id stringlengths 2 115 | lastModified stringlengths 24 24 | tags list | author stringlengths 2 42 ⌀ | description stringlengths 0 68.7k ⌀ | citation stringlengths 0 10.7k ⌀ | cardData null | likes int64 0 3.55k | downloads int64 0 10.1M | card stringlengths 0 1.01M |
|---|---|---|---|---|---|---|---|---|---|
marcel113/texttoimage | 2023-09-17T21:14:38.000Z | [
"region:us"
] | marcel113 | null | null | null | 0 | 0 | Entry not found |
barto17/gtzan_all_preprocessed | 2023-09-17T21:21:55.000Z | [
"region:us"
] | barto17 | null | null | null | 0 | 0 | ---
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
- split: test
path: data/test-*
dataset_info:
features:
- name: label
dtype:
class_label:
names:
'0': blues
'1': classical
'2': country
'3': disco
'4': hiphop
'5': jazz
'6': metal
'7': pop
'8': reggae
'9': rock
- name: input_values
sequence: float32
- name: attention_mask
sequence: int32
splits:
- name: train
num_bytes: 3452159816
num_examples: 899
- name: test
num_bytes: 384000696
num_examples: 100
download_size: 1923103923
dataset_size: 3836160512
---
# Dataset Card for "gtzan_all_preprocessed"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
Resizable/ToroInoue | 2023-09-17T21:29:31.000Z | [
"license:openrail",
"region:us"
] | Resizable | null | null | null | 0 | 0 | ---
license: openrail
---
|
open-llm-leaderboard/details_deepse__CodeUp-Llama-2-13b-chat-hf | 2023-09-17T21:43:00.000Z | [
"region:us"
] | open-llm-leaderboard | null | null | null | 0 | 0 | ---
pretty_name: Evaluation run of deepse/CodeUp-Llama-2-13b-chat-hf
dataset_summary: "Dataset automatically created during the evaluation run of model\
\ [deepse/CodeUp-Llama-2-13b-chat-hf](https://huggingface.co/deepse/CodeUp-Llama-2-13b-chat-hf)\
\ on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).\n\
\nThe dataset is composed of 3 configuration, each one coresponding to one of the\
\ evaluated task.\n\nThe dataset has been created from 1 run(s). Each run can be\
\ found as a specific split in each configuration, the split being named using the\
\ timestamp of the run.The \"train\" split is always pointing to the latest results.\n\
\nAn additional configuration \"results\" store all the aggregated results of the\
\ run (and is used to compute and display the agregated metrics on the [Open LLM\
\ Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).\n\
\nTo load the details from a run, you can for instance do the following:\n```python\n\
from datasets import load_dataset\ndata = load_dataset(\"open-llm-leaderboard/details_deepse__CodeUp-Llama-2-13b-chat-hf\"\
,\n\t\"harness_winogrande_5\",\n\tsplit=\"train\")\n```\n\n## Latest results\n\n\
These are the [latest results from run 2023-09-17T21:42:48.305634](https://huggingface.co/datasets/open-llm-leaderboard/details_deepse__CodeUp-Llama-2-13b-chat-hf/blob/main/results_2023-09-17T21-42-48.305634.json)(note\
\ that their might be results for other tasks in the repos if successive evals didn't\
\ cover the same tasks. You find each in the results and the \"latest\" split for\
\ each eval):\n\n```python\n{\n \"all\": {\n \"em\": 0.1782718120805369,\n\
\ \"em_stderr\": 0.003919630092588375,\n \"f1\": 0.2387195889261742,\n\
\ \"f1_stderr\": 0.003944947017182046,\n \"acc\": 0.448727630233375,\n\
\ \"acc_stderr\": 0.011074189612085313\n },\n \"harness|drop|3\": {\n\
\ \"em\": 0.1782718120805369,\n \"em_stderr\": 0.003919630092588375,\n\
\ \"f1\": 0.2387195889261742,\n \"f1_stderr\": 0.003944947017182046\n\
\ },\n \"harness|gsm8k|5\": {\n \"acc\": 0.15238817285822592,\n \
\ \"acc_stderr\": 0.009899572254794204\n },\n \"harness|winogrande|5\"\
: {\n \"acc\": 0.745067087608524,\n \"acc_stderr\": 0.012248806969376422\n\
\ }\n}\n```"
repo_url: https://huggingface.co/deepse/CodeUp-Llama-2-13b-chat-hf
leaderboard_url: https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
point_of_contact: clementine@hf.co
configs:
- config_name: harness_drop_3
data_files:
- split: 2023_09_17T21_42_48.305634
path:
- '**/details_harness|drop|3_2023-09-17T21-42-48.305634.parquet'
- split: latest
path:
- '**/details_harness|drop|3_2023-09-17T21-42-48.305634.parquet'
- config_name: harness_gsm8k_5
data_files:
- split: 2023_09_17T21_42_48.305634
path:
- '**/details_harness|gsm8k|5_2023-09-17T21-42-48.305634.parquet'
- split: latest
path:
- '**/details_harness|gsm8k|5_2023-09-17T21-42-48.305634.parquet'
- config_name: harness_winogrande_5
data_files:
- split: 2023_09_17T21_42_48.305634
path:
- '**/details_harness|winogrande|5_2023-09-17T21-42-48.305634.parquet'
- split: latest
path:
- '**/details_harness|winogrande|5_2023-09-17T21-42-48.305634.parquet'
- config_name: results
data_files:
- split: 2023_09_17T21_42_48.305634
path:
- results_2023-09-17T21-42-48.305634.parquet
- split: latest
path:
- results_2023-09-17T21-42-48.305634.parquet
---
# Dataset Card for Evaluation run of deepse/CodeUp-Llama-2-13b-chat-hf
## Dataset Description
- **Homepage:**
- **Repository:** https://huggingface.co/deepse/CodeUp-Llama-2-13b-chat-hf
- **Paper:**
- **Leaderboard:** https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
- **Point of Contact:** clementine@hf.co
### Dataset Summary
Dataset automatically created during the evaluation run of model [deepse/CodeUp-Llama-2-13b-chat-hf](https://huggingface.co/deepse/CodeUp-Llama-2-13b-chat-hf) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).
The dataset is composed of 3 configuration, each one coresponding to one of the evaluated task.
The dataset has been created from 1 run(s). Each run can be found as a specific split in each configuration, the split being named using the timestamp of the run.The "train" split is always pointing to the latest results.
An additional configuration "results" store all the aggregated results of the run (and is used to compute and display the agregated metrics on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).
To load the details from a run, you can for instance do the following:
```python
from datasets import load_dataset
data = load_dataset("open-llm-leaderboard/details_deepse__CodeUp-Llama-2-13b-chat-hf",
"harness_winogrande_5",
split="train")
```
## Latest results
These are the [latest results from run 2023-09-17T21:42:48.305634](https://huggingface.co/datasets/open-llm-leaderboard/details_deepse__CodeUp-Llama-2-13b-chat-hf/blob/main/results_2023-09-17T21-42-48.305634.json)(note that their might be results for other tasks in the repos if successive evals didn't cover the same tasks. You find each in the results and the "latest" split for each eval):
```python
{
"all": {
"em": 0.1782718120805369,
"em_stderr": 0.003919630092588375,
"f1": 0.2387195889261742,
"f1_stderr": 0.003944947017182046,
"acc": 0.448727630233375,
"acc_stderr": 0.011074189612085313
},
"harness|drop|3": {
"em": 0.1782718120805369,
"em_stderr": 0.003919630092588375,
"f1": 0.2387195889261742,
"f1_stderr": 0.003944947017182046
},
"harness|gsm8k|5": {
"acc": 0.15238817285822592,
"acc_stderr": 0.009899572254794204
},
"harness|winogrande|5": {
"acc": 0.745067087608524,
"acc_stderr": 0.012248806969376422
}
}
```
### Supported Tasks and Leaderboards
[More Information Needed]
### Languages
[More Information Needed]
## Dataset Structure
### Data Instances
[More Information Needed]
### Data Fields
[More Information Needed]
### Data Splits
[More Information Needed]
## Dataset Creation
### Curation Rationale
[More Information Needed]
### Source Data
#### Initial Data Collection and Normalization
[More Information Needed]
#### Who are the source language producers?
[More Information Needed]
### Annotations
#### Annotation process
[More Information Needed]
#### Who are the annotators?
[More Information Needed]
### Personal and Sensitive Information
[More Information Needed]
## Considerations for Using the Data
### Social Impact of Dataset
[More Information Needed]
### Discussion of Biases
[More Information Needed]
### Other Known Limitations
[More Information Needed]
## Additional Information
### Dataset Curators
[More Information Needed]
### Licensing Information
[More Information Needed]
### Citation Information
[More Information Needed]
### Contributions
[More Information Needed] |
chenqile09/llama2-chinese-couplet-770k | 2023-09-17T22:02:20.000Z | [
"region:us"
] | chenqile09 | null | null | null | 0 | 0 | ---
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
- split: validation
path: data/validation-*
dataset_info:
features:
- name: input
dtype: string
- name: output
dtype: string
- name: text
dtype: string
splits:
- name: train
num_bytes: 261365259
num_examples: 770491
- name: validation
num_bytes: 1358512
num_examples: 4000
download_size: 101554099
dataset_size: 262723771
---
# Dataset Card for "llama2-chinese-couplet-770k"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
OliBomby/ORS13402 | 2023-09-17T22:03:22.000Z | [
"license:mit",
"region:us"
] | OliBomby | null | null | null | 0 | 0 | ---
license: mit
---
|
josedanielaromi/Arg1998 | 2023-09-17T22:35:07.000Z | [
"region:us"
] | josedanielaromi | null | null | null | 0 | 0 | Entry not found |
LIXOLIXO/LIXOOOOOOOOOO | 2023-09-17T22:53:33.000Z | [
"region:us"
] | LIXOLIXO | null | null | null | 0 | 0 | Entry not found |
Albinator/Frankensteins-Monster-Grunts | 2023-09-17T23:07:36.000Z | [
"region:us"
] | Albinator | null | null | null | 0 | 0 | Entry not found |
LIXOLIXO/robsu | 2023-09-17T23:19:50.000Z | [
"region:us"
] | LIXOLIXO | null | null | null | 0 | 0 | Entry not found |
open-llm-leaderboard/details_w601sxs__b1ade-1b | 2023-09-17T23:22:35.000Z | [
"region:us"
] | open-llm-leaderboard | null | null | null | 0 | 0 | ---
pretty_name: Evaluation run of w601sxs/b1ade-1b
dataset_summary: "Dataset automatically created during the evaluation run of model\
\ [w601sxs/b1ade-1b](https://huggingface.co/w601sxs/b1ade-1b) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).\n\
\nThe dataset is composed of 3 configuration, each one coresponding to one of the\
\ evaluated task.\n\nThe dataset has been created from 1 run(s). Each run can be\
\ found as a specific split in each configuration, the split being named using the\
\ timestamp of the run.The \"train\" split is always pointing to the latest results.\n\
\nAn additional configuration \"results\" store all the aggregated results of the\
\ run (and is used to compute and display the agregated metrics on the [Open LLM\
\ Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).\n\
\nTo load the details from a run, you can for instance do the following:\n```python\n\
from datasets import load_dataset\ndata = load_dataset(\"open-llm-leaderboard/details_w601sxs__b1ade-1b\"\
,\n\t\"harness_winogrande_5\",\n\tsplit=\"train\")\n```\n\n## Latest results\n\n\
These are the [latest results from run 2023-09-17T23:22:23.773676](https://huggingface.co/datasets/open-llm-leaderboard/details_w601sxs__b1ade-1b/blob/main/results_2023-09-17T23-22-23.773676.json)(note\
\ that their might be results for other tasks in the repos if successive evals didn't\
\ cover the same tasks. You find each in the results and the \"latest\" split for\
\ each eval):\n\n```python\n{\n \"all\": {\n \"em\": 0.002936241610738255,\n\
\ \"em_stderr\": 0.0005541113054709937,\n \"f1\": 0.042455956375839043,\n\
\ \"f1_stderr\": 0.0012020859118158512,\n \"acc\": 0.2721723005338167,\n\
\ \"acc_stderr\": 0.00807495644821229\n },\n \"harness|drop|3\": {\n\
\ \"em\": 0.002936241610738255,\n \"em_stderr\": 0.0005541113054709937,\n\
\ \"f1\": 0.042455956375839043,\n \"f1_stderr\": 0.0012020859118158512\n\
\ },\n \"harness|gsm8k|5\": {\n \"acc\": 0.006065200909780136,\n \
\ \"acc_stderr\": 0.0021386703014604556\n },\n \"harness|winogrande|5\"\
: {\n \"acc\": 0.5382794001578532,\n \"acc_stderr\": 0.014011242594964125\n\
\ }\n}\n```"
repo_url: https://huggingface.co/w601sxs/b1ade-1b
leaderboard_url: https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
point_of_contact: clementine@hf.co
configs:
- config_name: harness_drop_3
data_files:
- split: 2023_09_17T23_22_23.773676
path:
- '**/details_harness|drop|3_2023-09-17T23-22-23.773676.parquet'
- split: latest
path:
- '**/details_harness|drop|3_2023-09-17T23-22-23.773676.parquet'
- config_name: harness_gsm8k_5
data_files:
- split: 2023_09_17T23_22_23.773676
path:
- '**/details_harness|gsm8k|5_2023-09-17T23-22-23.773676.parquet'
- split: latest
path:
- '**/details_harness|gsm8k|5_2023-09-17T23-22-23.773676.parquet'
- config_name: harness_winogrande_5
data_files:
- split: 2023_09_17T23_22_23.773676
path:
- '**/details_harness|winogrande|5_2023-09-17T23-22-23.773676.parquet'
- split: latest
path:
- '**/details_harness|winogrande|5_2023-09-17T23-22-23.773676.parquet'
- config_name: results
data_files:
- split: 2023_09_17T23_22_23.773676
path:
- results_2023-09-17T23-22-23.773676.parquet
- split: latest
path:
- results_2023-09-17T23-22-23.773676.parquet
---
# Dataset Card for Evaluation run of w601sxs/b1ade-1b
## Dataset Description
- **Homepage:**
- **Repository:** https://huggingface.co/w601sxs/b1ade-1b
- **Paper:**
- **Leaderboard:** https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
- **Point of Contact:** clementine@hf.co
### Dataset Summary
Dataset automatically created during the evaluation run of model [w601sxs/b1ade-1b](https://huggingface.co/w601sxs/b1ade-1b) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).
The dataset is composed of 3 configuration, each one coresponding to one of the evaluated task.
The dataset has been created from 1 run(s). Each run can be found as a specific split in each configuration, the split being named using the timestamp of the run.The "train" split is always pointing to the latest results.
An additional configuration "results" store all the aggregated results of the run (and is used to compute and display the agregated metrics on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).
To load the details from a run, you can for instance do the following:
```python
from datasets import load_dataset
data = load_dataset("open-llm-leaderboard/details_w601sxs__b1ade-1b",
"harness_winogrande_5",
split="train")
```
## Latest results
These are the [latest results from run 2023-09-17T23:22:23.773676](https://huggingface.co/datasets/open-llm-leaderboard/details_w601sxs__b1ade-1b/blob/main/results_2023-09-17T23-22-23.773676.json)(note that their might be results for other tasks in the repos if successive evals didn't cover the same tasks. You find each in the results and the "latest" split for each eval):
```python
{
"all": {
"em": 0.002936241610738255,
"em_stderr": 0.0005541113054709937,
"f1": 0.042455956375839043,
"f1_stderr": 0.0012020859118158512,
"acc": 0.2721723005338167,
"acc_stderr": 0.00807495644821229
},
"harness|drop|3": {
"em": 0.002936241610738255,
"em_stderr": 0.0005541113054709937,
"f1": 0.042455956375839043,
"f1_stderr": 0.0012020859118158512
},
"harness|gsm8k|5": {
"acc": 0.006065200909780136,
"acc_stderr": 0.0021386703014604556
},
"harness|winogrande|5": {
"acc": 0.5382794001578532,
"acc_stderr": 0.014011242594964125
}
}
```
### Supported Tasks and Leaderboards
[More Information Needed]
### Languages
[More Information Needed]
## Dataset Structure
### Data Instances
[More Information Needed]
### Data Fields
[More Information Needed]
### Data Splits
[More Information Needed]
## Dataset Creation
### Curation Rationale
[More Information Needed]
### Source Data
#### Initial Data Collection and Normalization
[More Information Needed]
#### Who are the source language producers?
[More Information Needed]
### Annotations
#### Annotation process
[More Information Needed]
#### Who are the annotators?
[More Information Needed]
### Personal and Sensitive Information
[More Information Needed]
## Considerations for Using the Data
### Social Impact of Dataset
[More Information Needed]
### Discussion of Biases
[More Information Needed]
### Other Known Limitations
[More Information Needed]
## Additional Information
### Dataset Curators
[More Information Needed]
### Licensing Information
[More Information Needed]
### Citation Information
[More Information Needed]
### Contributions
[More Information Needed] |
Marros/capes-geral-21 | 2023-09-28T13:19:44.000Z | [
"language:pt",
"license:mit",
"region:us"
] | Marros | null | null | null | 0 | 0 | ---
license: mit
language:
- pt
--- |
LIXOLIXO/robso | 2023-09-17T23:47:28.000Z | [
"region:us"
] | LIXOLIXO | null | null | null | 0 | 0 | Entry not found |
open-llm-leaderboard/details_mncai__SGPT-1.3B-insurance-epoch10 | 2023-09-18T00:09:18.000Z | [
"region:us"
] | open-llm-leaderboard | null | null | null | 0 | 0 | ---
pretty_name: Evaluation run of mncai/SGPT-1.3B-insurance-epoch10
dataset_summary: "Dataset automatically created during the evaluation run of model\
\ [mncai/SGPT-1.3B-insurance-epoch10](https://huggingface.co/mncai/SGPT-1.3B-insurance-epoch10)\
\ on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).\n\
\nThe dataset is composed of 3 configuration, each one coresponding to one of the\
\ evaluated task.\n\nThe dataset has been created from 1 run(s). Each run can be\
\ found as a specific split in each configuration, the split being named using the\
\ timestamp of the run.The \"train\" split is always pointing to the latest results.\n\
\nAn additional configuration \"results\" store all the aggregated results of the\
\ run (and is used to compute and display the agregated metrics on the [Open LLM\
\ Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).\n\
\nTo load the details from a run, you can for instance do the following:\n```python\n\
from datasets import load_dataset\ndata = load_dataset(\"open-llm-leaderboard/details_mncai__SGPT-1.3B-insurance-epoch10\"\
,\n\t\"harness_winogrande_5\",\n\tsplit=\"train\")\n```\n\n## Latest results\n\n\
These are the [latest results from run 2023-09-18T00:09:04.877490](https://huggingface.co/datasets/open-llm-leaderboard/details_mncai__SGPT-1.3B-insurance-epoch10/blob/main/results_2023-09-18T00-09-04.877490.json)(note\
\ that their might be results for other tasks in the repos if successive evals didn't\
\ cover the same tasks. You find each in the results and the \"latest\" split for\
\ each eval):\n\n```python\n{\n \"all\": {\n \"em\": 0.0,\n \"\
em_stderr\": 0.0,\n \"f1\": 1.99244966442953e-05,\n \"f1_stderr\"\
: 5.6438034448796525e-06,\n \"acc\": 0.25453827940015783,\n \"acc_stderr\"\
: 0.007025085047248852\n },\n \"harness|drop|3\": {\n \"em\": 0.0,\n\
\ \"em_stderr\": 0.0,\n \"f1\": 1.99244966442953e-05,\n \"\
f1_stderr\": 5.6438034448796525e-06\n },\n \"harness|gsm8k|5\": {\n \
\ \"acc\": 0.0,\n \"acc_stderr\": 0.0\n },\n \"harness|winogrande|5\"\
: {\n \"acc\": 0.5090765588003157,\n \"acc_stderr\": 0.014050170094497704\n\
\ }\n}\n```"
repo_url: https://huggingface.co/mncai/SGPT-1.3B-insurance-epoch10
leaderboard_url: https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
point_of_contact: clementine@hf.co
configs:
- config_name: harness_drop_3
data_files:
- split: 2023_09_18T00_09_04.877490
path:
- '**/details_harness|drop|3_2023-09-18T00-09-04.877490.parquet'
- split: latest
path:
- '**/details_harness|drop|3_2023-09-18T00-09-04.877490.parquet'
- config_name: harness_gsm8k_5
data_files:
- split: 2023_09_18T00_09_04.877490
path:
- '**/details_harness|gsm8k|5_2023-09-18T00-09-04.877490.parquet'
- split: latest
path:
- '**/details_harness|gsm8k|5_2023-09-18T00-09-04.877490.parquet'
- config_name: harness_winogrande_5
data_files:
- split: 2023_09_18T00_09_04.877490
path:
- '**/details_harness|winogrande|5_2023-09-18T00-09-04.877490.parquet'
- split: latest
path:
- '**/details_harness|winogrande|5_2023-09-18T00-09-04.877490.parquet'
- config_name: results
data_files:
- split: 2023_09_18T00_09_04.877490
path:
- results_2023-09-18T00-09-04.877490.parquet
- split: latest
path:
- results_2023-09-18T00-09-04.877490.parquet
---
# Dataset Card for Evaluation run of mncai/SGPT-1.3B-insurance-epoch10
## Dataset Description
- **Homepage:**
- **Repository:** https://huggingface.co/mncai/SGPT-1.3B-insurance-epoch10
- **Paper:**
- **Leaderboard:** https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
- **Point of Contact:** clementine@hf.co
### Dataset Summary
Dataset automatically created during the evaluation run of model [mncai/SGPT-1.3B-insurance-epoch10](https://huggingface.co/mncai/SGPT-1.3B-insurance-epoch10) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).
The dataset is composed of 3 configuration, each one coresponding to one of the evaluated task.
The dataset has been created from 1 run(s). Each run can be found as a specific split in each configuration, the split being named using the timestamp of the run.The "train" split is always pointing to the latest results.
An additional configuration "results" store all the aggregated results of the run (and is used to compute and display the agregated metrics on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).
To load the details from a run, you can for instance do the following:
```python
from datasets import load_dataset
data = load_dataset("open-llm-leaderboard/details_mncai__SGPT-1.3B-insurance-epoch10",
"harness_winogrande_5",
split="train")
```
## Latest results
These are the [latest results from run 2023-09-18T00:09:04.877490](https://huggingface.co/datasets/open-llm-leaderboard/details_mncai__SGPT-1.3B-insurance-epoch10/blob/main/results_2023-09-18T00-09-04.877490.json)(note that their might be results for other tasks in the repos if successive evals didn't cover the same tasks. You find each in the results and the "latest" split for each eval):
```python
{
"all": {
"em": 0.0,
"em_stderr": 0.0,
"f1": 1.99244966442953e-05,
"f1_stderr": 5.6438034448796525e-06,
"acc": 0.25453827940015783,
"acc_stderr": 0.007025085047248852
},
"harness|drop|3": {
"em": 0.0,
"em_stderr": 0.0,
"f1": 1.99244966442953e-05,
"f1_stderr": 5.6438034448796525e-06
},
"harness|gsm8k|5": {
"acc": 0.0,
"acc_stderr": 0.0
},
"harness|winogrande|5": {
"acc": 0.5090765588003157,
"acc_stderr": 0.014050170094497704
}
}
```
### Supported Tasks and Leaderboards
[More Information Needed]
### Languages
[More Information Needed]
## Dataset Structure
### Data Instances
[More Information Needed]
### Data Fields
[More Information Needed]
### Data Splits
[More Information Needed]
## Dataset Creation
### Curation Rationale
[More Information Needed]
### Source Data
#### Initial Data Collection and Normalization
[More Information Needed]
#### Who are the source language producers?
[More Information Needed]
### Annotations
#### Annotation process
[More Information Needed]
#### Who are the annotators?
[More Information Needed]
### Personal and Sensitive Information
[More Information Needed]
## Considerations for Using the Data
### Social Impact of Dataset
[More Information Needed]
### Discussion of Biases
[More Information Needed]
### Other Known Limitations
[More Information Needed]
## Additional Information
### Dataset Curators
[More Information Needed]
### Licensing Information
[More Information Needed]
### Citation Information
[More Information Needed]
### Contributions
[More Information Needed] |
open-llm-leaderboard/details_TheBloke__gpt4-alpaca-lora-30b-HF | 2023-09-18T00:20:32.000Z | [
"region:us"
] | open-llm-leaderboard | null | null | null | 0 | 0 | ---
pretty_name: Evaluation run of TheBloke/gpt4-alpaca-lora-30b-HF
dataset_summary: "Dataset automatically created during the evaluation run of model\
\ [TheBloke/gpt4-alpaca-lora-30b-HF](https://huggingface.co/TheBloke/gpt4-alpaca-lora-30b-HF)\
\ on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).\n\
\nThe dataset is composed of 3 configuration, each one coresponding to one of the\
\ evaluated task.\n\nThe dataset has been created from 1 run(s). Each run can be\
\ found as a specific split in each configuration, the split being named using the\
\ timestamp of the run.The \"train\" split is always pointing to the latest results.\n\
\nAn additional configuration \"results\" store all the aggregated results of the\
\ run (and is used to compute and display the agregated metrics on the [Open LLM\
\ Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).\n\
\nTo load the details from a run, you can for instance do the following:\n```python\n\
from datasets import load_dataset\ndata = load_dataset(\"open-llm-leaderboard/details_TheBloke__gpt4-alpaca-lora-30b-HF\"\
,\n\t\"harness_winogrande_5\",\n\tsplit=\"train\")\n```\n\n## Latest results\n\n\
These are the [latest results from run 2023-09-18T00:20:21.073173](https://huggingface.co/datasets/open-llm-leaderboard/details_TheBloke__gpt4-alpaca-lora-30b-HF/blob/main/results_2023-09-18T00-20-21.073173.json)(note\
\ that their might be results for other tasks in the repos if successive evals didn't\
\ cover the same tasks. You find each in the results and the \"latest\" split for\
\ each eval):\n\n```python\n{\n \"all\": {\n \"em\": 0.0016778523489932886,\n\
\ \"em_stderr\": 0.00041913301788269584,\n \"f1\": 0.06442533557047006,\n\
\ \"f1_stderr\": 0.0013970563636897643,\n \"acc\": 0.47865750583572136,\n\
\ \"acc_stderr\": 0.0105907760769931\n },\n \"harness|drop|3\": {\n\
\ \"em\": 0.0016778523489932886,\n \"em_stderr\": 0.00041913301788269584,\n\
\ \"f1\": 0.06442533557047006,\n \"f1_stderr\": 0.0013970563636897643\n\
\ },\n \"harness|gsm8k|5\": {\n \"acc\": 0.155420773313116,\n \
\ \"acc_stderr\": 0.009979689409499152\n },\n \"harness|winogrande|5\":\
\ {\n \"acc\": 0.8018942383583267,\n \"acc_stderr\": 0.01120186274448705\n\
\ }\n}\n```"
repo_url: https://huggingface.co/TheBloke/gpt4-alpaca-lora-30b-HF
leaderboard_url: https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
point_of_contact: clementine@hf.co
configs:
- config_name: harness_drop_3
data_files:
- split: 2023_09_18T00_20_21.073173
path:
- '**/details_harness|drop|3_2023-09-18T00-20-21.073173.parquet'
- split: latest
path:
- '**/details_harness|drop|3_2023-09-18T00-20-21.073173.parquet'
- config_name: harness_gsm8k_5
data_files:
- split: 2023_09_18T00_20_21.073173
path:
- '**/details_harness|gsm8k|5_2023-09-18T00-20-21.073173.parquet'
- split: latest
path:
- '**/details_harness|gsm8k|5_2023-09-18T00-20-21.073173.parquet'
- config_name: harness_winogrande_5
data_files:
- split: 2023_09_18T00_20_21.073173
path:
- '**/details_harness|winogrande|5_2023-09-18T00-20-21.073173.parquet'
- split: latest
path:
- '**/details_harness|winogrande|5_2023-09-18T00-20-21.073173.parquet'
- config_name: results
data_files:
- split: 2023_09_18T00_20_21.073173
path:
- results_2023-09-18T00-20-21.073173.parquet
- split: latest
path:
- results_2023-09-18T00-20-21.073173.parquet
---
# Dataset Card for Evaluation run of TheBloke/gpt4-alpaca-lora-30b-HF
## Dataset Description
- **Homepage:**
- **Repository:** https://huggingface.co/TheBloke/gpt4-alpaca-lora-30b-HF
- **Paper:**
- **Leaderboard:** https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
- **Point of Contact:** clementine@hf.co
### Dataset Summary
Dataset automatically created during the evaluation run of model [TheBloke/gpt4-alpaca-lora-30b-HF](https://huggingface.co/TheBloke/gpt4-alpaca-lora-30b-HF) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).
The dataset is composed of 3 configuration, each one coresponding to one of the evaluated task.
The dataset has been created from 1 run(s). Each run can be found as a specific split in each configuration, the split being named using the timestamp of the run.The "train" split is always pointing to the latest results.
An additional configuration "results" store all the aggregated results of the run (and is used to compute and display the agregated metrics on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).
To load the details from a run, you can for instance do the following:
```python
from datasets import load_dataset
data = load_dataset("open-llm-leaderboard/details_TheBloke__gpt4-alpaca-lora-30b-HF",
"harness_winogrande_5",
split="train")
```
## Latest results
These are the [latest results from run 2023-09-18T00:20:21.073173](https://huggingface.co/datasets/open-llm-leaderboard/details_TheBloke__gpt4-alpaca-lora-30b-HF/blob/main/results_2023-09-18T00-20-21.073173.json)(note that their might be results for other tasks in the repos if successive evals didn't cover the same tasks. You find each in the results and the "latest" split for each eval):
```python
{
"all": {
"em": 0.0016778523489932886,
"em_stderr": 0.00041913301788269584,
"f1": 0.06442533557047006,
"f1_stderr": 0.0013970563636897643,
"acc": 0.47865750583572136,
"acc_stderr": 0.0105907760769931
},
"harness|drop|3": {
"em": 0.0016778523489932886,
"em_stderr": 0.00041913301788269584,
"f1": 0.06442533557047006,
"f1_stderr": 0.0013970563636897643
},
"harness|gsm8k|5": {
"acc": 0.155420773313116,
"acc_stderr": 0.009979689409499152
},
"harness|winogrande|5": {
"acc": 0.8018942383583267,
"acc_stderr": 0.01120186274448705
}
}
```
### Supported Tasks and Leaderboards
[More Information Needed]
### Languages
[More Information Needed]
## Dataset Structure
### Data Instances
[More Information Needed]
### Data Fields
[More Information Needed]
### Data Splits
[More Information Needed]
## Dataset Creation
### Curation Rationale
[More Information Needed]
### Source Data
#### Initial Data Collection and Normalization
[More Information Needed]
#### Who are the source language producers?
[More Information Needed]
### Annotations
#### Annotation process
[More Information Needed]
#### Who are the annotators?
[More Information Needed]
### Personal and Sensitive Information
[More Information Needed]
## Considerations for Using the Data
### Social Impact of Dataset
[More Information Needed]
### Discussion of Biases
[More Information Needed]
### Other Known Limitations
[More Information Needed]
## Additional Information
### Dataset Curators
[More Information Needed]
### Licensing Information
[More Information Needed]
### Citation Information
[More Information Needed]
### Contributions
[More Information Needed] |
MyRebRIc/mcig | 2023-09-22T10:50:48.000Z | [
"region:us"
] | MyRebRIc | null | null | null | 0 | 0 | Entry not found |
v2ray/r-chatgpt-general-dump | 2023-09-18T23:08:08.000Z | [
"task_categories:conversational",
"size_categories:100K<n<1M",
"language:en",
"license:mit",
"not-for-all-audiences",
"region:us"
] | v2ray | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- conversational
language:
- en
tags:
- not-for-all-audiences
size_categories:
- 100K<n<1M
---
# r/ChatGPT General Dump
From [r/ChatGPT Discord #general channel](https://discord.gg/aRpD4pCw33). |
JuanKO/custom_tokenizer_littlestories | 2023-09-18T00:35:48.000Z | [
"license:openrail",
"region:us"
] | JuanKO | null | null | null | 0 | 0 | ---
license: openrail
dataset_info:
features:
- name: token
dtype: string
- name: embedding
sequence: float64
splits:
- name: train
num_bytes: 28247902
num_examples: 3443
download_size: 22867616
dataset_size: 28247902
---
|
forgetting-machine/embedded_faqs_medicare | 2023-09-18T00:49:49.000Z | [
"region:us"
] | forgetting-machine | null | null | null | 0 | 0 | Entry not found |
CVasNLPExperiments/cv-as-nlp-vqa-example-flan-xxl | 2023-09-18T01:35:24.000Z | [
"region:us"
] | CVasNLPExperiments | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: id
dtype: int64
- name: image
dtype: image
- name: prompt
dtype: string
- name: true_label
sequence: string
- name: prediction
dtype: string
splits:
- name: test
num_bytes: 585371.0
num_examples: 10
download_size: 587773
dataset_size: 585371.0
---
# Dataset Card for "cv-as-nlp-vqa-example-flan-xxl"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
PhoenixStormJr/EVC-just-in-case | 2023-09-18T01:27:46.000Z | [
"license:mit",
"region:us"
] | PhoenixStormJr | null | null | null | 0 | 0 | ---
license: mit
---
DO NOT USE THIS PROJECT!!! This is only just in case the original is taken down, there will be a backup. The original is listed here: https://huggingface.co/lj1995/VoiceConversionWebUI. ONLY USE THIS IF IT'S DELETED!!! |
open-llm-leaderboard/details_Andron00e__YetAnother_Open-Llama-3B-LoRA | 2023-09-18T01:32:29.000Z | [
"region:us"
] | open-llm-leaderboard | null | null | null | 0 | 0 | ---
pretty_name: Evaluation run of Andron00e/YetAnother_Open-Llama-3B-LoRA
dataset_summary: "Dataset automatically created during the evaluation run of model\
\ [Andron00e/YetAnother_Open-Llama-3B-LoRA](https://huggingface.co/Andron00e/YetAnother_Open-Llama-3B-LoRA)\
\ on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).\n\
\nThe dataset is composed of 3 configuration, each one coresponding to one of the\
\ evaluated task.\n\nThe dataset has been created from 1 run(s). Each run can be\
\ found as a specific split in each configuration, the split being named using the\
\ timestamp of the run.The \"train\" split is always pointing to the latest results.\n\
\nAn additional configuration \"results\" store all the aggregated results of the\
\ run (and is used to compute and display the agregated metrics on the [Open LLM\
\ Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).\n\
\nTo load the details from a run, you can for instance do the following:\n```python\n\
from datasets import load_dataset\ndata = load_dataset(\"open-llm-leaderboard/details_Andron00e__YetAnother_Open-Llama-3B-LoRA\"\
,\n\t\"harness_winogrande_5\",\n\tsplit=\"train\")\n```\n\n## Latest results\n\n\
These are the [latest results from run 2023-09-18T01:32:17.416050](https://huggingface.co/datasets/open-llm-leaderboard/details_Andron00e__YetAnother_Open-Llama-3B-LoRA/blob/main/results_2023-09-18T01-32-17.416050.json)(note\
\ that their might be results for other tasks in the repos if successive evals didn't\
\ cover the same tasks. You find each in the results and the \"latest\" split for\
\ each eval):\n\n```python\n{\n \"all\": {\n \"em\": 0.0,\n \"\
em_stderr\": 0.0,\n \"f1\": 0.0004886744966442953,\n \"f1_stderr\"\
: 8.997703088731367e-05,\n \"acc\": 0.2569060773480663,\n \"acc_stderr\"\
: 0.007023561458220211\n },\n \"harness|drop|3\": {\n \"em\": 0.0,\n\
\ \"em_stderr\": 0.0,\n \"f1\": 0.0004886744966442953,\n \"\
f1_stderr\": 8.997703088731367e-05\n },\n \"harness|gsm8k|5\": {\n \
\ \"acc\": 0.0,\n \"acc_stderr\": 0.0\n },\n \"harness|winogrande|5\"\
: {\n \"acc\": 0.5138121546961326,\n \"acc_stderr\": 0.014047122916440422\n\
\ }\n}\n```"
repo_url: https://huggingface.co/Andron00e/YetAnother_Open-Llama-3B-LoRA
leaderboard_url: https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
point_of_contact: clementine@hf.co
configs:
- config_name: harness_drop_3
data_files:
- split: 2023_09_18T01_32_17.416050
path:
- '**/details_harness|drop|3_2023-09-18T01-32-17.416050.parquet'
- split: latest
path:
- '**/details_harness|drop|3_2023-09-18T01-32-17.416050.parquet'
- config_name: harness_gsm8k_5
data_files:
- split: 2023_09_18T01_32_17.416050
path:
- '**/details_harness|gsm8k|5_2023-09-18T01-32-17.416050.parquet'
- split: latest
path:
- '**/details_harness|gsm8k|5_2023-09-18T01-32-17.416050.parquet'
- config_name: harness_winogrande_5
data_files:
- split: 2023_09_18T01_32_17.416050
path:
- '**/details_harness|winogrande|5_2023-09-18T01-32-17.416050.parquet'
- split: latest
path:
- '**/details_harness|winogrande|5_2023-09-18T01-32-17.416050.parquet'
- config_name: results
data_files:
- split: 2023_09_18T01_32_17.416050
path:
- results_2023-09-18T01-32-17.416050.parquet
- split: latest
path:
- results_2023-09-18T01-32-17.416050.parquet
---
# Dataset Card for Evaluation run of Andron00e/YetAnother_Open-Llama-3B-LoRA
## Dataset Description
- **Homepage:**
- **Repository:** https://huggingface.co/Andron00e/YetAnother_Open-Llama-3B-LoRA
- **Paper:**
- **Leaderboard:** https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
- **Point of Contact:** clementine@hf.co
### Dataset Summary
Dataset automatically created during the evaluation run of model [Andron00e/YetAnother_Open-Llama-3B-LoRA](https://huggingface.co/Andron00e/YetAnother_Open-Llama-3B-LoRA) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).
The dataset is composed of 3 configuration, each one coresponding to one of the evaluated task.
The dataset has been created from 1 run(s). Each run can be found as a specific split in each configuration, the split being named using the timestamp of the run.The "train" split is always pointing to the latest results.
An additional configuration "results" store all the aggregated results of the run (and is used to compute and display the agregated metrics on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).
To load the details from a run, you can for instance do the following:
```python
from datasets import load_dataset
data = load_dataset("open-llm-leaderboard/details_Andron00e__YetAnother_Open-Llama-3B-LoRA",
"harness_winogrande_5",
split="train")
```
## Latest results
These are the [latest results from run 2023-09-18T01:32:17.416050](https://huggingface.co/datasets/open-llm-leaderboard/details_Andron00e__YetAnother_Open-Llama-3B-LoRA/blob/main/results_2023-09-18T01-32-17.416050.json)(note that their might be results for other tasks in the repos if successive evals didn't cover the same tasks. You find each in the results and the "latest" split for each eval):
```python
{
"all": {
"em": 0.0,
"em_stderr": 0.0,
"f1": 0.0004886744966442953,
"f1_stderr": 8.997703088731367e-05,
"acc": 0.2569060773480663,
"acc_stderr": 0.007023561458220211
},
"harness|drop|3": {
"em": 0.0,
"em_stderr": 0.0,
"f1": 0.0004886744966442953,
"f1_stderr": 8.997703088731367e-05
},
"harness|gsm8k|5": {
"acc": 0.0,
"acc_stderr": 0.0
},
"harness|winogrande|5": {
"acc": 0.5138121546961326,
"acc_stderr": 0.014047122916440422
}
}
```
### Supported Tasks and Leaderboards
[More Information Needed]
### Languages
[More Information Needed]
## Dataset Structure
### Data Instances
[More Information Needed]
### Data Fields
[More Information Needed]
### Data Splits
[More Information Needed]
## Dataset Creation
### Curation Rationale
[More Information Needed]
### Source Data
#### Initial Data Collection and Normalization
[More Information Needed]
#### Who are the source language producers?
[More Information Needed]
### Annotations
#### Annotation process
[More Information Needed]
#### Who are the annotators?
[More Information Needed]
### Personal and Sensitive Information
[More Information Needed]
## Considerations for Using the Data
### Social Impact of Dataset
[More Information Needed]
### Discussion of Biases
[More Information Needed]
### Other Known Limitations
[More Information Needed]
## Additional Information
### Dataset Curators
[More Information Needed]
### Licensing Information
[More Information Needed]
### Citation Information
[More Information Needed]
### Contributions
[More Information Needed] |
DeepSpiral/ggs | 2023-09-18T01:45:40.000Z | [
"region:us"
] | DeepSpiral | null | null | null | 0 | 0 | Entry not found |
DaisyStar004/covid-llama2-500 | 2023-09-18T01:55:37.000Z | [
"region:us"
] | DaisyStar004 | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: text
dtype: string
splits:
- name: train
num_bytes: 317407
num_examples: 500
download_size: 181582
dataset_size: 317407
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
---
# Dataset Card for "covid-llama2-500"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
freefaller14/incometax | 2023-09-18T02:44:48.000Z | [
"region:us"
] | freefaller14 | null | null | null | 0 | 0 | Entry not found |
bongo2112/harmonize-SDxl-Comic-Style-Output | 2023-09-18T04:25:40.000Z | [
"region:us"
] | bongo2112 | null | null | null | 0 | 0 | Entry not found |
marketspace/real | 2023-09-26T21:55:50.000Z | [
"task_categories:text-generation",
"task_categories:translation",
"size_categories:1K<n<10K",
"license:openrail",
"code",
"region:us"
] | marketspace | null | null | null | 0 | 0 | ---
license: openrail
task_categories:
- text-generation
- translation
tags:
- code
size_categories:
- 1K<n<10K
---
## Real Language Model
---
license: other
--- |
CVasNLPExperiments/cv-as-nlp-vision-example-flan-xxl | 2023-09-18T03:00:18.000Z | [
"region:us"
] | CVasNLPExperiments | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: id
dtype: int64
- name: image
dtype: image
- name: prompt
dtype: string
- name: true_label
dtype: string
- name: prediction
dtype: string
splits:
- name: train
num_bytes: 119377.0
num_examples: 10
download_size: 119894
dataset_size: 119377.0
---
# Dataset Card for "cv-as-nlp-vision-example-flan-xxl"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
open-llm-leaderboard/details_clibrain__Llama-2-13b-ft-instruct-es | 2023-09-18T03:06:58.000Z | [
"region:us"
] | open-llm-leaderboard | null | null | null | 0 | 0 | ---
pretty_name: Evaluation run of clibrain/Llama-2-13b-ft-instruct-es
dataset_summary: "Dataset automatically created during the evaluation run of model\
\ [clibrain/Llama-2-13b-ft-instruct-es](https://huggingface.co/clibrain/Llama-2-13b-ft-instruct-es)\
\ on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).\n\
\nThe dataset is composed of 3 configuration, each one coresponding to one of the\
\ evaluated task.\n\nThe dataset has been created from 1 run(s). Each run can be\
\ found as a specific split in each configuration, the split being named using the\
\ timestamp of the run.The \"train\" split is always pointing to the latest results.\n\
\nAn additional configuration \"results\" store all the aggregated results of the\
\ run (and is used to compute and display the agregated metrics on the [Open LLM\
\ Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).\n\
\nTo load the details from a run, you can for instance do the following:\n```python\n\
from datasets import load_dataset\ndata = load_dataset(\"open-llm-leaderboard/details_clibrain__Llama-2-13b-ft-instruct-es\"\
,\n\t\"harness_winogrande_5\",\n\tsplit=\"train\")\n```\n\n## Latest results\n\n\
These are the [latest results from run 2023-09-18T03:06:46.998156](https://huggingface.co/datasets/open-llm-leaderboard/details_clibrain__Llama-2-13b-ft-instruct-es/blob/main/results_2023-09-18T03-06-46.998156.json)(note\
\ that their might be results for other tasks in the repos if successive evals didn't\
\ cover the same tasks. You find each in the results and the \"latest\" split for\
\ each eval):\n\n```python\n{\n \"all\": {\n \"em\": 0.0024119127516778523,\n\
\ \"em_stderr\": 0.0005023380498893348,\n \"f1\": 0.0655463506711411,\n\
\ \"f1_stderr\": 0.0014039891922809947,\n \"acc\": 0.42168315309067345,\n\
\ \"acc_stderr\": 0.009875785691028784\n },\n \"harness|drop|3\": {\n\
\ \"em\": 0.0024119127516778523,\n \"em_stderr\": 0.0005023380498893348,\n\
\ \"f1\": 0.0655463506711411,\n \"f1_stderr\": 0.0014039891922809947\n\
\ },\n \"harness|gsm8k|5\": {\n \"acc\": 0.08567096285064443,\n \
\ \"acc_stderr\": 0.007709218855882782\n },\n \"harness|winogrande|5\"\
: {\n \"acc\": 0.7576953433307024,\n \"acc_stderr\": 0.012042352526174787\n\
\ }\n}\n```"
repo_url: https://huggingface.co/clibrain/Llama-2-13b-ft-instruct-es
leaderboard_url: https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
point_of_contact: clementine@hf.co
configs:
- config_name: harness_drop_3
data_files:
- split: 2023_09_18T03_06_46.998156
path:
- '**/details_harness|drop|3_2023-09-18T03-06-46.998156.parquet'
- split: latest
path:
- '**/details_harness|drop|3_2023-09-18T03-06-46.998156.parquet'
- config_name: harness_gsm8k_5
data_files:
- split: 2023_09_18T03_06_46.998156
path:
- '**/details_harness|gsm8k|5_2023-09-18T03-06-46.998156.parquet'
- split: latest
path:
- '**/details_harness|gsm8k|5_2023-09-18T03-06-46.998156.parquet'
- config_name: harness_winogrande_5
data_files:
- split: 2023_09_18T03_06_46.998156
path:
- '**/details_harness|winogrande|5_2023-09-18T03-06-46.998156.parquet'
- split: latest
path:
- '**/details_harness|winogrande|5_2023-09-18T03-06-46.998156.parquet'
- config_name: results
data_files:
- split: 2023_09_18T03_06_46.998156
path:
- results_2023-09-18T03-06-46.998156.parquet
- split: latest
path:
- results_2023-09-18T03-06-46.998156.parquet
---
# Dataset Card for Evaluation run of clibrain/Llama-2-13b-ft-instruct-es
## Dataset Description
- **Homepage:**
- **Repository:** https://huggingface.co/clibrain/Llama-2-13b-ft-instruct-es
- **Paper:**
- **Leaderboard:** https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard
- **Point of Contact:** clementine@hf.co
### Dataset Summary
Dataset automatically created during the evaluation run of model [clibrain/Llama-2-13b-ft-instruct-es](https://huggingface.co/clibrain/Llama-2-13b-ft-instruct-es) on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard).
The dataset is composed of 3 configuration, each one coresponding to one of the evaluated task.
The dataset has been created from 1 run(s). Each run can be found as a specific split in each configuration, the split being named using the timestamp of the run.The "train" split is always pointing to the latest results.
An additional configuration "results" store all the aggregated results of the run (and is used to compute and display the agregated metrics on the [Open LLM Leaderboard](https://huggingface.co/spaces/HuggingFaceH4/open_llm_leaderboard)).
To load the details from a run, you can for instance do the following:
```python
from datasets import load_dataset
data = load_dataset("open-llm-leaderboard/details_clibrain__Llama-2-13b-ft-instruct-es",
"harness_winogrande_5",
split="train")
```
## Latest results
These are the [latest results from run 2023-09-18T03:06:46.998156](https://huggingface.co/datasets/open-llm-leaderboard/details_clibrain__Llama-2-13b-ft-instruct-es/blob/main/results_2023-09-18T03-06-46.998156.json)(note that their might be results for other tasks in the repos if successive evals didn't cover the same tasks. You find each in the results and the "latest" split for each eval):
```python
{
"all": {
"em": 0.0024119127516778523,
"em_stderr": 0.0005023380498893348,
"f1": 0.0655463506711411,
"f1_stderr": 0.0014039891922809947,
"acc": 0.42168315309067345,
"acc_stderr": 0.009875785691028784
},
"harness|drop|3": {
"em": 0.0024119127516778523,
"em_stderr": 0.0005023380498893348,
"f1": 0.0655463506711411,
"f1_stderr": 0.0014039891922809947
},
"harness|gsm8k|5": {
"acc": 0.08567096285064443,
"acc_stderr": 0.007709218855882782
},
"harness|winogrande|5": {
"acc": 0.7576953433307024,
"acc_stderr": 0.012042352526174787
}
}
```
### Supported Tasks and Leaderboards
[More Information Needed]
### Languages
[More Information Needed]
## Dataset Structure
### Data Instances
[More Information Needed]
### Data Fields
[More Information Needed]
### Data Splits
[More Information Needed]
## Dataset Creation
### Curation Rationale
[More Information Needed]
### Source Data
#### Initial Data Collection and Normalization
[More Information Needed]
#### Who are the source language producers?
[More Information Needed]
### Annotations
#### Annotation process
[More Information Needed]
#### Who are the annotators?
[More Information Needed]
### Personal and Sensitive Information
[More Information Needed]
## Considerations for Using the Data
### Social Impact of Dataset
[More Information Needed]
### Discussion of Biases
[More Information Needed]
### Other Known Limitations
[More Information Needed]
## Additional Information
### Dataset Curators
[More Information Needed]
### Licensing Information
[More Information Needed]
### Citation Information
[More Information Needed]
### Contributions
[More Information Needed] |
andrewsiah/se_cooking_preference_sft | 2023-09-18T03:30:15.000Z | [
"region:us"
] | andrewsiah | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: instruction
dtype: string
- name: input
dtype: string
- name: output
dtype: string
splits:
- name: train
num_bytes: 11095448
num_examples: 7262
download_size: 6879361
dataset_size: 11095448
---
# Dataset Card for "se_cooking_preference_sft"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
linhqyy/data_aug_new | 2023-09-18T03:32:22.000Z | [
"region:us"
] | linhqyy | null | null | null | 0 | 0 | Entry not found |
dangvinh77/Dataminer | 2023-09-18T03:48:22.000Z | [
"region:us"
] | dangvinh77 | null | null | null | 0 | 0 | # Selenium-bot
use pip install -r requirements.txt
|
BangumiBase/goblinslayer | 2023-09-29T08:35:36.000Z | [
"size_categories:1K<n<10K",
"license:mit",
"art",
"region:us"
] | BangumiBase | null | null | null | 0 | 0 | ---
license: mit
tags:
- art
size_categories:
- 1K<n<10K
---
# Bangumi Image Base of Goblin Slayer
This is the image base of bangumi Goblin Slayer, we detected 45 characters, 2771 images in total. The full dataset is [here](all.zip).
**Please note that these image bases are not guaranteed to be 100% cleaned, they may be noisy actual.** If you intend to manually train models using this dataset, we recommend performing necessary preprocessing on the downloaded dataset to eliminate potential noisy samples (approximately 1% probability).
Here is the characters' preview:
| # | Images | Download | Preview 1 | Preview 2 | Preview 3 | Preview 4 | Preview 5 | Preview 6 | Preview 7 | Preview 8 |
|:------|---------:|:---------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|
| 0 | 103 | [Download](0/dataset.zip) |  |  |  |  |  |  |  |  |
| 1 | 686 | [Download](1/dataset.zip) |  |  |  |  |  |  |  |  |
| 2 | 41 | [Download](2/dataset.zip) |  |  |  |  |  |  |  |  |
| 3 | 29 | [Download](3/dataset.zip) |  |  |  |  |  |  |  |  |
| 4 | 22 | [Download](4/dataset.zip) |  |  |  |  |  |  |  |  |
| 5 | 11 | [Download](5/dataset.zip) |  |  |  |  |  |  |  |  |
| 6 | 38 | [Download](6/dataset.zip) |  |  |  |  |  |  |  |  |
| 7 | 33 | [Download](7/dataset.zip) |  |  |  |  |  |  |  |  |
| 8 | 89 | [Download](8/dataset.zip) |  |  |  |  |  |  |  |  |
| 9 | 38 | [Download](9/dataset.zip) |  |  |  |  |  |  |  |  |
| 10 | 47 | [Download](10/dataset.zip) |  |  |  |  |  |  |  |  |
| 11 | 41 | [Download](11/dataset.zip) |  |  |  |  |  |  |  |  |
| 12 | 21 | [Download](12/dataset.zip) |  |  |  |  |  |  |  |  |
| 13 | 74 | [Download](13/dataset.zip) |  |  |  |  |  |  |  |  |
| 14 | 13 | [Download](14/dataset.zip) |  |  |  |  |  |  |  |  |
| 15 | 30 | [Download](15/dataset.zip) |  |  |  |  |  |  |  |  |
| 16 | 398 | [Download](16/dataset.zip) |  |  |  |  |  |  |  |  |
| 17 | 46 | [Download](17/dataset.zip) |  |  |  |  |  |  |  |  |
| 18 | 131 | [Download](18/dataset.zip) |  |  |  |  |  |  |  |  |
| 19 | 56 | [Download](19/dataset.zip) |  |  |  |  |  |  |  |  |
| 20 | 11 | [Download](20/dataset.zip) |  |  |  |  |  |  |  |  |
| 21 | 36 | [Download](21/dataset.zip) |  |  |  |  |  |  |  |  |
| 22 | 17 | [Download](22/dataset.zip) |  |  |  |  |  |  |  |  |
| 23 | 17 | [Download](23/dataset.zip) |  |  |  |  |  |  |  |  |
| 24 | 22 | [Download](24/dataset.zip) |  |  |  |  |  |  |  |  |
| 25 | 75 | [Download](25/dataset.zip) |  |  |  |  |  |  |  |  |
| 26 | 27 | [Download](26/dataset.zip) |  |  |  |  |  |  |  |  |
| 27 | 7 | [Download](27/dataset.zip) |  |  |  |  |  |  |  | N/A |
| 28 | 18 | [Download](28/dataset.zip) |  |  |  |  |  |  |  |  |
| 29 | 21 | [Download](29/dataset.zip) |  |  |  |  |  |  |  |  |
| 30 | 20 | [Download](30/dataset.zip) |  |  |  |  |  |  |  |  |
| 31 | 6 | [Download](31/dataset.zip) |  |  |  |  |  |  | N/A | N/A |
| 32 | 135 | [Download](32/dataset.zip) |  |  |  |  |  |  |  |  |
| 33 | 58 | [Download](33/dataset.zip) |  |  |  |  |  |  |  |  |
| 34 | 23 | [Download](34/dataset.zip) |  |  |  |  |  |  |  |  |
| 35 | 12 | [Download](35/dataset.zip) |  |  |  |  |  |  |  |  |
| 36 | 9 | [Download](36/dataset.zip) |  |  |  |  |  |  |  |  |
| 37 | 8 | [Download](37/dataset.zip) |  |  |  |  |  |  |  |  |
| 38 | 12 | [Download](38/dataset.zip) |  |  |  |  |  |  |  |  |
| 39 | 6 | [Download](39/dataset.zip) |  |  |  |  |  |  | N/A | N/A |
| 40 | 9 | [Download](40/dataset.zip) |  |  |  |  |  |  |  |  |
| 41 | 7 | [Download](41/dataset.zip) |  |  |  |  |  |  |  | N/A |
| 42 | 11 | [Download](42/dataset.zip) |  |  |  |  |  |  |  |  |
| 43 | 58 | [Download](43/dataset.zip) |  |  |  |  |  |  |  |  |
| noise | 199 | [Download](-1/dataset.zip) |  |  |  |  |  |  |  |  |
|
andrewsiah/anthropic_hh_sft | 2023-10-06T04:23:06.000Z | [
"region:us"
] | andrewsiah | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: instruction
dtype: string
- name: input
dtype: string
- name: output
dtype: string
splits:
- name: train
num_bytes: 13042104
num_examples: 20159
download_size: 7382066
dataset_size: 13042104
---
# Dataset Card for "anthropic_hh_sft"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
bongo2112/diamondplatnumz-SDxl-Comic-Style-Output | 2023-09-19T11:07:13.000Z | [
"region:us"
] | bongo2112 | null | null | null | 0 | 0 | Entry not found |
legacy107/qa_wikipedia_no_article | 2023-09-18T04:38:55.000Z | [
"region:us"
] | legacy107 | null | null | null | 0 | 0 | ---
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
- split: test
path: data/test-*
- split: validation
path: data/validation-*
dataset_info:
features:
- name: id
dtype: string
- name: title
dtype: string
- name: context
dtype: string
- name: question
dtype: string
- name: answer_start
dtype: int64
- name: answer
dtype: string
splits:
- name: train
num_bytes: 74337363
num_examples: 138712
- name: test
num_bytes: 9222514
num_examples: 17341
- name: validation
num_bytes: 9271740
num_examples: 17291
download_size: 25137600
dataset_size: 92831617
---
# Dataset Card for "qa_wikipedia_no_article"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
badbadal2000/Bgmiresults | 2023-09-18T04:39:03.000Z | [
"region:us"
] | badbadal2000 | null | null | null | 0 | 0 | Entry not found |
ravikant22/Test | 2023-09-18T05:02:24.000Z | [
"region:us"
] | ravikant22 | null | null | null | 0 | 0 | Entry not found |
eara4vu/zsxbtag | 2023-09-22T04:32:57.000Z | [
"region:us"
] | eara4vu | null | null | null | 0 | 0 | ***This branch is unmaintained!***
https://github.com/toriato/stable-diffusion-webui-wd14-tagger/issues/108
---
https://github.com/picobyte/stable-diffusion-webui-wd14-tagger
# Tagger for [Automatic1111's WebUI](https://github.com/AUTOMATIC1111/stable-diffusion-webui)
Interrogate booru style tags for single or multiple image files using various models, such as DeepDanbooru.
[한국어를 사용하시나요? 여기에 한국어 설명서가 있습니다!](README.ko.md)
## Disclaimer
I didn't make any models, and most of the code was heavily borrowed from the [DeepDanbooru](https://github.com/KichangKim/DeepDanbooru) and MrSmillingWolf's tagger.
## Installation
1. *Extensions* -> *Install from URL* -> Enter URL of this repository -> Press *Install* button
- or clone this repository under `extensions/`
```sh
$ git clone https://github.com/toriato/stable-diffusion-webui-wd14-tagger.git extensions/tagger
```
1. *(optional)* Add interrogate model
- #### [*Waifu Diffusion 1.4 Tagger by MrSmilingWolf*](docs/what-is-wd14-tagger.md)
Downloads automatically from the [HuggingFace repository](https://huggingface.co/SmilingWolf/wd-v1-4-vit-tagger) the first time you run it.
- #### *DeepDanbooru*
1. Various model files can be found below.
- [DeepDanbooru models](https://github.com/KichangKim/DeepDanbooru/releases)
- [e621 model by 🐾Zack🐾#1984](https://discord.gg/BDFpq9Yb7K)
*(link contains NSFW contents!)*
1. Move the project folder containing the model and config to `models/deepdanbooru`
1. The file structure should look like:
```
models/
└╴deepdanbooru/
├╴deepdanbooru-v3-20211112-sgd-e28/
│ ├╴project.json
│ └╴...
│
├╴deepdanbooru-v4-20200814-sgd-e30/
│ ├╴project.json
│ └╴...
│
├╴e621-v3-20221117-sgd-e32/
│ ├╴project.json
│ └╴...
│
...
```
1. Start or restart the WebUI.
- or you can press refresh button after *Interrogator* dropdown box.
- "You must close stable diffusion completely after installation and re-run it!"
## Model comparison
[Model comparison](docs/model-comparison.md)
## Screenshot

Artwork made by [hecattaart](https://vk.com/hecattaart?w=wall-89063929_3767)
## Copyright
Public domain, except borrowed parts (e.g. `dbimutils.py`)
|
dune10991/Ladozkie | 2023-09-18T05:45:15.000Z | [
"region:us"
] | dune10991 | null | null | null | 0 | 0 | Entry not found |
bongo2112/harmonize-SDxl-Many-Comic-Style-Output | 2023-09-18T12:51:21.000Z | [
"region:us"
] | bongo2112 | null | null | null | 0 | 0 | Entry not found |
Harish12/Llama-70b | 2023-09-18T06:16:57.000Z | [
"region:us"
] | Harish12 | null | null | null | 0 | 0 | Entry not found |
CyberHarem/miyamori_aoi_shirobako | 2023-09-18T06:52:19.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Miyamori Aoi
This is the dataset of Miyamori Aoi, containing 300 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 300 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 646 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 300 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 300 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 300 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 300 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 300 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 646 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 646 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 646 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
dune10991/Ladoga | 2023-09-18T15:25:55.000Z | [
"region:us"
] | dune10991 | null | null | null | 0 | 0 | ---
pretty_name: ladoga
--- |
CyberHarem/yasuhara_ema_shirobako | 2023-09-18T07:13:31.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Yasuhara Ema
This is the dataset of Yasuhara Ema, containing 300 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 300 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 639 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 300 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 300 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 300 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 300 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 300 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 639 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 639 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 639 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
linhqyy/data_aug_syllable | 2023-09-18T07:19:17.000Z | [
"region:us"
] | linhqyy | null | null | null | 0 | 0 | ---
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
- split: test
path: data/test-*
dataset_info:
features:
- name: sentence
dtype: string
- name: intent
dtype: string
- name: entities
list:
- name: type
dtype: string
- name: filler
dtype: string
- name: labels
dtype: string
splits:
- name: train
num_bytes: 2380518
num_examples: 11340
- name: test
num_bytes: 125890
num_examples: 597
download_size: 579180
dataset_size: 2506408
---
# Dataset Card for "data_aug_syllable"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
Admin08077/_U_ | 2023-09-18T07:24:20.000Z | [
"license:other",
"doi:10.57967/hf/1122",
"region:us"
] | Admin08077 | null | null | null | 0 | 0 | ---
license: other
---
|
CyberHarem/sakaki_shizuka_shirobako | 2023-09-18T07:27:33.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Sakaki Shizuka
This is the dataset of Sakaki Shizuka, containing 237 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 237 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 522 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 237 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 237 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 237 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 237 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 237 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 522 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 522 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 522 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
psm151/wav2vec2-large-xlsr-turkish-demo-colab | 2023-09-18T07:27:42.000Z | [
"license:openrail",
"region:us"
] | psm151 | null | null | null | 0 | 0 | ---
license: openrail
---
|
BangumiBase/paripikoumei | 2023-09-29T08:42:31.000Z | [
"size_categories:1K<n<10K",
"license:mit",
"art",
"region:us"
] | BangumiBase | null | null | null | 0 | 0 | ---
license: mit
tags:
- art
size_categories:
- 1K<n<10K
---
# Bangumi Image Base of Paripi Koumei
This is the image base of bangumi Paripi Koumei, we detected 33 characters, 2237 images in total. The full dataset is [here](all.zip).
**Please note that these image bases are not guaranteed to be 100% cleaned, they may be noisy actual.** If you intend to manually train models using this dataset, we recommend performing necessary preprocessing on the downloaded dataset to eliminate potential noisy samples (approximately 1% probability).
Here is the characters' preview:
| # | Images | Download | Preview 1 | Preview 2 | Preview 3 | Preview 4 | Preview 5 | Preview 6 | Preview 7 | Preview 8 |
|:------|---------:|:---------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|
| 0 | 34 | [Download](0/dataset.zip) |  |  |  |  |  |  |  |  |
| 1 | 344 | [Download](1/dataset.zip) |  |  |  |  |  |  |  |  |
| 2 | 65 | [Download](2/dataset.zip) |  |  |  |  |  |  |  |  |
| 3 | 42 | [Download](3/dataset.zip) |  |  |  |  |  |  |  |  |
| 4 | 34 | [Download](4/dataset.zip) |  |  |  |  |  |  |  |  |
| 5 | 36 | [Download](5/dataset.zip) |  |  |  |  |  |  |  |  |
| 6 | 68 | [Download](6/dataset.zip) |  |  |  |  |  |  |  |  |
| 7 | 201 | [Download](7/dataset.zip) |  |  |  |  |  |  |  |  |
| 8 | 19 | [Download](8/dataset.zip) |  |  |  |  |  |  |  |  |
| 9 | 235 | [Download](9/dataset.zip) |  |  |  |  |  |  |  |  |
| 10 | 32 | [Download](10/dataset.zip) |  |  |  |  |  |  |  |  |
| 11 | 14 | [Download](11/dataset.zip) |  |  |  |  |  |  |  |  |
| 12 | 15 | [Download](12/dataset.zip) |  |  |  |  |  |  |  |  |
| 13 | 41 | [Download](13/dataset.zip) |  |  |  |  |  |  |  |  |
| 14 | 10 | [Download](14/dataset.zip) |  |  |  |  |  |  |  |  |
| 15 | 32 | [Download](15/dataset.zip) |  |  |  |  |  |  |  |  |
| 16 | 19 | [Download](16/dataset.zip) |  |  |  |  |  |  |  |  |
| 17 | 8 | [Download](17/dataset.zip) |  |  |  |  |  |  |  |  |
| 18 | 12 | [Download](18/dataset.zip) |  |  |  |  |  |  |  |  |
| 19 | 102 | [Download](19/dataset.zip) |  |  |  |  |  |  |  |  |
| 20 | 451 | [Download](20/dataset.zip) |  |  |  |  |  |  |  |  |
| 21 | 11 | [Download](21/dataset.zip) |  |  |  |  |  |  |  |  |
| 22 | 121 | [Download](22/dataset.zip) |  |  |  |  |  |  |  |  |
| 23 | 48 | [Download](23/dataset.zip) |  |  |  |  |  |  |  |  |
| 24 | 6 | [Download](24/dataset.zip) |  |  |  |  |  |  | N/A | N/A |
| 25 | 8 | [Download](25/dataset.zip) |  |  |  |  |  |  |  |  |
| 26 | 7 | [Download](26/dataset.zip) |  |  |  |  |  |  |  | N/A |
| 27 | 6 | [Download](27/dataset.zip) |  |  |  |  |  |  | N/A | N/A |
| 28 | 36 | [Download](28/dataset.zip) |  |  |  |  |  |  |  |  |
| 29 | 17 | [Download](29/dataset.zip) |  |  |  |  |  |  |  |  |
| 30 | 18 | [Download](30/dataset.zip) |  |  |  |  |  |  |  |  |
| 31 | 9 | [Download](31/dataset.zip) |  |  |  |  |  |  |  |  |
| noise | 136 | [Download](-1/dataset.zip) |  |  |  |  |  |  |  |  |
|
CyberHarem/todo_misa_shirobako | 2023-09-18T07:40:04.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Tōdō Misa
This is the dataset of Tōdō Misa, containing 184 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 184 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 389 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 184 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 184 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 184 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 184 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 184 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 389 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 389 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 389 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
supramantest/hs_peer_support_chem | 2023-09-18T07:39:23.000Z | [
"region:us"
] | supramantest | null | null | null | 0 | 0 | ---
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
- split: test
path: data/test-*
- split: validation
path: data/validation-*
dataset_info:
features:
- name: text
dtype: string
splits:
- name: train
num_bytes: 2517979
num_examples: 4000
- name: test
num_bytes: 1748526
num_examples: 3000
- name: validation
num_bytes: 1736435
num_examples: 3000
download_size: 2214089
dataset_size: 6002940
---
# Dataset Card for "hs_peer_support_chem"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
Gautam18/mySQuAD | 2023-09-18T07:49:42.000Z | [
"region:us"
] | Gautam18 | null | null | null | 0 | 0 | Entry not found |
bongo2112/mooDewji-SDxl-Comic-Style-Outputs | 2023-09-18T09:24:44.000Z | [
"region:us"
] | bongo2112 | null | null | null | 0 | 0 | Entry not found |
CyberHarem/imai_midori_shirobako | 2023-09-18T07:56:37.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Imai Midori
This is the dataset of Imai Midori, containing 276 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 276 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 611 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 276 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 276 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 276 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 276 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 276 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 611 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 611 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 611 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
EleutherAI/bigbench | 2023-09-18T07:57:49.000Z | [
"region:us"
] | EleutherAI | null | null | null | 0 | 0 | Entry not found |
KnutJaegersberg/trilobite | 2023-09-18T09:26:51.000Z | [
"license:cc-by-nc-4.0",
"region:us"
] | KnutJaegersberg | null | null | null | 1 | 0 | ---
license: cc-by-nc-4.0
---
|
Cyclcrclicly/files | 2023-09-18T08:07:14.000Z | [
"region:us"
] | Cyclcrclicly | null | null | null | 0 | 0 | Entry not found |
CyberHarem/yano_erika_shirobako | 2023-09-18T08:16:57.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Yano Erika
This is the dataset of Yano Erika, containing 266 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 266 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 580 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 266 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 266 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 266 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 266 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 266 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 580 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 580 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 580 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
CyberHarem/tsubaki_ando_shirobako | 2023-09-18T08:21:10.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Tsubaki Ando
This is the dataset of Tsubaki Ando, containing 89 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 89 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 198 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 89 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 89 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 89 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 89 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 89 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 198 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 198 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 198 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
samyakmohelay/genai_dataset | 2023-09-18T08:45:28.000Z | [
"task_categories:summarization",
"task_ids:news-articles-summarization",
"annotations_creators:no-annotation",
"language_creators:found",
"multilinguality:monolingual",
"size_categories:100K<n<1M",
"source_datasets:original",
"language:en",
"license:apache-2.0",
"region:us"
] | samyakmohelay | null | null | null | 0 | 0 | ---
annotations_creators:
- no-annotation
language_creators:
- found
language:
- en
license:
- apache-2.0
multilinguality:
- monolingual
size_categories:
- 100K<n<1M
source_datasets:
- original
task_categories:
- summarization
task_ids:
- news-articles-summarization
paperswithcode_id: cnn-daily-mail-1
pretty_name: CNN / Daily Mail
train-eval-index:
- config: 3.0.0
task: summarization
task_id: summarization
splits:
eval_split: test
col_mapping:
article: text
highlights: target
dataset_info:
- config_name: 3.0.0
features:
- name: article
dtype: string
- name: highlights
dtype: string
- name: id
dtype: string
splits:
- name: train
num_bytes: 1261704133
num_examples: 287113
- name: validation
num_bytes: 57732436
num_examples: 13368
- name: test
num_bytes: 49925756
num_examples: 11490
download_size: 585439472
dataset_size: 1369362325
- config_name: 1.0.0
features:
- name: article
dtype: string
- name: highlights
dtype: string
- name: id
dtype: string
splits:
- name: train
num_bytes: 1261704133
num_examples: 287113
- name: validation
num_bytes: 57732436
num_examples: 13368
- name: test
num_bytes: 49925756
num_examples: 11490
download_size: 585439472
dataset_size: 1369362325
- config_name: 2.0.0
features:
- name: article
dtype: string
- name: highlights
dtype: string
- name: id
dtype: string
splits:
- name: train
num_bytes: 1261704133
num_examples: 287113
- name: validation
num_bytes: 57732436
num_examples: 13368
- name: test
num_bytes: 49925756
num_examples: 11490
download_size: 585439472
dataset_size: 1369362325
---
# Dataset Card for CNN Dailymail Dataset
## Table of Contents
- [Dataset Description](#dataset-description)
- [Dataset Summary](#dataset-summary)
- [Supported Tasks and Leaderboards](#supported-tasks-and-leaderboards)
- [Languages](#languages)
- [Dataset Structure](#dataset-structure)
- [Data Instances](#data-instances)
- [Data Fields](#data-fields)
- [Data Splits](#data-splits)
- [Dataset Creation](#dataset-creation)
- [Curation Rationale](#curation-rationale)
- [Source Data](#source-data)
- [Annotations](#annotations)
- [Personal and Sensitive Information](#personal-and-sensitive-information)
- [Considerations for Using the Data](#considerations-for-using-the-data)
- [Social Impact of Dataset](#social-impact-of-dataset)
- [Discussion of Biases](#discussion-of-biases)
- [Other Known Limitations](#other-known-limitations)
- [Additional Information](#additional-information)
- [Dataset Curators](#dataset-curators)
- [Licensing Information](#licensing-information)
- [Citation Information](#citation-information)
- [Contributions](#contributions)
## Dataset Description
- **Homepage:**
- **Repository:** [CNN / DailyMail Dataset repository](https://github.com/abisee/cnn-dailymail)
- **Paper:** [Abstractive Text Summarization Using Sequence-to-Sequence RNNs and Beyond](https://papers.nips.cc/paper/5945-teaching-machines-to-read-and-comprehend.pdf), [Get To The Point: Summarization with Pointer-Generator Networks](https://www.aclweb.org/anthology/K16-1028.pdf)
- **Leaderboard:** [Papers with Code leaderboard for CNN / Dailymail Dataset](https://paperswithcode.com/sota/document-summarization-on-cnn-daily-mail)
- **Point of Contact:** [Abigail See](mailto:abisee@stanford.edu)
### Dataset Summary
The CNN / DailyMail Dataset is an English-language dataset containing just over 300k unique news articles as written by journalists at CNN and the Daily Mail. The current version supports both extractive and abstractive summarization, though the original version was created for machine reading and comprehension and abstractive question answering.
### Supported Tasks and Leaderboards
- 'summarization': [Versions 2.0.0 and 3.0.0 of the CNN / DailyMail Dataset](https://www.aclweb.org/anthology/K16-1028.pdf) can be used to train a model for abstractive and extractive summarization ([Version 1.0.0](https://papers.nips.cc/paper/5945-teaching-machines-to-read-and-comprehend.pdf) was developed for machine reading and comprehension and abstractive question answering). The model performance is measured by how high the output summary's [ROUGE](https://huggingface.co/metrics/rouge) score for a given article is when compared to the highlight as written by the original article author. [Zhong et al (2020)](https://www.aclweb.org/anthology/2020.acl-main.552.pdf) report a ROUGE-1 score of 44.41 when testing a model trained for extractive summarization. See the [Papers With Code leaderboard](https://paperswithcode.com/sota/document-summarization-on-cnn-daily-mail) for more models.
### Languages
The BCP-47 code for English as generally spoken in the United States is en-US and the BCP-47 code for English as generally spoken in the United Kingdom is en-GB. It is unknown if other varieties of English are represented in the data.
## Dataset Structure
### Data Instances
For each instance, there is a string for the article, a string for the highlights, and a string for the id. See the [CNN / Daily Mail dataset viewer](https://huggingface.co/datasets/viewer/?dataset=cnn_dailymail&config=3.0.0) to explore more examples.
```
{'id': '0054d6d30dbcad772e20b22771153a2a9cbeaf62',
'article': '(CNN) -- An American woman died aboard a cruise ship that docked at Rio de Janeiro on Tuesday, the same ship on which 86 passengers previously fell ill, according to the state-run Brazilian news agency, Agencia Brasil. The American tourist died aboard the MS Veendam, owned by cruise operator Holland America. Federal Police told Agencia Brasil that forensic doctors were investigating her death. The ship's doctors told police that the woman was elderly and suffered from diabetes and hypertension, according the agency. The other passengers came down with diarrhea prior to her death during an earlier part of the trip, the ship's doctors said. The Veendam left New York 36 days ago for a South America tour.'
'highlights': 'The elderly woman suffered from diabetes and hypertension, ship's doctors say .\nPreviously, 86 passengers had fallen ill on the ship, Agencia Brasil says .'}
```
The average token count for the articles and the highlights are provided below:
| Feature | Mean Token Count |
| ---------- | ---------------- |
| Article | 781 |
| Highlights | 56 |
### Data Fields
- `id`: a string containing the heximal formated SHA1 hash of the url where the story was retrieved from
- `article`: a string containing the body of the news article
- `highlights`: a string containing the highlight of the article as written by the article author
### Data Splits
The CNN/DailyMail dataset has 3 splits: _train_, _validation_, and _test_. Below are the statistics for Version 3.0.0 of the dataset.
| Dataset Split | Number of Instances in Split |
| ------------- | ------------------------------------------- |
| Train | 287,113 |
| Validation | 13,368 |
| Test | 11,490 |
## Dataset Creation
### Curation Rationale
Version 1.0.0 aimed to support supervised neural methodologies for machine reading and question answering with a large amount of real natural language training data and released about 313k unique articles and nearly 1M Cloze style questions to go with the articles. Versions 2.0.0 and 3.0.0 changed the structure of the dataset to support summarization rather than question answering. Version 3.0.0 provided a non-anonymized version of the data, whereas both the previous versions were preprocessed to replace named entities with unique identifier labels.
### Source Data
#### Initial Data Collection and Normalization
The data consists of news articles and highlight sentences. In the question answering setting of the data, the articles are used as the context and entities are hidden one at a time in the highlight sentences, producing Cloze style questions where the goal of the model is to correctly guess which entity in the context has been hidden in the highlight. In the summarization setting, the highlight sentences are concatenated to form a summary of the article. The CNN articles were written between April 2007 and April 2015. The Daily Mail articles were written between June 2010 and April 2015.
The code for the original data collection is available at <https://github.com/deepmind/rc-data>. The articles were downloaded using archives of <www.cnn.com> and <www.dailymail.co.uk> on the Wayback Machine. Articles were not included in the Version 1.0.0 collection if they exceeded 2000 tokens. Due to accessibility issues with the Wayback Machine, Kyunghyun Cho has made the datasets available at <https://cs.nyu.edu/~kcho/DMQA/>. An updated version of the code that does not anonymize the data is available at <https://github.com/abisee/cnn-dailymail>.
Hermann et al provided their own tokenization script. The script provided by See uses the PTBTokenizer. It also lowercases the text and adds periods to lines missing them.
#### Who are the source language producers?
The text was written by journalists at CNN and the Daily Mail.
### Annotations
The dataset does not contain any additional annotations.
#### Annotation process
[N/A]
#### Who are the annotators?
[N/A]
### Personal and Sensitive Information
Version 3.0 is not anonymized, so individuals' names can be found in the dataset. Information about the original author is not included in the dataset.
## Considerations for Using the Data
### Social Impact of Dataset
The purpose of this dataset is to help develop models that can summarize long paragraphs of text in one or two sentences.
This task is useful for efficiently presenting information given a large quantity of text. It should be made clear that any summarizations produced by models trained on this dataset are reflective of the language used in the articles, but are in fact automatically generated.
### Discussion of Biases
[Bordia and Bowman (2019)](https://www.aclweb.org/anthology/N19-3002.pdf) explore measuring gender bias and debiasing techniques in the CNN / Dailymail dataset, the Penn Treebank, and WikiText-2. They find the CNN / Dailymail dataset to have a slightly lower gender bias based on their metric compared to the other datasets, but still show evidence of gender bias when looking at words such as 'fragile'.
Because the articles were written by and for people in the US and the UK, they will likely present specifically US and UK perspectives and feature events that are considered relevant to those populations during the time that the articles were published.
### Other Known Limitations
News articles have been shown to conform to writing conventions in which important information is primarily presented in the first third of the article [(Kryściński et al, 2019)](https://www.aclweb.org/anthology/D19-1051.pdf). [Chen et al (2016)](https://www.aclweb.org/anthology/P16-1223.pdf) conducted a manual study of 100 random instances of the first version of the dataset and found 25% of the samples to be difficult even for humans to answer correctly due to ambiguity and coreference errors.
It should also be noted that machine-generated summarizations, even when extractive, may differ in truth values when compared to the original articles.
## Additional Information
### Dataset Curators
The data was originally collected by Karl Moritz Hermann, Tomáš Kočiský, Edward Grefenstette, Lasse Espeholt, Will Kay, Mustafa Suleyman, and Phil Blunsom of Google DeepMind. Tomáš Kočiský and Phil Blunsom are also affiliated with the University of Oxford. They released scripts to collect and process the data into the question answering format.
Ramesh Nallapati, Bowen Zhou, Cicero dos Santos, and Bing Xiang of IMB Watson and Çağlar Gu̇lçehre of Université de Montréal modified Hermann et al's collection scripts to restore the data to a summary format. They also produced both anonymized and non-anonymized versions.
The code for the non-anonymized version is made publicly available by Abigail See of Stanford University, Peter J. Liu of Google Brain and Christopher D. Manning of Stanford University at <https://github.com/abisee/cnn-dailymail>. The work at Stanford University was supported by the DARPA DEFT ProgramAFRL contract no. FA8750-13-2-0040.
### Licensing Information
The CNN / Daily Mail dataset version 1.0.0 is released under the [Apache-2.0 License](http://www.apache.org/licenses/LICENSE-2.0).
### Citation Information
```
@inproceedings{see-etal-2017-get,
title = "Get To The Point: Summarization with Pointer-Generator Networks",
author = "See, Abigail and
Liu, Peter J. and
Manning, Christopher D.",
booktitle = "Proceedings of the 55th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers)",
month = jul,
year = "2017",
address = "Vancouver, Canada",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/P17-1099",
doi = "10.18653/v1/P17-1099",
pages = "1073--1083",
abstract = "Neural sequence-to-sequence models have provided a viable new approach for abstractive text summarization (meaning they are not restricted to simply selecting and rearranging passages from the original text). However, these models have two shortcomings: they are liable to reproduce factual details inaccurately, and they tend to repeat themselves. In this work we propose a novel architecture that augments the standard sequence-to-sequence attentional model in two orthogonal ways. First, we use a hybrid pointer-generator network that can copy words from the source text via pointing, which aids accurate reproduction of information, while retaining the ability to produce novel words through the generator. Second, we use coverage to keep track of what has been summarized, which discourages repetition. We apply our model to the CNN / Daily Mail summarization task, outperforming the current abstractive state-of-the-art by at least 2 ROUGE points.",
}
```
```
@inproceedings{DBLP:conf/nips/HermannKGEKSB15,
author={Karl Moritz Hermann and Tomás Kociský and Edward Grefenstette and Lasse Espeholt and Will Kay and Mustafa Suleyman and Phil Blunsom},
title={Teaching Machines to Read and Comprehend},
year={2015},
cdate={1420070400000},
pages={1693-1701},
url={http://papers.nips.cc/paper/5945-teaching-machines-to-read-and-comprehend},
booktitle={NIPS},
crossref={conf/nips/2015}
}
```
### Contributions
Thanks to [@thomwolf](https://github.com/thomwolf), [@lewtun](https://github.com/lewtun), [@jplu](https://github.com/jplu), [@jbragg](https://github.com/jbragg), [@patrickvonplaten](https://github.com/patrickvonplaten) and [@mcmillanmajora](https://github.com/mcmillanmajora) for adding this dataset. |
hungeni/mtet_dataset | 2023-09-18T09:00:31.000Z | [
"region:us"
] | hungeni | null | null | null | 0 | 0 |
Clone from https://github.com/vietai/mTet
---
license: cc-by-4.0
task_categories:
- translation
language:
- vi
license: cc-by-4.0
--- |
CyberHarem/priestess_goblinslayer | 2023-09-18T08:52:14.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Priestess
This is the dataset of Priestess, containing 300 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 300 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 653 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 300 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 300 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 300 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 300 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 300 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 653 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 653 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 653 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
dotan1111/MSA-amino-2-seq | 2023-09-18T11:46:50.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.
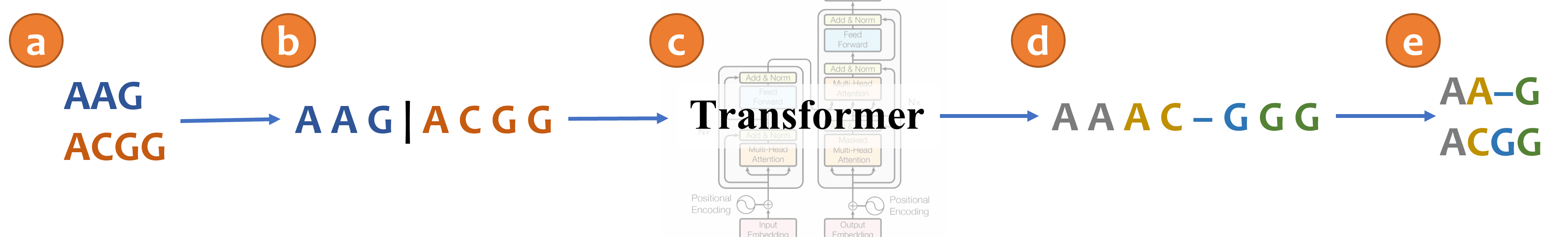
An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-amino-3-seq | 2023-09-18T11:46:25.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-amino-4-seq | 2023-09-18T11:47:10.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-amino-5-seq | 2023-09-18T11:47:27.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
hungeni/amrutaQA | 2023-09-20T11:47:41.000Z | [
"license:other",
"region:us"
] | hungeni | null | null | null | 0 | 0 | ---
license: other
---
This dataset crawl and re-organize from site: amruta.org, extracting questions and answers for fine-tuning tasks.
By the grace of H.H. Mother Shri Mataji Nirmala Devi |
dotan1111/MSA-amino-6-seq | 2023-09-18T11:47:37.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-amino-7-seq | 2023-09-18T11:47:47.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-amino-9-seq | 2023-09-18T11:48:30.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-amino-10-seq | 2023-09-18T11:48:44.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
yesenia09/comparison-data-mpt-small | 2023-09-18T08:55:55.000Z | [
"region:us"
] | yesenia09 | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: instruction
dtype: string
id: field
- name: answer_preferred
dtype: string
id: field
- name: answer_rejected
dtype: string
id: field
- name: choose-best
list:
- name: user_id
dtype: string
id: question
- name: value
dtype: int32
id: suggestion
- name: status
dtype: string
id: question
- name: choose-best-suggestion
dtype: int32
id: suggestion
- name: choose-best-suggestion-metadata
struct:
- name: type
dtype: string
id: suggestion-metadata
- name: score
dtype: float32
id: suggestion-metadata
- name: agent
dtype: string
id: suggestion-metadata
- name: external_id
dtype: string
id: external_id
- name: metadata
dtype: string
id: metadata
splits:
- name: train
num_bytes: 114617
num_examples: 147
download_size: 27484
dataset_size: 114617
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
---
# Dataset Card for "comparison-data-mpt-small"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
yesenia09/comparison-data-mpt | 2023-09-18T08:56:39.000Z | [
"region:us"
] | yesenia09 | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: instruction
dtype: string
id: field
- name: answer_preferred
dtype: string
id: field
- name: answer_rejected
dtype: string
id: field
- name: choose-best
list:
- name: user_id
dtype: string
id: question
- name: value
dtype: int32
id: suggestion
- name: status
dtype: string
id: question
- name: choose-best-suggestion
dtype: int32
id: suggestion
- name: choose-best-suggestion-metadata
struct:
- name: type
dtype: string
id: suggestion-metadata
- name: score
dtype: float32
id: suggestion-metadata
- name: agent
dtype: string
id: suggestion-metadata
- name: external_id
dtype: string
id: external_id
- name: metadata
dtype: string
id: metadata
splits:
- name: train
num_bytes: 114617
num_examples: 147
download_size: 27484
dataset_size: 114617
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
---
# Dataset Card for "comparison-data-mpt"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
CyberHarem/tsukimi_eiko_paripikoumei | 2023-09-18T09:07:03.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Tsukimi Eiko
This is the dataset of Tsukimi Eiko, containing 299 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 299 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 719 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 299 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 299 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 299 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 299 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 299 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 719 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 719 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 719 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
bvallegc/gtzan_all_preprocessed | 2023-09-18T09:09:33.000Z | [
"region:us"
] | bvallegc | null | null | null | 0 | 0 | ---
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
- split: test
path: data/test-*
dataset_info:
features:
- name: label
dtype:
class_label:
names:
'0': blues
'1': classical
'2': country
'3': disco
'4': hiphop
'5': jazz
'6': metal
'7': pop
'8': reggae
'9': rock
- name: input_values
sequence: float32
- name: attention_mask
sequence: int32
splits:
- name: train
num_bytes: 3452159816
num_examples: 899
- name: test
num_bytes: 384000696
num_examples: 100
download_size: 1923103923
dataset_size: 3836160512
---
# Dataset Card for "gtzan_all_preprocessed"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
haor/OpenMid-Dataset | 2023-09-18T10:46:50.000Z | [
"task_categories:text-to-image",
"size_categories:1M<n<10M",
"language:en",
"license:cc-by-nc-4.0",
"dataset",
"Nijijourney",
"Midjourney",
"doi:10.57967/hf/1123",
"region:us"
] | haor | null | null | null | 0 | 0 | ---
task_categories:
- text-to-image
language:
- en
tags:
- dataset
- Nijijourney
- Midjourney
size_categories:
- 1M<n<10M
license: cc-by-nc-4.0
---
Count:
- 2023-09-14 | 66371 | 2.97%
- 2023-09-15 | 595557 | 26.61%
- 2023-09-16 | 618586 | 27.64%
- 2023-09-17 | 566691 | 25.32%
- 2023-09-18 | 390878 | 17.46%
- Total items: 2238083 |
HugoGiddins/cleaned_ibmmq | 2023-09-18T09:13:13.000Z | [
"region:us"
] | HugoGiddins | null | null | null | 0 | 0 | Entry not found |
DigitalUmuganda/Monolingual_health_dataset | 2023-09-18T09:37:07.000Z | [
"size_categories:10K<n<100K",
"language:rw",
"language:en",
"license:cc-by-2.0",
"region:us"
] | DigitalUmuganda | null | null | null | 0 | 0 | ---
license: cc-by-2.0
language:
- rw
- en
size_categories:
- 10K<n<100K
---
# Monolingual Dataset
This a a malnutrition dataset in Kinyarwanda and English, it shall be translated using translators to make it a parallel corpus.
# Source of Data
1. Rwanda Biomedical Center (RBC) (26,390 sentences)
2. GPT-4 prompting (42,576 sentences) |
CyberHarem/high_elf_archer_goblinslayer | 2023-09-18T09:16:15.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of High Elf Archer
This is the dataset of High Elf Archer, containing 300 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 300 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 638 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 300 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 300 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 300 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 300 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 300 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 638 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 638 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 638 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
CyberHarem/kuon_nanami_paripikoumei | 2023-09-18T09:18:13.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Kuon Nanami
This is the dataset of Kuon Nanami, containing 153 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 153 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 358 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 153 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 153 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 153 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 153 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 153 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 358 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 358 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 358 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
CyberHarem/cow_girl_goblinslayer | 2023-09-18T09:25:25.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Cow Girl
This is the dataset of Cow Girl, containing 119 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 119 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 247 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 119 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 119 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 119 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 119 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 119 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 247 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 247 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 247 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
CyberHarem/guild_girl_goblinslayer | 2023-09-18T09:36:36.000Z | [
"task_categories:text-to-image",
"size_categories:n<1K",
"license:mit",
"art",
"not-for-all-audiences",
"region:us"
] | CyberHarem | null | null | null | 0 | 0 | ---
license: mit
task_categories:
- text-to-image
tags:
- art
- not-for-all-audiences
size_categories:
- n<1K
---
# Dataset of Guild Girl
This is the dataset of Guild Girl, containing 96 images and their tags.
Images are crawled from many sites (e.g. danbooru, pixiv, zerochan ...), the auto-crawling system is powered by [DeepGHS Team](https://github.com/deepghs)([huggingface organization](https://huggingface.co/deepghs)).
| Name | Images | Download | Description |
|:------------|---------:|:------------------------------------|:-------------------------------------------------------------------------|
| raw | 96 | [Download](dataset-raw.zip) | Raw data with meta information. |
| raw-stage3 | 221 | [Download](dataset-raw-stage3.zip) | 3-stage cropped raw data with meta information. |
| 384x512 | 96 | [Download](dataset-384x512.zip) | 384x512 aligned dataset. |
| 512x512 | 96 | [Download](dataset-512x512.zip) | 512x512 aligned dataset. |
| 512x704 | 96 | [Download](dataset-512x704.zip) | 512x704 aligned dataset. |
| 640x640 | 96 | [Download](dataset-640x640.zip) | 640x640 aligned dataset. |
| 640x880 | 96 | [Download](dataset-640x880.zip) | 640x880 aligned dataset. |
| stage3-640 | 221 | [Download](dataset-stage3-640.zip) | 3-stage cropped dataset with the shorter side not exceeding 640 pixels. |
| stage3-800 | 221 | [Download](dataset-stage3-800.zip) | 3-stage cropped dataset with the shorter side not exceeding 800 pixels. |
| stage3-1200 | 221 | [Download](dataset-stage3-1200.zip) | 3-stage cropped dataset with the shorter side not exceeding 1200 pixels. |
|
links-ads/grapevista-dataset | 2023-09-18T11:01:07.000Z | [
"task_categories:image-segmentation",
"size_categories:10K<n<100K",
"license:cc-by-2.0",
"region:us"
] | links-ads | null | null | null | 0 | 0 | ---
license: cc-by-2.0
pretty_name: GRAPEVISTA
size_categories:
- 10K<n<100K
task_categories:
- image-segmentation
---
# GRAPEVISTA - GRAPE Vineyard Imaging and Segmentation Technology Archive
## Description
This dataset contains two collections of high-resolution images captured in various vineyards called **VITIGEOSS** and **WGISD_Extension**. Each image is accompanied by either ground truth annotations or produced segmentation mask, providing valuable data for vineyard-related computer vision and machine learning tasks.
## Dataset Details
- **Source**:
- **VITIGEOSS**: These images were collected by infield cameras installed in 5 different vineyards across Italy, Spain and Portugal.
- **WGISD_Extension**: These images were collected during field visits to vineyards as mentioned in the [original work](https://github.com/thsant/wgisd).
- **Citation**: Please cite the dataset as follows:
``` latex
@inproceedings{blanco23automatic,
title={On the automatic detection and monitoring of Leaves and Grapes using in-field optical cameras},
author={Blanco, Giacomo and Oldani, Federico and Salza, Dario and Rossi, Claudio},
booktitle={2023 IEEE international workshop on metrology for agriculture and forestry (MetroAgriFor)},
year={2023},
organization={IEEE}
}
```
## Dataset Content
- **Number of Images**:
- **VITIGEOSS**: 4545
- **WGISD_Extension**: 8910 training + 1107 validation
- **File Format**: JPEG
- **Ground Truth Annotation Format**: PNG
## Data Fields/Columns
The two collections are provided with the following format:
- **VITIGEOSS**:
- `image_filename`: {CompanyCode}_{VineyardCode}_{CameraCode}_{Variety}_{YYYY-MM-DDTHH:MM:SS}.jpg
- `annotation_filename`: {CompanyCode}_{VineyardCode}_{CameraCode}_{Variety}_{YYYY-MM-DDTHH:MM:SS}.png
- **WGISD_Extension**:
- `image_filename`: {WGISDOriginalName}_{N}.jpg where N is the number of augmentation of the same image
- `annotation_filename`: {WGISDOriginalName}_{N}_labelTrainIds.jpg where N is the number of augmentation of the same image
## Ground Truth Annotation
For both collections semantic segmentation annotations are reported as images where each pixel indicates class among *background, leaves and grapes* for correspondent image
- **WGISD_Extension**: Ground truth annotations are obtained together with augmented images creation
- **VITIGEOSS**: Images are not provided with ground truth annotations but with the semgnation mask produced by the model developed in the aforemention work
## Dataset extraction
In order to extract GRAPEVISTA dataset for archive files, run the following commands
``` shell
cd data
cat grapevista.tar.*.gz.part > grapevista.tar.gz
tar -xvzf grapevista.tar.gz
```
## License Information
This dataset is provided under the CC BY-NC 2.0 license. See the [LICENSE](https://creativecommons.org/licenses/by-nc/2.0/) website for details. |
yicozy/dataset_pfs_by_arm | 2023-09-19T01:51:31.000Z | [
"region:us"
] | yicozy | null | null | null | 0 | 0 | ---
dataset_info:
features:
- name: instruction
dtype: string
- name: response
dtype: string
- name: text
dtype: string
splits:
- name: train
num_bytes: 1015503
num_examples: 1827
download_size: 0
dataset_size: 1015503
configs:
- config_name: default
data_files:
- split: train
path: data/train-*
---
# Dataset Card for "dataset_pfs_by_arm"
[More Information needed](https://github.com/huggingface/datasets/blob/main/CONTRIBUTING.md#how-to-contribute-to-the-dataset-cards) |
BangumiBase/bocchitherock | 2023-09-29T08:50:21.000Z | [
"size_categories:1K<n<10K",
"license:mit",
"art",
"region:us"
] | BangumiBase | null | null | null | 0 | 0 | ---
license: mit
tags:
- art
size_categories:
- 1K<n<10K
---
# Bangumi Image Base of Bocchi The Rock!
This is the image base of bangumi Bocchi the Rock!, we detected 23 characters, 2223 images in total. The full dataset is [here](all.zip).
**Please note that these image bases are not guaranteed to be 100% cleaned, they may be noisy actual.** If you intend to manually train models using this dataset, we recommend performing necessary preprocessing on the downloaded dataset to eliminate potential noisy samples (approximately 1% probability).
Here is the characters' preview:
| # | Images | Download | Preview 1 | Preview 2 | Preview 3 | Preview 4 | Preview 5 | Preview 6 | Preview 7 | Preview 8 |
|:------|---------:|:---------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|:-------------------------------|
| 0 | 538 | [Download](0/dataset.zip) |  |  |  |  |  |  |  |  |
| 1 | 54 | [Download](1/dataset.zip) |  |  |  |  |  |  |  |  |
| 2 | 35 | [Download](2/dataset.zip) |  |  |  |  |  |  |  |  |
| 3 | 13 | [Download](3/dataset.zip) |  |  |  |  |  |  |  |  |
| 4 | 286 | [Download](4/dataset.zip) |  |  |  |  |  |  |  |  |
| 5 | 108 | [Download](5/dataset.zip) |  |  |  |  |  |  |  |  |
| 6 | 8 | [Download](6/dataset.zip) |  |  |  |  |  |  |  |  |
| 7 | 88 | [Download](7/dataset.zip) |  |  |  |  |  |  |  |  |
| 8 | 14 | [Download](8/dataset.zip) |  |  |  |  |  |  |  |  |
| 9 | 439 | [Download](9/dataset.zip) |  |  |  |  |  |  |  |  |
| 10 | 66 | [Download](10/dataset.zip) |  |  |  |  |  |  |  |  |
| 11 | 8 | [Download](11/dataset.zip) |  |  |  |  |  |  |  |  |
| 12 | 257 | [Download](12/dataset.zip) |  |  |  |  |  |  |  |  |
| 13 | 29 | [Download](13/dataset.zip) |  |  |  |  |  |  |  |  |
| 14 | 6 | [Download](14/dataset.zip) |  |  |  |  |  |  | N/A | N/A |
| 15 | 9 | [Download](15/dataset.zip) |  |  |  |  |  |  |  |  |
| 16 | 14 | [Download](16/dataset.zip) |  |  |  |  |  |  |  |  |
| 17 | 6 | [Download](17/dataset.zip) |  |  |  |  |  |  | N/A | N/A |
| 18 | 14 | [Download](18/dataset.zip) |  |  |  |  |  |  |  |  |
| 19 | 12 | [Download](19/dataset.zip) |  |  |  |  |  |  |  |  |
| 20 | 13 | [Download](20/dataset.zip) |  |  |  |  |  |  |  |  |
| 21 | 9 | [Download](21/dataset.zip) |  |  |  |  |  |  |  |  |
| noise | 197 | [Download](-1/dataset.zip) |  |  |  |  |  |  |  |  |
|
josedanielaromi/Arg1996 | 2023-09-18T10:23:24.000Z | [
"region:us"
] | josedanielaromi | null | null | null | 0 | 0 | Entry not found |
dvrkdvys/SZA_Speaking | 2023-09-18T10:55:53.000Z | [
"license:openrail",
"region:us"
] | dvrkdvys | null | null | null | 0 | 0 | ---
license: openrail
---
|
ksh98/food_store | 2023-09-18T10:27:11.000Z | [
"license:other",
"region:us"
] | ksh98 | null | null | null | 0 | 0 | ---
license: other
---
|
dvrkdvys/SZA_Singing | 2023-09-18T10:28:58.000Z | [
"license:openrail",
"region:us"
] | dvrkdvys | null | null | null | 0 | 0 | ---
license: openrail
pretty_name: voice conversion
--- |
crumb/refinedweb-22mil-128clusters | 2023-09-18T16:28:08.000Z | [
"region:us"
] | crumb | null | null | null | 0 | 0 | Entry not found |
dotan1111/MSA-amino-8-seq | 2023-09-18T11:47:59.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-3-seq | 2023-09-18T11:49:24.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-2-seq | 2023-09-18T11:49:02.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-4-seq | 2023-09-18T11:49:37.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-5-seq | 2023-09-18T11:49:47.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-6-seq | 2023-09-18T11:49:55.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-7-seq | 2023-09-18T11:50:02.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-8-seq | 2023-09-18T11:50:10.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
dotan1111/MSA-nuc-9-seq | 2023-09-18T11:50:18.000Z | [
"sequence-to-sequence",
"bioinformatics",
"biology",
"region:us"
] | dotan1111 | null | null | null | 0 | 0 | ---
tags:
- sequence-to-sequence
- bioinformatics
- biology
---
# Multiple Sequence Alignment as a Sequence-to-Sequence Learning Problem
## Abstract:
The sequence alignment problem is one of the most fundamental problems in bioinformatics and a plethora of methods were devised to tackle it. Here we introduce BetaAlign, a methodology for aligning sequences using an NLP approach. BetaAlign accounts for the possible variability of the evolutionary process among different datasets by using an ensemble of transformers, each trained on millions of samples generated from a different evolutionary model. Our approach leads to alignment accuracy that is similar and often better than commonly used methods, such as MAFFT, DIALIGN, ClustalW, T-Coffee, PRANK, and MUSCLE.

An illustration of aligning sequences with sequence-to-sequence learning. (a) Consider two input sequences "AAG" and "ACGG". (b) The result of encoding the unaligned sequences into the source language (*Concat* representation). (c) The sentence from the source language is translated to the target language via a transformer model. (d) The translated sentence in the target language (*Spaces* representation). (e) The resulting alignment, decoded from the translated sentence, in which "AA-G" is aligned to "ACGG". The transformer architecture illustration is adapted from (Vaswani et al., 2017).
## Data:
We used SpartaABC (Loewenthal et al., 2021) to generate millions of true alignments. SpartaABC requires the following input: (1) a rooted phylogenetic tree, which includes a topology and branch lengths; (2) a substitution model (amino acids or nucleotides); (3) root sequence length; (4) the indel model parameters, which include: insertion rate (*R_I*), deletion rate (*R_D*), a parameter for the insertion Zipfian distribution (*A_I*), and a parameter for the deletion Zipfian distribution (*A_D*). MSAs were simulated along random phylogenetic tree topologies generated using the program ETE version 3.0 (Huerta-Cepas et al., 2016) with default parameters.
We generated 1,495,000, 2,000 and 3,000, protein MSAs with ten sequences that were used as training validation and testing data, respectively. We generated the same number of DNA MSAs. For each random tree, branch lengths were drawn from a uniform distribution in the range *(0.5,1.0)*. Next, the sequences were generated using SpartaABC with the following parameters: *R_I,R_D \in (0.0,0.05)*, *A_I, A_D \in (1.01,2.0)*. The alignment lengths as well as the sequence lengths of the tree leaves vary within and among datasets as they depend on the indel dynamics and the root length. The root length was sampled uniformly in the range *[32,44]*. Unless stated otherwise, all protein datasets were generated with the WAG+G model, and all DNA datasets were generated with the GTR+G model, with the following parameters: (1) frequencies for the different nucleotides *(0.37, 0.166, 0.307, 0.158)*, in the order "T", "C", "A" and "G"; (2) with the substitutions rate *(0.444, 0.0843, 0.116, 0.107, 0.00027)*, in the order "a", "b", "c", "d", and "e" for the substitution matrix.
## Example:
The following example correspond for the illustrated MSA in the figure above:
{"MSA": "AAAC-GGG", "unaligned_seqs": {"seq0": "AAG", "seq1": "ACGG"}}
## APA
```
Dotan, E., Belinkov, Y., Avram, O., Wygoda, E., Ecker, N., Alburquerque, M., Keren, O., Loewenthal, G., & Pupko T. (2023). Multiple sequence alignment as a sequence-to-sequence learning problem. The Eleventh International Conference on Learning Representations (ICLR 2023).
```
## BibTeX
```
@article{Dotan_multiple_2023,
author = {Dotan, Edo and Belinkov, Yonatan and Avram, Oren and Wygoda, Elya and Ecker, Noa and Alburquerque, Michael and Keren, Omri and Loewenthal, Gil and Pupko, Tal},
month = aug,
title = {{Multiple sequence alignment as a sequence-to-sequence learning problem}},
year = {2023}
}
``` |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.