path stringlengths 8 399 | content_id stringlengths 40 40 | detected_licenses list | license_type stringclasses 2 values | repo_name stringlengths 6 109 | repo_url stringlengths 25 128 | star_events_count int64 0 52.9k | fork_events_count int64 0 7.07k | gha_license_id stringclasses 9 values | gha_event_created_at timestamp[us] | gha_updated_at timestamp[us] | gha_language stringclasses 28 values | language stringclasses 1 value | is_generated bool 1 class | is_vendor bool 1 class | conversion_extension stringclasses 17 values | size int64 317 10.5M | script stringlengths 245 9.7M | script_size int64 245 9.7M |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
/RegressaoSimplesListaRedesNeurais.ipynb | e1c60a4ebbd2bad17c50335d850e60dad6434627 | [] | no_license | AlbertoRodrigues/DisciplinaRedesNeurais | https://github.com/AlbertoRodrigues/DisciplinaRedesNeurais | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 32,961 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pickle
import time
from ipypb import track
import WRCP_ALS3 as wrcp
from general_functions import sqrt_err_relative, check_coo_tensor, gen_coo_tensor
from general_functions import create_filter, hr
# -
# ### Zero launch of a function(for NUMBA):
init_shape = (100, 100, 10)
coo, vals = gen_coo_tensor(init_shape, density=0.002)
assert check_coo_tensor(coo)!= "Bad"
# +
# %%time
max_iter = 12
rank = 5
seed = 13
a, b, c, _, _ = wrcp.wrcp_als3(
coo, vals, init_shape,
rank=rank,
l2=0.25,
max_iter=max_iter,
seed=seed,
show_iter=False,
it_over=False,
)
rerr = sqrt_err_relative(
coo, vals, init_shape, a, b, c,
)
print(rerr)
# -
# ### Load the data:
# +
path_data = "Link_Prediction_Data/FB15K237/"
entity_list = pickle.load(open(path_data + 'entity_list', 'rb'))
relation_list = pickle.load(open(path_data + 'relation_list', 'rb'))
train_triples = pickle.load(open(path_data + 'train_triples', 'rb'))
valid_triples = pickle.load(open(path_data + 'valid_triples', 'rb'))
test_triples = pickle.load(open(path_data + 'test_triples', 'rb'))
train_valid_triples = pickle.load(open(path_data + 'train_valid_triples', 'rb'))
entity_map = pickle.load(open(path_data + 'entity_map', 'rb'))
relation_map = pickle.load(open(path_data + 'relation_map', 'rb'))
all_triples = train_valid_triples + test_triples
# -
print(entity_list[:3], "\n", len(entity_list))
print(relation_list[:3], "\n", len(relation_list))
len(train_triples)
len(valid_triples)
len(test_triples)
len(train_valid_triples)
len(all_triples)
list(entity_map.items())[:2]
list(relation_map.items())[:2]
test_filter = create_filter(test_triples, all_triples)
valid_filter = create_filter(valid_triples, all_triples)
# ### Create Sparse COO Train Tensor:
# +
values = [1] * len(train_triples)
values = np.array(values, dtype=np.float64)
coords = np.array(train_triples, dtype=np.int32)
nnz = len(train_triples)
data_shape = (len(entity_list), len(relation_list), len(entity_list))
data_shape
# -
# ## WRCP-ALS3:
max_iter = 2
rank = 25
l2 = 3e0
seed = 13
# %%time
a, b, c, err_arr, it = wrcp.wrcp_als3(
coords, values, data_shape,
rank=rank,
l2=l2,
max_iter=max_iter,
seed=seed,
show_iter=False,
it_over=False,
)
rerr = sqrt_err_relative(
coords, values, data_shape, a, b, c,
)
print(f"Relative error = {rerr}")
plt.xlabel("Iteration")
plt.ylabel("Relative error")
plt.title(f"FB15k-237 / WRCP-ALS3(R={rank})")
#plt.xticks(np.arange(it))
plt.yscale("log")
plt.plot(np.arange(1, it+1), err_arr[:it], '-*', c="#8b0a50")
# %%time
hr(valid_filter, valid_triples, a, b, c, [1, 3, 10])
# %%time
hr(test_filter, test_triples, a, b, c, [1, 3, 10])
# ## Find best parameters:
ranks = (25, 50, 100, 200, 300, 400, 600, 700)
l2 = 3e0
n_iter = 2
seed = 13
table_results = pd.DataFrame(
np.zeros((len(ranks), 14)),
index=[i for i in range(1, len(ranks) + 1)],
columns=("iter", "rank", "l2", "time_train",
"time_hr", "error", "hr1_valid", "hr1_test",
"hr3_valid", "hr3_test", "hr10_valid", "hr10_test",
"mrr_valid", "mrr_test"),
)
table_results
idx = 0
for rank in track(ranks):
idx += 1
# Train model
start_tr = time.time()
a, b, c, err_arr, it = wrcp.wrcp_als3(
coords, values, data_shape,
rank=rank,
l2=l2,
max_iter=n_iter,
seed=seed,
show_iter=False,
it_over=False,
)
end_tr = time.time()
# Find relative error
rerr = sqrt_err_relative(
coords, values, data_shape, a, b, c,
)
# Find Hit Rate on a validation set
start_hr = time.time()
hr1_valid, hr3_valid, hr10_valid, mrr_valid = hr(
valid_filter, valid_triples, a, b, c, [1, 3, 10]
)
# Find Hit Rate on a test set
hr1_test, hr3_test, hr10_test, mrr_test = hr(
test_filter, test_triples, a, b, c, [1, 3, 10]
)
end_hr = time.time()
table_results.loc[idx]["iter"] = n_iter
table_results.loc[idx]["rank"] = rank
table_results.loc[idx]["l2"] = l2
table_results.loc[idx]["time_train"] = end_tr - start_tr
table_results.loc[idx]["time_hr"] = end_hr - start_hr
table_results.loc[idx]["error"] = rerr
table_results.loc[idx]["hr1_valid"] = hr1_valid
table_results.loc[idx]["hr1_test"] = hr1_test
table_results.loc[idx]["hr3_valid"] = hr3_valid
table_results.loc[idx]["hr3_test"] = hr3_test
table_results.loc[idx]["hr10_valid"] = hr10_valid
table_results.loc[idx]["hr10_test"] = hr10_test
table_results.loc[idx]["mrr_valid"] = mrr_valid
table_results.loc[idx]["mrr_test"] = mrr_test
table_results
table_results.to_csv("Link_Prediction_Factors/FB15K237/results.csv")
plt.xlabel("Rank")
plt.ylabel("HR@10")
plt.title(f"FB15K237 / WRCP-ALS3")
plt.xticks(table_results['rank'])
#plt.yscale("log")
plt.plot(table_results['rank'], table_results['hr10_test'], '-*', c="#8b0a50")
# ## Merge train and valid sets:
# +
tv_values = [1] * len(train_valid_triples)
tv_values = np.array(tv_values, dtype=np.float64)
tv_coords = np.array(train_valid_triples, dtype=np.int32)
nnz = len(train_valid_triples)
data_shape = (len(entity_list), len(relation_list), len(entity_list))
data_shape, nnz
# -
max_iter = 2
rank = 700
l2 = 3e0
seed = 13
# %%time
a, b, c, err_arr, it = wrcp.wrcp_als3(
tv_coords, tv_values, data_shape,
rank=rank,
l2=l2,
max_iter=max_iter,
seed=seed,
show_iter=False,
it_over=False,
)
# %%time
hr(test_filter, test_triples, a, b, c, [1, 3, 10])
# ### Save Factors:
# +
path_factors = "Link_Prediction_Factors/FB15K237/"
pickle.dump(a, open(path_factors + 'A_factor', 'wb'))
pickle.dump(b, open(path_factors + 'B_factor', 'wb'))
pickle.dump(c, open(path_factors + 'C_factor', 'wb'))
pickle.dump(err_arr, open(path_factors + 'Errors', 'wb'))
# -
# ## We can load calculated factors:
# +
path_factors = "Link_Prediction_Factors/FB15K237/"
a = pickle.load(open(path_factors + 'A_factor', 'rb'))
b = pickle.load(open(path_factors + 'B_factor', 'rb'))
c = pickle.load(open(path_factors + 'C_factor', 'rb'))
# -
# %%time
hr(valid_filter, valid_triples, a, b, c, [1, 3, 10])
# %%time
hr(test_filter, test_triples, a, b, c, [1, 3, 10])
# ## Let's look at Tucker TD:
import polara.lib.hosvd as hosvd
# %%time
core_shape = (50, 50, 50)
p, q, r, g = hosvd.tucker_als(
idx=coords,
val=values,
shape=data_shape,
core_shape=core_shape,
iters=100,
growth_tol=0.01,
batch_run=False,
seed=15,
)
# +
from numba import jit
@jit(nopython=True)
def sqrt_err_tucker(coo_tensor, vals, shape, a, b, c, g):
result = 0.0
for item in range(coo_tensor.shape[0]):
coord = coo_tensor[item]
elem = 0.0
for p in range(a.shape[1]):
for q in range(b.shape[1]):
for r in range(c.shape[1]):
elem += g[p, q, r] * a[coord[0], p] * b[coord[1], q] * c[coord[2], r]
result += (vals[item] - elem)**2
return np.sqrt(result)
@jit(nopython=True)
def sqrt_err_relative_tucker(coo_tensor, vals, shape, a, b, c, g):
result = sqrt_err_tucker(coo_tensor, vals, shape, a, b, c, g)
return result / np.sqrt((vals**2).sum())
# -
rerr = sqrt_err_relative_tucker(
coords,
values,
data_shape,
p,
q,
r,
g,
)
print(f"Relative error = {rerr}")
# +
@jit(nopython=True)
def hit_rate_tucker_raw(test_triples, a, b, c, g,
how_many=10, iter_show=False, freq=3000):
total = len(test_triples)
hit = 0
iteration = 0
for entity in test_triples:
iteration += 1
p = entity[0]
q = entity[1]
r = entity[2]
candidate_values = np.sum(g[p, q, :] * a[p, :] * b[q, :] * c, axis=1)
candidate_values = sigmoid(candidate_values)
top = np.argsort(candidate_values)[::-1][:how_many]
for x in top:
if x == r:
hit += 1
if iter_show:
if iteration % freq == 0:
print(hit / iteration, hit, iteration)
return hit / total, hit, total
hit_rate_tucker_raw(
test_triples[:5],
p, q, r, g, 10,
iter_show=False,
freq=2,
)
# -
hit_rate_tucker_raw(
test_triples,
p, q, r, g, 10,
iter_show=True,
freq=1000,
)
| 8,755 |
/06/2. Ensemble learning.ipynb | ffcc02eb8df5138deb48a63c3f7d0f358dac3909 | [] | no_license | robom/vos2-2018 | https://github.com/robom/vos2-2018 | 4 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 571,545 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Obsah prednasky
#
# ### myslienka a zmysel ucenia suborom metod
#
# ### bagging
#
# #### hlasovanie
#
# #### random forest
#
# ### boosting
#
# ### stacking
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn
plt.rcParams['figure.figsize'] = 9, 6
from IPython.display import Image
# -
# # Zakladna myslienka
#
# natrenovat viacero klasifikatrov (alebo inych algoritmov pre ine ulohy analyzy dat) a ich vysledky spojit
#
# ked kombinujeme viacero nezavyslich a rozdielnych klasifikatorov, kde kazdy je aspon o trochu lepsi ako nahodne hadanie, tak nahodne chyby sa medzi sebou vyrusia a spravne predpovede sa posilnia
# # Bias - variance
#
# Ked trenujeme klasifikator, tak nasim cielom nie je naucit sa naspamat trenovacie data. Snazime sa naucit parametre procesy za datami, ktore tieto data generovali.
#
# toto je dolezite pre to, aby bol algoritmus dobre generalizovatelny tj. aby bol pouzitelny aj na datach, ktore nikdy nevidel a ktore boli generovane rovnakym procesom.
#
# Musime sa vysporiadat s kompromisom medzi bias a variance
#
# +
from IPython.display import YouTubeVideo
YouTubeVideo('W0NLs-A6hhQ')
# -
#
# # Bias (vychylenost)
# model, ktory je prilis jednoduchy alebo neflexibilny bude mat prilis velky bias
# bude mat velmi velku chybu na trenovacich aj testovacich datach. Tato chyba bude ale velmi stabilna. Ak mu dame data, ktore nikdy nevidel, tak bude fungovat priblizne rovnako dobre
#
# # Variance (premenlivost)
# zalezi od zlozitosti modelu
#
# zlozity model sa velmi dobre nauci trenovacie data a bude mat na nich malu chybu. Akonahle mu ale dame data, ktore nevidel, tak ta chyba bude velmi premenliva a moze byt obrovska.
#
# Obe tieto hodnoty potrebujeme znizit. Chceme presny klasifikator, ktory bude davat dobre vysledky aj na datach, ktore nikdy nevidel.
Image('./img/bias-variance.png', width='600px')
# # Na redukovanie biasu a variance sa casto pouziva ensemble learning
#
# **bagging** na redukovanie variancie
#
# **boosting** na redukovanie biasu
#
#
# # Bagging (Bootstrap aggregating)
#
# Priemerovanie redukuje varianciu. Priemer predikcii je stabilnejsi ako jednotlive predikcie.
#
# Pouzivame mnozstvo modelov.
#
# Otazka je: Ako ich natrenujeme na jednom datasete?
#
# Kazdy je trenovany na nahodnom vybere (with replacement) z dat.
#
# Vysledna predikcia je tvorena priemerom predikcii jednotlivych modelov
#
# Mozu sa pouzivat rovnake typy modelov. Dolezite ale je aby boli nekorelovane. Len nekorelovane modely sa totiz mozu doplnat. Ak by korelovali, tak sa budu navzajom podporovat a to len pokazi celu myslienku baggingu.
#
# Rozne postupy na zabezpecenie nekorelovanosti modelov:
# * rozne algoritmy na trenovanie modelov
# * sampling
# * feature bagging
# * zmena reprezentacie atributov
# * zapojenie nahody (sum, nahodne parametre ...)
# * ...
#
#
# Najcastejsi priklad je random forest (tu sa ale pouziva feature bagging, pretoze ked sa pouziva bagging na riadkoch, tak casto vznikaju silno korelovane stromy)
# # Metody kombinovania vysledkov modelov
#
# * vahovanie
# > uzitocne, ked jedlotlive modely maju porovnatelnu uspesnost
#
# * meta-learning (vid stacking)
# > uzitocne ak niektore modely konzistentne spravne predikuju niektore instancie
# # Rozne sposoby vahovania
#
# * **majority voting** - pri klasifikacii je trieda predikovanej instancie ta, ktoru oznacila vacsina modelov
# * **performance weighting** - vaha kazdeho modelu je pri hlasovani urcena jeho uspesnostou
# * **heuristic weighted voting** - optimalizacna metoda pouzita na natrenovanie vah (pozor na pretrenovanie)
# * **Bayesovska kombinacia** - vaha je pravdepodobnsot, ze data splnaju nejaku hypotezu (model/nastavenie modelu). Skusaju sa rozne nastavenia parametrov
# # Priklad vahovanie
#
# pouzijeme rozne algoitmy na zabezpecenie toho, ze kombinovane modely su na sebe nezavysle
#
# http://sebastianraschka.com/Articles/2014_ensemble_classifier.html
# +
from sklearn import datasets
iris = datasets.load_iris()
X, y = iris.data[:, 1:3], iris.target
# +
from sklearn import cross_validation
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
import numpy as np
np.random.seed(123)
clf1 = LogisticRegression()
clf2 = RandomForestClassifier()
clf3 = GaussianNB()
print('5-fold cross validation:\n')
for clf, label in zip([clf1, clf2, clf3], ['Logistic Regression', 'Random Forest', 'naive Bayes']):
scores = cross_validation.cross_val_score(clf, X, y, cv=5, scoring='accuracy')
print("Accuracy: %0.2f (+/- %0.2f) [%s]" % (scores.mean(), scores.std(), label))
# +
from sklearn.base import BaseEstimator
from sklearn.base import ClassifierMixin
import numpy as np
import operator
class EnsembleClassifier(BaseEstimator, ClassifierMixin):
"""
Ensemble classifier for scikit-learn estimators.
Parameters
----------
clf : `iterable`
A list of scikit-learn classifier objects.
weights : `list` (default: `None`)
If `None`, the majority rule voting will be applied to the predicted class labels.
If a list of weights (`float` or `int`) is provided, the averaged raw probabilities (via `predict_proba`)
will be used to determine the most confident class label.
"""
def __init__(self, clfs, weights=None):
self.clfs = clfs
self.weights = weights
def fit(self, X, y):
"""
Fit the scikit-learn estimators.
Parameters
----------
X : numpy array, shape = [n_samples, n_features]
Training data
y : list or numpy array, shape = [n_samples]
Class labels
"""
for clf in self.clfs:
clf.fit(X, y)
def predict(self, X):
"""
Parameters
----------
X : numpy array, shape = [n_samples, n_features]
Returns
----------
maj : list or numpy array, shape = [n_samples]
Predicted class labels by majority rule
"""
self.classes_ = np.asarray([clf.predict(X) for clf in self.clfs])
if self.weights:
avg = self.predict_proba(X)
maj = np.apply_along_axis(lambda x: max(enumerate(x), key=operator.itemgetter(1))[0], axis=1, arr=avg)
else:
maj = np.asarray([np.argmax(np.bincount(self.classes_[:,c])) for c in range(self.classes_.shape[1])])
return maj
def predict_proba(self, X):
"""
Parameters
----------
X : numpy array, shape = [n_samples, n_features]
Returns
----------
avg : list or numpy array, shape = [n_samples, n_probabilities]
Weighted average probability for each class per sample.
"""
self.probas_ = [clf.predict_proba(X) for clf in self.clfs]
avg = np.average(self.probas_, axis=0, weights=self.weights)
return avg
# +
np.random.seed(123)
eclf = EnsembleClassifier(clfs=[clf1, clf2, clf3], weights=[1,1,1])
for clf, label in zip([clf1, clf2, clf3, eclf], ['Logistic Regression', 'Random Forest', 'naive Bayes', 'Ensemble']):
scores = cross_validation.cross_val_score(clf, X, y, cv=5, scoring='accuracy')
print("Accuracy: %0.5f (+/- %0.5f) [%s]" % (scores.mean(), scores.std(), label))
# -
# #### Mozeme skusit ovahovat klasifikatory ich uspesnostou
weights = []
for clf, label in zip([clf1, clf2, clf3], ['Logistic Regression', 'Random Forest', 'naive Bayes']):
scores = cross_validation.cross_val_score(clf, X, y, cv=5, scoring='accuracy')
weights.append(scores.mean())
weights
# +
np.random.seed(123)
eclf = EnsembleClassifier(clfs=[clf1, clf2, clf3], weights=weights)
for clf, label in zip([clf1, clf2, clf3, eclf], ['Logistic Regression', 'Random Forest', 'naive Bayes', 'Ensemble']):
scores = cross_validation.cross_val_score(clf, X, y, cv=5, scoring='accuracy')
print("Accuracy: %0.5f (+/- %0.5f) [%s]" % (scores.mean(), scores.std(), label))
# -
# #### Mozeme tiez skusit implementaciu vahovacieho ensemblu priamo z sklearn
# +
np.random.seed(10)
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import VotingClassifier
# Training classifiers
clf1 = LogisticRegression()
clf2 = RandomForestClassifier()
clf3 = GaussianNB()
eclf = VotingClassifier(estimators=[('LR', clf1), ('RF', clf2), ('NB', clf3)], voting='hard', weights=[1,1,1])
scores = cross_validation.cross_val_score(eclf, X, y, cv=5, scoring='accuracy')
scores.mean()
# -
# #### Mozeme pouzit grid search na najdenie parametrov
# +
from sklearn.model_selection import GridSearchCV
clf1 = LogisticRegression(random_state=1)
clf2 = RandomForestClassifier(random_state=1)
clf3 = GaussianNB()
eclf = VotingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('gnb', clf3)], voting='soft')
params = {'lr__C': [1.0, 100.0], 'rf__n_estimators': [20, 200],
'weights': [[1,1,1], [0,1,1], [10,1,1]]}
grid = GridSearchCV(estimator=eclf, param_grid=params, cv=5)
grid = grid.fit(iris.data, iris.target)
# -
grid.grid_scores_
grid.cv_results_
# # á propos grid search
#
# parametre sa daju nastavovat aj nahodne ak je ich priestor prilis velky
#
# ```
# params_dist_grid = {
# 'max_depth': [1, 2, 3, 4],
# 'gamma': [0, 0.5, 1],
# 'n_estimators': randint(1, 1001), # uniform discrete random distribution
# 'learning_rate': uniform(), # gaussian distribution
# 'subsample': uniform(), # gaussian distribution
# 'colsample_bytree': uniform() # gaussian distribution
# }
# ```
#
# ```
# rs_grid = RandomizedSearchCV(
# estimator=RandomForest(**params_fixed, seed=seed),
# param_distributions=params_dist_grid,
# n_iter=10, # dolezite je nastavit pocet iteracii
# cv=cv,
# scoring='accuracy',
# random_state=seed
# )
# rs_grid.fit(X, y)
#
# ```
#
#
#
# ## GridSearch alternativy
#
# [https://sigopt.com/no-grid-search](https://sigopt.com/no-grid-search)
# # Priklad Bagging
#
# Pouzijeme ten isty algoritmus
#
# Pouzijeme sampling na to, aby sme zabezpecili, ze budu modely na sebe nezavysle
#
# priklad z: https://github.com/rasbt/python-machine-learning-book
Image(filename='./img/bagging.png', width=400)
# +
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier(criterion='entropy',
max_depth=None, # nastavime hlbku na maximum, takze mu nijak nebudeme branit v pretrenovani
random_state=1)
bag = BaggingClassifier(base_estimator=tree,
n_estimators=500,
max_samples=1.0,
max_features=1.0,
bootstrap=True,
bootstrap_features=False, # toto je rozdiel oproti random forest
n_jobs=1,
random_state=1)
# -
scores = cross_validation.cross_val_score(bag, X, y, cv=5, scoring='accuracy')
scores.mean()
# # ked si natrenujeme len jeden model, tak uspesnost asi bude mensia
scores = cross_validation.cross_val_score(tree, X, y, cv=5, scoring='accuracy')
scores.mean()
# Tato nizsia uspesnost je pravdepodobne sposobena tym, ze sa model pretrenoval. Baging mal dobru uspesnost, lebo pretrenovanie zmiernuje.
# # co sa stane ak budem stromu aspon trochu brannit v pretrenovani
tree2 = DecisionTreeClassifier(criterion='entropy',
max_depth=2, # nenecham ho naucit sa tie data uplne naspamat
random_state=1)
scores = cross_validation.cross_val_score(tree2, X, y, cv=5, scoring='accuracy')
scores.mean()
# baggingom teda viem bojovat s pretrenovanim, teda s varianciou modelu
# # Priklad Random forest
#
# tento algoritmus som uz spominal vela krat, takze to nebudem natahovat
#
# na rozdiel od klasickeho baggingu sa pouziva feature bagging
#
# kedze nie kazda vlastnost je pouzita na kazdy strom, tak sa da pozriet, ktore najviac prispievaju k presnosti klasifikacie (regresie)
#
# To sa da pouzit na vyber najuzitocnejsich atributov (ukazoval som minule)
#
# Dost casto sa to pouziva v spojeni so spracovanim obrazkov na zobrazenie ktore casti obrazku su najdolezitejsie.
#
# Pozor!!! aby ste toto mohli spravit, tak musite mat velmi kvalitne predpriprveny dataset. Vo vseobecnosti je lepsie nepozerat na konkretne pixely ale na vlastnosti vybrate z obrazku napriklad pomocou metody SIFT alebo Hough transformacie.
#
# Ak mate totiz nejake posunutie, rotaciu alebo inak transformovany obrazok, tak konkretne pixely vam toho vela nepovedia
#
# +
from sklearn.datasets import fetch_olivetti_faces
from sklearn.ensemble import ExtraTreesClassifier
# Number of cores to use to perform parallel fitting of the forest model
n_jobs = 1
# Load the faces dataset
data = fetch_olivetti_faces()
X = data.images.reshape((len(data.images), -1))
y = data.target
mask = y < 5 # Limit to 5 classes
X = X[mask]
y = y[mask]
# -
plt.imshow(data.images[0], cmap=plt.cm.gray)
# Tu mam dataset, kde su obrazky pekne centrovane, rovnakej velkosti a bez nejakej deformacie
#
# Cize dolezite pixely by mohli byt na rovnakom mieste
# +
# %%time
# Build a forest and compute the pixel importances
print("Fitting ExtraTreesClassifier on faces data with %d cores..." % n_jobs)
forest = ExtraTreesClassifier(n_estimators=1000,
max_features=128,
n_jobs=n_jobs,
random_state=0)
forest.fit(X, y)
importances = forest.feature_importances_
importances = importances.reshape(data.images[0].shape)
# Plot pixel importances
plt.matshow(importances, cmap=plt.cm.hot)
plt.title("Pixel importances with forests of trees")
# -
# # Boosting
#
# Inkrementalne vyrabame nove klasifiktory, ktore dostavaju tie pozorovania, ktore boli predtym zle klasifikovane s vacsou vahou.
#
# Typicky priklad AdaBoost (pekne vysvetlenie aj so vzorcami na vypocet vah http://machinelearningmastery.com/boosting-and-adaboost-for-machine-learning/)
# ## typicky postup
#
# 1. nastav vahu kazdemu trenovaciemu pripadu rovnomerne
# 2. natrenuj model slabim klasifikatorom
# 3. vypocitaj chybu natrenovaneho modelu na trenovacom datasete (tato chyba sa pouzije na vahovanie celeho modelu)
# 4. zmen vahy trenovacm pripadom. Zvys vahu tomu, ktory bol zle klasifikovany (ak algoritmus nevie pouzivat vahy, tak pouzi resampling)
# 5. skoc na krok 2 az dokedy nespravis maximalny pocet iteracii
# * vysledy model su vahovane predikcie ciastkovych modelov (vaha z kroku 3)
#
# Bagging ma problemy so sumom a outliermi. Snazi sa totiz spravne klasifikovat vsetko. Riziko pretrenovania
#
# tazsie sa paralelizuje ako bagging
#
# da sa pouzit early stop pri prdikcii ak si je niektory model v postupnosti dostatocne isty svojou predikciou
Image(filename='./img/boosting.png', width=600)
# +
df_wine = pd.read_csv('https://archive.ics.uci.edu/ml/'
'machine-learning-databases/wine/wine.data',
header=None)
df_wine.columns = ['Class label', 'Alcohol', 'Malic acid', 'Ash',
'Alcalinity of ash', 'Magnesium', 'Total phenols',
'Flavanoids', 'Nonflavanoid phenols', 'Proanthocyanins',
'Color intensity', 'Hue', 'OD280/OD315 of diluted wines',
'Proline']
# drop 1 class
df_wine = df_wine[df_wine['Class label'] != 1]
y = df_wine['Class label'].values
X = df_wine[['Alcohol', 'Hue']].values
# +
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
le = LabelEncoder() # toto mi opravi poradie tried. Ak som tam mal predtym napriklad -1, 7, 10 ..., tak potom budem mat 0,1,2 ...
y = le.fit_transform(y)
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.40,
random_state=1)
# +
from sklearn.ensemble import AdaBoostClassifier
tree = DecisionTreeClassifier(criterion='entropy',
max_depth=1, # nastavim hlbku na 1 aby sa trenovalo prave jedno pravidlo
random_state=0)
ada = AdaBoostClassifier(base_estimator=tree,
n_estimators=100,
learning_rate=0.1,
random_state=0)
# +
# teraz naschval nebudem pouzivat cross validaciu, aby sme sa mohli pozriet ako sa modelu dari na testovacej aj trenovacej vzorke
from sklearn.metrics import accuracy_score
tree = tree.fit(X_train, y_train)
y_train_pred = tree.predict(X_train)
y_test_pred = tree.predict(X_test)
tree_train = accuracy_score(y_train, y_train_pred)
tree_test = accuracy_score(y_test, y_test_pred)
print('Decision tree train/test accuracies %.3f/%.3f'
% (tree_train, tree_test))
ada = ada.fit(X_train, y_train)
y_train_pred = ada.predict(X_train)
y_test_pred = ada.predict(X_test)
ada_train = accuracy_score(y_train, y_train_pred)
ada_test = accuracy_score(y_test, y_test_pred)
print('AdaBoost train/test accuracies %.3f/%.3f'
% (ada_train, ada_test))
# -
# to, ze je uspesnost adaboost 100% na trenovacej sade moze zavanat pretrenovanim
#
# to, ze je chybovost na testovacej taka velka to potvrdzuje
# ### skusme si nakreslit ako sa v skutocnosti ten priestor rozdeluje
# +
x_min = X_train[:, 0].min() - 1
x_max = X_train[:, 0].max() + 1
y_min = X_train[:, 1].min() - 1
y_max = X_train[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.1),
np.arange(y_min, y_max, 0.1))
f, axarr = plt.subplots(nrows=1, ncols=2,
sharex='col',
sharey='row',
figsize=(12, 7))
for idx, clf, tt in zip([0, 1],
[tree, ada],
['Decision Tree', 'Boosting']):
clf.fit(X_train, y_train)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
axarr[idx].contourf(xx, yy, Z, alpha=0.3)
axarr[idx].scatter(X_train[y_train == 0, 0],
X_train[y_train == 0, 1],
c='blue', marker='^')
axarr[idx].scatter(X_train[y_train == 1, 0],
X_train[y_train == 1, 1],
c='red', marker='o')
axarr[idx].set_title(tt)
axarr[0].set_ylabel('Alcohol', fontsize=12)
plt.text(10.2, -1.2,
s='Hue',
ha='center', va='center', fontsize=12)
plt.tight_layout()
# -
# # naozaj ak je velky pocet modelov, tak sa to velmi rychlo pretrenuje
#
# skusme sa pohrat s tym, kolko modelov sa pouziva na boosting alebo s learning rate
# Inak je ale tato metoda velmi uzitocna ak mame naprikald nevyvazene triedy alebo je v datach viacero roznych pravidiel, na zaklade ktorych sa rozdeluju data
# # Stacking
#
# vsetky data su pouzite na natrenovanie roznych modelov
#
# Vysledky modelov su pridane k datam a pouzite na natrenovanie dalsej urovne modelov.
#
# Na kominovanie sa najcastejsie pouziva logisticka regresia.
# Sklearn nema implementaciu stackingu. Skusme si ale naimplementovat primitivny stacking sami.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.40, random_state=1) # X a y stale obsahuje dataset vin
# +
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.linear_model import LogisticRegression
base_algorithms = [('svm', SVC(probability=True)),
('RF', RandomForestClassifier(n_estimators=100)),
('ET', ExtraTreesClassifier(n_estimators=100))]
# stacking_train_dataset = pd.DataFrame()
# stacking_test_dataset = pd.DataFrame()
stacking_train_dataset = pd.DataFrame(X_train) # mozu sa pouzit povodne data ako zaklad a knim sa pridaju predikcie
stacking_test_dataset = pd.DataFrame(X_test)
for name, algorithm in base_algorithms: # mozu sa tu pridat aj pravdepodobnosti. Tiem odely to ale musia podporovat
stacking_train_dataset[name] = algorithm.fit(X_train, y_train).predict(X_train)
# stacking_train_dataset[name+"_proba"] = algorithm.fit(X_train, y_train).predict_proba(X_train)
stacking_test_dataset[name] = algorithm.predict(X_test)
# stacking_test_dataset[name+"_proba"] = algorithm.predict_proba(X_test)
# -
stacking_train_dataset.head()
combiner_algorithm = LogisticRegression(max_iter=500)
final_predictions = combiner_algorithm.fit(stacking_train_dataset, y_train).predict(stacking_test_dataset)
accuracy_score(y_test, final_predictions)
accuracy_score(y_test, stacking_test_dataset.ET)
# kasdy z jednotlivych klasifikatorov ma horsiu uspesnost ako ich spojenie
# # Sklearn nema priamo implementaciu pre stacking, ale balicek Brew ano
# +
import brew
from brew.base import Ensemble
from brew.combination.combiner import Combiner
from brew.stacking.stacker import EnsembleStack
from brew.stacking.stacker import EnsembleStackClassifier
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.linear_model import LogisticRegression
layer_1 = [SVC(probability=True),
RandomForestClassifier(n_estimators=100),
ExtraTreesClassifier(n_estimators=100)]
layer_2 = [SVC(probability=True), LogisticRegression(max_iter=500)]
stack = EnsembleStack(cv=10) # number of folds per layer
stack.add_layer(Ensemble(layer_1))
stack.add_layer(Ensemble(layer_2))
clf = EnsembleStackClassifier(stack, Combiner('mean'))
scores = cross_validation.cross_val_score(tree2, X, y, cv=5, scoring='accuracy')
scores.mean()
# -
# tuna som pouzil malicky dataset vin a strasne vela klasifikatorov
#
# pridal som do datasetu vela novych atributov a teda som sposobil, ze model ma celkom seriozny problem sa z toho mnozstva atributov nieco naucit. Uzitocnejsie by to bolo pouzit na nejaky vacsi dataset.
# # Dalsie citanie
#
# http://mlwave.com/kaggle-ensembling-guide/
# # Pipelining
#
# Uz len posledna vec na zaver.
#
# Tu je velmi pekny tutorial, takze budem prezentovat priamo z neho
#
# Nijak to ale neznizuje dolezitost tejto temy. Prave naopak, toto je velmi uzitocne.
#
# http://machinelearningmastery.com/automate-machine-learning-workflows-pipelines-python-scikit-learn/
#
# http://zacstewart.com/2014/08/05/pipelines-of-featureunions-of-pipelines.html
#
# http://scikit-learn.org/stable/modules/pipeline.html
# # Nevyvazene data
#
# [Imbalanced learn](https://imbalanced-learn.readthedocs.io/en/stable/)
#
# [priklad roznych metod](https://imbalanced-learn.readthedocs.io/en/stable/auto_examples/over-sampling/plot_comparison_over_sampling.html#sphx-glr-auto-examples-over-sampling-plot-comparison-over-sampling-py)
| 23,319 |
/abalone.ipynb | f144de67d1eb0f45092c5a98169af8d479a3da9e | [] | no_license | Attonasi/mach_learning_midproj | https://github.com/Attonasi/mach_learning_midproj | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 234,348 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from datetime import datetime
import matplotlib.pyplot as plt
plt.style.use('seaborn-darkgrid')
# -
# ## Glipse Data
# Read data
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
# btc = pd.read_csv('/kaggle/input/bitcoin-historical-data/bitstampUSD_1-min_data_2012-01-01_to_2020-09-14.csv')
btc = pd.read_csv('../input/bitcoin-historical-data/bitstampUSD_1-min_data_2012-01-01_to_2020-12-31.csv')
# Converting the Timestamp column from string to datetime
btc['Timestamp'] = [datetime.fromtimestamp(x) for x in btc['Timestamp']]
btc = btc.set_index('Timestamp')
# -
# #### Time resampling
btc_daily = btc.resample("24H").mean() #daily resampling
# #### Time Series Decomposition & Statistical Tests
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.tsa.stattools import kpss
from statsmodels.tsa.stattools import adfuller
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
# +
def fill_missing(df):
### function to impute missing values using interpolation ###
df['Open'] = df['Open'].interpolate()
df['Close'] = df['Close'].interpolate()
df['Weighted_Price'] = df['Weighted_Price'].interpolate()
df['Volume_(BTC)'] = df['Volume_(BTC)'].interpolate()
df['Volume_(Currency)'] = df['Volume_(Currency)'].interpolate()
df['High'] = df['High'].interpolate()
df['Low'] = df['Low'].interpolate()
print(df.head())
print(df.isnull().sum())
fill_missing(btc_daily)
# +
plt.figure(figsize=(15,12))
series = btc_daily.Weighted_Price
result = seasonal_decompose(series, model='additive',period=1)
result.plot()
# +
# Renaming the column names accroding to Prophet's requirements
daily_data_fb = btc_daily.reset_index()[['Timestamp','Weighted_Price']].rename({'Timestamp':'ds','Weighted_Price':'y'}, axis=1)
daily_data_fb.head()
# +
test_length = 365
split_date = "2020-01-01"
train_filt = daily_data_fb['ds'] <= split_date
test_filt = daily_data_fb['ds'] > split_date
train_fb = daily_data_fb[train_filt]
test_fb = daily_data_fb[test_filt]
print("train data shape :", train_fb.shape[0] / daily_data_fb.shape[0] * 100)
print("test data shape :", test_fb.shape[0] / daily_data_fb.shape[0] * 100)
# -
# 
#
# [Auto_TS: Auto_TimeSeries](https://github.com/AutoViML/Auto_TS)
# !pip install auto-ts
# # !pip install auto_ts --ignore-installed --no-cache-dir
# !pip install tscv
# +
from auto_ts import auto_timeseries as ATS
# -
train_fb = train_fb.reset_index()
train_fb.head()
test_fb = test_fb.reset_index()
test_fb.head()
# +
automl_model = ATS(score_type='rmse',
time_interval='D', forecast_period=30,
non_seasonal_pdq=None, seasonality=False, seasonal_period=12,
model_type=['best'],
verbose=2)
# -
ts_column = 'ds'
target = 'y'
sep = ','
automl_model.fit(
traindata=train_fb,
ts_column=ts_column,
target=target,
cv=5,
sep=',')
# ### Evaluate Results
automl_model.get_leaderboard()
# +
automl_model.plot_cv_scores()
# -
# There is a Bug with Prophet folds: https://github.com/AutoViML/Auto_TS/issues/25
automl_model.get_cv_scores()
results_dict = automl_model.get_ml_dict()
results_dict['Prophet']['forecast'][:5]
# +
# results_dict['auto_SARIMAX']['forecast'][:5]
# -
# #### Forecast Results
# Using Best Model
future_predictions = automl_model.predict(
testdata=test_fb
)
future_predictions
# Using specific model
future_predictions = automl_model.predict(
testdata=test_fb,
model='auto_SARIMAX'
)
future_predictions
# +
# Using specific model
future_predictions = automl_model.predict(
testdata=test_fb,
model='Prophet',
simple=False
)
future_predictions
# -
test_fb.head()
import copy
subm = test_fb[[target]]
print(subm.shape)
subm.head()
subm['predictions'] = future_predictions['yhat'].values
subm[[target,'predictions']].plot(figsize=(15,8))
start
# Ctrl-End go to cell end Ctrl-Down go to cell end
# Ctrl-Left go one word left Ctrl-Right go one word right
# Ctrl-Backspace delete word before Ctrl-Delete delete word after
# Esc command mode Ctrl-M command mode
# Shift-Enter run cell, select below Ctrl-Enter run cell
# Alt-Enter run cell, insert below Ctrl-Shift-Subtract split cell
# Ctrl-Shift-- split cell Ctrl-S Save and Checkpoint
# Ctrl-/ toggle comment on current or selected lines
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.preprocessing import OneHotEncoder
from sklearn import linear_model
from sklearn.cross_validation import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn import metrics
from sklearn.model_selection import ShuffleSplit
from sklearn import svm
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import accuracy_score
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import GradientBoostingRegressor
path = '/home/collin/machine_learning/CS6140ML18Fall-smith.colin/midterm/data/'
data_file = 'data_abalone.txt'
# -
data = pd.read_csv(path+data_file, delimiter=',', names=['Sex', 'Length', 'Diameter', 'Height', 'Whole Wt', 'Shucked Wt', 'Viscera Wt', 'Shell Wt', 'Rings'])
# This is a very clean dataset. Not much to add or manipulate.
#
# I am going to try two datasets. One with all of the groups and one with [Length, Shucked Wt, Viscera Wt] removed. Using the correlation table below the weights are all cross correlative though the Shell Weight is actually not that bad so I am leaving it in. Length and Diameter however approach 1. So I am dropping one of those in the second set.
#
# I also need to one hot the Sex column because it is not useful as an object and drop that column afterwords.
data.corr()
data.info()
# enc = OneHotEncoder(handle_unknown = 'ignore')
data_dummies = pd.get_dummies(data['Sex'])
data = pd.concat([data, data_dummies], axis=1)
data = data.drop(columns=['Sex'])
# I commented these scatter plots out but they gave some insight into their relation to the Rings column.
# +
# names=['Length', 'Diameter', 'Height', 'Whole Wt', 'Shucked Wt', 'Viscera Wt', 'Shell Wt', 'Rings']
# for name in names:
# data.plot(kind='scatter', x=name, y='Rings')
# -
reg = linear_model.LinearRegression()
log = linear_model.LogisticRegression()
# As I continue to look up things everyone already knows I got to StandardScalar. And normalizer.
# +
data_y = data['Rings']
scaler = StandardScaler()
cols = data.columns.tolist()
dataxX = pd.DataFrame(scaler.fit_transform(data))
dataxX.columns = cols
data1 = data.drop(columns=['Rings'])
data2 = data.drop(columns=['Shucked Wt', 'Viscera Wt', 'Length', 'Rings'])
datax = dataxX.drop(columns=['Rings'])
# -
# #### Being senstive to criticism from the other dataset about choosing baseline I am redoing the same assessment on this dataset.
def get_plusmin_23(list):
pm_2 = 0
pm_3 = 0
for i in range(len(hist_y)):
temp2 = 0
temp3 = 0
if(len(hist_y) < 6):
print ('The list is short. You can figure this out yourself.')
return 0, 0;
if i < 2:
temp2 = sum(hist_y[0:4])
elif i < 3:
temp2 = sum(hist_y[i-2:i+2])
temp3 = sum(hist_y[0:6])
else:
temp2 = sum(hist_y[i-2:i+3])
temp3 = sum(hist_y[i-3:i+4])
if temp2 > pm_2:
pm_2 = temp2
if temp3 > pm_3:
pm_3 = temp3
return pm_2, pm_3
# +
hist_y = []
for i in range (30):
hist_y.append(0)
for i in range (len(data_y)):
hist_y[data_y[i]] += 1
pm_2, pm_3 = get_plusmin_23(hist_y)
print (pm_2/len(data_y))
print (pm_3/len(data_y))
# -
# ### Given the above array we have 689, 634, 568, 487, 391 as the top 5 years. We can get 2769 +/-2 years by guessing 9 and including 10, 8, 11, 7 and get 66%.
#
# ### We will set that for baseline to judge usefulness.
(689+634+568+487+391) / 4177
x_train, x_test, y_train, y_test = train_test_split(data1, data_y, test_size=0.2, random_state=4)
x2_train, x2_test, y2_train, y2_test = train_test_split(data2, data_y, test_size=0.2, random_state=4)
# At this point the data is set up. The answer column was dropped. There are 2 data sets one with all of the columns and another with three columns that had strong cross correlation dropped. I have divided them into two test/training sets.
#
# I have a linear regression model and a logistic regression model set up. Time to fit and predict!
# +
reg.fit(x_train, y_train)
y_pred = reg.predict(x_test)
reg.fit(x2_train, y2_train)
y2_pred = reg.predict(x2_test)
log.fit(x_train, y_train)
log_y_pred = log.predict(x_test)
log.fit(x2_train, y2_train)
log2_y_pred = log.predict(x2_test)
# -
# I wrote a class called Result. I posted the code on Piazza and it is in a .py file right next to this notebook. I am importing it here and created a Result book for each of the regression predictions here.
# +
from result_lib import Result
lin_one_result = Result(y_pred, y_test.values, 1)
lin_two_result = Result(y2_pred, y2_test.values, 1)
log_one_result = Result(log_y_pred, y_test.values, 1)
log_two_result = Result(log2_y_pred, y2_test.values, 1)
# -
# The first tool the class I wrote gave me is a way to see basic numbers. What is the range, the number that are within that range of the predicted value, and most importantly the mean variance, i.e. how far off the guess was.
lin_two_result.display_totals()
lin_one_result.display_totals()
log_one_result.display_totals()
log_two_result.display_totals()
# While that will eventually be useful this next tool is at the moment the best one for taking next steps. I make a list of the natural values of the variances I plot on the X and I can plot them against the true values which I put on the Y. i.e. For each true value I can show how far off and in which direction the regression erred.
#
# After taking one look at the graphs the next step is not so obvious.
plt.plot(y2_test.values, lin_two_result.list_variances, 'o')
plt.plot(y_test.values, lin_one_result.list_variances, 'o')
plt.plot(y_test.values, log_one_result.list_variances, 'o')
plt.plot(y_test.values, log_two_result.list_variances, 'o')
#
plt.plot(log2_y_pred, log_two_result.list_variances, 'o')
plt.plot(y_test.values, log2_y_pred, 'o')
plt.plot(y_test.values, y_pred, 'o')
# After looking at the graphs and judging by performance here is the following observation:
#
# Abalone are all very much the same size.
#
# Deep I know.
#
# The graph just above this is a plot of the predicted values versus the actual values for the second linear regression. At the age of 5 we predict a little low on age. At the age of 7-12 we are very close. After that we predict they are 10 or 11. I think they just stop growing. Looking at the given data everything has to do with size.
#
# Below I use the get confidence tool and I see linear one drops us from +/-2 to +/-1.67 to get 66%. They both get to 73% at +/-2 so it is slightly better than noise.
#
# Both log regressions get 80% at +/-2 rather than 66%
#
# Judging these Linear Regression using the entire dataset seems to perform the best.
#
# Below I use the result class to get confidence intervals near +/-2.
lin_two_result.get_confidence_interval(73)
lin_one_result.get_confidence_interval(73)
log_two_result.get_confidence_interval(80)
log_one_result.get_confidence_interval(80)
cv_lin2 = cross_val_score(reg, data2, data_y, cv=5)
cv_lin1 = cross_val_score(reg, data1, data_y, cv=5)
# cv_log2 = cross_val_score(log, data2, data_y, cv=5)
# cv_log1 = cross_val_score(log, data1, data_y, cv=5)
# This section is going to be about implementing Support Vector Machines.
#
# Basic first and using the one-to-one .SCV classifier.
svm_lin1 = svm.SVC(kernel='linear', C=1).fit(x_train, y_train)
svm_lin2 = svm.SVC(kernel='linear', C=1).fit(x2_train, y2_train)
svm_lin1.score(x_test, y_test)
svm_lin2.score(x2_test, y2_test)
# This makes no sense. So Result.
# +
svm_pred1 = svm_lin1.predict(x_test)
svm_pred2 = svm_lin2.predict(x2_test)
svm1_result = Result(svm_pred1, y_test.values, 1)
svm2_result = Result(svm_pred2, y2_test.values, 1)
# -
svm1_result.display_totals()
svm2_result.display_totals()
plt.plot(y_test.values, svm_pred1, 'o')
plt.plot(y_test.values, svm1_result.list_variances, 'o')
# Same pattern. The predictions guess too high on young specimens and too low on old.
#
# Mostly too low on old.
#
# There are a lot of SVM options and a variety of ways to format them so I am going to try something a little different.
# +
x_trains = [x_train, x2_train]
y_trains = [y_train, y2_train]
kernals = ['linear', 'rbf', 'sigmoid']
for krnl in kernals:
for power in range(-5, 10):
svm_temp = svm.SVC(kernel=krnl, C=2**power).fit(x_train, y_train)
y_pred=svm_temp.predict(x_test)
res_temp = Result(y_pred, y_test.values, 1)
print('Kernal: ' + krnl + ' C: 2**' + str(power))
res_temp.display_totals()
# -
# Looking at the above results a couple things stick out. svm rbf mean variance was still descending at C 2^9 and sigmoid with C 2^-5 looks like a nutball outlier. It was also the best result. The linear SVM bottomed out around C 2^7.
#
# So.
# +
svm1_sig_n5 = svm.SVC(kernel='sigmoid', C=2**-5).fit(x_train, y_train)
svm2_sig_n5 = svm.SVC(kernel='sigmoid', C=2**-5).fit(x2_train, y2_train)
sig1_n5_pred = svm1_sig_n5.predict(x_test)
sig2_n5_pred = svm2_sig_n5.predict(x2_test)
# -
# Classic. The best variance we got was just predicting 9 every time. Turn the error to approach 0 and you get the mean.
#
# But you have to try.
plt.plot(sig1_n5_pred, y_test.values, 'o')
# Now for rbf. I want to see what error value approaches the bottom.
for power in range(7, 18):
svm_temp = svm.SVC(kernel='rbf', C=2**power).fit(x_train, y_train)
y_pred=svm_temp.predict(x_test)
res_temp = Result(y_pred, y_test.values, 1)
print('Kernal: ' + 'rbf' + ' C: 2**' + str(power))
res_temp.display_totals()
# The numbers are getting ridiculous. I am typing this while I wait. I see the start of a bottom around C 2^14. I might kill this off. I am going to dig into C 2^14. Even if C 2^20 pops up a magical result I don't know if I can explore it.
#
# It is still running... nm 20 isn't inclusive.
#
# I am goingto try both datasets with it as well.
#
# I got this result before I ran standard scalar on the data.
# +
rbf14_1 = svm.SVC(kernel='rbf', C=2**14).fit(x_train, y_train)
rbf14_y1_pred = rbf14_1.predict(x_test)
res1_rbf14 = Result(rbf14_y1_pred, y_test.values, 1)
res1_rbf14.display_totals()
# +
rbf14_2 = svm.SVC(kernel='rbf', C=2**14).fit(x2_train, y2_train)
rbf14_y2_pred = rbf14_2.predict(x2_test)
res2_rbf14 = Result(rbf14_y2_pred, y2_test.values, 1)
res2_rbf14.display_totals()
# -
plt.plot(y_test.values, res1_rbf14.list_variances, 'o')
plt.plot(y_test.values, res2_rbf14.list_variances, 'o')
res1_rbf14.get_confidence_interval(84)
# +
from sklearn.model_selection import cross_val_predict
cross_pred = cross_val_predict(rbf14_1, data1, data_y, cv=5)
# -
cross_pred
# cross_val_predict is not doing what it says it does in the docs.
#
# Dataset 2 with the cross correlations pulled out seems to not be as effective.
#
# Dataset 1 using the rbf kernel just comes closer. It seems to have pulled the predictions on the younger abalone closer to truth but still fails to identify specimens that live past 12 years old. If you give me a variance of 2 this model gives 84%.
#
# rbf14_1 = svm.SVC(kernel='rbf', C=2^14).fit(x_train, y_train)
#
# 2^14 is 16384 so the range of possible C's that may have a better variance is ~8150 to ~32000. Meh.
#
# Now to try poly. The first bunch of stuff here is just trying out all of the options.
poly_1 = svm.SVC(kernel='poly', C=1).fit(x_train, y_train)
poly_1_pred = poly_1.predict(x_test)
res_poly1 = Result(poly_1_pred, y_test.values, 1)
res_poly1.display_totals()
poly_1
poly_test = svm.SVC(kernel='poly', C=1, degree=1, probability=True, class_weight=None).fit(x_train, y_train)
poly_test_pred = poly_test.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
res_poly_test.display_totals()
plt.plot(y_test.values, res_poly_test.list_variances, 'o')
# So many options!
#
# The next cell might bork my computer. Time to cook something.
#
# As I watch results go by a couple observations. 1 degree polynomials are the best so far. Whatever probability does things are off the rails when it is true. Class Weight doesn't seem to change a lot.
#
# Variances are still descending at C 2^9.
#
# 1 degree and 2 degree are in possibly usful ranges. My guess is 1 degree is the best fit because we are really just measuring size and guessing age.
#
# The cell is commented just in case I hit run all cells again.
for power in range (-5, 10):
for deg in range(1, 10):
poly_test1 = svm.SVC(kernel='poly', C=2**power, degree=deg, probability=False, class_weight=None).fit(x_train, y_train)
poly_test_pred = poly_test1.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
print('Prob: False ClassW: None Power: 2**' + str(power) + ' Degree: ' + str(deg))
res_poly_test.display_totals()
poly_test2 = svm.SVC(kernel='poly', C=2**power, degree=deg, probability=False, class_weight='balanced').fit(x_train, y_train)
poly_test_pred = poly_test2.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
print('Prob: False ClassW: balanced Power: 2**' + str(power) + ' Degree: ' + str(deg))
res_poly_test.display_totals()
poly_test3 = svm.SVC(kernel='poly', C=2**power, degree=deg, probability=True, class_weight=None).fit(x_train, y_train)
poly_test_pred = poly_test3.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
print('Prob: True ClassW: None Power: 2**' + str(power) + ' Degree: ' + str(deg))
res_poly_test.display_totals()
poly_test4 = svm.SVC(kernel='poly', C=2**power, degree=deg, probability=True, class_weight='balanced').fit(x_train, y_train)
poly_test_pred = poly_test4.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
print('Prob: True ClassW: balanced Power: 2**' + str(power) + ' Degree: ' + str(deg))
res_poly_test.display_totals()
# I am hoping a 1 degree polynomial line can beat a linear regression which is the best result so far. I am going to run 1 and 2 degree just to make sure and use both balanced and None for class weight. I will run C's through 19 again.
#
#
for power in range (5, 20):
for deg in range(1, 3):
poly_test1 = svm.SVC(kernel='poly', C=2**power, degree=deg, class_weight=None).fit(x_train, y_train)
poly_test_pred = poly_test1.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
print('ClassW: None Power: 2**' + str(power) + ' Degree: ' + str(deg))
res_poly_test.display_totals()
poly_test2 = svm.SVC(kernel='poly', C=2**power, degree=deg, class_weight='balanced').fit(x_train, y_train)
poly_test_pred = poly_test2.predict(x_test)
res_poly_test = Result(poly_test_pred, y_test.values, 1)
print('ClassW: balanced Power: 2**' + str(power) + ' Degree: ' + str(deg))
res_poly_test.display_totals()
# The lowest total is around C 2^15 degree 2. For fun I am going to add 100 and subtract 100 from 2^15.
#
# +100 1.513
# -100 1.514
#
# Seems lucky that in a range between 16000 and 64000 2^15 was really close to the actual minimum.
poly1_p15d2 = svm.SVC(kernel='poly', C=2**15, degree=2, class_weight=None).fit(x_train, y_train)
poly1_p15d2_pred=poly1_p15d2.predict(x_test)
res1_polyp15d2 = Result(poly1_p15d2_pred, y_test.values, 1)
res1_polyp15d2.display_totals()
plt.plot(y_test.values, res1_polyp15d2.list_variances, 'o')
res1_polyp15d2.get_confidence_interval(83)
poly1_p15d2 = svm.SVC(kernel='poly', C=2**15-100, degree=2, class_weight=None).fit(x_train, y_train)
poly1_p15d2_pred=poly1_p15d2.predict(x_test)
res1_polyp15d2 = Result(poly1_p15d2_pred, y_test.values, 1)
res1_polyp15d2.display_totals
plt.plot(y_test.values, res1_polyp15d2.list_variances, 'o')
#
# #### I accidentally printed out the class weight 'balanced' answer and though it's variance is higher there is something to not about the variances. They don't slope down to the right indicating everything is being pulled to the center, particularly at the older ages.
#
# #### The problem it seems to have is that it has a tendancy to predict things in the middle ages too high. It is also randomly or probably not randomly the best performing balanced result.
#
# #### This should be exploitable. I don't know how. But he bottom kernel found a way to predict the ages of the oldest abalone fairly effectively.
#
# #### This is important because if there was money on the line and you had a week to dig into this you would look at this kernel. It did have a tendancy to predict high values on the middle ages and there are a few squirly's in the graph.
poly2_p15d2 = svm.SVC(kernel='poly', C=2**15, degree=2, class_weight='balanced').fit(x_train, y_train)
poly2_p15d2_pred=poly2_p15d2.predict(x_test)
res2_polyp15d2 = Result(poly2_p15d2_pred, y_test.values, 1)
res2_polyp15d2.display_totals()
plt.plot(y_test.values, res2_polyp15d2.list_variances, 'o')
# ## Gradient Boosting Regression
# +
from funkys_funcs import append_tts
t_sets = []
TEST_DESC = 0
X_TRAIN = 1
Y_TRAIN = 2
X_TEST = 3
Y_TEST = 4
a,b,c,d = train_test_split(data1, data_y, test_size=.2, random_state=17)
D1 = append_tts(a,b,c,d, 'L2 Norm Clip: ', t_sets)
a,b,c,d = train_test_split(data2, data_y, test_size=.2, random_state=19)
D2 = append_tts(a,b,c,d, 'L2 Norm Clip: ', t_sets)
a,b,c,d = train_test_split(datax, data_y, test_size=.2, random_state=23)
DX = append_tts(a,b,c,d, 'L2 Norm Clip: ', t_sets)
# +
grb = GradientBoostingRegressor(n_estimators=100, learning_rate=0.1, max_depth=1, random_state=0, loss='ls')
y_pred = grb.fit(t_sets[D1][X_TRAIN], t_sets[D1][Y_TRAIN]).predict(t_sets[D1][X_TEST])
res = Result(y_pred, t_sets[D1][Y_TEST].values, .1)
res.display_totals()
# -
plt.plot(t_sets[D1][Y_TEST], res.list_variances, 'o')
# #### The gradient boosting was really bad at guessing the ages of older abalone which is the source of most of the error. On the other hand the first try came out at MAE 1.75 which is much better than almost everything tried so far. Now that I know how to apply these in a few hours I could run a few hundred variations of this regressor and probably get something as good or better than SVM.
| 23,547 |
/Day3/.ipynb_checkpoints/House Rate Prediction-checkpoint.ipynb | b9f1741b04b88cf1f2cd89476469b2efe561ec92 | [] | no_license | rahdirs11/IEEE_ML | https://github.com/rahdirs11/IEEE_ML | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 215,228 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.datasets import load_boston
boston = load_boston()
type(boston)
boston
dir(boston)
boston.filename
boston.target
boston['target']
import pandas as pd
df = pd.DataFrame(boston.data)
# use DataFrame whenever you want to work with tables in python
df.head()
df.shape
df.tail()
boston.feature_names
df.columns = boston.feature_names
df.head()
dir(df)
df['PRICE'] = boston['target']
df.head()
df[3: 14]['AGE']
df.shape
type(df)
columnHeadings = boston['feature_names']
df.describe()
corr = df.corr()
corr.shape
# +
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(30, 30))
sns.heatmap(corr, annot=True)
# +
from sklearn.model_selection import train_test_split
X = df.drop(['PRICE'],axis=1)
y = df['PRICE']
xTrain, xTest, yTrain, yTest = train_test_split(X, y, test_size=0.2)
from sklearn.linear_model import LinearRegression
linearModel = LinearRegression()
linearModel.fit(xTrain, yTrain)
yPred = linearModel.predict(xTest)
# yPred
# print(xTest.describe())
linearModel.score(xTest, yTest)
# tests the accuracy of the model
# -
| 1,368 |
/1 Математика и Python для анализа данных/1.3 Центральная предельная теорема своими руками/task_1_3.ipynb | 4d17ffed94958ab0b18efcf681a25d5011f90fa4 | [] | no_license | romannovikov/yandex_mipt_DS_specialization | https://github.com/romannovikov/yandex_mipt_DS_specialization | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 84,245 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as sts
# Выберите ваше любимое непрерывное распределение (чем меньше оно будет похоже на нормальное, тем интереснее; попробуйте выбрать какое-нибудь распределение из тех, что мы не обсуждали в курсе):
#
# Для данной задачи было выбрано распределение Максвелла (http://mathworld.wolfram.com/MaxwellDistribution.html) с параметром a = 0.67.
summa = 0
for i in range(5, 12):
summa += 3**i * np.math.exp(-3) / np.math.factorial(i)
print(summa)
a = 0.67
maxwell_rv = sts.maxwell(scale=a)
# Сгенерируйте из него выборку объёма 1000, постройте гистограмму выборки и нарисуйте поверх неё теоретическую плотность распределения вашей случайной величины.
# код генерации выборки и отрисовки её гистограммы
sample = maxwell_rv.rvs(1000);
plt.hist(sample, normed=True, label='sample histogram');
# задаем масштаб оси х и получаем массив значений теоретической плотности распределения во всем масштабе значений
x = np.linspace(-1, 5, 100)
pdf = maxwell_rv.pdf(x)
# отрисовываем тпо на гистограмме, полученной выше, подписываем графики
plt.plot(x, pdf, label='theoretical pdf')
plt.legend()
plt.ylabel('$f(x)$')
plt.xlabel('$x$')
plt.show()
# Для нескольких значений n (например, 5, 10, 50) сгенерируйте 1000 выборок объёма n и постройте гистограммы распределений их выборочных средних. Используя информацию о среднем и дисперсии исходного распределения (её можно без труда найти в википедии), посчитайте значения параметров нормальных распределений, которыми, согласно центральной предельной теореме, приближается распределение выборочных средних. Обратите внимание: для подсчёта значений этих параметров нужно использовать именно теоретические среднее и дисперсию вашей случайной величины, а не их выборочные оценки. Поверх каждой гистограммы нарисуйте плотность соответствующего нормального распределения.
for n in [5, 25, 50]:
# код генерации выборки и отрисовки её гистограммы
sample_means = np.array([maxwell_rv.rvs(n).mean() for _ in range(1000)])
plt.hist(sample_means, 20, normed=True, range=[0.5, 2], label='sample means histogram')
# вычисляем pdf нормального распределения с параметрами, вычисленными из параметров распределения Максвелла
maxwell_mean = 2 * a * np.sqrt(2/np.pi)
maxwell_std = a**2 * (3*np.pi - 8) / np.pi
norm_rv = sts.norm(maxwell_mean, np.sqrt(maxwell_std / n))
x = np.linspace(0.5, 2, 100)
pdf = norm_rv.pdf(x)
# отрисовываем на графике
plt.plot(x, pdf, label='norm theoretical pdf')
# добавляем легенду на график
plt.legend()
plt.title("Sample size %d" % n)
plt.ylabel('fraction of samples $f(x)$')
plt.xlabel('sample mean $x$')
plt.show()
# Опишите разницу между полученными распределениями при различных значениях n. Как меняется точность аппроксимации распределения выборочных средних нормальным с ростом n?
# С увеличением объема выборки n все выборочные средние в полученных распределениях начинают концентрироваться вокруг среднего генеральной совокупности, при этом СКО уменьшается, что говорит о том, что точность аппроксимации с ростом n улучшается.
#
# Из также графиков видно, что полученные гистограммы сходятся с графиками теоретической плотности нормального распредения, построенными по параметрам среднего и СКО исходного распределения.
| 3,643 |
/mass_shootings.ipynb | 1139e60f7960a44659e875ddce4ed79b3371cbe6 | [] | no_license | subodhchhabra/mass-shootings | https://github.com/subodhchhabra/mass-shootings | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 261,407 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# -*- coding: utf-8 -*-
# %matplotlib inline
import pandas as pd
import numpy as np
import math
import matplotlib as mplstyle
import matplotlib.pyplot as plt
import locale
from locale import atof
from dateutil import parser
from datetime import datetime
# + deletable=true editable=true
# Import data
df = pd.read_csv('imported_data/mass_shootings.csv')
df_reworked = pd.read_csv('imported_data/mass_shootings_reworked.csv')
df_shooterData = pd.read_csv('imported_data/mass_shooters.csv')
df_ETupdated = pd.read_csv('updated_data/mass_shootings_updated.csv')
df_added = pd.read_csv('updated_data/mass_shootins_added.csv')
# + deletable=true editable=true
# Count number of occurances
df.State.count()
df_updated.State.count()
# + deletable=true editable=true
# Sort occurances by state
df_stateOccurances = pd.value_counts(df['State'].values, sort=True)
df_stateOccurances = df_stateOccurances.reset_index()
df_stateOccurances = df_stateOccurances.rename(columns = {'index':'state', 0:'count'})
# + deletable=true editable=true
# Graph by state
df_stateOccurances.plot(kind='bar',x='state',y='count')
# + deletable=true editable=true
# Count number of shooter suicides
# df_stateOccurances = pd.value_counts(df['State'].values, sort=True)
df_suicideCount = pd.value_counts(df['Shooter Suicide'].values)
# Graph
df_suicideCount.plot(kind='bar')
df_suicideCount
# + deletable=true editable=true
# Count number of guns used
# Make a new dataframe
df_weaponsUsed = pd.DataFrame()
# Add summed amounts of each gun type to dataframe
df_weaponsUsed['handgun'] = [df['Handgun'].sum()]
df_weaponsUsed['assult weapon'] = [df['Assault Weapon'].sum()]
df_weaponsUsed['other'] = [df['Other'].sum()]
df_weaponsUsed['unknown'] = [df['Unknown'].sum()]
df_weaponsUsed
# + deletable=true editable=true
# Count number of incidents by amount of victims
df_totalShot = pd.value_counts(df['TOTAL SHOT (Not Including Shooter)'].values, sort=True)
df_totalShot = df_totalShot.reset_index()
df_totalShot = df_totalShot.rename(columns = {'index':'num shot', 0:'count'})
df_totalShot.to_csv('exported_data/df_totalShot.csv')
# Graph
df_totalShot.plot(kind='bar',x='num shot',y='count')
# + deletable=true editable=true
# Average number of vitims per incident
df_totalShot['count'].mean()
# + deletable=true editable=true
# Function to remove unknowns and NaNs from cells
def removeUnknown(item):
if item == 'Unknown':
return 'NA'
elif item == 'nan':
return 'NA'
else:
return item
# Duplicate shooter data
df_shooterAgeData = df_shooterData
# Apply removeUnknown function to age column
df_shooterAgeData['Age'] = df_shooterData['Age'].apply(removeUnknown)
# df_shooterAgeData
# + deletable=true editable=true
# Filter out ages that are not numbers
df_shooterFilteredAge = df_shooterData[df_shooterData['Age'] != 'NA']
df_shooterFilteredAge.to_csv('exported_data/df_shooterFilteredAge.csv')
# + deletable=true editable=true
df
# + deletable=true editable=true
df['Number Killed [CALCULATED]'].sum() # 848
df['Children Killed (17 and Under)'].sum() # 211
# + deletable=true editable=true
# Breakdown gender data
df_shooterGenderData = pd.value_counts(df_shooterData['Gender'].values, sort=True)
df_shooterGenderData = df_shooterGenderData.reset_index()
df_shooterGenderData = df_shooterGenderData.rename(columns = {'index':'gender', 0:'count'})
# Graph
df_shooterGenderData.plot(kind='bar',x='gender',y='count')
df_shooterGenderData
# + deletable=true editable=true
# df[df['Killed Intimate Partner or Family [CALCULATED] '] == 'Yes'].count() #85
# df[df['Killed Intimate Partner or Family [CALCULATED] '] == 'No'].count() #71
df_dv = df
df_dv['idx'] = 1
df_dv.groupby('TOTAL SHOT (Not Including Shooter)').sum().reset_index()
# -
# # UPDATED DATA
# +
df_updated = df_ETupdated.merge(df_added,how='outer')
# UPDATE WHEN ADD NEW DATA
df_updated = df_updated[:168] # add one for each row
df_updated
# + deletable=true editable=true
# Count number of guns used
# Make a new dataframe
df_weaponsUsedUpd = pd.DataFrame()
# Add summed amounts of each gun type to dataframe
df_weaponsUsedUpd['handgun'] = [df_updated['Handgun'].sum()]
df_weaponsUsedUpd['assult weapon'] = [df_updated['Assault Weapon'].sum()]
df_weaponsUsedUpd['other'] = [df_updated['Other'].sum()]
df_weaponsUsedUpd['unknown'] = [df_updated['Unknown'].sum()]
df_weaponsUsedUpd
# + deletable=true editable=true
def getYear(date):
return date[-4:]
df_updated['Year'] = df_updated['Date'].apply(getYear)
# + deletable=true editable=true
df_updated.groupby('Year').sum().reset_index().drop(['Unknown','Unnamed: 47','Took place exclusively in private residence(s)'], axis=1)
# + deletable=true editable=true
# df_updated[df_updated['Killed Intimate Partner or Family [CALCULATED] '] == 'Yes'].count() #90
# df_updated[df_updated['Killed Intimate Partner or Family [CALCULATED] '] == 'No'].count() #75
df_dvUpd = df_updated
df_dvUpd['idx'] = 1
df_dvUpd.groupby('TOTAL SHOT (Not Including Shooter)').sum().reset_index().drop(['Other','Took place exclusively in private residence(s)','Unknown','Unnamed: 47','Handgun','Assault Weapon'],axis=1).to_csv('updated_data/dv.csv')
df_dvUpd.groupby('TOTAL SHOT (Not Including Shooter)').sum().reset_index().drop(['Other','Took place exclusively in private residence(s)','Unknown','Unnamed: 47','Handgun','Assault Weapon'],axis=1)
# + deletable=true editable=true
df_dvUpd_true = df_dvUpd[df_dvUpd['Killed Intimate Partner or Family [CALCULATED] '] == 'Yes']
df_dvUpd_false = df_dvUpd[df_dvUpd['Killed Intimate Partner or Family [CALCULATED] '] == 'No']
df_dvUpd_true.groupby('TOTAL SHOT (Not Including Shooter)').sum().reset_index().drop(['Other','Took place exclusively in private residence(s)','Unknown','Unnamed: 47','Handgun','Assault Weapon'],axis=1).to_csv('updated_data/dv_true.csv')
df_dvUpd_true.groupby('TOTAL SHOT (Not Including Shooter)').sum().reset_index().drop(['Other','Took place exclusively in private residence(s)','Unknown','Unnamed: 47','Handgun','Assault Weapon'],axis=1)
# + deletable=true editable=true
df_dvUpd_false.groupby('TOTAL SHOT (Not Including Shooter)').sum().reset_index().drop(['Other','Took place exclusively in private residence(s)','Unknown','Unnamed: 47','Handgun','Assault Weapon'],axis=1).to_csv('updated_data/dv_false.csv')
# + deletable=true editable=true
andn(2, 3, 3, 3)
b = np.random.randn(2,)
dout = np.random.randn(4, 2, 5, 5)
conv_param = {'stride': 1, 'pad': 1}
dx_num = eval_numerical_gradient_array(lambda x: conv_forward_naive(x, w, b, conv_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_forward_naive(x, w, b, conv_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_forward_naive(x, w, b, conv_param)[0], b, dout)
out, cache = conv_forward_naive(x, w, b, conv_param)
dx, dw, db = conv_backward_naive(dout, cache)
# Your errors should be around 1e-8'
print('Testing conv_backward_naive function')
print('dx error: ', rel_error(dx, dx_num))
print('dw error: ', rel_error(dw, dw_num))
print('db error: ', rel_error(db, db_num))
# + [markdown] deletable=true editable=true
# # Max pooling: Naive forward
# Implement the forward pass for the max-pooling operation in the function `max_pool_forward_naive` in the file `cs231n/layers.py`. Again, don't worry too much about computational efficiency.
#
# Check your implementation by running the following:
# + deletable=true editable=true
x_shape = (2, 3, 4, 4)
x = np.linspace(-0.3, 0.4, num=np.prod(x_shape)).reshape(x_shape)
pool_param = {'pool_width': 2, 'pool_height': 2, 'stride': 2}
out, _ = max_pool_forward_naive(x, pool_param)
correct_out = np.array([[[[-0.26315789, -0.24842105],
[-0.20421053, -0.18947368]],
[[-0.14526316, -0.13052632],
[-0.08631579, -0.07157895]],
[[-0.02736842, -0.01263158],
[ 0.03157895, 0.04631579]]],
[[[ 0.09052632, 0.10526316],
[ 0.14947368, 0.16421053]],
[[ 0.20842105, 0.22315789],
[ 0.26736842, 0.28210526]],
[[ 0.32631579, 0.34105263],
[ 0.38526316, 0.4 ]]]])
# Compare your output with ours. Difference should be around 1e-8.
print('Testing max_pool_forward_naive function:')
print('difference: ', rel_error(out, correct_out))
# + [markdown] deletable=true editable=true
# # Max pooling: Naive backward
# Implement the backward pass for the max-pooling operation in the function `max_pool_backward_naive` in the file `cs231n/layers.py`. You don't need to worry about computational efficiency.
#
# Check your implementation with numeric gradient checking by running the following:
# + deletable=true editable=true
np.random.seed(231)
x = np.random.randn(3, 2, 8, 8)
dout = np.random.randn(3, 2, 4, 4)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
dx_num = eval_numerical_gradient_array(lambda x: max_pool_forward_naive(x, pool_param)[0], x, dout)
out, cache = max_pool_forward_naive(x, pool_param)
dx = max_pool_backward_naive(dout, cache)
# Your error should be around 1e-12
print('Testing max_pool_backward_naive function:')
print('dx error: ', rel_error(dx, dx_num))
# + [markdown] deletable=true editable=true
# # Fast layers
# Making convolution and pooling layers fast can be challenging. To spare you the pain, we've provided fast implementations of the forward and backward passes for convolution and pooling layers in the file `cs231n/fast_layers.py`.
#
# The fast convolution implementation depends on a Cython extension; to compile it you need to run the following from the `cs231n` directory:
#
# ```bash
# python setup.py build_ext --inplace
# ```
#
# The API for the fast versions of the convolution and pooling layers is exactly the same as the naive versions that you implemented above: the forward pass receives data, weights, and parameters and produces outputs and a cache object; the backward pass recieves upstream derivatives and the cache object and produces gradients with respect to the data and weights.
#
# **NOTE:** The fast implementation for pooling will only perform optimally if the pooling regions are non-overlapping and tile the input. If these conditions are not met then the fast pooling implementation will not be much faster than the naive implementation.
#
# You can compare the performance of the naive and fast versions of these layers by running the following:
# + deletable=true editable=true
from cs231n.fast_layers import conv_forward_fast, conv_backward_fast
from time import time
np.random.seed(231)
x = np.random.randn(100, 3, 31, 31)
w = np.random.randn(25, 3, 3, 3)
b = np.random.randn(25,)
dout = np.random.randn(100, 25, 16, 16)
conv_param = {'stride': 2, 'pad': 1}
t0 = time()
out_naive, cache_naive = conv_forward_naive(x, w, b, conv_param)
t1 = time()
out_fast, cache_fast = conv_forward_fast(x, w, b, conv_param)
t2 = time()
print('Testing conv_forward_fast:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Difference: ', rel_error(out_naive, out_fast))
t0 = time()
dx_naive, dw_naive, db_naive = conv_backward_naive(dout, cache_naive)
t1 = time()
dx_fast, dw_fast, db_fast = conv_backward_fast(dout, cache_fast)
t2 = time()
print('\nTesting conv_backward_fast:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
print('dw difference: ', rel_error(dw_naive, dw_fast))
print('db difference: ', rel_error(db_naive, db_fast))
# + deletable=true editable=true
from cs231n.fast_layers import max_pool_forward_fast, max_pool_backward_fast
np.random.seed(231)
x = np.random.randn(100, 3, 32, 32)
dout = np.random.randn(100, 3, 16, 16)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
t0 = time()
out_naive, cache_naive = max_pool_forward_naive(x, pool_param)
t1 = time()
out_fast, cache_fast = max_pool_forward_fast(x, pool_param)
t2 = time()
print('Testing pool_forward_fast:')
print('Naive: %fs' % (t1 - t0))
print('fast: %fs' % (t2 - t1))
print('speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('difference: ', rel_error(out_naive, out_fast))
t0 = time()
dx_naive = max_pool_backward_naive(dout, cache_naive)
t1 = time()
dx_fast = max_pool_backward_fast(dout, cache_fast)
t2 = time()
print('\nTesting pool_backward_fast:')
print('Naive: %fs' % (t1 - t0))
print('speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
# + [markdown] deletable=true editable=true
# # Convolutional "sandwich" layers
# Previously we introduced the concept of "sandwich" layers that combine multiple operations into commonly used patterns. In the file `cs231n/layer_utils.py` you will find sandwich layers that implement a few commonly used patterns for convolutional networks.
# + deletable=true editable=true
from cs231n.layer_utils import conv_relu_pool_forward, conv_relu_pool_backward
np.random.seed(231)
x = np.random.randn(2, 3, 16, 16)
w = np.random.randn(3, 3, 3, 3)
b = np.random.randn(3,)
dout = np.random.randn(2, 3, 8, 8)
conv_param = {'stride': 1, 'pad': 1}
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
out, cache = conv_relu_pool_forward(x, w, b, conv_param, pool_param)
dx, dw, db = conv_relu_pool_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], b, dout)
print('Testing conv_relu_pool')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# + deletable=true editable=true
from cs231n.layer_utils import conv_relu_forward, conv_relu_backward
np.random.seed(231)
x = np.random.randn(2, 3, 8, 8)
w = np.random.randn(3, 3, 3, 3)
b = np.random.randn(3,)
dout = np.random.randn(2, 3, 8, 8)
conv_param = {'stride': 1, 'pad': 1}
out, cache = conv_relu_forward(x, w, b, conv_param)
dx, dw, db = conv_relu_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: conv_relu_forward(x, w, b, conv_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_relu_forward(x, w, b, conv_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_relu_forward(x, w, b, conv_param)[0], b, dout)
print('Testing conv_relu:')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# + [markdown] deletable=true editable=true
# # Three-layer ConvNet
# Now that you have implemented all the necessary layers, we can put them together into a simple convolutional network.
#
# Open the file `cs231n/classifiers/cnn.py` and complete the implementation of the `ThreeLayerConvNet` class. Run the following cells to help you debug:
# + [markdown] deletable=true editable=true
# ## Sanity check loss
# After you build a new network, one of the first things you should do is sanity check the loss. When we use the softmax loss, we expect the loss for random weights (and no regularization) to be about `log(C)` for `C` classes. When we add regularization this should go up.
# + deletable=true editable=true
model = ThreeLayerConvNet()
N = 50
X = np.random.randn(N, 3, 32, 32)
y = np.random.randint(10, size=N)
loss, grads = model.loss(X, y)
print('Initial loss (no regularization): ', loss)
model.reg = 0.5
loss, grads = model.loss(X, y)
print('Initial loss (with regularization): ', loss)
# + [markdown] deletable=true editable=true
# ## Gradient check
# After the loss looks reasonable, use numeric gradient checking to make sure that your backward pass is correct. When you use numeric gradient checking you should use a small amount of artifical data and a small number of neurons at each layer. Note: correct implementations may still have relative errors up to 1e-2.
# + deletable=true editable=true
num_inputs = 2
input_dim = (3, 16, 16)
reg = 0.0
num_classes = 10
np.random.seed(231)
X = np.random.randn(num_inputs, *input_dim)
y = np.random.randint(num_classes, size=num_inputs)
model = ThreeLayerConvNet(num_filters=3, filter_size=3,
input_dim=input_dim, hidden_dim=7,
dtype=np.float64)
loss, grads = model.loss(X, y)
for param_name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
param_grad_num = eval_numerical_gradient(f, model.params[param_name], verbose=False, h=1e-6)
e = rel_error(param_grad_num, grads[param_name])
print('%s max relative error: %e' % (param_name, rel_error(param_grad_num, grads[param_name])))
# + [markdown] deletable=true editable=true
# ## Overfit small data
# A nice trick is to train your model with just a few training samples. You should be able to overfit small datasets, which will result in very high training accuracy and comparatively low validation accuracy.
# + deletable=true editable=true
np.random.seed(231)
num_train = 100
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
model = ThreeLayerConvNet(weight_scale=1e-2)
solver = Solver(model, small_data,
num_epochs=15, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=1)
solver.train()
# + [markdown] deletable=true editable=true
# Plotting the loss, training accuracy, and validation accuracy should show clear overfitting:
# + deletable=true editable=true
plt.subplot(2, 1, 1)
plt.plot(solver.loss_history, 'o')
plt.xlabel('iteration')
plt.ylabel('loss')
plt.subplot(2, 1, 2)
plt.plot(solver.train_acc_history, '-o')
plt.plot(solver.val_acc_history, '-o')
plt.legend(['train', 'val'], loc='upper left')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.show()
# + [markdown] deletable=true editable=true
# ## Train the net
# By training the three-layer convolutional network for one epoch, you should achieve greater than 40% accuracy on the training set:
# + deletable=true editable=true
model = ThreeLayerConvNet(weight_scale=0.001, hidden_dim=500, reg=0.001)
solver = Solver(model, data,
num_epochs=1, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=20)
solver.train()
# + [markdown] deletable=true editable=true
# ## Visualize Filters
# You can visualize the first-layer convolutional filters from the trained network by running the following:
# + deletable=true editable=true
from cs231n.vis_utils import visualize_grid
grid = visualize_grid(model.params['W1'].transpose(0, 2, 3, 1))
plt.imshow(grid.astype('uint8'))
plt.axis('off')
plt.gcf().set_size_inches(5, 5)
plt.show()
# + [markdown] deletable=true editable=true
# # Spatial Batch Normalization
# We already saw that batch normalization is a very useful technique for training deep fully-connected networks. Batch normalization can also be used for convolutional networks, but we need to tweak it a bit; the modification will be called "spatial batch normalization."
#
# Normally batch-normalization accepts inputs of shape `(N, D)` and produces outputs of shape `(N, D)`, where we normalize across the minibatch dimension `N`. For data coming from convolutional layers, batch normalization needs to accept inputs of shape `(N, C, H, W)` and produce outputs of shape `(N, C, H, W)` where the `N` dimension gives the minibatch size and the `(H, W)` dimensions give the spatial size of the feature map.
#
# If the feature map was produced using convolutions, then we expect the statistics of each feature channel to be relatively consistent both between different imagesand different locations within the same image. Therefore spatial batch normalization computes a mean and variance for each of the `C` feature channels by computing statistics over both the minibatch dimension `N` and the spatial dimensions `H` and `W`.
# + [markdown] deletable=true editable=true
# ## Spatial batch normalization: forward
#
# In the file `cs231n/layers.py`, implement the forward pass for spatial batch normalization in the function `spatial_batchnorm_forward`. Check your implementation by running the following:
# + deletable=true editable=true
np.random.seed(231)
# Check the training-time forward pass by checking means and variances
# of features both before and after spatial batch normalization
N, C, H, W = 2, 3, 4, 5
x = 4 * np.random.randn(N, C, H, W) + 10
print('Before spatial batch normalization:')
print(' Shape: ', x.shape)
print(' Means: ', x.mean(axis=(0, 2, 3)))
print(' Stds: ', x.std(axis=(0, 2, 3)))
# Means should be close to zero and stds close to one
gamma, beta = np.ones(C), np.zeros(C)
bn_param = {'mode': 'train'}
out, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
print('After spatial batch normalization:')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(axis=(0, 2, 3)))
print(' Stds: ', out.std(axis=(0, 2, 3)))
# Means should be close to beta and stds close to gamma
gamma, beta = np.asarray([3, 4, 5]), np.asarray([6, 7, 8])
out, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
print('After spatial batch normalization (nontrivial gamma, beta):')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(axis=(0, 2, 3)))
print(' Stds: ', out.std(axis=(0, 2, 3)))
# + deletable=true editable=true
np.random.seed(231)
# Check the test-time forward pass by running the training-time
# forward pass many times to warm up the running averages, and then
# checking the means and variances of activations after a test-time
# forward pass.
N, C, H, W = 10, 4, 11, 12
bn_param = {'mode': 'train'}
gamma = np.ones(C)
beta = np.zeros(C)
for t in range(50):
x = 2.3 * np.random.randn(N, C, H, W) + 13
spatial_batchnorm_forward(x, gamma, beta, bn_param)
bn_param['mode'] = 'test'
x = 2.3 * np.random.randn(N, C, H, W) + 13
a_norm, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After spatial batch normalization (test-time):')
print(' means: ', a_norm.mean(axis=(0, 2, 3)))
print(' stds: ', a_norm.std(axis=(0, 2, 3)))
# + [markdown] deletable=true editable=true
# ## Spatial batch normalization: backward
# In the file `cs231n/layers.py`, implement the backward pass for spatial batch normalization in the function `spatial_batchnorm_backward`. Run the following to check your implementation using a numeric gradient check:
# + deletable=true editable=true
np.random.seed(231)
N, C, H, W = 2, 3, 4, 5
x = 5 * np.random.randn(N, C, H, W) + 12
gamma = np.random.randn(C)
beta = np.random.randn(C)
dout = np.random.randn(N, C, H, W)
bn_param = {'mode': 'train'}
fx = lambda x: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
fg = lambda a: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
fb = lambda b: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma, dout)
db_num = eval_numerical_gradient_array(fb, beta, dout)
_, cache = spatial_batchnorm_forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = spatial_batchnorm_backward(dout, cache)
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
# -
# # Extra Credit Description
# If you implement any additional features for extra credit, clearly describe them here with pointers to any code in this or other files if applicable.
| 24,411 |
/Python/python-machine-learning-book/chap3/Chapter3 A tour of Machine learning Clasifiers Using scikit-learn.ipynb | 0c1f6e885bb41dc20fb49010e008e4c609b1a99e | [] | no_license | dragstar328/NoteBooks | https://github.com/dragstar328/NoteBooks | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 118,640 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# hierarchical Clustering
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
dataset = pd.read_csv('Mall_Customers.csv')
X = dataset.iloc[:, [3, 4]].values
# -
# <h5>Scipy is an open source py lib to do import tools to build dendrogram <br>linkage is algo for hierrarchical clustering<br> ward method tries to minimize the variance within each cluster</h5>
# +
#using dendrogram to find optimal number of cluster
import scipy.cluster.hierarchy as sch
dendrogram = sch.dendrogram(sch.linkage(X, method = 'ward'))
plt.title('Dendrogram')
plt.xlabel('customers')
plt.ylabel('Eucledian dis')
plt.show
# -
# <h5>1) AgglomerativeClustering is a type of algorithm used in hierarchical clustering<br>2) affinity is the type of distance</h5>
#fitting hierarchical clustering to the data
from sklearn.cluster import AgglomerativeClustering
hc = AgglomerativeClustering(n_clusters = 5, affinity = 'euclidean', linkage = 'ward')
y_hc = hc.fit_predict(X)
print(y_hc)
# <h4>above numbers are showing to which cluster it belongs</h5>
#visualizing the cluster
plt.scatter(X[y_hc== 0, 0], X[y_hc== 0, 1], s = 70, c = 'red', label = 'Cluster 1')
plt.scatter(X[y_hc== 1, 0], X[y_hc== 1, 1], s = 70, c = 'blue', label = 'Cluster 2')
plt.scatter(X[y_hc== 2, 0], X[y_hc== 2, 1], s = 70, c = 'green', label = 'Cluster 3')
plt.scatter(X[y_hc== 3, 0], X[y_hc== 3, 1], s = 70, c = 'cyan', label = 'Cluster 4')
plt.scatter(X[y_hc== 4, 0], X[y_hc== 4, 1], s = 70, c = 'magenta', label = 'Cluster 5')
# plt.scatter(sch_[:, 0], sch_[:, 1], s = 300, c = 'yellow', label = 'Centroids')
plt.title('Clusters of customers')
plt.xlabel('Annual Income (k$)')
plt.ylabel('Spending Score (1-100)')
plt.legend()
plt.show()
())
# +
from sklearn.metrics import accuracy_score
print("Accuracy: {}".format(accuracy_score(y_test, y_pred)))
# +
from matplotlib.colors import ListedColormap
import warnings
def versiontuple(v):
return tuple(map(int, (v.split("."))))
def plot_decision_regions(X, y, classifier, test_idx=None, resolution=0.02):
#setup marker generator and color map
markers = ("s", "x", "o", "^", "v")
colors = ("red", "blue", "lightgreen", "gray", "cyan")
cmap = ListedColormap(colors[:len(np.unique(y))])
#plot the decision surface
x1_min, x1_max = X[:, 0].min() -1, X[:, 0].max() + 1
print("x1_min :{0}\tx1_max :{1}".format(x1_min, x1_max))
x2_min, x2_max = X[:, 1].min() -1, X[:, 1].max() + 1
print("x2_min :{0}\tx2_max :{1}".format(x2_min, x2_max))
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, resolution),
np.arange(x2_min, x2_max, resolution))
#print("xx1 :{0}\txx2 :{1}".format(xx1, xx2))
Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.4, cmap=cmap)
plt.xlim(xx1.min(), xx1.max())
plt.ylim(xx2.min(), xx2.max())
for idx, cl in enumerate(np.unique(y)):
plt.scatter(x=X[y == cl, 0],
y=X[y == cl, 1],
alpha = 0.6,
c=cmap(idx),
edgecolor='black',
marker=markers[idx],
label=cl)
# highlight test samples
if test_idx:
# plot all samples
if not versiontuple(np.__version__) >= versiontuple("1.9.0"):
X_test, y_test = X[list(test_idx), :], y[list(test_idx)]
warnings.warn("Please update to Numpy 1.9.0 or newer")
else:
X_test, y_test = X[test_idx, :], y[test_idx]
plt.scatter(X_test[:, 0],
X_test[:, 1],
c='',
alpha=1.0,
edgecolor='black',
linewidths='1',
marker='o',
s=55,
label='test set')
# +
X_combined_std = np.vstack((X_train_std, X_test_std))
y_combined_std = np.hstack((y_train, y_test))
plot_decision_regions(X=X_combined_std, y=y_combined_std, classifier=ppn, test_idx=range(105, 150))
plt.xlabel('petal length [standardized]')
plt.ylabel('petal width [standardized]')
plt.legend(loc="upper left")
plt.tight_layout()
# -
# ## Modeling class probabilities via Logistic Regression
def sigmoid(z):
return 1.0 / (1.0 + np.exp(-z))
# +
z = np.arange(-7, 7, 0.1)
phi_z = sigmoid(z)
plt.plot(z, phi_z)
plt.axvline(0.0, color="k")
plt.ylim(-0.1, 1.1)
plt.xlabel("z")
plt.ylabel("$\phi (z)$")
plt.yticks([0.0, 0.5, 1.0])
ax = plt.gca()
ax.yaxis.grid(True)
# -
# ## Leaning the weights of he logistic cost function
# +
def cost_1(z):
return -np.log(sigmoid(z))
def cost_0(z):
return -np.log(1 - sigmoid(z))
z = np.arange(-10, 10, 0.1)
phi_z = sigmoid(z)
c1 = [cost_1(x) for x in z]
plt.plot(phi_z, c1, label="J(w) if y=1")
c0 = [cost_0(x) for x in z]
plt.plot(phi_z, c0, linestyle="--", label="J(w) if y=0")
plt.ylim(0.0, 5.1)
plt.xlim(0, 1)
plt.xlabel("$\phi$(z)")
plt.ylabel("J(w)")
plt.legend(loc="best", frameon=True, edgecolor="blue")
# -
# ## Training a logistic regressin model with scikit-learn
from sklearn.linear_model import LogisticRegression
# +
lr = LogisticRegression(C=1000.0, random_state=0)
lr.fit(X_train_std, y_train)
plot_decision_regions(X_combined_std, y_combined_std, classifier=lr, test_idx=range(105, 150))
plt.xlabel("petal length [standardized]")
plt.ylabel("petal width [standardized]")
plt.legend(loc="upper left", frameon=True)
# -
if Version(sklearn_version) < '0.17':
lr.predict_proba(X_test_std[0, :])
else:
lr.predict_proba(X_test_std[0, :].reshape(1, -1))
# ## Tacking overfitting via regularization
| 6,012 |
/code/1_snapshotting/Untitled.ipynb | 123bb8160fb9b40b8b4926682a66270a6e092aec | [] | no_license | KinaMarie/siliconvalleyanalysis | https://github.com/KinaMarie/siliconvalleyanalysis | 0 | 0 | null | 2020-05-11T18:52:12 | 2016-02-18T17:53:42 | Jupyter Notebook | Jupyter Notebook | false | false | .py | 151,295 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Web Scraping - Lab
#
# ## Introduction
#
# Now that you've seen a more extensive example of developing a web scraping script, it's time to further practice and formalize that knowledge by writing functions to parse specific pieces of information from the web page and then synthesizing these into a larger loop that will iterate over successive web pages in order to build a complete dataset.
#
# ## Objectives
#
# You will be able to:
#
# * Navigate HTML documents using Beautiful Soup's children and sibling relations
# * Select specific elements from HTML using Beautiful Soup
# * Use regular expressions to extract items with a certain pattern within Beautiful Soup
# * Determine the pagination scheme of a website and scrape multiple pages
#
# ## Lab Overview
#
# This lab will build upon the previous lesson. In the end, you'll look to write a script that will iterate over all of the pages for the demo site and extract the title, price, star rating and availability of each book listed. Building up to that, you'll formalize the concepts from the lesson by writing functions that will extract a list of each of these features for each web page. You'll then combine these functions into the full script which will look something like this:
#
# ```python
# df = pd.DataFrame()
# for i in range(2,51):
# url = "http://books.toscrape.com/catalogue/page-{}.html".format(i)
# soup = BeautifulSoup(html_page.content, 'html.parser')
# new_titles = retrieve_titles(soup)
# new_star_ratings = retrieve_ratings(soup)
# new_prices = retrieve_prices(soup)
# new_avails = retrieve_avails(soup)
# ...
# ```
# ## Retrieving Titles
#
# To start, write a function that extracts the titles of the books on a given page. The input for the function should be the `soup` for the HTML of the page.
def retrieve_titles(soup):
warning = soup.find('div', class_="alert alert-warning")
book_container=warning.nextSibling.nextSibling
titles=[h3.find('a').attrs['title']for h3 in book_container.findAll('h3')]
return titles
# ## Retrieve Ratings
#
# Next, write a similar function to retrieve the star ratings on a given page. Again, the function should take in the `soup` from the given HTML page and return a list of the star ratings for the books. These star ratings should be formatted as integers.
def retrieve_ratings(soup):
warning = soup.find('div', class_="alert alert-warning")
book_container=warning.nextSibling.nextSibling
star_dict = {'One': 1, 'Two': 2, 'Three':3, 'Four': 4, 'Five':5}
star_ratings =[]
regex = re.compile("star-rating(.*)")
for p in book_container.findAll('p',{"class": regex}):
star_ratings.append(p.attrs['class'][-1])
star_ratings = [star_dict[s] for s in star_ratings]
return star_ratings
# ## Retrieve Prices
#
# Now write a function to retrieve the prices on a given page. The function should take in the `soup` from the given page and return a list of prices formatted as floats.
def retrieve_prices(soup):
warning = soup.find('div', class_="alert alert-warning")
book_container=warning.nextSibling.nextSibling
prices=[p.text for p in book_container.findAll('p', class_="price_color")]
prices=[float(p[1:]) for p in prices]
return prices
# ## Retrieve Availability
#
# Write a function to retrieve whether each book is available or not. The function should take in the `soup` from a given html page and return a list of the availability for each book.
def retrieve_availabilities(soup):
warning = soup.find('div', class_="alert alert-warning")
book_container=warning.nextSibling.nextSibling
avails = [a.text.strip() for a in book_container.findAll('p', class_="instock availability")]
return avails
# ## Create a Script to Retrieve All the Books From All 50 Pages
#
# Finally, write a script to retrieve all of the information from all 50 pages of the books.toscrape.com website.
# +
from bs4 import BeautifulSoup
import requests
import re
import pandas as pd
for i in range(1,51):
if i == 1:
url = 'http://books.toscrape.com'
html_page = requests.get(url)
soup = BeautifulSoup(html_page.content,'html.parser')
new_titles = retrieve_titles(soup)
new_star_ratings = retrieve_ratings(soup)
new_prices = retrieve_prices(soup)
new_avails = retrieve_availabilities(soup)
df = pd.DataFrame([new_titles, new_star_ratings, new_prices, new_avails]).transpose()
df.columns=['Title', 'Star_Rating', 'Price', 'Availability']
else:
url = "http://books.toscrape.com/catalogue/page-{}.html".format(i)
html_page = requests.get(url)
soup = BeautifulSoup(html_page.content,'html.parser')
new_titles = retrieve_titles(soup)
new_star_ratings = retrieve_ratings(soup)
new_prices = retrieve_prices(soup)
new_avails = retrieve_availabilities(soup)
df.append([new_titles, new_star_ratings, new_prices, new_avails]).transpose()
df.head()
# -
# ## Level-Up: Write a new version of the script you just wrote.
#
# If you used URL hacking to generate each successive page URL, instead write a function that retrieves the link from the `"next"` button at the bottom of the page. Conversely, if you already used this approach above, use URL-hacking (arguably the easier of the two methods in this case).
# +
def get_next_page(soup):
next_button=soup.find('li', class_="next")
if next_button:
return next_button.find('a').attrs['href']
else:
return None
get_next_page(soup)
def parse_url(url, new_titles = [], new_star_ratings = [], new_prices = [], new_avails = []):
html_page = requests.get(url)
soup = BeautifulSoup(html_page.content,'html.parser')
new_titles += retrieve_titles(soup)
new_star_ratings += retrieve_ratings(soup)
new_prices += retrieve_prices(soup)
new_avails += retrieve_availabilities(soup)
next_url_ext=get_next_page(soup)
if next_url_ext:
url = "/".join(url.split("/")[:-1]) + "/" + next_url_ext
return parse_url(next_url, new_titles, new_star_ratings, new_prices, new_avails)
else:
return new_titles, new_star_ratings, new_prices, new_avails
url = 'http://books.toscrape.com'
titles, star_ratings, prices, avails = parse_url (url, new_titles, new_star_ratings, new_prices, new_avails)
df = pd.DataFrame([titles, star_ratings, prices, avails]).transpose()
df.columns=['Title', 'Star_Rating', 'Price', 'Availability']
df.head()
# -
# ## Summary
#
# Well done! You just completed your first full web scraping project! You're ready to start harnessing the power of the web!
| 6,940 |
/plots_python/python_notebooks_Vanessa/check_CABLE_ccam_190227.ipynb | 39517e094c22395d7a6646bbddc588bc9aad00fd | [] | no_license | juergenknauer/CABLE_files | https://github.com/juergenknauer/CABLE_files | 0 | 2 | null | null | null | null | Jupyter Notebook | false | false | .py | 564,819 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
import netCDF4 as nc
import numpy as np
import re
import datetime
from matplotlib.ticker import FixedLocator
from matplotlib.backends.backend_pdf import PdfPages
import mpld3
#mpld3.enable_notebook()
import site
import importlib
import os
import matplotlib.gridspec as gridspecb
from struct import *
# read in MODIS land cover
lc_key = ['sparsely-vegetated','semi-arid','tundra & arctic-shrubland','grassland & cropland', \
'tropical forest','extra-tropical forest']
lcfilename = '/OSM/CBR/OA_GLOBALCABLE/work/CABLE-CCAM/ccam_100km.199912.modis_biome.nc'
f = nc.Dataset(lcfilename)
lc_index = np.squeeze(f.variables['climate_biome'][:,:])
cplant2 = np.squeeze(f.variables['cplant2'][:,:])
lat = f.variables['lat'][:]
lon = f.variables['lon'][:]
var_list = f.variables
for var in var_list:
print ('Name:',var)
lats = np.fliplr(np.repeat(lat[:,np.newaxis],720,axis=1))
lons = np.repeat(lon[np.newaxis,:],360,axis=0)
from mpl_toolkits.axes_grid1 import make_axes_locatable
ax = plt.subplot(111)
im = plt.contourf(lons,lats,cplant2)
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im, cax=cax)
years = np.arange(1980, 2000)
months = np.arange(12) +1
# get time series of biomass ,aggregated by land-cover class
file_path = '/flush3/rya17a/casacn/raijin/avg/'
file_stem = 'ccam_100km.'
count = 0
#time_vec = np.zeros(len(years)*len(months))
time_vec = np.array([])
for i in years:
for j in months:
file_name = file_path + file_stem + str(i) + str(j).zfill(2) + '.nc'
#print(file_name)
f = nc.Dataset(file_name)
time_vec = np.append(time_vec, (f.variables['time'][:]))
count = count + 1
# read land-cover map (1deg resolution)
lcfilename1 = '/OSM/CBR/OA_GLOBALCABLE/work/CABLE-CCAM/biome_MODIS_LC_1deg.nc'
f1 = nc.Dataset(lcfilename1)
lc1_index = np.squeeze(f1.variables['climate_biome'][:,:].filled(fill_value = 0))
lat1 = f1.variables['latitude'][:]
lon1 = f1.variables['longitude'][:]
lats1 = np.fliplr(np.repeat(lat1[:,np.newaxis],360,axis=1))
lons1 = np.repeat(lon1[np.newaxis,:],150,axis=0)
lc1_vec = np.reshape(lc1_index,[150*360])
lats1_vec = np.reshape(lats1,[150*360])
lons1_vec = np.reshape(lons1,[150*360])
lc1_index.shape
lat1.shape
len(lc1_vec[lc1_vec>0])
print(lats1.shape)
lats1_vec[0]
# create vector of indices relating the position in the collapsed vector to the position in the full array of grid-cells
file_name = '/OSM/CBR/OA_GLOBALCABLE/work/CABLE-CCAM/plume_out_cable_zero.nc'
f1 = nc.Dataset(file_name)
local_lat = f1.variables['local_lat'][:]
local_lon = f1.variables['local_lon'][:]
I1=list()
for i in np.arange(len(local_lat)):
I1.append((np.where((lats1_vec==local_lat[i]) & (lons1_vec==local_lon[i] ))))
for index, x in enumerate(np.nditer(lc1_vec, op_flags = ['readwrite'])):
lc1_vec[index] = np.int(x)
lc1_vec_reduced=np.array([])
lc1_vec_reduced = np.squeeze(lc1_vec[I1])
np.unique(lc1_vec_reduced)
lc1_mat = np.array([])
lc1_mat = lc1_vec.reshape([150,360])
np.unique(lc1_mat)
from mpl_toolkits.axes_grid1 import make_axes_locatable
from matplotlib import colors
cmap = colors.ListedColormap(['b','y','g','r','c','m'])
ax = plt.subplot(111)
#im = plt.contourf(lons1,lats1,lc1_mat, 7, cmap=cmap)
im = plt.contourf(lons,lats,lc_index, 6, cmap=cmap)
#im = plt.contourf(lons1,lats1,lons1)
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im, cax=cax)
from mpl_toolkits.axes_grid1 import make_axes_locatable
from matplotlib import colors
cmap = colors.ListedColormap(['k','b','y','g','r','c','m'])
ax = plt.subplot(111)
im = plt.contourf(lons1,lats1,lc1_mat, 7, cmap=cmap)
#im = plt.contourf(lons,lats,lc_index, 7, cmap=cmap)
#im = plt.contourf(lons1,lats1,lons1)
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im, cax=cax)
# read patchfrac from netcdf file
file_name = '/OSM/CBR/OA_GLOBALCABLE/work/CABLE-CCAM/plume_out_cable_zero.nc'
f1 = nc.Dataset(file_name)
patchfrac = f1.variables['patchfrac'][:,:].filled(fill_value = 0)
patchfrac = np.repeat(patchfrac[np.newaxis,:,:],f1.variables['time'].size,axis=0)
patchfrac.shape
vars = ['GPP', 'Evap', 'LAI','TotLivBiomass','SWE','LeafResp','Tair']
for v in vars:
mystring = v + '=f1.variables["' + v + '"][:,:].filled(fill_value = 0)'
exec(mystring)
mystring = v + 'tot = np.sum(patchfrac*' + v + ',1)'
exec(mystring)
print(mystring)
mystring = v + '_vec1 = np.zeros([6,240])'
exec(mystring)
for l in (np.arange(6)+1):
mystring = v + '_vec1[l-1,:] = ' + v + 'tot[:,lc1_vec_reduced==l ].mean(axis=1)'
print(mystring)
exec(mystring)
plt.plot(Tair_vec1.transpose()-273.15)
biomass_vec1 = TotLivBiomass_vec1.transpose()*1000
lai_vec1 = LAI_vec1.transpose()
evap_vec1 = Evap_vec1.transpose()*24*3600
snw_vec1 = SWE_vec1.transpose()
fpn_vec1 = (GPP_vec1.transpose() - LeafResp_vec1.transpose())*12.0 * 1e-6 * 3600 * 24
tas_vec1 = Tair_vec1.transpose() - 273.15
site_lat = [-35.57, 61.85]
site_lon = [148.15, 24.29]
site_lat = [ 61.85]
site_lon = [24.29]
Ilat = np.argmin(abs(lat-site_lat), axis=0)
Ilon = np.argmin(abs(lon-site_lon), axis=0)
print(lats[Ilat,Ilon])
print(lons[Ilat,Ilon])
file_path = '/flush3/rya17a/casacn/raijin/avg/'
file_stem = 'ccam_100km.'
count = 0
#time_vec = np.zeros(len(years)*len(months))
biomass_vec = np.array([])
lai_vec = np.array([])
evap_vec = np.array([])
snd_vec = np.array([])
snw_vec = np.array([])
fpn_vec = np.array([])
for i in years:
for j in months:
file_name = file_path + file_stem + str(i) + str(j).zfill(2) + '.nc'
#print(file_name)
f = nc.Dataset(file_name)
biomass_vec = np.append(biomass_vec, (f.variables['cplant2'][0,Ilat,Ilon]))
lai_vec = np.append(lai_vec, (f.variables['lai'][0,Ilat,Ilon]))
evap_vec = np.append(evap_vec, (f.variables['evap'][0,Ilat,Ilon]))
snd_vec = np.append(snd_vec, (f.variables['snd'][0,Ilat,Ilon]))
snw_vec = np.append(snw_vec, (f.variables['snw'][0,Ilat,Ilon]))
fpn_vec = np.append(fpn_vec, (f.variables['fpn_ave'][0,Ilat,Ilon]))
count = count + 1
file_path = '/flush3/rya17a/casacn/raijin/avg/'
file_stem = 'ccam_100km.'
count = 0
biomass_vec = np.zeros((240,6))
lai_vec = np.zeros((240,6))
evap_vec = np.zeros((240,6))
snd_vec = np.zeros((240,6))
snw_vec = np.zeros((240,6))
fpn_vec = np.zeros((240,6))
tas_vec = np.zeros((240,6))
for i in years:
for j in months:
file_name = file_path + file_stem + str(i) + str(j).zfill(2) + '.nc'
f = nc.Dataset(file_name)
for l in (np.arange(6)+1):
biomass_vec[count,l-1] = (f.variables['cplant1'][0,:,:])[lc_index==l].mean() + \
(f.variables['cplant2'][0,:,:])[lc_index==l].mean() + \
(f.variables['cplant3'][0,:,:])[lc_index==l].mean()
lai_vec[count,l-1] = (f.variables['lai'][0,:,:])[lc_index==l].mean()
evap_vec[count,l-1] = (f.variables['evap'][0,:,:])[lc_index==l].mean()*8.0 # mm/d
snd_vec[count,l-1] = (f.variables['snd'][0,:,:])[lc_index==l].mean()
snw_vec[count,l-1] = (f.variables['snw'][0,:,:])[lc_index==l].mean()
fpn_vec[count,l-1] = (f.variables['fpn_ave'][0,:,:])[lc_index==l].mean()*-1.0*24.*3600. # g C/m-2/d-1
tas_vec[count,l-1] = (f.variables['tas'][0,:,:])[lc_index==l].mean() - 273.15
count = count + 1
from textwrap import wrap
vars=['fpn_vec','evap_vec','lai_vec','biomass_vec','snw_vec', 'tas_vec']
varlabels = ['net photosyn', 'evap', 'lai', 'biomass','SWE','Tair']
fig=plt.figure(figsize=(15, 7),constrained_layout=True)
ncols = 6
nrows = 6
sindex=[]
x = np.arange(7)
gs = fig.add_gridspec(ncols=ncols, nrows=nrows)
[sindex.append((j,i)) for i in range(ncols) for j in range(nrows)]
index = 0
for index2, l in enumerate(lc_key):
for index1, v in enumerate(vars):
ax = fig.add_subplot(gs[sindex[index]])
mystring = 'y =' + v + '[:,index2]'
exec(mystring)
plt.plot(y, label='CCAM')
mystring = 'y =' + v + '1[:,index2]'
exec(mystring)
plt.plot(y, label='offline')
if (index1==0):
title = ax.set_title("\n".join(wrap(l, 15)))
#fig.tight_layout()
title.set_y(1.05)
#fig.subplots_adjust(top=0.8)
#plt.title(l)
if (index2==0):
ax.set_ylabel(varlabels[index1])
index = index + 1
if ((index2==0) and (index1==0)):
plt.legend(fontsize='small',frameon=True,loc=2)
plot_dir = '/OSM/CBR/OA_GLOBALCABLE/work/CABLE-CCAM/'
plot_fname = 'CCAM-CABLE-checks.pdf'
fig.savefig(os.path.join(plot_dir, plot_fname), bbox_inches='tight',
pad_inches=0.1)
bafile = '/OSM/CBR/OA_GLOBALCABLE/work/BIOS3_forcing/simfire_monthly_ba.nc'
f = nc.Dataset(bafile)
lat = f.variables['latitude'][:]
lon = f.variables['longitude'][:]
ba_monthly = f.variables['monthly_ba'][:]
lats = np.fliplr(np.repeat(lat[:,np.newaxis],720,axis=1))
lons = np.repeat(lon[np.newaxis,:],360,axis=0)
ba_monthly.shape
lat.shape
from mpl_toolkits.axes_grid1 import make_axes_locatable
ax = plt.subplot(111)
im = plt.contourf(lons,lats,np.squeeze(ba_monthly[:,:,10]))
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im, cax=cax)
site_lat = [-12.5]
site_lon = [131.15]
Ilat = np.argmin(abs(lat-site_lat), axis=0)
Ilon = np.argmin(abs(lon-site_lon), axis=0)
print(lats[Ilat,Ilon])
print(lons[Ilat,Ilon])
ba_monthly[Ilat, Ilon, :]
plt.plot(ba_monthly[Ilat, Ilon, :])
| 10,029 |
/DataScienceSeries_Lesson_1_EDA_Using_Python.ipynb | 297e7727aea7d8d87fea8b8fc5aa959704e6da46 | [] | no_license | nitarane/Python_Tutorial | https://github.com/nitarane/Python_Tutorial | 0 | 1 | null | null | null | null | Jupyter Notebook | false | false | .py | 221,292 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# This notebook, is to help the novice python learners to pick up python faster by showing them how to explore a dataset which aims to find patterns and relationships in data by using Python & its libraries. As this is the first notebook, we will cover the basic commands of Exploratory Data Analysis (EDA) which includes cleaning, munging, combining, reshaping, slicing and dicing, and transforming data for analysis purpose.
#
#
# In order to improve any model's accuracy, we need to spend significant time on exploring and analyzing the Data. EDA is a critical & first step in analyzing the data and we do this for below reasons :
#
# • Finding patterns in Data
# • Determining relationships in Data
# • Checking of assumptions
# • Preliminary selection of appropriate models
# • Detection of mistakes
#
#
# Data Exploration/Analysis is basically finding out more about the data we have.
# And for Data Analysis, Python has gathered a lot of interest recently as a choice of language. Python’s improved library support (primarily pandas) has made it a strong alternative for data manipulation tasks. Combined with Python’s strength in general purpose programming, it is an excellent choice as a single language for building data-centric applications.
#
# We will be using following set of libraries :
#
# Pandas: Extensively used for Structured data operations & manipulations and also for data munging & preparation
#
# Numpy : Numpy stands for Numerical Python. This library contains basic linear algebra functions, Fourier Transforms, Advanced Random Number Capabilities
#
# Matplotlib : Python based plotting library offers matplotlib with complete 2D & limited 3D graphic support
#
# SciPy : Collection of packages addressing a number of different standard problem domains in scientific computing. (Not used in this notebook but we will learn this later)
# +
# Importing Standard Library functions & Displaying Pandas Version
import pandas as pd
import numpy as np
import sys
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
#Here ast_note_interactivity kernel option make us see the value of multiple statements at once
print('Python version ' + sys.version)
print('Pandas version ' + pd.__version__)
print('NumPy version ' + np.__version__)
# -
# Just to begin with, we will analyse Chipotle Dataset for getting and knowing the Data which can help us drive valuable decisions for their business. You can download this Dataset from
# https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv
#
#
# For importing data we use pd.read_csv(filename) function.
# Here pd refers to pandas as imported above`;`
# df refers to pandas dataframe object
# +
# Read the chipotle dataset into pandas dataframe and see few sample records. This is in TSV(Tab Separated File) Form.
#location = r'C:\Users\SONY\Desktop\PANDAS\Python Pandas NumPy Jan 31\chipotle.tsv'
location = r'/Users/aurobindosarkar/Downloads/Nita/chipotle.tsv'
df = pd.read_csv(location, sep='\t')
df.head(5)
# -
# We can see few sample records & this helps to understand the following :
#
# • We are able to read the File
# • We have used correct function to read the file
# • Getting familiar with what values we might see. For e.g. item_price column is having `$` along with float values. So in case if we want to aggregate this column then we will have to get rid of the `$` sign.
#
# --------------------------------------------------------------------------------------------------------------------------------
#
#
# For inspecting Data we have below functions :
#
# df.head(n) -- First n rows of DataFrame
#
# df.tail(n) -- Last n rows of DataFrame
#
# df.shape -- Number of rows and columns
#
# df.info() -- Index, Datatype and Memory information
#
# df.dtypes -- Data Type of each Column
#
# df.describe() -- Summary statistics for numerical columns
#
# df.columns -- Prints the name of all columns
#
# df.ColumnName.value_counts(dropna=False) -- View unique values and counts
# dropna=False indicates Don’t include counts of NaN
#
# df.apply(pd.Series.value_counts) -- Unique values and counts for all columns
#
#
# Now Lets INSPECT Data by using above functions :
# +
#Its important to understand the volume of data i.e. the number of rows & columns present in the given dataset which is given by shape function
df.shape
print('Number of Rows: ' + str(df.shape[0]))
print('Number of Columns: ' + str(df.shape[1]))
# -
# As you can see, everything has been read in properly – we have 4622 rows and 5 columns. Also this attribute enables us to access the metadata. And help us to decide if we can work on entire dataset or we need to do sampling for EDA in case the dataset is really very huge.
#
# Sampling refers to the technique used to select, manipulate and analyze a representative subset of data in order to identify patterns and trends in the larger data set being examined. We will study this in detail in the upcoming notebook series.
# +
print('Index, Datatype and Memory information : \n')
df.info()
# Memory usage of each column can be found by calling the below method.
# It returns a Series with an index represented by column names and memory usage of each column shown in bytes
# And uses that as a launching point to improve the understanding of how each data types are stored in memory.
df.memory_usage()
# -
# This information is vital for our EDA as :
#
# Indexing enables us to store and manipulate data with an arbitrary number of dimensions in lower dimensional data structures like Series (1d) and DataFrame (2d). Indexing also helps in faster identification of rows.
#
#
# Memory Usage tells us about an approximate amount of RAM used to hold the DataFrame which can even help to write more efficient code.
# Also this helps us to avoid accidental copies of data as it is one of the major cause of slowness.
#
# When dealing with very large dataset /big data we need to keep a manageable in-memory data size and makes in-memory calculations fast & even multiple processing for cpu-intensive operations so understanding memory usage is critically important.
# And for this we have few Python libraries like H5py, PyTables & mmap for memory management. (Not a part of this notebook)
df.dtypes
# Knowing these data types is highly important as they help us in knowing the numeric & non numeric fields. In case if there are any objects present, we need to convert them into desired data types. This would help us to use the data for any aggregate/statistical function.
#
# For example, item_price, As we can see, data type for item_price is Object, we can convert it into Float by using lambda function as follows :
#
# Lambda Function :
#
# This is a way to create small anonymous functions, i.e. functions without a name. They are just needed where they have been created.
# Lets break it down :
#
# lambda : — this is a lambda function
# x : — the parameter name within the function
# float(x[1:-1]) : — what to do with the parameter
#
#
# The .apply() method is going through every record one-by-one in the df.item_price series, where ip is each record.
# The lambda function converts the value of each df.item_price record in float.
#
# They are especially convenient in data analysis because, as you’ll see, there are many cases where data transformation functions will take functions as arguments. It’s often less typing (and clearer) to pass a lambda function as opposed to writing a full-out function declaration or even assigning the lambda function to a local variable.
#
# This might be a strange pattern to see the first few times, but when you’re writing short functions, the lambda function allows you to work more quickly than the other function definition.
ip = lambda x: float(x[1:-1])
df.item_price = df.item_price.apply(ip)
# Now lets verify the data types again
df.dtypes
df.head(5)
# Yes. We have the data types for all columns in the required format. Also we got rid of $ from item_price
#
# --------------------------------------------------------------------------------------------------------------------------------
# --------------------------------------------------------------------------------------------------------------------------------
# +
# To displays a table of detailed distribution of Statistical information for each of the numerical attributes in our data frame, we use the below function
df.describe()
# -
# This dataset is not having much numerical fields to draw up any conclusion from above Statistical Info.
# At a time max 15 quantity of items are ordered. And below function (value_counts) shows the count of unique values of any column also indicates the same :)
# Lowest price of any item is `$`1.09, max price is `$`44.25
# Interpretation is the process of assigning meaning to the collected information and determining the conclusions, significance, and implications of the findings. So lets understand each of these statistical terms from EDA perspective.
#
# Mean –
# The mean score represents a numerical average for a set of responses.For a data set, the terms arithmetic mean, mathematical expectation, and sometimes average are used synonymously to refer to a central value of a discrete set of numbers. If the data set were based on a series of observations obtained by sampling from a statistical population, the arithmetic mean is termed the sample mean to distinguish it from the population mean.
# Here mean item_price is $7.46
#
# Standard deviation –
# The standard deviation represents the distribution of the responses around the mean. It indicates the degree of consistency among the responses. The standard deviation, in conjunction with the mean, provides a better understanding of the data. For example, if the mean is 7.46 with a standard deviation (StD) of 4.24, then two-thirds of the item_price lie between 3.22 (7.46 – 4.24) and 11.7 (7.46 + 4.24).
#
#
# Other values min, 25`%`, 50`%`, 75`%` & max helps us to gain maximum insight into the data set and its underlying structure.
#
#
# --------------------------------------------------------------------------------------------------------------------------------
#
#
# Lets learn few basic statistics functions
#
# df.describe() -- Summary statistics for numerical columns
#
# df.mean() -- Return the mean of all columns
#
# df.corr() -- Finds the correlation between columns in a DataFrame
#
# df.count() -- Counts the number of non-null values in each DataFrame column
#
# df.max() -- Finds the highest value in each column
#
# df.min() -- Finds the lowest value in each column
#
# df.median() -- Finds the median of each column
#
# df.std() -- Finds the standard deviation of each column
# +
# Statistical Information :
print('Display Mean of all available Columns : \n' + str(df.mean()) + '\n')
print('Display correlation between columns in a DataFrame : \n' + str(df.corr()) + '\n')
print('Display counts of non-null values in each DataFrame column : \n' + str(df.count()) + '\n')
print('Display highest value in each column : \n' + str(df.max()) + '\n')
print('Display lowest value in each column : \n' + str(df.min()) + '\n')
print('Display median of each column : \n' + str(df.median()) + '\n')
print('Display median of each column : \n' + str(df.std()))
# -
# As EDA is used to identify systematic relations between variables when there are no (or not complete) a expectations as to the nature of those relations, exploratory data analysis methods include both simple basic statistics and more advanced techniques designed to identify patterns in multivariate data sets.
#
# This statistical info helps to identify distribution of variables, skew or non-normal patterns in data, reviewing correlation (Correlation is a measure of the relation between two or more variables) for coefficients that meet certain thresholds, or examining data slice by slice systematically reviewing combinations of levels of control variables.
#
# --------------------------------------------------------------------------------------------------------------------------------
#
# As apparent from counts of non-null values in each column there are around (4622 - 3376 =) 1246 nulls present in choice_description column.
#
# Some conclusions might not report any information on outcomes of interest to the review. For example, there may be no information on quality, or on serious adverse effects. It is often difficult to determine whether this is because the outcome was not measured or because the outcome was not reported.
#
# Reasons for handling NaNs :
# • One might end up drawing an inaccurate inference about the data
# • Due to improper handling, the result obtained will differ from ones where the missing values are present
# • Probability cannot be predicted from the variables in the model which involves those types of missing values that are not randomly distributed across the observations
#
# Now we will check various functions to see if there are any null values present in any other columns
#
# df.isnull() -- Returns False for not null values in entire dataset
#
# df['column_name'].notnull() -- Returns True for not null columns else returns False for all rows in the mentioned column
#
#
#
# In case if you want to insert some null values in Dataset we have below function :
#
# df.iloc[3:5,0] = np.nan -- This will insert NaN in 3rd & 4th row's 0th (1st) column
# df.iloc[5:8,2] = np.nan -- This will insert NaN in 5th, 6th & 7th row's 2nd (3rd) column
#
# --------------------------------------------------------------------------------------------------------------------------------
#
# Points to Remember for NaNs :
#
# When summing data, NA (missing) values will be treated as zero
# If the data are all NA, the result will be NA
# Methods like cumsum and cumprod ignore NA values, but preserve them in the resulting arrays
# NA groups in GroupBy are automatically excluded
#
df['item_name'].notnull().tail(5)
df.isnull().head(5)
# As a general rule, most methodologists believe that missing data should not be used as a reason to exclude a study from a systematic review. It is more appropriate to include the study in the review, and to discuss the potential implications of its absence from a meta-analysis.
#
#
# Pandas objects are equipped with various data manipulation methods for dealing with missing data by either filling the missing data with some values ( constant value, mean value, value from previous row etc) or we can drop the missing value
#
# -------------------------------------------------------------------------------------------------------------------------------
#
# To simply exclude labels from a data set which refer to missing data, use the dropna method:
#
# df.dropna(axis=0) -- Drop entire row which contain NaN values
# df.dropna(axis=1) -- Drop entire column which contain NaN values
#
# By default, dropna() will drop all rows in which any null value is present
#
# But this drops some good data as well; one might rather be interested in dropping rows or columns with all NA values, or a majority of NA values. This can be specified through the how or thresh parameters, which allow fine control of the number of nulls to allow through.
#
# The default is how='any', such that any row or column (depending on the axis keyword) containing a null value will be dropped. You can also specify how='all', which will only drop rows/columns which are all null values.
#
# df.dropna(axis=1, how='all')
#
# For finer control, the thresh parameter lets us specify a minimum number of non-null values for the row/column to be kept.
#
# df.dropna(thresh=3)
#
# Here the only those row will be dropped which contain only two non-null values.
#
#
# -------------------------------------------------------------------------------------------------------------------------------
#
# Sometimes rather than dropping NA values, you’d rather replace them with a valid value. This value might be a single number like zero, or it might be some sort of imputation or interpolation from the good values
#
#
# Filling the Missing Values :
# The fillna function can “fill in” NA values with non-null data in a couple of ways & returns a copy of the array with the null values replaced :
#
# df.fillna(0) -- Fill all the NaN values with Zero
# df['column_name'].fillna('missing') -- Fill all the NaN values from mentioned column_name with "missing"
#
# We can propagate non-null values forward or backward
# df.fillna(method='pad', limit=1) -- pad / ffill Fill values forward; limit is for limiting the amount of fillings
# bfill / backfill Fill values backward
# dff.fillna(dff.mean()) -- Fill NaN from all columns with mean value
# dff.fillna(dff.mean()['B':'C']) -- Fill NaN from all column B & C with mean value
#
# We can specify a forward-fill to propagate the previous value forward : data.fillna(method='ffill')
#
# we can also specify an axis along-which a back-fill to propagate the next values backward : df.fillna(method='bfill', axis=1)
#
# Please Note :
# If a previous/next value is not available during a forward fill / backward fill, the NA value remains.
#
# --------------------------------------------------------------------------------------------------------------------------------
# --------------------------------------------------------------------------------------------------------------------------------
#
# Coming now are the ways to slice & dice the data.
#
# The axis labeling information in pandas objects serves many purposes:
#
# • Identifies data (i.e. provides metadata) using known indicators, important for analysis, visualization, and interactive console display
# • Enables automatic and explicit data alignment
# • Allows intuitive getting and setting of subsets of the data set
# • Preliminary selection of appropriate models
# • Detection of mistakes
# --------------------------------------------------------------------------------------------------------------------------------
#
# In Python, indexing operators [] and attribute operator . provide quick and easy access to pandas data structures across a wide range of use cases.
#
# Now we will see various ways to select the data. For this we have below functions :
#
# df['Column_Name'] -- Returns column with label Column_Name as Series
#
# df[[col1, col2]] -- Return Columns as a new DataFrame
#
# df.iloc[0] -- Selection by position (integer based indexing)
#
# df.loc[0] -- Selection by label
#
# df.iloc[0,:] -- Returns first row
#
# df.iloc[0,0] -- Returns first element of first column
#
# --------------------------------------------------------------------------------------------------------------------------------
#
# Few points to remember about Index :
#
# Add a date index :
# df.index = pd.date_range('1900/1/30', periods=df.shape[0])
# +
print('Display Column :\n' + str(df['item_name']))
print('Display any combination of Columns : \n' + str(df[['item_name','choice_description']]))
# Display only 2 columns
rev_left = df.loc[:, [ 'item_name', 'item_price' ]]
rev_left.tail(6)
print('Display any row from the Dataset by mentioning the position/row number :\n' + str(df.iloc[34]))
#this will display 34th row from the Dataset
print('Display any row from the Dataset by mentioning the index :\n' + str(df.loc[34]))
# Both functions, loc[9] & iloc[9] display same result
# -
# --------------------------------------------------------------------------------------------------------------------------------
# --------------------------------------------------------------------------------------------------------------------------------
#
#
# As we are analyzing this dataset for understanding customer's preference for food items and the quantity of items ordered which will help the Shop owner decide the requirements of basic raw materials they need to keep in stock, we will not skip the NaNs which are around 1/4th of total rows and that too only for choice_description column.
#
# Lets apply various Filter, Sort or Group By functions to analyse data which would help us to figure out the trends for customer's prefrence
print('Most ordered item in choice_description column: \n' + str(df.choice_description.value_counts().head(5)))
print('Most ordered item in item_name column: \n' + str(df.item_name.value_counts().head(5)))
--------------------------------------------------------------------------------------------------------------------------------
# value_counts function Returns object containing counts of unique values.
df.quantity.value_counts(dropna=False)
# This shows for quantity column we have 10 unique values for which various quantities are ordered. Majority of items are ordered only once. There is just 1 item ordered whose quantity is 15 (max quantity)
#
# This would help the owner know the frequency of the various items ordered.
# We will further explore the data before drawing any specific conclusion.
print('Total items ordered :\n' + str(df.quantity.sum()))
# +
print('TYPE 1 \n Total number of Orders : ' + str(df.order_id.value_counts().count()) + '\n')
orders = len((set([order for order in df.order_id])))
print('TYPE 2 \n Number of orders made in the period: ' + str(orders))
# -
print('Total revenue generated :\n' + str(df.item_price.sum()))
# +
print('TYPE 1 \n Number of different items sold: ' + str(len(df.item_name.unique())) + '\n')
print('TYPE 2 \n Number of different items sold: ' + str(df.item_name.value_counts().count()))
# +
# make the comparison
chipo10 = df[df['item_price'] > 10.00]
chipo10.head()
len(chipo10)
# +
#AS: Average Cost or Average Price?
avg_order = df.item_price.sum() / df.order_id.value_counts().count()
print "The average cost per order is: $%s" % round(avg_order, 2)
# +
#Slicing the dataset Column-wise
# delete the duplicates in item_name and quantity
chipo_filtered = df.drop_duplicates(['item_name','quantity'])
# select only the ones with quantity equals to 1
price_per_item = chipo_filtered[chipo_filtered.quantity == 1]
#
price_per_item = price_per_item[['item_name', 'item_price']]
# sort the values from the most to less expensive
price_per_item.sort_values(by = "item_price", ascending = False)
# -
print "Quantity of most expensive item ordered : "
df.sort_values(by = "item_price", ascending = False).head(1)
print "Number of times people orderd more than one Canned Soda"
# It can be any item
chipo_m1 = df[(df.item_name == "Canned Soda") & (df.quantity > 1)]
len(chipo_m1)
# +
print "Number of times Veggie Salad Bowl is ordered"
# It can be any item
chipo_salad = df[df.item_name == "Veggie Salad Bowl"]
len(chipo_salad)
# -
# This all analysis is a part of EDA and it helps to know the data by letting us know the customer prefernce which will eventually help the owner to increase the revenue
#
# For example, as people have ordered more than one canned soda 20 times, Owner can decide on how many canned soda they need to keep in stock.
#
# Also Veggie Salad Bowl is ordered 18 times, owner can think of improving the test by adding some extra sauce /more choices to it.
# +
# As we are aware that NaNs in group by are automatically excluded, we are not explicitly dropping NaN values
item_nm = df.groupby('item_name')
# -
# Grouping helps EDA in below ways :
#
# 1.Grouping data allows characteristics of the data to be more easily interpreted than would be true if the raw data were to be examined.
# 2.Improves the accuracy/efficiency of estimation.
# 3.Permits greater balancing of statistical power of tests of differences between strata by sampling equal numbers from strata varying widely in size.
#
# NOTE:
# Strata refers to sampling method in which the total population is divided into non-overlapping subgroups.
# To understand the total quantity ordered across each item & the total cost for that item we use below function.
dfn = item_nm.sum()
dfn
# Sorting the above list based on quantity, helps to know the highest/lowest preferred item along with its price
sort_list = dfn.sort_values(['quantity'], ascending=False)
sort_list.head(1)
# From we this we can help the owner decide the name of those items which they need to stock up as per the quantity based on customer's choice:
#
# Coke, Chicken, Fresh Tomato Salsa, Cheese, Black Beans are the top most choice so these need to be stored in higher quantity as compared to other items.
print('Most ordered item :' + str(df.item_name.value_counts().head(1)))
print('Count of Most ordered item :' + str(df.item_name.value_counts().max()))
cnt = df['item_name'].unique()
count = len(cnt)
print('Count of items ordered: ' + str(count))
# Lets group by choice_description column and then display the top 5 items. It is another way of slicing the data from another perspective (choice_description) to know more about the data
#
# We are doing all this as a part of EDA to get insights of our data which would help us to draw some conclusion about customer prefernce and accordingly we can advise the owner to help them grow their revenue.
#
# Now for grouping data based on choice_description.
# +
# Get rid of NaNs
# NaNs in group by are automatically excluded. This step can be excluded.
df2 = df.dropna(how='any')
print('After dropping NaNs : ' )
df2.head(2)
# Group Data by choice_description
ch_des = df2.groupby('choice_description')
# Do summation across groups. Again to know the most preferred choce and the cost.
dfn1 = ch_des.sum()
print('\nAfter grouping Data across choice_description : ')
dfn1.head(2)
# Sort data based on the quantity ordered to display top 'N' & bottom 'N' records
sort_list_cd = dfn1.sort_values(['quantity'], ascending=False)
print('\n Top 5 items after sorting the grouped Data by quantity : ')
sort_list_cd.head(50)
print('\n Last 5 items after sorting the grouped Data by quantity : ')
sort_list_cd.tail(50)
# Sort data based on the quantity ordered to display top 'N' & bottom 'N' records
sort_list_ip = dfn1.sort_values(['item_price'], ascending=False)
print('\n Top 5 items after sorting the grouped Data by item_price : ')
sort_list_ip.head(25)
print('\n Last 5 items after sorting the grouped Data by item_price : ')
sort_list_ip.tail(25)
# -
# Lets explore some more data by grouping on both item_name & choice_description
# +
# Grouping by 2 columns would help us to know the most preferred combination
item_tot = df.groupby(['item_name', 'choice_description']).sum()
it3 = item_tot.dropna(how='any')
#Sorting the grouped data based on item_price
sort_list_ic = it3.sort_values(['item_price'], ascending=False)
print('\n Top 10 items after sorting the grouped Data by item_price :')
sort_list_ic.head(100)
print('\n Last 10 items after sorting the grouped Data by item_price :')
sort_list_ic.tail(100)
# -
# From this we can make out
#
#
# 1. Any changes they can make to improve the taste/quality for the bottom most ordered items. Owner can experiment few of our suggestions as follows :
# For example for almost all top 5 item the preferred Salsa is Fresh Tomato Salsa, so if they can add this instead of just plain Salsa or Roasted Chili Corn Salsa, there is possibility for these bottom most items to go up the list or atleat to generate some more revenue
# For Burrito, there needs to be option for Black Beans & Fresh Tomato Salsa
# Roasted Chili Corn Salsa seems to be least fav so if possible can be replaced by other fav Salsa (e.g. Fresh Tomato Salsa) as per cutomer's choice
# Provide more choices for Crispy Tacos
#
# 2. Reduce the stocking quantity of raw materials which are less frequently used.
#
# 2. Along with these changes owner can reduce the price a bit for few items and then can rise it again based on customer's feedback
#
#
# All this will help the shop owner to increasse their revenue.
# I hope this notebook will help you maximize your efficiency when starting with basic data exploratory analysis in Python.
#
# Python is really a great tool, and is becoming an increasingly popular language as it is the easiest-to-use languages for rapidly whipping your data into shape. What people are increasingly finding is that Python is a suitable language not only for doingresearch and prototyping but also building the production systems, too.
#
# So, learn Python to perform the full life-cycle of any data science project. It includes reading, analyzing, visualizing and finally making predictions.
#
# Next notebook we will learn about making plots and static or interactive visualizations which is one of the most important tasks in data analysis
#
# If you come across any difficulty while practicing this, or you have any thoughts / suggestions / feedback on the post, please feel free to post them through comments below.
| 29,369 |
/02-hw-titanic-exercise.ipynb | 81cbb2332a29aa61de0b38c50f6133959e4aa283 | [
"MIT"
] | permissive | petarivanov9/Machine-Learning-Course-FMI | https://github.com/petarivanov9/Machine-Learning-Course-FMI | 1 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 67,832 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Домашнее задание
#
# 1. Разделить дата сет на трейн и тест в отношение 50:50 70:30 80:20 (с перемешиванием)
# 2. Обучать наши модели на трейне. Предсказывать и замерять метрику R^2 и на трейне и на тесте
# 3. Проверить следующие модели, для каждого разделения: а) sales ~ log_tv + radio б) sales ~ TV + radio в) sales ~ TV + radio + newspaper
#
#
# +
# Common imports
import pandas as pd
import numpy as np
import statsmodels.api as sm
import statsmodels.formula.api as smf
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
import matplotlib
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d
# %matplotlib inline
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
plt.rcParams['figure.figsize'] = (10, 5)
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
import math
# +
import os
import urllib
import shutil
def download_file(url, dir_path="data"):
if not os.path.exists(dir_path):
os.makedirs(dir_path)
file_name = os.path.split(url)[-1]
file_path = os.path.join(dir_path, file_name)
with urllib.request.urlopen(url) as response, open(file_path, 'wb') as out_file:
shutil.copyfileobj(response, out_file)
return file_path
# +
# # Nado?
from sklearn.preprocessing import StandardScaler, normalize, MinMaxScaler
# -
adv_df = pd.read_csv('data/Advertising.csv', usecols=[1,2,3,4])
adv_df.head()
# # Начало
# +
# Добавить кол-ку sq_tv
# adv_df['sq_log'] = adv_df.TV.apply(lambda x: math.log(x))
adv_df['sq_tv'] = adv_df.TV.apply(lambda x: math.pow(x, 0.4))
# Разделить дата сет на трейн и тест в отношение 50:50 70:30 80:20 (с перемешиванием
y = adv_df['sales'].copy()
adv_train, adv_test, y_train, y_test = train_test_split(adv_df, y, test_size=0.5, random_state=42)
adv_train.shape, adv_test.shape
# -
# #### sales ~ TV + radio + newspaper + sq_tv
# +
# Определяем данные
x_cols = adv_train.columns.drop("sales")
y_col = 'sales'
x_train = np.array(adv_train[x_cols])
x_test = np.array(adv_test[x_cols])
y_train = adv_train[y_col]
y_test = adv_test[y_col]
# Обучение на train LinearRegression
lm = LinearRegression().fit(
x_train,
y_train
)
# train model предсказать
y_train_preds = lm.predict(x_train)
# test model предсказать
y_test_preds = lm.predict(x_test)
# -
# минимизируем RSE и максимизируем 𝑅2. 𝑅2∈[0,1] - относительная величина, чем ближе к 1, тем лучше
# Посчитать R^2 train
lm.score(x_train, y_train)
# Посчитать R^2 test
lm.score(x_test, y_test)
# #### sales ~ sq_tv + radio
# +
# Определяем данные
x_cols = ['sq_tv','radio']
y_col = 'sales'
x_train = np.array(adv_train[x_cols])
x_test = np.array(adv_test[x_cols])
y_train = adv_train[y_col]
y_test = adv_test[y_col]
# Обучение на train LinearRegression
lm = LinearRegression().fit(
x_train,
y_train
)
# train model предсказать
y_train_preds = lm.predict(x_train)
# test model предсказать
y_test_preds = lm.predict(x_test)
# -
# Посчитать R^2 train
lm.score(x_train, y_train)
# Посчитать R^2 test
lm.score(x_test, y_test)
# #### sales ~ TV + radio
# +
# Определяем данные
x_cols = ['TV','radio']
y_col = 'sales'
x_train = np.array(adv_train[x_cols])
x_test = np.array(adv_test[x_cols])
y_train = adv_train[y_col]
y_test = adv_test[y_col]
# Обучение на train LinearRegression
lm = LinearRegression().fit(
x_train,
y_train
)
# train model предсказать
y_train_preds = lm.predict(x_train)
# test model предсказать
y_test_preds = lm.predict(x_test)
# -
# Посчитать R^2 train
lm.score(x_train, y_train)
# Посчитать R^2 test
lm.score(x_test, y_test)
# #### sales ~ TV + radio + newspaper
# +
# Определяем данные
x_cols = ['TV','radio','newspaper']
y_col = 'sales'
x_train = np.array(adv_train[x_cols])
x_test = np.array(adv_test[x_cols])
y_train = adv_train[y_col]
y_test = adv_test[y_col]
# Обучение на train LinearRegression
lm = LinearRegression().fit(
x_train,
y_train
)
# train model предсказать
y_train_preds = lm.predict(x_train)
# test model предсказать
y_test_preds = lm.predict(x_test)
# -
# Посчитать R^2 train
lm.score(x_train, y_train)
# Посчитать R^2 test
lm.score(x_test, y_test)
# Для 80:20: везде test значение R^2 выше, чем train.
# Наивысшее значени R^2 test = 0.942563909350695 для sales ~ sq_tv + radio. Good choice.
# Для 70:30: везде test значение R^2 ниже, чем train.
# Наивысшее значени R^2 test = 0.9271446043449548 для sales ~ sq_tv + radio.
# Для 50:50: везде test значение R^2 ниже, чем train, кроме sales ~ sq_tv + radio
# Наивысшее значени R^2 test = 0.9310741259261295 для sales ~ sq_tv + radio.
'Name', 'Age', 'SibSp', 'Parch', 'Ticket',
'Fare', 'FamilySize',
'Cabin'
], axis=1)
return data
data = prepare_data(data)
data.columns
# -
data.info()
# +
X = data.drop('Survived', axis=1)
y = data['Survived']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0, stratify=y)
model = RandomForestClassifier(random_state=0).fit(X_train, y_train)
print("train score: ", model.score(X_train, y_train))
print("test score: ", model.score(X_test, y_test))
search = GridSearchCV(model, {'n_estimators': [ 10, 20, 25, 30, 50, 60, 80, 100],
'max_depth': [2, 4, 5, 6, 8, 10, 12, 15],
'criterion': ['gini','entropy']
})
search.fit(X, y)
print(search.best_score_)
print(search.best_estimator_)
pd.DataFrame(search.cv_results_)[['rank_test_score', 'mean_test_score', 'params']].sort_values(by='rank_test_score').head(5)
# -
model = RandomForestClassifier(max_depth=4, random_state=0, n_estimators=70, criterion='gini')
model.fit(X_train, y_train)
print("train score: ", model.score(X_train, y_train))
print("test score: ", model.score(X_test, y_test))
# +
test = pd.read_csv('data/titanic/test.csv', index_col=['PassengerId'])
test = create_title(test)
test = add_age_by_title(test)
test = add_age_group(test)
test = add_fare_group(test)
test = prepare_data(test)
test.columns
test.info()
# +
predictions = model.predict(test)
frame = pd.DataFrame({
'PassengerId': pd.read_csv('data/titanic/test.csv').PassengerId,
'Survived': predictions
})
frame = frame.set_index('PassengerId')
frame.to_csv('data/titanic/predictions.csv')
frame.head()
# -
| 7,012 |
/ShEx_Python_Load_class.ipynb | 1863dd604e6772de175b1fc768754a83c9a29060 | [] | no_license | andrawaag/ShEx_Jupyter_kernel | https://github.com/andrawaag/ShEx_Jupyter_kernel | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 3,665 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ShEx
import pprint
input_url = "https://www.wikidata.org/wiki/Special:EntityData/Q42"
input_shex = "https://github.com/shexSpec/schemas/raw/master/Wikidata/pathways/Wikipathways/wikipathways.shex"
shex_start = "wd:Q42"
validation = ShEx.validate(input_shex, input_url)
pprint.pprint(validation.output["errors"])
ShEx.validate(input_shex, input_url)
pprint.pprint(validation.output["errors"])
| 668 |
/dog_app.ipynb | 9c57d2759ec7e17b01f0e13959129b989fca364a | [] | no_license | shwe87/project-dog-classification-udacity | https://github.com/shwe87/project-dog-classification-udacity | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 2,724,036 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ctrl.modeling import CTRLLMHeadModel as PDCTRLLMHeadModel
from ctrl import CTRLTokenizer
import paddle
paddle.set_grad_enabled(False)
pdmodel = PDCTRLLMHeadModel.from_pretrained("./pd/ctrl")
pdmodel.eval()
pd_tokenizer = CTRLTokenizer.from_pretrained("./pd/ctrl")
input_text_list = [
"Diet English : I lost 10 kgs! ; German : ",
"Reviews Rating: 5.0",
"Questions Q: What is the capital of India?",
"Books Weary with toil, I haste me to my bed,"
]
for inputs_text in input_text_list:
inputs = paddle.to_tensor([pd_tokenizer(inputs_text)["input_ids"]],dtype="int64")
outputs = pdmodel.generate(inputs,max_length=128-inputs.shape[1],repetition_penalty=1.2,temperature=0)[0][0]
decode_outputs = pd_tokenizer.convert_tokens_to_string(pd_tokenizer.convert_ids_to_tokens(outputs.cpu()))
print(f"{inputs_text}\n {decode_outputs}")
print("="*50)
e_highest = dfdate.groupby('acq_date').sum().sort_values(by='frp',ascending=False)
dfdate_highest.head(10)
plt.figure(figsize=(10,5))
sns.set_palette("pastel")
ax = sns.barplot(x='acq_date',y='frp',data=df)
for ind, label in enumerate(ax.get_xticklabels()):
if ind % 10 == 0: # every 10th label is kept
label.set_visible(True)
else:
label.set_visible(False)
ax.set_xticklabels(ax.get_xticklabels(), rotation=40, ha="right")
plt.xlabel("Date")
plt.ylabel('FRP (fire radiation power)')
plt.title("time line of bushfire in Australia")
plt.tight_layout()
from mpl_toolkits.mplot3d import Axes3D
from sklearn.preprocessing import StandardScaler
import os
sns.pairplot(data=dffam)
| 1,851 |
/LS_DS_422_Backprop_Assignment.ipynb | bdd572f43c4f13f3f67b336bca0e2db3821b5b26 | [] | no_license | mooglol/DS-Unit-4-Sprint-2-Neural-Networks | https://github.com/mooglol/DS-Unit-4-Sprint-2-Neural-Networks | 0 | 0 | null | 2019-12-10T22:24:54 | 2019-11-07T16:55:37 | null | Jupyter Notebook | false | false | .py | 15,161 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import necessary libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import yfinance as yf
# ## Scrape historic stock data
TD = yf.Ticker('TD.TO')
df = TD.history(start = "2001-01-02")
df
# Drop unnecessary column
df = df.drop(['Dividends','Stock Splits'], axis = 1)
df.head()
# ## Stock Price Analysis
# Close Price Movement
plt.figure(figsize =(15,10))
plt.plot(df['Close'])
plt.xlabel('Date')
plt.ylabel('Price($)')
plt.title('TD Stock Price 01/02/2001 - 06/29/2021', fontsize =20)
# ## Feature Generation
## 31 new features
# 6 originial features
new_df = pd.DataFrame()
def generate_features(df):
return(new_df)
new_df['open'] = df['Open']
new_df['open_1'] = df['Open'].shift(1)
new_df['high_1'] = df['High'].shift(1)
new_df['low_1'] = df['Low'].shift(1)
new_df['close_1'] = df['Close'].shift(1)
new_df['volume_1'] = df['Volume'].shift(1)
# Average closed price
new_df['avg_price_5'] = df['Close'].rolling(window=5).mean().shift(1)
new_df['avg_price_30'] = df['Close'].rolling(window=21).mean().shift(1)
new_df['avg_price_365'] = df['Close'].rolling(window=252).mean().shift(1)
new_df['ratio_avg_price_5_30'] = new_df['avg_price_5']/new_df['avg_price_30']
new_df['ratio_avg_price_5_365'] = new_df['avg_price_5']/new_df['avg_price_365']
new_df['ratio_avg_price_30_365'] = new_df['avg_price_30']/new_df['avg_price_365']
# Average volume
new_df['avg_vol_5'] = df['Volume'].rolling(window=5).mean().shift(1)
new_df['avg_vol_30'] = df['Volume'].rolling(window=21).mean().shift(1)
new_df['avg_vol_365'] = df['Volume'].rolling(window=252).mean().shift(1)
new_df['ratio_avg_vol_5_30'] = new_df['avg_vol_5']/new_df['avg_vol_30']
new_df['ratio_avg_vol_5_365'] = new_df['avg_vol_5']/new_df['avg_vol_365']
new_df['ratio_avg_vol_30_365'] = new_df['avg_vol_30']/new_df['avg_vol_365']
# Standard deviation of closed price
new_df['std_price_5'] = df['Close'].rolling(window=5).std().shift(1)
new_df['std_price_30'] = df['Close'].rolling(window=21).std().shift(1)
new_df['std_price_365'] = df['Close'].rolling(window=252).std().shift(1)
new_df['ratio_std_price_5_30'] = new_df['std_price_5']/new_df['std_price_30']
new_df['ratio_std_price_5_365'] = new_df['std_price_5']/new_df['std_price_365']
new_df['ratio_std_price_30_365'] = new_df['std_price_30']/new_df['std_price_365']
# Standard deviation of volumes
new_df['std_vol_5'] = df['Volume'].rolling(window=5).std().shift(1)
new_df['std_vol_30'] = df['Volume'].rolling(window=21).std().shift(1)
new_df['std_vol_365'] = df['Volume'].rolling(window=252).std().shift(1)
new_df['ratio_std_vol_5_30'] = new_df['std_vol_5']/new_df['std_vol_30']
new_df['ratio_std_vol_5_365'] = new_df['std_vol_5']/new_df['std_vol_365']
new_df['ratio_std_vol_30_365'] = new_df['std_vol_30']/new_df['std_vol_365']
# Return
new_df['return_1'] = ((df['Close']-df['Close'].shift(1))/df['Close'].shift(1)).shift(1)
new_df['return_5'] = ((df['Close']-df['Close'].shift(5))/df['Close'].shift(5)).shift(1)
new_df['return_30'] = ((df['Close']-df['Close'].shift(21))/df['Close'].shift(21)).shift(1)
new_df['return_365'] = ((df['Close']-df['Close'].shift(365))/df['Close'].shift(365)).shift(1)
# Moving average
new_df['moving_avg_5'] = new_df['return_1'].rolling(window=5).mean()
new_df['moving_avg_30'] = new_df['return_1'].rolling(window=21).mean()
new_df['moving_avg_365'] = new_df['return_1'].rolling(window=252).mean()
# target
new_df['close'] = df['Close']
new_df = generate_features(df)
new_df = new_df.dropna(axis=0)
new_df
# TD Stock Volatility Visualization
plt.figure(figsize=(20,10))
plt.plot(new_df['moving_avg_5'],'g-', label ='5-day Moving Average')
plt.plot(new_df['moving_avg_30'],'r-', label = '30-day Moving Average')
plt.plot(new_df['moving_avg_365'], 'y-', label = '1-year Moving Average')
plt.title("Stock Volatility 2001-2021", fontsize = 20)
plt.legend()
# Volatility in the past 200 trading days
plt.figure(figsize=(20,10))
plt.plot(new_df['moving_avg_5'].iloc[-200:],'g-', label ='5-day Moving Average')
plt.plot(new_df['moving_avg_30'].iloc[-200:],'r-', label = '30-day Moving Average')
plt.plot(new_df['moving_avg_365'].iloc[-200:], 'y-', label = '1-year Moving Average')
plt.title("Stock Volatility over the Past 200 Trading Days", fontsize = 20)
plt.xlabel("Date")
plt.legend()
# ## Linear Regression
# ### Train with Linear Regression
# Training Set
import datetime
start_training = datetime.date(2001,1,2)
end_training = datetime.date(2019,12,31)
features = new_df.drop(columns = ['close'],axis=1)
target = new_df['close']
X_train = features[start_training:end_training]
y_train = target[start_training:end_training]
X_train.shape
#Test Set
start_test = datetime.date(2020,1,2)
end_test = datetime.date(2021,6,29)
X_test = features[start_test:end_test]
y_test = target[start_test:end_test]
X_test.shape
# Standardization (due to price and ratio being in different scale)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_scaled_train = scaler.transform(X_train)
X_scaled_test = scaler.transform(X_test)
# Apply Stochastic Gradient Descent to train
from sklearn.linear_model import SGDRegressor
regressor = SGDRegressor(loss = 'squared_loss', learning_rate = 'constant', alpha = 0.0001, eta0 = 0.01, max_iter=1000)
regressor.fit(X_scaled_train, y_train)
predictions = regressor.predict(X_scaled_test)
predictions
# ### Evaluation
print('Training set score:{:.2f}'.format(regressor.score(X_scaled_train, y_train)))
print('Test set score:{:.2f}'.format(regressor.score(X_scaled_test, y_test)))
# ### Plotting the Prediction
| 5,839 |
/Question 2.ipynb | 64d0302d4d5758b0b49a11b0e67da2cfb9dd288b | [] | no_license | sulhicader/datathon_repo | https://github.com/sulhicader/datathon_repo | 0 | 1 | null | null | null | null | Jupyter Notebook | false | false | .r | 133,606 | # -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(dplyr)
library(ggplot2)
library(tidyr)
# # Read the tsv file and assign it to a variable
twitterdata <- read.table("sample_twitter_personal_data.tsv",header = TRUE)
head(twitterdata)
# # Q1. List all the headers in the dataset.
headers <- matrix(colnames(twitterdata))
headers
# # Q2. Order (ascending) the headers and assign numbers for the ordered headers.
ordered_headers <- matrix(sort(headers , decreasing=FALSE))
index <- seq(1,length(ordered_headers))
data.frame( index , ordered_headers )
# # Q3. List all Twitter users whose "followers_count" is greater (>) than 100.
twitterdata_great_1000 <- twitterdata %>%
filter( followers_count > 1000 )
head(twitterdata_great_1000)
# # Q4. List all the twitter MALE users whose "followers_count" is greater (>) than 100.
# +
twitterdata_male_100 <- twitterdata %>%
filter( followers_count > 100 ) %>%
filter(gender == "male")
head(twitterdata_male_100)
# -
# # Q5. It is a saying,
# “ Males always initiate to follow another Twitter account (female/male account) FIRST than
# Female ”
# To prove the above statement, analyze the data and provide some evidence.
boxplot(twitterdata$followers_count ~ twitterdata$gender, data = mtcars, xlab = "gender",ylab = "followers_count")
# According to the above boxplot we can see there are lot of outliers involve. Let's do a two sample t-test using two groups of data
male_numbers <- twitterdata %>%
drop_na(gender)%>%
filter( gender == "male" )
female_numbers <- twitterdata %>%
drop_na(gender)%>%
filter( gender == "female" )
t.test(male_numbers$followers_count,female_numbers$followers_count,paired=FALSE , alternative="greater")
# Our null hypothesis : mean (male) <= mean(female)
# alternative hypothesis : mean (male) > mean(female)
#
# according to the t-test result t-score = -28.565
#
# t-score lies within the confidence interval. So null hypothesis is rejected. So according to the alternative hypothesis above statement is false.
# # Q6.Your manager has requested study the data and asked you to provide a graphical representation for the following use cases,
# 1.HISTOGRAMS for day column in DOB (eg: dob_day)
dob_day <- twitterdata$dob_day
qplot(dob_day, geom="histogram" , bins=31 , width= 1 )
# 2.FREQUENCY PLOT for the followers count
# +
by_followers_count <- twitterdata %>%
count(followers_count)
ggplot(by_followers_count, aes(x=followers_count,y=n ))+
geom_col()+
coord_cartesian(xlim = c(0, 100), ylim = c(0, 2000))
#expand_limits(y=c(0,2000) , x = c(0,2000))
# -
# 3.FREQUENCY PLOT for the age of the users over the years in the sample dataset
# +
by_age <- twitterdata %>%
count(age)
ggplot(by_age, aes(x=age,y=n, width= 0.5 ))+
geom_col()+
expand_limits(y=0)
# -
# 4.Plot BAR CHART to indicate and show the total hearts given by male, female users
# +
by_gender <- twitterdata %>%
drop_na(gender)%>%
group_by( gender ) %>%
summarize(heart_count=sum(heart))
by_gender
# -
ggplot(by_gender, aes(x=gender,y=heart_count , width= 0.2 , height=0.1 ))+
geom_col()+
expand_limits(y=0)
# 5.Analyze and visualize in CHART to confirm that people use a mobile/web interface to
# experience twitter
# +
heart_mobile <- twitterdata %>%
summarize( heart_mobile = sum(mobile_app_heart))
heart_mobile
# +
heart_web <- twitterdata %>%
summarize( heart_web = sum(web_heart))
heart_web
# +
source_int <- data.frame( c("heart_web" ,"heart_mobile" ) , c(heart_web[1,1] , heart_mobile[1,1]) )
colnames(source_int) <- c("source" , "count")
ggplot(source_int, aes(x=source_int$source,y=source_int$count , width=0.1 ))+
geom_col()+
expand_limits(y=0)
# -
# # Q7. Based on your critical analysis write down in points your thoughts and suggestions to improve the twitter features.
head(twitterdata %>%count(heart))
# According to the above summarization there are 22308 twitter ids with 0 heart reaction. So increacing the numbers of reaction type may increace the interaction
# According to the bar chart in Q6. 4. we can see that men interaction is less that female. So add features whose can attract men.
head(twitterdata %>%count(age))
# According to above summarization there are no user with age grater than 18. Introduce feature such that people with age grater than 18 attract to twitter
# When we go through the dataframe we can't see location information. if there are location information, can suggest news based on locations.
| 4,722 |
/assignment2/FullyConnectedNets.ipynb | 80494edc97fc35b8c198bb4d3cb1640333e60640 | [] | no_license | minhcanh99/cs231n | https://github.com/minhcanh99/cs231n | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 373,553 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["pdf-title"]
# # Fully-Connected Neural Nets
# In the previous homework you implemented a fully-connected two-layer neural network on CIFAR-10. The implementation was simple but not very modular since the loss and gradient were computed in a single monolithic function. This is manageable for a simple two-layer network, but would become impractical as we move to bigger models. Ideally we want to build networks using a more modular design so that we can implement different layer types in isolation and then snap them together into models with different architectures.
# + [markdown] tags=["pdf-ignore"]
# In this exercise we will implement fully-connected networks using a more modular approach. For each layer we will implement a `forward` and a `backward` function. The `forward` function will receive inputs, weights, and other parameters and will return both an output and a `cache` object storing data needed for the backward pass, like this:
#
# ```python
# def layer_forward(x, w):
# """ Receive inputs x and weights w """
# # Do some computations ...
# z = # ... some intermediate value
# # Do some more computations ...
# out = # the output
#
# cache = (x, w, z, out) # Values we need to compute gradients
#
# return out, cache
# ```
#
# The backward pass will receive upstream derivatives and the `cache` object, and will return gradients with respect to the inputs and weights, like this:
#
# ```python
# def layer_backward(dout, cache):
# """
# Receive dout (derivative of loss with respect to outputs) and cache,
# and compute derivative with respect to inputs.
# """
# # Unpack cache values
# x, w, z, out = cache
#
# # Use values in cache to compute derivatives
# dx = # Derivative of loss with respect to x
# dw = # Derivative of loss with respect to w
#
# return dx, dw
# ```
#
# After implementing a bunch of layers this way, we will be able to easily combine them to build classifiers with different architectures.
#
# In addition to implementing fully-connected networks of arbitrary depth, we will also explore different update rules for optimization, and introduce Dropout as a regularizer and Batch/Layer Normalization as a tool to more efficiently optimize deep networks.
#
# + tags=["pdf-ignore"]
# As usual, a bit of setup
from __future__ import print_function
import time
import numpy as np
import matplotlib.pyplot as plt
from cs231n.classifiers.fc_net import *
from cs231n.data_utils import get_CIFAR10_data
from cs231n.gradient_check import eval_numerical_gradient, eval_numerical_gradient_array
from cs231n.solver import Solver
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# + tags=["pdf-ignore"]
# Load the (preprocessed) CIFAR10 data.
data = get_CIFAR10_data()
for k, v in list(data.items()):
print(('%s: ' % k, v.shape))
# -
# # Affine layer: foward
# Open the file `cs231n/layers.py` and implement the `affine_forward` function.
#
# Once you are done you can test your implementaion by running the following:
# +
# Test the affine_forward function
num_inputs = 2
input_shape = (4, 5, 6)
output_dim = 3
input_size = num_inputs * np.prod(input_shape)
weight_size = output_dim * np.prod(input_shape)
x = np.linspace(-0.1, 0.5, num=input_size).reshape(num_inputs, *input_shape)
w = np.linspace(-0.2, 0.3, num=weight_size).reshape(np.prod(input_shape), output_dim)
b = np.linspace(-0.3, 0.1, num=output_dim)
out, _ = affine_forward(x, w, b)
correct_out = np.array([[ 1.49834967, 1.70660132, 1.91485297],
[ 3.25553199, 3.5141327, 3.77273342]])
# Compare your output with ours. The error should be around e-9 or less.
print('Testing affine_forward function:')
print('difference: ', rel_error(out, correct_out))
# -
# # Affine layer: backward
# Now implement the `affine_backward` function and test your implementation using numeric gradient checking.
# +
# Test the affine_backward function
np.random.seed(231)
x = np.random.randn(10, 2, 3)
w = np.random.randn(6, 5)
b = np.random.randn(5)
dout = np.random.randn(10, 5)
dx_num = eval_numerical_gradient_array(lambda x: affine_forward(x, w, b)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: affine_forward(x, w, b)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: affine_forward(x, w, b)[0], b, dout)
_, cache = affine_forward(x, w, b)
dx, dw, db = affine_backward(dout, cache)
# The error should be around e-10 or less
print('Testing affine_backward function:')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# -
# # ReLU activation: forward
# Implement the forward pass for the ReLU activation function in the `relu_forward` function and test your implementation using the following:
# +
# Test the relu_forward function
x = np.linspace(-0.5, 0.5, num=12).reshape(3, 4)
out, _ = relu_forward(x)
correct_out = np.array([[ 0., 0., 0., 0., ],
[ 0., 0., 0.04545455, 0.13636364,],
[ 0.22727273, 0.31818182, 0.40909091, 0.5, ]])
# Compare your output with ours. The error shouldt be on the order of e-8
print('Testing relu_forward function:')
print('difference: ', rel_error(out, correct_out))
# -
# # ReLU activation: backward
# Now implement the backward pass for the ReLU activation function in the `relu_backward` function and test your implementation using numeric gradient checking:
# +
np.random.seed(231)
x = np.random.randn(10, 10)
dout = np.random.randn(*x.shape)
dx_num = eval_numerical_gradient_array(lambda x: relu_forward(x)[0], x, dout)
_, cache = relu_forward(x)
dx = relu_backward(dout, cache)
# The error should be on the order of e-12
print('Testing relu_backward function:')
print('dx error: ', rel_error(dx_num, dx))
# + [markdown] tags=["pdf-inline"]
# ## Inline Question 1:
#
# We've only asked you to implement ReLU, but there are a number of different activation functions that one could use in neural networks, each with its pros and cons. In particular, an issue commonly seen with activation functions is getting zero (or close to zero) gradient flow during backpropagation. Which of the following activation functions have this problem? If you consider these functions in the one dimensional case, what types of input would lead to this behaviour?
# 1. Sigmoid
# 2. ReLU
# 3. Leaky ReLU
#
# ## Answer:
# 1 & 2 will get saturated
#
# Sigmoid -> when the value of activate function nearly boundaries. Meaning reach to 1 or 0. Then the derivative is very small or close to 0.
#
# ReLU -> When the value of ReLU function is 0 or close 0. So the derivative is 0 or very close to 0.
#
# -
# # "Sandwich" layers
# There are some common patterns of layers that are frequently used in neural nets. For example, affine layers are frequently followed by a ReLU nonlinearity. To make these common patterns easy, we define several convenience layers in the file `cs231n/layer_utils.py`.
#
# For now take a look at the `affine_relu_forward` and `affine_relu_backward` functions, and run the following to numerically gradient check the backward pass:
# +
from cs231n.layer_utils import affine_relu_forward, affine_relu_backward
np.random.seed(231)
x = np.random.randn(2, 3, 4)
w = np.random.randn(12, 10)
b = np.random.randn(10)
dout = np.random.randn(2, 10)
out, cache = affine_relu_forward(x, w, b)
dx, dw, db = affine_relu_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: affine_relu_forward(x, w, b)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: affine_relu_forward(x, w, b)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: affine_relu_forward(x, w, b)[0], b, dout)
# Relative error should be around e-10 or less
print('Testing affine_relu_forward and affine_relu_backward:')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# -
# # Loss layers: Softmax and SVM
# You implemented these loss functions in the last assignment, so we'll give them to you for free here. You should still make sure you understand how they work by looking at the implementations in `cs231n/layers.py`.
#
# You can make sure that the implementations are correct by running the following:
# +
np.random.seed(231)
num_classes, num_inputs = 10, 50
x = 0.001 * np.random.randn(num_inputs, num_classes)
y = np.random.randint(num_classes, size=num_inputs)
dx_num = eval_numerical_gradient(lambda x: svm_loss(x, y)[0], x, verbose=False)
loss, dx = svm_loss(x, y)
# Test svm_loss function. Loss should be around 9 and dx error should be around the order of e-9
print('Testing svm_loss:')
print('loss: ', loss)
print('dx error: ', rel_error(dx_num, dx))
dx_num = eval_numerical_gradient(lambda x: softmax_loss(x, y)[0], x, verbose=False)
loss, dx = softmax_loss(x, y)
# Test softmax_loss function. Loss should be close to 2.3 and dx error should be around e-8
print('\nTesting softmax_loss:')
print('loss: ', loss)
print('dx error: ', rel_error(dx_num, dx))
# -
# # Two-layer network
# In the previous assignment you implemented a two-layer neural network in a single monolithic class. Now that you have implemented modular versions of the necessary layers, you will reimplement the two layer network using these modular implementations.
#
# Open the file `cs231n/classifiers/fc_net.py` and complete the implementation of the `TwoLayerNet` class. This class will serve as a model for the other networks you will implement in this assignment, so read through it to make sure you understand the API. You can run the cell below to test your implementation.
# +
np.random.seed(231)
N, D, H, C = 3, 5, 50, 7
X = np.random.randn(N, D)
y = np.random.randint(C, size=N)
std = 1e-3
model = TwoLayerNet(input_dim=D, hidden_dim=H, num_classes=C, weight_scale=std)
print('Testing initialization ... ')
W1_std = abs(model.params['W1'].std() - std)
b1 = model.params['b1']
W2_std = abs(model.params['W2'].std() - std)
b2 = model.params['b2']
assert W1_std < std / 10, 'First layer weights do not seem right'
assert np.all(b1 == 0), 'First layer biases do not seem right'
assert W2_std < std / 10, 'Second layer weights do not seem right'
assert np.all(b2 == 0), 'Second layer biases do not seem right'
print('Testing test-time forward pass ... ')
model.params['W1'] = np.linspace(-0.7, 0.3, num=D*H).reshape(D, H)
model.params['b1'] = np.linspace(-0.1, 0.9, num=H)
model.params['W2'] = np.linspace(-0.3, 0.4, num=H*C).reshape(H, C)
model.params['b2'] = np.linspace(-0.9, 0.1, num=C)
X = np.linspace(-5.5, 4.5, num=N*D).reshape(D, N).T
scores = model.loss(X)
correct_scores = np.asarray(
[[11.53165108, 12.2917344, 13.05181771, 13.81190102, 14.57198434, 15.33206765, 16.09215096],
[12.05769098, 12.74614105, 13.43459113, 14.1230412, 14.81149128, 15.49994135, 16.18839143],
[12.58373087, 13.20054771, 13.81736455, 14.43418138, 15.05099822, 15.66781506, 16.2846319 ]])
scores_diff = np.abs(scores - correct_scores).sum()
assert scores_diff < 1e-6, 'Problem with test-time forward pass'
print('Testing training loss (no regularization)')
y = np.asarray([0, 5, 1])
loss, grads = model.loss(X, y)
correct_loss = 3.4702243556
assert abs(loss - correct_loss) < 1e-10, 'Problem with training-time loss'
model.reg = 1.0
loss, grads = model.loss(X, y)
correct_loss = 26.5948426952
assert abs(loss - correct_loss) < 1e-10, 'Problem with regularization loss'
# Errors should be around e-7 or less
for reg in [0.0, 0.7]:
print('Running numeric gradient check with reg = ', reg)
model.reg = reg
loss, grads = model.loss(X, y)
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = eval_numerical_gradient(f, model.params[name], verbose=False)
print('%s relative error: %.2e' % (name, rel_error(grad_num, grads[name])))
# -
# # Solver
# In the previous assignment, the logic for training models was coupled to the models themselves. Following a more modular design, for this assignment we have split the logic for training models into a separate class.
#
# Open the file `cs231n/solver.py` and read through it to familiarize yourself with the API. After doing so, use a `Solver` instance to train a `TwoLayerNet` that achieves at least `50%` accuracy on the validation set.
# +
model = TwoLayerNet()
solver = None
##############################################################################
# TODO: Use a Solver instance to train a TwoLayerNet that achieves at least #
# 50% accuracy on the validation set. #
##############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
best_val = -1
max_count = 10
for count in range(max_count):
reg = 10**np.random.uniform(-4, 0)
lr = 10**np.random.uniform(-3, -4)
hidden_size = 50
model = TwoLayerNet(hidden_dim = hidden_size, reg= reg)
solver = Solver(model, data, update_rule='adam', optim_config={'learning_rate':lr},
lr_decay=0.95, num_epochs=5,batch_size=200, print_every=-1, verbose=False)
solver.train()
val_accuracy = solver.best_val_acc
if (best_val < val_accuracy):
best_val = val_accuracy
print('lr %e reg %e hid %d val accuracy: %f' % (
lr, reg, hidden_size, val_accuracy))
print('Best validation accuracy %f' % best_val)
#lr 1.078914e-04 reg 3.931614e-02 hid 50 val accuracy: 0.512000 -> rmsProp
#lr 1.525314e-04 reg 3.604910e-04 hid 50 val accuracy: 0.501000 -> Adam
#lr 1.676984e-04 reg 1.218733e-02 hid 50 val accuracy: 0.505000 -> Adam
#lr 1.906639e-04 reg 1.324207e-02 hid 50 val accuracy: 0.507000 -> Adam
#lr 1.164046e-04 reg 1.156979e-04 hid 50 val accuracy: 0.513000 -> Adam
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
##############################################################################
# END OF YOUR CODE #
##############################################################################
# +
# Run this cell to visualize training loss and train / val accuracy
lr = 1.164046e-04
reg = 1.156979e-04
hidden_size = 50
model = TwoLayerNet(hidden_dim = hidden_size, reg= reg)
solver = Solver(model, data, update_rule='adam', optim_config={'learning_rate':lr},
lr_decay=0.95, num_epochs=10,batch_size=200, print_every=-1, verbose=False)
solver.train()
plt.subplot(2, 1, 1)
plt.title('Training loss')
plt.plot(solver.loss_history, 'o')
plt.xlabel('Iteration')
plt.subplot(2, 1, 2)
plt.title('Accuracy')
plt.plot(solver.train_acc_history, '-o', label='train')
plt.plot(solver.val_acc_history, '-o', label='val')
plt.plot([0.5] * len(solver.val_acc_history), 'k--')
plt.xlabel('Epoch')
plt.legend(loc='lower right')
plt.gcf().set_size_inches(15, 12)
plt.show()
# -
# # Multilayer network
# Next you will implement a fully-connected network with an arbitrary number of hidden layers.
#
# Read through the `FullyConnectedNet` class in the file `cs231n/classifiers/fc_net.py`.
#
# Implement the initialization, the forward pass, and the backward pass. For the moment don't worry about implementing dropout or batch/layer normalization; we will add those features soon.
# ## Initial loss and gradient check
#
# As a sanity check, run the following to check the initial loss and to gradient check the network both with and without regularization. Do the initial losses seem reasonable?
#
# For gradient checking, you should expect to see errors around 1e-7 or less.
# +
np.random.seed(231)
N, D, H1, H2, C = 2, 15, 20, 30, 10
X = np.random.randn(N, D)
y = np.random.randint(C, size=(N,))
for reg in [0, 3.14]:
print('Running check with reg = ', reg)
model = FullyConnectedNet([H1, H2], input_dim=D, num_classes=C,
reg=reg, weight_scale=5e-2, dtype=np.float64)
loss, grads = model.loss(X, y)
print('Initial loss: ', loss)
# Most of the errors should be on the order of e-7 or smaller.
# NOTE: It is fine however to see an error for W2 on the order of e-5
# for the check when reg = 0.0
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = eval_numerical_gradient(f, model.params[name], verbose=False, h=1e-5)
print('%s relative error: %.2e' % (name, rel_error(grad_num, grads[name])))
# -
# As another sanity check, make sure you can overfit a small dataset of 50 images. First we will try a three-layer network with 100 units in each hidden layer. In the following cell, tweak the **learning rate** and **weight initialization scale** to overfit and achieve 100% training accuracy within 20 epochs.
# +
# TODO: Use a three-layer Net to overfit 50 training examples by
# tweaking just the learning rate and initialization scale.
num_train = 50
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
# max_count = 10
# for count in range(max_count):
# weight_scale = 10**np.random.uniform(-4, 0)
# learning_rate = 10**np.random.uniform(-3, -4)
weight_scale = 1e-1 # Experiment with this!
learning_rate = 7e-4 # Experiment with this!
model = FullyConnectedNet([100, 100],
weight_scale=weight_scale, dtype=np.float64)
solver = Solver(model, small_data,
print_every=10, num_epochs=20, batch_size=25,
update_rule='sgd',
optim_config={
'learning_rate': learning_rate,
}
)
solver.train()
plt.plot(solver.loss_history, 'o')
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.show()
# -
# Now try to use a five-layer network with 100 units on each layer to overfit 50 training examples. Again, you will have to adjust the learning rate and weight initialization scale, but you should be able to achieve 100% training accuracy within 20 epochs.
# +
# TODO: Use a five-layer Net to overfit 50 training examples by
# tweaking just the learning rate and initialization scale.
num_train = 50
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
# max_count = 10
# for count in range(max_count):
# weight_scale = 10**np.random.uniform(-4, 0)
# learning_rate = 10**np.random.uniform(-3, -4)
# print(weight_scale, learning_rate)
# print("-------------------------")
learning_rate = 3e-4 # Experiment with this!
weight_scale = 2e-1 # Experiment with this!
model = FullyConnectedNet([100, 100, 100, 100],
weight_scale=weight_scale, dtype=np.float64)
solver = Solver(model, small_data,
print_every=10, num_epochs=20, batch_size=25,
update_rule='sgd',
optim_config={
'learning_rate': learning_rate,
}
)
solver.train()
plt.plot(solver.loss_history, 'o')
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.show()
# + [markdown] tags=["pdf-inline"]
# ## Inline Question 2:
# Did you notice anything about the comparative difficulty of training the three-layer net vs training the five layer net? In particular, based on your experience, which network seemed more sensitive to the initialization scale? Why do you think that is the case?
#
# ## Answer:
# Training the five layer net is harder than the three layer net. Because the five layer is more sensitive to the initialization scale than the three layer. When we have a deeper network, a small scale leads to vanishing the gradients (small products), on the other hand a large scale leads to exploding the gradients (large products).
#
# -
# # Update rules
# So far we have used vanilla stochastic gradient descent (SGD) as our update rule. More sophisticated update rules can make it easier to train deep networks. We will implement a few of the most commonly used update rules and compare them to vanilla SGD.
# # SGD+Momentum
# Stochastic gradient descent with momentum is a widely used update rule that tends to make deep networks converge faster than vanilla stochastic gradient descent. See the Momentum Update section at http://cs231n.github.io/neural-networks-3/#sgd for more information.
#
# Open the file `cs231n/optim.py` and read the documentation at the top of the file to make sure you understand the API. Implement the SGD+momentum update rule in the function `sgd_momentum` and run the following to check your implementation. You should see errors less than e-8.
# +
from cs231n.optim import sgd_momentum
N, D = 4, 5
w = np.linspace(-0.4, 0.6, num=N*D).reshape(N, D)
dw = np.linspace(-0.6, 0.4, num=N*D).reshape(N, D)
v = np.linspace(0.6, 0.9, num=N*D).reshape(N, D)
config = {'learning_rate': 1e-3, 'velocity': v}
next_w, _ = sgd_momentum(w, dw, config=config)
expected_next_w = np.asarray([
[ 0.1406, 0.20738947, 0.27417895, 0.34096842, 0.40775789],
[ 0.47454737, 0.54133684, 0.60812632, 0.67491579, 0.74170526],
[ 0.80849474, 0.87528421, 0.94207368, 1.00886316, 1.07565263],
[ 1.14244211, 1.20923158, 1.27602105, 1.34281053, 1.4096 ]])
expected_velocity = np.asarray([
[ 0.5406, 0.55475789, 0.56891579, 0.58307368, 0.59723158],
[ 0.61138947, 0.62554737, 0.63970526, 0.65386316, 0.66802105],
[ 0.68217895, 0.69633684, 0.71049474, 0.72465263, 0.73881053],
[ 0.75296842, 0.76712632, 0.78128421, 0.79544211, 0.8096 ]])
# Should see relative errors around e-8 or less
print('next_w error: ', rel_error(next_w, expected_next_w))
print('velocity error: ', rel_error(expected_velocity, config['velocity']))
# -
# Once you have done so, run the following to train a six-layer network with both SGD and SGD+momentum. You should see the SGD+momentum update rule converge faster.
# +
num_train = 4000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
solvers = {}
for update_rule in ['sgd', 'sgd_momentum']:
print('running with ', update_rule)
model = FullyConnectedNet([100, 100, 100, 100, 100], weight_scale=5e-2)
solver = Solver(model, small_data,
num_epochs=5, batch_size=100,
update_rule=update_rule,
optim_config={
'learning_rate': 5e-3,
},
verbose=True)
solvers[update_rule] = solver
solver.train()
print()
plt.subplot(3, 1, 1)
plt.title('Training loss')
plt.xlabel('Iteration')
plt.subplot(3, 1, 2)
plt.title('Training accuracy')
plt.xlabel('Epoch')
plt.subplot(3, 1, 3)
plt.title('Validation accuracy')
plt.xlabel('Epoch')
for update_rule, solver in solvers.items():
plt.subplot(3, 1, 1)
plt.plot(solver.loss_history, 'o', label="loss_%s" % update_rule)
plt.subplot(3, 1, 2)
plt.plot(solver.train_acc_history, '-o', label="train_acc_%s" % update_rule)
plt.subplot(3, 1, 3)
plt.plot(solver.val_acc_history, '-o', label="val_acc_%s" % update_rule)
for i in [1, 2, 3]:
plt.subplot(3, 1, i)
plt.legend(loc='upper center', ncol=4)
plt.gcf().set_size_inches(15, 15)
plt.show()
# -
# # RMSProp and Adam
# RMSProp [1] and Adam [2] are update rules that set per-parameter learning rates by using a running average of the second moments of gradients.
#
# In the file `cs231n/optim.py`, implement the RMSProp update rule in the `rmsprop` function and implement the Adam update rule in the `adam` function, and check your implementations using the tests below.
#
# **NOTE:** Please implement the _complete_ Adam update rule (with the bias correction mechanism), not the first simplified version mentioned in the course notes.
#
# [1] Tijmen Tieleman and Geoffrey Hinton. "Lecture 6.5-rmsprop: Divide the gradient by a running average of its recent magnitude." COURSERA: Neural Networks for Machine Learning 4 (2012).
#
# [2] Diederik Kingma and Jimmy Ba, "Adam: A Method for Stochastic Optimization", ICLR 2015.
# +
# Test RMSProp implementation
from cs231n.optim import rmsprop
N, D = 4, 5
w = np.linspace(-0.4, 0.6, num=N*D).reshape(N, D)
dw = np.linspace(-0.6, 0.4, num=N*D).reshape(N, D)
cache = np.linspace(0.6, 0.9, num=N*D).reshape(N, D)
config = {'learning_rate': 1e-2, 'cache': cache}
next_w, _ = rmsprop(w, dw, config=config)
expected_next_w = np.asarray([
[-0.39223849, -0.34037513, -0.28849239, -0.23659121, -0.18467247],
[-0.132737, -0.08078555, -0.02881884, 0.02316247, 0.07515774],
[ 0.12716641, 0.17918792, 0.23122175, 0.28326742, 0.33532447],
[ 0.38739248, 0.43947102, 0.49155973, 0.54365823, 0.59576619]])
expected_cache = np.asarray([
[ 0.5976, 0.6126277, 0.6277108, 0.64284931, 0.65804321],
[ 0.67329252, 0.68859723, 0.70395734, 0.71937285, 0.73484377],
[ 0.75037008, 0.7659518, 0.78158892, 0.79728144, 0.81302936],
[ 0.82883269, 0.84469141, 0.86060554, 0.87657507, 0.8926 ]])
# You should see relative errors around e-7 or less
print('next_w error: ', rel_error(expected_next_w, next_w))
print('cache error: ', rel_error(expected_cache, config['cache']))
# +
# Test Adam implementation
from cs231n.optim import adam
N, D = 4, 5
w = np.linspace(-0.4, 0.6, num=N*D).reshape(N, D)
dw = np.linspace(-0.6, 0.4, num=N*D).reshape(N, D)
m = np.linspace(0.6, 0.9, num=N*D).reshape(N, D)
v = np.linspace(0.7, 0.5, num=N*D).reshape(N, D)
config = {'learning_rate': 1e-2, 'm': m, 'v': v, 't': 5}
next_w, _ = adam(w, dw, config=config)
expected_next_w = np.asarray([
[-0.40094747, -0.34836187, -0.29577703, -0.24319299, -0.19060977],
[-0.1380274, -0.08544591, -0.03286534, 0.01971428, 0.0722929],
[ 0.1248705, 0.17744702, 0.23002243, 0.28259667, 0.33516969],
[ 0.38774145, 0.44031188, 0.49288093, 0.54544852, 0.59801459]])
expected_v = np.asarray([
[ 0.69966, 0.68908382, 0.67851319, 0.66794809, 0.65738853,],
[ 0.64683452, 0.63628604, 0.6257431, 0.61520571, 0.60467385,],
[ 0.59414753, 0.58362676, 0.57311152, 0.56260183, 0.55209767,],
[ 0.54159906, 0.53110598, 0.52061845, 0.51013645, 0.49966, ]])
expected_m = np.asarray([
[ 0.48, 0.49947368, 0.51894737, 0.53842105, 0.55789474],
[ 0.57736842, 0.59684211, 0.61631579, 0.63578947, 0.65526316],
[ 0.67473684, 0.69421053, 0.71368421, 0.73315789, 0.75263158],
[ 0.77210526, 0.79157895, 0.81105263, 0.83052632, 0.85 ]])
# You should see relative errors around e-7 or less
print('next_w error: ', rel_error(expected_next_w, next_w))
print('v error: ', rel_error(expected_v, config['v']))
print('m error: ', rel_error(expected_m, config['m']))
# -
# Once you have debugged your RMSProp and Adam implementations, run the following to train a pair of deep networks using these new update rules:
# +
learning_rates = {'rmsprop': 1e-4, 'adam': 1e-3}
for update_rule in ['adam', 'rmsprop']:
print('running with ', update_rule)
model = FullyConnectedNet([100, 100, 100, 100, 100], weight_scale=5e-2)
solver = Solver(model, small_data,
num_epochs=5, batch_size=100,
update_rule=update_rule,
optim_config={
'learning_rate': learning_rates[update_rule]
},
verbose=True)
solvers[update_rule] = solver
solver.train()
print()
plt.subplot(3, 1, 1)
plt.title('Training loss')
plt.xlabel('Iteration')
plt.subplot(3, 1, 2)
plt.title('Training accuracy')
plt.xlabel('Epoch')
plt.subplot(3, 1, 3)
plt.title('Validation accuracy')
plt.xlabel('Epoch')
for update_rule, solver in list(solvers.items()):
plt.subplot(3, 1, 1)
plt.plot(solver.loss_history, 'o', label=update_rule)
plt.subplot(3, 1, 2)
plt.plot(solver.train_acc_history, '-o', label=update_rule)
plt.subplot(3, 1, 3)
plt.plot(solver.val_acc_history, '-o', label=update_rule)
for i in [1, 2, 3]:
plt.subplot(3, 1, i)
plt.legend(loc='upper center', ncol=4)
plt.gcf().set_size_inches(15, 15)
plt.show()
# + [markdown] tags=["pdf-inline"]
# ## Inline Question 3:
#
# AdaGrad, like Adam, is a per-parameter optimization method that uses the following update rule:
#
# ```
# cache += dw**2
# w += - learning_rate * dw / (np.sqrt(cache) + eps)
# ```
#
# John notices that when he was training a network with AdaGrad that the updates became very small, and that his network was learning slowly. Using your knowledge of the AdaGrad update rule, why do you think the updates would become very small? Would Adam have the same issue?
#
#
# ## Answer:
# AdaGrad updates become very small because every iteration cache accumulate with the squared gradients. The number of iteration increases, cache of course increases. When drivative divive to root square of cache, it will be small.
#
# Adam does not have this issue because it uses an exponential weighted average (EWA) of the squared gradients that comes from RMSProp. And it use momentum instead of the gradient (dw).
#
# -
# # Train a good model!
# Train the best fully-connected model that you can on CIFAR-10, storing your best model in the `best_model` variable. We require you to get at least 50% accuracy on the validation set using a fully-connected net.
#
# If you are careful it should be possible to get accuracies above 55%, but we don't require it for this part and won't assign extra credit for doing so. Later in the assignment we will ask you to train the best convolutional network that you can on CIFAR-10, and we would prefer that you spend your effort working on convolutional nets rather than fully-connected nets.
#
# You might find it useful to complete the `BatchNormalization.ipynb` and `Dropout.ipynb` notebooks before completing this part, since those techniques can help you train powerful models.
# +
best_model = None
best_val = -1
################################################################################
# TODO: Train the best FullyConnectedNet that you can on CIFAR-10. You might #
# find batch/layer normalization and dropout useful. Store your best model in #
# the best_model variable. #
################################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
for i in range(10):
ws = 10**np.random.uniform(-2, -3)
lr = 10**np.random.uniform(-3, -4)
reg = 10**np.random.uniform(-4, 0)
model = FullyConnectedNet([100, 100, 100], weight_scale=ws, reg=reg, dropout=1, normalization="batchnorm")
solver = Solver(model, data,
num_epochs=10, batch_size=200,
update_rule='adam',
optim_config={
'learning_rate': lr
},
verbose=False)
solver.train()
val_accuracy = solver.best_val_acc
if best_val < val_accuracy:
best_val = val_accuracy
best_model = model
# Print results
print('Try %d: lr %e ws %e reg %e val accuracy: %f' % (
(i + 1), lr, ws, reg, val_accuracy))
print('best validation accuracy achieved: %f' % best_val)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
################################################################################
# END OF YOUR CODE #
################################################################################
# -
# # Test your model!
# Run your best model on the validation and test sets. You should achieve above 50% accuracy on the validation set.
y_test_pred = np.argmax(best_model.loss(data['X_test']), axis=1)
y_val_pred = np.argmax(best_model.loss(data['X_val']), axis=1)
print('Validation set accuracy: ', (y_val_pred == data['y_val']).mean())
print('Test set accuracy: ', (y_test_pred == data['y_test']).mean())
| 32,700 |
/프로그래머스/Level_1/Level_1_x만큼_간격이_있는_n개의_숫자.ipynb | 7f55b51e952e11b5980027b87514fb96d259a0c5 | [] | no_license | Jpumpkin1223/CodingTest | https://github.com/Jpumpkin1223/CodingTest | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 1,767 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: aiffel
# language: python
# name: aiffel
# ---
# 함수 solution은 정수 x와 자연수 n을 입력 받아, x부터 시작해 x씩 증가하는 숫자를 n개 지니는 리스트를 리턴해야 합니다.
# 다음 제한 조건을 보고, 조건을 만족하는 함수, solution을 완성해주세요.
def solution(x, n):
i = 1
answer = []
while i <= n:
temp = x * i
answer.append(temp)
i += 1
return answer
x_1 = 2
n_1 = 5
answer_1 = [2,4,6,8,10]
x_2 = 4
n_2 = 3
answer_2 = [4,8,12]
x_3 = -4
n_3 = 2
answer_3 = [-4, -8]
solution(x_1, n_1)
ttp://ibmdecisionoptimization.github.io/docplex-doc/getting_started.html)
#
# Discover us [here](https://developer.ibm.com/docloud)
#
#
# Table of contents:
#
# - [Describe the business problem](#Describe-the-business-problem:--Games-Scheduling-in-the-National-Football-League)
# * [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
# * [Use decision optimization](#Use-decision-optimization)
# * [Step 1: Import the library](#Step-1:-Import-the-library)
# * [Step 2: Learn about constraint truth values](#Step-2:-Learn-about-constraint-truth-values)
# * [Step 3: Learn about equivalence constraints](#Step-3:-Learn-about-equivalence-constraints)
# * [Summary](#Summary)
# ****
# -
# Logical constraints let you use the _truth value_ of constraints inside the model. The truth value of a constraint
# is true when it is satisfied and false when not. Adding a constraint to a model ensures that it is always satisfied.
# However, with logical constraints, one can use the truth value of a constraint _inside_ the model, allowing to choose dynamically whether a constraint is to be satisfied (or not).
# + [markdown] deletable=true editable=true render=true
# ## How decision optimization can help
#
# * Prescriptive analytics (decision optimization) technology recommends actions that are based on desired outcomes. It takes into account specific scenarios, resources, and knowledge of past and current events. With this insight, your organization can make better decisions and have greater control of business outcomes.
#
# * Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
#
# * Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
# <br/>
#
# <u>With prescriptive analytics, you can:</u>
#
# * Automate the complex decisions and trade-offs to better manage your limited resources.
# * Take advantage of a future opportunity or mitigate a future risk.
# * Proactively update recommendations based on changing events.
# * Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
#
#
# + [markdown] deletable=true editable=true
# ## Use decision optimization
# + [markdown] deletable=true editable=true
# ### Step 1: Import the library
#
# Run the following code to import Decision Optimization CPLEX Modeling library. The *DOcplex* library contains the two modeling packages, Mathematical Programming and Constraint Programming, referred to earlier.
# -
import sys
try:
import docplex.mp
except:
raise Exception('Please install docplex. See https://pypi.org/project/docplex/')
# A restart of the kernel might be needed.
# + [markdown] deletable=true editable=true render=true
# ### Step 2: Learn about constraint truth values
#
# Any discrete linear constraint can be associated to a binary variable that holds the truth value of the constraint.
# But first, let's explain what a discrete constraint is
# -
# #### Discrete linear constraint
#
# A discrete linear constraint is built from discrete coefficients and discrete variables, taht is variables with type `integer` or `binary`. For example, assuming x and y are integer variables:
#
# - `2x+3y == 1` is discrete
# - `x+y = 3.14` is not (because of 3.14)
# - `1.1 x + 2.2 y <= 3` is not because of the non-integer coefficients 1.1 and 2.2
# #### The truth value of an added constraint is always 1
#
# The truth value of a constraint is accessed by the `status_var` property. This varianle is aplain Docplex decision variable that can be used anywhere a variable can. However, the value of the truth value variable and the constraint are linked, both ways:
#
# - a constraint is satisfied if and only if its truth value variable equals 1
# - a constraint is _not_ satisfied if and only if its truth value variable equals 0.
#
# In this toy model,we show that the truth value of a constraint which has been added to a model is always equal to 1.
# +
from docplex.mp.model import Model
m1 = Model()
x = m1.integer_var(name='ix')
y = m1.integer_var(name='iy')
ct = m1.add(x + y <= 3)
ct_truth = ct.status_var
m1.maximize(x+y)
assert m1.solve()
print('the truth value of [{0!s}] is {1}'.format(ct, ct_truth.solution_value))
# -
# #### The truth value of a constraint not added to a model is undefined
#
# A constraint that is not added to a model, has no effect. Its truth value is undefined: it can be either 1 or 0.
#
# In the following example, both `x` and `y` are set to their upper bound, so that the constraint is not satisfied; hence the truth value is 0.
m2 = Model()
x = m2.integer_var(name='ix', ub=4)
y = m2.integer_var(name='iy', ub=4)
ct = (x + y <= 3)
ct_truth = ct.status_var # not m2.add() here!
m2.maximize(x+y)
assert m2.solve()
m2.print_solution()
print('the truth value of [{0!s}] is {1}'.format(ct, ct_truth.solution_value))
# #### Using constraint truth values in modeling
#
# A constraint's truth value is actually a plain DOcplex decision variable, and as such, can be used with comparison operators and arithmetic operators.
# Let's experiment again with a toy model: in this model,
# we state that the truth value of `y == 4` is less than the truth value of `x ==3`.
# As we maximize y, y has value 4 in the optimal solution (it is the upper bound), and consequently the constraint `ct_y4` is satisfied. From the inequality between truth values,
# it follows that the truth value of `ct_x2` equals 1 and x is equal to 2.
#
# Using the constraints in the inequality has silently converted each constraint into its truth value.
m3 = Model()
x = m3.integer_var(name='ix', ub=4)
y = m3.integer_var(name='iy', ub=4)
ct_x2 = (x == 2)
ct_y4 = (y == 4)
m3.add( ct_y4 <= ct_x2 )
m3.maximize(y)
assert m3.solve()
m3.print_solution()
# Constraint truth values can be used with arithmetic operators, just as variables can. In th enext model, we express a more complex constraint:
# - either x is equal to 3, _or_ both y and z are equal to 5
#
# Let's see how we can express this easilty with truth values:
m31 = Model(name='m31')
x = m31.integer_var(name='ix', ub=4)
y = m31.integer_var(name='iy', ub=10)
z = m31.integer_var(name='iz', ub=10)
ct_x2 = (x == 3)
ct_y5 = (y == 5)
ct_z5 = (z == 5)
#either ct-x2 is true or -both- ct_y5 and ct_z5 mus be true
m31.add( 2 * ct_x2 + (ct_y5 + ct_z5) == 2)
# force x to be less than 2: it cannot be equal to 3!
m31.add(x <= 2)
# maximize sum of x,y,z
m31.maximize(x+y+z)
assert m31.solve()
# the expected solution is: x=2, y=5, z=5
assert m31.objective_value == 12
m31.print_solution()
# As we have seen, constraints can be used in expressions. This includes the `Model.sum()` and `Model.dot()` aggregation methods.
#
# In the next model, we define ten variables, one of which must be equal to 3 (we dpn't care which one, for now). As we maximize the sum of all `xs` variables, all will end up equal to their upper bound, except for one.
m4 = Model()
xs = m4.integer_var_list(10, ub=100)
cts = [xi==3 for xi in xs]
m4.add( m4.sum(cts) == 1)
m4.maximize(m4.sum(xs))
assert m4.solve()
m4.print_solution()
# As we can see, all variables but one are set to their upper bound of 100. We cannot predict which variable will be set to 3.
# However, let's imagine that we prefer variable with a lower index to be set to 3, how can we express this preference?
#
# The answer is to use an additional expression to the objective, using a scalar product of constraint truth value
preference = m4.dot(cts, (k+1 for k in range(len(xs))))
# we prefer lower indices for satisfying the x==3 constraint
# so the final objective is a maximize of sum of xs -minus- the preference
m4.maximize(m4.sum(xs) - preference)
assert m4.solve()
m4.print_solution()
# As expected, the `x` variable set to 3 now is the first one.
# #### Using truth values to state 'not equals' constraints.
#
# Truth values can be used to express elegantly 'not equal' constraints, by forcing the truth value of an equality constraint to 0.
#
# In the next model, we illustrate how an equality constraint can be negated by forcing its truth value to zero. This negation forbids y to be equal to 4, as it would be without this negation.
# Finally, the objective is 7 instead of 8.
# +
m5 = Model()
x = m5.integer_var(name='ix', ub=4)
y = m5.integer_var(name='iy', ub=4)
# this is the equality constraint we want to negate
ct_y4 = (y == 4)
# forcing truth value to zero means the constraint is not satisfied.
negation = m5.add( ct_y4 == 0)
# maximize x+y should yield both variables to 4, but y cannot be equal to 4
# as such we expect y to be equal to 3
m5.maximize(x + y)
assert m5.solve()
m5.print_solution()
# expecting 7 as objective, not 8
assert m5.objective_value == 7
# now remove the negation
m5.remove_constraint(negation)
# and solve again
assert m5.solve()
# the objective is 8 as expected: both x and y are equal to 4
assert m5.objective_value == 8
# -
# #### Summary
#
# We have seen that linear constraints have an associated binary variable, its _truth value_, whose value is linked to whether or not the constraint is satisfied.
# Moreover, this llink enables to express 'not equals' constraints.# now remove netation
# ### Step 3: Learn about equivalence constraints
#
# As we have seen, using a constraint in expressions automtically generates a truth value variable, whose value is linked to the status of the constraint.
#
# However, in some cases, it can be useful to relate the status of a constraint to an _existing_ binary variable. This is the purpose of equivalence constraints.
#
# An equiavelnec constraints relates an existing binary variable to the status of a discrete linear constraints. The syntax is:
#
# `Model.add_equivalence(bvar, linear_ct, active_value, name)`
#
# - `bvar` is the existing binary variable
# - `linear-ct` is a discrete linear constraint
# - `active_value` can take values 1 or 0 (the default is 1)
# - `name` is an optional string to name the equivalence.
# +
m6 = Model(name='m6')
size = 7
il = m6.integer_var_list(size, name='i', ub=10)
jl = m6.integer_var_list(size, name='j', ub=10)
bl = m6.binary_var_list(size, name='b')
for k in range(size):
# for each i, relate bl_k to il_k==5 *and* jl_k == 7
m6.add_equivalence(bl[k], il[k] == 5)
m6.add_equivalence(bl[k], jl[k] == 7)
# now maximize sum of bs
m6.maximize(m6.sum(bl))
assert m6.solve()
m6.print_solution()
# + [markdown] deletable=true editable=true
# ## Summary
#
#
# You learned how to set up and use the IBM Decision Optimization CPLEX Modeling for Python to formulate a Mathematical Programming model with logical constraints.
# + [markdown] deletable=true editable=true render=true
# #### References
# * [Decision Optimization CPLEX Modeling for Python documentation](http://ibmdecisionoptimization.github.io/docplex-doc/)
# * [Decision Optimization on Cloud](https://developer.ibm.com/docloud/)
# * Need help with DOcplex or to report a bug? Please go [here](https://developer.ibm.com/answers/smartspace/docloud)
# * Contact us at dofeedback@wwpdl.vnet.ibm.com"
#
# + [markdown] deletable=true editable=true
# Copyright © 2017-2018 IBM. Sample Materials.
| 12,188 |
/university ranking/shanghairanking.ipynb | f646460e0947f60adfbf0c79bdf2055f044f9f82 | [] | no_license | Chrislee3x7/CollegeInfo | https://github.com/Chrislee3x7/CollegeInfo | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 2,196 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="hE344njY9ozW" executionInfo={"status": "ok", "timestamp": 1632004198675, "user_tz": 240, "elapsed": 1749, "user": {"displayName": "Aa A", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "03231498138364491179"}} outputId="aa5be49d-5154-4518-d680-a21a07c9e766"
import csv
import requests
from bs4 import BeautifulSoup
response = requests.get('https://www.shanghairanking.com/rankings/arwu/2021')
soup=BeautifulSoup(response.text,"html.parser")
x=soup.find_all("td", class_="align-left")
for ele in x:
string=ele.text
string=str(string).strip()
string=string[:string.index(" ")]
print(string)
| 935 |
/01_mnist.ipynb | 114444b73c5984c8267dae40b6960d97e06e3669 | [] | no_license | PritikaRamu/Deep-Learning-Workshop | https://github.com/PritikaRamu/Deep-Learning-Workshop | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 62,159 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="./images/DLI_Header.png" style="width: 400px;">
# # Image Classification with the MNIST Dataset
# In this section we will do the "Hello World" of deep learning: training a deep learning model to correctly classify hand-written digits.
# ## Objectives
# * Understand how deep learning can solve problems traditional programming methods cannot
# * Learn about the [MNSIT handwritten digits dataset](http://yann.lecun.com/exdb/mnist/)
# * Use the [Keras API](https://keras.io/) to load the MNIST dataset and prepare it for training
# * Create a simple neural network to perform image classification
# * Train the neural network using the prepped MNIST dataset
# * Observe the performance of the trained neural network
# ## The Problem: Image Classification
# In traditional programming, the programmer is able to articulate rules and conditions in their code that their program can then use to act in the correct way. This approach continues to work exceptionally well for a huge variety of problems.
#
# Image classification, which asks a program to correctly classify an image it has never seen before into its correct class, is near impossible to solve with traditional programming techniques. How could a programmer possibly define the rules and conditions to correctly classify a huge variety of images, especially taking into account images that they have never seen?
# ## The Solution: Deep Learning
# Deep learning excels at pattern recognition by trial and error. By training a deep neural network with sufficient data, and providing the network with feedback on its performance via training, the network can identify, though a huge amount of iteration, its own set of conditions by which it can act in the correct way.
# ## The MNIST Dataset
# In the history of deep learning, the accurate image classification of the [MNSIT dataset](http://yann.lecun.com/exdb/mnist/), a collection of 70,000 grayscale images of handwritten digits from 0 to 9, was a major development. While today the problem is considered trivial, doing image classification with MNIST has become a kind of "Hello World" for deep learning.
# Here are 40 of the images included in the MNIST dataset:
# <img src="./images/mnist1.png" style="width: 600px;">
# ## Training and Validation Data and Labels
# When working with images for deep learning, we need both the images themselves, usually denoted as `X`, and also, correct [labels](https://developers.google.com/machine-learning/glossary#label) for these images, usually denoted as `Y`. Furthermore, we need `X` and `Y` values both for *training* the model, and then, a separate set of `X` and `Y` values for *validating* the performance of the model after it has been trained. Therefore, we need 4 segments of data for the MNIST dataset:
#
# 1. `x_train`: Images used for training the neural network
# 2. `y_train`: Correct labels for the `x_train` images, used to evaluate the model's predictions during training
# 3. `x_valid`: Images set aside for validating the performance of the model after it has been trained
# 4. `y_valid`: Correct labels for the `x_valid` images, used to evaluate the model's predictions after it has been trained
#
# The process of preparing data for analysis is called [Data Engineering](https://medium.com/@rchang/a-beginners-guide-to-data-engineering-part-i-4227c5c457d7). To learn more about the differences between training data and validation data (as well as test data), check out [this article](https://machinelearningmastery.com/difference-test-validation-datasets/) by Jason Brownlee.
# ## Loading the Data Into Memory (with Keras)
# There are many [deep learning frameworks](https://developer.nvidia.com/deep-learning-frameworks), each with their own merits. In this workshop we will be working with [Tensorflow 2](https://www.tensorflow.org/tutorials/quickstart/beginner), and specifically with the [Keras API](https://keras.io/). Keras has many useful built in functions designed for the computer vision tasks. It is also a legitimate choice for deep learning in a professional setting due to its [readability](https://blog.pragmaticengineer.com/readable-code/) and efficiency, though it is not alone in this regard, and it is worth investigating a variety of frameworks when beginning a deep learning project.
#
# One of the many helpful features that Keras provides are modules containing many helper methods for [many common datasets](https://www.tensorflow.org/api_docs/python/tf/keras/datasets), including MNIST.
#
# We will begin by loading the Keras dataset module for MNIST:
from tensorflow.keras.datasets import mnist
# With the `mnist` module, we can easily load the MNIST data, already partitioned into images and labels for both training and validation:
# the data, split between train and validation sets
(x_train, y_train), (x_valid, y_valid) = mnist.load_data()
# ## Exploring the MNIST Data
# We stated above that the MNIST dataset contained 70,000 grayscale images of handwritten digits. By executing the following cells, we can see that Keras has partitioned 60,000 of these images for training, and 10,000 for validation (after training), and also, that each image itself is a 2D array with the dimensions 28x28:
x_train.shape
x_valid.shape
# Furthermore, we can see that these 28x28 images are represented as a collection of unsigned 8-bit integer values between 0 and 255, the values corresponding with a pixel's grayscale value where `0` is black, `255` is white, and all other values are in between:
x_train.dtype
x_train.min()
x_train.max()
x_train[0]
# Using [Matplotlib](https://matplotlib.org/), we can render one of these grayscale images in our dataset:
# +
import matplotlib.pyplot as plt
image = x_train[0]
plt.imshow(image, cmap='gray')
# -
# In this way we can now see that this is a 28x28 pixel image of a 5. Or is it a 3? The answer is in the `y_train` data, which contains correct labels for the data. Let's take a look:
y_train[0]
# ## Preparing the Data for Training
# In deep learning, it is common that data needs to be transformed to be in the ideal state for training. For this particular image classification problem, there are 3 tasks we should perform with the data in preparation for training:
# 1. Flatten the image data, to simplify the image input into the model
# 2. Normalize the image data, to make the image input values easier to work with for the model
# 3. Categorize the labels, to make the label values easier to work with for the model
# ### Flattening the Image Data
# Though it's possible for a deep learning model to accept a 2-dimensional image (in our case 28x28 pixels), we're going to simplify things to start and [reshape](https://www.tensorflow.org/api_docs/python/tf/reshape) each image into a single array of 784 continuous pixels (note: 28x28 = 784). This is also called flattening the image.
#
# Here we accomplish this using the helper method `reshape`:
x_train = x_train.reshape(60000, 784)
x_valid = x_valid.reshape(10000, 784)
# We can confirm that the image data has been reshaped and is now a collection of 1D arrays containing 784 pixel values each:
x_train.shape
x_train[0]
# ### Normalizing the Image Data
# Deep learning models are better at dealing with floating point numbers between 0 and 1 (more on this topic later). Converting integer values to floating point values between 0 and 1 is called [normalization](https://developers.google.com/machine-learning/glossary#normalization), and a simple approach we will take here to normalize the data will be to divide all the pixel values (which if you recall are between 0 and 255) by 255:
x_train = x_train / 255
x_valid = x_valid / 255
# We can now see that the values are all floating point values between `0.0` and `1.0`:
x_train.dtype
x_train.min()
x_train.max()
# ### Categorical Encoding
# Consider for a moment, if we were to ask, what is 7 - 2? Stating that the answer was 4 is closer than stating that the answer was 9. However, for this image classification problem, we don't want the neural network to learn this kind of reasoning: we just want it to select the correct category, and understand that if we have an image of the number 5, that guessing 4 is just as bad as guessing 9.
#
# As it stands, the labels for the images are integers between 0 and 9. Because these values represent a numerical range, the model might try to draw some conclusions about its performance based on how close to the correct numerical category it guesses.
#
# Therefore, we will do something to our data called categorical encoding. This kind of transformation modifies the data so that each value is a collection of all possible categories, with the actual category that this particular value is set as true.
#
# As a simple example, consider if we had 3 categories: red, blue, and green. For a given color, 2 of these categories would be false, and the other would be true:
# |Actual Color| Is Red? | Is Blue? | Is Green?|
# |------------|---------|----------|----------|
# |Red|True|False|False|
# |Green|False|False|True|
# |Blue|False|True|False|
# |Green|False|False|True|
# Rather than use "True" or "False", we could represent the same using binary, either 0 or 1:
# |Actual Color| Is Red? | Is Blue? | Is Green?|
# |------------|---------|----------|----------|
# |Red|1|0|0|
# |Green|0|0|1|
# |Blue|0|1|0|
# |Green|0|0|1|
# This is what categorical encoding is, transforming values which are intended to be understood as categorical labels into a representation that makes their categorical nature explicit to the model. Thus, if we were using these values for training, we would convert...
# ```python
# values = ['red, green, blue, green']
# ```
# ... which a neural network would have a very difficult time making sense of, instead to:
# ```python
# values = [
# [1, 0, 0],
# [0, 0, 1],
# [0, 1, 0],
# [0, 0, 1]
# ]
# ```
# ### Categorically Encoding the Labels
# Keras provides a utility to [categorically encode values](https://www.tensorflow.org/api_docs/python/tf/keras/utils/to_categorical), and here we use it to perform categorical encoding for both the training and validation labels:
# +
import tensorflow.keras as keras
num_categories = 10
y_train = keras.utils.to_categorical(y_train, num_categories)
y_valid = keras.utils.to_categorical(y_valid, num_categories)
# -
# Here are the first 10 values of the training labels, which you can see have now been categorically encoded:
y_train[0:9]
# ## Creating the Model
# With the data prepared for training, it is now time to create the model that we will train with the data. This first basic model will be made up of several *layers* and will be comprised of 3 main parts:
#
# 1. An input layer, which will receive data in some expected format
# 2. Several [hidden layers](https://developers.google.com/machine-learning/glossary#hidden-layer), each comprised of many *neurons*. Each [neuron](https://developers.google.com/machine-learning/glossary#neuron) will have the ability to affect the network's guess with its *weights*, which are values that will be updated over many iterations as the network gets feedback on its performance and learns
# 3. An output layer, which will depict the network's guess for a given image
# ### Instantiating the Model
# To begin, we will use Keras's [Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) model class to instantiate an instance of a model that will have a series of layers that data will pass through in sequence:
# +
from tensorflow.keras.models import Sequential
model = Sequential()
# -
# ### Creating the Input Layer
# Next, we will add the input layer. This layer will be *densely connected*, meaning that each neuron in it, and its weights, will affect every neuron in the next layer. To do this with Keras, we use Keras's [Dense](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense) layer class.
from tensorflow.keras.layers import Dense
# The `units` argument specifies the number of neurons in the layer. We are going to use `512` which we have chosen from experimentation. Choosing the correct number of neurons is what puts the "science" in "data science" as it is a matter of capturing the statistical complexity of the dataset. Try playing around with this value later to see how it affects training and to start developing a sense for what this number means.
#
# We will learn more about activation functions later, but for now, we will use the `relu` activation function, which in short, will help our network to learn how to make more sophisticated guesses about data than if it were required to make guesses based on some strictly linear function.
#
# The `input_shape` value specifies the shape of the incoming data which in our situation is a 1D array of 784 values:
model.add(Dense(units=512, activation='relu', input_shape=(784,)))
# ### Creating the Hidden Layer
# Now we will add an additional densely connected layer. Again, much more will be said about these later, but for now know that these layers give the network more parameters to contribute towards its guesses, and therefore, more subtle opportunities for accurate learning:
model.add(Dense(units = 512, activation='relu'))
# ### Creating the Output Layer
# Finally, we will add an output layer. This layer uses the activation function `softmax` which will result in each of the layer's values being a probability between 0 and 1 and will result in all the outputs of the layer adding to 1. In this case, since the network is to make a guess about a single image belonging to 1 of 10 possible categories, there will be 10 outputs. Each output gives the model's guess (a probability) that the image belongs to that specific class:
model.add(Dense(units = 10, activation='softmax'))
# ### Summarizing the Model
# Keras provides the model instance method [summary](https://www.tensorflow.org/api_docs/python/tf/summary) which will print a readable summary of a model:
model.summary()
# Note the number of trainable parameters. Each of these can be adjusted during training and will contribute towards the trained model's guesses.
# ### Compiling the Model
# Again, more details are to follow, but the final step we need to do before we can actually train our model with data is to [compile](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential#compile) it. Here we specify a [loss function](https://developers.google.com/machine-learning/glossary#loss) which will be used for the model to understand how well it is performing during training. We also specify that we would like to track `accuracy` while the model trains:
model.compile(loss='categorical_crossentropy', metrics=['accuracy'])
# ## Training the Model
# Now that we have prepared training and validation data, and a model, it's time to train our model with our training data, and verify it with its validation data.
#
# "Training a model with data" is often also called "fitting a model to data." Put this latter way, it highlights that the shape of the model changes over time to more accurately understand the data that it is being given.
#
# When fitting (training) a model with Keras, we use the model's [fit](https://www.tensorflow.org/api_docs/python/tf/keras/Model#fit) method. It expects the following arguments:
#
# * The training data
# * The labels for the training data
# * The number of times it should train on the entire training dataset (called an *epoch*)
# * The validation or test data, and its labels
#
# Run the cell below to train the model. We will discuss its output after the training completes:
history = model.fit(
x_train, y_train, epochs=5, verbose=1, validation_data=(x_valid, y_valid)
)
# ### Observing Accuracy
# For each of the 5 epochs, notice the `accuracy` and `val_accuracy` scores. `accuracy` states how well the model did for the epoch on all the training data. `val_accuracy` states how well the model did on the validation data, which if you recall, was not used at all for training the model.
# The model did quite well! The accuracy quickly reached close to 100%, as did the validation accuracy. We now have a model that can be used to accurately detect and classify hand-written images.
#
# The next step would be to use this model to classify new not-yet-seen handwritten images. This is called [inference](https://blogs.nvidia.com/blog/2016/08/22/difference-deep-learning-training-inference-ai/). We'll explore the process of inference in a later exercise.
# ## Summary
# It's worth taking a moment to appreciate what we've done here. Historically, the expert systems that were built to do this kind of task were extremely complicated, and people spent their careers building them (check out the references on the [official MNIST page](http://yann.lecun.com/exdb/mnist/) and the years milestones were reached).
#
# MNIST is not only useful for its historical influence on Computer Vision, but it's also a great [benchmark](http://www.cs.toronto.edu/~serailhydra/publications/tbd-iiswc18.pdf) and debugging tool. Having trouble getting a fancy new machine learning architecture working? Check it against MNIST. If it can't learn on this dataset, chances are it won't learn on more complicated images and datasets.
# ## Clear the Memory
# Before moving on, please execute the following cell to clear up the GPU memory. This is required to move on to the next notebook.
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# ## Next
# In this section you learned how to build and train a simple neural network for image classification. In the next section, you will be asked to build your own neural network and perform data preparation to solve a different image classification problem.
# ## ☆ Bonus Exercise ☆
#
# Have time to spare? In the next section, we will talk about how we arrived at some of the numbers above, but we can try imagining what it was like to be a researcher developing the techniques commonly used today.
#
# Ultimately, each neuron is trying to fit a line to some data. Below, we have some datapoints and a randomly drawn line using the equation [y = mx + b](https://www.mathsisfun.com/equation_of_line.html).
#
# Try changing the `m` and the `b` in order to find the lowest possible loss. How did you find the best line? Can you make a program to follow your strategy?
# +
import numpy as np
from numpy.polynomial.polynomial import polyfit
import matplotlib.pyplot as plt
m = 6 # -2 to start, change me please
b = 10 # 40 to start, change me please
# Sample data
x = np.array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
y = np.array([10, 20, 25, 30, 40, 45, 40, 50, 60, 55])
y_hat = x * m + b
plt.plot(x, y, '.')
plt.plot(x, y_hat, '-')
plt.show()
print("Loss:", np.sum((y - y_hat)**2)/len(x))
# -
# Have an idea? Excellent! Please shut down the kernel before moving on.
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
| 19,266 |
/Preprocessing_LR_Coursera.ipynb | 7d2cd2c9d365625d8b12b6e501d3017b4c7eccd6 | [] | no_license | darrrya21/ml_projects | https://github.com/darrrya21/ml_projects | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 237,875 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# **Корректность проверена на Python 3.6:**
# + pandas 0.23.4
# + numpy 1.15.4
# + matplotlib 3.0.2
# + sklearn 0.20.2
# -
import warnings
warnings.filterwarnings('ignore')
# ## Предобработка данных и логистическая регрессия для задачи бинарной классификации
# ## Programming assignment
# В задании вам будет предложено ознакомиться с основными техниками предобработки данных, а так же применить их для обучения модели логистической регрессии. Ответ потребуется загрузить в соответствующую форму в виде 6 текстовых файлов.
# +
import pandas as pd
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
matplotlib.style.use('ggplot')
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
# ## Описание датасета
# Задача: по 38 признакам, связанных с заявкой на грант (область исследований учёных, информация по их академическому бэкграунду, размер гранта, область, в которой он выдаётся) предсказать, будет ли заявка принята. Датасет включает в себя информацию по 6000 заявкам на гранты, которые были поданы в университете Мельбурна в период с 2004 по 2008 год.
#
# Полную версию данных с большим количеством признаков можно найти на https://www.kaggle.com/c/unimelb.
data = pd.read_csv('data.csv')
data.shape
# Выделим из датасета целевую переменную Grant.Status и обозначим её за y
# Теперь X обозначает обучающую выборку, y - ответы на ней
X = data.drop('Grant.Status', 1)
y = data['Grant.Status']
# ## Теория по логистической регрессии
# После осознания того, какую именно задачу требуется решить на этих данных, следующим шагом при реальном анализе был бы подбор подходящего метода. В данном задании выбор метода было произведён за вас, это логистическая регрессия. Кратко напомним вам используемую модель.
#
# Логистическая регрессия предсказывает вероятности принадлежности объекта к каждому классу. Сумма ответов логистической регрессии на одном объекте для всех классов равна единице.
#
# $$ \sum_{k=1}^K \pi_{ik} = 1, \quad \pi_k \equiv P\,(y_i = k \mid x_i, \theta), $$
#
# где:
# - $\pi_{ik}$ - вероятность принадлежности объекта $x_i$ из выборки $X$ к классу $k$
# - $\theta$ - внутренние параметры алгоритма, которые настраиваются в процессе обучения, в случае логистической регрессии - $w, b$
#
# Из этого свойства модели в случае бинарной классификации требуется вычислить лишь вероятность принадлежности объекта к одному из классов (вторая вычисляется из условия нормировки вероятностей). Эта вероятность вычисляется, используя логистическую функцию:
#
# $$ P\,(y_i = 1 \mid x_i, \theta) = \frac{1}{1 + \exp(-w^T x_i-b)} $$
#
# Параметры $w$ и $b$ находятся, как решения следующей задачи оптимизации (указаны функционалы с L1 и L2 регуляризацией, с которыми вы познакомились в предыдущих заданиях):
#
# L2-regularization:
#
# $$ Q(X, y, \theta) = \frac{1}{2} w^T w + C \sum_{i=1}^l \log ( 1 + \exp(-y_i (w^T x_i + b ) ) ) \longrightarrow \min\limits_{w,b} $$
#
# L1-regularization:
#
# $$ Q(X, y, \theta) = \sum_{d=1}^D |w_d| + C \sum_{i=1}^l \log ( 1 + \exp(-y_i (w^T x_i + b ) ) ) \longrightarrow \min\limits_{w,b} $$
#
# $C$ - это стандартный гиперпараметр модели, который регулирует то, насколько сильно мы позволяем модели подстраиваться под данные.
# ## Предобработка данных
# Из свойств данной модели следует, что:
# - все $X$ должны быть числовыми данными (в случае наличия среди них категорий, их требуется некоторым способом преобразовать в вещественные числа)
# - среди $X$ не должно быть пропущенных значений (т.е. все пропущенные значения перед применением модели следует каким-то образом заполнить)
#
# Поэтому базовым этапом в предобработке любого датасета для логистической регрессии будет кодирование категориальных признаков, а так же удаление или интерпретация пропущенных значений (при наличии того или другого).
data.head()
# Видно, что в датасете есть как числовые, так и категориальные признаки. Получим списки их названий:
numeric_cols = ['RFCD.Percentage.1', 'RFCD.Percentage.2', 'RFCD.Percentage.3',
'RFCD.Percentage.4', 'RFCD.Percentage.5',
'SEO.Percentage.1', 'SEO.Percentage.2', 'SEO.Percentage.3',
'SEO.Percentage.4', 'SEO.Percentage.5',
'Year.of.Birth.1', 'Number.of.Successful.Grant.1', 'Number.of.Unsuccessful.Grant.1']
categorical_cols = list(set(X.columns.values.tolist()) - set(numeric_cols))
# Также в нём присутствуют пропущенные значения. Очевидны решением будет исключение всех данных, у которых пропущено хотя бы одно значение. Сделаем это:
data.dropna().shape
# Видно, что тогда мы выбросим почти все данные, и такой метод решения в данном случае не сработает.
#
# Пропущенные значения можно так же интерпретировать, для этого существует несколько способов, они различаются для категориальных и вещественных признаков.
#
# Для вещественных признаков:
# - заменить на 0 (данный признак давать вклад в предсказание для данного объекта не будет)
# - заменить на среднее (каждый пропущенный признак будет давать такой же вклад, как и среднее значение признака на датасете)
#
# Для категориальных:
# - интерпретировать пропущенное значение, как ещё одну категорию (данный способ является самым естественным, так как в случае категорий у нас есть уникальная возможность не потерять информацию о наличии пропущенных значений; обратите внимание, что в случае вещественных признаков данная информация неизбежно теряется)
# ## Задание 0. Обработка пропущенных значений.
# 1. Заполните пропущенные вещественные значения в X нулями и средними по столбцам, назовите полученные датафреймы X_real_zeros и X_real_mean соответственно. Для подсчёта средних используйте описанную ниже функцию calculate_means, которой требуется передать на вход вешественные признаки из исходного датафрейма. **Для подсчета среднего можно использовать функцию pandas.mean()**
# 2. Все категориальные признаки в X преобразуйте в строки, пропущенные значения требуется также преобразовать в какие-либо строки, которые не являются категориями (например, 'NA'), полученный датафрейм назовите X_cat.
#
# Для объединения выборок здесь и далее в задании рекомендуется использовать функции
#
# np.hstack(...)
# np.vstack(...)
def calculate_means(numeric_data):
means = np.zeros(numeric_data.shape[1])
for j in range(numeric_data.shape[1]):
to_sum = numeric_data.iloc[:,j]
indices = ~numeric_data.iloc[:,j].isnull().to_numpy().nonzero()[0]
correction = np.amax(to_sum[indices])
to_sum /= correction
for i in indices:
means[j] += to_sum[i]
means[j] /= indices.size
means[j] *= correction
return pd.Series(means, numeric_data.columns)
X_real_zeros = X[numeric_cols].fillna(value=0)
X_real_mean = X[numeric_cols].fillna(X[numeric_cols].mean())
X_cat = X[categorical_cols].fillna('NA').astype(str)
# ## Преобразование категориальных признаков.
# В предыдущей ячейке мы разделили наш датасет ещё на две части: в одной присутствуют только вещественные признаки, в другой только категориальные. Это понадобится нам для раздельной последующей обработке этих данных, а так же для сравнения качества работы тех или иных методов.
#
# Для использования модели регрессии требуется преобразовать категориальные признаки в вещественные. Рассмотрим основной способ преоборазования категориальных признаков в вещественные: one-hot encoding. Его идея заключается в том, что мы преобразуем категориальный признак при помощи бинарного кода: каждой категории ставим в соответствие набор из нулей и единиц.
#
# Посмотрим, как данный метод работает на простом наборе данных.
# +
from sklearn.linear_model import LogisticRegression as LR
from sklearn.feature_extraction import DictVectorizer as DV
categorial_data = pd.DataFrame({'sex': ['male', 'female', 'male', 'female'],
'nationality': ['American', 'European', 'Asian', 'European']})
print('Исходные данные:\n')
print(categorial_data)
encoder = DV(sparse = False)
encoded_data = encoder.fit_transform(categorial_data.T.to_dict().values())
print('\nЗакодированные данные:\n')
print(encoded_data)
# -
# Как видно, в первые три колонки оказалась закодированна информация о стране, а во вторые две - о поле. При этом для совпадающих элементов выборки строки будут полностью совпадать. Также из примера видно, что кодирование признаков сильно увеличивает их количество, но полностью сохраняет информацию, в том числе о наличии пропущенных значений (их наличие просто становится одним из бинарных признаков в преобразованных данных).
#
# Теперь применим one-hot encoding к категориальным признакам из исходного датасета. Обратите внимание на общий для всех методов преобработки данных интерфейс. Функция
#
# encoder.fit_transform(X)
#
# позволяет вычислить необходимые параметры преобразования, впоследствии к новым данным можно уже применять функцию
#
# encoder.transform(X)
#
# Очень важно применять одинаковое преобразование как к обучающим, так и тестовым данным, потому что в противном случае вы получите непредсказуемые, и, скорее всего, плохие результаты. В частности, если вы отдельно закодируете обучающую и тестовую выборку, то получите вообще говоря разные коды для одних и тех же признаков, и ваше решение работать не будет.
#
# Также параметры многих преобразований (например, рассмотренное ниже масштабирование) нельзя вычислять одновременно на данных из обучения и теста, потому что иначе подсчитанные на тесте метрики качества будут давать смещённые оценки на качество работы алгоритма. Кодирование категориальных признаков не считает на обучающей выборке никаких параметров, поэтому его можно применять сразу к всему датасету.
encoder = DV(sparse = False)
X_cat_oh = encoder.fit_transform(X_cat.T.to_dict().values())
X_cat_oh.shape
# Для построения метрики качества по результату обучения требуется разделить исходный датасет на обучающую и тестовую выборки.
#
# Обращаем внимание на заданный параметр для генератора случайных чисел: random_state. Так как результаты на обучении и тесте будут зависеть от того, как именно вы разделите объекты, то предлагается использовать заранее определённое значение для получение результатов, согласованных с ответами в системе проверки заданий.
# +
from sklearn.model_selection import train_test_split
(X_train_real_zeros,
X_test_real_zeros,
y_train, y_test) = train_test_split(X_real_zeros, y,
test_size=0.3,
random_state=0)
(X_train_real_mean,
X_test_real_mean) = train_test_split(X_real_mean,
test_size=0.3,
random_state=0)
(X_train_cat_oh,
X_test_cat_oh) = train_test_split(X_cat_oh,
test_size=0.3,
random_state=0)
# -
# ## Описание классов
# Итак, мы получили первые наборы данных, для которых выполнены оба ограничения логистической регрессии на входные данные. Обучим на них регрессию, используя имеющийся в библиотеке sklearn функционал по подбору гиперпараметров модели
#
# optimizer = GridSearchCV(estimator, param_grid)
#
# где:
# - estimator - обучающий алгоритм, для которого будет производиться подбор параметров
# - param_grid - словарь параметров, ключами которого являются строки-названия, которые передаются алгоритму estimator, а значения - набор параметров для перебора
#
# Данный класс выполняет кросс-валидацию обучающей выборки для каждого набора параметров и находит те, на которых алгоритм работает лучше всего. Этот метод позволяет настраивать гиперпараметры по обучающей выборке, избегая переобучения. Некоторые опциональные параметры вызова данного класса, которые нам понадобятся:
# - scoring - функционал качества, максимум которого ищется кросс валидацией, по умолчанию используется функция score() класса esimator
# - n_jobs - позволяет ускорить кросс-валидацию, выполняя её параллельно, число определяет количество одновременно запущенных задач
# - cv - количество фолдов, на которые разбивается выборка при кросс-валидации
#
# После инициализации класса GridSearchCV, процесс подбора параметров запускается следующим методом:
#
# optimizer.fit(X, y)
#
# На выходе для получения предсказаний можно пользоваться функцией
#
# optimizer.predict(X)
#
# для меток или
#
# optimizer.predict_proba(X)
#
# для вероятностей (в случае использования логистической регрессии).
#
# Также можно напрямую получить оптимальный класс estimator и оптимальные параметры, так как они является атрибутами класса GridSearchCV:
# - best\_estimator\_ - лучший алгоритм
# - best\_params\_ - лучший набор параметров
#
# Класс логистической регрессии выглядит следующим образом:
#
# estimator = LogisticRegression(penalty)
#
# где penalty принимает либо значение 'l2', либо 'l1'. По умолчанию устанавливается значение 'l2', и везде в задании, если об этом не оговорено особо, предполагается использование логистической регрессии с L2-регуляризацией.
# ## Задание 1. Сравнение способов заполнения вещественных пропущенных значений.
# 1. Составьте две обучающие выборки из вещественных и категориальных признаков: в одной вещественные признаки, где пропущенные значения заполнены нулями, в другой - средними. Рекомендуется записывать в выборки сначала вещественные, а потом категориальные признаки.
# 2. Обучите на них логистическую регрессию, подбирая параметры из заданной сетки param_grid по методу кросс-валидации с числом фолдов cv=3. В качестве оптимизируемой функции используйте заданную по умолчанию.
# 3. Постройте два графика оценок точности +- их стандратного отклонения в зависимости от гиперпараметра и убедитесь, что вы действительно нашли её максимум. Также обратите внимание на большую дисперсию получаемых оценок (уменьшить её можно увеличением числа фолдов cv).
# 4. Получите две метрики качества AUC ROC на тестовой выборке и сравните их между собой. Какой способ заполнения пропущенных вещественных значений работает лучше? В дальнейшем для выполнения задания в качестве вещественных признаков используйте ту выборку, которая даёт лучшее качество на тесте.
# 5. Передайте два значения AUC ROC (сначала для выборки, заполненной средними, потом для выборки, заполненной нулями) в функцию write_answer_1 и запустите её. Полученный файл является ответом на 1 задание.
#
# Информация для интересующихся: вообще говоря, не вполне логично оптимизировать на кросс-валидации заданный по умолчанию в классе логистической регрессии функционал accuracy, а измерять на тесте AUC ROC, но это, как и ограничение размера выборки, сделано для ускорения работы процесса кросс-валидации.
# +
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_auc_score
def plot_scores(optimizer):
scores=[]
for i in range(len(optimizer.cv_results_['params'])):
scores.append([optimizer.cv_results_['params'][i]['C'],
optimizer.cv_results_['mean_test_score'][i],
optimizer.cv_results_['std_test_score'][i]])
scores = np.array(scores)
plt.semilogx(scores[:,0], scores[:,1])
plt.fill_between(scores[:,0], scores[:,1]-scores[:,2],
scores[:,1]+scores[:,2], alpha=0.3)
plt.show()
def write_answer_1(auc_1, auc_2):
auc = (auc_1 + auc_2)/2
with open("preprocessing_lr_answer1.txt", "w") as fout:
fout.write(str(auc))
param_grid = {'C': [0.01, 0.05, 0.1, 0.5, 1, 5, 10]}
cv = 3
# place your code here
# -
X_train_cat_zeros = pd.DataFrame(data=np.hstack((X_train_cat_oh, X_train_real_zeros)))
X_test_cat_zeros = pd.DataFrame(data=np.hstack((X_test_cat_oh, X_test_real_zeros)))
logreg = LogisticRegression(solver='liblinear', penalty='l2')
# +
#zeros
grid1 = GridSearchCV(logreg, param_grid, cv=cv)
grid1.fit(X_train_cat_zeros, y_train)
# -
grid1.best_estimator_.predict_proba(X_test_cat_zeros)[:, 1:2]
answer1 = roc_auc_score(y_test, grid1.best_estimator_.predict_proba(X_test_cat_zeros)[:, 1:2])
print(answer1)
X_train_cat_means = pd.DataFrame(data=np.hstack((X_train_real_mean, X_train_cat_oh)))
X_test_cat_means = pd.DataFrame(data=np.hstack((X_test_real_mean, X_test_cat_oh)))
grid2 = GridSearchCV(logreg, param_grid, cv=cv)
grid2.fit(X_train_cat_means, y_train)
grid2.best_params_
answer2 = roc_auc_score(y_test, grid2.best_estimator_.predict_proba(X_test_cat_means)[:, 1])
print(answer1, answer2)
(answer1 + answer2) / 2
# ## Масштабирование вещественных признаков.
# Попробуем как-то улучшить качество классификации. Для этого посмотрим на сами данные:
# +
from pandas.plotting import scatter_matrix
data_numeric = pd.DataFrame(X_train_real_zeros, columns=numeric_cols)
list_cols = ['Number.of.Successful.Grant.1', 'SEO.Percentage.2', 'Year.of.Birth.1']
scatter_matrix(data_numeric[list_cols], alpha=0.5, figsize=(10, 10))
plt.show()
# -
# Как видно из графиков, разные признаки очень сильно отличаются друг от друга по модулю значений (обратите внимание на диапазоны значений осей x и y). В случае обычной регрессии это никак не влияет на качество обучаемой модели, т.к. у меньших по модулю признаков будут большие веса, но при использовании регуляризации, которая штрафует модель за большие веса, регрессия, как правило, начинает работать хуже.
#
# В таких случаях всегда рекомендуется делать стандартизацию (масштабирование) признаков, для того чтобы они меньше отличались друг друга по модулю, но при этом не нарушались никакие другие свойства признакового пространства. При этом даже если итоговое качество модели на тесте уменьшается, это повышает её интерпретабельность, потому что новые веса имеют смысл "значимости" данного признака для итоговой классификации.
#
# Стандартизация осуществляется посредством вычета из каждого признака среднего значения и нормировки на выборочное стандартное отклонение:
#
# $$ x^{scaled}_{id} = \dfrac{x_{id} - \mu_d}{\sigma_d}, \quad \mu_d = \frac{1}{N} \sum_{i=1}^l x_{id}, \quad \sigma_d = \sqrt{\frac{1}{N-1} \sum_{i=1}^l (x_{id} - \mu_d)^2} $$
# ## Задание 1.5. Масштабирование вещественных признаков.
#
# 1. По аналогии с вызовом one-hot encoder примените масштабирование вещественных признаков для обучающих и тестовых выборок X_train_real_zeros и X_test_real_zeros, используя класс
#
# StandardScaler
#
# и методы
#
# StandardScaler.fit_transform(...)
# StandardScaler.transform(...)
# 2. Сохраните ответ в переменные X_train_real_scaled и X_test_real_scaled соответственно
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train_real_zeros)
X_train_real_scaled = scaler.transform(X_train_real_zeros)
X_test_real_scaled = scaler.transform(X_test_real_zeros)
# -
# ## Сравнение признаковых пространств.
# Построим такие же графики для преобразованных данных:
data_numeric_scaled = pd.DataFrame(X_train_real_scaled, columns=numeric_cols)
list_cols = ['Number.of.Successful.Grant.1', 'SEO.Percentage.2', 'Year.of.Birth.1']
scatter_matrix(data_numeric_scaled[list_cols], alpha=0.5, figsize=(10, 10))
plt.show()
# Как видно из графиков, мы не поменяли свойства признакового пространства: гистограммы распределений значений признаков, как и их scatter-plots, выглядят так же, как и до нормировки, но при этом все значения теперь находятся примерно в одном диапазоне, тем самым повышая интерпретабельность результатов, а также лучше сочетаясь с идеологией регуляризации.
# ## Задание 2. Сравнение качества классификации до и после масштабирования вещественных признаков.
# 1. Обучите ещё раз регрессию и гиперпараметры на новых признаках, объединив их с закодированными категориальными.
# 2. Проверьте, был ли найден оптимум accuracy по гиперпараметрам во время кроссвалидации.
# 3. Получите значение ROC AUC на тестовой выборке, сравните с лучшим результатом, полученными ранее.
# 4. Запишите полученный ответ в файл при помощи функции write_answer_2.
# +
def write_answer_2(auc):
with open("preprocessing_lr_answer2.txt", "w") as fout:
fout.write(str(auc))
# place your code here
# -
X_train = np.hstack((X_train_real_scaled, X_train_cat_oh))
X_test = np.hstack((X_test_real_scaled, X_test_cat_oh))
grid3 = GridSearchCV(logreg, param_grid, cv=cv)
grid3.fit(X_train, y_train)
# +
c = [row['C'] for row in grid3.cv_results_['params']]
test_scores = grid3.cv_results_['mean_test_score']
test_std = grid3.cv_results_['std_test_score']
plt.fill_between(c, test_scores - test_std, test_scores + test_std, alpha=0.3)
plt.plot(c, test_scores)
plt.xscale('log')
# -
roc_auc_score(y_test, grid3.best_estimator_.predict_proba(X_test)[:, 1])
# ## Балансировка классов.
# Алгоритмы классификации могут быть очень чувствительны к несбалансированным классам. Рассмотрим пример с выборками, сэмплированными из двух гауссиан. Их мат. ожидания и матрицы ковариации заданы так, что истинная разделяющая поверхность должна проходить параллельно оси x. Поместим в обучающую выборку 20 объектов, сэмплированных из 1-й гауссианы, и 10 объектов из 2-й. После этого обучим на них линейную регрессию, и построим на графиках объекты и области классификации.
np.random.seed(0)
"""Сэмплируем данные из первой гауссианы"""
data_0 = np.random.multivariate_normal([0,0], [[0.5,0],[0,0.5]], size=40)
"""И из второй"""
data_1 = np.random.multivariate_normal([0,1], [[0.5,0],[0,0.5]], size=40)
"""На обучение берём 20 объектов из первого класса и 10 из второго"""
example_data_train = np.vstack([data_0[:20,:], data_1[:10,:]])
example_labels_train = np.concatenate([np.zeros((20)), np.ones((10))])
"""На тест - 20 из первого и 30 из второго"""
example_data_test = np.vstack([data_0[20:,:], data_1[10:,:]])
example_labels_test = np.concatenate([np.zeros((20)), np.ones((30))])
"""Задаём координатную сетку, на которой будем вычислять область классификации"""
xx, yy = np.meshgrid(np.arange(-3, 3, 0.02), np.arange(-3, 3, 0.02))
"""Обучаем регрессию без балансировки по классам"""
optimizer = GridSearchCV(LogisticRegression(), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train, example_labels_train)
"""Строим предсказания регрессии для сетки"""
Z = optimizer.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
"""Считаем AUC"""
auc_wo_class_weights = roc_auc_score(example_labels_test, optimizer.predict_proba(example_data_test)[:,1])
plt.title('Without class weights')
plt.show()
print('AUC: %f'%auc_wo_class_weights)
"""Для второй регрессии в LogisticRegression передаём параметр class_weight='balanced'"""
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced'), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train, example_labels_train)
Z = optimizer.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
auc_w_class_weights = roc_auc_score(example_labels_test, optimizer.predict_proba(example_data_test)[:,1])
plt.title('With class weights')
plt.show()
print('AUC: %f'%auc_w_class_weights)
# Как видно, во втором случае классификатор находит разделяющую поверхность, которая ближе к истинной, т.е. меньше переобучается. Поэтому на сбалансированность классов в обучающей выборке всегда следует обращать внимание.
#
# Посмотрим, сбалансированны ли классы в нашей обучающей выборке:
print(np.sum(y_train==0))
print(np.sum(y_train==1))
# Видно, что нет.
#
# Исправить ситуацию можно разными способами, мы рассмотрим два:
# - давать объектам миноритарного класса больший вес при обучении классификатора (рассмотрен в примере выше)
# - досэмплировать объекты миноритарного класса, пока число объектов в обоих классах не сравняется
# ## Задание 3. Балансировка классов.
# 1. Обучите логистическую регрессию и гиперпараметры с балансировкой классов, используя веса (параметр class_weight='balanced' регрессии) на отмасштабированных выборках, полученных в предыдущем задании. Убедитесь, что вы нашли максимум accuracy по гиперпараметрам.
# 2. Получите метрику ROC AUC на тестовой выборке.
# 3. Сбалансируйте выборку, досэмплировав в неё объекты из меньшего класса. Для получения индексов объектов, которые требуется добавить в обучающую выборку, используйте следующую комбинацию вызовов функций:
# np.random.seed(0)
# indices_to_add = np.random.randint(...)
# X_train_to_add = X_train[y_train.as_matrix() == 1,:][indices_to_add,:]
# После этого добавьте эти объекты в начало или конец обучающей выборки. Дополните соответствующим образом вектор ответов.
# 4. Получите метрику ROC AUC на тестовой выборке, сравните с предыдущим результатом.
# 5. Внесите ответы в выходной файл при помощи функции write_asnwer_3, передав в неё сначала ROC AUC для балансировки весами, а потом балансировки выборки вручную.
# +
def write_answer_3(auc_1, auc_2):
auc = (auc_1 + auc_2) / 2
with open("preprocessing_lr_answer3.txt", "w") as fout:
fout.write(str(auc))
# place your code here
# -
logreg_weight = LogisticRegression(solver='liblinear', class_weight='balanced', n_jobs=-1)
optimizator = GridSearchCV(logreg_weight, param_grid, cv=cv)
optimizator.fit(X_train, y_train)
optimizator.best_params_
answer1 = roc_auc_score(y_test, optimizator.best_estimator_.predict_proba(X_test)[:, 1])
# +
np.random.seed(42)
indices = np.random.randint(low=0, high=len(X_train[y_train == 1]), size=np.sum(y_train == 0) - np.sum(y_train == 1))
X_train_to_add = X_train[y_train == 1][indices]
X_train_over = np.vstack((X_train, X_train_to_add))
y_train_over = np.vstack((y_train.values.reshape(-1, 1), np.ones((len(X_train_to_add), 1))))
# -
optimizator_over = GridSearchCV(logreg_weight, param_grid, cv=cv)
optimizator_over.fit(X_train_over, y_train_over)
roc_auc_score(y_test, optimizator_over.best_estimator_.predict_proba(X_test)[:, 1])
# ## Стратификация выборок.
# Рассмотрим ещё раз пример с выборками из нормальных распределений. Посмотрим ещё раз на качество классификаторов, получаемое на тестовых выборках:
print('AUC ROC for classifier without weighted classes', auc_wo_class_weights)
print('AUC ROC for classifier with weighted classes: ', auc_w_class_weights)
# Насколько эти цифры реально отражают качество работы алгоритма, если учесть, что тестовая выборка так же несбалансирована, как обучающая? При этом мы уже знаем, что алгоритм логистический регрессии чувствителен к балансировке классов в обучающей выборке, т.е. в данном случае на тесте он будет давать заведомо заниженные результаты. Метрика классификатора на тесте имела бы гораздо больший смысл, если бы объекты были разделы в выборках поровну: по 20 из каждого класса на обучени и на тесте. Переформируем выборки и подсчитаем новые ошибки:
"""Разделим данные по классам поровну между обучающей и тестовой выборками"""
example_data_train = np.vstack([data_0[:20,:], data_1[:20,:]])
example_labels_train = np.concatenate([np.zeros((20)), np.ones((20))])
example_data_test = np.vstack([data_0[20:,:], data_1[20:,:]])
example_labels_test = np.concatenate([np.zeros((20)), np.ones((20))])
"""Обучим классификатор"""
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced'), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train, example_labels_train)
Z = optimizer.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
auc_stratified = roc_auc_score(example_labels_test, optimizer.predict_proba(example_data_test)[:,1])
plt.title('With class weights')
plt.show()
print('AUC ROC for stratified samples: ', auc_stratified)
# Как видно, после данной процедуры ответ классификатора изменился незначительно, а вот качество увеличилось. При этом, в зависимости от того, как вы разбили изначально данные на обучение и тест, после сбалансированного разделения выборок итоговая метрика на тесте может как увеличиться, так и уменьшиться, но доверять ей можно значительно больше, т.к. она построена с учётом специфики работы классификатора. Данный подход является частным случаем т.н. метода стратификации.
# ## Задание 4. Стратификация выборки.
#
# 1. По аналогии с тем, как это было сделано в начале задания, разбейте выборки X_real_zeros и X_cat_oh на обучение и тест, передавая в функцию
# train_test_split(...)
# дополнительно параметр
# stratify=y
# Также обязательно передайте в функцию переменную random_state=0.
# 2. Выполните масштабирование новых вещественных выборок, обучите классификатор и его гиперпараметры при помощи метода кросс-валидации, делая поправку на несбалансированные классы при помощи весов. Убедитесь в том, что нашли оптимум accuracy по гиперпараметрам.
# 3. Оцените качество классификатора метрике AUC ROC на тестовой выборке.
# 4. Полученный ответ передайте функции write_answer_4
# +
def write_answer_4(auc):
with open("preprocessing_lr_answer4.txt", "w") as fout:
fout.write(str(auc))
# place your code here
# -
X_zeros_train, X_zeros_test, X_cat_oh_train, X_cat_oh_test, y_train, y_test = train_test_split(X_real_zeros, X_cat_oh, y, test_size=0.3,
random_state=0, stratify=y)
# +
scaler = StandardScaler()
scaler.fit(X_zeros_train)
X_train_scaled = scaler.transform(X_zeros_train)
X_test_scaled = scaler.transform(X_zeros_test)
X_train = np.hstack((X_train_scaled, X_cat_oh_train))
X_test = np.hstack((X_test_scaled, X_cat_oh_test))
# -
logreg = LogisticRegression(class_weight='balanced', solver='liblinear', n_jobs=-1)
optimizator = GridSearchCV(logreg, param_grid, cv=cv)
optimizator.fit(X_train, y_train)
roc_auc_score(y_test, optimizator.best_estimator_.predict_proba(X_test)[:, 1])
# Теперь вы разобрались с основными этапами предобработки данных для линейных классификаторов.
# Напомним основные этапы:
# - обработка пропущенных значений
# - обработка категориальных признаков
# - стратификация
# - балансировка классов
# - масштабирование
#
# Данные действия с данными рекомендуется проводить всякий раз, когда вы планируете использовать линейные методы. Рекомендация по выполнению многих из этих пунктов справедлива и для других методов машинного обучения.
# ## Трансформация признаков.
#
# Теперь рассмотрим способы преобразования признаков. Существует достаточно много различных способов трансформации признаков, которые позволяют при помощи линейных методов получать более сложные разделяющие поверхности. Самым базовым является полиномиальное преобразование признаков. Его идея заключается в том, что помимо самих признаков вы дополнительно включаете набор все полиномы степени $p$, которые можно из них построить. Для случая $p=2$ преобразование выглядит следующим образом:
#
# $$ \phi(x_i) = [x_{i,1}^2, ..., x_{i,D}^2, x_{i,1}x_{i,2}, ..., x_{i,D} x_{i,D-1}, x_{i,1}, ..., x_{i,D}, 1] $$
#
# Рассмотрим принцип работы данных признаков на данных, сэмплированных их гауссиан:
# +
from sklearn.preprocessing import PolynomialFeatures
"""Инициализируем класс, который выполняет преобразование"""
transform = PolynomialFeatures(2)
"""Обучаем преобразование на обучающей выборке, применяем его к тестовой"""
example_data_train_poly = transform.fit_transform(example_data_train)
example_data_test_poly = transform.transform(example_data_test)
"""Обращаем внимание на параметр fit_intercept=False"""
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced', fit_intercept=False), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train_poly, example_labels_train)
Z = optimizer.predict(transform.transform(np.c_[xx.ravel(), yy.ravel()])).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
plt.title('With class weights')
plt.show()
# -
# Видно, что данный метод преобразования данных уже позволяет строить нелинейные разделяющие поверхности, которые могут более тонко подстраиваться под данные и находить более сложные зависимости. Число признаков в новой модели:
print(example_data_train_poly.shape)
# Но при этом одновременно данный метод способствует более сильной способности модели к переобучению из-за быстрого роста числа признаком с увеличением степени $p$. Рассмотрим пример с $p=11$:
transform = PolynomialFeatures(11)
example_data_train_poly = transform.fit_transform(example_data_train)
example_data_test_poly = transform.transform(example_data_test)
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced', fit_intercept=False), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train_poly, example_labels_train)
Z = optimizer.predict(transform.transform(np.c_[xx.ravel(), yy.ravel()])).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
plt.title('Corrected class weights')
plt.show()
# Количество признаков в данной модели:
print(example_data_train_poly.shape)
# ## Задание 5. Трансформация вещественных признаков.
#
# 1. Реализуйте по аналогии с примером преобразование вещественных признаков модели при помощи полиномиальных признаков степени 2
# 2. Постройте логистическую регрессию на новых данных, одновременно подобрав оптимальные гиперпараметры. Обращаем внимание, что в преобразованных признаках уже присутствует столбец, все значения которого равны 1, поэтому обучать дополнительно значение $b$ не нужно, его функцию выполняет один из весов $w$. В связи с этим во избежание линейной зависимости в датасете, в вызов класса логистической регрессии требуется передавать параметр fit_intercept=False. Для обучения используйте стратифицированные выборки с балансировкой классов при помощи весов, преобразованные признаки требуется заново отмасштабировать.
# 3. Получите AUC ROC на тесте и сравните данный результат с использованием обычных признаков.
# 4. Передайте полученный ответ в функцию write_answer_5.
# +
def write_answer_5(auc):
with open("preprocessing_lr_answer5.txt", "w") as fout:
fout.write(str(auc))
# place your code here
# +
transformer = PolynomialFeatures(2)
X_train_zeros_pol = transformer.fit_transform(X_zeros_train)
X_test_zeros_pol = transformer.transform(X_zeros_test)
scaler = StandardScaler()
X_train_zeros_pol = scaler.fit_transform(X_train_zeros_pol)
X_test_zeros_pol = scaler.transform(X_test_zeros_pol)
# -
X_train = np.hstack((X_train_zeros_pol, X_cat_oh_train))
X_test = np.hstack((X_test_zeros_pol, X_cat_oh_test))
logreg = LogisticRegression(solver='liblinear', class_weight='balanced', n_jobs=-1, fit_intercept=False)
optimizer = GridSearchCV(logreg, param_grid, cv=cv)
optimizer.fit(X_train, y_train)
optimizer.best_params_
roc_auc_score(y_test, optimizer.best_estimator_.predict_proba(X_test)[:, 1])
# ## Регрессия Lasso.
# К логистической регрессии также можно применить L1-регуляризацию (Lasso), вместо регуляризации L2, которая будет приводить к отбору признаков. Вам предлагается применить L1-регуляцию к исходным признакам и проинтерпретировать полученные результаты (применение отбора признаков к полиномиальным так же можно успешно применять, но в нём уже будет отсутствовать компонента интерпретации, т.к. смысловое значение оригинальных признаков известно, а полиномиальных - уже может быть достаточно нетривиально). Для вызова логистической регрессии с L1-регуляризацией достаточно передать параметр penalty='l1' в инициализацию класса.
# ## Задание 6. Отбор признаков при помощи регрессии Lasso.
# 1. Обучите регрессию Lasso на стратифицированных отмасштабированных выборках, используя балансировку классов при помощи весов. Для задания используем X_train_real_zeros.
# 2. Получите ROC AUC регрессии, сравните его с предыдущими результатами.
# 3. Найдите номера вещественных признаков, которые имеют нулевые веса в итоговой модели.
# 4. Передайте их список функции write_answer_6.
# +
def write_answer_6(features):
with open("preprocessing_lr_answer6.txt", "w") as fout:
fout.write(" ".join([str(num) for num in features]))
# place your code here
# -
X_train_zeros_scaled = scaler.fit_transform(X_zeros_train)
X_test_zeros_scaled = scaler.transform(X_zeros_test)
X_train = np.hstack((X_train_zeros_scaled, X_cat_oh_train))
X_test = np.hstack((X_test_zeros_scaled, X_cat_oh_test))
logreg = LogisticRegression(solver='liblinear', penalty='l1', n_jobs=-1, class_weight='balanced')
optimizer = GridSearchCV(logreg, param_grid, cv=cv)
optimizer.fit(X_train, y_train)
optimizer.best_params_
roc_auc_score(y_test, optimizer.best_estimator_.predict_proba(X_test)[:, 1])
np.where(optimizer.best_estimator_.coef_.T[:13] == 0)[0]
| 37,207 |
/AirBnB Vancouver - CRISP-DM.ipynb | 435581cb6dfff78e2331e4029c98b0d70f806e52 | [] | no_license | will-to-ride/Vancouver-AirBnB | https://github.com/will-to-ride/Vancouver-AirBnB | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 640,979 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# AirBnB Vancouver - CRISP-DM Model
#
# Using the CRISP-DM, the Vancouver AirBnB data set is collected, cleaned, and engineered in order to answer three questions:
#
# 1. How does price vary by date and neighbourhood?
# 2. How are review scores related to the price?
# 3. What are the most important features/ameninities that affect price?
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error
import seaborn as sns
# %matplotlib inline
pd.set_option('display.max_columns', None)
# -
#the review data
df_reviews = pd.read_csv('./data/reviews.csv')
df_reviews.head()
df_reviews.shape
#the listings data
df_listings = pd.read_csv('./data/listings.csv')
df_listings.head()
df_listings.columns
df_listings.shape
# There are 4273 listings, and 96 features
#neighbourhood data
df_nghbrhds = pd.read_csv('./data/neighbourhoods.csv')
df_nghbrhds.head(10)
#booking data
df_cal = pd.read_csv('./data/calendar.csv')
df_cal.head()
df_cal.shape
# First, look at how the price is distributed
# +
#clean the dollar signs from the listing prices
df_listings_price = df_listings[['price']].replace('[\$,]','',regex=True).astype(float)
ax = df_listings_price.plot.hist(bins=200, xlim=(0,800), figsize=(10,5))
ax.set(xlabel="Price", ylabel="Frequency")
# -
# The price distribution is right-skewed. Most listings are in the range of $\$$50 to $\$$150.
# # 1. How does price vary by date and neighbourhood?
#
# In order to examine the price variability by data, we will look at the calendar data
#clean the calendar data up
df_cal['date'] = pd.to_datetime(df_cal['date'])
df_cal[['price']] = df_cal[['price']].replace('[\$,]','',regex=True).astype(float)
df_cal = df_cal.dropna()
price_df = df_cal.groupby(['date'])['price'].mean()
ax = price_df.resample('D').mean().plot(figsize=(10,5))
ax.set(xlabel="Date", ylabel="Price")
# Noteables: is the seasonal fluctuation, christmas holiday increase, and weekend increases
# To analyze how price varies by neighbourhood, we will use the listings data
# +
df_listings[['price']] = df_listings[['price']].replace('[\$,]','',regex=True).astype(float)
price_df = df_listings.groupby(['neighbourhood_cleansed'])['price'].mean()
ax = price_df.sort_values().plot.bar(sort_columns=True, figsize=(10,5))
ax.set(xlabel="Neighbourhood", ylabel="Average Price")
# -
#Plot price on their corresponding longitude and latitude
ax = df_listings.plot.scatter(x="longitude", y="latitude", alpha=0.3, s=df_listings["price"]/15, label="price",
figsize=(12,8), c="price", cmap=plt.get_cmap("jet"), colorbar=True)
ax.set(xlabel="Longitude", ylabel="Latitude")
# We see that the downtown, and the neighbourhoods near downtown, have the highest prices. Also, downtown has the highest density of listings
# # 2. How are review scores and price related?
#plot review score ratings vs price
ax = df_listings.plot.scatter(x='review_scores_rating', y='price', color='DarkBlue',ylim=(0,2000),figsize=(10,5));
ax.set(xlabel="Review Scores", ylabel="Price")
# We see that there is no certainty that a high review score means a high price, though we do see that for all but one instance, all listings above $500 earn ratings greater than 80%
# # 3. What are the most important features/ameninities that affect price?
# Keep only columns that could be relevant to price. I assume these to be mainly features the potential customer would be interested in.
cols_to_keep = [ 'host_response_time', 'host_response_rate', 'host_acceptance_rate',
'host_is_superhost', 'host_identity_verified', 'neighbourhood_cleansed', 'city', 'neighbourhood_group_cleansed',
'latitude', 'longitude', 'property_type', 'room_type', 'accommodates', 'bathrooms',
'bedrooms', 'beds', 'bed_type', 'amenities', 'square_feet', 'price', 'weekly_price',
'monthly_price', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people',
'minimum_nights', 'maximum_nights', 'has_availability', 'availability_365', 'number_of_reviews',
'review_scores_rating', 'review_scores_accuracy', 'review_scores_cleanliness',
'review_scores_checkin', 'review_scores_communication', 'review_scores_value', 'requires_license',
'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'calculated_host_listings_count', 'reviews_per_month']
df_listings_reduced = df_listings[cols_to_keep]
# a look into the missing data
missing = pd.DataFrame(df_listings_reduced.isnull().sum()/df_listings_reduced.shape[0]*100, columns = ["missing (%)"])
#take only columns with missing data and sort
missing = missing[missing["missing (%)"]>0.0]
missing = missing.sort_values("missing (%)", ascending = False)
missing
# First we will drop columns that have more than 80% of the data missing.
to_drop = ['neighbourhood_group_cleansed','host_acceptance_rate','square_feet','weekly_price','monthly_price']
df_listings_reduced.drop(to_drop,axis=1,inplace=True)
df_listings_reduced.shape
#look at data types
df_listings_reduced.dtypes
# Separate out numeric columns and fill missing entries with the mean
# assign numeric features to separate dataframe
numeric_df = df_listings_reduced.select_dtypes(exclude=['O'])
numeric_df.head()
# Replace missing values with the median
numeric_df.fillna(numeric_df.median(), inplace=True)
#create separate dataframe for object datatypes
object_df = df_listings_reduced.select_dtypes(include=['O'])
object_df.head()
# +
# clean up the boolean columns by replace t and f, with True and False
bool_cols = ['host_is_superhost', 'host_identity_verified', 'has_availability', 'requires_license','instant_bookable',
'require_guest_profile_picture','require_guest_phone_verification']
for col in bool_cols:
object_df[col] = object_df[col].map(lambda x: True if x == "t" else False)
# -
# We will now clean up the amenities column by removing unnecessary chars and splitting them into separate columns while encoding them
# removing unnecessary characters from amenities
amenities = np.array(object_df['amenities'].map(lambda amns: amns.replace('"',"").replace('{',"").replace('}',"").split(',')))
# Create an array of all unique amenities
all_amenities = np.unique(np.concatenate(amenities))[1:]
# +
def available_amenties(all_amenities, amenities):
amenties_bool = np.empty((amenities.shape[0], all_amenities.shape[0]))
for i, row in enumerate(amenities):
amns = all_amenities.copy()
for j, amn in enumerate(amns):
if amn in amenities[i]:
amenties_bool[i,j] = True
else:
amenties_bool[i,j] = False
return amenties_bool
amenities = available_amenties(all_amenities, amenities)
# -
# Reset index of the train data frame and drop the old index
numeric_df = numeric_df.reset_index(drop=True)
# Combine the train data frame and the amenities data frame
df_listings_cleaned = pd.concat([numeric_df, pd.DataFrame(data=amenities, columns=all_amenities)], axis=1)
object_df.drop(['amenities'],axis=1,inplace=True)
#clean the percentage and dollar values
# object_df[['price']] = object_df[['price']].replace('[\$,]','',regex=True).astype(float)
object_df[['security_deposit']] = object_df[['security_deposit']].replace('[\$,]','',regex=True).astype(float)
object_df[['cleaning_fee']] = object_df[['cleaning_fee']].replace('[\$,]','',regex=True).astype(float)
object_df[['extra_people']] = object_df[['extra_people']].replace('[\$,]','',regex=True).astype(float)
object_df[['host_response_rate']] = object_df[['host_response_rate']].replace('[\%,]','',regex=True).astype(float)
object_df.isnull().sum()/object_df.shape[0]
#fill missing values with the mean
object_df.fillna(object_df.mean(), inplace=True)
# Getting dummy variables of categorical features
categorical_columns = ['host_response_time','city','neighbourhood_cleansed', 'property_type',
'room_type', 'bed_type','cancellation_policy']
object_df = pd.get_dummies(object_df,columns=categorical_columns)
object_df.head()
df_listings_clean = pd.concat([df_listings_cleaned,object_df],axis=1)
df_listings_clean.head()
# The data is now considered clean and we can proceed with analysis
#assign features and predictor (price)
X = df_listings_clean.drop(columns=['price'])
y = df_listings_clean['price']
#scale the features to mean 0 and std dev 1
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X)
X_scaled = scaler.transform(X)
#split the data into a training and test set. I chose a test size of 33% of the data.
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y, test_size=0.33, random_state=34)
# +
#First, try a decision tree regressor
from sklearn.tree import DecisionTreeRegressor
dt_model = DecisionTreeRegressor(random_state=34)
dt_model.fit(X_train, y_train)
#predict
dt_preds = dt_model.predict(X_train)
#get mean squared error
dt_mse = mean_squared_error(y_train, dt_preds)
dt_rmse = np.sqrt(dt_mse)
dt_rmse
# -
#Now try a random forest regressor
from sklearn.ensemble import RandomForestRegressor
rf_model = RandomForestRegressor(random_state=34)
rf_model.fit(X_train, y_train)
#predict and get mean squared error
rf_preds = rf_model.predict(X_train)
rf_mse = mean_squared_error(y_train, rf_preds)
rf_rmse = np.sqrt(rf_mse)
rf_rmse
# The decision tree appears to overfit so we will use the random forest regressor for the remainder of the analysis
# +
#perform a grid search to determmine the best parameters.
from sklearn.model_selection import GridSearchCV
param_grid = [
{'n_estimators': [3, 5, 10, 13, 15, 20, 30,35,40,60], 'max_features': [2, 4, 6, 8, 10,12,14]},
]
rf_model = RandomForestRegressor(random_state=34)
grid_search = GridSearchCV(rf_model, param_grid,
scoring='neg_mean_squared_error', return_train_score=True)
grid_search.fit(X_train, y_train)
# -
grid_search.best_params_
grid_search.best_estimator_
# +
#take the best model and calculate the RMSE
best_model = grid_search.best_estimator_
price_preds = best_model.predict(X_test)
final_mse = mean_squared_error(y_test, price_preds)
final_rmse = np.sqrt(final_mse)
final_rmse
# -
#important attributes from our model
imp_features = best_model.feature_importances_
attributes = df_listings_clean.drop(columns=['price']).columns
features = sorted(zip(imp_features, attributes), reverse=True)
#list of all attributes and their coefficient
att = []
coeff = []
for feature in features:
att.append(feature[1])
coeff.append(feature[0])
#plot top 25 important features
plt.figure(figsize=(10,5))
plt.bar(att[:25], height=coeff[:25])
plt.xticks(rotation=90)
plt.xlabel('Feature')
plt.ylabel('Feature Importance')
plt.show()
#Create list of all amenities and their coefficients
amenity= []
amn_coef = []
for feature in features:
if feature[1] in all_amenities:
amenity.append(feature[1])
amn_coef.append(feature[0])
#Plot 10 most important amenities
plt.figure(figsize=(10,5))
plt.bar(amenity[:10], height=amn_coef[:10])
plt.xticks(rotation=-90)
plt.xlabel('Amenity')
plt.ylabel('Amenity Importance')
plt.show()
# # Summary:
#
# We see that features such as accommodates, bedrooms, bathrooms, beds, and guests_included are important features for determining the price of an AirBnB. Usually, the more bedrooms, bathrooms, and guests, the higher the cost.
#
# Amongst ameninites, we see that safety cards and indoor fireplaces were important in determining price. It also appears that internet, a gym, and a hot tub have an affect on price
| 12,116 |
/Instructions/climate_starter.ipynb | a8a331cd5198b8e2398e15f788f892b3a1b786eb | [] | no_license | mjenkins15/Jenkinssqlalchemy_Challenge | https://github.com/mjenkins15/Jenkinssqlalchemy_Challenge | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 215,215 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pandas.plotting import table
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, inspect,func,desc
#Create an engine to connect to the database
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
conn = engine.connect()
# +
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# -
# We can view all of the classes that automap found
Base.classes.keys()
# +
# Save references to each table
# Assign the measurement class to a variable called "Measurement"
Measurement = Base.classes.measurement
# Assign the station class to a variabe called "Station"
Station = Base.classes.station
# -
# Create our session (link) from Python to the DB
session = Session(engine)
first_row = session.query(Measurement).first()
first_row
# +
# Display the row's columns and data in dictionary format (Measurement)
#first_row = session.query(Measurement).first()
#first_row.__dict__
# +
# Display the row's columns and data in dictionary format (Station)
#first_row = session.query(Station).first()
#first_row.__dict__
# -
# # Exploratory Climate Analysis
# + tags=["outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend"]
# DESIGN A QUERY TO RETRIEVE THE LAST 12 MONTHS OF PRECIPITATION DATA AND PLOT THE RESULTS
# Querying all records from the database
#precip = engine.execute("SELECT * FROM measurement")
#for record in precip:
# print (record)
# Don't need to use ("SELECT * FROM") because we are using the object version of sqlalchemy (from sqlalchemy.orm import Session), so we can just pass in the class.
# -
#Explore the database and print the table names using Inspector
inspector = inspect(engine)
inspector.get_table_names()
#Print the column names and types
columns = inspector.get_columns('measurement')
for column in columns:
print(column['name'], column['type'])
# Calculate the date 1 year ago from the last data point in the database
year_ago = dt.date(2017, 8, 23) - dt.timedelta(days = 365)
print ("The date 1 year ago was", year_ago)
# Query for the last 12 months of precipitation data selecting only the date and precipitation values
last_twelve_months=session.query(Measurement.date, Measurement.prcp).filter(Measurement.date >= year_ago).all()
last_twelve_months
# +
# Save the query results as a Pandas DataFrame and set the index to the date column
df = pd.DataFrame(last_twelve_months)
df.set_index('date')
# -
# Sort the dataframe by date
df = df.sort_values("date")
df
# Use Pandas Plotting with Matplotlib to plot the data
df.plot('date', 'prcp')
plt.xlabel("Date")
plt.xticks(rotation=90)
plt.ylabel("Rain (inches)")
plt.title("Precipitation from 8/23/2016 to 8/23/2017")
plt.show()
# Use Pandas to calcualte the summary statistics for the precipitation data
df.describe()
# Design a query to show how many stations are available in this dataset?
stations=session.query(Measurement.station).group_by(Measurement.station).count()
stations
# What are the most active stations? (i.e. what stations have the most rows)?
# List the stations and the counts in descending order.
most_active_station = session.query(Measurement.station, Station.name, func.count(Measurement.tobs)).filter(Measurement.station == Station.station).group_by(Measurement.station).order_by(func.count(Measurement.tobs).desc()).all()
most_active_station
# Using the station id from the previous query, calculate the lowest, highest, and average temperatures recorded from the most active station.
temp_ranges = session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).filter(Measurement.station == 'USC00519281').all()
temp_ranges
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station.
top_station_obs = session.query(Measurement.tobs).filter(Measurement.station == 'USC00519281').filter(Measurement.date >= year_ago).all()
top_station_obs
# +
#Example from matplotlib.org
#Create a bar chart based off of the group series from before
#N_points = 77
#n_bins = 12
# Generate a normal distribution, center at x=0 and y=5
#x = np.random.randn(N_points)
#y = .4 * x + np.random.randn(77) + 5
#fig, axs = plt.subplots(1, 2, sharey=True, tight_layout=True)
# We can set the number of bins with the `bins` kwarg
#axs[0].hist(x, bins=n_bins)
#axs[1].hist(y, bins=n_bins)
# -
df = pd.DataFrame(top_station_obs)
df.plot.hist(bins = 12)
plt.tight_layout()
plt.show()
# ## Bonus Challenge Assignment
# +
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# -
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
# +
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
# +
# Create a query that will calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
daily_normals("01-01")
# +
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
# Use the start and end date to create a range of dates
# Stip off the year and save a list of %m-%d strings
# Loop through the list of %m-%d strings and calculate the normals for each date
# -
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
# Plot the daily normals as an area plot with `stacked=False`
rain, y_train, cv=3, scoring='f1_macro', n_jobs=-1)
print(cv_results['test_score'].mean())
# -
adaboost.fit(X_train, y_train)
print(classification_report(adaboost.predict(X_test), y_test))
# ---
| 8,028 |
/notebooks/results/Results 300.ipynb | d4dc692aeaba5d3b71bdb75ad25faeb712c223eb | [] | no_license | murraymacmahon/Tennis-Data-Analysis | https://github.com/murraymacmahon/Tennis-Data-Analysis | 2 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 27,790 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Results 300
# ## Purpose
# In this notebook we will begin our analysis for research question 3 - 'Can we predict the next young Grand Slam winners?'. Focusing on analysing young players who fit between the mean and standard deviation of 6 match statistics of previous Grand Slam winners when they were under 23 y/o.
#
# ## Datasets
# * _Input_: GrandSlamWinners.csv, Under23_14_17.csv
# +
#importing relevant libraries
import os
import sys
import hashlib
import numpy as np
import pandas as pd
from datetime import datetime
import matplotlib.pyplot as plt
# %matplotlib inline
# -
stats_RQ3 = pd.read_csv("../data/stats_RQ3", low_memory = False)
# #### Names of players who appear between 6 and 5 of the standard deviations of the 6 match stats
# counting the number of times a name appears in the young player stats df created in Analysis 500
# sorted by w_ace as its the first column and they all have the same value
RQ3 = stats_RQ3.groupby(['winner_name']).count().sort_values(by ='w_ace', ascending=False)
RQ3.head(8)
# Three players achieved 5 out of 6 of the standard deviations of statistics from the Grand Slam winners.
# <br> Taylor Fritz ranked 68th
# <br> Dominic Thiem ranked 7th
# <br> Ryan Harrison ranked 59th
# Plot histogram of distribution of the young players by how many could reproduce
# similar statistics to previous Grand Slam winner
fig = plt.figure(figsize=(10,5))
RQ3['w_ace'].hist(bins=5, range=(1,6))
plt.xlabel('Young players, Under 23: 2014-2017', fontsize=10)
plt.ylabel('Number of statistics', fontsize=10)
plt.grid(True)
plt.title('Distribution of young players matching average Grand Slam winners statistics', fontsize=13)
# We see the distribution of the young players by how many could reproduce similar statistics to previous Grand Slam winner.
| 2,070 |
/site/en-snapshot/addons/tutorials/losses_triplet.ipynb | e3ec9b9bc0874cc0bf56110dd8b58219fe4fa8c7 | [
"Apache-2.0"
] | permissive | justaverygoodboy/docs-l10n | https://github.com/justaverygoodboy/docs-l10n | 0 | 0 | Apache-2.0 | 2020-08-29T16:06:29 | 2020-08-27T23:24:55 | null | Jupyter Notebook | false | false | .py | 11,302 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2020 The TensorFlow Authors.
#
# + cellView="form" colab={} colab_type="code" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # TensorFlow Addons Losses: TripletSemiHardLoss
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/addons/tutorials/losses_triplet"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/addons/blob/master/docs/tutorials/losses_triplet.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/addons/blob/master/docs/tutorials/losses_triplet.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/addons/docs/tutorials/losses_triplet.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# ## Overview
# This notebook will demonstrate how to use the TripletSemiHardLoss function in TensorFlow Addons.
#
# ### Resources:
# * [FaceNet: A Unified Embedding for Face Recognition and Clustering](https://arxiv.org/pdf/1503.03832.pdf)
# * [Oliver Moindrot's blog does an excellent job of describing the algorithm in detail](https://omoindrot.github.io/triplet-loss)
#
# + [markdown] colab_type="text" id="bQwBbFVAyHJ_"
# ## TripletLoss
#
# As first introduced in the FaceNet paper, TripletLoss is a loss function that trains a neural network to closely embed features of the same class while maximizing the distance between embeddings of different classes. To do this an anchor is chosen along with one negative and one positive sample.
# 
#
# **The loss function is described as a Euclidean distance function:**
#
# 
#
# Where A is our anchor input, P is the positive sample input, N is the negative sample input, and alpha is some margin we use to specify when a triplet has become too "easy" and we no longer want to adjust the weights from it.
# + [markdown] colab_type="text" id="wPJ5521HZHeL"
# ## SemiHard Online Learning
# As shown in the paper, the best results are from triplets known as "Semi-Hard". These are defined as triplets where the negative is farther from the anchor than the positive, but still produces a positive loss. To efficiently find these triplets we utilize online learning and only train from the Semi-Hard examples in each batch.
#
# + [markdown] colab_type="text" id="MUXex9ctTuDB"
# ## Setup
# -
# !pip install -U tensorflow-addons
# + colab={} colab_type="code" id="IqR2PQG4ZaZ0"
import io
import numpy as np
# + colab={} colab_type="code" id="WH_7-ZYZYblV"
import tensorflow as tf
import tensorflow_addons as tfa
import tensorflow_datasets as tfds
# + [markdown] colab_type="text" id="0_D7CZqkv_Hj"
# ## Prepare the Data
# + colab={} colab_type="code" id="iXvByj6wcT7d"
def _normalize_img(img, label):
img = tf.cast(img, tf.float32) / 255.
return (img, label)
train_dataset, test_dataset = tfds.load(name="mnist", split=['train', 'test'], as_supervised=True)
# Build your input pipelines
train_dataset = train_dataset.shuffle(1024).batch(32)
train_dataset = train_dataset.map(_normalize_img)
test_dataset = test_dataset.batch(32)
test_dataset = test_dataset.map(_normalize_img)
# + [markdown] colab_type="text" id="KR01t9v_fxbT"
# ## Build the Model
# + [markdown] colab_type="text" id="wvOPPuIKhLJi"
# 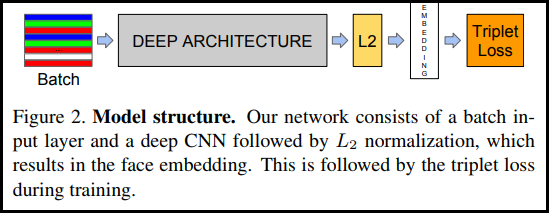
# + colab={} colab_type="code" id="djpoAvfWNyL5"
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(filters=64, kernel_size=2, padding='same', activation='relu', input_shape=(28,28,1)),
tf.keras.layers.MaxPooling2D(pool_size=2),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Conv2D(filters=32, kernel_size=2, padding='same', activation='relu'),
tf.keras.layers.MaxPooling2D(pool_size=2),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256, activation=None), # No activation on final dense layer
tf.keras.layers.Lambda(lambda x: tf.math.l2_normalize(x, axis=1)) # L2 normalize embeddings
])
# + [markdown] colab_type="text" id="HYE-BxhOzFQp"
# ## Train and Evaluate
# + colab={} colab_type="code" id="NxfYhtiSzHf-"
# Compile the model
model.compile(
optimizer=tf.keras.optimizers.Adam(0.001),
loss=tfa.losses.TripletSemiHardLoss())
# + colab={} colab_type="code" id="TGBYNGxgVDrj"
# Train the network
history = model.fit(
train_dataset,
epochs=5)
# + colab={} colab_type="code" id="1Y--0tK69SXf"
# Evaluate the network
results = model.predict(test_dataset)
# + colab={} colab_type="code" id="dqSuLdVZGNrZ"
# Save test embeddings for visualization in projector
np.savetxt("vecs.tsv", results, delimiter='\t')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for img, labels in tfds.as_numpy(test_dataset):
[out_m.write(str(x) + "\n") for x in labels]
out_m.close()
try:
from google.colab import files
files.download('vecs.tsv')
files.download('meta.tsv')
except:
pass
# + [markdown] colab_type="text" id="VAtj_m6Z_Uwe"
# ## Embedding Projector
# + [markdown] colab_type="text" id="Y4rjlG9rlbVA"
# The vector and metadata files can be loaded and visualized here: https://projector.tensorflow.org/
#
# You can see the results of our embedded test data when visualized with UMAP:
# 
#
| 6,867 |
/create password.ipynb | 435226fdf0e6ab197e775df0672cc758dd6bf3e8 | [] | no_license | Nisha-qureshi/Extra-Codes | https://github.com/Nisha-qureshi/Extra-Codes | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 1,213 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
from string import digits
from string import punctuation
from string import ascii_letters
symbols = ascii_letters + digits + punctuation
secure_random = random.SystemRandom()
password = "".join(secure_random.choice(symbols)for i in range(20))
print("This is highsecurity password ", password)
# -
| 584 |
/19_ene_cyp.ipynb | c9a0674d63ab7bfe06614963308786cca208aebb | [] | no_license | AndreStrong/cyp_2021 | https://github.com/AndreStrong/cyp_2021 | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 26,519 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# %config IPCompleter.greedy = True
from tensorflow import keras
import numpy as np
# +
model = keras.Sequential([keras.layers.Dense(units=1, input_shape=[1])])
model.compile(optimizer=keras.optimizers.SGD(),
loss=keras.losses.mean_squared_error)
xs = np.array([-1.0, 0.0, 1.0, 2.0, 3.0, 4.0], dtype=np.float)
ys = np.array([-3.0, -1.0, 1.0, 3.0, 5.0, 7.0], dtype=np.float)
model.fit(xs, ys, epochs=5000, verbose=False)
# -
model.predict([10.0])
/"} id="S-tTdHmNrU3e" outputId="f0f6f15a-c3fb-407f-8181-f17207cdba54"
frase = "Como hacer que la gente sea mas positiva? muy facil, haz que pierda un electron"
print(frase)
#Imprimir la palabra "ace" de la palabra hacer
print( frase[ 6 : 9 : 1] )
#valores por defecto
print(frase[ : 5 : 2])
print(len(frase))
#incremento
print(frase[ : : 2 ]) #toda la frase
print(frase [ -1 : : -1 ]) #inverso
#0 1 2 3 4 5 6 7 8
#-1 -2 -3 -4 -5 -6 -7 -8 -9
nombre = "Juan Jose"
print(nombre[ 1 : : -1 ])
primer = nombre [ -9 : : 1 ]
print(primer)
# + [markdown] id="I_kLiicTxPyj"
# # Metodos de la clase String (str)
#
# En python to se un objeto (P.O.O.)
# Esp signifoca que cada tipo de dato tre consigo metodos (funciones) incluidos.
#
# Para identificar los metodos disponibles se puede usar una de dos funciones:
#
# 1. dir(str) #Resumende los metodos de esa clase
# 2. help (str) #informacion completa de los metodos de esa clase
#
# + colab={"base_uri": "https://localhost:8080/"} id="zzHd-qp6yPFr" outputId="51336ebb-1bef-4458-82a6-07cf8281b31e"
nombre = "Juan Jose"
print(nombre)
print(dir(str))
# + [markdown] id="ex9uUHOiyudm"
# Los nombres de metodos que inician con doble guion bajo, significa que son metodos privados (POO), y no tendrias por que usarlos como programdos consumidor.
#
# Los demas metods son publicos y son de libre uso.
# + colab={"base_uri": "https://localhost:8080/"} id="MRW_VHKTzeau" outputId="9575cd07-31df-4dff-eda1-d9347aa8cf38"
print( nombre.islower())
nombre = "PEDRO FERNANDEZ"
print(nombre.capitalize())
# + colab={"base_uri": "https://localhost:8080/"} id="T5J5RRoUz73O" outputId="300d2cee-8b71-4ff4-b548-c6bdbdcaad7f"
print( help(str))
6,8)
plt.rcParams['figure.dpi'] = 150
sns.set()
from IPython.display import display, Latex, Markdown
# -
# ## `SciPy` and `special` ##
# Factorials and the *binomial coefficients* $\binom{n}{k} = \frac{n!}{k!(n-k)!}$ get large very quickly as $n$ gets large. One way to compute them is to use the `SciPy` module `special`. `SciPy` is a collection of Python-based software for math, probability, statistics, science, and engineering.
from scipy import special
# Below are some examples of `special.factorial`:
special.factorial(5), special.factorial(range(1, 6))
# Traditionally, subsets of $k$ individuals out of a population of $n$ individuals are called *combinations*, and so `special.comb(n, k)` evaluates to $\binom{n}{k}$.
#
# Note that we will always use the term *subsets* to mean un-ordered sets. We will use *permutations* in situations where we need to keep track of the order in which the elements appear.
#
# Look at the code and output below carefully (including types) before starting Question 1.
special.comb(5, 3), special.factorial(5) / (special.factorial(3) * special.factorial(2))
special.comb(5, range(6))
special.comb(100, 50), special.comb(100, 50, exact=True)
# ### Question 1
# Consider a population in which a proportion $p$ of individuals are called "successes" (or 1, if you prefer) and the remaining proportion are rudely called "failures" (or 0).
#
# As we saw in lecture, if you draw a sample of size $n$ (where $n$ is some positive integer) at random with replacement from the population, then the number of successes is a random variable that follows the binomial distribution. The probability of drawing $k$ successes and $n-k$ failures is $\binom{n}{k}p^k(1-p)^{n-k}$, for $0 \leq k \leq n$. To reduce writing, we will shorten "$k$ successes and $n-k$ failures" to "$k$ successes". In other words, "$k$ successes" means "exactly $k$ successes".
#
# To formalize notation: if $X$ is the number of successes we draw, then $P(X = k) = \binom{n}{k}p^k(1-p)^{n-k}$.
# + [markdown] deletable=false editable=false
# ### Question 1a) ###
# Suppose you sample 100 times at random with replacement from a population in which 26% of the individuals are successes. Write a Python expression that evaluates to the chance that the sample has 20 successes.
#
# **Computational note:** Don't import any other libraries; just use the ones already imported and plug into the formula above. It's far from the best way numerically, but it is fine for the numbers involved in this Homework.
#
# <!--
# BEGIN QUESTION
# name: q1a
# manual: false
# points: 1
# -->
# -
p = .26
prob_1a = special.comb(100, 20) * (p**20) * ((1-p)**80)
prob_1a
# + deletable=false editable=false
grader.check("q1a")
# + [markdown] deletable=false editable=false
# ### Question 1b) ###
# Complete the cell with a Python expression that evaluates to an array whose elements are the chances of $k$ successes for $k = 0, 1, 2, \ldots, 100$. That is, `all_probs[k]` should contain $P(X = k)$, where $X$ is the random variable describing the number of successes drawn from the scenario outlined above.
#
# <!--
# BEGIN QUESTION
# name: q1b
# points: 1
# -->
# -
k = np.arange(101)
all_probs = special.comb(100, k) * (p**k) * ((1-p)**(100-k))
np.round(all_probs, 4), sum(all_probs)
# + deletable=false editable=false
grader.check("q1b")
# + [markdown] deletable=false editable=false
# ### Question 1c) ###
#
#
# Complete the cell with an expression that evaluates to the chance that the number of successes in the sample is in the interval $26 \pm 10$ (inclusive on both sides). In other words, determine $P(16 \leq X \leq 36)$. **Hint: You might want to apply your Python indexing knowledge here.**
#
# **Note: Please assign your answer to `prob_1c`. Because we use an autograder, please make sure your answer is a proportion between 0 and 1, not a percent between 0 and 100.**
#
# <!--
# BEGIN QUESTION
# name: q1c
# manual: false
# points: 1
# -->
# -
prob_1c = sum(all_probs[16:37])
prob_1c
# + deletable=false editable=false
grader.check("q1c")
# -
# ### Question 2
# Consider (once again) a sample of size $n$ drawn at random with replacement from a population in which a proportion $p$ of the individuals are called successes.
#
# Let $S$ be the random variable that denotes the number of successes in our sample. (As stated above, $S$ follows the binomial distribution.) Then, the probability that the number of successes in our sample is **at most** $s$ (where $0 \leq s \leq n$) is
#
# $$P(S \leq s) = P(S = 0) + P(S = 1) + ... + P(S = s) = \sum_{k=0}^s \binom{n}{k}p^k(1-p)^{n-k}$$
#
# We obtain this by summing the probability that the number of successes is exactly $k$, for each value of $k = 0, 1, 2, ..., s$.
# + [markdown] deletable=false editable=false
# ### Question 2a) ###
#
# Please fill in the function `prob_at_most` which takes $n$, $p$, and $s$ and returns $P(S \le s)$ as defined above. If the inputs are invalid: for instance, if $p > 1$ OR $s > n$ then return 0."
#
# It might help to refer to the calculation in Question 1b.
#
# <!--
# BEGIN QUESTION
# name: q2a
# manual: false
# points: 1
# -->
# -
def prob_at_most(n, p, s):
"""
returns the probability of S <= s
Input n: sample size; p : proportion; s: number of successes at most"""
prob = []
if p > 1:
return 0
if s > n:
return 0
for i in range(s+1):
entry = special.comb(n,i) * (p**i) * ((1-p)**(n-i))
prob.append(entry)
return sum(prob)
# + deletable=false editable=false
grader.check("q2a")
# + [markdown] deletable=false editable=false
# ### Question 2b) ###
# In an election, supporters of Candidate C are in a minority. Only 45% of the voters in the population favor the candidate.
#
# Suppose a survey organization takes a sample of 200 voters at random with replacement from this population. Use `prob_at_most` to write an expression that evaluates to the chance that a majority (more than half) of the sampled voters favor Candidate C.
#
# <!--
# BEGIN QUESTION
# name: q2b
# manual: false
# points: 1
# -->
# -
total_voters = 200
c_prop = .45
p_majority = 1 - prob_at_most(200, .45, 100)
p_majority
# + deletable=false editable=false
grader.check("q2b")
# + [markdown] deletable=false editable=false
# ### Question 2c) ###
# Suppose each of five survey organizations takes a sample of voters at random with replacement from the population of voters in Part **b**, independently of the samples drawn by the other organizations.
#
# - Three of the organizations use a sample size of 200
# - One organization uses a sample size of 300
# - One organization uses a sample size of 400
#
# Write an expression that evaluates to the chance that in at least one of the five samples the majority of voters favor Candidate C. You can use any quantity or function defined earlier in this exercise.
#
# <!--
# BEGIN QUESTION
# name: q2c
# manual: false
# points: 2
# -->
# -
# 1 - prob(none of samples vote majority for C)
org_1 = prob_at_most(200,.45,100)
org_2 = prob_at_most(200,.45,100)
org_3 = prob_at_most(200,.45,100)
org_4 = prob_at_most(300,.45,150)
org_5 = prob_at_most(400,.45,200)
prob_2c = 1 - (org_1*org_2*org_3*org_4*org_5)
prob_2c
# + deletable=false editable=false
grader.check("q2c")
# -
# ### Question 3
# In Pennsylvania, 6,165,478 people voted in the 2016 Presidential election.
# Trump received 48.18% of the vote and Clinton recieved 47.46%.
# This doesn't add up to 100% because other candidates received votes.
# All together these other candidates received 100% - 48.18% - 47.46% = 4.36% of the vote.
#
# The table below displays the counts and proportions.
#
#
# | Voted for | Trump| Clinton| Other|
# |-----------|-----------|-----------|---------|
# | Probability | 0.4818 | 0.4746 | 0.0436 |
# | Number of people | 2,970,733 | 2,926,441 | 268,304 |
# + [markdown] deletable=false editable=false
# ### Question 3a) ###
# Suppose we pick a simple random sample of 20 of the 6,165,478 Pennsylvania (PA) voters.
#
# In the sample, let $N_T$ be the number of Trump voters, $N_C$ the number of Clinton voters, and $N_O$ the number of "other" voters. Then $N_T$, $N_C$, and $N_O$ are random: they depend on how the sample comes out. In Data 8 we called such quantities "statistics".
#
# Pick the correct option: $N_T + N_C + N_O$ is equal to
#
# (a) 3
#
# (b) 20
#
# (c) 6,165,478
#
# (d) a random quantity
#
# *Note*: if your answer is (d), put **ans_3a = 'd'** for the purpose of grading. Passing the test **does NOT** mean you answer is correct. The test only checks if your answer is in the correct format.
#
# <!--
# BEGIN QUESTION
# name: q3a
# manual: false
# points: 1
# -->
# -
ans_3a = 'b'
# + deletable=false editable=false
grader.check("q3a")
# + [markdown] deletable=false editable=false
# ### Question 3b) ###
# Pick the correct option.
#
# A simple random sample of 20 PA voters is like a sample drawn at random with replacement, because
#
# (a) that's the definition of "simple random sample"
#
# (b) there are only 3 categories of voters, which is small in comparison to 20
#
# (c) there are only 20 people in the sample, which is small in comparison to the total number of PA voters
#
# (d) all PA voters are equally likely to be selected
#
# *Note*: if your answer is (d), put **ans_3a = 'd'** for the purpose of grading. Passing the test **does NOT** mean you answer is correct. The test only checks if your answer is in the correct format.
#
# <!--
# BEGIN QUESTION
# name: q3b
# manual: false
# points: 1
# -->
# -
ans_3b = 'c'
# + deletable=false editable=false
grader.check("q3b")
# + [markdown] deletable=false editable=false
# ### Question 3c) ###
#
# Let $t$, $c$, and $o$ be any three non-negative integers. For the sample defined in Part **a**, the probability that the sample contains $t$ Trump voters, $c$ Clinton voters and $o$ "other" voters is denoted by $P(N_T = t, N_C = c, N_O = o)$.
#
# Define a function `prob_sample_counts` that takes any three non-negative integers $t$, $c$, and $o$, and returns $P(N_T = t, N_C = c, N_O = o)$. The probability is 0 for some choices of the arguments, and your function should return 0 in those cases.
#
# Remember that Part **b** implies you can use results for sampling with replacement.
#
# What should `prob_sample_counts(31, 8, 1)` evaluate to? Check that your function returns the right value (No extra code/cells required).
#
# <!--
# BEGIN QUESTION
# name: q3c
# manual: false
# points: 1
# -->
# -
trump = .4818
clinton = .4746
other = .0436
trump_voters = 2970733
clinton_voters = 2926441
other_voters = 268304
def prob_sample_counts(t, c, o):
"""
Input:
t - number of votes for Trump
c - number of votes for Clinton
o - number of votes for Other
Return: The probability of getting a such sample
"""
if (t + c + o) != 20:
return 0
part_1 = (special.factorial(t+c+o))/(special.factorial(t)*special.factorial(c)*special.factorial(o))
part_2 = (trump**t) * (clinton**c) * (other**o)
return part_1 * part_2
# + deletable=false editable=false
grader.check("q3c")
# + [markdown] deletable=false editable=false
# ### Question 3d) ###
#
# Check that your function determines a probability distribution, by summing all the positive terms. You know what the answer should be. **Hint: You might need to iterate through certain values of c and t to compute the sum of all positive probabilities.**
#
# <!--
# BEGIN QUESTION
# name: q3d
# manual: false
# points: 1
# -->
# -
def sum_probabilities():
total_probabilities = []
for t in range(21):
for c in range(21):
if t + c <= 20:
entry = prob_sample_counts(t, c, 20-t-c)
total_probabilities.append(entry)
return sum(total_probabilities)
prob_3d = sum_probabilities()
prob_3d
# + deletable=false editable=false
grader.check("q3d")
# + [markdown] deletable=false editable=false
# ### Question 3e) ###
# Use `prob_sample_counts` to find the chance that the sample consists of 11 Trump voters, 8 Clinton voters, and 1 "other" voter.
#
# <!--
# BEGIN QUESTION
# name: q3e
# manual: false
# points: 1
# -->
# -
prob_3e = prob_sample_counts(11,8,1)
prob_3e
# + deletable=false editable=false
grader.check("q3e")
# + [markdown] deletable=false editable=false
# ### Question 3f) ###
# Write an expression that evaluates the chance that the sample contains at least 7 Trump voters, at least 7 Clinton voters, and at least 1 "other" voter. **Hint: You might find 3d useful.**
#
# <!--
# BEGIN QUESTION
# name: q3f
# manual: false
# points: 2
# -->
# -
def at_least_7_7_1():
sample_list = []
for t in range(7,20):
for c in range(7,20):
if t+c < 20:
entry = prob_sample_counts(t, c, 20-t-c)
sample_list.append(entry)
return sum(sample_list)
prob_3f = at_least_7_7_1()
prob_3f
# + deletable=false editable=false
grader.check("q3f")
# + [markdown] nbgrader={"grade": false, "grade_id": "codebook", "locked": true, "schema_version": 2, "solution": false}
# ## Election Polling
#
# Political polling is a type of public opinion polling that can at best represent a snapshot of public opinion at the particular moment in time. Voter opinion shifts from week to week, even day to day, as candidates battle it out on the campaign field.
#
# Polls usually start with a "horse-race" question, where respondents are asked whom they would vote for in a head-to-head race if the election were tomorrow: Candidate A or Candidate B. The survey begins with this question so that the respondent is not influenced by any of the other questions asked in the survey. Some of these other questions are asked to help assess how likely is it that the respondent will vote. Other questions are asked about age, education, and sex in order to adjust the findings if one group appears overly represented in the sample.
#
# Pollsters typically use [random digit dialing](https://en.wikipedia.org/wiki/Random_digit_dialing) to contact people.
# -
# ### Question 4
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# #### Part 1 ####
#
# If we're trying to predict the results of the Clinton vs. Trump presidential race, what is the population of interest?
#
# <!--
# BEGIN QUESTION
# name: q4a
# manual: true
# points: 1
# -->
# -
# The population of interest is the U.S. citizens who can vote.
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-5a5f851db609367a", "locked": true, "schema_version": 2, "solution": false}
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# #### Part 2 ####
#
# What is the sampling frame?
#
# <!--
# BEGIN QUESTION
# name: q4b
# manual: true
# points: 1
# -->
# -
# The sampling frame is all people who have a registered phone number.
# + [markdown] nbgrader={"grade": false, "grade_id": "examine-contents", "locked": true, "schema_version": 2, "solution": false}
# <!-- END QUESTION -->
#
#
#
# ### How might the sampling frame differ from the population?
#
# After the fact, many experts have studied the 2016 election results. For example, according to the American Association for Public Opinion Research (AAPOR), predictions made before the election were flawed for three key reasons:
#
# 1. voters changed their preferences a few days before the election
# 2. those sampled were not representative of the voting population, e.g., some said that there was an overrepresentation of college graduates in some poll samples
# 3. voters kept their support for Trump to themselves (hidden from the pollsters)
#
# In the next two problems on this homework, we will do two things:
#
# # + HW Question 6: We will carry out a study of the sampling error when there is no bias. In other words, we will try to compute the chance that we get the election result wrong even if we collect our sample in a manner that is completely correct. In this case, any failure of our prediction is due entirely to random chance.
# # + HW Question 7: We will carry out a study of the sampling error when there is bias of the second type from the list above. In other words, we will try to compute the chance that we get the election result wrong if we have a small systematic bias. In this case, any failure of our prediction is due to a combination of random chance and our bias.
#
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# ### Question 5
#
# Why can't we assess the impact of the other two biases (voters changing preference and voters hiding their preference)?
#
# Note: You might find it easier to complete this question after you've completed the rest of the homework including the simulation study.
#
# <!--
# BEGIN QUESTION
# name: q5
# manual: true
# points: 1
# -->
# -
# We cannot assess the impact of the other two biases (voters changing preference and voters hiding their preference) because even if we use the best form of a sample, it cannot take in account for the two biases presented. Since the sample will only reflect the responses taken at the time the survey was taken, results will show the voting opinions of respondents at that time. Even if a respondent changes his mind after taking the survey, it will not show because results have already been recorded from the original response. Additionally, the sample will believe that respondents will be truthful with their responses and cannot tell whether they are lying. So even if a person says that he is in favor of Clinton on the survey, but in reality he is just embarrassed to say that he is a follower of Trump, the survey cannot take this into consideration.
# + [markdown] nbgrader={"grade": false, "grade_id": "examine-size", "locked": true, "schema_version": 2, "solution": false}
# <!-- END QUESTION -->
#
#
#
# ### How large was the sampling error?
#
# In some states the race was very close, and it may have been simply sampling error, i.e., random chance that the majority of the voters chosen for the sample voted for Clinton.
#
# One year after the 2016 election, Nate Silver wrote in
# *The Media Has A Probability Problem* that
# the "media’s demand for certainty -- and its lack of statistical rigor -- is a bad match for our complex world."
# FiveThirtyEight forecasted that Clinton had about a 70 percent chance of winning.
#
#
# A 2- or 3-point polling error in Trump’s favor (typical error historically) would likely be enough to tip the Electoral College to him.
#
# We will first carry out a simulation study to assess the impact of the sampling error on the predictions.
#
# + [markdown] nbgrader={"grade": false, "grade_id": "load-data", "locked": true, "schema_version": 2, "solution": false}
# ## The Electoral College
#
# The US president is chosen by the Electoral College, not by the
# popular vote. Each state is alotted a certain number of
# electoral college votes, as a function of their population.
# Whomever wins in the state gets all of the electoral college votes for that state.
#
# There are 538 electoral college votes (hence the name of the Nate Silver's site, FiveThirtyEight).
#
# Pollsters correctly predicted the election outcome in 46 of the 50 states.
# For these 46 states Trump received 231 and Clinton received 232 electoral college votes.
#
# The remaining 4 states accounted for a total of 75 votes, and
# whichever candidate received the majority of the electoral college votes in these states would win the election.
#
# These states were Florida, Michigan, Pennsylvania, and Wisconsin.
#
# |State |Electoral College Votes|
# | --- | --- |
# |Florida | 29 |
# |Michigan | 16 |
# |Pennsylvania | 20 |
# |Wisconsin | 10|
#
# For Donald Trump to win the election, he had to win either:
# * Florida + one (or more) other states
# * Michigan, Pennsylvania, and Wisconsin
#
#
# The electoral margins were very narrow in these four states, as seen below:
#
#
# |State | Trump | Clinton | Total Voters |
# | --- | --- | --- | --- |
# |Florida | 49.02 | 47.82 | 9,419,886 |
# |Michigan | 47.50 | 47.27 | 4,799,284|
# |Pennsylvania | 48.18 | 47.46 | 6,165,478|
# |Wisconsin | 47.22 | 46.45 | 2,976,150|
#
# Those narrow electoral margins can make it hard to predict the outcome given the sample sizes that the polls used.
# + [markdown] nbgrader={"grade": false, "grade_id": "q1", "locked": true, "schema_version": 2, "solution": false}
# ---
# ## Simulation Study of the Sampling Error
#
# Now that we know how people actually voted, we can carry
# out a simulation study that imitates the polling.
#
# Our ultimate goal in this problem is to understand the chance that we will incorrectly call the election for Hillary Clinton even if our sample was collected with absolutely no bias.
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "q1a", "locked": true, "schema_version": 2, "solution": false}
# ### Question 6
#
# #### Part 1
#
# For your convenience, the results of the vote in the four pivotal states is repeated below:
#
# |State | Trump | Clinton | Total Voters |
# | --- | --- | --- | --- |
# |Florida | 49.02 | 47.82 | 9,419,886 |
# |Michigan | 47.50 | 47.27 | 4,799,284|
# |Pennsylvania | 48.18 | 47.46 | 6,165,478|
# |Wisconsin | 47.22 | 46.45 | 2,976,150|
#
#
# Using the table above, write a function `draw_state_sample(N, state)` that returns a sample with replacement of N voters from the given state. Your result should be returned as a list, where the first element is the number of Trump votes, the second element is the number of Clinton votes, and the third is the number of Other votes. For example, `draw_state_sample(1500, "florida")` could return `[727, 692, 81]`. You may assume that the state name is given in all lower case.
#
# You might find `np.random.multinomial` useful.
#
# <!--
# BEGIN QUESTION
# name: q6a
# points: 2
# -->
# -
def draw_state_sample(N, state):
if state == 'florida':
return np.random.multinomial(N, [.4902, .4782, 1-(.4902+.4782)])
if state == 'michigan':
return np.random.multinomial(N, [.4750, .4727, 1-(.4750+.4727)])
if state == 'pennsylvania':
return np.random.multinomial(N, [.4818, .4746, 1-(.4818+.4746)])
if state == 'wisconsin':
return np.random.multinomial(N, [.4722, .4645, 1-(.4722+.4645)])
# + deletable=false editable=false
grader.check("q6a")
# + [markdown] deletable=false editable=false
# #### Part 2
#
# Now, create a function `trump_advantage` that takes in a sample of votes (like the one returned by `draw_state_sample`) and returns the difference in the proportion of votes between Trump and Clinton. For example `trump_advantage([100, 60, 40])` would return `0.2`, since Trump had 50% of the votes in this sample and Clinton had 30%.
#
# <!--
# BEGIN QUESTION
# name: q6b
# points: 1
# -->
# -
def trump_advantage(voter_sample):
return (voter_sample[0]/sum(voter_sample)) - (voter_sample[1]/sum(voter_sample))
# + deletable=false editable=false
grader.check("q6b")
# + [markdown] deletable=false editable=false
# #### Part 3
#
# Simulate Trump's advantage across 100,000 simple random samples of 1500 voters for the state of Pennsylvania and store the results of each simulation in a list called `simulations`.
#
# That is, `simulations[i]` should be Trump's proportion advantage for the `i+1`th simple random sample.
#
# <!--
# BEGIN QUESTION
# name: q6c
# points: 1
# -->
# +
def func():
simulations = []
for i in range(100000):
list_entry = trump_advantage(draw_state_sample(1500, 'pennsylvania'))
simulations.append(list_entry)
return simulations
simulations = func()
# + deletable=false editable=false
grader.check("q6c")
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "q1c", "locked": true, "schema_version": 2, "solution": false}
# <!-- BEGIN QUESTION -->
#
# #### Part 4
#
# Make a histogram of the sampling distribution of Trump's proportion advantage in Pennsylvania. Make sure to give your plot a title and add labels where appropriate.
# Hint: You should use the [`plt.hist`](https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.hist.html) function in your code.
#
# Make sure to include a title as well as axis labels. You can do this using `plt.title`, `plt.xlabel`, and `plt.ylabel`.
#
# <!--
# BEGIN QUESTION
# name: q6d
# manual: true
# points: 1
# -->
# -
plt.hist(simulations)
plt.title("Trump's Advantage in Pennsylvania over 100,000 Simulations")
plt.xlabel("Proportion Advantage")
plt.ylabel("Count")
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# #### Part 5
#
# Now write a function `trump_wins(N)` that creates a sample of N voters for each of the four crucial states (Florida, Michigan, Pennsylvania, and Wisconsin) and returns 1 if Trump is predicted to win based on these samples and 0 if Trump is predicted to lose.
#
# Recall that for Trump to win the election, he must either:
# * Win the state of Florida and 1 or more other states
# * Win Michigan, Pennsylvania, and Wisconsin
#
# <!--
# BEGIN QUESTION
# name: q6e
# manual: false
# points: 2
# -->
# -
def trump_wins(N):
florida = trump_advantage(draw_state_sample(N, 'florida'))
michigan = trump_advantage(draw_state_sample(N, 'michigan'))
pennsylvania = trump_advantage(draw_state_sample(N, 'pennsylvania'))
wisconsin = trump_advantage(draw_state_sample(N, 'wisconsin'))
if florida > 0 and michigan > 0:
return 1
if florida > 0 and pennsylvania > 0:
return 1
if florida > 0 and wisconsin > 0:
return 1
if michigan > 0 and pennsylvania > 0 and wisconsin > 0:
return 1
else:
return 0
# + deletable=false editable=false
grader.check("q6e")
# + [markdown] deletable=false editable=false
# #### Part 6
#
# If we repeat 100,000 simulations of the election, i.e. we call `trump_wins(1500)` 100,000 times, what proportion of these simulations predict a Trump victory? Give your answer as `proportion_trump`.
#
# This number represents the percent chance that a given sample will correctly predict Trump's victory *even if the sample was collected with absolutely no bias*.
#
# **Note: Many laypeople, even well educated ones, assume that this number should be 1. After all, how could a non-biased sample be wrong? This is the type of incredibly important intuition we hope to develop in you throughout this class and your future data science coursework.**
#
# <!--
# BEGIN QUESTION
# name: q6f
# manual: false
# points: 1
# -->
# +
def victory(x):
victories = 0
for i in range(100000):
if trump_wins(x) == 1:
victories += 1
return victories
proportion_trump = victory(1500)/100000
proportion_trump
# + deletable=false editable=false
grader.check("q6f")
# -
# We have just studied the sampling error, and found how
# our predictions might look if there was no bias in our
# sampling process.
# Essentially, we assumed that the people surveyed didn't change their minds,
# didn't hide who they voted for, and were representative
# of those who voted on election day.
# + [markdown] nbgrader={"grade": false, "grade_id": "q2", "locked": true, "schema_version": 2, "solution": false}
# ---
# ## Simulation Study of Selection Bias
#
# According to an article by Grotenhuis, Subramanian, Nieuwenhuis, Pelzer and Eisinga (https://blogs.lse.ac.uk/usappblog/2018/02/01/better-poll-sampling-would-have-cast-more-doubt-on-the-potential-for-hillary-clinton-to-win-the-2016-election/#Author):
#
# "In a perfect world, polls sample from the population of voters, who would state their political preference perfectly clearly and then vote accordingly."
#
# That's the simulation study that we just performed.
#
#
# It's difficult to control for every source of selection bias.
# And, it's not possible to control for some of the other sources of bias.
#
# Next we investigate the effect of small sampling bias on the polling results in these four battleground states.
#
# Throughout this problem, we'll examine the impacts of a 0.5 percent bias in favor of Clinton in each state. Such a bias has been suggested because highly educated voters tend to be more willing to participate in polls.
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "q2a", "locked": true, "schema_version": 2, "solution": false}
# ### Question 7
#
# Throughout this problem, adjust the selection of voters so that there is a 0.5% bias in favor of Clinton in each of these states.
#
# For example, in Pennsylvania, Clinton received 47.46 percent of the votes and Trump 48.18 percent. Increase the population of Clinton voters to 47.46 + 0.5 percent and correspondingly decrease the percent of Trump voters.
#
# #### Part 1
#
# Simulate Trump's advantage across 100,000 simple random samples of 1500 voters for the state of Pennsylvania and store the results of each simulation in a list called `biased_simulations`.
#
# That is, `biased_simulation[i]` should hold the result of the `i+1`th simulation.
#
# That is, your answer to this problem should be just like your answer from Question 7 part 3, but now using samples that are biased as described above.
#
# <!--
# BEGIN QUESTION
# name: q7a
# points: 1
# -->
# +
def draw_biased_state_sample(N, state):
if state == 'florida':
return np.random.multinomial(N, [.4902 - .005, .4782 + .005, 1-(.4902+.4782)])
if state == 'michigan':
return np.random.multinomial(N, [.4750 - .005, .4727 + .005, 1-(.4750+.4727)])
if state == 'pennsylvania':
return np.random.multinomial(N, [.4818 - .005, .4746 + .005, 1-(.4818+.4746)])
if state == 'wisconsin':
return np.random.multinomial(N, [.4722 - .005, .4645 + .005, 1-(.4722+.4645)])
def biased_trials(x):
results = []
for i in range(100000):
entry = trump_advantage(draw_biased_state_sample(x, 'pennsylvania'))
results.append(entry)
return results
biased_simulations = biased_trials(1500)
# + deletable=false editable=false
grader.check("q7a")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# #### Part 2
#
# Make a histogram of the new sampling distribution of Trump's proportion advantage now using these biased samples. That is, your histogram should be the same as in Q6.4, but now using the biased samples.
#
# Make sure to give your plot a title and add labels where appropriate.
#
#
# <!--
# BEGIN QUESTION
# name: q7b
# manual: true
# points: 1
# -->
# -
plt.hist(biased_simulations)
plt.title("Trump's Advantage in Pennsylvania over 100,000 Biased Simulations")
plt.xlabel("Proportion Advantage")
plt.ylabel("Count")
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# #### Part 3
#
# Compare the histogram you created in Q7.2 to that in Q6.4.
#
# <!--
# BEGIN QUESTION
# name: q7c
# manual: true
# points: 2
# -->
# -
# When looking at the two histograms, there seems to be higher frequencies of no advantages when we run simulations with the biased proportions. For the histogram in Q6.4 (no bias), it seems like more of the data is above 0.00, which indicates an advantage. Contrastingly, for the histogram in Q7.2 (yes bias), Trump's occurrences of advantages have decreased (height of bars are lower when Proportion Advantage is above 0.00), and occurrences of disadvantages have increased (there is more data on the left side of 0.00 Proportion Advantage and height of bars are higher when Proportion Advantage is below 0.00).
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# #### Part 4
#
# Now perform 100,000 simulations of all four states and return the proportion of these simulations that result in a Trump victory. This is the same fraction that you computed in Q6.6, but now using your biased samples.
#
# Give your answer as `proportion_trump_biased`.
#
# This number represents the chance that a sample biased 0.5% in Hillary Clinton's favor will correctly predict Trump's victory. You should observe that the chance is signficantly lower than with an unbiased sample, i.e. your answer in Q6.6.
#
# <!--
# BEGIN QUESTION
# name: q7d
# manual: false
# points: 1
# -->
# +
def trump_wins_biased(N):
florida = trump_advantage(draw_biased_state_sample(N, 'florida'))
michigan = trump_advantage(draw_biased_state_sample(N, 'michigan'))
pennsylvania = trump_advantage(draw_biased_state_sample(N, 'pennsylvania'))
wisconsin = trump_advantage(draw_biased_state_sample(N, 'wisconsin'))
if florida > 0 and michigan > 0:
return 1
if florida > 0 and pennsylvania > 0:
return 1
if florida > 0 and wisconsin > 0:
return 1
if michigan > 0 and pennsylvania > 0 and wisconsin > 0:
return 1
else:
return 0
def biased_wins(n):
wins = 0
for i in range(100000):
if trump_wins_biased(n) == 1:
wins += 1
return wins
proportion_trump_biased = biased_wins(1500)/100000
proportion_trump_biased
# + deletable=false editable=false
grader.check("q7d")
# -
# ## Further Study
#
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "q2c", "locked": true, "schema_version": 2, "solution": false}
# ### Question 8
#
# Would increasing the sample size have helped?
#
# #### Part 1
#
# Try a sample size of 5,000 and run 100,000 simulations of a sample with replacement. What proportion of the 100,000 times is Trump predicted to win the election in the unbiased setting? In the biased setting?
#
# Give your answers as `high_sample_size_unbiased_proportion_trump` and `high_sample_size_biased_proportion_trump`.
#
# <!--
# BEGIN QUESTION
# name: q8a
# manual: false
# points: 1
# -->
# -
high_sample_size_unbiased_proportion_trump = victory(5000)/100000
high_sample_size_biased_proportion_trump = biased_wins(5000)/100000
print(high_sample_size_unbiased_proportion_trump, high_sample_size_biased_proportion_trump)
# + deletable=false editable=false
grader.check("q8a")
# -
# #### Part 2
#
# What do your observations from Part 1 say about the impact of sample size
# on the sampling error and on the bias?
#
# Extra question for those who are curious: Just for fun, you might find it interesting to see what happens with even larger sample sizes (> 5000 voters) for both the unbiased and biased cases. Can you get them up to 99% success with sufficient large samples? How many? Why or why not? If you do this, include your observations in your answer.
# +
very_very_high_sample_size_unbiased_proportion_trump = victory(500000)/100000
very_very_high_sample_size_biased_proportion_trump = biased_wins(500000)/100000
print('Sample Size:500000 ', very_very_high_sample_size_unbiased_proportion_trump, very_very_high_sample_size_biased_proportion_trump)
very_high_sample_size_unbiased_proportion_trump = victory(15000)/100000
very_high_sample_size_biased_proportion_trump = biased_wins(15000)/100000
print('Sample Size:15000 ', very_high_sample_size_unbiased_proportion_trump, very_high_sample_size_biased_proportion_trump)
high_sample_size_unbiased_proportion_trump = victory(5000)/100000
high_sample_size_biased_proportion_trump = biased_wins(5000)/100000
print('Sample Size:5000 ', high_sample_size_unbiased_proportion_trump, high_sample_size_biased_proportion_trump)
small_sample_size_unbiased_proportion_trump = victory(1500)/100000
small_sample_size_biased_proportion_trump = biased_wins(1500)/100000
print('Sample Size:1500 ', small_sample_size_unbiased_proportion_trump, small_sample_size_biased_proportion_trump)
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# Write your answer in the cell below.
#
# <!--
# BEGIN QUESTION
# name: q8b
# manual: true
# points: 2
# -->
# -
# The output from above shows the results from having 500000 samples to 15000 samples to 5000 samples to 1500 samples for unbiased and biased Trump proportions. As we can see, increasing the sample size increases proportions favoring Trump (unbiased) to win, but it decreases proportions favoring Trump (biased) to win. From these observations, we can conclude that increasing the sample size reduces sampling error, and increases the effect of bias. Increasing the sample size also increased unbiased Trump's proportion of winning; in addition, when we increased sample size, we saw that the effects of bias were stronger because at a sample size of 500000, biased Trump's proportion for winning was almost 0, but at a sample size of 1500, biased Trump's proportion for winning was around .46
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# ### Question 9
#
# According to FiveThirtyEight: "... Polls of the November 2016 presidential election were about as accurate as polls of presidential elections have been on average since 1972."
#
# When the margin of victory may be relatively small as it was in 2016, why don't polling agencies simply gather significantly larger samples to bring this error close to zero?
#
# <!--
# BEGIN QUESTION
# name: q9
# manual: true
# points: 2
# -->
# -
# Polling agencies don't simply gather significantly larger samples to bring error close to zero because doing so will take much longer time and money, in addition to other factors polling agencies cannot control.
#
# As previously mentioned, voters changed their preferences a few days before the election. If a polling agency sampled about five million people, they would have no idea how many of these voters will change their preferences a few days before or even on the day of the election.
#
# Also, polling agencies may not have the best representation of the voting population. Since these surveys are voluntary, only those who are heavily interested in the election will respond to the survey. This group of respondents may only be college graduates, young people in the workforce, etc. Even if someone ends up voting, it does not necessarily mean that he/she will take part in a polling agency's survey.
# <!-- END QUESTION -->
#
#
#
# **Important**: To make sure the test cases run correctly, click `Kernel>Restart & Run All` and make sure all of the test cases are still passing.
# + [markdown] deletable=false editable=false
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false
grader.check_all()
# + [markdown] deletable=false editable=false
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zipfile for you to submit. **Please save before exporting!**
# + deletable=false editable=false
# Save your notebook first, then run this cell to export your submission.
grader.export()
# -
#
| 41,342 |
/Unit04 - GD/.ipynb_checkpoints/Gradient Descent-checkpoint.ipynb | e4e07dcc4375d49303c23ff9da6abe638b682598 | [] | no_license | J4RV13/IMLP343 | https://github.com/J4RV13/IMLP343 | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 63,126 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 梯度下降
# 梯度下降法的定義是:
#
# $w^{j+1}=w^j-\alpha\nabla E(w),j 為迭代次數,\alpha 為學習率。$
#
# 其中 $\nabla$ 就是梯度,一次微分的意思。<br>
# 若是單純的 $y=ax+b$ 的情況,一次微分就是斜率。<br>
# 只是在多變量的情境中,斜率這個名稱太狹隘,於是在數學上就稱為「梯度」。<br>
#
# 梯度的算法就是對每個變量作一次偏微分。
# 例如,給一函數為:$f(x,y)=x^2+2xy+y^2$。<br>
#
# 計算其梯度為:$\mathrm{\nabla}\ f\left(x_1,x_2\right)=\left(\frac{\partial f\left(x_1,x_2\right)}{\partial x_1},\ \frac{\partial f\left(x_1,x_2\right)}{\partial x_2}\right)=\left(2x_1+1,\ 2x_2+1\right)$ <br>
#
# 當 $(x_1, x_2)=(1, 2)$ 時,$\mathrm{\nabla}\ f\left(1,\ 2\right)=\left(3,\ 5\right)$。
# 上述結果用白話文來說,即 $x_1$ 方向上的斜率為 $3$;$x_2$ 方向上的斜率為 $5$ 的意思。
#
# 回過頭來看剛剛給的函數:$E(w)=w^2$,
# 我們這就來「手動」跑一次梯度下降法。
#
# 假定,第一次隨機初始化點位,$w=5$。
# 
# 在初始化點位 (5, 25) 上,經計算後的切線斜率,也就是梯度,為 +10,這會影響幾件事情:
#
# ### 1. 方向:將梯度的方向取負號,就是我們想要移動的方向。
# ### 2. 大小:由於學習率固定,因此梯度值愈大,每次移動的距離愈遠!
#
# 
#
# 
#
# ### 這個反覆迭代的過程會一直到終止條件出現為止,例如:
# #### 1. 迭代次數達到某個值。
# #### 2. 迭代後的 loss 值間的差異小於某個數。
# #### 3. 程式執行總時間限制。
# +
import numpy as np
import matplotlib.pyplot as plt
# 目標函數:y=x^2
def func(x): return np.square(x)
# 目標函數一階導數:dy/dx=2*x
def dfunc(x): return 2 * x
def GD(x_start, df, epochs, lr):
""" 梯度下降法。給定起始點與目標函數的一階導函數,求在epochs次反覆運算中x的更新值
:param x_start: x的起始點
:param df: 目標函數的一階導函數
:param epochs: 反覆運算週期
:param lr: 學習率
:return: x在每次反覆運算後的位置(包括起始點),長度為epochs+1
"""
xs = np.zeros(epochs+1)
x = x_start
xs[0] = x
for i in range(epochs):
dx = df(x)
# v表示x要改變的幅度
v = - dx * lr
x += v
xs[i+1] = x
return xs
# Main
# 起始權重
x_start = 10
# 執行週期數
epochs = 20
# 學習率
lr = 0.8
# 梯度下降法
x = GD(x_start, dfunc, epochs, lr=lr)
print (x)
# 輸出:[-5. -2. -0.8 -0.32 -0.128 -0.0512]
color = 'r'
#plt.plot(line_x, line_y, c='b')
from numpy import arange
t = arange(-6.0, 6.0, 0.01)
plt.plot(t, func(t), c='b')
plt.plot(x, func(x), c=color, label='lr={}'.format(lr))
plt.scatter(x, func(x), c=color, )
plt.legend()
plt.show()
# +
# Main
# 起始權重
x_start = 5
# 執行週期數
epochs = 15
# 學習率
lr = 0.1
# 梯度下降法
x = GD(x_start, dfunc, epochs, lr=lr)
print (x)
# 輸出:[-5. -2. -0.8 -0.32 -0.128 -0.0512]
color = 'r'
#plt.plot(line_x, line_y, c='b')
from numpy import arange
t = arange(-6.0, 6.0, 0.01)
plt.plot(t, func(t), c='b')
plt.plot(x, func(x), c=color, label='lr={}'.format(lr))
plt.scatter(x, func(x), c=color, )
plt.legend()
plt.show()
# -
| 2,846 |
/lesson_5/Classwork.ipynb | 1dfef1aa99938e3bfd1b0a878eeeff279756e1c3 | [] | no_license | sashasenna/pydat21 | https://github.com/sashasenna/pydat21 | 0 | 0 | null | 2021-04-11T17:55:56 | 2021-04-10T15:08:10 | null | Jupyter Notebook | false | false | .py | 32,378 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Про Поросенка
x = 0
y = 0
N = int(input('Сколько шагов? '))
for i in range(N):
order = input("Ход {}:".format(i))
order_parts = order.split()
direction = order_parts[0]
steps = int(order_parts[1])
if direction == 'Вверх':
y += steps
elif direction == 'Вниз':
y -= steps
elif direction == 'Влево':
x -= steps
else:
x += steps
print("Пришли в точку ({}, {})".format(x, y))
print("Могли дойти за {} шагов".format(abs(x) + abs(y)))
# +
# Про цифры и буквы
digits = 0
letters = 0
S = input()
for letter in S:
if letter.isdigit():
digits += 1
if letter.isalpha():
letters += 1
print("Цифр:", digits)
print("Букв:", letters)
# -
[i for i in range(10) if i % 3 == 0 or i % 5 == 0]
# ### Повторение
# +
# %%timeit
# project euler 1
S = 0
number = 0
while number < 1000:
if number % 3 == 0 or number % 5 == 0:
S += number
number += 1
# +
# %%timeit
S = 0
for number in range(1000):
if number % 3 == 0 or number % 5 == 0:
S += number
# -
# %%timeit
L = [i for i in range(1000) if i % 3 == 0 or i % 5 == 0]
# %%timeit
A = 3*int(999/3)*(1+int(999/3))/2 + 5*int(999/5)*(1+int(999/5))/2 - 15*int(999/15)*(1+int(999/15))/2
# ### Словари
# +
# списки
L = [1, 2, 3]
L1 = []
L2 = list("abcd")
L.append(4)
L.remove(4)
L[0]
del L[0]
L1 + L2
L1.sort()
# +
D = {}
print(type(D))
# -
# создаем словарь
D = {
"Россия": "Москва",
"Италия": "Рим",
"Украина": "Киев"
}
# Какая столица у России (Какое значение у словаря D по ключю "Россия")
D["Россия"]
# +
country = input('Введите страну:')
print("Столица страны {} - {}".format(country, D[country]))
# +
country = input('Введите страну:')
if country in D:
print("Столица страны {} - {}".format(country, D[country]))
else:
print('Такой страны нет?')
# -
print(D.keys())
print(D.values())
D["Венгрия"] = "Буддапешт"
D
D["Венгрия"] = "Будапешт"
D
A = {
"Америка": "Вашингтон",
"Канада": "Торонто"
}
# +
D.update(A)
D
# -
del D["Америка"]
D
for elem in [1, 2, 3, 4]:
print(elem)
for country in D: # D.keys()
print(country, D[country])
for capital in D.values():
print(capital)
for country, capital in D.items(): # for name, country in pythons
print(country, capital)
sorted(D)
list(D.items())
number_names = {
1: "Один",
1.1: "Один и еще чуть-чуть",
2: "Два"
}
number_names[1]
number_names[1.1]
forest = {
(0, 0): "Винни-Пух",
(10, 5): "Кролик",
(100, 100): "Осел",
}
# +
x = int(input())
y = int(input())
if (x, y) in forest:
print('Здесь живет {}'.format(forest[(x, y)]))
else:
print("Тут никто не живет")
# +
emails = ["A", "B", "A", "B", "A", "B", "C", "A", "E", "A"]
counter = {}
for email in emails:
if email in counter:
counter[email] = counter[email] + 1 # counter[email] += 1
else:
counter[email] = 1
print("Вижу {}, в словаре - {}".format(email, counter))
print(counter)
# -
value = counter.get("A")
print(value)
value = counter.get("sfsdf")
print(value, type(value))
counter.get("sdfsdf", 0)
# +
emails = ["A", "B", "A", "B", "A", "B", "C", "A", "E", "A"]
counter = {}
for email in emails:
counter[email] = counter.get(email, 0) + 1
print("Вижу {}, в словаре - {}".format(email, counter))
print(counter)
# +
names = input().split()
marks = input().split()
D = {}
for i in range(len(names)):
D[names[i]] = marks[i]
print(D)
# +
names = input().split()
marks = input().split()
D = dict(zip(names, marks))
print(D)
# -
# ### Обработка исключений
while True:
try:
income = int(input())
print(income * 0.13)
break
except:
print("Введите целое число")
# +
expression = input()
parts = expression.split()
A = int(parts[0])
sign = parts[1]
B = int(parts[2])
if sign == '+':
print(A + B)
elif sign == '-':
print(A - B)
elif sign == '*':
print(A * B)
elif sign == '/':
print(A / B)
# -
# 23 - 3
try:
expression = input()
parts = expression.split()
A = int(parts[0])
sign = parts[1]
B = int(parts[2])
if sign == '+':
print(A + B)
elif sign == '-':
print(A - B)
elif sign == '*':
print(A * B)
elif sign == '/':
print(A / B)
except:
print("Что то пошло не так")
# 23 - 3
try:
expression = input()
parts = expression.split()
A = int(parts[0])
sign = parts[1]
B = int(parts[2])
if sign == '+':
print(A + B)
elif sign == '-':
print(A - B)
elif sign == '*':
print(A * B)
elif sign == '/':
print(A / B)
except Exception as e:
print("Возникла ошибка", e)
# +
# 23 - 3
try:
expression = input()
parts = expression.split()
A = int(parts[0])
sign = parts[1]
B = int(parts[2])
if sign == '+':
print(A + B)
elif sign == '-':
print(A - B)
elif sign == '*':
print(A * B)
elif sign == '/':
print(A / B)
raise Exception("Неверная операция")
except ZeroDivisionError:
print("На ноль делить нельзя")
except ValueError:
print("Введите корректное выражение с пробелами")
except Exception as e:
print("Возникла ошибка", type(e), e)
# +
A = int(input())
if A > 10:
raise Exception("Да как вы смеете вводить число больше 10!!!")
print("Все ок")
# +
L = []
print(L[1])
# +
a = 3
b = 5
if a > b:
print(a)
else:
print(b)
# -
# ### Функции
# +
def say_hello(name):
print("Привет, {}!".format(name))
print("Рад вас видеть!")
say_hello("Василий")
# -
say_hello("Александр")
say_hello("Иван")
# +
def progression(step, last):
S = step*int(last/step)*(1+int(last/step))/2
return S
progression(3, 999)
# -
R = progression(5, 999)
print(R)
progression(15, 999)
progression(3, 999) + progression(5, 999) - progression(15, 999)
# +
A = int(input())
if A > 0:
B = 0
else:
B = 1
print(B)
# +
def F():
global B
A = 5
B += 4
print(A)
print(B)
F()
print(B)
print(A)
# -
| 6,363 |
/Partner_Assignment.ipynb | d7e6e2368432abf3cad145dd98bb19cf35439f3a | [] | no_license | jamesdiao/Dance-Partner-Assignment | https://github.com/jamesdiao/Dance-Partner-Assignment | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 21,258 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
### Declare variables, import packages
from random import *
from math import *
import csv
import numpy as np
from munkres import Munkres, print_matrix
generate = False
with open('cost_function.txt') as f:
content = f.readlines()
def cost_function(L, F):
return eval(content[0])
# +
### Get Cost Matrix
def random_ranking(numrow,numcol):
mat = [["O" for a in range(numcol)] for b in range(numrow)] # final combined preference matrix [numrow x numcol]
for row in range(numrow):
temp = []
for i in range(1, numcol + 1):
temp.append(i)
shuffle(temp)
mat[row] = temp
return np.array(mat)
def makelist(prefix, n):
values = []
for i in range(n):
values.append("%s%d" % (prefix,i))
return values
if (generate):
# Create a random set of preference for each leader, append into matrix1
seed(12345)
num_leaders = 8
num_follow = 12
matrix1 = random_ranking(num_leaders,num_follow)
matrix2 = random_ranking(num_follow,num_leaders)
leaderlist = dict(zip(range(numrow), makelist("L",numrow)))
followerlist = dict(zip(range(numcol), makelist("F",numcol)))
numrow = num_leaders
numcol = num_follow
else:
L_input = csv.reader(open("leader_matrix.csv","rb"),delimiter=',')
L_header = next(L_input)
L_header = [L_header[i] for i in range(1,len(L_header))]
matrix1 = np.array(list(L_input))[:, 1:].astype(np.integer)
F_input = csv.reader(open("follower_matrix.csv","rb"),delimiter=',')
F_header = next(F_input)
F_header = [F_header[i] for i in range(1,len(F_header))]
matrix2 = np.array(list(F_input))[:, 1:].astype(np.integer)
numrow = matrix1.shape[0]
numcol = matrix1.shape[1]
leaderlist = dict(zip(range(numrow), F_header))
followerlist = dict(zip(range(numcol), L_header))
matrix = cost_function(matrix1,matrix2.T)
matrix_original = matrix.copy()
# +
# Print Function
def printmatrix(matrix, dict1, dict2, sides):
left_bottom = np.array(np.c_[dict1.values(), matrix])
if (sides):
top = np.array([''] + dict2.values())[np.newaxis]
matrix = np.r_[top, left_bottom]
s = [[str(e) for e in row] for row in matrix]
lens = [max(map(len, col)) for col in zip(*s)]
fmt = '\t'.join('{{:{}}}'.format(x) for x in lens)
table = [fmt.format(*row) for row in s]
print '\n'.join(table)
print("\nPREFERENCE MATRIX FOR LEADERS (LEADERS LEFT X FOLLOWERS TOP)")
printmatrix(matrix1, leaderlist, followerlist, False)
print("\nPREFERENCE MATRIX FOR FOLLOWERS (FOLLOWERS LEFT X LEADERS TOP)")
printmatrix(matrix2, followerlist, leaderlist, False)
print("\nCOMBINED PREFERENCE MATRIX (LEADERS LEFT X FOLLOWERS TOP)")
printmatrix(matrix, leaderlist, followerlist, False)
# +
### Leader Scores
print("\nLEADER SCORES")
leader_scores = np.mean(matrix2, axis=1)
for i in np.argsort(leader_scores):
print '%s: %s' % (leaderlist.values()[i], round(leader_scores[i],2))
### Follower Scores
print("\nFOLLOWER SCORES")
follow_scores = np.mean(matrix1, axis=0)
for i in np.argsort(follow_scores):
print '%s: %s' % (followerlist.values()[i], round(follow_scores[i],2))
# -
m = Munkres()
indexes = m.compute(matrix)
print("\nFINAL PARTNER ASSIGNMENTS: (N[X] means Person N was given their Xth choice)")
print("------------------------------------------------------------")
csv_write = csv.writer(open('match_outputs.csv','wb'),delimiter = ',',quoting=csv.QUOTE_MINIMAL)
csv_write.writerow(['Leader_Name','Choice_Received','Follower_Name','Choice_Received','Penalty'])
total = 0
for row, col in indexes:
value = matrix_original[row][col]
total += value
csv_write.writerow([leaderlist[row], matrix1[row,col], followerlist[col], matrix2[col, row], value])
print '%s[%d] + %s[%d] >>> %d' % (leaderlist[row], matrix1[row,col], followerlist[col], matrix2[col, row], value)
print 'Total Penalty: %d' % total
csv_write.writerow(['\nTotal Penalty\n', total])
# STABLE MATCH ALGORITHM: FAVORS LEADERS
if numrow == numcol:
print("STABLE MATCH ALGORITHM:")
singlelist = [0 for a in range(numrow)] # 0=single
choicelist = [numrow + 1 for a in range(numcol)] # all high numbers
tempmatrix = matrix1.copy()
print('Starting State')
print(singlelist)
print(choicelist)
print
counter = 1
while sorted(singlelist)[0] == 0: # main algorithm loop, per round
print("\nROUND #%s" % counter)
for n in range(len(tempmatrix)): # For each suitor
if singlelist[n] == 0: # If they're single
p = np.argmin(tempmatrix[n])#tempmatrix[n].index(min(tempmatrix[n])) # col identifying their first choice
print("Suitor %s Proposes to Lady %s (His #%s choice)" % (n + 1, p + 1, min(tempmatrix[n])))
tempmatrix[n][p] = 99 # removes from consideration for next round
if choicelist[p] > matrix2[p][
n]: # hate of former suitor > hate of new suitor i.e. if you want to trade
if choicelist[p] != numrow + 1:
print("TRADE UP - Lady drops Suitor %s" % (1 + matrix2[p].tolist().index(choicelist[p])))
singlelist[matrix2[p].tolist().index(choicelist[p])] = 0 # set jinked suitor to single
singlelist[n] = min(tempmatrix[n]) - 1 # set new suitor to engaged
choicelist[p] = matrix2[p][n] # completes replacement
else:
print("REJECTION")
print(singlelist)
print(choicelist)
print
counter += 1
setsum = 0
print("STABLE MATCH COMPLETE")
print("N[X] means Person N was given their Xth choice")
print("------------------------------------------------------------")
for row in leaderlist:
followid = matrix1[row].tolist().index(singlelist[row])
print('%s[%s] + %s[%s] >>> %s' % (leaderlist[row], singlelist[row], followerlist[followid],
matrix2[followid,row], matrix_original[row,followid]))
setsum += matrix_original[row,followid]
print("PENALTY: " + str(setsum))
else:
print("SKIPPED: STABLE MATCH REQUIRES SQUARE COST MATRIX")
| 6,515 |
/zfel_pytorch/sase1d_run_input_part.ipynb | 9dacb80328d8e1f836caa51c6f6b9b7df47c2e0f | [] | no_license | xiaozhg/zfel_optim | https://github.com/xiaozhg/zfel_optim | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 453,251 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1.Import packages
import pandas as pd
import numpy as np
from numpy import genfromtxt
import matplotlib.pyplot as plt
# %matplotlib inline
import scipy.io
import random
import datetime
import time
#from zfel import sase1d_input_part
import sase1d_input_part
import torch
import torch.optim as optim
# 2.Input parameters
Nruns=1 # Number of runs
npart = 512 # n-macro-particles per bucket
s_steps = 31#200#31 # n-sample points along bunch length
z_steps = 20#200#20 # n-sample points along undulator
energy = 4313.34*1E6 # electron energy [eV]
eSpread = 0#1.0e-4 # relative rms energy spread [ ]
emitN = 1.2e-6 # normalized transverse emittance [m-rad]
currentMax = 3900 # peak current [Ampere]
beta = 26 # mean beta [meter]
unduPeriod = 0.03 # undulator period [meter]
#unduK = 3.5 # undulator parameter, K [ ]
unduK = np.ones(z_steps)*3.5 # tapered undulator parameter, K [ ]
if unduK.shape[0]!=z_steps:
print('Wrong! Number of steps should always be the same as z_steps')
unduL = 70#30 # length of undulator [meter]
radWavelength = 1.5e-9 # seed wavelength? [meter], used only in single-freuqency runs
dEdz = 0 # rate of relative energy gain or taper [keV/m], optimal~130
iopt = 'sase' # 'sase' or 'seeded'
P0 = 10000*0.0 # small seed input power [W]
constseed = 1 # whether we want to use constant random seed for reproducibility, 1 Yes, 0 No
particle_position=genfromtxt('./Inputs/particle_position.csv', delimiter=',') # or None
# particle information with positions in meter and eta,\
# if we want to load random particle positions and energy, then set None
hist_rule='square-root' # 'square-root' or 'sturges' or 'rice-rule' or 'self-design', number \
# of intervals to generate the histogram of eta value in a bucket
'''
Put input parameters into a inp_struct dict, for 1D FEL run
'''
inp_struct={'Nruns':Nruns,'npart':npart,'s_steps':s_steps,'z_steps':z_steps,'energy':energy,'eSpread':eSpread,\
'emitN':emitN,'currentMax':currentMax,'beta':beta,'unduPeriod':unduPeriod,'unduK':unduK,'unduL':\
unduL,'radWavelength':radWavelength,'dEdz':dEdz,'iopt':iopt,'P0':P0,'constseed':constseed,'particle_position':particle_position,'hist_rule':hist_rule}
# 3.1D FEL run
# Method 1: allow exploring all taper values
# +
# %load_ext autoreload
# %autoreload
unduK = np.ones(z_steps)*3.5 #np.arange(3.52,3.48,-0.04/(z_steps-1))# # tapered undulator parameter, K [ ]
if unduK.shape[0]!=z_steps:
print('Wrong! Number of steps should always be the same as z_steps')
unduK_1=torch.tensor(unduK[:int(z_steps/2)],dtype=torch.float64)
unduK_2=torch.tensor(unduK[int(z_steps/2):],dtype=torch.float64,requires_grad=True)
#unduK = torch.tensor(unduK, dtype=torch.float64, requires_grad=True)
#undu_K=torch.cat((unduK_1,unduK_2))
#params_K = unduK
#learning_rate = 5e-21
#optimizer = optim.SGD([unduK_2], lr=learning_rate)
lr = 5e-21*torch.arange(unduK_2.shape[0])**4
n_epochs = 3000
his_FEL_power=[]
# -
def training_loop(n_epochs, lr, unduK_1, unduK_2, his_FEL_power):
for epoch in range(1, n_epochs + 1):
print('epochs:',epoch)
'''
Use sase function in sase1d.py to run 1D FEL
'''
#with torch.autograd.set_detect_anomaly(True):
#inp_struct['unduK']=params_K
if unduK_2.grad is not None:
unduK_2.grad.zero_()
undu_K=torch.cat((unduK_1,unduK_2))
inp_struct['unduK']=undu_K
z,power_z,s,power_s,rho,detune,field,\
field_s,gainLength,resWavelength,\
thet_out,eta_out,bunching,spectrum,freq,\
Ns,history=sase1d_input_part.sase(inp_struct)
power_z[-1].backward()
print('gradient',unduK_2.grad)
unduK_2=(unduK_2 - lr * \
unduK_2.grad).detach().requires_grad_()
#optimizer.zero_grad()
#power_z[-1].backward()
#optimizer.step()
#print('gradient',unduK_2.grad)
his_FEL_power.append(power_z[-1].detach().numpy())
undu_K=torch.cat((unduK_1,unduK_2))
return undu_K, his_FEL_power
undu_K, his_FEL_power= training_loop(
n_epochs = n_epochs,
lr = lr,
#optimizer = optimizer,
unduK_1 = unduK_1,
unduK_2 = unduK_2,
his_FEL_power = his_FEL_power)
plt.figure()
plt.plot(undu_K.detach().numpy())
plt.title('final taper value')
plt.figure()
plt.plot(his_FEL_power,'.-')
plt.title('output FEL power vs. epochs')
# Method 2: fit to quadratic taper
# +
# %load_ext autoreload
# %autoreload
#params_K=torch.tensor([2.3,-0.001],requires_grad=True)
const_K=torch.tensor(2.3,requires_grad=True)
param_K=torch.tensor(-0.001,requires_grad=True)
lr_const = 1e-21 #1e-17
lr_param = 0#1e-21
#optimizer = optim.SGD([params_K], lr=learning_rate)
n_epochs = 100
his_FEL_power=[]
unduK_1=const_K*torch.ones(int(z_steps/2))
unduK_2=const_K+param_K*torch.arange(z_steps-int(z_steps/2))**2
undu_K=torch.cat((unduK_1,unduK_2))
plt.plot(undu_K.detach().numpy())
# -
def training_loop(n_epochs, const_K, param_K, lr_const, lr_param, z_steps, his_FEL_power):
for epoch in range(1, n_epochs + 1):
print('epochs:',epoch)
'''
Use sase function in sase1d.py to run 1D FEL
'''
if const_K.grad is not None:
const_K.grad.zero_()
if param_K.grad is not None:
param_K.grad.zero_()
#sase1d_input_part.sase().zero_grad()
#with torch.autograd.set_detect_anomaly(True):
#inp_struct['unduK']=params_K
#if unduK_2.grad is not None:
# unduK_2.grad.zero_()
unduK_1=const_K*torch.ones(int(z_steps/2))
unduK_2=const_K+param_K*torch.arange(z_steps-int(z_steps/2))**2
undu_K=torch.cat((unduK_1,unduK_2))
inp_struct['unduK']=undu_K
z,power_z,s,power_s,rho,detune,field,\
field_s,gainLength,resWavelength,\
thet_out,eta_out,bunching,spectrum,freq,\
Ns,history=sase1d_input_part.sase(inp_struct)
#optimizer = optim.SGD([params_K], lr=lr)
#optimizer.zero_grad()
if epoch==1:
(-power_z[-1]).backward(retain_graph=True)
else:
(-power_z[-1]).backward()
print('gradient',const_K.grad,param_K.grad)
const_K=(const_K + lr_const * const_K.grad).detach().requires_grad_()
param_K=(param_K + lr_param * param_K.grad).detach().requires_grad_()
#optimizer.step()
print('power',power_z[-1])
his_FEL_power.append(power_z[-1].detach().numpy())
unduK_1=const_K*torch.ones(int(z_steps/2))
unduK_2=const_K+param_K*torch.arange(z_steps-int(z_steps/2))**2
undu_K=torch.cat((unduK_1,unduK_2))
return undu_K, his_FEL_power
undu_K, his_FEL_power= training_loop(
n_epochs = n_epochs,
const_K = const_K,
param_K = param_K,
lr_const = lr_const,
lr_param = lr_param,
z_steps = z_steps,
his_FEL_power = his_FEL_power)
plt.figure()
plt.plot(undu_K.detach().numpy())
plt.title('final taper value')
plt.figure()
plt.plot(his_FEL_power,'.-')
plt.title('output FEL power vs. epochs')
hes: you should easily be able to achieve over 55% classification accuracy on the test set; our best model achieves about 60% classification accuracy.
print(X_train_feats.shape)
# +
from cs231n.classifiers.neural_net import TwoLayerNet
input_dim = X_train_feats.shape[1]
hidden_dim = 500
num_classes = 10
net = TwoLayerNet(input_dim, hidden_dim, num_classes)
best_net = None
################################################################################
# TODO: Train a two-layer neural network on image features. You may want to #
# cross-validate various parameters as in previous sections. Store your best #
# model in the best_net variable. #
################################################################################
pass
################################################################################
# END OF YOUR CODE #
################################################################################
# +
# Run your neural net classifier on the test set. You should be able to
# get more than 55% accuracy.
test_acc = (net.predict(X_test_feats) == y_test).mean()
print(test_acc)
# -
# # Bonus: Design your own features!
#
# You have seen that simple image features can improve classification performance. So far we have tried HOG and color histograms, but other types of features may be able to achieve even better classification performance.
#
# For bonus points, design and implement a new type of feature and use it for image classification on CIFAR-10. Explain how your feature works and why you expect it to be useful for image classification. Implement it in this notebook, cross-validate any hyperparameters, and compare its performance to the HOG + Color histogram baseline.
# # Bonus: Do something extra!
# Use the material and code we have presented in this assignment to do something interesting. Was there another question we should have asked? Did any cool ideas pop into your head as you were working on the assignment? This is your chance to show off!
vcard"}).text.replace("By ",""))
dates.append(post_soup.find('p',{'class':"topic single-topic"}))
# the first four articles
articles[:4]
# -
# This notebook was written for the Erdős Institute Cőde Data Science Boot Camp by Matthew Osborne, Ph. D., 2021.
#
# Redistribution of the material contained in this repository is conditional on acknowledgement of Matthew Tyler Osborne, Ph.D.'s original authorship and sponsorship of the Erdős Institute as subject to the license (see License.md)
| 10,390 |
/Chapter03/Activity03.ipynb | c0f6c53089db3fe75069dcdbcec329266e4fd67b | [
"MIT"
] | permissive | mcne65/Applied-Deep-Learning-with-Keras | https://github.com/mcne65/Applied-Deep-Learning-with-Keras | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 590,413 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: py3.7
# language: python
# name: py3.7
# ---
# # Activity 3
# import required packages from Keras
from keras.models import Sequential
from keras.layers import Dense, Activation
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
# import required packages for plotting
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
import matplotlib.patches as mpatches
# import the function for plotting decision boundary
from utils import plot_decision_boundary
# define a seed for random number generator so the result will be reproducible
seed = 1
# load the dataset, print the shapes of input and output and the number of examples
feats = pd.read_csv('data/outlier_feats.csv')
target = pd.read_csv('data/outlier_target.csv')
print("X size = ", feats.shape)
print("Y size = ", target.shape)
print("Number of examples = ", feats.shape[0])
# Plot the features and target
# +
# changing the size of the plots
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
class_1=plt.scatter(feats.loc[target['Class']==0,'feature1'], feats.loc[target['Class']==0,'feature2'], c="red", s=40, edgecolor='k')
class_2=plt.scatter(feats.loc[target['Class']==1,'feature1'], feats.loc[target['Class']==1,'feature2'], c="blue", s=40, edgecolor='k')
plt.legend((class_1, class_2),('Fail','Pass'))
plt.xlabel('Feature 1')
plt.ylabel('Feature 2')
# +
# Logistic Regression model
np.random.seed(seed)
model = Sequential()
model.add(Dense(1, activation='sigmoid', input_dim=2))
model.compile(optimizer='sgd', loss='binary_crossentropy')
# train the model for 100 epoches and batch size 5
model.fit(feats, target, batch_size=5, epochs=100, verbose=1, validation_split=0.2)
# -
# Plot the decision boundary
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plot_decision_boundary(lambda x: model.predict(x), feats, target)
plt.title("Logistic Regression")
# Evaluate the loss and accuracy on the test dataset
# Neural network with hidden layer size = 3
np.random.seed(seed)
model = Sequential()
model.add(Dense(3, activation='relu', input_dim=2))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='sgd', loss='binary_crossentropy')
# train the model for 200 epoches and batch size 5
model.fit(feats, target, batch_size=5, epochs=200, verbose=1, validation_split=0.2)
# Plot the decision boundary
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plot_decision_boundary(lambda x: model.predict(x), feats, target)
plt.title("Decision Boundary for Neural Network with hidden layer size 3")
# Create a neural network with hidden layer of size 6
np.random.seed(seed)
model = Sequential()
model.add(Dense(6, activation='relu', input_dim=2))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='sgd', loss='binary_crossentropy')
# train the model for 400 epoches
model.fit(feats, target, batch_size=5, epochs=400, verbose=1, validation_split=0.2)
# Plot the decision boundary
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plot_decision_boundary(lambda x: model.predict(x), feats, target)
plt.title("Decision Boundary for Neural Network with hidden layer size 6")
# Create a neural network with hidden layer of size 3 and tanh activation function
# Neural network with hidden layer size = 3 with tanh activation function
np.random.seed(seed)
model = Sequential()
model.add(Dense(3, activation='tanh', input_dim=2))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='sgd', loss='binary_crossentropy')
# train the model for 200 epoches and batch size 5
model.fit(feats, target, batch_size=5, epochs=200, verbose=1, validation_split=0.2)
# Plotting the decision boundary on the trainingdataset
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plot_decision_boundary(lambda x: model.predict(x), feats, target)
plt.title("Decision Boundary for Neural Network with hidden layer size 3")
# Neural network with hidden layer size = 6 with tanh activation function
np.random.seed(seed)
model = Sequential()
model.add(Dense(6, activation='tanh', input_dim=2))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='sgd', loss='binary_crossentropy')
# train the model for 400 epoches
model.fit(feats, target, batch_size=5, epochs=400, verbose=1, validation_split=0.2)
# Plot the decision boundary
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plot_decision_boundary(lambda x: model.predict(x), feats, target)
plt.title("Decision Boundary for Neural Network with hidden layer size 6")
ript
# urls = Array.from(document.querySelectorAll('.rg_di .rg_meta')).map(el=>JSON.parse(el.textContent).ou);
# window.open('data:text/csv;charset=utf-8,' + escape(urls.join('\n')));
# ```
# + [markdown] colab_type="text" id="MfMZFTXErnym"
# ### Create directory and upload urls file into your server
# + [markdown] colab_type="text" id="f5k3EqX4rnyo"
# Choose an appropriate name for your labeled images. You can run these steps multiple times to create different labels.
# + colab={} colab_type="code" id="tBiGXbUhrnyq"
folder = 'fire'
file = 'fire_urls.csv'
# + colab={} colab_type="code" id="5zQX7JEdrnyx"
folder = 'hurricane'
file = 'hurricane_urls.csv'
# + colab={} colab_type="code" id="z1ayfki8rny4"
folder = 'flooding'
file = 'flooding_urls.csv'
# + colab={} colab_type="code" id="TjDdwj7ry7kp"
folder = 'earthquake'
file = 'earthquake_urls.csv'
# + colab={} colab_type="code" id="92EeG78ftcas"
# folder = 'keanu'
# file = 'Keanu.csv'
# + colab={} colab_type="code" id="7TyPjLT9tpUJ"
# folder = 'danny'
# file = 'Danny.csv'
# + colab={} colab_type="code" id="awbAiQ0Ducq1"
# folder = 'stan'
# file = 'Stan.csv'
# + [markdown] colab_type="text" id="AuvONEIArny_"
# You will need to run this cell once per each category.
# + colab={} colab_type="code" id="ueNhbeQJrnzC"
path = Path('data/disasters')
dest = path/folder
dest.mkdir(parents=True, exist_ok=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 610, "status": "ok", "timestamp": 1573480931436, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 300} id="8wUfA-HLrnzI" outputId="d1371f60-d186-425d-de82-8cab4a611642"
path.ls()
# + [markdown] colab_type="text" id="0cpjvh5KrnzT"
# Finally, upload your urls file. You just need to press 'Upload' in your working directory and select your file, then click 'Upload' for each of the displayed files.
#
# 
# + [markdown] colab_type="text" id="OXjKbSBjrnzV"
# ## Download images
# + [markdown] colab_type="text" id="A2UdH5MSrnzX"
# Now you will need to download your images from their respective urls.
#
# fast.ai has a function that allows you to do just that. You just have to specify the urls filename as well as the destination folder and this function will download and save all images that can be opened. If they have some problem in being opened, they will not be saved.
#
# Let's download our images! Notice you can choose a maximum number of images to be downloaded. In this case we will not download all the urls.
#
# You will need to run this line once for every category.
# + colab={} colab_type="code" id="Mg0kOVi7rnzZ"
classes = ['earthquake', 'fire', 'flooding', 'hurricane']
# + colab={"base_uri": "https://localhost:8080/", "height": 320} colab_type="code" executionInfo={"elapsed": 655, "status": "error", "timestamp": 1573454837079, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 300} id="nZGuNvparnzh" outputId="0882f7b6-7ecb-4dc5-e598-ed95c9b76d27"
dest = path/folder
download_images(path/folder/file, dest, max_pics=500)
# + colab={} colab_type="code" id="8w4i2axZrnzs"
# If you have problems download, try with `max_workers=0` to see exceptions:
download_images(path/folder/file, dest, max_pics=20, max_workers=0)
# + [markdown] colab_type="text" id="sZ4dh2r8rnz3"
# Then we can remove any images that can't be opened:
# + colab={} colab_type="code" id="E_Etv3mqrnz5"
for c in classes:
print(c)
verify_images(path/c, delete=True, max_size=500)
# + [markdown] colab_type="text" id="79m0yhkJrn0A"
# ## View data
# + colab={} colab_type="code" id="UPBkedIYrn0B"
np.random.seed(42)
data = ImageDataBunch.from_folder(path, train=".", valid_pct=0.2,
ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
# + colab={} colab_type="code" id="RlPRxlF3rn0H"
# If you already cleaned your data, run this cell instead of the one before
# np.random.seed(42)
# data = ImageDataBunch.from_csv(path, folder=".", valid_pct=0.2, csv_labels='cleaned.csv',
# ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
# + [markdown] colab_type="text" id="GrxArUM5rn0M"
# Good! Let's take a look at some of our pictures then.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 533, "status": "ok", "timestamp": 1571924170846, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="CziTXe2frn0O" outputId="9aaf4efc-f972-42db-d7a5-c9fcd89e5d01"
data.classes
# + colab={} colab_type="code" id="RKl2SL4Ern0Z"
data.show_batch(rows=3, figsize=(7,8))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 500, "status": "ok", "timestamp": 1571924194424, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="Fv86w0ubrn0h" outputId="031ca67f-f667-4901-d697-8cf52e800bad"
data.classes, data.c, len(data.train_ds), len(data.valid_ds)
# + [markdown] colab_type="text" id="X1N7_Y7xrn0n"
# ## Train model
# + colab={} colab_type="code" id="CYLm7lQkrn0p"
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + colab={"base_uri": "https://localhost:8080/", "height": 412} colab_type="code" executionInfo={"elapsed": 343, "status": "error", "timestamp": 1571934576184, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="iWsIVGRqrn0u" outputId="ab28ec7b-14a3-4257-febf-a1a9cea9ae16"
learn.fit_one_cycle(4)
# + colab={} colab_type="code" id="62TnNvUTrn01"
learn.save('stage-1')
# + colab={} colab_type="code" id="6QoLAXUurn06"
learn.unfreeze()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 94563, "status": "ok", "timestamp": 1571924425305, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="4Xq71T8lrn0_" outputId="28ae60dd-8fc1-4ef1-a3a0-025fa2372399"
learn.lr_find()
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" executionInfo={"elapsed": 1062, "status": "ok", "timestamp": 1571924430115, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="nMU6pQzjrn1H" outputId="e5d6587f-f69f-451d-9aeb-3d95febef717"
learn.recorder.plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 307} colab_type="code" executionInfo={"elapsed": 480, "status": "error", "timestamp": 1571934505269, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="GAqF0CTgrn1N" outputId="78cc38fe-5960-46b6-8a85-e443e5da66b7"
learn.fit_one_cycle(1, max_lr=slice(3e-5,3e-4))
# + [markdown] colab_type="text" id="NGjtqxTxzafw"
# My data set must be pretty obvious if it gets a 3% error on the first try and couldn't impove on the second time around. Perhaps next time I should try to use a data set with more variation.
# + colab={} colab_type="code" id="TBe19aqPrn1Y"
learn.save('stage-2')
# + [markdown] colab_type="text" id="6g1v3YuKrn1e"
# ## Interpretation
# + colab={} colab_type="code" id="OPQ3VFVXrn1f"
learn.load('stage-2');
# + colab={"base_uri": "https://localhost:8080/", "height": 545} colab_type="code" executionInfo={"elapsed": 29, "status": "error", "timestamp": 1571934696505, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="I6K1Q4wQrn1n" outputId="50224874-329a-497a-82e6-c3b8d0d55678"
interp = ClassificationInterpretation.from_learner(learn)
# + colab={"base_uri": "https://localhost:8080/", "height": 311} colab_type="code" executionInfo={"elapsed": 793, "status": "ok", "timestamp": 1571934388238, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="9vdGc4bYrn1s" outputId="de068c9f-ed23-423c-e5c5-a758434d89dc"
interp.plot_confusion_matrix()
# + [markdown] colab_type="text" id="GXucekSkytrI"
# The most confused images are with Danny. He has many images with other people and has lived long enough to have pictures with his hair black or white. He also has pictures with or withour wrinkles, so that could be why he gets mixed up so much.
# + colab={"base_uri": "https://localhost:8080/", "height": 787} colab_type="code" executionInfo={"elapsed": 2799, "status": "ok", "timestamp": 1571934399284, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="BImAijJsyan1" outputId="35ffbcb1-3092-4923-e0c1-ff38e2ccc20b"
interp.plot_top_losses(9, figsize=(12,12))
# + [markdown] colab_type="text" id="OSp0EG7PzsJ0"
# A few of these images have several people which could mess things up. One of the Danny pictures is without his glasses, so that could be why that one is here. The red spots are focused on foreheads. Young Keanu and Danny both have big foreheads. There is a picture of a woman, so the NN is like wtf is this. Another picture has Keanu in a hat whose shadow kind of obscures his face.
# + [markdown] colab_type="text" id="bZZMX_N_rn1z"
# ## Cleaning Up
#
# Some of our top losses aren't due to bad performance by our model. There are images in our data set that shouldn't be.
#
# Using the `ImageCleaner` widget from `fastai.widgets` we can prune our top losses, removing photos that don't belong.
# + colab={} colab_type="code" id="cCu2Avm8rn11"
from fastai.widgets import *
# + [markdown] colab_type="text" id="gW4iEypjrn15"
# First we need to get the file paths from our top_losses. We can do this with `.from_toplosses`. We then feed the top losses indexes and corresponding dataset to `ImageCleaner`.
#
# Notice that the widget will not delete images directly from disk but it will create a new csv file `cleaned.csv` from where you can create a new ImageDataBunch with the corrected labels to continue training your model.
# + [markdown] colab_type="text" id="2XDJc8Ptrn16"
# In order to clean the entire set of images, we need to create a new dataset without the split. The video lecture demostrated the use of the `ds_type` param which no longer has any effect. See [the thread](https://forums.fast.ai/t/duplicate-widget/30975/10) for more details.
# + colab={} colab_type="code" id="JuN2b3Morn17"
db = (ImageList.from_folder(path)
.no_split()
.label_from_folder()
.transform(get_transforms(), size=224)
.databunch()
)
# + colab={} colab_type="code" id="RI3wHmbbrn2G"
# If you already cleaned your data using indexes from `from_toplosses`,
# run this cell instead of the one before to proceed with removing duplicates.
# Otherwise all the results of the previous step would be overwritten by
# the new run of `ImageCleaner`.
# db = (ImageList.from_csv(path, 'cleaned.csv', folder='.')
# .no_split()
# .label_from_df()
# .transform(get_transforms(), size=224)
# .databunch()
# )
# + [markdown] colab_type="text" id="PMVl4N21rn2Z"
# Then we create a new learner to use our new databunch with all the images.
# + colab={} colab_type="code" id="iOUDWzjprn2k"
learn_cln = cnn_learner(db, models.resnet34, metrics=error_rate)
learn_cln.load('stage-2');
# + colab={} colab_type="code" id="RS1TNMjyrn2p"
ds, idxs = DatasetFormatter().from_toplosses(learn_cln)
# + [markdown] colab_type="text" id="iGh18w2xrn2t"
# Make sure you're running this notebook in Jupyter Notebook, not Jupyter Lab. That is accessible via [/tree](/tree), not [/lab](/lab). Running the `ImageCleaner` widget in Jupyter Lab is [not currently supported](https://github.com/fastai/fastai/issues/1539).
# + colab={} colab_type="code" id="IrTk8DOTrn2u" outputId="286ac465-6c35-40a1-f0d9-9ca40e8b851b"
ImageCleaner(ds, idxs, path)
# + [markdown] colab_type="text" id="L24KWqd-rn22"
# Flag photos for deletion by clicking 'Delete'. Then click 'Next Batch' to delete flagged photos and keep the rest in that row. `ImageCleaner` will show you a new row of images until there are no more to show. In this case, the widget will show you images until there are none left from `top_losses.ImageCleaner(ds, idxs)`
# + [markdown] colab_type="text" id="t38eM0Rfrn24"
# You can also find duplicates in your dataset and delete them! To do this, you need to run `.from_similars` to get the potential duplicates' ids and then run `ImageCleaner` with `duplicates=True`. The API works in a similar way as with misclassified images: just choose the ones you want to delete and click 'Next Batch' until there are no more images left.
# + [markdown] colab_type="text" id="_CjRBtuhrn26"
# Make sure to recreate the databunch and `learn_cln` from the `cleaned.csv` file. Otherwise the file would be overwritten from scratch, loosing all the results from cleaning the data from toplosses.
# + colab={} colab_type="code" id="YJtjWtZIrn3B" outputId="1d7fa79a-d397-4bba-829d-33783eed7fb0"
ds, idxs = DatasetFormatter().from_similars(learn_cln)
# + colab={} colab_type="code" id="vmAKo0Msrn3H" outputId="d937a2d4-83e8-409c-a4fb-2acb6d678ef1"
ImageCleaner(ds, idxs, path, duplicates=True)
# + [markdown] colab_type="text" id="5qfJfFcvrn3N"
# Remember to recreate your ImageDataBunch from your `cleaned.csv` to include the changes you made in your data!
# + [markdown] colab_type="text" id="boCBVCq4rn3P"
# ## Putting your model in production
# + [markdown] colab_type="text" id="r359gCvCrn3R"
# First thing first, let's export the content of our `Learner` object for production:
# + colab={} colab_type="code" id="i17-vA6Trn3U"
learn.export()
# + [markdown] colab_type="text" id="qYREHkL4rn3Y"
# This will create a file named 'export.pkl' in the directory where we were working that contains everything we need to deploy our model (the model, the weights but also some metadata like the classes or the transforms/normalization used).
# + [markdown] colab_type="text" id="6H2zrz1Lrn3e"
# You probably want to use CPU for inference, except at massive scale (and you almost certainly don't need to train in real-time). If you don't have a GPU that happens automatically. You can test your model on CPU like so:
# + colab={} colab_type="code" id="5N9wYGXYrn3f"
defaults.device = torch.device('cpu')
# + colab={} colab_type="code" id="ATQp3CXIrn3i"
img = open_image(path/'flooding'/'00000121.jpg')
img
# + colab={} colab_type="code" id="OVPDuZBkERgA"
path = Path('.')
# + [markdown] colab_type="text" id="pil1lp4nrn3o"
# We create our `Learner` in production enviromnent like this, jsut make sure that `path` contains the file 'export.pkl' from before.
# + colab={} colab_type="code" id="WNumjBlArn3p"
learn = load_learner(path)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 575, "status": "ok", "timestamp": 1571934731163, "user": {"displayName": "Connor Grimberg", "photoUrl": "", "userId": "01720205661219630631"}, "user_tz": 240} id="Zuya2rnxrn3u" outputId="fec1cf45-e597-42fe-c871-98f46cb0e809"
pred_class,pred_idx,outputs = learn.predict(img)
pred_class
# + [markdown] colab_type="text" id="9zV58puzrn32"
# So you might create a route something like this ([thanks](https://github.com/simonw/cougar-or-not) to Simon Willison for the structure of this code):
#
# ```python
# @app.route("/classify-url", methods=["GET"])
# async def classify_url(request):
# bytes = await get_bytes(request.query_params["url"])
# img = open_image(BytesIO(bytes))
# _,_,losses = learner.predict(img)
# return JSONResponse({
# "predictions": sorted(
# zip(cat_learner.data.classes, map(float, losses)),
# key=lambda p: p[1],
# reverse=True
# )
# })
# ```
#
# (This example is for the [Starlette](https://www.starlette.io/) web app toolkit.)
# + [markdown] colab_type="text" id="jZUIdjvyrn35"
# ## Things that can go wrong
# + [markdown] colab_type="text" id="6aXjND_rrn36"
# - Most of the time things will train fine with the defaults
# - There's not much you really need to tune (despite what you've heard!)
# - Most likely are
# - Learning rate
# - Number of epochs
# + [markdown] colab_type="text" id="lk3Kndl9rn37"
# ### Learning rate (LR) too high
# + colab={} colab_type="code" id="bTIZHG3rrn38"
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + colab={} colab_type="code" id="1AQCjxW4rn4B" outputId="c5415026-2d1a-4abf-f861-2cace26afa20"
learn.fit_one_cycle(1, max_lr=0.5)
# + [markdown] colab_type="text" id="FEEuLL4Jrn4J"
# ### Learning rate (LR) too low
# + colab={} colab_type="code" id="qYxARJwIrn4K"
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + [markdown] colab_type="text" id="_lBp44L9rn4Q"
# Previously we had this result:
#
# ```
# Total time: 00:57
# epoch train_loss valid_loss error_rate
# 1 1.030236 0.179226 0.028369 (00:14)
# 2 0.561508 0.055464 0.014184 (00:13)
# 3 0.396103 0.053801 0.014184 (00:13)
# 4 0.316883 0.050197 0.021277 (00:15)
# ```
# + colab={} colab_type="code" id="4PsJt76yrn4R" outputId="6b247781-6f09-4d5e-bb9c-2a8c17d947d1"
learn.fit_one_cycle(5, max_lr=1e-5)
# + colab={} colab_type="code" id="C9dqz72lrn4X" outputId="bea7b01d-dc49-4ed7-9d55-fcadde6dd614"
learn.recorder.plot_losses()
# + [markdown] colab_type="text" id="QFOkQbDHrn4b"
# As well as taking a really long time, it's getting too many looks at each image, so may overfit.
# + [markdown] colab_type="text" id="SERSGNUgrn4c"
# ### Too few epochs
# + colab={} colab_type="code" id="it00A5ZErn4d"
learn = cnn_learner(data, models.resnet34, metrics=error_rate, pretrained=False)
# + colab={} colab_type="code" id="N4wjijz0rn4g" outputId="b37716ec-3809-4c8e-a97e-9eac08cc04ed"
learn.fit_one_cycle(1)
# + [markdown] colab_type="text" id="6J4EPoqjrn4l"
# ### Too many epochs
# + colab={} colab_type="code" id="svT1s13Yrn4m"
np.random.seed(42)
data = ImageDataBunch.from_folder(path, train=".", valid_pct=0.9, bs=32,
ds_tfms=get_transforms(do_flip=False, max_rotate=0, max_zoom=1, max_lighting=0, max_warp=0
),size=224, num_workers=4).normalize(imagenet_stats)
# + colab={} colab_type="code" id="U3xQesHPrn4p"
learn = cnn_learner(data, models.resnet50, metrics=error_rate, ps=0, wd=0)
learn.unfreeze()
# + colab={} colab_type="code" id="SS_BrM8Lrn4t" outputId="762b2d11-5a2c-4755-8b89-d981f7243e62"
learn.fit_one_cycle(40, slice(1e-6,1e-4))
}$$
# +
# GRADED FUNCTION: L2
def L2(yhat, y):
"""
Arguments:
yhat -- vector of size m (predicted labels)
y -- vector of size m (true labels)
Returns:
loss -- the value of the L2 loss function defined above
"""
### START CODE HERE ### (≈ 1 line of code)
loss = np.sum(np.dot(y-yhat,y-yhat))
### END CODE HERE ###
return loss
# -
yhat = np.array([.9, 0.2, 0.1, .4, .9])
y = np.array([1, 0, 0, 1, 1])
print("L2 = " + str(L2(yhat,y)))
# **Expected Output**:
# <table style="width:20%">
# <tr>
# <td> **L2** </td>
# <td> 0.43 </td>
# </tr>
# </table>
# Congratulations on completing this assignment. We hope that this little warm-up exercise helps you in the future assignments, which will be more exciting and interesting!
# <font color='blue'>
# **What to remember:**
# - Vectorization is very important in deep learning. It provides computational efficiency and clarity.
# - You have reviewed the L1 and L2 loss.
# - You are familiar with many numpy functions such as np.sum, np.dot, np.multiply, np.maximum, etc...
| 24,659 |
/Jupyter y Apuntes/C13 - Redes Neuronales/Redes Neuronales.ipynb | 0bd926a05ec0847bd447e460907d7cf0a456a465 | [] | no_license | ING559/Syllabus-2020-2 | https://github.com/ING559/Syllabus-2020-2 | 12 | 19 | null | null | null | null | Jupyter Notebook | false | false | .py | 242,587 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Redes Neuronales
#
# En este _notebook_ vamos a aprender a trabajar con Redes Neuronales, que es la técnica más usada en la actualidad para resolver problemas del área de _Machine Learning_. Para esto, vamos a usar Tensorflow y Keras para armar una red neuronal que usa la arquitectura _Multi Layer Perceptron_, que es uno de los modelos sencillo, pero muy útil. Así, vamos a aprender a:
#
# - Instanciar una red neuronal.
# - Agregar _hidden layers_ con un número determinado de neuronas.
# - Definir las funciones de activación de las capas.
# - Entrenar una red neuronal, definiendo el tamaño de los _batches_ de entrenamiento y el número de _epochs_.
# - Hacer regresión con una Red Neuronal.
#
# Para esto necesitamos tener instalado Tensorflow. Esta es una librería que nos permite definir grafos de computo. Nosotros vamos a usar una interfaz de alto nivel sobre esta librería llamada _Keras_, que nos permite definir Redes Neuronales de forma muy sencilla. Para instalar Tensorflow se puede hacer con `pip`:
#
# ```
# pip install tensorflow
# ```
#
# Ahora vamos a entrenar una red neuronal sobre el _dataset Iris_.
# +
from tensorflow import keras
from tensorflow.keras import layers
# Siempre es buena práctica reiniciar el modelo
keras.backend.clear_session()
# Para tener resultados replicables descomentar las líneas de abajo
# np.random.seed(42)
# tf.random.set_seed(42)
# Instanciamos un modelo "Secuencial", que corresponde a una secuencia de capas
model = keras.Sequential(
[
layers.Dense(5, activation='relu', input_shape=(2,), name='hidden_layer_1'),
layers.Dense(10, activation='relu', name='hidden_layer_2'),
layers.Dense(3, activation='softmax', name='output_layer'),
]
)
# Declaramos dos hidden layers: la primera con 5 neuronas y la segunda 10 neuronas cada una,
# en la primera capa declaramos el tamaño del input y en la última capa declaramos una
# una activación softmax para que el output represente una probabilidad
# Obtenemos un resumen del modelo
model.summary()
# -
# Vemos que al indicar el resumen del modelo, el número de parámetros calza según lo visto en clases (recordemos que cada neurona tiene asociado un _bias_). Ahora vamos a entrenar nuestro modelo.
# +
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
iris = load_iris()
X = iris.data[:,2:] # Nos quedamos con el largo y ancho del pétalo
y = iris.target
# Hacemos la división
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# Compilamos la red neuronal
# model.compile(loss='sparse_categorical_crossentropy',
# optimizer='sgd', metrics=['accuracy'])
# Compilamos la red neuronal pero entregando una learning rate
model.compile(loss='sparse_categorical_crossentropy',
optimizer=keras.optimizers.SGD(lr=0.01), metrics=['accuracy'])
# -
# Algunas acotaciones sobre el la compilación del modelo:
#
# - Dado que en este modelo estamos usando variables categóricas exclusivas (es decir, una flor no puede ser de dos tipos), usamos `sparse_categorical_cross_entropy`. Si queremos que las clases no sean exclusivas, utilizamos `categorical_cross_entropy`, pero las categorías deben estar con _encoding one hot_.
# - `sgd` es para utilizar _Stochastic Gradient Descent_.
# - `accuracy` es para que se nos muestre el _accuracy_ durante el entrenamiento.
# - Para clasificación binaria, en vez de una activación `softmax` se usa una activación `sigmoid`, además la función de pérdida pasa a ser `binary_crossentropy`.
#
# Ahora vamos a hacer `fit` del modelo.
model.fit(X_train, y_train, batch_size=16, epochs=200)
# Como vemos, en cada _epoch_ fue incrementando el _accuracy_ sobre el _dataset_ de entrenamiento. Ahora vamos a predecir sobre el _dataset_ de prueba.
# Aquí obtenemos un vector de probabilidades
y_proba = model.predict(X_test)
# Que redondeamos a dos decimales
y_proba.round(2)
# Y ahora para mostrar la clase
import numpy as np
y_pred = np.argmax(y_proba, axis=-1)
y_pred
# Vamos a comparar con nuestro _dataset_ de prueba:
np.sum(y_pred == y_test) / len(y_pred)
# Vamos a graficar ahora la frontera de decisión.
# +
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
# Colores
cmap_light = ListedColormap(['#f2cdc3', '#d0f2cd', '#c9f6ff'])
cmap_bold = ['#a72618', '#34892d', '#262ea7']
# Buscamos los límites del gráfico
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
# Generamos las combinaciones de x/y del plano en los rangos
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02),
np.arange(y_min, y_max, 0.02))
# Predecimos sobre el plano con la red neuronal
y_proba = model.predict(np.c_[xx.ravel(), yy.ravel()])
# Y lo convertimos a valores
Z = np.argmax(y_proba, axis=-1)
# Resultado de la frontera de decisión con color claro
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light, shading='auto')
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.scatter(X[y==0, 0], X[y==0, 1], c=cmap_bold[0], label='Iris Setosa')
plt.scatter(X[y==1, 0], X[y==1, 1], c=cmap_bold[1], label='Iris Versicolor')
plt.scatter(X[y==2, 0], X[y==2, 1], c=cmap_bold[2], label='Iris Virginica')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.legend()
plt.show()
# -
# Como vemos, la clasificación realizada por la red es muy distinta a la que nos habíamos encontrado hasta ahora. Ahora, un buen ejercicio es probar qué pasa con más neuronas en una de las capas.
# +
# Instanciamos un modelo "Secuencial"
# Siempre es buena práctica reiniciar el modelo
keras.backend.clear_session()
model = keras.Sequential(
[
layers.Dense(5, activation='relu', input_shape=(2,), name='hidden_layer_1'),
layers.Dense(20, activation='relu', name='hidden_layer_2'),
layers.Dense(3, activation='softmax', name='output_layer'),
]
)
# Compilamos la red neuronal
model.compile(loss='sparse_categorical_crossentropy',
optimizer='sgd', metrics=['accuracy'])
model.fit(X_train, y_train, batch_size=16, epochs=200, verbose=0)
# Colores
cmap_light = ListedColormap(['#f2cdc3', '#d0f2cd', '#c9f6ff'])
cmap_bold = ['#a72618', '#34892d', '#262ea7']
# Buscamos los límites del gráfico
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
# Generamos las combinaciones de x/y del plano en los rangos
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02),
np.arange(y_min, y_max, 0.02))
# Predecimos sobre el plano con la red neuronal
y_proba = model.predict(np.c_[xx.ravel(), yy.ravel()])
# Y lo convertimos a valores
Z = np.argmax(y_proba, axis=-1)
# Resultado de la frontera de decisión con color claro
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light, shading='auto')
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.scatter(X[y==0, 0], X[y==0, 1], c=cmap_bold[0], label='Iris Setosa')
plt.scatter(X[y==1, 0], X[y==1, 1], c=cmap_bold[1], label='Iris Versicolor')
plt.scatter(X[y==2, 0], X[y==2, 1], c=cmap_bold[2], label='Iris Virginica')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.legend()
plt.show()
# -
# Ahora, una buena idea también es entender como cambia la decisión con otras funciones de activación.
# +
# Instanciamos un modelo "Secuencial"
# Siempre es buena práctica reiniciar el modelo
keras.backend.clear_session()
model = keras.Sequential(
[
layers.Dense(5, activation='linear', input_shape=(2,), name='hidden_layer_1'),
layers.Dense(10, activation='linear', name='hidden_layer_2'),
layers.Dense(3, activation='softmax', name='output_layer'),
]
)
# Compilamos la red neuronal
model.compile(loss='sparse_categorical_crossentropy',
optimizer='sgd', metrics=['accuracy'])
model.fit(X_train, y_train, batch_size=16, epochs=200, verbose=0)
# Colores
cmap_light = ListedColormap(['#f2cdc3', '#d0f2cd', '#c9f6ff'])
cmap_bold = ['#a72618', '#34892d', '#262ea7']
# Buscamos los límites del gráfico
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
# Generamos las combinaciones de x/y del plano en los rangos
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02),
np.arange(y_min, y_max, 0.02))
# Predecimos sobre el plano con la red neuronal
y_proba = model.predict(np.c_[xx.ravel(), yy.ravel()])
# Y lo convertimos a valores
Z = np.argmax(y_proba, axis=-1)
# Resultado de la frontera de decisión con color claro
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light, shading='auto')
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.scatter(X[y==0, 0], X[y==0, 1], c=cmap_bold[0], label='Iris Setosa')
plt.scatter(X[y==1, 0], X[y==1, 1], c=cmap_bold[1], label='Iris Versicolor')
plt.scatter(X[y==2, 0], X[y==2, 1], c=cmap_bold[2], label='Iris Virginica')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.legend()
plt.show()
# -
# Como vemos, la activación lineal hace que la clasificación sea básicamente lineal en la separación de las clases.
# +
# Instanciamos un modelo "Secuencial"
# Siempre es buena práctica reiniciar el modelo
keras.backend.clear_session()
model = keras.Sequential(
[
layers.Dense(5, activation='sigmoid', input_shape=(2,), name='hidden_layer_1'),
layers.Dense(10, activation='sigmoid', name='hidden_layer_2'),
layers.Dense(3, activation='softmax', name='output_layer'),
]
)
# Compilamos la red neuronal
model.compile(loss='sparse_categorical_crossentropy',
optimizer='sgd', metrics=['accuracy'])
model.fit(X_train, y_train, batch_size=16, epochs=200, verbose=0)
# Colores
cmap_light = ListedColormap(['#f2cdc3', '#d0f2cd', '#c9f6ff'])
cmap_bold = ['#a72618', '#34892d', '#262ea7']
# Buscamos los límites del gráfico
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
# Generamos las combinaciones de x/y del plano en los rangos
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02),
np.arange(y_min, y_max, 0.02))
# Predecimos sobre el plano con la red neuronal
y_proba = model.predict(np.c_[xx.ravel(), yy.ravel()])
# Y lo convertimos a valores
Z = np.argmax(y_proba, axis=-1)
# Resultado de la frontera de decisión con color claro
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light, shading='auto')
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.scatter(X[y==0, 0], X[y==0, 1], c=cmap_bold[0], label='Iris Setosa')
plt.scatter(X[y==1, 0], X[y==1, 1], c=cmap_bold[1], label='Iris Versicolor')
plt.scatter(X[y==2, 0], X[y==2, 1], c=cmap_bold[2], label='Iris Virginica')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.legend()
plt.show()
# -
# Y como vemos, la función sigmoide no parece ser una buena elección para este problema. Ahora vamos a ver un segundo ejemplo sobre el _dataset_ Fashion MNIST.
#
# ## Fashion MNIST
#
# Este _dataset_ es similar al _dataset_ MNIST, solo que aquí vamos a clasificar prendas de ropa en distintos tipos.
# +
fashion_mnist = keras.datasets.fashion_mnist
# Cargamos los Datasets
(X_train_full, y_train_full), (X_test, y_test) = fashion_mnist.load_data()
X_train_full.shape
# -
X_train_full
# +
# Creamos datasets de entrenamiento, validación y prueba
# Las features se dividen en 255. para que los valores queden entre 0 y 1.
X_valid, X_train = X_train_full[:5000] / 255., X_train_full[5000:] / 255.
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
X_test = X_test / 255.
y_train
# -
# Las clases son números, su significado es el siguiente.
class_names = ["T-shirt/top", "Trouser", "Pullover", "Dress", "Coat",
"Sandal", "Shirt", "Sneaker", "Bag", "Ankle boot"]
n_rows = 4
n_cols = 10
plt.figure(figsize=(n_cols * 1.2, n_rows * 1.2))
for row in range(n_rows):
for col in range(n_cols):
index = n_cols * row + col
plt.subplot(n_rows, n_cols, index + 1)
plt.imshow(X_train[index], cmap="binary", interpolation="nearest")
plt.axis('off')
plt.title(class_names[y_train[index]], fontsize=12)
plt.subplots_adjust(wspace=0.2, hspace=0.5)
plt.show()
# +
keras.backend.clear_session()
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
# La capa flatten es para recibir la imagen de 28x28 sin transformar el dataset de entrenamiento
model.summary()
# +
model.compile(loss="sparse_categorical_crossentropy",
optimizer="sgd",
metrics=["accuracy"])
# El batch size default es de 32
history = model.fit(X_train, y_train, verbose=0, epochs=30,
validation_data=(X_valid, y_valid))
# -
# Ahora vamos a graficar cómo se comportó el modelo durante el entrenamiento.
# +
import pandas as pd
pd.DataFrame(history.history).plot(figsize=(8, 5))
plt.grid(True)
plt.gca().set_ylim(0, 1)
plt.show()
# -
# Ahora para evaluar sobre los datos de prueba.
model.evaluate(X_test, y_test)
# Que no está nada de mal para una tarea súper compleja. Uno además puede hacer _fine tune_ de varios hiperparámetros para mejorar el rendimiento. Si bien podemos usar `GridSearchCV` para encontrar buenos hiperparámetros, hay librerías optimizadas para Keras. Una de ellas es [la siguiente librería](https://www.tensorflow.org/tutorials/keras/keras_tuner).
#
# ## Regresión y redes neuronales.
#
# Como mencionamos antes, para hacer regresión con una red neuronal, la técnica que se usa en general es tener una única neurona de salida (y generalmente, sin una función de activación asociada), además de cambiar la función de pérdida por `mean_squared_error`. Vamos a ver un ejemplo.
# +
# Generamos el dataset
m = 100
X = 2 * np.random.rand(m, 1) - 1
y = (0.2 + 0.1 * X + 0.5 * X**2 + np.random.randn(m, 1)/10).ravel()
keras.backend.clear_session()
model = keras.models.Sequential([
keras.layers.Dense(100, activation="relu", input_shape=X.shape[1:]),
keras.layers.Dense(1)
])
model.compile(loss="mean_squared_error", optimizer=keras.optimizers.SGD(lr=1e-2))
history = model.fit(X, y, epochs=100, verbose=0)
# +
X_p = np.arange(-1.5, 1.5, 0.1)
X_p = X_p.reshape(-1, 1)
plt.plot(X_p, model.predict(X_p))
plt.plot(X, y, 'bs')
| 14,801 |
/ocsvm.ipynb | b692380b4d218d6fcdcafdac3c448b490224c16c | [] | no_license | watarium/time_ocsvm | https://github.com/watarium/time_ocsvm | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 62,736 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + [markdown] deletable=true editable=true
# # Data Science Callenge
# This challenge uses 4 years of power consumption measurements of a single household with one-minute intervals of sampling. The data is available here:
# http://archive.ics.uci.edu/ml/datasets/Individual+household+electric+power+consumption
#
# The challenge is accompanied by the following list of questions, for which responses are underneath:
#
# 1. How do you handle the data?
# * All data is wrangled with the `pandas` package in python. Analysis is done using various python packages. Although minute-by-minute data is provided, the challenge asks that we only make predictions using data upto the preceding hour. This constraint is conveniently satisfied by simply confining the data set to hourly intervals. Since there is a large amount of data available, the models we build are unlikely to be greatly affected by this approach of dropping data.
#
# 2. What analysis do you run to understand the features?
# * Basic stats (mean, std), time-series plots of the features/outputs, histograms, and correlation plots. This is important when verifying the validity of the imputing step. A very strong linear correlation was found between the Global_average_power and Global_intensity. Voltage measurements appear Gaussian with a peak just above 240 V, so it's likely the data was recorded from a UK home.
#
# 3. What do you do with missing or incorrect values?
# * Through trial-and-error, we find that missing values are stored as '?', which can be immediately picked up by `pandas`. They are then filled using the preceding value in the series, though they may be imputed using mean values as well. Using forward fill is better than mean imputation since the minute-by-minute variation is quite small (verified by the RMSE of a persistent model using minute-sampling data). The time-series of the Global_active_power is plotted before and after imputing to check for any outlying values when imputing from the mean.
#
# 4. What type of model do you use, and why?
# * Two models were investigated:
# * __ARMA__: This is a standard time-series model built on autogregression (using a number of preceding values (lag $\approx$ 50 below) as inputs) and moving averages. After experimenting a bit, we found the AR model worked best. It is quick to train and implement with no tuning parameters.
# * __LSTM Neural Network__: Neural nets are a much publicized learning algorithm which, in prinicple, require no a-priori modeling. In this analysis, we experimented with a few LSTM architectures with a casual approach to hyper-parameter tuning.
#
# 5. What metric do you use to evaluate the model?
# * The evaluation metric employed was the root mean square error:
# $$ \text{RMSE} = \frac{\sqrt{\sum_{n=1}^{n=N}(y_i - \hat{y}(t_i))^2}}{N-1}$$
#
# 6. What baseline do you compare it against?
# * The persistent model:
# $$\hat{y}(t_i+1 \text{ hour}) = y_i,$$
# which yields $\text{RMSE}_p = 1.041$. The RMSE of the trained models are
# * __ARMA__: 0.847
# * __LSTM NN__: 0.790 - 0.820
#
# Our model represents more than a 15% improvement over the baseline (presistent model), with the neural net performing a couple of percent better.
# + [markdown] deletable=true editable=true
# ## Load libraries and data
# + deletable=true editable=true
## load libraries
# # %matplotlib notebook
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import datetime as dt
# + deletable=true editable=true
## load data
file_name = 'household_power_consumption.zip'
# ensure the text is converted appropriately to floats
dtypes = {'Data': str, 'Time': str, 'Global_active_power': float, 'Global_reactive_power': float,
'Voltage': float, 'Global_intensity': float, 'Sub_metering_1': float,
'Sub_metering_2': float, 'Sub_metering_3': float}
data_df = pd.read_csv(file_name, compression='infer', sep=';', na_values='?', dtype=dtypes)
new_names = {'Sub_metering_1': 'Kitchen', 'Sub_metering_2': 'Laundry',
'Sub_metering_3': 'Water_AC'}
data_df.rename(columns=new_names, inplace=True)
# load various subsets of the data
data_df = data_df.iloc[::60, :] # get hourly data
# data_df = data_df.iloc[15000:, :] # the later half of the data
# data_df = data_df.iloc[::60*24, :] # get daily data
# data_df = data_df.iloc[-50*24*60:, :] # get last 15 days of data
# data_df = data_df.sample(frac=0.10) # get random sample of data
data_df.head(n=2)
# + deletable=true editable=true
## create datetime object as a feature
data_df['DateTime'] = data_df[['Date','Time']].apply(
lambda d: d[0] + ' ' + d[1], axis=1)
data_df['DateTime'] = data_df['DateTime'].apply(
lambda x: dt.datetime.strptime(x, '%d/%m/%Y %H:%M:%S'))
tmp_data = pd.DataFrame(data_df.pop('DateTime'))
data_df = tmp_data.join(data_df)
data_df.sort_values(by='DateTime', inplace=True)
# keep a subset of the data past some start date
# start_date = dt.datetime(year=2010, month=1, day=1)
# data_df = data_df[ data_df.DateTime > start_date ]
data_df.head(n=2)
# + deletable=true editable=true
## check how many missing values are around
count_nan = len(data_df) - data_df.count()
count_nan
# + [markdown] deletable=true editable=true
# ## Data/feature exploration
# + deletable=true editable=true
## check means and stds to see if they look like meaningful numbers
data_df.describe()
# + deletable=true editable=true
## plot Global_active_power and features against time
data_df.plot(x='DateTime', subplots=True, figsize=(10,10))
plt.tight_layout()
plt.show()
# + deletable=true editable=true
## impute missing values
from sklearn.preprocessing import Imputer
impute_method = 'forward_fill' # 'mean' or 'forward_fill' (preceding value)
if impute_method == 'mean':
feature_list = [c for c in data_df.columns if c not in ['Date', 'Time', 'DateTime']]
for feature in feature_list:
imputer = Imputer(missing_values=np.nan, strategy='mean')
data_df[feature] = imputer.fit_transform(data_df[feature].values.reshape(-1, 1))
elif impute_method == 'forward_fill':
data_df.fillna(method='ffill', inplace=True)
else:
print('Choose a proper imputing method.')
# + deletable=true editable=true
count_nan = len(data_df) - data_df.count()
if count_nan.sum() != 0:
print('There are {:d} more missing values to deal with.'.format(count_nan.sum()))
# + deletable=true editable=true
## plot Global_active_power again to make sure imputing didn't create any spurious behaviour
data_df.plot(x='DateTime', y='Global_active_power', figsize=(10,3))
plt.tight_layout()
plt.show()
# + [markdown] deletable=true editable=true
# ## Explore correlation between features/output
# + deletable=true editable=true
## plot histograms of each feature
feature_list = [c for c in data_df.columns if c not in ['Date', 'Time', 'DateTime']]
n_features = len(feature_list)
fig, ax = plt.subplots(nrows=int(np.ceil(n_features/4)),
ncols=4, figsize=(13,6), squeeze=True)
ax = ax.ravel()
nplot = 0
for feature in feature_list:
ax[nplot].hist(data_df[feature], bins=50, normed=False)
ax[nplot].set_title(feature)
nplot = nplot + 1
plt.tight_layout()
plt.show()
# + deletable=true editable=true
## scatter plot of each feature against the target
feature_list = [c for c in data_df.columns if c not in ['Date', 'Time', 'DateTime',
'Global_active_power']]
n_features = len(feature_list)
fig, ax = plt.subplots(nrows=int(np.ceil(n_features/3)),
ncols=3, figsize=(13,6), squeeze=True)
ax = ax.ravel()
nplot = 0
for feature in feature_list:
ax[nplot].scatter(data_df[feature], data_df.Global_active_power, s=20)
ax[nplot].set_title(feature)
nplot = nplot + 1
plt.tight_layout()
plt.show()
# + deletable=true editable=true
## heatmap of correlation matrix
corrmat = data_df.corr()
fig, ax = plt.subplots(figsize=(6, 4))
sns.heatmap(corrmat, vmax=.8, square=True)
plt.tight_layout()
plt.show()
# + [markdown] deletable=true editable=true
# ## Autoregression Models
# + deletable=true editable=true
## create time series for predicted output and prediction time stamps
data_ts = data_df['Global_active_power']
data_ts.index = data_df.DateTime
# + deletable=true editable=true
## check autocorrelation
from pandas.tools.plotting import autocorrelation_plot, lag_plot
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.stattools import adfuller
values = pd.DataFrame(data_ts)
df = pd.concat([values.shift(1), values], axis=1)
df.columns = ['t', 't+1']
result = df.corr()
print(result)
# lag_plot(data_ts)
# plt.show()
# autocorrelation_plot(data_ts)
# plt.show()
plot_acf(data_ts, lags=30)
plt.show()
plot_pacf(data_ts, lags=30)
plt.show()
# check if stationary
result = adfuller(data_ts)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
print('Critical Values:')
for key, value in result[4].items():
print('\t%s: %.3f' % (key, value))
# since adf statistics is less than the 1% critical value, we reject
# the null hypothesis. This means there is no unit run and the
# time series is stationary: there are not time-dependent structures
# one level of differencing, so d =>1 in ARIMA
# + deletable=true editable=true
## build and test the model
from statsmodels.tsa.ar_model import AR
from statsmodels.tsa.arima_model import ARIMA
from statsmodels.tsa.arima_model import ARMA
from sklearn.metrics import mean_squared_error
# experiment with AR and MA models
model = AR(data_ts, missing='raise') # rmse = 0.847
# model = ARMA(data_ts, (2,2)) # rmse = 0.889
# model = ARIMA(data_ts, order=(10, 0, 2), missing='raise')
model = model.fit()
lag = model.k_ar
print('Lag: %s' % lag)
# make predictions
pred_times = data_ts.index[lag:]
pred = model.predict(start=pred_times[0], end=pred_times[-1], dynamic=False)
rmse = np.sqrt(mean_squared_error(data_ts.values[lag:] , pred.values))
rmse_persistent = np.sqrt(mean_squared_error(data_ts.values[lag:-1],
data_ts.values[lag+1:]))
print('RMSE: {:0.3f}'.format(rmse))
print('RMSE Persistent: {:0.3f}'.format(rmse_persistent))
# + deletable=true editable=true
## plot the data
def plot_data(start_date, title='figure'):
ax = data_ts[ data_ts.index > start_date ].plot(
color='grey', alpha=0.8, figsize=(7,3), fontsize=12)
pred[ pred.index > start_date ].plot(color='red', alpha=0.4)
ax.legend(['data', 'forecast'], fontsize=12)
ax.set_ylabel('Global_active_power', fontsize=12)
ax.set_xlabel('')
ax.set_title(title, fontsize=12)
file_name = title.replace(' ','_').lower() + '.png'
plt.savefig(file_name, orientation='landscape',format='png')
plt.show()
start_date = dt.datetime(year=2005, month=1, day=1, hour=1, minute=1)
plot_data(start_date, title='Full data set')
start_date = dt.datetime(year=2010, month=11, day=15, hour=1, minute=1)
plot_data(start_date, title='Partial data set')
# + [markdown] deletable=true editable=true
# ## LSTM Net Model
# Tips available here:
# http://machinelearningmastery.com/time-series-prediction-lstm-recurrent-neural-networks-python-keras/
# + deletable=true editable=true
## load modules for building neural net
from keras.models import Sequential
from keras.layers import Dense, LSTM, Activation
from sklearn.preprocessing import MinMaxScaler
# + deletable=true editable=true
## pre-process data
# scale data for NN activation functions
scaler = MinMaxScaler(feature_range=(0,1))
# reshape : (-1,1) for a single feature. (1,-1) for single sample
data_values = data_ts.values.astype('float32').reshape(-1,1)
data_values = scaler.fit_transform(data_values)
# split into train and test set
train_size = int(len(data_values) * 1/2)
test_size = len(data_values) - train_size
train, test = data_values[0:train_size], data_values[train_size:]
print(len(train), len(test))
# + deletable=true editable=true
## create features to data set (preceding values for regression)
def create_dataset(dataset, look_back=1):
features, target = [], []
for i in range(0, len(dataset)-look_back-1):
features.append( dataset[i:(i+look_back),0] )
target.append( dataset[i+look_back,0] )
return np.array(features), np.array(target)
look_back = 24*21
trainX, trainY = create_dataset(train, look_back=look_back)
testX, testY = create_dataset(test, look_back=look_back)
# reshape for keras input: [samples, time steps, features]
# time step = look_back = number of previous steps to include
trainX = np.reshape(trainX, (trainX.shape[0], 1, trainX.shape[1]))
testX = np.reshape(testX, (testX.shape[0], 1, testX.shape[1]))
# + deletable=true editable=true
# ## create and fit LSTM network with 4 neurons in 1 layer
batch_size = 11 # must be factor of number of samples
model = Sequential()
# build/experiment with the network
# 4-neuron hidden layer, default LSTM activation = 'tanh'
# model.add(LSTM(4, input_dim=look_back, return_sequences=False))
# 3 hidden layers with drop out
model.add(LSTM(32, input_dim=look_back, return_sequences=True,
dropout_U=0.2, dropout_W=0.2))
model.add(LSTM(16, return_sequences=True, dropout_U=0.2, dropout_W=0.2))
model.add(LSTM(4, return_sequences=False))
model.add(Dense(1, activation='relu')) # default activation = linear
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(trainX, trainY, nb_epoch=20, batch_size=16, verbose=0)
## LSTM with memory (doesn't perform so well)
# model = Sequential()
# model.add(LSTM(64, batch_input_shape=(batch_size, 1, look_back),
# stateful=True, return_sequences=True,
# dropout_U=0.25, dropout_W=0.25))
# model.add(LSTM(32, batch_input_shape=(batch_size, 1, look_back),
# stateful=True, return_sequences=True,
# dropout_U=0.25, dropout_W=0.25))
# model.add(LSTM(8, batch_input_shape=(batch_size, 1, look_back),
# stateful=True))
# model.add(Dense(1))
# model.compile(loss='mean_squared_error', optimizer='adam')
# for i in range(40):
# model.fit(trainX, trainY, nb_epoch=1, batch_size=batch_size, verbose=2, shuffle=False)
# model.reset_states()
# + deletable=true editable=true
## make predictions and unscale
train_pred = model.predict(trainX, batch_size=batch_size)
train_pred = scaler.inverse_transform(train_pred)
trainY = scaler.inverse_transform([trainY])
# model.reset_states() # use if stateful memory
test_pred = model.predict(testX, batch_size=batch_size)
test_pred = scaler.inverse_transform(test_pred)
testY = scaler.inverse_transform([testY])
# + deletable=true editable=true
## test predictions
trainScore = np.sqrt(mean_squared_error(trainY[0,:], train_pred[:,0]))
print('Train Score: {:.3f} RMSE'.format(trainScore))
testScore = np.sqrt(mean_squared_error(testY[0,:], test_pred[:,0]))
print('Test Score: {:.3f} RMSE'.format(testScore))
# + deletable=true editable=true
## plot the data
# convert NN output to pandas time series with proper dates
train_plot = pd.Series(train_pred[:,0], index=data_ts.index[
look_back:len(train_pred)+look_back])
test_plot = pd.Series(test_pred[:,0], index=data_ts.index[
len(train_pred)+(2*look_back)+1:len(data_values)-1])
def plot_data(start_date, title='figure'):
ax = data_ts[ data_ts.index > start_date ].plot(
color='grey', alpha=0.8, figsize=(7,3), fontsize=12)
test_plot[ test_plot.index > start_date ].plot(color='red', alpha=0.4)
try:
train_plot[ train_plot.index > start_date ].plot(color='green', alpha=0.4)
except:
pass
ax.legend(['data', 'test set', 'training set'], fontsize=12)
ax.set_ylabel('Global_active_power', fontsize=12)
ax.set_xlabel('')
ax.set_title(title, fontsize=12)
file_name = title.replace(' ','_').lower() + '.png'
plt.savefig(file_name, orientation='landscape',format='png')
plt.show()
start_date = dt.datetime(year=2005, month=1, day=1, hour=1, minute=1)
plot_data(start_date, title='Full data set')
start_date = dt.datetime(year=2010, month=11, day=15, hour=1, minute=1)
plot_data(start_date, title='Partial data set')
| 16,619 |
/disjoint sets/Disjoint Set - I.ipynb | 2995c3d851b9cf13860e863b03249616990ecdad | [] | no_license | ShubhGOYAL5/data-structures-in-python | https://github.com/ShubhGOYAL5/data-structures-in-python | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 2,592 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class UnionFind:
def __init__(self, size):
self.root = [i for i in range(size)]
def find(self, x):
return self.root[x]
def union(self, x, y):
rootX = self.find(x)
rootY = self.find(y)
if rootX != rootY:
for i in range(len(self.root)):
if self.root[i] == rootY:
self.root[i] = rootX
def connected(self, x, y):
return self.find(x) == self.find(y)
# +
uf = UnionFind(10)
uf.union(1, 2)
uf.union(2, 5)
uf.union(5, 6)
uf.union(6, 7)
uf.union(3, 8)
uf.union(8, 9)
# 0-1-2-5-6-7 3-8-9 4
# -
print(uf.connected(1, 5))
print(uf.connected(5, 7))
print(uf.connected(4, 9))
print(uf.connected(0, 4))
uf.union(0, 4)
print(uf.connected(0, 4))
| 1,064 |
/Do_it_Algorithm_DataStructure/chap 5 재귀 알고리즘.ipynb | a6d7901e923f1b1bcac139fcaf3af0d68ed7c574 | [] | no_license | hayaseleu/leetcode_mysol | https://github.com/hayaseleu/leetcode_mysol | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 9,434 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="NXZYOHe0mZmy" colab_type="text"
# # Assignment 8
#
# Develop a model for 20 news groups dataset from scikit-learn. Select 20% of data for test set.
#
# Develop metric learning model with siamese network [3 points] and triplet loss [3 points] (from seminar).
# Use KNN and LSH (any library for approximate nearest neighbor search) for final prediction after the network was trained. [2 points]
#
# # # ! Remember, that LSH gives you a set of neighbor candidates, for which you have to calculate distances to choose top-k nearest neighbors.
#
# Your quality metric = accuracy score [2 points if acc > 0.8 ]
# + id="e9-P52zuDOVr" colab_type="code" colab={}
import numpy as np
import pandas as pd
import nltk
import torch as tt
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader, Dataset, TensorDataset
from torchtext import data
from sklearn.model_selection import train_test_split
from tqdm import tqdm_notebook
# + id="NVGRtYocvaZP" colab_type="code" outputId="6940f1e6-7ee0-4c23-9081-cd080107900c" colab={"base_uri": "https://localhost:8080/", "height": 51}
from sklearn.datasets import fetch_20newsgroups
all_data = fetch_20newsgroups(subset='train', remove=('headers', 'footers', 'quotes'))
# + id="JNUIlz95EF-S" colab_type="code" outputId="fd93d79f-c536-48ef-fb59-1913526999fe" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(all_data.data), len(all_data.target)
# + id="MYYlmqSvEQZc" colab_type="code" outputId="3bb1add2-87cd-4e05-b953-770186d977a3" colab={"base_uri": "https://localhost:8080/", "height": 88}
all_data.data[:3]
# + id="EXQIk4g6EU6j" colab_type="code" outputId="389b8d9a-2ad4-445d-c1cf-5f17d9e83ed1" colab={"base_uri": "https://localhost:8080/", "height": 34}
all_data.target[:3]
# + id="hxXk51hbAdhf" colab_type="code" outputId="d163dc57-4dbb-4a4d-b78f-9e4cea40f02a" colab={"base_uri": "https://localhost:8080/", "height": 204}
import pandas as pd
d = {'text': all_data.data, 'target': all_data.target}
df = pd.DataFrame(data = d)
df.head()
# + id="19fkruTQqbph" colab_type="code" colab={}
all_len = []
for row in df.text:
text_len = len(row)
all_len.append(text_len)
# + id="TQKCgUL8youc" colab_type="code" colab={}
df['text_len'] = all_len
# + id="JqLKik_6yuLM" colab_type="code" outputId="895988f7-6180-4bf1-b0d5-8fbe5f64737c" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + id="7FsqHQrAYjwi" colab_type="code" outputId="d9b305a5-70d5-46e5-91f8-3591be437d0b" colab={"base_uri": "https://localhost:8080/", "height": 204}
new_df = df[df.text_len < 1000].reset_index(drop=True)
new_df.drop('text_len', axis='columns', inplace=True)
new_df['target'] = new_df['target'].apply(float)
new_df.head()
# + id="jM0yZS1oEnof" colab_type="code" colab={}
pip install sentence_transformers
# + id="FKjRZWGMvOKP" colab_type="code" outputId="bd45fcbd-c002-42f2-8edf-306b574c9bd7" colab={"base_uri": "https://localhost:8080/", "height": 80}
from sentence_transformers import SentenceTransformer
model = SentenceTransformer('bert-base-wikipedia-sections-mean-tokens')
#модель была выбрана из представленных на сайте https://github.com/UKPLab/sentence-transformers
# + id="ue4KEU_MwFRE" colab_type="code" colab={}
sentences = ['This framework generates embeddings for each input sentence. Sentences are passed as a list of string. The quick brown fox jumps over the lazy dog.']
sentence_embeddings = model.encode(sentences)
# + id="FdlAQ3PO4ycH" colab_type="code" outputId="73da2a20-5fc3-43fb-d94b-ea26785ec47f" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(sentence_embeddings), sentence_embeddings[0].shape
# + id="TRx6c-EfyJFC" colab_type="code" outputId="e4bd6d43-95d9-4f22-9871-22afdee15f69" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="UKAmgQV1ykbU" colab_type="code" colab={}
import os
os.chdir('gdrive/My Drive/Colab Notebooks')
# + id="NUPsbxLP1y1r" colab_type="code" outputId="913167cd-bec4-4018-994f-1b8b7ca86cc3" colab={"base_uri": "https://localhost:8080/", "height": 34}
embs = model.encode(new_df.text, show_progress_bar=True)
# + id="QzdmiVJt9Zoc" colab_type="code" outputId="825fb205-2d24-4378-c026-c5960f15b0d8" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.externals import joblib
joblib.dump(embs, "embs.pkl")
# + id="Y_cwUMRR9ftb" colab_type="code" colab={}
from sklearn.externals import joblib
embs = joblib.load("embs.pkl")
# + id="I1jfKrhnTpO9" colab_type="code" outputId="44159ab0-2fda-4b84-83dc-4b9fe1d7ce56" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(embs)
# + id="JbxvPFHnsU5D" colab_type="code" colab={}
new_df['embs'] = embs
# + id="Rm2-FCw9salu" colab_type="code" outputId="336bc75a-4311-451d-9d2e-063a5d9525a5" colab={"base_uri": "https://localhost:8080/", "height": 204}
new_df.head()
# + id="r3nsVkAIuHkw" colab_type="code" colab={}
def pos_neg(row, df):
pos = np.random.choice(df[df['target'] == row['target']]['embs'])
neg = np.random.choice(df[df['target'] != row['target']]['embs'])
return pos, neg
# + id="LdUBGnmZ5XNt" colab_type="code" colab={}
pos_vars = []
neg_vars = []
for i, row in new_df.iterrows():
pos, neg = pos_neg(row, new_df)
pos_vars.append(pos)
neg_vars.append(neg)
# + id="kBjl4Y1c82Po" colab_type="code" outputId="76d145df-81b0-4355-95ad-96ed90e1e123" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(pos_vars), len(neg_vars)
# + id="Hr0o0sJg6EX1" colab_type="code" outputId="e7276ca2-7da8-407f-a79e-6ad56def5b83" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(embs[0]), len(pos_vars[0])
# + id="YBbWMjoq5Rjy" colab_type="code" outputId="e126ccd0-16fc-45ad-f901-cae9e0c78738" colab={"base_uri": "https://localhost:8080/", "height": 289}
new_df["pos"] = pos_vars
new_df["neg"] = neg_vars
new_df.head()
# + id="4q_2DQcG7GX1" colab_type="code" colab={}
df_train, df_test = train_test_split(new_df, test_size=0.2, random_state=42, shuffle=True)
df_train, df_val = train_test_split(df_train, test_size=0.1, random_state=42, shuffle=True)
# + id="5Xcm3c8P08de" colab_type="code" outputId="79bf7f0c-3fb1-4679-9fe5-85931ee45733" colab={"base_uri": "https://localhost:8080/", "height": 68}
print('Train shape = ', df_train.shape)
print('Validation shape = ', df_val.shape)
print('Test shape = ', df_test.shape)
# + id="NyrrQny29n1C" colab_type="code" colab={}
def triples(df):
anch = list(df.embs)
pos = list(df.pos)
neg = list(df.neg)
y = list(df.target)
return tt.FloatTensor(anch), tt.FloatTensor(pos), tt.FloatTensor(neg), tt.FloatTensor(y)
# + id="qSpV5o52-YLW" colab_type="code" colab={}
anch_train, pos_train, neg_train, y_train = triples(df_train)
anch_val, pos_val, neg_val, y_val = triples(df_val)
anch_test, pos_test, neg_test, y_test = triples(df_test)
# + id="SVsHZgKu13_z" colab_type="code" colab={}
batch_size = 32
train_loader = DataLoader(TensorDataset(anch_train, pos_train, neg_train, y_train), batch_size=batch_size)
valid_loader = DataLoader(TensorDataset(anch_val, pos_val, neg_val, y_val), batch_size=batch_size)
test_loader = DataLoader(TensorDataset(anch_test, pos_test, neg_test, y_test), batch_size=batch_size)
# + id="rlRzo94TQxoh" colab_type="code" colab={}
def _train_epoch(model, iterator, optimizer, curr_epoch):
model.train()
running_loss = 0
n_batches = len(iterator)
iterator = tqdm_notebook(iterator, total=n_batches, desc='epoch %d' % (curr_epoch), leave=True)
for i, batch in enumerate(iterator):
optimizer.zero_grad()
loss = model(batch)
loss.backward()
optimizer.step()
curr_loss = loss.data.cpu().detach().item()
loss_smoothing = i / (i+1)
running_loss = loss_smoothing * running_loss + (1 - loss_smoothing) * curr_loss
iterator.set_postfix(loss='%.5f' % running_loss)
return running_loss
# + id="pZiLJhhiQ2r1" colab_type="code" colab={}
def _test_epoch(model, iterator):
model.eval()
epoch_loss = 0
n_batches = len(iterator)
with tt.no_grad():
for batch in iterator:
loss = model(batch)
epoch_loss += loss.data.item()
return epoch_loss / n_batches
# + id="8ECZwKlXQ6DK" colab_type="code" colab={}
def nn_train(model, train_iterator, valid_iterator, optimizer, n_epochs=10,
scheduler=None, early_stopping=0):
prev_loss = 100500
es_epochs = 0
best_epoch = None
history = pd.DataFrame()
for epoch in range(n_epochs):
train_loss = _train_epoch(model, train_iterator, optimizer, epoch)
valid_loss = _test_epoch(model, valid_iterator)
valid_loss = valid_loss
print('validation loss %.5f' % valid_loss)
record = {'epoch': epoch, 'train_loss': train_loss, 'valid_loss': valid_loss}
history = history.append(record, ignore_index=True)
if early_stopping > 0:
if valid_loss > prev_loss:
es_epochs += 1
else:
es_epochs = 0
if es_epochs >= early_stopping:
best_epoch = history[history.valid_loss == history.valid_loss.min()].iloc[0]
print('Early stopping! best epoch: %d val %.5f' % (best_epoch['epoch'], best_epoch['valid_loss']))
break
prev_loss = min(prev_loss, valid_loss)
# + id="OvoAzWhYSNbu" colab_type="code" colab={}
def triplet_loss(anchor_embed, pos_embed, neg_embed, margin = 0.1):
res = F.cosine_similarity(anchor_embed, neg_embed) - F.cosine_similarity(anchor_embed, pos_embed) + margin
return tt.mean(res)
# + id="bbB6Wq5fQ_Gu" colab_type="code" colab={}
class Tripletnet(nn.Module):
def __init__(self):
super(Tripletnet, self).__init__()
self.fc = nn.Linear(768, 128)
def branch(self, x):
x = self.fc(x)
return x
def forward(self, batch):
anchor, pos, neg = batch[0], batch[1], batch[2]
anchor = self.branch(anchor)
pos = self.branch(pos)
neg = self.branch(neg)
return triplet_loss(anchor, pos, neg)
# + id="D0RJynBHRXyT" colab_type="code" colab={}
model = Tripletnet()
optimizer = optim.Adam(model.parameters())
# + id="hRi42_LYFK0s" colab_type="code" colab={}
nn_train(model, train_loader, test_loader, optimizer, n_epochs=100)
# + id="lqLuBdcQ5wbN" colab_type="code" colab={}
#from sklearn.externals import joblib
#joblib.dump(model, "trip_model.pkl")
# + id="YMYp04JSYt3_" colab_type="code" colab={}
#from sklearn.externals import joblib
#model = joblib.load("trip_model.pkl")
# + [markdown] id="diYIho9AVVB7" colab_type="text"
# Predictions
# + id="F8Xx35FaE8Iz" colab_type="code" colab={}
pip install annoy
# + id="7xvV53TZY6Sm" colab_type="code" outputId="38449d96-0f1a-490d-a268-3c511833637f" colab={"base_uri": "https://localhost:8080/", "height": 34}
from annoy import AnnoyIndex
a = AnnoyIndex(128, 'angular')
for i in range(anch_train.size()[0]):
v = model.branch(anch_train[i])
a.add_item(i, v)
a.build(1000)
# + id="rm6NmItabSpD" colab_type="code" outputId="163ca0d9-0270-43ee-ab1a-95ea6d0954a2" colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["ddf0b622d1c24d34a4444fd9df7c7ecc", "bb6dd0d6e14444de969fff93b167b28a", "1a0ea9cfefb3458e8dce784f047b88bd", "3e6a60a2d294446c8bca97d4fe5f179d", "68e14ebd7ab04a7eaea4901b02ef4796", "1ef335e77060417192100052a94d0cfd", "73d7030a43ab45feb1bebbe8a5076d6f", "04cfec16152542b8ae04f30817f78eab"]}
from sklearn.neighbors import KDTree
from scipy.spatial import cKDTree
preds = []
for i, vec in tqdm(enumerate(anch_val)):
top = a.get_nns_by_vector(model.branch(vec), 1500)
top_vecs = np.take(anch_train, np.array(top), axis=0).tolist()
top_y = np.take(y_train, np.array(top)).tolist()
t = cKDTree(top_vecs).query(vec, k=1)[1]
pred = top_vecs[t]
pred_idx = top_vecs.index(pred)
preds.append(top_y[pred_idx])
# + id="9yOpBFPRgcDa" colab_type="code" outputId="6737c608-f5b5-48c8-f914-1060632b5d63" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(y_val), len(preds)
# + id="XKGJ4r9WffLv" colab_type="code" outputId="82726abe-48e9-4a12-e561-bd51809ea74b" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.metrics import accuracy_score
accuracy_score(y_val, preds)
# + id="_mxPPoa94yx-" colab_type="code" colab={}
| 12,745 |
/C4/W1/Convolution_model_Application_v1a.ipynb | c80d513d1c7d5e7a0fabf9828daf022a0d4bbe97 | [] | no_license | Virusdoll/Stanford.CS230 | https://github.com/Virusdoll/Stanford.CS230 | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 182,955 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''cs230'': conda)'
# name: python3
# ---
# # Convolutional Neural Networks: Application
#
# Welcome to Course 4's second assignment! In this notebook, you will:
#
# - Implement helper functions that you will use when implementing a TensorFlow model
# - Implement a fully functioning ConvNet using TensorFlow
#
# **After this assignment you will be able to:**
#
# - Build and train a ConvNet in TensorFlow for a classification problem
#
# We assume here that you are already familiar with TensorFlow. If you are not, please refer the *TensorFlow Tutorial* of the third week of Course 2 ("*Improving deep neural networks*").
# ### <font color='darkblue'> Updates to Assignment <font>
#
# #### If you were working on a previous version
# * The current notebook filename is version "1a".
# * You can find your work in the file directory as version "1".
# * To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#
# #### List of Updates
# * `initialize_parameters`: added details about tf.get_variable, `eval`. Clarified test case.
# * Added explanations for the kernel (filter) stride values, max pooling, and flatten functions.
# * Added details about softmax cross entropy with logits.
# * Added instructions for creating the Adam Optimizer.
# * Added explanation of how to evaluate tensors (optimizer and cost).
# * `forward_propagation`: clarified instructions, use "F" to store "flatten" layer.
# * Updated print statements and 'expected output' for easier visual comparisons.
# * Many thanks to Kevin P. Brown (mentor for the deep learning specialization) for his suggestions on the assignments in this course!
# ## 1.0 - TensorFlow model
#
# In the previous assignment, you built helper functions using numpy to understand the mechanics behind convolutional neural networks. Most practical applications of deep learning today are built using programming frameworks, which have many built-in functions you can simply call.
#
# As usual, we will start by loading in the packages.
# +
import math
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
import tensorflow as tf
from tensorflow.python.framework import ops
from cnn_utils import *
# %matplotlib inline
np.random.seed(1)
# -
# Run the next cell to load the "SIGNS" dataset you are going to use.
# Loading the data (signs)
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# As a reminder, the SIGNS dataset is a collection of 6 signs representing numbers from 0 to 5.
#
# <img src="images/SIGNS.png" style="width:800px;height:300px;">
#
# The next cell will show you an example of a labelled image in the dataset. Feel free to change the value of `index` below and re-run to see different examples.
# Example of a picture
index = 6
plt.imshow(X_train_orig[index])
print ("y = " + str(np.squeeze(Y_train_orig[:, index])))
# In Course 2, you had built a fully-connected network for this dataset. But since this is an image dataset, it is more natural to apply a ConvNet to it.
#
# To get started, let's examine the shapes of your data.
X_train = X_train_orig/255.
X_test = X_test_orig/255.
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
conv_layers = {}
# ### 1.1 - Create placeholders
#
# TensorFlow requires that you create placeholders for the input data that will be fed into the model when running the session.
#
# **Exercise**: Implement the function below to create placeholders for the input image X and the output Y. You should not define the number of training examples for the moment. To do so, you could use "None" as the batch size, it will give you the flexibility to choose it later. Hence X should be of dimension **[None, n_H0, n_W0, n_C0]** and Y should be of dimension **[None, n_y]**. [Hint: search for the tf.placeholder documentation"](https://www.tensorflow.org/api_docs/python/tf/placeholder).
# +
# GRADED FUNCTION: create_placeholders
def create_placeholders(n_H0, n_W0, n_C0, n_y):
"""
Creates the placeholders for the tensorflow session.
Arguments:
n_H0 -- scalar, height of an input image
n_W0 -- scalar, width of an input image
n_C0 -- scalar, number of channels of the input
n_y -- scalar, number of classes
Returns:
X -- placeholder for the data input, of shape [None, n_H0, n_W0, n_C0] and dtype "float"
Y -- placeholder for the input labels, of shape [None, n_y] and dtype "float"
"""
### START CODE HERE ### (≈2 lines)
### END CODE HERE ###
return X, Y
# -
X, Y = create_placeholders(64, 64, 3, 6)
print ("X = " + str(X))
print ("Y = " + str(Y))
# **Expected Output**
#
# <table>
# <tr>
# <td>
# X = Tensor("Placeholder:0", shape=(?, 64, 64, 3), dtype=float32)
#
# </td>
# </tr>
# <tr>
# <td>
# Y = Tensor("Placeholder_1:0", shape=(?, 6), dtype=float32)
#
# </td>
# </tr>
# </table>
# ### 1.2 - Initialize parameters
#
# You will initialize weights/filters $W1$ and $W2$ using `tf.contrib.layers.xavier_initializer(seed = 0)`. You don't need to worry about bias variables as you will soon see that TensorFlow functions take care of the bias. Note also that you will only initialize the weights/filters for the conv2d functions. TensorFlow initializes the layers for the fully connected part automatically. We will talk more about that later in this assignment.
#
# **Exercise:** Implement initialize_parameters(). The dimensions for each group of filters are provided below. Reminder - to initialize a parameter $W$ of shape [1,2,3,4] in Tensorflow, use:
# ```python
# W = tf.get_variable("W", [1,2,3,4], initializer = ...)
# ```
# #### tf.get_variable()
# [Search for the tf.get_variable documentation](https://www.tensorflow.org/api_docs/python/tf/get_variable). Notice that the documentation says:
# ```
# Gets an existing variable with these parameters or create a new one.
# ```
# So we can use this function to create a tensorflow variable with the specified name, but if the variables already exist, it will get the existing variable with that same name.
#
# +
# GRADED FUNCTION: initialize_parameters
def initialize_parameters():
"""
Initializes weight parameters to build a neural network with tensorflow. The shapes are:
W1 : [4, 4, 3, 8]
W2 : [2, 2, 8, 16]
Note that we will hard code the shape values in the function to make the grading simpler.
Normally, functions should take values as inputs rather than hard coding.
Returns:
parameters -- a dictionary of tensors containing W1, W2
"""
tf.set_random_seed(1) # so that your "random" numbers match ours
### START CODE HERE ### (approx. 2 lines of code)
### END CODE HERE ###
parameters = {"W1": W1,
"W2": W2}
return parameters
# -
tf.reset_default_graph()
with tf.Session() as sess_test:
parameters = initialize_parameters()
init = tf.global_variables_initializer()
sess_test.run(init)
print("W1[1,1,1] = \n" + str(parameters["W1"].eval()[1,1,1]))
print("W1.shape: " + str(parameters["W1"].shape))
print("\n")
print("W2[1,1,1] = \n" + str(parameters["W2"].eval()[1,1,1]))
print("W2.shape: " + str(parameters["W2"].shape))
# ** Expected Output:**
#
# ```
# W1[1,1,1] =
# [ 0.00131723 0.14176141 -0.04434952 0.09197326 0.14984085 -0.03514394
# -0.06847463 0.05245192]
# W1.shape: (4, 4, 3, 8)
#
#
# W2[1,1,1] =
# [-0.08566415 0.17750949 0.11974221 0.16773748 -0.0830943 -0.08058
# -0.00577033 -0.14643836 0.24162132 -0.05857408 -0.19055021 0.1345228
# -0.22779644 -0.1601823 -0.16117483 -0.10286498]
# W2.shape: (2, 2, 8, 16)
# ```
# ### 1.3 - Forward propagation
#
# In TensorFlow, there are built-in functions that implement the convolution steps for you.
#
# - **tf.nn.conv2d(X,W, strides = [1,s,s,1], padding = 'SAME'):** given an input $X$ and a group of filters $W$, this function convolves $W$'s filters on X. The third parameter ([1,s,s,1]) represents the strides for each dimension of the input (m, n_H_prev, n_W_prev, n_C_prev). Normally, you'll choose a stride of 1 for the number of examples (the first value) and for the channels (the fourth value), which is why we wrote the value as `[1,s,s,1]`. You can read the full documentation on [conv2d](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d).
#
# - **tf.nn.max_pool(A, ksize = [1,f,f,1], strides = [1,s,s,1], padding = 'SAME'):** given an input A, this function uses a window of size (f, f) and strides of size (s, s) to carry out max pooling over each window. For max pooling, we usually operate on a single example at a time and a single channel at a time. So the first and fourth value in `[1,f,f,1]` are both 1. You can read the full documentation on [max_pool](https://www.tensorflow.org/api_docs/python/tf/nn/max_pool).
#
# - **tf.nn.relu(Z):** computes the elementwise ReLU of Z (which can be any shape). You can read the full documentation on [relu](https://www.tensorflow.org/api_docs/python/tf/nn/relu).
#
# - **tf.contrib.layers.flatten(P)**: given a tensor "P", this function takes each training (or test) example in the batch and flattens it into a 1D vector.
# * If a tensor P has the shape (m,h,w,c), where m is the number of examples (the batch size), it returns a flattened tensor with shape (batch_size, k), where $k=h \times w \times c$. "k" equals the product of all the dimension sizes other than the first dimension.
# * For example, given a tensor with dimensions [100,2,3,4], it flattens the tensor to be of shape [100, 24], where 24 = 2 * 3 * 4. You can read the full documentation on [flatten](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/flatten).
#
# - **tf.contrib.layers.fully_connected(F, num_outputs):** given the flattened input F, it returns the output computed using a fully connected layer. You can read the full documentation on [full_connected](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/fully_connected).
#
# In the last function above (`tf.contrib.layers.fully_connected`), the fully connected layer automatically initializes weights in the graph and keeps on training them as you train the model. Hence, you did not need to initialize those weights when initializing the parameters.
#
#
# #### Window, kernel, filter
# The words "window", "kernel", and "filter" are used to refer to the same thing. This is why the parameter `ksize` refers to "kernel size", and we use `(f,f)` to refer to the filter size. Both "kernel" and "filter" refer to the "window."
# **Exercise**
#
# Implement the `forward_propagation` function below to build the following model: `CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED`. You should use the functions above.
#
# In detail, we will use the following parameters for all the steps:
# - Conv2D: stride 1, padding is "SAME"
# - ReLU
# - Max pool: Use an 8 by 8 filter size and an 8 by 8 stride, padding is "SAME"
# - Conv2D: stride 1, padding is "SAME"
# - ReLU
# - Max pool: Use a 4 by 4 filter size and a 4 by 4 stride, padding is "SAME"
# - Flatten the previous output.
# - FULLYCONNECTED (FC) layer: Apply a fully connected layer without an non-linear activation function. Do not call the softmax here. This will result in 6 neurons in the output layer, which then get passed later to a softmax. In TensorFlow, the softmax and cost function are lumped together into a single function, which you'll call in a different function when computing the cost.
# +
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Note that for simplicity and grading purposes, we'll hard-code some values
such as the stride and kernel (filter) sizes.
Normally, functions should take these values as function parameters.
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "W2"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
W2 = parameters['W2']
### START CODE HERE ###
# CONV2D: stride of 1, padding 'SAME'
# RELU
# MAXPOOL: window 8x8, stride 8, padding 'SAME'
# CONV2D: filters W2, stride 1, padding 'SAME'
# RELU
# MAXPOOL: window 4x4, stride 4, padding 'SAME'
# FLATTEN
# FULLY-CONNECTED without non-linear activation function (not not call softmax).
# 6 neurons in output layer. Hint: one of the arguments should be "activation_fn=None"
### END CODE HERE ###
return Z3
# +
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(Z3, {X: np.random.randn(2,64,64,3), Y: np.random.randn(2,6)})
print("Z3 = \n" + str(a))
# -
# **Expected Output**:
#
# ```
# Z3 =
# [[-0.44670227 -1.57208765 -1.53049231 -2.31013036 -1.29104376 0.46852064]
# [-0.17601591 -1.57972014 -1.4737016 -2.61672091 -1.00810647 0.5747785 ]]
# ```
# ### 1.4 - Compute cost
#
# Implement the compute cost function below. Remember that the cost function helps the neural network see how much the model's predictions differ from the correct labels. By adjusting the weights of the network to reduce the cost, the neural network can improve its predictions.
#
# You might find these two functions helpful:
#
# - **tf.nn.softmax_cross_entropy_with_logits(logits = Z, labels = Y):** computes the softmax entropy loss. This function both computes the softmax activation function as well as the resulting loss. You can check the full documentation [softmax_cross_entropy_with_logits](https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits).
# - **tf.reduce_mean:** computes the mean of elements across dimensions of a tensor. Use this to calculate the sum of the losses over all the examples to get the overall cost. You can check the full documentation [reduce_mean](https://www.tensorflow.org/api_docs/python/tf/reduce_mean).
#
# #### Details on softmax_cross_entropy_with_logits (optional reading)
# * Softmax is used to format outputs so that they can be used for classification. It assigns a value between 0 and 1 for each category, where the sum of all prediction values (across all possible categories) equals 1.
# * Cross Entropy is compares the model's predicted classifications with the actual labels and results in a numerical value representing the "loss" of the model's predictions.
# * "Logits" are the result of multiplying the weights and adding the biases. Logits are passed through an activation function (such as a relu), and the result is called the "activation."
# * The function is named `softmax_cross_entropy_with_logits` takes logits as input (and not activations); then uses the model to predict using softmax, and then compares the predictions with the true labels using cross entropy. These are done with a single function to optimize the calculations.
#
# ** Exercise**: Compute the cost below using the function above.
# +
# GRADED FUNCTION: compute_cost
def compute_cost(Z3, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (number of examples, 6)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
### START CODE HERE ### (1 line of code)
### END CODE HERE ###
return cost
# +
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
cost = compute_cost(Z3, Y)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(cost, {X: np.random.randn(4,64,64,3), Y: np.random.randn(4,6)})
print("cost = " + str(a))
# -
# **Expected Output**:
# ```
# cost = 2.91034
# ```
# ## 1.5 Model
#
# Finally you will merge the helper functions you implemented above to build a model. You will train it on the SIGNS dataset.
#
# **Exercise**: Complete the function below.
#
# The model below should:
#
# - create placeholders
# - initialize parameters
# - forward propagate
# - compute the cost
# - create an optimizer
#
# Finally you will create a session and run a for loop for num_epochs, get the mini-batches, and then for each mini-batch you will optimize the function. [Hint for initializing the variables](https://www.tensorflow.org/api_docs/python/tf/global_variables_initializer)
# #### Adam Optimizer
# You can use `tf.train.AdamOptimizer(learning_rate = ...)` to create the optimizer. The optimizer has a `minimize(loss=...)` function that you'll call to set the cost function that the optimizer will minimize.
#
# For details, check out the documentation for [Adam Optimizer](https://www.tensorflow.org/api_docs/python/tf/train/AdamOptimizer)
# #### Random mini batches
# If you took course 2 of the deep learning specialization, you implemented `random_mini_batches()` in the "Optimization" programming assignment. This function returns a list of mini-batches. It is already implemented in the `cnn_utils.py` file and imported here, so you can call it like this:
# ```Python
# minibatches = random_mini_batches(X, Y, mini_batch_size = 64, seed = 0)
# ```
# (You will want to choose the correct variable names when you use it in your code).
# #### Evaluating the optimizer and cost
#
# Within a loop, for each mini-batch, you'll use the `tf.Session` object (named `sess`) to feed a mini-batch of inputs and labels into the neural network and evaluate the tensors for the optimizer as well as the cost. Remember that we built a graph data structure and need to feed it inputs and labels and use `sess.run()` in order to get values for the optimizer and cost.
#
# You'll use this kind of syntax:
# ```
# output_for_var1, output_for_var2 = sess.run(
# fetches=[var1, var2],
# feed_dict={var_inputs: the_batch_of_inputs,
# var_labels: the_batch_of_labels}
# )
# ```
# * Notice that `sess.run` takes its first argument `fetches` as a list of objects that you want it to evaluate (in this case, we want to evaluate the optimizer and the cost).
# * It also takes a dictionary for the `feed_dict` parameter.
# * The keys are the `tf.placeholder` variables that we created in the `create_placeholders` function above.
# * The values are the variables holding the actual numpy arrays for each mini-batch.
# * The sess.run outputs a tuple of the evaluated tensors, in the same order as the list given to `fetches`.
#
# For more information on how to use sess.run, see the documentation [tf.Sesssion#run](https://www.tensorflow.org/api_docs/python/tf/Session#run) documentation.
# +
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.009,
num_epochs = 100, minibatch_size = 64, print_cost = True):
"""
Implements a three-layer ConvNet in Tensorflow:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Arguments:
X_train -- training set, of shape (None, 64, 64, 3)
Y_train -- test set, of shape (None, n_y = 6)
X_test -- training set, of shape (None, 64, 64, 3)
Y_test -- test set, of shape (None, n_y = 6)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
train_accuracy -- real number, accuracy on the train set (X_train)
test_accuracy -- real number, testing accuracy on the test set (X_test)
parameters -- parameters learnt by the model. They can then be used to predict.
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
tf.set_random_seed(1) # to keep results consistent (tensorflow seed)
seed = 3 # to keep results consistent (numpy seed)
(m, n_H0, n_W0, n_C0) = X_train.shape
n_y = Y_train.shape[1]
costs = [] # To keep track of the cost
# Create Placeholders of the correct shape
### START CODE HERE ### (1 line)
### END CODE HERE ###
# Initialize parameters
### START CODE HERE ### (1 line)
### END CODE HERE ###
# Forward propagation: Build the forward propagation in the tensorflow graph
### START CODE HERE ### (1 line)
### END CODE HERE ###
# Cost function: Add cost function to tensorflow graph
### START CODE HERE ### (1 line)
### END CODE HERE ###
# Backpropagation: Define the tensorflow optimizer. Use an AdamOptimizer that minimizes the cost.
### START CODE HERE ### (1 line)
### END CODE HERE ###
# Initialize all the variables globally
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
with tf.Session() as sess:
# Run the initialization
sess.run(init)
# Do the training loop
for epoch in range(num_epochs):
minibatch_cost = 0.
num_minibatches = int(m / minibatch_size) # number of minibatches of size minibatch_size in the train set
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
"""
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the optimizer and the cost.
# The feedict should contain a minibatch for (X,Y).
"""
### START CODE HERE ### (1 line)
### END CODE HERE ###
minibatch_cost += temp_cost / num_minibatches
# Print the cost every epoch
if print_cost == True and epoch % 5 == 0:
print ("Cost after epoch %i: %f" % (epoch, minibatch_cost))
if print_cost == True and epoch % 1 == 0:
costs.append(minibatch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# Calculate the correct predictions
predict_op = tf.argmax(Z3, 1)
correct_prediction = tf.equal(predict_op, tf.argmax(Y, 1))
# Calculate accuracy on the test set
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print(accuracy)
train_accuracy = accuracy.eval({X: X_train, Y: Y_train})
test_accuracy = accuracy.eval({X: X_test, Y: Y_test})
print("Train Accuracy:", train_accuracy)
print("Test Accuracy:", test_accuracy)
return train_accuracy, test_accuracy, parameters
# -
# Run the following cell to train your model for 100 epochs. Check if your cost after epoch 0 and 5 matches our output. If not, stop the cell and go back to your code!
_, _, parameters = model(X_train, Y_train, X_test, Y_test)
# ## Use Tensorflow 2.x
def initialize_parameters():
"""
Initializes weight parameters to build a neural network with tensorflow. The shapes are:
W1 : [4, 4, 3, 8]
W2 : [2, 2, 8, 16]
Note that we will hard code the shape values in the function to make the grading simpler.
Normally, functions should take values as inputs rather than hard coding.
Returns:
parameters -- a dictionary of tensors containing W1, W2
"""
initializer = tf.initializers.GlorotUniform(0)
W1 = tf.Variable(initializer([4, 4, 3, 8])) # CONV2D 1
W2 = tf.Variable(initializer([2, 2, 8, 16])) # CONV2D 2
W3 = tf.Variable(initializer([2*2*16, 6])) # FC
b1 = tf.Variable(tf.zeros([8])) # CONV2D 1
b2 = tf.Variable(tf.zeros([16])) # CONV2D 2
b3 = tf.Variable(tf.zeros([6])) # FC
return {
'W1': W1,
'W2': W2,
'W3': W3,
'b1': b1,
'b2': b2,
'b3': b3
}
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Note that for simplicity and grading purposes, we'll hard-code some values
such as the stride and kernel (filter) sizes.
Normally, functions should take these values as function parameters.
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "W2"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
W1 = parameters['W1']
W2 = parameters['W2']
W3 = parameters['W3']
b1 = parameters['b1']
b2 = parameters['b2']
b3 = parameters['b3']
# CONV2D. Output shape: (n, 61, 61, 8)
X = tf.nn.conv2d(X, W1, strides=[1,1,1,1], padding='SAME')
X = tf.nn.bias_add(X, b1)
# RELU
X = tf.nn.relu(X)
# MAXPOOL. Output shape: (n, 8, 8, 8)
X = tf.nn.max_pool(X, ksize=[1,8,8,1], strides=[1,8,8,1], padding='SAME')
# CONV2D. Output shape: (n, 7, 7, 16)
X = tf.nn.conv2d(X, W2, strides=[1,1,1,1], padding='SAME')
X = tf.nn.bias_add(X, b2)
# RELU
X = tf.nn.relu(X)
# MAXPOOL. Output shape: (n, 2, 2, 16)
X = tf.nn.max_pool(X, ksize=[1,4,4,1], strides=[1,4,4,1], padding='SAME')
# FLATTEN Output shape: (n, 64)
X = tf.reshape(X, [-1, W3.get_shape().as_list()[0]])
# FULLYCONNECTED. Output shape: (n, 6)
X = tf.add(tf.matmul(X, W3), b3)
return X
def compute_cost(Y_hat, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (number of examples, 6)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(Y, Y_hat))
return cost
def run_optimization(X, Y, parameters, optimizer):
with tf.GradientTape() as tape:
Y_hat = forward_propagation(X, parameters)
cost = compute_cost(Y_hat, Y)
trainable_var = list(parameters.values())
gradients = tape.gradient(cost, trainable_var)
optimizer.apply_gradients(zip(gradients, trainable_var))
return cost
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.009,
num_epochs = 100, minibatch_size = 64, print_cost = True):
"""
Implements a three-layer ConvNet in Tensorflow:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Arguments:
X_train -- training set, of shape (None, 64, 64, 3)
Y_train -- test set, of shape (None, n_y = 6)
X_test -- training set, of shape (None, 64, 64, 3)
Y_test -- test set, of shape (None, n_y = 6)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
train_accuracy -- real number, accuracy on the train set (X_train)
test_accuracy -- real number, testing accuracy on the test set (X_test)
parameters -- parameters learnt by the model. They can then be used to predict.
"""
tf.random.set_seed(1) # to keep results consistent (tensorflow seed)
seed = 3 # to keep results consistent (numpy seed)
(m, n_H0, n_W0, n_C0) = X_train.shape
costs = [] # To keep track of the cost
# Initialize parameters
parameters = initialize_parameters()
# Initialize optimizer
optimizer = tf.optimizers.Adam(learning_rate)
# Do the training loop
for epoch in range(num_epochs):
minibatch_cost = 0.
# number of minibatches of size minibatch_size in the train set
num_minibatches = int(m / minibatch_size)
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
"""
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the optimizer and the cost.
# The feedict should contain a minibatch for (X,Y).
"""
### START CODE HERE ### (1 line)
temp_cost = run_optimization(minibatch_X, minibatch_Y, parameters, optimizer)
### END CODE HERE ###
minibatch_cost += temp_cost / num_minibatches
# Print the cost every epoch
if print_cost == True and epoch % 5 == 0:
print ("Cost after epoch %i: %f" % (epoch, minibatch_cost))
if print_cost == True and epoch % 1 == 0:
costs.append(minibatch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
Y_train_pred = forward_propagation(X_train, parameters)
train_accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(Y_train_pred, 1),
tf.argmax(Y_train, 1)), 'float'))
Y_test_pred = forward_propagation(X_test, parameters)
test_accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(Y_test_pred, 1),
tf.argmax(Y_test, 1)), 'float'))
print("Train Accuracy:", train_accuracy)
print("Test Accuracy:", test_accuracy)
return train_accuracy, test_accuracy, parameters
_, _, parameters = model(X_train, Y_train, X_test, Y_test)
# **Expected output**: although it may not match perfectly, your expected output should be close to ours and your cost value should decrease.
#
# <table>
# <tr>
# <td>
# **Cost after epoch 0 =**
# </td>
# <td>
# 1.917929
# </td>
# </tr>
# <tr>
# <td>
# **Cost after epoch 5 =**
# </td>
# <td>
# 1.506757
# </td>
# </tr>
# <tr>
# <td>
# **Train Accuracy =**
# </td>
# <td>
# 0.940741
# </td>
# </tr>
#
# <tr>
# <td>
# **Test Accuracy =**
# </td>
# <td>
# 0.783333
# </td>
# </tr>
# </table>
# Congratulations! You have finished the assignment and built a model that recognizes SIGN language with almost 80% accuracy on the test set. If you wish, feel free to play around with this dataset further. You can actually improve its accuracy by spending more time tuning the hyperparameters, or using regularization (as this model clearly has a high variance).
#
# Once again, here's a thumbs up for your work!
# +
fname = "images/thumbs_up.jpg"
image = Image.open(fname).resize((64, 64))
my_image = np.expand_dims(np.array(image), 0) / 255.
my_image_prediction = np.argmax(forward_propagation(my_image, parameters), axis=1)[0]
plt.imshow(image)
print("Your algorithm predicts: y = " + str(my_image_prediction))
# -
| 33,073 |
/Deep Learning/tensorflow_MNIST_-_Assaf_B._Spanier_ZogoZqs.ipynb | a40130ab6615514c377a81b7577c24dd8a1dcd30 | [] | no_license | shalomazar/Personal-Projects | https://github.com/shalomazar/Personal-Projects | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 10,612 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `pandas`
# %pylab inline
plt.style.use('ggplot')
import numpy as np
import pandas as pd
# ## Creating `pd.Series`
s = pd.Series(np.random.randn(10))
s
s = pd.Series(np.random.randn(10), name="random_series")
s
s = pd.Series(np.random.randn(10), name="random_series",
index=np.random.randint(23, size=(10,)))
s
s.index
s = pd.Series(np.random.randn(10), name="random_series",
index=pd.Index(np.random.randint(23, size=(10,)), name="main_index"))
s
s.index
s
# Index with a single occurence
s[8]
# Index with multiple occurences
s[[21]]
s
s[21].iloc[0]
s.iloc[1]
s = pd.Series({'a':3, 'c':6, 'b':2}, name="dict_series")
s
s.index
s[0], s['a'], s.iloc[0]
s['a':'c']
s.sort_index()['a':'c']
# ## Creating `pd.DataFrame`
df = pd.DataFrame(np.arange(20).reshape((5,4)))
df
df = pd.DataFrame(np.arange(20).reshape((5,4)),
columns=['a', 'b', 'c', 'd'])
df
import string
df = pd.DataFrame(np.arange(20).reshape((5,4)),
columns=['a', 'b', 'c', 'd'],
index=np.random.choice(list(string.ascii_lowercase), 5, replace=False))
df
df.columns
df.index
df['a']
df['z']
type(df['a'])
df.loc[["z"]]
df.loc[["z"]]
df.loc[["z"], "a"]
df
# +
# Indexing with right and wrong order
# df['n':'r']
df['r':'z']
# -
# # Indexing `pd.Series`
N_ELEMS = 20
s = pd.Series(np.random.randint(20, size=(N_ELEMS,)),
index=list(string.ascii_lowercase)[:N_ELEMS],
name='randint_series')
s
# ## Indexing with `[]`
s
s['i'] # But there's a caveat: it may be series or just an element
s[['i']]
s['a':'z']
s[['k', 'q', 'a', 'r']]
s[5:3:-1]
# ## Indexing with `.loc`
s_int_idx = pd.Series(np.random.randint(20, size=(N_ELEMS,)),
index=np.random.choice(N_ELEMS, N_ELEMS, replace=False),
name='randint_series')
s_int_idx
s_int_idx[2:15]
s_int_idx[2]
s_int_idx[2:5]
s_int_idx[s_int_idx.index.isin(range(2,6))]
s_int_idx.loc[2:5]
s_int_idx.iloc[2:5]
s_int_idx.loc[2:456]
s_int_idx.mean()
s_sorted = s_int_idx.sort_index()
s_sorted.index.is_monotonic
s_sorted.loc[2:5]
s_trick = pd.Series([1,2,3,4], index=[2,3,4,4])
s_trick.loc[2:4]
s_trick
s_trick.index.is_monotonic
s_int_idx.sort_index().loc[2:6]
s_int_idx
s_int_idx[s_int_idx.index!=11]
s_int_idx[(s_int_idx>15) | (s_int_idx<5)]
s_int_idx.loc[s_int_idx!=14]
s_int_idx
s_int_idx[s_int_idx.index!=11]
# ## Indexing with `.iloc`
s
s.iloc[5:8]
s.shape
s.iloc[15:20]
s.iloc[[3, 9, 8]]
s
s.iloc[[3, 9]]
s.loc[["d", "j"]]
# # Indexing `pd.DataFrame`
df = pd.DataFrame(np.arange(20).reshape((5,4)),
columns=['d', 'c', 'b', 'a'],
index=np.random.choice(list(string.ascii_lowercase), 5, replace=False))
df
df[2:5]
df['e'] # Nope, it doesn't work that way
df['a']
df[['b']]
df
df.columns
df[df.columns[2:]]
df.iloc[:, 2:]
df
df[:'e'] # Surprising!
df['s':'g'] # Not really surprising
df.sort_index()['s':'g']
df['g':]
df.sort_index()['g':]
df
df["s":'z'] # No, that won't work
df.sort_index()['k':'zjyyf']
df.index.to_series().rank()
df.sort_index().index.to_series().rank()
df
df['e':'s']['c'] # Not a very good idea
df[(df['a']>12) | (df['b']<3)]
df
# ## Indexing with `.loc`
df
df.loc['s']
df.loc['s', 'b']
df.loc['g':, 'b']
df.loc['e':, 'b':]
df
df.loc['e':, 'c':'d'].shape
df
df.columns.to_series().rank()
df.loc['e':, 'a'::-2]
df
df.sort_index(axis='columns').loc['e':, 'c':'d']
df.loc[:, ["a", "b"]]
df.loc["x", "b"] = 20
df.loc["h", "c"] = 30
df
df[["a", "b"]].rank()
df.loc[["e", "g", "s"]]
df.loc[["e", "g", "s"]].rank()
df = pd.DataFrame(np.arange(20).reshape((5,4)),
columns=['d', 'c', 'a', 'b'],
index=np.random.choice(list(string.ascii_lowercase), 5, replace=False))
df
df
df.loc['v':, 'b':'d']
df.columns.to_series().rank()
df.sort_index(axis='columns').loc['v':, 'c':'sflng']
df.sort_index(axis='columns').columns.to_series().rank()
df
df.loc[df['c']>10, 'c']
df.loc[[1,2], 'c'] # This won't work
# ## Indexing with `.iloc`
df.iloc[:3]
df.iloc[1:3]
df.iloc[1:3, 'b'] # This won't work
df.iloc[1:3, 2:]
df
df.values
df.iloc[(df['a']>10).values, 2:]
# ## `SettingWithCopyWarning`
df
df.loc[df['a']>10, 'c'] = 10
df
df.loc[df['a']>5]['c'] = 20.
df
df_1 = df.loc[df['a']>10]
df_1.is_copy
df_1['c'] = 25
df_1
# # Dataframe arithmetic
df_1 = pd.DataFrame(np.arange(40).reshape(10,4),
columns=['a', 'b', 'c', 'd'],
index=np.random.choice(list(string.ascii_lowercase), 10, replace=False))
df_1
df_2 = pd.DataFrame(np.arange(40).reshape(10,4),
columns=['a', 'e', 'c', 'd'],
index=np.random.choice(list(string.ascii_lowercase), 10, replace=False))
df_2
# A lot of missing values
df_1 + df_2
df_1.add(df_2, fill_value=0)
s_1 = pd.Series(np.arange(10),
name='f',
index=np.random.choice(list(string.ascii_lowercase), 10, replace=False))
s_1
df_1 + s_1
s_1 + df_1
s_1.add(df_1, axis='columns')
df1 = df.astype(np.float).copy()
df1.loc["h", "b"] = np.nan
df1
(df1 - df1.mean()) / df1.std()
# # Applying functions to dataframes
df
df.iloc[0]
df.apply(lambda x: np.sqrt(x.d), axis=1)
# %timeit df['d'].apply(lambda x: np.sqrt(x))
np.sqrt(df['d'])
# %timeit np.sqrt(df['d'])
# Better way
# %timeit np.sqrt(df['d'].values)
df
df.values
df.values.sum(axis=1)
df.apply(lambda x: x.sum(), axis=1)
np.sum(df, axis=1)
df.apply(lambda x: [x.d, x.b], axis=1)
dfm = df.apply(lambda x: pd.Series({'sum': x.sum(),
'sqrt': np.sqrt(x['d'])}),
axis=1)
dfm
dfm["sum"] = dfm["sum"].astype(np.int)
dfm.dtypes
dfm[dfm.columns[dfm.dtypes==np.int64]]
# # Reading CSV files
#
# We will use [Titanic dataset](https://www.kaggle.com/c/titanic/data), which is located in `data` directory.
titanic_train = pd.read_csv('data/train.csv')
titanic_train.head()
titanic_train.tail()
titanic_train.info()
titanic_train.describe()
# %timeit titanic_train[titanic_train.PassengerId==400]
titanic_train = pd.read_csv('data/train.csv', index_col='PassengerId')
titanic_test = pd.read_csv('data/test.csv', index_col='PassengerId')
# %timeit titanic_train.loc[400]
titanic = pd.concat([titanic_train, titanic_test])
titanic
titanic = pd.concat([titanic_train, titanic_test], sort=False)
titanic.head()
titanic.tail()
# # Dataframe statistics
titanic.index.is_unique, titanic.index.is_monotonic
titanic['Pclass'].value_counts()
titanic[titanic.Ticket=="CA 2144"]
titanic[titanic.SibSp==5]
titanic.SibSp.value_counts()
titanic.Embarked.value_counts() # S = Southampton, C = Cherbourg, Q = Queens Town
titanic.Sex.value_counts()
print("Average age: %2.2f" % titanic['Age'].mean())
print("STD of age: %2.2f" % titanic['Age'].std())
print("Minimum age: %2.2f" % titanic['Age'].min())
print("Maximum age: %2.2f" % titanic['Age'].max())
print("Average number of siblings/spouse: %2.2f" % titanic['SibSp'].mean())
print("Average number of siblings/spouse in class 1: %2.2f" % titanic.loc[titanic.Pclass==1, 'SibSp'].mean())
print("Average number of siblings/spouse in class 2: %2.2f" % titanic.loc[titanic.Pclass==2, 'SibSp'].mean())
print("Average number of siblings/spouse in class 3: %2.2f" % titanic.loc[titanic.Pclass==3, 'SibSp'].mean())
print("Minimum age (not survived): %2.2f" % titanic.loc[titanic.Survived==0, 'Age'].min())
print("Maximum age (not survived): %2.2f" % titanic.loc[titanic.Survived==0, 'Age'].max())
print("Minimum age (not survived): %2.2f" % titanic.loc[titanic.Survived==1, 'Age'].min())
print("Maximum age (not survived): %2.2f" % titanic.loc[titanic.Survived==1, 'Age'].max())
# # Announcing the future
titanic.groupby('Pclass')
titanic.groupby('Pclass').size()
titanic.Pclass.value_counts()
titanic["AgeGroup"] = 5 + 10*(titanic.Age//10)
group_counts = titanic.groupby(['Pclass', 'AgeGroup', 'Sex']).size()/titanic.groupby('Pclass').size()
group_counts[[(1, 5.0, "female"), (1, 15.0, "female")]]
100*group_counts
group_counts = group_counts.unstack()
group_counts.unstack(level=1)
group_counts
group_counts.loc[1]
# +
plt.figure(figsize=(15, 5))
for pclass in [1, 2, 3]:
plt.subplot(1, 3, pclass)
group_counts.loc[pclass].plot(ax=plt.gca())
plt.ylim(0, 0.25)
plt.title("Age distribution for Class %i" % pclass, fontsize=12)
plt.tight_layout()
# -
survival_groups = titanic.groupby(['Pclass', 'AgeGroup', 'Sex']).Survived.mean()
survival_groups = survival_groups.unstack()
# +
plt.figure(figsize=(15, 5))
for pclass in [1, 2, 3]:
plt.subplot(1, 3, pclass)
survival_groups.loc[pclass].plot(ax=plt.gca())
plt.ylim(0, 1)
plt.title("Survived in class %i" % pclass, fontsize=12)
plt.tight_layout()
# -
siblings_groups = titanic.groupby(['Pclass', 'AgeGroup', 'Sex']).SibSp.mean()
siblings_groups = siblings_groups.unstack()
# +
plt.figure(figsize=(15, 5))
for pclass in [1, 2, 3]:
plt.subplot(1, 3, pclass)
siblings_groups.loc[pclass].plot(ax=plt.gca())
plt.ylim(0, 5)
plt.title("Siblings in class %i" % pclass, fontsize=12)
plt.tight_layout()
# -
embark_counts = titanic.groupby(['Pclass', 'AgeGroup', 'Sex', 'Embarked']).size()/titanic.groupby('Pclass').size()
embark_counts = embark_counts.unstack([-1, -2])
embark_counts
embark_counts.loc[3, (('S', 'male'))]
# # Replacing and renaming
# +
# titanic.replace?
# -
titanic.replace(22, 122).head()
import re
titanic.replace(re.compile(r'\(.*\)'), '').head()
titanic.rename(lambda x: x.lower(), axis=1).head()
titanic.rename({'SibSp':'siblings_spouses'}, axis=1).head()
# ## String operations
titanic.head()
(titanic
.replace(re.compile(r'\(.*\)'), '')
.Name.str
.split(',', expand=True)
.rename({0:'family_name', 1:'first_name'}, axis=1)
.head())
# # Cleaning data
titanic.isnull().head()
titanic.isnull().any()
titanic.isnull().any(axis=1).head()
titanic.isnull().sum()
titanic.head(15)
fill_values = titanic[['Age', 'Fare']].mean()
titanic[titanic.Fare.isnull()]
titanic.fillna(fill_values).fillna({'AgeGroup':25.}).head(15)
# # Getting indicators and dummy variables
pd.get_dummies(titanic, columns=['Pclass', 'Sex', 'Embarked']).head()
0eea9594b589149d753a37edb4c", "56085ce0a5ca45f4a59ebf90561b2b21", "d814c04a15204256a70f36c5634b8ac9", "20152c8c04b44688b5c6032093a8db64", "133a3f00814543fab795470e284b24d9"]} id="fD2wnL9cAhnk" outputId="a2cff16f-7f27-4852-e0a2-bfe51e2f59f4"
# %%time
history += fit_one_cycle(epochs, max_lr, model, train_dl, valid_dl,
grad_clip=grad_clip,
weight_decay=weight_decay,
opt_func=opt_func)
# + id="tLLnBYWgAjxc"
model.unfreeze()
# + id="BTPy8l7AAlWq" colab={"base_uri": "https://localhost:8080/", "height": 378, "referenced_widgets": ["cfc206834b7f449dafc353705bfd1336", "50f90f286a4948bdb5fe401bf5806e2b", "8abd3b7f9bae47788015a1ee41eb9330", "454f13441a634ab1ad7d78abfba89b9b", "dae340c28a2c4e9f8cd941360287f6d2", "3550c051469348a09f5e789dab523e53", "55c47627bd4d4941ae6a99b6bd98e906", "d3512aaa700842a59b340569d79ee57c", "bde15bbee0c84b90b32ff07a5c327301", "4a3f64b472484dab8c32efda4253e101", "4ca42e9238814aba847bd35c04470c6f", "9f73b0f6fd67497194ed25e747d1a255", "97d09e3fae454524bbcd3f1ed1253b42", "4e7cd19bce7c4f568fc9174b7b274f08", "f423a15ca107429e9f0bb3eaecede0d8", "294df13022034a4ea1dcb8ca373b877a", "393ae52507d04cd38caa51d01cb87a5c", "df7ada302e4e49d2ae643b02ca59521e", "f8b6a45add4f40ae92888246153355a1", "09a67e6b48bd47c49a18fe6f7632e201", "4138101e915a4e8cb1b4280e08fe69ce", "f6477bd226eb4cb3a582a92757d2dcde", "e97745bd6e8f4867aa2764f4dfb559a7", "d47bfc1069bf472a85c0198802ef26d8", "e88fb540217545499540671a44c4a567", "f1ddd8cee959471eb076751dc5e9d685", "98dcec89825042dfbe1e87912b854bec", "d8988461651d4b5bb5b74a3ee241aa26", "4f4be63bbcbb41f082042bd5b554a6d2", "151771cf96de45a9b5d3820b209cd582", "02ec310ade4347a3ac0cb7994b097e63", "17b0a223a2bc42328681d502d1dd50fb", "a1e1012376654a1b8368a3948f8445c3", "3d32c7909b6f49639ad1172a881bbea9", "b8a1a9c289ae446d9e7bad3ddba6bb37", "211b014fa3444c3780be6394c74ade55", "a1f15ea47ea64c1f99c9ac1948317299", "77bacc2d99c24e8cb5ddd80ebe2db4b9", "119825eb9fc44b92b4126fc02bd94c43", "9240422cb99d4752b4e81f7b8406f28f"]} outputId="82878cf9-a921-47a0-9a76-9ec0db871758"
# %%time
history += fit_one_cycle(epochs, 0.001, model, train_dl, valid_dl,
grad_clip=grad_clip,
weight_decay=weight_decay,
opt_func=opt_func)
# + id="N1O8nh_PAoHV" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="92aea5b7-5661-4e6c-de0e-86e655c6b6df"
def plot_scores(history):
scores = [x['val_acc'] for x in history]
plt.plot(scores, '-x')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.title('F1 accuracy vs. No. of epochs');
plot_scores(history)
# + id="jWq14uuXAruI" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="735ec571-50b0-448d-cfe5-ee90a5a49a1a"
def plot_losses(history):
train_losses = [x.get('train_loss') for x in history]
val_losses = [x['val_loss'] for x in history]
plt.plot(train_losses, '-bx')
plt.plot(val_losses, '-rx')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['Training', 'Validation'])
plt.title('Loss vs. No. of epochs');
plot_losses(history)
# + id="NPfbYMoUAvO4" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="ceab8d51-832f-47f9-ee2e-30e84d76ffc3"
def plot_lrs(history):
lrs = np.concatenate([x.get('lrs', []) for x in history])
plt.plot(lrs)
plt.xlabel('Batch no.')
plt.ylabel('Learning rate')
plt.title('Learning Rate vs. Batch no.');
plot_lrs(history)
| 14,050 |
/starbucks_capstone/data_exploration.ipynb | 6fd4ec672ccbe83e8688a196dcea9df7b9491392 | [] | no_license | breakAwall/ml_lessons | https://github.com/breakAwall/ml_lessons | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 411,163 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
import pandas as pd
import numpy as np
import math
import json
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import scipy.stats as stats
import seaborn as sns
# %matplotlib inline
# read in the json files
portfolio = pd.read_json('data/portfolio.json', orient='records', lines=True).set_index('id')
profile = pd.read_json('data/profile.json', orient='records', lines=True).set_index('id')
transcript = pd.read_json('data/transcript.json', orient='records', lines=True)
# analyze profile table
print(profile.describe())
print(profile.gender.value_counts())
print(profile.isna().sum())
profile.became_member_on = pd.to_datetime(profile['became_member_on'], format="%Y%m%d")
# there is missing data, is it all in the same rows
print("Profile Unidentified Length")
unidentified = profile[profile.age > 102].copy()
print(unidentified.shape[0])
print(profile.shape[0])
print("Missing Percent: " + str(round(unidentified.shape[0]/profile.shape[0] * 100, 2)))
# verify that people with valid age have other fields as valid
identified = profile[profile.age <= 102]
identified.isna().sum()
# #### Plot "profile" distributions
# +
fig, ax = plt.subplots(3,2, figsize=(10,8))
fig.tight_layout()
ax[0, 0].hist(profile.age, bins=30)
ax[0, 0].set_title("Age")
ax[1, 0].hist(profile.income, bins=30)
ax[1, 0].set_title("Income")
ax[2, 0].hist(profile.became_member_on, bins=30)
ax[2, 0].set_title("Join Date")
years_fmt = mdates.DateFormatter('%Y')
ax[2, 0].xaxis.set_major_formatter(years_fmt)
ax[0, 1].hist(identified.age, bins=30)
ax[0, 1].set_title("Age")
ax[1, 1].hist(identified.income, bins=30)
ax[1, 1].set_title("Income")
ax[2, 1].hist(identified.became_member_on, bins=30)
ax[2, 1].set_title("Join Date")
ax[2, 1].xaxis.set_major_formatter(years_fmt)
plt.savefig("images/profile_hist.png")
# -
profile = profile.join(pd.get_dummies(profile.gender), how='inner')
profile.drop('gender', axis=1, inplace=True)
profile.became_member_on = pd.to_datetime(profile['became_member_on'], format="%Y%m%d")
identified = profile[profile.age <= 102]
profile.head(2)
fig, ax = plt.subplots()
ax.bar(['Female', 'Male', 'Other'], identified[['F', 'M', 'O']].sum(), color=['red', 'green', 'blue'])
plt.savefig("images/gender_distribution.png")
print(portfolio.describe())
portfolio.head(2)
# Lets break channels out into one-hot vectors
# +
email = {}
mobile = {}
social = {}
web = {}
for index, row in portfolio.iterrows():
if 'email' in row['channels']:
email[index] = 1
else:
email[index] = 0
if 'mobile' in row['channels']:
mobile[index] = 1
else:
mobile[index] = 0
if 'social' in row['channels']:
social[index] = 1
else:
social[index] = 0
if 'web' in row['channels']:
web[index] = 1
else:
web[index] = 0
email = pd.Series(email)
email.name = 'email'
mobile = pd.Series(mobile)
mobile.name = 'mobile'
social = pd.Series(social)
social.name = 'social'
web = pd.Series(web)
web.name = 'web'
portfolio = pd.concat([portfolio, email, mobile, social, web], axis=1)
# -
portfolio.drop('channels', axis=1, inplace=True)
portfolio.head(2)
print(f'We have {portfolio.shape[0]} different kinds of offers')
fig, ax = plt.subplots()
names = ['email', 'mobile', 'social', 'web']
ax.bar(names, portfolio[names].sum(), color=['red', 'green', 'blue', 'cyan'])
plt.savefig("images/promotion_channels.png")
# #### Plot the "portfolio" data distributions
fig, ax = plt.subplots(3, figsize=(6,8))
fig.tight_layout()
ax[0].hist(portfolio.reward, bins=30)
ax[0].set_title("reward")
ax[1].hist(portfolio.difficulty, bins=30)
ax[1].set_title("difficulty")
ax[2].hist(portfolio.duration, bins=30)
ax[2].set_title("duration (Days)")
plt.savefig("images/promotion_hist.png")
print(transcript.describe())
print(transcript.event.value_counts())
# modify trascript to include when transactions were in an offer period had been viewed
# this value will indicate wether the transaction in question happened due to a running offer.
def transaction_state(df, impression_colname, impression_source_colname):
""" input
df: transcript dataframe, has information on when users transacted and when they received offers
colname: name of new column that will have impression - transaction or not and if so is it because of offer
"""
customer_ids = set(df.person.values)
impression = {}
source = {}
# iterate over all customers
for customer in customer_ids:
customer_transcript = df[df.person == customer]
received_offer = {}
active_dict = {}
max_active_end = 0 # the end time of the event that is ending the latest
cur_max_offer_id = "" # id of the offer with max active end
for index, row in customer_transcript.iterrows():
time = row['time']
# eliminate adverts not active from active_dict
for key in list(active_dict.keys()):
if active_dict[key] < time:
del active_dict[key]
# if new offer, record when it will expire in received_offer
if row['event'] == "offer received":
new_event_end = row['time'] + portfolio.loc[
row['value']['offer id'] == portfolio.index]['duration'].values[0] * 24
received_offer[row['value']['offer id']] = new_event_end
# after viewing an offer we add it to the active_dict and possibly update the time until all transactions
# will be flagged as being due to an offer
if row['event'] == "offer viewed":
# error in data if element not in dict (viewed an offer that hasn't been received)
new_event_end = received_offer[row['value']['offer id']]
active_dict[row['value']['offer id']] = new_event_end
if new_event_end > max_active_end:
max_active_end = new_event_end
cur_max_offer_id = row['value']['offer id']
if row['event'] == 'offer completed':
offer_id = row['value']['offer_id']
# an unviewed offer could be completed
active_dict.pop(offer_id, None)
if offer_id == cur_max_offer_id:
# find next offer with highest end date
new_max_active_end = 0
new_max_offer_id = ""
for key, value in active_dict.items():
if value > new_max_active_end:
new_max_active_end = value
new_max_offer_id = key
max_active_end = new_max_active_end
new_max_offer_id = new_max_offer_id
# assign values in new col dict
if row['event'] == 'transaction':
if time <= max_active_end:
impression[index] = 1
else:
impression[index] = 0
this_source = []
for key, value in active_dict.items():
this_source.append(key)
source[index] = this_source
else:
impression[index] = -1
source[index] = []
df = df.copy()
df[impression_colname] = pd.Series(impression)
df[impression_source_colname] = pd.Series(source)
return df
impressions_df = transaction_state(transcript, 'because_of_offer', 'impression_source')
# +
# test the transaction state method.
test1 = transcript.loc[transcript.person == "78afa995795e4d85b5d9ceeca43f5fef"]
test1_return = transaction_state(test1, 'because_of_offer', 'impression_source')
test1_expected = {0: -1, 15561: -1, 47582: 1, 47583: -1, 49502: 0, 53176: -1, 85291: -1, 87134: 1,
92104: 1, 141566: 0, 150598: -1, 163375: -1, 201572: -1, 218393: 1, 218394: -1,
218395: -1, 230412: 0, 262138: -1}
test1_expected_source = {0: [], 15561: [], 47582: ['9b98b8c7a33c4b65b9aebfe6a799e6d9'], 47583: [], 49502: [],
53176: [], 85291: [], 87134: ['5a8bc65990b245e5a138643cd4eb9837'],
92104: ['5a8bc65990b245e5a138643cd4eb9837'], 141566: [], 150598: [], 163375: [],
201572: [], 218393: ['ae264e3637204a6fb9bb56bc8210ddfd'], 218394: [], 218395: [],
230412: [], 262138: []}
assert((test1_return.because_of_offer == pd.Series(test1_expected)).all())
assert((test1_return.impression_source == pd.Series(test1_expected_source)).all())
test2 = transcript.loc[transcript.person == "2e87ba0fba1a4d1a8614af771f07a94d"]
test2_return = transaction_state(test2, 'because_of_offer', 'impression_source')
test2_expected = {45: -1, 24223: -1, 31109: 1, 31110: -1, 42034: 0, 53222: -1, 65856: -1, 65857: 1,
65858: -1, 74812: 0, 106958: 0, 110873: -1, 123553: -1, 135232: 1, 135233: -1,
143579: 0, 150644: -1, 163387: -1, 174370: 1, 174371: -1, 187146: 0, 201615: -1,
221912: -1, 232884: 1, 235177: 1, 300661: 0}
assert((test2_return.because_of_offer == pd.Series(test2_expected)).all())
print("tests passed")
# -
impressions_df.to_csv("data/impressions.csv")
identified.to_csv("data/identified.csv")
impressions_df.tail()
# +
# see distribution of transactions overall when in offer and when not in offer
transactions_df = impressions_df.loc[impressions_df['because_of_offer'] > -1].copy()
transactions_df['value'] = transactions_df['value'].apply(lambda x: x['amount'])
without_offer = transactions_df[transactions_df.because_of_offer == 0]
with_offer = transactions_df[transactions_df.because_of_offer == 1]
without_offer_sum = without_offer.groupby('person').sum(1)
with_offer_sum = with_offer.groupby('person').sum(1)
with_offer_sum = with_offer_sum[stats.zscore(with_offer_sum.value) < 2]
without_offer_sum = without_offer_sum[stats.zscore(without_offer_sum.value) < 2]
fig, ax = plt.subplots(figsize=(8,8))
ax.hist([with_offer_sum.value.values, without_offer_sum.value.values], bins=40,
label=['with offer', 'without offer'])
ax.legend()
plt.savefig("images/revenue_diff.png")
# -
# As we would hope, it seems in general the availibility of offer does cause more revenue
def frequency_interaction(df, promotions_ids):
""" Get an encoding of individual and promotoins based on wether
the given individual saw/received the given promotion
Input:
df: The transcript table
promotoin_ids: An array containing id of all promotions - as given in the transcript table
"""
customer_ids = set(df.person.values)
viewed = {}
received = {}
raw_dict = {promo_id: 0 for promo_id in promotions_ids}
# iterate over all customers
for customer in customer_ids:
customer_transcript = df[df.person == customer]
person_dict_receive = raw_dict.copy()
person_dict_view = raw_dict.copy()
for index, row in customer_transcript.iterrows():
if row['event'] == "offer received":
person_dict_receive[row['value']['offer id']] += 1
if row['event'] == "offer viewed":
person_dict_view[row['value']['offer id']] += 1
viewed[customer] = person_dict_view
received[customer] = person_dict_receive
return pd.DataFrame(viewed), pd.DataFrame(received)
# +
test1 = transcript.loc[transcript.person == "78afa995795e4d85b5d9ceeca43f5fef"]
expected_receive1 = {'ae264e3637204a6fb9bb56bc8210ddfd': 1, '4d5c57ea9a6940dd891ad53e9dbe8da0': 0,
'3f207df678b143eea3cee63160fa8bed': 0, '9b98b8c7a33c4b65b9aebfe6a799e6d9': 1,
'0b1e1539f2cc45b7b9fa7c272da2e1d7': 0, '2298d6c36e964ae4a3e7e9706d1fb8c2': 0,
'fafdcd668e3743c1bb461111dcafc2a4': 0, '5a8bc65990b245e5a138643cd4eb9837': 1,
'f19421c1d4aa40978ebb69ca19b0e20d': 1, '2906b810c7d4411798c6938adc9daaa5': 0}
expected_view1 = {'ae264e3637204a6fb9bb56bc8210ddfd': 1, '4d5c57ea9a6940dd891ad53e9dbe8da0': 0,
'3f207df678b143eea3cee63160fa8bed': 0, '9b98b8c7a33c4b65b9aebfe6a799e6d9': 1,
'0b1e1539f2cc45b7b9fa7c272da2e1d7': 0, '2298d6c36e964ae4a3e7e9706d1fb8c2': 0,
'fafdcd668e3743c1bb461111dcafc2a4': 0, '5a8bc65990b245e5a138643cd4eb9837': 1,
'f19421c1d4aa40978ebb69ca19b0e20d': 1, '2906b810c7d4411798c6938adc9daaa5': 0}
view1, receive1 = frequency_interaction(test1, portfolio.index)
assert(view1.to_dict()['78afa995795e4d85b5d9ceeca43f5fef'] == expected_view1)
assert(receive1.to_dict()['78afa995795e4d85b5d9ceeca43f5fef'] == expected_receive1)
print('test passed')
# +
# determine if a given promotion influenced a certain individual
impressions_df_pos = impressions_df[impressions_df.because_of_offer == 1]
influenced = {}
customer_ids = set(impressions_df_pos.person.values)
raw_dict = {promo_id: 0 for promo_id in portfolio.index}
# iterate over all customers
for customer in customer_ids:
customer_impression = impressions_df_pos[impressions_df_pos.person == customer]
impression = raw_dict.copy()
for index, row in customer_impression.iterrows():
for promo_id in row['impression_source']:
impression[promo_id] = 1
influenced[customer] = impression
influenced_df = pd.DataFrame(influenced)
# -
viewed_df, received_df = frequency_interaction(transcript, portfolio.index)
# ### Investigate Relationships
# Lets see if distributions of influence, and receiving of promotions changes on characteristics of the individual/promotion
# +
fig, ax = plt.subplots(2,2, figsize=(8, 8))
received_df = received_df.T
received_counts = received_df.sum()
received_counts.name = 'received_counts'
received_total = pd.concat([portfolio, received_counts], axis=1)
_ = ax[0, 0].pie(received_total.received_counts)
_ = ax[0, 0].set_title("Frequency of Offers Received")
offer_type = received_total.groupby('offer_type').sum()
_ = ax[0, 1].pie(offer_type.received_counts, labels=offer_type.index, autopct='%1.1f%%')
_ = ax[0, 1].set_title("Type of Promotion")
reward = received_total.groupby('reward').sum()
_ = ax[1, 0].pie(reward.received_counts, labels=reward.index, autopct='%1.1f%%')
_ = ax[1, 0].set_title("Reward Amount")
difficulty = received_total.groupby('difficulty').sum()
_ = ax[1, 1].pie(difficulty.received_counts, labels=difficulty.index, autopct='%1.1f%%')
_ = ax[1, 1].set_title("Difficulty")
plt.savefig('images/general_promotion_pi.png')
# +
fig, ax = plt.subplots(2,2, figsize=(8, 8))
viewed_df = viewed_df.T
viewed_counts = viewed_df.sum()
viewed_counts.name = 'viewed_counts'
viewed_total = pd.concat([portfolio, viewed_counts], axis=1)
_ = ax[0, 0].pie(viewed_total.viewed_counts)
_ = ax[0, 0].set_title("Frequency of Offers viewed")
offer_type = viewed_total.groupby('offer_type').sum()
_ = ax[0, 1].pie(offer_type.viewed_counts, labels=offer_type.index, autopct='%1.1f%%')
_ = ax[0, 1].set_title("Type of Promotion")
reward = viewed_total.groupby('reward').sum()
_ = ax[1, 0].pie(reward.viewed_counts, labels=reward.index, autopct='%1.1f%%')
_ = ax[1, 0].set_title("Reward Amount")
difficulty = viewed_total.groupby('difficulty').sum()
_ = ax[1, 1].pie(difficulty.viewed_counts, labels=difficulty.index, autopct='%1.1f%%')
_ = ax[1, 1].set_title("Difficulty")
plt.savefig('images/viewed_promotion_pi.png')
# +
fig, ax = plt.subplots(2,2, figsize=(8, 8))
influenced_df = influenced_df.T
influenced_counts = influenced_df.sum()
influenced_counts.name = 'influence_counts'
influenced_total = pd.concat([portfolio, influenced_counts], axis=1)
_ = ax[0, 0].pie(influenced_total.influence_counts)
_ = ax[0, 0].set_title("ability to influence")
offer_type = influenced_total.groupby('offer_type').sum()
_ = ax[0, 1].pie(offer_type.influence_counts, labels=offer_type.index, autopct='%1.1f%%')
_ = ax[0, 1].set_title("Type of Promotion")
reward = influenced_total.groupby('reward').sum()
_ = ax[1, 0].pie(reward.influence_counts, labels=reward.index, autopct='%1.1f%%')
_ = ax[1, 0].set_title("Reward Amount")
difficulty = influenced_total.groupby('difficulty').sum()
_ = ax[1, 1].pie(difficulty.influence_counts, labels=difficulty.index, autopct='%1.1f%%')
_ = ax[1, 1].set_title("Difficulty")
plt.savefig('images/influenced_promotion_pi.png')
# -
# lets try combining influence with person data
influenced_df_summed = pd.DataFrame(influenced_df.sum(axis=1), columns=['influence_count'])
influence_person = influenced_df_summed.join(identified, how='inner')
influence_person.head(2)
# +
# lets bucket by index
fig, ax = plt.subplots(1, 2, figsize=(10, 10))
fig.tight_layout()
age_quantiles = list(np.quantile(influence_person.age, [0.2, 0.4, 0.6, 0.8, 1]))
influence_person['age_categ'] = pd.cut(influence_person.age, bins=([0] + age_quantiles))
age_categ = influence_person.groupby('age_categ').sum()
_ = ax[0].pie(age_categ.influence_count, labels=age_categ.index, autopct='%1f%%')
_ = ax[0].set_title("Influence by Age")
income_quantiles = list(np.quantile(influence_person.income, [0.2, 0.4, 0.6, 0.8, 1]))
influence_person['income_categ'] = pd.cut(influence_person.income, bins=([0] + income_quantiles))
income_categ = influence_person.groupby('income_categ').sum()
_ = ax[1].pie(income_categ.influence_count, labels=income_categ.index, autopct='%1.1f%%')
_ = ax[1].set_title("Influence by Income")
plt.savefig("images/profile_pi.png")
# +
fig, ax = plt.subplots(2,2, figsize=(8, 8))
low_income_df = influenced_df[influenced_df.index.isin(list(identified.loc[identified.income < 40000].id))]
influenced_counts = low_income_df.sum()
influenced_counts.name = 'influence_counts'
influenced_total = pd.concat([portfolio, influenced_counts], axis=1)
_ = ax[0, 0].pie(influenced_total.influence_counts)
_ = ax[0, 0].set_title("ability to influence")
offer_type = influenced_total.groupby('offer_type').sum()
_ = ax[0, 1].pie(offer_type.influence_counts, labels=offer_type.index, autopct='%1.1f%%')
_ = ax[0, 1].set_title("Type of Promotion")
reward = influenced_total.groupby('reward').sum()
_ = ax[1, 0].pie(reward.influence_counts, labels=reward.index, autopct='%1.1f%%')
_ = ax[1, 0].set_title("Reward Amount")
difficulty = influenced_total.groupby('difficulty').sum()
_ = ax[1, 1].pie(difficulty.influence_counts, labels=difficulty.index, autopct='%1.1f%%')
_ = ax[1, 1].set_title("Difficulty")
plt.savefig("images/low_income_pi.png")
# +
fig, ax = plt.subplots(2,2, figsize=(8, 8))
high_income_df = influenced_df[influenced_df.index.isin(list(identified.loc[identified.income > 80000].id))]
influenced_counts = high_income_df.sum()
influenced_counts.name = 'influence_counts'
influenced_total = pd.concat([portfolio, influenced_counts], axis=1)
_ = ax[0, 0].pie(influenced_total.influence_counts)
_ = ax[0, 0].set_title("ability to influence")
offer_type = influenced_total.groupby('offer_type').sum()
_ = ax[0, 1].pie(offer_type.influence_counts, labels=offer_type.index, autopct='%1.1f%%')
_ = ax[0, 1].set_title("Type of Promotion")
reward = influenced_total.groupby('reward').sum()
_ = ax[1, 0].pie(reward.influence_counts, labels=reward.index, autopct='%1.1f%%')
_ = ax[1, 0].set_title("Reward Amount")
difficulty = influenced_total.groupby('difficulty').sum()
_ = ax[1, 1].pie(difficulty.influence_counts, labels=difficulty.index, autopct='%1.1f%%')
_ = ax[1, 1].set_title("Difficulty")
plt.savefig("images/high_income_pi.png")
# +
# save any files for future use
impressions_df.to_csv("data/impressions.csv")
identified.to_csv("data/identified.csv")
viewed_df.to_csv("data/viewed.csv")
received_df.to_csv("data/received.csv")
influenced_df.to_csv("data/influenced.csv")
portfolio.to_csv("data/updated_portfolio.csv")
| 20,111 |
/.ipynb_checkpoints/starter_code-checkpoint.ipynb | a2e11c7d9c1418360d2c4f9944c0bf1d72a34e98 | [] | no_license | lidajav/Cryptocurrencies | https://github.com/lidajav/Cryptocurrencies | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 7,059,913 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] editable=true
# # Your first neural network
#
# In this project, you'll build your first neural network and use it to predict daily bike rental ridership. We've provided some of the code, but left the implementation of the neural network up to you (for the most part). After you've submitted this project, feel free to explore the data and the model more.
#
#
# + editable=true
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] editable=true
# ## Load and prepare the data
#
# A critical step in working with neural networks is preparing the data correctly. Variables on different scales make it difficult for the network to efficiently learn the correct weights. Below, we've written the code to load and prepare the data. You'll learn more about this soon!
# + editable=true
data_path = 'Bike-Sharing-Dataset/hour.csv'
rides = pd.read_csv(data_path)
# + editable=true
rides.head()
# + [markdown] editable=true
# ## Checking out the data
#
# This dataset has the number of riders for each hour of each day from January 1 2011 to December 31 2012. The number of riders is split between casual and registered, summed up in the `cnt` column. You can see the first few rows of the data above.
#
# Below is a plot showing the number of bike riders over the first 10 days or so in the data set. (Some days don't have exactly 24 entries in the data set, so it's not exactly 10 days.) You can see the hourly rentals here. This data is pretty complicated! The weekends have lower over all ridership and there are spikes when people are biking to and from work during the week. Looking at the data above, we also have information about temperature, humidity, and windspeed, all of these likely affecting the number of riders. You'll be trying to capture all this with your model.
# + editable=true
rides[:24*10].plot(x='dteday', y='cnt')
# + [markdown] editable=true
# ### Dummy variables
# Here we have some categorical variables like season, weather, month. To include these in our model, we'll need to make binary dummy variables. This is simple to do with Pandas thanks to `get_dummies()`.
# + editable=true
dummy_fields = ['season', 'weathersit', 'mnth', 'hr', 'weekday']
for each in dummy_fields:
dummies = pd.get_dummies(rides[each], prefix=each, drop_first=False)
rides = pd.concat([rides, dummies], axis=1)
fields_to_drop = ['instant', 'dteday', 'season', 'weathersit',
'weekday', 'atemp', 'mnth', 'workingday', 'hr']
data = rides.drop(fields_to_drop, axis=1)
data.head()
# + [markdown] editable=true
# ### Scaling target variables
# To make training the network easier, we'll standardize each of the continuous variables. That is, we'll shift and scale the variables such that they have zero mean and a standard deviation of 1.
#
# The scaling factors are saved so we can go backwards when we use the network for predictions.
# + editable=true
quant_features = ['casual', 'registered', 'cnt', 'temp', 'hum', 'windspeed']
# Store scalings in a dictionary so we can convert back later
scaled_features = {}
for each in quant_features:
mean, std = data[each].mean(), data[each].std()
scaled_features[each] = [mean, std]
data.loc[:, each] = (data[each] - mean)/std
# + [markdown] editable=true
# ### Splitting the data into training, testing, and validation sets
#
# We'll save the data for the last approximately 21 days to use as a test set after we've trained the network. We'll use this set to make predictions and compare them with the actual number of riders.
# + editable=true
# Save data for approximately the last 21 days
test_data = data[-21*24:]
# Now remove the test data from the data set
data = data[:-21*24]
# Separate the data into features and targets
target_fields = ['cnt', 'casual', 'registered']
features, targets = data.drop(target_fields, axis=1), data[target_fields]
test_features, test_targets = test_data.drop(target_fields, axis=1), test_data[target_fields]
# + [markdown] editable=true
# We'll split the data into two sets, one for training and one for validating as the network is being trained. Since this is time series data, we'll train on historical data, then try to predict on future data (the validation set).
# + editable=true
# Hold out the last 60 days or so of the remaining data as a validation set
train_features, train_targets = features[:-60*24], targets[:-60*24]
val_features, val_targets = features[-60*24:], targets[-60*24:]
# + [markdown] editable=true
# ## Time to build the network
#
# Below you'll build your network. We've built out the structure. You'll implement both the forward pass and backwards pass through the network. You'll also set the hyperparameters: the learning rate, the number of hidden units, and the number of training passes.
#
# <img src="assets/neural_network.png" width=300px>
#
# The network has two layers, a hidden layer and an output layer. The hidden layer will use the sigmoid function for activations. The output layer has only one node and is used for the regression, the output of the node is the same as the input of the node. That is, the activation function is $f(x)=x$. A function that takes the input signal and generates an output signal, but takes into account the threshold, is called an activation function. We work through each layer of our network calculating the outputs for each neuron. All of the outputs from one layer become inputs to the neurons on the next layer. This process is called *forward propagation*.
#
# We use the weights to propagate signals forward from the input to the output layers in a neural network. We use the weights to also propagate error backwards from the output back into the network to update our weights. This is called *backpropagation*.
#
# > **Hint:** You'll need the derivative of the output activation function ($f(x) = x$) for the backpropagation implementation. If you aren't familiar with calculus, this function is equivalent to the equation $y = x$. What is the slope of that equation? That is the derivative of $f(x)$.
#
# Below, you have these tasks:
# 1. Implement the sigmoid function to use as the activation function. Set `self.activation_function` in `__init__` to your sigmoid function.
# 2. Implement the forward pass in the `train` method.
# 3. Implement the backpropagation algorithm in the `train` method, including calculating the output error.
# 4. Implement the forward pass in the `run` method.
#
# + editable=true
#############
# In the my_answers.py file, fill out the TODO sections as specified
#############
from my_answers import NeuralNetwork
# + editable=true
def MSE(y, Y):
return np.mean((y-Y)**2)
# + [markdown] editable=true
# ## Unit tests
#
# Run these unit tests to check the correctness of your network implementation. This will help you be sure your network was implemented correctly befor you starting trying to train it. These tests must all be successful to pass the project.
# + editable=true
import unittest
inputs = np.array([[0.5, -0.2, 0.1]])
targets = np.array([[0.4]])
test_w_i_h = np.array([[0.1, -0.2],
[0.4, 0.5],
[-0.3, 0.2]])
test_w_h_o = np.array([[0.3],
[-0.1]])
class TestMethods(unittest.TestCase):
##########
# Unit tests for data loading
##########
def test_data_path(self):
# Test that file path to dataset has been unaltered
self.assertTrue(data_path.lower() == 'bike-sharing-dataset/hour.csv')
def test_data_loaded(self):
# Test that data frame loaded
self.assertTrue(isinstance(rides, pd.DataFrame))
##########
# Unit tests for network functionality
##########
def test_activation(self):
network = NeuralNetwork(3, 2, 1, 0.5)
# Test that the activation function is a sigmoid
self.assertTrue(np.all(network.activation_function(0.5) == 1/(1+np.exp(-0.5))))
def test_train(self):
# Test that weights are updated correctly on training
network = NeuralNetwork(3, 2, 1, 0.5)
network.weights_input_to_hidden = test_w_i_h.copy()
network.weights_hidden_to_output = test_w_h_o.copy()
network.train(inputs, targets)
self.assertTrue(np.allclose(network.weights_hidden_to_output,
np.array([[ 0.37275328],
[-0.03172939]])))
self.assertTrue(np.allclose(network.weights_input_to_hidden,
np.array([[ 0.10562014, -0.20185996],
[0.39775194, 0.50074398],
[-0.29887597, 0.19962801]])))
def test_run(self):
# Test correctness of run method
network = NeuralNetwork(3, 2, 1, 0.5)
network.weights_input_to_hidden = test_w_i_h.copy()
network.weights_hidden_to_output = test_w_h_o.copy()
self.assertTrue(np.allclose(network.run(inputs), 0.09998924))
suite = unittest.TestLoader().loadTestsFromModule(TestMethods())
unittest.TextTestRunner().run(suite)
# + [markdown] editable=true
# ## Training the network
#
# Here you'll set the hyperparameters for the network. The strategy here is to find hyperparameters such that the error on the training set is low, but you're not overfitting to the data. If you train the network too long or have too many hidden nodes, it can become overly specific to the training set and will fail to generalize to the validation set. That is, the loss on the validation set will start increasing as the training set loss drops.
#
# You'll also be using a method know as Stochastic Gradient Descent (SGD) to train the network. The idea is that for each training pass, you grab a random sample of the data instead of using the whole data set. You use many more training passes than with normal gradient descent, but each pass is much faster. This ends up training the network more efficiently. You'll learn more about SGD later.
#
# ### Choose the number of iterations
# This is the number of batches of samples from the training data we'll use to train the network. The more iterations you use, the better the model will fit the data. However, this process can have sharply diminishing returns and can waste computational resources if you use too many iterations. You want to find a number here where the network has a low training loss, and the validation loss is at a minimum. The ideal number of iterations would be a level that stops shortly after the validation loss is no longer decreasing.
#
# ### Choose the learning rate
# This scales the size of weight updates. If this is too big, the weights tend to explode and the network fails to fit the data. Normally a good choice to start at is 0.1; however, if you effectively divide the learning rate by n_records, try starting out with a learning rate of 1. In either case, if the network has problems fitting the data, try reducing the learning rate. Note that the lower the learning rate, the smaller the steps are in the weight updates and the longer it takes for the neural network to converge.
#
# ### Choose the number of hidden nodes
# In a model where all the weights are optimized, the more hidden nodes you have, the more accurate the predictions of the model will be. (A fully optimized model could have weights of zero, after all.) However, the more hidden nodes you have, the harder it will be to optimize the weights of the model, and the more likely it will be that suboptimal weights will lead to overfitting. With overfitting, the model will memorize the training data instead of learning the true pattern, and won't generalize well to unseen data.
#
# Try a few different numbers and see how it affects the performance. You can look at the losses dictionary for a metric of the network performance. If the number of hidden units is too low, then the model won't have enough space to learn and if it is too high there are too many options for the direction that the learning can take. The trick here is to find the right balance in number of hidden units you choose. You'll generally find that the best number of hidden nodes to use ends up being between the number of input and output nodes.
# + editable=true
import sys
####################
### Set the hyperparameters in you myanswers.py file ###
####################
from my_answers import iterations, learning_rate, hidden_nodes, output_nodes
N_i = train_features.shape[1]
network = NeuralNetwork(N_i, hidden_nodes, output_nodes, learning_rate)
losses = {'train':[], 'validation':[]}
for ii in range(iterations):
# Go through a random batch of 128 records from the training data set
batch = np.random.choice(train_features.index, size=128)
X, y = train_features.ix[batch].values, train_targets.ix[batch]['cnt']
network.train(X, y)
# Printing out the training progress
train_loss = MSE(network.run(train_features).T, train_targets['cnt'].values)
val_loss = MSE(network.run(val_features).T, val_targets['cnt'].values)
sys.stdout.write("\rProgress: {:2.1f}".format(100 * ii/float(iterations)) \
+ "% ... Training loss: " + str(train_loss)[:5] \
+ " ... Validation loss: " + str(val_loss)[:5])
sys.stdout.flush()
losses['train'].append(train_loss)
losses['validation'].append(val_loss)
# + editable=true
plt.plot(losses['train'], label='Training loss')
plt.plot(losses['validation'], label='Validation loss')
plt.legend()
_ = plt.ylim()
# + [markdown] editable=true
# ## Check out your predictions
#
# Here, use the test data to view how well your network is modeling the data. If something is completely wrong here, make sure each step in your network is implemented correctly.
# + editable=true
fig, ax = plt.subplots(figsize=(8,4))
mean, std = scaled_features['cnt']
predictions = network.run(test_features).T*std + mean
ax.plot(predictions[0], label='Prediction')
ax.plot((test_targets['cnt']*std + mean).values, label='Data')
ax.set_xlim(right=len(predictions))
ax.legend()
dates = pd.to_datetime(rides.ix[test_data.index]['dteday'])
dates = dates.apply(lambda d: d.strftime('%b %d'))
ax.set_xticks(np.arange(len(dates))[12::24])
_ = ax.set_xticklabels(dates[12::24], rotation=45)
# + [markdown] editable=true
# ## OPTIONAL: Thinking about your results(this question will not be evaluated in the rubric).
#
# Answer these questions about your results. How well does the model predict the data? Where does it fail? Why does it fail where it does?
#
# > **Note:** You can edit the text in this cell by double clicking on it. When you want to render the text, press control + enter
#
# #### Your answer below
# + [markdown] editable=true
# ## Submitting:
# Open up the 'jwt' file in the first-neural-network directory (which also contains this notebook) for submission instructions
# + editable=true
alue from a group of values, where each value has a probability of being picked.
# - Sampling allows us to generate random sequences of values.
# - Pick the next character's index according to the probability distribution specified by $\hat{y}^{\langle t+1 \rangle }$.
# - This means that if $\hat{y}^{\langle t+1 \rangle }_i = 0.16$, you will pick the index "i" with 16% probability.
# - You can use [np.random.choice](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.random.choice.html).
#
# Example of how to use `np.random.choice()`:
# ```python
# np.random.seed(0)
# probs = np.array([0.1, 0.0, 0.7, 0.2])
# idx = np.random.choice([0, 1, 2, 3] p = probs)
# ```
# - This means that you will pick the index (`idx`) according to the distribution:
#
# $P(index = 0) = 0.1, P(index = 1) = 0.0, P(index = 2) = 0.7, P(index = 3) = 0.2$.
#
# - Note that the value that's set to `p` should be set to a 1D vector.
# - Also notice that $\hat{y}^{\langle t+1 \rangle}$, which is `y` in the code, is a 2D array.
# ##### Additional Hints
# - [range](https://docs.python.org/3/library/functions.html#func-range)
# - [numpy.ravel](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ravel.html) takes a multi-dimensional array and returns its contents inside of a 1D vector.
# ```Python
# arr = np.array([[1,2],[3,4]])
# print("arr")
# print(arr)
# print("arr.ravel()")
# print(arr.ravel())
# ```
# Output:
# ```Python
# arr
# [[1 2]
# [3 4]]
# arr.ravel()
# [1 2 3 4]
# ```
#
# - Note that `append` is an "in-place" operation. In other words, don't do this:
# ```Python
# fun_hobbies = fun_hobbies.append('learning') ## Doesn't give you what you want
# ```
# - **Step 4**: Update to $x^{\langle t \rangle }$
# - The last step to implement in `sample()` is to update the variable `x`, which currently stores $x^{\langle t \rangle }$, with the value of $x^{\langle t + 1 \rangle }$.
# - You will represent $x^{\langle t + 1 \rangle }$ by creating a one-hot vector corresponding to the character that you have chosen as your prediction.
# - You will then forward propagate $x^{\langle t + 1 \rangle }$ in Step 1 and keep repeating the process until you get a "\n" character, indicating that you have reached the end of the dinosaur name.
# ##### Additional Hints
# - In order to reset `x` before setting it to the new one-hot vector, you'll want to set all the values to zero.
# - You can either create a new numpy array: [numpy.zeros](https://docs.scipy.org/doc/numpy/reference/generated/numpy.zeros.html)
# - Or fill all values with a single number: [numpy.ndarray.fill](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.fill.html)
# +
# GRADED FUNCTION: sample
def sample(parameters, char_to_ix, seed):
"""
Sample a sequence of characters according to a sequence of probability distributions output of the RNN
Arguments:
parameters -- python dictionary containing the parameters Waa, Wax, Wya, by, and b.
char_to_ix -- python dictionary mapping each character to an index.
seed -- used for grading purposes. Do not worry about it.
Returns:
indices -- a list of length n containing the indices of the sampled characters.
"""
# Retrieve parameters and relevant shapes from "parameters" dictionary
Waa, Wax, Wya, by, b = parameters['Waa'], parameters['Wax'], parameters['Wya'], parameters['by'], parameters['b']
vocab_size = by.shape[0]
n_a = Waa.shape[1]
### START CODE HERE ###
# Step 1: Create the a zero vector x that can be used as the one-hot vector
# representing the first character (initializing the sequence generation). (≈1 line)
x = np.zeros(shape=(vocab_size,1))
# Step 1': Initialize a_prev as zeros (≈1 line)
a_prev = np.zeros(shape=(n_a,1))
# Create an empty list of indices, this is the list which will contain the list of indices of the characters to generate (≈1 line)
indices = []
# idx is the index of the one-hot vector x that is set to 1
# All other positions in x are zero.
# We will initialize idx to -1
idx = -1
# Loop over time-steps t. At each time-step:
# sample a character from a probability distribution
# and append its index (`idx`) to the list "indices".
# We'll stop if we reach 50 characters
# (which should be very unlikely with a well trained model).
# Setting the maximum number of characters helps with debugging and prevents infinite loops.
counter = 0
newline_character = char_to_ix['\n']
while (idx != newline_character and counter != 50):
# Step 2: Forward propagate x using the equations (1), (2) and (3)
a = np.tanh(np.dot(Wax,x) + np.dot(Waa,a_prev) + b)
z = np.dot(Wya,a) + by
y = softmax(z)
# for grading purposes
np.random.seed(counter+seed)
# Step 3: Sample the index of a character within the vocabulary from the probability distribution y
# (see additional hints above)
idx = np.random.choice(list(range(vocab_size)), p=y.ravel())
# Append the index to "indices"
indices.append(idx)
# Step 4: Overwrite the input x with one that corresponds to the sampled index `idx`.
# (see additional hints above)
x = np.zeros(shape=(vocab_size,1))
x[idx] = 1
# Update "a_prev" to be "a"
a_prev = a
# for grading purposes
seed += 1
counter +=1
### END CODE HERE ###
if (counter == 50):
indices.append(char_to_ix['\n'])
return indices
# +
np.random.seed(2)
_, n_a = 20, 100
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
indices = sample(parameters, char_to_ix, 0)
print("Sampling:")
print("list of sampled indices:\n", indices)
print("list of sampled characters:\n", [ix_to_char[i] for i in indices])
# -
# ** Expected output:**
#
# ```Python
# Sampling:
# list of sampled indices:
# [12, 17, 24, 14, 13, 9, 10, 22, 24, 6, 13, 11, 12, 6, 21, 15, 21, 14, 3, 2, 1, 21, 18, 24, 7, 25, 6, 25, 18, 10, 16, 2, 3, 8, 15, 12, 11, 7, 1, 12, 10, 2, 7, 7, 11, 17, 24, 12, 13, 24, 0]
# list of sampled characters:
# ['l', 'q', 'x', 'n', 'm', 'i', 'j', 'v', 'x', 'f', 'm', 'k', 'l', 'f', 'u', 'o', 'u', 'n', 'c', 'b', 'a', 'u', 'r', 'x', 'g', 'y', 'f', 'y', 'r', 'j', 'p', 'b', 'c', 'h', 'o', 'l', 'k', 'g', 'a', 'l', 'j', 'b', 'g', 'g', 'k', 'q', 'x', 'l', 'm', 'x', '\n']
# ```
#
# * Please note that over time, if there are updates to the back-end of the Coursera platform (that may update the version of numpy), the actual list of sampled indices and sampled characters may change.
# * If you follow the instructions given above and get an output without errors, it's possible the routine is correct even if your output doesn't match the expected output. Submit your assignment to the grader to verify its correctness.
# ## 3 - Building the language model
#
# It is time to build the character-level language model for text generation.
#
#
# ### 3.1 - Gradient descent
#
# * In this section you will implement a function performing one step of stochastic gradient descent (with clipped gradients).
# * You will go through the training examples one at a time, so the optimization algorithm will be stochastic gradient descent.
#
# As a reminder, here are the steps of a common optimization loop for an RNN:
#
# - Forward propagate through the RNN to compute the loss
# - Backward propagate through time to compute the gradients of the loss with respect to the parameters
# - Clip the gradients
# - Update the parameters using gradient descent
#
# **Exercise**: Implement the optimization process (one step of stochastic gradient descent).
#
# The following functions are provided:
#
# ```python
# def rnn_forward(X, Y, a_prev, parameters):
# """ Performs the forward propagation through the RNN and computes the cross-entropy loss.
# It returns the loss' value as well as a "cache" storing values to be used in backpropagation."""
# ....
# return loss, cache
#
# def rnn_backward(X, Y, parameters, cache):
# """ Performs the backward propagation through time to compute the gradients of the loss with respect
# to the parameters. It returns also all the hidden states."""
# ...
# return gradients, a
#
# def update_parameters(parameters, gradients, learning_rate):
# """ Updates parameters using the Gradient Descent Update Rule."""
# ...
# return parameters
# ```
#
# Recall that you previously implemented the `clip` function:
#
# ```Python
# def clip(gradients, maxValue)
# """Clips the gradients' values between minimum and maximum."""
# ...
# return gradients
# ```
# #### parameters
#
# * Note that the weights and biases inside the `parameters` dictionary are being updated by the optimization, even though `parameters` is not one of the returned values of the `optimize` function. The `parameters` dictionary is passed by reference into the function, so changes to this dictionary are making changes to the `parameters` dictionary even when accessed outside of the function.
# * Python dictionaries and lists are "pass by reference", which means that if you pass a dictionary into a function and modify the dictionary within the function, this changes that same dictionary (it's not a copy of the dictionary).
# +
# GRADED FUNCTION: optimize
def optimize(X, Y, a_prev, parameters, learning_rate = 0.01):
"""
Execute one step of the optimization to train the model.
Arguments:
X -- list of integers, where each integer is a number that maps to a character in the vocabulary.
Y -- list of integers, exactly the same as X but shifted one index to the left.
a_prev -- previous hidden state.
parameters -- python dictionary containing:
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
b -- Bias, numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
learning_rate -- learning rate for the model.
Returns:
loss -- value of the loss function (cross-entropy)
gradients -- python dictionary containing:
dWax -- Gradients of input-to-hidden weights, of shape (n_a, n_x)
dWaa -- Gradients of hidden-to-hidden weights, of shape (n_a, n_a)
dWya -- Gradients of hidden-to-output weights, of shape (n_y, n_a)
db -- Gradients of bias vector, of shape (n_a, 1)
dby -- Gradients of output bias vector, of shape (n_y, 1)
a[len(X)-1] -- the last hidden state, of shape (n_a, 1)
"""
### START CODE HERE ###
# Forward propagate through time (≈1 line)
loss, cache = rnn_forward(X, Y, a_prev, parameters)
# Backpropagate through time (≈1 line)
gradients, a = rnn_backward(X, Y, parameters, cache)
# Clip your gradients between -5 (min) and 5 (max) (≈1 line)
gradients = clip(gradients, maxValue = 5)
# Update parameters (≈1 line)
parameters = update_parameters(parameters, gradients, learning_rate)
### END CODE HERE ###
return loss, gradients, a[len(X)-1]
# +
np.random.seed(1)
vocab_size, n_a = 27, 100
a_prev = np.random.randn(n_a, 1)
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
X = [12,3,5,11,22,3]
Y = [4,14,11,22,25, 26]
loss, gradients, a_last = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
print("Loss =", loss)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("np.argmax(gradients[\"dWax\"]) =", np.argmax(gradients["dWax"]))
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
print("a_last[4] =", a_last[4])
# -
# ** Expected output:**
#
# ```Python
# Loss = 126.503975722
# gradients["dWaa"][1][2] = 0.194709315347
# np.argmax(gradients["dWax"]) = 93
# gradients["dWya"][1][2] = -0.007773876032
# gradients["db"][4] = [-0.06809825]
# gradients["dby"][1] = [ 0.01538192]
# a_last[4] = [-1.]
# ```
# ### 3.2 - Training the model
# * Given the dataset of dinosaur names, we use each line of the dataset (one name) as one training example.
# * Every 100 steps of stochastic gradient descent, you will sample 10 randomly chosen names to see how the algorithm is doing.
# * Remember to shuffle the dataset, so that stochastic gradient descent visits the examples in random order.
#
# **Exercise**: Follow the instructions and implement `model()`. When `examples[index]` contains one dinosaur name (string), to create an example (X, Y), you can use this:
#
# ##### Set the index `idx` into the list of examples
# * Using the for-loop, walk through the shuffled list of dinosaur names in the list "examples".
# * If there are 100 examples, and the for-loop increments the index to 100 onwards, think of how you would make the index cycle back to 0, so that we can continue feeding the examples into the model when j is 100, 101, etc.
# * Hint: 101 divided by 100 is zero with a remainder of 1.
# * `%` is the modulus operator in python.
#
# ##### Extract a single example from the list of examples
# * `single_example`: use the `idx` index that you set previously to get one word from the list of examples.
# ##### Convert a string into a list of characters: `single_example_chars`
# * `single_example_chars`: A string is a list of characters.
# * You can use a list comprehension (recommended over for-loops) to generate a list of characters.
# ```Python
# str = 'I love learning'
# list_of_chars = [c for c in str]
# print(list_of_chars)
# ```
#
# ```
# ['I', ' ', 'l', 'o', 'v', 'e', ' ', 'l', 'e', 'a', 'r', 'n', 'i', 'n', 'g']
# ```
# ##### Convert list of characters to a list of integers: `single_example_ix`
# * Create a list that contains the index numbers associated with each character.
# * Use the dictionary `char_to_ix`
# * You can combine this with the list comprehension that is used to get a list of characters from a string.
# * This is a separate line of code below, to help learners clarify each step in the function.
# ##### Create the list of input characters: `X`
# * `rnn_forward` uses the `None` value as a flag to set the input vector as a zero-vector.
# * Prepend the `None` value in front of the list of input characters.
# * There is more than one way to prepend a value to a list. One way is to add two lists together: `['a'] + ['b']`
# ##### Get the integer representation of the newline character `ix_newline`
# * `ix_newline`: The newline character signals the end of the dinosaur name.
# - get the integer representation of the newline character `'\n'`.
# - Use `char_to_ix`
# ##### Set the list of labels (integer representation of the characters): `Y`
# * The goal is to train the RNN to predict the next letter in the name, so the labels are the list of characters that are one time step ahead of the characters in the input `X`.
# - For example, `Y[0]` contains the same value as `X[1]`
# * The RNN should predict a newline at the last letter so add ix_newline to the end of the labels.
# - Append the integer representation of the newline character to the end of `Y`.
# - Note that `append` is an in-place operation.
# - It might be easier for you to add two lists together.
# +
# GRADED FUNCTION: model
def model(data, ix_to_char, char_to_ix, num_iterations = 35000, n_a = 50, dino_names = 7, vocab_size = 27):
"""
Trains the model and generates dinosaur names.
Arguments:
data -- text corpus
ix_to_char -- dictionary that maps the index to a character
char_to_ix -- dictionary that maps a character to an index
num_iterations -- number of iterations to train the model for
n_a -- number of units of the RNN cell
dino_names -- number of dinosaur names you want to sample at each iteration.
vocab_size -- number of unique characters found in the text (size of the vocabulary)
Returns:
parameters -- learned parameters
"""
# Retrieve n_x and n_y from vocab_size
n_x, n_y = vocab_size, vocab_size
# Initialize parameters
parameters = initialize_parameters(n_a, n_x, n_y)
# Initialize loss (this is required because we want to smooth our loss)
loss = get_initial_loss(vocab_size, dino_names)
# Build list of all dinosaur names (training examples).
with open("dinos.txt") as f:
examples = f.readlines()
examples = [x.lower().strip() for x in examples]
# Shuffle list of all dinosaur names
np.random.seed(0)
np.random.shuffle(examples)
# Initialize the hidden state of your LSTM
a_prev = np.zeros((n_a, 1))
# Optimization loop
for j in range(num_iterations):
### START CODE HERE ###
# Set the index `idx` (see instructions above)
idx = j % len(examples)
# Set the input X (see instructions above)
single_example = examples[idx]
single_example_chars = [ c for c in single_example]
single_example_ix = [char_to_ix[c] for c in single_example_chars]
X = [None] + single_example_ix
# Set the labels Y (see instructions above)
ix_newline = char_to_ix['\n']
Y = X[1:] + [ix_newline]
# Perform one optimization step: Forward-prop -> Backward-prop -> Clip -> Update parameters
# Choose a learning rate of 0.01
curr_loss, gradients, a_prev = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
### END CODE HERE ###
# Use a latency trick to keep the loss smooth. It happens here to accelerate the training.
loss = smooth(loss, curr_loss)
# Every 2000 Iteration, generate "n" characters thanks to sample() to check if the model is learning properly
if j % 2000 == 0:
print('Iteration: %d, Loss: %f' % (j, loss) + '\n')
# The number of dinosaur names to print
seed = 0
for name in range(dino_names):
# Sample indices and print them
sampled_indices = sample(parameters, char_to_ix, seed)
print_sample(sampled_indices, ix_to_char)
seed += 1 # To get the same result (for grading purposes), increment the seed by one.
print('\n')
return parameters
# -
# Run the following cell, you should observe your model outputting random-looking characters at the first iteration. After a few thousand iterations, your model should learn to generate reasonable-looking names.
parameters = model(data, ix_to_char, char_to_ix)
# ** Expected Output**
#
# The output of your model may look different, but it will look something like this:
#
# ```Python
# Iteration: 34000, Loss: 22.447230
#
# Onyxipaledisons
# Kiabaeropa
# Lussiamang
# Pacaeptabalsaurus
# Xosalong
# Eiacoteg
# Troia
# ```
# ## Conclusion
#
# You can see that your algorithm has started to generate plausible dinosaur names towards the end of the training. At first, it was generating random characters, but towards the end you could see dinosaur names with cool endings. Feel free to run the algorithm even longer and play with hyperparameters to see if you can get even better results. Our implementation generated some really cool names like `maconucon`, `marloralus` and `macingsersaurus`. Your model hopefully also learned that dinosaur names tend to end in `saurus`, `don`, `aura`, `tor`, etc.
#
# If your model generates some non-cool names, don't blame the model entirely--not all actual dinosaur names sound cool. (For example, `dromaeosauroides` is an actual dinosaur name and is in the training set.) But this model should give you a set of candidates from which you can pick the coolest!
#
# This assignment had used a relatively small dataset, so that you could train an RNN quickly on a CPU. Training a model of the english language requires a much bigger dataset, and usually needs much more computation, and could run for many hours on GPUs. We ran our dinosaur name for quite some time, and so far our favorite name is the great, undefeatable, and fierce: Mangosaurus!
#
# <img src="images/mangosaurus.jpeg" style="width:250;height:300px;">
# ## 4 - Writing like Shakespeare
#
# The rest of this notebook is optional and is not graded, but we hope you'll do it anyway since it's quite fun and informative.
#
# A similar (but more complicated) task is to generate Shakespeare poems. Instead of learning from a dataset of Dinosaur names you can use a collection of Shakespearian poems. Using LSTM cells, you can learn longer term dependencies that span many characters in the text--e.g., where a character appearing somewhere a sequence can influence what should be a different character much much later in the sequence. These long term dependencies were less important with dinosaur names, since the names were quite short.
#
#
# <img src="images/shakespeare.jpg" style="width:500;height:400px;">
# <caption><center> Let's become poets! </center></caption>
#
# We have implemented a Shakespeare poem generator with Keras. Run the following cell to load the required packages and models. This may take a few minutes.
from __future__ import print_function
from keras.callbacks import LambdaCallback
from keras.models import Model, load_model, Sequential
from keras.layers import Dense, Activation, Dropout, Input, Masking
from keras.layers import LSTM
from keras.utils.data_utils import get_file
from keras.preprocessing.sequence import pad_sequences
from shakespeare_utils import *
import sys
import io
# To save you some time, we have already trained a model for ~1000 epochs on a collection of Shakespearian poems called [*"The Sonnets"*](shakespeare.txt).
# Let's train the model for one more epoch. When it finishes training for an epoch---this will also take a few minutes---you can run `generate_output`, which will prompt asking you for an input (`<`40 characters). The poem will start with your sentence, and our RNN-Shakespeare will complete the rest of the poem for you! For example, try "Forsooth this maketh no sense " (don't enter the quotation marks). Depending on whether you include the space at the end, your results might also differ--try it both ways, and try other inputs as well.
#
# +
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y, batch_size=128, epochs=1, callbacks=[print_callback])
# -
# Run this cell to try with different inputs without having to re-train the model
generate_output()
# The RNN-Shakespeare model is very similar to the one you have built for dinosaur names. The only major differences are:
# - LSTMs instead of the basic RNN to capture longer-range dependencies
# - The model is a deeper, stacked LSTM model (2 layer)
# - Using Keras instead of python to simplify the code
#
# If you want to learn more, you can also check out the Keras Team's text generation implementation on GitHub: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py.
#
# Congratulations on finishing this notebook!
# **References**:
# - This exercise took inspiration from Andrej Karpathy's implementation: https://gist.github.com/karpathy/d4dee566867f8291f086. To learn more about text generation, also check out Karpathy's [blog post](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
# - For the Shakespearian poem generator, our implementation was based on the implementation of an LSTM text generator by the Keras team: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py
| 39,965 |
/Capstone week1.ipynb | b2e701206bf46e77798cc4ee5911357602bc5bf2 | [] | no_license | duyennt97/Coursera_Capstone | https://github.com/duyennt97/Coursera_Capstone | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 1,145 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import log_loss, accuracy_score, confusion_matrix
# ### 特徴選択から再度実施
# ### Scikit-learnによる分類
# * 実際のデータで,説明変数が多次元の場合のモデルを構築
#
# 説明変数として考えられるもの
# * goal : 目標金額
# * deadline : 納期
# * launched : 開始日
# * country : PJ開催国
# * currency : 通貨の単位
# * deadline_days : 納期への近さ
# * goal_per_days : 1日あたりの目標金額
# * category_xxx : PJの小分類がxxxかどうか
# * name_length : PJ名の長さ
# * name_xxx : PJのタイトルにxxxが含まれているか
#CSVファイル読み込み
df = pd.read_csv("../1_data/ks-projects-201801.csv")
df.index.name="id"
#欠損値がある列を確認
df.isnull().sum()
#欠測値がある列を落とす
#stateが「live」のデータを落としていることを明示するためにあえて記載
df = df[df['state'] != 'live']
df = df[~df['name'].isnull()]
df = df[~df['usd pledged'].isnull()]
df['state'].loc[df['state'] == 'successful'] = 1
df['state'].loc[df['state'] != 1] = 0
#nameの長さを取得
df['name_length'] = df['name'].apply(lambda x: len(str(x).replace(' ', ''))).fillna(0)
#deadline,launchedを日付に変換するための関数
def period_to_date(x_date, format_date):
x_date_format = datetime.datetime.strptime(x_date, format_date)
return x_date_format
# +
#deadline と launched からプロジェクト納期への近さ(日数)を計算
df['deadline_period'] = df['deadline'].apply(lambda x : period_to_date(x, '%Y-%m-%d')) \
- df['launched'].apply(lambda x : period_to_date(x, '%Y-%m-%d %H:%M:%S'))
df['deadline_days'] = df['deadline_period'].apply(lambda x : x.days)
#goal に対して納期への日数1日あたりいくら必要かを計算
df['goal_per_days'] = df['goal'] / df['deadline_days']
df['goal_per_days'] = df['goal_per_days'].fillna(0)
df['goal_per_days'] [df['goal_per_days'] == np.inf]= 0
# +
#pairplot用の変数を揃える
#ここでは更に、goal_per_days(1日あたりの目標金額),name_length(PJ名の長さ),deadline_days(納期に対してどのくらいの近さでリリースしたか)を指標として加える
df['goal_log'] = df['goal'].apply(lambda x: np.log(x+1))
df['goal_per_days_log'] = df['goal_per_days'].apply(lambda x: np.log(x+1))
df['deadline_days_log'] = df['deadline_days'].apply(lambda x: np.log(x+1))
df['name_length_log'] = df['name_length'].apply(lambda x: np.log(x+1))
# +
#文字列を含む候補のリスト
name_first=['1st','First','FIRST','first','1ST']
#nameに特定の単語を含むか調べた列を追加
df['name_first'] = df['name'].str.contains('|'.join(name_first),na=False)
df['name_first'].loc[df['name_first'] == True] = 1
df['name_first'].loc[df['name_first'] == False] = 0
# +
#文字列を含む候補のリスト
name_quick=['quick','Quick','QUICK']
#nameに特定の単語を含むか調べた列を追加
df['name_quick'] = df['name'].str.contains('|'.join(name_quick),na=False)
df['name_quick'].loc[df['name_quick'] == True] = 1
df['name_quick'].loc[df['name_quick'] == False] = 0
# +
#文字列を含む候補のリスト
name_theatre=['theatre','Theatre','THERTRE','theater','Theater','THERTER']
#nameに特定の単語を含むか調べた列を追加
df['name_theatre'] = df['name'].str.contains('|'.join(name_theatre),na=False)
df['name_theatre'].loc[df['name_theatre'] == True] = 1
df['name_theatre'].loc[df['name_theatre'] == False] = 0
# +
#文字列を含む候補のリスト
name_movie=['movie','Movie','movie']
#nameに特定の単語を含むか調べた列を追加
df['name_movie'] = df['name'].str.contains('|'.join(name_movie),na=False)
df['name_movie'].loc[df['name_movie'] == True] = 1
df['name_movie'].loc[df['name_movie'] == False] = 0
# -
#category,main_category,currencyの値をダミー変数化
df = pd.get_dummies(df, columns = ['category','currency'])
# 相関係数をヒートマップにして可視化
import seaborn as sns
sns.heatmap(df[["goal_log", "deadline_days_log","goal_per_days_log", "category_Chiptune", "currency_USD","name_theatre","name_movie","name_length_log"]].corr())
plt.show()
# goalとgoal_per_daysに強い正の相関、どちらか一方のみを使うことが推奨される<br>
# country_USとcurrency_USDはほぼ同じ特徴量を持った列であり、どちらか一方でよい<br>
# その他はどの説明変数に関してもほぼ相関がない、つまりここで出た7つのうちいずれか5つの説明変数に関しては独立して扱える<br>
#goal_per_daysの値を標準化
df["goal_per_days_standardized"] = (df["goal_per_days"] - df["goal_per_days"].mean()) / df["goal_per_days"].std(ddof=0)
#deadline_daysの値を標準化
df["deadline_days_standardized"] = (df["deadline_days"] - df["deadline_days"].mean()) / df["deadline_days"].std(ddof=0)
#name_lengthの値を正規化
df["name_length_normalized"] = (df["name_length"] - df["name_length"].min()) / (df["name_length"].max() - df["name_length"].min())
# <h2>分類アルゴリズムに使う説明変数を決定する</h2>
y = df["state"].values
X = df[["goal_per_days_standardized","deadline_days_standardized","name_length_normalized","category_Anthologies","category_Chiptune","name_quick","name_theatre","currency_USD","name_first","name_movie"]].values
#テストデータの分割
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# +
clf = SGDClassifier(loss='log', penalty='none', max_iter=10000, fit_intercept=True, random_state=1234, tol=1e-3)
clf.fit(X_train, y_train)
# 重みを取得して表示
w0 = clf.intercept_[0]
w1 = clf.coef_[0, 0]
w2 = clf.coef_[0, 1]
w3 = clf.coef_[0, 2]
w4 = clf.coef_[0, 3]
w5 = clf.coef_[0, 4]
w6 = clf.coef_[0, 5]
w7 = clf.coef_[0, 6]
w8 = clf.coef_[0, 7]
w9 = clf.coef_[0, 8]
print("w0 = {:.3f}, w1 = {:.3f}, w2 = {:.3f}, w3 = {:.3f}, w4 = {:.3f}, w5 = {:.3f}, w6 = {:.3f}, w7 = {:.3f}, w8 = {:.3f}".format(w0, w1, w2, w3, w4, w5, w6, w7, w8))
# +
from sklearn.metrics import accuracy_score, precision_recall_fscore_support, confusion_matrix # 回帰問題における性能評価に関する関数
# ラベルを予測
y_est = clf.predict(X_test)
# 対数尤度を表示
print('対数尤度 = {:.3f}'.format(- log_loss(y_test, y_est)))
# 正答率を表示
print('正答率 = {:.3f}%'.format(100 * accuracy_score(y_test, y_est)))
# Precision, Recall, F1-scoreを計算
precision, recall, f1_score, _ = precision_recall_fscore_support(y_test, y_est)
# カテゴリ「state」に関するPrecision, Recall, F1-scoreを表示
print('適合率(Precision) = {:.3f}%'.format(100 * precision[0]))
print('再現率(Recall) = {:.3f}%'.format(100 * recall[0]))
print('F1値(F1-score) = {:.3f}%'.format(100 * f1_score[0]))
# -
# 予測値と正解のクロス集計
conf_mat = pd.DataFrame(confusion_matrix(y_test, y_est),
index=['正解 = fail', '正解 = successful'],
columns=['予測 = fail', '予測 = successful'])
conf_mat
# +
#ランダムフォレスト
from sklearn.ensemble import RandomForestClassifier
#ハイパーパラメータ調整
clf_rf = RandomForestClassifier(n_estimators=10,
max_depth=9,
criterion="gini",
min_samples_leaf=2,
min_samples_split=2,
random_state=1234)
clf_rf.fit(X_train, y_train)
#テストデータに対する予測データ作成
y_predict = clf_rf.predict(X_test)
# +
#正解率
print(accuracy_score(y_test, y_predict))
# 対数尤度を表示
print('対数尤度 = {:.3f}'.format(- log_loss(y_test, y_predict)))
# 正答率を表示
print('正答率 = {:.3f}%'.format(100 * accuracy_score(y_test, y_predict)))
# Precision, Recall, F1-scoreを計算
precision, recall, f1_score, _ = precision_recall_fscore_support(y_test, y_est)
# カテゴリ「state」に関するPrecision, Recall, F1-scoreを表示
print('適合率(Precision) = {:.3f}%'.format(100 * precision[0]))
print('再現率(Recall) = {:.3f}%'.format(100 * recall[0]))
print('F1値(F1-score) = {:.3f}%'.format(100 * f1_score[0]))
# 予測値と正解のクロス集計
conf_mat = pd.DataFrame(confusion_matrix(y_test, y_predict),
index=['正解 = fail', '正解 = successful'],
columns=['予測 = fail', '予測 = successful'])
conf_mat
# +
# 説明変数の重要度を出力する
# scikit-learnで算出される重要度は、ある説明変数による不純度の減少量合計である。
#特徴量の重要度
feature = clf_rf.feature_importances_
#特徴量の重要度を上から順に出力する
f = pd.DataFrame({'number': range(0, len(feature)),
'feature': feature[:]})
f2 = f.sort_values('feature',ascending=False)
f3 = f2.ix[:, 'number']
#特徴量の名前
label = df[["goal_per_days_standardized","deadline_days_standardized","name_length_normalized","category_Anthologies","category_Chiptune","name_quick","name_theatre","currency_USD","name_first","name_movie"]].columns[0:]
#特徴量の重要度順(降順)
indices = np.argsort(feature)[::-1]
for i in range(len(feature)):
print (str(i + 1) + " " + str(label[indices[i]]) + " " + str(feature[indices[i]]))
plt.title('Feature Importance')
plt.bar(range(len(feature)),feature[indices], color='lightblue', align='center')
plt.xticks(range(len(feature)), label[indices], rotation=90)
plt.xlim([-1, len(feature)])
plt.tight_layout()
plt.show()
# -
# goal_per_days,deadline_daysが重要だと評価されている
# +
#アダブースト
from sklearn.ensemble import AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import classification_report
clf_ad = AdaBoostClassifier(DecisionTreeClassifier(max_depth=3,
min_samples_leaf=2,
min_samples_split=2,
random_state=1234,
criterion="gini"))
clf_ad.fit(X_train, y_train)
#テストデータに対する予測データ作成
y_predict = clf_ad.predict(X_test)
# +
#正解率
print(accuracy_score(y_test, y_predict))
# 対数尤度を表示
print('対数尤度 = {:.3f}'.format(- log_loss(y_test, y_predict)))
# 正答率を表示
print('正答率 = {:.3f}%'.format(100 * accuracy_score(y_test, y_predict)))
# Precision, Recall, F1-scoreを計算
precision, recall, f1_score, _ = precision_recall_fscore_support(y_test, y_est)
# カテゴリ「state」に関するPrecision, Recall, F1-scoreを表示
print('適合率(Precision) = {:.3f}%'.format(100 * precision[0]))
print('再現率(Recall) = {:.3f}%'.format(100 * recall[0]))
print('F1値(F1-score) = {:.3f}%'.format(100 * f1_score[0]))
# 予測値と正解のクロス集計
conf_mat = pd.DataFrame(confusion_matrix(y_test, y_predict),
index=['正解 = fail', '正解 = successful'],
columns=['予測 = fail', '予測 = successful'])
conf_mat
# +
print(clf_ad.feature_importances_)
feature_name=df[["goal_per_days_standardized","deadline_days_standardized","name_length_normalized","category_Anthologies","category_Chiptune","name_quick","name_theatre","currency_USD","name_first","name_movie"]].columns[0:]
feature_name
# 説明変数の重要度を棒グラフに出力する。
pd.DataFrame(clf_ad.feature_importances_, feature_name).plot.barh(figsize=(10,10))
plt.ylabel("Importance")
plt.xlabel("Features")
plt.show()
# -
y = df["state"].values
X = df[["goal_per_days_standardized","deadline_days_standardized","name_length_normalized","category_Anthologies","category_Chiptune","name_quick","name_theatre","currency_USD","name_first","name_movie", \
"category_Residencies","category_Dance","category_Letterpress","category_Literary Spaces","category_Indie Rock"]].values
#テストデータの分割
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# +
#アダブースト
from sklearn.ensemble import AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import classification_report
from sklearn.feature_selection import RFECV
clf_ad = AdaBoostClassifier(DecisionTreeClassifier(max_depth=3,
min_samples_leaf=2,
min_samples_split=2,
random_state=1234,
criterion="gini"))
# RFECVは交差検証によってステップワイズ法による特徴選択を行う
# cvにはFold(=グループ)の数,scoringには評価指標を指定する
# 今回は回帰なのでneg_mean_absolute_errorを評価指標に指定(分類ならaccuracy)
rfecv = RFECV(clf_ad, cv=10, scoring='accuracy')
rfecv.fit(X_train, y_train)
# -
pd.to_pickle(rfecv,'../1_data/rfecv.model')
#テストデータに対する予測データ作成
y_predict = rfecv.predict(X_test)
# +
#正解率
print(accuracy_score(y_test, y_predict))
# 対数尤度を表示
print('対数尤度 = {:.3f}'.format(- log_loss(y_test, y_predict)))
# 正答率を表示
print('正答率 = {:.3f}%'.format(100 * accuracy_score(y_test, y_predict)))
# Precision, Recall, F1-scoreを計算
precision, recall, f1_score, _ = precision_recall_fscore_support(y_test, y_est)
# カテゴリ「state」に関するPrecision, Recall, F1-scoreを表示
print('適合率(Precision) = {:.3f}%'.format(100 * precision[0]))
print('再現率(Recall) = {:.3f}%'.format(100 * recall[0]))
print('F1値(F1-score) = {:.3f}%'.format(100 * f1_score[0]))
# 予測値と正解のクロス集計
conf_mat = pd.DataFrame(confusion_matrix(y_test, y_predict),
index=['正解 = fail', '正解 = successful'],
columns=['予測 = fail', '予測 = successful'])
conf_mat
# +
print(clf_ad.feature_importances_)
feature_name=df[["goal_per_days_standardized","deadline_days_standardized","name_length_normalized","category_Anthologies","category_Chiptune","name_quick","name_theatre","currency_USD","name_first","name_movie"]].columns[0:]
feature_name
# 説明変数の重要度を棒グラフに出力する。
pd.DataFrame(clf_ad.feature_importances_, feature_name).plot.barh(figsize=(10,10))
plt.ylabel("Importance")
plt.xlabel("Features")
plt.show()
# +
#XGBoost
import xgboost as xgb
from sklearn.model_selection import GridSearchCV
# xgboostモデルの作成
clf_xgb = xgb.XGBClassifier()
# ハイパーパラメータ探索
clf_xgn = GridSearchCV(clf_xgb, {'max_depth': [2,4,6], 'n_estimators': [50,100,200]}, verbose=1)
clf_xgn.fit(X_train, y_train)
print (clf_xgn.best_params_, clf_xgn.best_score_)
# 改めて最適パラメータで学習
clf_xgb = xgb.XGBClassifier(**clf_xgn.best_params_)
clf_xgb.fit(X_train, y_train)
# 学習モデルの保存、読み込み
import pickle
pickle.dump(clf_xgb, open("model_xgb.pkl", "wb"))
clf_xgb = pickle.load(open("model_xgb.pkl", "rb"))
# 学習モデルの評価
pred = clf_xgb.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
# +
#正解率
print(accuracy_score(y_test, pred))
# 対数尤度を表示
print('対数尤度 = {:.3f}'.format(- log_loss(y_test, pred)))
# 正答率を表示
print('正答率 = {:.3f}%'.format(100 * accuracy_score(y_test, pred)))
# Precision, Recall, F1-scoreを計算
precision, recall, f1_score, _ = precision_recall_fscore_support(y_test, pred)
# カテゴリ「state」に関するPrecision, Recall, F1-scoreを表示
print('適合率(Precision) = {:.3f}%'.format(100 * precision[0]))
print('再現率(Recall) = {:.3f}%'.format(100 * recall[0]))
print('F1値(F1-score) = {:.3f}%'.format(100 * f1_score[0]))
# 予測値と正解のクロス集計
conf_mat = pd.DataFrame(confusion_matrix(y_test, y_predict),
index=['正解 = fail', '正解 = successful'],
columns=['予測 = fail', '予測 = successful'])
conf_mat
# +
print(clf_xgb.feature_importances_)
feature_name=df[["goal_per_days_standardized","deadline_days_standardized","name_length_normalized","category_Anthologies","category_Chiptune","name_quick","name_theatre","currency_USD","name_first","name_movie"]].columns[0:]
feature_name
# 説明変数の重要度を棒グラフに出力する。
pd.DataFrame(clf_xgb.feature_importances_, feature_name).plot.barh(figsize=(10,10))
plt.ylabel("Importance")
plt.xlabel("Features")
plt.show()
# +
import lightgbm as lgb
lgb_train = lgb.Dataset(X_train, y_train)
lgb_eval = lgb.Dataset(X_test, y_test, reference=lgb_train)
# LightGBM parameters
params = {
'task': 'train',
'boosting_type': 'gbdt',
'objective': 'multiclass',
'metric': {'multi_logloss'},
'num_class': 3,
'learning_rate': 0.1,
'num_leaves': 23,
'min_data_in_leaf': 1,
'num_iteration': 200,
'verbose': 0
}
# train
gbm = lgb.train(params,
lgb_train,
num_boost_round=50,
valid_sets=lgb_eval,
early_stopping_rounds=10)
y_pred = gbm.predict(X_test, num_iteration=gbm.best_iteration)
y_pred = np.argmax(y_pred, axis=1)
# +
#正解率
print(accuracy_score(y_test, y_pred))
# 対数尤度を表示
print('対数尤度 = {:.3f}'.format(- log_loss(y_test, y_pred)))
# 正答率を表示
print('正答率 = {:.3f}%'.format(100 * accuracy_score(y_test, y_pred)))
# Precision, Recall, F1-scoreを計算
precision, recall, f1_score, _ = precision_recall_fscore_support(y_test, y_pred)
# カテゴリ「state」に関するPrecision, Recall, F1-scoreを表示
print('適合率(Precision) = {:.3f}%'.format(100 * precision[0]))
print('再現率(Recall) = {:.3f}%'.format(100 * recall[0]))
print('F1値(F1-score) = {:.3f}%'.format(100 * f1_score[0]))
# 予測値と正解のクロス集計
conf_mat = pd.DataFrame(confusion_matrix(y_test, y_predict),
index=['正解 = fail', '正解 = successful'],
columns=['予測 = fail', '予測 = successful'])
conf_mat
# -
| 16,021 |
/Lec12.ipynb | 128ede8301a6e0470d32dcb1ede1c136e2f10d47 | [] | no_license | keijak/udemy-pydata | https://github.com/keijak/udemy-pydata | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 84,753 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# 軸の目盛りになるアレイを用意
points = np.arange(-5,5,0.01)
# グリッドを作る
dx,dy=np.meshgrid(points,points)
dx
plt.imshow(dx)
dy
plt.imshow(dy)
# sinを使って、ちょっと計算
z = (np.sin(dx) + np.sin(dy))
z
plt.imshow(z)
#カラーバーを出す
plt.colorbar()
# タイトルを付ける
plt.title("Plot for sin(x)+sin(y)")
# +
# numpy 条件に合った値をとってくる
# 早くないやり方
A = np.array([1,2,3,4])
B= np.array([100,200,300,400])
# 真偽値のアレイ
condition = np.array([True,True,False,False])
# リスト内包表記を使った例
answer = [(a if cond else b) for a,b,cond in zip(A,B,condition)]
answer
# -
# numpy.whereを使う
answer2 = np.where(condition,A,B)
answer2
#np.whereは2次元のアレイにも使える
from numpy.random import randn
arr = randn(5,5)
arr
# 0より小さければ0を。そうでなければ、元の値を。
np.where(arr < 0,0,arr)
# その他の統計的な計算
arr = np.array([[1,2,3],[4,5,6],[7,8,9]])
arr
# 合計
arr.sum()
#計算を進める軸を指定出来る
arr.sum(0)
#平均
arr.mean()
#標準偏差
arr.std()
#分散
arr.var()
# any と all
bool_arr = np.array([True,False,True])
# 1つでもTrueがあるか
bool_arr.any()
# 全部Trueか?
bool_arr.all()
# アレイをソートする
# ランダムなアレイを作って、
arr = randn(5)
arr
# ソートする
arr.sort()
arr
# uniqueも便利
countries = np.array(['France', 'Japan', 'USA', 'Russia','USA','Mexico','Japan'])
np.unique(countries)
# in1d test values in one array
np.in1d(['France','USA','Sweden'],countries)
| 1,576 |
/example/audio_augmenter.ipynb | 7e80b12b838616eabb6c9c1b26f0ee551fbfd2a7 | [
"MIT"
] | permissive | avostryakov/nlpaug | https://github.com/avostryakov/nlpaug | 0 | 0 | MIT | 2019-10-08T12:16:30 | 2019-10-08T06:03:54 | null | Jupyter Notebook | false | false | .py | 222,584 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Given a string S, we can transform every letter individually to be lowercase or uppercase to create another string. Return a list of all possible strings we could create.
#
# **Examples:**
# ```
# Input: S = "a1b2"
# Output: ["a1b2", "a1B2", "A1b2", "A1B2"]
#
# Input: S = "3z4"
# Output: ["3z4", "3Z4"]
#
# Input: S = "12345"
# Output: ["12345"]
# ```
# **Note:**
# ```
# S will be a string with length at most 12.
# S will consist only of letters or digits.
# ```
# +
class Solution(object):
def letterCasePermutation(self, S):
"""
:type S: str
:rtype: List[str]
"""
result = ['']
for letter in S:
temp = []
for item in result:
if letter.isalpha():
temp.append(item + letter.upper())
temp.append(item + letter.lower())
else:
temp.append(item + letter)
result = temp
return result
s = Solution()
s.letterCasePermutation("")
ask Augmenter
# +
aug = naa.MaskAug(sampling_rate=sr, mask_with_noise=False)
augmented_data = aug.augment(data)
librosa_display.waveplot(data, sr=sr, alpha=0.5)
librosa_display.waveplot(augmented_data, sr=sr, color='r', alpha=0.25)
plt.tight_layout()
plt.show()
# -
# # Noise
# +
aug = naa.NoiseAug(noise_factor=0.03)
augmented_data = aug.augment(data)
librosa_display.waveplot(data, sr=sr, alpha=0.5)
librosa_display.waveplot(augmented_data, sr=sr, color='r', alpha=0.25)
plt.tight_layout()
plt.show()
# -
# # Pitch Augmenter
# +
aug = naa.PitchAug(sampling_rate=sr, pitch_factor=(2,3))
augmented_data = aug.augment(data)
librosa_display.waveplot(data, sr=sr, alpha=0.5)
librosa_display.waveplot(augmented_data, sr=sr, color='r', alpha=0.25)
plt.tight_layout()
plt.show()
# -
# # Shift Augmenter
# +
aug = naa.ShiftAug(sampling_rate=sr)
augmented_data = aug.augment(data)
librosa_display.waveplot(data, sr=sr, alpha=0.5)
librosa_display.waveplot(augmented_data, sr=sr, color='r', alpha=0.25)
plt.tight_layout()
plt.show()
# -
# # Speed Augmenter
# +
aug = naa.SpeedAug()
augmented_data = aug.augment(data)
librosa_display.waveplot(data, sr=sr, alpha=0.5)
librosa_display.waveplot(augmented_data, sr=sr, color='r', alpha=0.25)
plt.tight_layout()
plt.show()
| 2,577 |
/jupyter_notebooks/MSEplot_USAsondes_hvClickMap.ipynb | 8cef6af865915b2cc8b29bbb576b048d763e5f6d | [] | no_license | weiming9115/test- | https://github.com/weiming9115/test- | 0 | 3 | null | 2018-01-30T19:29:22 | 2018-01-23T18:41:06 | null | Jupyter Notebook | false | false | .py | 4,802,888 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/maylulu/hello-world/blob/master/Copy_of_00_Notebooks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bOChJSNXtC9g" colab_type="text"
# # Notebook Basics
# + [markdown] id="rSXwaU-ptNG6" colab_type="text"
# <img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/logo.png" width=150>
#
# Welcome to the very first lesson of practicalAI. In this lesson we will learn how to work with the notebook and saving it. If you already know how to use notebooks, feel free to skip this lesson.
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/colab.png" width=150>
#
# **Note**: To run the code in this notebook, follow these steps:
# 1. Sign into your Google account.
# 2. Click the **COPY TO DRIVE** button on the toolbar. This will open the notebook on a new tab.
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/copy_to_drive.png">
#
# 3. Rename this new notebook by removing the **Copy of** part in the title.
# 4. Run the code, make changes, etc. and it's all automatically saved to you personal Google Drive.
#
#
# + [markdown] id="cOEaLCZAu4JQ" colab_type="text"
# # Types of cells
# + [markdown] id="WcOgqq5xvtMn" colab_type="text"
# Notebooks are a great visual way of programming. We will use these notebooks to code in Python and learn the basics of machine learning. First, you need to know that notebooks are made up of cells. Each cell can either be a **code cell** or a **text cell**.
#
# * **text cells**: used for headers and paragraph text.
# * **code cells**: user for holding code.
#
#
#
# + [markdown] id="tBVFofpLutnn" colab_type="text"
# # Creating cells
#
# First, let's make a text cell. To create a cell at a particular location, just click on the spot and create a text cell by clicking on the **➕TEXT** below the *View* button up top. Once you made the cell, click on it and type the following inside it:
#
#
# ```
# ### This is a header
# Hello world!
# ```
# + [markdown] id="iXYgZpgpYS3N" colab_type="text"
# # Running cells
# Once you type inside the cell, press the **SHIFT** and **ENTER** together to run the cell.
# + [markdown] id="WKTbiBuvYexD" colab_type="text"
# # Editing cells
# To edit a cell, double click it and you should be able to replace what you've typed in there.
# + [markdown] id="Jv0ZSuhNYVIU" colab_type="text"
# # Moving cells
# Once you make the cell, you can move it with the ⬆️**CELL** and ⬇️**CELL** buttons above.
# + [markdown] id="B_VGiYf8YXiU" colab_type="text"
# # Deleting cells
# You can delete the cell by clicking on the cell and pressing the button with three vertical dots on the top right corner of the cell. Click **Delete cell**.
# + [markdown] id="hxl7Fk8LVQmR" colab_type="text"
# # Making a code cell
# Now let's take the same steps as above to make, edit and delete a code cell. You can create a code cell by clicking on the ➕CODE below the *File* button up top. Once you made the cell, click on it and type the following inside it:
#
# ```
# print ("hello world!")
# ```
#
# ⏰ - It may take a few seconds when you run your first code cell.
# + id="DfGf9KmQ3DJM" colab_type="code" outputId="973bea44-6c15-47a3-d789-69386e11b6f4" colab={"base_uri": "https://localhost:8080/", "height": 34}
print ("hello world!")
# + [markdown] id="GURvB6XzWN12" colab_type="text"
# **Note:** These Google colab notebooks timeout if you are idle for more than ~30 minutes which means you'll need to run all your code cells again.
# + [markdown] id="VoMq0eFRvugb" colab_type="text"
# # Saving the notebook
# + [markdown] id="nPWxXt5Hv7Ga" colab_type="text"
# Go to *File* and then click on **Save a copy in Drive**. Now you will have your own copy of each notebook in your own Google Drive. If you have a [Github](https://github.com/), you can explore saving it there or even downloading it as a .ipynb or .py file.
get_upper_air_data(t,station_code)
p = dataset['pressure'].values
T = dataset['temperature'].values
Td = dataset['dewpoint'].values
Z = dataset['height'].values
q = mixing_ratio(saturation_vapor_pressure(Td*units.degC),p*units.mbar)
q = specific_humidity_from_mixing_ratio(q)
# ax=mpt.msed_plots(p,T,q,Z,entrain=entrain,title=station_code+' '+str(t))
ax=mpt.msed_plots(p,T,q,Z,entrain=entrain)
label.value="Showing {} at {}".format(station_code,t)
except:
label.value="No data available at {} for this date {}".format(station_code,t)
# ### Setup additional interaction through widgets (time, loading status, checkbox for entraining parcels option)
#setup widgets to interact with energy plot
dates=pd.date_range(datetime(2013,1,1,0),datetime(2019,1,1,0),freq='D')
label=widgets.Label('')
sw=widgets.fixed('MFL')
entrainw=widgets.Checkbox(value=False,description='Entraining Plumes')
tw=widgets.SelectionSlider(options=dates,description="Time",continuous_update=False,layout=widgets.Layout(width='90%'))
# ### A method that responds to click on the interactive station map
def update_plot(x,y):
"""A call method which will be called when user clicks on the point on the map.
Routine finds the closest datapoint to the click location and updates the widget.
"""
if x:
try:
#this doesn't work because xarray cannot select nearest location for point data data structutre
station_code=str(da.sel(lon=x,lat=y,method='nearest').values)
label.value=str(station_code)
except:
#uses L1 degree distance (absoulte of distance) to find closest location to click
# below inline function finds the index of the nearest data location to click
l1dist=lambda x,y:np.argmin([abs(lo-x)+abs(la-y) for lo,la in list(zip(da.lon.values,da.lat.values))])
station_code=str(da[l1dist(x,y)].values)
stationlonlat=str(da[l1dist(x,y)].lonlat.values)
label.value='Loading '+str(station_code)+f' (lon,lat: {stationlonlat})'
else:
station_code='MFL'+str(abs(da.lon.values-x))
#label.value='Loading station {} at {}'.format(station_code,str(tw.value))
sw.value=station_code
label.value=''
tap_stream.add_subscriber(update_plot) #attach callback method for a tap stream
# ### Create an output area for all the interactive plots
output_area=widgets.Output() #create a area to stack all interactive plots in one cell
output_area #click on the points on the map to explore the energy mass plot
#makes changes to the cell area above, attaches matplotlib plot and widget interaction.
output_area.append_display_data(station_map) #put map in the cell area
with output_area:
display(widgets.VBox([tw,label,entrainw,widgets.interactive_output(energy_plot,{'station_code':sw,'t':tw,'entrain':entrainw})]))
#above line attaches energy plot with its time widget
| 7,368 |
/analysis_notebooks/8.CERAD_style_CNN_scores.ipynb | fc9974f728042080d27f28822fb1054a72e96ae9 | [] | no_license | gutmanlab/Emory_Plaquebox_Paper | https://github.com/gutmanlab/Emory_Plaquebox_Paper | 1 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 16,434 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 8) CERAD Style CNN Scores
#
# Modify the CNN scoring strategy for the confidence heatmaps. Original approach was to do blob detection on confidence heatmaps and divide the count of blobs by the number of pixels containing tissue (calculated by thresholding strategy on the image).
#
# Of interest is to generate these scores for small isolated regions of the images, which is similar to how pathologist score CERAD. In the traditional approach the pathologist only have a small field of view or FOV (viewed under a microscope) which they use for analyzing plaque burden. We want to know if generating scores in this fashion yields similar or different results then using the whole tissue region. Pathologist by protocol only assess the densest FOV, which we can identify here. We will call this strategy one.
# +
# %reset -f
# cell to load required data and imports
import matplotlib.pyplot as plt
from os.path import join as oj
import imageio
import numpy as np
import pandas as pd
import sys
import os
from collections import namedtuple
from tqdm import tqdm_notebook as tqdm
from scipy.stats import spearmanr
from skimage.transform import resize
from PIL import Image, ImageDraw
sys.path.append('..')
from modules import heatmap_analysis as ha, plotting
blob_mask_dir = '/mnt/data/outputs/blob_masks/'
fov_scores_dir = '/mnt/data/outputs/fov_scores/'
data_path = '/mnt/AB_Plaque_Box/CSVs/Emory_data.csv'
labels = ('cored', 'diffuse', 'caa')
fov_labels = ('cored_densest_fov', 'diffuse_densest_fov', 'caa_densest_fov')
# -
# #### Score FOV regions
#
# Slide a FOV region accross the blob mask (confidence heatmaps processed for blob detection) and score each FOV region by unique blobs in the FOV divide by the FOV area.
#
# (1) Ideally the FOV should mostly be containing tissue content and avoid non-tissue space. In reality the tissue threshold method is rife with noise and using this as a determination of tissue region would yield noisy results. Alternatively we assume that all region image is tissue and slide a small analysis region across the image with a small stride and report the highest blob count in the region regardless of what is under.
#
# (2) The best solution, though not done here, would be to carefully annotate the entire tissue region and use that as ground truth for what pixels contain tissue and what don't.
#
# (3) The output of this will be a csv file for each image containing columns: x, y, fov height, fov width, plaque type, count. Only FOV with at least one plaque will be saved, all others are assumed to be 0.
# +
# run for all images and save csv to a fov_scores dir
# run using default FOV shape of 128 by 128 and stride of 16
os.makedirs(fov_scores_dir, exist_ok=True)
for blob_mask_file in tqdm(os.listdir(blob_mask_dir)):
save_path = oj(fov_scores_dir, blob_mask_file.replace('.npy', '.csv'))
# don't run if save path already exists - might mean it was previously run
if not os.path.isfile(save_path):
ha.conf_heatmap_fov_scores(oj(blob_mask_dir, blob_mask_file), save_path, fov_shape=(251, 251))
# -
# #### Tissue CNN vs FOV CNN score correlations
#
# For the average of the n densest FOV (highest CNN scores) plot the correlation between that CNN score and the original CNN score (whole tissue).
#
# Side-by-side plot the box plots of the CNN scores grouped by CERAD-like categories.
# +
# load main data file
df = pd.read_csv(data_path)
Rectangle = namedtuple('Rectangle', 'xmin ymin xmax ymax')
def area(a, b): # returns None if rectangles don't intersect
dx = min(a.xmax, b.xmax) - max(a.xmin, b.xmin)
dy = min(a.ymax, b.ymax) - max(a.ymin, b.ymin)
if (dx>=0) and (dy>=0):
return dx*dy
else:
return 0
def fov_correlation(data_path, fov_dir, n=1, plot=False):
"""Plot correlation of the average n denset FOV in each image vs the whole tissue image.
:param data_path : str
path to data file for case info, CERAD categories, and whole tissue scores
:param fov_dir : str
the dir containing the fov csv files for each case
:param n : int (default of 1)
the n highest FOV scores to use
"""
labels=('cored', 'diffuse', 'caa')
fov_labels = ('cored_densest_fov', 'diffuse_densest_fov', 'caa_densest_fov')
tissue_labels = ('tissue_cored_score', 'tissue_diffuse_score', 'tissue_caa_score')
category_labels = ('Cored_MTG', 'Diffuse_MTG', 'CAA_MTG')
df = pd.read_csv(data_path)
# add a column for densest FOV count for each case's plaques types (cored, diffuse, caa)
densest_scores = [[], [], []]
for idx, row in df.iterrows():
# load the fov scores for this case
fov_df = pd.read_csv(oj(fov_scores_dir, row['WSI_ID'] + '.csv'))
for i, label in enumerate(labels):
# subset each type of plaque
df_subset = fov_df[fov_df['label']==label].reset_index(drop=True)
# sort by counts
df_subset = df_subset.sort_values(by=['count'], ascending=False).reset_index(drop=True)
# df subset is 0 then just return 0
if len(df_subset) == 0:
densest_scores[i].append(0)
continue
# take the n highest non-overlapping counts
x, y = df_subset.loc[0, 'x'], df_subset.loc[0, 'y']
w, h = df_subset.loc[0, 'width'], df_subset.loc[0, 'height']
scores = [df_subset.loc[0, 'count'] * 100 / (w * h)]
coords = [Rectangle(x, y, x + w, y + h)]
for j in range(1, len(df_subset)):
# if we already have the top n counts, then stop
if len(scores) == n:
break
# else check the this region to make sure it does not overlap with previous regions
x, y = df_subset.loc[j, 'x'], df_subset.loc[j, 'y']
w, h = df_subset.loc[j, 'width'], df_subset.loc[j, 'height']
r = Rectangle(x, y, x + w, y + h)
# for all already region check
flag = False
for coord in coords:
if area(coord, r) != 0:
flag = True
break
# if flag did not change to true then this region does not overlap, add it
if not flag:
coords.append(r)
scores.append(df_subset.loc[j, 'count'] * 100 / (w * h))
# convert scores to average and append to denset list - note that there may be less than n
# entries if not enough non-overlapping regions
densest_scores[i].append(sum(scores) / len(scores))
# add the densest fov column (scores)
for i in range(3):
df[fov_labels[i]] = densest_scores[i]
# sort the tissue scores and densest FOV scores and plot against each other
rhos = []
for i in range(3):
tissue_scores = df[tissue_labels[i]].tolist()
fov_scores = df[fov_labels[i]].tolist()
# return spearman correlation with the CERAD like scores
rho, p = spearmanr(fov_scores, df[category_labels[i]].tolist())
rhos.append(rho)
# calaculate spearman correlation between tissue and fov scores
rho, p = spearmanr(tissue_scores, fov_scores)
#
# sort the lists together before plotting
tissue_scores, fov_scores = (list(t) for t in zip(*sorted(zip(tissue_scores, fov_scores))))
if plot:
plt.figure(figsize=(7, 5))
plt.plot(tissue_scores, fov_scores, '.-')
plt.xlabel('Tissue CNN Scores', fontsize=16)
plt.ylabel('{} FOV CNN-Scores'.format(n), fontsize=16)
plt.title('r: {:0.3f}'.format(float(rho)))
plt.show()
x_param = {0: 'None', 1: 'Sparse', 2: 'Moderate', 3: 'Frequent'}
plotting.plot_cnn_scores(df, category_labels[i], fov_labels[i], x_param,
ylabel='{} Highest FOV CNN-Scores'.format(n),
xlabel='CERAD-like scores')
return rhos
output = []
rhos = fov_correlation(data_path, fov_scores_dir, n=1, plot=True)
# -
# #### Plot the effect of increasing FOV regions used to estimate CNN score vs Whole Tissue
# +
# for each plauqe get the spearman correlation coeff between the n average FOV vs CERAD categories
output = []
N = 16
for n in range(1, N):
rhos = fov_correlation(data_path, fov_scores_dir, n=n)
output.append(rhos)
output = np.array(output)
# add the spearman corrrelation coeff for whole tissue as last point
r_cored, p = spearmanr(df['Cored_MTG'].tolist(), df['tissue_cored_score'].tolist())
r_diffuse, p = spearmanr(df['Diffuse_MTG'].tolist(), df['tissue_diffuse_score'].tolist())
r_caa, p = spearmanr(df['CAA_MTG'].tolist(), df['tissue_caa_score'].tolist())
r_tissue = [r_cored, r_diffuse, r_caa]
fig, ax = plt.subplots(1, 1, figsize=(10, 7))
x = list(range(1, N+1))
for i in range(3):
fov_scores = list(output[:, i]) + [r_tissue[i]]
ax.plot(x, fov_scores, 'o-')
ax.set_xticks(x)
x_tick_labels = [str(_x) for _x in x[:-1]] + ['WT']
ax.set_xticklabels(x_tick_labels)
ax.legend(['Cored', 'Diffuse', 'CAA'], fontsize=16)
ax.set_ylabel('Spearman Coefficient', fontsize=20)
ax.xaxis.set_tick_params(labelsize=18, size=0, width=2)
ax.yaxis.set_tick_params(labelsize=18, size=5, width=2)
ax.set_xlabel('Avg. of n-highest FOV scores', fontsize=20)
fig.savefig('/mnt/data/figures/fov_correlations.png', bbox_inches='tight', dpi=300)
plt.show()
# -
# #### Use on example to show the three top FOV regions
# Re-using a lot of my code above.
# +
Rectangle = namedtuple('Rectangle', 'xmin ymin xmax ymax')
def area(a, b): # returns None if rectangles don't intersect
dx = min(a.xmax, b.xmax) - max(a.xmin, b.xmin)
dy = min(a.ymax, b.ymax) - max(a.ymin, b.ymin)
if (dx>=0) and (dy>=0):
return dx*dy
else:
return 0
# -
fov_df = pd.read_csv('/mnt/data/outputs/fov_scores/AB_case_4.csv')
hm = np.load('/mnt/data/outputs/heatmaps_emory/AB_case_4.npy')[0]
im = imageio.imread('/mnt/data/figures/AB_case_4.jpeg')
im = resize(im, hm.shape)
# +
# just show a cored example
df_subset = fov_df[fov_df['label']=='cored'].reset_index(drop=True)
# sort by counts
df_subset = df_subset.sort_values(by=['count'], ascending=False).reset_index(drop=True)
# take the n highest non-overlapping counts
x, y = df_subset.loc[0, 'x'], df_subset.loc[0, 'y']
w, h = df_subset.loc[0, 'width'], df_subset.loc[0, 'height']
scores = [df_subset.loc[0, 'count'] * 100 / (w * h)]
coords = [Rectangle(x, y, x + w, y + h)]
n = 3
for j in range(1, len(df_subset)):
# if we already have the top n counts, then stop
if len(scores) == n:
break
# else check the this region to make sure it does not overlap with previous regions
x, y = df_subset.loc[j, 'x'], df_subset.loc[j, 'y']
w, h = df_subset.loc[j, 'width'], df_subset.loc[j, 'height']
r = Rectangle(x, y, x + w, y + h)
# for all already region check
flag = False
for coord in coords:
if area(coord, r) != 0:
flag = True
break
# if flag did not change to true then this region does not overlap, add it
if not flag:
coords.append(r)
scores.append(df_subset.loc[j, 'count'] * 100 / (w * h))
# +
def draw_rectangle(draw, coordinates, color, width=1):
for i in range(width):
rect_start = (coordinates[0][0] - i, coordinates[0][1] - i)
rect_end = (coordinates[1][0] + i, coordinates[1][1] + i)
draw.rectangle((rect_start, rect_end), outline = color)
# draw the regions on the image
im_copy = Image.fromarray((im * 255).astype(np.uint8))
draw = ImageDraw.Draw(im_copy)
outline_width = 10
colors = ['red', 'orange', 'green']
for i, coord in enumerate(coords):
draw_rectangle(draw, [(coord.xmin, coord.ymin), (coord.xmax, coord.ymax)], color=colors[i],
width=outline_width)
del draw
im_copy.save('/mnt/data/figures/fov_boxes.png')
| 12,596 |
/DS18_422.ipynb | 72284df649e6ab9e94bd10eb2bb469c6bf5d14ec | [] | no_license | Jace-Hambrick/Assignments | https://github.com/Jace-Hambrick/Assignments | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 228,319 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NGGrt9EYlCqY"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# # Train Practice
#
# ## *Data Science Unit 4 Sprint 2 Assignment 2*
#
# Continue to use TensorFlow Keras & a sample of the [Quickdraw dataset](https://github.com/googlecreativelab/quickdraw-dataset) to build a sketch classification model. The dataset has been sampled to only 10 classes and 10000 observations per class. Please build a baseline classification model then run a few experiments with different optimizers and learning rates.
#
# *Don't forgot to switch to GPU on Colab!*
# -
import numpy as np
from sklearn.model_selection import train_test_split
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.optimizers import SGD
# + [markdown] id="ptJ2b3wk62Ud" colab_type="text"
# ### Write a function to load your data
#
# Wrap yesterday's preprocessing steps into a function that returns four items:
# * X_train
# * y_train
# * X_test
# * y_test
#
# Your function should accept a `path` to the data as a argument.
# + id="nJsIsrvp7O3e" colab_type="code" colab={}
def load_quickdraw10(path):
data = np.load('quickdraw10.npz')
X = data['arr_0']
y = data['arr_1']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
return X_train, y_train, X_test, y_test
# -
X_train, y_train, X_test, y_test = load_quickdraw10(r'U4W2\module2-Train')
X_train
# + [markdown] id="l-6PxI6H5__2" colab_type="text"
# ### Write a Model Function
# Using your model from yesterday, write a function called `create_model` which returns a compiled TensorFlow Keras Sequential Model suitable for classifying the QuickDraw-10 dataset. Include parameters for the following:
# * Learning Rate
# * Optimizer
# -
data = np.load('quickdraw10.npz')
X = data['arr_0']
y = data['arr_1']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# + colab_type="code" id="nEREYT-3wI1f" colab={}
##### Your Code Here #####
def create_model(lr,mizer):
opt = mizer(learning_rate=lr)
model = Sequential(
[Dense(32, activation='relu', input_dim=784),
Dense(32, activation='relu'),
Dense(10, activation='softmax')]
)
model.compile(optimizer=opt, loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
return model
# -
create_model(.01,SGD)
# + [markdown] id="f0pCkh8C7eGL" colab_type="text"
# ### Experiment with Batch Size
# * Run 5 experiments with various batch sizes of your choice.
# * Visualize the results
# * Write up an analysis of the experiments and select the "best" performing model among your experiments. Make sure to compare against your model's performance yesterday.
# -
from tensorflow.keras.layers import Dense
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import SGD
import seaborn as sns
import pandas as pd
model = create_model(.01,SGD)
batch1 = model.fit(X_train, y_train,
epochs=10,
batch_size=8,
validation_data=(X_test, y_test))
model = create_model(.01,SGD)
batch2 = model.fit(X_train, y_train,
epochs=10,
batch_size=16,
validation_data=(X_test, y_test))
model = create_model(.01,SGD)
batch3 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
model = create_model(.01,SGD)
batch4 = model.fit(X_train, y_train,
epochs=10,
batch_size=64,
validation_data=(X_test, y_test))
model = create_model(.01,SGD)
batch5 = model.fit(X_train, y_train,
epochs=10,
batch_size=128,
validation_data=(X_test, y_test))
# +
batch_sizes = []
for exp, result in zip([batch1,batch2,batch3,batch4,batch5], ["8_","16_","32_","64_","128_"]):
df = pd.DataFrame.from_dict(exp.history)
df['epoch'] = df.index.values
df['Batch Size'] = result
batch_sizes.append(df)
df = pd.concat(batch_sizes)
df['Batch Size'] = df['Batch Size'].astype('str')
df.head()
# -
sns.lineplot(x='epoch', y='val_accuracy', hue='Batch Size', data=df);
# + [markdown] colab_type="text" id="8b-r70o8p2Dm"
# ### Experiment with Learning Rate
# * Run 5 experiments with various learning rate magnitudes: 1, .1, .01, .001, .0001.
# * Use the "best" batch size from the previous experiment
# * Visualize the results
# * Write up an analysis of the experiments and select the "best" performing model among your experiments. Make sure to compare against the previous experiments and your model's performance yesterday.
# + id="_SA144xx8Luf" colab_type="code" colab={}
model = create_model(.0001,SGD)
learn1 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
# -
model = create_model(.001,SGD)
learn2 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
model = create_model(.01,SGD)
learn3 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
model = create_model(.1,SGD)
learn4 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
model = create_model(1,SGD)
learn5 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
# +
import seaborn as sns
import pandas as pd
learning_rates = []
for exp, result in zip([learn1,learn2,learn3,learn4,learn5], [".0001_",".001_",".01_",".1_","1_"]):
df = pd.DataFrame.from_dict(exp.history)
df['epoch'] = df.index.values
df['Learning Rate'] = result
learning_rates.append(df)
df = pd.concat(learning_rates)
df['Learning Rate'] = df['Learning Rate'].astype('str')
df.head()
# -
sns.lineplot(x='epoch', y='val_accuracy', hue='Learning Rate', data=df);
# + [markdown] id="gxMtSRhV9Q7I" colab_type="text"
# ### Experiment with different Optimizers
# * Run 5 experiments with various optimizers available in TensorFlow. See list [here](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers)
# * Visualize the results
# * Write up an analysis of the experiments and select the "best" performing model among your experiments. Make sure to compare against the previous experiments and your model's performance yesterday.
# * Repeat the experiment combining Learning Rate and different optimizers. Does the best performing model change?
# + id="ujLuzdNA91ip" colab_type="code" colab={}
model = create_model(.01,SGD)
optim1 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
# -
from tensorflow.keras.optimizers import Adam
model = create_model(.01,Adam)
optim2 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
from tensorflow.keras.optimizers import Adamax
model = create_model(.01,Adamax)
optim3 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
from tensorflow.keras.optimizers import Ftrl
model = create_model(.01,Ftrl)
optim4 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
from tensorflow.keras.optimizers import Adadelta
model = create_model(.01,Adadelta)
optim5 = model.fit(X_train, y_train,
epochs=10,
batch_size=32,
validation_data=(X_test, y_test))
# +
import seaborn as sns
import pandas as pd
optimizers = []
for exp, result in zip([optim1,optim2,optim3,optim4,optim5], ["SGD","Adam","Adamax","Ftrl","Adadelta"]):
df = pd.DataFrame.from_dict(exp.history)
df['epoch'] = df.index.values
df['optimizer'] = result
optimizers.append(df)
df = pd.concat(optimizers)
df['optimizer'] = df['optimizer'].astype('str')
df.head()
# -
sns.lineplot(x='epoch', y='val_accuracy', hue='optimizer', data=df);
# + [markdown] id="ydAqeY9S8uHA" colab_type="text"
# ### Additional Written Tasks
#
# 1. Describe the process of backpropagation in your own words:
# ```
# Your answer goes here.
# ```
#
#
# + [markdown] colab_type="text" id="FwlRJSfBlCvy"
# ## Stretch Goals:
#
# - Implement GridSearch on anyone of the experiments
# - On the learning rate experiments, implement [EarlyStopping](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping)
# - Review material on the math behind gradient descent:
#
# - Gradient Descent
# - Gradient Descent, Step-by-Step by StatQuest w/ Josh Starmer. This will help you understand the gradient descent based optimization that happens underneath the hood of neural networks. It uses a non-neural network example, which I believe is a gentler introduction. You will hear me refer to this technique as "vanilla" gradient descent.
# - Stochastic Gradient Descent, Clearly Explained!!! by StatQuest w/ Josh Starmer. This builds on the techniques in the previous video. This technique is the one that is actually implemented inside modern 'nets.
# These are great resources to help you understand tomorrow's material at a deeper level. I highly recommend watching these ahead of tomorrow.
#
# - Background Math
# - Dot products and duality by 3Blue1Brown. Explains the core linear algebra operation happening in today's perceptron.
# The paradox of the derivative by 3Blue1Brown. Does a great job explaining a derivative.
# - Visualizing the chain rule and product rule by 3Blue1Brown. Explains the black magic that happens within Stochastic Gradient Descent.
# These math resources are very much optional. They can be very heady, but I encourage you to explore. Your understanding of neural networks will greatly increase if you understand this math background.
#
#
#
| 10,839 |
/Linear Regression with G.D. (ml-coursera-dataset-ex1).ipynb | 41be15ab0096a56ef1244c671c3a46acbe38741b | [] | no_license | Budeestar/My-repo | https://github.com/Budeestar/My-repo | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 39,532 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df=pd.read_csv('/home/aasim/ex1data1.txt',delimiter=',')
newlist=df.values
m,n=newlist.shape
trainsize=round(0.7*m) #round() is used to round off
testsize=m-trainsize
#training set
X=newlist[0:trainsize,0].reshape(trainsize,1)
Y=newlist[0:trainsize,1].reshape(trainsize,1)
#test set
Xtest=newlist[trainsize:m+1,0].reshape(testsize,1)
Ytest=newlist[trainsize:m+1,0].reshape(testsize,1)
print(X.shape,Y.shape,Xtest.shape,Ytest.shape)
#training set plot
plt.scatter(X,Y,marker='.',color='red')
plt.xlabel('X')
plt.ylabel('Y')
plt.show()
# +
#begin training
np.random.seed(1)
theta=np.random.rand(2).reshape(2,1)
X=np.hstack((np.ones((trainsize,1)),X))
iters=2000
learning_rate=0.01
J=np.zeros(iters)
for i in range(iters):
h=np.sum((np.dot(X,theta)-Y)**2)
J[i]=h/(2*trainsize)
theta[0]-=(learning_rate/trainsize)*np.sum(np.dot(X,theta)-Y)
theta[1]-=(learning_rate/trainsize)*np.sum((np.dot(X,theta)-Y)*X)
print(J[0],J[-1])
iters_matrix=np.arange(iters)
plt.plot(iters_matrix,J)
plt.xlabel('Number of Iterations',color='red')
plt.ylabel('Cost J',color='red')
plt.title('Cost vs no. of iterations',color='red')
plt.axis([0,2000,3.5,5.2])
plt.show()
# -
plt.scatter(X[:,1],Y,marker='.',color='red')
plt.plot(X,np.dot(X,theta),color='green')
plt.axis([4,23,-0.5,25])
plt.xlabel('X',color='red')
plt.ylabel('Y',color='red')
plt.title('Line of best fit')
plt.show()
| 1,718 |
/0430.ipynb | f221a393b4092b9a68a82734763d4b743c7803cc | [] | no_license | daniuniuniuniu/spark | https://github.com/daniuniuniuniu/spark | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 72,558 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
crawled_list = []
created_headers = {}
dict_col_names = {'Pld':'Games_Played', 'W':'Games_Won', 'D': 'Games_Drew', 'L':'Games_Lost',\
'GF':'Goals_For','GA':'Goals_Against','GD':'Goal_Difference','Pts':'Total_Points'}
# +
def trimStrBy1 (str):
return str[:len(str)-1]
def getFullName (str):
str = str.strip().replace("\n","").replace(" ","_")
#
if "[" in str:
str = str[:str.find("[")]
if str not in dict_col_names:
return str
else:
return dict_col_names[str]
# +
from bs4 import BeautifulSoup
from urllib import request
from lxml.html import fromstring
import re
import csv
import pandas as pd
def getTableFrame(table):
tmp = table.find_all('tr')
first = tmp[0]
allRows = tmp[1:]
headers = [header.get_text() for header in first.find_all('th')]
new_headers = []
new_header = ''
all_headers = ''
for header in headers:
new_header = getFullName(header)
new_headers.append(new_header)
all_headers = all_headers + new_header + ","
all_headers = trimStrBy1(all_headers)
results = [[data.get_text() for data in row.find_all('td')] for row in allRows]
rowspan = []
for row_no, tr in enumerate(allRows):
tmp = []
for col_no, data in enumerate(tr.find_all('td')):
if data.has_attr("rowspan"):
rowspan.append((row_no, col_no, int(data["rowspan"]), data.get_text()))
if rowspan:
for i in rowspan:
for j in range(1, i[2]):
results[i[0]+j].insert(i[1], i[3])
df = pd.DataFrame(data=results, columns=new_headers)
return df, all_headers
# -
def insertIntoDB(league, league_year, success_count, df, all_headers):
conn = sqlite3.connect('top5tables.sqlite')
cur = conn.cursor()
table_name = league + '_' + league_year + '_' + success_count
df.to_sql(con=conn, name=table_name, if_exists='append', flavor='sqlite')
conn.commit()
# +
import wikipedia
from wikipedia import WikipediaPage
from lxml import html
from bs4 import BeautifulSoup
import xml.etree.ElementTree as ET
import sqlite3
import re
all_tables = []
single_table = []
single_row = []
leagues = [' FA Premier League', ' Premier League', ' La Liga', ' Serie A', ' Bundesliga', ' Ligue 1', ' French Division 1']
years = [1992, 1993, 1994, 1995, 1996, 1997, 1998, 1999, 2000, 2001, 2002, 2003, \
2004, 2005, 2006, 2007, 2008, 2009, 2010, 2011, 2012, 2013, 2014, 2015]
dashes = ['—','–','‐']
'1992–93 FA Premier League'
'1992–93 FA Premier League'
request_count = 0
success_count = 0
duplicate_count = 0
for year in years:
for league in leagues:
for dash in dashes:
if year == 1999: league_year = str(year) + dash + '2000'
else: league_year = str(year) + dash + str(year + 1)[2:]
title = league_year + league
try:
request_count += 1
print ("\n\nGetting page:",title)
parsed_html = BeautifulSoup(wikipedia.page(title).html(), "lxml")
page_title = wikipedia.page(title).title
print (" Wikipedia Page Title:", page_title)
except wikipedia.exceptions.PageError:
print (" Page Error:",title)
except:
print (" General Exception:",title)
if page_title not in crawled_list:
tables = parsed_html.findAll('table', attrs={'class':'wikitable'})
table_count = 0
for table in tables:
if len([header.get_text() for header in table.find_all('tr')[0].find_all('th')]) > 0:
df, all_headers = getTableFrame(table)
print ("\n\t",all_headers)
table_count += 1
else:
continue
#insertIntoDB(league, league_year, str(table_count), df, all_headers)
print (" Freshly crawled:",page_title)
success_count += 1
crawled_list.append(page_title)
else:
duplicate_count += 1
print (" Already crawled:",page_title)
print ("Page Requested:", request_count)
print ("Success:", success_count)
print ("Duplicate:", duplicate_count)
bler = VectorAssembler(inputCols=["subreddit_si", "controversiality", "no_follow", "collapsed"], outputCol="features")
### Split dataset randomly into Training and Test sets
(train, test) = df_tran.randomSplit([0.7, 0.3], seed = 100)
# Naivebayes Model
from pyspark.ml.classification import NaiveBayes
## Train a NaiveBayes model
nb = NaiveBayes(smoothing=1.0, modelType="multinomial",labelCol = 'score_bin_si',featuresCol='features')
# +
# Chain vecAssembler and NBmodel in a pipeline
pipeline = Pipeline(stages=[vecAssembler, nb])
# Run stages in pipeline and train model
model = pipeline.fit(train)
# +
# Make predictions on test data so we can measure the accuracy of our model on new data
predictions = model.transform(test)
predictions.printSchema()
# +
## see the result
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
evaluator = MulticlassClassificationEvaluator(labelCol="score_bin_si", predictionCol="prediction",
metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Model Accuracy: ",accuracy)
# -
# Random Forest
from pyspark.ml.classification import RandomForestClassifier
## using the RandomForestClassifier estimator to build model
rf = RandomForestClassifier(labelCol="score_bin_si", featuresCol="features")
# Chain vecAssembler and NBmodel in a pipeline
pipeline_rf = Pipeline(stages=[vecAssembler, rf])
# Run stages in pipeline and train model
model_rf = pipeline_rf.fit(train)
# +
# Make predictions on test data so we can measure the accuracy of our model on new data
predictions_rf = model_rf.transform(test)
evaluatorRF = MulticlassClassificationEvaluator(labelCol="score_bin_si", predictionCol="prediction", metricName="accuracy")
accuracy_rf = evaluatorRF.evaluate(predictions_rf)
print("Accuracy = %g" % accuracy_rf)
print("Test Error = %g" % (1.0 - accuracy_rf))
# -
spark.stop()
LG+bA7WNCZJl8jI+383fHW566gXNpoQDdI4YteSYb9kX2j5Vf0iF6wsDcVvEYPEo+Mb1by7XGEeRFPrL24MNVjNFl+REMcjFn7OwrBbbBtK8j2Ivu/FcSehak8EmuvTH56Bv2MU4bK3o+meF599gUbaI5Rv04DOYcd779nnMwrfUvLxP11rl/gGXOBm+zncNuzEHsK+ShXqt8yIx80BNLaHeOFDoO7WOeTaQjbB3aRqket//aa69FzTfnQc+p+mBzwyooAiIgAttIYMevXn4wnXXBfennr3s47Tp2Ip1/5IMNNDbTFxx7eXH9rAvuTRdeeWiwuqXJNScUmzgs6FAul4/rSLnNaEK2cvbUee/evRtPXGwBgbuy0YIOspGOrQ9bbMLjiQ3yLcXigycpti1Xj2UMOYZuOXZDZEZ1YF+0eBu7WCktJHxclfwNPTzrXH5k5xR5aK/mE5TzNnodYHMp/hAHpTLw4TrFqOlsb3O3DS1/EsUz4vO5F4w9OsHHpRgfeq0WN8ykdtzDDPHoxwL7+L19DN7/RXHny/Scg2mtf7HM06V/gKVnz7b6Y/DCuFial/haji/8nxtrWmKJ2xkS/zyGeXtxDruHyM/ZBtm9KdsLP/TKGFPe+qZ9PcWeULeOo2PaU10REAERWAUCO37xNx9cbKI/tvdYuuCRVzaeQGMDbekFj5xMH9v7d8k20R+//OBgvXmgz02gXjgWJ35Bh4nU5/v6Q9pkGVhU1NrhOnxcqw/7bFKN3iIOWZDDkzvbNtfECf1a/QV9e1PY17N4622jpXxJDyyaPOtcfkt7Q8qgvZpP4Lta7PKi1T/lM/0g53SMUYwj3K9qPhlSpyZzXa/jDfj2dN1vJuzp9U033dT99QjEY+tYgD7b48MS79b+BRmnU/8Ay1b2xgC8MC7W5iXwyrWB6zl/tvQ/1sHHZct5rm34vJSCobUTjaelukOvsb3ww1BZXG+O/s3ydSwCIiAC60xgxxdveCSddeF96aO/dzh94q+/n3YdPbFpI23nlv/R3z2czrrw3vQ7Nz02yl5MuLUNABrJlcdEVZvssCCzCW3I5IL2h9Q1G3L1bdLjt12WNtAmJ7J3iokTLzmy1CZMu5tsv8FsT+bsBTZY0LT6C37rTWFfbmHVK29o+ZIeOV/m8ofqUKuH9mo+wWKzxjRX7kyIUdheG0fYJ611uH+2LNx9mR6dWL9lHFs/wU+Beb2jc4vB1o+DY8z2cctj1bPPPrsYp77xjW+EY+MYBq3963TsH6XxD3PE4cOHFy/Dw5wFXpgjOe6Rx/7g6+ZjvIPC8g8ePLhxMyaqa3Ja+x+32Xo8hWwwtH6wrpvoOft3qy9UTgREQARWncCOBx7+Xtr56b9YvIH7l/70f73/NBof6T5iT6FfSZZ/zq9/fVHuwNHvjrKpZ5LCYsomIz+h8kRcmqjQnl+QtRiByXBIXZOfq+9/CsbbFukGWbywZgYsg7lFC9qWPNg8ZBONOqxrZBPnwT60y9f4GP5ssSEqU9MJekR1kcescz5gnac+xqK1tonmOGCdvT6Qx0+sT/cYBYMx8VSLVY4NxE5PWotV2IC0JXZL7bP/ITNKuR1jYG/qtu91+49x2qbLv9XbxyFir6RX6Zr1AejTyyuyzfKgU6l/na79AyxLzHENfMALvuW4R55nbWNT6SYMNui+np2jz07lb25jCtnMsLQ24XbHHrcwb22D9R/bv1vbVDkREAERWEcCO1784Y/Tf/vCgbTz1/alj+z5q/TLf/5s2nXs5WTfg7bUzi1/56f2Lcq9+MP/GGUnL+xLizabFDA55ybL2maNJ5ZSW5FBtkjas2fP4q54biEQ1UNerb7pbhNU69MZTGzMgu1jHZkxFjxRah+3tI9h4kVH9t1Q+7eXHeG7TWCMBRPsK6Wow7qWyts1s8Wegtu/X4xzXSxyInta8mo6gXNJFrPm8r0xxnb1HKNftPgEZXN2s/5+wXc6xyh4j4mn2iYabfSm0Cnns5w89mUpfnPXWuKXxxzTj39uKaeXr4M3dlt5xGdOJ+RjnLIU32G3scLah929vHL6Qqda/zod+wdYgjtSzBXGfv/+/RvzA/sQ4yL7G3kRayvnb7LYC/fwZDqqY3kt/aN1DoR9Ph0TS8zQj6k5m8bmtzKvtcNypujftfZ0XQREQATWmcCO//rZu+nPD30n7fzcA+nsT+1LH770wcVLxM678fFF+uFL38+361bOyo/9wyRoE1f0kyn2sWK+S52biHii9HL4SUFusWsfp7YFAb+Z0l6QwRN7aWE5tn4PR0zMPLnzhFdarPS048tiQ1xbUHI91GFd+fp2HCPmajpFnKEvFtfMGnlYhPE11LPUfIXf47RY5Y0El2s5RpstPvF9hDc9/NNnpThv0cnKROxWNUZhU2tcoDzbmRtXuOyQ4yE6DWkHddBfW2KA4ykX65DLKWLD+kluPOfyPceQXevbrTJ7+lerTCsX6bnq/aPFPvBCPMxtU0v/4DjF2NyTRrGEftIjp7Us2LXwjspMxZy59eiE2DZ7p+7fkb3KEwEREIFVILDDlPjp2z9L/+Nrz6Sdn92fzv61r6Wdn96XPvTZ+xfp4vyz+xfXrdxUf7yAz000LU9qefCO5JiM3KCOyT+qZ3n2kbLSU9Gx9XtYwk6e3KeaOEt6YOHQsmGDHNRhXXFtu9KWhZfpFnGGzvA3FhcoazF24MCBxacWchsrtI9YgwzI7kmhR6tPoCfa9unYTT10Rzvs91WNUegMv7DOuJZLYWfO17l6rflDdGqVHZVDf9Um+n06vf0rYhrlIW441la9f0R2+Dzwwpg2t00t/YM3g7n539tRO0c/8ePnFOdgV9Mhd30q5sytRyfEtrGYinfOVuWLgAiIwKoQWGyiTZm3fvqz9M2nT6aLf/9IOve/fz195HN/sUgvvu5I+p9Pv7K4PrXSNvDbU98rr7xy42UiNgjbR8fsCXFpA8u6mBx7Ksy/gWwLXNsE81NmrmPH9rTang76eviOny/vz8fW9/JK55ik1mEBhsUG61qybRnXWhZepkfEGfrxYpE/rm+bWV7EmN38xNfq83WL8Z4FCtpHCj1aN9FWzz5h4ftIbz9D+7k0Ysd2j7E516blI956eEBea1ygvKWw80zcRLM/ozhnTji2OojZOcYE+GMq2dB1SDzB5iiN9GSeq9g/Ijt8HnhB/7ltaumzvBk8EzZ1UzFnOavSv3286VwEREAEVoXAxiYaCv34Jz9NJ1//Sfr+K6fSKz/6SbJz/W0/gXVagGFTM/QO/dSLV/Ney8LLyoFzSXf72R7ceOGnd7xwsw1W9H136HE6LuzAjjczvCjDInvq3oR4GxI38EfJ37lrZ+Im2vcRY5C76YgXi/FN0jliIIq7qWNsCnmRnqveP1rs1ia6hdK8ZaaMI8SpjXur0L/nJSfpIiACIjCcwJZN9HBRqjknAUxs67BBwaYmt/mo5Q/ZDNXYY7PE/KI64FzS0V5uZJto3kBDFupHd/Hx9HoO+9D+dqZsO77zPeXiLmcb4m0IV8RFyd+5a2fqJtr8YL7m91bkGCHfWEU3lXI+7cmP4q6n/rLKRnquev9oYbPqm2jEYG86ZDxp4TVHmanjaJX69xy8JFMEREAEpiCgTfQUFLdJxtQTZ2TGmA1KJG+78mxRgLf6tn5NoKSrfTw692d+8V8jwAK69j37nMx1zVeMDvOcPcX1MTRMUr3WM888s+gbPV+hgVR7CaR9JcZuLPlNin1dwD61UXvbMmQNTdG3ajfIhsqfs97p2D/mtgk3vkr+5k8F+bhsPV+nTfRcMboK/Xsu2yRXBERABMYS0CZ6LEHVF4EGArawtI2R/kRABERABERABERABERABNabgDbR6+0/aS8CIiACIiACIiACIiACIiACIrBEAtpELxG2mhIBERABERABERABERABERABEVhvAtpEr7f/pL0IiIAIiIAIiIAIiIAIiIAIiMASCWgTvUTYakoEREAEREAEREAEREAEREAERGC9CWgTvd7+k/YiIAIiIAIiIAIiIAIiIAIiIAJLJKBN9BJhqykREAEREAEREAEREAEREAEREIH1JqBN9Hr7T9qLgAiIgAiIgAiIgAiIgAiIgAgskYA20UuEraZEQAREQAREQAREQAREQAREQATWm4A20evtP2kvAiIgAiIgAiIgAiIgAiIgAiKwRALaRC8RtpoSAREQAREQAREQAREQAREQARFYbwLaRK+3/6S9CIiACIiACIiACIiACIiACIjAEgloE71E2GpKBERABERABERABERABERABERgvQloE73e/pP2IiACIiACIiACIiACIiACIiACSySgTfQSYaspERABERABERABERABERABERCB9SagTfR6+0/ai4AIiIAIiIAIiIAIiIAIiIAILJGANtFLhK2mREAEREAEREAEREAEREAEREAE1puANtHr7T9pLwIiIAIiIAIiIAIiIAIiIAIisEQC2kQvEbaaEgEREAEREAEREAEREAEREAERWG8C2kSvt/+kvQiIgAiIgAiIgAiIgAiIgAiIwBIJaBO9RNhqSgREQAREQAREQAREQAREQAREYL0JaBO93v6T9iIgAiIgAiIgAiIgAiIgAiIgAkskoE30EmGrKREQAREQAREQAREQAREQAREQgfUmoE30evtP2ouACIiACIiACIiACIiACIiACCyRgDbRS4StpkRABERABERABERABERABERABNabgDbR6+0/aS8CIiACIiACIiACIiACIiACIrBEAtpELxG2mhIBERABERABERABERABERABEVhvAtpEr7f/pL0IiIAIiIAIiIAIiIAIiIAIiMASCWgTvUTYakoEREAEREAEREAEREAEREAERGC9CWgTvd7+k/YiIAIiIAIiIAIiIAIiIAIiIAJLJKBN9BJhqykREAEREAEREAEREAEREAEREIH1JqBN9Hr7T9qLgAiIgAiIgAiIgAiIgAiIgAgskYA20UuEraZEQAREQAREQAREQAREQAREQATWm4A20evtP2kvAiIgAiIgAiIgAiIgAiIgAiKwRALaRC8RtpoSAREQAREQAREQAREQAREQARFYbwLaRK+3/6S9CIiACIiACIiACIiACIiACIjAEgloE71E2GpKBERABERABERABERABERABERgvQloE73e/pP2IiACIiACIiACIiACIiACIiACSySgTfQSYaspERABERABERABERABERABERCB9SagTfR6+2+L9k8//XS65557NvJff/319KUvfSmdOnVqI2+qg5MnT6a33nprQ9w777yTXnvttY1zHYjAXHOX2pIAACAASURBVATeeOON5ONvirYshk2u4ngKmpIhAqtHQP179XwijURABERgHQloE13w2quvvpouueSSdPHFF29sQm2Rff3116fdu3ene++9t1B7+KUjR44s5N94441dQu6+++500UUXpSeeeGJRD/rfcsstmza7XUKDwszA2nv++ecTdJ6TS6DKrFmw6aqrrqryMyaHDh1KV1555cJ3xuGyyy5Lx44dK+poPrr11lvT7bffXm2jKChz8bnnnlvowzGcKbqS2Yhh1p/jb+o+2OPzocBW1Scter399ttpz549GzFucd7z39KXhnJddr1lxEpk03a1G+mybnktMT7GpinlW1/bu3dvuvrqq9Njjz22UGvOsc8amFL/Ho533XXXYhyJ1jy4YVqShzJ8Ux/l57RJfRGU82nEaMo4Nll33HHHop888MADC0WmlO8ti+zxZeY6nzOW59L5dJe77Ztoe3JqG1VbiEUDaM0B9qTVOhBkmBzb2N18883ppZdeqlUvXh+7gI90s43V/v37ixsmdNIeHta5sKE1o7DYzcmwScc23UO4gQsWz7aRwSRoebzhKQL+vxd5wIPMnnSuhTn8UJPveXjdS/VbB0VjZJt09tc111yzccMkx7lVvq8/xicch15u7znYckyxbrlNNOp5X+Cc5bFOrT7nOr3HQ33S2s6csYJxBRx701JfKNk3BzPIbLEhirMxscLjZa19z2xouxYXzzzzzGJjxuOItY+xJNqElPyyHddqfTvyFfSEz3P9H+WGpi3yc773OnFfG7qJZhklLrC3RX+UjVLEZhTTPo65Ppj49QrrDwZcz45r80HNJrQd6ezzvA2w1+d7Hf356dIXvV3RecSo5jO+7n3gY4TLIsajvEi3IXmRPTU5rI+3p3bOcV+L5UgPa/vw4cPhuG836Gr7kUim8j4gsG2baNtgXnvttZueYvjO8YGaW48sMGwTWApAL682+UIWOiLK8+TGnQHltmqXEt8cgFxOSxsNdFKvf9SO5UEnG8hx4wCdLXoSevDgwSI307Nkm7XJ7K2stYeFGfPK6cz50J/59BznJjBwbJEVsUb9nHyzATFibVxxxRXphRde2DCNYzySbwXhpxIzW0j4vsI22ScNcn8t8qO6Y3ySs5XtZf39Mcce+DIf1o3Lsh2o52XjnOVxvZrPeVEHWbk018ZQn7CeuePtiJWcLpxf48plo+M5mEFmzn+cH8XZGJt6Fu6+Pw1ptzYfwVabl6I5I/KJ5fXYgTaQlsbVXHuWX+vbka8gDz6P+mZP387pXpIPHXLMvE6sDxbSLWMf2rG0t3yL/izfHyM24WNOc8xMBpj4WI8Y+DZrNtZsQtusa+7Y6wd7S7Z5fde9L4JnjpHlI17N9ohRzWd83bfjfcBl0fejPPihRX9uEzJRP7IH13Ip68OyW46ZJXT3Y0Wu3YcffnhjXV5ry74Gug43UXO2blf+tmyiedCyDcBtt93W/SQaMmzSNxn8HUZ7ympP7fzmojb5IsgQtCjPAcudwXcuOBGBbvJscMXG1q6/+OKLGxui3EYandQPFpDvU55orE3TCzLs3DZ3+E409MdiiTvNiRMnqrpx2+ADXpDdM6GYPNQzXVtttnqwMdcerpvc2n/ULurn5LPeLWVKOnCMMWNuw3yGj+pbPt8MqcViTj63NfQYcWD25ThANveNHA+2BbJZf2bCZdHGmLTmc9/XcjZYPuvMOoFB7jqX7TlmLqsWKzWuNTvnYlZql3lGcTbWplLbHGcYX1G+t12Ut5i0ucDGEB73zU57Qs036iJ70T6nmIdL/SB3rTZWcDtTHZfiiJnndEZ+TveS/F4bWB/EQC0mfRu95afU3+tSOkcc+Xk4YuDl1Gwca1NJB/StXDx4XVHe4mhd+yJ4oi9EKeLV7IfNzKjmM8+tdB7JivIgo0V/toltydkD2S2p6Tb0fS7QvWXtwA+6crFma377ygjs9f2vxZ4zvcy2baLt42PYXOYG0Jxz0Clzm9BcvVJ+NFAOWcBz5+VBg9vmtqKghX3RNZaDY27TOoO1yxss5mRl/+zP/mzTIgpyLK3pxmXNb2wjOrgfdLhOdMz6t9pscsCJdYjk5/LgX2MWLRpr8rl+yWZwwUAVpblBETqwD9keXM/VR9u56yxryDHfVeebNb2yOO6YJRiz/hwvkd962+by4JmLKdYz13aNee0669NzDN3njBVmH8VxLS/HtWbnXMxK7bKtka/Be6hNpbZL9va0i/5jfrGxlTfPUfuQbeW5H0Zlh+ahjTm41XQqca3Vtes13cfKZx14rIEvajHJ9e04kuHL8PkQ/VmnWv+365Hfc2vAFv25/aifDrGplUktHljO6dIXe3lGjGo+Y26140hWlFeTw9fhq2gujezhurVj1I/WurV4b2WPctbfoj7hdbR1HPpuS3lf/0w+35ZNtAeeG0B9OTuvBVlUpyUv6jTI61nAo44FJCa+qH10JJaNcrgWdTKU8al1Gvs4tXV6a9c44QVo/om8r+vPa/6wjymbbNMdN0LwsZEhHZAHvB6bwSmalL1N0TkGmmigtPI1+bX6aBMxEbUDGVEcMJccV+4PUZmSfOg3JDXd+E7nUB+g7Rwj5DOfHBfELSaDnhR9tebzGm+zB8xr7bNN4DA0zTFheTXdoXdJL26nZl90fWictOjGtk5xzLZGfasWK0N1qPkJ7Xq+EVv0iehapB/b3FonklPKg/698hED3u7oPBprTSfIKMX4GN1z8mFzpKvPg+4cBxif2D9RTHrdIxm+DJ/n9Ocy/ph18rZE59Ecjzj111j/SJbPi5gMsQk2cvuR7Jxfo9iGjdE1tMcpc22tw/VbjqF/j/xenlEbbJvnivLet3zO+kayorwWHihTshH6sQ6o15Kivo91q8vxhj7PMkt6oRzbHrWBcj5FfA4dG728M+V87TbRpQAc47QoOHsW8Ggb+tUCEbJtYPCdBTJ6OgDanyJFZ8q1b5306NGjm55q2McBsaHu1WFopwenoYMZ7Mz5qiYfMYNFT85u+DoqBxmRDqV63BbsiPxVks8yWo/NV/4FZ2ZXz/coo7agp/clGDAfjheegMGBJ9vWY/TBms95kuO22SbYUmubbeL6Q47BKYoxlgdGQ2Mlx57bmOMYTKdkBpk1P9n1yNeIFV/fx3AvD/goZ2truy2xGukGLrVYiuq25EH/Xk7Qy/OOznPsICN3vaZ/TfecfNSLdPV54M7+w/jU2/9axwXYndMf14emsB+2eTmIeT8uMQPPKTqP+ukYm6BXLl5gl9fFxzbbEenoeeAcuue4odzQFPp7fUvyoFOOia8btVGKY5T3TPmc4ySSFeV5vUrn8Du3g/LQr4cZ6lpaqs9xgj7PdVvY12SwPD7uHSu47pl8vFabaO4YUYCNcSQCmzsNgooHC9YhGgwhh+tEekG2DQzeFshgXSIZc+QN7YBjdGGmPFC2Hg8ZzNjOHGf4ISe/5EPmgYGvZE8UL6gXXWP50DMq1yqD5UXH9t0Zews+22Af3/7KV76ykWcT/dA3PeYmLTBm2zheoj4Y6d+aB5Y5n3Pc5NquMa9db9WVy7XKhH3ME3JaZMzJHnpEaYtuUb1SHmRyTOeOI1+Dpa+Ti52SLrjGMv284MvU2kHfMf1ysiCTU9RrWbij33oGLec1/Vmn1mPokxvT4fMo/lvagH9yuo+VzzrwWAP/9fY/6NPiS2sb5YfyYf1xDJkWE1E/snI5v0UMIBdpjQna77UJvi71H5TJxQN0RJ8qyUJZTlGvxX9g2NL3fJma/qxTL8+IUc1n3B6Oc3Wi/CgPcmopx1wUr5E9NZl8vVSf20af57ot7BEzc8Ya63SmH2/aRP/o395MDx77l/SFPziazrv0wCK18x+98easnND5cxMfGkdwYDCMfqYJP9WBOi1prsP59kxWrizaQQeBjsj3KWRHgQ4ZNR5e5hTn8EXPoDq2XWbqB/eW8yG6YjAqTU7wQ04+691SpmRLFC+19sEdtkQyStdQP0rtxRdPPvnkppdOQH+8pAL1/EuJrBx+yo3fVo7yPi1NHOgnbBtzjyY5yLdy/qk59Mp9N7TGnHXNtV1jXrsO/XvSmt6QVWq7dA31mT3ioTfNcUMbUQrdettC+ahNyOTYitrO5bUyz9XnfHxFBvpGCyiUb20XfcdkluRBLlLUK42NKIv5Anr3pLkxE7J7U+6bOXvn9vlY+WxzZA/3vyimub4dI1Z8jLOcyGe+vJfbes7vzLD1TG7cRRz5NU/EwLfNtkRMen2ivugJbz7v5YkY5P5e89nmFt8/Qyz4cSmSFeVFMqM82OfbQdnIHlxrSUv1YSPGbLPD1mJPPfXU4pOe0K3UP72MFp2sTM+43yrzTCi32ES/++576dF/PJE+8dsPpXN2709nf/K+dPYnv7ZI7dzy7bqVm+MvN4D6thBA1hntLdf4SaVoEuj5HnAueJAfybe8aMDmOrmJ3OxCR4o6A675CcXzmPLcOiv8EOk0ZVvbLYsH2BJj+MH7n/mwv21zyZtGfts5TyBsP2KaZeI62s/VRTnIiAZ9XIvko75P0S7bbbLx2+vG784770y33nrrxtvCTYZ/06PVb2m3pCPzZX1wHPVB06VWL2Jl9WB7jjlPULm2S/ZYG7XrVqb3r6Y35KHtyH5cK/mM+w580JvmuEHHKIVuvW2hfNQmZJbsjXRBXitzlI9S48k3eswvta9GtLbbEquRTuASxYgvjzmjNI76OnOdg0vJn7AtKsO8EDe5NDc+lOTDbtuk2ad6eP3CL1pFOdYHawnuf1FMo66lXNb7kq9FNkZ8WHbt2OTzOzOsDZs/VnkTbTqrL9Y82z9/oV9GcWZ5tTiGRpDj+x7HMmRFeZBTSrlebkzL6VGSi3Ehx6Al38YAyCn1T7bBsyrpiLG8JLtU/0y9tuO991L6h3/+13TebzyQdn76/vShz+1PH/mtv0znfvGhRWrnlm/XrZyVn/oPzssFLdpD8H7pS19Ke/bsWbxECz/5Y2UwOSEg0aFQP5fm2q8txCP5LQHMk2NkM+yMruVsGJsPBn6yHSt3FeuDr8UJFieRnlwOMWWpH2RqcVJ6a3VpUET7tYEQMiLf4ZrXObIXeRbDN9xwQ7rpppvSQw89tOnn46wMx2+uDxw+fHjxFLvE12Rxf4nivcY2ap/rGHv+rj4/NY+YgDn7m31es91sqjGvXYcfelLovexY6dFx1cqO9UMr88ju6FNUPlajepbX0y7G9VpcoC3ujy11ID/qu5C5jDTql5yX68+sW6m8r59jU4spHpu8TDvn8bKmTzT2sT2+rZzOXKemP5fNHfPTZ7PJ5hHcMLA5KvrKTy6OmAGz4bY5ZiMmNZvUF5lm/bjG00vAeBXFu+VFPvMyvI85LlguZPnyXl7uHLZFaynUydkTrSdQB3JZ19rx1VdfnezfHlbYOsxuQEFOqS1rE+Va+bJNYAjdlZYJ7Pj3H/9n+vx1R9POz9yfzrn46+m8P/r79PEHv5t2Hf3BIrVzy7frn7/uSLLyU//lBlDfDju6FESQVyoD2TzR+EEa11hOS+fkALaJixfxc/xONGwZmoJraeAYKnvV6sGnGMDYt15XcGlZfFhc2F3sK6+8cuM7wrWPDlt7iJVIj9b2SzJK17y9rec8gY0dcKGf+SOKP/iL+dT6YK3/l/QHc8QHUrRfqgt+sAl1kI+0dh3lelLoXYvVUtu5a+AJFlOmNX17GPSWzdnLcmyBbR+ne/bZZ5PdGLJ3AFgft7milTnLs9j1PC3ua0+fWUZPu+g/5jPb6OaeBEI+Pz308yHKcApbtnsTDT24z3Ff9THL5WAPlx86rpViiuXzusBiAtx5DOTyXn87r+kIJtbWbbfdtpiXanVK+oNTlFo/4ae4ph/fFPIPOMxO+7Qg4hG6+jiqMfBcIvtyNqkvRp6s5+V45mpG41VtDveyfJu5uID/e+VbeyzTxyHrA3t87EVjCteb4thzKMnEmIK+aA8c0d+snjGyhwr6negSxfq1HY8fP5l2fmZ/2rl7X/qFP34i7frbE4v/84/84P3jh08s8u26lfvmP71Sl9pZIjeAejEcvOgsvoyd88KhtBDgjhZ1GsjhzsF1Sjr4u7G+w/Fk6W2AnZFOvuzYcx44SqzGtuPrw+eeyxTnuYU5+8744+54jjP8kJPnbZr6vHXAhJ4cp9ClVQbKt6QcM6U+UJPF/oDfPevePsgyS7qBmW8vlw9bWmwHc9iUSyN/oZ3eFG3WZMK+qFxOxnb01V77h5SHvTn/lPKHbqJNT8S0bTL4k1RDbGipA5+bPXZjzzbs0WLq2muv3bgBWOo73CZiIzeGctm5jtm+KK65Xfg8KtfSt1lWdFySX7rG4xZYsj6Ym7lcyUeIMfO51cV5ad1h9pR0jOy1zTHHDcdYrTzstHK5OGIGpf6IaxGTkk3gor4YeSvOK/GMaqB/8lzbGscmj2OAZaCtSFaUh/JRyuWjsYHrRPbw9VU6xk/Pon+U0nvuuWfTvLBKdqyyLjv+8E++tfj+889dfuj9J9DHXk7n/81LG/+7Hn45/cqD303nXn5oUe76r35rcntyA6hvCMFbmwi4Q0SDKuSi3Zw8DLDcqVplWxu4+4rNmgVwy9NJ2MmTDHSeOo1snLqNSB7Ylzr10GvRQGs6cJu2sMBkYO1EcQI/5ORFdk2ZB9/k4hNtwa4oXmAjxzDq+RRyhnIv1YsYgq/p9r3vfW/xFQ3vCzBg/Ut9kCdcLDy9nXaOtr1euXzIYPlRzFg5MC/xsGtsE+QPTcFpWbEyVM9VqtfiJ3ykzj6Sum/fvnT8+PGNhUYtVlbJ1tpNXcSqxU/PU/GxY0auD7Wyi3zo+zTLQvmo77X0bZYVHZfkI16icdpk4Tr0Z30wlpXGPujDZSDLrsFXke2oW9IfZXyKOvadZ34niD2dthcicZ9BXbvGN3KgW44N6g1JoV/J7iFyh9ZZ977Yy9PHtXHjGK2NAYgNjFG+fCQrysv5y8pyG+hrufKRPbmypXxwhF09aW2e53bNPny9jvci1p7Nb9HXK7i+jssEdvzq5QfTWRfcl37+uofTrmMn0vlHPthAYzN9wbGXF9fPuuDedOGVh8oSB1xFANcG0Nbg5Q6UkwlZFki+U8IELEx58GXZuXqoPzSFbjndh8qN6qEj82QbldvOPPAYoyNkeH/n8s1eXGttFyx7BkOUjQZFjrVcLPBCK4pH6MQxnPMl+iF0mjL1DKGXtYFJC7w5r7cP1njAdrTl9crlo16LfNiWY167jrZ60mXHSo9uQ8qC0ZQxCFlRPxmiYy1WajIRS1HfR1/0uubya23ZdYsRfHzPL6bwqxa8sWmRCX3Atjf19rW0iTK2GUF71o/5ZaM2Xka2IK6ivgl/mMyhepXkI15yYzmuY0xifTBGcj/P6Qg5Pq5YHtoAS6Ql/VGmNe2RhTjKsWltMyrXogfYeGYmD7p53rn8SAeft859sYUn24t45JhriWOTgbrml0cffXTj04PoD1YmkhXlsU445nKt/R46sT2Q15OCI8awnjSKU9+22WZfsfAvgPXlbI1lZW6//fZwzPTldb6ZwI5f/M0HF5voj+09li545JWNJ9DYQFt6wSMn08f2/l2yTfTHLz+4WcIEZxiMagMogq4WvNwxIpnoBBa0pbd49y7gJ0CxEAH9It2nagNyIhtxbVVS8Kj5Pacvfzck8jfizw+ive0iPnsGQ5TNDYrQoXY9WhQaD+iUu55jNmc+Ys5s9zEOX8BelGX9uX/7hY3pDRm5eMGCyfvb6oL3kLpgVmNeuw45vSl0BztfH9eZJZdp0YvZIXZ70pxurIcdQ5ce2a1lo5jx7becWxza96Vfe+21luJbyiC2IyaIYa9rLn+L8G3OgG3mE17wTqUWYtnkc1/luOF8tIvrUR/g2PbcUb+WluSXrvGYhjGR9QFDLhfpyDcWouvsl14+Ndv99ZK9viziGrb762POW/QAF/XFOukcT4yH9ukD+/UOewHw888/H86ptTg2LbiPI/6RZ35CXiQryvOW+a8iRP3F12G9ov4TlZ8yrxSnvp0WBlYn508vT+cxgR1fvOGRdNaF96WP/t7h9Im//n7adfTEpo20nVv+R3/3cDrrwnvT79z0WCxpRG7rAIoAiiZAbp4nH+4YFlS1DRXLidprDUyW449t4YV/+1kgG3TsN3ntTpC9hAwDxRwTitfF7LH27T+6c+/Lz3VuL+4xHfglbGgLPHoHLfaVLbaiDbS1EZUzFkPbhd4tKWIsmry9blYG36E0nQ8ePLjxJIbjnNsdM0DipUpTxgXs9Ytf6Mx91+y1j9DaUzPu8+yvyG5ug19uY220vp07F2usn9lQ+medYZ+lY3zCcvwxc5krVnrsj9jk4tzb0nrONkex0CoHc1Ckcy0v5+da24jTiAn08Tbl8mttLfs6bDN2WOxOoYP5234eCj6J+inGbSvj+3+p73Fse+6sO8ZFm6/sSY+9bO7SSy9dbBha5ZvemOvMJqxLOBZYHzAsxTvbXVo7mI74JEIPH2bQclxi4esjrkt6+zqt5y16IF6ZP+RDNx8TuXzUW5UUtk3VF8ETfTCXgiXikvtqKY6NG+qYbObO9eyaffWE81A2yoM/7Bq/BM/07PkKC3RjeyB77hS+BNtSeyUGXA/+HDqPsawz8XjHAw9/L+389F8s3sD9S3/6v95/Go2PdB+xp9CvJMs/59e/vih34Oh3J+eEwag2gHJQYFKJlEFQcKANueuEgOXgYh3QYU0HnvByg0otH/qik9Z4RLavY16OKWwBj55By3/viH0FuZyyDsZ9VTbRpqPFln95C8dS7uaA1UVf4Bhmu3PHzKPGLifD56M/me4lfayv2k/YWbuow+VbdDO7sVBkVjhGX/M61mKtp5+zztzOUJ+wjNzxdsRKThfOhx9z3Llsz3FLLLTIwxyE+OhJc36utQsmpba4701la02vKa6zbaW5uqetnjGdn8jaRvHUqVOLpkp9r6dve58hrkvyTQHm4mXYOS/mWR8wjGLA8rAJNxkt8ySPjxxjNf17/BXJwg0Ie4hgNyHsu5p2MwH9r7bmiZjUdDI97Luf9pUFxIGvU/OLcWVOkR+8zFU5Z9sQR2N0g199/Nr7foyzPQx6/PHHNz6hE82pOX42919//fUbN8mYOXTmutbvvv3tb2/UQXkugzyrbyxsbQHd/U0ktFFKI3tK5XPXIAe69KQYb3KyLT/HwNeBP4fOY17emXa+48Uf/jj9ty8cSDt/bV/6yJ6/Sr/858+mXcdeTvY9aEvt3PJ3fmrfotyLP/yPyRm1DqDWMAIv53AOHD+ZWF0LPp6oSsZg8OG2WD53Th7cS50BAw0Gdfsugv3boIMXcMDG2oRS0n2druWYwgbw8P7E9Si1gcE2UTZI2keKWv5MD34i39suypf8n7tWGxRNN757anLwPcaSbUMHyJpPSm3mrll/spsB1p/wFCZXFvk9fRB1kNqCzRaXvJmuvdgPPuyJNbTXmg71Sav8ZcdKi17wYy3OW2RxmTnilOWXjsf6EUxyY4Ll8xzD5eeMz5LNrddY1ykW7uiXxsRiCJ/IKelj/rF5lsf/ks9a5nBrm+du+/1We5kWPq1Tkg9dbaNgT9N5XLKx3I+JrA8Y5uIdNw3ship0QXu51PT46le/uulyi/6owPqVYrh0DeNB6xqQ2wQT6DMm5XjN6au+OIww+i6PWVEcWx5uBllclNbpVtY+Mm59OycLm3H2m1lg+pj8oS/UiuwZQgZycvFWyke/KbUbcYnK9/T5qP6Znrfjv372bvrzQ99JOz/3QDr7U/vShy99cPESsfNufHyRfvjS9/PtupWz8lP/tQ6g1i4PorZB4snUJgU8sWsJspodGFhbNtE1WT3X0bmWsYm2Qcgm8yl49djIZWud3a4P+e6hbaJaFxSsD47hBx78cS1KUb40+OWuzcV/6ABZ80lkf0ueye3xybL7IHzY6vMWm32Zkk+MDz6uyk/QvIw5zkt6oT0ef3OxXMqfOs7nilPYW0pbeJXqR7GN8pgTeQGIPPDla6iH+EWZKVNrb275sCNK7edaejaJkYyxPotkct6U8rmvYcNYineb78b+9ejP+rXEGT9AwIuM7OGBzQeI7dqah9sEk7E2W331xc1fTYrGlqGcMWbwnFqKY+vnfOOr1m4kK8qryWm9HtnTWpfLTSXHZEJWSz/sLTNlLLD9p8vxDjPkp2//LP2Prz2Tdn52fzr7176Wdn56X/rQZ+9fpIvzz+5fXLdyc/y1DqBoGwNeLhhsoTbFAIt2TtdNNA80xpIHObBeRsp6rFKHxcDUyqW3/DLY9iyKWJ9V8cmy++AyfFjyCa5hbFtmf0DbPN5xTNgxL2KhY0+qTfQHRKPYxlXMifA/yhq/AwcOLD6OGLFE/Pb4pLWs6TK3fNg/V9oS42PanlI+9zWsZ+Yel6fUv4cj4n0dNtHqiz2e/WCDx+uoKeM4khXl9WmdL40xkO3Jl85fmUqOtQBZrWN5TznMQXlLzuwri020IXjrpz9L33z6ZLr494+kc//719NHPvcXi/Ti646k//n0K4vrc6FqHUC5fesk/qOatqiwO9VD35bK8u0YgyUvKufsnGgfHaI2oaD80JRtsU41dlCYQo9V6rDwQyuX3vJDefXUG7oo4tjYTp8suw8uw4clnzB365PLZF/SCzEXLexxbTtS5rVMVmZrC68Skyi2UR5zotlkn7DC9/hsTmCbbW7yHwOGDKVbCYz12VaJm3OmlB/1Nfb9HPE+pf6byZTPEO+1NU/EpCy57ar6YhunIaWiOXXKOI5kRXlDdI/qRPZE5Wp5U8mptaPr8xLY2ESjmR//5Kfp5Os/Sd9/5VR65Uc/SXZ+pv5FA+ucnROc0blqEwrKj0lX7ePcPXfIuGzrRreHFfzQKhvlWa+e4zn8PXRRxHHeYwOXnWKRt+w+CB+2+rwnu6ONZwAAIABJREFUnlC2xScogydQqDtnijb5pqFvb65FrG+n9ZzjdIp4a23XyrXwKslDbHOf8cc33XTTxndnOSbZD3bzuPT9wZIOZ9q1sT6r8ZpSPvsY48Dc8T6l/jVWfH3IJtr3lZbz3ByrvsjemPY4mlOnjONIVpQ3lVWRPUNkQ05L3EZlcrE8RBfVGU5gyyZ6uKjTryYGVl5Uztk5QRCd60zpJMw0Gixa8niBCY5jU/ihVTbKt+gblZnD30MXRVP4ZIpNzbL7IHzY6vMhMVbzCZ48zhEPJX1relldXthHMdySNyVbjtMp4q3Ex19r4eXr8Dliu8TMXmJl76yImKG+nkYz1fLxWJ+VpY+/scLyua9pE/0+GWZS6je5a7kxFX0pV8/y1Rc5OtuPozl1ynE7khXltWtcLhnZU64RX4WcUsyVruViOW5NuXMR0Ca6k+ycnROqoHOpk4DI9qTwQ7SA3R6N+lude9HYr9H4Gsvog+O1zEso+QSLubEvUMq3nr9S0gu1xi5ibVEwZX+yWLC31tuLivglk9B37nTICw97dSq9MMrsn+rrS716rWP5lhgfY9fc8uce++bWP8e29Ul0rv6y8tUXpyE9dxzPKf90WBdO40VJMQLaRCsOREAERGBFCNjkbxsz/YmACIiACIiACIiACKwuAW2iV9c30kwEREAEREAEREAEREAEREAERGDFCGgTvWIOkToiIAIiIAIiIAIiIAIiIAIiIAKrS0Cb6NX1jTQTAREQAREQAREQAREQAREQARFYMQLaRK+YQ6SOCIiACIiACIiACIiACIiACIjA6hLQJnp1fSPNREAEREAEREAEREAEREAEREAEVoyANtEr5hCpIwIiIAIiIAIiIAIiIAIiIAIisLoEtIleXd9IMxEQAREQAREQAREQAREQAREQgRUjoE30ijlE6oiACIiACIiACIiACIiACIiACKwuAW2iV9c30kwEREAEREAEREAEREAEREAERGDFCGgTvWIOkToiIAIiIAIiIAIiIAIiIAIiIAKrS0Cb6NX1jTQTAREQAREQAREQAREQAREQARFYMQLaRK+YQ6SOCIiACIiACIiACIiACIiACIjA6hLQJnp1fSPNREAEREAEREAEREAEREAEREAEVoyANtEr5hCpIwIiIAIiIAIiIAIiIAIiIAIisLoEtIleXd9IMxEQAREQAREQAREQAREQAREQgRUjoE30ijlE6oiACIiACIiACIiACIiACIiACKwuAW2iV9c30kwEREAEREAEREAEREAEREAERGDFCGgTvWIOkToiIAIiIAIiIAIiIAIiIAIiIAKrS0Cb6NX1jTQTAREQAREQAREQAREQAREQARFYMQLaRK+YQ6SOCIiACIiACIiACIiACIiACIjA6hLQJnp1fSPNREAEREAEREAEREAEREAEREAEVoyANtEr5hCpIwIiIAIiIAIiIAIiIAIiIAIisLoEtIleXd9IMxEQAREQAREQAREQAREQAREQgRUjoE30ijlE6oiACIiACIiACIiACIiACIiACKwuAW2iV9c30kwEREAEREAEREAEREAEREAERGDFCGgTvWIOkToiIAIiIAIiIAIiIAIiIAIiIAKrS0Cb6NX1jTQTAREQAREQAREQAREQAREQARFYMQLaRK+YQ6SOCIiACIiACIiACIiACIiACIjA6hLQJnp1fSPNREAEREAEREAEREAEREAEREAEVoyANtEr5hCpIwIiIAIiIAIiIAIiIAIiIAIisLoEtIleXd9IMxEQAREQAREQAREQAREQAREQgRUjoE30ijlE6oiACIiACIiACIiACIiACIiACKwuAW2iV9c30kwEREAEREAEREAEREAEREAERGDFCGgTvWIOkToiIAIiIAIiIAIiIAIiIAIiIAKrS0Cb6NX1jTQTAREQAREQAREQAREQAREQARFYMQLaRK+YQ6SOCIiACIiACIiACIiACIiACIjA6hLQJnp1fSPNREAEREAEREAEREAEREAEREAEVoyANtEr5hCpIwIiIAIiIAIiIAIiIAIiIAIisLoEtIleXd9IMxEQAREQAREQAREQAREQAREQgRUjoE30ijlE6ojAOhA4efJkeu2119ZB1aXo+M477yRjMtXfG2+8Ib4pLZgqzj6IKsXZByx0JAIiIAIisD4EMH+dTnP6Wm6i77rrrrR79+507733rk/0rLmmzz333IL5xRdfnE6dOjW7NUeOHFm0d9VVV6W33npr9vZOlwaGcjtx4kR66qmn0vHjx6solh0LVYUmKDCUmzX96quvpksuuSTdeOONE2jyvgjos6pjHPTr7Z+Ks+Hj2irH2TPPPJPuuOOOdOWVVy7GbZuf8X/ZZZela665Jj300EPbMpa3xqot8K6//vqF3lP25d5BAX6+6KKL0vPPP7+ozrqtypjQMg+8/fbbac+ePRuxgJhoTXvHl17WvnxrrPh6redzy2/Vo7ecxeTevXsX/XgZ679e/aYqb/3szjvvTLfeeutG3+uVbf3i6quvrrLCPibXF6K1djQ29OrH5a1/ml9N38cee2xxac6xZmz8w/4eZmbU2HaZmT/e9k3066+/ng4cOLBl8rWJ96abbkpPPPGE1zkh+FZlMtmi4MAMDt5ckJTycxN/r9xo4mqZML3Z8FNJZ1zzbQ4NerPVFnQ2MNjGBvIttYWcxdO6bMqffvrpTf3CFlU333xzeumllzzqjfOh3HrqDYkFKNgbi+w/XlRCXpQukxsWidGEB91s42gbDI7Hmi+ZEyY3yJs6XSYvxdmwTXRLnPm44BiyfhTFEZeJrnuZ/txih+Oa+2vuuHfeZh1zMjnfz4OtMcfteBne7jnPsVDk8Y516+VX09VkHzp0aJMfMVeW6rbMA4hb9k/PsV8XlPTBtbnXHXPLhx251Px19913b/KXrZePHTs229oGMVma53L6cj6PF9vZx1gnPuZ4zY2HtTIt/cLarMVRxBp+4LGB9cdxTraXGdnSOtbY3s3W2ejPptMtt9xS/ARd61gMO3wK+9GmT719qD+2XciJ0m3bRJujbGHpIUTnV1xxxaa7QgiQqSeTCNAy8zh4Iw61vNyg1Cs3ktM6MDAv+Kmmt133bQ4Jeh6gS21aZ7cJp/Wvxw7f7pBFQIu/crE/hJtxgI0t+g6JBbBusc0zxLmPEchE2iJ7Lm6RXNOnZYyL6ppNmDByEwPsHpraBHrttdcWx+Ccboqz959YIjZ9Ohe3nNwoBuAj6JZbFA6NM4wDJt/G1P3794dfabCvOdgmjZ9S1/oy29PSr2GjpZ4RONTGNm6nRz/WNTpmTqwnH3N78AcvlFk3b1/UZmtebQywRXHuD3bNMT61+izSDXMZ880dM3eT1dLu3PIjm5BnG5fSk/1cjPNmKcfC5/N4gZgc6mvT2881nj1sjNIe5t6OHJOoHebE9nPZWpk5+wX8wGMD64bjHC/vv8iWlrHG1tqeM85LurX0L9gwZTpnu9uyifYDgd31fOGFFzbdRbPvBPLdUQ5oBMiUk8mUDhsqqyV4h8qu1UPntI7ArFFvjoEh6sBorzfoUd70t5su/omzsbUn1DyQt8YP4g2DRE/aM4Cb7RwDNhjxZt/6hN2BRvuRn5gDyllam7BgY62c6ThHLMDvnPJNkRrH7eBW4uD1sQ0Gfw+IFxWlSafHL8yuduz1K8VZ1E8QZzW/eD167Cnx9XLHnK9znOXsxnhuYyHGvGi8QP0ev1gdjp/WGLA6aKcU89CpJ4W9NtZ5O1tjlW1qGQdb9UMc83jsj7k92MKMWLeoP7bqwuVYprWFT/1Z/sGDBzfmmVx7sMsvzLmNocetPhsif8p1R9T+XPJZLvvLdOAxjGMJ+nFdH3u5c+5HiMkhvkaft3bspsxtt922iK1IT+jrU5aR0zeX3zo+WZvMie1nfWplcv2C+1tO11w+xgL4Aees15DjyBbWM+r7sM90NX/ik528psnFyZB+jTo5Nrl8ZgQZPbHQynPpm2h2mh8IIqXhUA5odKjIwZGMdcmDrRYUy7YNTHPBj46Tuz6EcUlmT9BjYDFuNjCjU+d0gmwrz3GVKz8kH230dlow4QHAtw/ZkS9wzQ8spQmL+2RL3EHHqH2v65Bz6wd8s6DFp9BpWdxqfRXXeZLxLFq4I7ZLdnm5LeeIk5JclIn8jGs98d1iL+sOn0btc7mhx6dDnEW2I/ZsDLCbI/ieb2ms642zXl9CT/h06rEX8RjFCq7VYpW5lcZL2NKawuZIt0hG5AvWrWWMjuT6PHDJjQG4ntO7xS7W289JLec1n3mbWs5LesPmMe3OJR+65fyFdsf0LfRr3wZiMhcLJe62trQHZfgaGtaaU/axqH3w6vEl7C8xrJWBHzyrMX0B/oAfcB7Z3ZMX2cJ6+rGGr0X+Y3nR9SE+QZ2W8YLLMCPI6ImFVo5L30SjA5WC1CtvjuOnOJDhHezrrds5BygHQ+vxUB7o9NZOTgbK+IFhKGPubFGbCHpve9QJEA/RtUg/5txaJ5JTyoP+PfJZr4gJ2mN2fmE8pF0MzMbaD3zwu/eDnU8VC7DL7Oc76tZGiQPXw0ahVH5KbuCSY2C2tLytuyV2Ucb7Bvb3pkPizHNVnOVfarnMOIt8D99YvLCv/Vjh6/bEGdto7dRuXKIt6MYLHFwbmrIuPk5NJtqsjcXMaqq+Zu3XxgpvN8ZjZsS6RTZ6GbXzFnk1ri12cTvRHFLLq/msZqe/zvpEPm6NFS8X53PJZ7kl//f0YejMKXzOsWfXEZO5+Y5l1I7H6liTj+tDfAn7a3HZcn0KVrAFKfzg/WPXYW+LbqjP9mJ+KMUa+jzqQy9OoUdkP655HaO+yDLHHqPdqccT02upm2gEgAEcAw2dsDSYjIFuQWQLYHtbsf+YeU7ukDpeFgevD7KW8yE8uBOVAgydJ+oY3o6Wc/gwJw9B7+32OrL+PfbDntJg0GJHrgz09/rmyls+24IBLSrPceJtHtIufAHW3DY44RqnOd9FOpfyzB7+6ga3gRemlOpvBzcwGzOOmU0t/oIPpuLdysv0y9kJvdlXdlyKd8hCHcVZ/VcHwKw1zuBbxAqPFcw76k+9ccYxYP00951o+xqK/ypNqz2Rnj4PesDm3HXEHVIfq8xqSv16uWKdxHMT6+bHfG9vy3nURlSvFH8tdk2td6RjTx50ZrZcH7GEGEHqY4Xr8PFc8lv9hfZzfYF1jY4xfng+aH+oXG6rFFNcbuwxfNnqO2sP9sPvY9IaK/v4c/SLBsYeX3H1DOAH7x8rB3tbdEZ9thfzQ6nPoo0SU+hoekAm7EB9r2PreGu62VrR3ibOMswee+mu7deiP7Rb0juq15K31E00DIEDWxSMyqATliYT/OSGwcZ/7m3faMMclHsRUG4xP6QO2vNpKXh92SnOuT0LyFKAjR2cWV/EQdTJUA5lSjpZ2VKHhawoRb2WWES8cadtPa7pz7pFAxpfxzH7zfeBVm6QBb8aB3tLvtllx37wQ3lLUac2SXAdf2xvrPYvG7K27TucNhDaBMNvfrRruUFy2dwQO6ZTiZO3OTpHbJUmkVb7IvlRXo+8nH6IM2PA/7l4R8woztrfzj0kzuAvxCWPFciLYsLyeuICMh5++OFNbwnmWMgdl77iALmtqX1yBe3k7GuNVWYFmb1ppANiv3W8hN95bmLd/JjfyorLteoEdpHuLTKm1ptt6D3m+M6Nt7DX+z03rrEOc8qHXpEfWIcodvh67ThXH/m19mvy7TrGqJwPIhmo4/3Sct7iO7TJPoz6spWrlWnpF/BnTX8bK/kPfuCxga/3Hke25Pos55d8x+X8WAW7e3wCm2B7LzOrj3Z93SF6QB+kmzbRP/q3N9ODx/4lfeEPjqbzLj2wSO38R2+8ifKjUnSEsYpDjneQKec/Duqh2XkUAOx4C9Dbb789Pfroo4vfjMNbRX3gDqlTAsjyIttKdXuv8UsAmFHONy0DQ00HaxMfu7U2c4OUyUHQ5/RBW9yxSvJQHinqeZ/iOqeIN+bUelzTn9tp9X808EEOuHn9opgHAyuL66hf4jI0FmzBjb7E+llbNlnYpz+effbZxSdA8P2paDNtdW2zjTLL5gb7S4zgj1Ja8iPXY/vgJ77ee8zySuMM6+fLIU5a4ltx9oGHwI3jn/vfByU/uFnVGmeIS/YJ+7o2PnLZnjizevhZwVz/ticr/uV6bGvvsbXZ8vIrkwvmzCVqj+33/mk9jxjDL60bEPQX9jvr5vtiZEstr5VJSffSNbTPercy9OWmsNfmEbzVuhQHrVxgH9K55bfqxWN2FIvQN5ci9nys5vJzckr5WE/1jDGo42Oj5bzkb69nC79amVq/AEvTPXowUHqBLOry2OBt6DmPbOE+y30vl+/b43Lex61xXJJptvsXB3tmrLfJQrs+Xnpiw+uE88Um+t1330uP/uOJ9Inffiids3t/OvuT96WzP/m1RWrnlm/XrdzQP3aWN7BXJjqUlwPnGWRblPP3qO0aT7p+gEHg54ITi/nnn39+Q90hdTYqBwfQ3zu69Tynu2/KFjz4bU+rYyx4EojkwFY/uHrZ0bnZxR/XNfn8NuCoDoK+FuRD4wr2RLZ6fRBvfkDw5aY4h90lvVAm8gWu+ZjxurO/PWPYazJ8HzMbwS5qv8QAE4DJjRbVJV9aDB0+fHjjTcNeL9g9NzezD3w8t5Lt0TXIaeGIsmPbhB49vCKmqF/TR3F2CsgXKbjV+qcV7vE5+o73Fc8pfs7bpNj/PelpM6q/jDyLKbxxPDdGsR5gXotVrjPlcWm85DHRxwT7kv3ox74hurYyge6sC9rDtdL4xXp7+1rPx9rL652SrmZXKxcwsHRu+dYG+qWfx1kPO2beJW658SLnU8RpjZ/XJzpvtYXrDqnD9VuPwcViMzdecplSDOdY5Rizjjk/wg++XT+22Rhpn6rFOt/K88vd0BbbAntzbXPZUmyZ7Jy/hvQvkwe7o3EItnC7nsfQdll27njHe++l9A///K/pvN94IO389P3pQ5/bnz7yW3+Zzv3iQ4vUzi3frls5Kz/kr8cBNflwkHekOd8W2nhC5eVwcPjBCJBzge9l2fmQOpEc5LF+vpO0nNcCzOTzW499eb84ueeeezZeGNPS8WEHUtwd4o7MTxBRLkrB1neGqCzioaWs1WfOLXUg38dMpMvYPNbN/MM3G8ATsYBBj9ts4caf1sjFO8eJ99mQWGAdc8djxohlcDO9uZ0x8QCG5svIj54R/Jrzly9fO2c7anHmx1mTDX1K/Udx9tgWN7Rws0rsn5Y4wxjlfcVyxsYZ2sD4M2VaiiNAtK+A8Fc8fNyinE9bmft6U52jr0d9F4vDiCXPz+xH7+MherYyge6sC9rDtcgulNnOFBsJsLV57NSpzTe1vH6tXKze3PJZN/S92ljQGieYa71f4VPfH3NxOsT3rbYMsZ/rDDkGF4uZ3HjJZRBbUZpjwyyHPon27bG/WL4vZ+dsV80WHmu4LOdHnHM+7ulfLBc2+XjlMnaMdpmH5Q9t18uPznf8+4//M33+uqNp52fuT+dc/PV03h/9ffr4g99Nu47+YJHaueXb9c9fdyRZ+SF/PQ6oyQeomiMjOYDpByPk15zEMofU4frLOraBlZ8EW0eyzhu9UZUHYSuHYMTgmhsY2BaTAR+hExtX3hBy+egYbNF+VAZ56GDWlvk1sgtlLeUNIg8oXIaPYYuPGS4z5bH1FX7SAoac5mK/xM38wt/5ry0qeBPE7fXEQg+XsWPEXNzYBu4fQ+OB45W5cjv+GMx7xicvw5/PxUtxFn+Cw/iX+if7pyfOEBvRWMlyWsY6yIriDOMgj0NTHUe6Gw/T389d1qbNX7mb5cyxh7mvN9U5mLbMndYmxgf2Afuxdcwo6d8ahyXdc9e2I07YVr+5tXhp/R5+C5e55bMtOAbT2pzTGieYaznGrC3Y79tBTPr+3hrTsMPSVlvG1uH6rcfgYnbmxstamVy/YB3A2fP0573fiWbdbEzFGGlxgXUv+5zL+7btnMea1tjicj6OYHduvGdGfMwyTf+hH+fubZd1yB3vePz4ybTzM/vTzt370i/88RNp19+eWPyff+QH7x8/fGKRb9et3Df/6ZWcrGI+O4sdU6yUuYhOWJNjd66ffPLJxfea7Q42f1/Lw2T9/JO3jBqbXjDQWicna6582whx5zA9LQBrf1bPPv6BO7ctAwPLxKDb2h7XHXKMzmm24iVwvJm2Tmgfu+LNaS1+oAfizQ8IuD5Xaj7gmLXBo7Z4BAcf39AR11sXFbZgOH78OKov0t5YAD+Ow6mOIzvn4AYAPJgPiQceZyLd0Y5PwZwnQF9m6PkcvBRnsTfApeb71jhDPOXiguXkFoWs6Zxxxu30HIOZjRm18S+Si/oR82WMTWDauuHA3Mk+ZT+2zlsRC+S16gR2ke45GctgCjs4tb7A71yxeOldf8DeKFbmls+2+GMwrc05rXGSGzfQjo8xxGQUB17X2jnaqNnCcobU4fqtx+DSuj6JxtRcv/A62P7Ef+Ta2rV+b2vvaJ0OP/DYwHJLbXNsgD3bC1u4HMcB56M+t41jLsf17Xqpf6F+LoXtNd/4Gw9j283pg/wdf/gn31p8//nnLj/0/hPoYy+n8//mpY3/XQ+/nH7lwe+mcy8/tCh3/Ve/hbpdKTvLg+0SRHeyIjnmQH7algMeDZIWgPzR45YJe0gdtheDQ07PMfmw0e5AYWPJbfcclzpnj5w5y9pmgP2XY2cDUM9T8bE+iuJ0Lg52o+DWW29dvMSHbyJwe/yuAM5vPe6NhbH8cn60fMR4q+65ci3crC5PEKWJJGqH6/YuRsA8N3lG7c2Z1zIZKs62emDqOEPfyo0xHHNYJG3V6oOcVYszaPbtb3+7+gkjlPWpvdDQfqHD5nM/JoJfaYwZeg1jE5i29nksFrmvsx9zvvZ2l86jNqLy4BONdb12RfKnzoNdtgnJ/dxNqc3auDa3/JxuJT9wndY4wXo8F2N+rIDdrTHMOvnjVlu4HuoM7YutfQZcWtvxnEznIf2iFndgAT+w33DNUsiJ+itfx9jE9sKWUgxBPupz2ziGjsYQMnEN9T3fnL6oh9R0s08mrdRPXP3q5QfTWRfcl37+uofTrmMn0vlHPthAYzN9wbGXF9fPuuDedOGVh2BPV8qOaQWWawAdyncMDghzkt2FtDeC2u8925t/bQKFE3NBYE/e+GmlyYm+kM+6DamD+rDFB9UU5zkb0XZPOmRgMPnwSdTpYbv3Yy6/RV+LM1uo2icP/IYad/f8QqomF/oM9Ym3r9beql+372Zbn7In1L0sV922mn5jxjHEUdQXau1i3JpiEVNrq+W6jXmIgZbyQ8oozq5ffIooN19iTC6N8xyvfkET+aQ1zljukHFxVeI4YjB1HvzUajMWoTxOMO8p5hOWl4svzN3m36jNXrum5nomycPcUerrxqPmMzBDOY4xu2Zjrq2V/R9isjWGfX0+hy25uOOyOEadIWNNLn4he+p0SL/AuMv+NT/Yvz2xtrnW/l988cXFutb7DTZATo4trqMdxIExwvzAY4Pv94iDXPumB9qIYgXXvB9z+sKusWnpRupY2Tt+8TcfWGyiP7b3WLrgkVc2nkBjA23pBY+cTB/b+3fJNtEfv/zg4DYBMILbIxQdyjsY+S0ORhDl2rVFIr/ExJwefUyA6w+pw/W34xgfKfF3dsxee3pd+23tms6lTgd/5fzo82ttLfs6bOMBaC4dbHKL7sDBT0NvELC+c8cCt7Ws4zm4IW5rYwjbiLGvNDZxeX88pE0vY1XOFWdtnqj5HDFlY0DvPxZMXpNamyjPC63etq380DXAmHZ7+ivsnCLNLarNFlsk28/6HThwYLHeML9gXuGxgu2eal5E/HA7bC+u53yVs4tl8CJ9SJzkdOM2cse8+cAmpDVtuUE8t3y2C76oxTDzzvVxk4tyrXwRk7lYYF1rxxhjptg4QS+LrZK9NZ2mvJ7rF7B7SD+wOub72iY617bZx2MI2CMOmB+X82MNX4MMZsfyouutccwyV/14xxdveCSddeF96aO/dzh94q+/n3YdPbFpI23nlv/R3z2czrrw3vQ7Nz062CYOeO+cHqEIRpbBzuN8L7fXiRY0+EK+BVpJNtoaUgd1l5WaL/wT91LntsG25yPQsAM+jwbryI9WL5cPmauSwjYegKbWzWKp5esJ8N0QPy0rFjwbs80+NfCVr3xl03e/2Ra7OTDk92Xn5IaJKoppb6OdY8wZGidmC77rF01MUZs9eehv4N6T9iyqFGfvf9+tdRytxRnHVY/PcnE4d5xZTMKmnrjhWGYde2xu6asm225U2tdhou8jsh49x7C5Rd9lbqKZpfGBzZbPPweaW/PArpIveV3WYr8v0+K3nC/mHtfmls92gXWNR2s5+MXLw9NP3Nyxdwp94xvf2LixU/I161s6Brcp5rJlrMFKtkTX4APPCnb7GI/O7aGW/dv6B1/Psxs7sNf7DXrArybTNt09LxbDTQgeF6K+D/usDX63Dn8i19sO/TBn1W4GoXyUcvsRu1JeTq+onda8HQ88/L2089N/kc759a+nX/rT4+8/jcZHuo/YU+hXFvl23co98Lffa5UdlkMg9RhjTuWXG0EGOxjBZQARDF4BDo5eJ6LNnnq9ddiGUiDkrrXqxu1YZ7Tvitn3h/xHc20g9W9GZeaeb3TObeX0ZpnsI86PZG93HtuWi7kxOjILY5d72oynrfwSslZ2bMPcscAs7OM1/uP2ufhAfvSdRpaJ47m5MbOa33nAb/UJ7EDKE2OtPdTpSTFOgXNP2jqOMzPFWdvNWGY21O89sdNTtie+uCz6Q2vccN3eY3tHhsWyxVsLPx43hvbVSEfY7PuVfdLLFsq2SH788ccT3iEAv5vezz///ELkXLqZz0s300ufvoNdc/gyYhCxLeXZHGNse/7x6cMWm+aWz7Zx3yxao3tRAAAgAElEQVTFJsZyXgsiz8df67ltduGPFi6sd3QMfdZhE83cW8YQ2GtrZ/Rn5E2Rwg88Nni5KJPzL9/EjexrGWswtkZtlHQ7LTfRL/7wx+m/feH/TTs/tS99ZM9fpV/+82fTrmMvJ/setKV2bvl23cq9+MP/8D7rOmcH23eWcackJ8TubthTGB440Ak5j+Xmgh0ONMfzIJNrm/PRZk+93jpsQxSctbwW3biD2IBY428MfB28sZv55I5bbMr5scWeXLvLyGfbcjE3Rg+OVx74cjLNT4i5lsnO+3XuWIDebJctJO1Js006/s9uDthHHflJfItdLH8ObqYnOJcWAhwfpcWot9ufz7lY9W21nvfopDh7n2pv/7RaLXFW8lm0SMqV7/Epy63NS9H1ln6c07OWb5zx6bHSgs7L4TjlOcmXm/sc4wbrPqduJtvfLMcN25KtPfFSkhNdixhE5abOm9Mm03WMfIwFub4DZtbfeD2CelE/5Dw8/cSNndtvv33xPVxbF0B2ru0eP0Cf0tzZKg96eZtb69fK8TjHTGv1Wq7b2sa+XtCy7oI82MtjA65xavsm/+Zv69O+rci+1rHGf33VdLJ1TunmAdZmY9b2Q/rQkDrMs3S8479+9m76f/76O2nn5w6ksz+1L3340gcXLxE774ZvLtIPX/rAIt+uWzkrP/bPDMJTKID3i2h8bw7leFJDJ+Q8drzJ5MUzAooHDHYiAikXAAhcq4+OP6TOWG65+j2BCb3NFuaXk418ZtAzmKBeNPhGfkQefBXpCHtRZsrU2ptbPpjWUrDgWK3VAW9jUvPTsmPBdOc2rS/5Tz/k7MMgaHZFMcH15uZmbUGfKK7tOvsBYwbr2HMMe0pybPzDzQa7Odlzo6tHF5St2Y9ylrLPa77jesywFstcz7d5OseZt9ufM/saw5Y4g3yWO2T8zfUbyB+a8i802LjpF40lubyG6InTkswh1xD3vFAu6bbsvg+bWsaAsXHCDNDunGmLTWPaHyOfWfoHUD7uW+fVVlsQk1P025ZxZlXWYMy8Nn62srRypf7cI2ds2ci+OXWDX3vWtN7GIX1oSB3fbu58h114+/97J92675m0c/f+dPavfS3t/PS+9KHP3r9IF+e79y+uW7mp/mxjW/oYEU/M/mOc6IR+ojNQ2HRzfRzbJjlyIgeSlbWBGx8Bwsd7kI+PVw2pMxU7LyeyyZfBOevt+aFMlGIQNQ49gwnqRYOv9yPKGn970YpnDr1gr12f+t+YzC0fdtRS8OkZcMCwxU/LjgWzt0c/5tOj69zcTK/aRNMbQ7nFInjlroMRJgn0h56+DRk9KdqL+rWX0+M7rgvbW2KZ69nx0Lo9uq5CnHm7/TnbUxq3wasWZ5DfKhfl50ytL/ImAn0gd1M8p0utT+fqTZ0f+aKkG/oi7J6778NetFsaAzhOoF9P2hqP0Gls2mLTmDbGykds5BjOdQMV7ZZ83coF42bppnDv/JnjEeX39A+O39L42Wo7ypX6M8osI43sm1M3+LVnTes5DOlDQ+r4dnPni020XXzrp/+V/v74D9PnrzuSfu7Xv57O/dz9i9TOHz9+cnE9J2RMvn/ijKC3j5Ts27cvvJOMThh1Bv8RA5PHH03K/U6n6VHa1Ps7f2bzkDpjWOXq9gQmdxAbEFvu1FsdMO8dREuDL2SaH81ve/bsWWyKbXAdomeOz7rmo+NbDPMnK3L29PppCOPeNryuPGgPfUJYm8zm5gab0E7UJ9AnMZ7V0txiEX2ktOAwfdiX1lY0NkLvKdKS7V4+67aMMcfaP1PizLP258b+zjvvXNwUxg1gX8bOW+MMdZlvrT+iztQp1g7ct2yu53dD2DWbu/HSrJIOHKdz95+SHpgzeUwo6cbXltH3oXvLGLAKcQJ9W9IWm1rk5MpMId++5mRfV+AHRvha1NRPoGEHYjKa61CmNe0da1rlzlFurvjlPrudY01k35y6YV10Wm6iEYD/8X9+mn74+k/S/z55Kv3w9f+T7PxM+rMAssmZfwqhNjANqTMl097AxIBoE65N1HixmNcJLxbjRUlvh+e2eLHDx/YzWpgQuHNxBzc9WzaS3oZ1PufBzHjhZpCPxzleLOa5TRELkIl4NZsw+fuvc1jZ6DvRHB+Q59NlcEObWBD09gvUL6XoOz0LFyzS5t7YoJ1W3WCL+XzuMQdMFWcgUU7hm1ZfmjQem+eONWhv/dpuguNrCzyH2BzGN4StXHRTHGMoZHLqxw2W33o8xTgAf7RuomED+uSy/IH2SnGzHXECHkPSFpuGyEWdueWjnalTxGTJ11O3uQryOH5bxwAul7v5PcVYk5Pdw43tw7jBuk0xnrE+mJNb1nFcj4/Rh5hz6/Ec8bvxJJqV1PF6ERgSmDYoRouMXDAO3cRi8M3JtXz71IFtoqOOhfoW/LxIWi8PDdfWBrRo0ZjjOcRPy4oFphB9/DJnE/Lt45n+BgLL5ONlcLP2MAlNPTjzRIbJje2LjvFpjikm10g+52Ei67Fbcfb+DYQhNwNXKc4sDqAP+uaQtDdOccMKbdlYV/vItvUJP37m5hLuc2ijN51i0Yk5r2cTvcy+j3GgZQyYIk6idQF0mDptsWlMm3PLH6NbqS5isme8L8lbl2tj4zc3xk0x1uRk97Bl+7DOYN2mGM9YnyF7Fa5vx+hDvWOzlZ8jfrWJ9h5aw/PcR9RbTMFH4mwj64PSnhLaU+KWj8K1tFUqY08dc3/WqUtv/MvVO53y8bQ55yc8YWndZEZstiMW8GSpZNdDDz3UvHn2di2DGxYYU0xq0B+TTeskBh16bjSgrSEpJrIhk5LibAjxD77nvZ1xBs158eXnjdbzXjssxi+99NLsJ6egW5TaHGJjjb3j5IEHHoiKrHReaWG77L4PUC1jwBRxssxNtLG09+HYm6nHzKVg5NO55fv2pjpHjA0Z76fSQXKWQ6A01ozVAOuaZfbpsTrX6msTXSOk6yIgAiJQIWATT/SR9Eq17GXb/PfcOJq6/axiurCtBKb2c2+cbavxajwkMHVMhI0oUwREQAREYAsBbaK3IFGGCIiACIiACIiACIiACIiACIiACMQEtImOuShXBERABERABERABERABERABERABLYQ0CZ6CxJliIAIiIAIiIAIiIAIiIAIiIAIiEBMQJvomItyRUAEREAEREAEREAEREAEREAERGALAW2ityBRhgiIgAiIgAiIgAiIgAiIgAiIgAjEBLSJjrkoVwREQAREQAREQAREQAREQAREQAS2ENAmegsSZYiACIiACIiACIiACIiACIiACIhATECb6JiLckVABERABERABERABERABERABERgCwFtorcgUYYIiIAIiIAIiIAIiIAIiIAIiIAIxAS0iY65KFcEREAEREAEREAEREAEREAEREAEthDQJnoLEmWIgAiIgAiIgAiIgAiIgAiIgAiIQExAm+iYi3JFQAREQAREQAREQAREQAREQAREYAsBbaK3IFGGCIiACIiACIiACIiACIiACIiACMQEtImOuShXBERABERABERABERABERABERABLYQ0CZ6CxJliIAIiIAIiIAIiIAIiIAIiIAIiEBMQJvomItyRUAEREAEREAEREAEREAEREAERGALAW2ityBRhgiIgAiIgAiIgAiIgAiIgAiIgAjEBLSJjrkoVwREQAREQAREQAREQAREQAREQAS2ENAmegsSZYiACIiACIiACIiACIiACIiACIhATECb6JiLckVABERABERABERABERABERABERgCwFtorcgUYYIiIAIiIAIiIAIiIAIiIAIiIAIxAS0iY65KFcEREAEREAEREAEREAEREAEREAEthDQJnoLEmWIgAiIgAiIgAiIgAiIgAiIgAiIQExAm+iYi3JFQAREQAREQAREQAREQAREQAREYAsBbaK3IFGGCIiACIiACIiACIiACIiACIiACMQEtImOuShXBERABERABERABERABERABERABLYQ0CZ6CxJliIAIiIAIiIAIiIAIiIAIiIAIiEBMQJvomItyRUAEREAEREAEREAEREAEREAERGALAW2ityBRhgiIgAiIgAiIgAiIgAiIgAiIgAjEBLSJjrkoVwREQAREQAREQAREQAREQAREQAS2ENAmegsSZYiACIiACIiACIiACIiACIiACIhATECb6JiLckVABERABERABERABERABERABERgCwFtorcgUYYIiIAIiIAIiIAIiIAIiIAIiIAIxAS0iY65KFcEREAEREAEREAEREAEREAEREAEthDQJnoLEmWIgAiIgAiIgAiIgAiIgAiIgAiIQExAm+iYi3JFQAREQAREQAREQAREQAREQAREYAsBbaK3IFGGCIiACIiACIiACIiACIiACIiACMQEtImOuShXBERABERABERABERABERABERABLYQ0CZ6CxJliIAIiIAIiIAIiIAIiIAIiIAIiEBMQJvomItyRUAEREAEREAEREAEREAEREAERGALAW2ityBZ/Yy333477dmzJ+3evTs99thjq6+wNBQBERABERABERABERABERCB04TAtm+iX3/99XTgwIF05ZVXLjaFtjG0/8suuyzddNNN6YknntiC+siRI4syV111VXrrrbe2XI8y7rrrrg35F198cTp16lRULMx79dVX0yWXXLJR/9577w3LLStz7Cba6l9//fULe7bblhKzd955Z0PPG2+8sVT0jLq2Lv6DU9iP6N8+vfrqq9P+/fub+zNkn47puvn3dPSBbBIBERABERABERCBEoFt20TbwvqOO+7Y2Jj6RTWfX3HFFen555/fsGPsJtpk9zzBRXvQabs3nmM30c8999wG994bChtOWMIBb760if4A+Lr4DxqzH9GHculFF120qa9DxpmUrpt/zyTfyFYREAEREAEREAERMALbsom2p8/4OLItpq+55pr0wgsvbHoK9cYbb6RDhw5tPAHmTS82tUOeRNsTL2uzdVOGDYAt7vG0fN030bYJv/baa/Ukek3HgHXxH/CiD+X6nV1/5plnNmLyTN9Ir5t/4WelIiACIiACIiACInCmEFj6JpqfotpiOfq4NsPHAnyqTbR9TNw+mt36BBYf5bbyX/7yl1di48kMmQtzOx2O4fvc5ut0sPFMsKHVjxzXrTe5zgR+3kYek3q+luLl6FwEREAEREAEREAERGAYgaVvovm7ya0bQFuEv/baaxsWjnkSbd8FxveBW9pHW7aoh+7r/iR6A+SKH7RuvlbcjDNevR4/or+13uQ6E+FqE30mel02i4AIiIAIiIAIrBKBpW6isfgb+2QRC+0hH+e2zTDq15528eLfNtzL2kRbu88++2x66qmntnzM3YKHn9jxjQBfb5UCbYguzL/mqyHyl1XnxIkTC1/yjaBltb0K7fT4Ed8HPh0/0m0cTp48me3Xrb7COLodNxrO9Fhu9ZHKiYAIiIAIiIAInN4ElrqJxuZ17AIZcoZuolsXob5cyybavttpL0yz717jP/eWcR9aTz/99Mb3ru1GA//by9VeeumlRRW/ibbvmN9www2L8vi4utU1zseOHfPNLM5hi9+c2kL/zjvvTLfeeuvGC57Mpr17927oY3JvvvnmDX3CBlJK9r32u+++e5NNrXVNZs/maw4den2JeDH2dnPDNoT8VndseqZgPLf/4DvW32ILb9C2F/1ZjOA8x7/Xj7VNNPTC+wkQ57l4NNb2bgXT9Vvf+tZCTd/PWmPSNpDWDyImpRskpkPuJYrGNOqj3r/cF3hc4OPceDhE75ZYxtdbENe5GGDdt/tTPDkdlS8CIiACIiACIiACPQQ2baJ/9G9vpgeP/Uv6wh8cTeddemCR2vmP3nizR2a2LBaGucVetqK7MHYT3bqog77YaOI8WgjawpwX17y4xTHkOHMWp7bZRDlLbXGNTTjywY030UePHl28pO2WW27Z9GI2fnkbP61G27DF68SyzU6vF3SxtLR4rvEobfChI/vJ64kypXSoDrV6ZnukD2889u3btxEP8CV4TcF4Tv+xHexvHFscHjx4cBGviMmSH3r8iL4NViy35pcoprjtWjxb/aivmIxSPzAuUdumO7dvZW6//fb06KOPLm5U4UaA5fOvD1g971+WAz/41PtijN4cA7lY/s53vrMR4xE3+A6yIjtRRqkIiIAIiIAIiIAIrBOBxSb63XffS4/+44n0id9+KJ2ze386+5P3pbM/+bVFaueWb9et3NA/v3EYKsfqYaHtF40lmX5RiideORmsLxaIkOE30Vjg2iLRNrP8VMquYcNhi17IYl1hj103ffDEmcvYBuL3f//3Fxtl1s3qRBs65hRdhy3+mpdt8u0puL09HX/2hBY3DHx9KwO2kT22ucd30muLanAt2QidfDpUB7Q5xJfYLJi+4Bb5cgrGc/mPdTMG/OI/851/oprrP+wPMK35kdv2csf6Ez6x1PdR3pxHm3ewtrr33HPPpptV9pQXb7q3675/Q+9crBtTe7pd20QzT8iMdOVyY/RuieVWv2J88z5lXXUsAiIgAiIgAiIgAutEYMd776X0D//8r+m833gg7fz0/elDn9ufPvJbf5nO/eJDi9TOLd+uWzkrP+SPF8h+E9orb8iiDAtKbPqgT25xi4UqL/wgw+tvi8nDhw+Hm1+zrbTY5MWq6fbWW29VcUB3W7SXFtKQHZWBLeCBRlm2yc/pBB942Vzfy0Ybluba5zIlblzOH4/RYSpf5uLKdGX9hjA2GTl+Y2VDbkl/+N505/7h/YDzmh/tI9q8kTW5vBllm3pjits2ub7vQkf0d18GfcjrhHqWchueB1j5fsL1o2P4IbIXupZkjtWb67fEQk4XZpNjH9mvPBEQAREQAREQARFYZQI7/v3H/5k+f93RtPMz96dzLv56Ou+P/j59/MHvpl1Hf7BI7dzy7frnrzuSrPyQP14Ij11MYWHqF6wlvaJFKfIifaJrUV6pTb4Gnf2iGPmlhSrLsWNm6eVxWZSLZMMWXx91bNOQWxhbG1hke9mwp1TX6rdsBHgB7vVkO/3xlDp42XYO+V4nMDF2/hrLGcvYZM3hP9Yr6hOwgf3S0ge5vLGp/duTYv4D7yExxW3XdAVTLhflsW44RjybbXwDALr7foJ6uRTtRnGEtko8UJ9tidqCLK/3kFiOYgZyeu2PdFWeCIiACIiACIiACKwKgR2PHz+Zdn5mf9q5e1/6hT9+Iu362xOL//OP/OD944dPLPLtupX75j+9Mkj31gV6i3AsTGsLRJaFRSUvSrGA9HKgq1/4QUa0WOS27COeTz755OJ7j/ZRTXz30Raq3FbPAp/lQz+/8OUydlwqB1uYh6/jr7F8yB7KqGVxzXxKurBedgzban5q0aHVl9Yu5PX4pWRXjjHb6OujjungrzEnlGP/tepvcnr6IPvR9Mr929cG+OPj0HeMP7ntWjxgPMDmtKcueJpt3A7n88sBYVsuhc2RD72eXsYUevfEAnTlsQ06IU4iO1BGqQiIgAiIgAiIgAisG4Edf/gn31p8//nnLj/0/hPoYy+n8//mpY3/XQ+/nH7lwe+mcy8/tCh3/Vfff8Ntr6G8mORFZq8cK4+FWbRoy8nDQo8Xc9CJNxJWH4tULx8yIv1t4eq/LxptFlgm2vcL75wNyOd6/NQL15GWysEW5mH1SnUgN1eO60a25/JyNvBmwOvJuvDxFDoM8WWOCeuGY9YxZ3tN3hz+Q9z7/gC9OUVZjme+zsclP2KzlmuTWeXiJ8oHV24beawbH8Mm6MJt1+pyOz5WTS7eIWC65t4kzrrk/GtloCc2+1zPjqfQu0cG9AE36MNMavxQR6kIiIAIiIAIiIAIrAOBHb96+cF01gX3pZ+/7uG069iJdP6RDzbQ2ExfcOzlxfWzLrg3XXjloUF28YLKLzJ7BU61ibZ2sVjljXGUlytr+bzgtEWyPXGyn/6x33m234W17zlHOnM9br/Gg+uVFqelcrDR+6JUh/WKynFetLHJ5eVsGBIzY3Xw9Vt96eMgZ9NU5ebwHzZDuc0Z+x9lx26iTWbOFs8qFz9RPvhzDCGP7eBj2ITNIMdCrS634/uUtWEvEOMXkJnO11xzTfY9CiUm0DPnpyn0HiqDxzHcIMnpyex1LAIiIAIiIAIiIALrRGDHL/7mA4tN9Mf2HksXPPLKxhNobKAtveCRk+lje/8u2Sb645cfHGwfNpJjF1WQ07KAh7K5RSkWpJCFxSMW0qhvKWTwQpHzozqoH+mMtmxB7WWiXpRyvdLivlQOtvgFf6kO6xKV47wee1guH9c2JlwWx2N1AJdeX1r73PZQv8COmjzoOaX/0BdKtkM/lEW/QX6U1vyIzZb1A8+NmQ6JKW7by/a6wibYz23X6nI73ifcDt7GzRt//x1wK5/zr12DnrlxdAq9e2SYTtH4hrwSD2ajYxEQAREQAREQARFYFwI7vnjDI+msC+9LH/29w+kTf/39tOvoiU0baTu3/I/+7uF01oX3pt+56dHBtvFieciCGA1jcdaygEed3KIUi0UsnP//9s7fxY4qiuP7V2SVqJVYCZY2JkRCEGSrLVKnimXANsVWAcso+NxEI5jCJuhWamURli22MRAIJqBZCCnXYBqjURn5Pvk+T87O3Dfv57wxn4Vl5s3ce358zp3HOXPn3XGCWifbMqLt7j+uEK6zuW3ibR+8jTpLyX2pnX3JCW6pj/VrW9duWn+i3Lg/jbxp+lhn9CnG2Oe9rYulzsX+08bFOsbJW0T8PPZ9LURb8n4Tg9xOn9vExP7k665N3zqdPhb7l2Ki9vbJxWnsWxoP6htjP66t2kt2fPd07mMe+fpUX8fJdupY/JuH3dGfcdykO7ePNrTpH+1nHwIQgAAEIAABCKw6gbVrX9+u1k9+Ur145mr1+ge7/85G+5HuG5qF/nl4XOfV7tpXt2fyyclhUwJYJ1wJ2e7u7uiUk92ccI8a1OxYb11S6nNKZON+FlN3Lt4YaEoWY0KZbbbMSXjkhDXb6c+ldtabeZT6WK62Te0sN/sZ+7bdj9yynSUZ09owayybmGRb59HOPmYus8iOfXNRF32IcWkT59g+22u5Jfb2tY0uy/O2jW61bWrXVrcL2zY3IGybtk3yfbyOl3WVvjPcfxwzy8p2x7HQ9L0W/Yi+yGbHs2Rj7s9nCEAAAhCAAAQg0BcCa3cPfqlefffTav3EoHpp84vqjQ/3qzd3fqr0O2ht9VnHdV7t7h48msk3J1eauW2zWq0ef7xw4cIzjzvPu4i2TbJnc3OzygmlHXZiGgsM9617FNX9bK/a5KTWSey4mWzL0rZtgltqZ19ykl7q08aG6E/b5DvKjftNhU1sU7c/rQ2zxnJWdtmXkrxFxc9ym64B2Rj55vGcfdDnNnGMbXLhFfVNOqai3LY+RR1txkTU0YZHZGTeuZ+P5+sz8s+cotxZ7S6Nvagn7jtOsmt7e3u4Cnud/bEP+xCAAAQgAAEIQKCPBNae/vl39dGXP1Trpz+rjp0YVMc3Ph8uIvba+98Mt8c3rg2P67zaqf2sf0q2vFqtElv9JlALcMU/vVpIq127XSxcXZTmxDP2z/ulpDQmwXWFrmVZRrQl9pUvOzs7bj5cTCiv2J1tjv2lWyy0EFn+29vbq86fPz881zbBLbWzLznJLfWJNjW1i/5kHrG/2ulx1qw/t9ENFHEptYt9tD+tDaV+upkzLpZNTLJ982i3qPjF4kvxi6+cEp/r168/84qqPJ6zrzkepTi6CMs3lEpxifrULo+p2FdyPSZ9jWWf6vwxa/W/fPnyM9envqe8YJhk37p1a2SS46xr+sGDB6Pj3omsMxfrzMfV15yk7/vv/3tbgr4j7t27Z/GjWe5J7ZYA266+8abCSHjNTh3ryKOmC4cgAAEIQAACEIBALwmsyerf//ir2hrsVeunrlTH3vq4Wj85qF54e3u4HX4+dWV4Xu3m9Ve3Wq0Strp/vRLGSa/0z7uIjjJLSaMT21hEq2+8KVBnv5Loks1KWJ2Iu79W7t3a2qouXrw4upHgBL9tgltqZ19ykl7qE2Nfaqdz0R8l+/LF//G92Vl/1JGTcrNp2saZuWltmCWWJSbRr3m0W2T8bt68ORpzdaw1Dl1Me0xG//J+jGPbeMdYSt608Yy6B4NB0S89iXJ4eJjNP/L75TomGuO52IxxVp94Hej98Zaj47nYbIqvWeiJGfeP22iDfI+/u47tvF9nd9YRZR6Bkw74e07y24yN1J2PEIAABCAAAQhAoBcEhkW0LP3tydPqu92D6p33blSvnLlavXx6e7jV52937w/PL8KjPOPs5O7cuXOVkt44s2L9TtQmSdJKSankelYoJ+/Wqa1l5CJa5+pW3VUh7Jk8zRKpiNSrr+INAcsfl/Aq2bWsmJyXEtxSO/uSi5pSH9uq7bh24/xRnMe9LzcWQB4XpW2O3bQ2TBvLcUzMbx7tFh0/z7z7SRBxP3v27Gj8TnINxjjm8WYm3nqWVfpy22niGXXrWmnyS0+Q1F2XtktbfVep+M1Mmmaa3SfeUMrjt+knLU3xtT1NY7Tu+3Iau9uOUdvjrb9H5Wfd96TbsYUABCAAAQhAAAJ9JjAqou3Eo8dPqoOHv1Y/3j+sDh4+rvSZv+USUNKrd0z7vy4xXq5Fs2nb398f+SKf7ty5M7ZgmU3j0d6rYMNRq/p7xEV0LnSX5VHbeOYieln2ZT2yI1/X44r2LKMPn118182u98F+bIQABCAAAQhAAAJtCBwpott0og0EIPB8Exg3U7oqdFaliF4VHou2wzdXJnlKaNE2IR8CEIAABCAAAQjMmwBF9LyJIg8C/3MCnm3swyO7FNHLG4yRNY9yL487miAAAQhAAAIQWD4BiujlM0cjBHpNwLONfXhkNxZ2pfUDeh2QFTHev2fPaxOsiHmYAQEIQAACEIAABOZGgCJ6bigRBIH+E9DCUBsbG8NXtUa9EBcAAAIqSURBVOXf7Kog9arcdQt/raL3FNHLiUpc0Z1Z6OUwRwsEIAABCEAAAt0RoIjujj2aIbByBOLqyiqUNduslfLjitQ63pffvFJEL26I5bHSp3GxOCpIhgAEIAABCEDgeSBAEf08RBkfITABAa0ifenSpSOFs4qk+Nq2CUR21pQienHocxGtV9blpxcWpx3JEIAABCAAAQhAoDsCFNHdsUczBCAAAQhAAAIQgAAEIAABCPSMAEV0zwKGuRCAAAQgAAEIQAACEIAABCDQHQGK6O7YoxkCEIAABCAAAQhAAAIQgAAEekaAIrpnAcNcCEAAAhCAAAQgAAEIQAACEOiOAEV0d+zRDAEIQAACEIAABCAAAQhAAAI9I0AR3bOAYS4EIAABCEAAAhCAAAQgAAEIdEeAIro79miGAAQgAAEIQAACEIAABCAAgZ4RoIjuWcAwFwIQgAAEIAABCEAAAhCAAAS6I0AR3R17NEMAAhCAAAQgAAEIQAACEIBAzwhQRPcsYJgLAQhAAAIQgAAEIAABCEAAAt0RoIjujj2aIQABCEAAAhCAAAQgAAEIQKBnBCiiexYwzIUABCAAAQhAAAIQgAAEIACB7ghQRHfHHs0QgAAEIAABCEAAAhCAAAQg0DMCFNE9CxjmQgACEIAABCAAAQhAAAIQgEB3BCiiu2OPZghAAAIQgAAEIAABCEAAAhDoGQGK6J4FDHMhAAEIQAACEIAABCAAAQhAoDsC/wDsuZW/kAW5iwAAAABJRU5ErkJggg==)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="-iBQStDFat-t" outputId="8ed21c45-c6e1-46ee-f21c-628af6102ce6"
breast_cancer.head()
# + id="f_zl7zYGauCC"
breast_cancer.columns = ["id_number", "clump_thickness", "unif_cell_size", "unif_cell_shape",
"marg_adhesion", "single_epith_cell_size","bare_nuclei", "bland_chromatin",
"normal_nucleoli", "mitoses", "class"]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="f-aK_5mmauF6" outputId="28b00bda-202c-4a20-ef73-1868dfc16732"
breast_cancer.head()
# + colab={"base_uri": "https://localhost:8080/"} id="6WjTrrlyauJY" outputId="ce0f1449-43b4-489f-a47c-22b3decd7359"
breast_cancer.info()
# + [markdown] id="OK7u7ca0eZaq"
# 누락값의 대체 및 클래스 레이블을 0과 1로 변환
# + id="lyj4hkrOauMD"
# "?"값을 null로 대체
breast_cancer.bare_nuclei = breast_cancer.bare_nuclei.replace("?", np.NaN)
# + colab={"base_uri": "https://localhost:8080/"} id="9iNygrxVauOj" outputId="d3e5aa13-bf48-4000-c84b-afc347490968"
breast_cancer.isnull().values.sum()
# + id="JBnl7nnIauSi"
#null값을 최빈값으로 대체
breast_cancer.bare_nuclei = breast_cancer.bare_nuclei.fillna(breast_cancer.bare_nuclei.value_counts().index[0])
# + colab={"base_uri": "https://localhost:8080/"} id="w98KA-ktauW1" outputId="8a3dc9e3-e8e3-4267-b714-f5a3a28d6cc0"
breast_cancer.isnull().values.sum()
# + id="7pt6ora8auZR"
#새로운 칼럼 cancer_ind을 생성하고 악성종양(4)의 데이터를 1로 설정
breast_cancer["cancer_ind"] = 0
breast_cancer.loc[breast_cancer["class"] == 4, "cancer_ind"] = 1
# + id="gRetZShIaued"
X = breast_cancer.drop(["id_number", "class", "cancer_ind"], axis=1)
y = breast_cancer.cancer_ind
# + [markdown] id="_-R27q5zgNcP"
# # 훈련용,테스트용 데이터 구성
# + id="UhIYOIK2auhG"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# + [markdown] id="aNJrz2nKdfzf"
# # 데이터 표준화 객체 생성
# + id="o0YNopEkaujZ"
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# + id="6nQgpflOaulw"
#훈련 데이터, 테스트 데이터를 표준화
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
# + [markdown] id="2piQ2JQ1dQpt"
# # KNN을 이용한 학습
# + id="F_jiajEtauoI"
from sklearn.neighbors import KNeighborsClassifier
# + colab={"base_uri": "https://localhost:8080/"} id="tZPK3lxNauqk" outputId="830db3d2-149a-485e-be6c-7a32c36db9df"
knn = KNeighborsClassifier(n_neighbors=3)
knn.fit(X_train_scaled, y_train)
# + [markdown] id="qX67RvmpdIUz"
# # 분류 모델의 혼동행렬,정확도,AUC
# + id="pqhC-KmVausm"
from sklearn.metrics import accuracy_score, confusion_matrix, roc_auc_score, roc_curve
# + id="VVgHbu9Rbzvw"
y_pred = knn.predict(X_test_scaled)
# + colab={"base_uri": "https://localhost:8080/"} id="Q8AnaJvNbzx6" outputId="2c42f434-014f-4ed9-95cc-7228472b2031"
accuracy_score(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="kgy29nW5bz0G" outputId="90e416b6-bdf7-43d2-c7ed-572f65938da6"
confusion_matrix(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="kygmCKqcbz2i" outputId="d2a2727a-55e6-4a8c-e360-a3797b6dd203"
roc_auc_score(y_test, y_pred)
# + [markdown] id="LsKhNRjVcupP"
# # 그리드 서치를 이용한 하이퍼파라미터의 최적 값 선택
# + id="209SzHUNbz5C"
from sklearn.model_selection import GridSearchCV
# + id="4fJCKwzWbz8G"
grid_search = GridSearchCV(knn, {"n_neighbors": [1, 2, 3, 4, 5]},n_jobs=-1, cv=7, scoring="roc_auc")
# + colab={"base_uri": "https://localhost:8080/"} id="g8DsWKTZauvm" outputId="8defcaa8-57db-4d47-eb84-a6e3c73988cc"
grid_search.fit(X_train_scaled, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="Cf0L3RxdcGJx" outputId="6155acf6-1fce-4bb8-d547-2ecb23340ac6"
grid_search.best_params_
# + id="sN0JkGXhaux9"
knn_best = grid_search.best_estimator_
# + id="GC3H3BHccQXL"
y_pred = knn_best.predict(X_test_scaled)
# + colab={"base_uri": "https://localhost:8080/"} id="TsaOs4BPcQaD" outputId="ee358aac-a361-40e7-d810-825b4b1926b3"
accuracy_score(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="HsVhTYuFcQcT" outputId="9dfea115-adee-461f-efb9-1ae624ce946b"
confusion_matrix(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="pwC1CRXvcQd4" outputId="15c372eb-4a56-48ef-b76a-7b1df20dda28"
roc_auc_score(y_test, y_pred)
# + [markdown] id="gKVzioTkjb1_"
# # Cross-Validation을 이용한 최적의 K 도출
# + id="CPl8AufTiAGo"
from sklearn.model_selection import cross_val_score
k_range = range(1,10) # k를 1~10까지 고려하여 최적의 k를 찾을 것이다.
k_scores = [] # 각 k들의 성능을 비교하기 위한 리스트생성이다.
for k in k_range :
knn = KNeighborsClassifier(k)
scores = cross_val_score(knn,X,y,cv=7,scoring = "accuracy") # 7-fold cross-validation
k_scores.append(scores.mean()) # 7-fold 각각 정확도의 평균으로 성능계산
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="9IOxDYjdjIMM" outputId="603d17de-ce7d-48aa-8f0c-c83dd7952127"
plt.plot(k_range, k_scores)
plt.xlabel('Value of K for KNN')
plt.ylabel('Cross-Validation Accuracy')
plt.show()
| 50,452 |
/forbes-list-2021.ipynb | 5ee8fd7ba2b250ba3c00a36c8140e97b9b2591c0 | [] | no_license | Ragnarstefanss/forbes-python | https://github.com/Ragnarstefanss/forbes-python | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 30,247 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
data=pd.read_csv("/kaggle/input/forbes-billionaires-of-2021/Billionaire.csv")
#data
# Used when printing all columns is unnecessary
column_filtering_one = ['Name', 'NetWorth', 'Source', 'Rank', 'Industry']
column_filtering_two = ['Name', 'NetWorth', 'Source', 'Industry']
column_filtering_three = ['NetWorth', 'Source', 'Industry']
column_filtering_country = ['Name', 'NetWorth', 'Source', 'Country']
# +
def get_person(data, name = ''):
return data[data['Name'].str.contains(name, regex=False)]
def get_from_country(data, name = ''):
return data[data['Country'] == name]
def count_values(data, value = ''):
return data[value].value_counts()
def net_worth_more_than(data, networth = 1):
return data[data['NewNetWorth'] >= networth]
def get_people_multiple_sources(data):
return data[data["Source"].str.contains(",", regex=False)]
def get_from_industry(data, name = ''):
return data[data['Industry'] == name]
# +
# All sources counted (even multiple sources)
sources = count_values(data, "Source")
# Convert string NetWorth to float without effecting the original column
data['NewNetWorth'] = pd.to_numeric(data.NetWorth.str.replace(r"[a-zA-Z\$]",''))
# All people with multiple sources
people_multiple_sources = get_people_multiple_sources(data)
# All people from <country>
from_us = get_from_country(data, 'United States')
from_china = get_from_country(data, 'China')
# All people from <country> with multiple sources
from_us_multiple_sources = get_people_multiple_sources(from_us)
from_china_multiple_sources = get_people_multiple_sources(from_china)
# -
# All people worth over <networth> billions
print(net_worth_more_than(people_multiple_sources, 2)[column_filtering_two])
# All people from <country> worth over <networth> billions
net_worth_more_than(from_us, 20)[column_filtering_country]
net_worth_more_than(from_china, 20)[column_filtering_country]
billion_dollars_number = 1000000000
print("US Billionaire Total Networth: ","{:,}".format(sum(from_us["NewNetWorth"])*billion_dollars_number))
print("China Billionaire Total Networth: ", "{:,}".format(sum(from_china["NewNetWorth"])*billion_dollars_number))
get_person(data, 'Jeff')
industries = count_values(data, 'Industry')
industries
for item in data['Industry'].unique():
industry = get_from_industry(data, item)
print(item, "networth: {:,}".format(int(sum(industry["NewNetWorth"]))), " billions")
print("---")
icelandic_billionaires = get_from_country(data, 'Iceland')
icelandic_billionaires
| 2,802 |
/3주차 과제.ipynb | 87b9bedeaf1df70480135fdf79840a7f03240821 | [] | no_license | namwoonKim/ML-Class | https://github.com/namwoonKim/ML-Class | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 701,274 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="oef8c2jr-PZY"
# 作業目標<br>
# 計算有缺失值的資料,統計量實作<br>
# 作業重點<br>
# 當遇到缺失值有函式可以處理,不須額外寫程式刪除<br>
# 計算統計量時不能出現缺失值
# + [markdown] id="4YgL8NF4-Ru3"
# 題目:<br>
# english_score = np.array([55,89,76,65,48,70])<br>
# math_score = np.array([60,85,60,68,np.nan,60])<br>
# chinese_score = np.array([65,90,82,72,66,77])<br>
# 上3列共六位同學的英文、數學、國文成績,第一個元素代表第一位同學,舉例第一位同學英文55分、數學60分、國文65分,今天第五位同學因某原因沒來考試,導致數學成績缺值,運用上列數據回答下列問題。<br>
# 1. 請計算各科成績平均、最大值、最小值、標準差,其中數學缺一筆資料可忽略?
# 2. 第五位同學補考數學後成績為55,請計算補考後數學成績平均、最大值、最小值、標準差?
# 3. 用補考後資料找出與國文成績相關係數最高的學科?
# + executionInfo={"elapsed": 617, "status": "ok", "timestamp": 1609854566856, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="QwsKU8qO-RK7"
import numpy as np
# + executionInfo={"elapsed": 1016, "status": "ok", "timestamp": 1609854567261, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="B5BqDQAe_czh"
english_score = np.array([55,89,76,65,48,70])
math_score = np.array([60,85,60,68,np.nan,60])
chinese_score = np.array([65,90,82,72,66,77])
# + executionInfo={"elapsed": 1005, "status": "ok", "timestamp": 1609854567262, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="K3tUA6u9aI3N"
#1. 請計算各科成績平均、最大值、最小值、標準差,其中數學缺一筆資料可忽略?
#平均
english_mean = np.nanmean(english_score)
math_mean = np.nanmean(math_score)
chinese_mean = np.nanmean(chinese_score)
#最大值
english_max = np.nanmax(english_score)
math_max = np.nanmax(math_score)
chinese_max = np.nanmax(chinese_score)
#最小值
english_min = np.nanmin(english_score)
math_min = np.nanmin(math_score)
chinese_min = np.nanmin(chinese_score)
#標準差
english_std = np.nanstd(english_score)
math_std = np.nanstd(math_score)
chinese_std = np.nanstd(chinese_score)
print("英文成績平均:",english_mean,"最大值:",english_max,"最小值:",english_min,"標準差:",english_std)
print("數學成績平均:",math_mean,"最大值:",math_max,"最小值:",math_min,"標準差:",math_std)
print("國文成績平均:",chinese_mean,"最大值:",chinese_max,"最小值:",chinese_min,"標準差:",chinese_std)
# + executionInfo={"elapsed": 996, "status": "ok", "timestamp": 1609854567262, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="mXjDzvSrbLlf"
#2. 第五位同學補考數學後成績為55,請計算補考後數學成績平均、最大值、最小值、標準差?
math_score = np.array([60,85,60,68,55,60])
math_mean = np.nanmean(math_score)
math_max = np.nanmax(math_score)
math_min = np.nanmin(math_score)
math_std = np.nanstd(math_score)
print("補考後數學成績平均:",math_mean,"最大值:",math_max,"最小值:",math_min,"標準差:",math_std)
# + executionInfo={"elapsed": 989, "status": "ok", "timestamp": 1609854567264, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="SaGolKx0cKoU"
#3. 用補考後資料找出與國文成績相關係數最高的學科?
print("國文和數學的相關係數:",np.corrcoef(chinese_score,math_score))
print("國文和英文的相關係數:",np.corrcoef(chinese_score,english_score))
# + executionInfo={"elapsed": 987, "status": "ok", "timestamp": 1609854567264, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="fTNlLZ9Pzyiy"
y = mglearn.datasets.make_wave(n_samples=40)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0, test_size=0.2)
reg = KNeighborsRegressor(n_neighbors=3)
reg.fit(X_train, y_train)
print("테스트 세트 예측:\n{}".format(reg.predict(X_test)))
print("테스트 세트 R^2: {:.2f}".format(reg.score(X_test, y_test)))
# +
from sklearn.neighbors import KNeighborsRegressor
X, y = mglearn.datasets.make_wave(n_samples=40)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0, test_size=0.15)
reg = KNeighborsRegressor(n_neighbors=3)
reg.fit(X_train, y_train)
print("테스트 세트 예측:\n{}".format(reg.predict(X_test)))
print("테스트 세트 R^2: {:.2f}".format(reg.score(X_test, y_test)))
# test_size를 0.1, 0.2, 0.15를 사용해서 정확도를 측정을 했음(위의 2개의 cell 포함)
# test_size에 따라 정확도가 모두 변하는 것을 확인할 수 있었음
# +
fig, axes = plt.subplots(1, 3, figsize=(15, 4))
for n_neighbors, ax in zip([1, 3, 9], axes):
reg = KNeighborsRegressor(n_neighbors=n_neighbors)
print("X train.shape: {}".format(X_train.shape), "X_test.shape: {}".format(X_test.shape))
reg.fit(X_train, y_train)
ax.plot(X_test, reg.predict(X_test), 'v', c=mglearn.cm3(2), markersize=8)
ax.plot(X_train, y_train, '^', c=mglearn.cm2(0), markersize=8)
ax.plot(X_test, y_test, 'v', c=mglearn.cm2(1), markersize=8)
ax.set_title(
"{} 이웃의 훈련 스코어: {:.2f} 테스트 스코어: {:.2f}".format(
n_neighbors, reg.score(X_train, y_train), reg.score(X_test, y_test)))
ax.set_xlabel("특성")
ax.set_ylabel("타깃")
axes[0].legend(["모델 예측", "훈련 데이터/타깃", "테스트 데이터/타깃"], loc="best")
# 이웃을 바꿔가면서 출력
# 처음에는 강의 내용과 다르게 출력이 되었음
# 위에서 계속 test_size를 변경해서 정확도를 측정했기 때문이라고 생각을 했음
# 그런데 test_size가 0.4인 것으로 해봐도 똑같이 나오지 않았음
# 그래서 더 위쪽의 test_size를 설정하지 않은 것으로 실행을 했고, 결과가 같은 것이 나옴
# 1번째 그래프는 주황색에서 가장 가까운 파란색을 통해서 타깃 값을 정함
# 2번째 그래프에서는 주황색에서 가까운 파란색 3개의 평균을 타깃 값으로 정함
# 3번째 그래프에서도 주황색에서 가까운 파란색 9개의 평균을 타깃 값으로 정함
# 초록색을 선을 그려보면 9개로 한 것은 부드럽게 결정 경계가 정해짐 - 다음 셀에서 확인 가능
# +
fig, axes = plt.subplots(1, 3, figsize=(15, 4))
line = np.linspace(-3, 3, 1000).reshape(-1, 1)
for n_neighbors, ax in zip([1, 3, 9], axes):
reg = KNeighborsRegressor(n_neighbors=n_neighbors)
print("X train.shape: {}".format(X_train.shape), "X_test.shape: {}".format(X_test.shape))
reg.fit(X_train, y_train)
ax.plot(line, reg.predict(line))
ax.plot(X_train, y_train, '^', c=mglearn.cm2(0), markersize=8)
ax.plot(X_test, y_test, 'v', c=mglearn.cm2(1), markersize=8)
ax.set_title(
"{} 이웃의 훈련 스코어: {:.2f} 테스트 스코어: {:.2f}".format(
n_neighbors, reg.score(X_train, y_train), reg.score(X_test, y_test)))
ax.set_xlabel("특성")
ax.set_ylabel("타깃")
axes[0].legend(["모델 예측", "훈련 데이터/타깃", "테스트 데이터/타깃"], loc="best")
# k의 수가 증가함에 따라 훈련에 대한 정확도는 감소, 테스트에 대해서는 더 안정된 예측이 가능함
# 강의 내용에서는 line에 대해서 나와있지 않기 때문에 line에 대해서 추가를 했음
# 결정 경계가 갈수록 부드러워 지는 것을 확인할 수 있음
# +
# x 값과 y 값
x=[2, 4, 6, 8]
y=[81, 93, 91, 97]
# x와 y의 평균값
mx = np.mean(x)
my = np.mean(y)
print("x의 평균값:", mx)
print("y의 평균값:", my)
# 기울기 공식의 분모
divisor = sum([(mx - i)**2 for i in x])
# 기울기 공식의 분자
def top(x, mx, y, my):
d = 0
for i in range(len(x)):
d += (x[i] - mx) * (y[i] - my)
return d
dividend = top(x, mx, y, my)
print("분모:", divisor)
print("분자:", dividend)
# 기울기와 y 절편 구하기
a = dividend / divisor
b = my - (mx*a)
# 출력으로 확인
print("기울기 a =", a)
print("y 절편 b =", b)
# np.mean을 처음 봤는데, 평균을 바로 구해주므로 유용하게 쓰일 것 같다.
# for i in x 문장이 i에다가 x의 값을 하나씩 넣어 반복하는 것이다.
# +
x=[10, 22, 41, 38]
y=[22, 40, 60, 44]
mx = np.mean(x)
my = np.mean(y)
print("x의 평균값:", mx)
print("y의 평균값:", my)
divisor = sum([(mx - i)**2 for i in x])
def top(x, mx, y, my):
d = 0
for i in range(len(x)):
d += (x[i] - mx) * (y[i] - my)
return d
dividend = top(x, mx, y, my)
print("분모:", divisor)
print("분자:", dividend)
a = dividend / divisor
b = my - (mx*a)
print("기울기 a =", a)
print("y 절편 b =", b)
# 리스트 x, y의 원소들을 바꿔서 실행을 하였다.
# x, y의 평균이 실수가 나오게 바꿧는데, 이 경우 a, b가 정확히 떨어지지 않았고,
# 소수점 이하로 계속해서 나오게 되었다.
# 이런 것을 보면 딱 떨어지는 숫자는 구하기가 쉽겠지만 딱 떨어지는 숫자가 아니면
# 계산하기 힘들 것 같다고 생각이 들었다.
# +
import numpy as np
# 기울기 a와 y 절편 b
ab = [3, 76]
# x, y의 데이터 값
data = [[2, 81], [4, 93], [6, 91], [8, 97]]
x = [i[0] for i in data]
y = [i[1] for i in data]
# y = ax + b에 a와 b 값을 대입하여 결과를 출력하는 함수
def predict(x):
return ab[0]*x + ab[1]
def rmse(p, a):
return np.sqrt(((p - a) ** 2).mean())
# RMSE 함수를 각 y 값에 대입하여 최종 값을 구하는 함수
def rmse_val(predict_result,y):
return rmse(np.array(predict_result), np.array(y))
# 예측 값이 들어갈 빈 리스트
predict_result = []
# 모든 x 값을 한 번씩 대입하여
for i in range(len(x)):
# predict_result 리스트를 완성한다.
predict_result.append(predict(x[i]))
print("공부한 시간 = %.f, 실제 점수 = %.f, 예측 점수 = %.f" % (x[i], y[i], predict(x[i])))
# 최종 RMSE 출력
print("rmse 최종값: " + str(rmse_val(predict_result,y)))
# 설명으로만 들었을 때는 엄청 간단하게 보였지만 프로그램으로 구현을 하니
# 복잡한 부분도 있었다. 이렇듯 간단한 것도 프로그램으로 구현을 하면 복잡한 부분이
# 생기는데 복잡한 것을 프로그램으로 구현을 하면 얼마나 복잡하게 프로그램을 만들어야
# 할지 감이 잡히지가 않았다.
# +
from sklearn.linear_model import LinearRegression
X, y = mglearn.datasets.make_wave(n_samples=60)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# 선형 모델을 사용해야 하므로 필요한 모듈을 import함
# 이번엔 샘플의 수가 60개임
# seed를 42로 해서 X, y를 train, test로 분할
lr = LinearRegression().fit(X_train, y_train)
# 선형 모델에서 X_train, y_train 으로 훈련
# +
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
# 과소적합됨 = 선 하나로 표시하므로 좋은 값이 나오지 않음
# -
print("lr.coef_:", lr.coef_)
print("lr.intercept_:", lr.intercept_)
# 기울기와 절편을 함수로 간단하게 보여줌
# 위에서 처럼 여러 함수들을 정의하고 구하지 않고, 두 함수로 간단하게 보여줌
# 여기서는 특성이 1개이므로 직선임
# 특성이 2개면 평면이고, n개면 초평면임
# 이러한 함수들이 있는지 처음 알았고, 빠르게 할 수 있다는 측면에서 좋은 것 같다.
# 이런 함수들을 잘 기억을 해놔야겠다.
# +
X, y = mglearn.datasets.make_wave(n_samples=100)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
lr = LinearRegression().fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
print("lr.coef_:", lr.coef_)
print("lr.intercept_:", lr.intercept_)
# +
X, y = mglearn.datasets.make_wave(n_samples=500)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
lr = LinearRegression().fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
print("lr.coef_:", lr.coef_)
print("lr.intercept_:", lr.intercept_)
# 샘플의 수도 점수들과 절편, 기울기에 영향을 줄 수 있다는 것을 알 수 있었다.
# 또한 점수들이 0.5~0.6 정도로 나오는 것을 보고 선 하나로 하는 것이 좋지 않아 보였다.
# +
plt.figure(figsize=(8, 8))
plt.plot(line, lr.predict(line))
plt.plot(X_train, y_train, 'o', c=mglearn.cm2(0))
plt.plot(X_test, y_test, 'v', c=mglearn.cm2(1), markersize=8)
# KNN 결과 출력
#1차원 특징이라서 정확률이 낮다. 샘플수가 커질수록 정확률이낮다. 데이터의 분포가 퍼져서
plt.plot(X_test, reg.predict(X_test),'v', c=mglearn.cm3(2), markersize=8)
ax = plt.gca()
ax.spines['left'].set_position('center')
ax.spines['right'].set_color('none')
ax.spines['bottom'].set_position('center')
ax.spines['top'].set_color('none')
ax.set_ylim(-3, 3)
#ax.set_xlabel("Feature")
#ax.set_ylabel("Target")
ax.legend(["모델", "훈련 데이터", "테스트", "KNN"], loc="best")
ax.grid(True)
ax.set_aspect('equal')
# 오른쪽 부분을 봤을 때 KNN과 선이 비슷하다는 느낌을 받았다.
# 왼쪽은 선과 비슷하지는 않지만 이웃의 수가 증가하면 오른쪽과 같이
# 선처럼 될 수 있을 것 같다고 생각을 했다.
# +
from sklearn.datasets import load_boston
boston = load_boston()
print("boston.keys(): {}\n".format(boston.keys()))
print("Data의 크기: {}\n".format(boston['data'].shape))
print("특성이름:\n{}\n".format(boston.feature_names))
print(boston['DESCR'] + "\n...")
df = pd.DataFrame(boston.data, columns=boston.feature_names)
print(df.head)
# boston의 키, data의 크기, 특성 이름, DESCR을 출력
# boston의 데이터를 DataFrame으로 출력
# -
import matplotlib.pyplot as plt
plt.scatter(boston.data[:, 5], boston.target, color='r', s=10)
# 방의 개수와 주택 가격의 산점도
# 방의 개수에 따라 주택 가격이 어떻게 되는가
# 대체적으로 방의 개수가 늘어나면 주택 가격이 늘어난다
# 몇몇 예외를 제외하면 늘어나는 것을 볼 수 있음
# +
# 속성 하나로 선형 회귀 실험
X = boston.data[:, 5]
print(X.shape)
# LinearRegression의 함수에 인자로 전달하기 위해서는
# 각 원소의 값이 담긴 2차원 배열을 (n, 1)의 형태로 만들어야 함
# atleast_2d 함수는 x 데이터를 2차원 배열을 (1, n) 형태로 만들어준다.
X_1 = np.atleast_2d(X)
X_2 = np.transpose(X_1)
print(X_2.shape)
y = boston.target
lr.fit(X_2, y)
y_predicted = lr.predict(X_2)
print("w[0]: %f b: %f" % (lr.coef_[0], lr.intercept_))
# 특징 하나로 2차원 배열로 만들고 lr을 통해 예측을 함
# 그 후에 기울기와 절편을 구함
# -
plt.plot(X_2, y_predicted)
plt.plot(X_2, y, 'o', color='r', markersize=3)
# 선형 회귀 모델을 먼저 그리고, 그 다음에 산점도 모델을 그림
X_train, X_test, y_train, y_test = train_test_split(boston.data, boston.target, random_state=0)
print("X_train.shape: {}".format(X_train.shape), "X_test.shape: {}".format(X_test.shape))
lr = LinearRegression().fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
# 선형 회귀 모델의 성능 측정
# 훈련, 테스트의 점수를 출력
X_train, X_test, y_train, y_test = train_test_split(boston.data, boston.target, random_state=0)
print("X_train.shape: {}".format(X_train.shape), "X_test.shape: {}".format(X_test.shape))
lr = KNeighborsRegressor().fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
# 이번엔 KNN으로 성능을 측정
# 선형 회귀 모델보다 점수들이 낮은 것을 볼 수 있음
# 나는 지금까지 선형 모델이 선 하나로 하기 때문에 KNN보다 낮은 점수가 나올 것이라
# 생각을 해왔었다. 하지만 여기서 선형 회귀 모델과 KNN을 비교해봤더니 이번엔 선형
# 회귀 모델의 점수가 더 좋은 것을 볼 수 있었다. 그래서 둘 중 어느 것이 낫다가
# 아니라 상황에 따라 적절한 모델이 있으므로 모델을 적절하게 사용해야 한다고
# 생각을 하였다.
# +
X, y = mglearn.datasets.load_extended_boston()
print("X.shape: {}".format(X.shape))
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
print("X_train.shape: {}".format(X_train.shape), "X_test.shape: {}".format(X_test.shape))
lr = LinearRegression().fit(X_train, y_train)
# 이번엔 보스톤 데이터를 확장 한 것(특성이 104개)으로 두 모델 비교
# 먼저 선형 회귀 모델을 만들고 훈련선형 회귀 모델을 만들고 훈련
# -
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
# 선형 회귀 모델에서 훈련 세트 점수는 엄청 높지만 테스트 세트 점수는 낮다.
# 즉, 과대적합이라고 생각할 수 있다. 따라서 조정이 필요하다.
knn = KNeighborsRegressor(n_neighbors=3).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(knn.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(knn.score(X_test, y_test)))
# KNN도 위와 비슷한 결과가 나옴
# 이것도 과대적합이라 볼 수 있음
knn = KNeighborsRegressor(n_neighbors=5).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(knn.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(knn.score(X_test, y_test)))
knn = KNeighborsRegressor(n_neighbors=9).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(knn.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(knn.score(X_test, y_test)))
knn = KNeighborsRegressor(n_neighbors=33).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(knn.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(knn.score(X_test, y_test)))
# 이웃의 수를 증가시켜서 해봤지만 훈련 세트 점수와 테스트 세트 점수 모두 떨어지는
# 것을 볼 수 있었다.(위의 셀들 포함) 그래서 이 보스턴 데이터를 확장한 것은
# KNN에 적합하지 않은 모델인 것 같다고 생각을 하였다.
# +
from sklearn.linear_model import Ridge
ridge = Ridge().fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge.score(X_test, y_test)))
# default 값인 1.0일 경우의 점수
# 이 값을 크게하면 계수의 편차가 적음(과소적합)
# 작게하면 계수 편차가 크다(과대적합)
# 아래 두 셀과 비교를 해보면 linear, KNN 두 모델 모두 테스트 점수가 낮지만
# ridge에서 테스트 점수가 높아진 것을 볼 수 있다.
# -
print("훈련 세트 점수: {:.2f}".format(lr.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lr.score(X_test, y_test)))
knn = KNeighborsRegressor(n_neighbors=3).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(knn.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(knn.score(X_test, y_test)))
ridge10 = Ridge(alpha=10).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
# alpha가 높을수록 과소적합
# 계수의 편차가 적다
# 계수를 0에 가깝게
# 훈련 데이터의 성능이 나빠짐
# 일반화에 도움이 된다
# 모델 복잡도가 낮아진다
ridge10 = Ridge(alpha=5).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
ridge10 = Ridge(alpha=0.5).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
ridge10 = Ridge(alpha=0.3).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
ridge10 = Ridge(alpha=0.2).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
ridge10 = Ridge(alpha=0.1).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
# +
ridge10 = Ridge(alpha=0.01).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(ridge10.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(ridge10.score(X_test, y_test)))
# alpha가 작을수록 과대적합
# 계수 편차가 크다
# 계수에 거의 제한을 두지 않는다
# 선형모델과 비슷해진다
# 그러면 alpha를 어느 정도로 해야 하는가?
# 테스트 점수가 어느 정도 있고, 훈련 점수도 어느 정도 있는 정도를 사용하면 됨
# 여기서는 alpha=0.5 정도가 좋다
# +
from sklearn.linear_model import Lasso
lasso = Lasso().fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lasso.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lasso.score(X_test, y_test)))
print("사용한 특성의 개수: {}".format(np.sum(lasso.coef_ !=0)))
# ahpha가 기본 값인 1
# 낮은 성능, 적은 특성 사용, 과소적합
# +
lasso001 = Lasso(alpha=0.01, max_iter=100000).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lasso001.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lasso001.score(X_test, y_test)))
print("사용한 특성의 개수: {}".format(np.sum(lasso001.coef_ !=0)))
# ahpha를 0.01로 해서 다시 실행
# 패널티 효과 감소, 복잡도 증가
# 선형 모델과 유사하게 나옴(ridge10과 비교)
# alpha를 아주 작게하면 서형 모델과 유사
# +
lasso00001 = Lasso(alpha=0.0001, max_iter=100000).fit(X_train, y_train)
print("훈련 세트 점수: {:.2f}".format(lasso00001.score(X_train, y_train)))
print("테스트 세트 점수: {:.2f}".format(lasso00001.score(X_test, y_test)))
print("사용한 특성의 개수: {}".format(np.sum(lasso00001.coef_ !=0)))
# ahpha를 0;0001로 해서 다시 실행
# ahpah=0.01보다 테스트 점수가 떨어짐
# 일반화가 잘 안되고 있음
# -
print("계수값:\n", lasso.coef_)
print(np.where(lasso.coef_ !=0))
# lasso에 사용된 특성을 출력
# 그런데 4개 밖에 사용이 안됨(0이 아닌 값들)
# 사용된 특성들의 index를 출력
# where 함수도 처음 봤는데, 검색을 해보니 조건에 해당하는 인덱스를 출력을 해준다.
# +
plt.plot(lasso.coef_, 's', label="Lasso alpha=1")
plt.plot(lasso001.coef_, '^', label="Lasso alpha=0.01")
plt.plot(lasso00001.coef_, 'v', label="Lasso alpha=0.0001")
plt.plot(ridge10.coef_, 'o', label="Lidge alpha=0.1")
plt.legend(ncol=2, loc=(0, 1.05))
plt.ylim(-25, 25)
plt.xlabel("계수 목록")
plt.ylabel("계수 크기")
# 라쏘 회귀
# alpha = 1일 경우, 계수 대부분이 0 ,나머지도 크기가 작음
# alpha = 0.01의 경우, 많은 특성이 0이 된다.(0이 아닌 것도 있다.)
# alpha = 0.0001의 경우, 값이 커지고 규제를 받지 않음
# 라쏘 회귀와 릿지 회귀를 잘 구분해야 함
# +
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
# iris 로지스틱회귀를 하기 위해 필요한 모듈을 import함
# +
iris = datasets.load_iris()
print(list(iris.keys()))
print("타깃의 이름: {}".format(iris['target_names']))
print("특성의 이름: {}".format(iris['feature_names']))
X = iris["data"][:, 3:]
print(X.shape)
y=(iris["target"]==2).astype(np.int)
print(y.shape)
# datasets에서 iris를 불러옴
# iris의 키의 리스트를 출력하고
# 타깃의 이름과 특성의 이름을 출력
# data에서 pental width만 가져옴
# target에서도 virginica(1 아니면 0만 있음)만 가져옴
# +
log_reg = LogisticRegression()
log_reg.fit(X, y)
X_new = np.linspace(0, 3, 1000).reshape(-1, 1)
y_proba = log_reg.predict_proba(X_new)
plt.plot(X, y, "b.")
plt.plot(X_new,y_proba[:,1],"g-",label="iris-Virginica")
plt.plot(X_new,y_proba[:,0],"b--",label="Not iris-Virginica")
plt.xlabel("Petal width", fontsize=14)
plt.ylabel("Probability", fontsize=14)
plt.legend(loc="upper left", fontsize=14)
plt.show()
# 꽃잎의 너비가 0~3cm인 꽃에 대해 모델의 추정 확률
# X, y는 점으로 표현 - 출력이 0 아니면 1로 되어있음
# Iris-Virginica는 녹색 실선, Not iris-Virginica는 파란 점선
# 두 선이 만나는 부분이 결정경계임(아래 쉘을 통해 1.7이라는 것을 알 수 있음)
# +
log_reg.predict([[1.7], [1.5], [1.3], [2.0]])
# Pental width이 결정경계가 1.7인 것을 알 수 있음
# 결정경계보다 크거나 같으면 1이고 작으면 0임
# +
log_reg = LogisticRegression(solver='liblinear',penalty='l1')
log_reg.fit(X, y)
X_new = np.linspace(0, 3, 1000).reshape(-1, 1)
y_proba = log_reg.predict_proba(X_new)
plt.plot(X, y, "b.")
plt.plot(X_new,y_proba[:,1],"g-",label="iris-Virginica")
plt.plot(X_new,y_proba[:,0],"b--",label="Not iris-Virginica")
plt.xlabel("Petal width", fontsize=14)
plt.ylabel("Probability", fontsize=14)
plt.legend(loc="upper left", fontsize=14)
plt.show()
# 강의 내용에 나온 log_reg = LogisticRegression(solver='liblinear',penalty='l1')
# 으로 실행을 해봤다.
# 하지만 결과는 같았다.
# 아마 기본 값이 같아서 같은 결과가 나오지 않았을까 하는 생각을 했다.
# +
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
X, y = mglearn.datasets.make_forge()
fig, axes = plt.subplots(1, 2, figsize=(10, 3))
for model, ax in zip([LinearSVC(), LogisticRegression()], axes):
clf = model.fit(X, y)
mglearn.plots.plot_2d_separator(clf, X, fill=False, eps=0.5,
ax=ax, alpha=.7)
mglearn.discrete_scatter(X[:, 0], X[:, 1], y, ax=ax)
ax.set_title("{}".format(clf.__class__.__name__))
ax.set_xlabel("특성 0")
ax.set_ylabel("특성 1")
axes[0].legend
# make_forge로 예전에 만들었었는데, LinearSVC, LogisticRegression을 적용
# 둘을 살펴보면 결정 경계가 다름
# 세모, 동그라미 하나씩 잘못 되어있음
# 나는 아래에 빨간 경고문구가 뜨고, LogisticRegression 그래프가 강의와 다르게 나오는데
# 아직도 왜 경고가 뜨고, 다르게 나오는지 해결하지 못하고 있다.
# +
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, stratify=cancer.target, random_state=42)
logreg = LogisticRegression(max_iter=5000).fit(X_train, y_train)
print("훈련 세트 점수: {:.3f}".format(logreg.score(X_train, y_train)))
print("테스트 세트 점수: {:.3f}".format(logreg.score(X_test, y_test)))
# cancer 데이터로 malignant와 benign을 불러서 사용
# 샘플 569개, 특성 30개
# 먼저 기본 값(1.0)으로 실행
# +
logreg100 = LogisticRegression(C=100, max_iter=5000).fit(X_train, y_train)
print("훈련 세트 점수: {:.3f}".format(logreg100.score(X_train, y_train)))
print("테스트 세트 점수: {:.3f}".format(logreg100.score(X_test, y_test)))
# 이번에는 C(규제)에 100을 줌
# inverse이므로 1/100임
# 패널티가 작아지고, 규제가 완화됨, 훈련세트에 맞추려 함
# 개개의 포인터에 맞추려 함
# +
logreg001 = LogisticRegression(C=0.01, max_iter=5000).fit(X_train, y_train)
print("훈련 세트 점수: {:.3f}".format(logreg001.score(X_train, y_train)))
print("테스트 세트 점수: {:.3f}".format(logreg001.score(X_test, y_test)))
# 이번에는 C(규제)에 0.01을 줌
# inverse이므로 100임
# 패널티가 커지고, 규제가 강화됨, 계수를 0에 가깝게
# 다수의 포인터에 맞추려 함, 일반화가 됨
# 아직 데이터가 크거나 일반화되지 않아서 별로 달라지지 않음
# 섞어서 하거나 새로운 데이터에 더 잘맞을 수도 있음
# +
for C, marker in zip([0.001, 1, 100], ['o', '^', 'v']):
lr_l1 = LogisticRegression(solver='liblinear', C=C, penalty='l1', max_iter=1000).fit(X_train, y_train)
print("C={:.3f} 인 l1 로지스틱 회귀의 훈련 정확도: {:.2f}".format(
C, lr_l1.score(X_train, y_train)))
print("C={:.3f} 인 l1 로지스틱 회귀의 테스트 정확도: {:.2f}".format(
C, lr_l1.score(X_test, y_test)))
plt.plot(lr_l1.coef_.T, marker, label="C={:.3f}".format(C))
plt.xticks(range(cancer.data.shape[1]), cancer.feature_names, rotation=90)
xlims = plt.xlim()
plt.hlines(0, xlims[0], xlims[1])
plt.xlim(xlims)
plt.xlabel("특성")
plt.ylabel("계수 크기")
plt.ylim(-5, 5)
plt.legend(loc=3)
# 앞에서는 L2 규제를 해봤음, 이번에는 L1 규제(앞에서는 L1 규제를 할 수 없음)
# 규제가 100이면 0.99이므로 과적합이 L2보다 커짐
# 이것은 L1의 특징이 전체를 다 쓰지 않고 선별적으로 쓰므로 너무 작게쓰면 과소적합이
# 될 수 있고, 너무 크게 쓰면 과대 적합이 될 수 있음
# L1은 L2보다 0으로 되는 것이 많음(C가 작을수록)
# +
for C, marker in zip([0.001, 1, 100], ['o', '^', 'v']):
lr_l1 = LogisticRegression(solver='liblinear', C=C, max_iter=1000).fit(X_train, y_train)
print("C={:.3f} 인 l2 로지스틱 회귀의 훈련 정확도: {:.2f}".format(
C, lr_l1.score(X_train, y_train)))
print("C={:.3f} 인 l2 로지스틱 회귀의 테스트 정확도: {:.2f}".format(
C, lr_l1.score(X_test, y_test)))
plt.plot(lr_l1.coef_.T, marker, label="C={:.3f}".format(C))
plt.xticks(range(cancer.data.shape[1]), cancer.feature_names, rotation=90)
xlims = plt.xlim()
plt.hlines(0, xlims[0], xlims[1])
plt.xlim(xlims)
plt.xlabel("특성")
plt.ylabel("계수 크기")
plt.ylim(-5, 5)
plt.legend(loc=3)
# 나는 패널티를 디폴트로 했는데 정확도가 강의와 다르게 나타났다.
# 그래도 전체적으로는 비슷하게 나타나서 그대로 사용을 하였다.
# 주황색은 조금 살아남았고, 초록색은 많이 살아남았다.
# mean parameter는 전부 0에 가까우므로 특정 클래스에 영향을 주지 않고
# 양 클래스 모두에 영향을 줄 수 있는 특징임
# texture error의 경우는 양수에 치우쳐져 있으므로 음수보다 양수쪽으로 계수가
# 맞춰져 있다는 것은 어떤 한 클래스와 구분하게 영향을 주는 특징이다.
# +
for C, marker in zip([0.001, 1, 100], ['o', '^', 'v']):
lr_l1 = LogisticRegression(solver='liblinear', C=C, penalty='l1', max_iter=1000).fit(X_train, y_train)
print("C={:.3f} 인 l1 로지스틱 회귀의 훈련 정확도: {:.2f}".format(
C, lr_l1.score(X_train, y_train)))
print("C={:.3f} 인 l1 로지스틱 회귀의 테스트 정확도: {:.2f}".format(
C, lr_l1.score(X_test, y_test)))
print("사용한 특성의 개수: {}".format(np.sum(lr_l1.coef_ != 0)))
print("사용한 특성: {}\n".format(lr_l1.coef_))
plt.plot(lr_l1.coef_.T, marker, label="C={:.3f}".format(C))
plt.xticks(range(cancer.data.shape[1]), cancer.feature_names, rotation=90)
xlims = plt.xlim()
plt.hlines(0, xlims[0], xlims[1])
plt.xlim(xlims)
plt.xlabel("특성")
plt.ylabel("계수 크기")
plt.ylim(-5, 5)
plt.legend(loc=3)
# 사용된 특성의 개수와 어떤 특성을 사용했는지 출력
# C가 커질수록 사용된 특성의 개수가 많아짐
# C가 큰 경우와 작은 경우 어떻게 되는지 알고 있어야 함
| 24,643 |
/preprocessing/propertyvalues/fix_tax_addresses_master.ipynb | 8b7b20f47b021adf1445389c5e64b3f4a87849b9 | [] | no_license | asensio-lab/albany-housing-analytics | https://github.com/asensio-lab/albany-housing-analytics | 0 | 0 | null | 2022-12-08T05:19:37 | 2019-08-01T02:47:41 | Jupyter Notebook | Jupyter Notebook | false | false | .py | 197,292 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pymongo as pm
from bson.objectid import ObjectId
import logging
logging.basicConfig(level=logging.DEBUG)
log = logging.getLogger(__name__)
# +
# Initialize PyMongo to work with MongoDBs
conn = 'mongodb://localhost:27017'
client = pm.MongoClient(conn, maxPoolSize=200)
# define db
DB_NAME = 'FINALP'
collection = 'WashTimes'
db = client[DB_NAME]
# +
# rename collection
# db.dailycaller.rename('r_'+'dailycaller')
# -
sorted(db.collection_names())
# extract text field and save to a file
i = 0
PATH = 'l'
for collection in ['l_atlantic',
'l_bbc',
'l_guardian',
'l_huffpost',
'l_mjones',
'l_newrep',
'l_nytimes',
'l_politoco',
'l_slate',
'l_thedailybeast',
'l_theintercept',
'l_washpost']:
for doc in db[collection].find():
i+=1
filename = PATH + '/' + str(i) + '.txt'
log.debug(f'Processing {doc["_id"]}: {filename}')
try:
with open(filename,"w+") as f:
f.write(str(doc['meta']['text']))
except Exception as e:
log.debug(e)
# extract text field and save to a file
i = 0
PATH = 'r'
for collection in ['r_amconser',
'r_breitbart',
'r_dailycaller',
'r_dailywire',
'r_economist',
'r_fiscaltimes',
'r_foxnews',
'r_nypost',
'r_reason',
'r_thehill',
'r_washtimes']:
for doc in db[collection].find():
i+=1
filename = PATH + '/' + str(i) + '.txt'
log.debug(f'Processing {doc["_id"]}: {filename}')
try:
with open(filename,"w+") as f:
f.write(str(doc['meta']['text']))
except Exception as e:
log.debug(e)
# LEFT = !ls l/
len(LEFT)
# +
bad_string = 'Copyright © 2018 The Washington Times, LLC. Click here for reprint permission.'
def substract(a, b):
return "".join(a.rsplit(b))
for doc in db[collection].find():
log.debug(doc['_id'])
# remove unneeded text sentence from article
text = substract(doc['meta']['text'],bad_string)
db[collection].update_one(
{
'_id':ObjectId(doc['_id'])
},
{
'$set':
{
'meta.text' : text
}
}
,
upsert=True
)
# -
doc = db[collection].find_one()
log.debug(doc['_id'])
doc = db[collection].find_one({'_id':ObjectId('5b5ba39dd1c45dd7bcdc9e9e')})
doc
| 3,035 |
/ML-Ensemble/BAGGING-RandomForest.ipynb | 80e362daf610fbfacb891e0dfc3f4c650a696253 | [] | no_license | Aprilcui11/Python_notes | https://github.com/Aprilcui11/Python_notes | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 2,709 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Random Forest
# Random forest is an extension of bagged decision trees.
#
# Samples of the training dataset are taken with replacement, but the trees are constructed in a way that reduces the correlation between individual classifiers. Specifically, rather than greedily choosing the best split point in the construction of the tree, only a random subset of features are considered for each split.
#
# You can construct a Random Forest model for classification using the RandomForestClassifier class.
#
# The example below provides an example of Random Forest for classification with 100 trees and split points chosen from a random selection of 3 features.
# Random Forest Classification
import pandas
from sklearn import cross_validation
from sklearn.ensemble import RandomForestClassifier
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/pima-indians-diabetes/pima-indians-diabetes.data"
names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class']
dataframe = pandas.read_csv(url, names=names)
array = dataframe.values
X = array[:,0:8]
Y = array[:,8]
num_folds = 10
num_instances = len(X)
seed = 7
num_trees = 100
max_features = 3
kfold = cross_validation.KFold(n=num_instances, n_folds=num_folds, random_state=seed)
model = RandomForestClassifier(n_estimators=num_trees, max_features=max_features)
results = cross_validation.cross_val_score(model, X, Y, cv=kfold)
print(results.mean())
| 1,700 |
/assignment2/ConvolutionalNetworks.ipynb | 89683f984f6f29f521f69b7bf06a8eff7bb87279 | [] | no_license | MellonGuan/cs231n | https://github.com/MellonGuan/cs231n | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 213,853 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##################################
### Import the usual libraries ###
##################################
### Numpy
import numpy as np
### Astropy
import astropy
#Astropy FITS/Table handling
from astropy.io import fits, ascii
from astropy.table import Table, Column
#astropy coorindates/units
from astropy.coordinates import SkyCoord
import astropy.constants as const
import astropy.units as u
### Pandas
import pandas as pd
### fitting
import statsmodels.api as sm
### PCA
from sklearn.decomposition import PCA, KernelPCA
### Model generation
import itertools as it
### Matplotlib
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
matplotlib.rcParams.update({'font.size': 18}) #make plots more readable
# ### Yumi's RA/DEC ---> Magellanic Stream Coorindates
import MagellanicStream as ms
# ### Functions used in this code
import age_functions as afunc
### Stuff Writtten by A Really Cool Professor
# import dlnpyutils as dlpu
### Other
import warnings
warnings.filterwarnings('ignore')
from tqdm import tqdm_notebook
### Age of Universe
H0 = 74.03*(u.km/u.s)/u.Mpc
hertz = H0.to(u.km/u.s/u.pc).to(u.km/u.s/u.km)
tage = (1/hertz).to(u.yr)
ageU = tage.value
# +
###################
### Import Data ###
###################
# parsec
''' Ages used 8 to 10.1 steps 0.15'''
''' Metallicity used -2.6 to 0.1 steps 0.099'''
parsecall = ascii.read('/Users/joshuapovick/Desktop/Research/parsec/parsec3_3.dat', \
format='basic', delimiter='\s')
rgb = np.where(parsecall['label']==3)
parsec = parsecall[rgb]
# lmc
lmc = fits.getdata('/Users/joshuapovick/Desktop/Research/fits/lmc_rgbmembers.r13-l33-58672.fits.gz')
cln = np.where((lmc['FE_H']>-9999.0)&(lmc['AK_TARG']>-100.0)&(lmc['LOGG']>=0.0)&
(lmc['M_H_ERR']>-100.0)&(lmc['C_FE']>-100.0)&(lmc['N_FE']>-100.0))
lmc = lmc[cln]
# APOKASC
# kasc = fits.getdata('/Users/joshpovick/Desktop/Research/LMC_Ages/APOKASC_cat_v6.6.1.fits.gz')
# Diane Feuillet
pdfout = fits.getdata('/Users/joshuapovick/Desktop/Research/fits/LMC_DR16_all_PDF.fits.gz', 1)
_, r13_ind, pdfout_ind = np.intersect1d(lmc['APOGEE_ID'],pdfout['OBJ'],return_indices=True)
# -
plt.hist(lmc['LOGG'])
lowgrav = parsec[np.where(parsec['logg']<=1.5)]
# +
###############################
### Some Calcuations Needed ###
###############################
### Convert RA DEC to MS L B
c_icrs = SkyCoord(ra=lmc['RA']*u.degree, dec=lmc['DEC']*u.degree)
c_ms = c_icrs.transform_to(ms.MagellanicStream)
ms_lg,ms_bg = c_ms.MSLongitude.degree, c_ms.MSLatitude.degree
for i in range(len(ms_lg)):
if ms_lg[i]-360<-180:
ms_lg[i] += 360.0
### Get Solar Fractions
abd = {'C':8.39,'N':7.78,'O':8.66,'Mg':7.53,'Ca':6.31,'S':7.14,'Si':7.51,'Fe':7.45}
keys = list(abd.keys())
sol_2_H = []
for i in keys:
sol_2_H.append(abd[i]-12.0)
sol_xm = 10**np.asarray(sol_2_H)/sum(10**np.asarray(sol_2_H))
# +
### Setup
data = np.array([lowgrav['logTe'],lowgrav['Ksmag'],lowgrav['MH'],lowgrav['logg']]).T
### Determine Number of factors
pca = PCA(n_components=4).fit(data)#_rescaled)
plt.rcParams["figure.figsize"] = (12,6)
fig, ax = plt.subplots()
y = np.cumsum(pca.explained_variance_ratio_)
xi = np.arange(1, len(pca.explained_variance_ratio_)+1, step=1)
plt.ylim(0.0,1.1)
plt.plot(xi, y, marker='o', linestyle='--', color='b')
plt.xlabel('Number of Components')
plt.xticks(np.arange(0, 5, step=1)) #change from 0-based array index to 1-based human-readable label
plt.ylabel('Cumulative variance (%)')
plt.title('The number of components needed to explain variance')
plt.axhline(y=0.99, color='r', linestyle='-')
plt.text(0.5, 0.85, '99% cut-off threshold', color = 'red', fontsize=16)
ax.grid(axis='x')
plt.show()
# +
### Find all models
#transform data to new basis
new_data = PCA(n_components=3).fit(data).transform(data)
#create new variables
# linear terms
x1 = new_data[:,0]
x2 = new_data[:,1]
x3 = new_data[:,2]
# linear int
x12 = np.multiply(x1,x2)
x13 = np.multiply(x1,x3)
x23 = np.multiply(x2,x3)
# squares
x1sq = x1**2
x2sq = x2**2
x3sq = x3**2
# cubes
x1cu = x1**3
x2cu = x2**3
x3cu = x3**3
#find all possible models
models = []
models_str = []
all_var_str = ['x1','x2','x3','x12','x13','x23','x1sq','x2sq','x3sq','x1cu','x2cu','x3cu']
all_var = [x1,x2,x3,x12,x13,x23,x1sq,x2sq,x3sq,x1cu,x2cu,x3cu]
for i in range(1,len(all_var)+1):
for subset in it.combinations(all_var,i):
models.append(subset)
for subset_str in it.combinations(all_var_str,i):
models_str.append(np.array(subset_str))
models = np.array(models)
models_str = np.array(models_str)
### Fit All Models
import statsmodels.api as sm
all_params = []
summaries = []
max_resid = []
mads = []
resids = []
predict = []
ll = []
for i in tqdm_notebook(range(len(models)),desc='Done?'):
pmodl = np.array(models[i]).T
pmodl = sm.add_constant(pmodl)
model = sm.OLS(lowgrav['logAge'],pmodl).fit()
summaries.append(model.summary())
predictions = model.predict(pmodl)
predict.append(predictions)
residual = predictions - lowgrav['logAge']
resids.append(residual)
all_params.append(np.asarray(model.params))
max_resid.append(np.max(np.absolute(residual)))
mads.append(afunc.mad(residual))
ll.append(model.llf)
# -
plt.scatter(mads,ll)
plt.scatter(ll,max_resid)
print(r'MAD')
print(models_str[np.asarray(mads).argmin()])
print(all_params[np.asarray(mads).argmin()])
print(r'LL')
print(models_str[np.asarray(ll).argmax()])
print(all_params[np.asarray(ll).argmax()])
print(r'max resid')
print(models_str[np.asarray(max_resid).argmin()])
print(all_params[np.asarray(max_resid).argmin()])
print(r'Current')
print('''['x1' 'x2' 'x3' 'x12' 'x13' 'x23' 'x1sq' 'x2sq' 'x3sq' 'x2cu' 'x3cu']''')
parsimonious = np.array([x1,x2,x3,x12,x13,x23,x1sq,x2sq,x3sq,x2cu,x3cu]).T
parsimonious = sm.add_constant(parsimonious)
parsimonious_model = sm.OLS(lowgrav['logAge'],parsimonious).fit()
parsimonious_predictions = parsimonious_model.predict(parsimonious)
parsimonious_residual = parsimonious_predictions - lowgrav['logAge']
print(afunc.mad(parsimonious_residual))
parsimonious_summary = parsimonious_model.summary()
print(parsimonious_summary)
print(np.asarray(parsimonious_model.params))
# +
def mad_age(x1,x2,x3):
#'x1' 'x2' 'x3' 'x12' 'x13' 'x1sq' 'x2sq' 'x3sq' 'x1cu' 'x2cu'
p = [9.62380331e+00,-4.37017699e-02,-8.10474922e-02,-3.01927839e+00,3.66311343e-02,1.13899497e-01,
-9.24659108e-03,2.08472233e-02,6.99173508e-01,3.40709987e-03,2.41138017e-03]
age = (p[0]+
p[1]*x1+p[2]*x2+p[3]*x3+
p[4]*np.multiply(x1,x2)+p[5]*np.multiply(x1,x3)+
p[6]*(x1**2)+p[7]*(x2**2)+p[8]*(x3**2)+
p[9]*(x1**3)+p[10]*(x2**3))
return age
def ll_age(x1,x2,x3):
#'x1' 'x2' 'x3' 'x12' 'x13' 'x23' 'x1sq' 'x2sq' 'x3sq' 'x1cu' 'x2cu' 'x3cu'
p = [9.62345764e+00,-4.30768983e-02,-8.06826028e-02,-3.01563712e+00,3.65508436e-02,1.17097905e-01,
-2.59253472e-02,-8.98018594e-03,2.10598963e-02,7.16404583e-01,3.32627539e-03,1.34864314e-03,
-3.07136600e-02]
age = (p[0]+
p[1]*x1+p[2]*x2+p[3]*x3+
p[4]*np.multiply(x1,x2)+p[5]*np.multiply(x1,x3)+p[6]*np.multiply(x2,x3)+
p[7]*(x1**2)+p[8]*(x2**2)+p[9]*(x3**2)+
p[10]*(x1**3)+p[11]*(x2**3)+p[12]*(x3**3))
return age
def max_resid_age(x1,x2,x3):
#'x1' 'x2' 'x3' 'x12' 'x13' 'x1sq' 'x2sq' 'x3cu'
p = [9.62671618e+00,-3.76516348e-02,-7.73334895e-02,-3.03980891e+00,3.45423253e-02,8.77085003e-02,
-6.14819060e-03,2.43493988e-02,1.29436784e+00]
age = (p[0]+
p[1]*x1+p[2]*x2+p[3]*x3+
p[4]*np.multiply(x1,x2)+p[5]*np.multiply(x1,x3)+
p[6]*(x1**2)+p[7]*(x2**2)+
p[8]*(x3**3))
return age
def current_age(x1,x2,x3):
#'x1' 'x2' 'x3' 'x12' 'x13' 'x23' 'x1sq' 'x2sq' 'x3sq' 'x2cu' 'x3cu'
p = [9.62160564e+00,-3.73279863e-02,-8.03408138e-02,-3.01132874e+00,3.47946523e-02,1.07568186e-01,
-2.82038911e-02,-6.71667916e-03,2.32342481e-02,7.17749242e-01,5.41993609e-04,-3.22534784e-02]
age = (p[0]+
p[1]*x1+p[2]*x2+p[3]*x3+
p[4]*np.multiply(x1,x2)+p[5]*np.multiply(x1,x3)+p[6]*np.multiply(x2,x3)+
p[7]*(x1**2)+p[8]*(x2**2)+p[9]*(x3**2)+
p[10]*(x2**3)+p[11]*(x3**3))
return age
# +
### Uncertainty Functions
def add_noise(quant,quant_err,distribution='normal'):
'''
Add noise to data and return new values
Parameters:
----------
quant: 1d array-like data to add noise to
quant_err: 1d array-like object of errors for quant
distribution: which distribution to use 'normal', 'poisson', 'uniform'
return:
------
1d array-like object of data with added noise
'''
if distribution == 'normal':
return np.random.normal(quant,quant_err)
if distribution == 'poisson':
return quant + np.random.poisson(quant_err)
if distribution == 'uniform':
return np.random.uniform(-quant_err+quant,quant+quant_err)
# +
mad_age_dist = []
ll_age_dist = []
max_resid_age_dist = []
current_age_dist = []
pca_transform = PCA(n_components=3).fit(data)
for i in tqdm_notebook(range(len(lmc)),desc='Done?'):
# temperature
teff = lmc['TEFF'][i]
teff_err = lmc['TEFF_ERR'][i]
# photometry
ra = lmc['RA'][i]
dec = lmc['DEC'][i]
ks = lmc['K'][i]
ks_err = lmc['K_ERR'][i]
ak = lmc['AK_TARG'][i]
# abundances
mh = lmc['M_H'][i]
mh_err = lmc['M_H_ERR'][i]
cfe = lmc['C_FE'][i]
cfe_err = lmc['C_FE_ERR'][i]
nfe = lmc['N_FE'][i]
nfe_err = lmc['N_FE_ERR'][i]
feh = lmc['FE_H'][i]
feh_err = lmc['FE_H_ERR'][i]
aM = lmc['ALPHA_M'][i]
aM_err = lmc['ALPHA_M_ERR'][i]
# surface gravity
lgg = lmc['LOGG'][i]
lgg_err = lmc['LOGG_ERR'][i]
noise_mad_ages = []
noise_ll_ages = []
noise_max_resid_ages = []
noise_current_ages = []
new = []
for j in range(750):
# calculate noisy temperature
teff_new = add_noise(teff,teff_err)
lgteff_new = np.log10(teff_new)
# Calculate noisy K magnitude
_, _, dist = afunc.LMCdisk_cart(ra,dec)
ks_new = add_noise(ks,ks_err)
absK_new = afunc.absmag(ks_new,dist) - ak
# Calculate noisy Salaris correction
x_C_new = sol_xm[0]*10**(add_noise(cfe,cfe_err)+
add_noise(feh,feh_err)-add_noise(mh,mh_err))
x_N_new = sol_xm[1]*10**(add_noise(nfe,nfe_err)+
add_noise(feh,feh_err)-add_noise(mh,mh_err))
ffac_new = (x_C_new+x_N_new)/sum(sol_xm[0:2])
aM_new = add_noise(aM,aM_err)
mh_new = add_noise(mh,mh_err)
# Calculate noisy surface gravity
lgg_new = add_noise(lgg,lgg_err)
# calculate noisy age
new.append(np.array([lgteff_new,absK_new,afunc.sal(mh_new,aM_new),lgg_new]))
xs = pca_transform.transform(np.asarray(np.squeeze(new)))
for l in range(len(xs)):
noise_mad_ages.append(mad_age(xs[l][0],xs[l][1],xs[l][2]))
noise_ll_ages.append(ll_age(xs[l][0],xs[l][1],xs[l][2]))
noise_max_resid_ages.append(max_resid_age(xs[l][0],xs[l][1],xs[l][2]))
noise_current_ages.append(current_age(xs[l][0],xs[l][1],xs[l][2]))
mad_age_dist.append(noise_mad_ages)
ll_age_dist.append(noise_ll_ages)
max_resid_age_dist.append(noise_max_resid_ages)
current_age_dist.append(noise_current_ages)
# +
mad_parages = []
ll_parages = []
max_resid_parages = []
current_parages = []
mad_devs = []
ll_devs = []
max_resid_devs = []
current_devs = []
for k in tqdm_notebook(range(len(mad_age_dist))):
mad_devs.append(afunc.mad(mad_age_dist[k]))
mad_parages.append(np.median(mad_age_dist[k]))
ll_devs.append(afunc.mad(ll_age_dist[k]))
ll_parages.append(np.median(ll_age_dist[k]))
max_resid_devs.append(afunc.mad(max_resid_age_dist[k]))
max_resid_parages.append(np.median(max_resid_age_dist[k]))
current_devs.append(afunc.mad(current_age_dist[k]))
current_parages.append(np.median(current_age_dist[k]))
### MAD
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(mad_parages)[lmc_ind],10**pdfout['P_MEAN'][pdfout_ind],c=lmc['LOGG'][lmc_ind],
marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.xscale('log')
plt.yscale('log')
plt.plot(10**np.array([7,8,9,10,11,12,13]),10**np.array([7,8,9,10,11,12,13]),c='k')
plt.ylim(10**7,10**11)
plt.xlim(10**7,10**13)
plt.xlabel(r'Polynomial Interpolation')
plt.ylabel(r'Bayesian')
plt.show()
## #ll
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(ll_parages)[lmc_ind],10**pdfout['P_MEAN'][pdfout_ind],c=lmc['LOGG'][lmc_ind],
marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.xscale('log')
plt.yscale('log')
plt.plot(10**np.array([7,8,9,10,11,12,13]),10**np.array([7,8,9,10,11,12,13]),c='k')
plt.ylim(10**7,10**11)
plt.xlim(10**7,10**13)
plt.xlabel(r'Polynomial Interpolation')
plt.ylabel(r'Bayesian')
plt.show()
### max_resid
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(max_resid_parages)[lmc_ind],10**pdfout['P_MEAN'][pdfout_ind],c=lmc['LOGG'][lmc_ind],
marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.xscale('log')
plt.yscale('log')
plt.plot(10**np.array([7,8,9,10,11,12,13]),10**np.array([7,8,9,10,11,12,13]),c='k')
plt.ylim(10**7,10**11)
plt.xlim(10**7,10**13)
plt.xlabel(r'Polynomial Interpolation')
plt.ylabel(r'Bayesian')
plt.show()
### current
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(current_parages)[lmc_ind],10**pdfout['P_MEAN'][pdfout_ind],c=lmc['LOGG'][lmc_ind],
marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.xscale('log')
plt.yscale('log')
plt.plot(10**np.array([7,8,9,10,11,12,13]),10**np.array([7,8,9,10,11,12,13]),c='k')
plt.ylim(10**7,10**11)
plt.xlim(10**7,10**13)
plt.xlabel(r'Polynomial Interpolation')
plt.ylabel(r'Bayesian')
plt.show()
# +
plt.figure(figsize=[12,10])
plt.scatter(range(len(np.asarray(mad_parages)[r13_ind])),
np.asarray(mad_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.scatter(range(len(np.asarray(ll_parages)[r13_ind])),
np.asarray(ll_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.scatter(range(len(np.asarray(max_resid_parages)[r13_ind])),
np.asarray(max_resid_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.scatter(range(len(np.asarray(current_parages)[r13_ind])),
np.asarray(current_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.ylim(-1,2)
plt.ylabel(r'Polynomial - Bayesian')
plt.show()
plt.figure(figsize=[12,10])
plt.scatter(range(len(np.asarray(mad_parages)[r13_ind])),
np.asarray(mad_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:blue')
plt.scatter(range(len(np.asarray(ll_parages)[r13_ind])),
np.asarray(ll_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:orange')
plt.scatter(range(len(np.asarray(max_resid_parages)[r13_ind])),
np.asarray(max_resid_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:green')
plt.scatter(range(len(np.asarray(current_parages)[r13_ind])),
np.asarray(current_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:red')
plt.ylim(-0.5,0.5)
plt.ylabel(r'Polynomial - Bayesian')
plt.show()
print(np.median(np.asarray(mad_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind]))
print(np.median(np.asarray(ll_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind]))
print(np.median(np.asarray(max_resid_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind]))
print(np.median(np.asarray(current_parages)[r13_ind]-pdfout['P_MEAN'][pdfout_ind]))
# +
### MAD
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(mad_parages),mad_devs,c=lmc['LOGG'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(mad_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'MAD; Color as LOGG')
plt.xlabel(r'Lowest MAD Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### ll
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(ll_parages),ll_devs,c=lmc['LOGG'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(ll_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'll; Color as LOGG')
plt.xlabel(r'Lowest ll Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### max_resid
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(max_resid_parages),max_resid_devs,c=lmc['LOGG'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(max_resid_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.ylim(top=0.215)
plt.title(r'max_resid; Color as LOGG')
plt.xlabel(r'Lowest max resid Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### current
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(current_parages),current_devs,c=lmc['LOGG'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(current_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'Current; Color as LOGG')
plt.xlabel(r'Lowest current Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
# +
### MAD
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(mad_parages),mad_devs,c=lmc['LOGG_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(mad_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'MAD; Color as LOGG_ERR')
plt.xlabel(r'Lowest MAD Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### ll
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(ll_parages),ll_devs,c=lmc['LOGG_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(ll_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'll; Color as LOGG_ERR')
plt.xlabel(r'Lowest ll Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### max_resid
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(max_resid_parages),max_resid_devs,c=lmc['LOGG_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(max_resid_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.ylim(top=0.215)
plt.title(r'max_resid; Color as LOGG_ERR')
plt.xlabel(r'Lowest max resid Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### current
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(current_parages),current_devs,c=lmc['LOGG_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(current_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'Current; Color as LOGG_ERR')
plt.xlabel(r'Lowest current Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
# +
### MAD
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(mad_parages),mad_devs,c=lmc['FE_H'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(mad_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'MAD; Color as FE_H')
plt.xlabel(r'Lowest MAD Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### ll
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(ll_parages),ll_devs,c=lmc['FE_H'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(ll_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'll; Color as FE_H')
plt.xlabel(r'Lowest ll Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### max_resid
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(max_resid_parages),max_resid_devs,c=lmc['FE_H'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(max_resid_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.ylim(top=0.215)
plt.title(r'max_resid; Color as FE_H')
plt.xlabel(r'Lowest max resid Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### current
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(current_parages),current_devs,c=lmc['FE_H'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(current_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'Current; Color as FE_H')
plt.xlabel(r'Lowest current Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
# +
### MAD
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(mad_parages),mad_devs,c=lmc['FE_H_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(mad_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'MAD; Color as FE_H_ERR')
plt.xlabel(r'Lowest MAD Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### ll
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(ll_parages),ll_devs,c=lmc['FE_H_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(ll_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'll; Color as FE_H_ERR')
plt.xlabel(r'Lowest ll Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### max_resid
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(max_resid_parages),max_resid_devs,c=lmc['FE_H_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(max_resid_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.ylim(top=0.215)
plt.title(r'max_resid; Color as FE_H_ERR')
plt.xlabel(r'Lowest max resid Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
### current
plt.figure(figsize=[12,10])
plt.scatter(10**np.asarray(current_parages),current_devs,c=lmc['FE_H_ERR'],marker='+',cmap='nipy_spectral')
plt.colorbar()
plt.axvline(ageU)
plt.axvline(10**(np.log10(ageU)+np.median(current_devs)),c='r')
plt.xscale('log')
plt.xlim(10**7,10**13)
plt.title(r'Current; Color as FE_H_ERR')
plt.xlabel(r'Lowest current Age')
plt.ylabel(r'Log(Age) Error')
plt.show()
# +
######################
### Residual Plots ###
######################
# +
cat = Table()
col_id = Column(np.asarray(r13['APOGEE_ID']),name='APOGEE_ID')
col_mad_age = Column(np.asarray(mad_parages),name='MAD_AGE')
col_mad_dev = Column(np.asarray(mad_devs),name='MAD_AGE_ERR')
col_ll_age = Column(np.asarray(ll_parages),name='LL_AGE')
col_ll_dev = Column(np.asarray(ll_devs),name='LL_AGE_ERR')
col_max_resid_age = Column(np.asarray(max_resid_parages),name='MAX_RESID_AGE')
col_max_resid_dev = Column(np.asarray(max_resid_devs),name='MAX_RESID_AGE_ERR')
col_current_age = Column(np.asarray(current_parages),name='CURRENT_AGE')
col_current_dev = Column(np.asarray(current_devs),name='CURRENT_AGE_ERR')
cat.add_column(col_id)
cat.add_column(col_mad_age)
cat.add_column(col_mad_dev)
cat.add_column(col_ll_age)
cat.add_column(col_ll_dev)
cat.add_column(col_max_resid_age)
cat.add_column(col_max_resid_dev)
cat.add_column(col_current_age)
cat.add_column(col_current_dev)
# cat.write('pca_ols_loggs.fits',format='fits',overwrite=True)
# +
plt.figure(figsize=[12,10])
plt.scatter(range(len(np.asarray(mad_parages)[r13_ind])),
10**np.asarray(mad_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.scatter(range(len(np.asarray(ll_parages)[r13_ind])),
10**np.asarray(ll_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.scatter(range(len(np.asarray(max_resid_parages)[r13_ind])),
10**np.asarray(max_resid_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.scatter(range(len(np.asarray(current_parages)[r13_ind])),
10**np.asarray(current_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],c=r13['LOGG'][r13_ind],
marker='+',cmap='nipy_spectral')
plt.colorbar()
# plt.ylim(-1,2)
plt.ylabel(r'Polynomial - Bayesian')
plt.show()
plt.figure(figsize=[12,10])
plt.scatter(range(len(np.asarray(mad_parages)[r13_ind])),
10**np.asarray(mad_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:blue')
plt.scatter(range(len(np.asarray(ll_parages)[r13_ind])),
10**np.asarray(ll_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:orange')
plt.scatter(range(len(np.asarray(max_resid_parages)[r13_ind])),
10**np.asarray(max_resid_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:green')
plt.scatter(range(len(np.asarray(current_parages)[r13_ind])),
10**np.asarray(current_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind],
marker='+',c='tab:red')
plt.ylim(0,20*10**9)
plt.ylabel(r'Polynomial - Bayesian')
plt.show()
print(np.median(10**np.asarray(mad_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind])/10**9)
print(np.median(10**np.asarray(ll_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind])/10**9)
print(np.median(10**np.asarray(max_resid_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind])/10**9)
print(np.median(10**np.asarray(current_parages)[r13_ind]-10**pdfout['P_MEAN'][pdfout_ind])/10**9)
# -
parsec.columns
plt.hist(parsec['Xo'])
sol_xm[2]
parsec.columns
| 26,066 |
/jupyter_notebooks/Websim.ipynb | c5b29a284eb96387d8623f48bda4b3a2928d4296 | [] | no_license | d07s1d0s4d1/alphatron | https://github.com/d07s1d0s4d1/alphatron | 1 | 1 | null | null | null | null | Jupyter Notebook | false | false | .py | 13,530 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import json
import time
import logging
class WebSim(object):
'''
Example:
ws = websim.WebSim(login = 'gosha-bor@yandex.ru', password = '...')
ws.authorise()
ws.send_alpha('-returns')
'''
def __init__(self, login = None, password = None, UID = None, WSSID = None, account_id = None):
self.login = login
self.password = password
self.UID = UID
self.WSSID = WSSID
self.account_id = account_id
self.headers = {
'Host': 'websim.worldquantchallenge.com',
'User-Agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10.11; rv:52.0) Gecko/20100101 Firefox/52.0',
'Accept': 'application/json',
'Accept-Language':'en-US,en;q=0.5',
'Cache-Control': 'max-age=0',
'Content-Type': 'application/x-www-form-urlencoded',
'Origin': 'https://websim.worldquantchallenge.com',
'Referer': 'https://websim.worldquantchallenge.com/result?q=9190149',
'Connection': 'keep-alive',
}
def send_alpha(self, code, delay = "0", decay = 10, opneut = "subindustry", optrunc = "0.1", univid = "TOP3000", backdays = 256):
cookies = {
'django_language': 'en',
# 'sessionid': 'p50gmiem9hykheowx9vqp0o2k15ls3gh',
# '_xsrf': '2|c1d41832|6d1377ea652ff73e906cd52084aa5255|1491915833',
# 'UID': 'd6f1c3b48fe0f907e06f3e4555c2a5af1c23c56e4ee7f4a9e84f864d13d1955e',
# 'WSSID':'"2|1:0|10:1507563160|5:WSSID|60:RlNXbzRKTDFIdEc2OStJMzExR05WVHBPK3p6UEI1Z0IxKzdGb3pCSjNnMD0=|7f9c94134a9b5b4852f082537e50c91faed0fea4a801491017006bbe01ac5098"',
'UID': self.UID,
'WSSID': self.WSSID,
}
# data = json.dump()
r = requests.post('https://websim.worldquantchallenge.com/simulate',
headers = self.headers,
cookies = cookies,
data = 'args=' + json.dumps([{
'delay': delay,
'unitcheck': 'off',
'univid': univid,
'opcodetype': 'EXPRESSION',
'opassetclass': 'EQUITY',
'optrunc': optrunc,
'code': code,
'region': 'USA',
'opneut': opneut,
#'IntradayType':'null',
'tags': 'equity',
'decay': decay,
'DataViz': 0,
'backdays': backdays,
'simtime': 'Y5',
}])
)
# 'IntradayType': 'null',
# print r
print('\nsimulate...')
print(r.text)
job = json.loads(r.text)
if not job['status']:
raise ValueError(job)
job_id = job['result'][0]
# print 'sending all'
r = requests.post('https://websim.worldquantchallenge.com/simulate/settings/all',
headers = self.headers,
cookies = cookies,
)
print('\nsettings/all:')
print(r.text)
all_ = json.loads(r.text)
# print 'details'
r = requests.post('https://websim.worldquantchallenge.com/job/details/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
print('\ndetails:')
print(r.text)
details = json.loads(r.text)
while True:
time.sleep(3)
print('\nrequest progress...')
r = requests.post('https://websim.worldquantchallenge.com/job/progress/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
print(r.text)
if r.text == "\"DONE\"":
break
if r.text == "\"ERROR\"":
#raise ValueError('Got ERROR in process request')
return 'ERROR'
# print 'details after done'
r = requests.post('https://websim.worldquantchallenge.com/job/details/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
# print r
# print r.text
details2 = json.loads(r.text)
alphaID = details2['result']['clientAlphaId']
r = requests.post('https://websim.worldquantchallenge.com/job/pnlchart/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
pnl_chart = json.loads(r.text)
r = requests.post('https://websim.worldquantchallenge.com/job/grade/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
grade = json.loads(r.text)
r = requests.post('https://websim.worldquantchallenge.com/job/simsummary/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
simsummary = json.loads(r.text)
r = requests.post('https://websim.worldquantchallenge.com/job/simsummary/{}'.format(job_id),
headers = self.headers,
cookies = cookies,
)
simsummary = json.loads(r.text)
return {
'alphaID': alphaID,
'grade': grade,
'pnl_chart': pnl_chart,
'simsummary': simsummary,
}
def authorise(self):
cookies = {
}
r = requests.post('https://websim.worldquantchallenge.com/login/process',
headers = self.headers,
cookies = cookies,
data = {
'EmailAddress': self.login,
'Password': self.password,
'next':'/dashboard',
'g-recaptcha-response': '',
}
)
print(r)
print(r.text)
#print(r.cookies)
cookies_ = r.cookies.get_dict(domain = '.worldquantchallenge.com')
#print r.cookies
print("COOKIES:", cookies_)
self.UID = cookies_['UID']
self.WSSID = cookies_['WSSID']
# -
ws = WebSim(login = 'dsd_kem@mail.ru', password = 'NBSn6ZNe#1')
ws.authorise()
ws.send_alpha(code = '(rank((vwap-close))/rank((vwap+close)))')
ws.correlation(['04fdf7f95bd64e0eb0c0e32b913ddf2a','c6d760aac3ba434c8dbc63536c8c1ab4'],'inner_corr')
| 6,465 |
/notebooks/AttentiveLSTMs.ipynb | 87b60ec79b627b4a60f93cb1ec1d76eaa6f81bd4 | [] | no_license | FASLADODO/SentimentClassification_TreeLSTM | https://github.com/FASLADODO/SentimentClassification_TreeLSTM | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 165,286 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data science in Python
#
# - Course GitHub repo: https://github.com/pycam/python-data-science
# - Python website: https://www.python.org/
#
# ## Session 2.1: Working with Pandas
#
# - [Reading CSV Data Using Pandas](#Reading-CSV-Data-Using-Pandas)
# - [Exploring our data](#Exploring-our-data)
# - [DataFrame manipulation](#DataFrame-manipulation)
# - [Exercise 2.1.1](#Exercise-2.1.1)
# ## Mind map
#
# <img src="img/mind_maps/mind_maps.004.jpeg">
# ## Reading CSV Data Using Pandas
#
# ### Importing the `pandas` package
#
# [`pandas`](http://pandas.pydata.org/) is a widely-used external Python package for data analysis, particularly useful for tabular data (see [documentation](http://pandas.pydata.org/pandas-docs/stable/)).
#
# The `pandas` DataFrame object borrows many features from R's `data.frame`. Both are 2-dimentional tables whose columns can contain different data types (e.g. boolean, integer, float, categorical/factor). Both the rows and columns are indexed, and can be referred to by number or name.
#
# Because `pandas` is an external third-party package, it is not included in Python by default. You can install it using `pip install pandas`, or if using conda, `conda install pandas`.
#
# Once installed, we load the package with the `import` command:
import pandas
# ### Reading CSV data
#
# To read a Comma Separated Values (CSV) data file with `pandas`, we use the `.read_csv()` command.
#
# We are going to load a slightly different Gapminder dataset for Oceania, where each column represents the GDP per capita for different years and each row represents a country in Oceania. In this case, our file is in a sub-directory called `data/`, and the name of the file is `gapminder_gdp_oceania.csv`.
#
# We store the resulting DataFrame object and give it the variable name `data`:
data = pandas.read_csv('data/gapminder_gdp_oceania.csv')
print(data)
# If you forget to include `data/`, or if you include it but your copy of the file is saved somewhere else, you will get a runtime error that ends with a line like this:
# ```
# FileNotFoundError: File b'gapminder_gdp_oceania.csv' does not exist
# ```
# Note that `pandas` uses a backslash `\` to show wrapped lines when the output is too wide to fit on the screen. Looking at the index column on the far left, you can see that our DataFrame has two rows (automatically 0 and 1, since Python uses 0-based indexing).
#
# Generally, columns in a DataFrame are the observed variables, and the rows are the observations.
#
# Instead of treating the `'country'` column like any other observed variable, we can tell `pandas` to use this column as the row index. We can then refer to the rows by country name:
data = pandas.read_csv('data/gapminder_gdp_oceania.csv', index_col='country')
print(data)
# Note that the `.read_csv()` command is not limited to reading CSV files. For example, you can read Tab Separated Value (TSV) files by adding the argument `sep='\t'`.
# ## Exploring our data
#
# A `pandas` DataFrame is a 2-dimensional object that can store columns with different data types (including string, boolean, integer, float, categorical/factor, etc). It is similar to a spreadsheet or an SQL table or the `data.frame` structure in R.
#
# A DataFrame always has an index (0-based). An index refers to the position of an element in the data structure. Using the `.info()` method, we can view basic information about our DataFrame object:
data.info()
# As expected, our object is a `DataFrame` (or, to use the full name that Python uses to refer to it internally, a `pandas.core.frame.DataFrame`).
#
# It has 2 rows (named Australia and New Zealand) and 12 columns. The columns consist of 64-bit floating point values. It uses about 200 bytes of memory.
# As mentioned, a DataFrame is a Python object, which means it can have **object attributes** and **object methods**.
#
# **Object attributes** contain information about the object. You can access [DataFrame attributes](https://pandas.pydata.org/pandas-docs/stable/reference/series.html#attributes) to learn more about the contents of your DataFrame. To do this, use the object variable name followed by the attribute name, separated by a `.`. Do not use any `()` to access attributes.
#
# For example, let's access our column data types, which are stored in the `.dtypes` attribute:
data.dtypes
# You can access the dimensions of your DataFrame using the `.shape` attribute. The first value is the number of rows, and the second the number of columns:
data.shape
# The row and column names can be accessed using the attributes `.index.values` and `.columns.values`:
data.index.values
data.columns.values
# **Object methods** are functions that are associated with an object. Because they are functions, you do use `()` to call them, and can add arguments inside the parentheses to control the behaviour of the method. For example, the `.info()` command we executed previously was a method.
#
# Let's now download the European Gapminder data and explore a few of these (see the [DataFrame API](https://pandas.pydata.org/pandas-docs/stable/reference/index.html) for more).
eu_data = pandas.read_csv('data/gapminder_gdp_europe.csv', index_col='country')
print(eu_data)
eu_data.info()
# The `.head()` method prints the first few rows of the table, while the `.tail()` method prints the last few lines:
eu_data.head()
eu_data.head(3)
eu_data.tail(2)
# The `.describe` method computes some summary statistics for the columns (including the count, mean, median, and std):
eu_data.describe()
# We often want to calculate summary statistics grouped by subsets or attributes within fields of our data. For example, we might want to calculate the average GDP per capita for 1962.
#
# There are two ways to access columns in a DataFrame. The first is using a `.` followed by the name of the column, while the second is using square brackets:
eu_data.gdpPercap_1962.head()
eu_data['gdpPercap_1962'].head()
# We can also compute metrics on specific columns:
eu_data['gdpPercap_1962'].mean()
eu_data['gdpPercap_1962'].std()
# Note: these statistics can only be computed on numeric columns. They can be particularly useful when your DataFrame contains thousands of entries.
# ## DataFrame manipulation
#
# ### Pandas Cheat Sheet
#
# The [Pandas Cheat Sheet](http://pandas.pydata.org/Pandas_Cheat_Sheet.pdf) can be very helpful for recalling basic `pandas` operations.
#
# Let's start by creating a small DataFrame object and storing it using the variable name `df`. One way to do this is from a Python dictionary:
import pandas
df = pandas.DataFrame({'gene': ['BRCA2', 'TNFAIP3', 'TCF7'],
'chrom': ['13', '6', '5'],
'length': [84195, 16099, 37155]}
)
print(df)
# ### Selecting columns and rows
# To select rows and columns in `pandas`, we use square brackets `[]`. There are several ways to select or "slice" a DataFrame objects. It's important to distinguish between **positional indexing**, which uses index numbers, and **label-based indexing** which uses column or row names.
#
# Let's start by selecting one column, using its name:
df['length']
# Now we select the first two rows using their numeric index:
df[:2]
# We can combine these to get the first 2 rows of the column `'length'`:
df[:2]['length']
# To do **positional indexing** for both rows and columns, use `.iloc[]`. The first argument is the numeric index of the rows, and the second the numeric index of the columns:
df.iloc[:2]
df.iloc[:2,2]
# We can also select the first 2 rows, and all of the columns from index 1 until the end, as follows:
df.iloc[:2,1:]
# For **label-based indexing**, use `.loc[]` with the column and row names:
df.loc[:1,'chrom':]
# Note: because the rows have numeric indices in this DataFrame, we might assume that selecting rows with `.iloc[]` and `.loc[]` would be the same. As we see below, this is not the case.
# ### Sorting columns
# To sort the entire DataFrame according to one of the columns, we can use the `.sort_values()` method. Note that this method returns a DataFrame object, so we need to store that object using a new variable name such as `sorted_df`:
sorted_df = df.sort_values('length')
print(sorted_df)
# Going back to positional versus label-based indexing, we see that `.iloc[0]` will return the first row in the DataFrame, while `.loc[0]` will return the row with index 0:
sorted_df.iloc[0]
sorted_df.loc[0]
# We can also sort the DataFrame in descending order:
sorted_df = df.sort_values('length', ascending=False)
print(sorted_df)
# If you would like to reset the row indices after sorting, use `.reset_index()` and store the result in a new variable. Adding the argument `drop=True` will prevent `pandas` from adding the old index as a column:
sorted_df_reindexed = sorted_df.reset_index(drop=True)
print(sorted_df_reindexed)
# ## Exercise 2.1.1
#
# - Read the data in `gapminder_gdp_americas.csv` (which should be in the same directory as `gapminder_gdp_oceania.csv`) into a variable called `americas_data` and display its summary statistics.
# - As well as the `.read_csv()` function for reading data from a file, Pandas provides a `.to_csv()` function to write DataFrames to files. Applying what you’ve learned about reading from files, write one of your DataFrame to a file called `processed.csv`. You can use help to get information on how to use `.to_csv()`.
# ## Manipulating data with Pandas (live coding session)
#
# Let's now open a new Jupyter notebook, and explore another dataset `GRCm38.gff3` from the `data/` folder.
#
# [GFF is a standard file format](http://gmod.org/wiki/GFF3) for storing genomic features in a text file. GFF stands for Generic Feature Format. GFF files are plain text, 9 column, tab-delimited files.
#
# The 9 columns of the annotation section are as follows:
#
# - Column 1: "seqid" - The ID of the landmark used to establish the coordinate system for the current feature.
# - Column 2: "source" - The source is a free text qualifier intended to describe the algorithm or operating procedure that generated this feature.
# - Column 3: "type" - The type of the feature.
# - Columns 4 & 5: "start" and "end" - The start and end of the feature.
# - Column 6: "score" - The score of the feature, a floating point number.
# - Column 7: "strand" - The strand of the feature.
# - Column 8: "phase" - For features of type "CDS", the phase indicates where the feature begins with reference to the reading frame.
# - Column 9: "attributes" - A list of feature attributes in the format tag=value.
#
# We have modified these files and added a 10th column "gbid" which is the GenBank ID of each feature, and taken a random subset of these features for each species.
# ## Next session
#
# Go to our next notebook: [Session 2.2: Data visualisation with Matplotlib](22_python_data.ipynb)
olab only.*
# + colab={} colab_type="code" id="qKQMGtkR5KWr"
# http://pytorch.org/
from os.path import exists
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
import torch
from torch import optim
from torch import nn
# + colab={} colab_type="code" id="BhiRqhTM5V4c"
# this should result in the PyTorch version
torch.__version__
# + colab={} colab_type="code" id="BYt8uTyGCKc7"
# PyTorch can run on CPU or on Nvidia GPU (video card) using CUDA
# This cell selects the GPU if one is available.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device
# + colab={} colab_type="code" id="2d1VMOOYx1Bw"
# Seed manually to make runs reproducible
# You need to set this again if you do multiple runs of the same model
torch.manual_seed(0)
# When running on the CuDNN backend two further options must be set for reproducibility
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# + [markdown] colab_type="text" id="uWBTzkuE3CtZ"
# # BOW
# + [markdown] colab_type="text" id="TBAjYYySOA5W"
# Our first model is a super simple neural **bag-of-words (BOW) model**.
# Unlike the bag-of-words model that you used in the previous lab, here we associate each word with a vector of size 5, exactly our number of sentiment classes.
#
# To make a classification, we simply **sum** the vectors of the words in our sentence and a bias vector. Because we sum the vectors, we lose word order: that's why we call this a neural bag-of-words model.
#
# ```
# this [0.0, 0.1, 0.1, 0.1, 0.0]
# movie [0.1, 0.1, 0.1, 0.2, 0.1]
# is [0.0, 0.1, 0.0, 0.0, 0.0]
# stupid [0.9, 0.5, 0.1, 0.0, 0.0]
#
# bias [0.0, 0.0, 0.0, 0.0, 0.0]
# --------------------------------
# sum [0.9, 0.8, 0.3, 0.3, 0.1]
#
# argmax: 0 (very negative)
# ```
#
# Now, the **argmax** of this sum is our predicted label.
#
# We initialize all vectors *randomly* and train them using cross-entropy loss.
# + [markdown] colab_type="text" id="rLtBAIQGynkB"
# #### Model class
# + colab={} colab_type="code" id="QZfNklWf3tvs"
class BOW(nn.Module):
"""A simple bag-of-words model"""
def __init__(self, vocab_size, embedding_dim, vocab):
super(BOW, self).__init__()
self.vocab = vocab
# this is a trained look-up table with word embeddings
self.embed = nn.Embedding(vocab_size, embedding_dim)
# this is a trained bias term
self.bias = nn.Parameter(torch.zeros(embedding_dim), requires_grad=True)
def forward(self, inputs):
# this is the forward pass of the neural network
# given inputs, it computes the output
# this looks up the embeddings for each word ID in inputs
# the result is a sequence of word embeddings
embeds = self.embed(inputs)
# the output is the sum across the time dimension (1)
# with the bias term added
logits = embeds.sum(1) + self.bias
return logits
# + colab={} colab_type="code" id="eKHvBnoBAr6z"
# Let's create a model.
vocab_size = len(v.w2i)
n_classes = len(t2i)
bow_model = BOW(vocab_size, n_classes, v)
print(bow_model)
# + [markdown] colab_type="text" id="vfCx-HvMH1qQ"
# > **Hey, wait, where is the bias vector?**
# > PyTorch does not print Parameters, only Modules!
#
# > We can print it ourselves though, to check that it is there.
# + colab={} colab_type="code" id="Fhvk5HenAroT"
# Here we print each parameter name, shape, and if it is trainable.
def print_parameters(model):
total = 0
for name, p in model.named_parameters():
total += np.prod(p.shape)
print("{:24s} {:12s} requires_grad={}".format(name, str(list(p.shape)), p.requires_grad))
print("\nTotal parameters: {}\n".format(total))
print_parameters(bow_model)
# + [markdown] colab_type="text" id="WSAw292WxuP4"
# #### Preparing an example for input
#
# To feed sentences to our PyTorch model, we need to convert a sequence of tokens to a sequence of IDs. The `prepare_example` function below takes care of this for us. With these IDs we index the word embedding table.
# + colab={} colab_type="code" id="YWeGTC_OGReV"
def prepare_example(example, vocab):
"""
Map tokens to their IDs for 1 example
"""
# vocab returns 0 if the word is not there
x = [vocab.w2i.get(t, 0) for t in example.tokens]
x = torch.LongTensor([x])
x = x.to(device)
y = torch.LongTensor([example.label])
y = y.to(device)
return x, y
# + [markdown] colab_type="text" id="oKNQjEc0yXnJ"
# #### Evaluation
# We will need one more thing: an evaluation metric.
# How many predictions do we get right? The accuracy will tell us.
# Make sure that you understand this code block.
#
# + colab={} colab_type="code" id="yGmQLcVYKZsh"
def simple_evaluate(model, data, prep_fn=prepare_example, **kwargs):
"""Accuracy of a model on given data set."""
correct = 0
total = 0
model.eval() # disable dropout (explained later)
for example in data:
# convert the example input and label to PyTorch tensors
x, target = prep_fn(example, model.vocab)
# forward pass
# get the output from the neural network for input x
with torch.no_grad():
logits = model(x)
# get out the prediction
prediction = logits.argmax(dim=-1)
# add the number of correct predictions to the total correct
correct += (prediction == target).sum().item()
total += 1
return correct, total, correct / float(total)
# + [markdown] colab_type="text" id="dIk6OtSdzGRP"
# #### Example feed
# For stochastic gradient descent (SGD) we will need a random training example for every update.
# We implement this by shuffling the training data and returning examples one by one using `yield`.
#
# Shuffling is optional so that we get to use this to get validation and test examples, too.
# + colab={} colab_type="code" id="dxDFOZLfCXvJ"
def get_examples(data, shuffle=True, **kwargs):
"""Shuffle data set and return 1 example at a time (until nothing left)"""
if shuffle:
print("Shuffling training data")
random.shuffle(data) # shuffle training data each epoch
for example in data:
yield example
# + [markdown] colab_type="text" id="g09SM8yb2cjx"
# #### Exercise: Training function
#
# Your task is now to complete the training loop below.
# Before you do so, please read the section about optimization.
# + [markdown] colab_type="text" id="TVfUukVdM_1c"
# **Optimization**
#
# As mentioned in the "Introduction to PyTorch" notebook, one of the perks of using PyTorch are its automatic differentiation abilities. We will use these to train our BOW model.
#
# We train our model by feeding it an input and performing **forward** pass, which results in an output for which we obtain a **loss** with our loss function.
# After the gradients are calculated in the **backward** pass, we can take a step on the loss surface towards more optimal parameter settings (e.g. gradient descent).
#
# The package we will use to do this optimization is [torch.optim](https://pytorch.org/docs/stable/optim.html). Besides implementations of stochastic gradient descent (SGD), this package also implements the optimization algorithm Adam, which we'll be using in this practical.
# For the purposes of this assignment you do not need to know what Adam does besides that it uses gradient information to update our model parameters by calling:
#
# ```
# optimizer.step()
# ```
# Remember when we updated our parameters in the PyTorch tutorial in a loop?
#
#
# ```python
# # update weights
# learning_rate = 0.5
# for f in net.parameters():
# # for each parameter, take a small step in the opposite dir of the gradient
# p.data = p.data - p.grad.data * learning_rate
#
# ```
# The function call optimizer.step() does effectively the same thing.
#
# *(If you want to know more about optimization algorithms using gradient information [this blog](http://ruder.io/optimizing-gradient-descent/.) gives a nice intuitive overview.)*
# + colab={} colab_type="code" id="ktFnKBux25lD"
def train_model(model, optimizer, num_iterations=10000,
print_every=1000, eval_every=1000,
batch_fn=get_examples,
prep_fn=prepare_example,
eval_fn=simple_evaluate,
batch_size=1, eval_batch_size=None):
"""Train a model."""
iter_i = 0
train_loss = 0.
print_num = 0
start = time.time()
criterion = nn.CrossEntropyLoss() # loss function
best_eval = 0.
best_iter = 0
# store train loss and validation accuracy during training
# so we can plot them afterwards
losses = []
accuracies = []
if eval_batch_size is None:
eval_batch_size = batch_size
while True: # when we run out of examples, shuffle and continue
for batch in batch_fn(train_data, batch_size=batch_size):
# forward pass
model.train()
x, targets = prep_fn(batch, model.vocab)
logits = model(x)
B = targets.size(0) # later we will use B examples per update
# compute cross-entropy loss (our criterion)
# note that the cross entropy loss function computes the softmax for us
loss = criterion(logits.view([B, -1]), targets.view(-1))
train_loss += loss.item()
# backward pass
# Tip: check the Introduction to PyTorch notebook.
# erase previous gradients
model.zero_grad()
loss.backward()
# update weights - take a small step in the opposite dir of the gradient
optimizer.step()
print_num += 1
iter_i += 1
# print info
if iter_i % print_every == 0:
print("Iter %r: loss=%.4f, time=%.2fs" %
(iter_i, train_loss, time.time()-start))
losses.append(train_loss)
print_num = 0
train_loss = 0.
# evaluate
if iter_i % eval_every == 0:
_, _, accuracy = eval_fn(model, dev_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
accuracies.append(accuracy)
print("iter %r: dev acc=%.4f" % (iter_i, accuracy))
# save best model parameters
if accuracy > best_eval:
print("new highscore")
best_eval = accuracy
best_iter = iter_i
path = "{}.pt".format(model.__class__.__name__)
ckpt = {
"state_dict": model.state_dict(),
"optimizer_state_dict": optimizer.state_dict(),
"best_eval": best_eval,
"best_iter": best_iter
}
torch.save(ckpt, path)
# done training
if iter_i == num_iterations:
print("Done training")
# evaluate on train, dev, and test with best model
print("Loading best model")
path = "{}.pt".format(model.__class__.__name__)
ckpt = torch.load(path)
model.load_state_dict(ckpt["state_dict"])
_, _, train_acc = eval_fn(
model, train_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
_, _, dev_acc = eval_fn(
model, dev_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
_, _, test_acc = eval_fn(
model, test_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
print("best model iter {:d}: "
"train acc={:.4f}, dev acc={:.4f}, test acc={:.4f}".format(
best_iter, train_acc, dev_acc, test_acc))
return losses, accuracies
# + [markdown] colab_type="text" id="XEPsLvI-3D5b"
# ### Training the BOW model
# + [markdown] colab_type="text" id="E9mB1_XhMPNN"
# # CBOW
# + [markdown] colab_type="text" id="pWk78FvNMw4o"
# We now continue with a **continuous bag-of-words (CBOW)** model.
# It is similar to the BOW model, but now embeddings can have a dimension of *arbitrary size*.
# This means that we can make them bigger and learn more aspects of each word. We will still sum the vectors to get a sentence representation, but now the result is no longer conveniently the number of sentiment classes.
#
# *Note that this is not the same as word2vec CBOW.* Both models take a few words as input, sum and make a prediction. However, in word2vec the prediction is the word in the middle of the input words. Here, the prediction is a sentiment class (1 out of 5 possible numbers 0-4).
#
# So what can we do? We can *learn* a parameter matrix $W$ that turns our summed vector into the number of output classes, by matrix-multiplying it: $$Wx$$
# If the size of $W$ is `5 x d`, and $x$ is `d x 1` and , then the result will be `5x1`, and then we can again find our prediction using an argmax.
# + [markdown] colab_type="text" id="gIjrCPfCwsXI"
# ## Exercise: write CBOW class & train it
#
# Write a class `CBOW` that:
#
# - has word embeddings with size 300
# - sums the word vectors for the input words (just like in `BOW`)
# - projects the resulting vector down to 5 units using a linear layer (including a bias term)
#
# Train your CBOW model and plot the validation accuracy and training loss over time.
# + colab={} colab_type="code" id="PEV22aR2MP0Q"
# YOUR CODE HERE
class CBOW(nn.Module):
"""A continuos bag-of-words model"""
def __init__(self, vocab_size, embedding_dim, output_dim, vocab):
super(CBOW, self).__init__()
self.vocab = vocab
# this is a trained look-up table with word embeddings
self.embed = nn.Embedding(vocab_size, embedding_dim)
self.applyW = nn.Linear(embedding_dim, output_dim)
# this is a trained bias term
self.bias = nn.Parameter(torch.zeros(output_dim), requires_grad=True)
def forward(self, inputs):
# this is the forward pass of the neural network
# given inputs, it computes the output
# this looks up the embeddings for each word ID in inputs
# the result is a sequence of word embeddings
embeds = self.embed(inputs)
trans_embeds = self.applyW(embeds)
# the output is the sum across the time dimension (1)
# with the bias term added
logits = trans_embeds.sum(1) + self.bias
return logits
# +
# cbow_model = CBOW(len(v.w2i), 300, len(t2i), vocab=v)
# print(cbow_model)
# cbow_model = cbow_model.to(device)
# optimizer = optim.Adam(cbow_model.parameters(), lr=0.0005)
# cbow_losses, cbow_accuracies = train_model(
# cbow_model, optimizer, num_iterations=300000,
# print_every=10000, eval_every=10000, batch_size = 64)
# +
# plt.plot(cbow_accuracies)
# +
# plt.plot(cbow_losses)
# + [markdown] colab_type="text" id="zpFt_Fo2TdN0"
# # Deep CBOW
# + [markdown] colab_type="text" id="iZanOMesTfEZ"
# To see if we can squeeze some more performance out of the CBOW model, we can make it deeper and non-linear: add more layers and tanh-activations.
# By using more parameters we can learn more aspects of the data, and by using more layers and non-linearities, we can try to learn a more complex function.
# This is not something that always works. If the input-output mapping of your data is simple, then a complicated function could easily overfit on your training set, which will lead to poor generalization.
#
# We will use E for the size of the word embeddings (use: 300) and D for the size of a hidden layer (use: 100).
#
# #### Exercise: write Deep CBOW class and train it
#
# Write a class `DeepCBOW`.
#
# In your code, make sure that your `output_layer` consists of the following:
# - A linear transformation from E units to D units.
# - A Tanh activation
# - A linear transformation from D units to D units
# - A Tanh activation
# - A linear transformation from D units to 5 units (our output classes).
#
# We recommend using [nn.Sequential](https://pytorch.org/docs/stable/nn.html?highlight=sequential#torch.nn.Sequential) to implement this exercise.
# + colab={} colab_type="code" id="l8Z1igvpTrZq"
# YOUR CODE HERE
class DeepCBOW(nn.Module):
"""A continuos bag-of-words model"""
def __init__(self, vocab_size, E, D, output_dim, vocab):
super(DeepCBOW, self).__init__()
self.vocab = vocab
# this is a trained look-up table with word embeddings
self.embed = nn.Embedding(vocab_size, E)
self.output_layer = nn.Sequential(OrderedDict([
('linear1', nn.Linear(E, D)),
('tanh1', nn.Tanh()),
('dropout1', nn.Dropout(0.1)),
('linear2', nn.Linear(D,D)),
('relu2', nn.Tanh()),
('dropout2', nn.Dropout(0.1)),
('linear3', nn.Linear(D,output_dim))
]))
# this is a trained bias term
self.bias = nn.Parameter(torch.zeros(output_dim), requires_grad=True)
def forward(self, inputs):
# this is the forward pass of the neural network
# given inputs, it computes the output
# this looks up the embeddings for each word ID in inputs
# the result is a sequence of word embeddings
embeds = self.embed(inputs)
for layer in self.output_layer:
embeds = layer(embeds)
# the output is the sum across the time dimension (1)
# with the bias term added
logits = embeds.sum(1) + self.bias
return logits
# +
# deep_cbow_model = DeepCBOW(len(v.w2i), 300, 100, len(t2i), vocab=v)
# print(deep_cbow_model)
# deep_cbow_model = deep_cbow_model.to(device)
# optimizer = optim.Adam(deep_cbow_model.parameters(), lr=0.002)
# deep_cbow_losses, deep_cbow_accuracies = train_model(
# deep_cbow_model, optimizer, num_iterations=300000,
# print_every=1000, eval_every=1000, batch_size = 64)
# +
# plt.plot(deep_cbow_accuracies)
# +
# plt.plot(deep_cbow_losses)
# + [markdown] colab_type="text" id="MQZ5flHwiiHY"
# # Pre-trained word embeddings
# + [markdown] colab_type="text" id="9NX35vecmHy6"
# The Stanford sentiment treebank is a very small data set, since it was manually annotated. This makes it difficult for the model to learn good word embeddings, to learn the "meaning" of the words in our vocabulary.
#
# Think for a moment about the fact that the only error signal that the network receives is from predicting the sentiment!
#
# To mitigate our data sparsity, we can download **pre-trained word embeddings**.
# You can choose which pre-trained word embeddings to use:
#
# - **Glove**. The "original" Stanford Sentiment classification [paper](http://aclweb.org/anthology/P/P15/P15-1150.pdf) used Glove embeddings, which are just another method (like *word2vec*) to get word embeddings from unannotated text. Glove is described in the following paper which you should cite if you use them:
# > Jeffrey Pennington, Richard Socher, and Christopher Manning. ["Glove: Global vectors for word representation."](https://nlp.stanford.edu/pubs/glove.pdf) EMNLP 2014.
#
# - **word2vec**. This is the method that you learned about in class, described in:
# > Mikolov, Tomas, et al. ["Distributed representations of words and phrases and their compositionality."](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) Advances in neural information processing systems. 2013.
#
# With these pre-trained word embeddings, we can initialize our word embedding lookup table and start form a point where similar words are already close to one another.
#
# You can choose to keep the word embeddings **fixed** or to train them further.
# We will keep them fixed for now.
#
# For the purposes of this lab, it is enough if you understand how word2vec works (whichever vectors you use). If you are interested you may also check out the Glove paper.
#
# You can either download the word2vec vectors, or the Glove vectors.
# If you want to compare your results to the Stanford paper later on, then you should use Glove.
# **At the end of this lab you have the option to compare which vectors give you the best performance. For now, simply choose one of them and continue with that.**
# + colab={} colab_type="code" id="lGYr02WWO993"
# This downloads the Glove 840B 300d embeddings.
# The original file is at http://nlp.stanford.edu/data/glove.840B.300d.zip
# Since that file is 2GB, we provide you with a *filtered version*
# which contains all the words you need for this data set.
# You only need to do this once.
# Please comment this cell out after downloading.
# # !wget https://gist.githubusercontent.com/bastings/b094de2813da58056a05e8e7950d4ad1/raw/3fbd3976199c2b88de2ae62afc0ecc6f15e6f7ce/glove.840B.300d.sst.txt
# + colab={} colab_type="code" id="6NLsgFGiTjmI"
# This downloads the word2vec 300D Google News vectors
# The file has been truncated to only contain words that appear in our data set.
# You can find the original file here: https://code.google.com/archive/p/word2vec/
# You only need to do this once.
# Please comment this out after downloading.
# # !wget https://gist.githubusercontent.com/bastings/4d1c346c68969b95f2c34cfbc00ba0a0/raw/76b4fefc9ef635a79d0d8002522543bc53ca2683/googlenews.word2vec.300d.txt
# + colab={} colab_type="code" id="GXBITzPRQUQb"
# Mount Google Drive (to save the downloaded files)
# from google.colab import drive
# drive.mount('/gdrive')
# + colab={} colab_type="code" id="uFvzPuiKSCbl"
# Copy word vectors *to* Google Drive
# You only need to do this once.
# Please comment this out after running it.
# # !cp "glove.840B.300d.sst.txt" "/gdrive/My Drive/"
# # !cp "googlenews.word2vec.300d.txt" "/gdrive/My Drive/"
# + colab={} colab_type="code" id="kUMH0bM6BuY9"
# If you copied the word vectors to your Drive before,
# here is where you copy them back to the Colab notebook.
# Copy Glove vectors *from* Google Drive
# # !cp "/gdrive/My Drive/glove.840B.300d.sst.txt" .
# # !cp "/gdrive/My Drive/googlenews.word2vec.300d.txt" .
# + [markdown] colab_type="text" id="MX2GJVHILM8n"
# At this point you have the pre-trained word embedding files, but what do they look like?
# + colab={} colab_type="code" id="ChsChH14Ruxn"
# Exercise: Print the first 4 lines of the files that you downloaded.
# What do you see?
with open("glove.840B.300d.sst.txt", "r") as f:
glove_init_lines = f.readlines()
print(glove_init_lines[2].split(" ")[0])
# + [markdown] colab_type="text" id="WIVCkUkE_IjR"
# #### Exercise: New Vocabulary
#
# Since we now use pre-trained word embeddings, we need to create a new vocabulary.
# This is because of two reasons:
#
# 1. Not all words in our training set are in the set of pre-trained word embeddings. We do not want words in our vocabulary that are not in that set.
# 2. We get to look up the pre-trained word embedding for words in the validation and test set, even if we don't know them from training.
#
# Now, create a new vocabulary object `v` and load the pre-trained vectors into a `vectors`.
#
# The vocabulary `v` should consist of:
# - a `<unk>` token at position 0,
# - a `<pad>` token at position 1,
# - and then all words in the pre-trained vector set.
#
#
# After storing each vector in a list `vectors`, turn in into a numpy matrix like this:
# ```python
# vectors = np.stack(vectors, axis=0)
# ```
#
#
#
# + colab={} colab_type="code" id="ITyyCvDnCL4U"
# YOUR CODE HERE
v = Vocabulary()
vectors = list()
vectors.append(np.zeros(300, dtype=np.float32)) # <unk>
vectors.append(np.zeros(300, dtype=np.float32)) # <pad>
for line in glove_init_lines:
v.count_token(line.split(" ")[0])
vectors.append(np.array([float(x) for x in line.split(" ")[1:]], dtype=np.float32))
vectors = np.stack(vectors, axis=0)
v.build()
# + [markdown] colab_type="text" id="xC-7mRyYNG9b"
# #### Exercise: words not in our pre-trained set
#
# How many words in the training, dev, and test set are also in your vector set?
# How many words are not there?
#
# Store the words that are not in the word vector set in the set below.
# + colab={} colab_type="code" id="K6MA3-wF_X5M"
words_not_found = set()
# YOUR CODE HERE
for data_set in (train_data,):
for ex in data_set:
for token in ex.tokens:
if v.w2i.get(token, 0) == 0:
words_not_found.update([token])
print(words_not_found)
print(len(words_not_found))
# + [markdown] colab_type="text" id="BfEd38W0NnAI"
# #### Exercise: train Deep CBOW with (fixed) pre-trained embeddings
#
# Now train Deep CBOW again using the pre-trained word vectors.
#
# + colab={} colab_type="code" id="z_6ooqgEsB20"
# We define a dummy class so that we save the model to a different file.
class PTDeepCBOW(DeepCBOW):
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, vocab):
super(PTDeepCBOW, self).__init__(
vocab_size, embedding_dim, hidden_dim, output_dim, vocab)
# + colab={} colab_type="code" id="JfIh4Ni6yuAh"
# YOUR CODE HERE
# pt_deep_cbow_model = PTDeepCBOW(len(v.w2i), 300, 100, 5, v)
# # copy pre-trained word vectors into embeddings table
# pt_deep_cbow_model.embed.weight.data.copy_(torch.from_numpy(vectors))
# disable training the pre-trained embeddings
# pt_deep_cbow_model.embed.weight.requires_grad = False
# move model to specified device
# pt_deep_cbow_model = pt_deep_cbow_model.to(device)
# train the model
# pt_deep_cbow_model = pt_deep_cbow_model.to(device)
# optimizer = optim.Adam(pt_deep_cbow_model.parameters(), lr=0.0001)
# pt_deep_cbow_losses, pt_deep_cbow_accuracies = train_model(
# pt_deep_cbow_model, optimizer, num_iterations=30000,
# print_every=1000, eval_every=1000, batch_size = 64)
# + colab={} colab_type="code" id="Ufujv3x31ufD"
# plot dev accuracies
# plt.plot(pt_deep_cbow_accuracies)
# + colab={} colab_type="code" id="YTJtKBzd7Qjr"
# plot train loss
# plt.plot(pt_deep_cbow_losses)
# + [markdown] colab_type="text" id="yFu8xzCy9XDW"
# **It looks like we've hit what is possible with just using words.**
# Let's move on by incorporating word order!
# + [markdown] colab_type="text" id="g41yW4PL9jG0"
# # LSTM
# + [markdown] colab_type="text" id="ODzXEH0MaGpa"
# It is time to get more serious. Even with pre-trained word embeddings and multiple layers, we seem to do pretty badly at sentiment classification here.
# The next step we can take is to introduce word order again, and to get a representation of the sentence as a whole, without independence assumptions.
#
# We will get this representation using an **Long Short-Term Memory** (LSTM). As an exercise, we will code our own LSTM cell, so that we get comfortable with its inner workings.
# Once we have an LSTM cell, we can call it repeatedly, updating its hidden state one word at a time:
#
# ```python
# rnn = MyLSTMCell(input_size, hidden_size)
#
# hx = torch.zeros(1, hidden_size) # initial state
# cx = torch.zeros(1, hidden_size) # initial memory cell
# output = [] # to save intermediate LSTM states
#
# # feed one word at a time
# for i in range(n_timesteps):
# hx, cx = rnn(input[i], (hx, cx))
# output.append(hx)
# ```
#
# If you need some more help understanding LSTMs, then check out these resources:
# - Blog post (highly recommended): http://colah.github.io/posts/2015-08-Understanding-LSTMs/
# - Paper covering LSTM formulas in detail: https://arxiv.org/abs/1503.04069
#
# #### Exercise: Finish the LSTM cell below.
# You will need to implement the LSTM formulas:
#
# $$
# \begin{array}{ll}
# i = \sigma(W_{ii} x + b_{ii} + W_{hi} h + b_{hi}) \\
# f = \sigma(W_{if} x + b_{if} + W_{hf} h + b_{hf}) \\
# g = \tanh(W_{ig} x + b_{ig} + W_{hg} h + b_{hg}) \\
# o = \sigma(W_{io} x + b_{io} + W_{ho} h + b_{ho}) \\
# c' = f * c + i * g \\
# h' = o \tanh(c') \\
# \end{array}
# $$
#
# where $\sigma$ is the sigmoid function.
#
# *Note that the LSTM formulas can differ slightly between different papers. We use the PyTorch LSTM formulation here.*
# + colab={} colab_type="code" id="zJ9m5kLMd7-v"
class MyLSTMCell(nn.Module):
"""Our own LSTM cell"""
def __init__(self, input_size, hidden_size, bias=True):
"""Creates the weights for this LSTM"""
super(MyLSTMCell, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.bias = bias
self.tanh_act = nn.Tanh()
self.sigmoid_act = nn.Sigmoid()
# self.input_gate = nn.Linear(input_size + hidden_size, hidden_size)
# self.forget_gate = nn.Linear(input_size + hidden_size, hidden_size)
# self.candidate_gate = nn.Linear(input_size + hidden_size, hidden_size)
# self.output_gate = nn.Linear(input_size + hidden_size, hidden_size)
self.combined_gate = nn.Linear(input_size + hidden_size, 4 * hidden_size)
self.reset_parameters()
def reset_parameters(self):
"""This is PyTorch's default initialization method"""
stdv = 1.0 / math.sqrt(self.hidden_size)
for weight in self.parameters():
weight.data.uniform_(-stdv, stdv)
def forward(self, input_, hx, mask=None):
"""
input is (batch, input_size)
hx is ((batch, hidden_size), (batch, hidden_size))
"""
prev_h, prev_c = hx
# project input and prev state
cat_input = torch.cat([input_,prev_h], dim=1)
# main LSTM computation
# i = self.sigmoid_act(self.input_gate(cat_input))
# f = self.sigmoid_act(self.forget_gate(cat_input))
# g = self.tanh_act(self.candidate_gate(cat_input))
# o = self.sigmoid_act(self.output_gate(cat_input))
combined_output = self.combined_gate(cat_input)
i = self.sigmoid_act(combined_output[:,0 * self.hidden_size:1 * self.hidden_size])
f = self.sigmoid_act(combined_output[:,1 * self.hidden_size:2 * self.hidden_size])
g = self.tanh_act(combined_output[:,2 * self.hidden_size:3 * self.hidden_size])
o = self.sigmoid_act(combined_output[:,3 * self.hidden_size:4 * self.hidden_size])
c = f * prev_c + i * g
h = o * self.tanh_act(c)
return h, c
def __repr__(self):
return "{}({:d}, {:d})".format(
self.__class__.__name__, self.input_size, self.hidden_size)
# + [markdown] colab_type="text" id="4JM7xPhkQeE5"
# #### Optional: Efficient Matrix Multiplication
#
# It is more efficient to do a few big matrix multiplications than to do many smaller ones.
#
# It is possible to implement the above cell using just **two** linear layers.
#
# This is because the eight linear transformations from one forward pass through an LSTM cell can be done in just two:
# $$W_h h + b_h$$
# $$W_i x + b_i $$
#
# with $h = $ `prev_h` and $x = $ `input_`.
#
# and where:
#
# $W_h = \begin{pmatrix}
# W_{hi}\\
# W_{hf}\\
# W_{hg}\\
# W_{ho}
# \end{pmatrix}$, $b_h = \begin{pmatrix}
# b_{hi}\\
# b_{hf}\\
# b_{hg}\\
# b_{ho}
# \end{pmatrix}$, $W_i = \begin{pmatrix}
# W_{ii}\\
# W_{if}\\
# W_{ig}\\
# W_{io}
# \end{pmatrix}$ and $b_i = \begin{pmatrix}
# b_{ii}\\
# b_{if}\\
# b_{ig}\\
# b_{io}
# \end{pmatrix}$.
#
# Convince yourself that, after chunking with [torch.chunk](https://pytorch.org/docs/stable/torch.html?highlight=chunk#torch.chunk), the output of those two linear transformations is equivalent to the output of the eight linear transformations in the LSTM cell calculations above.
# + [markdown] colab_type="text" id="X9gA-UcqSBe0"
# #### LSTM Classifier
#
# Having an LSTM cell is not enough: we still need some code that calls it repeatedly, and then makes a prediction from the final hidden state.
# You will find that code below. Make sure that you understand it.
# + colab={} colab_type="code" id="3iuYZm5poEn5"
class LSTMClassifier(nn.Module):
"""Encodes sentence with an LSTM and projects final hidden state"""
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, vocab):
super(LSTMClassifier, self).__init__()
self.vocab = vocab
self.hidden_dim = hidden_dim
self.embed = nn.Embedding(vocab_size, embedding_dim, padding_idx=1)
self.rnn = MyLSTMCell(embedding_dim, hidden_dim)
self.output_layer = nn.Sequential(
nn.Dropout(p=0.1), # explained later
nn.Linear(hidden_dim, output_dim)
)
def forward(self, x):
B = x.size(0) # batch size (this is 1 for now, i.e. 1 single example)
T = x.size(1) # time (the number of words in the sentence)
input_ = self.embed(x)
# here we create initial hidden states containing zeros
# we use a trick here so that, if input is on the GPU, then so are hx and cx
hx = input_.new_zeros(B, self.rnn.hidden_size)
cx = input_.new_zeros(B, self.rnn.hidden_size)
# process input sentences one word/timestep at a time
# input is batch-major, so the first word(s) is/are input_[:, 0]
outputs = []
for i in range(T):
hx, cx = self.rnn(input_[:, i], (hx, cx))
outputs.append(hx)
# if we have a single example, our final LSTM state is the last hx
if B == 1:
final = hx
else:
#
# This part is explained in next section, ignore this else-block for now.
#
# we processed sentences with different lengths, so some of the sentences
# had already finished and we have been adding padding inputs to hx
# we select the final state based on the length of each sentence
# two lines below not needed if using LSTM form pytorch
outputs = torch.stack(outputs, dim=0) # [T, B, D]
outputs = outputs.transpose(0, 1).contiguous() # [B, T, D]
# to be super-sure we're not accidentally indexing the wrong state
# we zero out positions that are invalid
pad_positions = (x == 1).unsqueeze(-1)
outputs = outputs.contiguous()
outputs = outputs.masked_fill_(pad_positions, 0.)
mask = (x != 1) # true for valid positions [B, T]
lengths = mask.sum(dim=1) # [B, 1]
indexes = (lengths - 1) + torch.arange(B, device=x.device, dtype=x.dtype) * T
final = outputs.view(-1, self.hidden_dim)[indexes] # [B, D]
# we use the last hidden state to classify the sentence
logits = self.output_layer(final)
return logits
# + [markdown] colab_type="text" id="FxFoVpvMPB6g"
# #### Dropout
#
# Besides not being able to learn meaningful word embeddings, there is another negative effect that can follow from data sparsity and a small data set: *overfitting*. This is a phenomenom that is very likely to occur when fitting strong and expressive models, like LSTMs, to small data. In practice, if your model overfits, this means that it will be very good at predicting (or: 'remembering') the sentiment of the training set, but unable to generalize to new, unseen data in the test set. This is undesirable and one technique to mitigate it is *dropout*.
#
# A dropout layer is defined by the following formula: $\mathbf{d} \in \{0, 1\}^n$, with $d_j \sim \text{Bernoulli}(p)$, and can be applied to for example a linear layer:
#
# $$\text{tanh}(W(\mathbf{h}\odot \mathbf{d}) + \mathbf{b})$$
#
#
# These formulas simply mean that we *drop* certain parameters during training (by setting them to zero). Which parameters we drop is stochastically determined by a Bernoulli distribution and the probability of each parameter being dropped is set to $p = 0.5$ in our experiments (see the previous cell of code where we define our output layer). A dropout layer can be applied at many different places in our models. This technique helps against the undesirable effect where our model relies on single parameters for prediction (e.g. if $h^{\prime}_j$ is large, always predict positive). If we use dropout, the model needs to learn to rely on different parameters, which is desirable when generalizing to unseen data.
# + [markdown] colab_type="text" id="XQjEjLt9z0XW"
# **Let's train our LSTM! ** Note that is will be a lot slower, because we need to do many more computations per sentence!
#
# #### Training
# + colab={} colab_type="code" id="LgZoSPD4fsf_"
"""
lstm_model = LSTMClassifier(len(v.w2i), 300, 168, len(t2i), v)
# copy pre-trained word vectors into embeddings table
with torch.no_grad():
lstm_model.embed.weight.data.copy_(torch.from_numpy(vectors))
lstm_model.embed.weight.requires_grad = False
print(lstm_model)
print_parameters(lstm_model)
lstm_model = lstm_model.to(device)
optimizer = optim.Adam(lstm_model.parameters(), lr=3e-4)
lstm_losses, lstm_accuracies = train_model(
lstm_model, optimizer, num_iterations=25000,
print_every=250, eval_every=1000)
"""
# + colab={} colab_type="code" id="2BKVnyg0Hq5E"
# plot validation accuracy
# plt.plot(lstm_accuracies)
# + colab={} colab_type="code" id="ZowTV0EBTb3z"
# plot training loss
# plt.plot(lstm_losses)
# + [markdown] colab_type="text" id="YEw6XHQY_AAQ"
# # Mini-batching
#
#
# + [markdown] colab_type="text" id="FPf96wGzBTQJ"
# **Why is the LSTM so slow?** Despite our best efforts, we still need to make a lot of matrix multiplications per example (linear in the length of the example) just to get a single classification, and we can only process the 2nd word once we have computed the hidden state for the 1st word (sequential computation).
#
# GPUs are more efficient if we do a few big matrix multiplications, rather than lots of small ones. If we could process multiple examples at the same time, then we could exploit that. We still process the input sequentially, but now we can do so for multiple sentences at the same time.
#
# Up to now our "minibatch" consisted of a single example. This was for a reason: the sentences in our data sets have **different lengths**, and this makes it difficult to process them at the same time.
#
# Consider a batch of 2 sentences:
#
# ```
# this movie is bad
# this movie is super cool !
# ```
#
# Let's say the IDs for these sentences are:
#
# ```
# 2 3 4 5
# 2 3 4 6 7 8
# ```
#
# We cannot feed PyTorch an object with variable length rows! We need to turn this into a matrix.
#
# The solution is to add **padding values** to our mini-batch:
#
# ```
# 2 3 4 5 1 1
# 2 3 4 6 7 8
# ```
#
# Whenever a sentence is shorter than the longest sentence in a mini-batch, we just use a padding value (here: 1) to fill the matrix.
#
# In our computation, we should **ignore** the padding positions (e.g. mask them out). Paddings should not contribute to the loss.
#
# #### Mini-batch feed
# We will now code a `get_minibatch` function that will replace our `get_example` function, and returns a mini-batch of the requested size.
# + colab={} colab_type="code" id="IoAE2JBiXJ3P"
def get_minibatch(data, batch_size=25, shuffle=True):
"""Return minibatches, optional shuffling"""
if shuffle:
# print("Shuffling training data")
random.shuffle(data) # shuffle training data each epoch
batch = []
# yield minibatches
for example in data:
batch.append(example)
if len(batch) == batch_size:
yield batch
batch = []
# in case there is something left
if len(batch) > 0:
yield batch
# + [markdown] colab_type="text" id="DwZM-XYkT8Zx"
# #### Pad function
# We will need a function that adds padding 1s to a sequence of IDs so that
# it becomes as long as the longest sequencen in the minibatch.
# + colab={} colab_type="code" id="sp0sK1ghw4Ft"
def pad(tokens, length, pad_value=1):
"""add padding 1s to a sequence to that it has the desired length"""
return tokens + [pad_value] * (length - len(tokens))
# example
tokens = [2, 3, 4]
pad(tokens, 5)
# + [markdown] colab_type="text" id="SL2iixMYUgfh"
# #### New prepare function
#
# We will also need a new function that turns a mini-batch into PyTorch tensors.
# + colab={} colab_type="code" id="ZID0cqozWks8"
def prepare_minibatch(mb, vocab):
"""
Minibatch is a list of examples.
This function converts words to IDs and returns
torch tensors to be used as input/targets.
"""
batch_size = len(mb)
maxlen = max([len(ex.tokens) for ex in mb])
# vocab returns 0 if the word is not there
x = [pad([vocab.w2i.get(t, 0) for t in ex.tokens], maxlen) for ex in mb]
x = torch.LongTensor(x)
x = x.to(device)
y = [ex.label for ex in mb]
y = torch.LongTensor(y)
y = y.to(device)
return x, y
# + colab={} colab_type="code" id="OwDAtCv1x2hB"
# Let's test our new function.
# This should give us 3 examples.
mb = next(get_minibatch(train_data, batch_size=3, shuffle=False))
print(len(mb))
for ex in mb:
print(ex)
# + colab={} colab_type="code" id="dg8zEK8zyUCH"
# We should find 1s at the end where padding is.
x, y = prepare_minibatch(mb, v)
print("x", x)
print("y", y)
# + [markdown] colab_type="text" id="xYBJEoSNUwI0"
# #### Evaluate (mini-batch version)
#
# We can now update our evaluation function to use minibatches
# + colab={} colab_type="code" id="eiZZpEghzqou"
def evaluate(model, data,
batch_fn=get_minibatch, prep_fn=prepare_minibatch,
batch_size=16):
"""Accuracy of a model on given data set (using minibatches)"""
correct = 0
total = 0
model.eval() # disable dropout
for mb in batch_fn(data, batch_size=batch_size, shuffle=False):
x, targets = prep_fn(mb, model.vocab)
with torch.no_grad():
logits = model(x)
predictions = logits.argmax(dim=-1).view(-1)
# add the number of correct predictions to the total correct
correct += (predictions == targets.view(-1)).sum().item()
total += targets.size(0)
print("Evaluation on " + str(total) + " elements. Correct: " + str(correct))
return correct, total, correct / float(total)
# + [markdown] colab_type="text" id="23wAZomozh_2"
# # LSTM (Mini-batched)
# + [markdown] colab_type="text" id="B-gkPU7jzBe2"
# With this, let's run the LSTM again but now using minibatches!
# + colab={} colab_type="code" id="226Xg9OPzFbA"
"""
lstm_model = LSTMClassifier(
len(v.w2i), 300, 168, len(t2i), v)
# copy pre-trained vectors into embeddings table
with torch.no_grad():
lstm_model.embed.weight.data.copy_(torch.from_numpy(vectors))
lstm_model.embed.weight.requires_grad = False
print(lstm_model)
print_parameters(lstm_model)
lstm_model = lstm_model.to(device)
batch_size = 512
optimizer = optim.Adam(lstm_model.parameters(), lr=2e-4)
lstm_losses, lstm_accuracies = train_model(
lstm_model, optimizer, num_iterations=30000,
print_every=250, eval_every=250,
batch_size=batch_size,
batch_fn=get_minibatch,
prep_fn=prepare_minibatch,
eval_fn=evaluate)
"""
# + colab={} colab_type="code" id="ymj1rLDMvyhp"
# plot validation accuracy
# plt.plot(lstm_accuracies)
# + colab={} colab_type="code" id="1je5S1RHVC5R"
# plot training loss
# plt.plot(lstm_losses)
# -
# + [markdown] colab_type="text" id="q7WjcxXntMi5"
# # Tree LSTM
# + [markdown] colab_type="text" id="jyj_UD6GtO5M"
# In the final part of this lab we will exploit the tree-structure of our data.
# Until now we only used the surface tokens, but remember that our data examples include trees with a sentiment score at every node.
#
# In particular, we will implement **N-ary Tree-LSTMs** which are described in:
#
# > Kai Sheng Tai, Richard Socher, and Christopher D. Manning. [Improved Semantic Representations From Tree-Structured Long Short-Term Memory Networks](http://aclweb.org/anthology/P/P15/P15-1150.pdf) ACL 2015.
#
# Since our trees are binary, N=2, and we can refer to these as *Binary Tree-LSTMs*.
#
# You should read this paper carefully and make sure that you understand the approach. You will also find our LSTM baseline there.
# Note however that Tree LSTMs were also invented around the same time by two other groups:
#
# > Phong Le and Willem Zuidema. [Compositional distributional semantics with long short term memory](http://anthology.aclweb.org/S/S15/S15-1002.pdf). *SEM 2015.
#
# > Xiaodan Zhu, Parinaz Sobihani, and Hongyu Guo. [Long short-term memory over recursive structures](http://proceedings.mlr.press/v37/zhub15.pdf). ICML 2015.
#
# It is good scientific practice to cite all three papers in your report.
#
# If you study equations (9) to (14) in the paper, you will find that they are not all too different from the original LSTM that you already have.
#
#
#
# + [markdown] colab_type="text" id="1rDzvSos3JFp"
# ## Computation
#
# Do you remember the `transitions_from_treestring` function all the way in the beginning of this lab? Every example contains a **transition sequence** made by this function. Let's look at it again:
#
#
# + colab={} colab_type="code" id="5pg0Xumc3ZUS"
ex = next(examplereader("trees/dev.txt"))
print(TreePrettyPrinter(ex.tree))
print("Transitions:")
print(ex.transitions)
# + [markdown] colab_type="text" id="ceBFe9fU4BI_"
# Note that the tree is **binary**. Every node has two children, except for pre-terminal nodes.
#
# A tree like this can be described by a sequence of **SHIFT (0)** and **REDUCE (1)** actions.
#
# We can use the transitions like this to construct the tree:
# - **reverse** the sentence (a list of tokens) and call this the **buffer**
# - the first word is now on top (last in the list), and we would get it when calling pop() on the buffer
# - create an empty list and call it the **stack**
# - iterate through the transition sequence:
# - if it says SHIFT(0), we pop a word from the buffer, and push it to the stack
# - if it says REDUCE(1), we pop the **top two items** from the stack, and combine them (e.g. with a tree LSTM!), creating a new node that we push back on the stack
#
# Convince yourself that going through the transition sequence above will result in the tree that you see.
# For example, we would start by putting the following words on the stack (by shifting 5 times, starting with `It`):
#
# ```
# Top of the stack:
# -----------------
# film
# lovely
# a
# 's
# It
# ```
# Now we find a REDUCE in the transition sequence, so we get the top two words (film and lovely), and combine them, so our new stack becomes:
# ```
# Top of the stack:
# -----------------
# lovely film
# a
# 's
# It
# ```
#
# We will use this approach when encoding sentences with our Tree LSTM.
# Now, our sentence is a (reversed) list of word embeddings.
# When we shift, we move a word embedding to the stack.
# When we reduce, we apply a Tree LSTM to the top two vectors, and the result is a single vector that we put back on the stack.
# After going through the whole transition sequence, we will have the root node on our stack! We can use that to classify the sentence.
#
#
# + [markdown] colab_type="text" id="pDWKShm1AfmR"
# ## Obtaining the transition sequence
#
# + [markdown] colab_type="text" id="fO7VKWVpAbWj"
#
# So what goes on in the `transitions_from_treestring` function?
#
# The idea ([explained in this blog post](https://devblogs.nvidia.com/recursive-neural-networks-pytorch/)) is that, if we had a tree, we could traverse through the tree, and every time that we find a node containing only a word, we output a SHIFT.
# Every time **after** we have finished visiting the children of a node, we output a REDUCE.
# (What is this tree traversal called?)
#
# However, our `transitions_from_treestring` function operates directly on the string representation. It works as follows.
#
# We start with the representation:
#
# ```
# (3 (2 It) (4 (4 (2 's) (4 (3 (2 a) (4 (3 lovely) (2 film))) (3 (2 with) (4 (3 (3 lovely) (2 performances)) (2 (2 by) (2 (2 (2 Buy) (2 and)) (2 Accorsi))))))) (2 .)))
# ```
#
# First we remove pre-terminal nodes (and add spaces before closing brackets):
#
# ```
# (3 It (4 (4 's (4 (3 a (4 lovely film ) ) (3 with (4 (3 lovely performances ) (2 by (2 (2 Buy and ) Accorsi ) ) ) ) ) ) . ) )
# ```
#
# Then we remove node labels:
#
# ```
# ( It ( ( 's ( ( a ( lovely film ) ) ( with ( ( lovely performances) ( by ( ( Buy and ) Accorsi ) ) ) ) ) ) . ) )
# ```
#
# Then we remove opening brackets:
#
# ```
# It 's a lovely film ) ) with lovely performances ) by Buy and ) Accorsi ) ) ) ) ) ) . ) )
# ```
#
# Now we replace words by S (for SHIFT), and closing brackets by R (for REDUCE):
#
# ```
# S S S S S R R S S S R S S S R S R R R R R R S R R
# 0 0 0 0 0 1 1 0 0 0 1 0 0 0 1 0 1 1 1 1 1 1 0 1 1
# ```
#
# Et voila. We just obtained the transition sequence!
# + colab={} colab_type="code" id="1y069gM4_v64"
# for comparison
seq = ex.transitions
s = " ".join(["S" if t == 0 else "R" for t in seq])
print(s)
print(" ".join(map(str, seq)))
# + [markdown] colab_type="text" id="d-qOuKbDAiBn"
# ## Coding the Tree LSTM
#
# The code below contains a Binary Tree LSTM cell.
# It is used in the TreeLSTM class below it, which in turn is used in the TreeLSTMClassifier.
# The job of the TreeLSTM class is to encode a complete sentence and return the root node.
# The job of the TreeLSTMCell is to return a new state when provided with two children (a reduce action). By repeatedly calling the TreeLSTMCell, the TreeLSTM will encode a sentence. This can be done for multiple sentences at the same time.
#
#
# #### Exercise
# Check the `forward` function and complete the Tree LSTM formulas.
# You can see that we defined a large linear layer for you, that projects the *concatenation* of the left and right child into the input gate, left forget gate, right forget gate, candidate, and output gate.
# + colab={} colab_type="code" id="J9b9mjMlN7Pb"
class TreeLSTMCell(nn.Module):
"""A Binary Tree LSTM cell"""
def __init__(self, input_size, hidden_size, bias=True):
"""Creates the weights for this LSTM"""
super(TreeLSTMCell, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.bias = bias
self.reduce_layer = nn.Linear(2 * hidden_size, 5 * hidden_size)
self.dropout_layer = nn.Dropout(p=0.25)
self.tanh_act = nn.Tanh()
self.sigmoid_act = nn.Sigmoid()
self.reset_parameters()
def reset_parameters(self):
"""This is PyTorch's default initialization method"""
stdv = 1.0 / math.sqrt(self.hidden_size)
for weight in self.parameters():
weight.data.uniform_(-stdv, stdv)
def forward(self, hx_l, hx_r, mask=None):
"""
hx_l is ((batch, hidden_size), (batch, hidden_size))
hx_r is ((batch, hidden_size), (batch, hidden_size))
"""
prev_h_l, prev_c_l = hx_l # left child
prev_h_r, prev_c_r = hx_r # right child
B = prev_h_l.size(0)
# we concatenate the left and right children
# you can also project from them separately and then sum
children = torch.cat([prev_h_l, prev_h_r], dim=1)
# project the combined children into a 5D tensor for i,fl,fr,g,o
# this is done for speed, and you could also do it separately
proj = self.reduce_layer(children) # shape: B x 5D
# each shape: B x D
i, f_l, f_r, g, o = torch.chunk(proj, 5, dim=-1)
# main Tree LSTM computation
# YOUR CODE HERE
# You only need to complete the commented lines below.
# The shape of each of these is [batch_size, hidden_size]
i = self.sigmoid_act(i)
f_l = self.sigmoid_act(f_l)
f_r = self.sigmoid_act(f_r)
g = self.tanh_act(g)
o = self.sigmoid_act(o)
c = i * g + f_l * prev_c_l + f_r * prev_c_r
h = o * self.tanh_act(c)
return h, c
def __repr__(self):
return "{}({:d}, {:d})".format(
self.__class__.__name__, self.input_size, self.hidden_size)
# + [markdown] colab_type="text" id="Dj5dYSGh_643"
# ## Explanation of the TreeLSTM class
#
#
# The code below contains the TreeLSTM class, which implements everything we need in order to encode a sentence from word embeddings. The calculations are the same as in the paper, implemented such that the class `TreeLSTMCell` above is as general as possible and only takes two children to reduce them into a parent.
#
#
# **Initialize $\mathbf{h}$ and $\mathbf{c}$ outside of the cell for the leaves**
#
# At the leaves of each tree the children nodes are **empty**, whereas in higher levels the nodes are binary tree nodes that *do* have a left and right child (but no input $x$). By initializing the leaf nodes outside of the cell class (`TreeLSTMCell`), we avoid if-else statements in the forward pass.
#
# The `TreeLSTM` class (among other things) pre-calculates an initial $h$ and $c$ for every word in the sentence. Since the initial left and right child are 0, the only calculations we need to do are based on $x$, and we can drop the forget gate calculation (`prev_c_l` and `prev_c_r` are zero). The calculations we do in order to initalize $h$ and $c$ are then:
#
# $$
# c_1 = W^{(u)}x_1 \\
# o_1 = \sigma (W^{(i)}x_1) \\
# h_1 = o_1 \odot \text{tanh}(c_1)$$
# *NB: note that these equations are chosen as initializations of $c$ and $h$, other initializations are possible and might work equally well.*
#
# **Sentence Representations**
#
# All our leaf nodes are now initialized, so we can start processing the sentence in its tree form. Each sentence is represented by a buffer (initially a list with a concatenation of $[h_1, c_1]$ for every word in the reversed sentence), a stack (initially an empty list) and a transition sequence. To encode our sentence, we construct the tree from its transition sequence as explained earlier.
#
# *A short example that constructs a tree:*
#
# We loop over the time dimension of the batched transition sequences (i.e. row by row), which contain values of 0's, 1's and 2's (representing SHIFT, REDUCE and padding respectively). If we have a batch of size 2 where the first example has a transition sequence given by [0, 0, 1, 0, 0, 0, 1] and the second by [0, 0, 1, 0, 0, 1], our transition batch will be given by the following two-dimensional numpy array:
#
# $$
# \text{transitions} =
# \begin{pmatrix}
# 0 & 0\\
# 0 & 0\\
# 1 & 1\\
# 0 & 0\\
# 0 & 0\\
# 0 & 1\\
# 1 & 2
# \end{pmatrix}
# $$
# The inner loop (`for transition, buffer, stack in zip(t_batch, buffers, stacks)`) goes over each example in the batch and updates its buffer and stack. The nested loop for this example will then do roughy the following:
#
# ```
# Time = 0: t_batch = [0, 0], the inner loop performs 2 SHIFTs.
#
# Time = 1: t_batch = [0, 0], ".."
#
# Time = 2: t_batch = [1, 1], causing the inner loop to fill the list child_l and child_r for both examples in the batch. Now the statement if child_l will return True, triggering a REDUCE action to be performed by our Tree LSTM cell with a batch size of 2.
#
# Time = 3: t_batch = [0, 0], "..".
#
# Time = 4: t_batch = [0, 0], ".."
#
# Time = 5: t_batch = [0, 1], one SHIFT will be done and another REDUCE action will be performed by our Tree LSTM, this time of batch size 1.
#
# Time = 6: t_batch = [1, 2], triggering another REDUCE action with batch size 1.
# ```
# *NB: note that this was an artificial example for the purpose of demonstrating parts of the code, the transition sequences do not necessarily represent actual trees.*
#
# **Batching and Unbatching**
#
# Within the body of the outer loop over time, we use the functions for batching and unbatching.
#
# *Batching*
#
# Before passing two lists of children to the reduce layer (an instance of `TreeLSTMCell`), we batch the children as they are at this point a list of tensors of variable length based on how many REDUCE actions there are to perform at a certain time step across the batch (let's call the length `L`). To do an efficient forward pass we want to transform the list to a pair of tensors of shape `([L, D], [L, D])`, which the function `batch` achieves.
#
# *Unbatching*
#
# In the same line where we batched the children, we unbatch the output of the forward pass to become a list of states of length `L` again. We do this because we need to loop over each example's transition at the current time step and push the children that are reduced into a parent to the stack.
#
# *The batch and unbatch functions let us switch between the "PyTorch world" (Tensors) and the Python world (easy to manipulate lists).*
#
# + colab={} colab_type="code" id="5PixvTd4AqsQ"
# Helper functions for batching and unbatching states
# For speed we want to combine computations by batching, but
# for processing logic we want to turn the output into lists again
# to easily manipulate.
def batch(states):
"""
Turns a list of states into a single tensor for fast processing.
This function also chunks (splits) each state into a (h, c) pair"""
return torch.cat(states, 0).chunk(2, 1)
def unbatch(state):
"""
Turns a tensor back into a list of states.
First, (h, c) are merged into a single state.
Then the result is split into a list of sentences.
"""
return torch.split(torch.cat(state, 1), 1, 0)
# + [markdown] colab_type="text" id="CynltDasaLPt"
# Take some time to understand the class below, having read the explanation above.
# + colab={} colab_type="code" id="rQOqMXG4gX5G"
class TreeLSTM(nn.Module):
"""Encodes a sentence using a TreeLSTMCell"""
def __init__(self, input_size, hidden_size, bias=True):
"""Creates the weights for this LSTM"""
super(TreeLSTM, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.bias = bias
self.reduce = TreeLSTMCell(input_size, hidden_size)
# project word to initial c
self.proj_x = nn.Linear(input_size, hidden_size)
self.proj_x_gate = nn.Linear(input_size, hidden_size)
self.buffers_dropout = nn.Dropout(p=0.5)
def forward(self, x, transitions):
"""
WARNING: assuming x is reversed!
:param x: word embeddings [B, T, E]
:param transitions: [2T-1, B]
:return: root states
"""
B = x.size(0) # batch size
T = x.size(1) # time
# compute an initial c and h for each word
# Note: this corresponds to input x in the Tai et al. Tree LSTM paper.
# We do not handle input x in the TreeLSTMCell itself.
buffers_c = self.proj_x(x)
buffers_h = buffers_c.tanh()
buffers_h_gate = self.proj_x_gate(x).sigmoid()
buffers_h = buffers_h_gate * buffers_h
# concatenate h and c for each word
buffers = torch.cat([buffers_h, buffers_c], dim=-1)
D = buffers.size(-1) // 2
# we turn buffers into a list of stacks (1 stack for each sentence)
# first we split buffers so that it is a list of sentences (length B)
# then we split each sentence to be a list of word vectors
buffers = buffers.split(1, dim=0) # Bx[T, 2D]
buffers = [list(b.squeeze(0).split(1, dim=0)) for b in buffers] # BxTx[2D]
# create B empty stacks
stacks = [[] for _ in buffers]
# t_batch holds 1 transition for each sentence
for t_batch in transitions:
child_l = [] # contains the left child for each sentence with reduce action
child_r = [] # contains the corresponding right child
# iterate over sentences in the batch
# each has a transition t, a buffer and a stack
for transition, buffer, stack in zip(t_batch, buffers, stacks):
if transition == SHIFT:
stack.append(buffer.pop())
elif transition == REDUCE:
assert len(stack) >= 2, \
"Stack too small! Should not happen with valid transition sequences"
child_r.append(stack.pop()) # right child is on top
child_l.append(stack.pop())
# if there are sentences with reduce transition, perform them batched
if child_l:
reduced = iter(unbatch(self.reduce(batch(child_l), batch(child_r))))
for transition, stack in zip(t_batch, stacks):
if transition == REDUCE:
stack.append(next(reduced))
final = [stack.pop().chunk(2, -1)[0] for stack in stacks]
final = torch.cat(final, dim=0) # tensor [B, D]
return final
# + [markdown] colab_type="text" id="s4EzbVzqaXkw"
# Just like the LSTM before, we will need an extra class that does the classifications.
# + colab={} colab_type="code" id="nLxpYRvtQKge"
class TreeLSTMClassifier(nn.Module):
"""Encodes sentence with a TreeLSTM and projects final hidden state"""
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, vocab):
super(TreeLSTMClassifier, self).__init__()
self.vocab = vocab
self.hidden_dim = hidden_dim
self.embed = nn.Embedding(vocab_size, embedding_dim, padding_idx=1)
self.treelstm = TreeLSTM(embedding_dim, hidden_dim)
self.output_layer = nn.Sequential(
nn.Dropout(p=0.5),
nn.Linear(hidden_dim, output_dim, bias=True)
)
def forward(self, x):
# x is a pair here of words and transitions; we unpack it here.
# x is batch-major: [B, T], transitions is time major [2T-1, B]
x, transitions = x
emb = self.embed(x)
# we use the root/top state of the Tree LSTM to classify the sentence
root_states = self.treelstm(emb, transitions)
# we use the last hidden state to classify the sentence
logits = self.output_layer(root_states)
return logits
# + [markdown] colab_type="text" id="gh9RbhGwaiLg"
# ## Special prepare function for Tree LSTM
#
# We need yet another prepare function. For our implementation our sentences to be *reversed*. We will do that here.
# + colab={} colab_type="code" id="DiqH-_2xdm9H"
def prepare_treelstm_minibatch(mb, vocab):
"""
Returns sentences reversed (last word first)
Returns transitions together with the sentences.
"""
batch_size = len(mb)
maxlen = max([len(ex.tokens) for ex in mb])
# vocab returns 0 if the word is not there
# NOTE: reversed sequence!
x = [pad([vocab.w2i.get(t, 0) for t in ex.tokens], maxlen)[::-1] for ex in mb]
x = torch.LongTensor(x)
x = x.to(device)
y = [ex.label for ex in mb]
y = torch.LongTensor(y)
y = y.to(device)
maxlen_t = max([len(ex.transitions) for ex in mb])
transitions = [pad(ex.transitions, maxlen_t, pad_value=2) for ex in mb]
transitions = np.array(transitions)
transitions = transitions.T # time-major
return (x, transitions), y
# + [markdown] colab_type="text" id="IMUsrlL9ayVe"
# ## Training
# + colab={} colab_type="code" id="IpOYUdg2D3v0"
# Now let's train the Tree LSTM!
"""
tree_model = TreeLSTMClassifier(
len(v.w2i), 300, 150, len(t2i), v)
with torch.no_grad():
tree_model.embed.weight.data.copy_(torch.from_numpy(vectors))
tree_model.embed.weight.requires_grad = False
def do_train(model):
print(model)
print_parameters(model)
model = model.to(device)
optimizer = optim.Adam(model.parameters(), lr=2e-4)
return train_model(
model, optimizer, num_iterations=30000,
print_every=250, eval_every=250,
prep_fn=prepare_treelstm_minibatch,
eval_fn=evaluate,
batch_fn=get_minibatch,
batch_size=32, eval_batch_size=32)
results = do_train(tree_model)
"""
# + colab={} colab_type="code" id="DHcHHaLtguUg"
# plot
# plt.plot(results[0])
# +
# plt.plot(results[1])
# + [markdown] colab_type="text" id="f7QZZH86eHqu"
# # Further experiments and report
#
# For your report, you are expected to answer research questions by doing further experiments.
#
# ## Research Questions
#
# Make sure you cover at least the following:
#
# - How important is word order for this task?
# - Does the tree structure help to get a better accuracy?
# - How does performance depend on the sentence length? Compare the various models. Is there a model that does better on longer sentences? If so, why?
# - Do you get better performance if you supervise the sentiment **at each node in the tree**? You can extract more training examples by treating every node in each tree as a separate tree. You will need to write a function that extracts all subtrees given a treestring.
# - Warning: NLTK's Tree function seems to result in invalid trees in some cases, so be careful if you want to parse the string to a tree structure before extraction the phrases.
#
# Optionally, you can also investigate the following:
#
# - When making a wrong prediction, can you figure out at what point in the tree (sentence) the model fails? You can make a prediction at each node to investigate.
# - How does N-ary Tree LSTM compare to the Child-Sum Tree LSTM?
# - How do the Tai et al. Tree LSTMs compare to Le & Zuidema's formulation?
# - Or.. your own research question!
#
# In general:
#
# - ***When you report numbers, please report the mean accuracy across 3 (or more) runs with different random seed, together with the standard deviation.*** This is because the final performance may vary per random seed.
#
# ## Report instructions
#
# Your report needs to be written in LaTeX. You are required to use the ACL 2018 template [(zip)](https://acl2018.org/downloads/acl18-latex.zip) which you can edit directly on [Overleaf](https://www.overleaf.com/latex/templates/instructions-for-acl-2018-proceedings/xzmhqgnmkppc). Make sure your names and student numbers are visible at the top. (Tip: you need to uncomment `/aclfinalcopy`).
# You can find some general tips about writing a research paper [here](https://www.microsoft.com/en-us/research/academic-program/write-great-research-paper/), but note that you need to make your own judgment about what is appropriate for this project.
#
# We expect you to use the following structure:
# 1. Introduction (~1 page) - describe the problem, your research questions and goals, a summary of your findings and contributions. Please cite related work (models, data set) as part of your introduction here, since this is a short paper.
# - Introduce the task and the main goal
# - Clear research questions
# - Motivating the importance of the questions and explaining the expectations
# - How are these addressed or not addressed in the literature
# - What is your approach
# - Short summary of your findings
# 2. Background (~1/2-1 page) -
# cover the main techniques ("building blocks") used in your project (e.g. word embeddings, LSTM, Tree LSTM) and intuitions behind them. Be accurate and concise.
# - How each technique that you use works (don't just copy the formulas)
# - The relation between the techniques
# 3. Models (~1/2 page) - Cover the models that you used.
# - The architecture of the final models (how do you use LSTM or Tree LSTM for the sentiment classification task. what layers you have, how do you do classification? What is your loss function?)
# 4. Experiments (~1/2 page) - Describe your experimental setup. The information here should allow someone else to re-create your experiments. Describe how you evaluate the models.
# - Explain the task and the data
# - Training the models (model, data, parameters and hyper parameters if the models, training algorithms, what supervision signals you use, etc.)
# - Evaluation (e.g. metrics)
# 5. Results and Analysis (~1 page). Go over the results and analyse your findings.
# - Answer each of the research questions you raised in the introduction.
# - Plots and figures highlighting interesting patterns
# - What are the factors that makes model A better than model B in task C? investigate to prove their effect!
# 6. Conclusion (~1/4 page). The main conclusions of your experiments.
# - What you learned from you experiments? how does it relate to what is already known in the literature?
# - Where the results as expected ? any surprising results? why?
# - Based on what you learned what would you suggest to do next?
#
#
# General Tips:
#
# - Math notation – define each variable (either in running text, or in a pseudo-legenda after or before the equation)
# - Define technical terminology you need
# - Avoid colloquial language – everything can be said in a scientific-sounding way
# - Avoid lengthy sentences, stay to the point!
# - Do not spend space on "obvious" things!
#
#
# An ideal report:
# - Precise, scientific-sounding, technical, to the point
# - Little general “waffle”/chit-chat
# - Not boring – because you don’t explain obvious things too much
# - Efficient delivery of (only) the facts that we need to know to understand/reimplement
# - Results visually well-presented and described with the correct priority of importance of sub-results
# - Insightful analysis – speculation should connect to something interesting and not be too much; the reader “learns something new”
# - No typos, no colloquialisms – well-considered language
# - This normally means several re-draftings (re-orderings of information)
# + [markdown] colab={} colab_type="code" id="uCINIXV1q1oe"
# # Iterative attention LSTMs
# +
def augment_example(ex):
tokens = ex.tokens
rand_num = random.uniform(0,1)
if rand_num < 0.0:
random.shuffle(tokens)
elif rand_num < 0.0:
# Two words switching their place
index_1 = random.randint(0,len(tokens)-1)
index_2 = random.randint(0,len(tokens)-1)
tmp = tokens[index_1]
tokens[index_1] = tokens[index_2]
tokens[index_2] = tmp
# for i in range(len(tokens)):
# if random.uniform(0,1) < 1.0/(len(tokens)):
# tokens[i] = "<unk>"
return tokens
def prepare_minibatch(mb, vocab, is_eval=False):
"""
Minibatch is a list of examples.
This function converts words to IDs and returns
torch tensors to be used as input/targets.
"""
batch_size = len(mb)
maxlen = max([len(ex.tokens) for ex in mb])
if not is_eval:
aug_ex = [augment_example(ex) for ex in mb]
else:
aug_ex = [ex.tokens for ex in mb]
# vocab returns 0 if the word is not there
x = [pad([vocab.w2i.get(t, 0) for t in ex_tokens], maxlen) for ex_tokens in aug_ex]
x = torch.LongTensor(x)
x = x.to(device)
y = [ex.label for ex in mb]
y = torch.LongTensor(y)
y = y.to(device)
return x, y
# -
def evaluate(model, data,
batch_fn=get_minibatch, prep_fn=prepare_minibatch,
batch_size=16):
"""Accuracy of a model on given data set (using minibatches)"""
correct = 0
total = 0
model.eval() # disable dropout
for mb in batch_fn(data, batch_size=batch_size, shuffle=False):
x, targets = prep_fn(mb, model.vocab, is_eval=True)
with torch.no_grad():
logits = model(x)
predictions = logits.argmax(dim=-1).view(-1)
# add the number of correct predictions to the total correct
correct += (predictions == targets.view(-1)).sum().item()
total += targets.size(0)
print("Evaluation on " + str(total) + " elements. Correct: " + str(correct))
return correct, total, correct / float(total)
def train_model(model, optimizer, num_iterations=10000,
print_every=1000, eval_every=1000,
batch_fn=get_examples,
prep_fn=prepare_example,
eval_fn=simple_evaluate,
batch_size=1, eval_batch_size=None):
"""Train a model."""
iter_i = 0
train_loss = 0.
print_num = 0
start = time.time()
criterion = nn.CrossEntropyLoss() # loss function
best_eval = 0.
best_iter = 0
# store train loss and validation accuracy during training
# so we can plot them afterwards
losses = []
accuracies = []
if eval_batch_size is None:
eval_batch_size = batch_size
while True: # when we run out of examples, shuffle and continue
for batch in batch_fn(train_data, batch_size=batch_size):
# forward pass
model.train()
x, targets = prep_fn(batch, model.vocab)
logits = model(x)
B = targets.size(0) # later we will use B examples per update
# compute cross-entropy loss (our criterion)
# note that the cross entropy loss function computes the softmax for us
loss = criterion(logits.view([B, -1]), targets.view(-1))
train_loss += loss.item()
# backward pass
# Tip: check the Introduction to PyTorch notebook.
# erase previous gradients
model.zero_grad()
loss.backward()
# update weights - take a small step in the opposite dir of the gradient
optimizer.step()
print_num += 1
iter_i += 1
# print info
if iter_i % print_every == 0:
print("Iter %r: loss=%.4f, time=%.2fs" %
(iter_i, train_loss, time.time()-start))
losses.append(train_loss)
print_num = 0
train_loss = 0.
# evaluate
if iter_i % eval_every == 0:
_, _, accuracy = eval_fn(model, dev_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
accuracies.append(accuracy)
print("iter %r: dev acc=%.4f" % (iter_i, accuracy))
# save best model parameters
if accuracy > best_eval:
print("new highscore")
best_eval = accuracy
best_iter = iter_i
path = "{}.pt".format(model.__class__.__name__)
ckpt = {
"state_dict": model.state_dict(),
"optimizer_state_dict": optimizer.state_dict(),
"best_eval": best_eval,
"best_iter": best_iter
}
torch.save(ckpt, path)
# done training
if iter_i == num_iterations:
print("Done training")
# evaluate on train, dev, and test with best model
print("Loading best model")
path = "{}.pt".format(model.__class__.__name__)
ckpt = torch.load(path)
model.load_state_dict(ckpt["state_dict"])
_, _, train_acc = eval_fn(
model, train_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
_, _, dev_acc = eval_fn(
model, dev_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
_, _, test_acc = eval_fn(
model, test_data, batch_size=eval_batch_size,
batch_fn=batch_fn, prep_fn=prep_fn)
print("best model iter {:d}: "
"train acc={:.4f}, dev acc={:.4f}, test acc={:.4f}".format(
best_iter, train_acc, dev_acc, test_acc))
return losses, accuracies
class AttentionModule(nn.Module):
"""Realizes a very simple attention module. Takes last state and word into account with two-layer feedforward"""
def __init__(self, embedding_dim, state_dim, hidden_dim):
super(AttentionModule, self).__init__()
self.embedding_dim = embedding_dim
self.hidden_dim = hidden_dim
self.state_dim = state_dim
self.output_layer = nn.Sequential(OrderedDict([
('fc1', nn.Linear(embedding_dim+state_dim, hidden_dim)),
('tanh', nn.Tanh()),
('dropout', nn.Dropout(0.5)),
('fc2', nn.Linear(hidden_dim, 1)),
# ('fc', nn.Linear(embedding_dim+state_dim, 1)),
('sigmoid', nn.Sigmoid())
]))
self.reset_parameters()
def reset_parameters(self):
"""This is PyTorch's default initialization method"""
stdv = 1.0 / math.sqrt(self.hidden_dim)
for weight in self.parameters():
weight.data.uniform_(-stdv, stdv)
def forward(self, word_embed, last_state):
input_tensor = torch.cat([word_embed, last_state], dim=1)
for layer in self.output_layer:
input_tensor = layer(input_tensor)
return input_tensor
class IterativeLSTMClassifier(nn.Module):
"""Encodes sentence with an LSTM and projects final hidden state"""
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, vocab, iterations):
super(IterativeLSTMClassifier, self).__init__()
self.vocab = vocab
self.hidden_dim = hidden_dim
self.embedding_size = embedding_dim
self.embed = nn.Embedding(vocab_size, embedding_dim, padding_idx=1)
self.rnn = MyLSTMCell(embedding_dim, hidden_dim)
self.attention_module = AttentionModule(embedding_dim, hidden_dim, embedding_dim)
self.iterations = iterations
self.output_layer = nn.Sequential(
nn.Dropout(p=0.5),
nn.Linear(hidden_dim, output_dim)
)
self.state_dropout = nn.Dropout(p=0.0)
def forward(self, x):
B = x.size(0) # batch size (this is 1 for now, i.e. 1 single example)
T = x.size(1) # time (the number of words in the sentence)
input_ = self.embed(x)
# here we create initial hidden states containing zeros
# we use a trick here so that, if input is on the GPU, then so are hx and cx
hx = input_.new_zeros(B, self.rnn.hidden_size)
cx = input_.new_zeros(B, self.rnn.hidden_size)
# process input sentences one word/timestep at a time
# input is batch-major, so the first word(s) is/are input_[:, 0]
for iter_ind in range(self.iterations):
if iter_ind > -1:
# hx has dimension B,hidden_size -> need to be B,T,hidden_size
exp_hx = hx.unsqueeze(dim=1).repeat([1,T,1])
attent_hx_input = exp_hx.view((B * T, self.rnn.hidden_size))
attent_word_input = input_.view((B * T, self.embedding_size))
attention_scores = self.attention_module(attent_word_input, attent_hx_input)
attention_scores = attention_scores.reshape(B,T,1)
outputs = []
for i in range(T):
hx = self.state_dropout(hx)
cx = self.state_dropout(cx)
hx_new, cx_new = self.rnn(input_[:, i], (hx, cx))
if iter_ind == -1:
hx = hx_new
cx = cx_new
else:
hx = attention_scores[:, i] * hx_new + (1 - attention_scores[:, i]) * hx
cx = attention_scores[:, i] * cx_new + (1 - attention_scores[:, i]) * cx
# print(hx.shape)
# print(cx.shape)
hx
outputs.append(hx)
# if we have a single example, our final LSTM state is the last hx
if B == 1:
last_final_states = hx
else:
#
# This part is explained in next section, ignore this else-block for now.
#
# we processed sentences with different lengths, so some of the sentences
# had already finished and we have been adding padding inputs to hx
# we select the final state based on the length of each sentence
# two lines below not needed if using LSTM form pytorch
outputs = torch.stack(outputs, dim=0) # [T, B, D]
outputs = outputs.transpose(0, 1).contiguous() # [B, T, D]
# to be super-sure we're not accidentally indexing the wrong state
# we zero out positions that are invalid
pad_positions = (x == 1).unsqueeze(-1)
outputs = outputs.contiguous()
outputs = outputs.masked_fill_(pad_positions, 0.)
mask = (x != 1) # true for valid positions [B, T]
lengths = mask.sum(dim=1) # [B, 1]
indexes = (lengths - 1) + torch.arange(B, device=x.device, dtype=x.dtype) * T
last_final_states = outputs.view(-1, self.hidden_dim)[indexes] # [B, D]
hx = last_final_states
cx = input_.new_zeros(B, self.rnn.hidden_size)
if iter_ind == 0:
# we use the last hidden state to classify the sentence
logits = self.output_layer(last_final_states) / self.iterations
else:
logits += self.output_layer(last_final_states) / self.iterations
return logits
# +
lstm_model = IterativeLSTMClassifier(
len(v.w2i), 300, 168, len(t2i), v, 1) # 168
# # copy pre-trained vectors into embeddings table
with torch.no_grad():
lstm_model.embed.weight.data.copy_(torch.from_numpy(vectors))
lstm_model.embed.weight.requires_grad = False
print(lstm_model)
print_parameters(lstm_model)
lstm_model = lstm_model.to(device)
batch_size = 4096
optimizer = optim.Adam(lstm_model.parameters(), lr=2e-4)
lstm_losses, lstm_accuracies = train_model(
lstm_model, optimizer, num_iterations=30000,
print_every=250, eval_every=50,
batch_size=batch_size,
batch_fn=get_minibatch,
prep_fn=prepare_minibatch,
eval_fn=evaluate)
# -
plt.plot(lstm_losses)
plt.plot(lstm_accuracies)
| 93,236 |
/teaching/DLinMedicine20/Labs/lab1/jupyter_demo.ipynb | 6b40a4649c9c0db0b7e438773bf2f098b5d54273 | [
"MIT"
] | permissive | nhungle714/nhungle714.github.io | https://github.com/nhungle714/nhungle714.github.io | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 14,255 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Operations On Set - Part 2
# Tutorial By **Tech Blooded**
# ### Join Two Sets
# There are several ways to join two or more sets in Python.
# We can use the **union()** method that returns a new set containing all items from both sets,
#
# or
#
# the **update()** method that inserts all the items from one set into another
a = {1,2,3}
b = {4,5,6}
c = a.union(b)
print(c)
a.update(b)
print(a)
# Both union() and update() will exclude any duplicate items.
# ### Keep ONLY the Duplicates - Intersection
# The **intersection_update()** method will keep only the items that are present in both sets.
print(a)
print(b)
b.intersection_update(a)
print(b)
# The **intersection()** method will return a new set, that only contains the items that are present in both sets.
c = a.intersection(b)
print(c)
# ### Keep All, But NOT the Duplicates
# The **symmetric_difference_update()** method will keep only the elements that are NOT present in both sets.
a = {1,2,3,4}
b = {3,4,5,6}
a.symmetric_difference_update(b)
print(a)
# The **symmetric_difference()** method will return a new set, that contains only the elements that are NOT present in both sets.
a = {1,2,3,4}
b = {3,4,5,6}
c = a.symmetric_difference(b)
print(c)
# # Thank You
| 1,517 |
/_notebooks/2021-06-18-bitcoin-technical-analysis.ipynb | 8a12a2bd22c0cd57b0a74619e972b8988704cb01 | [
"Apache-2.0"
] | permissive | namanmanchanda09/mindAI | https://github.com/namanmanchanda09/mindAI | 1 | 0 | Apache-2.0 | 2021-05-27T16:11:09 | 2021-05-27T16:04:37 | Jupyter Notebook | Jupyter Notebook | false | false | .py | 306,023 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.019153, "end_time": "2021-06-18T11:02:33.586946", "exception": false, "start_time": "2021-06-18T11:02:33.567793", "status": "completed"}
# <center>
# <div class="alert alert-warning">
# <div>
# <h2 style="color:purple">Bitcoin Technical Analysis</h2>
# </div>
# </div>
# </center>
# + [markdown] papermill={"duration": 0.017734, "end_time": "2021-06-18T11:02:33.623484", "exception": false, "start_time": "2021-06-18T11:02:33.605750", "status": "completed"}
# **Introdution**
#
# This notebook includes conducting a technical analysis on Bitcoin Data. Cryptocurrency markets are highly volatile and although, 70-80% of it is based upon sentiments and social media, an extensive technical analysis might provide some insights into how the trend works. For now, 2 datasets have been used, one of which contains 1-minute data of Bitcoin and the other one contains daily Bitcoin data.
#
# The technical analysis metrics consists of
#
# 1. `Moving Averages` - This metric can be used to get a decent idea of the asset's established trend. This metric can't be used to predict future trends.
#
# 2. `Average True Range` - It is the average of the true ranges over the specified period. It measures the volatility in the market. A typical ATR calculation is of 14 days.
#
# 3. `Relative Strength Index` - The tool measures the strength of the dynamics and the trend of the digital asset. In other terms, it can be used to predict the momentum and the divergence of the asset.
#
# 4. `Bollinger Bands` - It is useful to in measuring volatility and trends. The upper band is 2 standard deviation above mean and the lower band is 2 standard deviation below mean.
# + [markdown] papermill={"duration": 0.017715, "end_time": "2021-06-18T11:02:33.660076", "exception": false, "start_time": "2021-06-18T11:02:33.642361", "status": "completed"}
# Table of Contents:
#
# 1. [Imports](#1)
# - 1.1 [Packages](#2)
# - 1.2 [Dataset](#3)
# 2. [Understanding data](#4)
# - 2.1 [The 1-min data](#5)
# - 2.2 [Daily data](#6)
# - 2.3 [Heads after preprocessing](#7)
# 3. [Analysing Daily data](#8)
# - 3.1 [Candlestick plot](#9)
# - 3.2 [Moving Average](#10)
# - 3.3 [Average True Range](#11)
# + [markdown] papermill={"duration": 0.018194, "end_time": "2021-06-18T11:02:33.696200", "exception": false, "start_time": "2021-06-18T11:02:33.678006", "status": "completed"}
# ### 1. Imports <a id=1></a>
# + [markdown] papermill={"duration": 0.017639, "end_time": "2021-06-18T11:02:33.731517", "exception": false, "start_time": "2021-06-18T11:02:33.713878", "status": "completed"}
# #### 1.1 Packages <a id=2></a>
# + _kg_hide-input=true papermill={"duration": 2.143634, "end_time": "2021-06-18T11:02:35.892945", "exception": false, "start_time": "2021-06-18T11:02:33.749311", "status": "completed"}
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import plotly.express as px
from termcolor import colored
import matplotlib.pyplot as plt
import plotly.graph_objects as go
warnings.filterwarnings("ignore")
from plotly.subplots import make_subplots
print('Packages successfully imported...')
# + [markdown] papermill={"duration": 0.01922, "end_time": "2021-06-18T11:02:35.931008", "exception": false, "start_time": "2021-06-18T11:02:35.911788", "status": "completed"}
# #### 1.2 Dataset <a id=3></a>
# + _kg_hide-input=true papermill={"duration": 2.115261, "end_time": "2021-06-18T11:02:38.065649", "exception": false, "start_time": "2021-06-18T11:02:35.950388", "status": "completed"}
# The 1-minute data for Bitcoin
df = pd.read_parquet('/kaggle/input/binance-full-history/BTC-USDT.parquet')
# The 1-day data for Bitcoin
daily = pd.read_csv('../input/cryptocurrency-data/BTC-USD-2.csv', parse_dates = [0])
print('Data successfully imported...')
# + [markdown] papermill={"duration": 0.018982, "end_time": "2021-06-18T11:02:38.104151", "exception": false, "start_time": "2021-06-18T11:02:38.085169", "status": "completed"}
# ### 2. Understanding Data <a id=4></a>
# + [markdown] papermill={"duration": 0.019835, "end_time": "2021-06-18T11:02:38.142859", "exception": false, "start_time": "2021-06-18T11:02:38.123024", "status": "completed"}
# #### 2.1 The 1-min data <a id=5></a>
# + [markdown] papermill={"duration": 0.018325, "end_time": "2021-06-18T11:02:38.180805", "exception": false, "start_time": "2021-06-18T11:02:38.162480", "status": "completed"}
# ##### 2.1.1 Head
# + _kg_hide-input=true papermill={"duration": 0.045444, "end_time": "2021-06-18T11:02:38.244691", "exception": false, "start_time": "2021-06-18T11:02:38.199247", "status": "completed"}
df.head()
# + [markdown] _kg_hide-input=false papermill={"duration": 0.01902, "end_time": "2021-06-18T11:02:38.282809", "exception": false, "start_time": "2021-06-18T11:02:38.263789", "status": "completed"}
# ##### 2.1.2 Preprocessing
# + _kg_hide-input=true papermill={"duration": 0.067978, "end_time": "2021-06-18T11:02:38.370019", "exception": false, "start_time": "2021-06-18T11:02:38.302041", "status": "completed"}
df.index.rename('date',inplace=True)
df.drop(['quote_asset_volume','taker_buy_base_asset_volume','taker_buy_quote_asset_volume'],inplace = True,axis=1)
# + [markdown] papermill={"duration": 0.018547, "end_time": "2021-06-18T11:02:38.407801", "exception": false, "start_time": "2021-06-18T11:02:38.389254", "status": "completed"}
# ##### 2.1.3 Checking Null Values
# + _kg_hide-input=true papermill={"duration": 0.051876, "end_time": "2021-06-18T11:02:38.478942", "exception": false, "start_time": "2021-06-18T11:02:38.427066", "status": "completed"}
df.isnull().sum()
# + [markdown] papermill={"duration": 0.019124, "end_time": "2021-06-18T11:02:38.518048", "exception": false, "start_time": "2021-06-18T11:02:38.498924", "status": "completed"}
# #### 2.2 Daily Data <a id=6></a>
# + [markdown] papermill={"duration": 0.018602, "end_time": "2021-06-18T11:02:38.556157", "exception": false, "start_time": "2021-06-18T11:02:38.537555", "status": "completed"}
# ##### 2.2.1 Head
# + _kg_hide-input=true papermill={"duration": 0.039452, "end_time": "2021-06-18T11:02:38.614905", "exception": false, "start_time": "2021-06-18T11:02:38.575453", "status": "completed"}
daily.head()
# + [markdown] papermill={"duration": 0.018979, "end_time": "2021-06-18T11:02:38.653882", "exception": false, "start_time": "2021-06-18T11:02:38.634903", "status": "completed"}
# ##### 2.2.2 Set date as index
# + _kg_hide-input=true papermill={"duration": 0.027914, "end_time": "2021-06-18T11:02:38.701068", "exception": false, "start_time": "2021-06-18T11:02:38.673154", "status": "completed"}
daily.set_index('Date',inplace=True)
# + [markdown] papermill={"duration": 0.018998, "end_time": "2021-06-18T11:02:38.739579", "exception": false, "start_time": "2021-06-18T11:02:38.720581", "status": "completed"}
# ##### 2.2.3 Checking for null values
# + _kg_hide-input=true papermill={"duration": 0.030879, "end_time": "2021-06-18T11:02:38.790199", "exception": false, "start_time": "2021-06-18T11:02:38.759320", "status": "completed"}
daily.isnull().sum()
# + [markdown] papermill={"duration": 0.020014, "end_time": "2021-06-18T11:02:38.829992", "exception": false, "start_time": "2021-06-18T11:02:38.809978", "status": "completed"}
# ##### 2.2.4 Removing null values
# + _kg_hide-input=true papermill={"duration": 0.035456, "end_time": "2021-06-18T11:02:38.885491", "exception": false, "start_time": "2021-06-18T11:02:38.850035", "status": "completed"}
daily.fillna(method='ffill',inplace=True, axis=0)
daily.isnull().sum()
# + [markdown] papermill={"duration": 0.01975, "end_time": "2021-06-18T11:02:38.925547", "exception": false, "start_time": "2021-06-18T11:02:38.905797", "status": "completed"}
# #### 2.3 Heads after preprocessing <a id=7></a>
# + [markdown] papermill={"duration": 0.020535, "end_time": "2021-06-18T11:02:38.968870", "exception": false, "start_time": "2021-06-18T11:02:38.948335", "status": "completed"}
# ##### 2.3.1 Daily data
# + _kg_hide-input=true papermill={"duration": 0.035515, "end_time": "2021-06-18T11:02:39.024507", "exception": false, "start_time": "2021-06-18T11:02:38.988992", "status": "completed"}
daily.head()
# + [markdown] papermill={"duration": 0.020989, "end_time": "2021-06-18T11:02:39.066728", "exception": false, "start_time": "2021-06-18T11:02:39.045739", "status": "completed"}
# ##### 2.3.2 1-minute data
# + _kg_hide-input=true papermill={"duration": 0.03846, "end_time": "2021-06-18T11:02:39.127244", "exception": false, "start_time": "2021-06-18T11:02:39.088784", "status": "completed"}
df.head()
# + [markdown] papermill={"duration": 0.0211, "end_time": "2021-06-18T11:02:39.169075", "exception": false, "start_time": "2021-06-18T11:02:39.147975", "status": "completed"}
# ### 3. Analysing Daily data <a id=8></a>
# + [markdown] papermill={"duration": 0.020986, "end_time": "2021-06-18T11:02:39.211010", "exception": false, "start_time": "2021-06-18T11:02:39.190024", "status": "completed"}
# #### 3.1 Candlestick plot <a id=9></a>
# + _kg_hide-input=true papermill={"duration": 1.095236, "end_time": "2021-06-18T11:02:40.327800", "exception": false, "start_time": "2021-06-18T11:02:39.232564", "status": "completed"}
fig = go.Figure(data=[go.Candlestick(x=daily.index,
open=daily['Open'],
high=daily['High'],
low=daily['Low'],
close=daily['Close'])],
layout = go.Layout({
'title':{
'text':'Bitcoin Daily data',
'font':{
'size':25
}
}
})
)
fig.update_layout({"template":"plotly_dark"})
fig.show()
# + [markdown] papermill={"duration": 0.021649, "end_time": "2021-06-18T11:02:40.372042", "exception": false, "start_time": "2021-06-18T11:02:40.350393", "status": "completed"}
# #### 3.2 Moving Averages <a id=10></a>
# + [markdown] papermill={"duration": 0.023062, "end_time": "2021-06-18T11:02:40.417272", "exception": false, "start_time": "2021-06-18T11:02:40.394210", "status": "completed"}
# $$
# MA = 1/n * \sum_{n=x_i}^{x_n} Prices
# $$
#
# Moving average cuts down the noise in the price chart. The *direction* of the `MA` might give an idea of the trend of the chart. A subtle observation can be made from the plot that is whenever the candlestick cuts the `MA` threshold level to rise above it, the price is likely to go high for some duration of time before which there's a change in direction of price to due to other factors. In the same way, when the candlestick cuts the `MA` threshold level to rise below it, the price is likely to fall for some duration. At the rising cut-point, a trader can enter the market and at the falling cut-point, a trader might sell. This is also called as a crossover.
# + _kg_hide-input=true papermill={"duration": 0.101311, "end_time": "2021-06-18T11:02:40.540897", "exception": false, "start_time": "2021-06-18T11:02:40.439586", "status": "completed"}
avg_20 = daily.Close.rolling(window=20, min_periods=1).mean()
avg_50 = daily.Close.rolling(window=50, min_periods=1).mean()
avg_100 = daily.Close.rolling(window=100, min_periods=1).mean()
set_1 = {
'x':daily.index,
'open': daily.Open,
'close':daily.Close,
'high':daily.High,
'low':daily.Low,
'type':'candlestick'
}
set_2 = {
'x':daily.index,
'y':avg_20,
'type':'scatter',
'mode':'lines',
'line':{
'width':2,
'color':'white'
},
'name':'Moving average of 20 days'
}
set_3 = {
'x':daily.index,
'y':avg_50,
'type':'scatter',
'mode':'lines',
'line':{
'width':2,
'color':'yellow'
},
'name':'Moving average of 50 days'
}
set_4 = {
'x':daily.index,
'y':avg_100,
'type':'scatter',
'mode':'lines',
'line':{
'width':2,
'color':'orange'
},
'name':'Moving average of 100 days'
}
data = [set_1, set_2, set_3, set_4]
layout = go.Layout({
'title':{
'text':'Moving averages',
'font':{
'size':25
}
}
})
fig = go.Figure(data=data, layout=layout)
fig.update_layout({"template":"plotly_dark"})
fig.show()
# + [markdown] papermill={"duration": 0.023842, "end_time": "2021-06-18T11:02:40.592101", "exception": false, "start_time": "2021-06-18T11:02:40.568259", "status": "completed"}
# #### 3.3 Average True Range <a id=11></a>
# + [markdown] papermill={"duration": 0.023558, "end_time": "2021-06-18T11:02:40.639731", "exception": false, "start_time": "2021-06-18T11:02:40.616173", "status": "completed"}
# $$
# TR = MAX([(H-L), Abs(H-Cp), Abs(L-Cp)])
# $$
#
#
#
# $$
# ATR = 1/n * \sum_{n=x_i}^{x_n} TR
# $$
#
# The true range indicator is taken as the greatest of the following: current high less the current low; the absolute value of the current high less the previous close; and the absolute value of the current low less the previous close. The ATR is then a moving average, generally using 14 days, of the true ranges. Simply put, a stock experiencing a high level of volatility has a higher ATR, and a low volatility stock has a lower ATR.
#
#
# + _kg_hide-input=true papermill={"duration": 0.268978, "end_time": "2021-06-18T11:02:40.933705", "exception": false, "start_time": "2021-06-18T11:02:40.664727", "status": "completed"}
high_low = daily['High'] - daily['Low']
high_close = np.abs(daily['High'] - daily['Close'].shift())
low_close = np.abs(daily['Low'] - daily['Close'].shift())
ranges = pd.concat([high_low, high_close, low_close], axis=1)
true_range = np.max(ranges, axis=1)
atr = true_range.rolling(14).sum()/14
atr_daily = daily
atr_daily['atr'] = atr
atr_daily['High_atr'] = atr_daily['High'] + atr_daily['atr']
atr_daily['Low_atr'] = atr_daily['Low'] - atr_daily['atr']
fig = make_subplots(rows=2, cols=1, row_heights=[0.5, 0.5], vertical_spacing=.3, subplot_titles=["Bitcoin Daily Data", "Average True Range"])
fig.append_trace(go.Candlestick(x=daily.index,
open=atr_daily['Open'],
high=atr_daily['High'],
low=atr_daily['Low'],
close=atr_daily['Close']),
row=1,col=1)
fig.append_trace(go.Scatter(x=atr_daily.index,
y=atr_daily.atr,
line_color='Orange'
),row=2,col=1)
fig.update_layout(
margin=dict(l=20, r=20, t=20, b=20),
)
fig.update_layout({"template":"plotly_dark"})
fig.show()
# + [markdown] papermill={"duration": 0.025508, "end_time": "2021-06-18T11:02:40.985732", "exception": false, "start_time": "2021-06-18T11:02:40.960224", "status": "completed"}
# ##### 3.3.1 ATR Significance
#
# + [markdown] papermill={"duration": 0.02541, "end_time": "2021-06-18T11:02:41.037108", "exception": false, "start_time": "2021-06-18T11:02:41.011698", "status": "completed"}
# Though ATR doesn't give much of an idea about the trend of the chart, but it can be used by traders to put stop losses. An ATR value above `close` or `high` can be used as a potential exit point. An ATR value below `open` or `low` can be used as a potential exit point as well in case of down fall of the price.
# + _kg_hide-input=true papermill={"duration": 0.090558, "end_time": "2021-06-18T11:02:41.153252", "exception": false, "start_time": "2021-06-18T11:02:41.062694", "status": "completed"}
set_1 = go.Candlestick(x=daily.index,
open=atr_daily['Open'],
high=atr_daily['High'],
low=atr_daily['Low'],
close=atr_daily['Close'])
set_2 = go.Scatter(x=atr_daily.index,
y=atr_daily.High_atr,
line_color='Orange')
set_3 = go.Scatter(x=atr_daily.index,
y=atr_daily.Low_atr,
line_color='Yellow')
data = [set_1, set_2, set_3]
layout = go.Layout({
'title':{
'text':'Upper/Lower limit of ATR values for trading',
'font':{
'size':25
}
}
})
fig = go.Figure(data=data, layout=layout)
fig.update_layout({"template":"plotly_dark"})
fig.show()
# + papermill={"duration": 0.027238, "end_time": "2021-06-18T11:02:41.209061", "exception": false, "start_time": "2021-06-18T11:02:41.181823", "status": "completed"}
| 16,547 |
/School/2. Basic Practice.ipynb | b80ecfddf8011ceceb7006b4f865e104c18f9c9b | [] | no_license | jameslee5656/WebScraping | https://github.com/jameslee5656/WebScraping | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 4,901 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding:utf-8 -*-
import requests
from bs4 import BeautifulSoup as bsp
resp = requests.get('http://www.ntpu.edu.tw/chinese/')
resp.encoding='big5'
soup = bsp(resp.text, 'html.parser')
lis = soup.find_all('a',{'class':'font-color-focus'})
lis += soup.find_all('a',{'class':'font-black-13'})
# print(soup.dt.text)
# -
divs = soup.find_all('div')
s = []
for div in divs:
s.append(x for x in div.stripped_strings)
print([x for x in s[0]])
resp2 = requests.get('http://lms.ntpu.edu.tw/')
resp2.encoding='utf-8'
soup2 = bsp(resp2.text,'html.parser')
rows = soup2.find('table','table').find_all('tr')
for row in rows:
content = row.find_all('td')
print(content[1].text, content[3].text)
print("\n")
# links = soup.find_all('a')
# for link in links:
# print(link.parent)
# print('2',link.parent.previous_sibling.text)
all_href = []
for row in rows:
#method 1: find_all('td')
tds = (row.find_all('td'))
#method 2: find row (<tr>)`s all direct children
# tds = ([td for td in row.children]) =>> became list type =>> not useful
# if 'herf' in tds[1].a:
# print(tds[3].a['href'])
# else:
# pass
# all_href.append(tds)
ext(texto_procesado, lowercase=True, no_punct=True)
doc = textacy.Doc(texto_procesado)
list(textacy.extract.ngrams(doc, 3, filter_stops=True, filter_punct=True, filter_nums=False))
list(textacy.extract.ngrams(doc, 2, min_freq=2))
list(textacy.extract.named_entities(doc, drop_determiners=True))
def encontrar_personajes(doc):
"""
Devuelve una lista de los personajes de un `doc` con su cantidad de
ocurrencias
:param doc: NLP documento parseado por Spacy
:return: Lista de Tuplas con la forma
[('winston', 686), ("o'brien", 135), ('julia', 85),]
"""
personajes = Counter()
for ent in doc.ents:
if ent.label_ == 'PERSON':
personajes[ent.lemma_] += 1
return personajes.most_common()
# Extrayendo los personajes principales del texto y contando cuantas veces
# son nombrados.
print(encontrar_personajes(doc)[:20])
| 2,364 |
/Chapter_9.ipynb | 037b57e041cbb6f8e0d0ec807c06bb57ba269044 | [] | no_license | Mohsina-Jannat/Bioinformatics_Rosalind | https://github.com/Mohsina-Jannat/Bioinformatics_Rosalind | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 12,498 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="niaJFwj4QCph" colab_type="code" outputId="e063bb04-5a86-4406-df58-e09c28784626" colab={"base_uri": "https://localhost:8080/", "height": 34}
#ba9i
words = "GCGTGCCTGGTCA$"
a = words
list = []
str = ""
for i in range(len(words)):
word = a[-1] + a[:-1]
new = ''.join(word)
a = new
list.append(new)
i += 1
list.sort()
for i in range(len(list)):
str += list[i][-1]
print(str)
# + id="TyCfle0EQrUN" colab_type="code" outputId="5ed3ba04-b4d1-4cd4-86b2-f1dc37d71ba7" colab={"base_uri": "https://localhost:8080/", "height": 34}
#ba9j
bwt = "TTCCTAACG$A"
lst = sorted(bwt)
lst1 = [('$',1),('A',1),('A',2),('A',3),('C',1),('C',2),('C',3),('G',1),('T',1),('T',2),('T',3)]
lst2 = [('T',1),('T',2),('C',1),('C',2),('T',3),('A',1),('A',2),('C',3),('G',1),('$',1),('A',3)]
dct = {('$',1):('T',1),('A',1):('T',2),('A',2):('C',1),('A',3):('C',2),('C',1):('T',3),('C',2):('A',1),('C',3):('A',2),('G',1):('C',3),('T',1):('G',1),('T',2):('$',1),('T',3):('A',3)}
p = dct[('$',1)]
ans = []
str = ""
for i in range(len(dct)):
if p == ('$',1):
break
str += p[0]
p = dct[p]
print(str[::-1])
# + id="9mDpOFVvYFZf" colab_type="code" outputId="0c29f16f-b090-4173-b3c5-29e7bcdc0cc7" colab={"base_uri": "https://localhost:8080/", "height": 34}
#ba9h
text = "AATCGGGTTCAATCGGGGT"
p = ["ATCG","GGGT"]
pos = []
def suffixes(string):
for idx in range(len(string)):
yield string[idx:]
def create_suffix_array(text):
d = {}
for idx, suffix in enumerate(suffixes(text)):
d[suffix] = idx
#return OrderedDict(sorted(d.items()))
#return sorted(d.items())
for item in p:
k = len(item)
for key,value in d.items():
if key[0:k] == item:
pos.append(value)
print(sorted(pos))
create_suffix_array(text)
# + id="1y7MiltgQ-hR" colab_type="code" outputId="462b5c02-6d11-4307-c3d7-209839f95ae2" colab={"base_uri": "https://localhost:8080/", "height": 34}
#ba9k
a = 'T$GACCA'
indx = 3
map = {}
str = ''.join(sorted(a))
#print(str)
for i in range(len(str)):
if str[i] == '$':
map[str[i]] = 0
else:
map[str[i]] = i - 1
#print(map)
print(map[a[indx]])
# + id="cGZmZEtrRAD1" colab_type="code" outputId="6ad1773f-6549-43ac-ddac-3695bcdbdcd6" colab={"base_uri": "https://localhost:8080/", "height": 34}
#ba9n
s = "AATCGGGTTCAATCGGGGT"
s1 = 'ATCG'
s2 = 'GGGT'
lst = []
leng = len(s)
k = len(s1)
for i in range(leng-k+1):
if s[i:i+k] == s1 or s[i:i+k] == s2:
lst.append(i)
print(lst)
# + id="2qTwQhxFS1Og" colab_type="code" outputId="b42ece79-e774-40bf-d346-1a8faeed905b" colab={"base_uri": "https://localhost:8080/", "height": 34}
#ba9o
def hamming_dist(s1,s2):
cnt = 0
for c1,c2 in zip(s1,s2):
if c1!=c2:
cnt += 1
return cnt
s = "ACATGCTACTTT"
p = ["ATT","GCC","GCTA","TATT"]
d = 1
pos = []
leng = len(s)
for i in range(len(p)):
k = len(p[i])
for j in range(leng-k+1):
ss = s[j:j+k]
dist = hamming_dist(ss,p[i])
if dist <= d:
pos.append(j)
pos.sort()
print(pos)
# + id="pWC4M4ABS_XQ" colab_type="code" outputId="05c8a2ea-fb68-46f1-c28f-9bff2bb791d9" colab={"base_uri": "https://localhost:8080/", "height": 54}
#BA9G
text = "PANAMABANANAS$"
def suffixes(text):
for i in range(len(text)):
yield text[i:]
def suffix_array(text):
d = {}
for i,item in enumerate(suffixes(text)):
d[item] = i
lst = sorted(d.items())
return lst
print(suffix_array(text))
# lst = suffix_array(text)
# for i in range(len(lst)):
# if lst[i][1]%5 == 0:
# print(i,lst[i][1])
# + id="Njy6KRSS1b0y" colab_type="code" outputId="4428c95f-3a92-437e-f16e-0dee1805a0db" colab={"base_uri": "https://localhost:8080/", "height": 105}
#BA9q
text = "PANAMABANANAS$"
def suffixes(text):
for i in range(len(text)):
yield text[i:]
def suffix_array(text):
d = {}
for i,item in enumerate(suffixes(text)):
d[item] = i
lst = sorted(d.items())
return lst
print(suffix_array(text))
lst = suffix_array(text)
for i in range(len(lst)):
if lst[i][1]%5 == 0:
print(i,lst[i][1])
# + id="-AdSLNeh465q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="b9278eb1-cd61-45ad-ab0e-23708f778a65"
#ba9a
patterns = ["ATAGA","ATC","GAT"]
tree = []
tree.append({})
cnt = 1
for pattern in patterns:
current_node = 0
for char in pattern:
if char not in tree[current_node]:
tree[current_node][char] = cnt
print(str(current_node) + "->" + str(cnt) + ":" + char)
tree.append({})
cnt += 1
current_node = tree[current_node][char]
# + id="r3Rf7UN8UX5i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2064c2fd-1627-4cf1-eb24-35a047408e4e"
#ba9b
text = "AATCGGGTTCAATCGGGGT"
patterns = ["ATCG","GGGT"]
def constructTrieFrom(patterns):
tree = []
tree.append({})
cnt = 1
for pattern in patterns:
current_node = 0
for ch in pattern:
if ch not in tree[current_node]:
tree[current_node][ch] = cnt
tree.append({})
cnt += 1
current_node = tree[current_node][ch]
return tree
trie = constructTrieFrom(patterns)
ans = []
for i in range(len(text)):
pos = i
current_node = 0
ch = text[pos]
while(ch in trie[current_node] and len(trie[current_node]) != 0):
current_node = trie[current_node][ch]
pos += 1
if pos == len(text):
break
ch = text[pos]
if len(trie[current_node]) == 0:
ans.append(i)
print(*ans)
# + id="xmY27OhjUgOj" colab_type="code" colab={}
| 5,682 |
/AlgorithmicToolbox/JdbelloroAlgorithmicToolboxCertificate.ipynb | f733e14e455af56a6bdc4baf4951953ecd9d629b | [] | no_license | jdbelloro/AlgorithmsUN2021I | https://github.com/jdbelloro/AlgorithmsUN2021I | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 117,567 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="NIu0iMGYVuDm"
# #Coursera Certificate for Algorithmic Toolbox - University of California San Diego - HSE Moscow (National Research University Higher School of Economics
# + [markdown] id="X-v7imIhV2Ja"
# ##Linkedin Link:
# + [markdown] id="4WZ8t9jHWGvi"
# https://www.linkedin.com/feed/update/urn:li:activity:6828874604510109696/
# + [markdown] id="JVrS8lbzWCmY"
# ##Screenshot:
# 
| 117,068 |
/model/RecSys_0.2.ipynb | fa1c34068664644b1e696adcd33d9b7379e587a4 | [
"MIT"
] | permissive | zxzhaixiang/QMDB | https://github.com/zxzhaixiang/QMDB | 2 | 0 | MIT | 2022-12-10T06:20:42 | 2019-12-09T17:48:53 | Jupyter Notebook | Jupyter Notebook | false | false | .py | 1,282,457 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="WyzSN_bDVkZA" outputId="a4ca0d49-8469-4ccb-ab50-f5900bd0bf21" colab={"base_uri": "https://localhost:8080/", "height": 163}
# !git clone https://github.com/zxzhaixiang/QMDB
# + colab_type="code" id="IXTnOznGbXHT" colab={}
datapath = './QMDB/data/processed_data/'
# + colab_type="code" id="dyTFcL_uzaBX" outputId="bdc553d8-08da-4987-92cf-d959406cdd8c" colab={"base_uri": "https://localhost:8080/", "height": 90}
# !pip install hiddenlayer
# + colab_type="code" id="m-S0kpbEV5IX" outputId="f096735a-cbfa-4b24-8dc8-6ccba7861e4f" colab={"base_uri": "https://localhost:8080/", "height": 74}
import os
import numpy as np
import matplotlib.pyplot as plt
import time
import sys
import pickle
import random
import scipy.signal as sp_signal
import pandas as pd
import seaborn as sns
from datetime import datetime
import re
from imblearn.over_sampling import RandomOverSampler
from sklearn import decomposition
from sklearn.preprocessing import StandardScaler
from sklearn.manifold import TSNE
import scipy.stats
from IPython.display import HTML, display
import hiddenlayer as hl
# %matplotlib inline
# + colab_type="code" id="0nJY2mv4WB3P" colab={}
import torch
from torch import nn
from torch.utils.data import Dataset, DataLoader, TensorDataset
from torch.utils.data.sampler import SubsetRandomSampler, WeightedRandomSampler
from torch.autograd import Variable
from torchvision import transforms
# + colab_type="code" id="yu1GAW9wWJOv" outputId="bdfa8a48-14fd-4ee7-f760-3bfd082ffdf4" colab={"base_uri": "https://localhost:8080/", "height": 35}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print("Device is ", device)
# + [markdown] colab_type="text" id="EMut62-EqdhV"
# ## Loading data and prepare data loader
# + colab_type="code" id="xMKqS3zzvoRe" colab={}
def get_date_time_info(timestamps):
dt_objs = [datetime.fromtimestamp(ts) for ts in timestamps]
is_weekend = [1 if dt_obj.weekday()>=5 else 0 for dt_obj in dt_objs]
hr_of_day = [dt_obj.hour for dt_obj in dt_objs]
return is_weekend, hr_of_day
# + colab_type="code" id="j0bOPwl-dSr0" outputId="65685ab7-11ba-45b8-f415-2ffb2460ad1a" colab={"base_uri": "https://localhost:8080/", "height": 35}
print('loading raw rating data..',end='')
rating_df = pd.read_csv(os.path.join(datapath,'ratings_with_kws.csv'), compression='gzip')
#print('Done\n {} total rating record loaded\n'.format(rating_df.shape[0]))
userIds = rating_df['userId'].unique()
# + colab_type="code" id="GKHjFADWl1p3" colab={}
user_df = rating_df.groupby('userId', as_index=False).agg({
'movieId': [('num_movie_rated','count')],
'rating': [('user_ave_rating','mean'),('user_std_rating','std')]}).fillna(0)
user_df.columns.set_levels([item if item!='' else 'userId' for item in user_df.columns.levels[1]],level=1,inplace=True)
user_df.columns = user_df.columns.droplevel(0) #drop first level column name (old column name before aggregation)
# + colab_type="code" id="5Zz8CjCkmEED" colab={}
rating_by_movie_df = rating_df.groupby('movieId', as_index = False).agg({
'userId': [('num_rating', 'count')],
'rating': [('movie_ave_rating','mean'),('movie_std_rating', 'std')]})
rating_by_movie_df.columns.set_levels([item if item!='' else 'movieId' for item in rating_by_movie_df.columns.levels[1]],level=1,inplace=True)
rating_by_movie_df.columns = rating_by_movie_df.columns.droplevel(0)#drop first level column name (old column name before aggregation)
# + colab_type="code" id="cHlPoXLKdiW7" colab={}
movie_df = pd.read_csv(os.path.join(datapath,'keywords_all.csv'))
movie_df = movie_df.merge(rating_by_movie_df, how='left',on='movieId').reset_index()
#movie_df.drop(columns='genres',inplace=True)
movie_df['genre_kws'] = movie_df['genre_kws'].apply(lambda s: re.sub(r'\[|\]|\'| ', '', s).lower())
#movie_df['description_kws'] = movie_df['description_kws'].apply(lambda s: re.sub(r'\[|\]|\'| |"', '', s).lower())
#movie_df['cast_kws'] = movie_df['cast_kws'].apply(lambda s: re.sub(r'\[|\]|\'| ', '', s))
#genres = {a for s in movie_df['genres'] for a in s.split('|')}
#print('{} number of movie genres exist. They are '.format(len(genres)), genres)
#genres_dict = dict(zip(genres, range(len(genres))))
# + colab_type="code" id="8omwpdTMb20_" outputId="a8e77553-407d-42c8-c891-b5d795b9dcc6" colab={"base_uri": "https://localhost:8080/", "height": 55}
genres_list = {a.lower() for s in movie_df['genre_kws'] for a in s.split(',')}
genres_list = list(genres_list)
print('{} number of movie genres exist. They are '.format(len(genres_list)), genres_list)
genres_dict = dict(zip(genres_list, range(len(genres_list))))
# + colab_type="code" id="8N01e0TR6Qpw" colab={}
genres = np.zeros((movie_df.shape[0], len(genres_list)))
for irow, s in enumerate(movie_df['genre_kws']):
for j in [genres_dict[e] for e in s.split(',')]:
genres[irow,j]=1
# + id="rAt-rvmVOiki" colab_type="code" colab={}
with open(os.path.join(datapath,'keyword_embedding.pkl'), 'rb') as f:
kw_embedding = pickle.load(f)
# + id="OrhLrFeISE7E" colab_type="code" colab={}
kw_default_embedding = torch.zeros(1,1,64)
n=0
for key in kw_embedding:
kw_embedding[key] = torch.FloatTensor(kw_embedding[key]).unsqueeze(dim=0).unsqueeze(dim=0)
kw_default_embedding += kw_embedding[key]
n+=1
kw_default_embedding/=n
# + id="gJ8-GzE6PEsW" colab_type="code" colab={}
kw_set = set(kw_embedding.keys())
# + id="D2BfcZf3PPgp" colab_type="code" colab={}
kw_by_movie = [[kw for kw in kws.split(' ')] for kws in movie_df['kws_all']]
# + colab_type="code" id="3h6kyZV2jLBO" colab={}
nM = movie_df.shape[0]
nU = user_df.shape[0]
nR = rating_df['rating'].unique().shape[0]
nRecord = rating_df['rating'].shape[0]
# + colab_type="code" id="oiQAAmbIv-1C" colab={}
is_weekend, hr_of_day = get_date_time_info(rating_df['timestamp'].values)
# + colab_type="code" id="irK4xqn-jX0U" colab={}
U = torch.tensor(rating_df['userId'].values, dtype=torch.long).reshape(-1,1)
M = torch.tensor(rating_df['qmdbId'].values, dtype=torch.long).reshape(-1,1)
R = torch.tensor(rating_df['rating'].values, dtype=torch.float).reshape(-1,1)
G = torch.tensor(genres, dtype=torch.float)
# + colab_type="code" id="Cfb1coTFt4EY" colab={}
tensor_weekend = torch.tensor(is_weekend, dtype=torch.float).reshape(-1,1)
tensor_hr = torch.tensor(hr_of_day, dtype=torch.float).reshape(-1,1)
# + colab_type="code" id="cHHoWOLoPK_Q" colab={}
batch_size = 4096
validation_split = .1
shuffle_dataset = True
random_seed = 42
# + colab_type="code" id="GFvfSgChPNT4" colab={}
dataset_size = nRecord
indices = list(range(dataset_size))
split = int(np.floor(validation_split * dataset_size))
if shuffle_dataset :
np.random.seed(random_seed)
np.random.shuffle(indices)
train_idx, val_idx = indices[split:], indices[:split]
# + colab_type="code" id="bDm6phvmnnKX" colab={}
ros = RandomOverSampler(random_state=random_seed)
train_idx_resampled, _ = ros.fit_resample(np.array(train_idx).reshape(-1,1), (R[train_idx].numpy()*2).reshape(-1).astype(int))
train_idx_resampled = list(train_idx_resampled.reshape(-1))
# + colab_type="code" id="ui0jUTIKoJxA" outputId="a9c9b0fd-cdaf-493c-d5fe-77d9a72d045c" colab={"base_uri": "https://localhost:8080/", "height": 341}
plt.figure(figsize=(15,5))
plt.subplot(121)
sns.distplot(R[train_idx])
plt.subplot(122)
sns.distplot(R[train_idx_resampled])
# + colab_type="code" id="Xa-uP2GCpk1a" outputId="f8f0d1ed-147c-45f0-81b1-77255b7db0b9" colab={"base_uri": "https://localhost:8080/", "height": 35}
print('Check if resampled indices covers all the original indices: {}'.format(set(train_idx)==set(train_idx_resampled)))
# + colab_type="code" id="tXCjfNrDW7gy" colab={}
train_dataset = torch.utils.data.TensorDataset(
U[train_idx_resampled], M[train_idx_resampled], R[train_idx_resampled],
tensor_weekend[train_idx_resampled], tensor_hr[train_idx_resampled])
train_loader = DataLoader(
train_dataset, batch_size=batch_size, shuffle = True)
# + colab_type="code" id="FR-K0WiUW7ZY" colab={}
val_dataset = torch.utils.data.TensorDataset(
U[val_idx], M[val_idx], R[val_idx],
tensor_weekend[val_idx], tensor_hr[val_idx])
val_loader = DataLoader(
val_dataset, batch_size=batch_size, shuffle = True)
# + [markdown] colab_type="text" id="SLcT_MG5TaTb"
# ## Build model
# + colab_type="code" id="eoivviDu0Vrl" colab={}
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
# + colab_type="code" id="Yufd8ny9Tbnw" colab={}
class NCF(nn.Module):
def __init__(self, config):
super(NCF, self).__init__()
self.config = config
self.user_dim = config['nUsers']
self.movie_dim = config['nMovies']
self.genres_dim = config['nGenres']
self.g_latent_dim = config['genre_latent_dim']
self.u_latent_dim = config['user_latent_dim']
self.i_latent_dim = config['item_latent_dim']
self.query_dim = config['Query_latent_dim']
self.user_embedding = nn.Embedding(self.user_dim, self.u_latent_dim,
max_norm=1, scale_grad_by_freq = True)
self.movie_embedding = nn.Embedding(self.movie_dim, self.i_latent_dim,
max_norm=1, scale_grad_by_freq = True)
self.genres_layers = nn.Sequential(
nn.Linear(self.genres_dim, self.g_latent_dim)
)
self.fc_layers = nn.Sequential(
nn.Linear(self.u_latent_dim + self.g_latent_dim + self.query_dim + 2, self.u_latent_dim*2),
nn.Tanh(),
nn.Linear(self.u_latent_dim*2,self.u_latent_dim),
nn.Tanh(),
nn.Linear(self.u_latent_dim,self.u_latent_dim),
nn.Tanh())
self.cos_sim = nn.CosineSimilarity(dim=2, eps=1e-6)
def forward(self, user_id, movie_id, genres, em_query, weekend, hr):
user_em = self.user_embedding(user_id)
movie_em = self.movie_embedding(movie_id)
genres_em = self.genres_layers(genres)
total_em = torch.cat((user_em, genres_em, em_query, weekend.unsqueeze(dim=1), hr.unsqueeze(dim=1)), dim=2)
final_em = self.fc_layers(total_em)
return self.cos_sim(user_em,movie_em)*5, self.cos_sim(final_em,movie_em)*5
# + colab_type="code" id="3SJ-CGqLTeZ3" outputId="aaac6011-29ee-4550-d0f8-b6705533690d" colab={"base_uri": "https://localhost:8080/", "height": 35}
config = {'nUsers': nU, 'nMovies': nM,
'nGenres': len(genres_list),
'user_latent_dim': 128, 'item_latent_dim': 128,
'genre_latent_dim': 32, 'Query_latent_dim': 64
}
print(config)
# + colab_type="code" id="qXosorBOTg_u" colab={}
CF = NCF(config).to(device)
# + colab_type="code" id="sMXAhTZNTnzq" colab={}
CF_criterion = nn.MSELoss()
CF_optimizer = torch.optim.Adam(CF.parameters(), lr = 0.001, betas = (0.5, 0.999))
# + colab_type="code" id="0VuamkRtztlM" outputId="7c742c6b-f94b-4344-afe7-e210b7a29440" colab={"base_uri": "https://localhost:8080/", "height": 35}
count_parameters(CF)
# + colab_type="code" id="PXBQ5cPT1oE1" outputId="db2a84a4-a42b-469d-bdb0-ca6537c61ff8" colab={"base_uri": "https://localhost:8080/", "height": 1000}
hl.build_graph(CF, (torch.LongTensor(np.zeros((2,1))).to(device), #uids
torch.LongTensor(np.zeros((2,1))).to(device), #mids
torch.FloatTensor(np.zeros((2,1,len(genres_list)))).to(device), #gids
torch.FloatTensor(np.zeros((2,1,config['Query_latent_dim']))).to(device), #query
torch.FloatTensor(np.zeros((2,1))).to(device), #weekend
torch.FloatTensor(np.zeros((2,1))).to(device))) #hr
# + colab_type="code" id="_Tzj83p80aej" outputId="186a8437-4e30-4635-b75f-54fe64c029ef" colab={"base_uri": "https://localhost:8080/", "height": 90}
CF(torch.LongTensor(np.zeros((2,1))).to(device), #uids
torch.LongTensor(np.zeros((2,1))).to(device), #mids
torch.FloatTensor(np.zeros((2,1,len(genres_list)))).to(device), #gids
torch.FloatTensor(np.zeros((2,1,config['Query_latent_dim']))).to(device), #query
torch.FloatTensor(np.zeros((2,1))).to(device), #weekend
torch.FloatTensor(np.zeros((2,1))).to(device)) #h
# + [markdown] colab_type="text" id="Yso166D5Tsc9"
# ## Training
# + colab_type="code" id="BjkB6kWpTtt7" colab={}
class progress():
def __init__(self,max = 100):
self.mrange = max
self.pbar = display(self.progress(0,M=self.mrange, loss=0), display_id = True)
def update(self, i, loss=0):
self.pbar.update(self.progress(i,M=self.mrange, loss=loss))
def progress(self, value, loss=0, M=100):
return HTML("""Training Mini Batch [{value}/{M}] Loss={loss:0.5f} <progress
value='{value}'
max='{M}',
style='width: 80%'
>
90
</progress>
""".format(value=value,M=M,loss=loss))
# + colab_type="code" id="93SWS597Tut2" colab={}
train_losses = []
val_losses = []
train_losses_epoch = []
val_losses_epoch = []
# + colab_type="code" id="xuFt1veqTwdL" outputId="b972554f-30db-4628-ae6b-84b9bc08302f" colab={"base_uri": "https://localhost:8080/", "height": 35}
nBatch_train = train_loader.__len__()
nBatch_val = val_loader.__len__()
print('{} mini-batches in training dataset, {} mini-batches in validation dataset'.format(nBatch_train, nBatch_val))
# + colab_type="code" id="6TkUb4g3T541" colab={}
epoch_num = 50
# + colab_type="code" id="5AyLjo2OT757" colab={}
def lr_schedular(optimizer, epoch):
if(epoch<2):
lr = 0.001
elif(epoch<10):
lr = 0.0005
elif (epoch<20):
lr = 0.00025
else:
lr = 0.00001
for param_group in optimizer.param_groups:
param_group['lr']=lr
return optimizer
# + colab_type="code" id="X-r3vX8T2Bcu" colab={}
G = G.to(device)
# + colab_type="code" id="uzu40QET_0Lh" colab={}
def get_query_tensor(movieIds):
def get_embedding(kw_list):
if len(kw_list)==0:
return kw_default_embedding
else:
res = torch.zeros((1,1,config['Query_latent_dim']))
for kw in kw_list:
# if kw in kw_set:
res+=kw_embedding[kw]
return res/len(kw_list)
res = []
for i in movieIds.detach().cpu().numpy().flatten():
kws = random.sample(kw_by_movie[i], random.randrange(len(kw_by_movie[i])))
res.append(get_embedding(kws))
return torch.cat(res,dim=0).to(device)
# + colab_type="code" id="kpR-ihn9T89b" outputId="1b828008-adfe-4e50-c12b-24d293cbb478" colab={"base_uri": "https://localhost:8080/", "height": 857}
for epoch in range(epoch_num):
print('Training epoch {}/{}'.format(epoch, epoch_num))
pbar = progress(train_loader.__len__())
train_loss = 0
val_loss = 0
tmp_loss = 0
CF_optimizer = lr_schedular(CF_optimizer, epoch)
for i,(users, movies, ratings, weekend, hr) in enumerate(train_loader):
if i%5==0:
pbar.update(i+1, tmp_loss)
U, I, R = users.to(device), movies.to(device), ratings.to(device)
W, H = weekend.to(device), hr.to(device)
Q = get_query_tensor(movies)
rhat_final, rhat_user = CF(U,I, G[movies], Q, W, H)
loss = CF_criterion(rhat_final, R)+CF_criterion(rhat_user, R)
CF_optimizer.zero_grad()
loss.backward()
CF_optimizer.step()
tmp_loss = loss.item()
train_loss += loss.item()
train_losses.append(loss.item())
train_loss/=nBatch_train
train_losses_epoch.append(train_loss)
for i, (u,m,r, w, h) in enumerate(val_loader):
U, I, R = u.to(device), m.to(device), r.to(device)
W, H = w.to(device), h.to(device)
Q = get_query_tensor(m)
rhat_final, rhat_user = CF(U, I, G[m], Q, W, H)
loss = CF_criterion(rhat_final, R) + CF_criterion(rhat_user, R)
val_loss += loss.item()
val_losses.append(loss.item())
val_loss/=nBatch_val
val_losses_epoch.append(val_loss)
print('epoch[{}/{}], train loss = {}, val loss = {}'.
format(epoch, epoch_num, train_loss, val_loss))
if (epoch%5==0):
plt.figure(figsize=(15,5))
plt.subplot(121)
plt.plot(train_losses, label='train loss')
plt.xlabel('# batch'), plt.ylabel('loss'), plt.legend(), plt.yscale('log')
plt.subplot(122)
plt.plot(val_losses, label='val loss')
plt.xlabel('# batch'), plt.ylabel('loss'), plt.legend(), plt.yscale('log')
plt.show()
# + id="Q3thhgFGWPhO" colab_type="code" colab={}
torch.save(CF.state_dict(), './cfmodel')
# + id="UH0PTGZiWod-" colab_type="code" colab={}
Movie_Em = CF.movie_embedding.weight.data.detach().cpu().numpy()
User_Em = CF.user_embedding.weight.data.detach().cpu().numpy()
# + id="-ckWrM_MWpje" colab_type="code" colab={}
ave_User_Em = User_Em.mean(axis=0, keepdims=True)
# + id="JV-IlnlYW-0H" colab_type="code" colab={}
with open('./movie_embedding', 'wb') as f:
pickle.dump(Movie_Em, f)
with open('./ave_user_embedding', 'wb') as f:
pickle.dump(ave_User_Em, f)
# + colab_type="code" id="1-Y2YIJvjzc5" outputId="ae936d0d-e24e-422a-fb42-205a3b982fbb" colab={"base_uri": "https://localhost:8080/", "height": 392}
plt.figure(figsize=(18,6))
plt.plot(np.linspace(0,epoch_num-1, len(train_losses)),train_losses,label='train_loss')
plt.plot(np.linspace(0,epoch_num-1, len(val_losses)),val_losses,label='val_loss')
plt.plot(train_losses_epoch,label='train_loss by epoch')
plt.plot(val_losses_epoch,label='val_loss by epoch')
plt.yscale('log')
plt.legend(),plt.xlabel('epoch'), plt.ylabel('loss')
plt.grid(True)
#plt.ylim([0.01,0.05])
# + colab_type="code" id="TGm0dNOXyXPL" outputId="07b5d6ba-6859-4445-d3ce-add26f89f8fa" colab={"base_uri": "https://localhost:8080/", "height": 108}
r_real = []
r_pred = []
for i, (u,m,r, w, h) in enumerate(val_loader):
U, I, R = u.to(device), m.to(device), r.to(device)
W, H = w.to(device), h.to(device)
Q = get_query_tensor(m)
rhat_final, rhat_user = CF(U,I, G[m], Q, W, H)
rhat = .9*rhat_final+ .1*rhat_user
r_real+=list(r.detach().cpu().numpy().reshape(-1))
r_pred+=list(rhat.detach().cpu().numpy().reshape(-1))
if(i%20==0):
print('{}/{}'.format(i,nBatch_val))
# + colab_type="code" id="sqQ1nPT8yx8u" colab={}
r_real = np.array(r_real) #, dtype=np.int)
r_pred = np.array(r_pred) #, dtype=np.int)
e = r_pred - r_real
# + colab_type="code" id="STT3nEMey33N" colab={}
r_pearson = scipy.stats.pearsonr(r_real, r_pred)[0]
r_spearman = scipy.stats.spearmanr(r_real, r_pred).correlation
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(r_real, r_pred)
# + colab_type="code" id="c8NsrcAVzYhN" outputId="96b0be40-ef92-4840-c40a-3686477ad8fc" colab={"base_uri": "https://localhost:8080/", "height": 108}
print('average error = {}, std = {}'.format(e.mean(), e.std()))
print('mae = {}'.format(np.abs(e).mean()))
print('rmse = {}'.format(np.sqrt(np.square(e).mean())))
print('slope={}, r2={}'.format(slope, r_value))
print('pearson = {}, spearman = {}'.format(r_pearson, r_spearman))
# + colab_type="code" id="z1hsVwdCzaHM" outputId="d8810e19-b33b-452e-d7e8-955a07d641b7" colab={"base_uri": "https://localhost:8080/", "height": 301}
plt.figure(figsize=(8,4))
sns.distplot(r_pred-r_real, kde=True)
plt.xlabel('predict error'), plt.ylabel('count')
# + colab_type="code" id="flvYXlseze_x" colab={}
real = r.view(-1).detach().cpu().numpy()
pred = rhat.view(-1).detach().cpu().numpy()
users = U.detach().cpu().numpy().reshape(-1)
movies = I.detach().cpu().numpy().reshape(-1)
pd_df = pd.DataFrame({'real': real,'pred':pred, 'userId': users, 'movieId': movies})
pd_df = pd_df.merge(movie_df, how='left', on='movieId')
pd_df = pd_df.merge(user_df, how='left', on='userId')
#pd_df = pd_df.drop(columns = ML_Data.movie_genres)
# + colab_type="code" id="i-g50vc0zgtb" outputId="4bd499ee-9a1b-4fc9-8e8e-88369da2a2b9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
pd_df.head(50)
# + colab_type="code" id="tdS05u8ezq2e" outputId="91023102-a7d3-4fbb-d724-95868f65987f" colab={"base_uri": "https://localhost:8080/", "height": 899}
plt.figure(figsize=(15,15))
sns.categorical.boxplot(x="real", y="pred", data=pd_df)
# + colab_type="code" id="liXs5dazzun4" outputId="d5aa0287-e525-4594-f84c-bdc1fa6f0f2a" colab={"base_uri": "https://localhost:8080/", "height": 627}
plt.figure(figsize=(15,10))
sns.categorical.violinplot(x="real", y="pred", split=True, data=pd_df, palette="pastel")
# + colab_type="code" id="8Rykb8rwzzck" outputId="7f87f752-241d-4f13-bad6-d4daedf86769" colab={"base_uri": "https://localhost:8080/", "height": 591}
rs = list(pd_df['real'].unique())
rs.sort()
plt.figure(figsize=(16,8))
for r in rs:
sns.distplot(pd_df['pred'][pd_df['real']==r], label=str(int(r*2)/2))
plt.legend()
# + colab_type="code" id="-OuxJ-9VMRsC" colab={}
genre_weight = CF.genres_layers[0].weight.detach().cpu().numpy().T
pca = decomposition.PCA(n_components = 5)
G_std = StandardScaler().fit_transform(genre_weight)
G_pca = pca.fit_transform(G_std)
# + colab_type="code" id="4OMMJRavz8j4" outputId="c27166a1-04dd-4153-d777-6e16dfafd39c" colab={"base_uri": "https://localhost:8080/", "height": 617}
plt.figure(figsize=(10,10))
xs = []
ys = []
for key in genres_dict:
i = genres_dict[key]
x,y = G_pca[i,0], G_pca[i,1]
xs+=[x]
ys+=[y]
plt.text(x,y,key,color='b', fontsize=10)
plt.xlim([min(xs),max(xs)])
plt.ylim([min(ys),max(ys)])
# + colab_type="code" id="Si6XLdwMz1-y" colab={}
Movie_Em = CF.movie_embedding.weight.data.detach().cpu().numpy()
# + colab_type="code" id="aKj6B6xPz4_s" colab={}
L = np.linalg.norm(Movie_Em,axis=1,keepdims=True)
# + colab_type="code" id="UzaeGSPTz71A" outputId="ebe02032-bec9-4351-c8c0-e81e6e42fde0" colab={"base_uri": "https://localhost:8080/", "height": 287}
sns.distplot(L,kde=False)
# + colab_type="code" id="ffvAKzcXz8zx" colab={}
pca = decomposition.PCA(n_components = 5)
M_std = StandardScaler().fit_transform(Movie_Em)
M_pca = pca.fit_transform(M_std)
# + colab_type="code" id="OBNVAuzsz_E7" outputId="e8778469-1bb6-4ff6-829f-fd54482d3e73" colab={"base_uri": "https://localhost:8080/", "height": 613}
plt.figure(figsize=(16,10))
ss = (movie_df['num_rating'].values - movie_df['num_rating'].min())/60
plt.scatter(M_pca[:,0], M_pca[:,1], c = movie_df['movie_ave_rating'].values, s = ss, cmap='jet')
plt.colorbar()
plt.axis('equal')
# + colab_type="code" id="p7ii2Sxt0BRj" outputId="b5d7df71-df68-47bf-9546-fa9df3173700" colab={"base_uri": "https://localhost:8080/", "height": 287}
plt.plot(M_pca[:,0], movie_df['movie_ave_rating'],'.')
# + colab_type="code" id="eaV8jBz70Nmf" outputId="7f73850c-13c3-4e68-f763-b92932a81527" colab={"base_uri": "https://localhost:8080/", "height": 505}
plt.figure(figsize=(8,8))
#for g in ML_Dataset.movie_genres:
for g in ['horror','romance','documentary']:
i_g = genres_dict[g]
ind = np.where(genres[:,i_g]==1)[0].tolist()
plt.scatter(M_pca[ind,2], M_pca[ind,1],label=g,s=20, alpha=0.9)
plt.legend()
# + colab_type="code" id="SI-UsY4k0P6E" outputId="32944192-5382-461c-93f5-cbaa5aec2a7e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
TopMovies_id = movie_df['num_rating'].nlargest(200).index.values
plt.figure(figsize=(45,45))
#plt.scatter(M_pca[:,0], M_pca[:,1])
xs = []
ys = []
for i in TopMovies_id:
t = movie_df['title'][i]
x,y = M_pca[i,2], M_pca[i,1]
xs+=[x]
ys+=[y]
#if(x>xlim[0] and x<xlim[1] and y>ylim[0] and y<ylim[1]):
plt.text(x,y,t,color='b', fontsize=10)
plt.xlim([min(xs),max(xs)])
plt.ylim([min(ys),max(ys)])
# + colab_type="code" id="k0P4XsoG0V4H" colab={}
#Movie_Em = Movie_Em/L
# + colab_type="code" id="SDl1bJHf0ddb" colab={}
Movie_S = np.matmul(Movie_Em, Movie_Em.T)
# + colab_type="code" id="IDoqFpL-0fNY" outputId="ecf643c7-af5f-4488-ddc6-59afebc02fe2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for i in range(50):
movie_id = TopMovies_id[i]
t = movie_df['title'][movie_id]
print('Movie:'+t)
tmp_S = Movie_S[movie_id]
tmp_ids = tmp_S.argsort()[::-1]
n = 0
for tmp_id in tmp_ids[1:]:
if(movie_df['movie_ave_rating'][tmp_id]>0.0):
t = movie_df['title'][tmp_id]
print(' ',t)
n+=1
if (n>10):
break
print()
# + colab_type="code" id="uspAQqL74LgR" colab={}
| 24,957 |
/T1P2-Traffic-Sign-Classifier/Traffic_Sign_Classifier.ipynb | 27b77fa8af9ed0c2cadd3935aa7d965d05c29727 | [
"MIT"
] | permissive | rajeshb/SelfDrivingCar | https://github.com/rajeshb/SelfDrivingCar | 1 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 1,365,345 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Summer Academy 2017 Analysis
# ## By: Andre Williams
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Goal: Provide practical insights and analysis to help Make School
# - I'm going to answer 7 questions and do statistical analysis
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Import Packages
# + slideshow={"slide_type": "fragment"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
df = pd.read_csv('SA-Feedback-Surveys_FINAL/2017/Student Feedback Surveys-Superview.csv')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Basic Analysis
# + slideshow={"slide_type": "fragment"}
df.head()
# -
df = df.dropna()
# + slideshow={"slide_type": "fragment"}
df.shape
# + slideshow={"slide_type": "fragment"}
df.info()
# + slideshow={"slide_type": "fragment"}
df.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ## What Was the number of students in each track ?
# + slideshow={"slide_type": "subslide"}
table = pd.crosstab(df['Track'], df['Week'])
table
# + [markdown] slideshow={"slide_type": "slide"}
# ## Where did the majority of the students come from?
# + slideshow={"slide_type": "subslide"}
df.groupby(['Location']).count().plot(kind='bar')
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is the total number of students in the program?
# + slideshow={"slide_type": "fragment"}
apps = dict(pd.crosstab(df['Week'], df['Track']))['Apps'].values
max(apps)
# + slideshow={"slide_type": "fragment"}
explorer_apps = dict(pd.crosstab(df['Week'], df['Track']))['Apps, Explorer'].values
max(explorer_apps)
# + slideshow={"slide_type": "fragment"}
games = dict(pd.crosstab(df['Week'], df['Track']))['Games'].values
max(games)
# + slideshow={"slide_type": "fragment"}
explorer_games = dict(pd.crosstab(df['Week'], df['Track']))['Games, Explorer'].values
max(explorer_games)
# + slideshow={"slide_type": "fragment"}
vr = dict(pd.crosstab(df['Week'], df['Track']))['VR'].values
max(vr)
# + slideshow={"slide_type": "subslide"}
students = {'Apps': 183,
'Explorer Apps': 42,
'Games': 36,
'Explorer Games': 7,
'VR': 10}
print('The total number of students in the 2017 Academy: ' + str(sum(students.values())))
# + [markdown] slideshow={"slide_type": "slide"}
# ## How many more promoters are there than detractors across our 2017 data?
# + [markdown] slideshow={"slide_type": "fragment"}
# ### NPS = (Promoters - Detractors) (Promoters + Passives + Detractors)
# - Promoter (9 – 10)
# - Passive (7 – 8)
# - Detractor (1 – 6)
# + slideshow={"slide_type": "slide"}
df = df.sort_values("Rating (Num)") # sorts values in descending order
# + slideshow={"slide_type": "fragment"}
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3 columns had #Error in our dataset and gets rid of them which changes our response data for Detractors
# + slideshow={"slide_type": "fragment"}
df = df[~df['Rating (Num)'].str.contains("#")]
# + slideshow={"slide_type": "fragment"}
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Find Number of Promoters, Passives and Detractors
# + slideshow={"slide_type": "fragment"}
Promoters = df[pd.to_numeric(df['Rating (Num)']) >= 9]
Passive = df[(pd.to_numeric(df['Rating (Num)']) >= 7) & (pd.to_numeric(df['Rating (Num)']) <= 8)]
Detractors = df[pd.to_numeric(df['Rating (Num)']) < 7]
# + slideshow={"slide_type": "fragment"}
print('The number of promoters: ' + str(len(Promoters)))
print('The number of passives: ' + str(len(Passive)))
print('The number of detractors: ' + str(len(Detractors)))
# + slideshow={"slide_type": "fragment"}
Answer = Promoters - Detractors
print('There are ' + str(len(Answer)) + ' more promoters responses than detractors')
# -
# NPS = (Promoters - Detractors) (Promoters + Passives + Detractors)
NPS_overall = round((len(Promoters) - len(Detractors)) / (len(Promoters) + len(Passive) + len(Detractors))*100, 5)
print('NPS_Overall: ' + str(NPS_overall))
# ## Conclusion: The overall NPS score is good compared to average NPS scores
# ## Week 1 NPS Scores
one_promoters = df[(pd.to_numeric(df['Rating (Num)']) >= 9) & (df['Week']=='Week 1')]
print('Week 1 promoters: ' + str(len(one_promoters)))
one_passive = df[(pd.to_numeric(df['Rating (Num)']) >= 7) & (pd.to_numeric(df['Rating (Num)']) <= 8) & (df['Week']=='Week 1')]
print('Week 1 Passives: ' + str(len(one_passive)))
one_detractors = df[(pd.to_numeric(df['Rating (Num)']) < 7) & (df['Week']=='Week 1')]
#len(df[(df['Rating (Num)'] <= '6') & (df['Week']=='Week 1')])
print('Week 1 detractors: ' + str(len(one_promoters)))
# NPS = (Promoters - Detractors) (Promoters + Passives + Detractors)
NPS = (len(one_promoters - one_detractors)) / (len(one_promoters) + len(one_passive) + len(one_detractors)) * 100
print('NPS score for week one: ' + str(NPS))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Is there a linear relationship in our data?
# + slideshow={"slide_type": "subslide"}
sns.pairplot(df)
# + [markdown] slideshow={"slide_type": "fragment"}
# There is no linear relationship and the data is not normally distributed so we can't use Pearsonr correlation method. We must try a different method to see if a correlation exist in the data because Pearsonr works for linear relationships for continuous variables.
# + [markdown] slideshow={"slide_type": "slide"}
# # Non-parametric Methods for Correlation
# + [markdown] slideshow={"slide_type": "fragment"}
# This is used to find a correlation between categorical non-linearly and non-normally distributed variables.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why Spearman's Rank Correlation?
# + [markdown] slideshow={"slide_type": "fragment"}
# - Use Spearman rank correlation when you have two ranked variables, and you want to see whether the two variables covary; whether, as one variable increases, the other variable tends to increase or decrease
# + [markdown] slideshow={"slide_type": "fragment"}
# ## This is a technique to summarize the strength of data on an ordinal scale. Ordinal variables are variables that are able to be categorized. This conveys the strength of a link between two sets of data.
#
# ### 1+ = Perfect Positive Correlation
# ### 0 = No Correlation
# ### -1 = Perfect Negative Correlation
# + slideshow={"slide_type": "slide"}
import scipy
from scipy.stats import spearmanr
from pylab import rcParams
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
rcParams['figure.figsize']=5, 4
sns.set_style('whitegrid')
# + slideshow={"slide_type": "fragment"}
df.head()
# + [markdown] slideshow={"slide_type": "fragment"}
# - We have NAN values so lets get rid of them
# + slideshow={"slide_type": "fragment"}
df.dropna()
# + slideshow={"slide_type": "slide"}
location = df['Location']
track = df['Track']
week = df['Week']
rating = df['Rating (Num)']
schedule = df['Schedule Pacing']
spearmanr_coefficient, p_value = spearmanr(week, rating)
print('Spearman Rank Correlation Coefficient: ' + str(spearmanr_coefficient))
# + slideshow={"slide_type": "slide"}
spearmanr_coefficient, p_value = spearmanr(week, schedule)
print('Spearman Rank Correlation Coefficient: ' + str(spearmanr_coefficient))
# + slideshow={"slide_type": "slide"}
spearmanr_coefficient, p_value = spearmanr(rating, schedule)
print('Spearman Rank Correlation Coefficient: ' + str(spearmanr_coefficient))
# + slideshow={"slide_type": "skip"}
spearmanr_coefficient, p_value = spearmanr(week, location)
print('Spearman Rank Correlation Coefficient: ' + str(spearmanr_coefficient))
# + slideshow={"slide_type": "skip"}
spearmanr_coefficient, p_value = spearmanr(location, schedule)
print('Spearman Rank Correlation Coefficient: ' + str(spearmanr_coefficient))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Chi-square test for independence
# + [markdown] slideshow={"slide_type": "fragment"}
# - We use the tables to check independence between variables
# + [markdown] slideshow={"slide_type": "fragment"}
# - P < 0.05: Reject null hypothesis and conclude variables are correlated
# - P > 0.05: Reject null hypothesis and conclude variables are independent of each other
# + [markdown] slideshow={"slide_type": "fragment"}
# P_value (probability value) = the probability of obtaining the observed results of a test assuming the null is correct. It is evidence against the null hypothesis.
# + slideshow={"slide_type": "slide"}
table = pd.crosstab(rating, schedule)
from scipy.stats import chi2_contingency
chi2, p, dof, expected = chi2_contingency(table.values)
print('Ch-square Statistic %0.3f p_value %0.3f' % (chi2, p))
# + [markdown] slideshow={"slide_type": "fragment"}
# Correlated
# + slideshow={"slide_type": "slide"}
table = pd.crosstab(week, rating)
from scipy.stats import chi2_contingency
chi2, p, dof, expected = chi2_contingency(table.values)
print('Ch-square Statistic %0.3f p_value %0.3f' % (chi2, p))
# + [markdown] slideshow={"slide_type": "fragment"}
# Independent
# + slideshow={"slide_type": "slide"}
table = pd.crosstab(df['Week'], df['Schedule Pacing'])
from scipy.stats import chi2_contingency
chi2, p, dof, expected = chi2_contingency(table.values)
print('Ch-square Statistic %0.3f p_value %0.3f' % (chi2, p))
# + [markdown] slideshow={"slide_type": "fragment"}
# Correlated
# + [markdown] slideshow={"slide_type": "slide"}
# ## Conclusion
# + [markdown] slideshow={"slide_type": "fragment"}
# For these test we need a p_value > .05 in order to conclude that the variables are independent of each other. This means we would fail to reject the null and the variables are independent. If the p_value < .05 then we reject the null hypothesis and conclude that the two variables are correlated and not independent of each other.
# + slideshow={"slide_type": "fragment"}
sns.pairplot(df)
# -
x = df[['Location','Track','Week','Rating (Num)', 'Schedule Pacing']]
sns.pairplot(x)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Students from which location are more likely to give higher ratings?
# + slideshow={"slide_type": "fragment"}
ratloc = pd.crosstab(df['Location'], df['Rating (Num)'])
ratloc
# + [markdown] slideshow={"slide_type": "slide"}
# # Conclusion
# + [markdown] slideshow={"slide_type": "fragment"}
# ## 1. San Francisco is where the majority of the students came from and Hong Kong is where the smallest number of students came from.
# ## 2. The most popular track was Apps and the least popular track was Explorer Games. 65% of students were in Apps and 3% were in Explorer Games.
# ## 3. The total number of students in the program is 278, under the assumption that the majority of the students didn't take the survey more than once on the same day.
# ## 4. After cleaning the data the majority of the negative responses came from New York City. So they should think about limiting the amount of New Yorkers into their program in the future.
# ## 5. There were 113 more detractors responses than promoters.
# ## 6. There is a weak positive correlation between the increase in the week and the feeling of pace in the program.
# ## 7. There is a weak negative correlation between the week and rating
# ## 8. There is a weak negative correlation between rating and schedule
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Advice: Give each student a unique ID to determine more about each individual student. Obtain more data on preferences and student performance to help improve the number of returning students and a stronger curriculum.
# -
nal samples to mitigate imbalanced distribution of the training data.
# - Added 3x training samples, with slight rotation (-10 to 10 degrees) of the images, to traffic signs with less than 500 samples in the dataset.
# - Normalized training and test data.
# - StartifiedShuffleSplit used for splitting the training and validation samples, for each of the EPOC. This step enabled the use of entire training data, while keeping specified amount of data (5%) for cross validation.
# ### ARCHITECTURE
# 
# +
### Define your architecture here.
### Feel free to use as many code cells as needed.
import tensorflow as tf
from tensorflow.contrib.layers import flatten
# reset graph
tf.reset_default_graph()
keep_prob = tf.placeholder(tf.float32, name='KeepProb')
def LeNet(x):
# Layer 1: Convolutional. Input = 32x32x3. Output = 32x32x16.
conv1_W = tf.get_variable("conv1_W", shape=[3, 3, 3, 16],
initializer=tf.contrib.layers.xavier_initializer())
conv1_b = tf.get_variable("conv1_b", shape=[16],
initializer=tf.contrib.layers.xavier_initializer())
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='SAME') + conv1_b
# Activation.
conv1 = tf.nn.relu(conv1)
# Dropout.
conv1 = tf.nn.dropout(conv1, keep_prob)
# Pooling. Input = 32x32x16. Output = 16x16x16.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME')
# Layer 2: Convolutional. Output = 16x16x64.
conv2_W = tf.get_variable("conv2_W", shape=[3, 3, 16, 64],
initializer=tf.contrib.layers.xavier_initializer())
conv2_b = tf.get_variable("conv2_b", shape=[64],
initializer=tf.contrib.layers.xavier_initializer())
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='SAME') + conv2_b
# Activation.
conv2 = tf.nn.relu(conv2)
# Dropout.
conv2 = tf.nn.dropout(conv2, keep_prob)
# Pooling. Input = 16x16x64. Output = 8x8x64.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME')
# Flatten. Input = 8x8x64. Output = 4096.
fc0 = flatten(conv2)
# Layer 3: Fully Connected. Input = 4096. Output = 512.
fc1_W = tf.get_variable("fc1_W", shape=[4096, 512],
initializer=tf.contrib.layers.xavier_initializer())
fc1_b = tf.get_variable("fc1_b", shape=[512],
initializer=tf.contrib.layers.xavier_initializer())
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# Activation.
fc1 = tf.nn.relu(fc1)
# Layer 4: Fully Connected. Input = 512. Output = 144.
fc2_W = tf.get_variable("fc2_W", shape=[512, 144],
initializer=tf.contrib.layers.xavier_initializer())
fc2_b = tf.get_variable("fc2_b", shape=[144],
initializer=tf.contrib.layers.xavier_initializer())
fc1 = tf.nn.dropout(fc1, keep_prob)
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# Activation.
fc2 = tf.nn.relu(fc2)
# Layer 5: Fully Connected. Input = 144. Output = 43.
fc3_W = tf.get_variable("fc3_W", shape=[144, 43],
initializer=tf.contrib.layers.xavier_initializer())
fc3_b = tf.get_variable("fc3_b", shape=[43],
initializer=tf.contrib.layers.xavier_initializer())
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
# -
# ### Question 3
#
# _What does your final architecture look like? (Type of model, layers, sizes, connectivity, etc.) For reference on how to build a deep neural network using TensorFlow, see [Deep Neural Network in TensorFlow
# ](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/b516a270-8600-4f93-a0a3-20dfeabe5da6/concepts/83a3a2a2-a9bd-4b7b-95b0-eb924ab14432) from the classroom._
#
# **Answer:**
#
# - Re-purposed LeNet architecture from MINST data classification.
# - Architecture: 5 layers architecture, 2 convolutional and 3 fully connected layers.
# - After the activation of convolutional layers, dropouts and max pooling techniques are used.
# - Added TensorBoard visualization as part of the project submission.
# ### TRAINING MODEL
# +
### Train your model here.
### Feel free to use as many code cells as needed.
EPOCHS = 25
BATCH_SIZE = 128
LEARNING_RATE = 0.001
LOG_PATH = './log'
# Split the train and validation, different dataset for each of the EPOCH
from sklearn.model_selection import StratifiedShuffleSplit
sss = StratifiedShuffleSplit(n_splits=EPOCHS, test_size=0.05, random_state=450)
X_train_org = X_train
y_train_org = y_train
x = tf.placeholder(tf.float32, (None, 32, 32, 3), name="InputData")
y = tf.placeholder(tf.int32, (None), name="LabelData")
one_hot_y = tf.one_hot(y, 43)
logits = LeNet(x)
with tf.name_scope('Model'):
prediction = tf.nn.softmax(logits)
with tf.name_scope('Loss'):
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits, one_hot_y)
loss_operation = tf.reduce_mean(cross_entropy)
with tf.name_scope('Train'):
optimizer = tf.train.AdamOptimizer(learning_rate = LEARNING_RATE)
training_operation = optimizer.minimize(loss_operation)
with tf.name_scope('Accuracy'):
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
init = tf.global_variables_initializer()
# create a summary for our cost and accuracy
tf.summary.scalar("loss", loss_operation)
tf.summary.scalar("accuracy", accuracy_operation)
# merge all summaries into a single operation
summary_operation = tf.summary.merge_all()
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
total_loss = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy, loss = sess.run([accuracy_operation, loss_operation] ,
feed_dict={x: batch_x, y: batch_y, keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
total_loss += (loss * len(batch_x))
return total_accuracy / num_examples, total_loss / num_examples
with tf.Session() as sess:
sess.run(init)
# Initialize logger for Tensorboard
writer = tf.summary.FileWriter(LOG_PATH, graph=tf.get_default_graph())
print("Training...")
print()
epoch = 0
for train_index, test_index in sss.split(X_train_org, y_train_org):
# Generate Training and Validation datasets for each of the EPOCH
X_train, X_validation = X_train_org[train_index], X_train_org[test_index]
y_train, y_validation = y_train_org[train_index], y_train_org[test_index]
num_examples = len(X_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
_, summary = sess.run([training_operation, summary_operation], feed_dict={x: batch_x, y: batch_y, keep_prob: 0.5})
writer.add_summary(summary, epoch * BATCH_SIZE + offset)
validation_accuracy, validation_loss = evaluate(X_validation, y_validation)
print("EPOCH {} Validation Accuracy = {:.3f} Loss = {:.3f}".format(epoch+1, validation_accuracy, validation_loss))
epoch += 1
saver.save(sess, 'lenet')
print("Model saved")
# -
# ### Question 4
#
# _How did you train your model? (Type of optimizer, batch size, epochs, hyperparameters, etc.)_
#
# ### **Answer:**
#
# - Optimizer: AdamOptimizer (over GradientDescent), as it uses the momentum (moving average of the parameters) and requires less turning of hyper-parameters.
# - Batch size: 128
# - EPOCH size: 25, higher the number doesn't add much to the validation accuracy
# - Learning rate: 0.001, tried a few higher/lower values resulted with lower validation accuracy
# ### Question 5
#
#
# _What approach did you take in coming up with a solution to this problem? It may have been a process of trial and error, in which case, outline the steps you took to get to the final solution and why you chose those steps. Perhaps your solution involved an already well known implementation or architecture. In this case, discuss why you think this is suitable for the current problem._
# **Answer:**
#
# - The goal was to get higher validation accuracy before running against test samples.
# - Started with 95% validation accuracy using LeNet architecture.
# - After improving the pre-process steps and with data augmentation, the accuracy gotten better to 97%
# - Fine tuning the architecture output, epoch size, batchsize, learning rate parameters using trial-and-error to get the validation accuracy above 99%
# #### TEST RESULTS
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy, test_loss = evaluate(X_test, y_test)
print("Test accuracy of the model = {:.3f}".format(test_accuracy))
# ---
#
# ## Step 3: Test a Model on New Images
#
# Take several pictures of traffic signs that you find on the web or around you (at least five), and run them through your classifier on your computer to produce example results. The classifier might not recognize some local signs but it could prove interesting nonetheless.
#
# You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Implementation
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# +
from PIL import Image
print("SETTING UP NEW IMAGES")
print('=====================')
IMAGE_FOLDER = './new_images'
random_images = [
'35_ahead.jpg',
'14_stop_sign.png',
'5_speed_limit_80.png',
'22_bumpy_road.png',
'36_straight_or_right.png']
y_test_new = np.array([35,14,5,22,36])
cols = len(random_images)
rows = 1
fig = plt.figure(figsize=(cols, rows))
X_test_new = []
for idx, image_name in enumerate(random_images):
im = Image.open(IMAGE_FOLDER + '/' + image_name)
X_test_new.append(np.array(im.getdata(),
np.uint8).reshape(im.size[1], im.size[0], 3))
plt.subplot(rows, cols, idx+1)
plt.imshow(im)
plt.xlabel('{}'.format(y_test_new[idx]))
plt.xticks([])
plt.yticks([])
plt.show()
X_test_new = np.asarray(X_test_new, dtype=np.float32)
print("Normalizing new images ...")
X_test_new = normalize(X_test_new)
print('Done')
# -
# ### Question 6
#
# _Choose five candidate images of traffic signs and provide them in the report. Are there any particular qualities of the image(s) that might make classification difficult? It could be helpful to plot the images in the notebook._
#
#
# **Answer:**
#
# - Picked 5 random traffic sign images from the web and plotted above, both in PNG and JPEG formats.
# - Most of the new images are much cleaner looking images, compared to the sample images, which may pose some challenge in prediction.
# - A couple of images (stop sign and speed limit 80) are pictures with background, similar to sample images, should be easier to predict, I guess.
# - It would be interesting to see if the model will be able to predict all of these new traffic signs.
# ### Question 7
#
# _Is your model able to perform equally well on captured pictures when compared to testing on the dataset? The simplest way to do this check the accuracy of the predictions. For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate._
#
# _**NOTE:** You could check the accuracy manually by using `signnames.csv` (same directory). This file has a mapping from the class id (0-42) to the corresponding sign name. So, you could take the class id the model outputs, lookup the name in `signnames.csv` and see if it matches the sign from the image._
#
# ### **Answer:**
#
# - Test accuracy is **94.5%** for the model.
# - Prediction accuracy is **100%**, in some runs the prediction accuracy was **80%** (4 out 5 images).
# - In those runs with 80% prediction accuracy, '22 - bumpy road' traffic sign was incorrectly predicted as '29 - Bicycles crossing' traffic sign. However, '22 - bumpy road' sign came up second or third in the top-k certainities.
# +
### Run the predictions for the new images.
### Feel free to use as many code cells as needed.
print("PREDICTIONS FOR NEW IMAGES")
print("==========================")
print("Test accuracy for the model = {:.3f}".format(test_accuracy))
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
new_images_accuracy = sess.run(accuracy_operation, feed_dict={x: X_test_new, y: y_test_new, keep_prob: 1.0})
new_images_prediction = sess.run(tf.argmax(tf.nn.softmax(logits),1), feed_dict={x: X_test_new, keep_prob: 1.0})
new_images_certainity = sess.run(tf.nn.top_k(logits, 3), feed_dict={x: X_test_new, keep_prob: 1.0})
print("Prediction accuracy for new images = {:.3f}".format(new_images_accuracy))
print("Expected result (E) =", y_test_new)
print("Predicted result (P) =", new_images_prediction)
print()
for i, (predictions, probabilities, expected_class) in enumerate(zip(new_images_certainity.indices, new_images_certainity.values, y_test_new)):
print("Expected class ({}) Predictions ({}) Probabilities ({})".format(expected_class, predictions, probabilities))
# -
# ### Question 8
#
# *Use the model's softmax probabilities to visualize the **certainty** of its predictions, [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here. Which predictions is the model certain of? Uncertain? If the model was incorrect in its initial prediction, does the correct prediction appear in the top k? (k should be 5 at most)*
#
# `tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example:
#
# ```
# # (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
# ```
#
# Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
#
# ```
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
# ```
#
# Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
# **Answer:**
#
# - While tuning the parameters to improve performance, I have noticed at least one of the predictions go wrong ('22 - bumpy road' was incorrectly predicted as '29 - Bicycles crossing' or '17 - No entry' traffic sign classes).
# - Top K certainity will be helpful, to check if an image falls in top 2 or 3 certainities, especially if the certainities are too close to call.
# - Based on the results, it looks that the model works resonably good. However, the lower certainities on predicting new images suggest that there should be a continuous way to improve the model with newer images.
# +
# plot softmax probabilities to visualize the certainty
for i, (predictions, probabilities, expected_image, expected_class) in enumerate(zip(new_images_certainity.indices, new_images_certainity.values, X_test_new, y_test_new)):
fig = plt.figure(figsize=(15, 2))
plt.bar(predictions, probabilities)
plt.ylabel('Certainity in %')
#plt.yticks(np.arange(0.0, 50.0, 10), np.arange(0.0, 50.0, 10))
plt.xticks(np.arange(0.5, 43.5, 1.0), list(traffic_sign_names.values.flatten()), ha='right', rotation=45)
ax = plt.axes([0.75,0.25,0.5,0.5], frameon=True)
ax.imshow(expected_image)
ax.set_xlabel('E:{} P:{}'.format(expected_class, predictions[0]))
ax.set_xticks([])
ax.set_yticks([])
plt.show()
# -
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
| 29,166 |
/meetup3/occupation-Exercise_with_Solution.ipynb | 30c373a984126302bd002f65c56cd32c468de2f2 | [] | no_license | Sdoof/Workshop | https://github.com/Sdoof/Workshop | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 28,266 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="k3KV4rsm_1zS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="eeae0161-5167-42dd-858d-44511cedcc06"
### Pythonのバージョン確認
import sys
print(sys.version)
# + id="BV5svbrCuM5m" colab_type="code" colab={}
import matplotlib.pyplot as plt
import math
# + [markdown] id="ewQuq0ipzVoS" colab_type="text"
# # フィールドの初期化
# + id="vX4b2JZJr2ZX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="9e37bad1-a778-4d55-d656-205a2baaf6db"
### 障害物座標 (x, y) を保持する配列
obs_xs = []
obs_ys = []
max_x = 50
max_y = 50
### 下壁
for i in range(0, max_x):
obs_xs.append(i)
obs_ys.append(0)
### 上壁
for i in range(0, max_x):
obs_xs.append(i)
obs_ys.append(max_y-1)
### 左壁
for i in range(0, max_y):
obs_xs.append(0)
obs_ys.append(i)
### 右壁
for i in range(0, max_y):
obs_xs.append(max_x-1)
obs_ys.append(i)
### 障害物(縦線1)
for i in range(30, 41):
obs_xs.append(10)
obs_ys.append(i)
### 障害物(横線1)
for i in range(10, 31):
obs_xs.append(i)
obs_ys.append(30)
### 障害物(縦線2)
for i in range(20, 31):
obs_xs.append(30)
obs_ys.append(i)
### 障害物(横線2)
for i in range(30, 36):
obs_xs.append(i)
obs_ys.append(20)
print('obs_xs =', obs_xs)
print('obs_ys =', obs_ys)
# + id="-SLiNtPFyajd" colab_type="code" colab={}
### スタート位置とゴール位置の座標 (x, y) を保持する変数
start_x, start_y = 10, 10
goal_x, goal_y = 40, 40
# + id="oqUame_cwFuv" colab_type="code" colab={}
### フィールドを描画する関数
def DrawField(obs_xs, obs_ys, start_x, start_y, goal_x, goal_y):
plt.scatter(obs_xs, obs_ys, marker='.', color='black')
plt.scatter(start_x, start_y, marker='x', color='blue')
plt.scatter(goal_x, goal_y, marker='x', color='red')
plt.xlabel('X')
plt.ylabel('Y')
plt.axis('equal')
plt.grid(True)
return
# + id="Pca58XTM0v93" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="4824d403-0ceb-4f3c-f3c3-f5d249fadd35"
DrawField(obs_xs, obs_ys, start_x, start_y, goal_x, goal_y)
# + [markdown] id="ELncEK6UUR5e" colab_type="text"
# # 障害物マップとロボットのモーションモデルの作成
# + id="xA9y0pWqNv-F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 986} outputId="981525b1-df26-4ccb-d1f9-68f9ea82fafa"
### 0なら障害物なし / 1なら障害物あり
NOT_OBS = 0
IS_OBS = 1
### 2次元配列の全マスをとりあえず「障害物なし」で初期化
obs_map = []
for v in range(max_y):
obs_map_oneline = [NOT_OBS for u in range(max_x)] ### [NOT_OBS, NOT_OBS, NOT_OBS, ... , NOT_OBS]
obs_map.append(obs_map_oneline)
print('len(obs_map) =', len(obs_map))
print('len(obs_map[0]) =', len(obs_map[0]))
print()
### 障害物の総数
assert len(obs_xs) == len(obs_ys)
num_obs = len(obs_xs)
print('num_obs =', num_obs)
print()
### 障害物がある座標は「障害物あり」に上書き
for i in range(num_obs):
u = obs_xs[i]
v = (max_y - 1) - obs_ys[i] ### 注:Y軸(上向き正 : 左下原点)とV軸(下向き正 : 左上原点)は上下反転する
obs_map[v][u] = IS_OBS
### 画面に表示して動作確認
print('obs_map = [')
for v in range(max_y):
obs_map_oneline = obs_map[v]
print(' ', obs_map_oneline)
print(']')
# + id="XdJ40CKe0GiC" colab_type="code" colab={}
'''
1.41 1 1.41
↖ ↑ ↗
1 ← * → 1
↙ ↓ ↘
1.41 1 1.41
'''
### motion_dx, motion_dy, motion_cost
motion_model = [
[ 1, 0, 1],
[ 0, 1, 1],
[-1, 0, 1],
[ 0, -1, 1],
[-1, -1, math.sqrt(2)],
[-1, 1, math.sqrt(2)],
[ 1, -1, math.sqrt(2)],
[ 1, 1, math.sqrt(2)]
]
# + [markdown] id="0WLMvtywBp0L" colab_type="text"
# # 経路探索アルゴリズムに使うデータ構造や関数の準備
# + id="jA_WiSpc198k" colab_type="code" colab={}
'''
(x, y) : 左下原点
[
[(0,3), (1,3), (2,3), (3,3)],
[(0,2), (1,2), (2,2), (3,2)],
[(0,1), (1,1), (2,1), (3,1)],
[(0,0), (1,0), (2,0), (3,0)]
]
↑
↓
(v, u) : 左上原点
[
[(0,0), (0,1), (0,2), (0,3)],
[(1,0), (1,1), (1,2), (1,3)],
[(2,0), (2,1), (2,2), (2,3)],
[(3,0), (3,1), (3,2), (3,3)]
]
↑
↓
id : 左上原点
[
[ 0, 1, 2, 3],
[ 4, 5, 6, 7],
[ 8, 9, 10, 11],
[12, 13, 14, 15],
]
'''
### (x, y) から ---> id に変換する関数
def XY2ID(x, y, max_x, max_y):
u = x
v = (max_y - 1) - y ### 注:Y軸(上向き正 : 左下原点)とV軸(下向き正 : 左上原点)は上下反転する
id = v*max_x + u
return id
# + id="BoJXMoEU5pK1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="36063403-99bb-4b30-d306-cb7040dfc127"
### 画面に表示して動作確認
print('id =', XY2ID(x=0, y=3, max_x=4, max_y=4))
print('id =', XY2ID(x=0, y=2, max_x=4, max_y=4))
print('id =', XY2ID(x=0, y=1, max_x=4, max_y=4))
print('id =', XY2ID(x=0, y=0, max_x=4, max_y=4))
# + id="cTpuGGlKNtI7" colab_type="code" colab={}
class Node:
def __init__(self, x, y, cost, parent_id):
self.x = x
self.y = y
self.cost = cost
self.parent_id = parent_id
# + id="h15cXt479dK3" colab_type="code" colab={}
### ここがダイクストラ法とA*の違い
### ヒューリスティックコスト(今回は単に注目ノードからゴールノードまでの距離)を計算して返す関数
def CalcHeuristicCost(goal_node, open_node):
dist_x = goal_node.x - open_node.x
dist_y = goal_node.y - open_node.y
heuristic_cost = math.sqrt(dist_x**2 + dist_y**2)
return heuristic_cost
'''
open_nodes_dict = {
0: Node(x0, y0, cost0, parent_id0),
1: Node(x1, y1, cost1, parent_id1),
2: Node(x2, y2, cost2, parent_id2),
3: Node(x3, y3, cost3, parent_id3),
:
:
}
'''
### ここがダイクストラ法とA*の違い
### ノードリスト(辞書型)の中からヒューリスティックコスト込みでトータルコストが最小のノードを探してそのノードIDを返す関数
def FindMinTotalCostNodeID(open_nodes_dict, goal_node):
min_total_cost = sys.float_info.max
min_total_cost_node_id = None
for id, open_node in open_nodes_dict.items():
heuristic_cost = CalcHeuristicCost(goal_node, open_node)
total_cost = open_node.cost + heuristic_cost
if total_cost < min_total_cost:
min_total_cost = total_cost
min_total_cost_node_id = id
return min_total_cost_node_id
# + id="O75k8Hk-GliA" colab_type="code" colab={}
def IsObsNode(obs_map, node, max_x, max_y):
u = node.x
v = (max_y - 1) - node.y ### 注:Y軸(上向き正 : 左下原点)とV軸(下向き正 : 左上原点)は上下反転する
### 注目ノードが障害物ノードならアウト
if obs_map[v][u] == IS_OBS:
return True
### 注目ノードがフィールド外ならアウト(念のため確認)
if node.x < 0 or (max_x - 1) < node.x:
return True
if node.y < 0 or (max_y - 1) < node.y:
return True
### どの条件にも引っ掛からなかったらセーフ
return False
# + id="DSlPd912Qj1W" colab_type="code" colab={}
### オープンノードリストとクローズノードリストを描画する関数
### (コストが低いほど青に近い色 〜 コストが高いほど赤に近い色)
def DrawOpenAndCloseNodes(open_nodes_dict, close_nodes_dict, max_x, max_y):
### コストの値を 0~1 の間に正規化するために最大値(大体の値)を用意
max_cost = math.sqrt(max_x**2 + max_y**2)
for open_node in open_nodes_dict.values():
if open_node.parent_id != None:
norm_node_cost = open_node.cost/max_cost
rgb = (norm_node_cost, 0, 1 - norm_node_cost) ### rgb = (0~1, 0, 0~1)
plt.scatter(open_node.x, open_node.y, facecolors='none', edgecolors=rgb, s=20) ### オープンノードは○で描画
for close_node in close_nodes_dict.values():
if close_node.parent_id != None:
norm_node_cost = close_node.cost/max_cost
rgb = (norm_node_cost, 0, 1 - norm_node_cost) ### rgb = (0~1, 0, 0~1)
plt.scatter(close_node.x, close_node.y, marker='o', color=rgb, s=20) ### クローズノードは●で描画
# + [markdown] id="LQf3lpblYgK6" colab_type="text"
# # A*による経路探索
# + id="daPVwHmMWt01" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 905} outputId="1eae1829-1b48-4e6e-e219-377a5b71606c"
### スタートノードとゴールノードを初期化
start_node = Node(start_x, start_y, cost=0.0, parent_id=None)
goal_node = Node(goal_x, goal_y, cost=0.0, parent_id=None)
### オープンノードリストとクローズノードリストを辞書型で初期化
open_nodes_dict = dict()
close_nodes_dict = dict()
### まず最初にオープンノードリストにスタートノードを追加
start_node_id = XY2ID(start_node.x, start_node.y, max_x, max_y)
open_nodes_dict[start_node_id] = start_node
### ゴールノードに到達まで繰り返し探索
step = 0
draw_step = 300
while True:
### オープンノードリストが空なら(残念ながら)ゴールまでの経路が発見できなかったということなので終了する
if (len(open_nodes_dict) == 0):
print('open_nodes_dict is empty...')
break
### ここがダイクストラ法とA*の違い
### オープンノードリストの中でヒューリスティックコスト込みのトータルコストが最小のノードを現在ノードに設定
curr_node_id = FindMinTotalCostNodeID(open_nodes_dict, goal_node)
curr_node = open_nodes_dict[curr_node_id]
### ゴールノードに到達したら,ノード情報(コスト,親ノードID)を保存してi経路探索を終了する
if curr_node.x == goal_node.x and curr_node.y == goal_node.y:
print('Goal !!! \n')
goal_node.cost = curr_node.cost
goal_node.parent_id = curr_node.parent_id
break
### 現在ノードをオープンノードリストから削除してクローズノードリストに追加
del open_nodes_dict[curr_node_id]
close_nodes_dict[curr_node_id] = curr_node
### 現在ノードの8近傍(8隣接)ノードをしらみ潰しに探索
for motion_dx, motion_dy, motion_cost in motion_model:
### 隣接ノードとそのノードIDを設定
neighbor_node = Node(
curr_node.x + motion_dx,
curr_node.y + motion_dy,
cost = curr_node.cost + motion_cost,
parent_id = curr_node_id
)
neighbor_node_id = XY2ID(neighbor_node.x, neighbor_node.y, max_x, max_y)
### もし隣接ノードが既にクローズノードリストの中にあるなら,もう再度探索はしない
if neighbor_node_id in close_nodes_dict.keys():
continue
### もし隣接ノードが障害物なら,探索しない
if IsObsNode(obs_map, neighbor_node, max_x, max_y):
continue
### もし隣接ノードがオープンノードリストの中にまだなければ,そのまま追加
### もし隣接ノードが既にオープンノードリストの中にあったとしてもコストが新たに下回るなら,上書き
if neighbor_node_id not in open_nodes_dict.keys():
open_nodes_dict[neighbor_node_id] = neighbor_node
elif neighbor_node.cost < open_nodes_dict[neighbor_node_id].cost:
open_nodes_dict[neighbor_node_id] = neighbor_node
### End of for
### 繰り返しのステップ数が draw_step 回ごとに探索の途中経過を描画
if step % draw_step == 0:
DrawOpenAndCloseNodes(open_nodes_dict, close_nodes_dict, max_x, max_y)
DrawField(obs_xs, obs_ys, start_x, start_y, goal_x, goal_y)
plt.pause(0.001)
step += 1
### End of while
print('step =', step)
print()
print('len(open_nodes_dict) =', len(open_nodes_dict))
print('len(close_nodes_dict) =', len(close_nodes_dict))
# + id="fAIM7x-rzOS7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="5e6ab0d0-c161-466b-e520-8afafea5525e"
### 探索した最終経路の始まりはゴールノードから
path_x = [goal_node.x]
path_y = [goal_node.y]
next_node_id = goal_node.parent_id
### ゴールノードから親ノードIDを辿ってスタートノードに辿り着くまで(つまり next_node_id が None になるまで)繰り返し
while next_node_id != None:
next_node = close_nodes_dict[next_node_id]
path_x.append(next_node.x)
path_y.append(next_node.y)
next_node_id = next_node.parent_id
### 探索した最終経路とフィールドを描画
plt.scatter(path_x, path_y, marker='.', color='cyan')
DrawField(obs_xs, obs_ys, start_x, start_y, goal_x, goal_y)
# + id="Q7UVVj8zPtLu" colab_type="code" colab={}
| 10,938 |
/SAMPLING TECHNIQUE 30.08.2021.ipynb | e19c7691d7ee9cf27329e0443d222584116d63c1 | [] | no_license | AadeshY/KAGGLE-Dataset-Analysis | https://github.com/AadeshY/KAGGLE-Dataset-Analysis | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 115,214 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sampling technique is part of classification algorithm
# Data imbalance:
# There are 2 types of sampling technique :-
# 1. Over sampling
# 2. Under sampling
# 
#
# If you are basically reducing the majority class that is known as under sampling
# if you are increasing the minority class that is known as over sampling .
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# dataset
df = pd.read_csv('telecom_churn.csv')
df.head()
# To see all the columns or rows we use
#
# pd.set_option('display.max_columns',None)
#
# pd.set_option('display.max_rows',None)
# +
# customer churn can be termed as loss of custormers or customer defection
# predict whether the customer will churn
# +
# preprocessing the data
# drop the unwanted columns from the data set
df.drop('customerID',axis=1,inplace=True)
df.head()
# -
# checking null values
sns.heatmap(df.isnull())
plt.show()
df.info()
# total charges which is int but has dtype as object
df['TotalCharges'].value_counts()
# the first data is space where there is no input
# filling the void
df['TotalCharges'].replace(' ',np.nan,inplace=True)
df.isnull().sum()
# change data type of TotalCharges
df['TotalCharges'] = df['TotalCharges'].astype(float)
# checking the change
df.dtypes
# filling the null with mean
df['TotalCharges'].fillna(df['TotalCharges'].mean(),inplace=True)
# checking the above changes
df.isnull().sum()
df['Churn'].value_counts()
# the Yes is nearly one third of no data which shows data imbalance
sns.countplot(data=df,x='Churn')
c=df['Churn'].value_counts()
plt.yticks(c)
plt.show()
# using label encoder
# object type data
df_cat = df.select_dtypes('object')
# numeric type data
df_num = df.select_dtypes(['float64','int64'])
from sklearn.preprocessing import LabelEncoder
# object creation
for col in df_cat:
le = LabelEncoder()
df_cat[col]=le.fit_transform(df[[col]]) # input always 2D
df_cat.head()
# concatenating the two datasets
df_new = pd.concat([df_num,df_cat],axis=1)
df_new.head()
# +
# install package for data imbalance handling
# #!pip install imblearn
# -
from imblearn.under_sampling import RandomUnderSampler
from imblearn.over_sampling import RandomOverSampler
# seperating input and output variable from the df_new
X = df_new.drop('Churn',axis=1) # input
Y = df_new['Churn'] # target
# train_test_split
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,test_size=0.3,random_state=1)
# function create
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
def create_model(model):
model.fit(X_train,Y_train)
Y_pred=model.predict(X_test)
print(classification_report(Y_test,Y_pred))
print('Confusion Matrix')
print(confusion_matrix(Y_test,Y_pred))
return model
#Base Line Model - Logistic Regression
from sklearn.linear_model import LogisticRegression
#creating object of LogisticRegression class
lr=LogisticRegression()
lr = create_model(lr)
# applying sampling technique
ros = RandomOverSampler()
# increasing the record of minority class
pd.Series(Y_train).value_counts() # check for balance
# applying random over sampling
X_train1,Y_train1 = ros.fit_resample(X_train,Y_train)
#after apply randomoversampler
pd.Series(Y_train1).value_counts() #check if not balance
# apply LR
#create object of LogisticRegression class
lr=LogisticRegression()
#Train the model
lr.fit(X_train1,Y_train1)
#before apply randomoversampler ,check Y_test
pd.Series(Y_test).value_counts() #check if not balance
#apply randomoversampling for balance
X_test1,Y_test1=ros.fit_resample(X_test,Y_test)
#after apply randomoversampler
pd.Series(Y_test1).value_counts() #check if not balance
#Test the model
Y_pred=lr.predict(X_test1)
print(classification_report(Y_test1,Y_pred))
# +
# DecisionTreeClassifier
from sklearn.tree import DecisionTreeClassifier
# -
# creating object for Decision Tree
dt = DecisionTreeClassifier()
# train
dt.fit(X_train1,Y_train1)
#test
Y_pred=dt.predict(X_test1)
print(classification_report(Y_test1,Y_pred))
# +
# applying pruning for overfitting and so on to get answer
dt = DecisionTreeClassifier(max_depth=10)
# -
dt = create_model(dt)
#RandomUnderSampling : -
from imblearn.under_sampling import RandomUnderSampler
#we delete record randomly from majority class
#create object of RandomUnderSampler class
rus=RandomUnderSampler()
#before apply randomoversampler
pd.Series(Y_train).value_counts() #check if not balance
X_train1,Y_train1=rus.fit_resample(X_train,Y_train)
pd.Series(Y_train1).value_counts()
#before apply randomoversampler ,check Y_test
pd.Series(Y_test).value_counts() #check if not balance
#apply randomoversampling for balance
X_test1,Y_test1=rus.fit_resample(X_test,Y_test)
#before apply randomoversampler ,check Y_test
pd.Series(Y_test1).value_counts() #check if not balance
# apply LR
#create object of LogisticRegression class
lr=LogisticRegression()
#Train the model
lr.fit(X_train1,Y_train1)
# test
Y_pred=lr.predict(X_test1)
print(classification_report(Y_test1,Y_pred))
| 5,473 |
/lab 4.ipynb | 1629abb0d803da560c814dd92daefd2adfb2f460 | [] | no_license | vipinethical/pythonworkshop | https://github.com/vipinethical/pythonworkshop | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 5,222 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
var1= (1,2,3,4)
print(var1[0:3],var1[-1])
# ## ques1
#
tuple=(1,2,3)
tuple
# ## ques2
tuple1=(1,"v","VIPIN")
tuple1
# ## ques3
tuple3=(1,2,3)
tuple3=[1]
# +
tuple3
# -
# ## ques4
tuple4=(1,2,3,4)
print(tuple4)
v1,v2, *v3=tuple4
print(v1,v2,v3)
# ## ques5
tuple5=(1,2,3)
# +
print(tuple5)
# -
tuple6 = tuple5 + (9,)
print(tuple6)
# ## ques6
tuple3=("vipin","chaudhary")
str= ''.join(tuple3)
print(str)
# ## ques7
| 717 |
/model_building_4_29_Feature_Reduction.ipynb | e3062aef84fff37cbe36efb847afa51686dcf23e | [] | no_license | LinChiGong/COVID19-Forecasting | https://github.com/LinChiGong/COVID19-Forecasting | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 221,203 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python (py365)
# language: python
# name: py365
# ---
# # Inception model explanations, t0
# ## First we need dataframes for every subject
import sys
sys.path.append('../')
import warnings
warnings.filterwarnings('ignore')
# +
import pandas as pd
import numpy as np
import utils
import keras
# %matplotlib inline
# DEFINE GRAPH
from importlib import reload
reload(utils)
pd.set_option('max_colwidth', 800)
# +
# Data paths
rgb_path = '../data/jpg_320_180_1fps/'
of_path = 'no'
# Hyperparameters
input_width = 320
input_height = 180
seq_length = 10
seq_stride = 1
batch_size = 10
COLOR = True
nb_labels = 2
# Data augmentation
aug_flip = 0
aug_crop = 0
aug_light = 0
nb_input_dims = 4
subject_ids = pd.read_csv('../metadata/horse_subjects.csv')['Subject'].values
channels = 3
# -
from utils import ArgsProxy
args = ArgsProxy(rgb_path, of_path, input_height, input_width,
seq_length, seq_stride, batch_size, nb_labels,
aug_flip, aug_crop, aug_light, nb_input_dims)
import data_handler as dathand
dh = dathand.DataHandler(path=args.data_path,
of_path=args.of_path,
clip_list_file='videos_overview_missingremoved.csv',
data_columns=['Pain'], # Here one can append f. ex. 'Observer'
image_size=(args.input_height, args.input_width),
seq_length=args.seq_length,
seq_stride=args.seq_stride,
batch_size=args.batch_size,
color=COLOR,
nb_labels=args.nb_labels,
aug_flip=args.aug_flip,
aug_crop=args.aug_crop,
aug_light=args.aug_light,
nb_input_dims=args.nb_input_dims)
import utils
subject_dfs = utils.read_or_create_subject_dfs(dh, args, subject_ids)
sequence_df = utils.get_sequence(args=args, subject_dfs=subject_dfs, subject=1)
y = sequence_df['Pain'].values
image_paths = sequence_df['Path'].values
image_paths
y
# +
# from keras.utils import np_utils
# label_onehot = np_utils.to_categorical(y, num_classes=args.nb_labels)
# batch_label = label_onehot.reshape(args.batch_size, -1, args.nb_labels)
# batch_img = np.concatenate(utils.read_images_and_return_list(args, image_paths, 'local'), axis=1)
# -
# ## Now that we have frames from all subjects, we can load the model we want to test.
# Choose a model to work on
best_model_path = '../models/BEST_MODEL_inception_4d_input_from_scratch_rmsprop_LSTMunits_None_CONVfilters_None_dense512_320x180jpg2fps_bs100_rmsprop_all_aug_v4_t0_run1_gc_rerun.h5'
# +
from keras import backend as K
import tensorflow as tf
tf.reset_default_graph()
K.clear_session()
images = tf.placeholder(tf.float32, [batch_size, args.input_width, args.input_height, channels])
labels = tf.placeholder(tf.float32, [batch_size, 2])
model = utils.InceptionNetwork(images, from_scratch=0, path=best_model_path)
# +
sess = K.get_session() # Grab the Keras session where the weights are initialized.
print('model preds: ', model.preds)
cost = (-1) * tf.reduce_sum(tf.multiply(labels, tf.log(model.preds)), axis=1)
print('cost: ', cost)
y_c = tf.reduce_sum(tf.multiply(model.dense_2, labels), axis=1)
print('y_c: ', y_c)
# target_conv_layer = model.c_86 # Choose which layer to study
# target_conv_layer = model.c_94 # Here: last conv.
target_conv_layer = model.mixed10
print('TYPE tcl: ', type(target_conv_layer))
print('tcl: ', target_conv_layer)
target_conv_layer_grad = tf.gradients(y_c, target_conv_layer)[0]
print('tcl grad: ', target_conv_layer_grad)
print('tclg: ', target_conv_layer_grad)
gb_grad = tf.gradients(cost, images)[0] # Guided backpropagation back to input layer
# +
def run_on_one_sequence(sess, model, batch_img, args):
with sess.as_default():
prob = sess.run(model.preds,
feed_dict={images: batch_img,
K.learning_phase(): 0})
print(prob)
gb_grad_value, target_conv_layer_value, target_conv_layer_grad_value = sess.run([gb_grad, target_conv_layer, target_conv_layer_grad],
feed_dict={images: batch_img,
labels: batch_label,
K.learning_phase(): 0})
# print('tcl: ')
# print(target_conv_layer_value)
# print('')
# print('tclg: ')
# print(target_conv_layer_grad_value)
utils.visualize_overlays(batch_img, target_conv_layer_value,
target_conv_layer_grad_value, args)
# -
# # Inception predictions for subject 1
# +
batch_img, batch_label = utils.data_for_one_random_sequence_4D(args, subject_dfs, computer='local', subject=0)
batch_img = np.array(batch_img, dtype=np.float32)
batch_label = np.array(batch_label, dtype=np.uint8)
run_on_one_sequence(sess, model, batch_img, args)
# +
batch_img, batch_label = utils.data_for_one_random_sequence_4D(args, subject_dfs, computer='local', subject=0)
batch_img = np.array(batch_img, dtype=np.float32)
batch_label = np.array(batch_label, dtype=np.uint8)
run_on_one_sequence(sess, model, batch_img, args)
# +
batch_img, batch_label = utils.data_for_one_random_sequence_4D(args, subject_dfs, computer='local', subject=0)
batch_img = np.array(batch_img, dtype=np.float32)
batch_label = np.array(batch_label, dtype=np.uint8)
run_on_one_sequence(sess, model, batch_img, args)
# +
batch_img, batch_label = utils.data_for_one_random_sequence_4D(args, subject_dfs, computer='local', subject=0)
batch_img = np.array(batch_img, dtype=np.float32)
batch_label = np.array(batch_label, dtype=np.uint8)
run_on_one_sequence(sess, model, batch_img, args)
# +
batch_img, batch_label = utils.data_for_one_random_sequence_4D(args, subject_dfs, computer='local', subject=0)
batch_img = np.array(batch_img, dtype=np.float32)
batch_label = np.array(batch_label, dtype=np.uint8)
run_on_one_sequence(sess, model, batch_img, args)
# +
batch_img, batch_label = utils.data_for_one_random_sequence_4D(args, subject_dfs, computer='local', subject=0, start_index=684)
batch_img = np.array(batch_img, dtype=np.float32)
batch_label = np.array(batch_label, dtype=np.uint8)
run_on_one_sequence(sess, model, batch_img, args)
# -
| 6,536 |
/analysis/.ipynb_checkpoints/study_4_bdes-checkpoint.ipynb | a281bd65e9452f77249b771069aecb8ebcb31090 | [
"BSD-3-Clause"
] | permissive | johngigantic/supramodal_arousal | https://github.com/johngigantic/supramodal_arousal | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 288,688 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
# <h4 align='Left'>Data Merging</h4>
# +
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
# Required data saved into a single file
data=pd.read_csv('tabl1.txt',names=["ID","mag_i","vartype"])
data1=pd.read_csv('tabl2.txt', sep=",",header=None ,names=["ID","Mag","MJD"])
#result = pd.merge(data, data1, on='ID',how='left')
result = data[['ID','vartype']].merge(data1[['ID','Mag','MJD']], on = 'ID',how = 'left')
#result.drop(result.index[[1]])
result.to_csv('mergex.txt', encoding='utf-8',index=False,header=None)
print(result)
#header=none..removes the name of columns from txt file
#index=true..writes row names in txt file
# -
# <h3 align='Left'>dmdt mapping</h3>
# +
ID,vartype,MJD,Mag=np.loadtxt('mergex1.txt', unpack=True, delimiter=',')
#Finding out dm, dt values and hence dmdt pairs
c=[]
e=[]
for i in range(0,len(Mag)):
for j in range(i+1,len(Mag)):
c.append(Mag[i]-Mag[j])
#print(c)
for i in range(0,len(MJD)):
for j in range(i+1,len(MJD)):
e.append(MJD[j]-MJD[i])
#print(e)
s1 = pd.Series(c, name='dm_y')
s2 = pd.Series(e,name='dt_x')
#Assigning bins to all dmdt values
bins=[-8,-5,-3,-2.5,-2,-1.5,-1,-0.5,-0.3,-0.2,-0.1,0,0.1,0.2,0.3,0.5,1,1.5,2,2.5,3,5,8]
bins1=[1/145,2/145,3/145,4/145,1/25,2/25,3/25,1.5,2.5,3.5,4.5,5.5,7,10,20,30,60,90,120,240,600,960,2000,4000] #bins of x axis
return_bins_of_s2= np.digitize(s2, bins1) # np.digitize gives the bin number to which a particular value of dm, dt belongs to.
return_bins_of_s1 = np.digitize(s1, bins)
ybins = pd.Series(return_bins_of_s1, name='ybin')
xbins = pd.Series(return_bins_of_s2, name='xbin')
dmdt=pd.concat([s1, s2,ybins,xbins], axis=1)
dmdt.to_csv('checkxbins.txt',encoding='utf-8')
print(dmdt)
# exclude all the values from dataframe dmdt that are outside the bin boundries i.e. bin number 0, 23 and 24
dmdt=dmdt[dmdt.ybin != 23]
dmdt=dmdt[dmdt.xbin != 0]
dmdt=dmdt[dmdt.ybin != 0]
dmdt=dmdt[dmdt.xbin != 24]
dmdt['bins_xaxis']=pd.cut(dmdt['dt_x'],bins1) #pd.cut gives range of bin for all dt_x values
dmdt['bins_yaxis']=pd.cut(dmdt['dm_y'],bins)
dmdt['merged_dmdt']= dmdt.dm_y.map(str) + ',' + dmdt.dt_x.map(str)
dmdt['dmdt_bin_number']= dmdt.ybin.map(str) + ',' + dmdt.xbin.map(str)
dmdt.dropna() # drops all the values with NaN in dataframe dmdt
print(dmdt)
dmdtpairs=len(dmdt.dm_y) # number of dmdtpairs for a light curve of length n that falls under the given bin range
print(dmdtpairs)
#grouping and counting all the values that fall in all individual bins
df=dmdt.groupby('dmdt_bin_number',sort=False).merged_dmdt.agg(['count']) #counts how many merged_dmdt benlongs to each 'dmdt_bin_number' range
print(df)
df.to_csv('count1.txt',encoding='utf-8')
data_count=pd.read_csv('count1.txt')
ldf=dmdt.set_index('dmdt_bin_number').join(data_count.set_index('dmdt_bin_number'))
abcc=ldf.dropna()
final=abcc.drop_duplicates(['ybin','xbin']) # drops all the rows where both of values of ybin and xbin are duplicate
final1=final.drop(['bins_xaxis','bins_yaxis','merged_dmdt'],axis=1)
print(final1)
final1.to_csv('final_table11.txt',encoding='utf-8',header=None,index=False)
# +
import os, numpy, PIL
from PIL import Image
# Access all PNG files in directory
allfiles=os.listdir("images_sv6")
imlist=[filename for filename in allfiles if filename[-4:] in [".png",".PNG"]]
N=len(imlist)
print(N)
im = Image.open('Var6_fig0.png')
w,h = im.size
# Alternative method using PIL blend function
def average_img_1(imlist):
# Assuming all images are the same size, get dimensions of first image
w,h=Image.open(imlist[0]).size
N=len(imlist)
# Create a numpy array of floats to store the average (assume RGB images)
arr=np.zeros((h,w),np.float)
# Build up average pixel intensities, casting each image as an array of floats
for im in imlist:
imarr=np.array(Image.open(im),dtype=np.float)
arr=arr+imarr/N
out = Image.fromarray(arr)
return out
def average_img_2(imlist):
# Alternative method using PIL blend function
N = len(imlist)
avg=Image.open(imlist[0])
for i in xrange(1,N):
img=Image.open(imlist[i])
avg=Image.blend(avg,img,1.0/float(i+1))
return avg
def average_img_3(imlist):
# Alternative method using numpy mean function
images = np.array([np.array(Image.open(fname)) for fname in imlist])
arr = np.array(np.mean(images, axis=(0)), dtype=np.uint8)
out = Image.fromarray(arr)
return out
# -
# +
import os, numpy, PIL
from PIL import Image
# Access all PNG files in directory
allfiles=("images_sv6")
imlist=[filename for filename in allfiles if filename[-4:] in [".png",".PNG"]]
N=len(imlist)
print(N)
# Assuming all images are the same size, get dimensions of first image
#with Image.open(imlist[0]) as img:
# w,h = img.size
#img = Image.open(imlist[0])
#width, height = img.size
w,h=Image.open("Var6_fig0").size
N=len(imlist)
# Create a numpy array of floats to store the average (assume RGB images)
arr=numpy.zeros((h,w,3),numpy.float)
# Build up average pixel intensities, casting each image as an array of floats
for im in imlist:
imarr=numpy.array(Image.open(im),dtype=numpy.float)
arr=arr+imarr/N
# Round values in array and cast as 8-bit integer
arr=numpy.array(numpy.round(arr),dtype=numpy.uint8)
# Generate, save and preview final image
out=Image.fromarray(arr,mode="RGB")
out.save("Average.png")
out.show()
# -
#
# <h1 align='Left'>Heat Map</h1>
# +
dm,dt,xbin,ybin,count=np.loadtxt('final_table11.txt',unpack=True,delimiter=',',dtype=int)
# assignig intensity values to each bin. Bins are in the form of 23x24 numpy array
aa = np.zeros(shape=(22,23))
for gg in range(0,len(xbin)):
l1=xbin[gg]
m1=ybin[gg]
o1=count[gg]
i1=(255 * o1)/(dmdtpairs + 0.99999)
print(i1)
aa[l1-1,m1-1]=i1
print (aa)
#df2 = pd.DataFrame(data,dtype=float)
#print(df2)
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace = go.Heatmap(aa)
data=[trace]
py.iplot(data, filename='basic-heatmap')
# +
import matplotlib.pyplot as plt
import numpy as np
plt.imshow(aa,cmap='jet')
plt.show()
# -
# <h2 align='Left'>Heat Map Plot</h2>
import seaborn as sns
sns.set()
image11=sns.heatmap(aa,xticklabels=False, yticklabels=False,cmap='jet')
#image11=sns.heatmap(aa,cbar=False,xticklabels=False, yticklabels=False)
#vmin=0, vmax=255,
#cmap='RdYlBu'// colormap of heatmap
#sns_plot.figure.savefig("output.png")
image11.figure.savefig("output_image1.png")
| 6,901 |
/MNIST.ipynb | 9393bdb1480006ec5956cc64f4aacdeed0619f23 | [] | no_license | gunny-probablistically/amlb | https://github.com/gunny-probablistically/amlb | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 218,068 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tpot import TPOTClassifier
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
digits = load_digits()
X_train, X_test, y_train, y_test = train_test_split(digits.data, digits.target,
train_size=0.75, test_size=0.25)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
from config.classifier_models_only import classifier_config_dict
time_allocated = 60
tpot = TPOTClassifier(max_time_mins=time_allocated, config_dict=classifier_config_dict, verbosity=3)
tpot.fit(X_train, y_train)
print(tpot.score(X_test, y_test))
tpot.export('tpot_mnist_pipeline.py')
# +
# # %load tpot_mnist_pipeline.py
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
# NOTE: Make sure that the class is labeled 'class' in the data file
tpot_data = np.recfromcsv('PATH/TO/DATA/FILE', delimiter='COLUMN_SEPARATOR', dtype=np.float64)
features = np.delete(tpot_data.view(np.float64).reshape(tpot_data.size, -1), tpot_data.dtype.names.index('class'), axis=1)
training_features, testing_features, training_classes, testing_classes = \
train_test_split(features, tpot_data['class'], random_state=42)
exported_pipeline = KNeighborsClassifier(n_neighbors=4, p=2, weights="distance")
exported_pipeline.fit(training_features, training_classes)
results = exported_pipeline.predict(testing_features)
# -
# # XGBOOST Experiment
# +
import warnings
warnings.filterwarnings('ignore')
from sklearn.model_selection import train_test_split
from src.hypersearch import xgbc_random
from config.xgboost import config_dict
# -
xgbc_random(
config_dict,
3600,
X_train,
y_train,
X_test,
y_test)
rop_duplicates())
# ## Make sure that risks magnitude (which is going to be used as an entry label) has a single value
cwr_agg.magnitude = cwr_agg.magnitude.apply(lambda x: 'Extremely serious' if 'Extremely serious' in x else ('Serious' if 'Serious' in x else list(x)[0]))
cwr_agg.magnitude.unique()
# ## Merge (outer) datasets and find the columns that are set (list_columns)
# filling the non-merged rows with {np.nan} on list_columns to make sure there is contingency in each column
cw_data = pd.merge(cwa_agg, cwr_agg, on=id_columns, how='outer')
list_columns = cw_data.iloc[0].apply(lambda x: isinstance(x, set))
list_columns = list_columns[list_columns].index
cw_data[list_columns] = cw_data[list_columns].applymap(lambda x: {np.nan} if pd.isna(x) else x)
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
cw_data.head()
# ## Extract longitude and latitude from city_location
cw_data['city_location'] = cw_data['city_location'].apply(eval)
cw_data['latitude'] = cw_data['city_location'].apply(lambda x: x[0])
cw_data['longitude'] = cw_data['city_location'].apply(lambda x: x[1])
id_columns = [x for x in id_columns] + ['latitude','longitude']
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
# ## Find list columns that are singular (every element contains a length 1 list)
# +
cw_value_columns = [ x for x in cw_data.columns if x not in id_columns]
# Values that can be converted back to strings from sets
singular_cols = cw_data.set_index(id_columns)[list_columns].applymap(lambda x: len(x)==1).product() > 0
singular_cols
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
# -
# ## Convert singular columns to non list ones
# magnitude is nicely singular per id
cw_data.loc[:, singular_cols[singular_cols].index] = cw_data.loc[:, singular_cols[singular_cols].index].applymap(lambda x: list(x)[0])
list_columns = [x for x in list_columns if x not in singular_cols[singular_cols].index]
# ## Remove entries that are invalid or in different language than English from the list columns
# **also convert sets to lists**
def try_lang_detect_except(x):
try:
return (langdetect(x)=='en')
except:
return False
cw_data = cw_data.applymap(lambda x: [t for t in x if not pd.isnull(t) and (t!='') and (t!='ERROR: #NAME?') and try_lang_detect_except(t)] if isinstance(x,set) else x)
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
# ## Fix specific risks annotations by inspection
cw_data.risks_to_citys_water_supply.unique()
cw_data.risks_to_citys_water_supply = cw_data.risks_to_citys_water_supply.apply(lambda x: x.replace('Inadequate or ageing infrastructure','Inadequate or aging infrastructure'))
cw_data['risks_to_citys_water_supply'] = cw_data['risks_to_citys_water_supply'].apply(lambda x: x.replace('Declining water quality: Declining water quality', 'Declining water quality'))
cw_data.risks_to_citys_water_supply.unique()
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
cw_data.drop_duplicates(subset=id_columns,inplace=True)
cw_data.head()
to_drop = ['timescale','current_population','population_year']
cw_data['population_year'] = cw_data['population_year'].fillna(0)
t = cw_data.sort_values('population_year' ).groupby([x for x in cw_data.columns if (x !='population_year') and x not in list_columns],dropna=False).last().reset_index()
cw_data
# ## Drop metadata columns that will not be used
cw_data.drop(columns=['timescale','population_year'],inplace=True)
cw_data.rename(columns={'current_population':'population'},inplace=True)
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
# ## Generate for each of the list columns a new one that contains the number of elements
# **Create value_columns that contains the names of all the list columns + the created ones**
value_columns = set([x for x in cw_value_columns if x not in to_drop] + ['risks_to_citys_water_supply'])
list_columns = cw_data.iloc[0].apply(lambda x: isinstance(x, list))
list_columns = list_columns[list_columns].index
len_columns = set()
for x in value_columns.copy():
if x in list_columns:
cw_data[x+'_n'] = cw_data[x].apply(lambda x: len(x) if x else 1)
len_columns.add(x+'_n')
value_columns = value_columns.union(len_columns)
value_columns
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
# # Filter by C40
# **We are making the assumption that, given that those cities participate in a specific program, they will also have better structured data. So this subset will be used as a reference for the imputation**
c40_data = cw_data[cw_data['c40']].copy()
c40_data.shape
# ## Find the null percentage of list columns
list_nulls = c40_data[list_columns].applymap(lambda x: len(x) == 0)
list_nulls.mean()
c40_data['magnitude'].unique()
# ## Create a report of the values of magnitude
c40_risks = c40_data[id_columns+['magnitude']].copy()
c40_risks['low'] = c40_risks['magnitude'] == 'Less Serious'
c40_risks['medium'] = c40_risks['magnitude'] == 'Serious'
c40_risks['high'] = c40_risks['magnitude'] == 'Extremely serious'
c40_risks['unknown'] = c40_risks['magnitude'].isna()
c40_risks = c40_risks.drop(columns=['magnitude']).groupby([x for x in id_columns if x!="risks_to_citys_water_supply"]).agg(sum).reset_index()
c40_risks
from sklearn.preprocessing import LabelEncoder
from utils.nlp import SimilarityAnalysis, create_sim_vector
c40_data.risks_to_citys_water_supply = c40_data.risks_to_citys_water_supply.apply(lambda x: np.nan if x=='nan' else x)
c40_data[c40_data.risks_to_citys_water_supply.isnull()]
c40_data.risks_to_citys_water_supply.unique()
# ## Fit Similarity Analysis
# **per string list column of the value columns, by concatenating strings together. Vectorize all the string list columns based on their in between similarities**
#
analyses = {x:SimilarityAnalysis() for x in value_columns if x in list_columns}
vectorized = np.hstack([analyses[x].fit_transform(c40_data[x]) for x in analyses])
# ## Encode magnitude and risks into numerical labels
cols_to_encode = [x for x in value_columns if x not in list_columns and x not in len_columns]
c40_data[cols_to_encode] = c40_data[cols_to_encode].fillna('nan')
encoders = {x:LabelEncoder() for x in cols_to_encode}
encoded = np.array([encoders[x].fit_transform(c40_data[x]) for x in cols_to_encode]).T.astype(float)
for t in range(len(cols_to_encode)):
encoded[encoded[:,t] == np.where(encoders[cols_to_encode[t]].classes_=='nan')[0],t] = np.nan
lab_freqs = c40_data[cols_to_encode].groupby(cols_to_encode,as_index=False).size()
lab_freqs
# ## Create dataset that contains the encoded labels, the vectorized columns and the columns referring the lists lengths
whole = np.hstack([encoded, vectorized, c40_data[len_columns]])
import sys
sys.path.append('..')
from labeled_preprocessing.imputation import LabeledDatasetImputer
# ## Fit LabeledDatasetImputer and transform dataset
# **(LabeledDatasetImputer is a union of two Iterative Imputers , one for labels and one for features)**
imputer = LabeledDatasetImputer(verbose=2,k_features_per_label=0, seed=seed)
continuous_imputed, y = imputer.fit_transform(whole[:,2:], whole[:,:2], ret_imputed_x=True)
# ## Use the prediction power on the imputed data as measurement of coherent imputation.
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
simple_model = make_pipeline(StandardScaler(), SVC(kernel='linear', C=0.01, random_state=seed))
scoring = 'accuracy'
scores = cross_val_score(simple_model, continuous_imputed, y[:,cols_to_encode.index('magnitude')], cv=5,scoring=scoring)
np.mean(scores)
# ## Plot features correlation
import matplotlib.pyplot as plt
flg = np.all(~np.isnan(whole),axis=1), np.hstack([[True, True], imputer.selection_mask[:]])
corr = np.corrcoef(whole[flg[0],:][:,flg[1]].T)
print(corr.shape)
plt.matshow(corr)
# ## Retrieve the categorical imputed labels
imputed = np.array([
encoders[x].inverse_transform(
y[:, c].astype(int)
) for c,x in enumerate(cols_to_encode)]).T
c40_data_imputed = c40_data.copy()
c40_data_imputed[cols_to_encode] = imputed
c40_data_imputed.head()
# ## Find which labels have been updated after imputation
imp_lab_freqs = c40_data_imputed[cols_to_encode].groupby(cols_to_encode,as_index=False).size()
imp_ret = pd.merge(lab_freqs, imp_lab_freqs,suffixes=('','_imp'), how='right',on=cols_to_encode)
imp_ret['increase'] = (imp_ret['size_imp'] - imp_ret['size'])/imp_ret['size']
imp_ret
# ## Make sure that all risks have been encoded and are imputation outcome candidates
set([x for x in cw_data['risks_to_citys_water_supply'] if x not in encoders['risks_to_citys_water_supply'].classes_])
# # Impute whole dataset
cw_data['risks_to_citys_water_supply'].value_counts()
# ## Apply encoding and vectorization
# **based on fitted encoders and SimilarityAnalysis objects**
cw_data[cols_to_encode] = cw_data[cols_to_encode].fillna('nan')
encoded = np.array([encoders[x].transform(cw_data[x]) for x in cols_to_encode]).T.astype(float)
for t in range(len(cols_to_encode)):
encoded[encoded[:,t] == np.where(encoders[cols_to_encode[t]].classes_=='nan')[0],t] = np.nan
all_vectorized = np.hstack([analyses[x].transform(cw_data[x]) for x in value_columns if x in list_columns])
# ## Apply imputation
# **using the trained imputer**
all_imputed_x, all_imputed_y = imputer.transform(np.hstack([all_vectorized,cw_data[len_columns]]), encoded, ret_imputed_x=True)
all_imputed_y_dec = np.array([
encoders[x].inverse_transform(
all_imputed_y[:, c].astype(int)
) for c,x in enumerate(cols_to_encode)]).T
len(cw_data[id_columns]), len(cw_data[id_columns].drop_duplicates())
cw_data_imputed = cw_data.copy()
cw_data_imputed[cols_to_encode] = all_imputed_y_dec
cw_data_imputed.drop_duplicates(id_columns,inplace=True)
# ## Plot features correlation
import matplotlib.pyplot as plt
corr = np.corrcoef(np.hstack([all_imputed_y, all_imputed_x]).T)
print(corr.shape)
plt.matshow(corr)
scores = cross_val_score(simple_model, all_imputed_x, all_imputed_y[:,cols_to_encode.index('magnitude')], cv=5,scoring=scoring)
np.mean(scores)
# **The score did not have significant drop, so we can assume that the imputation was cohesive across data**
# ## Create output dataset
final_labeled_data = cw_data_imputed[['city','latitude','longitude','country','population','c40','magnitude','risks_to_citys_water_supply']].copy()
final_labeled_data.rename(columns={'magnitude': 'risk','risks_to_citys_water_supply': 'description'},inplace=True)
final_labeled_data[['city','latitude','longitude','c40', 'population','description','risk']]
cw_data_imputed[id_columns].drop_duplicates().shape
# ## Pivoting risks description, so that each row is unique per city
risks = final_labeled_data.description.unique()
risks_description = {risk: f'risk{c}' for c,risk in enumerate(risks)}
risks_df = pd.Series(risks_description).to_frame()
risks_df.reset_index(inplace=True)
risks_df.columns=['description','code']
final_labeled_data['description'] = final_labeled_data['description'].apply(lambda x: risks_description[x])
risks_df
main_index=['city','latitude','longitude', 'country']
# ## Check if there are cities that are listed to be both in and not in C40
# and drop the ones that are not
c40_check = final_labeled_data.groupby(main_index)['c40'].nunique()
c40_check[c40_check>1]
final_labeled_data.city.nunique()
final_labeled_data[final_labeled_data.city == 'Santiago']
final_labeled_data = final_labeled_data.sort_values(by='c40').groupby(main_index + ['description']).last().reset_index()
final_labeled_data = final_labeled_data.drop(columns=['c40']).merge(final_labeled_data.groupby(main_index)['c40'].max().reset_index(),on=main_index)
final_labeled_data[final_labeled_data.city=='Santiago']
final_labeled_data = final_labeled_data.sort_values(by='c40').groupby(main_index + ['description']).last().reset_index()
# ## Fix population
pop_df = final_labeled_data.groupby(index)[['population']
].max().reset_index().drop_duplicates()
final_labeled_data = final_labeled_data.drop(columns=['population']).merge(pop_df.reset_index(), on=index)
final_labeled_data = final_labeled_data.pivot(index= index + ['c40','population'], columns='description', values='risk').reset_index()
# ## Encode risks from 0 to 1
severity_mapping = {'Less Serious':1, "Serious":2, 'Extremely serious':3}
pd.DataFrame([severity_mapping]).to_csv(SEVERITY_MAPPING_PATH,index=False)
nrisks = list(risks_description.values())
final_labeled_data[nrisks] = final_labeled_data[nrisks].replace(severity_mapping)
# ### Add 0 to risks belonging to C40, assuming that those have reported no risks because indeed they were none, not because they were unknown
final_labeled_data.loc[final_labeled_data['c40'], nrisks] = final_labeled_data.loc[final_labeled_data['c40'], nrisks].fillna(0)
final_labeled_data.to_csv(LABELED_CITIES_PATH,index=False)
risks_df.to_csv(RISKS_MAPPING_PATH,index=False)
final_labeled_data.head()
# ## See a report of the filled labels
cw_lab_freqs = cw_data[cols_to_encode].groupby(cols_to_encode,as_index=False).size()
imp_cw_lab_freqs = cw_data_imputed[cols_to_encode].groupby(cols_to_encode,as_index=False).size()
imp_cw_ret = pd.merge(cw_lab_freqs, imp_cw_lab_freqs,suffixes=('','_imp'), how='right',on=cols_to_encode)
imp_cw_ret['increase'] = (imp_cw_ret['size_imp'] - imp_cw_ret['size'])/imp_cw_ret['size']
imp_cw_ret
imp_report = pd.merge(imp_cw_ret, imp_ret, suffixes=('','_c40'), on=('magnitude','risks_to_citys_water_supply'))
imp_report.to_csv(IMPUTATION_REPORT_PATH,index=False)
imp_report
| 15,906 |
/pyDS_KHTN/VI_DU/B6_MATPLOTLIB/waffle_plot.ipynb | 57053c2654a26c9e7e558ec66f23f70183a923e8 | [] | no_license | danhtran2k20/pyData | https://github.com/danhtran2k20/pyData | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 46,459 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numpy高级
# 初始化环境
import numpy as np
# ## Numpy广播机制
a = np.arange(3)
a
b = 1
a + b
a = np.arange(6).reshape(2, -1)
a
b = [0.1, 0.2, 0.3]
a + b
# 广播机制也是有要求的
a = np.arange(6).reshape(2, 3)
a
b = np.array([0.1, 0.2, 0.3, 0.4])
# a + b
# ## Where条件
a = np.array([1, 2, 3, 5, 7, 6, 6, 8, 6])
a == 6
# 统一把6换成9 => a==6条件,9符合条件设置的值, a不符合条件设置的值
np.where(a == 6, 9, a)
# ## Numpy矩阵运算
# ### 矩阵的乘法
# +
a = np.array([
[20, 5],
[25, 10],
])
b = np.array([2, 1])
# 矩阵点乘
# a @ b
# a.dot(b)
np.dot(a, b)
# +
# 解决鸡兔同笼问题
a = np.array([
[1, 1],
[2, 4]
])
b = np.array([35, 94])
np.linalg.solve(a, b)
# -
# ### 矩阵的逆
# +
# 解决鸡兔同笼问题
a = np.array([
[1, 1],
[2, 4]
])
b = np.array([35, 94])
np.linalg.inv(a) @ b
# đơn giá trung bình theo mỗi nước
df = df.head(10)
df['price'].fillna(0.1, inplace=True)
fig = plt.figure(
FigureClass=Waffle,
rows=df.shape[0],
values=df.price,
labels=list(df.index),
figsize=(10, 5),
legend={'loc': 'upper left', 'bbox_to_anchor': (1, 1)
}
)
plt.show()
| 1,308 |
/docs/user_guide/main_usage/lagrange_polynomials.ipynb | 0f52bd53b5e5039a17813375b615386b763b6e01 | [
"MIT"
] | permissive | jonathf/chaospy | https://github.com/jonathf/chaospy | 405 | 87 | MIT | 2023-05-18T11:52:46 | 2023-05-09T17:26:01 | Python | Jupyter Notebook | false | false | .py | 45,047 | # ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# ## Lagrange Polynomials
#
# This tutorial uses the same example as the [problem formulation](./problem_formulation.ipynb).
#
# [Lagrange polynomials](https://en.wikipedia.org/wiki/Lagrange_polynomial) are not a method for creating orthogonal polynomials.
# Instead it is an interpolation method for creating an polynomial expansion that has the property that each polynomial interpolates exactly one point in space with the value 1 and has the value 0 for all other interpolation values.
#
# To summarize, we need to do the following:
#
# * Generate $Q_1, ..., Q_N = (\alpha_1, \beta_1), ..., (\alpha_N, \beta_N)$, using (pseudo-)random samples or otherwise.
# * Construct a Lagrange polynomial expansion $\Phi_1, ..., \Phi_M$ from the samples $Q_1, ..., Q_N$.
# * Evaluate model predictor $U_1, ..., U_N$ for each sample.
# * Analyze model approximation $u(t; \alpha, \beta) = \sum_m U_m(t) \Phi_n(\alpha, \beta)$ instead of actual model solver.
# %% [markdown]
# ### Evaluation samples
#
# In chaospy, creating low-discrepancy sequences can be done using the `distribution.sample` method.
# Creating quadrature points can be done using `chaospy.generate_quadrature`.
# For example:
# %%
from problem_formulation import joint
samples = joint.sample(5, rule="halton")
# %%
from matplotlib import pyplot
pyplot.scatter(*samples)
pyplot.rc("figure", figsize=[6, 4])
pyplot.xlabel("Parameter $I$ (normal)")
pyplot.ylabel("Parameter $a$ (uniform)")
pyplot.axis([1, 2, 0.1, 0.2])
pyplot.show()
# %% [markdown]
# ### Lagrange basis
#
# Creating an expansion of Lagrange polynomial terms can be done using the following constructor:
# %%
import chaospy
polynomial_expansion = chaospy.expansion.lagrange(samples)
polynomial_expansion[0].round(2)
# %% [markdown]
# On can verify that the returned polynomials follows the property of evaluating 0 for all but one of the samples used in the construction as follows:
# %%
polynomial_expansion(*samples).round(8)
# %% [markdown]
# ### Model approximation
#
# Fitting the polynomials to the evaluations does not need to involve regression.
# Instead it is enough to just multiply the Lagrange polynomial expansion with the evaluations, and sum it up:
# %%
from problem_formulation import model_solver
model_evaluations = numpy.array([
model_solver(sample) for sample in samples.T])
model_approximation = chaospy.sum(
model_evaluations.T*polynomial_expansion, axis=-1).T
model_approximation.shape
# %% [markdown]
# ### Assess statistics
#
# The results is close to what we are used to from the other methods:
# %%
expected = chaospy.E(model_approximation, joint)
variance = chaospy.Var(model_approximation, joint)
# %%
from problem_formulation import coordinates
pyplot.fill_between(coordinates, expected-variance**0.5,
expected+variance**0.5, alpha=0.3)
pyplot.plot(coordinates, expected)
pyplot.axis([0, 10, 0, 2])
pyplot.xlabel("coordinates $t$")
pyplot.ylabel("Model evaluations $u$")
pyplot.show()
# %% [markdown]
# ### Avoiding matrix inversion issues
#
# It is worth noting that the construction of Lagrange polynomials are not always numerically stable.
# For example when using grids along , most often the expansion construction fails:
# %%
nodes, _ = chaospy.generate_quadrature(1, joint)
try:
chaospy.expansion.lagrange(nodes)
except numpy.linalg.LinAlgError as err:
error = err.args[0]
error
# %% [markdown]
# It is impossible to avoid the issue entirely, but in the case of structured grid, it is often possible to get around the problem.
# To do so, the following steps can be taken:
#
# * Create multiple Lagrange polynomials, one for each marginal distribution.
# * Rotate dimensions so each expansion get a dimension corresponding to the marginal it was created for.
# * Use an outer product of the expansions to combine them into a single expansion.
#
# So we begin with making marginal Lagrange polynomials:
# %%
nodes, _ = list(zip(*[chaospy.generate_quadrature(1, marginal)
for marginal in joint]))
expansions = [chaospy.expansion.lagrange(nodes_)
for nodes_ in nodes]
[expansion_.round(4) for expansion_ in expansions]
# %% [markdown]
# Then we rotate the dimensions:
# %%
vars_ = chaospy.variable(len(joint))
expansions = [
expans(q0=var)
for expans, var in zip(expansions, vars_)]
[expans.round(4) for expans in expansions]
# %% [markdown]
# And finally construct the final expansion using an outer product:
# %%
expansion = chaospy.outer(*expansions).flatten()
expansion.round(2)
# %% [markdown]
# The end results can be verified to have the properties we are looking for:
# %%
nodes, _ = chaospy.generate_quadrature(1, joint)
expansion(*nodes).round(8)
| 5,053 |
/Principal Component Analysis(PCA)/PCA_on_Breast_Cancer_Dataset.ipynb | d1034d82ecef2874bcf30fc72a48e9aa27aa8576 | [] | no_license | gourab-sinha/Machine_Learning | https://github.com/gourab-sinha/Machine_Learning | 1 | 1 | null | null | null | null | Jupyter Notebook | false | false | .py | 5,903 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gourab-sinha/Machine_Learning/blob/master/Principal%20Component%20Analysis(PCA)/PCA_on_Breast_Cancer_Dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PZLsDzCWo1UG" colab_type="code" colab={}
from sklearn import decomposition,datasets,linear_model
import numpy as np
import time
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
# + id="pOQIUhVXpkUt" colab_type="code" outputId="b1b76083-0d78-414f-fd32-11359dce5b1b" colab={"base_uri": "https://localhost:8080/", "height": 34}
bcancer = datasets.load_breast_cancer()
x = bcancer.data
x.shape
# + id="DwZ_SavtqCYj" colab_type="code" colab={}
sc = StandardScaler()
X_std = sc.fit_transform(x)
x_train,x_test,y_train,y_test = train_test_split(X_std,bcancer.target,random_state=0)
# + id="BI_kUI03qnWz" colab_type="code" colab={}
pca = decomposition.PCA(n_components=15)
x_train_pca = pca.fit_transform(x_train)
x_test_pca = pca.fit_transform(x_test)
# + id="OH1hb27Eq_dv" colab_type="code" outputId="8a7e073a-29b6-490a-bc45-c4cadd2540df" colab={"base_uri": "https://localhost:8080/", "height": 51}
lr = linear_model.LogisticRegression()
start = time.time()
lr.fit(x_train,y_train)
end = time.time()
print(end-start)
print(lr.score(x_test,y_test))
# + id="ioKgq_YOppjM" colab_type="code" outputId="a3bcd52c-9711-4932-a126-beace2babf77" colab={"base_uri": "https://localhost:8080/", "height": 51}
lr = linear_model.LogisticRegression()
start = time.time()
lr.fit(x_train_pca,y_train)
end = time.time()
print(end-start)
print(lr.score(x_test_pca,y_test))
# + [markdown] id="OG2qEq3igC85" colab_type="text"
# # Choose No of Component
# #### Using varrience we can find the no of components i.e no of features. In this example I have take 95% varrience and for which I will need only 10 components i.e features.
# ##### NOTE: We aren't picking only 10 features from the given set of features we are making new 10 features using provided features of N.
# + id="8dGfjhU1eepG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e67700d6-6491-41e3-a5c1-48dbc5c01af8"
pca = decomposition.PCA()
pca.fit_transform(x_train)
total = sum(pca.explained_variance_)
k = 0
current_varrience = 0
while current_varrience/total<0.95:
current_varrience+= pca.explained_variance_[k]
k+=1
k
| 2,751 |
/models/text_classification.ipynb | 99ec6f02044f6a9b0b1965842b69da54411494bf | [
"MIT"
] | permissive | taswani/ML_capstone | https://github.com/taswani/ML_capstone | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 10,830 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NLP Sentiment Analysis notebook
# Importing all necessary modules
import data_pipeline as dp
from textblob import TextBlob
# Running all the functions from the data pipeline to get us the results dataframe and the processed features
result_df = dp.prepare_data(dp.price_csv, dp.headline_csv)
processed_features = dp.vectorization(result_df)
result_df
# Look to use pre-trained model, aggregrate the predictions and append to result_df
def sentiment_analysis(result_df, processed_features):
# List for taking in the sentiments associated with the processed features
sentiments = []
polarity = []
# Using a pre-trained model to analyze sentiment for each headline
for feature in processed_features:
sentence = TextBlob(feature)
polarity.append(sentence.sentiment.polarity)
if sentence.sentiment.polarity > 0:
sentiments.append(1)
else:
sentiments.append(0)
# Adding a new column in the dataframe and returning it
result_df['Sentiment'] = sentiments
result_df['Polarity'] = polarity
# pushing df to a csv
result_df.to_csv("final_amazon.csv")
return result_df
how to get a Carbonate account, how to set up ReD, and how to get started using the Jupyter Notebook on ReD.
# ### Run CyberDH environment
# The code in the cell below points to a Python environment specifically for use with the Python Jupyter Notebooks created by Cyberinfrastructure for Digital Humanities. It allows for the use of the different packages in our notebooks and their subsequent data sets.
#
# ##### Packages
# - **sys:** Provides access to some variables used or maintained by the interpreter and to functions that interact strongly with the interpreter. It is always available.
# - **os:** Provides a portable way of using operating system dependent functionality.
#
# #### NOTE: This cell is only for use with Research Desktop. You will get an error if you try to run this cell on your personal device!!
import sys
import os
sys.path.insert(0,"/N/u/cyberdh/Carbonate/dhPyEnviron/lib/python3.6/site-packages")
os.environ["NLTK_DATA"] = "/N/u/cyberdh/Carbonate/dhPyEnviron/nltk_data"
# ### Include necessary packages for notebook
#
# Python's extensibility comes in large part from packages. Packages are groups of functions, data, and algorithms that allow users to easily carry out processes without recreating the wheel. Some packages are included in the basic installation of Python, others created by Python users are available for download. Make sure to have the following packages installed before beginning so that they can be accessed while running the scripts.
#
# In your terminal, packages can be installed by simply typing `pip install nameofpackage --user`. However, since you are using ReD and our Python environment, you will not need to install any of the packages below to use this notebook. Anytime you need to make use of a package, however, you need to import it so that Python knows to look in these packages for any functions or commands you use. Below is a brief description of the packages we are using in this notebook:
#
#
# - **nltk:** Platform for building Python programs to work with human language data.
#
# - **string:** contains a number of useful constants and classes, as well as some deprecated legacy functions that are also available as methods on strings.
#
# - **collections:** Implements specialized container datatypes providing alternatives to Python's general purpose built-in containers: dict, list, set, and tuple.
#
# - **pandas:** An open source, BSD-licensed library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language.
#
# - **gensim:** Python library for topic modelling, document indexing and similarity retrieval with large corpora.
#
# - **spacy:** A library for advanced Natural Language Processing in Python and Cython.
#
# - **operator:** Exports a set of functions corresponding to the intrinsic operators of Python.
#
# - **plotly:** Graphing library that makes interactive, publication-quality graphs.
#
# - **re:** Provides regular expression matching operations similar to those found in Perl.
#
# - **math:** Provides access to the mathematical functions defined by the C standard.
#
# Notice we import some of the packages differently. In some cases we just import the entire package when we say `import XYZ`. For some packages which are small, or, from which we are going to use a lot of the functionality it provides, this is fine.
#
# Sometimes when we import the package directly we say `import XYZ as X`. All this does is allow us to type `X` instead of `XYZ` when we use certain functions from the package. So we can now say `X.function()` instead of `XYZ.function()`. This saves time typing and eliminates errors from having to type out longer package names. I could just as easily type `import XYZ as potato` and whenever I use a function from the `XYZ` package I would need to type `potato.function()`. What we import the package as is up to you, but some commonly used packages have abbreviations that are standard amongst Python users such as `import pandas as pd` or `import matplotlib.pyplot as plt`. You do not need to us `pd` or `plt`, however, these are widely used and using something else could confuse other users and is generally considered bad practice.
#
# Other times we import only specific elements or functions from a package. This is common with packages that are very large and provide a lot of functionality, but from which we are only using a couple functions or a specific subset of the package that contains the functionality we need. This is seen when we say `from XYZ import ABC`. This is saying I only want the `ABC` function from the `XYZ` package. Sometimes we need to point to the specific location where a function is located within the package. We do this by adding periods in between the directory names, so it would look like `from XYZ.123.A1B2 import LMN`. This says we want the `LMN` function which is located in the `XYZ` package and then the `123` and `A1B2` directory in that package.
#
# You can also import more than one function from a package by separating the functions with commas like this `from XYZ import ABC, LMN, QRS`. This imports the `ABC`, `LMN` and `QRS` functions from the `XYZ` package.
from nltk.corpus import stopwords
import string
from collections import defaultdict
import pandas as pd
import gensim
from gensim.models.phrases import Phrases, Phraser
from gensim.utils import simple_preprocess
import spacy
import operator
import plotly as py
import plotly.express as px
import re
import math
# #### File paths
# Here we are saving as variables different file paths that we need in our code.
#
# First we use the `os` package above to find our 'HOME' directory using the `environ` function. This will work for any operating system, so if you decide to try this out on your personal computer instead of ReD, the `homePath` variable will still be the path to your 'home' directory, so no changes are needed.
#
# Next, we combine the `homePath` file path with the folder name that leads to where our various data directories are stored. Note that we do not use any file names yet, just the path to the folder. This is because we have different data folders in this directory and we may change which one we are using. We assign the path to a variable named `dataHome`.
#
# Now we use the `dataHome` variable as part of two more file paths. The first file path points to a directory containing multiple files. This file path we assign to the variable `corpusRoot`. The next file path points to a directory that contains other directories, all of which contain ".txt" files. This is for if our data is divided into multiple directories and we need to use all of the files in all of the directories. We assign this file path to variable `directRoot`.
#
# Lastly, we add the `homePath` file path to other folder names that lead to a folder where we will want to save any output generated by this code. We save this file path as the variable `dataResults`.
#
# If you are using these notebooks for your own research then your data is stored in a different location you will need to change these file paths to point to where your data resides. Additionally, if you wish for the output generated by this notebook to be saved to a different location, you will need to assign the appropriate file path to the `dataResults` variable.
homePath = os.environ['HOME']
dataHome = os.path.join(homePath, 'Text-Analysis-master', 'data')
corpusRoot = os.path.join(dataHome, 'shakespeareFolger')
directRoot = os.path.join(dataHome, 'starTrek')
dataResults = os.path.join(homePath, 'Text-Analysis-master', 'Output')
# ### Set needed variables
# This is where you will make some decisions about your data and set the necessary variables. We again do this so you will not need to make as many changes to the code later.
#
# **corpusLevel**<br>
# Your data needs to be in a '.txt' file format to work with this notebook. If it is then you need to specify if you want to read in a single document, multiple documents from a single directory, or multiple documents located in multiple directories. If you want to read in a single document, then assign `"lines"` (keep the quotes) to `corpusLevel`. If you want to read in multiple documents from a single directory then assign `"files"` (again, keep quotes) to `corpusLevel`. Lastly, if you want to read in multiple documents stored in multiple directories, then assign `"direct"` to `corpusLevel`.
#
# **nltkStop**<br>
# The `nltkStop` is where you determine if you want to use the built in stopword list provided by the NLTK package. They provide stopword lists in multiple languages. If you wish to use this then set `nltkStop` equal to `True`. If you do not, then set `nltkStop` equal to `False`.
#
# **customStop**<br>
# The `customStop` variable is for if you have a dataset that contains additional stopwords that you would like to read in and have added to the existing `stopWords` list. You do **NOT** need to use the NLTK stopwords list in order to add your own custom list of stopwords. **NOTE: Your custom stopwords file needs to have one word per line as it reads in a line at a time and the full contents of the line is read in and added to the existing stopwords list.** If you have a list of your own then set `customStop` equal to `True`. If you do not have your own custom stopwords list then set `customStop` equal to `False`.
#
# **spacyLem**<br>
# `spacyLem` is where we decide if we want to use the spaCy package lemmatization function. What is lemmatization? Lemmatization is the process of grouping together the inflected forms of a word so they can be analysed as a single item, and identified by the word's lemma, or dictionary form. In computational linguistics, lemmatisation is the algorithmic process of determining the lemma of a word based on its intended meaning. Unlike stemming, lemmatisation depends on correctly identifying the intended part of speech and meaning of a word in a sentence, where as stemming does not take the context of the word into account. For example, if we lemmatize the word "running" or "ran" it will become the word "run". If we stem the word "running" most stemmers will convert it to "runn" only removing the "ing" and leaving the second "n". Stemming will also change "police" and "policy" both to "polic" and they will be considered the same word and will be counted together by the wordcloud algorithm. The lemmatizer will leave both words as "police" and "policy". This is useful for word frequency algorithms as it allows the algorithm to just count "walk" instead of "walking", "walked", and "walk" and thereby can give further insight into the corpus that may not have been evident otherwise. To use the spacy lemmatizer set `spacyLem` equal to `True`. If you do not wish to use the lemmatizer set `spacyLem` equal to `False`.
#
# **stopLang**<br>
# The `stopLang` variable is to choose the language of the nltk stopword list you wish to use. It is currently set to `'english'`. If you need a different language, simply change `'english'` to the anglicized name of the language you wish to use (e.g. 'spanish' instead of 'espanol' or 'german' instead of 'deutsch').
#
# **lemLang**<br>
# Now we choose the language for our lemmatizer. The languages available for spacy include the list below and the abbreviation spacy uses for that language:
#
# - **English:** `"en"`
# - **Spanish:** `"es"`
# - **German:** `"de"`
# - **French:** `"fr"`
# - **Italian:** `"it"`
# - **Portuguese:** `"pt"`
# - **Dutch:** `"nl"`
# - **Multi-Language:** `"xx"`
#
# To choose a language simply type the two letter code following the angliscized language name in the list above and assign it to the variable `lemLang`. So for Spanish it would be `"es"` (with the quotes) and for German `"de"` and so on.
#
# **encoding/errors**<br>
# The variable `encoding` is where you determine what type encoding to use (ascii, ISO-8850-1, utf-8, et cetera). We have it set to `"UTF-8"` at the moment as we have found it is less likely to have any problems. However, errors do occur, but the encoding errors rarely impact our results and it causes the Python code to exit. So instead of dealing with unhelpful errors we ignore the ones dealing with encoding by assigning `"ignore"` to the `errors` variable. If you want to see any encoding errors then change `'ignore'` to `None` without the quotes.
#
# **stopWords**<br>
# The `stopWords = []` is an empty list that will contain the final list of stop words to be removed form your dataset. What ends up in the list depends on whether you set `nltkStop` and/or `customStop` equal to `True` or `False` and if you add any additional words to the list.
corpusLevel = "lines"
nltkStop = True
customStop = True
spacyLem = True
stopLang = 'english'
lemLang = "en"
encoding = "utf-8"
errors = "ignore"
stopWords = []
# ### Stopwords
# If you set `nltkStop` equal to `True` above then this will add the nltk stopwords list to the empty list named `stopWords`. You should already have chosen your language above, so there is no need to do that here.
#
# If you wish to add additional words to the `stopWords` list, add the word in quotes to the list in `stopWords.extend(['the', 'words', 'you', 'want', 'to', 'add'])`.
# NLTK Stop words
if nltkStop is True:
stopWords.extend(stopwords.words(stopLang))
stopWords.extend(['would', 'said', 'says', 'also', 'good', 'lord', 'come', 'let', 'say', 'speak', 'know', 'hamlet'])
# #### Add own stopword list
#
# Here is where your own stopwords list is added if you selected `True` in `customStop` above. Here you will need to change the folder names and file name to match your folders and file. Remember to put each folder name in quotes and in the correct path order, always putting the file name including the file extension ('.txt') last.
if customStop is True:
stopWordsFilepath = os.path.join(homePath, "Text-Analysis-master", "data", "earlyModernStopword.txt")
with open(stopWordsFilepath, "r",encoding = encoding) as stopfile:
stopWordsCustom = [x.strip() for x in stopfile.readlines()]
stopWords.extend(stopWordsCustom)
# ### Functions
# We need to create a few functions in order to calculate and create a bargraph. Any time you see `def` that means we are *DE*claring a *F*unction. The `def` is usually followed by the name of the function being created and then in parentheses are the parameters the function requires. After the parentheses is a colon, which closes the declaration, then a bunch of code below which is indented. The indented code is the program statement or statements to be executed. Once you have created your function all you need to do in order to run it is call the function by name and make sure you have included all the required parameters in the parentheses. This allows you to do what the function does without having to write out all the code in the function every time you wish to perform that task.
# #### Text Cleaning
# The first function does some text cleaning for us and requires the parameter: sentences (as in the sentences that make up your corpus)
#
# We are using the gensim package's `simple_preprocess` function which lowercases and removes punctuation from our text. We use a for loop to iteratively apply the function to the whole corpus.
def sentToWords(sentences):
for sentence in sentences:
yield(gensim.utils.simple_preprocess(str(sentence), deacc=True)) # deacc=True removes punctuations
# #### Get Ngrams
#
# This code will most likely not need to be adjusted. It creates a model of bigrams and trigrams in your dataset that occur frequently and then connects them with an underscore so the word counting algorithm will later consider them as one word. This is a good idea for items like 'new york' or 'new zealand' or 'Ho Chi Minh'. If we do not combine these frequently occuring phrases then 'new' and 'york' will be considered independently and give us less accurate results.
#
# However, you may wish to adjust two settings depending on your data. The first is `min_count` and we assign a number to a variable `minCount` that will be used for `min_count` in the code. The other setting you might want to adjust is `threshold` and for this we assign a number to a variable `tHold` that will be used for `threshold` in the code.
#
# Right now we have a `min_count` of 5 and a `threshold` of 100. The `min_count` is simply the minimum number of times the bigram or trigram needs to occur in order to be combined with an underscore. The `threshold` is a score that the bigram or trigram needs to exceed in order to be combined with an underscore. The score is determined by using this formula: (bigram_count - min_count)\*vocab_count/(wordA_count \* wordB_count). So let's say we have the bigram "good_lord" and it appears 30 times in a text of 10,000 words where "good" appears 60 times total and "lord" appears 40. With our `min_count` set to 5 we get the following: (30 - 5)\*10000/(60 \* 40) = 104.167 which means since our `threshold` is set to 100 "good_lord" will be combined with an underscore and made into a bigram. If the resulting score is above your `threshold` then the ngram is considered important enough to combine with an underscore and will be viewed as one word for the word counting later. Therefore, if you increase the `threshold`, you will get fewer bigrams and trigrams. If our threshold was set to 110, then "good" and "lord" would not be combined into "good_lord".
#
# The Phraser function takes the model you built with the Phrases function and cuts down memory consumption of Phrases, by discarding model state not strictly needed for the bigram detection task.
# +
# Variables
minCount = 5
tHold = 100
def get2gramPhrases(tokens):
# Build the bigram and trigram models
bigram = Phrases(tokens, min_count=minCount, threshold=tHold) # higher threshold fewer phrases.
# Removes model state from Phrases thereby reducing memory use.
bigramMod = Phraser(bigram)
return bigramMod
def get3gramPhrases(tokens):
bigram = Phrases(tokens, min_count=minCount, threshold=tHold) # higher threshold fewer phrases.
trigram = Phrases(bigram[tokens], threshold=tHold)
trigramMod = Phraser(trigram)
return trigramMod
# -
# Below are functions we are creating that perform certain tasks. First we are creating a function to remove the stopwords that are in our stopword list we created previously. Then we create functions to apply our bigram and trigram code from above.
# +
# Define functions for stopwords, bigrams, trigrams and lemmatization
def removeStopwords(texts):
return [[word for word in simple_preprocess(str(doc)) if word not in stopWords] for doc in texts]
def makeBigrams(tokens):
bigrams = get2gramPhrases(tokens)
return [bigrams[doc] for doc in tokens]
def makeTrigrams(tokens):
bigrams = get2gramPhrases(tokens)
trigrams = get3gramPhrases(tokens)
return [trigrams[bigrams[doc]] for doc in tokens]
# -
# Next we create a function that will lemmatize words if you set `spacyLem` equal to `True` above. If you set it equal to `False` then the function will simply find commonly occuring ngrams and add them to the token list.
# Form ngrams
def makeLemma(tokens):
dataWordsNgrams = makeTrigrams(tokens)
if spacyLem is True:
def lemmatization(tokens):
"""https://spacy.io/api/annotation"""
textsOut = []
for sent in tokens:
doc = nlp(" ".join(sent))
textsOut.append([token.lemma_ for token in doc if token.lemma_ != '-PRON-'])
return textsOut
# Initialize spacy language model, eliminating the parser and ner components
nlp = spacy.load(lemLang, disable=['parser', 'ner'])
# Do lemmatization
dataLemmatized = lemmatization(dataWordsNgrams)
return dataLemmatized
else:
return dataWordsNgrams
# #### Frequency count
#
# Now we need a function to count all the words and put them in order from most frequent to least frequent. The first four lines say that we are creating a function that creates a dictionary that adds '1' next to a word in that dictionary every time that word is used. The last line says that we are sorting that dictionary from the highest number to the lowest number.
def getFreq(tokens):
freq = defaultdict(int)
for t in tokens:
freq[t] += 1
# sorted frequency in descending order
return sorted(freq.items(), key = operator.itemgetter(1), reverse = True)
# #### Plot Graph Function
#
# This next function describes how we want to plot or visualize our results. We name this function `plotTopTen` and it begins by pulling only the top words from our sorted frequency above. You will be able to adjust the number of plotted words further down in another cell. After you choose how many words to plot, we create a data frame called `df`. Next we determine what percent each word makes up of the total text (not including stopwords) and assign it to a newly created column labeled "Pct". Then we assign the first "n" rows of our data frame to the the variable `dfPct`.
#
# Next we assign numbers to the variables `high` and `low` which will be used to determine the highest and lowest points on the y axis of the bar graph. Then it says we want a barplot (`fig = px.bar`) and the rest describes different aspects of the barplot, such as what we want for the x and y axis, the centering of the graph, the colors of the bars, the labels and titles of the graph, and how to save a html file of the visualization so we can use it in future presentations or publications. We will make changes to some of these later in the code.
def plotTopTen(sortedFreq, title, imgFilepath):
topn = n
df = pd.DataFrame(sortedFreq, columns = ["Words", "Count"])
df["Pct"] = ((df["Count"]/df["Count"].sum())*100).round(3)
df["Pct"] = df["Pct"].astype(str) + "%"
dfPct = df[0 : topn]
high = max(df["Count"])
low = 0
fig = px.bar(dfPct, x = "Words", y = "Count",hover_data=[dfPct["Pct"]],text = "Count", color = "Words",
title = title, color_discrete_sequence=colors,
labels = {"Words":Xlabel,"Count":Ylabel,"Pct":Zlabel})
fig.update_layout(title={'y':0.90, 'x':0.5, 'xanchor': 'center', 'yanchor':'top'},
font={"color": labCol}, width = wide, height = tall, showlegend=False)
fig.update_xaxes(tickangle = angle)
fig.update_yaxes(range = [low,math.ceil(high + 0.1 * (high - low))])
py.offline.plot(fig, filename=imgFilepath, auto_open = False)
fig.show()
# #### Pull from a single file
#
# This function opens the file, reads it, then applies the cleaning functions we created above. If you assigned `False` to `spacyLem` then the functions will not lemmatize your corpus, but will still include commonly occuring bigrams and trigrams.
def getTokensFrom1File(textFilepath):
docs=[]
with open(textFilepath, "r", encoding = encoding, errors = errors) as f:
for line in f:
stripLine = line.strip()
if len(stripLine) == 0:
continue
docs.append(stripLine.split())
words = list(sentToWords(docs))
stop = removeStopwords(words)
lemma = makeLemma(stop)
tokens = [item for sublist in lemma for item in sublist]
return tokens
# #### Pull from a directory
#
# This function is similar to the one above that reads a single text, except this one reads every file in a directory and applies the text cleaning functions to all of them.
def getTokensFromManyFiles(corpusRoot):
docs = []
for root, subdirs, files in os.walk(corpusRoot):
for filename in files:
# skip hidden file
if filename.startswith('.'):
continue
textFilepath = os.path.join(root, filename)
with open(textFilepath, "r", encoding = encoding, errors = errors) as f:
docs.append(f.read().strip('\n').splitlines())
words = list(sentToWords(docs))
stop = removeStopwords(words)
lemma = makeLemma(stop)
tokens = [item for sublist in lemma for item in sublist]
return tokens
# #### Pull from multiple directories
# This function reads in every ".txt" file in multiple directories and applies the text cleaning functions to all of them.
def getTokensFromDirect(directRoot):
paths = []
docs = []
dataPath = os.path.join(directRoot)
for folder in sorted(os.listdir(dataPath)):
if not os.path.isdir(os.path.join(dataPath, folder)):
continue
for file in sorted(os.listdir(os.path.join(dataPath, folder))):
paths.append(((dataPath, folder, file)))
df = pd.DataFrame(paths, columns = ["Root", "Folder", "File"])
df["Paths"] = df["Root"].astype(str) +"/" + df["Folder"].astype(str) + "/" + df["File"].astype(str)
for path in df["Paths"]:
if not path.endswith(".txt"):
continue
with open(path, "r", encoding = encoding, errors = errors) as f:
docs.extend(f.read().strip().split())
words = list(sentToWords(docs))
stop = removeStopwords(words)
lemma = makeLemma(stop)
tokens = [item for sublist in lemma for item in sublist]
return tokens
# ### Plot Top Ten
#
# Here we have an 'if else' statement that differentiates between creating a bargraph from a single document, an entire directory, or multiple directories depending on if you set `corpusLevel` above equal to `"lines"`, `"files"` or `"direct"`.
#
# #### First set variables
# Before we run the cell below, make sure the variables in the beginning of the cell are set correctly. The `n` variable should have a number assigned to it that is the number of words you wish to plot in your graph.
#
# The `singleDocName` variable should contain the single file you wish to run if you are only interested in a single document. If you set `corpusLevel` above to `"files"` or `"direct"` then don't worry about the `singleDocName` variable.
#
# Next we name the output file produced by the code below. Right now the name is a generic `"topTenPlainText"` but you may wish to change it to something that matches your data.
#
# Then we set our file format. We assign this format to the variable `fmt`. At the moment the only supported format is ".html", however, plotly has an option to download a static ".png" file when you open the ".html" file in a browser.
#
# Now we choose the labels for our x and y axes and assign them to the variables `Xlabel` and `Ylabel` respectively. Then we choose the label for our "Pct" column which will appear when we hover over the bars and we assign this label to the variable `Zlabel`.
#
# Next we choose our figure size and assign it to the variables `wide` and `tall`.
#
# Then we determine at what angle we want the x tick labels to be. The angle can be anywhere from -360 to 360. We assign this number to the variable `angle`.
#
# Next, we need to give our graph a title. We assign the title to the variable `title`.
#
# Lastly, we need to choose some color(s) for our graph. First we choose the bar colors. This is currently set so each bar has it's own color. The variable `color`determines the color palette of the bar graph. To change the color palette you only need to change the name of the palette which is located after the last period in `px.colors.qualitative.XXXXX`. To find other color options you can go here: [https://plotly.com/python/discrete-color/](https://plotly.com/python/discrete-color/). Then we choose the color for all our text. We assign our chosen color to the variable `labCol`. This is just the name of a color in quotes.
#
# #### From a single text
#
# If you set `corpusLevel` above equal to `"lines"`, then the first part of the script in the box below uses the functions and file paths we have created to get word frequencies and plot a bar graph from a single '.txt' file.
#
# We start by pointing to the specific file. Then we break the file down into tokens uisng the single file function from above.
#
# Next we count word frequencies using our word frequency function we made earlier.
#
# Then we save the output file based on what we set in the `#Variables` section at the beginning of the cell.
#
# Then we implement the plotting function we created above to give us a bargraph of the most frequent thematic words in our chosen text.
#
# #### Bargraph from multiple files in a directory
#
# If you set `corpusLevel` equal to `"files"`, then the second part of the script in the box below after `elif corpusLevel == "files":` now uses the functions and file paths we have created above to get word frequencies and plot a bargraph from multiple '.txt' files in a directory.
#
# We start by breaking the documents in our folder down into tokens using the function for getting tokens from a directory of documents we made earlier.
#
# Then we get the word frequencies using our function for counting words from above.
#
# Next we say where we want the output file saved based on our setting in the `#Variables` section at the beginning of the cell.
#
# Then we implement the plotting function we created above to give us a bargraph of the most frequent thematic words in our chosen corpus.
#
# #### Bargraph from multiple files in a multiple directories
#
# If you set `corpusLevel` equal to `"direct"`, then the third part of the script in the box below after `elif corpusLevel == "direct":` now uses the functions and file paths we have created above to get word frequencies and plot a bargraph from multiple '.txt' files in multiple directories.
#
# We start by breaking the documents in our directories down into tokens using the function for getting tokens from multiple directories of documents we made earlier.
#
# Then we get the word frequencies using our function for counting words from above.
#
# Next we say where we want the output file saved based on our setting in the `#Variables` section at the beginning of the cell.
#
# Then we implement the plotting function we created above to give us a bargraph of the most frequent thematic words in our chosen corpus.
# +
n = 10
singleDocName = 'Hamlet.txt'
outputFile = "topTenPlainText"
fmt = '.html'
Xlabel = "Word"
Ylabel = "Count"
Zlabel = "Percent"
wide = 750
tall = 550
angle = -45
title = 'Top 10 Words, Hamlet'
colors = px.colors.qualitative.Dark24
labCol = "crimson"
if corpusLevel == "lines":
# Use case one, analyze top 10 most frequent words from a single text
textFilepath = os.path.join(corpusRoot, singleDocName)
# get tokens
tokens = getTokensFrom1File(textFilepath)
# get frequency
freq = getFreq(tokens)
imgFilepath = os.path.join(dataResults, outputFile + fmt)
plotTopTen(freq, title, imgFilepath)
elif corpusLevel == "files":
# Use case two, analyze top 10 most frequent words from a corpus root
tokens = getTokensFromManyFiles(corpusRoot)
# get frequency
freq = getFreq(tokens)
imgFilepath = os.path.join(dataResults, outputFile +fmt)
plotTopTen(freq, title, imgFilepath)
elif corpusLevel == "direct":
tokens = getTokensFromDirect(directRoot)
# get frequency
freq = getFreq(tokens)
imgFilepath = os.path.join(dataResults, outputFile +fmt)
plotTopTen(freq, title, imgFilepath)
else:
None
# -
# ## VOILA!
| 32,993 |
/extract features/04 Transfer Learning C VGG16 RF.ipynb | cfce8dfb957c0e04cce47b22a1154738571e92ef | [] | no_license | ma17852003/TensorFlow_Keras | https://github.com/ma17852003/TensorFlow_Keras | 1 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 124,067 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="aJGx5zoxjRfa"
# -*- coding: utf-8 -*-
#-*- coding: cp950 -*-
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from PIL import Image
from pathlib import Path
from typing import *
import torch
import torch.optim as optim
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai import *
from fastai.vision import *
from fastai.text import *
from fastai.callbacks import *
from fastai.core import *
import tensorflow as tf
import random as rn
np.random.seed(0)
rn.seed(0)
import keras
from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
from keras.applications.vgg16 import preprocess_input
# Importing sklearn libraries
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, accuracy_score
import pickle
from keras.preprocessing import image
from sklearn.model_selection import train_test_split
# Importing Keras libraries
from keras.utils import np_utils
from keras.models import Sequential
from keras.applications import VGG16
from keras.applications import imagenet_utils
from keras.callbacks import ModelCheckpoint
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.layers import Dense, Conv2D, MaxPooling2D
from keras.layers import Dropout, Flatten, GlobalAveragePooling2D
import warnings
warnings.filterwarnings('ignore')
import cv2
# + colab={} colab_type="code" id="7DlryhdFjRfd" outputId="b8387445-32d6-4fd5-fcbc-142b259f249c"
os.listdir()
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 4145, "status": "ok", "timestamp": 1590209387771, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="Ee82pcWgjRfg" outputId="72fa89fe-6ad6-4e32-e057-735d1771c98a"
#在colab下接入google drive
from google.colab import drive
drive.mount('/content/gdrive',force_remount = True)
base_dir = '/content/gdrive/My Drive'
#path = Path(base_dir +'/imgs4') #imgs
path = Path(base_dir +'/DLtry')
path.mkdir(parents=True,exist_ok=True)
os.chdir(path)
# + colab={} colab_type="code" id="67XDOYhejRfi"
#分割數據與設定參數
input_shape = (224, 224, 3)
batch_size = 128
epochs =12 # 100
num_classes = 2 #######
x = []
y = []
for f in os.listdir("sakura"):
x.append(image.img_to_array(image.load_img("sakura/"+f, target_size=input_shape[:2])))
y.append(0)
for f in os.listdir("cosmos"):
x.append(image.img_to_array(image.load_img("cosmos/"+f, target_size=input_shape[:2])))
y.append(1)
x = np.asarray(x)
x /= 255
y = np.asarray(y)
y = keras.utils.to_categorical(y, num_classes)
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.33, random_state= 3)
# + colab={} colab_type="code" id="yJMl3DrXjRfk"
#VGG16のモデル・パラメータを読み込みます。畳み込み層、プーリング層の出力を特徴量として使用するために、
#末尾の全結合層は除き、最後に平均値でプーリングします。
from keras.models import Sequential #必須適時呼叫***
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras.applications.vgg16 import VGG16
from keras.models import Model
#base_model = VGG16(weights='imagenet', include_top=False)
base_model = VGG16(weights='imagenet', include_top=False, pooling="avg")
#訓練データをVGG16に処理させたものを、新しい特徴量とします。
x_train_vgg16 = base_model.predict(x_train)
x_test_vgg16 = base_model.predict(x_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 369} colab_type="code" executionInfo={"elapsed": 635, "status": "error", "timestamp": 1590209537559, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="TEXbd1K6jRfm" outputId="05f56ade-c317-402b-c1f2-f85eaa708e99"
#線形SVMに訓練データを学習させます。
from sklearn.svm import SVC
clf = SVC(kernel="linear").fit(x_train_vgg16, y_train)
#精度
from sklearn.metrics import accuracy_score
print("acc = {}".format(accuracy_score(clf.predict(x_train_vgg16), y_train)))
print("val_acc = {}".format(accuracy_score(clf.predict(x_test_vgg16), y_test)))
# + colab={} colab_type="code" id="DuEKKM8ejRfo"
訓練データ、バリデーションデータともに、100%の精度を達成できています
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 1839, "status": "ok", "timestamp": 1590210050653, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="H3OjnPa5jRfv" outputId="5a21ece0-4a38-4eee-a475-475e19ded9c8"
#######################
#RF
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(max_depth=45,n_estimators=220,random_state=230).fit(x_train_vgg16, y_train)
#精度
from sklearn.metrics import accuracy_score
print("acc = {}".format(accuracy_score(clf.predict(x_train_vgg16), y_train)))
print("val_acc = {}".format(accuracy_score(clf.predict(x_test_vgg16), y_test)))
# + colab={"base_uri": "https://localhost:8080/", "height": 302} colab_type="code" executionInfo={"elapsed": 615, "status": "error", "timestamp": 1590210180192, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="gTv46Mzm50xf" outputId="cecb1c63-b7be-4c21-b7bf-88ce59c94bcc"
print(clf.predict([[0, 0]]))
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 649, "status": "ok", "timestamp": 1590210505671, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="FKYpn8Lb51Ek" outputId="567c0b53-b839-4ea9-efba-289961409734"
clf.feature_importances_
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 667, "status": "ok", "timestamp": 1590210628747, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="h0m_HAK87u2n" outputId="664c5278-7710-4bd5-8e2c-f058c9e614ee"
list(zip(x_train, clf.feature_importances_))
# + colab={"base_uri": "https://localhost:8080/", "height": 644} colab_type="code" executionInfo={"elapsed": 45318, "status": "ok", "timestamp": 1590210736211, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="5_cYaZBS8MT8" outputId="ae446687-7d0d-46c8-8066-04d4142073f5"
pd.DataFrame(list(zip(x_train, clf.feature_importances_))) #PCA,NMF
# + colab={"base_uri": "https://localhost:8080/", "height": 369} colab_type="code" executionInfo={"elapsed": 617, "status": "error", "timestamp": 1590209558667, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="EJBztP6YjRfy" outputId="18040b90-c15f-469b-da97-9ef2ae41fc1c"
##################################
#logistic
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(class_weight='balanced', multi_class="auto", max_iter=200, random_state=1).fit(x_train_vgg16, y_train)
#精度
from sklearn.metrics import accuracy_score
print("acc = {}".format(accuracy_score(clf.predict(x_train_vgg16), y_train)))
print("val_acc = {}".format(accuracy_score(clf.predict(x_test_vgg16), y_test)))
# + colab={"base_uri": "https://localhost:8080/", "height": 369} colab_type="code" executionInfo={"elapsed": 660, "status": "error", "timestamp": 1590209819464, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="WYtAB4vnjRf3" outputId="177981fe-bb4f-4000-a564-cdc0f67dda43"
#################
#GB
from sklearn.ensemble import GradientBoostingClassifier
clf = GradientBoostingClassifier(random_state=230).fit(x_train_vgg16, y_train)
#精度
from sklearn.metrics import accuracy_score
print("acc = {}".format(accuracy_score(clf.predict(x_train_vgg16), y_train)))
print("val_acc = {}".format(accuracy_score(clf.predict(x_test_vgg16), y_test)))
# + colab={} colab_type="code" id="hBrzfhmfjRf4"
from sklearn.tree import DecisionTreeClassifier ## decision tree
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from xgboost import XGBClassifier
import xgboost
# + colab={"base_uri": "https://localhost:8080/", "height": 369} colab_type="code" executionInfo={"elapsed": 633, "status": "error", "timestamp": 1590209990360, "user": {"displayName": "Mike Wu", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhpqrLyHPmWGWl0C8pNFquPQiDSPUKVzkeZTK_hzA=s64", "userId": "02480068086185773948"}, "user_tz": -480} id="wkD3Ifd6jRf6" outputId="75f20073-8573-46a8-85bf-1b701039d96b"
clf = XGBClassifier(random_state=230).fit(x_train_vgg16, y_train)
#精度
from sklearn.metrics import accuracy_score
print("acc = {}".format(accuracy_score(clf.predict(x_train_vgg16), y_train)))
print("val_acc = {}".format(accuracy_score(clf.predict(x_test_vgg16), y_test)))
def partition(arr,s,e):
# initialize pivot index to the left
pivotindex = s
# initialize pivot value to the value of the rightmost element
pivotvalue = arr[e]
# for all elements if their value is lower than pivot value
# put the value to the current pivotindex (left at beginning, and increment pivotindex)
# when this is done put the pivotvalue at the pivotindex position
# return that position so we can have the next iteration on left
# and right parts relative to the pivotindex.
for i in range (s,e):
if arr[i] < pivotvalue:
swap(arr,i,pivotindex)
pivotindex +=1
swap(arr,pivotindex,e)
return pivotindex
size = 500_000
arr = [random.random() for i in range(size)]
size = len(arr)
start_time = time.time()
quicksort(arr, 0, size-1 )
end_time = time.time()
duration = end_time - start_time
print(f"Total execution time: {duration:.1f} seconds")
# -
# call the quicksort with a list of sizes, build a list of durations, plot
# +
import random
import time
import matplotlib.pyplot as plt
def swap(arr,i,j):
### swaps i and j values in array a
t = arr[i]
arr[i] = arr[j]
arr[j] = t
def quicksort(arr, start, end):
if start >= end:
# done
return
# calculate pivot index with the partition function
index = partition(arr, start, end)
# call quicksort for both sides of the pivot index.
# left part
quicksort(arr,start,index - 1)
# right part
quicksort(arr,index + 1, end)
def partition(arr,s,e):
# initialize pivot index to the left
pivotindex = s
# initialize pivot value to the value of the rightmost element
pivotvalue = arr[e]
# for all elements if their value is lower than pivot value
# put the value to the current pivotindex (left at beginning, and increment pivotindex)
# when this is done put the pivotvalue at the pivotindex position
# return that position so we can have the next iteration on left
# and right parts relative to the pivotindex.
for i in range (s,e):
if arr[i] < pivotvalue:
swap(arr,i,pivotindex)
pivotindex +=1
swap(arr,pivotindex,e)
return pivotindex
def get_quicksort_time(n):
arr = [random.random() for i in range(n)]
start_time = time.time()
quicksort(arr,0,n-1)
end_time = time.time()
return end_time - start_time
times = []
sizes = [50000,100000,200000,300000,400000,500000,600000,700000]
for n in sizes:
times.append(get_quicksort_time(n))
print(times)
plt.plot(sizes,times,color="blue")
# -
# Finally, see what the Python included sort() function does
# +
import random
import time
import matplotlib.pyplot as plt
def get_python_sort_time(n):
arr = [random.random() for i in range(n)]
start_time = time.time()
arr.sort()
end_time = time.time()
return end_time - start_time
times = []
sizes = [50000,100000,200000,300000,400000,500000,600000,700000]
for n in sizes:
times.append(get_quicksort_time(n))
print(times)
plt.plot(sizes,times,color="blue")
# -
# ## C-2 Decimals of $\pi$
# ## C-2.1 Decimals of $\pi$ with Monte-Carlo simulation
# +
import time
from random import random
start_time = time.time()
#---> calculation starts here
iterations = 5_000_000 # change this for accuracy vs duration
hit = 0 # counter of random hits with module < 1
for i in range(iterations):
if random()**2 + random()**2 < 1: # random(a,b) returns a pseudo random float
hit += 1 # between a & b. Default : between 0 and 1
#---> calulation ends here
end_time = time.time()
duration = end_time - start_time
print(f"Total execution time: {duration:.1f} seconds")
print(f"estimation of PI = {4 * hit / iterations}")
# -
# ## C-2.2 Decimals of $\pi$ with better maths
# ### C-2.2.1 Viète formula 1593 (after [François Viète](https://en.wikipedia.org/wiki/Fran%C3%A7ois_Vi%C3%A8te), 1540-1603):
#
# \begin{equation}
# \pi = 2\times\frac{2}{\sqrt{2}} \times \frac{2}{\sqrt{2 + \sqrt{2}}} \times \frac{2}{\sqrt{2 + \sqrt{2 + \sqrt{2}}}} \times ... \times \frac{2}{\sqrt{2+\sqrt{2+\sqrt{2+\sqrt{2 + ...}}}}}
# \end{equation}
# +
# -
# ### C-2.2.2 Wallis formula 1656 (after [John Wallis](https://en.wikipedia.org/wiki/John_Wallis), 1616-1703)
#
# \begin{equation}
# {\displaystyle {\pi }=2 \times \prod _{n=1}^{\infty }{\frac {4n^{2}}{4n^{2}-1}}}
# \end{equation}
# +
# -
# ## C-2.2.3 Using Euler formula and Huffini-Horner to calculate decimals of $\pi$
#
# ### Part I: From Euler development of Arctan
# \begin{equation}
# \forall x\in [-1,1] \quad \arctan(x)=\sum _{n=0}^{\infty }{\frac {2^{2n}(n!)^{2}}{(2n+1)!}}{\frac {x^{2n+1}}{(1+x^{2})^{n+1}}}
# \end{equation}
#
#
# With $ \frac{\pi}{4} = \arctan(1)$, we can plug x=1 in the Euler development :
#
# \begin{equation}
# \frac{\pi}{4} = \sum _{n=0}^{\infty }{\frac {2^{2n}(n!)^{2}}{(2n+1)!}}{\frac {1^{2n+1}}{(1+1^{2})^{n+1}}}
# \end{equation}
#
# With $1+1^2=1$ and $1^k = 1$, we can also reduce de power of 2 up and down to get:
#
# \begin{equation}
# \frac{\pi}{4} = \sum _{n=0}^{\infty }{\frac {2^{2n}(n!)^{2}}{(2n+1)!}}{\frac {1}{2^{n+1}}} = \frac{1}{2}\sum _{n=0}^{\infty }{\frac {2^n(n!)^{2}}{(2n+1)!}}
# \end{equation}
#
# Multiply left and right by 4, and moving out the term for n=0 :
#
# \begin{equation}
# \pi = 2\sum _{n=0}^{\infty }{\frac {2^n(n!)^{2}}{(2n+1)!}} = 2 + 2 \sum _{n=1}^{\infty }{\frac {2^n(n!)^{2}}{(2n+1)!}}
# \end{equation}
#
#
# note that:
#
# \begin{equation}
# \frac{2^n(n!)^2}{(2n+1)!} = \frac{n!\times (2^n \times n!)}{(2n+1)!} = n!\times\frac{(2\times1)\times(2\times2)\times(2\times3)\times(2\times4)\times(2\times5)\times ... \times(2\times n)}{1\times2\times3\times4\times5 ... \times(2n+1)}
# \end{equation}
#
# which simplifies into :
#
# \begin{equation}
# n!\times\frac{1}{1\times3\times5\times7 ... \times(2n+1)} = \frac{1\times2\times3\times4 ... \times n}{1\times3\times5\times7 ... \times(2n+1)}
# \end{equation}
#
# Now we have :
#
# \begin{equation}
# \pi = 2 + 2 \sum _{n=1}^{\infty }{\frac{1\times2\times3\times4 ... \times n}{1\times3\times5\times7 ... \times(2n+1)} }
# \end{equation}
#
#
# At this point we can calculate an estimation for $\pi\$ based on Euler's formula:
# +
# -
# ## Calcul des décimales de $\pi$
#
# ### Algorithme de Ruffini-Horner
#
# On a:
#
# \begin{equation}
# \pi = 2 + 2 \sum _{n=1}^{\infty }{\frac{1\times2\times3\times4 ... \times n}{1\times3\times5\times7 ... \times(2n+1)} }
# \end{equation}
#
# C'est à dire (en écrivant les termes de la sommation) :
#
#
# \begin{equation}
# \pi = 2 + 2\frac{1}{3} + 2\frac{2}{3\times5} + 2\frac{2\times3}{3\times5\times7} + 2\frac{2\times3\times4}{3\times5\times7\times9} + 2\frac{2\times3\times4 ... \times n}{3\times5\times7\times9 ... \times(2n+1)}
# \end{equation}
#
# Que l'on peut factoriser:
#
# \begin{equation}
# \pi = 2 + \frac{1}{3}\bigl(2 + \frac{2}{5}\bigl(2 + \frac{3}{7}\bigl( 2 + \frac{4}{9}\bigl(2 + \frac{5}{11}\bigl(2+ ... + \frac{n}{2n+1}(2) \bigr)\bigr)\bigr)\bigr)\bigr)
# \end{equation}
#
#
# Et donc : $\pi$ peut être écrit $22222....22222$ dans la base à pas variable $\big[1;\frac{1}{3};\frac{2}{5};\frac{3}{7};\frac{4}{9};....; \frac{n}{2n+1}\bigr];$
#
# +
# Source : José Ouin - Algorithmique programmation par la pratique. Ed Ellipses
entree = input("entrer le nombre de décimales de 𝝅 à calculer :")
decimales = int(entree)
N = 4 * decimales
DCMPI = [0] * (decimales+2)
P = [0] * (N + 1)
Q = [0] * (N + 1)
RTN = [0] * (N + 1)
SMM = [0] * (N + 1)
RST = [2] * (N + 1)
for i in range(1, N+1):
P[i] = i
Q[0] = 10
for i in range(1,N+1):
Q[i] = 2*i + 1
for j in range(0,decimales+2):
VINI = [10 * u for u in RST]
RTN[N] = 0
for k in range(N,-1,-1):
SMM[k] = VINI[k] + RTN[k]
RST[k] = SMM[k]%Q[k]
if k !=0:
RTN[k-1] = P[k]*(SMM[k] - RST[k])/Q[k]
else:
DCMPI[j] = (SMM[0]-RST[0])/Q[0]
if DCMPI[j] >= 10:
TMP = DCMPI[j]%10
DCMPI[j-1] = DCMPI[j-1] + (TMP - DCMPI[j])/10
DCMPI = [int(u) for u in DCMPI]
DCMPI = DCMPI[:-1]
print("Décimales de 𝝅:",DCMPI)
| 18,086 |
/lab_2_stable_baselines.ipynb | 72d9e817fc4291264da9da38bccb5f8fd384231a | [
"MIT"
] | permissive | jbarsce/AprendizajePorRefuerzos | https://github.com/jbarsce/AprendizajePorRefuerzos | 0 | 0 | MIT | 2018-10-09T18:59:19 | 2018-10-09T18:22:07 | null | Jupyter Notebook | false | false | .py | 60,130 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: diplodatos_rl
# language: python
# name: diplodatos_rl
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/DiploDatos/AprendizajePorRefuerzos/blob/master/lab_2_stable_baselines.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="SytTR8K1oII-"
# # Introducción
# + [markdown] id="8BFrF74foIJD"
# Créditos:
#
# * Documentación y repo de Stable-baselines https://stable-baselines3.readthedocs.io.
# * Tutorial sobre SB3: https://github.com/araffin/rl-tutorial-jnrr19.
# * Documentación y repo de OpenAI Gym https://github.com/openai/gym/blob/master/docs/.
# * Crear un entorno https://github.com/openai/gym/blob/master/docs/creating-environments.md.
# + [markdown] id="HlZ0ehGZoIJD"
# Stable-baselines3: framework de deep RL que provee interfaces para ejecutar y adaptar algoritmos de RL "al estilo scikit-learn". Permite utilizar agentes abstrayéndonos de los detalles de bajo nivel de abstracción referentes a la implementación del algoritmo$^1$
#
# Además, ofrece herramientas muy útiles como
#
# * Monitores que permiten ver el rendimiento del agente según se desempeña en el entorno, sin tener que esperar a que finalice de entrenar.
# * Callbacks que permiten accionar eventos cuando se cumplen algunas condiciones en el entrenamiento de nuestro agente (por ejemplo, detenerlo si la recompensa recibida es menor a cierto umbral tras un cierto período de tiempo).
#
#
# Documentación https://stable-baselines3.readthedocs.io
#
# Es un fork activamente mantenido de [OpenAI baselines](https://github.com/openai/baselines)
#
# La versión 3 cambia el framework subyacente de Tensorflow a Pytorch y está activamente en desarrollo; no obstante la versión 2 es completamente funcional
#
# $^1$ no obstante, al igual que sucede generalmente con librerías de ML:
#
# * Siempre es bueno tener en mente las características, ventajas y desventajas del algoritmo utilizado, pues de eso depende mucho la convergencia de nuestra solución, especialmente cuando se emplean entornos adaptados para nuestras necesidades.
#
# * Esta librería, al igual que demás frameworks generales de RL, están muy probadas en entornos estándares de RL como Atari o PyBullet. No obstante, es posible que nuestro entorno o nuestras necesidades difieran significativamente, lo que hace que en algunos casos haya que meter mano directo en el código de los algoritmos/librería.
# + [markdown] id="W4SWX4wuoIJE"
# # Interfaz básica stable-baselines
# + [markdown] id="SD-FSSuKoIJF"
# ### Instalación de Stable-baselines
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/"} id="ttqOLol_oIJG" outputId="e7b26d16-a6d3-45af-ca5f-332a2330427a"
#@title Instalación (no modificar)
# !pip install stable-baselines3[extra,tests,docs]
# -
# Desde Windows, además, instalar:
# * Microsoft Visual C++ desde https://visualstudio.microsoft.com/visual-cpp-build-tools/
# * PyType, mediante `conda install -c conda-forge pytype`
# + [markdown] id="uUUkScRxoIJQ"
# ### Instalación de RLBaselinesZoo
# + [markdown] id="hiKPJUfdoIJQ"
# Desde Google Colab
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="ZM_pM0mIoIJQ" outputId="8b3aabec-28d0-4caf-93e7-4f4d6cd8725f"
#@title Instalación de RLBaselinesZoo (no modificar)
# Estamos en Colab?
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
# !git clone --recursive https://github.com/DLR-RM/rl-baselines3-zoo
# !cd rl-baselines3-zoo/
# !apt-get install swig cmake ffmpeg
# !pip install -r /content/rl-baselines3-zoo/requirements.txt
# + [markdown] id="SzBEoyPzoIJR"
# Desde Linux, ejecutando
#
# git clone --recursive https://github.com/DLR-RM/rl-baselines3-zoo
# cd rl-baselines3-zoo/
# conda install swig
# pip install -r requirements.txt
# + [markdown] id="GPUpOonyoIJH"
# ## Ejecución de un algoritmo de RL
# + [markdown] id="Morjd1dFoIJH"
# ### Importaciones/inicializaciones
# + id="OG7i44kqoIJH" jupyter={"source_hidden": true}
import os
from subprocess import Popen, PIPE
import numpy as np
import matplotlib.pyplot as plt
import gym
from gym import spaces
#from gym.envs.registration import register
from stable_baselines3 import DQN, PPO
from stable_baselines3.common.monitor import Monitor
from stable_baselines3.common.vec_env import DummyVecEnv
from stable_baselines3.common.callbacks import EvalCallback, StopTrainingOnRewardThreshold
from stable_baselines3.common.env_util import make_vec_env
os.makedirs('logs', exist_ok=True)
cwd = os.getcwd()
# %matplotlib inline
# %load_ext tensorboard
# + [markdown] id="is2ytf-coIJH"
# ### Ejemplo básico
# + colab={"base_uri": "https://localhost:8080/"} id="h8SqFh7noIJI" outputId="1d56db34-53bd-450a-8fe8-825f7201ffcf"
env = gym.make('CartPole-v1')
# MlpPolicy es una política "estándar" que aprende con perceptron multicapa
# (es decir sin capas convolucionales o demás variantes),
# 2 capas ocultas con 64 neuronas cada una
model = DQN('MlpPolicy', env)
model.learn(total_timesteps=10000)
# + [markdown] id="0f5JFC7AoIJK"
# ### Renderización
# + id="5nIL0TuwoIJK"
if not IN_COLAB:
obs = env.reset()
for i in range(1000):
action, _states = model.predict(obs, deterministic=True)
obs, reward, done, info = env.step(action)
env.render()
if done:
obs = env.reset()
env.close()
# + [markdown] id="iTtXPFd2oIJK"
# ### Logging
# + [markdown] id="BFYyMrvboIJK"
# #### Ver rendimiento del agente en tensorboard
# + colab={"base_uri": "https://localhost:8080/"} id="pC1hqla2oIJK" outputId="77c1b83c-9952-4ee7-b4eb-2c8bfa0d2024"
venv = make_vec_env(lambda: gym.make('CartPole-v1'), n_envs=1)
model = DQN('MlpPolicy', venv, tensorboard_log='tensorboard/')
model.learn(total_timesteps=100000)
# + [markdown] id="rldIL1u4oIJL"
# Para verlo en tensorboard, correr
# + [markdown] id="v4Db-mCxoIJL"
# `tensorboard --logdir=tensorboard/`
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="2AfqEmCWxWFv" outputId="e2297f30-400f-4480-b04e-90f1b4dc2549"
# %tensorboard --logdir=tensorboard
# + [markdown] id="2u0_1dk5oIJL"
# ### Monitor
#
# Vamos a crear un monitor para loguear nuestro agente en la carpeta logs. Nuestro monitor guardará datos de recompensa (r), duración (l) y tiempo total (t)
# + colab={"base_uri": "https://localhost:8080/"} id="r7uF7sQdoIJL" outputId="ea0fb222-58aa-4c18-9c8c-ae97132ee3ee"
env = gym.make('CartPole-v1')
env = Monitor(env, 'logs/') # reemplazamos env por su monitor
model = DQN('MlpPolicy', env, )
model.learn(total_timesteps=10000)
# + [markdown] id="4yVVSGjvoIJM"
# ### Callbacks
# + colab={"base_uri": "https://localhost:8080/"} id="47XdjvaSoIJM" outputId="8eb9dd8d-ca66-4aee-cdbf-bce8a510ecea"
env = gym.make('CartPole-v1')
callbacks = [] # lista de callbacks a usar, pueden ser varios
# callback para detener entrenamiento al alcanzar recompensa de 9.8
# (es una recompensa muy baja, pero la establecemos a fines demostrativos)
stop_training_callback = StopTrainingOnRewardThreshold(reward_threshold=9.8)
# al crear EvalCallback, se asocia el mismo con stop_training_callback
callbacks.append(EvalCallback(Monitor(env, 'logs/'),
eval_freq=1000,
callback_on_new_best=stop_training_callback))
# la semilla aleatoria hace que las ejecuciones sean determinísticas
model = DQN('MlpPolicy', env, seed=42)
model.learn(total_timesteps=10000, callback=callbacks)
# + [markdown] id="QCPD7oaNoIJM"
# ### Ejecutar agente RL en múltiples ambientes
# + [markdown] id="PNiTT2AzoIJM"
# Esta librería provee una interfaz para ejecutar agentes en varias instancias de un mismo entorno a la vez (*vectorized environments*), de modo tal que se habilite la ejecución paralela y de otras funcionalidades útiles.
#
# Para ello, varios de sus algoritmos implementan cambios que consideren la posibilidad de que haya múltiples entornos subyacentes, por ejemplo `step(accion)` cambia a `step(lista_acciones)`, aplicando acciones a todos los entornos, recibiendo ahora múltiples observaciones y recompensas.
#
# Otro cambio: se aplica `reset()` automáticamente a cada entorno que llega a un estado final.
# + [markdown] id="9GGZjjcDoIJN"
# SB brinda dos formas de utilizar entornos vectorizados:
#
# * **DummyVecEnv**, el cuál consiste en un *wrapper* de varios entornos, los cuáles funcionarán en un sólo hilo. Este wrapper es útil como entrada de algoritmos que requieren los entornos de esta forma, y habilita los procesamientos y operaciones comunes de los entornos vectorizados.
# * **SubprocVecEnv**, el cuál paraleliza multiples entornos pero en procesos de ejecucíon separados. Cada proceso tiene su propia memoria y puede adquirir derechos sobre las CPUs de la computadora donde se ejecuta. Se utiliza cuando el entorno del agente es computacionalemente complejo. Atención! **Puede comer mucha RAM**.
#
# Vemos un ejemplo:
# + colab={"base_uri": "https://localhost:8080/"} id="ovc6JDmYoIJN" outputId="6f36d2d0-bc4e-4299-b612-58103bc73f6e"
# ejemplo de ambiente dummy
venv = DummyVecEnv([lambda: gym.make('CartPole-v1')]*4)
model = PPO('MlpPolicy', venv, )
model.learn(total_timesteps=10000)
# + [markdown] id="kkadM0S1oIJO"
# También puede hacerse con un una función de SB a tal efecto
# + colab={"base_uri": "https://localhost:8080/"} id="wez-ZY_IoIJO" outputId="bc30a171-8081-4b97-ba76-063551b2a200"
venv = make_vec_env(lambda: env, n_envs=4)
model = PPO('MlpPolicy', venv, )
model.learn(total_timesteps=10000)
# + [markdown] id="BjH4ZQ4NoIJO"
# ### Ejecutar agente con políticas personalizadas
# + colab={"base_uri": "https://localhost:8080/"} id="jp_aPhIdoIJO" outputId="19b3f2a7-2821-4fe9-a749-b32be258aee8"
# Creamos una clase con una red neuronal de 128x128 neuronas
model = PPO('MlpPolicy', policy_kwargs=dict(net_arch=[128,128]), env='CartPole-v1', verbose=1)
model.learn(total_timesteps=10000)
# -
# [Buen post](https://medium.com/aureliantactics/understanding-ppo-plots-in-tensorboard-cbc3199b9ba2) en donde se explica qué significan varias de estas métricas. Para verlas en detalle podemos consultar directamente el [código fuente](https://github.com/DLR-RM/stable-baselines3/blob/master/stable_baselines3/ppo/ppo.py), que está bien documentado y no es muy difícil de seguir.
# + [markdown] id="WbwkBi87oIJO"
# ### Utilizar un entorno personalizado
#
# Antes que nada, además de la interfaz que ya vimos de Gym, hay otras nociones que tenemos que tener en cuenta en este contexto:
#
# * Los entornos definen un espacio de estados y de acciones, a partir de los cuáles los modelos asumen y respetan la "forma" de observaciones y acciones. Por ejemplo, algunos algoritmos están diseñados para espacios de acciones discretos (DQN), continuos (DDPG) o bien poseen implementaciones particulares pueden usarse en ambos (PPO, en el repo de SB3). En cuanto a los espacios, algunos algoritmos asumen explícitamente un espacio discreto (y pequeño), como Q-Learning, mientras que otros como PPO asumen cualquier tipo de espacio.
# * Los dos tipos más comunes de estados o acciones son los espacios discretos `gym.spaces.Discrete` y los continuos `gym.spaces.Box`.
# * Los espacios discretos definen un conjunto de $n$ estados/acciones $\{ 0, 1, \dots, n-1 \}$, mientras que los espacios continuos definen un espacio $\mathbb{R}^d$, de una de las siguientes 4 formas: $[a, b], (-\infty, b], [a, \infty), (-\infty, \infty)$, en donde $a,b$ son las cotas superior e inferior (de existir).
# * Ejemplos: un espacio de acciones `Discrete(4)` tiene 4 acciones: $\{0,1,2,3\}$; un espacio de estados `Discrete(16)` tiene 16 estados. Un espacio de estados ALTURA, ANCHO, N_CANALES que represente una imagen RGB acotada en $[a=0, b=255]$ se puede crear como
#
# `observation_space = spaces.Box(low=0, high=255, shape=(HEIGHT, WIDTH, N_CHANNELS), dtype=np.uint8)`
# + [markdown] id="YFLZieLNoIJO"
# Para usar un entorno compatible por esta librería, el mismo tiene que heredar de *gym.Env*. Vemos un ejemplo (crédito: Antonin Raffin https://colab.research.google.com/github/araffin/rl-tutorial-jnrr19/blob/sb3/5_custom_gym_env.ipynb)
# + id="str0v6DUoIJO"
class GoLeftEnv(gym.Env):
"""
Ambiente personalizado que sigue la interfaz de gym.
Es un entorno simple en el cuál el agente debe aprender a ir siempre
hacia la izquierda.
"""
# Dado que podemos estar en colab, no podemos implementar la salida por interfaz
# gráfica ('human' render mode)
metadata = {'render.modes': ['console']}
# Definimos las constantes
LEFT = 0
RIGHT = 1
def __init__(self, grid_size=10):
super(GoLeftEnv, self).__init__()
# Tamaño de la grilla de 1D
self.grid_size = grid_size
# Inicializamos en agente a la derecha de la grilla
self.agent_pos = grid_size - 1
# Definimos el espacio de acción y observaciones
# Los mismos deben ser objetos gym.spaces
# En este ejemplo usamos dos acciones discretas: izquierda y derecha
n_actions = 2
self.action_space = spaces.Discrete(n_actions)
# La observación será la coordenada donde se encuentra el agente
# puede ser descrita tanto por los espacios Discrete como Box
self.observation_space = spaces.Box(low=0, high=self.grid_size,
shape=(1,), dtype=np.float32)
def reset(self):
"""
Importante: la observación devuelta debe ser un array de numpy
:return: (np.array)
"""
# Se inicializa el agente a la derecha de la grilla
self.agent_pos = self.grid_size - 1
# convertimos con astype a float32 (numpy) para hacer más general el agente
# (en caso de que querramos usar acciones continuas)
return np.array([self.agent_pos]).astype(np.float32)
def step(self, action):
if action == self.LEFT:
self.agent_pos -= 1
elif action == self.RIGHT:
self.agent_pos += 1
else:
raise ValueError("Se recibió una acción inválida={} que no es parte del espacio de acciones".format(action))
# Evitamos que el agente se salga de los límites de la grilla
self.agent_pos = np.clip(self.agent_pos, 0, self.grid_size)
# Llegó el agente a su estado objetivo (izquierda) de la grilla?
done = bool(self.agent_pos == 0)
# Asignamos recompensa sólo cuando el agente llega a su objetivo
# (recompensa = 0 en todos los demás estados)
reward = 1 if self.agent_pos == 0 else 0
# gym también nos permite devolver información adicional, ej. en Atari:
# las vidas restantes del agente (no usaremos esto por ahora)
info = {}
return np.array([self.agent_pos]).astype(np.float32), reward, done, info
def render(self, mode='console'):
if mode != 'console':
raise NotImplementedError()
# en nuestra interfaz de consola, representamos el agente como una cruz, y
# el resto como un punto
print("." * self.agent_pos, end="")
print("x", end="")
print("." * (self.grid_size - self.agent_pos))
def close(self):
pass
# + colab={"base_uri": "https://localhost:8080/"} id="flB9-j1SoIJP" outputId="67ccb533-c91c-441a-8f85-72346c10e1ac"
env = GoLeftEnv(grid_size=10)
env = make_vec_env(lambda: env, n_envs=1)
model = PPO('MlpPolicy', env, verbose=1).learn(20000)
# -
# Ver agente entrenado
obs = env.reset()
for i in range(10):
action, _states = model.predict(obs)
obs, rewards, dones, info = env.step(action)
env.render(mode='console')
if dones[0]:
obs = env.reset()
# + [markdown] id="sgrWKJlsoIJT"
# ## Normalización de features y recompensas
# + [markdown] id="BIfn3o99oIJT"
# Stable-Baselines3 tiene predefinidos un conjunto de **wrappers** genericos que pueden utilizarse para preprocesar las observaciones que llegan al agente RL, desacoplando del mismo el prepocesamiento.
#
# Entre las funcionalidades disponibles tenemos:
# * **VecFrameStack**: Se utiliza cuando la observación que percibe el agente es una imagen. Sirve para expandir el espacio de estados apilando N frames de manera conjunta (ver [paper de DQN](https://arxiv.org/abs/1312.5602) para más detalle).
# * **VecNormalize**: Se utiliza para normalizar las observaciónes y/o las recompenzas que percibe el agente, a $\mu=0$ y $\sigma=1$. También permite cortar valores de observaciones y/o recompensas que excedan un rango establecido.
# * **VecCheckNan**: Se utiliza para trackear los estados del entorno que generan que los gradientes de la RNN se hagan NaN.
# * **VecVideoRecorder**: Se utiliza para exportar el funcionamiento de la política aprendida por el agente a un video (MP4).
#
# Ademas, se pueden crear **wrappers** personalizados extendiendo la clase **VecEnvWrapper**:
#
# ```
# class MiWrapper(VecEnvWrapper):
# [...]
# ```
# ### Ejemplo (VecNormalize)
# + colab={"base_uri": "https://localhost:8080/"} id="3VOh1HQCVJNA" outputId="b9f70740-a806-4978-988d-aa296af53c58"
from stable_baselines3.common.vec_env import DummyVecEnv, VecNormalize
env = gym.make('CartPole-v1')
env = DummyVecEnv([lambda: env]) # Multiples entornos vectorizados
# Normalizamos tanto observaciones como recompensas, y recortamos ambas cuando exceden 10
env = VecNormalize(env, norm_obs=True, norm_reward=True, clip_obs=10.0, clip_reward=10.0)
model = DQN('MlpPolicy', env)
model.learn(total_timesteps=10000)
# + [markdown] id="exel4Gr8oIJQ"
# # RL-baselines3-zoo
#
# Colección de agentes RL y herramientas útiles para ejecutarlos, evaluarlos e incluso hacer videos con ellos. Los agentes de este repo están preparados con la configuración requerida para los distintos tipos de entornos, incluyendo Atari, PyBullet y entornos clásicos, incluyendo configuraciones e híper-parámetros que producen buenas políticas para tales entornos.
#
# Esta librería ofrece un muy buen punto de partida para utilizar agentes / entornos personalizados, ya que ofrece una [interfaz](https://github.com/DLR-RM/rl-baselines3-zoo/blob/master/train.py) fácilmente adaptable a nuestras necesidades.
#
# Si se usan entornos personalizados con rl-baselines3-zoo, debe tenerse en cuenta que se deben definir todos los híper-parámetros de antemano, sea al instanciar el agente, o en la carpeta */rl-baselines3-zoo/hyperparams*; de lo contrario arrojará error por no encontrar qué híper-parámetro usar.
# + [markdown] id="p6TCyu_UoIJR"
# ## Ejecución
#
# Los agentes pueden ser llamados desde la consola mediante comandos como
#
# `python train.py --algo algo_name --env env_id`
#
# Los cuales pueden ser llamados usando
# + id="jBLMVxhIoIJR"
os.chdir('rl-baselines3-zoo/')
args = [
'-n', str(100000),
'--algo', 'ppo',
'--env', 'CartPole-v1'
]
p = Popen(['python', 'train.py'] + args,
stdin=PIPE, stdout=PIPE, stderr=PIPE)
output, err = p.communicate()
rc = p.returncode
os.chdir(cwd)
assert rc == 0
# + [markdown] id="40H4hNcqoIJR"
# Ver en acción el agente entrenado (nota: no disponible en Google Colab, requiere ffmpeg)
# + id="cNija96loIJR"
if not IN_COLAB:
os.chdir('rl-baselines3-zoo/')
args = [
'--algo', 'ppo',
'--env', 'CartPole-v1',
'--folder', 'logs/'
]
p = Popen(['python', 'enjoy.py'] + args,
stdin=PIPE, stdout=PIPE, stderr=PIPE)
output, err = p.communicate()
rc = p.returncode
os.chdir(cwd)
assert rc == 0
# + [markdown] id="MSfIitK7oIJR"
# También es posible grabar un video! Ver https://stable-baselines3.readthedocs.io/en/master/guide/examples.html#record-a-video
# + [markdown] id="Phq975BYoIJR"
# Ver curva de aprendizaje obtenida por el agente desde *utils.plot*
# + id="Q8EA41oQoIJS"
os.chdir('rl-baselines3-zoo/')
args = [
'--algo', 'ppo',
'--env', 'CartPole-v1',
'--exp-folder', 'logs/'
]
p = Popen(['python', '-m', 'scripts.plot_train'] + args, stdout=PIPE)
output, err = p.communicate()
rc = p.returncode
os.chdir(cwd)
assert rc == 0
# + colab={"base_uri": "https://localhost:8080/"} id="__1mgFR0oIJT" outputId="2c355f37-24ae-4e78-ea4c-ac66d3003c23"
print(output)
# + [markdown] id="BWzAHj6qoIJT"
# ## Híper-parámetros
#
# rl-baselines-zoo provee híper-parámetros que resultan en curvas de aprendizaje que convergen en buena cantidad de entornos. Estos híper-parámetros pueden verse en cada uno de los archivos YAML de cada algoritmo, [acá](https://github.com/DLR-RM/rl-baselines3-zoo/tree/master/hyperparams).
#
# También provee funcionalidad para optimizar los híper-parámetros con la librería [Optuna]( https://github.com/optuna/optuna). En los mismos se incluyen rangos de híper-parámetros que se usaron para optimizar entornos como los de PyBullet, y son fácilmente modificables para adaptarlo a nuestros propios entornos. Para ver cómo se llama a la interfaz de Optuna ver [este código](https://github.com/DLR-RM/rl-baselines3-zoo/blob/master/utils/hyperparams_opt.py).
#
# Nota: **Optuna come muchos recursos!**
# -
# # Entrenando otros agentes
# ### Algunos conjuntos de entornos
#
# CartPole es una excelente línea base (de hecho suele ser la prueba preliminar de todo nuevo algoritmo), porque tiene recompensas constantes pero requiere cierta solidez por parte del algoritmo para hacerlo converger al óptimo.
#
# No obstante, al implementar un algoritmo, es deseable que el mismo pueda desenvolverse de forma consistente en varios grupos de entornos. A continuación va una lista con varios entornos que sirven como prueba:
#
# | Entornos | Estados | Acciones | Dificultad | Implementado por |
# |--------------------------------------------------------------------------------------------------------------------|--------------------|---------------------|-----------------|-----------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
# | Clásicos (CartPole, MountainCar, Pendulum, Deep sea (testea exploración), umbrella (testea asignación de crédito)) | Discretos/Continuos | Continuas/discretas | Baja/Media | [Gym](https://github.com/openai/gym/wiki/Table-of-environments) y [BSuite](https://github.com/deepmind/bsuite) |
# | Grilla (desde pequeñas donde hay que salir hasta grandes con muchas habitaciones y subproblemas) | Discretos/imágenes | Discretas | Baja/Media/Alta | [gym-minigrid](https://github.com/maximecb/gym-minigrid) |
# | Grilla/plataforma/estilo Atari, generados proceduralmente | Imágenes | Discretas | Media/Alta | [Gym](https://github.com/openai/procgen) |
# | Plataforma 2D, como LunarLander o BipedalWalker | Continuos | Continuas/discretas | Media/Alta | [Gym](https://github.com/openai/gym/wiki/Table-of-environments) |
# | Primera persona en 3D | Imágenes | Continuas/discretas | Media/Alta | [Deepmind](https://github.com/deepmind/lab) |
# | Simulación física de pequeños robots | Continuos | Continuas | Media/Alta | Gym (con el motor MuJoCo o su versión open-source, [PyBullet](https://docs.google.com/document/d/10sXEhzFRSnvFcl3XxNGhnD4N2SedqwdAvK3dsihxVUA/edit#heading=h.wz5to0x8kqmr)) |
# | Atari | Continuos | Discretas | Media/Alta | [Gym](https://github.com/openai/gym/wiki/Table-of-environments) (son todos aquellos entornos que terminan en "*-v4*" |
#
# ## Resumen de algunos algoritmos
#
# Se resumen ahora varios algoritmos del estado del arte de aprendizaje por refuerzos profundo
#
#
# | Algoritmo | Tipo | Espacio de acciones | Resumen rápido | Artículo |
# |-----------|------------|---------------------|---------------------------------------------------------------------------------------------------------------------------------------------------------------|----------------------------------|
# | DQN | Off-policy | Discretas | Extiende Q-Learning a deep learning. En Stable-baselines, DQN incluye todas las mejoras ya incorporadas. | https://arxiv.org/abs/1312.5602 |
# | ACER | Off-policy | Discretas | Combina una arquitectura actor-critic con un buffer y repetición de experiencia. | https://arxiv.org/abs/1611.01224 |
# | A3C | On-policy | Ambos | Múltiples agentes corriendo en múltiples instancias del ambiente, acumulando sus gradientes y actualizándolos tras un cierto tiempo. | https://arxiv.org/abs/1602.01783 |
# | PPO | On-policy | Ambos | Punta de lanza de policy gradient, incluye mecanismo para que el gradiente actualice de forma acotada, mejorando drásticamente la estabilidad. | https://arxiv.org/abs/1707.06347 |
# | DDPG | Off-policy | Continuas | Como en espacios de acciones continuos es muy difícil encontrar $\max_a Q(s,a)$, se aproxima via $Q(s, a(s \mid \theta_a))$, siendo $a$ un actor estocástico. | https://arxiv.org/abs/1509.02971 |
# | TD3 | Off-policy | Continuas | Mejora DDPG utilizando dos funciones $Q$ y retrasando la actualización para reducir la sobreestimación de $Q$. | https://arxiv.org/abs/1802.09477 |
# | SAC | Off-policy | Continuas | Usa dos funciones $Q$, introduce el bonus por entropía y usa un actor estocástico que muestrea acciones según una política $\pi$. | https://arxiv.org/abs/1801.01290 |
# ## Resumen de algunas herramientas/trucos comúnmente usados
#
# * Experience replay/buffer de experiencia: guarda las experiencias en un buffer para poder usarlas repetidamente durante el entrenamiento.
# * Ventajas: experiencias raras pero muy relevantes (ej: que tienen mucho error de actualización) quedan guardadas en memoria, pudiendo ser usadas repetidamente para aprender sin necesidad de esperar a que se repitan.
#
# En métodos de gradiente de política, en cambio, el aprendizaje queda reflejado en los pesos, lo cuál puede hacer que un agente no se desenvuelva correctamente en entornos de recompensa escasa como MountainCar.
#
# * Desventajas: requiere considerable RAM, son usables solamente en algoritmos off-policy y su muestreo no necesariamente refleja la probabilidad real de tener esas experiencias en el entorno (añadiendo sesgo), por lo que es recomendable usarlo junto con importance sampling.
#
# * Importance sampling: aplica un descuento a las actualizaciones a partir de experience replay relacionado a cuán probable era realizar esa transición.
# * Entropía: añade un bonus a la función de recompensa para que bonifique políticas $\pi(s \mid a)$ que tengan mayor entropía que otras, motivando la exploración del agente.
# * Juntar varias secuencias de imágenes. Usado principalmente en entornos de Atari para poder evaluar la dirección de movimientos.
# * Normalización de recompensas/estados. Normaliza las recompensas y observaciones usualmente con una media móvil, de modo tal que las observaciones/recompensas reflejen su relación y varianza con respecto a las demás.
# * Clipping (recorte) de recompensas. Se usaba principalmente en entornos de Atari con algoritmos tipo DQN para recortar el impacto que las distintas recompensas tenían, a una constante (ej: 1). Suele usarse como una cota máxima de recompensas/observaciones normalizadas.
# # Lab 2
#
# 1. Crear tu propio entorno y entrenar agentes RL en el mismo. Analizar la convergencia con distintos algoritmos* (ej: PPO, DQN), resultados con distintas funciones de recompensa e híper-parámetros.
#
# Algunas ideas:
#
# * Transformar GoLeftEnv en una grilla 2D, añadir paredes / trampas / agua.
# * Crear un entorno que juegue a algún juego como el ta-te-ti.
# * Crea un entorno totalmente nuevo que sea de tu interés!
#
# 2. Entrena agentes en entornos más complejos con stable-baselines/rl-baselines-zoo. Tener en cuenta:
#
# * Google Colab tiene una limitante en cuanto a cantidad de recursos de CPU/GPU (incluido un "rendimiento decreciente silencioso"), lo cuál reduce la capacidad de entrenar distintos entornos.
# * Si el entorno no está implementado en stable-baselines, debe hacerse un wrapper a mano, lo que puede ser sencillo o puede llevar algo más de trabajo, teniendo que tocar código subyacente de la librería.
#
# \* pueden ser usando stable-baselines/rl-baselines-zoo o bien utilizando algún otro algoritmo (incluso tabular)
# + [markdown] id="P5YbRAbyoIJT"
# # Recursos adicionales
#
# * [Excelente recurso para aprender más de deep RL](https://spinningup.openai.com/en/latest/spinningup/spinningup.html)
# * [Framework adicional de aprendizaje por refuerzos a gran escala](https://docs.ray.io/en/master/rllib.html)
# * [Awesome Deep RL](https://github.com/kengz/awesome-deep-rl)
# * [Comunidad de Bots de RL para Rocket League](https://rlbot.org/)
# * [Blog de Lilian Weng de Deep RL, robótica y NLP](https://lilianweng.github.io/lil-log/)
# * [The Nuts and Bolts of Deep RL Research](http://joschu.net/docs/nuts-and-bolts.pdf)
# + [markdown] id="rwXMoJy9oIJT"
# FIN
| 31,096 |
/Assignments/HW2-solution.ipynb | ff90987ea38f60f36982be53bbd5f6da4c51c1ba | [] | no_license | coder-happy/CS273a | https://github.com/coder-happy/CS273a | 1 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 106,542 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
np.random.seed(0)
import mltools as ml
reload(ml)
reload(ml.base)
import matplotlib.pyplot as plt # use matplotlib for plotting with inline plots
# %matplotlib inline
# ## P1: Linear Regression
# First, we'll load the data, and split it into training and evaluation blocks:
curve = np.genfromtxt("data/curve80.txt",delimiter=None)
print curve.shape
X = curve[:,0] # extract features
X = X[:,np.newaxis] # force X to be shape (M,1)
Y = curve[:,1] # extract target values
print X.shape, Y.shape # check shapes
Xt,Xe,Yt,Ye = ml.splitData(X,Y,0.75)
# Now, let's train a simple linear regression model on the training data, and plot the training data, evaluation data, and our linear fit:
# +
import mltools.linear
reload(ml.linear)
lr = ml.linear.linearRegress(Xt,Yt, reg=0) # init and train the model
# to plot the prediction, we'll evaluate our model at a dense set of locations:
xs = np.linspace(0,10,200)
xs = xs[:,np.newaxis]
ys = lr.predict(xs)
plt.rcParams['figure.figsize'] = (5.0, 3.0)
lines = plt.plot(xs,ys,'k-',Xt,Yt,'r.',Xe,Ye,'g.', linewidth=3,markersize=12)
plt.legend(['Prediction','Train','Test'],loc='lower right');
# -
print "MSE (Train) = ", lr.mse(Xt,Yt)
print "MSE (Test) = ", lr.mse(Xe,Ye)
# ### Polynomial features
# As mentioned in the homework, you can create additional features manually, e.g.,
Xt2 = np.zeros((Xt.shape[0],2))
print Xt2.shape
print Xt.shape
Xt2[:,0] = Xt[:,0]
Xt2[:,1] = Xt[:,0]**2
print Xt2[0:6,:] # look at a few data points to check:
# or, you can create them using the provided "fpoly" function:
import mltools.transforms as xform
Xt2 = xform.fpoly(Xt,2, bias=False)
print Xt2[0:6,:] # look at the same data points -- same values
# Now, let's try fitting a linear regression model using different numbers of polynomial features of x:
# +
degrees = np.array(range(0,19))
learners = [ [] ]*19
errT = np.zeros((19,))
errV = np.zeros((19,))
plt.rcParams['figure.figsize'] = (8.0, 5.0)
fig,ax = plt.subplots(2,3)
axFlat = [a for row in ax for a in row] # 2x3 subplots as simple list
i=0
for d in degrees:
XtP = xform.fpoly(Xt, d, bias=False) # generate polynomial features up to degree d
XtP,params = xform.rescale(XtP) # normalize scale & save transform parameters
Phi = lambda X: xform.rescale(xform.fpoly(X,d,bias=False),params)[0] # and define feature transform f'n
learners[d] = ml.linear.linearRegress( Phi(Xt), Yt , reg=1e-3)
if d in [1,3,5,7,10,18]:
axFlat[i].plot(Xt,Yt,'r.',Xe,Ye,'g.',markersize=8)
axFlat[i].set_title("Degree {}".format(d))
axisSize = axFlat[i].axis()
axFlat[i].plot(xs,learners[d].predict(Phi(xs)),'k-',linewidth=3)
axFlat[i].axis(axisSize) # restore original Y-range
i += 1
errT[d] = learners[d].mse(Phi(Xt),Yt)
errV[d] = learners[d].mse(Phi(Xe),Ye)
# -
# Finally, let's plot the training and test error as a function of the polyomial degree we used:
# +
plt.rcParams['figure.figsize'] = (5.0, 4.0)
plt.semilogy(degrees,errT,'r-',degrees,errV,'g-',linewidth=2);
# -
# From this particular plot, it looks like degree 12 is the lowest validation error, and hence the model to pick (based on this information), although probably one could argue for anything between about 6 and 14.
# ## P2: Cross-validation
# Cross validation is another method of model complexity assessment. We use it only to determine the correct setting of complexity parameters ("hyperparameters"), such as how many and which features to use, or parameters like "k" in KNN, for which training error alone provides little information. In particular, cross validation will not produce a model -- only a setting of the parameter values that cross-validation thinks will lead to a model with low test error.
#
# We imagine that we do not have our "test" data $X_E, Y_E$, that we used for assessment in the previous problem, and will try to assess the quality of fit of our polynomial regressions using only the "training" data $X_T, Y_T$. To this end, we split (again) the data multiple times, one for eack of the $K$ partitions (nFolds in the code), and repeat our entire training and evaluation procedure on each split:
# +
nFolds = 5
errX = np.zeros((19,5))
# Run over all degrees, and each fold for each degree
for d in degrees:
for iFold in range(nFolds):
# Extract the ith cross-validation fold (training/validation split)
[Xti,Xvi,Yti,Yvi] = ml.crossValidate(Xt,Yt,nFolds,iFold)
# Build the feature transform on these training data
XtiP = xform.fpoly(Xti, d, bias=False)
XtiP,params = xform.rescale(XtiP)
Phi = lambda X: xform.rescale(xform.fpoly(X,d,bias=False),params)[0]
# Create and train the model, and save the error for this degree & split:
lr = ml.linear.linearRegress( Phi(Xti),Yti )
errX[d,iFold] = lr.mse( Phi(Xvi),Yvi )
# Now, to estimate the test performance of each degree, take the average error across the folds:
errX = np.mean(errX, axis=1)
plt.rcParams['figure.figsize'] = (5.0, 4.0)
plt.semilogy(degrees,errT,'r-', # training error (from P1)
degrees,errV,'g-', # validation error (from P1)
degrees,errX,'m-', # cross-validation estimate of validation error
linewidth=2);
plt.axis([0,18,0,1e2]);
# -
# The resulting plot shows a pretty similar characterization of performance as a function of degree to the validation data used in the previous problem (which, remember, we didn't access in this problem). A noticable difference is that moderate to high degrees begin to degrade the performance (and begin overfitting) earlier than in the previous problem -- mainly, this is due to the 20% decrease in training data within our cross-validation process, which causes slighly lower degrees to begin overfitting.
#
# (If you like, you can plot a learning curve to see how performance is changing for these degrees as a function of the number of training data.)
#
# This illustrates how cross-validation can be used for model selection, as a pretty good surrogate for having additional validation / test data.
# ## P3: Kaggle
# Here is an illustration of how to load & predict data for our Kaggle competition.
Xt = np.genfromtxt("../../kaggle/kaggle.X1.train.txt",delimiter=',')
Yt = np.genfromtxt("../../kaggle/kaggle.Y.train.txt",delimiter=',')
print Xt.shape, Yt.shape # 60k training examples
lr = ml.linear.linearRegress(Xt,Yt, reg=1e-3)
print lr.mse(Xt,Yt) # how does linear regression do (training error)?
# +
Xe = np.genfromtxt('../../kaggle/kaggle.X1.test.txt',delimiter=',')
YeHat = lr.predict(Xe)
print YeHat.shape # 40k test data points for upload
# Note: the next code expects a flat vector, not a matrix, so we need to reshape it:
YeHat = YeHat[:,0]
print YeHat.shape
# +
# Output our predictions to a file:
fh = open('predictions.csv','w') # open file for upload
fh.write('ID,Prediction\n') # output header line
for i,yi in enumerate(YeHat):
fh.write('{},{}\n'.format(i+1,yi)) # output each prediction
fh.close() # close the file
# and then upload them!
| 7,502 |
/cardfraud.ipynb | babc01ee7c1e5dbf992c7c65db5b28a98ef7f06b | [] | no_license | ateyodin/week3 | https://github.com/ateyodin/week3 | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 136,532 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ateyodin/week3/blob/master/cardfraud.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="0JLGSEC1KGHK" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + id="VQQYXvGyKI1t" colab_type="code" outputId="11e5f010-a8f3-4714-826a-81e40c354bf5" colab={"base_uri": "https://localhost:8080/", "height": 51}
train_transaction = pd.read_csv('./train_transaction.csv')
test_transaction = pd.read_csv('./test_transaction.csv')
print('train transaction size:', len(train_transaction))
print('test transaction size:', len(test_transaction))
# + id="xlrZpjoeKkrp" colab_type="code" colab={}
train_identity = pd.read_csv('./train_identity.csv')
train_transaction = pd.read_csv('./train_transaction.csv')
test_identity = pd.read_csv('./test_identity.csv')
test_transaction = pd.read_csv('./test_transaction.csv')
sub= pd.read_csv('sample_submission.csv')
# + id="VVRuRpf1LNXs" colab_type="code" colab={}
train = pd.merge(train_transaction, train_identity, on='TransactionID', how='left')
test = pd.merge(test_transaction, test_identity, on='TransactionID', how='left')
# + id="XfAkj1WUMC2a" colab_type="code" outputId="f42ae55c-29f2-4661-d5e5-af207300a618" colab={"base_uri": "https://localhost:8080/", "height": 287}
train.head()
# + id="yr6u5v2vMGQh" colab_type="code" outputId="3f238780-e651-4880-b91f-cda9254c6a78" colab={"base_uri": "https://localhost:8080/", "height": 253}
test.head()
# + id="bQNtmfsBM6OX" colab_type="code" outputId="6f5ceb46-a9b9-4d94-910a-3c0f5e5cf082" colab={"base_uri": "https://localhost:8080/", "height": 300}
sns.scatterplot(x='TransactionID', y='isFraud', data=train)
# + id="vLh4yde09Hhg" colab_type="code" outputId="f2fb233b-1f44-4c0e-9df7-697f7cb74a82" colab={"base_uri": "https://localhost:8080/", "height": 386}
sns.lmplot(x='TransactionID', y='TransactionAmt', hue='isFraud', data=train)
# + id="aS3P49ZXCrkv" colab_type="code" colab={}
# + id="ocSosVnVNhGm" colab_type="code" outputId="3032ea01-6ac9-45c5-f99c-9068932afe73" colab={"base_uri": "https://localhost:8080/", "height": 300}
sns.countplot(x='isFraud', data=train)
# + id="r-svbMXtYxaB" colab_type="code" outputId="c0b6b459-a9b1-4ae6-de60-317d1a9c0c29" colab={"base_uri": "https://localhost:8080/", "height": 253}
train_new = pd.DataFrame(train, columns=train.isnull().sum().sort_values()[:185].index)
train_new = train_new.drop(columns=['TransactionID', 'TransactionDT', 'card6', 'card4', 'P_emaildomain'])
train_new_label = train_new.isFraud
train_new = train_new.drop(columns=['isFraud'])
train_new.head()
# + id="fXZRc077aM6-" colab_type="code" outputId="e4b32538-242c-4ea4-fd5f-2e9a9ceffb87" colab={"base_uri": "https://localhost:8080/", "height": 253}
test_new = pd.DataFrame(test, columns=train.isnull().sum().sort_values()[:185].index)
test_new = test_new.drop(columns=['isFraud', 'TransactionID', 'TransactionDT', 'card6', 'card4', 'P_emaildomain'])
test_new.head()
# + id="ine7zed1an4m" colab_type="code" colab={}
from sklearn.preprocessing import LabelEncoder
labelencoder = LabelEncoder()
# + id="4-dElFOyarwe" colab_type="code" outputId="6ee4b00f-753e-4933-eec1-6753ad25a523" colab={"base_uri": "https://localhost:8080/", "height": 119}
train_new['ProductCD'] = labelencoder.fit_transform(train_new['ProductCD'])
train_new.ProductCD[:5]
# + id="sI18IlCsa1Ea" colab_type="code" outputId="98e3c122-2066-43e0-9aeb-14c3480db712" colab={"base_uri": "https://localhost:8080/", "height": 119}
test_new['ProductCD'] = labelencoder.fit_transform(test_new['ProductCD'])
test_new.ProductCD[:5]
# + id="3RMM-VEsa_Os" colab_type="code" colab={}
train_new = train_new.fillna(-9999)
test_new = test_new.fillna(-9999)
# + id="VTeLJ9fl06Mg" colab_type="code" outputId="363d3d63-83e5-4e40-90b5-919c19cc419d" colab={"base_uri": "https://localhost:8080/", "height": 102}
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(train_new, train_new_label, test_size=0.35)
logistic = linear_model.LogisticRegression(C=1e5)
logistic.fit(x_train, y_train)
print('Score:', logistic.score(x_test, y_test))
# + id="WcgMAs5dHKw-" colab_type="code" outputId="6d0a388b-3bdf-4660-8bb7-17bbfece8f94" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_predicted = np.array(logistic.predict(x_test))
print(y_predicted)
| 4,806 |
/26.Intervals/Confidential_intervals.ipynb | b255502a089fd7a1332ad7776a631da4e9c027c7 | [] | no_license | iDreika/pyda-11-homework | https://github.com/iDreika/pyda-11-homework | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 2,749,804 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.display import Image
import numpy as np
import pandas as pd
import scipy.stats
import numpy as np
Image("netology-logo.png")
# +
def mean_confidence_interval(data, confidence=0.95):
n = len(data)
m, se = np.mean(data), scipy.stats.sem(data)
h = se * scipy.stats.t.ppf((1 + confidence)/2, n)
return m-h,m, m+h#,h
a = [5,7,7,9,10,16,7,14,13,15,16,15,16,17,18,13,14,15,6,13]
print(mean_confidence_interval(a, 0.95))
# -
# # Выборы в президенты или зачем нам доверительные интерваллы?!
Image("votting_usa.png")
df = pd.read_csv('2012_US_elect_county.csv', sep=',') # Откроем датасет
df.dropna(how='any',inplace=True) #Удалим пустые значения
df=df[df['%']!='#DIV/0!'][['State Postal','County Name','%']] #Удалим мусор из данных и оставим только нужное
df['%'] = df['%'].astype(float) # Приведем к численному значению результаты голосования
df['%'].hist(bins=100)
print('Среднее по всей выборке: ',df['%'].mean())
# +
some_states_1= ['AK','OH','WV','ME','CA','MD']
some_states_2 = ['WA','AK','DE']
some_states_3 = ['AZ','RI','MA']
sub_df = df.loc[df['State Postal'].isin(some_states_2)] # Выборка по нескольким штатам
l,s,r=mean_confidence_interval(sub_df['%'], 0.95)
l,s,r
# +
#df['State Postal'].value_counts() Расскоментить, если интересно соотношение штатов.
# -
Image("ArticleImage.png")
# ### Расмотрим теперь другой пример. Допустим, вы журнались и оказались в 2016 году в предверии выборов презедента в США и хотите по результатам своих исследований достаточно точно определить, кто победит на этих выборах. Сколько вам необходимо опросить людей, чтобы назвать будущего президента с точность в 95%???
# 95%. Результаты предшествующих исследований свидетельствуют, что стандартное отклонение генеральной совокупности приближенно равно 25 долл. Таким образом, е = 5, σ = 25 и Z = 1,96
Image("formuls.png")
# +
sigma = 15
Z = 1.96 # 95%
e = 5 #%
N = ((sigma*Z)/e)**2
N
# -
# # Проверим:
df = df['%']
print('Стандартное отклонение: ',df.std())
print('Среднее: ',df.mean())
df.hist(bins=80)
MEAN = df.sample(n = int(N)).mean() # Возьмем выборку из случайных n значений
print('Среднее выборки: ', MEAN)
print('Среднее генеральной совокупности: ',df.mean())
MEAN-e, MEAN+e
# from scipy.stats import norm
# norm.ppf(0.95, loc=0, scale=1)
# +
#Test with sample with identical means:
# +
from scipy import stats
np.random.seed(12345678)
rvs1 = stats.norm.rvs(loc=50,scale=10,size=500)
rvs2 = stats.norm.rvs(loc=49,scale=10, size=1500)
stats.ttest_ind(rvs1,rvs2)
# stats.ttest_ind(rvs1,rvs2, equal_var = False)
#pd.DataFrame(rvs1).hist()
#pd.DataFrame(rvs2).hist()
# -
pd.DataFrame(rvs2).hist()
# # Проверка гипотез
# ### t-Тест Стьюдента
# +
from scipy import stats
## Определим 2 случайных распределения
N = 10 #Размер выборок
b = np.random.randn(N) #Нормальное распределение с mean = 0 and var = 1
a = np.random.randn(N) + 1.5 #Нормальное распределение с mean = 2 and var = 1
### мы получаем хорошее значение p, равное 0,0005, и, таким образом, мы отклоняем нулевую гипотезу и, таким образом,
### это доказывает, что средние значения двух распределений различны и различие статистически значимо.
t , p = stats.ttest_ind(a,b)
print("t = " + str(t))
print("p = " + str(p))
# A large t-score tells you that the groups are different.
# A small t-score tells you that the groups are similar.
# -
# # a. One-sample T-test with Python
# Давайте попробуем это на одном образце. Тест покажет нам, отличаются ли средние значения выборки и генеральной совокупности. Рассмотрим количество голосующих в Индии и население всего Мира. Отличается ли средний возраст избирателей Индии от возраста населения? Давай выясним.
# +
import numpy as np
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
np.random.seed(6)
population_ages1=stats.norm.rvs(loc=18,scale=45,size=150000) #Мир
population_ages2=stats.norm.rvs(loc=18,scale=10,size=100000)
population_ages=np.concatenate((population_ages1,population_ages2))
india_ages1=stats.norm.rvs(loc=18,scale=45,size=3000) # Индия
india_ages2=stats.norm.rvs(loc=18,scale=35,size=3000)
india_ages=np.concatenate((india_ages1,india_ages2))
population_ages.mean()
# -
india_ages.mean()
stats.ttest_1samp(a=india_ages,popmean=population_ages.mean())
#Теперь это значение 1.807 говорит нам, насколько отклоняется выборочное среднее от нулевой гипотезы.
pd.DataFrame(population_ages).hist(bins=120)
pd.DataFrame(india_ages).hist(bins=120)
# # b. Two-sample T-test With Python
# Такой тест показывает, имеют ли две выборки данных разные средние значения. Здесь мы принимаем нулевую гипотезу о том, что обе группы имеют равные средние. Для этого нам не нужен известный параметр численности.
# +
np.random.seed(12)
maharashtra_ages1=stats.norm.rvs(loc=17,scale=5,size=3000)
maharashtra_ages2=stats.norm.rvs(loc=17,scale=15,size=2000)
pd.DataFrame(maharashtra_ages2).hist(bins=120)
# maharashtra_ages=np.concatenate((maharashtra_ages1,maharashtra_ages2))
# maharashtra_ages.mean()
pd.DataFrame(maharashtra_ages).hist(bins=120)
# +
stats.ttest_ind(a=india_ages,b=maharashtra_ages,equal_var=False)
#The value of 0.152 tells us there’s a 15.2% chance that the sample data is such far apart
#for two identical groups. This is greater than the 5% confidence level.
#Значение 0,152 говорит о том, что существует вероятность 7,2% того, что выборочные данные для двух
#идентичных групп сильно различаются. Это больше, чем уровень достоверности 5%.
# -
pd.DataFrame(india_ages).hist(bins=120)
pd.DataFrame(maharashtra_ages).hist(bins=120)
# # c. Paired T-test With Python
# T-критерий парной выборки, иногда называемый t-критерием зависимой выборки, представляет собой статистическую процедуру, используемую для определения того, равна ли нулю средняя разница между двумя наборами наблюдений. В парном выборочном t-тесте каждый субъект или объект измеряется дважды, в результате чего получаются пары наблюдений. Общие применения парного t-критерия выборки включают исследования случай-контроль или планы повторных измерений.
#
# Предположим, вас интересует оценка эффективности программы обучения компании. Один из подходов, который вы можете рассмотреть, - это измерение производительности выборки сотрудников до и после завершения программы и анализ различий с использованием парного выборочного t-критерия.
np.random.seed(11)
before=stats.norm.rvs(scale=30,loc=250,size=100)
after=before+stats.norm.rvs(scale=5,loc=-1.25,size=100)
weight_df=pd.DataFrame({"weight_before":before,
"weight_after":after,
"weight_change":after-before})
weight_df.describe()
stats.ttest_rel(a=before,b=after)
# Итак, мы видим, что у нас есть только 1% шансов найти такие огромные различия между образцами.
# # Практическое применение t-Тест Стьюдента
# Представим, что вы следователь какой-то крупной рекламной компаннии.
# Ваша задача иследовать рыннок манго в течени последних нескольких лет.
# У вас уже есть предварительные данные. Ваши подчиненные решили схалтурить и принести вам совсем другие отчеты.
# Сможели ли вы их разоблачить с помощью t-Тест Стьюдента и уволить?
Image("prof_redaktor.jpg")
# +
df1 = pd.read_csv('avocado.csv', sep=',', index_col=0).AveragePrice[:1000]
df2 = pd.read_csv('StudentsPerformance.csv', sep=',', index_col=0)['math score']
t, p = stats.ttest_ind(df1 ,df2)
t,p
# -
# и действительно выборки пренадлежать разным распределениям
df2.hist(bins=50)
df1.hist(bins=50)
# # $Хи^{2}- Пирсона$
# +
import scipy as sp
# Сгенерируем случайную матрицу 2х2
X = np.random.randint(2, size=50).reshape(10, 5)
y = np.random.randint(2, size=10)
contingency_table = sp.sparse.coo_matrix( (np.ones_like(y), (X[:, 0], y)),
shape=(np.unique(X[:, 0]).shape[0],
np.unique(y).shape[0])).A
print(contingency_table)
chi2, p, do,expected = sp.stats.chi2_contingency(contingency_table)
print(chi2, p, do)
print(expected)
# Возвращает:
# chi2 : The test statistic.
# p : The p-value of the test
# dof : Degrees of freedom
# expected : Ожидаемые частоты, основанные на предельных суммах таблицы.
# +
# Example of the Chi-Squared Test
from scipy.stats import chi2_contingency
table = [[10, 20, 30],[6, 9, 17]]
print(np.array(table))
stat, p, dof, expected = chi2_contingency(table)
print('stat=%.3f, p=%.3f' % (stat, p))
if p > 0.05:
print('Probably independent')
else:
print('Probably dependent')
# -
subjects = pd.DataFrame(
[
[25,46,15],
[15,44,15],
[10,10,20]
],
index=['Biology','Chemistry','Physics'],
columns=['Math SL AA','Math SL AI','Math HL'])
subjects
# Если рассчитанный хи-квадрат больше критического значения, мы отклоняем нулевую гипотезу.
# +
chi, pval, dof, exp = scipy.stats.chi2_contingency(subjects) #
print('p-value is: ', pval)
significance = 0.05
p = 1 - significance
critical_value = scipy.stats.chi2.ppf(p, dof)
print('chi=%.6f, critical value=%.6f\n' % (chi, critical_value))
if chi > critical_value:
print("""At %.2f level of significance, we reject the null hypotheses and accept H1.
They are not independent.""" % (significance))
else:
print("""At %.2f level of significance, we accept the null hypotheses.
They are independent.""" % (significance))
# -
# В качестве альтернативы мы можем сравнить p-значение и уровень значимости. Если значение p <уровня значимости, мы отклоняем нулевую гипотезу.
# +
chi, pval, dof, exp = scipy.stats.chi2_contingency(subjects)
significance = 0.05
print('p-value=%.6f, significance=%.2f\n' % (pval, significance))
if pval < significance:
print("""At %.2f level of significance, we reject the null hypotheses and accept H1.
They are not independent.""" % (significance))
else:
print("""At %.2f level of significance, we accept the null hypotheses.
They are independent.""" % (significance))
# -
# # точный критерий Фишера
# +
import scipy as sp
x = [[107,93],[74,45]]
oddsratio, pvalue = sp.stats.fisher_exact(x)
pvalue
# -
# # Проанализируем теперь рынок жилья в New York City
# New York City Airbnb Open Data
# Airbnb listings and metrics in NYC, NY, USA (2019)
Image("BARRIOS_NY.jpg")
df = pd.read_csv('AB_NYC_2019.csv')
df.dropna(how='any',inplace=True)
df.head(4)
# +
# df.neighbourhood_group.hist()
# -
df[df.price<500].price.hist(bins=50)
# +
print(df[df.price<400].price.mean())
df = pd.read_csv('AB_NYC_2019.csv')
df.dropna(how='any',inplace=True)
data = df[(df.neighbourhood_group == 'Brooklyn') | (df.neighbourhood_group=='Staten Island') | (df.neighbourhood_group=='Queens')].price
mean_confidence_interval(data, confidence=0.95) # Возможно вам понадобиться эта функция.
# -
# # Задания для самостоятельного решения
#
# 1. Найдите минимально необходимый объем выборки для построения интервальной оценки среднего с точностью ∆ = 3, дисперсией σ^2 = 225 и уровнем доверия β = 0.95.
# +
### Ваш код...
# -
# 2. Вам даны две выборки роста мужчин и женщин. Докажите, используя t-Тест Стьдента, что различия между выборками незначительно, если уровень значимости равен 0.001
# +
import scipy.stats as stats
population_men =stats.norm.rvs(loc=19,scale=171,size=11000000) # Выборка мужчин со средним ростом 171
population_women=stats.norm.rvs(loc=16,scale=165,size=12000) # Выборка женщин со средним ростом 165
### Ваш код...
# -
# 3. Определите объем необходимой выборки для исследования среднего чека за кофе в случайном городе, если известно, что в этом городе стандартное отклонение = 150, уровень доверия = 95%. Погрешность 50 рублей.
# +
### Ваш код...
# -
# 4. Представьте, что вы хотите разоблачить "волшебника", который считает, что умеет предсказывать погоду на завтра. Отвечая просто: дождь или солнце. Вы пронаблюдали за ответами "волшебника" в течении какого периода времени и получили такие результаты (см.ниже). Можно ли сказать, что маг действительно умеет предсказывать погоду,
# если уровнь значимости принять за 0.05 ?
observations = pd.DataFrame([[25,36],[15,44]],
index=['Дождь','Солнце'],
columns=['Ответ волшебника','Реальность'])
observations
# +
import scipy as sp
oddsratio, pvalue = sp.stats.fisher_exact('Таблица') # Возможно вам пригодится эта функция.
### Ваш код...
# -
# 5. Используя функцию mean_confidence_interval(data, confidence), постройте доверительный интервал с уровнем доверия 90% для выборки: data = [1,5,8,9,6,7,5,6,7,8,5,6,7,0,9,8,4,6,7,9,8,6,5,7,8,9,6,7,5,8,6,7,9,5]
#
# +
data = [4,5,8,9,6,7,5,6,7,8,5,6,7,0,9,8,4,6,7,9,8,6,5,7,8,9,6,7,5,8,6,7,9,5,10]
### Ваш код...
# -
# 6. Принадлежит ли выборка data_1 и data_2 одному множеству? Оцените это с помощью известных вам тестов проверки гипотез.
# +
data_1 = [4,5,8,9,6,7,5,6,7,8,5,6,7,0,9,8,4,6,7,9,8,6,5,7,8,9,6,7,5,8,6,7,9,5,10]
data_2 = [8,5,6,7,0,1,8,4,6,7,0,2,6,5,7,5,3,5,3,5,3,5,5,8,7,6,4,5,3,5,4,6,4,5,3,2,6,4,2,6,1,0,4,3,5,4,3,4,5,4,3,4,5,4,3,4,5,3,4,4,1,2,4,3,1,2,4,3,2,1,5,3,4,6,4,5,3,2,4,5,6,4,3,1,3,5,3,4,4,4,2,5,3]
#Ваш код:
# -
# 7. На примере датасета про жилье в New York City, мы сталкивались с примером, когда переменная имеет не совсем нормальное распределение.
#
# Предположим, Вы сформировали две гипотезы:
# Нулевая гипотеза - распределение нормальное,
# Альтернативная гипотеза - распределение не нормальное.
#
#
# Допустим, вы применили какой-то тест (сейчас неважно какой), который показал уровень значимости (p-value) = 0.03.
# Каковы будут ваши выводы? Будем считать что у нас нормальное распределение или все-таки нет? Вопрос без подвоха)
# 8. Первая выборка — это пациенты, которых лечили препаратом А.
# Вторая выборка — пациенты, которых лечили препаратом Б. Значения в выборках — это некоторая характеристика эффективности лечения (уровень метаболита в крови, температура через три дня после начала лечения, срок выздоровления, число койко-дней, и т.д.)
#
# а) Требуется выяснить, имеется ли значимое различие эффективности препаратов А и Б, или различия являются чисто случайными и объясняются «естественной» дисперсией выбранной характеристики? (уровень значимости принять за 5% или 0.05)
#
# b) При каком минимальном P-values различия были бы уже значимы?
# +
np.random.seed(11)
A = stats.norm.rvs(scale=50,loc=10,size=300)
B = A+stats.norm.rvs(scale=10,loc=-1.25,size=300)
#Ваш код:
# Подсказка, обратити внимание на Two-sample T-test
# -
| 14,770 |
/archive/3D-spatial-privacy-testing.ipynb | 79eb30ff570ac15047446772cbfe6fbf1849e32a | [
"MIT"
] | permissive | spatial-privacy/3d-spatial-privacy | https://github.com/spatial-privacy/3d-spatial-privacy | 2 | 3 | MIT | 2021-11-03T22:50:41 | 2021-11-03T22:50:04 | Jupyter Notebook | Jupyter Notebook | false | false | .py | 57,962 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python [conda env:p35_env]
# language: python
# name: conda-env-p35_env-py
# ---
# +
# %matplotlib inline
import numpy as np
import sys
import os
import matplotlib.pyplot as plt
import math
import pickle
import pandas as pd
import scipy.io
import time
import h5py
import bz2
from mpl_toolkits.mplot3d import Axes3D
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
from mpl_toolkits.axes_grid1.axes_divider import make_axes_locatable
from numpy import linalg as LA
from scipy.spatial import Delaunay
from sklearn.neighbors import NearestNeighbors
#sys.path.insert(0, "../")
from info3d import *
from nn_matchers import *
# -
# # EXTRACTING the existing sample data
# +
with open('point_collection/new_contiguous_point_collection.pickle','rb') as f:
new_contiguous_point_collection = pickle.load(f)
with open('descriptors/new_complete_res5_4by5_descriptors.pickle','rb') as f:
descriptors = pickle.load(f)
"""
Parameters
"""
# We used a radius range of 0.25 to 5.0 in increments of 0.25
radius_range = np.arange(0.5,1.6,0.5)
# Extracting the bz2 to regular pickle
for radius in radius_range:
with bz2.BZ2File('testing_samples/{}_successive_point_cloud.pickle.bz2'.format(radius), 'r') as bz2_f:
successive_point_collection = pickle.load(bz2_f)
with open('testing_samples/{}_successive_point_cloud.pickle'.format(radius), 'wb') as f:
pickle.dump(successive_point_collection,f)
with bz2.BZ2File('testing_samples/{}_partial_point_cloud.pickle.bz2'.format(radius), 'r') as bz2_f:
partial_point_collection = pickle.load(bz2_f)
with open('testing_samples/{}_partial_point_cloud.pickle'.format(radius), 'wb') as f:
pickle.dump(partial_point_collection,f)
# -
# # Step 1.1: Testing with Partial Spaces
# # 1.1 Testing our partial samples: Raw spaces
for radius in radius_range:
t0 = time.time()
with open('testing_samples/{}_partial_point_cloud.pickle'.format(radius), 'rb') as f:
partial_point_collection = pickle.load(f)
samples = len(partial_point_collection)
partial_scores = []
for obj_meta, partial_pointcloud, partial_triangles in partial_point_collection:
# ROTATION
random_theta = (2*np.pi)*np.random.random()# from [0, 2pi)
random_axis = np.random.choice(np.arange(0,3))
rotated_pointCloud = rotatePointCloud(partial_pointcloud, random_theta, random_axis)
# TRANSLATION
t_pointCloud = np.asarray(rotated_pointCloud)
random_tx_axis = np.random.choice(np.arange(0,3))
random_translation = np.random.random()
t_pointCloud[:,random_tx_axis] = t_pointCloud[:,random_tx_axis] + random_translation
t_triangles = np.asarray(partial_triangles)#-vertices_length
t1 = time.time()
try:
p_descriptors, p_keypoints, p_d_c = getSpinImageDescriptors(
t_pointCloud,
down_resolution = 5,
cylindrical_quantization = [4,5]
)
except Exception as ex:
print("Error getting descriptors of",obj_meta)
print("Error Message:",ex)
continue
# Resetting the diff_Ratio matrix
diff_scores = np.ones((p_descriptors.shape[0],len(descriptors),2))
diff_ratios = np.ones((p_descriptors.shape[0],len(descriptors)))
diff_indexs = np.ones((p_descriptors.shape[0],len(descriptors),2))
#print(diff_ratios.shape)
local_keypoint_matches = []
for i_r, ref_descriptor in enumerate(descriptors):
r_descriptors = ref_descriptor[1]
r_keypoints = ref_descriptor[2]
matching_range = np.arange(r_descriptors.shape[1])
try:
f_nearestneighbor, diff = getMatches(p_descriptors,r_descriptors,2,range_to_match=matching_range)
diff = diff/np.amax(diff) # max-normalization of differences
diff_ratio = diff[:,0]/diff[:,1]
diff_ratios[:,i_r] = diff_ratio
diff_scores[:,i_r] = diff
diff_indexs[:,i_r] = f_nearestneighbor
# Taking note of the matched keypoints
local_keypoint_matches.append([
obj_meta,
p_keypoints,
r_keypoints[f_nearestneighbor[:,0]]
])
except Exception as ex:
print(rotation,"Error Matching:",ex)
# Accumulating the diff_ratio matrix for every partial (rotated) object
partial_scores.append([
obj_meta,
np.asarray(diff_ratios),
np.asarray(diff_indexs),
np.asarray(diff_scores),
local_keypoint_matches
])
if len(partial_scores) % 10 == (samples-1)%10:
#print('Test')
print(" radius = {}: Done with {} iterations. Time to match {:.3f} seconds.".format(
radius,
len(partial_scores),
time.time()-t0)
)
t0 = time.time()
current_errors = NN_matcher(partial_scores)
print(" Error Rate:",np.sum(current_errors[:,1]/len(partial_scores)))
partial_errors = NN_matcher(partial_scores)
print(radius,"Error Rate:",np.sum(partial_errors[:,1]/len(partial_scores)))
with open('testing_results/partials/radius_{}_RAW_scores.pickle'.format(radius), 'wb') as f:
pickle.dump(partial_scores,f)
with open('testing_results/partials/radius_{}_RAW_errors.pickle'.format(radius), 'wb') as f:
pickle.dump(partial_errors,f)
# # 1.2 Testing our partial samples: RANSAC spaces
for radius in radius_range:
t0 = time.time()
with open('testing_samples/{}_partial_point_cloud.pickle'.format(radius), 'rb') as f:
partial_point_collection = pickle.load(f)
samples = len(partial_point_collection)
partial_scores = []
partial_properties = []
for obj_meta, partial_pointcloud, partial_triangles in partial_point_collection:
# ROTATION
random_theta = (2*np.pi)*np.random.random()# from [0, 2pi)
random_axis = np.random.choice(np.arange(0,3))
rotated_pointCloud = rotatePointCloud(partial_pointcloud, random_theta, random_axis)
# TRANSLATION
t_pointCloud = np.asarray(rotated_pointCloud)
random_tx_axis = np.random.choice(np.arange(0,3))
random_translation = np.random.random()
t_pointCloud[:,random_tx_axis] = t_pointCloud[:,random_tx_axis] + random_translation
t_triangles = np.asarray(partial_triangles)#-vertices_length
#if object_name % 13 == 0:
# print('{}: Processing iteration {} of object {}.'.format(radius,iteration,object_name))
t1 = time.time()
# GETTING GENERALIZATION
gen_planes = getRansacPlanes(
t_pointCloud
)
p_pointcloud, p_triangles = getGeneralizedPointCloud(
planes=gen_planes,
)
try:
p_descriptors, p_keypoints, p_d_c = getSpinImageDescriptors(
p_pointcloud,
down_resolution = 5,
cylindrical_quantization = [4,5]
)
except Exception as ex:
print("Error getting descriptors of",obj_meta)
print("Error Message:",ex)
continue
# Resetting the diff_Ratio matrix
diff_scores = np.ones((p_descriptors.shape[0],len(descriptors),2))
diff_ratios = np.ones((p_descriptors.shape[0],len(descriptors)))
diff_indexs = np.ones((p_descriptors.shape[0],len(descriptors),2))
#print(diff_ratios.shape)
local_keypoint_matches = []
for i_r, ref_descriptor in enumerate(descriptors):
#o_, r_descriptors, r_keypoints, r_d_c
r_descriptors = ref_descriptor[1]
r_keypoints = ref_descriptor[2]
matching_range = np.arange(r_descriptors.shape[1])
try:
f_nearestneighbor, diff = getMatches(p_descriptors,r_descriptors,2,range_to_match=matching_range)
diff = diff/np.amax(diff) # max-normalization of differences
diff_ratio = diff[:,0]/diff[:,1]
diff_ratios[:,i_r] = diff_ratio
diff_scores[:,i_r] = diff
diff_indexs[:,i_r] = f_nearestneighbor
# Taking note of the matched keypoints
local_keypoint_matches.append([
obj_meta,
p_keypoints,
r_keypoints[f_nearestneighbor[:,0]]
])
except Exception as ex:
print(rotation,"Error Matching:",ex)
# Accumulating the diff_ratio matrix for every partial (rotated) object
partial_scores.append([
obj_meta,
np.asarray(diff_ratios),
np.asarray(diff_indexs),
np.asarray(diff_scores),
local_keypoint_matches
])
if len(partial_scores) % 10 == (samples-1)%10:
#print('Test')
print(" radius = {}: Done with {} iterations. Time to match {:.3f} seconds.".format(
radius,
len(partial_scores),
time.time()-t0)
)
t0 = time.time()
current_errors = NN_matcher(partial_scores)
print(" Error Rate:",np.sum(current_errors[:,1]/len(partial_scores)))
partial_errors = NN_matcher(partial_scores)
print(radius,"Error Rate:",np.sum(partial_errors[:,1]/len(partial_scores)))
with open('testing_results/partials/radius_{}_RANSAC_scores.pickle'.format(radius), 'wb') as f:
pickle.dump(partial_scores,f)
with open('testing_results/partials/radius_{}_RANSAC_errors.pickle'.format(radius), 'wb') as f:
pickle.dump(partial_errors,f)
# # 1.3 Results of partial spaces
# +
fig=plt.figure(figsize=(9, 3))
RawNN = []
RansacGeneralizedNN = []
RawNN_intra_errors = []
RansacGeneralizedNN_intra_errors = []
for radius in radius_range:
try:
with open('testing_results/partials/radius_{}_RAW_errors.pickle'.format(radius), 'rb') as f:
partial_errors = pickle.load(f)
RawNN.append([
radius,
np.mean(partial_errors[:,1]),
np.std(partial_errors[:,1]),
])
correct_interspace_labels_idxs = np.where(partial_errors[:,1]==0)[0]
intraspace_errors = partial_errors[correct_interspace_labels_idxs,2]
RawNN_intra_errors.append([
radius,
np.nanmean(intraspace_errors),
np.nanstd(intraspace_errors)
])
except:
pass
try:
with open('testing_results/partials/radius_{}_RANSAC_errors.pickle'.format(radius), 'rb') as f:
partial_errors = pickle.load(f)
RansacGeneralizedNN.append([
radius,
np.nanmean(partial_errors[:,1]),
np.nanstd(partial_errors[:,1]),
])
correct_interspace_labels_idxs = np.where(partial_errors[:,1]==0)[0]
intraspace_errors = partial_errors[correct_interspace_labels_idxs,2]
RansacGeneralizedNN_intra_errors.append([
radius,
np.nanmean(intraspace_errors),
np.nanstd(intraspace_errors)
])
except:
pass
RansacGeneralizedNN = np.asarray(RansacGeneralizedNN)
RawNN = np.asarray(RawNN)
RawNN_intra_errors = np.asarray(RawNN_intra_errors)
RansacGeneralizedNN_intra_errors = np.asarray(RansacGeneralizedNN_intra_errors)
ax1 = fig.add_subplot(121)
ax1.grid(alpha = 0.7)
ax1.set_ylim(-0.025,1.025)
ax1.set_xlim(radius_range[0]-0.25,radius_range[-1]+0.25)
markersize = 8
ax1.set_ylabel("INTER-space Privacy")
ax1.set_xlabel("Partial Radius")
#ax1.set_yticklabels(fontsize = 16)
#ax1.set_xticklabels(fontsize = 16)
ax1.plot(
RawNN[:,0],RawNN[:,1],
"-o",
linewidth = 2,
mew = 2,markersize = markersize,
label = "Raw"
)
ax1.plot(
RansacGeneralizedNN[:,0],RansacGeneralizedNN[:,1],
"-s",
linewidth = 2,
mew = 2,markersize = markersize,
label = "RANSAC"
)
ax1.legend(loc = "lower left")
ax2 = fig.add_subplot(122)
ax2.grid(alpha = 0.7)
ax2.set_ylim(-0.25,10.25)
ax2.set_xlim(radius_range[0]-0.25,radius_range[-1]+0.25)
ax2.set_ylabel("INTRA-space Privacy")
ax2.set_xlabel("Partial Radius")
#ax2.set_yticklabels(fontsize = 16)
#ax2.set_xticklabels(fontsize = 16)
plt.minorticks_on()
ax2.plot(
RawNN_intra_errors[:,0],
RawNN_intra_errors[:,1],
linewidth = 2,
marker = 'o',fillstyle = 'none',
mew = 2,markersize = markersize,
label = "Raw"
)
ax2.plot(
RansacGeneralizedNN_intra_errors[:,0],
RansacGeneralizedNN_intra_errors[:,1],
linewidth = 2,
marker = 's',fillstyle = 'none',
mew = 2,markersize = markersize,
label = "RANSAC"
)
ax2.legend(loc = "lower left");
# -
# # Step 2 Testing with successively released partial spaces
# +
"""
Parameters
"""
# We used a radius range of 0.25 to 5.0 in increments of 0.25.
radius_range = radius_range[:2]
# For our work, we orignally used 50 samples with further 100 successive releases for our investigation.
# Below are lower parameters, change as desired.
#samples = 50
#releases = 50
# For demonstration purposes, we skip testing some successive samples but we still accumulate them.
skip = 3
# -
# # 2.1 Testing the successive case: RAW and RANSAC
for radius in radius_range:
t0 = time.time()
try:
with open('testing_samples/{}_successive_point_cloud.pickle'.format(radius), 'rb') as f:
successive_point_collection = pickle.load(f)
samples = len(successive_point_collection)
except:
pass
successive_scores = []
successive_errors = []
g_successive_scores = []
g_successive_errors = []
for obj_, growing_point_collection in successive_point_collection:
iteration_scores = []
g_iteration_scores = []
# ROTATION param
random_theta = (2*np.pi)*np.random.random()# from [0, 2pi)
random_axis = np.random.choice(np.arange(0,3))
# TRANSLATION param
random_tx_axis = np.random.choice(np.arange(0,3))
random_translation = np.random.random()
growing_point_cloud = []
growing_p_point_cloud = []
growing_p_triangles = []
releases = len(growing_point_collection)
release_count = 0
for obj_meta, partial_pointcloud, partial_triangles in growing_point_collection:
#i_obj, released_growing_point_collection in enumerate(growing_point_collection):
rotated_pointCloud = rotatePointCloud(partial_pointcloud, random_theta, random_axis)
# TRANSLATION
t_pointCloud = np.asarray(rotated_pointCloud)
t_pointCloud[:,random_tx_axis] = t_pointCloud[:,random_tx_axis] + random_translation
t_triangles = np.asarray(partial_triangles)#-vertices_length
#Regular Accumulation
if len(growing_point_cloud) == 0:
growing_point_cloud = t_pointCloud
else:
growing_point_cloud = np.concatenate(
(growing_point_cloud,t_pointCloud),
axis=0
)
#RANSAC generalizations
if len(growing_p_point_cloud) == 0:
gen_planes = getLOCALIZEDRansacPlanes(
pointCloud = t_pointCloud,
original_vertex = obj_meta[-1]
)
else:
gen_planes = updatePlanesWithSubsumption(
new_pointCloud=t_pointCloud,
existing_pointCloud=growing_p_point_cloud,
planes_to_find = max(min(release_count,50),30),
#verbose=True
)
if len(gen_planes) == 0:
print("No gen planes after release",release_count,growing_point_cloud.shape)
continue
try:
updated_point_cloud, updated_triangles = getGeneralizedPointCloud(
planes = gen_planes,
triangle_area_threshold = 0.2,#2.0*np.amax(getTriangleAreas(partial_pointCloud, partial_triangles))
#verbose = True
)
growing_p_point_cloud = updated_point_cloud
growing_p_triangles = updated_triangles
#print(" Successful:",release_count,len(growing_p_point_cloud), len(growing_p_triangles),partial_pointCloud.shape)
except Exception as ex:
print("Error getting updated point cloud in release",release_count+1)
print(" ",growing_p_point_cloud.shape, growing_p_triangles.shape,partial_pointCloud.shape)
#print(ex)
continue
if len(growing_p_point_cloud) == 0:
continue
release_count += 1
if release_count % skip != 1: # skip
continue
#Regular Processing
growing_point_cloud = np.unique(growing_point_cloud,axis=0)
try:
p_descriptors, p_keypoints, p_d_c = getSpinImageDescriptors(
growing_point_cloud,
down_resolution = 5,
cylindrical_quantization = [4,5]
)
except:
p_descriptors = []
p_keypoints = []
# Resetting the diff_Ratio matrix
diff_scores = np.ones((p_descriptors.shape[0],len(descriptors),2))
diff_ratios = np.ones((p_descriptors.shape[0],len(descriptors)))
diff_indexs = np.ones((p_descriptors.shape[0],len(descriptors),2))
local_keypoint_matches = []
for i_r, ref_descriptor in enumerate(descriptors):
r_descriptors = ref_descriptor[1]
r_keypoints = ref_descriptor[2]
matching_range = np.arange(r_descriptors.shape[1])
try:
f_nearestneighbor, diff = getMatches(p_descriptors,r_descriptors,2,range_to_match=matching_range)
diff = diff/np.amax(diff) # max-normalization of differences
diff_ratio = diff[:,0]/diff[:,1]
diff_ratios[:,i_r] = diff_ratio
diff_scores[:,i_r] = diff
diff_indexs[:,i_r] = f_nearestneighbor
# Taking note of the matched keypoints
local_keypoint_matches.append([
obj_meta,
p_keypoints,
r_keypoints[f_nearestneighbor[:,0]]
])
except Exception as ex:
print(rotation,"Error Matching:",ex)
# Accumulating the diff_ratio matrix for every partial (rotated) object
iteration_scores.append([
obj_meta,
diff_ratios,
diff_indexs,
diff_scores,
local_keypoint_matches
])
if release_count % 2 == 0:
#print('Test')
print(" radius = {}: Done with {} releases ({}) ({} samples). Time to match {:.3f} seconds. ({})".format(
radius,
len(iteration_scores),
release_count,
len(successive_scores),
time.time()-t0,
growing_point_cloud.shape
)
)
t0 = time.time()
current_errors = NN_matcher(iteration_scores)
print(" Error Rate:",np.sum(current_errors[:,1]/len(iteration_scores)))
#RANSAC Processing
try:
p_descriptors, p_keypoints, p_d_c = getSpinImageDescriptors(
growing_p_point_cloud,
down_resolution = 5,
cylindrical_quantization = [4,5]
)
except:
p_descriptors = []
p_keypoints = []
print("Error getting descriptors at release",release_count,"; using next release as this release.")
#release_count -= 1
continue
# Resetting the diff_Ratio matrix
diff_scores = np.ones((p_descriptors.shape[0],len(descriptors),2))
diff_ratios = np.ones((p_descriptors.shape[0],len(descriptors)))
diff_indexs = np.ones((p_descriptors.shape[0],len(descriptors),2))
local_keypoint_matches = []
for i_r, ref_descriptor in enumerate(descriptors):
#o_, r_descriptors, r_keypoints, r_d_c
r_descriptors = ref_descriptor[1]
r_keypoints = ref_descriptor[2]
matching_range = np.arange(r_descriptors.shape[1])
try:
f_nearestneighbor, diff = getMatches(p_descriptors,r_descriptors,2,range_to_match=matching_range)
diff = diff/np.amax(diff) # max-normalization of differences
diff_ratio = diff[:,0]/diff[:,1]
diff_ratios[:,i_r] = diff_ratio
diff_scores[:,i_r] = diff
diff_indexs[:,i_r] = f_nearestneighbor
# Taking note of the matched keypoints
local_keypoint_matches.append([
obj_meta,
p_keypoints,
r_keypoints[f_nearestneighbor[:,0]]
])
except Exception as ex:
print(rotation,"Error Matching:",ex)
# Accumulating the diff_ratio matrix for every partial (rotated) object
g_iteration_scores.append([
obj_meta,
diff_ratios,
diff_indexs,
diff_scores,
local_keypoint_matches
])
if release_count % 2 == 0:
#print('Test')
print(" radius = {} [G]: Done with {} releases ({}) ({} samples). Time to match {:.3f} seconds. ({})".format(
radius,
len(g_iteration_scores),
release_count,
len(g_successive_scores),
time.time()-t0,
growing_p_point_cloud.shape
)
)
t0 = time.time()
current_errors = NN_matcher(g_iteration_scores)
print(" G_Error Rate:",np.sum(current_errors[:,1]/len(g_iteration_scores)))
iteration_errors = NN_matcher(iteration_scores)
g_iteration_errors = NN_matcher(g_iteration_scores)
if len(successive_scores) % 5 == (samples-1)%5 :
try:
print(radius,len(successive_scores),"Error Rate:",np.sum(iteration_errors[:,1]/len(iteration_scores)))
print(radius,len(g_successive_scores),"G_Error Rate:",np.sum(g_iteration_errors[:,1]/len(g_iteration_scores)))
except:
pass
successive_scores.append([
obj_,
iteration_scores
])
successive_errors.append([
obj_,
iteration_errors
])
g_successive_scores.append([
obj_,
g_iteration_scores
])
g_successive_errors.append([
obj_,
g_iteration_errors
])
with open('testing_results/successive/radius_{}_RAW_successive_scores.pickle'.format(radius), 'wb') as f:
pickle.dump(successive_scores,f)
with open('testing_results/successive/radius_{}_RAW_successive_errors.pickle'.format(radius), 'wb') as f:
pickle.dump(successive_errors,f)
with open('testing_results/successive/radius_{}_RANSAC_successive_scores.pickle'.format(radius), 'wb') as f:
pickle.dump(g_successive_scores,f)
with open('testing_results/successive/radius_{}_RANSAC_successive_errors.pickle'.format(radius), 'wb') as f:
pickle.dump(g_successive_errors,f)
# # 2.2 Results of the successive case
# +
succ_RawNN_errors = []
succ_RawNN_partial_errors = []
succ_RansacGeneralizedNN_errors = []
succ_RansacGeneralizedNN_partial_errors = []
t0 = time.time()
for radius in radius_range:
succ_RawNN_per_iteration_errors = []
succ_RansacGeneralizedNN_per_iteration_errors = []
try:
with open('testing_results/successive/radius_{}_RAW_successive_scores.pickle'.format(radius), 'rb') as f:
successive_scores = pickle.load(f)
with open('testing_results/successive/radius_{}_RAW_successive_errors.pickle'.format(radius), 'rb') as f:
successive_errors = pickle.load(f)
for obj_, iteration_errors in successive_errors:
#print(" RAW",radius,iteration_errors.shape)
if iteration_errors.shape[0] < int(releases/skip):
continue
else:
succ_RawNN_per_iteration_errors.append(iteration_errors[:int(releases/skip)])
succ_RawNN_errors.append([
radius,
np.asarray(succ_RawNN_per_iteration_errors)
])
print(np.asarray(succ_RawNN_per_iteration_errors).shape)
except:
pass
try:
with open('testing_results/successive/radius_{}_RANSAC_successive_scores.pickle'.format(radius), 'rb') as f:
successive_scores = pickle.load(f)
with open('testing_results/successive/radius_{}_RANSAC_successive_errors.pickle'.format(radius), 'rb') as f:
successive_errors = pickle.load(f)
for obj_, iteration_scores in successive_scores:#[:-1]:
#print(" RANSAC",radius,iteration_errors.shape)
iteration_errors = NN_matcher(iteration_scores)
if iteration_errors.shape[0] < int(releases/skip):
continue
else:
succ_RansacGeneralizedNN_per_iteration_errors.append(iteration_errors[:int(releases/skip)])
succ_RansacGeneralizedNN_errors.append([
radius,
np.asarray(succ_RansacGeneralizedNN_per_iteration_errors)
])
print(np.asarray(succ_RansacGeneralizedNN_errors).shape)
except:# Exception as ex:
#print(radius,": successive RansacNN\n ", ex)
pass
print("Done with radius = {:.2f} in {:.3f} seconds".format(radius,time.time() - t0))
t0 = time.time()
for radius, per_iteration_errors in succ_RawNN_errors:
#print(per_iteration_errors.shape)
succ_RawNN_partial_errors_per_rel = []
for rel_i in np.arange(per_iteration_errors.shape[1]):
correct_interspace_labels_idxs = np.where(per_iteration_errors[:,rel_i,1]==0)[0]
intraspace_errors = per_iteration_errors[correct_interspace_labels_idxs,rel_i,2]
succ_RawNN_partial_errors_per_rel.append([
rel_i,
np.mean(intraspace_errors),
np.std(intraspace_errors)
])
succ_RawNN_partial_errors.append([
radius,
np.asarray(succ_RawNN_partial_errors_per_rel)
])
for radius, per_iteration_errors in succ_RansacGeneralizedNN_errors:
#print(radius,per_iteration_errors.shape)
succ_RansacGeneralizedNN_errors_per_rel = []
for rel_i in np.arange(per_iteration_errors.shape[1]):
correct_interspace_labels_idxs = np.where(per_iteration_errors[:,rel_i,1]==0)[0]
intraspace_errors = per_iteration_errors[correct_interspace_labels_idxs,rel_i,2]
succ_RansacGeneralizedNN_errors_per_rel.append([
rel_i,
np.mean(intraspace_errors),
np.std(intraspace_errors)
])
succ_RansacGeneralizedNN_partial_errors.append([
radius,
np.asarray(succ_RansacGeneralizedNN_errors_per_rel)
])
# +
fig=plt.figure(figsize=(15, 5))
ax1 = fig.add_subplot(121)
ax1.grid(alpha = 0.7)
ax1.set_ylim(-0.025,1.025)
ax1.set_xlim(0,releases-skip)
markersize = 8
ax1.set_ylabel("INTER-space Privacy")
ax1.set_xlabel("Releases")
for radius, RawNN_per_iteration_errors in succ_RawNN_errors:
print(RawNN_per_iteration_errors.shape)
ax1.plot(
np.arange(1,releases-skip,skip),#[:RawNN_per_iteration_errors.shape[1]],
np.mean(RawNN_per_iteration_errors[:,:,1], axis = 0),
':o',
label = "r ="+ str(radius) + " Raw"
)
for radius, RansacNN_per_iteration_errors in succ_RansacGeneralizedNN_errors:
print(RansacNN_per_iteration_errors.shape)
ax1.plot(
np.arange(1,releases-skip,skip),
np.mean(RansacNN_per_iteration_errors[:,:,1], axis = 0),
'-s',
label = "r ="+ str(radius) + " RANSAC"
)
ax1.legend(loc = "best", ncol = 2)
ax2 = fig.add_subplot(122)
ax2.grid(alpha = 0.7)
ax2.set_ylim(-0.25,12.25)
ax2.set_xlim(0,releases-skip)
ax2.set_ylabel("INTRA-space Privacy")
ax2.set_xlabel("Partial Radius")
#ax2.set_yticklabels(fontsize = 16)
#ax2.set_xticklabels(fontsize = 16)
#plt.minorticks_on()
for radius, errors_per_rel in succ_RansacGeneralizedNN_partial_errors:
ax2.plot(
np.arange(1,releases-skip,skip),
errors_per_rel[:,1],
#errors_per_rel[:,2],
'-s',
linewidth = 2, #capsize = 4.0,
#marker = markers[0],
#fillstyle = 'none',
mew = 2, markersize = markersize,
label = "r ="+ str(radius)+", RANSAC"
)
ax2.legend(loc = "best")
# -
# # Step 3.1 Testing with conservative plane releasing
# +
"""
Parameters:
Also, we use the same successive samples from successive releasing for direct comparability of results.
"""
# We used a radius range of 0.25 to 5.0 in increments of 0.25.
radius_range = radius_range[:2]
# For our work, we orignally used 50 samples with further 100 successive releases for our investigation.
# Below are lower parameters, change as desired.
samples = 25
releases = 50
planes = np.arange(1,30,3)
# For demonstration purposes, we skip testing some successive samples but we still accumulate them.
skip = 3
# -
for radius in radius_range:
t0 = time.time()
try:
with open('testing_samples/{}_successive_point_cloud.pickle'.format(radius), 'rb') as f:
successive_point_collection = pickle.load(f)
samples = len(successive_point_collection)
except:
pass
conservative_scores = []
conservative_errors = []
for obj_, growing_point_collection in successive_point_collection:
# ROTATION param
random_theta = (2*np.pi)*np.random.random()# from [0, 2pi)
random_axis = np.random.choice(np.arange(0,3))
# TRANSLATION param
random_tx_axis = np.random.choice(np.arange(0,3))
random_translation = np.random.random()
per_plane_scores = []
per_plane_errors = []
latest_plane = 0
for plane_number in planes:
print(radius," {} planes, ({} Done)".format(plane_number,len(per_plane_scores)))
g_iteration_scores = []
growing_point_cloud = []
growing_p_point_cloud = []
growing_p_triangles = []
release_count = 0
for obj_meta, partial_pointcloud, partial_triangles in growing_point_collection:
#i_obj, released_growing_point_collection in enumerate(growing_point_collection):
releases = len(growing_point_collection)
# ROTATION
rotated_pointCloud = rotatePointCloud(partial_pointcloud, random_theta, random_axis)
# TRANSLATION
t_pointCloud = np.asarray(rotated_pointCloud)
t_pointCloud[:,random_tx_axis] = t_pointCloud[:,random_tx_axis] + random_translation
t_triangles = np.asarray(partial_triangles)#-vertices_length
#RANSAC generalizations
try:
if len(growing_p_point_cloud) == 0:
gen_planes = getLOCALIZEDRansacPlanes(
pointCloud = t_pointCloud,#np.where(hPointConnectivity < 5)[0],axis = 0),
original_vertex = obj_meta[-1],
planes_to_find=plane_number
#verbose=True
#threshold = threshold # the point-plane distance threshold
)
else:
gen_planes = updatePlanesWithSubsumption(
new_pointCloud=t_pointCloud,
existing_pointCloud=growing_p_point_cloud,
planes_to_find = plane_number
#verbose=True
)
except:
print("Error getting planes",release_count,growing_point_cloud.shape)
continue
if len(gen_planes) == 0:
print("No gen planes after release",release_count,growing_point_cloud.shape)
continue
try:
updated_point_cloud, updated_triangles = getGeneralizedPointCloud(
planes = gen_planes,
triangle_area_threshold = 0.2,#2.0*np.amax(getTriangleAreas(partial_pointCloud, partial_triangles))
#verbose = True
)
growing_p_point_cloud = updated_point_cloud
growing_p_triangles = updated_triangles
#print(" Successful:",release_count,len(growing_p_point_cloud), len(growing_p_triangles),partial_pointCloud.shape)
except Exception as ex:
print("Error getting updated point cloud in release",release_count+1)
print(" ",growing_p_point_cloud.shape, growing_p_triangles.shape,partial_pointCloud.shape)
#print(ex)
continue
if len(growing_p_point_cloud) == 0:
continue
release_count += 1
if release_count % skip != 1: # skip
continue
#RANSAC Processing
try:
p_descriptors, p_keypoints, p_d_c = getSpinImageDescriptors(
growing_p_point_cloud,
down_resolution = 5,
cylindrical_quantization = [4,5]
)
except:
p_descriptors = []
p_keypoints = []
print("Error getting descriptors at release",release_count,"; using next release as this release.")
#release_count -= 1
continue
# Resetting the diff_Ratio matrix
diff_scores = np.ones((p_descriptors.shape[0],len(descriptors),2))
diff_ratios = np.ones((p_descriptors.shape[0],len(descriptors)))
diff_indexs = np.ones((p_descriptors.shape[0],len(descriptors),2))
local_keypoint_matches = []
for i_r, ref_descriptor in enumerate(descriptors):
#o_, r_descriptors, r_keypoints, r_d_c
r_descriptors = ref_descriptor[1]
r_keypoints = ref_descriptor[2]
matching_range = np.arange(r_descriptors.shape[1])
try:
f_nearestneighbor, diff = getMatches(p_descriptors,r_descriptors,2,range_to_match=matching_range)
diff = diff/np.amax(diff) # max-normalization of differences
diff_ratio = diff[:,0]/diff[:,1]
diff_ratios[:,i_r] = diff_ratio
diff_scores[:,i_r] = diff
diff_indexs[:,i_r] = f_nearestneighbor
# Taking note of the matched keypoints
local_keypoint_matches.append([
obj_meta,
p_keypoints,
r_keypoints[f_nearestneighbor[:,0]]
])
except Exception as ex:
print(rotation,"Error Matching:",ex)
# Accumulating the diff_ratio matrix for every partial (rotated) object
g_iteration_scores.append([
obj_meta,
diff_ratios,
diff_indexs,
diff_scores,
local_keypoint_matches
])
# --- !
if release_count % 2 == 0:
#print('Test')
print(" radius = {} [G]: Done with {} releases ({}) ({} samples). Time to match {:.3f} seconds. ({})".format(
radius,
len(g_iteration_scores),
release_count,
len(conservative_scores),
time.time()-t0,
growing_p_point_cloud.shape
)
)
t0 = time.time()
current_errors = NN_matcher(g_iteration_scores)
print(" ({} actual planes) G_Error Rate: {}".format(
len(gen_planes),
np.sum(current_errors[:,1]/len(g_iteration_scores))
))
g_iteration_errors = NN_matcher(g_iteration_scores)
try:
print(radius,plane_number,len(conservative_scores),"G_Error Rate:",np.sum(g_iteration_errors[:,1]/len(g_iteration_scores)))
except:
pass
per_plane_scores.append([
plane_number,
g_iteration_scores
])
per_plane_errors.append([
plane_number,
g_iteration_errors
])
conservative_scores.append([
obj_,
per_plane_scores
])
conservative_errors.append([
obj_,
per_plane_errors
])
if len(conservative_scores) % 5 == (samples-1)%5 :
try:
print("Radius = {}, {} objects done".format(radius,len(g_successive_scores)))
except:
pass
with open('testing_results/conservative/radius_{}_RANSAC_conservative_scores.pickle'.format(radius), 'wb') as f:
pickle.dump(conservative_scores,f)
with open('testing_results/conservative/radius_{}_RANSAC_conservative_errors.pickle'.format(radius), 'wb') as f:
pickle.dump(conservative_errors,f)
# # 3.2 Results with conservative plane releasing
# +
conservative_RANSAC_error_results = []
t0 = time.time()
for radius in radius_range:
succ_RansacGeneralizedNN_per_iteration_errors = []
try:
with open('testing_results/conservative/radius_{}_RANSAC_conservative_scores.pickle'.format(radius), 'rb') as f:
conservative_scores = pickle.load(f)
for obj_, per_plane_scores in conservative_scores:#[:-1]:
per_plane_errors = []
skipped= False
for max_plane, iteration_scores in per_plane_scores:
iteration_errors = NN_matcher(iteration_scores)
if iteration_errors.shape[0] >= int(releases/skip):
per_plane_errors.append(iteration_errors[:int(releases/skip)])
else:
skipped = True
#print("RANSAC: skipped",iteration_errors.shape)
if not skipped:
succ_RansacGeneralizedNN_per_iteration_errors.append(per_plane_errors)
conservative_RANSAC_error_results.append([
radius,
np.asarray(succ_RansacGeneralizedNN_per_iteration_errors)
])
print(np.asarray(succ_RansacGeneralizedNN_per_iteration_errors).shape)
except Exception as ex:
print(radius,": conservative RansacNN\n ", ex)
pass
print("Done with radius = {:.2f} in {:.3f} seconds".format(radius,time.time() - t0))
t0 = time.time()
"""
# Uncomment below if you want to overwrite the existing results.
"""
#with open('testing_results/conservative/conservative_RANSAC_error_results.pickle', 'wb') as f:
# pickle.dump(conservative_RANSAC_error_results,f)
# +
"""
Preparing the results of the case with *Conservative Releasing*.
"""
releases_range = np.arange(1,releases-skip,skip)
X, Y = np.meshgrid(releases_range, planes)
test_vp_cn_05 = np.asarray(conservative_RANSAC_error_results[0][1])
mean_vp_cn_05 = np.mean(test_vp_cn_05[:,:,:,1],axis = 0)
#test_vp_cn_10 = np.asarray(conservative_RANSAC_error_results[1][1])
#mean_vp_cn_10 = np.mean(test_vp_cn_10[:,:,:,1],axis = 0)
intra_vp_cn_05 = np.zeros(test_vp_cn_05.shape[1:])
#intra_vp_cn_10 = np.zeros(test_vp_cn_10.shape[1:])
for plane_i, plane in enumerate(planes):
for rel_i, rel in enumerate(releases_range):
correct_interspace_labels_idxs_05 = np.where(test_vp_cn_05[:,plane_i,rel_i,1]==0)[0]
#correct_interspace_labels_idxs_10 = np.where(test_vp_cn_10[:,plane_i,rel_i,1]==0)[0]
intraspace_errors_05 = test_vp_cn_05[correct_interspace_labels_idxs_05,plane_i,rel_i,2]
#intraspace_errors_10 = test_vp_cn_10[correct_interspace_labels_idxs_10,plane_i,rel_i,2]
intra_vp_cn_05[plane_i,rel_i] = np.asarray([
np.mean(intraspace_errors_05),
np.std(intraspace_errors_05),
0,
np.nan
])
# +
fig = plt.figure(figsize=(11,8))
ax = plt.axes(projection='3d')
surf = ax.plot_surface(
X, Y,
mean_vp_cn_05,
cmap=plt.cm.plasma,
)
surf.set_clim(0.0,1.0)
ax.set_title("r = 0.5", fontsize = 24)
ax.set_xlabel("Releases", labelpad=10, fontsize = 24)
ax.set_xlim(0,releases)
ax.set_xticklabels(releases_range,fontsize = 16)
ax.set_zlabel("INTER-space Privacy", labelpad=10, fontsize = 24)
ax.set_zlim(0,1)
ax.set_zticklabels([0,0.2,0.4,0.6,0.8,1.0],fontsize = 16)
ax.set_ylabel("Max number of planes", labelpad=10, fontsize = 22)#, offset = 1)
ax.set_ylim(0,30)
ax.set_yticklabels(np.arange(0,35,5),fontsize = 16)
cbar = fig.colorbar(surf, aspect=30, ticks = np.arange(0.0,1.1,0.25))
cbar.ax.set_yticklabels(np.arange(0.0,1.1,0.25),fontsize = 16)
ax.view_init(25,135);
# -
| 43,861 |
/ground_truth_labeling_jobs/3d_dense_point_cloud_downsampling_tutorial/ground_truth_annotation_dense_point_cloud_tutorial.ipynb | fa1728ae792b55f0f0a8be9cec39e2cd5ac825a3 | [
"Apache-2.0"
] | permissive | vidyaravipati/amazon-sagemaker-examples | https://github.com/vidyaravipati/amazon-sagemaker-examples | 0 | 1 | Apache-2.0 | 2021-05-04T22:10:52 | 2021-05-04T20:22:00 | null | Jupyter Notebook | false | false | .py | 35,592 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Annotation of Dense Point Clouds Using Amazon SageMaker Ground Truth
#
# This notebook walks through how to annotate dense point clouds using SageMaker Ground Truth. We define dense point clouds as point clouds with a high number of points per scene (> 500k) and have enough points relative to the feature size of a scene that labelers can still make out objects in the scene (cars, pedestrians, etc) while only looking at a subset of the scene's total points. When these conditions are met, you can often improve the labeling experience and throughput by reducing your scene's point cloud size through downsampling techniques. After labeling, you can either directly use labels on the full-size scene for some modalities (like object detection bounding boxes), or you can "upsample" your labels by doing some in-sample prediction of the unlabeled points.
#
# Recommended Kernel: `conda_python3`
#
# Let's start by importing required libraries and initializing session and other variables used in this notebook. By default, the notebook uses the default Amazon S3 bucket in the same AWS Region you use to run this notebook. If you want to use a different S3 bucket, make sure it is in the same AWS Region you use to complete this tutorial, and specify the bucket name for `bucket`. It is also recommended to use an instance size with at least 16 GB of RAM.
# +
# %pylab inline
import boto3
import json
import uuid
import zlib
from array import array
import sagemaker as sm
import scipy.stats
from sklearn.neighbors import KNeighborsClassifier
from mpl_toolkits.mplot3d import Axes3D
# -
# ## Prerequisites
#
# You will create some of the resources you need to launch a Ground Truth audit labeling job in this notebook. You must create the following resources before executing this notebook:
#
# * A work team. A work team is a group of workers that complete labeling tasks. If you want to preview the worker UI and execute the labeling task you will need to create a private work team, add yourself as a worker to this team, and provide the work team ARN below. This [GIF](images/create-workteam-loop.gif) demonstrates how to quickly create a private work team on the Amazon SageMaker console. To learn more about private, vendor, and Amazon Mechanical Turk workforces, see [Create and Manage Workforces
# ](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-workforce-management.html).
# +
WORKTEAM_ARN = '<<ADD WORK TEAM ARN HERE>>'
# Make sure workteam arn is populated if private work team is chosen
assert (WORKTEAM_ARN != '<<ADD WORK TEAM ARN HERE>>')
# -
# * The IAM execution role you used to create this notebook instance must have the following permissions:
# * If you do not require granular permissions for your use case, you can attach [AmazonSageMakerFullAccess](https://console.aws.amazon.com/iam/home?#/policies/arn:aws:iam::aws:policy/AmazonSageMakerFullAccess) to your IAM user or role. If you are running this example in a SageMaker notebook instance, this is the IAM execution role used to create your notebook instance.If you need granular permissions, please see [Assign IAM Permissions to Use Ground Truth](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-security-permission.html#sms-security-permissions-get-started) for granular policy to use Ground Truth.
# * AWS managed policy [AmazonSageMakerGroundTruthExecution](https://console.aws.amazon.com/iam/home#policies/arn:aws:iam::aws:policy/AmazonSageMakerGroundTruthExecution). Run the following code-block to see your IAM execution role name. This [GIF](images/add-policy-loop.gif) demonstrates how to attach this policy to an IAM role in the IAM console. You can also find instructions in the IAM User Guide: [Adding and removing IAM identity permissions](https://docs.aws.amazon.com/IAM/latest/UserGuide/access_policies_manage-attach-detach.html#add-policies-console).
# * When you create your role, you specify Amazon S3 permissions. Make sure that your IAM role has access to the S3 bucket that you plan to use in this example. If you do not specify an S3 bucket in this notebook, the default bucket in the AWS region you are running this notebook instance will be used. If you do not require granular permissions, you can attach [AmazonS3FullAccess](https://console.aws.amazon.com/iam/home#policies/arn:aws:iam::aws:policy/AmazonS3FullAccess) to your role.
# +
role = sm.get_execution_role()
role_name = role.split('/')[-1]
print('IMPORTANT: Make sure this IAM role has one or more IAM policies with the permissions described above attached.')
print('********************************************************************************')
print('The IAM execution role name:', role_name)
print('The IAM execution role ARN:', role)
print('********************************************************************************')
sagemaker_cl = boto3.client('sagemaker')
# Make sure the bucket is in the same region as this notebook.
bucket = '<< YOUR S3 BUCKET NAME >>'
sm_session = sm.Session()
s3 = boto3.client('s3')
if bucket == '<< YOUR S3 BUCKET NAME >>':
bucket=sm_session.default_bucket()
region = boto3.session.Session().region_name
bucket_region = s3.head_bucket(Bucket=bucket)['ResponseMetadata']['HTTPHeaders']['x-amz-bucket-region']
assert bucket_region == region, f'Your S3 bucket {bucket} and this notebook need to be in the same region.'
print(f'IMPORTANT: make sure the role {role_name} has the access to read and write to this bucket.')
print('********************************************************************************************************')
print(f'This notebook will use the following S3 bucket: {bucket}')
print('********************************************************************************************************')
print("If you don't have a CORS policy attached, you will be unable to label data in the task, use the following policy.")
print("""
[
{
"AllowedHeaders": [
"*"
],
"AllowedMethods": [
"GET",
"HEAD",
"PUT"
],
"AllowedOrigins": [
"*"
],
"ExposeHeaders": [
"Access-Control-Allow-Origin"
],
"MaxAgeSeconds": 3000
}
]
""")
# -
# ## Section 1: Download Data
#
# You can download from the following location. There is an object called `rooftop_12_49_41.xyz` that you will visualize, downsample, and then label. This data is a scan of an apartment building rooftop generated using the 3d Scanner App on an iPhone12 Pro. The app allows users to scan a given area and then export a pointcloud file. In this case the pointcloud data is in xyzrgb format, an accepted format for SageMaker Ground Truth point cloud, for more information on the data types allowed in SageMaker Ground Truth point cloud refer to the following documentation:
#
# https://docs.aws.amazon.com/sagemaker/latest/dg/sms-point-cloud-raw-data-types.html
# !wget https://aws-ml-blog.s3.amazonaws.com/artifacts/annotating-dense-point-cloud-data/rooftop_12_49_41.xyz -O pointcloud.xyz
# +
# Let's read our dataset into a numpy file, we'll call our pointclouds "pc" for short.
# Our point cloud consists of 3D euclidean coordinates of each point and RGB color values for each point.
# For example:
# p1_x, p1_y, p1_z, p1_r, p1_g, p1_b
# p2_x, p2_y, p2_z, p2_r, p2_g, p2_b
# p3_x, p3_y, p3_z, p3_r, p3_g, p3_b
# ...
pc = np.loadtxt("pointcloud.xyz", delimiter=",")
print(f"Loaded points of shape {pc.shape}")
# -
# ### Visualize our pointcloud
#
# You can visualize the point cloud using [matplotlib's scatter3d](https://matplotlib.org/stable/gallery/mplot3d/scatter3d.html) function. The pointcloud file contains all of the correct points but in this case is not rotated correctly. You can rotate the object around it's axis by multiplying the pointcloud by a rotation matrix. You can obtain a rotation matrix using [scipy](https://docs.scipy.org/doc/scipy/reference/generated/scipy.spatial.transform.Rotation.html) and specifying the degree changes you would like to make to each axis using the from_euler method.
# +
# playing with view of 3D scene
from scipy.spatial.transform import Rotation
def plot_pointcloud(pc, rot = [[0,0,0]], color=True, title="Simple Downsampling 1", figsize=(50,25), verbose=False):
rot1 = Rotation.from_euler('zyx', rot, degrees=True)
R1 = rot1.as_matrix()
if verbose:
print('Rotation matrix:','\n',R1)
# matrix multiplication between our rotation matrix and pointcloud
pc_show = np.matmul(R1, pc.copy()[:,:3].transpose() ).transpose()
if color:
rot_color = pc.copy()[:,3:]
fig = plt.figure( figsize=figsize)
ax = fig.add_subplot(111, projection="3d")
ax.set_title(title, fontdict={'fontsize':20})
if color:
# need to create list of tuples for matplotlib to plot RGB color values
color_tuple_list = []
for i in range(pc_show.shape[0]):
color_tuple = (int(np.abs(rot_color[i,0]))/255, int(np.abs(rot_color[i,1]))/255, int(np.abs(rot_color[i,2]))/255)
color_tuple_list.append(color_tuple)
ax.scatter(pc_show[:,0], pc_show[:,1], pc_show[:,2], c=color_tuple_list, s=0.05)
else:
ax.scatter(pc_show[:,0], pc_show[:,1], pc_show[:,2], c='blue', s=0.05)
# rotate in z direction 30 degrees, y direction 90 degrees, and x direction 60 degrees
# only run this once!
rot = Rotation.from_euler('zyx', [[30,90,60]], degrees=True)
R = rot.as_matrix()
print('Rotation matrix:','\n', R)
pc_rot = np.matmul(R, pc[:,:3].transpose() ).transpose().squeeze()
pc_color = pc.copy()[:,3:]
pc = np.zeros_like(pc)
pc[:,:3] = pc_rot
pc[:,3:] = pc_color
# +
# view full pointcloud, will take ~10 seconds to render depending on the instance type you are using
plot_pointcloud(pc, rot = [[0,0,0]], color=True, title="Full Pointcloud", figsize=(50,30))
# -
# ## Section 2: Downsample Approaches
#
# Just like in image processing, there are many different approaches to performing downsampling in a point cloud.
# Certain approaches may preserve more information relevant for labelers, or may have different noise characteristics. You'll walk through two approaches, a very basic linear sampling and a more advanced interpolation method.
#
# Other alternatives include 3d box filtering, median filtering, and even random sampling.
#
#
# ### I: Basic Approach
# The simplest form of downsampling is to choose values using a fixed step size based on how large we want our resulting point cloud to be.
target_num_pts = 500_000
subsample = int(np.ceil(len(pc) / target_num_pts))
pc_downsample_simple = pc[::subsample]
print(f"We've subsampled to {len(pc_downsample_simple)} points")
# ### II: Advanced Approach
#
# A more advanced approach is to break the input space into cubes, otherwise known as voxels, and choose a single point per box using an averaging function.
#
# In this example you will use cube sizes of 1.3 cm, which will partition the entire pointcloud scene into 1.3 cm by 1.3 cm cubes. Then for each cube, you will get a list of the points contained within the cube. Given the coordinates of this list of points, you take the centroid, or mean across each coordinate dimension to get a mean point that is representative of the overall set of points within the voxel. You run the operation twice, once over the xyz values then again over the color channel RGB values since you want to use the mins and maxes of the xyz values, but also preserve the color.
#
# You will use scipy's `binned_statistic_dd` to efficiently compute these representative points per voxel, then construct a new point cloud using these representative points and use it for downstream labeling.
# +
boxsize = 0.013 # 1.3 cm box size.
mins = pc[:,:3].min(axis=0)
maxes = pc[:,:3].max(axis=0)
volume = maxes - mins
num_boxes_per_axis = np.ceil(volume / boxsize).astype('int32').tolist()
print('Number of boxes per axis', num_boxes_per_axis)
# For each voxel or "box", use the mean of the box to chose which points are in the box.
means, _, _ = scipy.stats.binned_statistic_dd(
pc[:,:3],
[pc[:,0], pc[:,1], pc[:,2]],
statistic="mean",
bins=num_boxes_per_axis,
)
# run a second operation over the color channels
means2, _, _ = scipy.stats.binned_statistic_dd(
pc[:,:3],
[pc[:,3], pc[:,4], pc[:,5]],
statistic="mean",
bins=num_boxes_per_axis,
)
x_means = means[0,~np.isnan(means[0])].flatten()
y_means = means[1,~np.isnan(means[1])].flatten()
z_means = means[2,~np.isnan(means[2])].flatten()
r_means = means2[0,~np.isnan(means2[0])].flatten()
b_means = means2[1,~np.isnan(means2[1])].flatten()
g_means = means2[2,~np.isnan(means2[2])].flatten()
pc_downsample_adv = np.column_stack([x_means, y_means, z_means, r_means, b_means, g_means])
print('downsampled pointcloud shape:',pc_downsample_adv.shape)
del x_means, y_means, z_means, r_means, b_means, g_means, means, num_boxes_per_axis
# -
# ## Section 3: Visualize 3D Rendering
#
# You can visualize pointclouds using a 3D scatter plot of the points. You will see that the advanced voxel mean method creates a "smoother" point cloud as averaging has a noise reduction effect.
# +
fig = plt.figure( figsize=(50, 30))
ax = fig.add_subplot(121, projection="3d")
ax.set_title("Simple Downsampling 1", fontdict={'fontsize':20})
color_tuple_list = []
for i in range(pc_downsample_simple.shape[0]):
color_tuple = (int(pc_downsample_simple[i,3])/255, int(pc_downsample_simple[i,4])/255, int(pc_downsample_simple[i,5])/255)
color_tuple_list.append(color_tuple)
ax.scatter(pc_downsample_simple[:,0], pc_downsample_simple[:,1], pc_downsample_simple[:,2], c=color_tuple_list, s=0.05)
ax = fig.add_subplot(122, projection="3d")
ax.set_title("Voxel Mean Downsampling 1", fontdict={'fontsize':20})
color_tuple_list = []
for i in range(pc_downsample_adv.shape[0]):
color_tuple = (int(pc_downsample_adv[i,3])/255, int(pc_downsample_adv[i,4])/255, int(pc_downsample_adv[i,5])/255)
color_tuple_list.append(color_tuple)
ax.scatter(pc_downsample_adv[:,0], pc_downsample_adv[:,1], pc_downsample_adv[:,2], c=color_tuple_list, s=0.05)
# -
# Below you can look at the point clouds from different perspectives by running the following function, which will multiply the pointclouds by the specified rotation matrices. The current setting will rotate the pointcloud 90 degrees to the right.
#
# try some different angles
plot_pointcloud(pc_downsample_adv, rot = [[90,0,0]], color=True, title="Advanced Downsampling", figsize=(50,30))
# ## Section 4: Amazon SageMaker Ground Truth (SMGT) Labeling Job
#
# You can submit this downsampled point cloud as a SageMaker Ground Truth job by converting to SMGT labeling format, then creating a labeling job using this point cloud as input. You only save the points themselves since you are going to launch a semantic segmentation labeling task where you will label the points using distinct colors.
# +
# Save in Ground Truth format
# See: https://docs.aws.amazon.com/sagemaker/latest/dg/sms-point-cloud-raw-data-types.html
np.savetxt("pointcloud.txt", pc_downsample_adv[:,:3], delimiter=" ")
# -
# !aws s3 cp pointcloud.txt s3://{bucket}/input/pointcloud.txt
# In the following cell, you generate an input manifest file. This file tells Ground Truth where 3D point cloud input data is located and contains metadata about your input data. To learn more, see [Create an Input Manifest File for a 3D Point Cloud Labeling Job](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-point-cloud-input-manifest.html).
# +
# Generate an input manifest.
input_manifest = json.dumps({
"source-ref": f"s3://{bucket}/input/pointcloud.txt",
"source-ref-metadata": {
"format": "text/xyz",
"unix-timestamp": 1566861644.759115,
}
})
input_manifest_name = "input.manifest"
with open(input_manifest_name, "w") as f:
f.write(input_manifest)
input_manifest_s3_uri = f"s3://{bucket}/input/input.manifest"
# !aws s3 cp {input_manifest_name} {input_manifest_s3_uri}
# -
# In the following cell, you generate a label category configuration file. This file contains the label categories workers use to label the 3D point cloud, worker instructions, and other information about your task. To learn more about all of the options you have when you create a label category configuration file, see [Create a Labeling Category Configuration File with Label Category and Frame Attributes](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-label-cat-config-attributes.html).
# +
# Generate label category configuration.
label_category_config = json.dumps({
"labels": [
{"label": "Floor", "attributes": []},
{"label": "Wall", "attributes": []},
{"label": "Furniture", "attributes": []},
{"label": "Plant", "attributes": []}
],
"frameAttributes": [],
"categoryGlobalAttributes": [],
"instructions": {
"shortInstruction": "Label",
"fullInstruction": '<ul><li>After you’ve painted an object, use the <strong>Unpaint</strong> category to remove paint. </li><li>If you need to paint an object that is partially blocked by another object that you have painted, select the Paint cloud as background tool before you start painting.</li><li>Use the <strong>Shortcuts</strong> menu to see keyboard shortcuts that you can use to label objects faster.</li><li>Use the <strong>View</strong> menu to modify your view of the 3D point cloud and the worker portal.</li><li>Use the <strong>3D Point Cloud </strong>menu to modify the perspective of and pixel-attributes you see in the 3D point cloud.</li><li>Use this <a href="https://docs.aws.amazon.com/sagemaker/latest/dg/sms-point-cloud-worker-instructions-semantic-segmentation.html" rel="noopener noreferrer" target="_blank">resource</a> to learn about worker portal navigation, tools available to complete your task, icons, and view options.</li></ul>',
},
"document-version": "2020-03-01",
})
label_category_config_filename = "labels.json"
with open(label_category_config_filename, "w") as f:
f.write(label_category_config)
label_category_config_s3_uri = f"s3://{bucket}/input/labels.json"
# !aws s3 cp {label_category_config_filename} {label_category_config_s3_uri}
# -
# In the following cell you configure the labeling job request. To learn more about the parameters used to configure a labeling job, see [Create a Labeling Job (API)](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-create-labeling-job-api.html).
# +
hash_str = str(uuid.uuid4())[:4]
job_name = f"downsample-demo-{hash_str}"
lambda_account_id = {
"us-east-1": "432418664414",
"us-east-2": "266458841044",
"us-west-2": "081040173940",
"ca-central-1": "918755190332",
"eu-west-1": "568282634449",
"eu-west-2": "487402164563",
"eu-central-1": "203001061592",
"ap-northeast-1": "477331159723",
"ap-northeast-2": "845288260483",
"ap-south-1": "565803892007",
"ap-southeast-1": "377565633583",
"ap-southeast-2": "454466003867",
}[region]
job_creation_request = {
"LabelingJobName": job_name,
"LabelAttributeName": f"{job_name}-ref",
"InputConfig": {
"DataSource": {
"S3DataSource": {
"ManifestS3Uri": input_manifest_s3_uri,
}
},
"DataAttributes": {"ContentClassifiers": []},
},
"OutputConfig": {
"S3OutputPath": f"s3://{bucket}/output",
"KmsKeyId": "",
},
"RoleArn": role,
"LabelCategoryConfigS3Uri": label_category_config_s3_uri,
"StoppingConditions": {"MaxPercentageOfInputDatasetLabeled": 100},
"HumanTaskConfig": {
"WorkteamArn": WORKTEAM_ARN,
"UiConfig": {
"HumanTaskUiArn": f"arn:aws:sagemaker:{region}:394669845002:human-task-ui/PointCloudSemanticSegmentation"
},
"PreHumanTaskLambdaArn": f"arn:aws:lambda:{region}:{lambda_account_id}:function:PRE-3DPointCloudSemanticSegmentation",
"TaskKeywords": ["Point cloud", "segmentation"],
"TaskTitle": job_name,
"TaskDescription": "Create a semantic segmentation mask by painting objects in a 3D point cloud",
"NumberOfHumanWorkersPerDataObject": 1,
"TaskTimeLimitInSeconds": 604800,
"TaskAvailabilityLifetimeInSeconds": 864000,
"MaxConcurrentTaskCount": 1000,
"AnnotationConsolidationConfig": {
"AnnotationConsolidationLambdaArn": f"arn:aws:lambda:{region}:{lambda_account_id}:function:ACS-3DPointCloudSemanticSegmentation"
},
},
}
# Review the request for any errors before submitting.
print(json.dumps(job_creation_request, indent=2))
# -
sagemaker_cl.create_labeling_job(**job_creation_request)
# ### Label Data in the Worker Portal
# Run the following cell to see the worker portal URL of your private workforce. Use this URL to login to the worker portal to see your labeling tasks. See the [3D point cloud worker instructions](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-point-cloud-worker-instructions.html) to learn more about the feature you'll see in the worker UI.
workforce = sagemaker_cl.describe_workforce(
WorkforceName="default"
)
worker_portal_url = workforce["Workforce"]["SubDomain"]
print(f"Sign-in by going here: {worker_portal_url}")
# ### Label Data in the Worker Portal
# After submitting the job in the worker portal, wait until the labeling job has status "complete"
# +
job = sagemaker_cl.describe_labeling_job(LabelingJobName=job_name)
status = job["LabelingJobStatus"]
print(f"Status: '{status}'")
assert status == "Completed"
label_attribute_name = job["LabelAttributeName"]
output_manifest_s3_uri = job["LabelingJobOutput"]["OutputDatasetS3Uri"]
# !aws s3 cp {output_manifest_s3_uri} output.manifest
with open("output.manifest") as f:
manifest_line = f.read().splitlines()[0]
data_object = json.loads(manifest_line)
labels_s3_uri = data_object[label_attribute_name]
# !aws s3 cp {labels_s3_uri} pointcloud.zlib
# -
# ### Load Annotations From S3
#
# +
def semantic_segmentation_to_classes(filename):
"""Convert from class map labels to numpy array."""
with open(filename, "rb") as f:
binary_content = zlib.decompress(f.read())
my_classes = array("B", binary_content)
return np.array(my_classes)
annotations = semantic_segmentation_to_classes("pointcloud.zlib")
# Verify we have a label for each point in the downsampled point cloud.
assert len(annotations) == len(pc_downsample_adv)
# -
# ## Section 5: KNN Upsample Approach
# Now that we have the downsampled labels, we will train a K-NN classifier from SKLearn to predict the full dataset labels by treating our annotated points as training data and performing inference on the remainder of the unlabeled points in our full size point cloud. Note that you can tune the number of points used as well as the distance metric and weighting scheme to influence how label inference is performed. If you label a few tiles in the full-size dataset, you can use those labeled tiles as ground truth to evaluate the accuracy of the K-NN predictions. This accuracy metric can then be used for hyperparameter tuning of K-NN or to try different inference algorithms to reduce your number of misclassified points between object boundaries, resulting in the lowest possible in-sample error rate.
# +
# There's a lot of possibility to tune KNN further
# 1) Prevent classification of points far away from all other points (random unfiltered ground point)
# 2) Perform a non-uniform weighted vote
# 3) Tweak number of neighbors
knn = KNeighborsClassifier(n_neighbors=3)
print(f"Training on {len(pc_downsample_adv)} labeled points")
knn.fit(pc_downsample_adv[:,:3], annotations)
print(f"Upsampled to {len(pc)} labeled points")
annotations_full = knn.predict(pc[:,:3])
# -
# ## Section 6: Visualize 3D Rendering
#
# Now that you've performed upsampling of the labeled data, you can visualize a tile of the original full size point cloud.
# +
pc_downsample_annotated = np.column_stack((pc_downsample_adv[:,:3], annotations))
pc_annotated = np.column_stack((pc[:,:3], annotations_full))
labeled_area = pc_downsample_annotated[pc_downsample_annotated[:,3] != 255]
min_bounds = np.min(labeled_area, axis=0)
max_bounds = np.max(labeled_area, axis=0)
min_bounds = [-2, -2, -5.5, -1]
max_bounds = [2, 2, 0, 256]
def extract_tile(point_cloud, min_bounds, max_bounds):
return point_cloud[
(point_cloud[:,0] > min_bounds[0])
& (point_cloud[:,1] > min_bounds[1])
& (point_cloud[:,2] > min_bounds[2])
& (point_cloud[:,0] < max_bounds[0])
& (point_cloud[:,1] < max_bounds[1])
& (point_cloud[:,2] < max_bounds[2])
]
tile_downsample_annotated = extract_tile(pc_downsample_annotated, min_bounds, max_bounds)
tile_annotated = extract_tile(pc_annotated, min_bounds, max_bounds)
# +
down_rot = tile_downsample_annotated.copy()
down_rot_color = tile_downsample_annotated.copy()[:,3:].astype('int32')
# change the color scheme to clearly display classes
down_rot_color[down_rot_color==0]=50
down_rot_color[down_rot_color==1]=100
down_rot_color[down_rot_color==2]=150
down_rot_color[down_rot_color==3]=200
full_rot = tile_annotated.copy()
full_rot_color = tile_annotated.copy()[:,3:].astype('int32')
full_rot_color[full_rot_color==0]=50
full_rot_color[full_rot_color==1]=100
full_rot_color[full_rot_color==2]=150
full_rot_color[full_rot_color==3]=200
fig = plt.figure(figsize=(50, 20))
ax = fig.add_subplot(121, projection="3d")
ax.set_title("Downsampled Annotations", fontdict={'fontsize':25})
ax.scatter(down_rot[:,0], down_rot[:,1], down_rot[:,2], c=down_rot_color[:,0], s=0.05)
ax = fig.add_subplot(122, projection="3d")
ax.set_title("Upsampled Annotations", fontdict={'fontsize':25})
ax.scatter(full_rot[:,0], full_rot[:,1], full_rot[:,2], c=full_rot_color[:,0], s=0.05)
# -
# ## Conclusion
#
# This notebook demonstrated a couple approaches to performing downsampling on pointclouds for data labeling use cases. These approaches can also be applied as preprocessing steps before training and inference. The notebook demonstrated a simple step based approach as well as a a more complex approach where you broke the pointcloud bounds into voxels and replaced each voxel with a summary point created by taking the mean of all points in the voxel.
#
# You then learned how to convert these pointclouds into a SageMaker Ground Truth Semantic Segmentation job. Finally you learned how to take the resulting labels and apply them to the full-size point cloud using some in-sample prediction based on the K Nearest Neighbors algorithm.
#
# Using these approaches, you can save time and effort labeling high-density pointclouds by avoiding tiling and instead labeling downsampled pointclouds.
#
# ## Cleanup
#
# If for some reason you didn't complete the labeling job, you can run the following command to stop it.
sagemaker_cl.stop_labeling_job(job_name)
| 27,198 |
/generatePatterns.ipynb | 57b2cc422bcab4ea4792ae5e0a54ab7542ac3bad | [] | no_license | Silvidheer/DataScience_Python3Assignment3.2 | https://github.com/Silvidheer/DataScience_Python3Assignment3.2 | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 3,502 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
arr = [6, 7, 8, 9]
print(type(arr))
# -
a = np.array(arr)
print(type(a))
print(a.shape)
print(a.dtype)
# +
#get the dimension of a with ndim
print(a.ndim)
b = np.array([[1, 2, 3], [4, 5, 6]])
print(b)
# -
# a 2x3 array with random values
np.random.random((2, 3))
#a 2x3 array of zeros
np.zeros((2, 3))
#a 2x3 array of ones
np.ones((2, 3))
#a 3x3 identity matrix
np.zeros((3, 3, 3))
# +
c = np.array([[9.0, 8.0, 7.0], [1.0, 2.0, 3.0]])
d = np.array([[4.0, 5.0, 6.0], [9.0, 8.0, 7.0]])
c + d
# +
c = np.array([[9.0, 8.0, 7.0], [1.0, 2.0, 3.0]])
d = np.array([[4.0, 5.0, 6.0], [9.0, 8.0, 7.0]])
c * d
# +
c = np.array([[9.0, 8.0, 7.0], [1.0, 2.0, 3.0]])
d = np.array([[4.0, 5.0, 6.0], [9.0, 8.0, 7.0]])
5/d
# +
c = np.array([[9.0, 8.0, 7.0], [1.0, 2.0, 3.0]])
d = np.array([[4.0, 5.0, 6.0], [9.0, 8.0, 7.0]])
c ** 2
# -
a[0]
d[1, 0:2]
# +
e = np.array([[10, 11, 12], [13, 14, 15],
[16, 17, 18], [19, 20, 21]])
#slicing
e[:3, :2]
# -
e[[2, 0, 3, 1], [2, 1, 0, 2]]
e[e>15]
3IgaW1wbGllZC4KLy8gU2VlIHRoZSBMaWNlbnNlIGZvciB0aGUgc3BlY2lmaWMgbGFuZ3VhZ2UgZ292ZXJuaW5nIHBlcm1pc3Npb25zIGFuZAovLyBsaW1pdGF0aW9ucyB1bmRlciB0aGUgTGljZW5zZS4KCi8qKgogKiBAZmlsZW92ZXJ2aWV3IEhlbHBlcnMgZm9yIGdvb2dsZS5jb2xhYiBQeXRob24gbW9kdWxlLgogKi8KKGZ1bmN0aW9uKHNjb3BlKSB7CmZ1bmN0aW9uIHNwYW4odGV4dCwgc3R5bGVBdHRyaWJ1dGVzID0ge30pIHsKICBjb25zdCBlbGVtZW50ID0gZG9jdW1lbnQuY3JlYXRlRWxlbWVudCgnc3BhbicpOwogIGVsZW1lbnQudGV4dENvbnRlbnQgPSB0ZXh0OwogIGZvciAoY29uc3Qga2V5IG9mIE9iamVjdC5rZXlzKHN0eWxlQXR0cmlidXRlcykpIHsKICAgIGVsZW1lbnQuc3R5bGVba2V5XSA9IHN0eWxlQXR0cmlidXRlc1trZXldOwogIH0KICByZXR1cm4gZWxlbWVudDsKfQoKLy8gTWF4IG51bWJlciBvZiBieXRlcyB3aGljaCB3aWxsIGJlIHVwbG9hZGVkIGF0IGEgdGltZS4KY29uc3QgTUFYX1BBWUxPQURfU0laRSA9IDEwMCAqIDEwMjQ7Ci8vIE1heCBhbW91bnQgb2YgdGltZSB0byBibG9jayB3YWl0aW5nIGZvciB0aGUgdXNlci4KY29uc3QgRklMRV9DSEFOR0VfVElNRU9VVF9NUyA9IDMwICogMTAwMDsKCmZ1bmN0aW9uIF91cGxvYWRGaWxlcyhpbnB1dElkLCBvdXRwdXRJZCkgewogIGNvbnN0IHN0ZXBzID0gdXBsb2FkRmlsZXNTdGVwKGlucHV0SWQsIG91dHB1dElkKTsKICBjb25zdCBvdXRwdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQob3V0cHV0SWQpOwogIC8vIENhY2hlIHN0ZXBzIG9uIHRoZSBvdXRwdXRFbGVtZW50IHRvIG1ha2UgaXQgYXZhaWxhYmxlIGZvciB0aGUgbmV4dCBjYWxsCiAgLy8gdG8gdXBsb2FkRmlsZXNDb250aW51ZSBmcm9tIFB5dGhvbi4KICBvdXRwdXRFbGVtZW50LnN0ZXBzID0gc3RlcHM7CgogIHJldHVybiBfdXBsb2FkRmlsZXNDb250aW51ZShvdXRwdXRJZCk7Cn0KCi8vIFRoaXMgaXMgcm91Z2hseSBhbiBhc3luYyBnZW5lcmF0b3IgKG5vdCBzdXBwb3J0ZWQgaW4gdGhlIGJyb3dzZXIgeWV0KSwKLy8gd2hlcmUgdGhlcmUgYXJlIG11bHRpcGxlIGFzeW5jaHJvbm91cyBzdGVwcyBhbmQgdGhlIFB5dGhvbiBzaWRlIGlzIGdvaW5nCi8vIHRvIHBvbGwgZm9yIGNvbXBsZXRpb24gb2YgZWFjaCBzdGVwLgovLyBUaGlzIHVzZXMgYSBQcm9taXNlIHRvIGJsb2NrIHRoZSBweXRob24gc2lkZSBvbiBjb21wbGV0aW9uIG9mIGVhY2ggc3RlcCwKLy8gdGhlbiBwYXNzZXMgdGhlIHJlc3VsdCBvZiB0aGUgcHJldmlvdXMgc3RlcCBhcyB0aGUgaW5wdXQgdG8gdGhlIG5leHQgc3RlcC4KZnVuY3Rpb24gX3VwbG9hZEZpbGVzQ29udGludWUob3V0cHV0SWQpIHsKICBjb25zdCBvdXRwdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQob3V0cHV0SWQpOwogIGNvbnN0IHN0ZXBzID0gb3V0cHV0RWxlbWVudC5zdGVwczsKCiAgY29uc3QgbmV4dCA9IHN0ZXBzLm5leHQob3V0cHV0RWxlbWVudC5sYXN0UHJvbWlzZVZhbHVlKTsKICByZXR1cm4gUHJvbWlzZS5yZXNvbHZlKG5leHQudmFsdWUucHJvbWlzZSkudGhlbigodmFsdWUpID0+IHsKICAgIC8vIENhY2hlIHRoZSBsYXN0IHByb21pc2UgdmFsdWUgdG8gbWFrZSBpdCBhdmFpbGFibGUgdG8gdGhlIG5leHQKICAgIC8vIHN0ZXAgb2YgdGhlIGdlbmVyYXRvci4KICAgIG91dHB1dEVsZW1lbnQubGFzdFByb21pc2VWYWx1ZSA9IHZhbHVlOwogICAgcmV0dXJuIG5leHQudmFsdWUucmVzcG9uc2U7CiAgfSk7Cn0KCi8qKgogKiBHZW5lcmF0b3IgZnVuY3Rpb24gd2hpY2ggaXMgY2FsbGVkIGJldHdlZW4gZWFjaCBhc3luYyBzdGVwIG9mIHRoZSB1cGxvYWQKICogcHJvY2Vzcy4KICogQHBhcmFtIHtzdHJpbmd9IGlucHV0SWQgRWxlbWVudCBJRCBvZiB0aGUgaW5wdXQgZmlsZSBwaWNrZXIgZWxlbWVudC4KICogQHBhcmFtIHtzdHJpbmd9IG91dHB1dElkIEVsZW1lbnQgSUQgb2YgdGhlIG91dHB1dCBkaXNwbGF5LgogKiBAcmV0dXJuIHshSXRlcmFibGU8IU9iamVjdD59IEl0ZXJhYmxlIG9mIG5leHQgc3RlcHMuCiAqLwpmdW5jdGlvbiogdXBsb2FkRmlsZXNTdGVwKGlucHV0SWQsIG91dHB1dElkKSB7CiAgY29uc3QgaW5wdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQoaW5wdXRJZCk7CiAgaW5wdXRFbGVtZW50LmRpc2FibGVkID0gZmFsc2U7CgogIGNvbnN0IG91dHB1dEVsZW1lbnQgPSBkb2N1bWVudC5nZXRFbGVtZW50QnlJZChvdXRwdXRJZCk7CiAgb3V0cHV0RWxlbWVudC5pbm5lckhUTUwgPSAnJzsKCiAgY29uc3QgcGlja2VkUHJvbWlzZSA9IG5ldyBQcm9taXNlKChyZXNvbHZlKSA9PiB7CiAgICBpbnB1dEVsZW1lbnQuYWRkRXZlbnRMaXN0ZW5lcignY2hhbmdlJywgKGUpID0+IHsKICAgICAgcmVzb2x2ZShlLnRhcmdldC5maWxlcyk7CiAgICB9KTsKICB9KTsKCiAgY29uc3QgY2FuY2VsID0gZG9jdW1lbnQuY3JlYXRlRWxlbWVudCgnYnV0dG9uJyk7CiAgaW5wdXRFbGVtZW50LnBhcmVudEVsZW1lbnQuYXBwZW5kQ2hpbGQoY2FuY2VsKTsKICBjYW5jZWwudGV4dENvbnRlbnQgPSAnQ2FuY2VsIHVwbG9hZCc7CiAgY29uc3QgY2FuY2VsUHJvbWlzZSA9IG5ldyBQcm9taXNlKChyZXNvbHZlKSA9PiB7CiAgICBjYW5jZWwub25jbGljayA9ICgpID0+IHsKICAgICAgcmVzb2x2ZShudWxsKTsKICAgIH07CiAgfSk7CgogIC8vIENhbmNlbCB1cGxvYWQgaWYgdXNlciBoYXNuJ3QgcGlja2VkIGFueXRoaW5nIGluIHRpbWVvdXQuCiAgY29uc3QgdGltZW91dFByb21pc2UgPSBuZXcgUHJvbWlzZSgocmVzb2x2ZSkgPT4gewogICAgc2V0VGltZW91dCgoKSA9PiB7CiAgICAgIHJlc29sdmUobnVsbCk7CiAgICB9LCBGSUxFX0NIQU5HRV9USU1FT1VUX01TKTsKICB9KTsKCiAgLy8gV2FpdCBmb3IgdGhlIHVzZXIgdG8gcGljayB0aGUgZmlsZXMuCiAgY29uc3QgZmlsZXMgPSB5aWVsZCB7CiAgICBwcm9taXNlOiBQcm9taXNlLnJhY2UoW3BpY2tlZFByb21pc2UsIHRpbWVvdXRQcm9taXNlLCBjYW5jZWxQcm9taXNlXSksCiAgICByZXNwb25zZTogewogICAgICBhY3Rpb246ICdzdGFydGluZycsCiAgICB9CiAgfTsKCiAgaWYgKCFmaWxlcykgewogICAgcmV0dXJuIHsKICAgICAgcmVzcG9uc2U6IHsKICAgICAgICBhY3Rpb246ICdjb21wbGV0ZScsCiAgICAgIH0KICAgIH07CiAgfQoKICBjYW5jZWwucmVtb3ZlKCk7CgogIC8vIERpc2FibGUgdGhlIGlucHV0IGVsZW1lbnQgc2luY2UgZnVydGhlciBwaWNrcyBhcmUgbm90IGFsbG93ZWQuCiAgaW5wdXRFbGVtZW50LmRpc2FibGVkID0gdHJ1ZTsKCiAgZm9yIChjb25zdCBmaWxlIG9mIGZpbGVzKSB7CiAgICBjb25zdCBsaSA9IGRvY3VtZW50LmNyZWF0ZUVsZW1lbnQoJ2xpJyk7CiAgICBsaS5hcHBlbmQoc3BhbihmaWxlLm5hbWUsIHtmb250V2VpZ2h0OiAnYm9sZCd9KSk7CiAgICBsaS5hcHBlbmQoc3BhbigKICAgICAgICBgKCR7ZmlsZS50eXBlIHx8ICduL2EnfSkgLSAke2ZpbGUuc2l6ZX0gYnl0ZXMsIGAgKwogICAgICAgIGBsYXN0IG1vZGlmaWVkOiAkewogICAgICAgICAgICBmaWxlLmxhc3RNb2RpZmllZERhdGUgPyBmaWxlLmxhc3RNb2RpZmllZERhdGUudG9Mb2NhbGVEYXRlU3RyaW5nKCkgOgogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAnbi9hJ30gLSBgKSk7CiAgICBjb25zdCBwZXJjZW50ID0gc3BhbignMCUgZG9uZScpOwogICAgbGkuYXBwZW5kQ2hpbGQocGVyY2VudCk7CgogICAgb3V0cHV0RWxlbWVudC5hcHBlbmRDaGlsZChsaSk7CgogICAgY29uc3QgZmlsZURhdGFQcm9taXNlID0gbmV3IFByb21pc2UoKHJlc29sdmUpID0+IHsKICAgICAgY29uc3QgcmVhZGVyID0gbmV3IEZpbGVSZWFkZXIoKTsKICAgICAgcmVhZGVyLm9ubG9hZCA9IChlKSA9PiB7CiAgICAgICAgcmVzb2x2ZShlLnRhcmdldC5yZXN1bHQpOwogICAgICB9OwogICAgICByZWFkZXIucmVhZEFzQXJyYXlCdWZmZXIoZmlsZSk7CiAgICB9KTsKICAgIC8vIFdhaXQgZm9yIHRoZSBkYXRhIHRvIGJlIHJlYWR5LgogICAgbGV0IGZpbGVEYXRhID0geWllbGQgewogICAgICBwcm9taXNlOiBmaWxlRGF0YVByb21pc2UsCiAgICAgIHJlc3BvbnNlOiB7CiAgICAgICAgYWN0aW9uOiAnY29udGludWUnLAogICAgICB9CiAgICB9OwoKICAgIC8vIFVzZSBhIGNodW5rZWQgc2VuZGluZyB0byBhdm9pZCBtZXNzYWdlIHNpemUgbGltaXRzLiBTZWUgYi82MjExNTY2MC4KICAgIGxldCBwb3NpdGlvbiA9IDA7CiAgICB3aGlsZSAocG9zaXRpb24gPCBmaWxlRGF0YS5ieXRlTGVuZ3RoKSB7CiAgICAgIGNvbnN0IGxlbmd0aCA9IE1hdGgubWluKGZpbGVEYXRhLmJ5dGVMZW5ndGggLSBwb3NpdGlvbiwgTUFYX1BBWUxPQURfU0laRSk7CiAgICAgIGNvbnN0IGNodW5rID0gbmV3IFVpbnQ4QXJyYXkoZmlsZURhdGEsIHBvc2l0aW9uLCBsZW5ndGgpOwogICAgICBwb3NpdGlvbiArPSBsZW5ndGg7CgogICAgICBjb25zdCBiYXNlNjQgPSBidG9hKFN0cmluZy5mcm9tQ2hhckNvZGUuYXBwbHkobnVsbCwgY2h1bmspKTsKICAgICAgeWllbGQgewogICAgICAgIHJlc3BvbnNlOiB7CiAgICAgICAgICBhY3Rpb246ICdhcHBlbmQnLAogICAgICAgICAgZmlsZTogZmlsZS5uYW1lLAogICAgICAgICAgZGF0YTogYmFzZTY0LAogICAgICAgIH0sCiAgICAgIH07CiAgICAgIHBlcmNlbnQudGV4dENvbnRlbnQgPQogICAgICAgICAgYCR7TWF0aC5yb3VuZCgocG9zaXRpb24gLyBmaWxlRGF0YS5ieXRlTGVuZ3RoKSAqIDEwMCl9JSBkb25lYDsKICAgIH0KICB9CgogIC8vIEFsbCBkb25lLgogIHlpZWxkIHsKICAgIHJlc3BvbnNlOiB7CiAgICAgIGFjdGlvbjogJ2NvbXBsZXRlJywKICAgIH0KICB9Owp9CgpzY29wZS5nb29nbGUgPSBzY29wZS5nb29nbGUgfHwge307CnNjb3BlLmdvb2dsZS5jb2xhYiA9IHNjb3BlLmdvb2dsZS5jb2xhYiB8fCB7fTsKc2NvcGUuZ29vZ2xlLmNvbGFiLl9maWxlcyA9IHsKICBfdXBsb2FkRmlsZXMsCiAgX3VwbG9hZEZpbGVzQ29udGludWUsCn07Cn0pKHNlbGYpOwo=", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 91} outputId="e149212d-b186-4d99-80fd-238d4cfab17b"
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# + id="5NeqTxuNpLEj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="80fe10e2-f170-4ca5-8edc-bf979a758bd5"
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
import io
data = pd.read_csv(io.StringIO(uploaded['datasets_483_982_spam.csv'].decode('latin-1')))
del data['Unnamed: 2'],data['Unnamed: 3'],data['Unnamed: 4']
data.head()
# + id="4Dx7gv3-pYqA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="2d4d023e-fb47-4280-876b-a055d1429ad2"
sns.countplot(data.v1)
plt.xlabel('Label')
plt.title('Number of Spam and Ham')
plt.show()
spam,ham = data['v1'].value_counts()
print("Spam :",spam)
print("Ham :",ham)
# + id="GoanECZ6vtkt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="bb2b284a-b8ae-4260-a9cd-4238b639c4e3"
X = data.v2
Y = data.v1
print(Y.shape)
Y
# + id="PEww7fLPqJET" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="6c43d119-4d90-42fc-81d6-8f9f7de32e1b"
#Preprocessing data and splite
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
obj_le = LabelEncoder()
y = obj_le.fit_transform(Y)
y = y.reshape(-1,1)
print(y.shape)
X_train,X_test,y_train,y_test = train_test_split = train_test_split(X,y,test_size=0.15)
X_train.head()
# + id="wNAryQV9xoZd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="4322497e-1f4f-47f2-966e-5b7e0ceb7929"
#Text Preprocessing of text data
from keras.preprocessing.text import Tokenizer
from keras_preprocessing.sequence import pad_sequences
max_words = 1000
max_sent_len= 150
emb_dim = 50
tok = Tokenizer(num_words=max_words)
tok.fit_on_texts(X_train)
#print(tok.word_index)
sequences = tok.texts_to_sequences(X_train)
embedded_matrix = pad_sequences(sequences,padding='pre',maxlen=max_sent_len)
embedded_matrix
# + id="hFcsJaiM6CjC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 425} outputId="4e0820dd-a16c-454d-daf9-ddd2a3eb58e2"
#Working on DL Keras
from keras.models import Model
from keras.layers import LSTM,Dense,Dropout,Input,Activation,Embedding
from keras.optimizers import RMSprop
from keras.utils import to_categorical
from keras.callbacks import EarlyStopping
def RNN():
inputs = Input(name='inputs',shape = [max_sent_len])
layers = Embedding(max_words,emb_dim,input_length=max_sent_len)(inputs)
layers = LSTM(64)(layers)
layers = Dense(256,name='FC1')(layers)
layers = Activation('relu')(layers)
layers = Dropout(0.5)(layers)
layers = Dense(1,name='out_layer')(layers)
layers = Activation('sigmoid')(layers)
obj_model = Model(inputs=inputs,outputs=layers)
return obj_model
obj_model = RNN()
obj_model.summary()
obj_model.compile(loss='binary_crossentropy',optimizer=RMSprop(),metrics=['accuracy'])
# + id="4r9XeP-pZDoQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 479} outputId="95aad678-e037-4b3a-c858-c4a38ba4b1fc"
obj_model.fit(embedded_matrix,y_train,epochs=10,batch_size=128,validation_split=0.2,callbacks=[EarlyStopping(monitor='var_loss',min_delta=0.0001)])
# + id="4_2srkE1dX8G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="6c0c4764-1d81-43c3-925e-72351edf329b"
#tok.fit_on_texts(X_test)
X_test_sequences = tok.texts_to_sequences(X_test)
X_test_embedded_matrix = pad_sequences(X_test_sequences,padding='pre',maxlen=max_sent_len)
X_test_embedded_matrix
# + id="kmEXAf8FfeTY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="d9bd841b-fe1a-48f2-c62e-fec3da4dc7c1"
accurcy = obj_model.evaluate(X_test_embedded_matrix,y_test)
print('Test set\n Loss: {:0.3f}\n Accuracy: {:0.3f}'.format(accurcy[0],accurcy[1]))
# + id="omSq2-v9fn73" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c44adcee-7905-4dd3-d7b0-6b74fe1f3bb1"
Testing_text = ["win price for watching this ..:)"]
test_text = tok.texts_to_sequences(Testing_text)
text_embed_matrix = pad_sequences(test_text,maxlen=max_sent_len)
preds = obj_model.predict(text_embed_matrix)
print(preds)
# + id="gFLiDcBtiTvM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6575f6b4-f220-4be6-d0ce-09edc84dfefb"
Testing_text = ["Oh k...i'm watching here:)"]
test_text = tok.texts_to_sequences(Testing_text)
text_embed_matrix = pad_sequences(test_text,maxlen=max_sent_len)
preds = obj_model.predict(text_embed_matrix)
print(preds)
# + id="iaFQUPw3it6T" colab_type="code" colab={}
d try all possible pairs of values for $C$ and $\sigma$ (e.g., $C = 0.3$ and $\sigma = 0.1$). For example, if you try each of the 8 values listed above for $C$ and for $\sigma^2$, you would end up training and evaluating (on the cross validation set) a total of $8^2 = 64$ different models. After you have determined the best $C$ and $\sigma$ parameters to use, you should modify the code in `dataset3Params`, filling in the best parameters you found. For our best parameters, the SVM returned a decision boundary shown in the figure below.
#
# 
#
# <div class="alert alert-block alert-warning">
# **Implementation Tip:** When implementing cross validation to select the best $C$ and $\sigma$ parameter to use, you need to evaluate the error on the cross validation set. Recall that for classification, the error is defined as the fraction of the cross validation examples that were classified incorrectly. In `numpy`, you can compute this error using `np.mean(predictions != yval)`, where `predictions` is a vector containing all the predictions from the SVM, and `yval` are the true labels from the cross validation set. You can use the `utils.svmPredict` function to generate the predictions for the cross validation set.
# </div>
# <a id="dataset3Params"></a>
def dataset3Params(X, y, Xval, yval):
"""
Returns your choice of C and sigma for Part 3 of the exercise
where you select the optimal (C, sigma) learning parameters to use for SVM
with RBF kernel.
Parameters
----------
X : array_like
(m x n) matrix of training data where m is number of training examples, and
n is the number of features.
y : array_like
(m, ) vector of labels for ther training data.
Xval : array_like
(mv x n) matrix of validation data where mv is the number of validation examples
and n is the number of features
yval : array_like
(mv, ) vector of labels for the validation data.
Returns
-------
C, sigma : float, float
The best performing values for the regularization parameter C and
RBF parameter sigma.
Instructions
------------
Fill in this function to return the optimal C and sigma learning
parameters found using the cross validation set.
You can use `svmPredict` to predict the labels on the cross
validation set. For example,
predictions = svmPredict(model, Xval)
will return the predictions on the cross validation set.
Note
----
You can compute the prediction error using
np.mean(predictions != yval)
"""
# You need to return the following variables correctly.
C = 1
sigma = 0.3
# ====================== YOUR CODE HERE ======================
possible = [0.01, 0.03, 0.1, 0.3, 1, 3, 10, 30]
smallestError = 10000000000
for Ctry in possible:
for sigmatry in possible:
error = np.mean(utils.svmPredict(utils.svmTrain(X, y, Ctry, gaussianKernel, args=(sigmatry,)), Xval) != yval)
if error < smallestError:
smallestError = error
C = Ctry
sigma = sigmatry
# ============================================================
return C, sigma
# The provided code in the next cell trains the SVM classifier using the training set $(X, y)$ using parameters loaded from `dataset3Params`. Note that this might take a few minutes to execute.
# +
# Try different SVM Parameters here
C, sigma = dataset3Params(X, y, Xval, yval)
# Train the SVM
# model = utils.svmTrain(X, y, C, lambda x1, x2: gaussianKernel(x1, x2, sigma))
model = utils.svmTrain(X, y, C, gaussianKernel, args=(sigma,))
utils.visualizeBoundary(X, y, model)
print(C, sigma)
# -
# One you have computed the values `C` and `sigma` in the cell above, we will submit those values for grading.
#
# *You should now submit your solutions.*
grader[2] = lambda : (C, sigma)
grader.grade()
# <a id="section3"></a>
# ## 2 Spam Classification
#
# Many email services today provide spam filters that are able to classify emails into spam and non-spam email with high accuracy. In this part of the exercise, you will use SVMs to build your own spam filter.
#
# You will be training a classifier to classify whether a given email, $x$, is spam ($y = 1$) or non-spam ($y = 0$). In particular, you need to convert each email into a feature vector $x \in \mathbb{R}^n$ . The following parts of the exercise will walk you through how such a feature vector can be constructed from an email.
#
# The dataset included for this exercise is based on a a subset of the [SpamAssassin Public Corpus](http://spamassassin.apache.org/old/publiccorpus/). For the purpose of this exercise, you will only be using the body of the email (excluding the email headers).
# ### 2.1 Preprocessing Emails
#
# Before starting on a machine learning task, it is usually insightful to take a look at examples from the dataset. The figure below shows a sample email that contains a URL, an email address (at the end), numbers, and dollar
# amounts.
#
# <img src="Figures/email.png" width="700px" />
#
# While many emails would contain similar types of entities (e.g., numbers, other URLs, or other email addresses), the specific entities (e.g., the specific URL or specific dollar amount) will be different in almost every
# email. Therefore, one method often employed in processing emails is to “normalize” these values, so that all URLs are treated the same, all numbers are treated the same, etc. For example, we could replace each URL in the
# email with the unique string “httpaddr” to indicate that a URL was present.
#
# This has the effect of letting the spam classifier make a classification decision based on whether any URL was present, rather than whether a specific URL was present. This typically improves the performance of a spam classifier, since spammers often randomize the URLs, and thus the odds of seeing any particular URL again in a new piece of spam is very small.
#
# In the function `processEmail` below, we have implemented the following email preprocessing and normalization steps:
#
# - **Lower-casing**: The entire email is converted into lower case, so that captialization is ignored (e.g., IndIcaTE is treated the same as Indicate).
#
# - **Stripping HTML**: All HTML tags are removed from the emails. Many emails often come with HTML formatting; we remove all the HTML tags, so that only the content remains.
#
# - **Normalizing URLs**: All URLs are replaced with the text “httpaddr”.
#
# - **Normalizing Email Addresses**: All email addresses are replaced with the text “emailaddr”.
#
# - **Normalizing Numbers**: All numbers are replaced with the text “number”.
#
# - **Normalizing Dollars**: All dollar signs ($) are replaced with the text “dollar”.
#
# - **Word Stemming**: Words are reduced to their stemmed form. For example, “discount”, “discounts”, “discounted” and “discounting” are all replaced with “discount”. Sometimes, the Stemmer actually strips off additional characters from the end, so “include”, “includes”, “included”, and “including” are all replaced with “includ”.
#
# - **Removal of non-words**: Non-words and punctuation have been removed. All white spaces (tabs, newlines, spaces) have all been trimmed to a single space character.
#
# The result of these preprocessing steps is shown in the figure below.
#
# <img src="Figures/email_cleaned.png" alt="email cleaned" style="width: 600px;"/>
#
# While preprocessing has left word fragments and non-words, this form turns out to be much easier to work with for performing feature extraction.
# #### 2.1.1 Vocabulary List
#
# After preprocessing the emails, we have a list of words for each email. The next step is to choose which words we would like to use in our classifier and which we would want to leave out.
#
# For this exercise, we have chosen only the most frequently occuring words as our set of words considered (the vocabulary list). Since words that occur rarely in the training set are only in a few emails, they might cause the
# model to overfit our training set. The complete vocabulary list is in the file `vocab.txt` (inside the `Data` directory for this exercise) and also shown in the figure below.
#
# <img src="Figures/vocab.png" alt="Vocab" width="150px" />
#
# Our vocabulary list was selected by choosing all words which occur at least a 100 times in the spam corpus,
# resulting in a list of 1899 words. In practice, a vocabulary list with about 10,000 to 50,000 words is often used.
# Given the vocabulary list, we can now map each word in the preprocessed emails into a list of word indices that contains the index of the word in the vocabulary dictionary. The figure below shows the mapping for the sample email. Specifically, in the sample email, the word “anyone” was first normalized to “anyon” and then mapped onto the index 86 in the vocabulary list.
#
# <img src="Figures/word_indices.png" alt="word indices" width="200px" />
#
# Your task now is to complete the code in the function `processEmail` to perform this mapping. In the code, you are given a string `word` which is a single word from the processed email. You should look up the word in the vocabulary list `vocabList`. If the word exists in the list, you should add the index of the word into the `word_indices` variable. If the word does not exist, and is therefore not in the vocabulary, you can skip the word.
#
# <div class="alert alert-block alert-warning">
# **python tip**: In python, you can find the index of the first occurence of an item in `list` using the `index` attribute. In the provided code for `processEmail`, `vocabList` is a python list containing the words in the vocabulary. To find the index of a word, we can use `vocabList.index(word)` which would return a number indicating the index of the word within the list. If the word does not exist in the list, a `ValueError` exception is raised. In python, we can use the `try/except` statement to catch exceptions which we do not want to stop the program from running. You can think of the `try/except` statement to be the same as an `if/else` statement, but it asks for forgiveness rather than permission.
#
# An example would be:
# <br>
#
# ```
# try:
# do stuff here
# except ValueError:
# pass
# # do nothing (forgive me) if a ValueError exception occured within the try statement
# ```
# </div>
# <a id="processEmail"></a>
def processEmail(email_contents, verbose=True):
"""
Preprocesses the body of an email and returns a list of indices
of the words contained in the email.
Parameters
----------
email_contents : str
A string containing one email.
verbose : bool
If True, print the resulting email after processing.
Returns
-------
word_indices : list
A list of integers containing the index of each word in the
email which is also present in the vocabulary.
Instructions
------------
Fill in this function to add the index of word to word_indices
if it is in the vocabulary. At this point of the code, you have
a stemmed word from the email in the variable word.
You should look up word in the vocabulary list (vocabList).
If a match exists, you should add the index of the word to the word_indices
list. Concretely, if word = 'action', then you should
look up the vocabulary list to find where in vocabList
'action' appears. For example, if vocabList[18] =
'action', then, you should add 18 to the word_indices
vector (e.g., word_indices.append(18)).
Notes
-----
- vocabList[idx] returns a the word with index idx in the vocabulary list.
- vocabList.index(word) return index of word `word` in the vocabulary list.
(A ValueError exception is raised if the word does not exist.)
"""
# Load Vocabulary
vocabList = utils.getVocabList()
# Init return value
word_indices = []
# ========================== Preprocess Email ===========================
# Find the Headers ( \n\n and remove )
# Uncomment the following lines if you are working with raw emails with the
# full headers
# hdrstart = email_contents.find(chr(10) + chr(10))
# email_contents = email_contents[hdrstart:]
# Lower case
email_contents = email_contents.lower()
# Strip all HTML
# Looks for any expression that starts with < and ends with > and replace
# and does not have any < or > in the tag it with a space
email_contents =re.compile('<[^<>]+>').sub(' ', email_contents)
# Handle Numbers
# Look for one or more characters between 0-9
email_contents = re.compile('[0-9]+').sub(' number ', email_contents)
# Handle URLS
# Look for strings starting with http:// or https://
email_contents = re.compile('(http|https)://[^\s]*').sub(' httpaddr ', email_contents)
# Handle Email Addresses
# Look for strings with @ in the middle
email_contents = re.compile('[^\s]+@[^\s]+').sub(' emailaddr ', email_contents)
# Handle $ sign
email_contents = re.compile('[$]+').sub(' dollar ', email_contents)
# get rid of any punctuation
email_contents = re.split('[ @$/#.-:&*+=\[\]?!(){},''">_<;%\n\r]', email_contents)
# remove any empty word string
email_contents = [word for word in email_contents if len(word) > 0]
# Stem the email contents word by word
stemmer = utils.PorterStemmer()
processed_email = []
for word in email_contents:
# Remove any remaining non alphanumeric characters in word
word = re.compile('[^a-zA-Z0-9]').sub('', word).strip()
word = stemmer.stem(word)
processed_email.append(word)
if len(word) < 1:
continue
# Look up the word in the dictionary and add to word_indices if found
# ====================== YOUR CODE HERE ======================
try:
word_indices.append(vocabList.index(word))
except ValueError:
pass
# =============================================================
if verbose:
print('----------------')
print('Processed email:')
print('----------------')
print(' '.join(processed_email))
return word_indices
# Once you have implemented `processEmail`, the following cell will run your code on the email sample and you should see an output of the processed email and the indices list mapping.
# +
# To use an SVM to classify emails into Spam v.s. Non-Spam, you first need
# to convert each email into a vector of features. In this part, you will
# implement the preprocessing steps for each email. You should
# complete the code in processEmail.m to produce a word indices vector
# for a given email.
# Extract Features
with open(os.path.join('Data', 'emailSample1.txt')) as fid:
file_contents = fid.read()
word_indices = processEmail(file_contents)
#Print Stats
print('-------------')
print('Word Indices:')
print('-------------')
print(word_indices)
# -
# *You should now submit your solutions.*
grader[3] = processEmail
grader.grade()
# <a id="section4"></a>
# ### 2.2 Extracting Features from Emails
#
# You will now implement the feature extraction that converts each email into a vector in $\mathbb{R}^n$. For this exercise, you will be using n = # words in vocabulary list. Specifically, the feature $x_i \in \{0, 1\}$ for an email corresponds to whether the $i^{th}$ word in the dictionary occurs in the email. That is, $x_i = 1$ if the $i^{th}$ word is in the email and $x_i = 0$ if the $i^{th}$ word is not present in the email.
#
# Thus, for a typical email, this feature would look like:
#
# $$ x = \begin{bmatrix}
# 0 & \dots & 1 & 0 & \dots & 1 & 0 & \dots & 0
# \end{bmatrix}^T \in \mathbb{R}^n
# $$
#
# You should now complete the code in the function `emailFeatures` to generate a feature vector for an email, given the `word_indices`.
# <a id="emailFeatures"></a>
def emailFeatures(word_indices):
"""
Takes in a word_indices vector and produces a feature vector from the word indices.
Parameters
----------
word_indices : list
A list of word indices from the vocabulary list.
Returns
-------
x : list
The computed feature vector.
Instructions
------------
Fill in this function to return a feature vector for the
given email (word_indices). To help make it easier to process
the emails, we have have already pre-processed each email and converted
each word in the email into an index in a fixed dictionary (of 1899 words).
The variable `word_indices` contains the list of indices of the words
which occur in one email.
Concretely, if an email has the text:
The quick brown fox jumped over the lazy dog.
Then, the word_indices vector for this text might look like:
60 100 33 44 10 53 60 58 5
where, we have mapped each word onto a number, for example:
the -- 60
quick -- 100
...
Note
----
The above numbers are just an example and are not the actual mappings.
Your task is take one such `word_indices` vector and construct
a binary feature vector that indicates whether a particular
word occurs in the email. That is, x[i] = 1 when word i
is present in the email. Concretely, if the word 'the' (say,
index 60) appears in the email, then x[60] = 1. The feature
vector should look like:
x = [ 0 0 0 0 1 0 0 0 ... 0 0 0 0 1 ... 0 0 0 1 0 ..]
"""
# Total number of words in the dictionary
n = 1899
# You need to return the following variables correctly.
x = np.zeros(n)
# ===================== YOUR CODE HERE ======================
for wordIndex in word_indices:
x[wordIndex] = 1
# ===========================================================
return x
# Once you have implemented `emailFeatures`, the next cell will run your code on the email sample. You should see that the feature vector had length 1899 and 45 non-zero entries.
# +
# Extract Features
with open(os.path.join('Data', 'emailSample1.txt')) as fid:
file_contents = fid.read()
word_indices = processEmail(file_contents)
features = emailFeatures(word_indices)
# Print Stats
print('\nLength of feature vector: %d' % len(features))
print('Number of non-zero entries: %d' % sum(features > 0))
# -
# *You should now submit your solutions.*
grader[4] = emailFeatures
grader.grade()
# ### 2.3 Training SVM for Spam Classification
#
# In the following section we will load a preprocessed training dataset that will be used to train a SVM classifier. The file `spamTrain.mat` (within the `Data` folder for this exercise) contains 4000 training examples of spam and non-spam email, while `spamTest.mat` contains 1000 test examples. Each
# original email was processed using the `processEmail` and `emailFeatures` functions and converted into a vector $x^{(i)} \in \mathbb{R}^{1899}$.
#
# After loading the dataset, the next cell proceed to train a linear SVM to classify between spam ($y = 1$) and non-spam ($y = 0$) emails. Once the training completes, you should see that the classifier gets a training accuracy of about 99.8% and a test accuracy of about 98.5%.
# +
# Load the Spam Email dataset
# You will have X, y in your environment
data = loadmat(os.path.join('Data', 'spamTrain.mat'))
X, y= data['X'].astype(float), data['y'][:, 0]
print('Training Linear SVM (Spam Classification)')
print('This may take 1 to 2 minutes ...\n')
C = 0.1
model = utils.svmTrain(X, y, C, utils.linearKernel)
# +
# Compute the training accuracy
p = utils.svmPredict(model, X)
print('Training Accuracy: %.2f' % (np.mean(p == y) * 100))
# -
# Execute the following cell to load the test set and compute the test accuracy.
# +
# Load the test dataset
# You will have Xtest, ytest in your environment
data = loadmat(os.path.join('Data', 'spamTest.mat'))
Xtest, ytest = data['Xtest'].astype(float), data['ytest'][:, 0]
print('Evaluating the trained Linear SVM on a test set ...')
p = utils.svmPredict(model, Xtest)
print('Test Accuracy: %.2f' % (np.mean(p == ytest) * 100))
# -
# ### 2.4 Top Predictors for Spam
#
# To better understand how the spam classifier works, we can inspect the parameters to see which words the classifier thinks are the most predictive of spam. The next cell finds the parameters with the largest positive values in the classifier and displays the corresponding words similar to the ones shown in the figure below.
#
# <div style="border-style: solid; border-width: 1px; margin: 10px 10px 10px 10px; padding: 10px 10px 10px 10px">
# our click remov guarante visit basenumb dollar pleas price will nbsp most lo ga hour
# </div>
#
# Thus, if an email contains words such as “guarantee”, “remove”, “dollar”, and “price” (the top predictors shown in the figure), it is likely to be classified as spam.
#
# Since the model we are training is a linear SVM, we can inspect the weights learned by the model to understand better how it is determining whether an email is spam or not. The following code finds the words with the highest weights in the classifier. Informally, the classifier 'thinks' that these words are the most likely indicators of spam.
# +
# Sort the weights and obtin the vocabulary list
# NOTE some words have the same weights,
# so their order might be different than in the text above
idx = np.argsort(model['w'])
top_idx = idx[-15:][::-1]
vocabList = utils.getVocabList()
print('Top predictors of spam:')
print('%-15s %-15s' % ('word', 'weight'))
print('----' + ' '*12 + '------')
for word, w in zip(np.array(vocabList)[top_idx], model['w'][top_idx]):
print('%-15s %0.2f' % (word, w))
# -
# ### 2.5 Optional (ungraded) exercise: Try your own emails
#
# Now that you have trained a spam classifier, you can start trying it out on your own emails. In the starter code, we have included two email examples (`emailSample1.txt` and `emailSample2.txt`) and two spam examples (`spamSample1.txt` and `spamSample2.txt`). The next cell runs the spam classifier over the first spam example and classifies it using the learned SVM. You should now try the other examples we have provided and see if the classifier gets them right. You can also try your own emails by replacing the examples (plain text files) with your own emails.
#
# *You do not need to submit any solutions for this optional (ungraded) exercise.*
# +
filename = os.path.join('Data', 'emailSample1.txt')
with open(filename) as fid:
file_contents = fid.read()
word_indices = processEmail(file_contents, verbose=False)
x = emailFeatures(word_indices)
p = utils.svmPredict(model, x)
print('\nProcessed %s\nSpam Classification: %s' % (filename, 'spam' if p else 'not spam'))
# -
# ### 2.6 Optional (ungraded) exercise: Build your own dataset
#
# In this exercise, we provided a preprocessed training set and test set. These datasets were created using the same functions (`processEmail` and `emailFeatures`) that you now have completed. For this optional (ungraded) exercise, you will build your own dataset using the original emails from the SpamAssassin Public Corpus.
#
# Your task in this optional (ungraded) exercise is to download the original
# files from the public corpus and extract them. After extracting them, you should run the `processEmail` and `emailFeatures` functions on each email to extract a feature vector from each email. This will allow you to build a dataset `X`, `y` of examples. You should then randomly divide up the dataset into a training set, a cross validation set and a test set.
#
# While you are building your own dataset, we also encourage you to try building your own vocabulary list (by selecting the high frequency words that occur in the dataset) and adding any additional features that you think
# might be useful. Finally, we also suggest trying to use highly optimized SVM toolboxes such as [`LIBSVM`](https://www.csie.ntu.edu.tw/~cjlin/libsvm/) or [`scikit-learn`](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.svm).
#
# *You do not need to submit any solutions for this optional (ungraded) exercise.*
| 37,430 |
/TASK_18.ipynb | c77b318df84a16fa33eb12df24afc7f5ddebf350 | [] | no_license | Alen-8080/Learn.py | https://github.com/Alen-8080/Learn.py | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 7,528 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class mylist:
l=[]
def __init__(s,n):
#n=int(input("How many data in the list?"))
temp=[]
for i in range(1,n+1):
x=int(input("Enter the data:"))
temp.append(x)
s.l=temp
def sum(s):
sum=0
for i in s.l:
sum=sum+i
return sum
def count(s):
count=0
for i in s.l:
count=count+1
return count
def average(s):
sum=0
count=0
for i in s.l:
sum=sum+i
count=count+1
average=sum/count
return average
def interchange(s):
temp=s.l[0]
s.l[0]=s.l[-1]
s.l[-1]=temp
return s.l
def max(s):
max=s.l[0]
for i in s.l:
if(i>max):
max=i
return max
def min(s):
min=s.l[0]
for i in s.l:
if(i<min):
min=i
return min
def isitem(s,item):
for x in s.l:
if(x==item):
return True
return False
# -
markdown] id="QuoucvjxBPmX"
# 2. Create a DataFrame by passing a random numpy array,with a datetie index and labelled columns.and
# - Dislplay the index from the 1 to 3 date and the columns A and B of respective data.
# - Display the datas at the 3rd index.
# - Display datas from 2nd to 5th.
# - Display the datas from the index 1,2,4 and the column 0 and 2.
# + id="qV_wzsp1DUKI" outputId="1c32649f-6981-4a70-bb15-92f6cc0340e4" colab={"base_uri": "https://localhost:8080/"}
#please provide your answer below thins line.
import numpy as np
import pandas as pd
dates = pd.date_range('20210829', periods=6)
df = pd.DataFrame(np.random.randn(6, 4), index=dates, columns=list('ABCD'))
print("DataFrame:")
print(df)
print("\n")
print("index from 1-3 dates and A and B columns :")
print(df.iloc[1:4,0:2])
print("\n")
print("Data at 3rd index:")
print(df.iloc[3,0:4])
print("\n")
print("Datas from 2-5th:")
print(df.iloc[2:6,0:4])
print("\n")
print("Datas from the index 1,2,4 and the column 0 and 2: ")
print(df.iloc[0:6:2,0:3:2])
| 2,421 |
/Tasks/Task6_DecisionTreeML.ipynb | 1b272de0fc46e48f0d54e44aee0e89b036f06bb9 | [] | no_license | DIRGH712/ML-Algorithms-Basics | https://github.com/DIRGH712/ML-Algorithms-Basics | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 112,470 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="QfX8RBRFIAtb"
#
# <br></br>
# <br></br>
# # <center>Data Science and Business Analytics
# ## ✨Task-6: Prediction using Decision Tree Algorithm
#
# ## Name : DIRGH PATEL
# <br></br>
# <br></br>
#
# + id="JOUWz3tHD3d6"
import sklearn.datasets as datasets
import pandas as pd
# -
import warnings
warnings.filterwarnings('ignore')
from IPython.display import HTML
HTML('''<script>
code_show_err=false;
function code_toggle_err() {
if (code_show_err){
$('div.output_stderr').hide();
} else {
$('div.output_stderr').show();
}
code_show_err = !code_show_err
}
$( document ).ready(code_toggle_err);
</script>
To toggle on/off output_stderr, click <a href="javascript:code_toggle_err()">here</a>.''')
# + id="p784TCaAusVG"
iris = datasets.load_iris()
df = pd.DataFrame(data = iris.data, columns = iris.feature_names)
y = iris.target
# + colab={"base_uri": "https://localhost:8080/"} id="4BsnNclXwj_i" outputId="e44ac820-aeb2-4c35-c4a8-6e72637911ee"
from sklearn.tree import DecisionTreeClassifier
dtree = DecisionTreeClassifier()
dtree.fit(df,y)
# + id="eI_mu_6mzZ34"
from sklearn import tree
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 575} id="hzM-4LIDznDE" outputId="c21aff12-1d6a-4a66-8690-099301119ae4"
plt.figure(figsize=(15,10))
tree.plot_tree(dtree, feature_names=iris.feature_names, class_names=iris.target_names, filled=True)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="eK30BH2x5JlG" outputId="ad444572-c5c6-4cc3-af1f-ab4b93cc3cec"
iris
# -
# <br></br>
# # Completed
| 1,841 |
/Emotion Classification.ipynb | 28eced4197e46fce9a0603a99d8d6334fba8786f | [] | no_license | ajay-del-bot/Emotion-Detection | https://github.com/ajay-del-bot/Emotion-Detection | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 112,228 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# ## Load the dataset
# +
train = pd.read_csv('data/train.txt',
header=None, sep=';',
names=['Text', 'Emotion'],
encoding='utf-8')
test = pd.read_csv('data/test.txt',
header=None, sep=';',
names=['Text', 'Emotion'],
encoding='utf-8')
val = pd.read_csv('data/val.txt',
header=None, sep=';',
names=['Text', 'Emotion'],
encoding='utf-8')
# -
train.head()
# ## Encoding
emotions_to_labels = {'anger': 0, 'love': 1, 'fear': 2,
'joy': 3, 'sadness': 4,'surprise': 5}
train['Label'] = train['Emotion'].replace(emotions_to_labels)
test['Label'] = test['Emotion'].replace(emotions_to_labels)
val['Label'] = val['Emotion'].replace(emotions_to_labels)
train.head()
train.isnull().sum()
train.Emotion.value_counts()
# ## Analyzing the Data
sns.countplot(x='Emotion', data=train)
sns.countplot(x='Emotion', data=test)
sns.countplot(x='Emotion', data=val)
# ## Text Preprocessing
# +
import nltk
nltk.download('punkt')
nltk.download('stopwords')
from nltk.corpus import stopwords
STOPWORDS = set(stopwords.words('english'))
from nltk.tokenize import word_tokenize
# -
#Accepts a line and returns preprocessed text
def text_preprocess(text, stop_words=False):
#remove non words from text
text = re.sub(r'\W+', ' ', text).lower()
tokens = word_tokenize(text)
if stop_words:
tokens = [token for token in tokens if token not in STOPWORDS]
return tokens
# +
print('Before: ')
print(train.head())
x_train = [text_preprocess(t, stop_words=True) for t in train['Text']]
y_train = train['Label'].values
print('\nAfter:')
for x in list(zip(x_train[:5], y_train[:5])):
print(x)
# +
x_test = [text_preprocess(t, stop_words=True) for t in test['Text']]
y_test = test['Label'].values
x_val = [text_preprocess(t, stop_words=True) for t in val['Text']]
y_val = val['Label'].values
# -
# ## Word Embeddings
# load pre-trained model
import gensim.downloader as api
model_wiki = api.load('fasttext-wiki-news-subwords-300')
# +
from gensim.models import Word2Vec
model_w2v = Word2Vec(x_train + x_test + x_val, # data for model to train on
vector_size = 300, # embedding vector size
min_count = 2).wv
# -
# ## Vectorization
from tensorflow.keras.preprocessing.text import Tokenizer
DICT_SIZE = 15000
#max size of columns
tokenizer = Tokenizer(num_words=DICT_SIZE)
total = x_train + x_train + x_val
tokenizer.fit_on_texts(total)
list(tokenizer.word_index.items())[:5]
len(tokenizer.word_index)
# +
#max length of sentence
x_train_max_len = max([len(i) for i in x_train])
x_test_max_len = max([len(i) for i in x_test])
x_val_max_len = max([len(i) for i in x_val])
MAX_LEN = max(x_train_max_len, x_test_max_len, x_val_max_len)
# -
MAX_LEN
from tensorflow.keras.preprocessing.sequence import pad_sequences
# +
#change size of vectors to MAX_LEN and pad indexes
X_train = tokenizer.texts_to_sequences(x_train)
X_train_pad = pad_sequences(X_train, maxlen=MAX_LEN)
X_test = tokenizer.texts_to_sequences(x_test)
X_test_pad = pad_sequences(X_test, maxlen=MAX_LEN)
X_val = tokenizer.texts_to_sequences(x_val)
X_val_pad = pad_sequences(X_val, maxlen=MAX_LEN)
# -
X_train[0]
X_train_pad[0]
# ## Embedding Matrix
def create_weight_matrix(model, second_model=False):
vector_size = model.get_vector('like').shape[0]
w_matrix = np.zeros((DICT_SIZE, vector_size))
skipped_words = []
for word, index in tokenizer.word_index.items():
if index < DICT_SIZE:
if word in model.key_to_index:
w_matrix[index] = model.get_vector(word)
else:
if second_model:
if word in second_model.key_to_index:
w_matrix[index] = second_model.get_vector(word)
else:
skipped_words.append(word)
else:
skipped_words.append(word)
print(f'{len(skipped_words)} words were skipped. Some of them:')
print(skipped_words[:50])
return w_matrix
weight_matrix = create_weight_matrix(model_wiki, model_w2v)
weight_matrix.shape
# ## Creating the layers
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding, LSTM, Bidirectional, Dense, Dropout
from tensorflow.keras.optimizers import Adam
#early stopping
from tensorflow.keras.callbacks import EarlyStopping
stop = EarlyStopping(monitor='val_loss',
mode='min', verbose=1, patience=3)
# initialize sequential model
model = Sequential()
model.add(Embedding(input_dim = DICT_SIZE, # the whole vocabulary size
output_dim = weight_matrix.shape[1], # vector space dimension
input_length = X_train_pad.shape[1], # max_len of text sequence
weights=[weight_matrix], # assign the embedding weight with embedding marix
trainable=False)) # set the weight to be not trainable (static)
model.add(Bidirectional(LSTM(128, return_sequences=True)))
model.add(Dropout(0.2))
model.add(Bidirectional(LSTM(256, return_sequences=True)))
model.add(Dropout(0.2))
model.add(Bidirectional(LSTM(128, return_sequences=False)))
model.add(Dense(6, activation = 'softmax'))
model.compile(loss='sparse_categorical_crossentropy',
optimizer='adam', metrics=['accuracy'])
history = model.fit(X_train_pad, y_train,
validation_data = (X_val_pad, y_val),
batch_size = 8,
epochs = 3, callbacks=[stop])
print()
def plot_history(history):
loss = history.history['loss']
accuracy = history.history['acc']
val_loss = history.history['val_loss']
val_accuracy = history.history['val_acc']
x = range(1, len(loss) + 1)
plt.figure(figsize=(12, 5))
plt.subplot(1, 2, 1)
plt.plot(x, accuracy, label='Training acc', color='#707bfb')
plt.plot(x, val_accuracy, label='Validation acc', color='#fbcbff')
plt.title('Training and validation accuracy')
plt.grid(True)
plt.legend()
plt.subplot(1, 2, 2)
plt.plot(x, loss, label='Training loss', color='#707bfb')
plt.plot(x, val_loss, label='Validation loss', color='#fbcbff')
plt.title('Training and validation loss')
plt.grid(True)
plt.legend()
plot_history(history)
model.evaluate(X_test_pad, y_test)
y_pred = np.argmax(model.predict(X_test_pad), axis=1)
from sklearn import metrics
print(metrics.classification_report(y_test, y_pred))
model.save('my_model.h5')
import tensorflow as tf
model = tf.keras.models.load_model('my_model.h5')
model.summary()
def plot_confusion_matrix(matrix, fmt=''):
plt.figure(figsize=(6, 5))
sns.heatmap(matrix, annot=True,
cmap='inferno',
fmt=fmt,
xticklabels=emotions_to_labels.keys(),
yticklabels=emotions_to_labels.keys())
plt.ylabel('True labels')
plt.xlabel('Predicted labels')
plt.show()
matrix = metrics.confusion_matrix(y_test, y_pred)
plot_confusion_matrix(matrix)
# +
# create new confusion matrix
# where values are normed by row
matrix_new = np.zeros(matrix.shape)
for row in range(len(matrix)):
sum = np.sum(matrix[row])
for element in range(len(matrix[row])):
matrix_new[row][element] = matrix[row][element] / sum
plot_confusion_matrix(matrix_new, fmt='.2')
# -
# ## Test On Custom Data
def predict(texts):
texts_prepr = [text_preprocess(t) for t in texts]
sequences = tokenizer.texts_to_sequences(texts_prepr)
pad = pad_sequences(sequences, maxlen=MAX_LEN)
predictions = model.predict(pad)
labels = np.argmax(predictions, axis=1)
for i, lbl in enumerate(labels):
print(f'\'{texts[i]}\' --> {labels_to_emotions[lbl]}')
labels_to_emotions = {j:i for i,j in emotions_to_labels.items()}
# +
test_texts = ['I am so happy', 'The man felt lonely', 'The guests felt satisfied']
predict(test_texts)
# -
model.save('my_model.h5')
| 8,517 |
/Livemint.ipynb | 6e526f2077c87650244fc9557a15c71922fc21d8 | [] | no_license | sroy263/Data-Visualization | https://github.com/sroy263/Data-Visualization | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 3,645 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import copy
# #### Initialising all the parametric values
# +
theta=0.01
delta =1
grid = np.random.rand(4,4) # random initialisation of the grid world
grid[0,0]=grid[3,3]=0
r=-1 # reward parameter
gamma=0.999 # discounting factor
pi = np.zeros((4,4,4)) # policy matrix
directions= ['UP','DOWN','LEFT','RIGHT']
dir_values=[-1,1,-1,1]
dir_l=list(zip(directions,dir_values)) # list of action along with their values -1 for UP etc.
# -
# #### Maximising state values
# +
while delta > theta: # loop condition
delta =0
count+=1
for k in range (0,4): # running loop within all state values
for l in range (0,4):
if (k==0 and l==0) or (k==3 and l==3):
continue
grid_eval=np.zeros((4,1)) # evaluation grid
v=grid[k,l] # temporary value
for m in range(0,2): # storing values for actions UP and DOWN
if k==0 and m==0:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
continue
elif k==3 and m ==1:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
else:
grid_eval[m,0] = (r+gamma*(grid[k+dir_l[m][1],l]))
for m in range(2,4): # storing values for actions LEFT and RIGHT
if l==0 and m==2:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
continue
elif l==3 and m==3:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
else:
grid_eval[m,0] = (r+gamma*(grid[k,l+dir_l[m][1]]))
grid[k,l] = np.max(grid_eval) # assigning max value to the current state grid[k,l]
delta = max(delta ,abs(v-grid[k,l])) # updating the delta value
print(grid) #final state after setting maximising state values
# -
# #### Finding optimal policy
# +
for k in range(0,4):
for l in range(0,4):
if (k==0 and l==0) or (k==3 and l==3):
continue
grid_eval=np.zeros((4,1))
for m in range(0,2):
if k==0 and m==0:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
continue
elif k==3 and m ==1:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
else:
grid_eval[m,0] = (r+gamma*(grid[k+dir_l[m][1],l]))
for m in range(2,4):
if l==0 and m==2:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
continue
elif l==3 and m==3:
grid_eval[m,0] = (r+gamma*(grid[k,l]))
else:
grid_eval[m,0] = (r+gamma*(grid[k,l+dir_l[m][1]]))
pi[k,l,np.argmax(grid_eval,axis=0)]=1 # setting action which leads to max state value as 1
print(pi)
# the optimal policy matrix is a 3D 4,4,4 matrix with first two indexes representing rows and columns and last
#index representing the action taken in order UP, DOWN, LEFT and RIGHT
# -
| 3,918 |
/HomeWork/Day_090_color_histogram_HW.ipynb | 0c174f15e7d94b2dbc1b88b8daebf611d928d037 | [] | no_license | fyang109/3rd-ML100Days | https://github.com/fyang109/3rd-ML100Days | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 134,605 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 作業目標
# * 畫出 16 個 bin 的顏色直方圖,並嘗試回答每個 channel 在 [16, 32] 這個 bin 中有多少個 pixel?
# * 確保同學知道如何改變參數以及理解 cv2.calcHist 這個函數輸出的資料含義
# +
import os
import keras
os.environ["CUDA_VISIBLE_DEVICES"] = "" # 使用 CPU
import cv2 # 載入 cv2 套件
import matplotlib.pyplot as plt
train, test = keras.datasets.cifar10.load_data()
# +
image = train[0][0] # 讀取圖片
plt.imshow(image)
plt.show()
# -
# 把彩色的圖片轉為灰度圖
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
plt.imshow(gray)
plt.show()
# 通過調用 __cv2.calcHist(images, channels, mask, histSize, ranges)__ 函數來得到統計後的直方圖值
#
# * images (list of array):要分析的圖片
# * channels:產生的直方圖類型。例:[0]→灰度圖,[0, 1, 2]→RGB三色。
# * mask:optional,若有提供則僅計算 mask 部份的直方圖。
# * histSize:要切分的像素強度值範圍,預設為256。每個channel皆可指定一個範圍。例如,[32,32,32] 表示RGB三個channels皆切分為32區段。
# * ranges:像素的範圍,預設為[0,256],表示<256。
# 調用 cv2.calcHist 函數,回傳值就是 histogram
hist = cv2.calcHist([gray], [0], None, [256], [0, 256])
plt.figure()
plt.title("Grayscale Histogram")
plt.xlabel("Bins")
plt.ylabel("# of Pixels")
plt.plot(hist)
plt.xlim([0, 256])
plt.show()
print("hist shape:", hist.shape, "\n直方圖中前兩個值:", hist[:2]) # 1 表示該灰度圖中,只有 1 個 pixel 的值是 0,0 個 pixel 的值是 1
# +
chans = cv2.split(image) # 把圖像的 3 個 channel 切分出來
colors = ("r", "g", "b")
plt.figure()
plt.title("'Flattened' Color Histogram")
plt.xlabel("Bins")
plt.ylabel("# of Pixels")
# 對於所有 channel
for (chan, color) in zip(chans, colors):
# 計算該 channel 的直方圖
hist = cv2.calcHist([chan], [0], None, [256], [0, 256])
# 畫出該 channel 的直方圖
plt.plot(hist, color = color)
plt.xlim([0, 256])
plt.show()
# +
#畫出 16 個 bin 的顏色直方圖,並嘗試回答每個 channel 在 [16, 32] 這個 bin 中有多少個 pixel?
chans = cv2.split(image) # 把圖像的 3 個 channel 切分出來
colors = ("r", "g", "b")
plt.figure()
plt.title("'Flattened' Color Histogram")
plt.xlabel("Bins")
plt.ylabel("# of Pixels")
# 對於所有 channel
for (chan, color) in zip(chans, colors):
# 計算該 channel 的直方圖
hist = cv2.calcHist([chan], [0], None, [16], [0, 256])
print('{0}:{1}個pixels'.format(color,sum(hist[1])))
plt.plot(hist, color = color)
plt.xlim([0,15])
plt.show()
| 2,314 |
/code/candidates.ipynb | 8eee31dcb90216106215b3760a81a5abeb5379a6 | [] | no_license | sahilchinoy/econ191 | https://github.com/sahilchinoy/econ191 | 0 | 1 | null | null | null | null | Jupyter Notebook | false | false | .py | 6,427 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 直接调用XGBoost
# ## 导入必要的工具包
# +
# 导入必要的工具包
import xgboost as xgb
# 计算分类正确率
from sklearn.metrics import accuracy_score
# -
# ## 数据读取
# XGBoost可以加载libsvm格式的文本数据,libsvm的文件格式(稀疏特征)如下:
# 1 101:1.2 102:0.03
# 0 1:2.1 10001:300 10002:400
# ...
#
# 每一行表示一个样本,第一行的开头的“1”是样本的标签。“101”和“102”为特征索引,'1.2'和'0.03' 为特征的值。
# 在两类分类中,用“1”表示正样本,用“0” 表示负样本。也支持[0,1]表示概率用来做标签,表示为正样本的概率。
#
# 下面的示例数据需要我们通过一些蘑菇的若干属性判断这个品种是否有毒。
# UCI数据描述:http://archive.ics.uci.edu/ml/machine-learning-databases/mushroom/ ,
# 每个样本描述了蘑菇的22个属性,比如形状、气味等等(将22维原始特征用加工后变成了126维特征,
# 并存为libsvm格式),然后给出了这个蘑菇是否可食用。其中6513个样本做训练,1611个样本做测试。
#
# XGBoost加载的数据存储在对象DMatrix中
# XGBoost自定义了一个数据矩阵类DMatrix,优化了存储和运算速度
# DMatrix文档:http://xgboost.readthedocs.io/en/latest/python/python_api.html
# read in data,数据在xgboost安装的路径下的demo目录,现在我们将其copy到当前代码下的data目录
my_workpath = './data/'
dtrain = xgb.DMatrix(my_workpath + 'agaricus.txt.train')
dtest = xgb.DMatrix(my_workpath + 'agaricus.txt.test')
# 查看数据情况
dtrain.num_col()
dtrain.num_row()
dtest.num_row()
# ## 训练参数设置
# max_depth: 树的最大深度。缺省值为6,取值范围为:[1,∞]
# eta:为了防止过拟合,更新过程中用到的收缩步长。在每次提升计算之后,算法会直接获得新特征的权重。
# eta通过缩减特征的权重使提升计算过程更加保守。缺省值为0.3,取值范围为:[0,1]
# silent:取0时表示打印出运行时信息,取1时表示以缄默方式运行,不打印运行时信息。缺省值为0
# objective: 定义学习任务及相应的学习目标,“binary:logistic” 表示二分类的逻辑回归问题,输出为概率。
#
# 其他参数取默认值。
# specify parameters via map
param = {'max_depth':2, 'eta':1, 'silent':0, 'objective':'binary:logistic' }
print param
# ## 训练模型
# 有了参数列表和数据就可以训练模型了
# +
# 设置boosting迭代计算次数
num_round = 2
import time
starttime = time.clock()
bst = xgb.train(param, dtrain, num_round)
endtime = time.clock()
print (endtime - starttime)
# -
# 查看模型在训练集上的分类性能
# XGBoost预测的输出是概率。这里蘑菇分类是一个二类分类问题,输出值是样本为第一类的概率。
# 我们需要将概率值转换为0或1。
train_preds = bst.predict(dtrain)
train_predictions = [round(value) for value in train_preds]
y_train = dtrain.get_label()
train_accuracy = accuracy_score(y_train, train_predictions)
print ("Train Accuary: %.2f%%" % (train_accuracy * 100.0))
# ## 测试
# 模型训练好后,可以用训练好的模型对测试数据进行预测
# make prediction
preds = bst.predict(dtest)
# 检查模型在测试集上的正确率
# XGBoost预测的输出是概率,输出值是样本为第一类的概率。我们需要将概率值转换为0或1。
predictions = [round(value) for value in preds]
y_test = dtest.get_label()
test_accuracy = accuracy_score(y_test, predictions)
print("Test Accuracy: %.2f%%" % (test_accuracy * 100.0))
# ## 模型可视化
# 调用XGBoost工具包中的plot_tree,在显示
# 要可视化模型需要安装graphviz软件包(请见预备0)
#
# plot_tree()的三个参数:
# 1. 模型
# 2. 树的索引,从0开始
# 3. 显示方向,缺省为竖直,‘LR'是水平方向
# +
from matplotlib import pyplot
import graphviz
xgb.plot_tree(bst, num_trees=0, rankdir= 'LR' )
pyplot.show()
#xgb.plot_tree(bst,num_trees=1, rankdir= 'LR' )
#pyplot.show()
#xgb.to_graphviz(bst,num_trees=0)
#xgb.to_graphviz(bst,num_trees=1)
| 2,933 |
/Multi-Model-professional-association.ipynb | 5998253c5a4dc46fd4d82e751c7e7c199f31d2d1 | [] | no_license | sara-kassani/Accounting-Agents-ML | https://github.com/sara-kassani/Accounting-Agents-ML | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 22,378 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings('always')
warnings.filterwarnings('ignore')
# +
import numpy as np
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split, KFold , cross_val_score, GridSearchCV
from sklearn.neighbors import KNeighborsClassifier
from xgboost import XGBClassifier, plot_importance
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.metrics import classification_report
import matplotlib.pyplot as plt
# %matplotlib inline
# -
df_all = pd.read_csv("data/first data.csv")
df_all.shape
df = df_all.drop('Questionnaire Number',axis=1)
df.drop('human-resources-actor',axis=1, inplace=True)
df.drop('technology-actor',axis=1, inplace=True)
df.drop('regulation-actor',axis=1, inplace=True)
df.drop('administrative-dimension-actor',axis=1, inplace=True)
df.drop('external-users',axis=1, inplace=True)
df.drop('state-institution',axis=1, inplace=True)
df.drop('scientific-community',axis=1, inplace=True)
# df.drop('professional-association',axis=1, inplace=True)
df.drop('economic-actor',axis=1, inplace=True)
df.drop('technical-dimension',axis=1, inplace=True)
df.drop('political-social',axis=1, inplace=True)
df.info()
X = df.drop("professional-association", axis = 1)
y = df["professional-association"]
X.shape
y.shape
X.head()
# ____
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state=42)
X_train.shape, y_train.shape, X_test.shape, y_test.shape
# +
seed = 42
models = []
models.append(('XGBoost', XGBClassifier(seed = seed) ))
models.append(('SVC', SVC(random_state=seed)))
models.append(('RF', RandomForestClassifier(random_state=seed, n_jobs=-1 )))
tree = DecisionTreeClassifier(max_depth=4, random_state=seed)
models.append(('KNN', KNeighborsClassifier(n_jobs=-1)))
results, names = [], []
num_folds = 10
scoring = 'accuracy'
for name, model in models:
kfold = KFold(n_splits=num_folds, random_state=seed)
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring = scoring, n_jobs= -1)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# -
fig = plt.figure(figsize=(8,6))
fig.suptitle("Algorithms comparison")
ax = fig.add_subplot(1,1,1)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
# Update current m and b
m_updated = m_current - alpha * m_gradient
b_updated = b_current - alpha * b_gradient
#Return updated parameters
return b_updated, m_updated
def gradient_descent(data, starting_b, starting_m, learning_rate, num_iterations):
"""runs gradient descent
Args:
data (np.array): training data, containing x,y
starting_b (float): initial value of b (random)
starting_m (float): initial value of m (random)
learning_rate (float): hyperparameter to adjust the step size during descent
num_iterations (int): hyperparameter, decides the number of iterations for which gradient descent would run
Returns:
list : the first and second item are b, m respectively at which the best fit curve is obtained, the third and fourth items are two lists, which store the value of b,m as gradient descent proceeded.
"""
# initial values
b = starting_b
m = starting_m
# to store the cost after each iteration
cost_graph = []
# to store the value of b -> bias unit, m-> slope of line after each iteration (pred = m*x + b)
b_progress = []
m_progress = []
# For every iteration, optimize b, m and compute its cost
for i in range(num_iterations):
cost_graph.append(compute_cost(b, m, data))
b, m = step_gradient(b, m, array(data), learning_rate)
b_progress.append(b)
m_progress.append(m)
return [b, m, cost_graph,b_progress,m_progress]
# + [markdown] id="5v-y-qI3-ep7"
# #### Run gradient_descent() to get optimized parameters b and m
# + id="WV6KIdPq-ep7"
b, m, cost_graph,b_progress,m_progress = gradient_descent(data, initial_b, initial_m, learning_rate, num_iterations)
#Print optimized parameters
print ('Optimized b:', b)
print ('Optimized m:', m)
#Print error with optimized parameters
print ('Minimized cost:', compute_cost(b, m, data))
# + [markdown] id="wnmqepye-ep_"
# #### Plotting the cost per iterations
# + id="x1VREIE7-eqA"
plt.plot(cost_graph)
plt.xlabel('No. of iterations')
plt.ylabel('Cost')
plt.title('Cost per iteration')
plt.show()
# + [markdown] id="YWQAkFE8-eqD"
# Gradient descent converges to local minimum after 5 iterations
# + [markdown] id="OT2wKbHu-eqE"
# #### Plot line of best fit
# + id="hD531y8K-eqF"
#Plot dataset
plt.scatter(x, y)
#Predict y values
pred = m * x + b
#Plot predictions as line of best fit
plt.plot(x, pred, c='r')
plt.xlabel('Hours of study')
plt.ylabel('Test scores')
plt.title('Line of best fit')
plt.show()
# + [markdown] id="XeXdBWc-Cic-"
# ### Gradient descent's progress with num of iterations
# + id="pHGbi8nY-eqI"
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import numpy as np
fig = plt.figure()
ax = fig.add_subplot(111)
b = b_progress[0]
m = m_progress[0]
pred = m*x + b
line = ax.plot(x,pred, '-',c='r')[0]
def animate(i,b_prog,m_prog):
pred = m_prog[i] * x + b_prog[i]
line.set_data(x,pred)
return line,
ani = animation.FuncAnimation(fig, animate, frames=len(b_progress), fargs=(b_progress,m_progress,))
ax.scatter(x,y)
HTML(ani.to_jshtml())
| 5,996 |
/generalization_MDP_driver/DDPG2.ipynb | 07d3e81454fa7ad8f45a31d484bd08162b5e866c | [] | no_license | zidane89/experiment | https://github.com/zidane89/experiment | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 96,777 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
from tensorflow import keras
import os
import math
import random
import pickle
import glob
import matplotlib.pyplot as plt
from collections import deque
from tensorflow.keras import layers
import time
from vehicle_model_DDPG2 import Environment
from cell_model import CellModel
from driver_MDP import Driver_MDP
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
# +
drving_cycle = '../../OC_SIM_DB/OC_SIM_DB_Cycles/Highway/01_FTP72_fuds.mat'
battery_path = "../../OC_SIM_DB/OC_SIM_DB_Bat/OC_SIM_DB_Bat_nimh_6_240_panasonic_MY01_Prius.mat"
motor_path = "../../OC_SIM_DB/OC_SIM_DB_Mot/OC_SIM_DB_Mot_pm_95_145_X2.mat"
cell_model = CellModel()
# env = Environment(cell_model, drving_cycle, battery_path, motor_path, 10)
driver = Driver_MDP(0.02)
num_states = 4
# -
class OUActionNoise:
def __init__(self, mean, std_deviation, theta=0.15, dt=1e-2, x_initial=None):
self.theta = theta
self.mean = mean
self.std_dev = std_deviation
self.dt = dt
self.x_initial = x_initial
self.reset()
def reset(self):
if self.x_initial is not None:
self.x_prev = self.x_initial
else:
self.x_prev = 0
def __call__(self):
x = (
self.x_prev + self.theta * (self.mean - self.x_prev) * self.dt
+ self.std_dev * np.sqrt(self.dt) * np.random.normal()
)
self.x_prev = x
return x
class Buffer:
def __init__(self, buffer_capacity=100000, batch_size=64):
self.buffer_capacity = buffer_capacity
self.batch_size = batch_size
self.buffer_counter = 0
self.state_buffer = np.zeros((self.buffer_capacity, num_states))
self.action_buffer = np.zeros((self.buffer_capacity, 1))
self.reward_buffer = np.zeros((self.buffer_capacity, 1))
self.next_state_buffer = np.zeros((self.buffer_capacity, num_states))
def record(self, obs_tuple):
index = self.buffer_counter % self.buffer_capacity
self.state_buffer[index] = obs_tuple[0]
self.action_buffer[index] = obs_tuple[1]
self.reward_buffer[index] = obs_tuple[2]
self.next_state_buffer[index] = obs_tuple[3]
self.buffer_counter += 1
def learn(self):
record_range = min(self.buffer_counter, self.buffer_capacity)
batch_indices = np.random.choice(record_range, self.batch_size)
state_batch = tf.convert_to_tensor(self.state_buffer[batch_indices])
action_batch = tf.convert_to_tensor(self.action_buffer[batch_indices])
reward_batch = tf.convert_to_tensor(self.reward_buffer[batch_indices])
reward_batch = tf.cast(reward_batch, dtype=tf.float32)
next_state_batch = tf.convert_to_tensor(self.next_state_buffer[batch_indices])
with tf.GradientTape() as tape:
target_actions = target_actor(next_state_batch)
y = reward_batch + gamma * target_critic([next_state_batch, target_actions])
critic_value = critic_model([state_batch, action_batch])
critic_loss = tf.math.reduce_mean(tf.square(y - critic_value))
critic_grad = tape.gradient(critic_loss, critic_model.trainable_variables)
critic_optimizer.apply_gradients(
zip(critic_grad, critic_model.trainable_variables)
)
with tf.GradientTape() as tape:
actions = actor_model(state_batch)
critic_value = critic_model([state_batch, actions])
actor_loss = - tf.math.reduce_mean(critic_value)
actor_grad = tape.gradient(actor_loss, actor_model.trainable_variables)
actor_optimizer.apply_gradients(
zip(actor_grad, actor_model.trainable_variables)
)
def update_target(tau):
new_weights = []
target_variables = target_critic.weights
for i, variable in enumerate(critic_model.weights):
new_weights.append(target_variables[i] * (1 - tau) + tau * variable)
target_critic.set_weights(new_weights)
new_weights = []
target_variables = target_actor.weights
for i, variable in enumerate(actor_model.weights):
new_weights.append(target_variables[i] * (1 - tau) + tau * variable)
target_actor.set_weights(new_weights)
def get_actor():
last_init = tf.random_uniform_initializer(minval=-0.003, maxval=0.003)
inputs = layers.Input(shape=(num_states))
inputs_batchnorm = layers.BatchNormalization()(inputs)
out = layers.Dense(512, activation="relu")(inputs_batchnorm)
# out = layers.BatchNormalization()(out)
out = layers.Dense(512, activation="relu")(out)
# out = layers.BatchNormalization()(out)
outputs = layers.Dense(1, activation="sigmoid",
kernel_initializer=last_init)(out)
model = tf.keras.Model(inputs, outputs)
return model
def get_critic():
state_input = layers.Input(shape=(num_states))
state_input_batchnorm = layers.BatchNormalization()(state_input)
state_out = layers.Dense(16, activation="relu")(state_input_batchnorm)
# state_out = layers.BatchNormalization()(state_out)
state_out = layers.Dense(32, activation="relu")(state_out)
# state_out = layers.BatchNormalization()(state_out)
action_input = layers.Input(shape=(1))
action_out = layers.Dense(32, activation="relu")(action_input)
# action_out = layers.BatchNormalization()(action_out)
concat = layers.Concatenate()([state_out, action_out])
out = layers.Dense(512, activation="relu")(concat)
# out = layers.BatchNormalization()(out)
out = layers.Dense(512, activation="relu")(out)
# out = layers.BatchNormalization()(out)
outputs = layers.Dense(1)(out)
model = tf.keras.Model([state_input, action_input], outputs)
return model
def policy(state, noise_object):
j_min = state[0][2].numpy()
j_max = state[0][3].numpy()
sampled_action = tf.squeeze(actor_model(state))
noise = noise_object()
sampled_action = sampled_action.numpy() + noise
legal_action = sampled_action * j_max
legal_action = np.clip(legal_action, j_min, j_max)
# print(j_min, j_max, legal_action, noise)
return legal_action
def policy_epsilon_greedy(state, eps):
j_min = state[0][-2].numpy()
j_max = state[0][-1].numpy()
if random.random() < eps:
a = random.randint(0, 9)
return np.linspace(j_min, j_max, 10)[a]
else:
sampled_action = tf.squeeze(actor_model(state)).numpy()
legal_action = sampled_action * j_max
legal_action = np.clip(legal_action, j_min, j_max)
return legal_action
# +
std_dev = 0.2
ou_noise = OUActionNoise(mean=0, std_deviation=0.2)
critic_lr = 0.0005
actor_lr = 0.00025
critic_optimizer = tf.keras.optimizers.Adam(critic_lr)
actor_optimizer = tf.keras.optimizers.Adam(actor_lr)
total_episodes = 1000
gamma = 0.95
tau = 0.001
MAX_EPSILON = 1.0
MIN_EPSILON = 0.01
DECAY_RATE = 0.00002
BATCH_SIZE = 32
DELAY_TRAINING = 5000
# -
def initialization():
actor_model = get_actor()
critic_model = get_critic()
target_actor = get_actor()
target_critic = get_critic()
target_actor.set_weights(actor_model.get_weights())
target_critic.set_weights(critic_model.get_weights())
# actor_model.load_weights("./DDPG1_trial1/actor_model_checkpoint")
# critic_model.load_weights("./DDPG1_trial1/critic_model_checkpoint")
# target_actor.load_weights("./DDPG1_trial1/target_actor_checkpoint")
# target_critic.load_weights("./DDPG1_trial1/target_critic_checkpoint")
buffer = Buffer(500000, BATCH_SIZE)
return actor_model, critic_model, target_actor, target_critic, buffer
def save_weights(actor_model, critic_model, target_actor, target_critic, root):
actor_model.save_weights("./{}/actor_model_checkpoint".format(root))
critic_model.save_weights("./{}/critic_model_checkpoint".format(root))
target_actor.save_weights("./{}/target_actor_checkpoint".format(root))
target_critic.save_weights("./{}/target_critic_checkpoint".format(root))
print("model is saved..")
def initialization_env(driving_path, reward_factor):
env = Environment(cell_model, driving_path, battery_path, motor_path, reward_factor)
return env
def test_agent(actor_model, reward_factor):
test_cycle = driver.get_cycle()
env = initialization_env(test_cycle, reward_factor)
total_reward = 0
state = env.reset()
while True:
tf_state = tf.expand_dims(tf.convert_to_tensor(state), 0)
action = policy_epsilon_greedy(tf_state, -1)
next_state, reward, done = env.step(action)
state = next_state
total_reward += reward
if done:
break
SOC_deviation_history = np.sum(np.abs(np.array(env.history["SOC"]) - 0.6))
print("******************* Test is start *****************")
# print(test_cycle)
print('Total reward: {}'.format(total_reward),
"SOC: {:.4f}".format(env.SOC),
"Cumulative_SOC_deviation: {:.4f}".format(SOC_deviation_history),
"Fuel Consumption: {:.4f}".format(env.fuel_consumption))
print("******************* Test is done *****************")
print("")
plt.plot(test_cycle)
plt.show()
return env.history
# +
# print(env.version)
num_trials = 1
results_dict = {}
# driving_cycle_paths = glob.glob("../data/driving_cycles/city/*.mat")[:1]
for trial in range(num_trials):
print("")
print("Trial {}".format(trial))
print("")
actor_model, critic_model, target_actor, target_critic, buffer = initialization()
eps = MAX_EPSILON
steps = 0
episode_rewards = []
episode_SOCs = []
episode_FCs = []
episode_test_history = []
episode_num_test = []
for ep in range(total_episodes):
driving_cycle = driver.get_cycle()
env = initialization_env(driving_cycle, 10)
start = time.time()
state = env.reset()
episodic_reward = 0
while True:
tf_state = tf.expand_dims(tf.convert_to_tensor(state), 0)
action = policy_epsilon_greedy(tf_state, eps)
# print(action)
next_state, reward, done = env.step(action)
if done:
next_state = [0] * num_states
buffer.record((state, action, reward, next_state))
episodic_reward += reward
if steps > DELAY_TRAINING:
buffer.learn()
update_target(tau)
eps = MIN_EPSILON + (MAX_EPSILON - MIN_EPSILON) * np.exp(-DECAY_RATE * (steps
-DELAY_TRAINING))
steps += 1
if done:
break
state = next_state
elapsed_time = time.time() - start
print("elapsed_time: {:.3f}".format(elapsed_time))
episode_rewards.append(episodic_reward)
episode_SOCs.append(env.SOC)
episode_FCs.append(env.fuel_consumption)
# print("Episode * {} * Avg Reward is ==> {}".format(ep, avg_reward))
SOC_deviation_history = np.sum(np.abs(np.array(env.history["SOC"]) - 0.6))
print(
'Episode: {}'.format(ep + 1),
"Exploration P: {:.4f}".format(eps),
'Total reward: {}'.format(episodic_reward),
"SOC: {:.4f}".format(env.SOC),
"Cumulative_SOC_deviation: {:.4f}".format(SOC_deviation_history),
"Fuel Consumption: {:.4f}".format(env.fuel_consumption),
)
print("")
if (ep + 1) % 10 == 0:
history = test_agent(actor_model, 10)
episode_test_history.append(history)
episode_num_test.append(ep + 1)
root = "DDPG2_trial{}".format(trial+1)
save_weights(actor_model, critic_model, target_actor, target_critic, root)
results_dict[trial + 1] = {
"rewards": episode_rewards,
"SOCs": episode_SOCs,
"FCs": episode_FCs,
"test_history": episode_test_history,
"test_episode_num": episode_num_test,
}
# -
with open("DDPG2.pkl", "wb") as f:
pickle.dump(results_dict, f, pickle.HIGHEST_PROTOCOL)
# +
# results_dict
# -
| 12,783 |
/PythonCode/U41A2.ipynb | dfc88763b775c705e0a484bf4a0f1caaab6bdd56 | [] | no_license | JoKaBus/VEH2020 | https://github.com/JoKaBus/VEH2020 | 0 | 0 | null | 2020-09-29T18:35:51 | 2020-09-29T18:34:47 | Jupyter Notebook | Jupyter Notebook | false | false | .py | 247,896 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 300
# +
def f(y):
return np.sqrt(y-1)
def g(m):
return m+1
def h(z):
return z/(z**2+1)
# +
plot_min = 1
plot_max = 100
X = np.linspace(plot_min,plot_max,1000)
fig, ax = plt.subplots()
ax.set_ylim(-2.5,max(f(X))+1)
ax.plot(X,f(X), label = '$y\mapsto \sqrt{y-1}$')
ax.legend()
props = dict(boxstyle='round', facecolor='wheat', alpha=0.5)
textstr = 'Nicht bijektiv, da $f^{-1}(z) = \emptyset $ for $z<0$'
ax.text(0.05, 0.05, textstr, transform=ax.transAxes, fontsize=14,
verticalalignment='bottom', bbox=props)
# plt.savefig('Plots/PresPlots/U41A2asolve.png')
# +
plot_min = 1
plot_max = 10
X = np.arange(plot_min,plot_max+1,2)
fig, ax = plt.subplots()
ax.set_ylim(-0.3,max(g(X))+1)
ax.scatter(X,g(X), label = '$m \mapsto m+1$ for $m\equiv 0$ (mod $2$)')
ax.scatter([1],[1], color = 'red')
ax.legend()
props = dict(boxstyle='round', facecolor='wheat', alpha=0.5)
textstr = 'Nicht bijektiv, da $f^{-1}(1) = \emptyset $'
ax.text(0.15, 0.07, textstr, transform=ax.transAxes, fontsize=14,
verticalalignment='bottom', bbox=props)
# plt.savefig('Plots/PresPlots/U41A2bsolve.png')
# +
plot_min = 1
plot_max = 100
X = np.linspace(plot_min,plot_max,1000)
fig, ax = plt.subplots()
ax.plot(X,h(X), label = '$z\mapsto z/(z^2+1)$')
ax.legend()
# plt.savefig('Plots/PresPlots/U41A2c.png')
# +
# emptybox
plot_min = 1
plot_max = 100
X = np.linspace(plot_min,plot_max,1000)
fig, ax = plt.subplots()
ax.plot(X,f(X), color = 'white')
# plt.savefig('Plots/PresPlots/U41A2empty.png')
: Why aren't we using the [merge](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.merge.html) method to combine the dataframes?) If you don’t get it, I’ll show you how afterwards. Make sure to save your work in this notebook! You'll come back to this later.
# +
# append dataframes
wine_df = red_df.append(white_df)
# view dataframe to check for success
wine_df.head()
# -
# ## Save Combined Dataset
# Save your newly combined dataframe as `winequality_edited.csv`. Remember, set `index=False` to avoid saving with an unnamed column!
wine_df.to_csv('winequality_edited.csv', index=False)
{A:.3f}'
summary.append(res)
if bestBIC is None or B < bestBIC:
bestBIC = B
besti = len(summary) - 1
if bestAIC is None or A < bestAIC:
bestAIC = A
bestj = len(summary) - 1
for i in range(len(summary)):
if i == besti:
if i == bestj:
pre = f'{i+1:3} **'
else:
pre = f'{i+1:3} B*'
elif i == bestj:
pre = f'{i+1:3} A*'
else:
pre = f'{i+1:3} '
print(f'{pre} {summary[i]}')
# ## List of Pareto solutions: model specifications
# For each model in the Pareto set, we provide a description of the model specification.
counter = 0
for p in pareto.pareto:
counter += 1
print(f'*************** Model {counter} ************************')
print(p)
print('\n')
# ## List of Pareto solutions: illustration
# The plot below illustrates all models considered by the algorithm. Each model corresponds to one point in the graph.
#
# - The x-coordinate corresponds to the negative log likelihood of the model, and the y-coordinate to the number of parameters.
# - The larger circles correspond to all models that are not dominated. They are in the Pareto set.
# - The crosses corresponds to model that happened to be non dominated at some point during the course of the algorithm, but have been removed from the Pareto set afterwards, as a dominating model has been identified.
# - Finally, the small dots corresponds to models that have been considered, but rejected because dominated by another model already in the set.
objectives = list(pareto.pareto)[0].objectivesNames
objectives
par_obj = [p.objectives for p in pareto.pareto]
par_x, par_y = zip(*par_obj)
con_obj = [p.objectives for p in pareto.considered]
con_x, con_y = zip(*con_obj)
rem_obj = [p.objectives for p in pareto.removed]
rem_x, rem_y = zip(*rem_obj)
x_buffer = 10
y_buffer = 0.1
plt.axis([min(par_x)-x_buffer,
max(par_x)+x_buffer,
min(par_y)-y_buffer,
max(par_y)+y_buffer])
plt.plot(par_x, par_y, 'o', label='Pareto')
plt.plot(rem_x, rem_y, 'x', label='Removed')
plt.plot(con_x, con_y, ',', label='Considered')
plt.xlabel(objectives[0])
plt.ylabel(objectives[1])
plt.legend()
| 4,699 |
/AAAI/Learnability/hard_attention_sdc/non-linear/dataset_4/hard_attention_9 (1).ipynb | 11f27f4c1fa7264a45f0207fb4722ffec584415b | [
"MIT"
] | permissive | lnpandey/DL_explore_synth_data | https://github.com/lnpandey/DL_explore_synth_data | 3 | 3 | null | null | null | null | Jupyter Notebook | false | false | .py | 130,084 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="0vlCAi2JLSzD"
# from google.colab import drive
# drive.mount('/content/drive')
# + id="PiPNZm1iTgHy"
# path="/content/drive/MyDrive/Research/Hard_Attention/dataset_2/m_5_size_100/run_"
# + id="SrZgZMlK-GDe"
# + colab={"base_uri": "https://localhost:8080/"} id="N2_J4Rw2r0SQ" outputId="d0a95b24-e155-4cda-b6e4-2d258f70ae1e"
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
# %matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
from tqdm import tqdm as tqdm
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
# + id="-Dmy2iPWlgnc"
from torch.utils.tensorboard import SummaryWriter
writer = SummaryWriter('runs/hard_attention')
# + [markdown] id="F6fjud_Fr0Sa"
# # Generate dataset
# + colab={"base_uri": "https://localhost:8080/"} id="CqdXHO0Cr0Sd" outputId="db15d4ad-6140-4e92-d2f5-603b3331a687"
np.random.seed(1)
y = np.concatenate((np.zeros(500),np.ones(500),np.ones(500)*2,np.ones(500)*3,np.ones(500)*4,
np.ones(500)*5,np.ones(500)*6,np.ones(500)*7,np.ones(500)*8,np.ones(500)*9))
#y = np.random.randint(0,3,6000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
# + id="ddhXyODwr0Sk"
x = np.zeros((5000,2))
# + id="DyV3N2DIr0Sp"
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [4,6.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [5.5,6],cov=[[0.01,0],[0,0.01]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [4.5,4.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [3,3.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [2.5,5.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [3.5,8],cov=[[0.01,0],[0,0.01]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [5.5,8],cov=[[0.01,0],[0,0.01]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [7,6.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [6.5,4.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [5,3],cov=[[0.01,0],[0,0.01]],size=sum(idx[9]))
# + colab={"base_uri": "https://localhost:8080/"} id="qh1mDScsU07I" outputId="f73961e3-241f-45c8-a945-07ef97c11b4b"
x[idx[0]][0], x[idx[2]][5]
# + colab={"base_uri": "https://localhost:8080/"} id="9Vr5ErQ_wSrV" outputId="7d0a7057-4082-4613-bffb-f2fdc05290cd"
print(x.shape,y.shape)
# + id="NG-3RpffwU_i"
idx= []
for i in range(10):
idx.append(y==i)
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="hJ8Jm7YUr0St" outputId="ddac3265-6d37-4da7-adaf-3ce093bb23d6"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="fNWgnhUJnWLV"
x = ( x - np.mean(x,axis=0,keepdims=True) ) / np.std(x,axis=0,keepdims=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="8-VLhUfDDeHt" outputId="fdc08cdd-4576-48c3-8f84-4f5cf3dc1298"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="UfFHcZJOr0Sz"
foreground_classes = {'class_0','class_1' }
background_classes = {'bg_classes',}
# + colab={"base_uri": "https://localhost:8080/"} id="jqbvfbwVr0TN" outputId="cd2f9995-d35a-4973-a06d-0cedc5baf95f"
desired_num = 3000
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
m = 2000
for j in tqdm(range(desired_num)):
np.random.seed(j)
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(np.reshape(a,(m,2)))
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
# + id="BOsFmWfMr0TR"
# mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
# + colab={"base_uri": "https://localhost:8080/"} id="2aIPMgLXNiXW" outputId="877c8a57-644e-4732-86f0-ad442adf4095"
len(mosaic_list_of_images), mosaic_list_of_images[0],mosaic_list_of_images[0].shape
# + id="iPoIwbMHx44n"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
# + id="fOPAJQJeW8Ah"
batch = 50
msd1 = MosaicDataset(mosaic_list_of_images[0:2000], mosaic_label[0:2000] , fore_idx[0:2000])
train_loader = DataLoader( msd1 ,batch_size= batch ,shuffle=True)
# + id="aWBIcyvGApLt"
data,_,_=iter(train_loader).next()
# + colab={"base_uri": "https://localhost:8080/"} id="cauJIvKEAxKM" outputId="efbb609d-f831-45c1-e0da-6d410d1550b3"
data.shape
# + id="qjNiQgxZW8bA"
batch = 250
msd2 = MosaicDataset(mosaic_list_of_images[2000:], mosaic_label[2000:] , fore_idx[2000:])
test_loader = DataLoader( msd2 ,batch_size= batch ,shuffle=True)
# + id="yda1E5ApiKpH"
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.fc1 = nn.Linear(2,50, bias=False)
self.fc2 = nn.Linear(50,1,bias=False)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.xavier_normal_(self.fc2.weight)
#self.fc2 = nn.Linear(64, 1, bias=False)
#torch.nn.init.xavier_normal_(self.fc2.weight)
def forward(self,z):
#print("data",z)
batch = z.size(0)
patches = z.size(1)
z = z.view(batch,patches,2*1)
alp1,ft1 = self.helper(z)
alpha = F.softmax(alp1,dim=1)
#print(self.training)
if self.training:
alpha =alpha[:,:,0]
y = ft1
return alpha,y
else:
#alpha_cumsum = torch.cumsum(alpha, dim = 1)
#print(alpha_cumsum)
#len_batch = alpha_cumsum.size(0)
#patches = alpha_cumsum.size(1)
#rand_prob = torch.rand(len_batch,patches, 1).to(device)
#alpha_relu = F.relu(rand_prob-alpha_cumsum)
#print(alpha_relu)
#alpha_index = torch.count_nonzero(alpha_relu,dim=1)
#alpha_hard = F.one_hot(alpha_index,num_classes=patches)
#print(alpha_hard)
#alpha_hard = torch.transpose(alpha_hard,dim0=1,dim1=2)
#print(ft1,"alpha_hard",alpha_hard)
#y = torch.sum(alpha_hard*ft1,dim=1)
#print(alpha,alpha.shape)
index = torch.argmax(alpha,dim=1)
hard_alpha = torch.nn.functional.one_hot(index[:,0], patches)
y = torch.sum(hard_alpha[:,:,None]*ft1,dim=1)
alpha = alpha[:,:,0]
return alpha,y
def helper(self, x):
x1 = x
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x,x1
# + id="0dYXnywAD-4l"
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.fc1 = nn.Linear(2, 50)
self.fc2 = nn.Linear(50,3)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.zeros_(self.fc1.bias)
torch.nn.init.xavier_normal_(self.fc2.weight)
torch.nn.init.zeros_(self.fc2.bias)
def forward(self, x):
#print(x.shape)
#x = x.view(-1, 1)
#print(x.shape)
x = F.relu(self.fc1(x))
x = self.fc2(x)
# print(x.shape)
return x
# + id="lSa6O9f6XNf4"
torch.manual_seed(13)
focus_net = Focus().double()
focus_net = focus_net.to(device)
# + id="36k3H2G-XO9A"
torch.manual_seed(13)
classify = Classification().double()
classify = classify.to(device)
# + id="bK78aII8-GDl"
import torch.optim as optim
optimizer_classify = optim.Adam(classify.parameters(), lr=0.01 ) #, momentum=0.9)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.01 ) #, momentum=0.9)
# + id="h0mjWiFG-GDl"
def my_cross_entropy(output,target,alpha):
criterion = nn.CrossEntropyLoss(reduce=False)
batch = output.size(0)
#print(batch)
patches = output.size(1)
classes = output.size(2)
output = torch.reshape(output,(batch*patches,classes))
target = target.repeat_interleave(patches)
loss = criterion(output,target)
#print(loss,loss.shape)
loss = torch.reshape(loss,(batch,patches))
#print(loss.size())
final_loss = torch.sum(torch.mul(loss,alpha),dim=1)
#print(final_loss.shape)
final_loss = torch.mean(final_loss,dim=0)
#print(final_loss)
return final_loss
# + id="pjD2VZuV9Ed4"
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
# + id="eEVrBg7d-GDl"
def plot_attended_data(trainloader,net,epoch):
attd_data =[]
lbls = []
for data in trainloader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to(device),labels.to(device), fore_idx.to(device)
alphas, avg_images = focus_net(inputs)
attd_data.append(avg_images.numpy())
lbls.append(labels)
attd_data = np.concatenate(attd_data,axis=0)
lbls = np.concatenate(lbls,axis=0)
plt.figure(figsize=(6,8))
plt.scatter(attd_data[:,0],attd_data[:,1],c=lbls)
plt.title("EPOCH_"+str(epoch))
# + id="uALi25pmzQHV" colab={"base_uri": "https://localhost:8080/"} outputId="80b21f6d-d052-459e-f80c-9e592cfec958"
correct = 0
total = 0
count = 0
flag = 1
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to(device),labels.to(device), fore_idx.to(device)
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
loss = my_cross_entropy(outputs,labels,alphas)
print(loss)
# print(outputs.shape)
_, predicted = torch.max(outputs.data, 1)
print("="*100)
# + id="4vmNprlPzTjP"
correct = 0
total = 0
count = 0
flag = 1
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to(device),labels.to(device), fore_idx.to(device)
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# + id="_nvicAzw-GDm" colab={"base_uri": "https://localhost:8080/"} outputId="78c5058b-1733-4ed8-ddc0-e664157d1446"
for param in focus_net.named_parameters():
print(param)
# + id="Yl41sE8vFERk" colab={"base_uri": "https://localhost:8080/"} outputId="eac683c7-3f5a-4e10-d709-0f988e31f8fe"
nos_epochs = 2000
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
batch = inputs.size(0)
inputs, labels = inputs.to(device), labels.to(device)
inputs = inputs.double()
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = my_cross_entropy(outputs, labels,alphas)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 40
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if(np.mean(epoch_loss) <= 0.01):
break;
#plot_attended_data(train_loader,focus_net,epoch)
print('Finished Training')
# + id="3xPsiBtU-GDn" colab={"base_uri": "https://localhost:8080/"} outputId="52064a85-a27e-459c-9ef7-1578ab06042d"
for param in focus_net.named_parameters():
print(param)
# + id="jhvhkEAyeRpt" colab={"base_uri": "https://localhost:8080/"} outputId="d26f93aa-05b9-4d93-c9c4-755fa52570f7"
correct = 0
total = 0
focus_net.eval()
classify.eval()
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to(device), labels.to(device)
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
#print(outputs.shape)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the train images: %f %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="OKcmpKwGeS8M" colab={"base_uri": "https://localhost:8080/"} outputId="ce40f96c-e2c4-4e1f-a44e-80ec30d6e84d"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to(device), labels.to(device)
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the test images: %f %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="Dsa6zNVf75Yr"
| 15,109 |
/python/sklearnbook01.ipynb | 4db8ebbd76c20170c5248f260dcab19f49efd8e2 | [] | no_license | terazawa/study | https://github.com/terazawa/study | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 66,313 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# * https://github.com/amueller/introduction_to_ml_with_python
# * https://github.com/amueller/mglearn
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
import matplotlib.pyplot as plt
import seaborn
seaborn.set(style='ticks', context='talk', font='Osaka')
seaborn.set_palette("tab10", n_colors=24)
plt.rcParams['figure.figsize'] = [10.0, 7.0]
import numpy as np
x= np.array([[1,2,3], [4,5,6]])
print("x: \n{}".format(x))
from scipy import sparse
eye = np.eye(4)
print("NumPy array:\n{}".format(eye))
sparse_matrix = sparse.csr_matrix(eye)
print(sparse_matrix)
data = np.ones(4)
row_indices = np.arange(4)
col_indices = np.arange(4)
eye_coo = sparse.coo_matrix((data, (row_indices, col_indices)))
print(eye_coo)
x = np.linspace(-10, 10, 100)
y = np.sin(x)
plt.plot(x, y, marker='X')
import pandas as pd
data = {
'Name' : ["John", "Anna", "Peter", "Linda"],
'Location' : ["New Yowrk", "Paris", "Berlin", "London"],
'Age' : [24, 13, 53, 33]
}
df = pd.DataFrame(data)
df
df[df.Age > 30]
from sklearn.datasets import load_iris
iris_dataset = load_iris()
iris_dataset.keys()
print(iris_dataset['DESCR'][:193])
print(iris_dataset['target_names'])
print(iris_dataset['feature_names'])
x = iris_dataset['data']
print(type(x), x.shape)
pd.DataFrame(iris_dataset['data'][:5])
x = iris_dataset['target']
print(type(x), x.shape)
print(iris_dataset['target'])
print(iris_dataset['target_names'])
print(iris_dataset.target_names[iris_dataset.target])
# * https://docs.scipy.org/doc/numpy-1.13.0/user/basics.indexing.html
| 1,824 |
/machinelearning.ipynb | d3d01cdfc35b748993030987248727e997bc8740 | [] | no_license | eliasmaia/machinelearning-alura | https://github.com/eliasmaia/machinelearning-alura | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 4,818 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
##LECTURA DEL FICHERO SPSS Y SUS METADATOS
##Debe estar instalado el paquete de pyreadstat que permite leer ficheros SPSS y SAS
import pyreadstat
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from factor_analyzer import FactorAnalyzer
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn import preprocessing
# %matplotlib inline
matplotlib.style.use('ggplot')
pd.options.display.max_colwidth = 500
pd.options.display.max_rows = 999
pd.options.display.max_columns = 9999
# +
#d1 = pd.read_csv('DatosOtrasFuentes.csv',sep=';')
d2 = pd.read_csv('PCAReguntasESC.csv')
d2 = d2.reindex(sorted(d2.columns), axis=1)
d3 = pd.read_csv('PCAReguntasPRO.csv')
d3 = d3.reindex(sorted(d3.columns), axis=1)
d4 = pd.read_csv('PCAReguntasEST.csv')
d4 = d4.reindex(sorted(d4.columns), axis=1)
# -
d2[d2['CNTSCHID']==72400134]
datos = pd.merge(d2, d3,on='CNTSCHID')
datos2 = pd.merge(datos, d4,on='CNTSCHID')
datos2
# +
#datos2.to_csv('Datosmod.csv')
# -
min_max_scaler = preprocessing.MinMaxScaler()
#datos['CNTSCHID'] = d1['CNTSCHID']
#datos['CNTSTUID'] = d1['CNTSTUID']
#datos['LECTURA1'] = min_max_scaler.fit_transform(d1['LECTURA1'].values.reshape(-1, 1))*100
#datos['MATES1'] = min_max_scaler.fit_transform(d1['MATES1'].values.reshape(-1, 1))*100
#datos['CIENCIA1'] = min_max_scaler.fit_transform(d1['CIENCIA1'].values.reshape(-1, 1))*100
datos2['GLOBAL1'] = min_max_scaler.fit_transform(datos2['GLOBAL1'].values.reshape(-1, 1))*100
#datos['LECTURA2'] = min_max_scaler.fit_transform(d1['LECTURA2'].values.reshape(-1, 1))*100
#datos['MATES2'] = min_max_scaler.fit_transform(d1['MATES2'].values.reshape(-1, 1))*100
#datos['CIENCIA2'] = min_max_scaler.fit_transform(d1['CIENCIA2'].values.reshape(-1, 1))*100
datos2['GLOBAL2'] = min_max_scaler.fit_transform(datos2['GLOBAL2'].values.reshape(-1, 1))*100
#datos = datos.groupby(['CNTSCHID']).mean()
pd.DataFrame(d2[d2['CNTSCHID']==72400134].columns).to_csv('col')
#pd.DataFrame(d4[d4['CNTSCHID']==72400134].columns).to_csv('col')
#pd.DataFrame(d3[d3['CNTSCHID']==72400134].columns).to_csv('col')
dataMod = datos2.drop(['GLOBAL1', 'GLOBAL2','Unnamed: 0','CNTSCHID', 'SC167Q06HA_0'],axis=1)
dataMod
pd.DataFrame(d4[d4['CNTSCHID']==72400134].columns).sort_values(0).to_csv('col')
# +
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.preprocessing import StandardScaler
std_sc = StandardScaler()
#x = std_sc.fit_transform(dataMod.values)
x = dataMod.values
y = datos2['GLOBAL2'].values
#y = data2['LECTURA1'].values
#y = data2['MATES1'].values
#y = data['CIENCIA1'].values
#y = data['GLOBAL2'].values
#y = data['LECTURA2'].values
#y = data['MATES2'].values
#y = data['CIENCIA2'].values
lr_m = Ridge()
#lr_m = ElasticNet()
#ajuste sobre la muestra completa, sin split train-test
lr_m.fit(x, y)
print("model intercept: %f" % lr_m.intercept_)
nticks = len(lr_m.coef_)
plt.xticks(range(len(dataMod.columns.to_list())), dataMod.columns.to_list(), rotation=45)
plt.title('Linear Regression coefs')
_ = plt.plot(lr_m.coef_.T, '-', lr_m.coef_.T, 'r*')
# -
#Dataframe con el coeficiente y nombre columna
VarCoef = pd.DataFrame(lr_m.coef_.T,dataMod.columns.to_list()).sort_values(0)
VarCoef ['Cols']=VarCoef.index
VarCoef['GRUPO'] = str(VarCoef['Cols'].values).split('_')[0]
str(VarCoef['Cols'].values).split('_')
VarCoef2 = VarCoef.sort_values('Cols')
a =[]
for i in VarCoef['Cols']:
print(i.split('_'))
a.append(i.split('_')[0])
a.sort()
VarCoef2['Grupo'] = a
VarCoef2.groupby('Grupo').sum()
# +
from sklearn.metrics import mean_squared_error, mean_absolute_error
y_pred = np.clip(lr_m.predict(x), 0, 100.)
print("MAE: %.3f\tR^2: %.3f" % (mean_absolu | 4,096 |
/Python Bootcamp From Zero to Hero in Python/01. Python Object and Data Structure Basics/04 - Print Formatting with Strings.ipynb | 4b3e2ca2fba1efb9f8cc06254243d834b65c0437 | [] | no_license | nicorema/Backend-Trainee | https://github.com/nicorema/Backend-Trainee | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 2,447 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print('Hello Sr {}'.format('Nicolás'))
print('The {2} {1} is {0}'.format('running','fox','red'))
print('The {color} {animal} is {action}'.format(action='running',animal='fox',color='red'))
result = 100/3
print('The result is {r:1.3f}'.format(r=result))
name = 'Nicolás'
title='Sr.'
print(f'Hello {title} {name}')
| 588 |
/.ipynb_checkpoints/projeto-checkpoint.ipynb | 3ff75a85b2a1fd902701eeaeacca50482ec8252c | [] | no_license | Gabriel1955/Using_pytesseract_identify_in_PCB | https://github.com/Gabriel1955/Using_pytesseract_identify_in_PCB | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 166,327 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#primeira abordagem
import numpy as np
import cv2
import cv2 as cv
import matplotlib.pyplot as plt
img = cv.imread('placa.jpg')
img = cv.medianBlur(img,5)
for y in range(0, img.shape[0], 1): #percorre as linhas
for x in range(0, img.shape[1], 1): #percorre as colunas imagem[y, x] = (0,(x*y)%256,0)
(b, g ,r) = img[y,x]
media = (np.float16(b) + np.float16(g) +np.float16(r))/3
erro = abs(media - b)+abs(media - g)+abs(media - r)
# print(erro)
if erro < 36 and media > 120:
img[y,x]= (255,255,255)
else:
img[y,x]=(0,0,0)
plt.imshow(img)
plt.show()
cv2.imwrite("saida3.jpg", img)
# -
# +
#segunda abordagem
import cv2 as cv
import numpy as np
from matplotlib import pyplot as plt
from skimage.util import random_noise
i = 1
while i < 15:
img = cv.imread("p/placa"+str(i)+".jpg",0)
img = cv.medianBlur(img,5)
th3 = cv.adaptiveThreshold(img,255,cv.ADAPTIVE_THRESH_GAUSSIAN_C,\
cv.THRESH_BINARY,11,-2)
printImg = th3
plt.imshow(printImg)
plt.show()
cv2.imwrite("p/saida"+str(i)+".jpg", printImg)
i = i+1
# +
#segunda abordagem
import cv2 as cv
import numpy as np
from matplotlib import pyplot as plt
from skimage.util import random_noise
img = cv.imread('placa2.jpg',0)
img = cv.medianBlur(img,5)
th3 = img
vis = th3.copy()
# detect regions in gray scale image
mser = cv2.MSER_create(-2, 200, 600, 5, 0.2, 200, 1.01, 0.003, 5)
regions, _ = mser.detectRegions(th3)
hulls = [cv2.convexHull(p.reshape(-1, 1, 2)) for p in regions]
cv2.polylines(vis, hulls, 1, (0, 255, 0))
# teste
mask = np.zeros((th3.shape[0], th3.shape[1], 1), dtype=np.uint8)
for contour in hulls:
cv2.drawContours(mask, [contour], -1, (255 , 255, 255), -1)
#this is used to find only text regions, remaining are ignored
text_only = cv2.bitwise_and(th3, th3, mask=mask)
printImg = text_only
plt.imshow(printImg)
plt.show()
cv2.imwrite("saida4.jpg", printImg)
# +
import cv2
import pytesseract
import numpy as np
from matplotlib import pyplot as plt
img = cv2.imread('saida1.jpg')
h, w, c = img.shape
boxes = pytesseract.image_to_boxes(img)
for b in boxes.splitlines():
b = b.split(' ')
img = cv2.rectangle(img, (int(b[1]), h - int(b[2])), (int(b[3]), h - int(b[4])), (0, 255, 0), 2)
printImg = img
plt.imshow(printImg)
plt.show()
cv2.imwrite("saida5.jpg", printImg)
# +
import cv2
import pytesseract
from pytesseract import Output
img = cv2.imread('saida3.jpg')
d = pytesseract.image_to_data(img, output_type=Output.DICT)
print(d.keys())
n_boxes = len(d['text'])
for i in range(n_boxes):
if int(d['conf'][i]) > 60:
(x, y, w, h) = (d['left'][i], d['top'][i], d['width'][i], d['height'][i])
img = cv2.rectangle(img, (x, y), (x + w, y + h), (0, 255, 0), 2)
printImg = img
plt.imshow(printImg)
plt.show()
cv2.imwrite("saida6.jpg", printImg)
# +
import pytesseract
from PIL import Image
import numpy as np
import cv2
img = cv2.imread('saida3.jpg')
(T, img) = cv2.threshold(img, 190, 255, cv2.THRESH_BINARY)
img = cv2.medianBlur(img, 7)
text = pytesseract.image_to_string(img, lang='myFont2')
print(text)
# +
#segunda abordagem
import cv2 as cv
import numpy as np
from matplotlib import pyplot as plt
from skimage.util import random_noise
i = 14
while i < 15:
img = cv.imread("capture.png",0)
#th3 = cv.adaptiveThreshold(img,255,cv.ADAPTIVE_THRESH_GAUSSIAN_C,\
# cv.THRESH_BINARY,111,10)
(T, img) = cv.threshold(img, 190, 255, cv.THRESH_BINARY)
img = cv.medianBlur(img,7)
printImg = img
plt.imshow(printImg)
plt.show()
cv.imwrite("saida_capture.png", printImg)
i = i+1
# -
'.format(decay[i]))
plt.ylim([0, 1.1])
plt.legend(loc='best')
plt.show()
# -
# # Result
# 衰減越大,學習率衰減地越快。
# 衰減確實能夠對震盪起到減緩的作用
# # Momentum (動量)
# 如何用“動量”來解決:
#
# (1)學習率較小時,收斂到極值的速度較慢。
#
# (2)學習率較大時,容易在搜索過程中發生震盪。
#
# 當使用動量時,則把每次w的更新量v考慮為本次的梯度下降量 (-dx*lr), 與上次w的更新量v乘上一個介於[0, 1]的因子momentum的和
#
#
# w ← x − α ∗ dw (x沿負梯度方向下降)
#
# v = ß ∗ v − α ∗ d w
#
# w ← w + v
#
# (ß 即momentum係數,通俗的理解上面式子就是,如果上一次的momentum(即ß )與這一次的負梯度方向是相同的,那這次下降的幅度就會加大,所以這樣做能夠達到加速收斂的過程
#
# 如果上一次的momentum(即ß )與這一次的負梯度方向是相反的,那這次下降的幅度就會縮減,所以這樣做能夠達到減速收斂的過程
#
#
# +
line_x = np.linspace(-5, 5, 100)
line_y = func(line_x)
plt.figure('Gradient Desent: Decay')
'''
x= w_init
epochs = 10
lr = [.......]
decay = [.......]
'''
x_start = 5
lr = [.1,.01,.001]
color = ['k', 'r', 'g', 'y']
row = len(lr)
col = len(decay)
size = np.ones(epochs + 1) * 10
size[-1] = 70
plt.figure(figsize = (15,6))
for i in range(row):
for j in range(col):
x = GD_decay(x_start, dfunc, epochs, lr=lr[i], decay=decay[j])
plt.subplot(row, col, i * col + j + 1)
plt.plot(line_x, line_y, c='b')
plt.plot(x, func(x), c=color[i], label='lr={}, de={}'.format(lr[i], decay[j]))
plt.scatter(x, func(x), c=color[i], s=size)
plt.legend(loc=0)
plt.show()
# -
| 5,205 |
/APetryashev_Class_HW.ipynb | 05b219286dfd03b5c533f397c29813d045dda297 | [] | no_license | apetryashev/PYDA_18_HW | https://github.com/apetryashev/PYDA_18_HW | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 14,844 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Домашнее задание к лекции «Понятие класса»
# А.Петряшев (DS-28)
import requests
import json
valute = requests.get('https://www.cbr-xml-daily.ru/daily_json.js')
# Задание 1.
#
# Напишите функцию, которая возвращает название валюты (поле ‘Name’) с максимальным значением курса с помощью сервиса https://www.cbr-xml-daily.ru/daily_json.js
#
#
# Не совсем понял, какое максимальное значение нужно получить, поэтому сделал 2 решения.
# 1.1.
# Максимальное значение из предыдущего и текущего курсов конкретной валюты или всех валют сервиса.
# +
user_input = input()
def valute_max(valuta=''):
currency = valute.json()['Valute']
for val, des in currency.items():
if valuta != '':
if val == valuta.upper():
name = des['Name']
max_v = max([des['Value'], des['Previous']])
print(name, max_v)
else: pass
else:
name = des['Name']
max_v = max([des['Value'], des['Previous']])
print(name, max_v)
valute_max(user_input)
# -
# 1.2.
# Максимальное значение текущего курса валюты из всех валют сервиса.
# +
def max_value():
cur = valute.json()['Valute']
all_rates ={}
for val, des in cur.items():
current_rate = des['Value']
val_name = des['Name']
all_rates[current_rate] = val_name
#print(all_rates)
max_rate = max(all_rates)
name = all_rates[max_rate]
print(name, max_rate)
max_value()
# -
# Задание 2.
#
# Добавьте в класс Rate параметр diff (со значениями True или False), который в случае значения True в методах курсов валют (eur, usd итд) будет возвращать не курс валюты, а изменение по сравнению в прошлым значением. Считайте, self.diff будет принимать значение True только при возврате значения курса. При отображении всей информации о валюте он не используется.
#
class Rate:
def __init__(self, format_='value', diff=True):
self.format = format_
self.diff = diff
def exchange_rates(self):
"""
Возвращает ответ сервиса с информацией о валютах в виде:
{
'AMD': {
'CharCode': 'AMD',
'ID': 'R01060',
'Name': 'Армянских драмов',
'Nominal': 100,
'NumCode': '051',
'Previous': 14.103,
'Value': 14.0879
},
...
}
"""
self.r = requests.get('https://www.cbr-xml-daily.ru/daily_json.js')
return self.r.json()['Valute']
def make_format(self, currency):
"""
Возвращает информацию о валюте currency в вариантах:
- полная информация о валюте при self.format = 'full':
Rate('full').make_format('EUR')
{
'CharCode': 'EUR',
'ID': 'R01239',
'Name': 'Евро',
'Nominal': 1,
'NumCode': '978',
'Previous': 79.6765,
'Value': 79.4966
}
Rate('value').make_format('EUR')
79.4966
если diff=True - находится разница значений 'Value'-'Previous' и выводится вместо курса валюты,
в противном случае выводится текущий курс;
diff не используется при выводе поной информации о валюте
"""
response = self.exchange_rates()
if currency in response:
if self.format == 'full':
return response[currency]
if self.format == 'value':
if self.diff:
return response[currency]['Value'] - response[currency]['Previous']
else:
return response[currency]['Value']
return 'Error'
def eur(self):
"""Возвращает курс евро на сегодня в формате self.format"""
return self.make_format('EUR')
def usd(self):
"""Возвращает курс доллара на сегодня в формате self.format"""
return self.make_format('USD')
def brl(self):
"""Возвращает курс бразильского реала на сегодня в формате self.format"""
return self.make_format('BRL')
r = Rate(format_='value', diff=True)
r.eur()
# Задание 3.
#
# Напишите класс Designer, который учитывает количество международных премий для дизайнеров (из презентации: “Повышение на 1 грейд за каждые 7 баллов. Получение международной премии – это +2 балла”). Считайте, что при выходе на работу сотрудник уже имеет две премии и их количество не меняется со стажем (конечно если хотите это можно вручную менять).
#
# Класс Designer пишется по аналогии с классом Developer из материалов занятия. Комментарий про его условия Вика написала выше: “Повышение на 1 грейд за каждые 7 баллов. Получение международной премии – это +2 балла”
class Employee:
def __init__(self, name, seniority):
self.name = name
self.seniority = seniority
self.grade = 1
def grade_up(self):
"""Повышает уровень сотрудника"""
self.grade += 1
def publish_grade(self):
"""Публикация результатов аккредитации сотрудников"""
print(self.name, self.grade)
def check_if_it_is_time_for_upgrade(self):
pass
# Для решения предположил, что премиии могут быть в любой год.
# Так как балов за премию дают 2, а повышение грейда идет за каждые 7 баллов, то постарался реализовать такую логику:
#
# количество премий в год выбирается случайно, но не более одной в год - random.randint(0,1);
#
# сначала повышается грейд за стаж, затем за балллы - учитывается накопление баллов и шкала для повышения (7, 14, 21,...).
import random
# +
class Designer(Employee):
def __init__(self, name, seniority, raiting = 4, prom_raiting=[0]):
super().__init__(name, seniority)
self.raiting = raiting
self.prom_raiting = prom_raiting
def check_if_it_is_time_raiting_for_upgrade(self):
"""
prize_points = random.randint(0,1)*2 - подсчет квалификационных баллов за примемии, случайное определение кол-ва премий
diff = self.raiting - self.prom_raiting[-1] - разница баллов последнего повышения и текущих
points_for_grade = (self.raiting // 7) * 7 - баллы за которые должны повышаться грейды
"""
prize_points = random.randint(0,1)*2
self.raiting +=prize_points
self.seniority += 1
diff = self.raiting - self.prom_raiting[-1]
points_for_grade = (self.raiting // 7) * 7
if self.seniority % 5 == 0:
self.grade_up()
elif diff >= 7 or self.raiting % 7 == 0:
self.prom_raiting.append(points_for_grade)
self.grade_up()
#print('баллов за премию: ', prize_points, '//', 'всего баллов: ', self.raiting, '//', 'баллы на момент прома: ', self.prom_raiting, '//', 'разница: ', diff)
return self.publish_grade()
# -
alex= Designer('alex', 0)
for i in range(10):
alex.check_if_it_is_time_raiting_for_upgrade()
# такую логику использовал для повышения грейда
# +
t = 2
k =[0]
seniority = 0
for i in range(0,20):
seniority +=1
t+=2
d = t - k[-1] #разница баллов последнего повышения и текущих
p_p = (t//7)*7 #баллы за которые должны повышаться грейды
if seniority % 5 == 0:
print('grade up +5year', 'in company', seniority, 'year', '//', p_p)
elif d >= 7 or t % 7 == 0:
k.append(p_p)
print('grade up +7points', t, 'overall points', '//', p_p)
else: print('still', t)
print(k)
| 7,947 |
/06_Pandas/01_Series/10_Series_abs.ipynb | 603afad7f0911cfba2ab3f799ff7797ff973fb95 | [] | no_license | smahaboob2/Data_Science | https://github.com/smahaboob2/Data_Science | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 3,714 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# importing pandas module
import pandas as pd
# creating lists
lst = [2, -10.87, -3.14, 0.12]
lst2 = [-10.87 + 4j]
ser = pd.Series(lst)
ser1 = pd.Series(lst2)
# printing values explaining abs()
print(ser1.abs(), '\n\n', ser.abs())
# +
# importing pandas module
import pandas as pd
df = pd.DataFrame({'Name': ['John', 'Hari', 'Peter', 'Loani'],
'Age': [31, 29, 57, 40],
'val': [98, 48, -80, -14]})
df['ope'] = (df.val - 87).abs()
df
# -
| 786 |
/week8/lab8-with-code.ipynb | 0622fd9b79564ec1382e9bb386cf8eec4feafd28 | [] | no_license | keqpan/skoltech-gcv-course-2021 | https://github.com/keqpan/skoltech-gcv-course-2021 | 9 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 1,969,060 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: lab9
# language: python
# name: lab9
# ---
import meshplot as mp
import numpy as np
import igl
import yaml
from yaml import CLoader as Loader
# ## Reading CAD Data
# +
def read_model(obj_path, feat_path):
v, _, n, f, _, ni = igl.read_obj(obj_path)
with open(feat_path) as fi:
feat = yaml.load(fi, Loader=Loader)
m = {"vertices": v, "face_indices": f, "normals": n,
"normal_indices": ni, "features": feat}
return m
m = read_model("data/test_trimesh.obj", "data/test_features.yml")
v, f, feat = m["vertices"], m["face_indices"], m["features"]
print(v.shape, f.shape)
print(list(feat.keys()))
# -
# ## CAD Features: Surface Normals
# +
from data.utils import get_averaged_normals
# Average normals at vertices with multiple normals
av_normals = get_averaged_normals(m)
p = mp.plot(v, f, c=np.abs(av_normals))
# Add normals to the plot
p.add_lines(m["vertices"], m["vertices"] + av_normals,
shading={"line_color": "black"})
# Determine normals with uniform weighting in libigl
normals = igl.per_vertex_normals(v, f)
p.add_lines(m["vertices"], m["vertices"] + normals,
shading={"line_color": "red"})
# -
# ## CAD Features: Sharp Edges/Curves
feat["curves"]
# Retrieve the sharp features
lines = []
for i, fe in enumerate(feat["curves"]):
if fe["sharp"]:
for j in range(len(fe['vert_indices'])-1):
lines.append([fe['vert_indices'][j], fe['vert_indices'][j+1]])
################################
### your code here
################################
lines
# Visualize the sharp features
p = mp.plot(v, f)
p.add_edges(v, np.array(lines))
# ## CAD Features: Sharp Edges/Curves¶
# +
# Retrieve the sharp features
v_class = np.zeros((v.shape[0], 1))
for i, fe in enumerate(feat["curves"]):
if fe["sharp"]:
v_class[fe['vert_indices']]=1
################################
### your code here
################################
# Visualize the sharp features
mp.plot(v, c=-v_class, shading={"point_size": 2.})
# -
# ## CAD Features: Surface Patch Types
feat["surfaces"]
# +
# Retrieve the surface patch types
t_map = {"Plane": 0, "Cylinder": 1,
"Cone": 2, "Sphere": 3,
"Torus": 4, "Bezier": 5,
"BSpline": 6, "Revolution": 7,
"Extrusion": 8, "Other": 9}
c1 = np.zeros(f.shape[0])
for fe in feat["surfaces"]:
t = t_map[fe["type"]]
for j in fe['face_indices']:
c1[j] = t
################################
### your code here
################################
# Visualize the patch types
mp.plot(v, f, -c1)
# -
# ## CAD Features: Surface Patch Types¶
# +
# Retrieve the surface patch types per vertex
c2 = np.zeros(v.shape[0])
for fe in feat["surfaces"]:
t = t_map[fe["type"]]
for j in fe['vert_indices']:
c2[j] = t
################################
### your code here
################################
# Visualize the vertices
mp.plot(v, c=-c2, shading={"point_size": 2.})
# -
# # Machine Learning Setup
# +
import numpy as np
import meshplot as mp
import torch
import torch.nn.functional as F
from torch.nn import Sequential, Dropout, Linear
import torch_geometric.transforms as T
from torch_geometric.data import DataLoader
from torch_geometric.nn import DynamicEdgeConv
from data.utils import MLP
from data.utils import ABCDataset
# -
# ## Loading the CAD Data
# +
tf_train = T.Compose([
T.FixedPoints(5000, replace=False),
T.RandomTranslate(0.002),
T.RandomRotate(15, axis=0),
T.RandomRotate(15, axis=1),
T.RandomRotate(15, axis=2)
])
tf_test = T.Compose([T.FixedPoints(10000, replace=False)])
pre = T.NormalizeScale()
train_dataset_n = ABCDataset("data/ml/ABC", "Normals", True, tf_train, pre)
test_dataset_n = ABCDataset("data/ml/ABC", "Normals", False, tf_test, pre)
train_dataset_e = ABCDataset("data/ml/ABC", "Edges", True, tf_train, pre)
test_dataset_e = ABCDataset("data/ml/ABC", "Edges", False, tf_test, pre)
train_dataset_t = ABCDataset("data/ml/ABC", "Types", True, tf_train, pre)
test_dataset_t = ABCDataset("data/ml/ABC", "Types", False, tf_test, pre)
# -
# ## Statistics and Visualization
# +
dataset = train_dataset_n
print("Number of models:", len(dataset))
print("Number of classes:", dataset.num_classes)
counts = [0]*dataset.num_classes
total = 0
for d in dataset:
y = d.y.numpy()
for i in range(dataset.num_classes):
counts[i] += np.sum(y==i)
total += y.shape[0]
for i, c in enumerate(counts):
print("%0.2f%% labels are of class %i."%(c/total, i))
# +
d = test_dataset_e[3]
v = d.pos.numpy()
y = d.y.numpy()
print("Shape of model:", v.shape)
print("Shape of labels:", y.shape)
mp.plot(v, c=-y, shading={"point_size": 0.1})
# -
d = train_dataset_n[8]
v = d.pos.numpy()
y = d.y.numpy()
mp.plot(v, c=np.abs(y), shading={"point_size": 0.1})
d = train_dataset_t[8]
v = d.pos.numpy()
y = d.y.numpy()
mp.plot(v, c=-y, shading={"point_size": 0.1})
# ## Defining the Network (DGCNN)
class Net(torch.nn.Module):
def __init__(self, out_channels, k=30, aggr='max',
typ='Edges'):
super(Net, self).__init__()
self.typ = typ
self.conv1 = DynamicEdgeConv(MLP([2 * 3, 64, 64]), k, aggr)
self.conv2 = DynamicEdgeConv(MLP([2* 64, 64, 64]), k, aggr)
self.conv3 = DynamicEdgeConv(MLP([2* 64, 64, 64]), k, aggr)
self.lin1 = MLP([3 * 64, 1024])
self.mlp = Sequential(MLP([1024, 256]), Dropout(0.5),
MLP([256, 128]), Dropout(0.5),
Linear(128, out_channels))
def forward(self, data):
pos, batch = data.pos, data.batch
x1 = self.conv1(pos, batch)
x2 = self.conv2(x1, batch)
x3 = self.conv3(x2, batch)
out = self.lin1(torch.cat([x1, x2, x3], dim=1))
out = self.mlp(out)
if self.typ == "Edges" or self.typ == "Types":
return F.log_softmax(out, dim=1)
if self.typ == "Normals":
return F.normalize(out, p=2, dim=-1)
# ## Visualizing the Nearest Neighbour Graph
# +
tf_pre = T.Compose([
T.FixedPoints(1000),
T.NormalizeScale(),
T.KNNGraph(k=6)
])
dataset = ABCDataset("data/ml/ABC_graph", "Edges", pre_transform=tf_pre)
vd = dataset[0].pos.numpy()
p = mp.plot(vd, shading={"point_size": 0.1})
p.add_edges(vd, dataset[0].edge_index.numpy().T)
# -
# ## Defining the Loss Function (Normals)
# +
class Cosine_Loss(torch.nn.Module):
def __init__(self):
super(Cosine_Loss,self).__init__()
def forward(self, x, y):
dotp = torch.mul(x,y).sum(1)
loss = torch.sum(1 - dotp.pow(2))/x.shape[0]
angle = torch.sum(torch.acos(torch.clamp(torch.abs(dotp), 0.0, 1.0)))/x.shape[0]
##################### ###########
### your code here
################################
return loss, angle
cosine_loss = Cosine_Loss()
# -
# ## Defining the Training Procedure
def train(loader, typ="Edges"):
model.train()
for i, data in enumerate(loader):
total_loss = correct_nodes = total_nodes = 0
data = data.to(device)
optimizer.zero_grad()
out = model(data)
if typ == "Edges" or typ == "Types":
loss = F.nll_loss(out, data.y)
if typ == "Normals":
loss, angle = cosine_loss(out, data.y)
loss.backward()
optimizer.step()
total_loss += loss.item()
if typ == "Edges" or typ == "Types":
pred = out.max(dim=1)[1]
correct_nodes += pred.eq(data.y).sum().item()
total_nodes += data.num_nodes
acc = correct_nodes / total_nodes
if typ == "Normals":
acc = angle.item()*180/np.pi
print('[Train {}/{}] Loss: {:.4f}, Accuracy: {:.4f}'.format(
i + 1, len(loader), total_loss / loader.batch_size, acc))
# ## Defining the Testing Procedure
def test(loader, typ="Edges"):
model.eval()
correct_nodes = total_nodes = 0
for data in loader:
data = data.to(device)
with torch.no_grad():
out = model(data)
if typ == "Edges" or typ == "Types":
pred = out.max(dim=1)[1]
correct_nodes += pred.eq(data.y).sum().item()
total_nodes += data.num_nodes
if typ == "Normals":
_, angle = cosine_loss(out, data.y)
correct_nodes += angle.item() * 180 / np.pi
total_nodes += 1
return correct_nodes / total_nodes
# ## Running the Training
# +
typ = "Edges"
if typ == "Edges":
train_dataset = train_dataset_e
test_dataset = test_dataset_e
if typ == "Normals":
train_dataset = train_dataset_n
test_dataset = test_dataset_n
if typ == "Types":
train_dataset = train_dataset_t
test_dataset = test_dataset_t
train_loader = DataLoader(train_dataset, batch_size=10, shuffle=True)
test_loader = DataLoader(test_dataset, batch_size=1, shuffle=False)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model = Net(train_dataset.num_classes, k=30, typ=typ).to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
scheduler = torch.optim.lr_scheduler.StepLR(optimizer,
step_size=20, gamma=0.8)
for epoch in range(1, 2):
train(train_loader, typ=typ)
acc = test(test_loader, typ=typ)
print('Test: {:02d}, Accuracy: {:.4f}'.format(epoch, acc))
torch.save(model.state_dict(), "%02i_%.2f.dat"%(epoch, acc))
scheduler.step()
# -
# ## Loading a Pretrained Model - Edges
# +
typ = "Edges"
test_dataset = test_dataset_e
state_file = "Edges_72_0.96.dat"
#typ = "Normals"
#test_dataset = test_dataset_n
#state_file = "Normals_44_12.52.dat"
#typ = "Types"
#test_dataset = test_dataset_t
#state_file = "Types_57_0.76.dat"
model = Net(test_dataset.num_classes, k=30, typ=typ)
if torch.cuda.is_available():
state = torch.load("data/ml/ABC/models/%s"%state_file)
device = torch.device('cuda')
else:
state = torch.load("data/ml/ABC/models/%s"%state_file,
map_location=torch.device('cpu'))
device = torch.device('cpu')
model.load_state_dict(state)
model.to(device);
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
scheduler = torch.optim.lr_scheduler.StepLR(optimizer,
step_size=20, gamma=0.8)
for epoch in range(1, 2):
train(train_loader, typ=typ)
acc = test(test_loader, typ=typ)
print('Test: {:02d}, Accuracy: {:.4f}'.format(epoch, acc))
# -
# ## Visualizing the Predicted Results
test_loader = DataLoader(test_dataset, batch_size=1, shuffle=True)
loader = iter(test_loader)
# +
d = loader.next()
with torch.no_grad():
out = model(d.to(device))
v = d.pos.cpu().numpy()
y = d.y.cpu().numpy()
# Calculate accuracy
acc = test(test_loader)
print('Accuracy: {:.4f}'.format(acc))
# -
# Plot groundtruth
mp.plot(v, c=-y, shading={"point_size":0.15})
e = out.max(dim=1)[1].cpu().numpy()
# Plot estimation
mp.plot(v, c=-e, shading={"point_size": 0.15})
# ## Loading a Pretrained Model - Normals
# +
typ = "Normals"
test_dataset = test_dataset_n
state_file = "Normals_44_12.52.dat"
model = Net(test_dataset.num_classes, k=30, typ=typ)
if torch.cuda.is_available():
state = torch.load("data/ml/ABC/models/%s"%state_file)
device = torch.device('cuda')
else:
state = torch.load("data/ml/ABC/models/%s"%state_file,
map_location=torch.device('cpu'))
device = torch.device('cpu')
model.load_state_dict(state)
model.to(device);
# -
# ## Visualizing the Predicted Results
test_loader = DataLoader(test_dataset, batch_size=1, shuffle=True)
loader = iter(test_loader)
# +
d = loader.next()
with torch.no_grad():
out = model(d.to(device))
v = d.pos.cpu().numpy()
y = d.y.cpu().numpy()
# Calculate accuracy
_, angle = cosine_loss(out, d.y)
print(angle.item() * 180 / np.pi)
# -
# Plot groundtruth
c1 = np.abs(y)
mp.plot(v, c=c1, shading={"point_size":0.15})
n = out.cpu().numpy()
c2 = np.abs(n)
# Plot estimation
mp.plot(v, c=c2, shading={"point_size": 0.15})
| 12,342 |
/05-Object Oriented Programming/02-Object Oriented Programming Homework.ipynb | 8064df30d7b5280ee16b5044e22d060c698248d7 | [] | no_license | EJAZ-IOT/COMPLETE-PYTHON3-BOOTCAMP | https://github.com/EJAZ-IOT/COMPLETE-PYTHON3-BOOTCAMP | 1 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 3,843 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Oriented Programming
# ## Homework Assignment
#
# #### Problem 1
# Fill in the Line class methods to accept coordinates as a pair of tuples and return the slope and distance of the line.
class Line:
def __init__(self,coor1,coor2):
self.coor1=coor1
self.coor2=coor2
def distance(self):
return ((self.coor2[0]-self.coor1[0])**2 + (self.coor2[1]-self.coor1[1])**2 )**0.5
def slope(self):
return ((self.coor2[1]-self.coor1[1])/(self.coor2[0]-self.coor1[0]))
# +
coordinate1 = (5,2)
coordinate2 = (2,10)
li = Line(coordinate1,coordinate2)
# -
li.distance()
li.slope()
# ________
# #### Problem 2
# Fill in the class
class Cylinder:
pi=3.14
def __init__(self,height=1,radius=1):
self.height=height
self.radius=radius
def volume(self):
return Cylinder.pi*self.radius**2*self.height
def surface_area(self):
return (2*Cylinder.pi*self.radius**2)+(2*Cylinder.pi*self.radius*self.height)
c = Cylinder(6,2)
c.volume()
c.surface_area()
ython has an [enumerate()](https://docs.python.org/3/library/functions.html#enumerate) function that can help us with this, though. The [enumerate()](https://docs.python.org/3/library/functions.html#enumerate) function allows us to have two variables in the body of a for loop -- an index, and the value.
#
# >```python
# animals = ["Dog", "Tiger", "SuperLion", "Cow", "Panda"]
# viciousness = [1, 5, 10, 10, 1]
# for i, animal in enumerate(animals):
# print("Animal")
# print(animal)
# print("Viciousness")
# print(viciousness[i])
# ```
# +
ships = ["Andrea Doria", "Titanic", "Lusitania"]
cars = ["Ford Edsel", "Ford Pinto", "Yugo"]
for i,item in enumerate(ships):
print(item)
print(cars[i])
# +
#adding columns
things = [["apple", "monkey"], ["orange", "dog"], ["banana", "cat"]]
trees = ["cedar", "maple", "fig"]
for i, item in enumerate(things):
item.append(trees[i])
print(things)
# -
# ## 2. List Comprehensions
#
# We've written many short for loops to manipulate lists. Here's an example:
#
# >```python
# animals = ["Dog", "Tiger", "SuperLion", "Cow", "Panda"]
# animal_lengths = []
# for animal in animals:
# animal_lengths.append(len(animal))
# ```
#
# It takes three lines to calculate the length of each string <span style="background-color: #F9EBEA; color:##C0392B">animals</span> this way. However, we can condense this down to one line with a list comprehension:
#
# >```python
# animal_lengths = [len(animal) for animal in animals]
# ```
#
# List comprehensions are much more compact notation, and can save space when you need to write multiple for loops.
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Use list comprehension to create a new list called <span style="background-color: #F9EBEA; color:##C0392B">apple_prices_doubled</span>, where you multiply each item in <span style="background-color: #F9EBEA; color:##C0392B">apple_prices</span> by <span style="background-color: #F9EBEA; color:##C0392B">2</span>.
# 2. Use list comprehension to create a new list called <span style="background-color: #F9EBEA; color:##C0392B">apple_prices_lowered</span>, where you subtract <span style="background-color: #F9EBEA; color:##C0392B">100</span> from each item in <span style="background-color: #F9EBEA; color:##C0392B">apple_prices</span>.
#
# >```python
# apple_prices = [100, 101, 102, 105]
# ```
# ## 3. Numpy
#
# In the previous notebooks, we used nested lists in Python to represent datasets. Python lists offer a few advantages when representing data:
#
# - lists can contain mixed types
# - lists can shrink and grow dynamically
#
# Using Python lists to represent and work with data also has a few key disadvantages:
#
# - to support their flexibility, lists tend to consume lots of memory
# - they struggle to work with medium and larger sized datasets
#
# While there are many different ways to classify programming languages, an important way that keeps performance in mind is the difference between **low-level** and **high-level** languages. Python is a high-level programming language that allows us to quickly write, prototype, and test our logic. The C programming language, on the other hand, is a low-level programming language that is highly performant but has a much slower human workflow.
#
# <span style="background-color: #F9EBEA; color:##C0392B">NumPy</span> is a library that combines the flexibility and ease-of-use of Python with the speed of C. In this mission, we'll start by getting familiar with the core NumPy data structure and then build up to using NumPy to work with the dataset <span style="background-color: #F9EBEA; color:##C0392B">world_alcohol.csv</span>, which contains data on how much alcohol is consumed per capita in each country.
#
#
# ### 3.1 Creating Arrays
#
# The core data structure in NumPy is the <span style="background-color: #F9EBEA; color:##C0392B">ndarray</span> object, which stands for **N-dimensional array**. An **array** is a collection of values, similar to a list. **N-dimensional** refers to the number of indices needed to select individual values from the object.
#
# <img width="500" alt="creating a repo" src="https://drive.google.com/uc?export=view&id=0BxhVm1REqwr0X0VuT3NoZGZ0UlU">
#
# A **1-dimensional** array is often referred to as a vector while a **2-dimensional** array is often referred to as a **matrix**. Both of these terms are both borrowed from a branch of mathematics called linear algebra. They're also often used in data science literature, so we'll use these words throughout this course.
#
# To use <span style="background-color: #F9EBEA; color:##C0392B">NumPy</span>, we first need to import it into our environment. NumPy is commonly imported using the alias <span style="background-color: #F9EBEA; color:##C0392B">np</span>:
#
# >```python
# import numpy as np
# ```
#
# We can directly construct arrays from lists using the <span style="background-color: #F9EBEA; color:##C0392B">numpy.array()</span> function. To construct a vector, we need to pass in a single list (with no nesting):
#
# >```python
# matrix = np.array([[5, 10, 15], [20, 25, 30], [35, 40, 45]])
# ```
# +
import numpy as np
vector = np.array([10, 20, 30])
matrix = np.array([[5, 10, 15], [20, 25, 30], [35, 40, 45]])
print(vector[0])
print(matrix[0])
print(matrix[0][1])
# -
# ### 3.2 Array shape
#
# It's often useful to know how many elements an array contains. We can use the [ndarray.shape](http://docs.scipy.org/doc/numpy-1.10.1/reference/generated/numpy.ndarray.shape.html) property to figure out how many elements are in the array.
#
#
# +
vector = np.array([1, 2, 3, 4])
print(vector.shape)
matrix = np.array([[5, 10, 15], [20, 25, 30]])
print(matrix.shape)
# -
# ### 3.3 Using numpy
#
# We can read in datasets using the [numpy.genfromtxt()](http://docs.scipy.org/doc/numpy-1.10.0/reference/generated/numpy.genfromtxt.html) function. Our dataset, <span style="background-color: #F9EBEA; color:##C0392B">world_alcohol.csv</span> is a comma separated value dataset. We can specify the delimiter using the delimiter parameter:
#
# >```python
# import numpy
# data = numpy.genfromtxt("data.csv", delimiter=",")
# ```
#
# **"world_alcohol.csv'**
#
# Here's what each column represents:
#
# - Year -- the year the data in the row is for.
# - WHO Region -- the region in which the country is located.
# - Country -- the country the data is for.
# - Beverage Types -- the type of beverage the data is for.
# - Display Value -- the number of liters, on average, of the beverage type a citizen of the country drank in the given year.
#
#
import numpy as np
world_alcohol = np.genfromtxt("data/world_alcohol.csv", delimiter=',')
print(type(world_alcohol))
# Each value in a NumPy array has to have the same data type. NumPy data types are similar to Python data types, but have slight differences. You can find a full list of NumPy data types [here](http://docs.scipy.org/doc/numpy-1.10.1/user/basics.types.html).
print(world_alcohol.dtype)
# ### 3.4 Inspecting the data
#
# Here's how NumPy represents the first few rows of the dataset:
#
# >```python
# array([[ nan, nan, nan, nan, nan],
# [ 1.98600000e+03, nan, nan, nan, 0.00000000e+00],
# [ 1.98600000e+03, nan, nan, nan, 5.00000000e-01]])
# ```
world_alcohol
# There are a few concepts we haven't been introduced to yet that we'll dive into into:
#
# - Many items in <span style="background-color: #F9EBEA; color:##C0392B">world_alcohol</span> are <span style="background-color: #F9EBEA; color:##C0392B">nan</span>, including the entire first row. <span style="background-color: #F9EBEA; color:##C0392B">nan</span>, which stands for **"not a number"**, is a data type used to represent missing values.
# - Some of the numbers are written like <span style="background-color: #F9EBEA; color:##C0392B">1.98600000e+03</span>.
#
# The data type of <span style="background-color: #F9EBEA; color:##C0392B">world_alcohol</span> is float. Because all of the values in a **NumPy array have to have the same data type**, NumPy attempted to convert all of the columns to floats when they were read in. The <span style="background-color: #F9EBEA; color:##C0392B">numpy.genfromtxt()</span> function will attempt to guess the correct data type of the array it creates.
#
# In this case, the **WHO Region**, **Country**, and **Beverage Types** columns are actually <span style="background-color: #F9EBEA; color:##C0392B">strings</span>, and couldn't be converted to <span style="background-color: #F9EBEA; color:##C0392B">floats</span>. When NumPy can't convert a value to a numeric data type like float or integer, it uses a special nan value that stands for **"not a number"**. NumPy assigns an na value, which stands for "not available", when the value doesn't exist. <span style="background-color: #F9EBEA; color:##C0392B">nan</span> and <span style="background-color: #F9EBEA; color:##C0392B">na</span> values are types of missing data. We'll dive more into how to deal with missing data in later missions.
#
# The whole first row of <span style="background-color: #F9EBEA; color:##C0392B">world_alcohol.csv</span> is a header row that contains the names of each column. This is not actually part of the data, and consists entirely of strings. Since the strings couldn't be converted to floats properly, NumPy uses nan values to represent them.
#
# ### 3.5 Reading the data correctly
#
# When reading in the data using the [numpy.genfromtxt()](http://docs.scipy.org/doc/numpy-1.10.0/reference/generated/numpy.genfromtxt.html) function, we can use parameters to customize how we want the data to be read in. While we're at it, we can also specify that we want to skip the header row of <span style="background-color: #F9EBEA; color:##C0392B">world_alcohol.csv</span>.
#
# To specify the data type for the entire NumPy array, we use the keyword argument dtype and set it to <span style="background-color: #F9EBEA; color:##C0392B">"U75"</span>. This specifies that we want to read in each value as a 75 byte unicode data type. We'll dive more into unicode and bytes later on, but for now, it's enough to know that this will read in our data properly.
#
# To skip the header when reading in the data, we use the skip_header parameter. The <span style="background-color: #F9EBEA; color:##C0392B">skip_header</span>skip_header parameter accepts an integer value, specifying the number of lines from the top of the file we want NumPy to ignore.
#
world_alcohol = np.genfromtxt("data/world_alcohol.csv", delimiter=",", dtype="U75", skip_header=1)
print(world_alcohol)
#slicing
print(world_alcohol[1,:])
print(world_alcohol[1,0:2])
# ### 3.5 Array Comparisons
#
# One of the most powerful aspects of the NumPy module is the ability to make comparisons across an entire array. These comparisons result in Boolean values.
#
#
#
vector = np.array([5, 10, 15, 20])
vector == 10
matrix = np.array([[5, 10, 15],
[20, 25, 30],
[35, 40, 45]]
)
matrix == 25
# ### 3.6 Selecting elements
#
# We mentioned that comparisons are very powerful, but it may not have been obvious why on the last screen. Comparisons give us the power to select elements in arrays using Boolean vectors. This allows us to conditionally select certain elements in vectors, or certain rows in matrices.
#
#
# +
vector = np.array([5, 10, 15, 20])
equal_to_ten = (vector == 10)
print(vector[equal_to_ten])
# +
vector = np.array([5, 10, 15, 20])
equal_to_ten_and_five = (vector == 10) | (vector == 5)
print(equal_to_ten_and_five)
# -
vector = np.array([5, 10, 15, 20])
equal_to_ten_or_five = (vector == 10) | (vector == 5)
vector[equal_to_ten_or_five] = 50
print(vector)
# ### 3.7 Computing with NumPy
#
# Now that alcohol_consumption consists of numeric values, we can perform computations on it. NumPy has a few built-in methods that operate on arrays. You can view all of them in the documentation. For now, here are a few important ones:
#
# - sum() -- Computes the sum of all the elements in a vector, or the sum along a dimension in a matrix
# - mean() -- Computes the average of all the elements in a vector, or the average along a dimension in a matrix
# - max() -- Identifies the maximum value among all the elements in a vector, or the maximum along a dimension in a matrix
#
# Here's an example of how we'd use one of these methods on a vector:
vector = np.array([5, 10, 15, 20])
vector.sum()
# With a matrix, we have to specify an additional keyword argument, axis. The axis dictates which dimension we perform the operation on. 1 means that we want to perform the operation on each row, and 0 means on each column. The example below performs an operation across each row:
#
#
#
matrix = np.array([
[5, 10, 15],
[20, 25, 30],
[35, 40, 45]
])
matrix.sum(axis=1)
# ### 3.8 NumPy Strengths And Weaknesses
# You should now have a good foundation in NumPy, and in handling issues with your data. NumPy is much easier to work with than lists of lists, because:
#
# - It's easy to perform computations on data.
# - Data indexing and slicing is faster and easier.
# - We can convert data types quickly.
# Overall, NumPy makes working with data in Python much more efficient. It's widely used for this reason, especially for machine learning.
#
# You may have noticed some limitations with NumPy as you worked through the past two missions, though. For example:
#
# - All of the items in an array must have the same data type. For many datasets, this can make arrays cumbersome to work with.
# - Columns and rows must be referred to by number, which gets confusing when you go back and forth from column name to column number.
# - In the next few missions, we'll learn about the Pandas library, one of the most popular data analysis libraries. Pandas builds on NumPy, but does a better job addressing the limitations of NumPy.
#
#
| 15,491 |
/Optimization.ipynb | ac759102eea61c78f8dfb11b33e0a196055033d9 | [] | no_license | sblml/deeplearning | https://github.com/sblml/deeplearning | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 7,653 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Objective : 10. Styling Pandas Table
# <hr>
#
# 1. Applymap for applying entire table
# 2. Apply for applying column wise
# 3. Highlighting Null
# 4. Applying colors to subset of data
#
# <hr>
import pandas as pd
import numpy as np
df = pd.read_csv('../Data/online-sales/TrainingDataset.csv')
df = df.iloc[:100,list(range(10))]
df
# ### 1. Applymap
# * Apply styling on complete data
# * function returns a css parameter
# +
def color_product_sales(val):
if val > 10000:
color = 'green'
elif val < 2000:
color = 'red'
else:
color = 'black'
return 'color: %s' % color
df.style.applymap(color_product_sales)
# -
# ### 2. Apply
# 1. In case, we want to do series wise check apply can be used.
# 2. Function argument will be series.
# 3. By default, columns.
# 4. If assigned axis = 1, rows
# +
def highlight_max(s):
is_max = s == s.max()
return ['background-color: yellow' if v else '' for v in is_max]
df.style.apply(highlight_max)
# -
# * Chaining is also supported
df.style.applymap(color_product_sales).apply(highlight_max)
# ### 3. Highlighting Null Values
df.style.highlight_null(null_color='red')
# ### 4. Dealing with subset of Data
# * Selecting subset of columns
df.style.apply(highlight_max, subset=['Outcome_M8','Outcome_M9'])
# * Selecting subset of rows
df.style.apply(highlight_max, subset=pd.IndexSlice[2:14,:], axis=1)
df.style.apply(highlight_max, subset=pd.IndexSlice[2:14, ['Outcome_M6','Outcome_M7','Outcome_M8','Outcome_M9']], axis=1)
la carga y solo hay un campo eléctrico que viene dado por:
#
# $$
# \vec E' = \frac{q}{r'^3}\vec{r}'
# $$
# donde $\vec r'=\Delta x' \hat{e}_x + \Delta y' \hat{e}_y + \Delta z' \hat{e}_z$.
# Según las reglas de transformación el campo en el sistema de referencia del observador quedaría:
#
# $$
# \begin{array}{rcl}
# \vec E & = & \vec E'+(\gamma-1) \vec E'_\perp\\
# \vec B & = & \gamma \vec u_L \times \vec E'\\
# \end{array}
# $$
# Teniendo en cuenta que $\vec E'-\vec E'_\perp = \vec E'_\parallel$ y que $\vec u_L\times \vec E'=\vec u_L\times \vec E'_\perp$, el resultado se puede escribir como:
#
# $$
# \begin{array}{rcl}
# \vec E & = & \vec E'_\parallel+\gamma \vec E'_\perp\\
# \vec B & = & \vec u_L \times \vec E\\
# \end{array}
# $$
# Aquí notamos un interesante efecto: el campo eléctrico en dirección del movimiento no se modifica, pero si lo hace en dirección perpendicular donde es incrementado por un factor $\gamma$.
# Ahora bien, el valor del campo dado por la ley de Coulomb tiene una modificación debido a la contracción de longitudes en dirección de propagación. Si fijamos el eje x en la dirección de la velocidad, entonces:
#
# $$
# \vec r' = \gamma \Delta x \hat{e}_x + \Delta y \hat{e}_y + \Delta z \hat{e}_z
# $$
# de donde:
#
# $$
# r'^2 = \gamma \Delta x^2 + \Delta y^2 + \Delta z^2
# $$
# que se puede escribir como:
#
# $$
# r'^2 = \gamma^2 r^2 (1-v_L^2 \sin^2\theta)
# $$
# donde $\sin\theta=\sqrt{\Delta y^2+\Delta z^2}/\Delta x$ es el ángulo entre la dirección al punto en la que se esta calculando el campo y la dirección e propagación. Este resultado es general independientemente de si se usa o no la configuración estándar.
# Con este resultado el campo eléctrico en el sistema de referencia del observaor se puede escribir como:
#
# $$
# \vec E=\frac{1}{\gamma^2 (1-v_L^2\sin^2\theta)^{3/2}}\frac{q}{r^3}\vec{r}
# $$
# El campo magnético por otro lado esta dado por:
#
# $$
# \vec B=\frac{1}{\gamma^2 (1-v_L^2\sin^2\theta)^{3/2}}\frac{q}{r^3}\vec{v}_L\times \vec{r}
# $$
# Como vemos en cada momento ambos campos siguen disminuyendo como $1/r^2$, pero su magnitud depende de la dirección en la que mida el campo respecto a su dirección de propragación.
# Cuando $\theta=0, \pi$ (en la dirección de movimiento o en dirección contraria) la magnitud del campo eléctrico es:
#
# $$
# E(\theta=0,\pi)=\frac{q}{(\gamma r)^2}
# $$
# Es decir, en dirección del campo la contracción de longitudes hace que en el sistema de referencia del observador el campo a una distancia $r$ dada tenga una intensidad equivalente a la que vería la carga a una distancia $\gamma r$.
# En dirección perpendicular al campo $\theta=\pm\pi/2$ el campo por otro lado es:
#
# $$
# E(\theta=\pm\pi/2)=\frac{(\gamma q)}{r^2}
# $$
# En dirección perpendicular el efecto es curioso. En este caso es como si el campo fuera producido por una carga más intensa (carga relativista) $q_r=\gamma q$. En realidad el efecto es debido a los efectos en el espacio-tiempo en los que el campo magnético que ahora esta presente juega también un papel importante.
# En síntesis a la misma distancia hacia adelante el campo es $\gamma^3$ más débil que en dirección perpendicular.
# Podemos visualizar este campo de dos maneras diferentes, por un lado mostrando el vector de campo a una distancia determinada.
# Fijemos las propiedades de la partícula.
# + codelabel="" codeplot=0
q=1
vL=0.7
gamma=1/(1-vL**2)**0.5
# -
# Calculamos las componentes del campo eléctrico (tanto en reposo como en movimiento) a una distancia constante:
# + codelabel="" codeplot=0
#Angulos en los que evaluaremos el campo
from numpy import linspace,concatenate,pi,zeros_like
tetas=concatenate((linspace(-pi,0,10),
linspace(0,pi,10)))
#Vector de posición
xs=zeros_like(tetas)
ys=zeros_like(tetas)
#Vector de campo eléctrico
from numpy import zeros_like
Exs=zeros_like(tetas)
Eys=zeros_like(tetas)
Erxs=zeros_like(tetas)
Erys=zeros_like(tetas)
#Distancia constante
r=0.3
from numpy import cos,sin
for i,teta in enumerate(tetas):
xs[i]=r*cos(teta)
ys[i]=r*sin(teta)
# Campo eléctrico en reposo
Erxs[i]=q/r**3*xs[i]
Erys[i]=q/r**3*ys[i]
# Campo eléctrico en movimiento
Exs[i]=q/(r**3*gamma**2*(1-vL**2*sin(teta)**2)**1.5)*xs[i]
Eys[i]=q/(r**3*gamma**2*(1-vL**2*sin(teta)**2)**1.5)*ys[i]
# -
# Ahora podemos graficar el campo:
# + codelabel="" codeplot=0
# %matplotlib nbagg
# + codelabel="fig:01.08.04.00.RelatividadEspecial.Electrodinamica.Ejemplos_30" codeplot=1 figcaption="<b>Figura 1.30.</b> "
import matplotlib.pyplot as plt
fig=plt.figure(figsize=(5,5))
ax=fig.gca()
#Gráfico de los puntos
ax.plot(xs,ys,'b--')
#Gráfico de los vectores de campo
ax.quiver(xs,ys,Erxs,Erys,
color='g',alpha=0.5,scale=70,
label="Carga en reposo")
ax.quiver(xs,ys,Exs,Eys,
scale=70,
label="Carga en movimiento")
#Posición de la carga
ax.plot([0],[0],'ro')
#Decoración
ax.legend();
ax.set_xlim((-1,1));
ax.set_ylim((-1,1));
ax.set_xlabel("$x$");
ax.set_ylabel("$y$");
# -
# <a id='fig:01.08.04.00.RelatividadEspecial.Electrodinamica.Ejemplos_30'></a><center><b>Figura 1.30.</b> </center>
# #### 1.17.11.2. Campo producido por un alambre recto
# <a id='campo_alambre'></a>
# + [markdown] tags=["navigation"]
# [Indice](index.ipynb) | Previo: [RelatividadEspecial.Electrodinamica.TensorFaraday](01.08.03.00.RelatividadEspecial.Electrodinamica.TensorFaraday.ipynb) | Siguiente: [RelatividadEspecial.ProblemasSeleccionados](01.09.00.00.RelatividadEspecial.ProblemasSeleccionados.ipynb)
| 7,361 |
/notebooks/2_NYC_Mapping_MatPlotLib.ipynb | add8e00229592db97ae1d5061975772f3a6e9cd6 | [] | no_license | makingw/Geospatial_Mapping_In_Python | https://github.com/makingw/Geospatial_Mapping_In_Python | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 668,596 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Now let's look at this data plotted geospatially with MatPlotLib
# ### ---------------------------------------------------------------------------------------
#
#
# ## Matplotlib is straight up python plotting. It has a Basemap library. But it will not get as pretty in the User Interface as people are used to like a Google Map. It will work if something quick is needed. Using Geopandas or Folium with their basemap tiles is better for finished products. But those also require different language adaptations.
#
# ### ------------------
# #!/usr/bin/python
import json
import requests
import pandas as pd
import numpy as np
import geopandas as gpd
import matplotlib.pyplot as plt
import seaborn as sns
from mpl_toolkits.basemap import Basemap
import io
# %matplotlib inline
bestPizza = pd.read_csv("../data/PizzaEssentials.csv",index_col=False)
restPandI = pd.read_csv("../data/restPandI.csv")
print(restPandI.shape)
allPizza = pd.read_csv("../data/restPizza.csv")
print(allPizza.shape)
subway = pd.read_csv("../data/subwayStations.csv", index_col=0)
print(subway.shape)
# ## Geo Mapping with Matplotlib
# +
# The most simple way to see you data is like this!
restPandI.plot(kind="scatter", x="Longitude", y="Latitude", alpha=0.4)
plt.show()
# +
# And to add fun things like a color map and scale bar
restPandI.plot(kind="scatter", x="Longitude", y="Latitude",
#s=housing['population']/100, label="population",
c="ZIPCODE", cmap=plt.get_cmap("jet"),
colorbar=True, alpha=0.4, figsize=(10,7),
)
plt.legend()
plt.show()
# -
# ## Now with the Basemaps Library
# + jupyter={"outputs_hidden": true}
help(Basemap)
# +
# read in data to use for plotted points
lat = restPandI['Latitude'].values
lon = restPandI['Longitude'].values
# determine range to print based on min, max lat and lon of the data
margin = .1 # buffer to add to the range
lat_min = min(lat) - margin
lat_max = max(lat) + margin
lon_min = min(lon) - margin
lon_max = max(lon) + margin
# create map using BASEMAP
bmap = Basemap(llcrnrlon=lon_min,
llcrnrlat=lat_min,
urcrnrlon=lon_max,
urcrnrlat=lat_max,
lat_0=(lat_max - lat_min)/2,
lon_0=(lon_max-lon_min)/2,
epsg='4326',
resolution = 'h',
area_thresh=1000.,
)
bmap.drawstates(color='gray',linewidth=0.25)
bmap.drawcoastlines(linewidth=0.25)
bmap.drawcountries(linewidth=0.25)
bmap.fillcontinents(lake_color='aqua')
bmap.drawmapboundary(fill_color='lightblue')
# convert lat and lon to map projection coordinates
x,y = bmap(lon, lat)
bmap.plot(x, y, ".",'bo', markersize=1)
plt.title('NYC Pizza Places', fontsize=25)
# SETTING THE LEGEND
plt.legend(loc="upper left",title='Legend', markerscale=5,
frameon=True,facecolor='white', edgecolor='k',fancybox=True,framealpha=1,fontsize=7,title_fontsize=7)
plt.rcParams['figure.dpi'] = 400
plt.show()
# -
# ### Below we try to change the basemap to shadedrelief but we are too zoomed in to see it
# +
# read in data to use for plotted points
lat = restPandI['Latitude'].values
lon = restPandI['Longitude'].values
population = restPandI['ZIPCODE'].values
fig = plt.figure(figsize=(8, 8))
m = Basemap(llcrnrlon=lon_min,
llcrnrlat=lat_min,
urcrnrlon=lon_max,
urcrnrlat=lat_max,
lat_0=(lat_max - lat_min)/2,
lon_0=(lon_max-lon_min)/2,
projection='merc',
resolution = 'h',
area_thresh=1000.,
)
#width=1.05E6,
#height=1.2E6)
# trying with shaded relief basemap
m.shadedrelief()
m.drawcoastlines(color='gray',linewidth=1)
m.drawcountries(color='gray',linewidth=1)
#m.drawstates(color='gray')
lons, lats = m(lon, lat)
# scatter city data, with c reflecting population
m.scatter(lon,lat, latlon=True,
c=population, marker='o', zorder=5,
cmap='gist_heat', alpha=0.5)
#create colorbar
#plt.colorbar(label=r'Population')
plt.clim(300000, 4000000)
# -
# ## Here's a full list of colormap (cmap) options
#
# ValueError: Colormap Perceptually Uniform Sequential is not recognized. Possible values are: Accent, Accent_r, Blues, Blues_r, BrBG, BrBG_r, BuGn, BuGn_r, BuPu, BuPu_r, CMRmap, CMRmap_r, Dark2, Dark2_r, GnBu, GnBu_r, Greens, Greens_r, Greys, Greys_r, OrRd, OrRd_r, Oranges, Oranges_r, PRGn, PRGn_r, Paired, Paired_r, Pastel1, Pastel1_r, Pastel2, Pastel2_r, PiYG, PiYG_r, PuBu, PuBuGn, PuBuGn_r, PuBu_r, PuOr, PuOr_r, PuRd, PuRd_r, Purples, Purples_r, RdBu, RdBu_r, RdGy, RdGy_r, RdPu, RdPu_r, RdYlBu, RdYlBu_r, RdYlGn, RdYlGn_r, Reds, Reds_r, Set1, Set1_r, Set2, Set2_r, Set3, Set3_r, Spectral, Spectral_r, Wistia, Wistia_r, YlGn, YlGnBu, YlGnBu_r, YlGn_r, YlOrBr, YlOrBr_r, YlOrRd, YlOrRd_r, afmhot, afmhot_r, autumn, autumn_r, binary, binary_r, bone, bone_r, brg, brg_r, bwr, bwr_r, cividis, cividis_r, cool, cool_r, coolwarm, coolwarm_r, copper, copper_r, cubehelix, cubehelix_r, flag, flag_r, gist_earth, gist_earth_r, gist_gray, gist_gray_r, gist_heat, gist_heat_r, gist_ncar, gist_ncar_r, gist_rainbow, gist_rainbow_r, gist_stern, gist_stern_r, gist_yarg, gist_yarg_r, gnuplot, gnuplot2, gnuplot2_r, gnuplot_r, gray, gray_r, hot, hot_r, hsv, hsv_r, icefire, icefire_r, inferno, inferno_r, jet, jet_r, magma, magma_r, mako, mako_r, nipy_spectral, nipy_spectral_r, ocean, ocean_r, pink, pink_r, plasma, plasma_r, prism, prism_r, rainbow, rainbow_r, rocket, rocket_r, seismic, seismic_r, spring, spring_r, summer, summer_r, tab10, tab10_r, tab20, tab20_r, tab20b, tab20b_r, tab20c, tab20c_r, terrain, terrain_r, twilight, twilight_r, twilight_shifted, twilight_shifted_r, viridis, viridis_r, vlag, vlag_r, winter, winter_r
| 5,958 |
/CP3_Catboost.ipynb | 2e28df5948d29836d85c701a9346ce1634fdee1d | [] | no_license | WeiDong-Sanderson/Zillow_prediction | https://github.com/WeiDong-Sanderson/Zillow_prediction | 0 | 0 | null | null | null | null | Jupyter Notebook | false | false | .py | 231,755 | # ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.15.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # KNN Model
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.metrics import plot_confusion_matrix
from sklearn.metrics import roc_auc_score
plt.rcParams['figure.figsize'] = (15, 15)
# -
# ## Retrieve all of the features and labels
npzfile = np.load('../data/melspects_128.npz', allow_pickle=True)
X_train, y_train = npzfile['X_train'], npzfile['y_train']
X_test, y_test = npzfile['X_test'], npzfile['y_test']
# ## Normalize the Data
#
# We need to reshape the data so that we can use sklearn's `StandardScaler` becasue it only works with 2D arrays. Our data is currently stored as a 3D array.
X_train = X_train.reshape(X_train.shape[0], X_train.shape[1]*X_train.shape[2])
X_test = X_test.reshape(X_test.shape[0], X_test.shape[1]*X_test.shape[2])
# +
scaler = StandardScaler()
scaler.fit(X_train)
X_train_sc = scaler.transform(X_train)
X_test_sc = scaler.transform(X_test)
# -
# ## PCA
#
# So currently we have 61440 features, I'm going to use PCA to reduce the dimensionality significantly.
X_train_sc.shape
# +
pca = PCA(n_components=18)
pca.fit(X_train_sc)
X_train_reduced = pca.transform(X_train_sc)
X_test_reduced = pca.transform(X_test_sc)
# -
X_train_reduced.shape
# So after performing PCA, only keeping 18 features. Considerably less than we started with. I found that this yielded the most accurate models when testing different values for `n_components`.
# ## Training the Model
#
# I'm going to use sklearn's `GridSearchCV` for hyperparameter tuning. The two parameters I'm going to tune are the `KNeighborsClassifier`'s `n_neighbors` and `weights` parameters. The two weight options I will consider are the uniform and distance option. I'll be looking in a range of 5 to 15 for the number of neighbors.
#
# The nice thing about `GridSearchCV` is it will perform K-Fold crossvalidation for me. By default it will use 5 folds, which is plenty.
# +
parameters = {'weights':('uniform', 'distance'), 'n_neighbors': range(5, 16)}
knn = KNeighborsClassifier()
clf = GridSearchCV(knn, parameters)
# -
clf.fit(X_train_reduced, y_train)
# +
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test_reduced)
print(classification_report(y_true, y_pred))
print()
# -
# So for the KNN model the best parameter set was to use 5 for `n_neighbors` and distance for `weights`. The classification report is telling us that the overall accuracy of the model was 0.49, which is honestly better than I expected from the KNN model. The genre of music that performs the worst is 3 (disco). We are misclassifying it everytime, the precision is 0.00 and the recall is 0.00. The second worst genre was 0 (blues).
# +
class_names = ['blues', 'classical', 'country', 'disco',
'hiphop', 'jazz', 'metal', 'pop', 'reggae', 'rock']
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
titles_options = [("Confusion matrix, without normalization", None),
("Normalized confusion matrix", 'true')]
for title, normalize in titles_options:
disp = plot_confusion_matrix(clf, X_test_reduced, y_test,
display_labels=class_names,
cmap=plt.cm.Blues,
normalize=normalize)
disp.ax_.set_title(title)
plt.show()
# -
# So the genres that were the easiest to classify were metal, pop, and classical. This makes sense becasue those genres have very distinct sounds.
# +
y_prob = clf.predict_proba(X_test_reduced)
macro_roc_auc_ovo = roc_auc_score(y_test, y_prob, multi_class="ovo", average="macro")
weighted_roc_auc_ovo = roc_auc_score(y_test, y_prob, multi_class="ovo", average="weighted")
macro_roc_auc_ovr = roc_auc_score(y_test, y_prob, multi_class="ovr", average="macro")
weighted_roc_auc_ovr = roc_auc_score(y_test, y_prob, multi_class="ovr", average="weighted")
print("One-vs-One ROC AUC scores:\n{:.6f} (macro)".format(macro_roc_auc_ovo, weighted_roc_auc_ovo))
print("One-vs-Rest ROC AUC scores:\n{:.6f} (macro)".format(macro_roc_auc_ovr))
re_list].astype(str)
num_ensembles = 5
y_pred = 0.0
for i in tqdm(range(num_ensembles)):
model = CatBoostRegressor(
iterations=630, learning_rate=0.03,
depth=6, l2_leaf_reg=3,
loss_function='MAE',
eval_metric='MAE',
random_seed=i)
model.fit(
X_train, y_train,
cat_features=cat_features)
y_pred += model.predict(X_test)
y_pred /= num_ensembles
# +
submission = pd.DataFrame({
'ParcelId': test_df['ParcelId'],
})
test_dates = {
'201610': pd.Timestamp('2016-09-30'),
'201611': pd.Timestamp('2016-10-31'),
'201612': pd.Timestamp('2016-11-30'),
'201710': pd.Timestamp('2017-09-30'),
'201711': pd.Timestamp('2017-10-31'),
'201712': pd.Timestamp('2017-11-30')
}
for label, test_date in test_dates.items():
print("Predicting for: %s . " % (label))
submission[label] = y_pred
submission.to_csv('CatBoost_prediction.csv', float_format='%.6f',index=False)
# -
submission
| 6,063 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.