""Find all metrics matching some criteria."""
query_stmt = select(self._metric_table)
if not allow_archived:
query_stmt = query_stmt.where(self._metric_table.c.archived.is_(False))
if filter_ref_ids:
query_stmt = query_stmt.where(self._metric_table.c.ref_id.in_(fi.as_int() for fi in filter_ref_ids))
if filter_keys:
query_stmt = query_stmt.where(
self._metric_table.c.the_key.in_(str(k) for k in filter_keys))
results = self._connection.execute(query_stmt)
return [self._row_to_entity(row) for row in results]
def remove(self, ref_id: EntityId) -> Metric:
"""Hard remove a metric - an irreversible operation."""
query_stmt = select(self._metric_table).where(self._metric_table.c.ref_id == ref_id.as_int())
result = self._connection.execute(query_stmt).first()
if result is None:
raise MetricNotFoundError(f"Metric with id {ref_id} does not exist")
self._connection.execute(delete(self._metric_table).where(self._metric_table.c.ref_id == ref_id.as_int()))
return self._row_to_entity(result)
@staticmethod
def _row_to_entity(row: Result) -> Metric:
return Metric(
_ref_id=EntityId.from_raw(str(row["ref_id"])),
_archived=row["archived"],
_created_time=Timestamp.from_db(row["created_time"]),
_archived_time=Timestamp.from_db(row["archived_time"])
if row["archived_time"] else None,
_last_modified_time=Timestamp.from_db(row["last_modified_time"]),
_events=[],
_key=MetricKey.from_raw(row["the_key"]),
_name=EntityName.from_raw(row["name"]),
_collection_params=RecurringTaskGenParams(
project_ref_id=EntityId.from_raw(str(row["collection_project_ref_id"])),
period=RecurringTaskPeriod.from_raw(row["collection_period"]),
eisen=[Eisen.from_raw(e) for e in row["collection_eisen"]],
difficulty=Difficulty.from_raw(row["collection_difficulty"])
if row["collection_difficulty"] else None,
actionable_from_day=RecurringTaskDueAtDay(row["collection_actionable_from_day"])
if row["collection_actionable_from_day"] is not None else None,
actionable_from_month=RecurringTaskDueAtMonth(row["collection_actionable_from_month"])
if row["collection_actionable_from_month"] is not None else None,
due_at_time=RecurringTaskDueAtTime.from_raw(row["collection_due_at_time"])
if row["collection_due_at_time"] is not None else None,
due_at_day=RecurringTaskDueAtDay(row["collection_due_at_day"])
if row["collection_due_at_day"] is not None else None,
due_at_month=RecurringTaskDueAtMonth(row["collection_due_at_month"])
if row["collection_due_at_month"] is not None else None)
if row["collection_project_ref_id"] is not None and row["collection_period"] is not None else None,
_metric_unit=MetricUnit.from_raw(row["metric_unit"])
if row["metric_unit"] else None)
class SqliteMetricEntryRepository(MetricEntryRepository):
"""A repository of metric entries."""
_connection: Final[Connection]
_metric_entry_table: Final[Table]
_metric_entry_event_table: Final[Table]
def __init__(self, connection: Connection, metadata: MetaData) -> None:
"""Constructor."""
self._connection = connection
self._metric_entry_table = Table(
'metric_entry',
metadata,
Column('ref_id', Integer, primary_key=True, autoincrement=True),
Column('archived', Boolean, nullable=False),
Column('created_time', DateTime, nullable=False),
Column('last_modified_time', DateTime, nullable=False),
Column('archived_time', DateTime, nullable=True, index=True),
Column('metric_ref_id', ForeignKey('metric.ref_id'), nullable=False),
Column('collection_time', DateTime, nullable=False),
Column('value', Float, nullable=False),
Column('notes', UnicodeText, nullable=True),
keep_existing=True)
self._metric_entry_event_table = build_event_table(self._metric_entry_table, metadata)
def create(self, metric_entry: MetricEntry) -> MetricEntry:
"""Create a metric entry."""
result = self._connection.execute(insert(self._metric_entry_table).values(
ref_id=metric_entry.ref_id.as_int() if metric_entry.ref_id != BAD_REF_ID else None,
archived=metric_entry.archived,
created_time=metric_entry.created_time.to_db(),
last_modified_time=metric_entry.last_modified_time.to_db(),
archived_time=metric_entry.archived_time.to_db() if metric_entry.archived_time else None,
metric_ref_id=metric_entry.metric_ref_id.as_int(),
collection_time=metric_entry.collection_time.to_db(),
value=metric_entry.value,
notes=metric_entry.notes))
metric_entry.assign_ref_id(EntityId(str(result.inserted_primary_key[0])))
upsert_events(self._connection, self._metric_entry_event_table, metric_entry)
return metric_entry
def save(self, metric_entry: MetricEntry) -> MetricEntry:
"""Save a metric entry - it should already exist."""
result = self._connection.execute(
update(self._metric_entry_table)
.where(self._metric_entry_table.c.ref_id == metric_entry.ref_id.as_int())
.values(
archived=metric_entry.archived,
created_time=metric_entry.created_time.to_db(),
last_modified_time=metric_entry.last_modified_time.to_db(),
archived_time=metric_entry.archived_time.to_db() if metric_entry.archived_time else None,
metric_ref_id=metric_entry.metric_ref_id.as_int(),
collection_time=metric_entry.collection_time.to_db(),
value=metric_entry.value,
notes=metric_entry.notes))
if result.rowcount == 0:
raise MetricEntryNotFoundError(f"Metric entry with id {metric_entry.ref_id} does not exist")
upsert_events(self._connection, self._metric_entry_event_table, metric_entry)
return metric_entry
def load_by_id(self, ref_id: EntityId, allow_archived: bool = False) -> MetricEntry:
"""Load a given metric entry."""
query_stmt = select(self._metric_entry_table).where(self._metric_entry_table.c.ref_id == ref_id.as_int())
if not allow_archived:
query_stmt = query_stmt.where(self._metric_entry_table.c.archived.is_(False))
result = self._connection.execute(query_stmt).first()
if result is None:
raise MetricEntryNotFoundError(f"Metric entry with id {ref_id} does not exist")
return self._row_to_entity(result)
def find_all_for_metric(self, metric_ref_id: EntityId, allow_archived: bool = False) -> List[MetricEntry]:
"""Retrieve all metric entries for a given metric."""
query_stmt = select(self._metric_entry_table)\
.where(self._metric_entry_table.c.metric_ref_id == metric_ref_id.as_int())
if not allow_archived:
query_stmt = query_stmt.where(self._metric_entry_table.c.archived.is_(False))
results = self._connection.execute(query_stmt)
return [self._row_to_entity(row) for row in results]
def find_all(
self,
allow_archived: bool = False,
filter_ref_ids: Optional[Iterable[EntityId]] = None,
filter_metric_ref_ids: Optional[Iterable[EntityId]] = None) -> List[MetricEntry]:
"""Find all metric entries matching some criteria."""
query_stmt = select(self._metric_entry_table)
if not allow_archived:
query_stmt = query_stmt.where(self._metric_entry_table.c.archived.is_(False))
if filter_ref_ids:
query_stmt = query_stmt.where(self._metric_entry_table.c.ref_id.in_(fi.as_int() for fi in filter_ref_ids))
if filter_metric_ref_ids:
query_stmt = query_stmt\
.where(self._metric_entry_table.c.metric_ref_id.in_(fi.as_int() for fi in filter_metric_ref_ids))
results = self._connection.execute(query_stmt)
return [self._row_to_entity(row) for row in results]
def remove(self, ref_id: EntityId) -> MetricEntry:
"""Hard remove a metric entry - an irreversible operation."""
query_stmt = select(self._metric_entry_table).where(self._metric_entry_table.c.ref_id == ref_id.as_int())
result = self._connection.execute(query_stmt).first()
if result is None:
raise MetricEntryNotFoundError(f"Metric entry with id {ref_id} does not exist")
self._connection.execute(
delete(self._metric_entry_table).where(self._metric_entry_table.c.ref_id == ref_id.as_int()))
return self._row_to_entity(result)
@staticmethod
def _row_to_entity(row: Result) -> MetricEntry:
return MetricEntry(
_ref_id=EntityId.from_raw(str(row["ref_id"])),
_archived=row["archived"],
_created_time=Timestamp.from_db(row["created_time"]),
_archived_time=Timestamp.from_db(row["archived_time"])
if row["archived_time"] else None,
_last_modified_time=Timestamp.from_db(row["last_modified_time"]),
_events=[],
_metric_ref_id=EntityId.from_raw(str(row["metric_ref_id"])),
_collection_time=ADate.from_db(row["collection_time"]),
_value=row["value"],
_notes=row["notes"])
|
<filename>differdb.py
'''
Copyright (c) 2012 Lolapps, Inc. All rights reserved.
Redistribution and use in source and binary forms, with or without modification, are
permitted provided that the following conditions are met:
1. Redistributions of source code must retain the above copyright notice, this list of
conditions and the following disclaimer.
2. Redistributions in binary form must reproduce the above copyright notice, this list
of conditions and the following disclaimer in the documentation and/or other materials
provided with the distribution.
THIS SOFTWARE IS PROVIDED BY LOLAPPS, INC. ''AS IS'' AND ANY EXPRESS OR IMPLIED
WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND
FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL LOLAPPS, INC. OR
CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR
CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR
SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON
ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING
NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF
ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
The views and conclusions contained in the software and documentation are those of the
authors and should not be interpreted as representing official policies, either expressed
or implied, of Lolapps, Inc..
--------------------------------------------------------------------------------------------
differdb.py
Putting all the various functions inside of here for interacting
with the differ database and tables.
'''
import sqlalchemy
import time
import util
from settings import *
LEN_CODE_METHOD = 32
LEN_PRODUCT = 8
DEBUG, INFO, WARNING, ERROR, CRITICAL = range(10, 51, 10) # based on logging
def get_log_type(error_msg):
if 'WARNING' in error_msg:
return WARNING
return ERROR
class LolflyError(object):
def __init__(self, filename, differ_db):
self.differ_db = differ_db
self.file_name = filename # name of the file where the error occurred
self.initialize()
def initialize(self):
self.timestamp = time.strftime('%Y%m%d %H:%M:%S', time.localtime())
self.product = None # attempt to figure out what we are
self.revision = None # revision info, get this from the filesystem
self.error_msg = None # the full error message
self.line_number = None # the line number of the error
self.location = None # the location of the error
self.method = None # the method where the error occurred
def print_pretty(self):
print ("%s,%s,%s,%s,%s,%s" % (self.file_name,
self.product,
self.location,
self.method,
self.error_msg,
self.exception))
def differ_db_inject(self):
self.differ_db.add_differ_error(self.file_name,
self.product,
self.location,
self.method,
self.error_msg,
self.exception,
int(time.time()),
util.get_differ_hostname().strip())
class DifferDB(object):
""" Basic DB class for interacting with our database.
Provides an initial database object so that we can easily re-use
the database connection.
"""
def __init__(self, dbhost=DIFFERDBHOST, dbuser=DIFFERDBUSER, dbpasswd=<PASSWORD>, db=DIFFERDB):
""" Init our database object
"""
engine_def = 'mysql://%s:%s@%s/%s' % (dbuser, dbpasswd, dbhost, db)
self.engine = sqlalchemy.create_engine(engine_def, echo=False)
def add_differ_error(self, logfile, product, code_location, code_method, error_message,
exception, timestamp, host):
""" add_differ_error is intended to be used by the various differ
clients. Once they encounter an error and do their parsing, this
function is used for entering that data into the database.
"""
query_dict = {}
query_dict['logfile'] = logfile
if product:
product = product[:LEN_PRODUCT]
query_dict['product'] = product
query_dict['code_location'] = code_location
if code_method:
code_method = code_method[:LEN_CODE_METHOD]
query_dict['code_method'] = code_method
query_dict['error_message'] = error_message
query_dict['exception'] = exception
query_dict['timestamp'] = timestamp
query_dict['host'] = host
table_name = ('differ_errors' if get_log_type(error_message) >= ERROR
else 'differ_warnings')
query = sqlalchemy.sql.text("""
INSERT INTO %s (timestamp, host,
logfile, product, code_location, code_method, error_message, exception)
VALUES (:timestamp, :host, :logfile, :product, :code_location, :code_method,
:error_message, :exception)
""" % table_name)
attempts = 0
while attempts < 5:
attempts += 1
try:
self.engine.execute(query, query_dict)
break
except sqlalchemy.exceptions.OperationalError, e:
util.write_log('%s' % e)
time.sleep(attempts*2)
def get_grouped_unfiled_exceptions(self):
""" get_grouped_unfiled_exceptions
same as get_unfiled_exceptions, except we try to do some grouping here
"""
query = """
SELECT count(*) as count,product,code_location,code_method,
exception,error_message,logfile,host
FROM differ_errors
WHERE fbz_case IS NULL
AND exception IS NOT NULL
GROUP BY product,code_location,code_method,exception
"""
results = self.engine.execute(query)
return results
def get_unfiled_nonexceptions(self, limit=1):
""" get_unfiled_nonexceptions
Basic query function to return values from differ entries that
do *not* have an exception value and do *not* have a fbz_case
associated with it.
Takes a single argument which is the number of entries to return.
"""
query = sqlalchemy.sql.text("""
SELECT id, timestamp, host, logfile, product,
code_location, code_method, exception, error_message
FROM differ_errors
WHERE fbz_case IS NULL AND exception IS NULL
LIMIT :limit""")
query_dict = {'limit': limit}
results = self.engine.execute(query, query_dict)
return results
def get_grouped_filed_errors(self, start, product='%%'):
""" get_grouped_filed_errors
Returns the different errors returned since the specified start time
that have been filed and groups them.
Takes two args:
start : epoch time of the start
product : the product you want to summarize [optional]
"""
query = sqlalchemy.sql.text("""
SELECT count(*) as count, product, code_location, code_method,
exception, fbz_case, error_message
FROM differ_errors
WHERE product LIKE :product AND fbz_case IS NOT NULL AND timestamp > :start
GROUP BY fbz_case
ORDER BY count desc
""")
query_dict = {'start': start, 'product': product}
results = self.engine.execute(query, query_dict)
return results
def get_grouped_warnings(self, start, product='%%'):
'''Return warnings from ``start``, grouped by code location and product.
Args:
start (int): Unix epoch timestamp
product (string): the product, default to ``%%``
Returns:
An SQL Alchemy result set
'''
query = sqlalchemy.sql.text("""
SELECT count(*) as count,product,code_location,code_method,
exception,error_message,logfile,host
FROM differ_warnings
WHERE product LIKE :product AND timestamp > :start
GROUP BY product,code_location,code_method,exception
ORDER BY count DESC
""")
query_dict = {'start': start, 'product': product}
results = self.engine.execute(query, query_dict)
return results
def update_case_id(self, errorid, fbz_case):
""" update_case_id
Give it an id, and it will update the differ table with that id
with the fogbugz case number
"""
query = sqlalchemy.sql.text("""
UPDATE differ_errors
SET fbz_case=:fbz_case
WHERE id=:errorid
""")
query_dict = {}
query_dict['errorid'] = errorid
query_dict['fbz_case'] = fbz_case
self.engine.execute(query, query_dict)
def update_group_case_id(self, fbz_case, code_location, code_method, exception):
query = sqlalchemy.sql.text("""
UPDATE differ_errors
SET fbz_case=:fbz_case
WHERE code_location=:code_location AND code_method=:code_method
AND exception=:exception AND fbz_case IS NULL
""")
query_dict = {}
query_dict['fbz_case'] = fbz_case
query_dict['code_location'] = code_location
query_dict['code_method'] = code_method
query_dict['exception'] = exception
self.engine.execute(query, query_dict)
def update_group_product(self, product, fbz_case, code_location, code_method, exception):
query = sqlalchemy.sql.text("""
UPDATE differ_errors
SET product=:product
WHERE code_location=:code_location AND code_method=:code_method
AND exception=:exception AND fbz_case=:fbz_case
""")
query_dict = {}
query_dict['fbz_case'] = fbz_case
query_dict['code_location'] = code_location
query_dict['code_method'] = code_method
query_dict['exception'] = exception
if product:
product = product[:LEN_PRODUCT]
query_dict['product'] = product
self.engine.execute(query, query_dict)
def error_count(self, duration):
""" just return the number of errors over the past X seconds
"""
current = int(time.time())
start = current - duration
query = sqlalchemy.sql.text("""
SELECT count(*) FROM differ_errors WHERE timestamp > :start
""")
query_dict = {}
query_dict['start'] = start
results = self.engine.execute(query, query_dict).scalar()
return results
def close_connection(self):
self.engine.dispose()
|
<gh_stars>1-10
# Train multiple images per person
# Find and recognize faces in an image using a SVC with scikit-learn
"""
Structure:
<test_image>.jpg
<train_dir>/
<person_1>/
<person_1_face-1>.jpg
<person_1_face-2>.jpg
.
.
<person_1_face-n>.jpg
<person_2>/
<person_2_face-1>.jpg
<person_2_face-2>.jpg
.
.
<person_2_face-n>.jpg
.
.
<person_n>/
<person_n_face-1>.jpg
<person_n_face-2>.jpg
.
.
<person_n_face-n>.jpg
"""
import face_recognition
from sklearn import svm
import os
# Training the SVC classifier
# The training data would be all the face encodings from all the known images and the labels are their names
# Test directory
test_dir_path = './test_dir/'
output_dir = "./out_dir/"
import pickle
from PIL import Image, ImageDraw
import numpy as np
def draw_name_on_face(image_path, person):
known_image = face_recognition.load_image_file(image_path)
# Find all the faces and face encodings in the unknown image
face_locations = face_recognition.face_locations(known_image)
face_encodings = face_recognition.face_encodings(known_image, face_locations)
# Convert the image to a PIL-format image so that we can draw on top of it with the Pillow library
# See http://pillow.readthedocs.io/ for more about PIL/Pillow
pil_image = Image.fromarray(known_image)
# Create a Pillow ImageDraw Draw instance to draw with
draw = ImageDraw.Draw(pil_image)
# Loop through each face found in the unknown image
for (top, right, bottom, left), face_encoding in zip(face_locations, face_encodings):
# See if the face is a match for the known face(s)
#matches = face_recognition.compare_faces(known_face_encodings, face_encoding)
name = person[0].replace("pins_","")
print(name)
# If a match was found in known_face_encodings, just use the first one.
# if True in matches:
# first_match_index = matches.index(True)
# name = known_face_names[first_match_index]
# Or instead, use the known face with the smallest distance to the new face
#face_distances = face_recognition.face_distance(known_face_encodings, face_encoding)
#best_match_index = np.argmin(face_distances)
#if matches[best_match_index]:
# name = known_face_names[best_match_index]
left = left -10
right = right + 10
bottom = bottom + 25
top = top - 20
# Draw a box around the face using the Pillow module
draw.rectangle(((left, top), (right, bottom)), outline=(0, 0, 255))
# Draw a label with a name below the face
text_width, text_height = draw.textsize(name)
draw.rectangle(((left, bottom - text_height - 10), (right, bottom)), fill=(0, 0, 255), outline=(0, 0, 255))
draw.text((left + 6, bottom - text_height - 5), name, fill=(255, 255, 255, 255))
# Remove the drawing library from memory as per the Pillow docs
del draw
# Display the resulting image
pil_image.show()
# You can also save a copy of the new image to disk if you want by uncommenting this line
#get image name
image_file_array = image_path.split("/")
image_file_name = image_file_array[len(image_file_array) - 1]
pil_image.save(output_dir + image_file_name)
modelfilename = 'finalized_model.sav'
clf = pickle.load(open(modelfilename, 'rb'))
# Load the test image with unknown faces into a numpy array
for subdir, dirs, files in os.walk(test_dir_path):
for file in files:
image_path = os.path.join(subdir, file)
if (image_path.find(".jpg") == -1):
continue
test_image = face_recognition.load_image_file(image_path)
# Find all the faces in the test image using the default HOG-based model
face_locations = face_recognition.face_locations(test_image)
no = len(face_locations)
print("Number of faces detected: ", no)
# Predict all the faces in the test image using the trained classifier
print("Found faces in image " + image_path)
for i in range(no):
test_image_enc = face_recognition.face_encodings(test_image)[i]
name = clf.predict([test_image_enc])
draw_name_on_face(image_path, name)
#print(*name)
|
import xarray as xr
import glob
import numpy as np
import pandas as pd
import os
import json
from netCDF4 import Dataset, stringtochar
from .aux.file_to_radar_object import file_to_radar_object
from .aux.get_var_arrays_from_radar_object import get_var_arrays_from_radar_object
from .iah_filter import iah_filter_ppi, iah_filter_rhi
from .rh_filter import rh_filter
# filters.py
# create filters for a day using all day filters
# use functions: iah_filter, rh_filter
def filters(radar_config_file, date, met_path):
"""
filters loops through a day's worth of radar files (specify PPI or HSRHI, dual or horizontal polarization),
and calculates IAH and RH filters (if desired) for every file in a day.
A netCDF for each day with a filter-passing array is stored.
Parameters
----------
radar_config_file: str
path to JSON file containing specifications: data directory, file extension, clutter map directory, baseline directory, baseline date, daily CSV directory, scan type, polarization, site, instrument, range limit
date: str
YYYYMMDD specifying date of interest
met_path: str
path to cormet files
"""
os.environ['HDF5_USE_FILE_LOCKING'] = 'FALSE'
print(date)
config_vars = json.load(open(radar_config_file))
datadir = config_vars["data_directory"]
extension = config_vars["file_extension"]
cluttermap_dir = config_vars["cluttermap_directory"]
baseline_dir = config_vars["baseline_directory"]
baseline_date = config_vars["baseline_date"]
dailycsvdir = config_vars["daily_csv_dir"]
scantype = config_vars["scan_type"]
polarization = config_vars["polarization"]
site = config_vars["site_abbrev"]
inst = config_vars["instrument_abbrev"]
range_limit = config_vars["range_limit"]
# Identify which radar band you are using (change if statement as needed)
# Most important to identify Ka-band radars
if inst == "kasacr":
radar_band = "ka"
else:
radar_band = inst[0]
# Read in clutter map netCDF and baseline value netCDF
dataset = Dataset(
cluttermap_dir
+ "cluttermap_"
+ scantype
+ "_"
+ site
+ inst
+ "_"
+ "composite"
+ ".nc"
)
dataset_b = Dataset(
baseline_dir
+ "baseline_"
+ scantype
+ "_"
+ site
+ inst
+ "_"
+ baseline_date
+ ".nc"
)
if scantype == "ppi":
clutter_map_mask_h = dataset.variables["clutter_map_mask_zh"][:, :]
baseline_dbz95_h = dataset_b.variables["baseline_dbz95_zh"][:]
elif scantype == "rhi":
clutter_map_mask_h = dataset.variables["clutter_map_mask_zh"][:, :, :]
baseline_dbz95_h = dataset_b.variables["baseline_dbz95_zh"][:]
if polarization == "dual" and scantype == "ppi":
clutter_map_mask_v = dataset.variables["clutter_map_mask_zv"][:, :]
baseline_dbz95_v = dataset_b.variables["baseline_dbz95_zv"][:]
elif polarization == "dual" and scantype == "rhi":
clutter_map_mask_v = dataset.variables["clutter_map_mask_zv"][:, :, :]
baseline_dbz95_v = dataset_b.variables["baseline_dbz95_zv"][:]
dataset.close()
dataset_b.close()
# Get all the cormet files for a specified day
met_files = glob.glob(met_path+'*'+date+'*.cdf')
met_files.sort()
met = xr.open_mfdataset(met_files)
met = met.to_dataframe()
# Set RH (%) threshold
rh_thresh = 90
# Get all files in a day
day_files = []
#for f in glob.glob(os.path.join(datadir, "*" + date + ".*.??")):
for f in glob.glob(os.path.join(datadir, "*" + date + ".20*.??")):
day_files.append(f)
# Sort files chronologically
day_files.sort()
# For each time file, calculate filters
iah_filter_list = []
rh_filter_list = []
rh_list = []
datetime_list = []
for idx_f, f in enumerate(day_files):
# print(f)
extension = f[-3:]
radar = file_to_radar_object(f, extension)
var_dict = get_var_arrays_from_radar_object(radar, radar_config_file)
date_time_iah, pass_filter_iah = iah_filter_rhi(var_dict,
polarization,
range_limit,
radar_band,
clutter_map_mask_h,
clutter_mask_v=None
)
iah_filter_list.append(pass_filter_iah)
date_time_rh, rh, pass_filter_rh = rh_filter(date,
var_dict,
met,
rh_thresh
)
datetime_list.append(date_time_iah[0:19])
rh_list.append(rh)
rh_filter_list.append(pass_filter_rh)
# Convert lists to arrays
#datetime_array = np.asarray(datetime_list)
#rh_array = np.array(rh_list)
iah_filter_array = np.array(iah_filter_list)
rh_filter_array = np.array(rh_filter_list)
filter_array = np.zeros(iah_filter_array.shape, dtype=int)
iah_pass = iah_filter_array > 0
rh_pass = rh_filter_array > 0
pass_filter = np.logical_and(iah_pass,rh_pass)
filter_array[pass_filter] = 1
print(type(rh_filter_array), type(iah_filter_array), type(filter_array))
print(rh_filter_array.dtype, iah_filter_array.dtype, filter_array.dtype)
print(filter_array.dtype.str)
# Convert date_time string to character
#date_time_str_out = stringtochar(datetime_array, 'S18')
# Write filter arrays to netCDF
d = Dataset(
dailycsvdir+'/filters/'
+ "filters_"
+ scantype
+ "_"
+ site
+ inst
+ "_"
+ date
+ ".nc",
"w",
format="NETCDF4",
)
print(d.data_model)
print(len(filter_array))
arr_len = d.createDimension("len",len(filter_array))
print(d.dimensions)
#scalar_example = d.createVariable("scalar","")
total_filter = d.createVariable("iah_and_rh_filter",np.int64,("len",))
#total_filter = d.createVariable("iah_and_rh_filter",rh_filter_array.dtype.str,("len",))
#print(total_filter)
print(type(filter_array), filter_array.dtype)
d.close()
#arr_len = d.createDimension("arr_len", len(filter_array))
#str_len = d.createDimension("nchar", len(datetime_array))
#total_filter = d.createVariable("iah_and_rh_filter", 'f4', ("arr_len",))
#iah_fil = d.createVariable("iah_filter", 'f4', ("arr_len",))
#rh_fil = d.createVariable("rh_filter", 'f4', ("arr_len",))
#rh_value = d.createVariable("rh_value", 'f4', ("arr_len",))
#datetime = d.createVariable("datetime", 'S4', ("arr_len",))
#total_filter.long_name = "Combined filter using IAH and RH filters"
#iah_fil.long_name = "IAH filter"
#rh_fil.long_name = "RH filter"
#rh_value.long_name = "RH value"
#datetime.long_name = "Datetime"
#print(type(filter_array), type(datetime_array))
#print(filter_array.dtype, datetime_array.dtype)
#total_filter[:] = filter_array
#iah_fil[:] = iah_filter_array
#rh_fil[:] = rh_filter_array
#rh_value[:] = rh_array
#datetime[:] = datetime_array
#datetime[:] = date_time_str_out
#d.close()
|
"""
Test extensions
"""
import os
import unittest
from cement.core import handler
from cement.utils import shell
from scilifelab.pm.core.production import ProductionController
from test_default import PmTest
filedir = os.path.abspath(os.path.dirname(os.path.realpath(__file__)))
class PmShellTest(PmTest):
def test_wait(self):
"""Test that submitted shell jobs are run sequentially"""
print "running first sleep"
out = shell.exec_cmd(["sleep", "3"])
print "finishing first sleep"
print "running second sleep"
shell.exec_cmd(["sleep", "3"])
print "finishing second sleep"
class PmHsMetricsTest(PmTest):
def test_hsmetrics(self):
"""Run hs metrics"""
self.app = self.make_app(argv=['production', 'hs-metrics', 'J.Doe_00_01', '-f', '120829_SN0001_0001_AA001AAAXX', '--targets', os.path.join(filedir, 'regionfile'), '--force', '-n'], extensions=[])
handler.register(ProductionController)
self._run_app()
hsmetrics_str = "(DRY_RUN): java -Xmx3g -jar {}/CalculateHsMetrics.jar INPUT={}/120829_SN0001_0001_AA001AAAXX/1_120829_AA001AAAXX_nophix_8-sort-dup.bam TARGET_INTERVALS={}/regionfile BAIT_INTERVALS={}/regionfile OUTPUT={}/120829_SN0001_0001_AA001AAAXX/1_120829_AA001AAAXX_nophix_8-sort-dup.hs_metrics VALIDATION_STRINGENCY=SILENT".format(os.getenv("PICARD_HOME"), self.app.config.get("production", "root"), filedir, filedir, self.app.config.get("production", "root"))
self.eq(hsmetrics_str, str(sorted(self.app._output_data['stderr'].getvalue().rstrip().split("\n"))[-1]))
def test_hsmetrics_empty(self):
"""Run hs metrics when no files present"""
self.app = self.make_app(argv=['production', 'hs-metrics', 'J.Doe_00_02', '-f', '120829_SN0001_0001_AA001AAAXX','--targets', os.path.join(filedir, 'regionfile'), '--force', '-n'], extensions=[])
handler.register(ProductionController)
self._run_app()
## Shouldn't produce any output
self.eq([''], self.app._output_data['stdout'].getvalue().split("\n"))
@unittest.skipIf(not os.getenv("DRMAA_LIBRARY_PATH"), "not running production test: no $DRMAA_LIBRARY_PATH")
def test_hsmetrics_drmaa(self):
"""Run hs metrics over drmaa"""
self.app = self.make_app(argv=['production', 'hs-metrics', 'J.Doe_00_01', '-f', '120829_SN0001_0001_AA001AAAXX', '--targets', os.path.join(filedir, 'regionfile'), '--force', '-A', 'jobaccount', '--jobname', 'jobname', '--partition', 'node', '--time', '10:00:00', '--drmaa', '-n'], extensions=['scilifelab.pm.ext.ext_distributed'])
handler.register(ProductionController)
self._run_app()
hsmetrics_str = "(DRY_RUN): java -Xmx3g -jar {}/CalculateHsMetrics.jar INPUT={}/120829_SN0001_0001_AA001AAAXX/1_120829_AA001AAAXX_nophix_8-sort-dup.bam TARGET_INTERVALS={}/regionfile BAIT_INTERVALS={}/regionfile OUTPUT={}/120829_SN0001_0001_AA001AAAXX/1_120829_AA001AAAXX_nophix_8-sort-dup.hs_metrics VALIDATION_STRINGENCY=SILENT".format(os.getenv("PICARD_HOME"), self.app.config.get("production", "root"), filedir, filedir, self.app.config.get("production", "root"))
self.eq(hsmetrics_str, str(sorted(self.app._output_data['stderr'].getvalue().rstrip().split("\n"))[-1]))
|
<filename>streaming_event_compliance/services/build_automata/case_thread.py<gh_stars>1-10
from streaming_event_compliance import app
from streaming_event_compliance.objects.variable.globalvar import gVars, CL, T, C
from streaming_event_compliance.objects.automata import automata
from streaming_event_compliance.objects.exceptions.exception import ThreadException
from streaming_event_compliance.objects.logging.server_logging import ServerLogging
import threading
import traceback
import sys
check_executing_order = {}
WINDOW_SIZE = app.config['WINDOW_SIZE']
MAXIMUN_WINDOW_SIZE = app.config['MAXIMUN_WINDOW_SIZE']
def run_build(case_id):
func_name = sys._getframe().f_code.co_name
ServerLogging().log_info(func_name, str(threading.current_thread()))
try:
if C.lock_List.get(case_id).acquire():
ServerLogging().log_info(func_name, "server", case_id, "Acquiring lock")
windows_memory = C.dictionary_cases.get(case_id)[0: MAXIMUN_WINDOW_SIZE + 1]
C.dictionary_cases.get(case_id).pop(0)
C.lock_List.get(case_id).release()
ServerLogging().log_info(func_name, "server", case_id, "Released lock")
executing_order4test(case_id, windows_memory)
calculate_connection_for_different_prefix_automata(windows_memory)
ServerLogging().log_info(func_name, "server", case_id, "Calculating connections")
except Exception:
ServerLogging().log_error(func_name, "server", case_id, "Error with Caselock")
raise ThreadException(traceback.format_exc())
def calculate_connection_for_different_prefix_automata(windowsMemory):
"""
Description:
This function will calculate the connections with different size for the windowsMemory.
:param windowsMemory: :`list` a list of activities from the same case_id of current event(another event),
size is maximum_window_size, and the current event is in the last position of the
windowsMemory (i.e. event == windowsMemory[maximum_window_size]).
"""
for ws in WINDOW_SIZE:
source_node = ','.join(windowsMemory[MAXIMUN_WINDOW_SIZE - ws: MAXIMUN_WINDOW_SIZE])
sink_node = ','.join(windowsMemory[MAXIMUN_WINDOW_SIZE - ws + 1: MAXIMUN_WINDOW_SIZE + 1])
if CL.lock_list.get((source_node, sink_node)):
if CL.lock_list.get((source_node, sink_node)).acquire():
try:
if windowsMemory[MAXIMUN_WINDOW_SIZE] == '~!@#$%' and source_node.find('*') == -1:
gVars.autos.get(ws).update_automata(automata.Connection(source_node, '~!@#$%', 0))
elif source_node.find('*') == -1:
gVars.autos.get(ws).update_automata(automata.Connection(source_node, sink_node, 1))
elif source_node.find('*') != -1 and sink_node.find('*') == -1:
gVars.autos.get(ws).update_automata(automata.Connection('NONE', sink_node, 1))
CL.lock_list.get((source_node, sink_node)).release()
except Exception as ec:
raise ec
else:
lock = threading.RLock()
CL.lock_list[source_node, sink_node] = lock
if CL.lock_list.get((source_node, sink_node)).acquire():
try:
if windowsMemory[MAXIMUN_WINDOW_SIZE] == '~!@#$%' and source_node.find('*') == -1:
gVars.autos.get(ws).update_automata(automata.Connection(source_node, '~!@#$%', 0))
elif source_node.find('*') == -1:
gVars.autos.get(ws).update_automata(automata.Connection(source_node, sink_node, 1))
CL.lock_list.get((source_node, sink_node)).release()
except Exception as ec:
raise ec
def executing_order4test(case_id, windows_memory):
global check_executing_order
'''--------For Testing: Before releasing lock, which thread used it will be stored-------'''
if check_executing_order.get(case_id):
check_executing_order.get(case_id).append(windows_memory[MAXIMUN_WINDOW_SIZE])
else:
check_executing_order[case_id] = []
check_executing_order.get(case_id).append(windows_memory[MAXIMUN_WINDOW_SIZE])
'''--------For Testing: Before releasing lock, which thread used it will be stored-------'''
|
# Copyright (c) 2006-2008 The Chromium Authors. All rights reserved.
# Use of this source code is governed by a BSD-style license that can be
# found in the LICENSE file.
"""Some utility methods for getting paths used by run_webkit_tests.py.
"""
import errno
import os
import platform_utils
import subprocess
import sys
import google.path_utils
class PathNotFound(Exception): pass
# Save some paths here so we don't keep re-evaling.
_webkit_root = None
_layout_data_dir = None
_layout_tests_dir = None
# A map from platform description to directory list.
_platform_results_dirs = {}
# An instance of the PlatformUtility for use by methods mapped from there.
_platform_util = None
# TODO this should probably be moved into path_utils as ToUnixPath().
def WinPathToUnix(path):
"""Convert a windows path to use unix-style path separators (a/b/c)."""
return path.replace('\\', '/')
def WebKitRoot():
"""Returns the full path to the directory containing webkit.gyp. Raises
PathNotFound if we're unable to find webkit.gyp."""
global _webkit_root
if _webkit_root:
return _webkit_root
webkit_gyp_path = google.path_utils.FindUpward(google.path_utils.ScriptDir(),
'webkit.gyp')
_webkit_root = os.path.dirname(webkit_gyp_path)
return _webkit_root
def LayoutDataDir():
"""Gets the full path to the tests directory. Raises PathNotFound if
we're unable to find it."""
global _layout_data_dir
if _layout_data_dir:
return _layout_data_dir
_layout_data_dir = google.path_utils.FindUpward(WebKitRoot(), 'webkit',
'data', 'layout_tests')
return _layout_data_dir
def LayoutTestsDir(path = None):
"""Returns the full path to the directory containing layout tests, based on
the supplied relative or absolute path to a layout tests. If the path contains
"LayoutTests" directory, locates this directory, assuming it's either in
in webkit/data/layout_tests or in third_party/WebKit."""
if path != None and path.find('LayoutTests') == -1:
return LayoutDataDir()
global _layout_tests_dir
if _layout_tests_dir:
return _layout_tests_dir
if os.path.exists(os.path.join(LayoutDataDir(), 'LayoutTests')):
_layout_tests_dir = LayoutDataDir()
else:
_layout_tests_dir = google.path_utils.FindUpward(
google.path_utils.ScriptDir(), 'third_party', 'WebKit')
return _layout_tests_dir
def ChromiumPlatformResultsEnclosingDir():
"""Returns the full path to the directory containing Chromium platform
result directories.
"""
# TODO(pamg): Once we move platform/chromium-* into LayoutTests/platform/,
# remove this and use PlatformResultsEnclosingDir() for everything.
return os.path.join(LayoutDataDir(), 'platform')
def WebKitPlatformResultsEnclosingDir():
"""Gets the full path to just above the platform results directory."""
return os.path.join(LayoutTestsDir(), 'LayoutTests', 'platform')
def PlatformResultsEnclosingDir(platform):
"""Gets the path to just above the results directory for this platform."""
if platform.startswith('chromium'):
return ChromiumPlatformResultsEnclosingDir()
return WebKitPlatformResultsEnclosingDir()
def ExpectedFilename(filename, suffix, platform):
"""Given a test name, returns an absolute path to its expected results.
The result will be sought in the hierarchical platform directories, in the
corresponding WebKit platform directories, in the WebKit platform/mac/
directory, and finally next to the test file.
Suppose that the |platform| is 'chromium-win-xp'. In that case, the
following directories are searched in order, if they exist, and the first
match is returned:
LayoutTests/platform/chromium-win-xp/
LayoutTests/platform/chromium-win/
LayoutTests/platform/chromium/
LayoutTests/platform/win-xp/
LayoutTests/platform/win/
LayoutTests/platform/mac/
the directory in which the test itself is located
If the |platform| is 'chromium-mac-leopard', the sequence will be as follows:
LayoutTests/platform/chromium-mac-leopard/
LayoutTests/platform/chromium-mac/
LayoutTests/platform/chromium/
LayoutTests/platform/mac-leopard/
LayoutTests/platform/mac/
the directory in which the test itself is located
A platform may optionally fall back to the Windows results if its own
results are not found, by returning True from its platform-specific
platform_utils.IsNonWindowsPlatformTargettingWindowsResults(). Supposing
that Linux does so, the search sequence for the |platform| 'chromium-linux'
will be
LayoutTests/platform/chromium-linux/
LayoutTests/platform/chromium/
LayoutTests/platform/linux/
LayoutTests/platform/chromium-win/
LayoutTests/platform/win/
LayoutTests/platform/mac/
the directory in which the test itself is located
If no expected results are found in any of the searched directories, the
directory in which the test itself is located will be returned.
Args:
filename: absolute filename to test file
suffix: file suffix of the expected results, including dot; e.g. '.txt'
or '.png'. This should not be None, but may be an empty string.
platform: a hyphen-separated list of platform descriptors from least to
most specific, matching the WebKit format, that will be used to find
the platform/ directories to look in. For example, 'chromium-win' or
'chromium-mac-leopard'.
Returns:
An absolute path to the most specific matching result file for the given
test, following the search rules described above.
"""
testname = os.path.splitext(RelativeTestFilename(filename))[0]
# While we still have tests in LayoutTests/, chrome/, and pending/, we need
# to strip that outer directory.
# TODO(pamg): Once we upstream all of chrome/ and pending/, clean this up.
testdir, testname = testname.split('/', 1)
results_filename = testname + '-expected' + suffix
# Use the cached directory list if we have one.
global _platform_results_dirs
platform_dirs = _platform_results_dirs.get(platform, [])
if len(platform_dirs) == 0:
# Build the list of platform directories: chromium-foo-bar, chromium-foo,
# chromium.
segments = platform.split('-')
for length in range(len(segments), 0, -1):
platform_dirs.append('-'.join(segments[:length]))
# Append corresponding WebKit platform directories too.
if platform.startswith('chromium-'):
for length in range(len(segments), 1, -1):
platform_dirs.append('-'.join(segments[1:length]))
if platform_utils.IsNonWindowsPlatformTargettingWindowsResults():
if platform.startswith('chromium'):
platform_dirs.append('chromium-win')
platform_dirs.append('win')
# Finally, append platform/mac/ to all searches.
if 'mac' not in platform_dirs:
platform_dirs.append('mac')
platform_dirs = [os.path.join(PlatformResultsEnclosingDir(x), x)
for x in platform_dirs]
_platform_results_dirs[platform] = platform_dirs
for platform_dir in platform_dirs:
# TODO(pamg): Clean this up once we upstream everything in chrome/ and
# pending/.
if os.path.basename(platform_dir).startswith('chromium'):
platform_file = os.path.join(platform_dir, testdir, results_filename)
else:
platform_file = os.path.join(platform_dir, results_filename)
if os.path.exists(platform_file):
return platform_file
# If it wasn't found in a platform directory, return the expected result
# in the test's directory, even if no such file actually exists.
return os.path.join(os.path.dirname(filename),
os.path.basename(results_filename))
def TestShellBinaryPath(target):
"""Gets the full path to the test_shell binary for the target build
configuration. Raises PathNotFound if the file doesn't exist"""
platform_util = platform_utils.PlatformUtility('')
full_path = os.path.join(WebKitRoot(), target,
platform_util.TestShellBinary())
if not os.path.exists(full_path):
# try output directory from either Xcode or chrome.sln
full_path = platform_util.TestShellBinaryPath(target)
if not os.path.exists(full_path):
raise PathNotFound('unable to find test_shell at %s' % full_path)
return full_path
def LayoutTestHelperBinaryPath(target):
"""Gets the full path to the layout test helper binary for the target build
configuration. Raises PathNotFound if the file doesn't exist"""
platform_util = platform_utils.PlatformUtility('')
# try output directory from either Xcode or chrome.sln
full_path = platform_util.LayoutTestHelperBinaryPath(target)
if not os.path.exists(full_path):
raise PathNotFound('unable to find layout_test_helper at %s' % full_path)
return full_path
def RelativeTestFilename(filename):
"""Provide the filename of the test relative to the layout data
directory as a unix style path (a/b/c)."""
return WinPathToUnix(filename[len(LayoutTestsDir(filename)) + 1:])
def GetPlatformUtil():
"""Returns a singleton instance of the PlatformUtility."""
global _platform_util
if not _platform_util:
# Avoid circular import by delaying it.
import layout_package.platform_utils
_platform_util = (
layout_package.platform_utils.PlatformUtility(WebKitRoot()))
return _platform_util
# Map platform specific path utility functions. We do this as a convenience
# so importing path_utils will get all path related functions even if they are
# platform specific.
def GetAbsolutePath(path):
return GetPlatformUtil().GetAbsolutePath(path)
def FilenameToUri(path):
return GetPlatformUtil().FilenameToUri(path)
def TestListPlatformDir():
return GetPlatformUtil().TestListPlatformDir()
def PlatformDir():
return GetPlatformUtil().PlatformDir()
def PlatformNewResultsDir():
return GetPlatformUtil().PlatformNewResultsDir()
|
<reponame>bopopescu/fantastico
# -*- coding: utf-8 -*-
# MySQL Connector/Python - MySQL driver written in Python.
# Copyright (c) 2009, 2013, Oracle and/or its affiliates. All rights reserved.
# MySQL Connector/Python is licensed under the terms of the GPLv2
# <http://www.gnu.org/licenses/old-licenses/gpl-2.0.html>, like most
# MySQL Connectors. There are special exceptions to the terms and
# conditions of the GPLv2 as it is applied to this software, see the
# FOSS License Exception
# <http://www.mysql.com/about/legal/licensing/foss-exception.html>.
#
# This program is free software; you can redistribute it and/or modify
# it under the terms of the GNU General Public License as published by
# the Free Software Foundation.
#
# This program is distributed in the hope that it will be useful,
# but WITHOUT ANY WARRANTY; without even the implied warranty of
# MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# GNU General Public License for more details.
#
# You should have received a copy of the GNU General Public License
# along with this program; if not, write to the Free Software
# Foundation, Inc., 51 Franklin St, Fifth Floor, Boston, MA 02110-1301 USA
"""Unittests for examples
"""
import sys
import tests
import mysql.connector
class TestExamples(tests.MySQLConnectorTests):
def setUp(self):
config = self.getMySQLConfig()
self.cnx = mysql.connector.connect(**config)
def tearDown(self):
self.cnx.close()
def _exec_main(self, example):
try:
return example.main(self.getMySQLConfig())
except Exception as e:
self.fail(e)
def test_dates(self):
"""examples/dates.py"""
try:
import examples.dates as example
except Exception as e:
self.fail(e)
output = example.main(self.getMySQLConfig())
exp = [' 1 | 1977-06-14 | 1977-06-14 21:10:00 | 21:10:00 |',
' 2 | None | None | 0:00:00 |',
' 3 | None | None | 0:00:00 |']
self.assertEqual(output, exp)
example.DATA.append(('0000-00-00', None, '00:00:00'),)
self.assertRaises(mysql.connector.errors.IntegrityError,
example.main, self.getMySQLConfig())
def test_engines(self):
"""examples/engines.py"""
try:
import examples.engines as example
except:
self.fail()
output = self._exec_main(example)
# Can't check output as it might be different per MySQL instance
# We check only if MyISAM is present
found = False
for s in output:
if s.find('MyISAM') > -1:
found = True
break
self.assertTrue(found,'MyISAM engine not found in output')
def test_inserts(self):
"""examples/inserts.py"""
try:
import examples.inserts as example
except Exception as e:
self.fail(e)
output = self._exec_main(example)
exp = ['1 | Geert | 30\nInfo: c..\n',
'2 | Jan | 30\nInfo: c..\n', '3 | Michel | 30\nInfo: c..\n']
self.assertEqual(output,exp,'Output was not correct')
def test_transactions(self):
"""examples/transactions.py"""
db = mysql.connector.connect(**self.getMySQLConfig())
r = self.haveEngine(db,'InnoDB')
db.close()
if not r:
return
try:
import examples.transaction as example
except Exception as e:
self.fail(e)
output = self._exec_main(example)
exp = ['Inserting data', 'Rolling back transaction',
'No data, all is fine.', 'Data before commit:',
'4 | Geert', '5 | Jan', '6 | Michel', 'Data after commit:',
'4 | Geert', '5 | Jan', '6 | Michel']
self.assertEqual(output,exp,'Output was not correct')
def test_unicode(self):
"""examples/unicode.py"""
try:
import examples.unicode as example
except Exception as e:
self.fail(e)
output = self._exec_main(example)
exp = ['Unicode string: ¿Habla español?',
'Unicode string coming from db: ¿Habla español?']
self.assertEqual(output,exp)#,'Output was not correct')
def test_warnings(self):
"""examples/warnings.py"""
try:
import examples.warnings as example
except Exception as e:
self.fail(e)
output = self._exec_main(example)
exp = ["Executing 'SELECT 'abc'+1'",
"1292: Truncated incorrect DOUBLE value: 'abc'"]
self.assertEqual(output,exp,'Output was not correct')
example.STMT = "SELECT 'abc'"
self.assertRaises(Exception, example.main, self.getMySQLConfig())
def test_multi_resultsets(self):
"""examples/multi_resultsets.py"""
try:
import examples.multi_resultsets as example
except Exception as e:
self.fail(e)
output = self._exec_main(example)
exp = ['Inserted 1 row', 'Number of rows: 1', 'Inserted 2 rows',
'Names in table: <NAME>']
self.assertEqual(output,exp,'Output was not correct')
def test_microseconds(self):
"""examples/microseconds.py"""
try:
import examples.microseconds as example
except Exception as e:
self.fail(e)
output = self._exec_main(example)
if self.cnx.get_server_version() < (5,6,4):
exp = "does not support fractional precision for timestamps."
self.assertTrue(output[0].endswith(exp))
else:
exp = [
' 1 | 1 | 0:00:47.510000',
' 1 | 2 | 0:00:47.020000',
' 1 | 3 | 0:00:47.650000',
' 1 | 4 | 0:00:46.060000',
]
self.assertEqual(output, exp, 'Output was not correct')
|
import sys
import spotipy
import yaml
import spotipy.util as util
from pprint import pprint
import json
import argparse
import matplotlib.pyplot as plt
import numpy as np
def load_config():
global user_config
stream = open('config.yaml')
user_config = yaml.load(stream, Loader=yaml.FullLoader)
def get_playlist_info(username, playlist_uri):
playlist_id = uri.split(':')[2]
results = sp.user_playlist(username, playlist_id)
playlist_name = results['name']
return playlist_name, results
def get_features_for_playlist(plists, username, uri, label):
playlist_name, results = get_playlist_info(username, uri)
plists[playlist_name] = {}
plists[playlist_name]['name'] = []
plists[playlist_name]['label'] = label
plists[playlist_name]['track uri'] = []
plists[playlist_name]['acousticness'] = []
plists[playlist_name]['danceability'] = []
plists[playlist_name]['energy'] = []
plists[playlist_name]['instrumentalness'] = []
plists[playlist_name]['liveness'] = []
plists[playlist_name]['loudness'] = []
plists[playlist_name]['speechiness'] = []
plists[playlist_name]['tempo'] = []
plists[playlist_name]['valence'] = []
plists[playlist_name]['popularity'] = []
for track in results['tracks']['items']:
# print(json.dumps(track, indent=4)) # DEBUG STATEMENT
# save metadata stuff
name = track['track']['name']
track_uri = track['track']['uri']
plists[playlist_name]['name'].append(name)
plists[playlist_name]['track uri'].append(track_uri)
# extract features
features = sp.audio_features(track_uri)
plists[playlist_name]['acousticness'].append(features[0]['acousticness'])
plists[playlist_name]['danceability'].append(features[0]['danceability'])
plists[playlist_name]['energy'].append(features[0]['energy'])
plists[playlist_name]['instrumentalness'].append(features[0]['instrumentalness'])
plists[playlist_name]['liveness'].append(features[0]['liveness'])
plists[playlist_name]['loudness'].append(features[0]['loudness'])
plists[playlist_name]['speechiness'].append(features[0]['speechiness'])
plists[playlist_name]['tempo'].append(features[0]['tempo'])
plists[playlist_name]['valence'].append(features[0]['valence'])
return plists
def print_audio_feature_stats(plists):
"""manually inspect all of the values to determine whether the median or mean is a better metric to plot"""
for playlist in plists:
print("––––––––––––––––––––––––––––––––––––––––––––––––––––––––––––")
print(playlist)
for feature in plists[playlist]:
if feature != 'name' and feature != 'track uri':
print(feature.upper(), "| median:", np.median(plists[playlist][feature]), "| mean:", np.mean(plists[playlist][feature]))
# Helper function to plot each playlist on the radar chart.
def add_to_radar(ax, angles, pdict, color):
values = [np.median(pdict['acousticness']), np.median(pdict['danceability']), np.median(pdict['energy']),
np.median(pdict['valence']), np.mean(pdict['instrumentalness']), np.median(pdict['tempo']),
np.median(pdict['speechiness'])]
# tempo values typically range from 50-220, so I divided by 220 to get a number between 0 and 1
values[-2] = values[-2]/220
# speechiness values values are highly concentrated between 0 and 0.25-ish, so I multiplied by 4. Adjust this if needed
values[-1] = values[-1]*4
values += values[:1]
ax.plot(angles, values, color=color, linewidth=1, label=pdict['label'])
ax.fill(angles, values, color=color, alpha=0.25)
return ax
def make_radar_graph(plists):
# print_audio_feature_stats(plists)
fig, ax = plt.subplots(figsize=(6, 6), subplot_kw=dict(polar=True))
labels = ['acousticness', 'danceability', 'energy', 'valence', 'instrumentalness', 'tempo', 'speechiness']
num_vars = len(labels)
colors = ['red', 'green', 'blue']
# Split the circle into even parts and save the angles so we know where to put each axis.
angles = np.linspace(0, 2 * np.pi, num_vars, endpoint=False).tolist()
angles += angles[:1]
# Add each additional playlist to the chart.
for i, pname in enumerate(plists.keys()):
print(i, pname)
ax = add_to_radar(ax, angles, plists[pname], colors[i])
# polar coordinates math stuff
ax.set_theta_offset(np.pi / 2)
ax.set_theta_direction(-1)
# Draw axis lines for each angle and label.
ax.set_thetagrids(np.degrees(angles), labels)
# Go through labels and adjust alignment based on where it is in the circle.
for label, angle in zip(ax.get_xticklabels(), angles):
if angle in (0, np.pi):
label.set_horizontalalignment('center')
elif 0 < angle < np.pi:
label.set_horizontalalignment('left')
else:
label.set_horizontalalignment('right')
# Set position of y-labels (0-100) to be in the middle of the first two axes.
ax.set_ylim(0, 1)
ax.set_rlabel_position(180 / num_vars)
# Add some custom styling.
ax.tick_params(colors='#222222') # color of tick labels
ax.tick_params(axis='y', labelsize=8) # y-axis labels
ax.grid(color='#AAAAAA') # color of circular gridlines
ax.spines['polar'].set_color('#222222') # color of outermost gridline (spine)
ax.set_facecolor('#FAFAFA') # background color inside the circle itself
#Lastly, give the chart a title and a legend
ax.set_title('Playlist Comparison', y=1.08)
ax.legend(loc='best', bbox_to_anchor=(1.1, 1.1))
fig.savefig('playlist_comp.png')
if __name__ == '__main__':
global sp
global user_config
load_config()
token = util.prompt_for_user_token(user_config['username'], scope='playlist-read-private', client_id=user_config['client_id'], client_secret=user_config['client_secret'], redirect_uri=user_config['redirect_uri'])
if token:
sp = spotipy.Spotify(auth=token)
uris = []
labels = []
for i in range(2):
uris.append(input("URI " + str(i+1) + ": "))
labels.append(input("Label " + str(i+1) + ": "))
plists = {}
for i, uri in enumerate(uris):
plists = get_features_for_playlist(plists, user_config['username'], uri, labels[i])
make_radar_graph(plists)
else:
print ("Can't get token for", user_config['username']) |
<filename>app/main.py
import base64
from pathlib import Path
import dash
import dash_core_components as dcc
import dash_html_components as html
from dash.dependencies import Input, Output
import pandas as pd
import dash_bootstrap_components as dbc
from dotenv import load_dotenv
from apps.dataset_page import generate_dataset_page
from apps.utils import generate_kpi_card, DATASET_COLUMNS, generate_badge, MLJAR_INFO_DICT
BASE_PATH = Path(__file__).parent.resolve()
DATA_PATH = BASE_PATH.joinpath("assets/datasets").resolve()
from apps.banner import get_banner
load_dotenv(verbose=True)
from apps.utils import add_nb_features_category, add_nb_lines_category, NB_FEATURES_CATEGORIES, NB_LINES_CATEGORIES, \
get_dataset_info
app = dash.Dash(
__name__,
external_stylesheets=[dbc.themes.BOOTSTRAP],
suppress_callback_exceptions=True,
url_base_pathname="/dgml/",
meta_tags=[{"name": "viewport", "content": "width=device-width, initial-scale=1"}],
)
app.title = "DGML"
server = app.server
# Read data
encoded_image_validated = base64.b64encode(open(DATA_PATH.parent.joinpath("quality.png"), 'rb').read()).decode()
df = pd.read_csv(DATA_PATH.joinpath("dgml_datasets.csv"))
task_list = df["task"].unique()
nb_features_bins = list(NB_FEATURES_CATEGORIES.keys())
nb_lines_bins = list(NB_LINES_CATEGORIES.keys())
topic_list = df["topic"].unique()
add_nb_features_category(df)
add_nb_lines_category(df)
def check_if_resource(df):
dataset_folders = [Path(paths).stem for paths in DATA_PATH.joinpath(f"resources/").glob('*')]
filtered_df = df[df['dgf_resource_id'].isin(dataset_folders)]
return filtered_df
df = check_if_resource(df)
def description_card():
"""
The section dealing with the description of the website
:return: A Div containing dashboard title & descriptions.
"""
return html.Div(
id="description-card",
children=[
html.H3("Bienvenue sur DGML!"),
html.Div(
id="intro",
children=["Data Gouv pour le Machine Learning (DGML) est le catalogue des jeux de données de",
html.A(" data.gouv.fr", href="https://www.data.gouv.fr",
target="_blank"),
" pour le Machine Learning. ",
html.Br(),
"Cliquez sur un jeu de données pour voir: ses statistiques, les résultats ",
"de l'entraînement et test automatique d'algorithmes de Machine Learning sur les données, ainsi que des ",
"exemples de code et des réutilisations qui vont vous guider dans la mise en oeuvre de votre modèle de Machine Learning avec ces données.",
html.Br(), "DGML a été développé par le ",
html.A("Lab IA d'Etalab: ",
href="https://www.etalab.gouv.fr/datasciences-et-intelligence-artificielle",
target="_blank"),
"visitez notre Github pour en savoir plus sur le projet, sur le choix des jeux de données, pour mieux comprendre les résultats ou nous contacter."],
),
],
)
def generate_control_card():
"""
The section that presents the control filtering options
:return: A Div containing controls for graphs.
"""
return html.Div(
id="control-card",
children=[
html.P("Tâche"),
dcc.Checklist(
id="task-select",
options=[{"label": f" {i}", "value": i} for i in task_list],
value=task_list,
),
html.Br(),
html.P("Nombre de colonnes"),
dcc.Checklist(
id="features-select",
options=[{"label": f" {i}", "value": i} for i in nb_features_bins],
value=nb_features_bins,
),
html.Br(),
html.P("Nombre de lignes"),
dcc.Checklist(
id="lines-select",
options=[{"label": f" {i}", "value": i} for i in nb_lines_bins],
value=nb_lines_bins,
),
html.Br(),
html.P("Validation"),
dcc.Checklist(
id="valid-select",
options=[{'label': f" {l}", "value": l} for l in ["Sélectionné", "Automatique"]],
value=["Sélectionné", "Automatique"],
),
html.Br(),
html.P("Thème"),
dcc.Dropdown(
id="topic-select",
options=[{"label": f" {i}", "value": i} for i in topic_list],
multi=True,
value=topic_list,
clearable=False
),
html.Br(),
html.P("Filtrer par:"),
dcc.Dropdown(
id="sort-by",
options=[{"label": f" {i}", "value": i} for i in DATASET_COLUMNS],
value="Validé",
clearable=False
),
html.Br(),
dcc.RadioItems(
id="sort-by-order",
options=[{"label": i, "value": i} for i in ["Ascendant", "Descendant"]],
value="Ascendant",
),
html.Br(),
html.Div(
id="reset-btn-outer",
children=html.Button(id="reset-btn", children="Reset", n_clicks=0, hidden=True),
),
],
style={"width": "100%"}
)
app_layout = html.Div(
id="app-container",
children=[
# Banner
get_banner(),
html.Div(id='app-page-content',
children=[
# Left column
html.Div(
id="left-column",
className="four columns",
children=[description_card(), generate_control_card()]
),
# Right column
html.Div(
id="right-column",
className="seven columns",
children=[
html.Div(id="dataset-card-div")
],
)
])
],
)
url_bar_and_content_div = html.Div([
dcc.Location(id='url', refresh=False),
html.Div(id='page-content')
])
app.layout = url_bar_and_content_div
# app.validation_layout = html.Div([
# url_bar_and_content_div,
# generate_dataset_page("test")
# ])
def generate_dataset_block(tasks, features, lines, valid, topics, sort_by, sort_order, reset_click):
curated_dict = {"Sélectionné": True, "Automatique": False}
chosen_tasks_df = df[df.task.isin(tasks)]
chosen_features_df = chosen_tasks_df[chosen_tasks_df.nb_features_cat.isin(features)]
chosen_lines_df = chosen_features_df[chosen_features_df.nb_lines_cat.isin(lines)]
chosen_topics_df = chosen_lines_df[chosen_lines_df.topic.isin(topics)]
chosen_validation = chosen_topics_df[chosen_topics_df["is_validated"].isin([curated_dict[v] for v in valid])]
chosen_sort_by_df = chosen_validation.sort_values(by=DATASET_COLUMNS[sort_by],
ascending=True if sort_order == 'Ascendant' and sort_by != "Validé"
else False)
cards_list = []
for index, dataset_row in chosen_sort_by_df.iterrows():
dataset_dict = get_dataset_info(dataset_row)
main_dataset_card = html.Div(dbc.Card(
[
dbc.CardBody(
[
html.H4(
[
dcc.Link(f"{dataset_dict['title']}", href=f"{app.config['url_base_pathname']}"
f"{dataset_dict['dgf_resource_id']}"),
html.Img(id="validated-img",
src="data:image/png;base64,{}".format(encoded_image_validated),
style={'height': '3%', 'width': '3%', "float": "right"},
hidden=not dataset_dict["is_validated"]),
dbc.Tooltip("Ce jeu de données a été sélectionné et analysé manuellement.",
target="validated-img",
style={'font-size': 13}
)
],
className="card-title"),
dbc.CardDeck([
# profiling
generate_kpi_card("Tâche proposée", f"{dataset_dict['task']}"),
generate_kpi_card("Thème", f"{dataset_dict['topic']}"),
generate_kpi_card("Colonnes", dataset_dict['nb_features']),
generate_kpi_card("Lignes", dataset_dict['nb_lines']),
]),
]
),
],
),
style={"marginTop": "20px"}
)
cards_list.append(main_dataset_card)
return cards_list
# Index callbacks
@app.callback(Output('page-content', 'children'),
Input('url', 'pathname'))
def display_page(pathname):
if pathname == app.config['url_base_pathname']:
return app_layout
else:
return generate_dataset_page(pathname, df)
#
@app.callback([Output('mljar-div', 'children'),
Output('mljar-link-badge', 'children')],
[Input('target-var-model-select', 'value'),
Input('url', 'pathname')])
def update_automl_model(target_var, pathname):
if pathname == app.config['url_base_pathname']:
return None, None
dataset_id = pathname.split("/")[-1]
if dataset_id not in MLJAR_INFO_DICT or target_var not in MLJAR_INFO_DICT[dataset_id]:
return html.P("On n'a pas des modèles AutoML pour ce dataset"), None
models_table, mljlar_profile_url = MLJAR_INFO_DICT[dataset_id][target_var]
mljar_profile_badge = generate_badge("Full Mljar Profile", url=mljlar_profile_url.as_posix(),
background_color="#6d92ad",
new_tab=True)
return models_table, mljar_profile_badge
@app.callback(
Output("dataset-card-div", "children"),
[
Input("task-select", "value"),
Input("features-select", "value"),
Input("lines-select", "value"),
Input("valid-select", "value"),
Input("topic-select", "value"),
Input("sort-by", "value"),
Input("sort-by-order", "value"),
Input("reset-btn", "n_clicks"),
],
)
def update_dataset_block(task, feature, line, valid, topic, sort_by, sort_order, reset_click):
reset = False
# Find which one has been triggered
ctx = dash.callback_context
if ctx.triggered:
prop_id = ctx.triggered[0]["prop_id"].split(".")[0]
if prop_id == "reset-btn":
reset = True
# Return to original hm(no colored annotation) by resetting
return generate_dataset_block(task, feature, line, valid, topic, sort_by, sort_order, reset_click)
# Run the server
if __name__ == "__main__":
app.run_server(debug=True, port=8050)
|
# Copyright (c) OpenMMLab. All rights reserved.
import itertools
import logging
import os.path as osp
from collections import OrderedDict
from typing import Dict, List, Optional, Sequence, Union
import mmcv
import numpy as np
from mmcv.utils import print_log
from mmdet.datasets.api_wrappers import COCO, COCOeval
from mmdet.datasets.builder import DATASETS
from mmdet.datasets.coco import CocoDataset
from terminaltables import AsciiTable
from .base import BaseFewShotDataset
# pre-defined classes split for few shot setting
COCO_SPLIT = dict(
ALL_CLASSES=('person', 'bicycle', 'car', 'motorcycle', 'airplane', 'bus',
'train', 'truck', 'boat', 'traffic light', 'fire hydrant',
'stop sign', 'parking meter', 'bench', 'bird', 'cat', 'dog',
'horse', 'sheep', 'cow', 'elephant', 'bear', 'zebra',
'giraffe', 'backpack', 'umbrella', 'handbag', 'tie',
'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
'kite', 'baseball bat', 'baseball glove', 'skateboard',
'surfboard', 'tennis racket', 'bottle', 'wine glass', 'cup',
'fork', 'knife', 'spoon', 'bowl', 'banana', 'apple',
'sandwich', 'orange', 'broccoli', 'carrot', 'hot dog',
'pizza', 'donut', 'cake', 'chair', 'couch', 'potted plant',
'bed', 'dining table', 'toilet', 'tv', 'laptop', 'mouse',
'remote', 'keyboard', 'cell phone', 'microwave', 'oven',
'toaster', 'sink', 'refrigerator', 'book', 'clock', 'vase',
'scissors', 'teddy bear', 'hair drier', 'toothbrush'),
NOVEL_CLASSES=('person', 'bicycle', 'car', 'motorcycle', 'airplane', 'bus',
'train', 'boat', 'bird', 'cat', 'dog', 'horse', 'sheep',
'cow', 'bottle', 'chair', 'couch', 'potted plant',
'dining table', 'tv'),
BASE_CLASSES=('truck', 'traffic light', 'fire hydrant', 'stop sign',
'parking meter', 'bench', 'elephant', 'bear', 'zebra',
'giraffe', 'backpack', 'umbrella', 'handbag', 'tie',
'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
'kite', 'baseball bat', 'baseball glove', 'skateboard',
'surfboard', 'tennis racket', 'wine glass', 'cup', 'fork',
'knife', 'spoon', 'bowl', 'banana', 'apple', 'sandwich',
'orange', 'broccoli', 'carrot', 'hot dog', 'pizza', 'donut',
'cake', 'bed', 'toilet', 'laptop', 'mouse', 'remote',
'keyboard', 'cell phone', 'microwave', 'oven', 'toaster',
'sink', 'refrigerator', 'book', 'clock', 'vase', 'scissors',
'teddy bear', 'hair drier', 'toothbrush'))
@DATASETS.register_module()
class FewShotCocoDataset(BaseFewShotDataset, CocoDataset):
"""COCO dataset for few shot detection.
Args:
classes (str | Sequence[str] | None): Classes for model training and
provide fixed label for each class. When classes is string,
it will load pre-defined classes in :obj:`FewShotCocoDataset`.
For example: 'BASE_CLASSES', 'NOVEL_CLASSES` or `ALL_CLASSES`.
num_novel_shots (int | None): Max number of instances used for each
novel class. If is None, all annotation will be used.
Default: None.
num_base_shots (int | None): Max number of instances used for each base
class. If is None, all annotation will be used. Default: None.
ann_shot_filter (dict | None): Used to specify the class and the
corresponding maximum number of instances when loading
the annotation file. For example: {'dog': 10, 'person': 5}.
If set it as None, `ann_shot_filter` will be
created according to `num_novel_shots` and `num_base_shots`.
min_bbox_area (int | float | None): Filter images with bbox whose
area smaller `min_bbox_area`. If set to None, skip
this filter. Default: None.
dataset_name (str | None): Name of dataset to display. For example:
'train dataset' or 'query dataset'. Default: None.
test_mode (bool): If set True, annotation will not be loaded.
Default: False.
"""
def __init__(self,
classes: Optional[Union[str, Sequence[str]]] = None,
num_novel_shots: Optional[int] = None,
num_base_shots: Optional[int] = None,
ann_shot_filter: Optional[Dict[str, int]] = None,
min_bbox_area: Optional[Union[int, float]] = None,
dataset_name: Optional[str] = None,
test_mode: bool = False,
**kwargs) -> None:
if dataset_name is None:
self.dataset_name = 'Test dataset' \
if test_mode else 'Train dataset'
else:
self.dataset_name = dataset_name
self.SPLIT = COCO_SPLIT
assert classes is not None, f'{self.dataset_name}: classes in ' \
f'`FewShotCocoDataset` can not be None.'
# `ann_shot_filter` will be used to filter out excess annotations
# for few shot setting. It can be configured manually or generated
# by the `num_novel_shots` and `num_base_shots`
self.num_novel_shots = num_novel_shots
self.num_base_shots = num_base_shots
self.min_bbox_area = min_bbox_area
self.CLASSES = self.get_classes(classes)
if ann_shot_filter is None:
if num_novel_shots is not None or num_base_shots is not None:
ann_shot_filter = self._create_ann_shot_filter()
else:
assert num_novel_shots is None and num_base_shots is None, \
f'{self.dataset_name}: can not config ann_shot_filter and ' \
f'num_novel_shots/num_base_shots at the same time.'
# these values would be set in `self.load_annotations_coco`
self.cat_ids = []
self.cat2label = {}
self.coco = None
self.img_ids = None
super().__init__(
classes=None,
ann_shot_filter=ann_shot_filter,
dataset_name=dataset_name,
test_mode=test_mode,
**kwargs)
def get_classes(self, classes: Union[str, Sequence[str]]) -> List[str]:
"""Get class names.
It supports to load pre-defined classes splits.
The pre-defined classes splits are:
['ALL_CLASSES', 'NOVEL_CLASSES', 'BASE_CLASSES']
Args:
classes (str | Sequence[str]): Classes for model training and
provide fixed label for each class. When classes is string,
it will load pre-defined classes in `FewShotCocoDataset`.
For example: 'NOVEL_CLASSES'.
Returns:
list[str]: list of class names.
"""
# configure few shot classes setting
if isinstance(classes, str):
assert classes in self.SPLIT.keys(
), f'{self.dataset_name} : not a pre-defined classes or split ' \
f'in COCO_SPLIT.'
class_names = self.SPLIT[classes]
if 'BASE_CLASSES' in classes:
assert self.num_novel_shots is None, \
f'{self.dataset_name}: BASE_CLASSES do not have ' \
f'novel instances.'
elif 'NOVEL_CLASSES' in classes:
assert self.num_base_shots is None, \
f'{self.dataset_name}: NOVEL_CLASSES do not have ' \
f'base instances.'
elif isinstance(classes, (tuple, list)):
class_names = classes
else:
raise ValueError(f'Unsupported type {type(classes)} of classes.')
return class_names
def _create_ann_shot_filter(self) -> Dict:
"""Generate `ann_shot_filter` for novel and base classes.
Returns:
dict[str, int]: The number of shots to keep for each class.
"""
ann_shot_filter = {}
# generate annotation filter for novel classes
if self.num_novel_shots is not None:
for class_name in self.SPLIT['NOVEL_CLASSES']:
ann_shot_filter[class_name] = self.num_novel_shots
# generate annotation filter for base classes
if self.num_base_shots is not None:
for class_name in self.SPLIT['BASE_CLASSES']:
ann_shot_filter[class_name] = self.num_base_shots
return ann_shot_filter
def load_annotations(self, ann_cfg: List[Dict]) -> List[Dict]:
"""Support to Load annotation from two type of ann_cfg.
- type of 'ann_file': COCO-style annotation file.
- type of 'saved_dataset': Saved COCO dataset json.
Args:
ann_cfg (list[dict]): Config of annotations.
Returns:
list[dict]: Annotation infos.
"""
data_infos = []
for ann_cfg_ in ann_cfg:
if ann_cfg_['type'] == 'saved_dataset':
data_infos += self.load_annotations_saved(ann_cfg_['ann_file'])
elif ann_cfg_['type'] == 'ann_file':
data_infos += self.load_annotations_coco(ann_cfg_['ann_file'])
else:
raise ValueError(f'not support annotation type '
f'{ann_cfg_["type"]} in ann_cfg.')
return data_infos
def load_annotations_coco(self, ann_file: str) -> List[Dict]:
"""Load annotation from COCO style annotation file.
Args:
ann_file (str): Path of annotation file.
Returns:
list[dict]: Annotation info from COCO api.
"""
self.coco = COCO(ann_file)
# to keep the label order equal to the order in CLASSES
if len(self.cat_ids) == 0:
for i, class_name in enumerate(self.CLASSES):
cat_id = self.coco.get_cat_ids(cat_names=[class_name])[0]
self.cat_ids.append(cat_id)
self.cat2label[cat_id] = i
else:
# check categories id consistency between different files
for i, class_name in enumerate(self.CLASSES):
cat_id = self.coco.get_cat_ids(cat_names=[class_name])[0]
assert self.cat2label[cat_id] == i, \
'please make sure all the json files use same ' \
'categories id for same class'
self.img_ids = self.coco.get_img_ids()
data_infos = []
total_ann_ids = []
for i in self.img_ids:
info = self.coco.load_imgs([i])[0]
info['filename'] = info['file_name']
info['ann'] = self._get_ann_info(info)
# to support different version of coco since some annotation file
# contain images from train2014 and val2014 at the same time
if 'train2014' in info['filename']:
info['filename'] = 'train2014/' + info['filename']
elif 'val2014' in info['filename']:
info['filename'] = 'val2014/' + info['filename']
elif 'instances_val2017' in ann_file:
info['filename'] = 'val2017/' + info['filename']
elif 'instances_train2017' in ann_file:
info['filename'] = 'train2017/' + info['filename']
data_infos.append(info)
ann_ids = self.coco.get_ann_ids(img_ids=[i])
total_ann_ids.extend(ann_ids)
assert len(set(total_ann_ids)) == len(
total_ann_ids
), f'{self.dataset_name}: Annotation ids in {ann_file} are not unique!'
return data_infos
def _get_ann_info(self, data_info: Dict) -> Dict:
"""Get COCO annotation by index.
Args:
data_info(dict): Data info.
Returns:
dict: Annotation info of specified index.
"""
img_id = data_info['id']
ann_ids = self.coco.get_ann_ids(img_ids=[img_id])
ann_info = self.coco.load_anns(ann_ids)
return self._parse_ann_info(data_info, ann_info)
def get_cat_ids(self, idx: int) -> List[int]:
"""Get category ids by index.
Overwrite the function in CocoDataset.
Args:
idx (int): Index of data.
Returns:
list[int]: All categories in the image of specified index.
"""
return self.data_infos[idx]['ann']['labels'].astype(np.int64).tolist()
def _filter_imgs(self,
min_size: int = 32,
min_bbox_area: Optional[int] = None) -> List[int]:
"""Filter images that do not meet the requirements.
Args:
min_size (int): Filter images with length or width
smaller than `min_size`. Default: 32.
min_bbox_area (int | None): Filter images with bbox whose
area smaller `min_bbox_area`. If set to None, skip
this filter. Default: None.
Returns:
list[int]: valid indices of data_infos.
"""
valid_inds = []
valid_img_ids = []
if min_bbox_area is None:
min_bbox_area = self.min_bbox_area
for i, img_info in enumerate(self.data_infos):
# filter empty image
if self.filter_empty_gt and img_info['ann']['labels'].size == 0:
continue
# filter images smaller than `min_size`
if min(img_info['width'], img_info['height']) < min_size:
continue
# filter image with bbox smaller than min_bbox_area
# it is usually used in Attention RPN
if min_bbox_area is not None:
skip_flag = False
for bbox in img_info['ann']['bboxes']:
bbox_area = (bbox[2] - bbox[0]) * (bbox[3] - bbox[1])
if bbox_area < min_bbox_area:
skip_flag = True
if skip_flag:
continue
valid_inds.append(i)
valid_img_ids.append(img_info['id'])
# update coco img_ids
self.img_ids = valid_img_ids
return valid_inds
def evaluate(self,
results: List[Sequence],
metric: Union[str, List[str]] = 'bbox',
logger: Optional[object] = None,
jsonfile_prefix: Optional[str] = None,
classwise: bool = False,
proposal_nums: Sequence[int] = (100, 300, 1000),
iou_thrs: Optional[Union[float, Sequence[float]]] = None,
metric_items: Optional[Union[List[str], str]] = None,
class_splits: Optional[List[str]] = None) -> Dict:
"""Evaluation in COCO protocol and summary results of different splits
of classes.
Args:
results (list[list | tuple]): Testing results of the dataset.
metric (str | list[str]): Metrics to be evaluated. Options are
'bbox', 'proposal', 'proposal_fast'. Default: 'bbox'
logger (logging.Logger | None): Logger used for printing
related information during evaluation. Default: None.
jsonfile_prefix (str | None): The prefix of json files. It includes
the file path and the prefix of filename, e.g., "a/b/prefix".
If not specified, a temp file will be created. Default: None.
classwise (bool): Whether to evaluating the AP for each class.
proposal_nums (Sequence[int]): Proposal number used for evaluating
recalls, such as recall@100, recall@1000.
Default: (100, 300, 1000).
iou_thrs (Sequence[float] | float | None): IoU threshold used for
evaluating recalls/mAPs. If set to a list, the average of all
IoUs will also be computed. If not specified, [0.50, 0.55,
0.60, 0.65, 0.70, 0.75, 0.80, 0.85, 0.90, 0.95] will be used.
Default: None.
metric_items (list[str] | str | None): Metric items that will
be returned. If not specified, ``['AR@100', 'AR@300',
'AR@1000', 'AR_s@1000', 'AR_m@1000', 'AR_l@1000' ]`` will be
used when ``metric=='proposal'``, ``['mAP', 'mAP_50', 'mAP_75',
'mAP_s', 'mAP_m', 'mAP_l']`` will be used when
``metric=='bbox'``.
class_splits: (list[str] | None): Calculate metric of classes split
in COCO_SPLIT. For example: ['BASE_CLASSES', 'NOVEL_CLASSES'].
Default: None.
Returns:
dict[str, float]: COCO style evaluation metric.
"""
if class_splits is not None:
for k in class_splits:
assert k in self.SPLIT.keys(), 'please define classes split.'
metrics = metric if isinstance(metric, list) else [metric]
allowed_metrics = ['bbox', 'proposal', 'proposal_fast']
for metric in metrics:
if metric not in allowed_metrics:
raise KeyError(f'metric {metric} is not supported')
if iou_thrs is None:
iou_thrs = np.linspace(
.5, 0.95, int(np.round((0.95 - .5) / .05)) + 1, endpoint=True)
if metric_items is not None:
if not isinstance(metric_items, list):
metric_items = [metric_items]
result_files, tmp_dir = self.format_results(results, jsonfile_prefix)
eval_results = OrderedDict()
cocoGt = self.coco
for metric in metrics:
msg = f'Evaluating {metric}...'
if logger is None:
msg = '\n' + msg
print_log(msg, logger=logger)
iou_type = 'bbox' if metric == 'proposal' else metric
if metric not in result_files:
raise KeyError(f'{metric} is not in results')
try:
predictions = mmcv.load(result_files[metric])
cocoDt = cocoGt.loadRes(predictions)
except IndexError:
print_log(
'The testing results of the whole dataset is empty.',
logger=logger,
level=logging.ERROR)
break
# eval each class splits
if class_splits is not None:
class_splits = {k: COCO_SPLIT[k] for k in class_splits}
for split_name in class_splits.keys():
split_cat_ids = [
self.cat_ids[i] for i in range(len(self.CLASSES))
if self.CLASSES[i] in class_splits[split_name]
]
self._evaluate_by_class_split(
cocoGt,
cocoDt,
iou_type,
proposal_nums,
iou_thrs,
split_cat_ids,
metric,
metric_items,
eval_results,
False,
logger,
split_name=split_name + ' ')
# eval all classes
self._evaluate_by_class_split(cocoGt, cocoDt, iou_type,
proposal_nums, iou_thrs,
self.cat_ids, metric, metric_items,
eval_results, classwise, logger)
if tmp_dir is not None:
tmp_dir.cleanup()
return eval_results
def _evaluate_by_class_split(self,
cocoGt: object,
cocoDt: object,
iou_type: str,
proposal_nums: Sequence[int],
iou_thrs: Union[float, Sequence[float]],
cat_ids: List[int],
metric: str,
metric_items: Union[str, List[str]],
eval_results: Dict,
classwise: bool,
logger: object,
split_name: str = '') -> Dict:
"""Evaluation a split of classes in COCO protocol.
Args:
cocoGt (object): coco object with ground truth annotations.
cocoDt (object): coco object with detection results.
iou_type (str): Type of IOU.
proposal_nums (Sequence[int]): Number of proposals.
iou_thrs (float | Sequence[float]): Thresholds of IoU.
cat_ids (list[int]): Class ids of classes to be evaluated.
metric (str): Metrics to be evaluated.
metric_items (str | list[str]): Metric items that will
be returned. If not specified, ``['AR@100', 'AR@300',
'AR@1000', 'AR_s@1000', 'AR_m@1000', 'AR_l@1000' ]`` will be
used when ``metric=='proposal'``, ``['mAP', 'mAP_50', 'mAP_75',
'mAP_s', 'mAP_m', 'mAP_l']`` will be used when
``metric=='bbox'``.
eval_results (dict[str, float]): COCO style evaluation metric.
classwise (bool): Whether to evaluating the AP for each class.
split_name (str): Name of split. Default:''.
Returns:
dict[str, float]: COCO style evaluation metric.
"""
cocoEval = COCOeval(cocoGt, cocoDt, iou_type)
cocoEval.params.imgIds = self.img_ids
cocoEval.params.maxDets = list(proposal_nums)
cocoEval.params.iouThrs = iou_thrs
cocoEval.params.catIds = cat_ids
# mapping of cocoEval.stats
coco_metric_names = {
'mAP': 0,
'mAP_50': 1,
'mAP_75': 2,
'mAP_s': 3,
'mAP_m': 4,
'mAP_l': 5,
'AR@100': 6,
'AR@300': 7,
'AR@1000': 8,
'AR_s@1000': 9,
'AR_m@1000': 10,
'AR_l@1000': 11
}
if metric_items is not None:
for metric_item in metric_items:
if metric_item not in coco_metric_names:
raise KeyError(
f'metric item {metric_item} is not supported')
if split_name is not None:
print_log(f'\n evaluation of {split_name} class', logger=logger)
if metric == 'proposal':
cocoEval.params.useCats = 0
cocoEval.evaluate()
cocoEval.accumulate()
cocoEval.summarize()
if metric_items is None:
metric_items = [
'AR@100', 'AR@300', 'AR@1000', 'AR_s@1000', 'AR_m@1000',
'AR_l@1000'
]
for item in metric_items:
val = float(f'{cocoEval.stats[coco_metric_names[item]]:.3f}')
eval_results[split_name + item] = val
else:
cocoEval.evaluate()
cocoEval.accumulate()
cocoEval.summarize()
if classwise: # Compute per-category AP
# Compute per-category AP
# from https://github.com/facebookresearch/detectron2/
precisions = cocoEval.eval['precision']
# precision: (iou, recall, cls, area range, max dets)
assert len(self.cat_ids) == precisions.shape[2], \
f'{self.cat_ids},{precisions.shape}'
results_per_category = []
for idx, catId in enumerate(self.cat_ids):
# area range index 0: all area ranges
# max dets index -1: typically 100 per image
nm = self.coco.loadCats(catId)[0]
precision = precisions[:, :, idx, 0, -1]
precision = precision[precision > -1]
if precision.size:
ap = np.mean(precision)
else:
ap = float('nan')
results_per_category.append(
(f'{nm["name"]}', f'{float(ap):0.3f}'))
num_columns = min(6, len(results_per_category) * 2)
results_flatten = list(itertools.chain(*results_per_category))
headers = [split_name + 'category', split_name + 'AP'] * (
num_columns // 2)
results_2d = itertools.zip_longest(*[
results_flatten[i::num_columns] for i in range(num_columns)
])
table_data = [headers]
table_data += [result for result in results_2d]
table = AsciiTable(table_data)
print_log('\n' + table.table, logger=logger)
if metric_items is None:
metric_items = [
'mAP', 'mAP_50', 'mAP_75', 'mAP_s', 'mAP_m', 'mAP_l'
]
for metric_item in metric_items:
key = f'{metric}_{metric_item}'
val = float(
f'{cocoEval.stats[coco_metric_names[metric_item]]:.3f}')
eval_results[split_name + key] = val
ap = cocoEval.stats[:6]
eval_results[split_name + f'{metric}_mAP_copypaste'] = (
f'{ap[0]:.3f} {ap[1]:.3f} {ap[2]:.3f} {ap[3]:.3f} '
f'{ap[4]:.3f} {ap[5]:.3f}')
return eval_results
@DATASETS.register_module()
class FewShotCocoCopyDataset(FewShotCocoDataset):
"""Copy other COCO few shot datasets' `data_infos` directly.
This dataset is mainly used for model initialization in some meta-learning
detectors. In their cases, the support data are randomly sampled
during training phase and they also need to be used in model
initialization before evaluation. To copy the random sampling results,
this dataset supports to load `data_infos` of other datasets via `ann_cfg`
Args:
ann_cfg (list[dict] | dict): contain `data_infos` from other
dataset. Example: [dict(data_infos=FewShotCocoDataset.data_infos)]
"""
def __init__(self, ann_cfg: Union[List[Dict], Dict], **kwargs) -> None:
super().__init__(ann_cfg=ann_cfg, **kwargs)
def ann_cfg_parser(self, ann_cfg: Union[List[Dict], Dict]) -> List[Dict]:
"""Parse annotation config from a copy of other dataset's `data_infos`.
Args:
ann_cfg (list[dict] | dict): contain `data_infos` from other
dataset. Example:
[dict(data_infos=FewShotCocoDataset.data_infos)]
Returns:
list[dict]: Annotation information.
"""
data_infos = []
if isinstance(ann_cfg, dict):
assert ann_cfg.get('data_infos', None) is not None, \
'ann_cfg of FewShotCocoCopyDataset require data_infos.'
# directly copy data_info
data_infos = ann_cfg['data_infos']
elif isinstance(ann_cfg, list):
for ann_cfg_ in ann_cfg:
assert ann_cfg_.get('data_infos', None) is not None, \
'ann_cfg of FewShotCocoCopyDataset require data_infos.'
# directly copy data_info
data_infos += ann_cfg_['data_infos']
return data_infos
@DATASETS.register_module()
class FewShotCocoDefaultDataset(FewShotCocoDataset):
"""FewShot COCO Dataset with some pre-defined annotation paths.
:obj:`FewShotCocoDefaultDataset` provides pre-defined annotation files
to ensure the reproducibility. The pre-defined annotation files provide
fixed training data to avoid random sampling. The usage of `ann_cfg' is
different from :obj:`FewShotCocoDataset`. The `ann_cfg' should contain
two filed: `method` and `setting`.
Args:
ann_cfg (list[dict]): Each dict should contain
`method` and `setting` to get corresponding
annotation from `DEFAULT_ANN_CONFIG`.
For example: [dict(method='TFA', setting='1shot')].
"""
coco_benchmark = {
f'{shot}SHOT': [
dict(
type='ann_file',
ann_file=f'data/few_shot_ann/coco/benchmark_{shot}shot/'
f'full_box_{shot}shot_{class_name}_trainval.json')
for class_name in COCO_SPLIT['ALL_CLASSES']
]
for shot in [10, 30]
}
# pre-defined annotation config for model reproducibility
DEFAULT_ANN_CONFIG = dict(
TFA=coco_benchmark,
FSCE=coco_benchmark,
Attention_RPN={
**coco_benchmark, 'Official_10SHOT': [
dict(
type='ann_file',
ann_file='data/few_shot_ann/coco/attention_rpn_10shot/'
'official_10_shot_from_instances_train2017.json')
]
},
MPSR=coco_benchmark,
MetaRCNN=coco_benchmark,
FSDetView=coco_benchmark)
def __init__(self, ann_cfg: List[Dict], **kwargs) -> None:
super().__init__(ann_cfg=ann_cfg, **kwargs)
def ann_cfg_parser(self, ann_cfg: List[Dict]) -> List[Dict]:
"""Parse pre-defined annotation config to annotation information.
Args:
ann_cfg (list[dict]): Each dict should contain
`method` and `setting` to get corresponding
annotation from `DEFAULT_ANN_CONFIG`.
For example: [dict(method='TFA', setting='1shot')]
Returns:
list[dict]: Annotation information.
"""
new_ann_cfg = []
for ann_cfg_ in ann_cfg:
assert isinstance(ann_cfg_, dict), \
f'{self.dataset_name} : ann_cfg should be list of dict.'
method = ann_cfg_['method']
setting = ann_cfg_['setting']
default_ann_cfg = self.DEFAULT_ANN_CONFIG[method][setting]
ann_root = ann_cfg_.get('ann_root', None)
if ann_root is not None:
for i in range(len(default_ann_cfg)):
default_ann_cfg[i]['ann_file'] = \
osp.join(ann_root, default_ann_cfg[i]['ann_file'])
new_ann_cfg += default_ann_cfg
return super(FewShotCocoDataset, self).ann_cfg_parser(new_ann_cfg)
|
<gh_stars>0
import argparse
import csv
from datetime import datetime, timedelta
import json
import os
from progress.spinner import Spinner
import requests
import time
def lookup_channel_id_by_name(token, channel_name):
r = requests.get("https://slack.com/api/channels.list?token=" +
token)
channel_list_parsed = r.json()["channels"]
for channel in channel_list_parsed:
if channel["name"] == channel_name:
return channel["id"]
return ""
def fetch_from_slack(token, channel, offset):
results = []
newest_timestamp = offset
more_results = True
spinner = Spinner('Fetching history for ' +
channel + ' from ' + str(datetime.fromtimestamp(int(offset))) + ' ')
while more_results == True:
print(str(datetime.fromtimestamp(float(newest_timestamp))))
r = requests.get("https://slack.com/api/channels.history?token=" +
token + "&channel=" + channel + "&count=100&inclusive=true&oldest=" + newest_timestamp)
channel_parsed = r.json()
if not channel_parsed['ok']:
raise ValueError("Error fetching channel history from Slack: ",
channel_parsed["error"])
more_results = channel_parsed["has_more"]
message_data = channel_parsed['messages']
if len(results) == 0:
results = message_data
else:
results = results + message_data
newest_timestamp = message_data[0].get('ts')
time.sleep(2)
spinner.next()
return results
def main():
parser = argparse.ArgumentParser(description='slack2csv')
parser.add_argument('--text', help='text to search for', default='')
parser.add_argument(
'--past_days', help='days to go back', default='1')
requiredNamed = parser.add_argument_group('required named arguments')
requiredNamed.add_argument(
'--token', help='Slack API token', required=True)
requiredNamed.add_argument(
'--channel', help='Slack channel id or name', required=True)
requiredNamed.add_argument(
'--filename', help='CSV filename', required=True)
args = parser.parse_args()
channel_id = args.channel
# Check if this is an id or a name
if not channel_id.startswith("C"):
id = lookup_channel_id_by_name(args.token, args.channel)
if id == "":
print(channel_id, " was not found in the Slack channel list. Exiting...")
return False
channel_id = id
time_diff = str((datetime.now() - timedelta(days=int(args.past_days))
).timestamp()).split('.')[0]
messages = fetch_from_slack(args.token, channel_id, time_diff)
# open a file for writing
csv_file = open(args.filename, 'w')
# create the csv writer object
csvwriter = csv.writer(csv_file)
count = 0
last_timestamp = 0
for msg in messages:
try:
msgText = msg.get('text')
except:
raise
if msg.get('subtype') != 'bot_message':
msgUser = msg.get('user')
msg.setdefault('subtype', '')
if msgUser != None and msgText.find(args.text) == 0:
# Write the header if first row
if count == 0:
header = msg.keys()
del msg['subtype']
csvwriter.writerow(header)
count += 1
last_timestamp = datetime.fromtimestamp(
int(msg.get('ts').split('.')[0]))
# write the csv row
csvwriter.writerow(msg.values())
csv_file.close()
if __name__ == "__main__":
# execute only if run as a script
main()
|
# -*- coding: utf-8 -*-
# Max-Planck-Gesellschaft zur Förderung der Wissenschaften e.V. (MPG) is
# holder of all proprietary rights on this computer program.
# You can only use this computer program if you have closed
# a license agreement with MPG or you get the right to use the computer
# program from someone who is authorized to grant you that right.
# Any use of the computer program without a valid license is prohibited and
# liable to prosecution.
#
# Copyright©2019 Max-Planck-Gesellschaft zur Förderung
# der Wissenschaften e.V. (MPG). acting on behalf of its Max Planck Institute
# for Intelligent Systems. All rights reserved.
#
# Contact: <EMAIL>
import math
import trimesh
import pyrender
import numpy as np
from pyrender.constants import RenderFlags
class WeakPerspectiveCamera(pyrender.Camera):
def __init__(self,
scale,
translation,
znear=pyrender.camera.DEFAULT_Z_NEAR,
zfar=None,
name=None):
super(WeakPerspectiveCamera, self).__init__(
znear=znear,
zfar=zfar,
name=name,
)
self.scale = scale
self.translation = translation
def get_projection_matrix(self, width=None, height=None):
P = np.eye(4)
P[0, 0] = self.scale[0]
P[1, 1] = self.scale[1]
P[0, 3] = self.translation[0] * self.scale[0]
P[1, 3] = -self.translation[1] * self.scale[1]
P[2, 2] = -1
return P
class Renderer:
def __init__(self, face, resolution=(224,224), orig_img=False, wireframe=False):
self.resolution = resolution
self.faces = face
self.orig_img = orig_img
self.wireframe = wireframe
self.renderer = pyrender.OffscreenRenderer(
viewport_width=self.resolution[0],
viewport_height=self.resolution[1],
point_size=1.0
)
# set the scene
self.scene = pyrender.Scene(bg_color=[0.0, 0.0, 0.0, 0.0], ambient_light=(0.3, 0.3, 0.3))
# light = pyrender.PointLight(color=[1.0, 1.0, 1.0], intensity=0.8)
light = pyrender.DirectionalLight(color=[1.0, 1.0, 1.0], intensity=0.8)
light_pose = np.eye(4)
light_pose[:3, 3] = [0, -1, 1]
self.scene.add(light, pose=light_pose)
light_pose[:3, 3] = [0, 1, 1]
self.scene.add(light, pose=light_pose)
light_pose[:3, 3] = [1, 1, 2]
self.scene.add(light, pose=light_pose)
def render(self, img, verts, cam, angle=None, axis=None, mesh_filename=None, color=[1.0, 1.0, 0.9], rotate=False):
mesh = trimesh.Trimesh(vertices=verts, faces=self.faces, process=False)
Rx = trimesh.transformations.rotation_matrix(math.radians(180), [1, 0, 0])
mesh.apply_transform(Rx)
if rotate:
rot = trimesh.transformations.rotation_matrix(
np.radians(60), [0, 1, 0])
mesh.apply_transform(rot)
if mesh_filename is not None:
mesh.export(mesh_filename)
if angle and axis:
R = trimesh.transformations.rotation_matrix(math.radians(angle), axis)
mesh.apply_transform(R)
sx, sy, tx, ty = cam
camera = WeakPerspectiveCamera(
scale=[sx, sy],
translation=[tx, ty],
zfar=1000.
)
material = pyrender.MetallicRoughnessMaterial(
metallicFactor=0.0,
alphaMode='OPAQUE',
smooth=True,
wireframe=True,
roughnessFactor=1.0,
emissiveFactor=(0.1, 0.1, 0.1),
baseColorFactor=(color[0], color[1], color[2], 1.0)
)
# material = pyrender.MetallicRoughnessMaterial(
# metallicFactor=0.2,
# alphaMode='OPAQUE',
# baseColorFactor=(0.8, 0.3, 0.3, 1.0))
mesh = pyrender.Mesh.from_trimesh(mesh, material=material)
mesh_node = self.scene.add(mesh, 'mesh')
camera_pose = np.eye(4)
cam_node = self.scene.add(camera, pose=camera_pose)
if self.wireframe:
render_flags = RenderFlags.RGBA | RenderFlags.ALL_WIREFRAME
else:
render_flags = RenderFlags.RGBA
rgb, depth = self.renderer.render(self.scene, flags=render_flags)
valid_mask = (depth > 0)[:, :, np.newaxis]
output_img = rgb * valid_mask + (1 - valid_mask) * img
image = output_img.astype(np.uint8)
self.scene.remove_node(mesh_node)
self.scene.remove_node(cam_node)
return image
|
<filename>lib/svtplay_dl/service/viaplay.py
# ex:ts=4:sw=4:sts=4:et
# -*- tab-width: 4; c-basic-offset: 4; indent-tabs-mode: nil -*-
# pylint has issues with urlparse: "some types could not be inferred"
# pylint: disable=E1103
from __future__ import absolute_import
import re
import json
import copy
import os
from svtplay_dl.utils import filenamify
from svtplay_dl.utils.urllib import urlparse
from svtplay_dl.service import Service, OpenGraphThumbMixin
from svtplay_dl.log import log
from svtplay_dl.fetcher.rtmp import RTMP
from svtplay_dl.fetcher.hds import hdsparse
from svtplay_dl.fetcher.hls import hlsparse
from svtplay_dl.subtitle import subtitle
from svtplay_dl.error import ServiceError
class Viaplay(Service, OpenGraphThumbMixin):
supported_domains = [
'tv3play.se', 'tv6play.se', 'tv8play.se', 'tv10play.se',
'tv3play.no', 'tv3play.dk', 'tv6play.no', 'viasat4play.no',
'tv3play.ee', 'tv3play.lv', 'tv3play.lt', 'tvplay.lv', 'viagame.com',
'juicyplay.se', 'viafree.se', 'viafree.dk', 'viafree.no',
'play.tv3.lt', 'tv3play.tv3.ee', 'tvplay.skaties.lv'
]
def _get_video_id(self):
"""
Extract video id. It will try to avoid making an HTTP request
if it can find the ID in the URL, but otherwise it will try
to scrape it from the HTML document. Returns None in case it's
unable to extract the ID at all.
"""
html_data = self.get_urldata()
match = re.search(r'data-video-id="([0-9]+)"', html_data)
if match:
return match.group(1)
match = re.search(r'data-videoid="([0-9]+)', html_data)
if match:
return match.group(1)
clips = False
match = re.search('params":({.*}),"query', self.get_urldata())
if match:
jansson = json.loads(match.group(1))
if "seasonNumberOrVideoId" in jansson:
season = jansson["seasonNumberOrVideoId"]
match = re.search("\w-(\d+)$", season)
if match:
season = match.group(1)
else:
return False
if "videoIdOrEpisodeNumber" in jansson:
videp = jansson["videoIdOrEpisodeNumber"]
match = re.search('(\w+)-(\d+)', videp)
if match:
episodenr = match.group(2)
else:
episodenr = videp
clips = True
match = re.search('(s\w+)-(\d+)', season)
if match:
season = match.group(2)
else:
# sometimes videoIdOrEpisodeNumber does not work.. this is a workaround
match = re.search('(episode|avsnitt)-(\d+)', self.url)
if match:
episodenr = match.group(2)
else:
episodenr = season
if clips:
return episodenr
else:
match = re.search('"ContentPageProgramStore":({.*}),"ApplicationStore', self.get_urldata())
if match:
janson = json.loads(match.group(1))
for i in janson["format"]["videos"].keys():
for n in janson["format"]["videos"][i]["program"]:
if str(n["episodeNumber"]) and int(episodenr) == n["episodeNumber"] and int(season) == n["seasonNumber"]:
return n["id"]
elif n["id"] == episodenr:
return episodenr
parse = urlparse(self.url)
match = re.search(r'/\w+/(\d+)', parse.path)
if match:
return match.group(1)
match = re.search(r'iframe src="http://play.juicyplay.se[^\"]+id=(\d+)', html_data)
if match:
return match.group(1)
return None
def get(self):
vid = self._get_video_id()
if vid is None:
yield ServiceError("Can't find video file for: %s" % self.url)
return
url = "http://playapi.mtgx.tv/v3/videos/%s" % vid
self.options.other = ""
data = self.http.request("get", url)
if data.status_code == 403:
yield ServiceError("Can't play this because the video is geoblocked.")
return
dataj = json.loads(data.text)
if "msg" in dataj:
yield ServiceError(dataj["msg"])
return
if dataj["type"] == "live":
self.options.live = True
if self.exclude():
yield ServiceError("Excluding video")
return
streams = self.http.request("get", "http://playapi.mtgx.tv/v3/videos/stream/%s" % vid)
if streams.status_code == 403:
yield ServiceError("Can't play this because the video is geoblocked.")
return
streamj = json.loads(streams.text)
if "msg" in streamj:
yield ServiceError("Can't play this because the video is either not found or geoblocked.")
return
if self.options.output_auto:
directory = os.path.dirname(self.options.output)
self.options.service = "tv3play"
basename = self._autoname(dataj)
title = "%s-%s-%s" % (basename, vid, self.options.service)
if len(directory):
self.options.output = os.path.join(directory, title)
else:
self.options.output = title
if dataj["sami_path"]:
if dataj["sami_path"].endswith("vtt"):
subtype = "wrst"
else:
subtype = "sami"
yield subtitle(copy.copy(self.options), subtype, dataj["sami_path"])
if dataj["subtitles_webvtt"]:
yield subtitle(copy.copy(self.options), "wrst", dataj["subtitles_webvtt"])
if dataj["subtitles_for_hearing_impaired"]:
if dataj["subtitles_for_hearing_impaired"].endswith("vtt"):
subtype = "wrst"
else:
subtype = "sami"
if self.options.get_all_subtitles:
yield subtitle(copy.copy(self.options), subtype, dataj["subtitles_for_hearing_impaired"],"-SDH")
else:
yield subtitle(copy.copy(self.options), subtype, dataj["subtitles_for_hearing_impaired"])
if streamj["streams"]["medium"]:
filename = streamj["streams"]["medium"]
if ".f4m" in filename:
streams = hdsparse(self.options, self.http.request("get", filename, params={"hdcore": "3.7.0"}), filename)
if streams:
for n in list(streams.keys()):
yield streams[n]
else:
parse = urlparse(filename)
match = re.search("^(/[^/]+)/(.*)", parse.path)
if not match:
yield ServiceError("Can't get rtmpparse info")
return
filename = "%s://%s:%s%s" % (parse.scheme, parse.hostname, parse.port, match.group(1))
path = "-y %s" % match.group(2)
self.options.other = "-W http://flvplayer.viastream.viasat.tv/flvplayer/play/swf/player.swf %s" % path
yield RTMP(copy.copy(self.options), filename, 800)
if streamj["streams"]["hls"]:
streams = hlsparse(self.options, self.http.request("get", streamj["streams"]["hls"]), streamj["streams"]["hls"])
if streams:
for n in list(streams.keys()):
yield streams[n]
def find_all_episodes(self, options):
videos = []
match = re.search('"ContentPageProgramStore":({.*}),"ApplicationStore', self.get_urldata())
if match:
janson = json.loads(match.group(1))
seasons = []
for i in janson["format"]["seasons"]:
seasons.append(i["seasonNumber"])
for i in seasons:
for n in janson["format"]["videos"][str(i)]["program"]:
videos.append(n["sharingUrl"])
n = 0
episodes = []
for i in videos:
if n == options.all_last:
break
episodes.append(i)
n += 1
return episodes
def _autoname(self, dataj):
program = dataj["format_slug"]
season = dataj["format_position"]["season"]
episode = None
if season:
if len(dataj["format_position"]["episode"]) > 0:
episode = dataj["format_position"]["episode"]
name = filenamify(program)
if season:
name = "{}.s{:02d}".format(name, int(season))
if episode:
name = "{}e{:02d}".format(name, int(episode))
return name |
<reponame>ArtObr/indy-node
import json
from collections import OrderedDict
from plenum.common.constants import TXN_TYPE, TARGET_NYM, \
DATA, ENC, RAW, HASH, ALIAS, TXN_ID, TRUSTEE, STEWARD, \
TXN_TIME, VERKEY
from plenum.common.types import f
from indy_common.constants import NYM, ATTRIB, GET_ATTR, \
ROLE, REF, TRUST_ANCHOR, SIGNATURE_TYPE
def AddNym(target, role=None):
return newTxn(txnType=NYM, target=target, role=role)
def AddAttr(target, attrData, role=None):
return newTxn(txnType=ATTRIB, target=target, role=role,
enc=attrData)
def GetAttr(target, attrName, role=None):
queryData = json.dumps({"name": attrName})
return newTxn(txnType=GET_ATTR, target=target, role=role,
data=queryData)
# TODO: Change name to txn or some thing else after discussion
def newTxn(txnType, target=None, data=None, enc=None, raw=None,
hash=None, role=None):
txn = {
TXN_TYPE: txnType
}
if target:
txn[TARGET_NYM] = target
if data:
txn[DATA] = data
if enc:
txn[ENC] = enc
if raw:
txn[RAW] = raw
if hash:
txn[HASH] = hash
if role:
txn[ROLE] = role
return txn
def getGenesisTxns():
return [{ALIAS: "Trustee1",
TARGET_NYM: "9XNVHKtucEZWh7GrS9S8nRWtVuFQwYLfzGD7pQ7Scjtc",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4a",
TXN_TYPE: NYM,
ROLE: TRUSTEE},
{ALIAS: "Steward1",
TARGET_NYM: "5rArie7XKukPCaEwq5XGQJnM9Fc5aZE3M9HAPVfMU2xC",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4b",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward2",
TARGET_NYM: "2btLJAAb1S3x6hZYdVyAePjqtQYi2ZBSRGy4569RZu8h",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4c",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward3",
TARGET_NYM: "CECeGXDi6EHuhpwz19uyjjEnsRGNXodFYqCRgdLmLRkt",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4d",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward4",
TARGET_NYM: "3znAGhp6Tk4kmebhXnk9K3jaTMffu82PJfEG91AeRkq2",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4e",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward5",
TARGET_NYM: "4AdS22kC7xzb4bcqg9JATuCfAMNcQYcZa1u5eWzs6cSJ",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4f",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward6",
TARGET_NYM: "4Yk9HoDSfJv9QcmJbLcXdWVgS7nfvdUqiVcvbSu8VBru",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b50",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward7",
TARGET_NYM: "FR5pWwinRBn35GNhg7bsvw8Q13kRept2pm561DwZCQzT",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b51",
TXN_TYPE: NYM,
ROLE: STEWARD},
{TXN_TYPE: NYM,
TARGET_NYM: 'EGRf6ho37aqg5ZZpAyD2mesS6XrNUeSkoVUAbpL6bmJ9',
ROLE: STEWARD,
TXN_ID: '6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4b'},
{TXN_TYPE: NYM,
f.IDENTIFIER.nm: 'EGRf6ho37aqg5ZZpAyD2mesS6XrNUeSkoVUAbpL6bmJ9',
TARGET_NYM: 'C2AafyXuDBbcdiHJ8pdJ14PJ17X5KEBjbyfPPJWZFA4b',
ROLE: TRUST_ANCHOR,
TXN_ID: '6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4c'},
{TXN_TYPE: NYM,
TARGET_NYM: '4qU9QRZ79CbWuDKUtTvpDUnUiDnkLkwd1i8p2B3gJNU3',
TXN_ID: '50c2f66f7fda2ece684d1befc667e894b4460cb782f5387d864fa7d5f14c4066',
ROLE: TRUST_ANCHOR,
f.IDENTIFIER.nm: 'EGRf6ho37aqg5ZZpAyD2mesS6XrNUeSkoVUAbpL6bmJ9'},
{TXN_TYPE: NYM,
TARGET_NYM: 'adityastaging',
TXN_ID: '77c2f66f7fda2ece684d1befc667e894b4460cb782f5387d864fa7d5f14c4066',
f.IDENTIFIER.nm: '4qU9QRZ79CbWuDKUtTvpDUnUiDnkLkwd1i8p2B3gJNU3'},
{TXN_TYPE: NYM,
TARGET_NYM: 'iosstaging',
TXN_ID: '91c2f66f7fda2ece684d1befc667e894b4460cb782f5387d864fa7d5f14c4066',
f.IDENTIFIER.nm: '4qU9QRZ79CbWuDKUtTvpDUnUiDnkLkwd1i8p2B3gJNU3'},
{ALIAS: "Steward8",
TARGET_NYM: "6vAQkuCgTm7Jeki3vVhZm1FTAQYCeLE5mSvVRQdiwt1w",
TXN_ID: "4770beb7e45bf623bd9987af4bd6d6d8eb8b68a4d00fa2a4c6b6f3f0c1c036f8",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward9",
TARGET_NYM: "6hbecbh36EMK6yAi5NZ9bLZEuRsWFt6qLa2SyMQGXs7H",
TXN_ID: "4770beb7e45bf623bd9987af4bd6d6d8eb8b68a4d00fa2a4c6b6f3f0c1c036f9",
TXN_TYPE: NYM,
ROLE: STEWARD},
]
def getGenesisTxnsForLocal():
return [{ALIAS: "Steward1",
TARGET_NYM: "5rArie7XKukPCaEwq5XGQJnM9Fc5aZE3M9HAPVfMU2xC",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4b",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward2",
TARGET_NYM: "3NhxuJKShrpnhxG8VYGkum6mv3HeXWUDfj7ktn5NbeymHoDX",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4c",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward3",
TARGET_NYM: "CECeGXDi6EHuhpwz19uyjjEnsRGNXodFYqCRgdLmLRkt",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4d",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Steward4",
TARGET_NYM: "3znAGhp6Tk4kmebhXnk9K3jaTMffu82PJfEG91AeRkq2",
TXN_ID: "6b86b273ff34fce19d6b804eff5a3f5747ada4eaa22f1d49c01e52ddb7875b4e",
TXN_TYPE: NYM,
ROLE: STEWARD},
{ALIAS: "Alice",
TARGET_NYM: "4AdS22kC7xzb4bcqg9JATuCfAMNcQYcZa1u5eWzs6cSJ",
"identifier": "<KEY>",
TXN_ID: "e7f6c011776e8db7cd330b54174fd76f7d0216b612387a5ffcfb81e6f0919683",
TXN_TYPE: NYM},
{ALIAS: "Jason",
TARGET_NYM: "46Kq4hASUdvUbwR7s7Pie3x8f4HRB3NLay7Z9jh9eZsB",
"identifier": "<KEY>",
TXN_ID: "e7f6c011776e8db7cd330b54174fd76f7d0216b612387a5ffcfb81e6f0919684",
TXN_TYPE: NYM},
{ALIAS: "John",
TARGET_NYM: "3wpYnGqceZ8DzN3guiTd9rrYkWTwTHCChBSuo6cvkXTG",
"identifier": "<KEY>",
TXN_ID: "e7f6c011776e8db7cd330b54174fd76f7d0216b612387a5ffcfb81e6f0919685",
TXN_TYPE: NYM},
{ALIAS: "Les",
TARGET_NYM: "4Yk9HoDSfJv9QcmJbLcXdWVgS7nfvdUqiVcvbSu8VBru",
"identifier": "<KEY>",
TXN_ID: "e7f6c011776e8db7cd330b54174fd76f7d0216b612387a5ffcfb81e6f0919686",
TXN_TYPE: NYM}]
def getTxnOrderedFields():
return OrderedDict([
(f.IDENTIFIER.nm, (str, str)),
(f.REQ_ID.nm, (str, int)),
(f.SIG.nm, (str, str)),
(TXN_TIME, (str, int)),
(TXN_TYPE, (str, str)),
(TARGET_NYM, (str, str)),
(VERKEY, (str, str)),
(DATA, (str, str)),
(ALIAS, (str, str)),
(RAW, (str, str)),
(ENC, (str, str)),
(HASH, (str, str)),
(ROLE, (str, str)),
(REF, (str, str)),
(SIGNATURE_TYPE, (str, str))
])
|
<gh_stars>0
import time
from datetime import datetime, date
import pandas as pd
import sqlalchemy as sa
from sqlalchemy.exc import InternalError
from analysis.utils.db import engine, session
from analysis.utils.db import DailyDiagnosticChangeModel
from analysis.utils.db import IndividualReportModel
from analysis.utils import db_enum as enum
def calculate(report):
"""Calculate symptom factor per report / row."""
S = sum(
column.value
for column in (
enum.Scale3[report.temp],
enum.Scale4[report.cough],
enum.Scale4[report.breathless],
enum.Energy[report.energy],
)
)
return S
def map_calculate(collection_size: int):
"""Calcalate symptom factor S for the whole DB where analysis_done = 0"""
start_time_analysis = time.time()
# load the next collection of reports to analyse
next_reports = pd.read_sql(
(
"SELECT * FROM individual_report WHERE analysis_done = 0 "
"ORDER BY timestamp LIMIT "
)
+ str(collection_size),
con=engine,
index_col="document_id",
)
S = next_reports.apply(calculate, axis=1)
query = """
UPDATE individual_report AS old, temp_table AS new
SET old.S = new.0, old.analysis_done = 1
WHERE old.document_id = new.document_id
"""
with engine.begin() as con:
S.to_sql(
"temp_table",
con,
if_exists="replace",
dtype={
"document_id": sa.String(30),
"S": sa.Integer,
}
)
try:
con.execute(query)
except:
print("ERROR: while executing query: {}".format(query))
try:
con.execute("DROP TABLE temp_table")
except InternalError:
print("WARNING: no temp_table to drop")
session.commit()
spend_time = time.time() - start_time_analysis
print('Analysed {} samples in {} s'.format(collection_size, spend_time))
def group_reports_by_location():
start_time_analysis = time.time()
# load all analysed reports
df = pd.read_sql(
"SELECT * FROM individual_report WHERE analysis_done = 1",
con=engine,
index_col="document_id",
)
# diagnosis using S factors
# healthy = df.query('S == 0')
# sick_guess_no_corona = df.query('0 < S < 4')
# sick_guess_corona = df.query('S >= 4')
# diagnosis from report
# healthy = df.query('diagnostic == 0')
# sick_guess_no_corona = df.query('diagnostic == 1')
# sick_guess_corona = df.query('diagnostic == 2')
# sick_corona_confirmed = df.query('diagnostic == 3')
# recovered_confirmed = df.query('diagnostic == 4')
# recovered_not_confirmed = df.query('diagnostic == 5')
def group(df_diagnosis):
"""Get number of reports by location"""
return df_diagnosis.groupby('locator').sum()
def to_sql(totals, column):
query = """
UPDATE locations AS old, temp_totals AS new
SET old.{column} = new.analysis_done
WHERE old.postal_code = new.locator
""".format(column=column)
with engine.begin() as con:
totals.to_sql(
"temp_totals",
con,
if_exists="replace",
dtype={
"locator": sa.Integer,
}
)
try:
con.execute(query)
except:
print("ERROR: while executing query: {}".format(query))
try:
con.execute("DROP TABLE temp_totals")
except InternalError:
print("WARNING: no temp_totals to drop")
# total_healthy = group(healthy)
# print(total_healthy)
# to_sql(total_healthy, "total_healthy")
columns = (
"healthy",
"sick_guess_no_corona",
"sick_guess_corona",
"sick_corona_confirmed",
"recovered_confirmed",
"recovered_not_confirmed",
)
for diagnostic, column in enumerate(columns):
df_diagnosis = df.query('diagnostic == {}'.format(diagnostic))
df_totals = group(df_diagnosis)
to_sql(df_totals, "total_{}".format(column))
session.commit()
spend_time = time.time() - start_time_analysis
print('Grouped {} samples by location in {} s'.format(len(df), spend_time))
if __name__ == "__main__":
map_calculate(10)
group_reports_by_location()
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.