|
|
--- |
|
|
license: mit |
|
|
tags: |
|
|
- rna |
|
|
- biology |
|
|
- rna-design |
|
|
- biomolecule-design |
|
|
- 3d-design |
|
|
- inverse-folding |
|
|
- inverse-design |
|
|
--- |
|
|
|
|
|
# gRNAde Datasets |
|
|
|
|
|
[](https://www.biorxiv.org/content/10.1101/2025.11.29.691298) |
|
|
[](https://github.com/chaitjo/geometric-rna-design/blob/main/LICENSE) |
|
|
[](https://github.com/chaitjo/geometric-rna-design) |
|
|
|
|
|
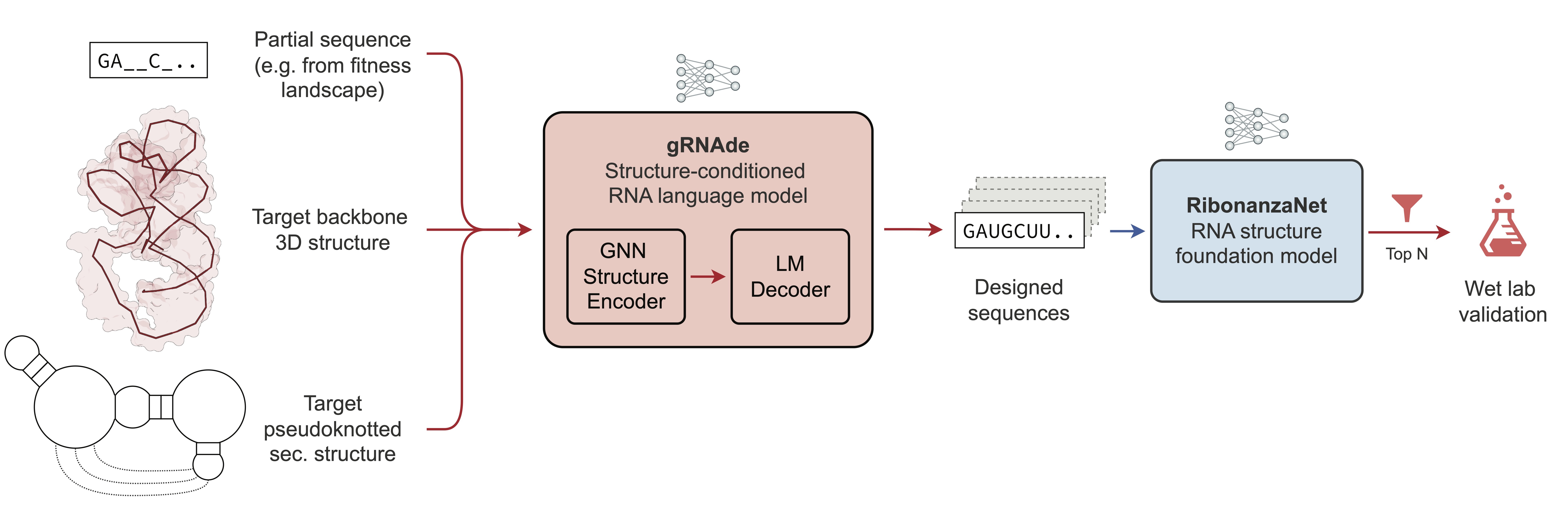
This repository contains datasets and wet-lab experimental data for **gRNAde**, a generative AI framework for RNA inverse design. |
|
|
|
|
|
 |
|
|
|
|
|
## 📦 Datasets and Experimental Data |
|
|
|
|
|
``` |
|
|
. |
|
|
├── RNASolo_31102023_processed.pt # Pre-processed training dataset (ready for ML) |
|
|
├── RNASolo_31102023_raw.tar.gz # Raw PDB structures from RNASolo |
|
|
└── projects # Data for reproducing design campaigns in the paper |
|
|
├── openknot_benchmark # Eterna OpenKnot Benchmark for psuedoknotted RNA design |
|
|
└── rna_polymerase_ribozyme # Generative Design of RNA polymerase ribozymes |
|
|
``` |
|
|
|
|
|
### Pre-processed Training Dataset |
|
|
- **File**: `RNASolo_31102023_processed.pt` |
|
|
- **Description**: ML-ready dataset of RNA 3D structures from the Protein Data Bank |
|
|
- **Source**: RNASolo database (October 31, 2023 cutoff) |
|
|
- **Resolution**: ≤4.0 Å |
|
|
- **Format**: PyTorch tensors with sequences, 3D coordinates, secondary structures, and metadata |
|
|
- **Use case**: Training new gRNAde models or fine-tuning for specific RNA families |
|
|
|
|
|
### Raw PDB Structures |
|
|
- **File**: `RNASolo_31102023_raw.tar.gz` |
|
|
- **Description**: Raw PDB files for all RNA structures from RNASolo |
|
|
- **Contents**: Thousands of experimentally determined RNA structures |
|
|
- **Use case**: Custom data processing pipelines or structure visualization |
|
|
|
|
|
### Eterna OpenKnot Competition Data |
|
|
- **Directory**: `projects/openknot_benchmark/` |
|
|
- **Description**: Designs and experimental validation data from the Eterna OpenKnot Benchmark |
|
|
- **Contents**: Chemical probing data, OpenKnot scores, and designed sequences |
|
|
- **Targets**: 40 pseudoknotted RNA puzzles including riboswitches, ribozymes, and viral elements |
|
|
|
|
|
### RNA Polymerase Ribozyme Data |
|
|
- **Directory**: `projects/rna_polymerase_ribozyme/` |
|
|
- **Description**: Functional design campaign data for engineering RNA polymerase ribozymes |
|
|
- **Contents**: Generated sequences, activity assays, fitness landscapes |
|
|
|
|
|
## 🚀 Quick Start |
|
|
|
|
|
After setting up the [gRNAde codebase](https://github.com/chaitjo/geometric-rna-design), datasets can be downloaded manually or using HuggingFace CLI (recommended): |
|
|
|
|
|
```bash |
|
|
# Ensure you are in the base directory |
|
|
cd ~/geometric-rna-design |
|
|
|
|
|
# Install HuggingFace CLI (https://huggingface.co/docs/huggingface_hub/main/en/guides/cli) |
|
|
pip install -U "huggingface_hub[cli]" |
|
|
# alternate: curl -LsSf https://hf.co/cli/install.sh | bash |
|
|
# alternate: brew install huggingface-cli |
|
|
|
|
|
# Download all datasets to data/ directory |
|
|
huggingface-cli download chaitjo/gRNAde_datasets --local-dir data/ --repo-type dataset |
|
|
``` |
|
|
|
|
|
Alternately, download specific files: |
|
|
|
|
|
```bash |
|
|
# Download only the pre-processed training dataset |
|
|
huggingface-cli download chaitjo/gRNAde_datasets RNASolo_31102023_processed.pt --local-dir data/ --repo-type dataset |
|
|
|
|
|
# Download only raw PDB structures |
|
|
huggingface-cli download chaitjo/gRNAde_datasets RNASolo_31102023_raw.tar.gz --local-dir data/ --repo-type dataset |
|
|
``` |
|
|
|
|
|
## Citations |
|
|
|
|
|
``` |
|
|
@article{joshi2025generative, |
|
|
title={Generative inverse design of RNA structure and function with g{RNA}de}, |
|
|
author={Joshi, Chaitanya K and Gianni, Edoardo and Kwok, Samantha LY and Mathis, Simon V and Lio, Pietro and Holliger, Philipp}, |
|
|
journal={bioRxiv}, |
|
|
year={2025}, |
|
|
publisher={Cold Spring Harbor Laboratory} |
|
|
} |
|
|
|
|
|
@inproceedings{joshi2025grnade, |
|
|
title={g{RNA}de: Geometric Deep Learning for 3D RNA inverse design}, |
|
|
author={Joshi, Chaitanya K and Jamasb, Arian R and Vi{\~n}as, Ramon and Harris, Charles and Mathis, Simon V and Morehead, Alex and Anand, Rishabh and Li{\`o}, Pietro}, |
|
|
booktitle={International Conference on Learning Representations (ICLR)}, |
|
|
year={2025}, |
|
|
} |
|
|
|
|
|
@incollection{joshi2024grnade, |
|
|
title={g{RNA}de: A Geometric Deep Learning pipeline for 3D RNA inverse design}, |
|
|
author={Joshi, Chaitanya K and Li{\`o}, Pietro}, |
|
|
booktitle={RNA Design: Methods and Protocols}, |
|
|
pages={121--135}, |
|
|
year={2024}, |
|
|
publisher={Springer} |
|
|
} |
|
|
``` |
|
|
|