repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
neuml/txtai | nlp | 456 | Check for empty queue before attempting to convert inputs to dictionaries | Related to https://github.com/neuml/paperai/issues/67
A check should be added to ensure the extractor input queue has content before attempting to convert inputs to dictionaries. | closed | 2023-03-29T19:59:45Z | 2023-03-29T20:01:23Z | https://github.com/neuml/txtai/issues/456 | [
"bug"
] | davidmezzetti | 0 |
jupyterlab/jupyter-ai | jupyter | 1,103 | Chat window works, but IPython magics do not |
## Description
I installed jupyter, jupyterai and langchain-openai in a clean environment (using pixi). I entered my openAI api key and the chat window seems to work. however, using the %%ai magic in the notebook doesn't work
## Reproduce
### screenshot showing that the chat window works
<img width="1285"... | open | 2024-11-12T00:13:06Z | 2025-02-09T14:16:36Z | https://github.com/jupyterlab/jupyter-ai/issues/1103 | [
"bug"
] | sg-s | 8 |
flasgger/flasgger | api | 10 | Fix regex to add missing type to rules | Currently rules only work if type is defined
bad
```
@route('/<param>/<another>')
```
good
```
@route('/<string:param>/<string:another>')
```
But we should accept the first, so regex have to be fixed.
| closed | 2016-01-11T12:49:59Z | 2016-01-14T00:34:21Z | https://github.com/flasgger/flasgger/issues/10 | [] | rochacbruno | 7 |
skforecast/skforecast | scikit-learn | 391 | Prophet | Hi, is Prophet supported?
I see they declare sklearn API.
https://facebook.github.io/prophet/docs/quick_start.html
| closed | 2023-04-08T04:55:37Z | 2023-04-09T19:46:02Z | https://github.com/skforecast/skforecast/issues/391 | [
"help wanted",
"question"
] | AVPokrovsky | 1 |
WZMIAOMIAO/deep-learning-for-image-processing | pytorch | 549 | Up大大,运行predict时导入模型出现错误 | up大大您好,我使用了您的Faster R-CNN代码,我使用规范的VOC数据集保存的方式组织了自己的文件夹,同时运行train_res50_fpn.py这个脚本成功生成了15个epoch的权重文件(已正确指定检测的类别数),但是在使用predict.py预测脚本时,在导入自己的模型这一步出现了问题,我查询了很多资料也没能解决该问题,所以通过这种方式打扰您

The docstring for `.from_json()` says that it takes a dict, but when I... | closed | 2024-03-13T09:55:09Z | 2024-03-26T13:29:42Z | https://github.com/explosion/spaCy/issues/13374 | [
"feat / serialize",
"feat / doc"
] | undercertainty | 1 |
reiinakano/scikit-plot | scikit-learn | 119 | Bug: Failing to import Scikitplot fails due to ImportError | Hey @reiinakano,
first, your work was a great addition to the open-source ML community and is still used. 👍
Unfortunately, importing a freshly installed scikit-plot now fails with the following ImportErrror:
```
import scikitplot
print("Hello World")
```
>>> import scikitplot
Traceback (most recent cal... | open | 2024-01-22T10:19:08Z | 2024-04-10T13:18:08Z | https://github.com/reiinakano/scikit-plot/issues/119 | [] | radlfabs | 4 |
SALib/SALib | numpy | 643 | Grouping of parameters yields NaN sigma values. | Hello there,
firstly thank you for setting up SALIB, it's a great and intuitive tool!
Currently, I am working on a model which includes wind and solar profiles as well as electricity market prices which change hourly. I made dynamic bounds for every hour for each parameter. In my analysis I group these paramete... | closed | 2024-12-12T08:46:28Z | 2024-12-12T14:21:55Z | https://github.com/SALib/SALib/issues/643 | [] | JMH-gif | 2 |
TencentARC/GFPGAN | deep-learning | 245 | error: (-215:Assertion failed) !buf.empty() in function 'imdecode_' | First I met the error as follow:

Then I add int at ’quality‘ as [https://github.com/TencentARC/GFPGAN/issues/93](url)
But I got another error as follow:

### Expected behavior
Different image data shows differen... | closed | 2022-10-11T05:08:11Z | 2022-10-11T14:57:23Z | https://github.com/jmcnamara/XlsxWriter/issues/913 | [
"bug",
"awaiting user feedback"
] | ye-xue | 15 |
anselal/antminer-monitor | dash | 132 | [ERROR] Invalid username or password | It gives me error on v0.5
Any idea how to solve it?
| closed | 2018-10-05T10:15:07Z | 2018-10-05T10:19:17Z | https://github.com/anselal/antminer-monitor/issues/132 | [
":octocat: help wanted"
] | papampi | 2 |
polakowo/vectorbt | data-visualization | 320 | Best practice to install vectorbt on Apple silicon (M1) | Dear Oleg,
Thank you for this excellent tool!
You mentioned in the docs that you are running vectorbt on an M1 Macbok Air.
I usually use pyenv with the venv module from the standard library and pip.
Since numba/llvmlite is not pip installable on the M1, I would be very interested in how you set up your python en... | closed | 2022-01-05T09:56:04Z | 2025-02-06T21:23:49Z | https://github.com/polakowo/vectorbt/issues/320 | [] | 1081 | 5 |
zappa/Zappa | flask | 549 | [Migrated] Unable to access json event data | Originally from: https://github.com/Miserlou/Zappa/issues/1458 by [joshlsullivan](https://github.com/joshlsullivan)
Hi there, when I deploy Zappa, I'm unable to access json data from the Lambda event. If I print the event data, this is what I get:
`[DEBUG] 2018-03-24T14:40:37.991Z 517bfc13-2f71-11e8-9ff3-ed7722cf9e... | closed | 2021-02-20T12:22:36Z | 2024-04-13T16:37:17Z | https://github.com/zappa/Zappa/issues/549 | [
"no-activity",
"auto-closed"
] | jneves | 2 |
sergree/matchering | numpy | 8 | CAN'T SAVE TO A DIFFERENT FOLDER in DJANGO |
Request Method: | GET
-- | --
http://127.0.0.1:8000/dashboard/track/8/master
3.0.2
RuntimeError
Error opening './media/goody/mastered/my_song_master_16bit.wav': System error.
/home/goodness/.local/share/virtualenvs/MeshakProj--GI6wqXg/lib/python3.6/site-packages/soundfile.py in _error_check, line 1357
/home/go... | closed | 2020-01-30T09:55:34Z | 2020-02-03T13:59:18Z | https://github.com/sergree/matchering/issues/8 | [] | GoodnessEzeokafor | 8 |
smarie/python-pytest-cases | pytest | 29 | pytest_fixture_plus does not seem to work with pytest.param parameters | First of all let me thank you for the nice `pytest_fixture_plus`, I am really excited about it.
The problem I am having can be seen in the following example.
This works:
```
from pytest_cases import pytest_fixture_plus as fixture
import pytest
@fixture
@pytest.mark.parametrize("arg1, arg2", [
(1,2),... | closed | 2019-03-21T20:07:42Z | 2019-03-22T16:57:47Z | https://github.com/smarie/python-pytest-cases/issues/29 | [] | Sup3rGeo | 4 |
awtkns/fastapi-crudrouter | fastapi | 119 | Custom ORM | So. I have a ORM which I'd like to use, and to intigrate.
It's a homegrown ORM for the fun of it, but I like my work, so I wanna use it here.
I however couldn't find any section in the docs on how to implement an adapter for your own. | open | 2021-11-19T14:22:45Z | 2021-11-19T14:22:55Z | https://github.com/awtkns/fastapi-crudrouter/issues/119 | [] | luckydonald | 0 |
serpapi/google-search-results-python | web-scraping | 61 | SSLCertVerificationError [SSL: CERTIFICATE_VERIFY_FAILED] error | A user reported receiving this error:
```
SSLCertVerificationError Traceback (most recent call last)
/opt/anaconda3/lib/python3.8/site-packages/urllib3/connectionpool.py in urlopen(self, method, url, body, headers, retries, redirect, assert_same_host, timeout, pool_timeout, release_conn, chunked, body_pos, **respo... | closed | 2023-11-22T01:26:54Z | 2023-11-22T01:27:01Z | https://github.com/serpapi/google-search-results-python/issues/61 | [] | hilmanski | 1 |
shibing624/text2vec | nlp | 66 | 是否可以商用? | RT, 模型是否可以商用?打算用在搜索领域,服务于我们自身的业务。
| closed | 2023-05-10T01:26:43Z | 2023-05-10T02:25:35Z | https://github.com/shibing624/text2vec/issues/66 | [
"question"
] | bh4ffu | 1 |
ploomber/ploomber | jupyter | 783 | SQLUpload constructor should validate that the client is a sqlalchemyclient | otherwise, it'll fail when trying to access the `engine` attribute - which isn't a clear error | closed | 2022-05-17T17:31:49Z | 2022-09-06T01:57:19Z | https://github.com/ploomber/ploomber/issues/783 | [] | edublancas | 0 |
qubvel-org/segmentation_models.pytorch | computer-vision | 668 | ImportError: cannot import name 'get_source_inputs' from 'keras.engine' (/usr/local/lib/python3.7/dist-packages/keras/engine/__init__.py) | When I try to import the library in Google Colab, it raises an error. | closed | 2022-10-09T23:43:24Z | 2022-12-17T01:57:16Z | https://github.com/qubvel-org/segmentation_models.pytorch/issues/668 | [
"Stale"
] | Abecadarian | 2 |
plotly/plotly.py | plotly | 4,699 | Declare `kaleido` as an optional dependency | When calling `write_image` on a figure, it is required that `kaleido` is installed in the active environment. It would be great if this dependency were declared in pyproject.toml as an optional group.
For example, here is the approach taken in pyvista:
https://github.com/pyvista/pyvista/blob/e1401a34cbd281bbe74bc... | open | 2024-07-29T09:26:51Z | 2024-08-13T13:26:53Z | https://github.com/plotly/plotly.py/issues/4699 | [
"feature",
"P3",
"infrastructure"
] | tpgillam | 0 |
snarfed/granary | rest-api | 23 | instagram: implement location | [evidently i left it as a TODO](https://github.com/snarfed/activitystreams-unofficial/blob/master/instagram.py#L207) and never came back to it. :P
ideally, fixing this should make IG photo map locations show up visibly in my feed reader via https://instagram-atom.appspot.com/
| closed | 2015-01-06T18:00:34Z | 2015-01-07T04:09:37Z | https://github.com/snarfed/granary/issues/23 | [] | snarfed | 2 |
home-assistant/core | asyncio | 140,510 | SolaX Power 'measurement' but 'last_reset' is missing | ### The problem
The SolaX integration itself works fine but reporting to energy the energy dashboard seems to have the following error in the energy dashboard configuration that "last_reset" is missing. The same problem as #127805
The X3.0 stick has been tested with various firmware versions. Unfortunately, the resul... | open | 2025-03-13T12:25:05Z | 2025-03-13T12:25:16Z | https://github.com/home-assistant/core/issues/140510 | [
"integration: solax"
] | Tonestep | 1 |
cvat-ai/cvat | pytorch | 8,386 | Delay to check my assessment | Hello, it a month since i submitted my training assessment and i have not heard anything from you. | closed | 2024-09-02T06:09:21Z | 2024-09-02T06:26:33Z | https://github.com/cvat-ai/cvat/issues/8386 | [] | peterizcrisp | 0 |
microsoft/nni | deep-learning | 5,685 | Removing redundant string format in the final experiment log | ### This is a small but very simple request.
In the final experiment summary JSON generated through the NNI WebUI, there are some fields that were originally dictionaries that have been reformatted into strings. This is a small but annoying detail and probably easy to fix.
Most notably, this happens for values i... | open | 2023-09-26T12:19:49Z | 2023-09-26T12:28:22Z | https://github.com/microsoft/nni/issues/5685 | [] | olive004 | 0 |
pydata/bottleneck | numpy | 125 | Where should we host the sphinx manual? | I'd like to move the sphinx manual. Where should we host it? Github? Readthedocs?
| closed | 2016-06-01T19:21:08Z | 2016-09-30T20:57:39Z | https://github.com/pydata/bottleneck/issues/125 | [] | kwgoodman | 5 |
skypilot-org/skypilot | data-science | 4,348 | [Serve] Fall back to latest ready version when detects unrecoverable failure | > Hi @alita-moore ! Thanks for reporting this. We do have some checks against unrecoverable error and stop scaling after detect such error. However, the case you described in the PR is a little bit complicated - it turns to READY first, and then NOT_READY, making it very similar to network transient error.
> https:/... | open | 2024-11-13T20:59:01Z | 2024-12-19T23:08:53Z | https://github.com/skypilot-org/skypilot/issues/4348 | [] | cblmemo | 0 |
Netflix/metaflow | data-science | 1,919 | Silent failure to trigger Argo Workflow from CLI | If the workflow exists in the given namespace, it seems that `python flow.py argo-workflows trigger` can yield this output: `Workflow 'foo' triggered on Argo Workflows (run-id 'bar').`, even if the workflow was not triggered.
We use port forwarding to connect to our Argo Workflows server. If someone forget to port ... | closed | 2024-07-15T19:36:22Z | 2024-07-30T22:21:54Z | https://github.com/Netflix/metaflow/issues/1919 | [] | notablegrace | 2 |
widgetti/solara | jupyter | 868 | Solara Dev Documentation is Buggy | **Issue:**
When I go to [solara docs](https://solara.dev/documentation/), I cannot immediately scroll on the web page. I see the left side panel open and the content, but I can not scroll. Sometimes, when the page is loading, I noticed that I could scroll, but then a quick "flash" of a grey popup shows and disappears... | open | 2024-11-21T16:03:56Z | 2024-11-22T09:47:55Z | https://github.com/widgetti/solara/issues/868 | [
"documentation"
] | jonkimdev | 1 |
scikit-learn-contrib/metric-learn | scikit-learn | 98 | Can this library be used for similarity metric learning | I have a set of vectors and for each pair of these vectors I have a distance (which is not Euclidean). I would like to embed the vectors into Euclidean space so that they are more likely to be close in R^d if they are close under the original measure of distance. I believe this is called similarity metric learning.
... | closed | 2018-06-26T08:41:07Z | 2018-07-04T08:51:05Z | https://github.com/scikit-learn-contrib/metric-learn/issues/98 | [] | lesshaste | 5 |
manbearwiz/youtube-dl-server | rest-api | 109 | Update function fails due to color in pip output | Update function fails due to color in pip output that is not expected since `print(output.decode("ascii"))` is used:
```
File "/usr/src/app/./youtube-dl-server.py", line 141, in <module>
update()
File "/usr/src/app/./youtube-dl-server.py", line 75, in update
print(output.decode("ascii"))
UnicodeDecode... | closed | 2022-08-24T23:06:10Z | 2022-08-25T02:44:40Z | https://github.com/manbearwiz/youtube-dl-server/issues/109 | [] | 0xE1 | 1 |
lepture/authlib | django | 444 | Confusing behavior with OAuth2Session and state not being checked | **Describe the bug**
In the [documentation](https://docs.authlib.org/en/latest/client/oauth2.html#fetch-token) for how to use OAuth2Session client, it says that by supplying state when instantiating the object, then state will be checked when making the `fetch_token` request. In addition, the [docstring](https://git... | closed | 2022-03-22T16:34:48Z | 2022-07-02T19:31:51Z | https://github.com/lepture/authlib/issues/444 | [
"bug"
] | rorour | 2 |
SALib/SALib | numpy | 35 | Is there an API for Method of Morris? | This isn't an issue but a question or two. Is there an API for interfacing with SALib.sample.morris and SALib.analyze.morris, without the need to read and write parameter files, sample files, and model results? Is it possible to get the analysis results directly without using the operating system to write it to a file ... | closed | 2015-01-23T03:09:57Z | 2015-01-29T16:21:08Z | https://github.com/SALib/SALib/issues/35 | [
"enhancement"
] | kdavies4 | 3 |
vitalik/django-ninja | rest-api | 333 | Dynamic response schema based on request parameters | Hello,
Is it currently possible to implement functionality similar to [djangorestframework-queryfields](https://github.com/wimglenn/djangorestframework-queryfields) with Django Ninja? That is, given a request parameter parameter `?fields=field1,field2` or `?exclude_fields=field3`, would it be possible to dynamicall... | open | 2022-01-22T00:03:38Z | 2025-03-08T08:16:03Z | https://github.com/vitalik/django-ninja/issues/333 | [] | dralley | 10 |
slackapi/bolt-python | fastapi | 1,092 | Can't seem to pass back errors for modal field during `.action()` callback method execution | I'm trying to build a modal with a simple input block, and pass back errors to the user after an API call. It looks something like this:
```python
# This is what the input is described in the view
## Note: this is an excerpt, the modal + input renders fine. These are
## built with the slack_sdk's models here: htt... | closed | 2024-06-12T18:57:17Z | 2024-06-14T00:52:02Z | https://github.com/slackapi/bolt-python/issues/1092 | [
"question"
] | macintacos | 5 |
sqlalchemy/alembic | sqlalchemy | 677 | Add --version argument to cli | I was looking for the alembic version in an env, and the cli does not seem to expose it.
There is special action of argparse for this. and is supported also in [python2](https://docs.python.org/2/library/argparse.html#action), so it should be fairly easy to add | closed | 2020-04-06T18:16:48Z | 2020-04-20T17:29:40Z | https://github.com/sqlalchemy/alembic/issues/677 | [
"easy",
"use case"
] | CaselIT | 1 |
Zeyi-Lin/HivisionIDPhotos | machine-learning | 192 | 调用api:idphoto_crop报错ValueError: could not broadcast input array from shape (835,596,3) into shape (835,596,4) | 
| open | 2024-10-14T08:28:19Z | 2024-10-16T09:10:42Z | https://github.com/Zeyi-Lin/HivisionIDPhotos/issues/192 | [] | snowhahaha | 1 |
LAION-AI/Open-Assistant | python | 2,769 | 中文回答过程 | 中文的回答过程中有乱码,回答完毕后,部分文本能正常展示中文,部分展示异常 | closed | 2023-04-20T05:45:38Z | 2023-04-20T06:02:53Z | https://github.com/LAION-AI/Open-Assistant/issues/2769 | [] | TotheAnts | 2 |
Ehco1996/django-sspanel | django | 11 | 大神,看下是我姿势的问题吗 | 随机端口那个我提过了
签到按钮,点击签到之后,弹出签到成功页面,点击确定。
然后刷新页面,会弹出签到失败,刷新一次,弹出一次。
资料编辑
修改协议之后,点击提交按钮,然后刷新页面,总是弹出已经修改成功的页面。刷新一次,弹出一次。
修改混淆,修改加密方式,都这样
希望刷新之后,不弹出各种提示,,因为点击之后,已经弹出过一次提示框了,刷新的话,不应该再次弹出的。 | closed | 2017-10-06T22:00:54Z | 2017-10-08T03:55:31Z | https://github.com/Ehco1996/django-sspanel/issues/11 | [] | cheapssr | 2 |
babysor/MockingBird | deep-learning | 269 | 运行synthesizer_preprocess_audio.py进行预处理时报错 | 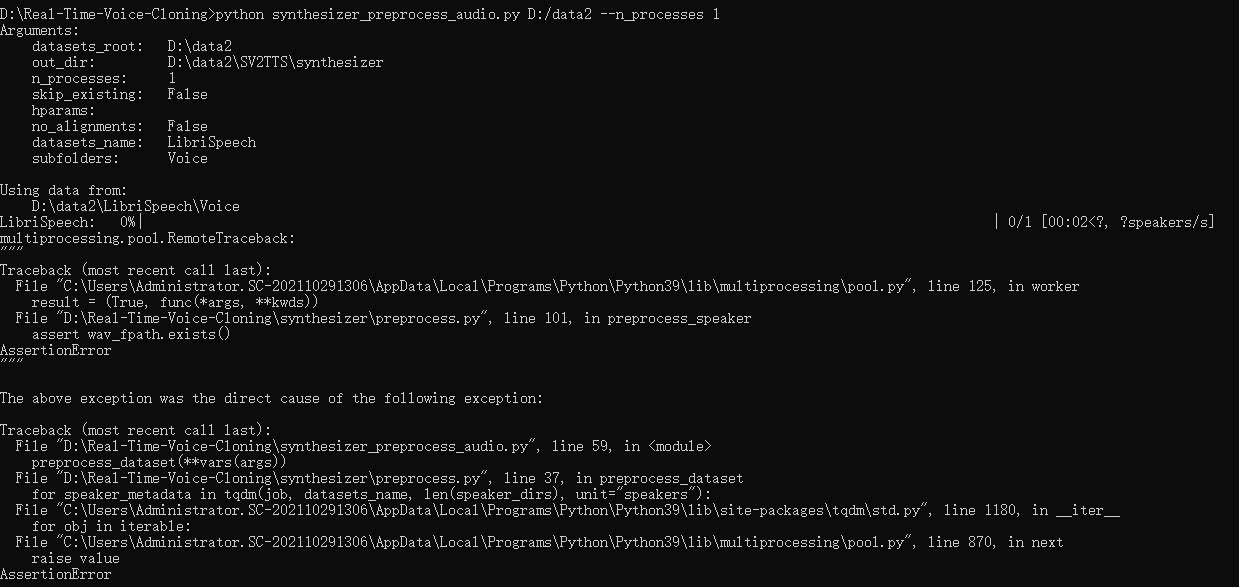
结构如图,也有了对应的alignment对齐文本
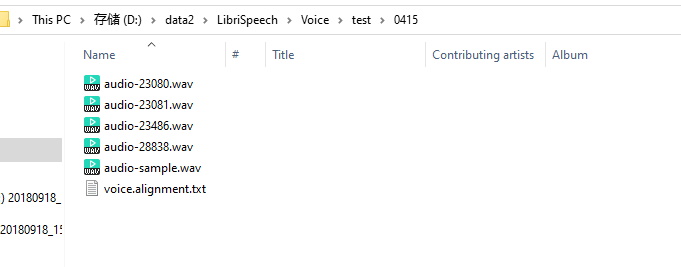
数据对齐文本
, so in some systems `pytest-xdist` is harder to install than it should be.
I suggest we make `psutil` dependency optional, falling back to using `multiprocessing` to detect the n... | closed | 2020-08-17T12:42:53Z | 2020-08-25T12:46:52Z | https://github.com/pytest-dev/pytest-xdist/issues/585 | [
"enhancement"
] | nicoddemus | 1 |
kennethreitz/records | sqlalchemy | 131 | Would it be possible to issue a new release? | The lazy connection feature added in fe0ed3199dd952d57bfa12ecdb6c69acd1c98ece is critical for many use case (e.g. an API). Would you be so kind as to release a new version of records that contains it?
Thanks, Kenneth.
Charles | closed | 2018-03-13T09:46:49Z | 2019-03-17T15:08:18Z | https://github.com/kennethreitz/records/issues/131 | [
"question"
] | charlax | 7 |
pytest-dev/pytest-xdist | pytest | 462 | Test suite memory leak? | While attempting to run the tests with Python 3.7 there seem to be a memory leak. I did not have the issue on Python 2.7.
```sh
Installing collected packages: pytest-xdist
Successfully installed pytest-xdist-1.28.0
/build/pytest-xdist-1.28.0
post-installation fixup
shrinking RPATHs of ELF executables and librarie... | open | 2019-08-18T07:17:53Z | 2019-08-18T10:26:16Z | https://github.com/pytest-dev/pytest-xdist/issues/462 | [] | FRidh | 6 |
JaidedAI/EasyOCR | deep-learning | 1,186 | Inconsistent color conversion, BGR or RGB | Looking at the source code, depending on what data type the input image is supplied as, it will get converted to either RGB or BGR.
If the image is a bytestring, then it is converted to RGB:
https://github.com/JaidedAI/EasyOCR/blob/master/easyocr/utils.py#L745
If the image is a numpy array with 4 channels (RGBA)... | open | 2023-12-15T00:16:33Z | 2024-03-29T06:33:33Z | https://github.com/JaidedAI/EasyOCR/issues/1186 | [] | dkbarn | 4 |
aleju/imgaug | machine-learning | 616 | normalize_shape causes ambiguities | I just spent a good hour tracking down this bug.
It turns out that when you pass the `shape` keyword argument to `KeypointsOnImage` or `BoundingBoxesOnImage` they use `normalize_shape` to preprocess the input. If `shape` is a tuple then there is no problem.
However, the issue happens when I pass `shape` as a n... | closed | 2020-02-16T23:21:28Z | 2020-02-29T15:34:26Z | https://github.com/aleju/imgaug/issues/616 | [
"TODO"
] | Erotemic | 1 |
odoo/odoo | python | 202,420 | [16.0] point_of_sale: It does not allow you to deselect the product when splitting. Ticket Odoo 4656992 | ### Odoo Version
- [x] 16.0
- [ ] 17.0
- [ ] 18.0
- [ ] Other (specify)
### Steps to Reproduce
1. Create a product of type good/stockable.
2. Add one or more variants to the product from step 1.
3. Open the point of sale (verify that the product is visible).
4. Open an order.
5. Select the products created in step 1... | open | 2025-03-18T20:51:48Z | 2025-03-18T21:08:52Z | https://github.com/odoo/odoo/issues/202420 | [] | luandryperez | 0 |
AUTOMATIC1111/stable-diffusion-webui | deep-learning | 16,589 | [Bug]: OSError: Cannot find empty port in range: 7860-7860 with EC2 in Auto scaling group | ### Checklist
- [ ] The issue exists after disabling all extensions
- [ ] The issue exists on a clean installation of webui
- [ ] The issue is caused by an extension, but I believe it is caused by a bug in the webui
- [ ] The issue exists in the current version of the webui
- [ ] The issue has not been reported before... | open | 2024-10-25T02:48:22Z | 2024-10-25T07:21:13Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/16589 | [
"asking-for-help-with-local-system-issues"
] | PiPyL | 1 |
davidteather/TikTok-Api | api | 207 | Why can't i get object user ? | Hi, can you help me? Why does not it work?
code
```
from TikTokApi import TikTokApi
api = TikTokApi()
tiktoks = api.byUsername('donelij')
print(tiktoks)
for tiktok in tiktoks:
print(tiktok)
```
error
> `https://m.tiktok.com/api/user/detail/?uniqueId=donelij&language=en&verifyFp=verify_kdf52cly_U2L... | closed | 2020-08-03T23:24:39Z | 2020-08-09T18:22:42Z | https://github.com/davidteather/TikTok-Api/issues/207 | [
"bug",
"question"
] | markdrrr | 7 |
Buuntu/fastapi-react | fastapi | 45 | Documentation for deployment options | Maybe start with Heroku and Docker Swarm? https://dockerswarm.rocks/ | closed | 2020-05-27T13:21:25Z | 2020-08-05T14:49:46Z | https://github.com/Buuntu/fastapi-react/issues/45 | [
"documentation",
"enhancement",
"good first issue"
] | Buuntu | 0 |
laughingman7743/PyAthena | sqlalchemy | 45 | Handle InvalidRequestException errors raised | I have a script using Athena + SQLAlchemy to run a query and have the results of that query read in as a pandas DataFrame. However, for some queries (which are long), I get this error:
```
botocore.errorfactory.InvalidRequestException: An error occurred (InvalidRequestException) when calling the StartQueryExecution o... | closed | 2018-08-09T21:15:54Z | 2018-08-13T16:52:09Z | https://github.com/laughingman7743/PyAthena/issues/45 | [] | koshy1123 | 3 |
mljar/mljar-supervised | scikit-learn | 640 | How to select models for more SHAP plots? | Hello MLJAR Team! I followed the attached tutorial, and my question is how to use a specific model for predictions and more detailed Shapley Values? After completing the following tutorial:
```python
import pandas as pd
import numpy as np
from sklearn import datasets
from sklearn.model_selection import train_te... | open | 2023-07-28T23:01:34Z | 2023-10-18T07:34:07Z | https://github.com/mljar/mljar-supervised/issues/640 | [] | michael-mazzucco | 5 |
strawberry-graphql/strawberry | fastapi | 3,412 | enhancing robustness of int/float castings for string values containing commas | I run into this problem again and again and it is always annoying to fix it
## Feature Request Type
- [x] Core functionality
- [x] Alteration (enhancement/optimization) of existing feature(s)
- [ ] New behavior
## Description
```
ValueError: could not convert string to float: '3,3'
File "graphql/type/... | closed | 2024-03-19T14:14:27Z | 2025-03-20T15:56:37Z | https://github.com/strawberry-graphql/strawberry/issues/3412 | [] | Speedy1991 | 1 |
deepinsight/insightface | pytorch | 2,060 | Should I change brightness for best results in face recognition? | If it "yes" HOW should I do it? Maybe I should normalize brightness or something like this?! | open | 2022-07-28T11:48:58Z | 2022-08-07T15:04:07Z | https://github.com/deepinsight/insightface/issues/2060 | [] | IamSVP94 | 1 |
streamlit/streamlit | python | 10,750 | Support expandable blocks in markdown | ### Checklist
- [x] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar feature requests.
- [x] I added a descriptive title and summary to this issue.
### Summary
Support expandable content blocks in markdown similar to how its supported in Notion and Github, e.g.:
<deta... | open | 2025-03-12T16:49:06Z | 2025-03-12T17:01:12Z | https://github.com/streamlit/streamlit/issues/10750 | [
"type:enhancement",
"feature:markdown"
] | lukasmasuch | 1 |
python-gitlab/python-gitlab | api | 2,630 | Add support for compliance_frameworks | ## Description of the problem, including code/CLI snippet
Please add support for compliance_frameworks
https://docs.gitlab.com/ee/api/projects.html
## Expected Behavior
## Actual Behavior
## Specifications
- python-gitlab version:
- API version you are using (v3/v4):
- Gitlab server version ... | closed | 2023-08-02T00:03:40Z | 2024-08-26T01:34:07Z | https://github.com/python-gitlab/python-gitlab/issues/2630 | [
"upstream"
] | coffeecoco | 1 |
microsoft/nni | machine-learning | 5,318 | How to use DDP in multi-trial NAS? | Hi, is there a easy way to use DDP in multi-trial NAS?
I tried multi-trial NAS based on this example: https://github.com/microsoft/nni/blob/master/examples/nas/multi-trial/mnist/search.py. Is it possible to wrap it with DDP? | closed | 2023-01-17T02:56:54Z | 2023-03-08T09:34:32Z | https://github.com/microsoft/nni/issues/5318 | [] | heibaidaolx123 | 3 |
bmoscon/cryptofeed | asyncio | 866 | deribit L2_BOOK raising ValueError Authenticated channel | **Describe the bug**
I am trying to subscribe to Deribit L2_BOOK channel (public data) and Cryptofeed tels me that the channel is authenticated and needs auth keys.
**To Reproduce**
f = FeedHandler()
f.add_feed(Deribit(symbols=['BTC-USD-PERP'], channels=[L2_BOOK], callbacks={L2_BOOK: book}))
f.run()
**Expecte... | closed | 2022-06-29T11:21:17Z | 2022-06-29T13:23:05Z | https://github.com/bmoscon/cryptofeed/issues/866 | [
"bug"
] | docek | 1 |
skypilot-org/skypilot | data-science | 4,861 | [Doc] document how to deploy multiple API servers and deploy server using existing ingress | Deploying our helm chart to k8s cluster created by `sky local up` raises the following errors:
```
Error: Unable to continue with install: IngressClass "nginx" in namespace "" exists and cannot be imported into the current release: invalid ownership metadata; label validation error: missing key "app.kubernetes.io/mana... | open | 2025-03-01T09:20:24Z | 2025-03-01T09:20:34Z | https://github.com/skypilot-org/skypilot/issues/4861 | [] | aylei | 0 |
pydantic/pydantic-core | pydantic | 1,437 | Is possible to publish pydantic-core to crates.io | Would it be possible to publish pydantic-core to crates.io? This would allow Rust developers to directly use and benefit from pydantic-core's functionality in their Rust projects. | closed | 2024-09-04T16:03:23Z | 2024-09-17T12:02:07Z | https://github.com/pydantic/pydantic-core/issues/1437 | [] | Folyd | 2 |
ets-labs/python-dependency-injector | flask | 56 | Review docs: Providers | closed | 2015-05-08T14:39:31Z | 2015-07-13T07:31:58Z | https://github.com/ets-labs/python-dependency-injector/issues/56 | [
"docs"
] | rmk135 | 0 | |
pallets-eco/flask-sqlalchemy | flask | 1,189 | __bind_key__ not working | Hello,
the bind key is not working for me. Is this a bug, or a problem with my code?
All data is written to `database.db`, but shold be seperated to the two databases. The `database_logging.db` was created but is empty.
The relevant extract of the code. I need the declarative_base because I want to seperate th... | closed | 2023-04-08T17:22:21Z | 2023-04-23T01:10:28Z | https://github.com/pallets-eco/flask-sqlalchemy/issues/1189 | [] | Laserlicht | 2 |
pyg-team/pytorch_geometric | deep-learning | 9,393 | Error converting "to_dense_adj" from "from_scipy_sparse_matrix" | ### 🐛 Describe the bug
I'm trying to convert to a dense matrix but going via network x is too memory intensive with the real file. I was hoping that I could do it via the csr representations rather than via networkx as they are more efficient. Oddly this is occuring even with very small files (example attached)
``... | closed | 2024-06-04T16:57:09Z | 2024-06-11T11:27:20Z | https://github.com/pyg-team/pytorch_geometric/issues/9393 | [
"bug"
] | timdnewman | 2 |
numpy/numpy | numpy | 27,994 | ENH: Support of new str.format() syntax instead of the old %-formatting in numpy.savetxt | ### Proposed new feature or change:
Currently `numpy.savetxt` uses old %-formatting to format strings. But old formatting doesn't support some new features. For example: a control of negative zeros ([PEP-682](https://peps.python.org/pep-0682/)), or a binary output (#20755). It would be great to support new formattin... | open | 2024-12-13T14:45:13Z | 2024-12-19T15:32:27Z | https://github.com/numpy/numpy/issues/27994 | [
"01 - Enhancement"
] | PavelStishenko | 5 |
encode/databases | asyncio | 422 | DatabaseUrl bug when using Unix domain socket | I'm deploying a FastAPI application on Google Cloud Run which connects to a Cloud SQL instance using this package. The crux of the issue is that connecting with:
```python
db = databases.Database(url)
await db.connect()
```
fails whereas connecting through sqlalchemy's `create_engine` with
```python
engin... | closed | 2021-11-15T11:57:51Z | 2021-11-16T09:22:47Z | https://github.com/encode/databases/issues/422 | [
"bug"
] | dbatten5 | 13 |
huggingface/datasets | deep-learning | 6,640 | Sign Language Support | ### Feature request
Currently, there are only several Sign Language labels, I would like to propose adding all the Signed Languages as new labels which are described in this ISO standard: https://www.evertype.com/standards/iso639/sign-language.html
### Motivation
Datasets currently only have labels for several signe... | open | 2024-02-02T21:54:51Z | 2024-02-02T21:54:51Z | https://github.com/huggingface/datasets/issues/6640 | [
"enhancement"
] | Merterm | 0 |
graphql-python/gql | graphql | 222 | RequestsHTTPTransport: retries option with POST method does not effect | - RequestsHTTPTransport's retries option uses requests library.
- RequestsHTTPTransport's HTTP(S) request uses POST method by default.
- `requests`'s max_retries option does not effects with POST method by default.
- requests.adapters.Retry == urllib3.util.retry.Retry
- https://stackoverflow.com/a/35707701
-... | closed | 2021-07-30T12:53:05Z | 2021-10-26T08:46:27Z | https://github.com/graphql-python/gql/issues/222 | [
"type: bug"
] | khiker | 4 |
ghtmtt/DataPlotly | plotly | 55 | Translation updates | With the new UI all the new strings have to be pushed to transifex | closed | 2017-11-26T11:42:37Z | 2017-11-27T10:09:09Z | https://github.com/ghtmtt/DataPlotly/issues/55 | [
"docs"
] | ghtmtt | 1 |
pytorch/pytorch | machine-learning | 149,616 | intermittent toch.compiler failures when running gemma model | ### 🐛 Describe the bug
Hi, i'm trying to fix a intermittent torch.compiler failures with cpp wrapper when running gemma model and wonder if someone can help providing some clue for debug or get a minial reproducer? The error is not specific to arm but also reproducible on intel machines.
```TORCHINDUCTOR_CPP_WRAPPER... | closed | 2025-03-20T10:50:48Z | 2025-03-24T17:16:10Z | https://github.com/pytorch/pytorch/issues/149616 | [
"module: cpu",
"triaged",
"oncall: pt2",
"module: inductor"
] | taoye9 | 2 |
streamlit/streamlit | data-visualization | 10,415 | Multi-Index Columns with brackets get renamed to square brackets | ### Checklist
- [x] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar issues.
- [x] I added a very descriptive title to this issue.
- [x] I have provided sufficient information below to help reproduce this issue.
### Summary
Hey guys,
When using brackets in column titl... | open | 2025-02-17T12:01:30Z | 2025-03-06T16:13:31Z | https://github.com/streamlit/streamlit/issues/10415 | [
"type:bug",
"feature:st.dataframe",
"status:confirmed",
"priority:P4"
] | itsToggle | 3 |
vchaptsev/cookiecutter-django-vue | graphql | 28 | Subdomain support | Hi, cool project. Do you have plans to support subdomains? I'm working on a similar project and I am working on how to set up subdomain support. From what I have seen, I think it can be done with a middleware class on the Django side, some processing of the URL in Vue's router and editing nginx to catch the subdomains. | closed | 2018-10-07T23:48:16Z | 2019-03-15T13:58:58Z | https://github.com/vchaptsev/cookiecutter-django-vue/issues/28 | [
"question"
] | briancaffey | 2 |
FujiwaraChoki/MoneyPrinter | automation | 149 | [BUG] Error: float division by zero | it does this

here's my .env settings because i think it can maybe be this

| closed | 2024-02-10T14:24:13Z | 2024-02-11T09:01:28Z | https://github.com/FujiwaraChoki/MoneyPrinter/issues/149 | [] | N3xi0 | 7 |
taverntesting/tavern | pytest | 875 | Try to use the external function in the URL | Hi
I want to get an `ID` value from another program and splice it into the URL.
Similar to: http://localhost:7200/api/manager/system/organizations/{id}
so I try to use external function,
- Tavern `2.2.0`
```
- name: Delete a user
request:
method: DELETE
url:
$ext:
func... | closed | 2023-07-04T11:39:08Z | 2023-09-18T14:50:52Z | https://github.com/taverntesting/tavern/issues/875 | [] | IrisBlume-dot | 1 |
nolar/kopf | asyncio | 684 | Scoping and liveness in embedded mode? | ## Question
<!-- What problem do you currently face and see no solution for it? -->
Hi, so I recently refactored some of my code to use [embedding](https://kopf.readthedocs.io/en/stable/embedding/) but now that I don't have access to the `kopf run` CLI flags, I feel a bit like a second-class citizen 😞
I'm curr... | closed | 2021-02-16T20:07:17Z | 2021-02-17T04:21:00Z | https://github.com/nolar/kopf/issues/684 | [
"question"
] | OmegaVVeapon | 2 |
healthchecks/healthchecks | django | 289 | psycopg2-binary warnining | When I run the project I get the following warning from psycopg2
````
/usr/local/lib/python3.7/dist-packages/psycopg2/__init__.py:144: UserWarning: The psycopg2 wheel package will be renamed from release 2.8; in order to keep installing from binary please use "pip install psycopg2-binary" instead. For details see: <h... | closed | 2019-09-29T14:38:35Z | 2019-09-30T13:08:49Z | https://github.com/healthchecks/healthchecks/issues/289 | [] | SuperSandro2000 | 1 |
keras-team/keras | deep-learning | 20,437 | Add `ifft2` method to ops | I'm curious why there is no `ops.ifft2`. Given that there is already `fft` and `fft2`, implementing one is trivial.
Here is an example of what an `ifft2` would look like:
```python
import keras
def keras_ops_ifft2(fft_real,fft_imag):
"""
Inputs are real and imaginary parts of array
of shape [.... | closed | 2024-11-01T18:32:58Z | 2024-11-05T00:14:56Z | https://github.com/keras-team/keras/issues/20437 | [
"stat:contributions welcome",
"type:feature"
] | markveillette | 1 |
mljar/mercury | jupyter | 275 | cant donwload files from output directory for private notebooks and s3 storage | Steps to reproduce:
- create private Site,
- create notebook with OutputDir,
- use s3 as storage,
When downloading a file you will get an error because token is added to requests to s3 server. We should send request to s3 without authentication token. | closed | 2023-05-16T10:21:00Z | 2023-05-16T11:03:35Z | https://github.com/mljar/mercury/issues/275 | [
"bug"
] | pplonski | 0 |
pallets-eco/flask-sqlalchemy | flask | 377 | Doc site missing link to PDF download | I was able to figure out the link is http://flask-sqlalchemy.pocoo.org/2.1/.latex/Flask-SQLAlchemy.pdf and successfully downloaded it.
Looking at the source, the intent was clearly to have the link in the sidebar.
(Don't have a chance to figure out how to make the change myself at the moment. I'll look into it as s... | closed | 2016-02-24T23:44:18Z | 2020-12-05T21:31:06Z | https://github.com/pallets-eco/flask-sqlalchemy/issues/377 | [] | vassudanagunta | 3 |
slackapi/bolt-python | fastapi | 520 | How to know team_id in app_home_opened event listeners | Unable to see view in app_home_opened event:
- When someone opens a home tab, I am fetching the team_id from the Event payload (inside view) to identify the workspace and sending the corresponding home tab but It seems like there is no view section in the Event payload.
```python
@bolt_app.event("app_home_open... | closed | 2021-11-10T12:10:38Z | 2021-11-10T12:44:28Z | https://github.com/slackapi/bolt-python/issues/520 | [
"question"
] | Cyb-Nikh | 2 |
amidaware/tacticalrmm | django | 1,764 | 0.17.5 fails to install properly and requires some manual changes to shape it | **Server Info (please complete the following information):**
- OS: Ubuntu 22.04.3 LTS
- RMM Version (as shown in top left of web UI): 0.17.5
**Installation Method:**
- [x] Standard
- [ ] Docker
**Describe the bug**
A clear and concise description of what the bug is.
1. it's a fresh-new ubuntu vps
2... | closed | 2024-02-21T15:34:48Z | 2024-02-21T17:07:47Z | https://github.com/amidaware/tacticalrmm/issues/1764 | [] | optiproplus | 2 |
pyeventsourcing/eventsourcing | sqlalchemy | 138 | Rebuilding Aggregate root | Hey @johnbywater
First of all a big thanks, this library is awesome!
I've got a question about rebuilding the aggregate root.
I've got this simple hangman web API and I do different calls to that API for guessing letters. I just noticed that every time that I do a call to the API, I will get new Aggregate Root... | closed | 2018-02-15T22:51:44Z | 2018-02-15T23:17:18Z | https://github.com/pyeventsourcing/eventsourcing/issues/138 | [] | weemen | 1 |
CorentinJ/Real-Time-Voice-Cloning | deep-learning | 1,172 | Is this lib capable of TTS? | Hi, I think this is the only lib out here that can synthesize hq voice . I was wondering if it can also generate tts | open | 2023-03-10T18:15:56Z | 2023-03-10T18:15:56Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/1172 | [] | destlaver | 0 |
microsoft/hummingbird | scikit-learn | 305 | Generate automatically the schema for ONNX models. | ONNX models contains all the input information therefore it should be possible to automatically generate the schema definition \ inputs starting just from the model.
If this is true, we can remove `ONNX_INITIAL_TYPES ` from the support input parameters. | closed | 2020-09-22T23:31:11Z | 2020-10-20T23:10:51Z | https://github.com/microsoft/hummingbird/issues/305 | [] | interesaaat | 0 |
keras-rl/keras-rl | tensorflow | 388 | Value error when running DQN.fit | I tried teaching AI how to play breakout but my code crashes when I try to teach DQN model.
``
import gym
import numpy as np
import tensorflow as tf
from rl.agents.dqn import DQNAgent
from rl.policy import LinearAnnealedPolicy, EpsGreedyQPolicy
from rl.memory import SequentialMemory
from keras.layers import Den... | open | 2022-04-16T19:25:24Z | 2022-09-29T17:56:02Z | https://github.com/keras-rl/keras-rl/issues/388 | [] | GravermanDev | 2 |
tiangolo/uvicorn-gunicorn-fastapi-docker | fastapi | 43 | Support for nvidia | Hello,
Thanks for your work ! Do you plan to add a docker image starting from nvidia/cuda to have cuda toolkit installed ?
| closed | 2020-05-15T08:05:04Z | 2020-06-15T12:58:48Z | https://github.com/tiangolo/uvicorn-gunicorn-fastapi-docker/issues/43 | [
"answered"
] | griseau | 1 |
Gozargah/Marzban | api | 1,021 | Marzban-node "unable to get local issuer certificate" | I have server with few normally worked marzban-nodes.
I try to add new, do it same, but Marzban central server can not connect to node.
I get error:
`[SSL: CERTIFICATE_VERIFY_FAILED] certificate verify failed: unable to get local issuer certificate (_ssl.c:1007)`
Can you help me to understand? Certificates issued... | open | 2024-05-30T18:31:16Z | 2024-05-30T18:31:16Z | https://github.com/Gozargah/Marzban/issues/1021 | [
"Bug"
] | ultraleks | 0 |
albumentations-team/albumentations | deep-learning | 1,698 | Unexpected Kernel Size Application in `Blur` Transformation | I've discovered what seems to be a bug where the `Blur` transformation applies a smaller kernel size than specified by the `blur_limit`. This behavior occurs even when large values are set for the `blur_limit`, leading to blurring effects that do not match the expected intensity.
### Steps to Reproduce
I observed... | closed | 2024-05-03T12:36:39Z | 2024-05-04T05:34:23Z | https://github.com/albumentations-team/albumentations/issues/1698 | [
"bug"
] | LeLub | 4 |
ARM-DOE/pyart | data-visualization | 1,037 | Remove numpy import in setup.py | When installing pyart in a brand new virtual env, errors appear when installing it along with other requirements (with pip).
The reason is that pyart needs `numpy` installed prior to its installation. We can see that `numpy` is required within the `setup.py` file 2 times :
`from numpy.distutils.misc_util import Co... | closed | 2022-02-23T08:56:57Z | 2022-02-23T09:01:29Z | https://github.com/ARM-DOE/pyart/issues/1037 | [] | Vforcell | 0 |
deepspeedai/DeepSpeed | deep-learning | 6,987 | model.parameters() return [Parameter containing: tensor([], device='cuda:0', dtype=torch.bfloat16, requires_grad=True)] when using zero3 | **Describe the bug**
Try to print model.parameters() in transfomers trainer(), but get Parameter containing: tensor([], device='cuda:0', dtype=torch.bfloat16, requires_grad=True) for all layers
In fact, I am trying to return the correct model.parameters() in DeepSpeed Zero-3 mode and use the EMA model. Could you sugges... | closed | 2025-01-31T16:48:27Z | 2025-02-14T00:48:09Z | https://github.com/deepspeedai/DeepSpeed/issues/6987 | [
"bug",
"training"
] | fanfanffff1 | 1 |
robinhood/faust | asyncio | 324 | Autodiscovery: venusian.scan should ignore `__main__.py` | If venusian.scan imports `__main__.py` it causes the app to be created twice | closed | 2019-03-28T16:48:58Z | 2019-03-28T16:59:48Z | https://github.com/robinhood/faust/issues/324 | [] | ask | 0 |
AirtestProject/Airtest | automation | 346 | 为什么script中的图片和实际程序的截图识别吻合度相差很大? | **(重要!问题分类)**
* 图像识别、设备控制相关问题 -> 按下面的步骤
**描述问题bug**
使用airtestIDE直接连接整个windows屏幕。
分别在不同分辨率下进行测试图像识别(用了[#4](https://github.com/AirtestProject/Airtest/issues/4)提到的方法之后,使用运行很好,没有问题)
之前发现如果连接整个屏幕的话,script框内的图片也会被识别到
但是在不同分辨率下运行之后,我发现对于**其他程序识别度99.9%的图片,在script框内的识别度只有40-60%**
明明script框里的图片和实际程序的截图应该是一样的,为什么... | closed | 2019-04-08T07:02:43Z | 2019-07-10T09:15:35Z | https://github.com/AirtestProject/Airtest/issues/346 | [] | niuniuprice | 1 |
matplotlib/matplotlib | data-visualization | 29,659 | [Bug]: Unnecessary double start of `Animation` in some circumstances | ### Bug summary
When a a figure has already had a draw event happen, then an Animation object is intialized, after calling `animation.save` the animation will loop again after the save has finished.
This is due to this line: https://github.com/matplotlib/matplotlib/blob/964355130c5389926641a03154f56f8e081fbfd3/lib/m... | open | 2025-02-22T07:02:28Z | 2025-02-24T08:07:36Z | https://github.com/matplotlib/matplotlib/issues/29659 | [] | ianhi | 1 |
apify/crawlee-python | automation | 721 | Implement option for persistent context to PlaywrightCrawler | - Implement an option for persistent context (`user_data_dir`) to PlaywrightCrawler in a similar way as it is in the Crawlee JS.
- https://crawlee.dev/api/browser-pool/interface/BrowserPluginOptions#userDataDir
- Before implementation sync with @barjin, as he can provide further context and also suggest potential imp... | closed | 2024-11-21T19:51:51Z | 2025-02-25T09:12:11Z | https://github.com/apify/crawlee-python/issues/721 | [
"enhancement",
"t-tooling"
] | vdusek | 0 |
onnx/onnx | tensorflow | 6,380 | Invalid protobuf error when loading successfully exported onnx model | # Bug Report
### Is the issue related to model conversion?
<!-- If the ONNX checker reports issues with this model then this is most probably related to the converter used to convert the original framework model to ONNX. Please create this bug in the appropriate converter's GitHub repo (pytorch, tensorflow-onnx, sk... | closed | 2024-09-21T00:27:56Z | 2024-11-01T14:57:29Z | https://github.com/onnx/onnx/issues/6380 | [] | DagonArises | 2 |
pallets/quart | asyncio | 274 | can't load variables with render_template | The bug stops me from even trying to load any kind of variable
1. Setup a basic app
2. make a html page in the templates folder
3. Import `render_template` and try to load a variable into the template
4. It will error out after running and visiting the page
`from quart import Quart, render_template`
`app ... | closed | 2023-09-20T16:00:15Z | 2023-10-05T00:17:27Z | https://github.com/pallets/quart/issues/274 | [] | Arcader717 | 2 |
vitalik/django-ninja | rest-api | 834 | [BUG] `servers` are null, but should be empty list/non-existent | **Describe the bug**
When exporting OpenAPI Schema, some tools (e.g. [openapi-generator-cli](https://github.com/OpenAPITools/openapi-generator-cli)) can't handle null value. Handling null is not documented, but empty list/lack of this property is ([OpenAPI specs](https://spec.openapis.org/oas/v3.1.0#oasServers)).
*... | closed | 2023-08-28T10:56:46Z | 2023-08-28T11:50:31Z | https://github.com/vitalik/django-ninja/issues/834 | [] | pgronkievitz | 2 |
AntonOsika/gpt-engineer | python | 596 | Run benchmark for `--steps tdd` and compare to "default" benchmark results | If --steps tdd doesn't work well, we should remove it.
If we see obvious ways to improve it, we can of course first consider fixing and see if it helps as part of the investigation.
| closed | 2023-08-15T21:19:46Z | 2023-09-06T11:08:55Z | https://github.com/AntonOsika/gpt-engineer/issues/596 | [
"good first issue"
] | AntonOsika | 5 |
comfyanonymous/ComfyUI | pytorch | 6,493 | Didn't see Nvidia Cosmos workflow | ### Feature Idea
Nvidia Cosmos 7B and 14B: text to video and image to video diffusion model support.
### Existing Solutions
_No response_
### Other
_No response_ | closed | 2025-01-17T01:51:14Z | 2025-01-18T17:43:45Z | https://github.com/comfyanonymous/ComfyUI/issues/6493 | [
"Feature"
] | IAFFeng | 1 |
vanna-ai/vanna | data-visualization | 447 | Why is "add_question_sql" storing vectors as question+sql? SQL isn't natural language, so it wouldn't affect the semantic matching of input queries? | 
Why is "add_question_sql" storing vectors as question+sql? SQL isn't natural language, so it wouldn't affect the semantic matching of input queries? | closed | 2024-05-17T01:50:31Z | 2024-05-23T02:32:05Z | https://github.com/vanna-ai/vanna/issues/447 | [] | qingwu11 | 2 |
pytest-dev/pytest-html | pytest | 750 | Captured stdout in a subtest (pytest-subtests) is not displayed properly | Captured stdout in a subtest is not displayed as expected. Tested with v3.2.0 and v4.0.2 - both have the issue in a different way. Also the issue behavior changes depending on the `--capture` option value. I tried `fd` (the default value) and `tee-sys`. Only v3.2.0 with `--capture tee-sys` seems to work as expected.
... | closed | 2023-10-20T18:27:23Z | 2023-10-23T15:36:25Z | https://github.com/pytest-dev/pytest-html/issues/750 | [] | yugokato | 2 |
cvat-ai/cvat | tensorflow | 8,494 | Error in django request | Hi,
I have installed cvat v12.6.2.
Everything else is working fine, but I am getting these errors in Cvat_opa container logs
```typescript
{"level":"error","msg":"Bundle load failed: request failed: Get \"http://cvat-server:8080/api/auth/rules\": dial tcp: lookup cvat-server on 127.0.0.11:53: no such host","name":"... | open | 2024-10-01T11:56:12Z | 2024-10-08T16:02:50Z | https://github.com/cvat-ai/cvat/issues/8494 | [
"bug"
] | ShreenidhiH | 7 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.