code stringlengths 114 1.05M | path stringlengths 3 312 | quality_prob float64 0.5 0.99 | learning_prob float64 0.2 1 | filename stringlengths 3 168 | kind stringclasses 1
value |
|---|---|---|---|---|---|
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk
import constructs
@jsii.enum(
jsii_type="@renovosolutions/cdk-aspects-library-security-group.AnnotationType"
)
class AnnotationType(enum.Enum):
'''The supported annotation types.
Only error will stop deployment of restricted resources.
'''
WARNING = "WARNING"
ERROR = "ERROR"
INFO = "INFO"
@jsii.data_type(
jsii_type="@renovosolutions/cdk-aspects-library-security-group.AspectPropsBase",
jsii_struct_bases=[],
name_mapping={
"annotation_text": "annotationText",
"annotation_type": "annotationType",
},
)
class AspectPropsBase:
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''The base aspect properties available to any aspect.
JSII doesn't support an Omit when extending interfaces, so we create a base class to extend from.
This base class meets the needed properties for all non-base aspects.
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
self._values: typing.Dict[str, typing.Any] = {}
if annotation_text is not None:
self._values["annotation_text"] = annotation_text
if annotation_type is not None:
self._values["annotation_type"] = annotation_type
@builtins.property
def annotation_text(self) -> typing.Optional[builtins.str]:
'''The annotation text to use for the annotation.'''
result = self._values.get("annotation_text")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def annotation_type(self) -> typing.Optional[AnnotationType]:
'''The annotation type to use for the annotation.'''
result = self._values.get("annotation_type")
return typing.cast(typing.Optional[AnnotationType], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AspectPropsBase(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-aspects-library-security-group.AspectPropsExtended",
jsii_struct_bases=[AspectPropsBase],
name_mapping={
"annotation_text": "annotationText",
"annotation_type": "annotationType",
"any_source": "anySource",
"ports": "ports",
"restricted_cidrs": "restrictedCidrs",
"restricted_s_gs": "restrictedSGs",
},
)
class AspectPropsExtended(AspectPropsBase):
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
any_source: typing.Optional[builtins.bool] = None,
ports: typing.Optional[typing.Sequence[jsii.Number]] = None,
restricted_cidrs: typing.Optional[typing.Sequence[builtins.str]] = None,
restricted_s_gs: typing.Optional[typing.Sequence[builtins.str]] = None,
) -> None:
'''The extended aspect properties available only to the base security aspects.
These additional properties shouldn't be changed in aspects that already have clearly defined goals.
So, this extended properties interface is applied selectively to the base aspects.
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
:param any_source: Whether any source is valid. This will ignore all other restrictions and only check the port. Default: false
:param ports: The restricted port. Defaults to restricting all ports and only checking sources. Default: undefined
:param restricted_cidrs: The restricted CIDRs for the given port. Default: ['0.0.0.0/0', '::/0']
:param restricted_s_gs: The restricted source security groups for the given port. Default: undefined
'''
self._values: typing.Dict[str, typing.Any] = {}
if annotation_text is not None:
self._values["annotation_text"] = annotation_text
if annotation_type is not None:
self._values["annotation_type"] = annotation_type
if any_source is not None:
self._values["any_source"] = any_source
if ports is not None:
self._values["ports"] = ports
if restricted_cidrs is not None:
self._values["restricted_cidrs"] = restricted_cidrs
if restricted_s_gs is not None:
self._values["restricted_s_gs"] = restricted_s_gs
@builtins.property
def annotation_text(self) -> typing.Optional[builtins.str]:
'''The annotation text to use for the annotation.'''
result = self._values.get("annotation_text")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def annotation_type(self) -> typing.Optional[AnnotationType]:
'''The annotation type to use for the annotation.'''
result = self._values.get("annotation_type")
return typing.cast(typing.Optional[AnnotationType], result)
@builtins.property
def any_source(self) -> typing.Optional[builtins.bool]:
'''Whether any source is valid.
This will ignore all other restrictions and only check the port.
:default: false
'''
result = self._values.get("any_source")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def ports(self) -> typing.Optional[typing.List[jsii.Number]]:
'''The restricted port.
Defaults to restricting all ports and only checking sources.
:default: undefined
'''
result = self._values.get("ports")
return typing.cast(typing.Optional[typing.List[jsii.Number]], result)
@builtins.property
def restricted_cidrs(self) -> typing.Optional[typing.List[builtins.str]]:
'''The restricted CIDRs for the given port.
:default: ['0.0.0.0/0', '::/0']
'''
result = self._values.get("restricted_cidrs")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
@builtins.property
def restricted_s_gs(self) -> typing.Optional[typing.List[builtins.str]]:
'''The restricted source security groups for the given port.
:default: undefined
'''
result = self._values.get("restricted_s_gs")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AspectPropsExtended(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-aspects-library-security-group.RuleCheckArgs",
jsii_struct_bases=[AspectPropsExtended],
name_mapping={
"annotation_text": "annotationText",
"annotation_type": "annotationType",
"any_source": "anySource",
"ports": "ports",
"restricted_cidrs": "restrictedCidrs",
"restricted_s_gs": "restrictedSGs",
"node": "node",
},
)
class RuleCheckArgs(AspectPropsExtended):
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
any_source: typing.Optional[builtins.bool] = None,
ports: typing.Optional[typing.Sequence[jsii.Number]] = None,
restricted_cidrs: typing.Optional[typing.Sequence[builtins.str]] = None,
restricted_s_gs: typing.Optional[typing.Sequence[builtins.str]] = None,
node: constructs.IConstruct,
) -> None:
'''The arguments for the checkRules function.
Extends the IAspectPropsBase interface which includes additional properties that can be used as args.
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
:param any_source: Whether any source is valid. This will ignore all other restrictions and only check the port. Default: false
:param ports: The restricted port. Defaults to restricting all ports and only checking sources. Default: undefined
:param restricted_cidrs: The restricted CIDRs for the given port. Default: ['0.0.0.0/0', '::/0']
:param restricted_s_gs: The restricted source security groups for the given port. Default: undefined
:param node: The node to check.
'''
self._values: typing.Dict[str, typing.Any] = {
"node": node,
}
if annotation_text is not None:
self._values["annotation_text"] = annotation_text
if annotation_type is not None:
self._values["annotation_type"] = annotation_type
if any_source is not None:
self._values["any_source"] = any_source
if ports is not None:
self._values["ports"] = ports
if restricted_cidrs is not None:
self._values["restricted_cidrs"] = restricted_cidrs
if restricted_s_gs is not None:
self._values["restricted_s_gs"] = restricted_s_gs
@builtins.property
def annotation_text(self) -> typing.Optional[builtins.str]:
'''The annotation text to use for the annotation.'''
result = self._values.get("annotation_text")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def annotation_type(self) -> typing.Optional[AnnotationType]:
'''The annotation type to use for the annotation.'''
result = self._values.get("annotation_type")
return typing.cast(typing.Optional[AnnotationType], result)
@builtins.property
def any_source(self) -> typing.Optional[builtins.bool]:
'''Whether any source is valid.
This will ignore all other restrictions and only check the port.
:default: false
'''
result = self._values.get("any_source")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def ports(self) -> typing.Optional[typing.List[jsii.Number]]:
'''The restricted port.
Defaults to restricting all ports and only checking sources.
:default: undefined
'''
result = self._values.get("ports")
return typing.cast(typing.Optional[typing.List[jsii.Number]], result)
@builtins.property
def restricted_cidrs(self) -> typing.Optional[typing.List[builtins.str]]:
'''The restricted CIDRs for the given port.
:default: ['0.0.0.0/0', '::/0']
'''
result = self._values.get("restricted_cidrs")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
@builtins.property
def restricted_s_gs(self) -> typing.Optional[typing.List[builtins.str]]:
'''The restricted source security groups for the given port.
:default: undefined
'''
result = self._values.get("restricted_s_gs")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
@builtins.property
def node(self) -> constructs.IConstruct:
'''The node to check.'''
result = self._values.get("node")
assert result is not None, "Required property 'node' is missing"
return typing.cast(constructs.IConstruct, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "RuleCheckArgs(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.implements(aws_cdk.IAspect)
class SecurityGroupAspectBase(
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.SecurityGroupAspectBase",
):
'''The base class for all security group aspects in the library.
By default this will not restrict anything.
'''
def __init__(
self,
*,
any_source: typing.Optional[builtins.bool] = None,
ports: typing.Optional[typing.Sequence[jsii.Number]] = None,
restricted_cidrs: typing.Optional[typing.Sequence[builtins.str]] = None,
restricted_s_gs: typing.Optional[typing.Sequence[builtins.str]] = None,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param any_source: Whether any source is valid. This will ignore all other restrictions and only check the port. Default: false
:param ports: The restricted port. Defaults to restricting all ports and only checking sources. Default: undefined
:param restricted_cidrs: The restricted CIDRs for the given port. Default: ['0.0.0.0/0', '::/0']
:param restricted_s_gs: The restricted source security groups for the given port. Default: undefined
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsExtended(
any_source=any_source,
ports=ports,
restricted_cidrs=restricted_cidrs,
restricted_s_gs=restricted_s_gs,
annotation_text=annotation_text,
annotation_type=annotation_type,
)
jsii.create(self.__class__, self, [props])
@jsii.member(jsii_name="visit")
def visit(self, node: constructs.IConstruct) -> None:
'''All aspects can visit an IConstruct.
:param node: -
'''
return typing.cast(None, jsii.invoke(self, "visit", [node]))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="annotationText")
def annotation_text(self) -> builtins.str:
return typing.cast(builtins.str, jsii.get(self, "annotationText"))
@annotation_text.setter
def annotation_text(self, value: builtins.str) -> None:
jsii.set(self, "annotationText", value)
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="annotationType")
def annotation_type(self) -> AnnotationType:
return typing.cast(AnnotationType, jsii.get(self, "annotationType"))
@annotation_type.setter
def annotation_type(self, value: AnnotationType) -> None:
jsii.set(self, "annotationType", value)
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="anySource")
def any_source(self) -> builtins.bool:
return typing.cast(builtins.bool, jsii.get(self, "anySource"))
@any_source.setter
def any_source(self, value: builtins.bool) -> None:
jsii.set(self, "anySource", value)
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ports")
def ports(self) -> typing.Optional[typing.List[jsii.Number]]:
return typing.cast(typing.Optional[typing.List[jsii.Number]], jsii.get(self, "ports"))
@ports.setter
def ports(self, value: typing.Optional[typing.List[jsii.Number]]) -> None:
jsii.set(self, "ports", value)
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="restrictedCidrs")
def restricted_cidrs(self) -> typing.Optional[typing.List[builtins.str]]:
return typing.cast(typing.Optional[typing.List[builtins.str]], jsii.get(self, "restrictedCidrs"))
@restricted_cidrs.setter
def restricted_cidrs(
self,
value: typing.Optional[typing.List[builtins.str]],
) -> None:
jsii.set(self, "restrictedCidrs", value)
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="restrictedSGs")
def restricted_s_gs(self) -> typing.Optional[typing.List[builtins.str]]:
return typing.cast(typing.Optional[typing.List[builtins.str]], jsii.get(self, "restrictedSGs"))
@restricted_s_gs.setter
def restricted_s_gs(
self,
value: typing.Optional[typing.List[builtins.str]],
) -> None:
jsii.set(self, "restrictedSGs", value)
class NoIngressCommonManagementPortsAspect(
SecurityGroupAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoIngressCommonManagementPortsAspect",
):
'''Aspect to restrict any access to common management ports.
22 - SSH
3389 - RDP
5985 - WinRM
5986 - WinRM HTTPS
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoIngressCommonRelationalDBPortsAspect(
SecurityGroupAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoIngressCommonRelationalDBPortsAspect",
):
'''Aspect to restrict any access to common relational DB ports.
3306 - MySQL
5432 - PostgreSQL
1521 - Oracle
1433 - SQL Server
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoIngressCommonWebPortsAspect(
SecurityGroupAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoIngressCommonWebPortsAspect",
):
'''Aspect to restrict any access to common web ports.
80 - HTTP
443 - HTTPS
8080 - HTTP
8443 - HTTPS
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
@jsii.implements(aws_cdk.IAspect)
class NoPublicIngressAspectBase(
SecurityGroupAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressAspectBase",
):
'''The base aspect to determine if a security group allows inbound traffic from the public internet to any port.
This inherits everything from the base SecurityGroupAspectBase class and sets a default set of CIDRS that match allowing all IPs on AWS.
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoPublicIngressCommonManagementPortsAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressCommonManagementPortsAspect",
):
'''Aspect to restrict public access to common management ports.
22 - SSH
3389 - RDP
5985 - WinRM
5986 - WinRM HTTPS
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoPublicIngressCommonRelationalDBPortsAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressCommonRelationalDBPortsAspect",
):
'''Aspect to restrict public access to common relational DB ports.
3306 - MySQL
5432 - PostgreSQL
1521 - Oracle
1433 - SQL Server
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoPublicIngressCommonWebPortsAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressCommonWebPortsAspect",
):
'''Aspect to restrict public access to common web ports.
80 - HTTP
443 - HTTPS
8080 - HTTP
8443 - HTTPS
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoPublicIngressRDPAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressRDPAspect",
):
'''Aspect to determine if a security group allows inbound traffic from the public internet to the RDP port.'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class NoPublicIngressSSHAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressSSHAspect",
):
'''Aspect to determine if a security group allows inbound traffic from the public internet to the SSH port.'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class AWSRestrictedCommonPortsAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.AWSRestrictedCommonPortsAspect",
):
'''Restricted common ports based on AWS Config rule https://docs.aws.amazon.com/config/latest/developerguide/restricted-common-ports.html.'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class CISAwsFoundationBenchmark4Dot1Aspect(
NoPublicIngressSSHAspect,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.CISAwsFoundationBenchmark4Dot1Aspect",
):
'''CIS AWS Foundations Benchmark 4.1.
CIS recommends that no security group allow unrestricted ingress access to port 22. Removing unfettered connectivity to remote console services, such as SSH, reduces a server's exposure to risk.
This aspect uses the NoPublicIngressSSHAspect with an alternate annotation text.
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
class CISAwsFoundationBenchmark4Dot2Aspect(
NoPublicIngressRDPAspect,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.CISAwsFoundationBenchmark4Dot2Aspect",
):
'''CIS AWS Foundations Benchmark 4.2.
CIS recommends that no security group allow unrestricted ingress access to port 3389. Removing unfettered connectivity to remote console services, such as RDP, reduces a server's exposure to risk.
This aspect uses the NoPublicIngressRDPAspect with an alternate annotation text.
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
@jsii.implements(aws_cdk.IAspect)
class NoPublicIngressAspect(
NoPublicIngressAspectBase,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-aspects-library-security-group.NoPublicIngressAspect",
):
'''The same as the base NoPublicIngressAspectBase but with a more descriptive annotation.
Blocks the ANY port from the public internet.
'''
def __init__(
self,
*,
annotation_text: typing.Optional[builtins.str] = None,
annotation_type: typing.Optional[AnnotationType] = None,
) -> None:
'''
:param annotation_text: The annotation text to use for the annotation.
:param annotation_type: The annotation type to use for the annotation.
'''
props = AspectPropsBase(
annotation_text=annotation_text, annotation_type=annotation_type
)
jsii.create(self.__class__, self, [props])
__all__ = [
"AWSRestrictedCommonPortsAspect",
"AnnotationType",
"AspectPropsBase",
"AspectPropsExtended",
"CISAwsFoundationBenchmark4Dot1Aspect",
"CISAwsFoundationBenchmark4Dot2Aspect",
"NoIngressCommonManagementPortsAspect",
"NoIngressCommonRelationalDBPortsAspect",
"NoIngressCommonWebPortsAspect",
"NoPublicIngressAspect",
"NoPublicIngressAspectBase",
"NoPublicIngressCommonManagementPortsAspect",
"NoPublicIngressCommonRelationalDBPortsAspect",
"NoPublicIngressCommonWebPortsAspect",
"NoPublicIngressRDPAspect",
"NoPublicIngressSSHAspect",
"RuleCheckArgs",
"SecurityGroupAspectBase",
]
publication.publish() | /renovosolutions.aws_cdk_aspects_security_group-2.0.199-py3-none-any.whl/aspects-security-group/__init__.py | 0.752468 | 0.205276 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk
import aws_cdk.aws_ec2
import constructs
class Ipam(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-ipam.Ipam",
):
'''Creates an IPAM.
PAM is a VPC feature that you can use to automate your IP address management workflows including
assigning, tracking, troubleshooting, and auditing IP addresses across AWS Regions and accounts
throughout your AWS Organization. For more information, see What is IPAM? in the Amazon VPC IPAM
User Guide.
:see: https://docs.aws.amazon.com/vpc/latest/ipam/what-is-it-ipam.html
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
description: typing.Optional[builtins.str] = None,
operating_regions: typing.Optional[typing.Sequence[builtins.str]] = None,
tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param description: The description for the IPAM.
:param operating_regions: The operating Regions for an IPAM. Operating Regions are AWS Regions where the IPAM is allowed to manage IP address CIDRs. IPAM only discovers and monitors resources in the AWS Regions you select as operating Regions. For more information about operating Regions, see Create an IPAM in the Amazon VPC IPAM User Guide. Default: Stack.of(this).region
:param tags: The key/value combination of tags to assign to the resource.
'''
props = IpamProps(
description=description, operating_regions=operating_regions, tags=tags
)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipam")
def ipam(self) -> aws_cdk.aws_ec2.CfnIPAM:
'''The underlying IPAM resource.'''
return typing.cast(aws_cdk.aws_ec2.CfnIPAM, jsii.get(self, "ipam"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamId")
def ipam_id(self) -> builtins.str:
'''The ID of the resulting IPAM resource.'''
return typing.cast(builtins.str, jsii.get(self, "ipamId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="privateDefaultScopeId")
def private_default_scope_id(self) -> builtins.str:
'''The default private scope ID.'''
return typing.cast(builtins.str, jsii.get(self, "privateDefaultScopeId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="publicDefaultScopeId")
def public_default_scope_id(self) -> builtins.str:
'''The default public scope ID.'''
return typing.cast(builtins.str, jsii.get(self, "publicDefaultScopeId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="scopeCount")
def scope_count(self) -> jsii.Number:
'''The number of scopes in this IPAM.'''
return typing.cast(jsii.Number, jsii.get(self, "scopeCount"))
class IpamAllocation(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamAllocation",
):
'''An IPAM Allocation.
In IPAM, an allocation is a CIDR assignment from an IPAM pool to another resource or IPAM pool.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
ipam_pool: "IpamPool",
cidr: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
netmask_length: typing.Optional[jsii.Number] = None,
) -> None:
'''
:param scope: -
:param id: -
:param ipam_pool: The IPAM pool from which you would like to allocate a CIDR.
:param cidr: The CIDR you would like to allocate from the IPAM pool. Note the following:. If there is no DefaultNetmaskLength allocation rule set on the pool, you must specify either the NetmaskLength or the CIDR. If the DefaultNetmaskLength allocation rule is set on the pool, you can specify either the NetmaskLength or the CIDR and the DefaultNetmaskLength allocation rule will be ignored.
:param description: A description of the pool allocation.
:param netmask_length: The netmask length of the CIDR you would like to allocate from the IPAM pool. Note the following:. If there is no DefaultNetmaskLength allocation rule set on the pool, you must specify either the NetmaskLength or the CIDR. If the DefaultNetmaskLength allocation rule is set on the pool, you can specify either the NetmaskLength or the CIDR and the DefaultNetmaskLength allocation rule will be ignored.
'''
props = IpamAllocationProps(
ipam_pool=ipam_pool,
cidr=cidr,
description=description,
netmask_length=netmask_length,
)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="allocation")
def allocation(self) -> aws_cdk.aws_ec2.CfnIPAMAllocation:
'''The underlying IPAM Allocation resource.'''
return typing.cast(aws_cdk.aws_ec2.CfnIPAMAllocation, jsii.get(self, "allocation"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamPoolAllocationId")
def ipam_pool_allocation_id(self) -> builtins.str:
'''The ID of the allocation.'''
return typing.cast(builtins.str, jsii.get(self, "ipamPoolAllocationId"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamAllocationProps",
jsii_struct_bases=[],
name_mapping={
"ipam_pool": "ipamPool",
"cidr": "cidr",
"description": "description",
"netmask_length": "netmaskLength",
},
)
class IpamAllocationProps:
def __init__(
self,
*,
ipam_pool: "IpamPool",
cidr: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
netmask_length: typing.Optional[jsii.Number] = None,
) -> None:
'''Properties of an IPAM Allocation.
:param ipam_pool: The IPAM pool from which you would like to allocate a CIDR.
:param cidr: The CIDR you would like to allocate from the IPAM pool. Note the following:. If there is no DefaultNetmaskLength allocation rule set on the pool, you must specify either the NetmaskLength or the CIDR. If the DefaultNetmaskLength allocation rule is set on the pool, you can specify either the NetmaskLength or the CIDR and the DefaultNetmaskLength allocation rule will be ignored.
:param description: A description of the pool allocation.
:param netmask_length: The netmask length of the CIDR you would like to allocate from the IPAM pool. Note the following:. If there is no DefaultNetmaskLength allocation rule set on the pool, you must specify either the NetmaskLength or the CIDR. If the DefaultNetmaskLength allocation rule is set on the pool, you can specify either the NetmaskLength or the CIDR and the DefaultNetmaskLength allocation rule will be ignored.
'''
self._values: typing.Dict[str, typing.Any] = {
"ipam_pool": ipam_pool,
}
if cidr is not None:
self._values["cidr"] = cidr
if description is not None:
self._values["description"] = description
if netmask_length is not None:
self._values["netmask_length"] = netmask_length
@builtins.property
def ipam_pool(self) -> "IpamPool":
'''The IPAM pool from which you would like to allocate a CIDR.'''
result = self._values.get("ipam_pool")
assert result is not None, "Required property 'ipam_pool' is missing"
return typing.cast("IpamPool", result)
@builtins.property
def cidr(self) -> typing.Optional[builtins.str]:
'''The CIDR you would like to allocate from the IPAM pool. Note the following:.
If there is no DefaultNetmaskLength allocation rule set on the pool, you must
specify either the NetmaskLength or the CIDR.
If the DefaultNetmaskLength allocation rule is set on the pool, you can specify
either the NetmaskLength or the CIDR and the DefaultNetmaskLength allocation rule
will be ignored.
'''
result = self._values.get("cidr")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''A description of the pool allocation.'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def netmask_length(self) -> typing.Optional[jsii.Number]:
'''The netmask length of the CIDR you would like to allocate from the IPAM pool. Note the following:.
If there is no DefaultNetmaskLength allocation rule set on the pool, you must specify either the
NetmaskLength or the CIDR.
If the DefaultNetmaskLength allocation rule is set on the pool, you can specify either the
NetmaskLength or the CIDR and the DefaultNetmaskLength allocation rule will be ignored.
'''
result = self._values.get("netmask_length")
return typing.cast(typing.Optional[jsii.Number], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "IpamAllocationProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class IpamPool(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamPool",
):
'''An IPAM Pool.
In IPAM, a pool is a collection of contiguous IP addresses CIDRs. Pools enable you to organize your IP addresses
according to your routing and security needs. For example, if you have separate routing and security needs for
development and production applications, you can create a pool for each.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
address_family: "IpamPoolAddressFamily",
ipam_scope_id: builtins.str,
allocation_default_netmask_length: typing.Optional[jsii.Number] = None,
allocation_max_netmask_length: typing.Optional[jsii.Number] = None,
allocation_min_netmask_length: typing.Optional[jsii.Number] = None,
allocation_resource_tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
auto_import: typing.Optional[builtins.bool] = None,
description: typing.Optional[builtins.str] = None,
locale: typing.Optional[builtins.str] = None,
provisioned_cidrs: typing.Optional[typing.Sequence[aws_cdk.aws_ec2.CfnIPAMPool.ProvisionedCidrProperty]] = None,
source_ipam_pool_id: typing.Optional[builtins.str] = None,
tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param address_family: The address family of the pool, either IPv4 or IPv6.
:param ipam_scope_id: The IPAM scope this pool is associated with.
:param allocation_default_netmask_length: The default netmask length for allocations added to this pool. If, for example, the CIDR assigned to this pool is 10.0.0.0/8 and you enter 16 here, new allocations will default to 10.0.0.0/16.
:param allocation_max_netmask_length: The maximum netmask length possible for CIDR allocations in this IPAM pool to be compliant. The maximum netmask length must be greater than the minimum netmask length. Possible netmask lengths for IPv4 addresses are 0 - 32. Possible netmask lengths for IPv6 addresses are 0 - 128.
:param allocation_min_netmask_length: The minimum netmask length required for CIDR allocations in this IPAM pool to be compliant. The minimum netmask length must be less than the maximum netmask length. Possible netmask lengths for IPv4 addresses are 0 - 32. Possible netmask lengths for IPv6 addresses are 0 - 128.
:param allocation_resource_tags: Tags that are required for resources that use CIDRs from this IPAM pool. Resources that do not have these tags will not be allowed to allocate space from the pool. If the resources have their tags changed after they have allocated space or if the allocation tagging requirements are changed on the pool, the resource may be marked as noncompliant.
:param auto_import: If selected, IPAM will continuously look for resources within the CIDR range of this pool and automatically import them as allocations into your IPAM. The CIDRs that will be allocated for these resources must not already be allocated to other resources in order for the import to succeed. IPAM will import a CIDR regardless of its compliance with the pool's allocation rules, so a resource might be imported and subsequently marked as noncompliant. If IPAM discovers multiple CIDRs that overlap, IPAM will import the largest CIDR only. If IPAM discovers multiple CIDRs with matching CIDRs, IPAM will randomly import one of them only. A locale must be set on the pool for this feature to work.
:param description: The description of the pool.
:param locale: The locale of the IPAM pool. In IPAM, the locale is the AWS Region where you want to make an IPAM pool available for allocations.Only resources in the same Region as the locale of the pool can get IP address allocations from the pool. You can only allocate a CIDR for a VPC, for example, from an IPAM pool that shares a locale with the VPC’s Region. Note that once you choose a Locale for a pool, you cannot modify it. If you choose an AWS Region for locale that has not been configured as an operating Region for the IPAM, you'll get an error.
:param provisioned_cidrs: The CIDRs provisioned to the IPAM pool. A CIDR is a representation of an IP address and its associated network mask (or netmask) and refers to a range of IP addresses
:param source_ipam_pool_id: The ID of the source IPAM pool. You can use this option to create an IPAM pool within an existing source pool.
:param tags: The key/value combination of tags to assign to the resource.
'''
props = IpamPoolProps(
address_family=address_family,
ipam_scope_id=ipam_scope_id,
allocation_default_netmask_length=allocation_default_netmask_length,
allocation_max_netmask_length=allocation_max_netmask_length,
allocation_min_netmask_length=allocation_min_netmask_length,
allocation_resource_tags=allocation_resource_tags,
auto_import=auto_import,
description=description,
locale=locale,
provisioned_cidrs=provisioned_cidrs,
source_ipam_pool_id=source_ipam_pool_id,
tags=tags,
)
jsii.create(self.__class__, self, [scope, id, props])
@jsii.member(jsii_name="provisionCidr")
def provision_cidr(self, cidr: builtins.str) -> None:
'''Adds a CIDR to the pool.
:param cidr: The CIDR to add to the pool.
'''
return typing.cast(None, jsii.invoke(self, "provisionCidr", [cidr]))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="arn")
def arn(self) -> builtins.str:
'''The ARN of the resulting IPAM Pool resource.'''
return typing.cast(builtins.str, jsii.get(self, "arn"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamArn")
def ipam_arn(self) -> builtins.str:
'''The ARN of the IPAM this pool belongs to.'''
return typing.cast(builtins.str, jsii.get(self, "ipamArn"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamPoolId")
def ipam_pool_id(self) -> builtins.str:
'''The ID of the resulting IPAM Pool resource.'''
return typing.cast(builtins.str, jsii.get(self, "ipamPoolId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamScopeArn")
def ipam_scope_arn(self) -> builtins.str:
'''The ARN of the scope of the IPAM Pool.'''
return typing.cast(builtins.str, jsii.get(self, "ipamScopeArn"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamScopeType")
def ipam_scope_type(self) -> builtins.str:
'''The IPAM scope type (public or private) of the scope of the IPAM Pool.'''
return typing.cast(builtins.str, jsii.get(self, "ipamScopeType"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="pool")
def pool(self) -> aws_cdk.aws_ec2.CfnIPAMPool:
'''The underlying IPAM Pool resource.'''
return typing.cast(aws_cdk.aws_ec2.CfnIPAMPool, jsii.get(self, "pool"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="poolDepth")
def pool_depth(self) -> jsii.Number:
'''The depth of pools in your IPAM pool.'''
return typing.cast(jsii.Number, jsii.get(self, "poolDepth"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="provisionedCidrs")
def provisioned_cidrs(
self,
) -> typing.List[aws_cdk.aws_ec2.CfnIPAMPool.ProvisionedCidrProperty]:
'''The provisioned CIDRs for this pool.'''
return typing.cast(typing.List[aws_cdk.aws_ec2.CfnIPAMPool.ProvisionedCidrProperty], jsii.get(self, "provisionedCidrs"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="state")
def state(self) -> builtins.str:
'''The state of the IPAM pool.'''
return typing.cast(builtins.str, jsii.get(self, "state"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="stateMessage")
def state_message(self) -> builtins.str:
'''A message related to the failed creation of an IPAM pool.'''
return typing.cast(builtins.str, jsii.get(self, "stateMessage"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="allocationDefaultNetmaskLength")
def allocation_default_netmask_length(self) -> typing.Optional[jsii.Number]:
'''The default netmask length for allocations added to this pool.'''
return typing.cast(typing.Optional[jsii.Number], jsii.get(self, "allocationDefaultNetmaskLength"))
@jsii.enum(jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamPoolAddressFamily")
class IpamPoolAddressFamily(enum.Enum):
IPV4 = "IPV4"
IPV6 = "IPV6"
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamPoolProps",
jsii_struct_bases=[],
name_mapping={
"address_family": "addressFamily",
"ipam_scope_id": "ipamScopeId",
"allocation_default_netmask_length": "allocationDefaultNetmaskLength",
"allocation_max_netmask_length": "allocationMaxNetmaskLength",
"allocation_min_netmask_length": "allocationMinNetmaskLength",
"allocation_resource_tags": "allocationResourceTags",
"auto_import": "autoImport",
"description": "description",
"locale": "locale",
"provisioned_cidrs": "provisionedCidrs",
"source_ipam_pool_id": "sourceIpamPoolId",
"tags": "tags",
},
)
class IpamPoolProps:
def __init__(
self,
*,
address_family: IpamPoolAddressFamily,
ipam_scope_id: builtins.str,
allocation_default_netmask_length: typing.Optional[jsii.Number] = None,
allocation_max_netmask_length: typing.Optional[jsii.Number] = None,
allocation_min_netmask_length: typing.Optional[jsii.Number] = None,
allocation_resource_tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
auto_import: typing.Optional[builtins.bool] = None,
description: typing.Optional[builtins.str] = None,
locale: typing.Optional[builtins.str] = None,
provisioned_cidrs: typing.Optional[typing.Sequence[aws_cdk.aws_ec2.CfnIPAMPool.ProvisionedCidrProperty]] = None,
source_ipam_pool_id: typing.Optional[builtins.str] = None,
tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
) -> None:
'''Properties of an IPAM Pool.
:param address_family: The address family of the pool, either IPv4 or IPv6.
:param ipam_scope_id: The IPAM scope this pool is associated with.
:param allocation_default_netmask_length: The default netmask length for allocations added to this pool. If, for example, the CIDR assigned to this pool is 10.0.0.0/8 and you enter 16 here, new allocations will default to 10.0.0.0/16.
:param allocation_max_netmask_length: The maximum netmask length possible for CIDR allocations in this IPAM pool to be compliant. The maximum netmask length must be greater than the minimum netmask length. Possible netmask lengths for IPv4 addresses are 0 - 32. Possible netmask lengths for IPv6 addresses are 0 - 128.
:param allocation_min_netmask_length: The minimum netmask length required for CIDR allocations in this IPAM pool to be compliant. The minimum netmask length must be less than the maximum netmask length. Possible netmask lengths for IPv4 addresses are 0 - 32. Possible netmask lengths for IPv6 addresses are 0 - 128.
:param allocation_resource_tags: Tags that are required for resources that use CIDRs from this IPAM pool. Resources that do not have these tags will not be allowed to allocate space from the pool. If the resources have their tags changed after they have allocated space or if the allocation tagging requirements are changed on the pool, the resource may be marked as noncompliant.
:param auto_import: If selected, IPAM will continuously look for resources within the CIDR range of this pool and automatically import them as allocations into your IPAM. The CIDRs that will be allocated for these resources must not already be allocated to other resources in order for the import to succeed. IPAM will import a CIDR regardless of its compliance with the pool's allocation rules, so a resource might be imported and subsequently marked as noncompliant. If IPAM discovers multiple CIDRs that overlap, IPAM will import the largest CIDR only. If IPAM discovers multiple CIDRs with matching CIDRs, IPAM will randomly import one of them only. A locale must be set on the pool for this feature to work.
:param description: The description of the pool.
:param locale: The locale of the IPAM pool. In IPAM, the locale is the AWS Region where you want to make an IPAM pool available for allocations.Only resources in the same Region as the locale of the pool can get IP address allocations from the pool. You can only allocate a CIDR for a VPC, for example, from an IPAM pool that shares a locale with the VPC’s Region. Note that once you choose a Locale for a pool, you cannot modify it. If you choose an AWS Region for locale that has not been configured as an operating Region for the IPAM, you'll get an error.
:param provisioned_cidrs: The CIDRs provisioned to the IPAM pool. A CIDR is a representation of an IP address and its associated network mask (or netmask) and refers to a range of IP addresses
:param source_ipam_pool_id: The ID of the source IPAM pool. You can use this option to create an IPAM pool within an existing source pool.
:param tags: The key/value combination of tags to assign to the resource.
'''
self._values: typing.Dict[str, typing.Any] = {
"address_family": address_family,
"ipam_scope_id": ipam_scope_id,
}
if allocation_default_netmask_length is not None:
self._values["allocation_default_netmask_length"] = allocation_default_netmask_length
if allocation_max_netmask_length is not None:
self._values["allocation_max_netmask_length"] = allocation_max_netmask_length
if allocation_min_netmask_length is not None:
self._values["allocation_min_netmask_length"] = allocation_min_netmask_length
if allocation_resource_tags is not None:
self._values["allocation_resource_tags"] = allocation_resource_tags
if auto_import is not None:
self._values["auto_import"] = auto_import
if description is not None:
self._values["description"] = description
if locale is not None:
self._values["locale"] = locale
if provisioned_cidrs is not None:
self._values["provisioned_cidrs"] = provisioned_cidrs
if source_ipam_pool_id is not None:
self._values["source_ipam_pool_id"] = source_ipam_pool_id
if tags is not None:
self._values["tags"] = tags
@builtins.property
def address_family(self) -> IpamPoolAddressFamily:
'''The address family of the pool, either IPv4 or IPv6.'''
result = self._values.get("address_family")
assert result is not None, "Required property 'address_family' is missing"
return typing.cast(IpamPoolAddressFamily, result)
@builtins.property
def ipam_scope_id(self) -> builtins.str:
'''The IPAM scope this pool is associated with.'''
result = self._values.get("ipam_scope_id")
assert result is not None, "Required property 'ipam_scope_id' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def allocation_default_netmask_length(self) -> typing.Optional[jsii.Number]:
'''The default netmask length for allocations added to this pool.
If, for example, the CIDR assigned to this pool is 10.0.0.0/8 and you enter 16 here,
new allocations will default to 10.0.0.0/16.
'''
result = self._values.get("allocation_default_netmask_length")
return typing.cast(typing.Optional[jsii.Number], result)
@builtins.property
def allocation_max_netmask_length(self) -> typing.Optional[jsii.Number]:
'''The maximum netmask length possible for CIDR allocations in this IPAM pool to be compliant.
The maximum netmask length must be greater than the minimum netmask length.
Possible netmask lengths for IPv4 addresses are 0 - 32. Possible netmask lengths for IPv6 addresses are 0 - 128.
'''
result = self._values.get("allocation_max_netmask_length")
return typing.cast(typing.Optional[jsii.Number], result)
@builtins.property
def allocation_min_netmask_length(self) -> typing.Optional[jsii.Number]:
'''The minimum netmask length required for CIDR allocations in this IPAM pool to be compliant.
The minimum netmask length must be less than the maximum netmask length.
Possible netmask lengths for IPv4 addresses are 0 - 32. Possible netmask lengths for IPv6 addresses are 0 - 128.
'''
result = self._values.get("allocation_min_netmask_length")
return typing.cast(typing.Optional[jsii.Number], result)
@builtins.property
def allocation_resource_tags(self) -> typing.Optional[typing.List[aws_cdk.CfnTag]]:
'''Tags that are required for resources that use CIDRs from this IPAM pool.
Resources that do not have these tags will not be allowed to allocate space from the pool.
If the resources have their tags changed after they have allocated space or if the allocation
tagging requirements are changed on the pool, the resource may be marked as noncompliant.
'''
result = self._values.get("allocation_resource_tags")
return typing.cast(typing.Optional[typing.List[aws_cdk.CfnTag]], result)
@builtins.property
def auto_import(self) -> typing.Optional[builtins.bool]:
'''If selected, IPAM will continuously look for resources within the CIDR range of this pool and automatically import them as allocations into your IPAM.
The CIDRs that will be allocated for these resources must not already be allocated
to other resources in order for the import to succeed. IPAM will import a CIDR regardless of its compliance with the
pool's allocation rules, so a resource might be imported and subsequently marked as noncompliant. If IPAM discovers
multiple CIDRs that overlap, IPAM will import the largest CIDR only. If IPAM discovers multiple CIDRs with matching
CIDRs, IPAM will randomly import one of them only.
A locale must be set on the pool for this feature to work.
'''
result = self._values.get("auto_import")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''The description of the pool.'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def locale(self) -> typing.Optional[builtins.str]:
'''The locale of the IPAM pool.
In IPAM, the locale is the AWS Region where you want to make an IPAM pool available
for allocations.Only resources in the same Region as the locale of the pool can get IP address allocations from the pool.
You can only allocate a CIDR for a VPC, for example, from an IPAM pool that shares a locale with the VPC’s Region.
Note that once you choose a Locale for a pool, you cannot modify it. If you choose an AWS Region for locale that has
not been configured as an operating Region for the IPAM, you'll get an error.
'''
result = self._values.get("locale")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def provisioned_cidrs(
self,
) -> typing.Optional[typing.List[aws_cdk.aws_ec2.CfnIPAMPool.ProvisionedCidrProperty]]:
'''The CIDRs provisioned to the IPAM pool.
A CIDR is a representation of an IP address and its associated network mask
(or netmask) and refers to a range of IP addresses
'''
result = self._values.get("provisioned_cidrs")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_ec2.CfnIPAMPool.ProvisionedCidrProperty]], result)
@builtins.property
def source_ipam_pool_id(self) -> typing.Optional[builtins.str]:
'''The ID of the source IPAM pool.
You can use this option to create an IPAM pool within an existing source pool.
'''
result = self._values.get("source_ipam_pool_id")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def tags(self) -> typing.Optional[typing.List[aws_cdk.CfnTag]]:
'''The key/value combination of tags to assign to the resource.'''
result = self._values.get("tags")
return typing.cast(typing.Optional[typing.List[aws_cdk.CfnTag]], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "IpamPoolProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamProps",
jsii_struct_bases=[],
name_mapping={
"description": "description",
"operating_regions": "operatingRegions",
"tags": "tags",
},
)
class IpamProps:
def __init__(
self,
*,
description: typing.Optional[builtins.str] = None,
operating_regions: typing.Optional[typing.Sequence[builtins.str]] = None,
tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
) -> None:
'''Properties of the IPAM.
:param description: The description for the IPAM.
:param operating_regions: The operating Regions for an IPAM. Operating Regions are AWS Regions where the IPAM is allowed to manage IP address CIDRs. IPAM only discovers and monitors resources in the AWS Regions you select as operating Regions. For more information about operating Regions, see Create an IPAM in the Amazon VPC IPAM User Guide. Default: Stack.of(this).region
:param tags: The key/value combination of tags to assign to the resource.
'''
self._values: typing.Dict[str, typing.Any] = {}
if description is not None:
self._values["description"] = description
if operating_regions is not None:
self._values["operating_regions"] = operating_regions
if tags is not None:
self._values["tags"] = tags
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''The description for the IPAM.'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def operating_regions(self) -> typing.Optional[typing.List[builtins.str]]:
'''The operating Regions for an IPAM.
Operating Regions are AWS Regions where the IPAM is allowed to manage IP address CIDRs. IPAM only
discovers and monitors resources in the AWS Regions you select as operating Regions.
For more information about operating Regions, see Create an IPAM in the Amazon VPC IPAM User Guide.
:default: Stack.of(this).region
:see: https://vpc/latest/ipam/create-ipam.html
'''
result = self._values.get("operating_regions")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
@builtins.property
def tags(self) -> typing.Optional[typing.List[aws_cdk.CfnTag]]:
'''The key/value combination of tags to assign to the resource.'''
result = self._values.get("tags")
return typing.cast(typing.Optional[typing.List[aws_cdk.CfnTag]], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "IpamProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class IpamScope(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamScope",
):
'''An IPAM Scope.
In IPAM, a scope is the highest-level container within IPAM. An IPAM contains two default scopes.
Each scope represents the IP space for a single network. The private scope is intended for all private
IP address space. The public scope is intended for all public IP address space. Scopes enable you to
reuse IP addresses across multiple unconnected networks without causing IP address overlap or conflict.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
ipam: Ipam,
description: typing.Optional[builtins.str] = None,
tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param ipam: The IPAM for which you're creating the scope.
:param description: The description of the scope.
:param tags: The key/value combination of tags to assign to the resource.
'''
props = IpamScopeProps(ipam=ipam, description=description, tags=tags)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="arn")
def arn(self) -> builtins.str:
'''The ARN of the resulting IPAM Scope resource.'''
return typing.cast(builtins.str, jsii.get(self, "arn"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamArn")
def ipam_arn(self) -> builtins.str:
'''The ARN of the IPAM this scope belongs to.'''
return typing.cast(builtins.str, jsii.get(self, "ipamArn"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="ipamScopeId")
def ipam_scope_id(self) -> builtins.str:
'''The ID of the resulting IPAM Scope resource.'''
return typing.cast(builtins.str, jsii.get(self, "ipamScopeId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="isDefault")
def is_default(self) -> aws_cdk.IResolvable:
'''Indicates whether the scope is the default scope for the IPAM.'''
return typing.cast(aws_cdk.IResolvable, jsii.get(self, "isDefault"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="poolCount")
def pool_count(self) -> jsii.Number:
'''The number of pools in the scope.'''
return typing.cast(jsii.Number, jsii.get(self, "poolCount"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="scope")
def scope(self) -> aws_cdk.aws_ec2.CfnIPAMScope:
'''The underlying IPAM Scope resource.'''
return typing.cast(aws_cdk.aws_ec2.CfnIPAMScope, jsii.get(self, "scope"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-ipam.IpamScopeProps",
jsii_struct_bases=[],
name_mapping={"ipam": "ipam", "description": "description", "tags": "tags"},
)
class IpamScopeProps:
def __init__(
self,
*,
ipam: Ipam,
description: typing.Optional[builtins.str] = None,
tags: typing.Optional[typing.Sequence[aws_cdk.CfnTag]] = None,
) -> None:
'''Properties of an IPAM Scope.
:param ipam: The IPAM for which you're creating the scope.
:param description: The description of the scope.
:param tags: The key/value combination of tags to assign to the resource.
'''
self._values: typing.Dict[str, typing.Any] = {
"ipam": ipam,
}
if description is not None:
self._values["description"] = description
if tags is not None:
self._values["tags"] = tags
@builtins.property
def ipam(self) -> Ipam:
'''The IPAM for which you're creating the scope.'''
result = self._values.get("ipam")
assert result is not None, "Required property 'ipam' is missing"
return typing.cast(Ipam, result)
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''The description of the scope.'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def tags(self) -> typing.Optional[typing.List[aws_cdk.CfnTag]]:
'''The key/value combination of tags to assign to the resource.'''
result = self._values.get("tags")
return typing.cast(typing.Optional[typing.List[aws_cdk.CfnTag]], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "IpamScopeProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"Ipam",
"IpamAllocation",
"IpamAllocationProps",
"IpamPool",
"IpamPoolAddressFamily",
"IpamPoolProps",
"IpamProps",
"IpamScope",
"IpamScopeProps",
]
publication.publish() | /renovosolutions.aws-cdk-aws-ipam-0.1.107.tar.gz/renovosolutions.aws-cdk-aws-ipam-0.1.107/src/ipam/__init__.py | 0.71103 | 0.212538 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk
import aws_cdk.aws_iam
import aws_cdk.custom_resources
import constructs
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.AccountProps",
jsii_struct_bases=[],
name_mapping={
"email": "email",
"name": "name",
"allow_move": "allowMove",
"disable_delete": "disableDelete",
"import_on_duplicate": "importOnDuplicate",
},
)
class AccountProps:
def __init__(
self,
*,
email: builtins.str,
name: builtins.str,
allow_move: typing.Optional[builtins.bool] = None,
disable_delete: typing.Optional[builtins.bool] = None,
import_on_duplicate: typing.Optional[builtins.bool] = None,
) -> None:
'''The properties of an Account.
:param email: The email address of the account. Most be unique.
:param name: The name of the account.
:param allow_move: Whether or not to allow this account to be moved between OUs. If importing is enabled this will also allow imported accounts to be moved. Default: false
:param disable_delete: Whether or not attempting to delete an account should raise an error. Accounts cannot be deleted programmatically, but they can be removed as a managed resource. This property will allow you to control whether or not an error is thrown when the stack wants to delete an account (orphan it) or if it should continue silently. Default: false
:param import_on_duplicate: Whether or not to import an existing account if the new account is a duplicate. If this is false and the account already exists an error will be thrown. Default: false
'''
self._values: typing.Dict[str, typing.Any] = {
"email": email,
"name": name,
}
if allow_move is not None:
self._values["allow_move"] = allow_move
if disable_delete is not None:
self._values["disable_delete"] = disable_delete
if import_on_duplicate is not None:
self._values["import_on_duplicate"] = import_on_duplicate
@builtins.property
def email(self) -> builtins.str:
'''The email address of the account.
Most be unique.
'''
result = self._values.get("email")
assert result is not None, "Required property 'email' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def name(self) -> builtins.str:
'''The name of the account.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def allow_move(self) -> typing.Optional[builtins.bool]:
'''Whether or not to allow this account to be moved between OUs.
If importing is enabled
this will also allow imported accounts to be moved.
:default: false
'''
result = self._values.get("allow_move")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def disable_delete(self) -> typing.Optional[builtins.bool]:
'''Whether or not attempting to delete an account should raise an error.
Accounts cannot be deleted programmatically, but they can be removed as a managed resource.
This property will allow you to control whether or not an error is thrown
when the stack wants to delete an account (orphan it) or if it should continue
silently.
:default: false
:see: https://aws.amazon.com/premiumsupport/knowledge-center/close-aws-account/
'''
result = self._values.get("disable_delete")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def import_on_duplicate(self) -> typing.Optional[builtins.bool]:
'''Whether or not to import an existing account if the new account is a duplicate.
If this is false and the account already exists an error will be thrown.
:default: false
'''
result = self._values.get("import_on_duplicate")
return typing.cast(typing.Optional[builtins.bool], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AccountProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.AccountResourceProps",
jsii_struct_bases=[AccountProps],
name_mapping={
"email": "email",
"name": "name",
"allow_move": "allowMove",
"disable_delete": "disableDelete",
"import_on_duplicate": "importOnDuplicate",
"parent": "parent",
"provider": "provider",
},
)
class AccountResourceProps(AccountProps):
def __init__(
self,
*,
email: builtins.str,
name: builtins.str,
allow_move: typing.Optional[builtins.bool] = None,
disable_delete: typing.Optional[builtins.bool] = None,
import_on_duplicate: typing.Optional[builtins.bool] = None,
parent: typing.Union[builtins.str, "OrganizationOU"],
provider: aws_cdk.custom_resources.Provider,
) -> None:
'''The properties of an OrganizationAccount custom resource.
:param email: The email address of the account. Most be unique.
:param name: The name of the account.
:param allow_move: Whether or not to allow this account to be moved between OUs. If importing is enabled this will also allow imported accounts to be moved. Default: false
:param disable_delete: Whether or not attempting to delete an account should raise an error. Accounts cannot be deleted programmatically, but they can be removed as a managed resource. This property will allow you to control whether or not an error is thrown when the stack wants to delete an account (orphan it) or if it should continue silently. Default: false
:param import_on_duplicate: Whether or not to import an existing account if the new account is a duplicate. If this is false and the account already exists an error will be thrown. Default: false
:param parent: The parent OU id.
:param provider: The provider to use for the custom resource that will create the OU. You can create a provider with the OrganizationOuProvider class
'''
self._values: typing.Dict[str, typing.Any] = {
"email": email,
"name": name,
"parent": parent,
"provider": provider,
}
if allow_move is not None:
self._values["allow_move"] = allow_move
if disable_delete is not None:
self._values["disable_delete"] = disable_delete
if import_on_duplicate is not None:
self._values["import_on_duplicate"] = import_on_duplicate
@builtins.property
def email(self) -> builtins.str:
'''The email address of the account.
Most be unique.
'''
result = self._values.get("email")
assert result is not None, "Required property 'email' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def name(self) -> builtins.str:
'''The name of the account.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def allow_move(self) -> typing.Optional[builtins.bool]:
'''Whether or not to allow this account to be moved between OUs.
If importing is enabled
this will also allow imported accounts to be moved.
:default: false
'''
result = self._values.get("allow_move")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def disable_delete(self) -> typing.Optional[builtins.bool]:
'''Whether or not attempting to delete an account should raise an error.
Accounts cannot be deleted programmatically, but they can be removed as a managed resource.
This property will allow you to control whether or not an error is thrown
when the stack wants to delete an account (orphan it) or if it should continue
silently.
:default: false
:see: https://aws.amazon.com/premiumsupport/knowledge-center/close-aws-account/
'''
result = self._values.get("disable_delete")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def import_on_duplicate(self) -> typing.Optional[builtins.bool]:
'''Whether or not to import an existing account if the new account is a duplicate.
If this is false and the account already exists an error will be thrown.
:default: false
'''
result = self._values.get("import_on_duplicate")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def parent(self) -> typing.Union[builtins.str, "OrganizationOU"]:
'''The parent OU id.'''
result = self._values.get("parent")
assert result is not None, "Required property 'parent' is missing"
return typing.cast(typing.Union[builtins.str, "OrganizationOU"], result)
@builtins.property
def provider(self) -> aws_cdk.custom_resources.Provider:
'''The provider to use for the custom resource that will create the OU.
You can create a provider with the OrganizationOuProvider class
'''
result = self._values.get("provider")
assert result is not None, "Required property 'provider' is missing"
return typing.cast(aws_cdk.custom_resources.Provider, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AccountResourceProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.interface(
jsii_type="@renovosolutions/cdk-library-aws-organization.IPAMAdministratorProps"
)
class IPAMAdministratorProps(typing_extensions.Protocol):
'''The properties of an OrganizationAccount custom resource.'''
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="delegatedAdminAccountId")
def delegated_admin_account_id(self) -> builtins.str:
'''The account id of the IPAM administrator.'''
...
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="provider")
def provider(self) -> aws_cdk.custom_resources.Provider:
'''The provider to use for the custom resource that will handle IPAM admin delegation.
You can create a provider with the IPAMAdministratorProvider class
'''
...
class _IPAMAdministratorPropsProxy:
'''The properties of an OrganizationAccount custom resource.'''
__jsii_type__: typing.ClassVar[str] = "@renovosolutions/cdk-library-aws-organization.IPAMAdministratorProps"
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="delegatedAdminAccountId")
def delegated_admin_account_id(self) -> builtins.str:
'''The account id of the IPAM administrator.'''
return typing.cast(builtins.str, jsii.get(self, "delegatedAdminAccountId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="provider")
def provider(self) -> aws_cdk.custom_resources.Provider:
'''The provider to use for the custom resource that will handle IPAM admin delegation.
You can create a provider with the IPAMAdministratorProvider class
'''
return typing.cast(aws_cdk.custom_resources.Provider, jsii.get(self, "provider"))
# Adding a "__jsii_proxy_class__(): typing.Type" function to the interface
typing.cast(typing.Any, IPAMAdministratorProps).__jsii_proxy_class__ = lambda : _IPAMAdministratorPropsProxy
class IPAMAdministratorProvider(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-organization.IPAMAdministratorProvider",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
props: "IPAMAdministratorProviderProps",
) -> None:
'''
:param scope: -
:param id: -
:param props: -
'''
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="provider")
def provider(self) -> aws_cdk.custom_resources.Provider:
return typing.cast(aws_cdk.custom_resources.Provider, jsii.get(self, "provider"))
@jsii.interface(
jsii_type="@renovosolutions/cdk-library-aws-organization.IPAMAdministratorProviderProps"
)
class IPAMAdministratorProviderProps(typing_extensions.Protocol):
'''The properties of an IPAM administrator custom resource provider.'''
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="role")
def role(self) -> typing.Optional[aws_cdk.aws_iam.IRole]:
'''The role the custom resource should use for working with the IPAM administrator delegation if one is not provided one will be created automatically.'''
...
class _IPAMAdministratorProviderPropsProxy:
'''The properties of an IPAM administrator custom resource provider.'''
__jsii_type__: typing.ClassVar[str] = "@renovosolutions/cdk-library-aws-organization.IPAMAdministratorProviderProps"
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="role")
def role(self) -> typing.Optional[aws_cdk.aws_iam.IRole]:
'''The role the custom resource should use for working with the IPAM administrator delegation if one is not provided one will be created automatically.'''
return typing.cast(typing.Optional[aws_cdk.aws_iam.IRole], jsii.get(self, "role"))
# Adding a "__jsii_proxy_class__(): typing.Type" function to the interface
typing.cast(typing.Any, IPAMAdministratorProviderProps).__jsii_proxy_class__ = lambda : _IPAMAdministratorProviderPropsProxy
class IPAMdministrator(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-organization.IPAMdministrator",
):
'''The construct to create or update the delegated IPAM administrator for an organization.
This relies on the custom resource provider IPAMAdministratorProvider.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
props: IPAMAdministratorProps,
) -> None:
'''
:param scope: -
:param id: -
:param props: -
'''
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="resource")
def resource(self) -> aws_cdk.CustomResource:
return typing.cast(aws_cdk.CustomResource, jsii.get(self, "resource"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.OUObject",
jsii_struct_bases=[],
name_mapping={
"children": "children",
"properties": "properties",
"accounts": "accounts",
"id": "id",
},
)
class OUObject:
def __init__(
self,
*,
children: typing.Sequence["OUObject"],
properties: "OUProps",
accounts: typing.Optional[typing.Sequence[AccountProps]] = None,
id: typing.Optional[builtins.str] = None,
) -> None:
'''The structure of an OrgObject.
:param children: OUs that are children of this OU.
:param properties: The OU object properties.
:param accounts: Accounts that belong to this OU.
:param id: The unique id of the OUObject. This is used as the unique identifier when instantiating a construct object. This is important for the CDK to be able to maintain a reference for the object when utilizing the processOUObj function rather then using the name property of an object which could change. If the id changes the CDK treats this as a new construct and will create a new construct resource and destroy the old one. Not strictly required but useful when using the processOUObj function. If the id is not provided the name property will be used as the id in processOUObj. You can create a unique id however you like. A bash example is provided.
'''
if isinstance(properties, dict):
properties = OUProps(**properties)
self._values: typing.Dict[str, typing.Any] = {
"children": children,
"properties": properties,
}
if accounts is not None:
self._values["accounts"] = accounts
if id is not None:
self._values["id"] = id
@builtins.property
def children(self) -> typing.List["OUObject"]:
'''OUs that are children of this OU.'''
result = self._values.get("children")
assert result is not None, "Required property 'children' is missing"
return typing.cast(typing.List["OUObject"], result)
@builtins.property
def properties(self) -> "OUProps":
'''The OU object properties.'''
result = self._values.get("properties")
assert result is not None, "Required property 'properties' is missing"
return typing.cast("OUProps", result)
@builtins.property
def accounts(self) -> typing.Optional[typing.List[AccountProps]]:
'''Accounts that belong to this OU.'''
result = self._values.get("accounts")
return typing.cast(typing.Optional[typing.List[AccountProps]], result)
@builtins.property
def id(self) -> typing.Optional[builtins.str]:
'''The unique id of the OUObject.
This is used as the unique identifier when instantiating a construct object.
This is important for the CDK to be able to maintain a reference for the object when utilizing
the processOUObj function rather then using the name property of an object which could change.
If the id changes the CDK treats this as a new construct and will create a new construct resource and
destroy the old one.
Not strictly required but useful when using the processOUObj function. If the id is not provided
the name property will be used as the id in processOUObj.
You can create a unique id however you like. A bash example is provided.
Example::
echo "ou-$( echo $RANDOM | md5sum | head -c 8 )"
'''
result = self._values.get("id")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OUObject(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.OUProps",
jsii_struct_bases=[],
name_mapping={
"name": "name",
"allow_recreate_on_update": "allowRecreateOnUpdate",
"import_on_duplicate": "importOnDuplicate",
},
)
class OUProps:
def __init__(
self,
*,
name: builtins.str,
allow_recreate_on_update: typing.Optional[builtins.bool] = None,
import_on_duplicate: typing.Optional[builtins.bool] = None,
) -> None:
'''The properties of an OU.
:param name: The name of the OU.
:param allow_recreate_on_update: Whether or not a missing OU should be recreated during an update. If this is false and the OU does not exist an error will be thrown when you try to update it. Default: false
:param import_on_duplicate: Whether or not to import an existing OU if the new OU is a duplicate. If this is false and the OU already exists an error will be thrown. Default: false
'''
self._values: typing.Dict[str, typing.Any] = {
"name": name,
}
if allow_recreate_on_update is not None:
self._values["allow_recreate_on_update"] = allow_recreate_on_update
if import_on_duplicate is not None:
self._values["import_on_duplicate"] = import_on_duplicate
@builtins.property
def name(self) -> builtins.str:
'''The name of the OU.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def allow_recreate_on_update(self) -> typing.Optional[builtins.bool]:
'''Whether or not a missing OU should be recreated during an update.
If this is false and the OU does not exist an error will be thrown when you try to update it.
:default: false
'''
result = self._values.get("allow_recreate_on_update")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def import_on_duplicate(self) -> typing.Optional[builtins.bool]:
'''Whether or not to import an existing OU if the new OU is a duplicate.
If this is false and the OU already exists an error will be thrown.
:default: false
'''
result = self._values.get("import_on_duplicate")
return typing.cast(typing.Optional[builtins.bool], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OUProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.OUResourceProps",
jsii_struct_bases=[OUProps],
name_mapping={
"name": "name",
"allow_recreate_on_update": "allowRecreateOnUpdate",
"import_on_duplicate": "importOnDuplicate",
"parent": "parent",
"provider": "provider",
},
)
class OUResourceProps(OUProps):
def __init__(
self,
*,
name: builtins.str,
allow_recreate_on_update: typing.Optional[builtins.bool] = None,
import_on_duplicate: typing.Optional[builtins.bool] = None,
parent: typing.Union[builtins.str, "OrganizationOU"],
provider: aws_cdk.custom_resources.Provider,
) -> None:
'''The properties of an OrganizationOU custom resource.
:param name: The name of the OU.
:param allow_recreate_on_update: Whether or not a missing OU should be recreated during an update. If this is false and the OU does not exist an error will be thrown when you try to update it. Default: false
:param import_on_duplicate: Whether or not to import an existing OU if the new OU is a duplicate. If this is false and the OU already exists an error will be thrown. Default: false
:param parent: The parent OU id.
:param provider: The provider to use for the custom resource that will create the OU. You can create a provider with the OrganizationOuProvider class
'''
self._values: typing.Dict[str, typing.Any] = {
"name": name,
"parent": parent,
"provider": provider,
}
if allow_recreate_on_update is not None:
self._values["allow_recreate_on_update"] = allow_recreate_on_update
if import_on_duplicate is not None:
self._values["import_on_duplicate"] = import_on_duplicate
@builtins.property
def name(self) -> builtins.str:
'''The name of the OU.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def allow_recreate_on_update(self) -> typing.Optional[builtins.bool]:
'''Whether or not a missing OU should be recreated during an update.
If this is false and the OU does not exist an error will be thrown when you try to update it.
:default: false
'''
result = self._values.get("allow_recreate_on_update")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def import_on_duplicate(self) -> typing.Optional[builtins.bool]:
'''Whether or not to import an existing OU if the new OU is a duplicate.
If this is false and the OU already exists an error will be thrown.
:default: false
'''
result = self._values.get("import_on_duplicate")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def parent(self) -> typing.Union[builtins.str, "OrganizationOU"]:
'''The parent OU id.'''
result = self._values.get("parent")
assert result is not None, "Required property 'parent' is missing"
return typing.cast(typing.Union[builtins.str, "OrganizationOU"], result)
@builtins.property
def provider(self) -> aws_cdk.custom_resources.Provider:
'''The provider to use for the custom resource that will create the OU.
You can create a provider with the OrganizationOuProvider class
'''
result = self._values.get("provider")
assert result is not None, "Required property 'provider' is missing"
return typing.cast(aws_cdk.custom_resources.Provider, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OUResourceProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class OrganizationAccount(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-organization.OrganizationAccount",
):
'''The construct to create or update an Organization account.
This relies on the custom resource provider OrganizationAccountProvider.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
parent: typing.Union[builtins.str, "OrganizationOU"],
provider: aws_cdk.custom_resources.Provider,
email: builtins.str,
name: builtins.str,
allow_move: typing.Optional[builtins.bool] = None,
disable_delete: typing.Optional[builtins.bool] = None,
import_on_duplicate: typing.Optional[builtins.bool] = None,
) -> None:
'''
:param scope: -
:param id: -
:param parent: The parent OU id.
:param provider: The provider to use for the custom resource that will create the OU. You can create a provider with the OrganizationOuProvider class
:param email: The email address of the account. Most be unique.
:param name: The name of the account.
:param allow_move: Whether or not to allow this account to be moved between OUs. If importing is enabled this will also allow imported accounts to be moved. Default: false
:param disable_delete: Whether or not attempting to delete an account should raise an error. Accounts cannot be deleted programmatically, but they can be removed as a managed resource. This property will allow you to control whether or not an error is thrown when the stack wants to delete an account (orphan it) or if it should continue silently. Default: false
:param import_on_duplicate: Whether or not to import an existing account if the new account is a duplicate. If this is false and the account already exists an error will be thrown. Default: false
'''
props = AccountResourceProps(
parent=parent,
provider=provider,
email=email,
name=name,
allow_move=allow_move,
disable_delete=disable_delete,
import_on_duplicate=import_on_duplicate,
)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="resource")
def resource(self) -> aws_cdk.CustomResource:
return typing.cast(aws_cdk.CustomResource, jsii.get(self, "resource"))
class OrganizationAccountProvider(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-organization.OrganizationAccountProvider",
):
'''The provider for account custom resources.
This creates a lambda function that handles custom resource requests for creating/updating accounts.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
role: typing.Optional[aws_cdk.aws_iam.IRole] = None,
) -> None:
'''
:param scope: -
:param id: -
:param role: The role the custom resource should use for taking actions on OUs if one is not provided one will be created automatically.
'''
props = OrganizationOUProviderProps(role=role)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="provider")
def provider(self) -> aws_cdk.custom_resources.Provider:
return typing.cast(aws_cdk.custom_resources.Provider, jsii.get(self, "provider"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.OrganizationAccountProviderProps",
jsii_struct_bases=[],
name_mapping={"role": "role"},
)
class OrganizationAccountProviderProps:
def __init__(self, *, role: typing.Optional[aws_cdk.aws_iam.IRole] = None) -> None:
'''The properties for the account custom resource provider.
:param role: The role the custom resource should use for taking actions on OUs if one is not provided one will be created automatically.
'''
self._values: typing.Dict[str, typing.Any] = {}
if role is not None:
self._values["role"] = role
@builtins.property
def role(self) -> typing.Optional[aws_cdk.aws_iam.IRole]:
'''The role the custom resource should use for taking actions on OUs if one is not provided one will be created automatically.'''
result = self._values.get("role")
return typing.cast(typing.Optional[aws_cdk.aws_iam.IRole], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OrganizationAccountProviderProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class OrganizationOU(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-organization.OrganizationOU",
):
'''The construct to create or update an Organization OU.
This relies on the custom resource provider OrganizationOUProvider.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
parent: typing.Union[builtins.str, "OrganizationOU"],
provider: aws_cdk.custom_resources.Provider,
name: builtins.str,
allow_recreate_on_update: typing.Optional[builtins.bool] = None,
import_on_duplicate: typing.Optional[builtins.bool] = None,
) -> None:
'''
:param scope: -
:param id: -
:param parent: The parent OU id.
:param provider: The provider to use for the custom resource that will create the OU. You can create a provider with the OrganizationOuProvider class
:param name: The name of the OU.
:param allow_recreate_on_update: Whether or not a missing OU should be recreated during an update. If this is false and the OU does not exist an error will be thrown when you try to update it. Default: false
:param import_on_duplicate: Whether or not to import an existing OU if the new OU is a duplicate. If this is false and the OU already exists an error will be thrown. Default: false
'''
props = OUResourceProps(
parent=parent,
provider=provider,
name=name,
allow_recreate_on_update=allow_recreate_on_update,
import_on_duplicate=import_on_duplicate,
)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="resource")
def resource(self) -> aws_cdk.CustomResource:
return typing.cast(aws_cdk.CustomResource, jsii.get(self, "resource"))
class OrganizationOUProvider(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-organization.OrganizationOUProvider",
):
'''The provider for OU custom resources.
This creates a lambda function that handles custom resource requests for creating/updating/deleting OUs.
'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
role: typing.Optional[aws_cdk.aws_iam.IRole] = None,
) -> None:
'''
:param scope: -
:param id: -
:param role: The role the custom resource should use for taking actions on OUs if one is not provided one will be created automatically.
'''
props = OrganizationOUProviderProps(role=role)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="provider")
def provider(self) -> aws_cdk.custom_resources.Provider:
return typing.cast(aws_cdk.custom_resources.Provider, jsii.get(self, "provider"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-organization.OrganizationOUProviderProps",
jsii_struct_bases=[],
name_mapping={"role": "role"},
)
class OrganizationOUProviderProps:
def __init__(self, *, role: typing.Optional[aws_cdk.aws_iam.IRole] = None) -> None:
'''The properties for the OU custom resource provider.
:param role: The role the custom resource should use for taking actions on OUs if one is not provided one will be created automatically.
'''
self._values: typing.Dict[str, typing.Any] = {}
if role is not None:
self._values["role"] = role
@builtins.property
def role(self) -> typing.Optional[aws_cdk.aws_iam.IRole]:
'''The role the custom resource should use for taking actions on OUs if one is not provided one will be created automatically.'''
result = self._values.get("role")
return typing.cast(typing.Optional[aws_cdk.aws_iam.IRole], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OrganizationOUProviderProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"AccountProps",
"AccountResourceProps",
"IPAMAdministratorProps",
"IPAMAdministratorProvider",
"IPAMAdministratorProviderProps",
"IPAMdministrator",
"OUObject",
"OUProps",
"OUResourceProps",
"OrganizationAccount",
"OrganizationAccountProvider",
"OrganizationAccountProviderProps",
"OrganizationOU",
"OrganizationOUProvider",
"OrganizationOUProviderProps",
]
publication.publish() | /renovosolutions.aws_cdk_aws_organization-0.4.69-py3-none-any.whl/organization/__init__.py | 0.66356 | 0.224374 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from typeguard import check_type
from ._jsii import *
import aws_cdk as _aws_cdk_ceddda9d
import aws_cdk.aws_iam as _aws_cdk_aws_iam_ceddda9d
import aws_cdk.aws_sso as _aws_cdk_aws_sso_ceddda9d
import constructs as _constructs_77d1e7e8
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.AssignmentAttributes",
jsii_struct_bases=[],
name_mapping={},
)
class AssignmentAttributes:
def __init__(self) -> None:
'''Attributes for an assignment of which there are none.'''
self._values: typing.Dict[builtins.str, typing.Any] = {}
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AssignmentAttributes(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.AssignmentOptions",
jsii_struct_bases=[],
name_mapping={
"principal": "principal",
"target_id": "targetId",
"target_type": "targetType",
},
)
class AssignmentOptions:
def __init__(
self,
*,
principal: typing.Union["PrincipalProperty", typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional["TargetTypes"] = None,
) -> None:
'''The options for creating an assignment.
:param principal: The principal to assign the permission set to.
:param target_id: The target id the permission set will be assigned to.
:param target_type: The entity type for which the assignment will be created. Default: TargetTypes.AWS_ACCOUNT
'''
if isinstance(principal, dict):
principal = PrincipalProperty(**principal)
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__7ae439f39faaa114233bc0b1a16b1904736952c8fbf1932dc6475eed1f3d817a)
check_type(argname="argument principal", value=principal, expected_type=type_hints["principal"])
check_type(argname="argument target_id", value=target_id, expected_type=type_hints["target_id"])
check_type(argname="argument target_type", value=target_type, expected_type=type_hints["target_type"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"principal": principal,
"target_id": target_id,
}
if target_type is not None:
self._values["target_type"] = target_type
@builtins.property
def principal(self) -> "PrincipalProperty":
'''The principal to assign the permission set to.'''
result = self._values.get("principal")
assert result is not None, "Required property 'principal' is missing"
return typing.cast("PrincipalProperty", result)
@builtins.property
def target_id(self) -> builtins.str:
'''The target id the permission set will be assigned to.'''
result = self._values.get("target_id")
assert result is not None, "Required property 'target_id' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def target_type(self) -> typing.Optional["TargetTypes"]:
'''The entity type for which the assignment will be created.
:default: TargetTypes.AWS_ACCOUNT
'''
result = self._values.get("target_type")
return typing.cast(typing.Optional["TargetTypes"], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AssignmentOptions(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.AssignmentProps",
jsii_struct_bases=[AssignmentOptions],
name_mapping={
"principal": "principal",
"target_id": "targetId",
"target_type": "targetType",
"permission_set": "permissionSet",
},
)
class AssignmentProps(AssignmentOptions):
def __init__(
self,
*,
principal: typing.Union["PrincipalProperty", typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional["TargetTypes"] = None,
permission_set: "IPermissionSet",
) -> None:
'''The properties of a new assignment.
:param principal: The principal to assign the permission set to.
:param target_id: The target id the permission set will be assigned to.
:param target_type: The entity type for which the assignment will be created. Default: TargetTypes.AWS_ACCOUNT
:param permission_set: The permission set to assign to the principal.
'''
if isinstance(principal, dict):
principal = PrincipalProperty(**principal)
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__3677dd84d66e98c8bbf6b4b630ea271a32c589d1f12d4383871d2614207cad7b)
check_type(argname="argument principal", value=principal, expected_type=type_hints["principal"])
check_type(argname="argument target_id", value=target_id, expected_type=type_hints["target_id"])
check_type(argname="argument target_type", value=target_type, expected_type=type_hints["target_type"])
check_type(argname="argument permission_set", value=permission_set, expected_type=type_hints["permission_set"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"principal": principal,
"target_id": target_id,
"permission_set": permission_set,
}
if target_type is not None:
self._values["target_type"] = target_type
@builtins.property
def principal(self) -> "PrincipalProperty":
'''The principal to assign the permission set to.'''
result = self._values.get("principal")
assert result is not None, "Required property 'principal' is missing"
return typing.cast("PrincipalProperty", result)
@builtins.property
def target_id(self) -> builtins.str:
'''The target id the permission set will be assigned to.'''
result = self._values.get("target_id")
assert result is not None, "Required property 'target_id' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def target_type(self) -> typing.Optional["TargetTypes"]:
'''The entity type for which the assignment will be created.
:default: TargetTypes.AWS_ACCOUNT
'''
result = self._values.get("target_type")
return typing.cast(typing.Optional["TargetTypes"], result)
@builtins.property
def permission_set(self) -> "IPermissionSet":
'''The permission set to assign to the principal.'''
result = self._values.get("permission_set")
assert result is not None, "Required property 'permission_set' is missing"
return typing.cast("IPermissionSet", result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AssignmentProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.CustomerManagedPolicyReference",
jsii_struct_bases=[
_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.CustomerManagedPolicyReferenceProperty
],
name_mapping={"name": "name", "path": "path"},
)
class CustomerManagedPolicyReference(
_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.CustomerManagedPolicyReferenceProperty,
):
def __init__(
self,
*,
name: builtins.str,
path: typing.Optional[builtins.str] = None,
) -> None:
'''
:param name: The name of the IAM policy that you have configured in each account where you want to deploy your permission set.
:param path: The path to the IAM policy that you have configured in each account where you want to deploy your permission set. The default is ``/`` . For more information, see `Friendly names and paths <https://docs.aws.amazon.com/IAM/latest/UserGuide/reference_identifiers.html#identifiers-friendly-names>`_ in the *IAM User Guide* .
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__d76e58f544c6bf87eb4ed6045f0b6948c3216099d824187a7cc57942e89eb9fe)
check_type(argname="argument name", value=name, expected_type=type_hints["name"])
check_type(argname="argument path", value=path, expected_type=type_hints["path"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"name": name,
}
if path is not None:
self._values["path"] = path
@builtins.property
def name(self) -> builtins.str:
'''The name of the IAM policy that you have configured in each account where you want to deploy your permission set.
:link: http://docs.aws.amazon.com/AWSCloudFormation/latest/UserGuide/aws-properties-sso-permissionset-customermanagedpolicyreference.html#cfn-sso-permissionset-customermanagedpolicyreference-name
'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def path(self) -> typing.Optional[builtins.str]:
'''The path to the IAM policy that you have configured in each account where you want to deploy your permission set.
The default is ``/`` . For more information, see `Friendly names and paths <https://docs.aws.amazon.com/IAM/latest/UserGuide/reference_identifiers.html#identifiers-friendly-names>`_ in the *IAM User Guide* .
:link: http://docs.aws.amazon.com/AWSCloudFormation/latest/UserGuide/aws-properties-sso-permissionset-customermanagedpolicyreference.html#cfn-sso-permissionset-customermanagedpolicyreference-path
'''
result = self._values.get("path")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "CustomerManagedPolicyReference(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.interface(jsii_type="@renovosolutions/cdk-library-aws-sso.IAssignment")
class IAssignment(_aws_cdk_ceddda9d.IResource, typing_extensions.Protocol):
'''The resource interface for an AWS SSO assignment.
This interface has no attributes because the resulting resource has none.
'''
pass
class _IAssignmentProxy(
jsii.proxy_for(_aws_cdk_ceddda9d.IResource), # type: ignore[misc]
):
'''The resource interface for an AWS SSO assignment.
This interface has no attributes because the resulting resource has none.
'''
__jsii_type__: typing.ClassVar[str] = "@renovosolutions/cdk-library-aws-sso.IAssignment"
pass
# Adding a "__jsii_proxy_class__(): typing.Type" function to the interface
typing.cast(typing.Any, IAssignment).__jsii_proxy_class__ = lambda : _IAssignmentProxy
@jsii.interface(jsii_type="@renovosolutions/cdk-library-aws-sso.IPermissionSet")
class IPermissionSet(_aws_cdk_ceddda9d.IResource, typing_extensions.Protocol):
'''The resource interface for an AWS SSO permission set.'''
@builtins.property
@jsii.member(jsii_name="permissionSetArn")
def permission_set_arn(self) -> builtins.str:
'''The permission set ARN of the permission set.
Such as
``arn:aws:sso:::permissionSet/ins-instanceid/ps-permissionsetid``.
:attribute: true
'''
...
@builtins.property
@jsii.member(jsii_name="ssoInstanceArn")
def sso_instance_arn(self) -> builtins.str:
'''The SSO instance ARN of the permission set.'''
...
@jsii.member(jsii_name="grant")
def grant(
self,
id: builtins.str,
*,
principal: typing.Union["PrincipalProperty", typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional["TargetTypes"] = None,
) -> "Assignment":
'''Grant this permission set to a given principal for a given targetId (AWS account identifier) on a given SSO instance.
:param id: -
:param principal: The principal to assign the permission set to.
:param target_id: The target id the permission set will be assigned to.
:param target_type: The entity type for which the assignment will be created. Default: TargetTypes.AWS_ACCOUNT
'''
...
class _IPermissionSetProxy(
jsii.proxy_for(_aws_cdk_ceddda9d.IResource), # type: ignore[misc]
):
'''The resource interface for an AWS SSO permission set.'''
__jsii_type__: typing.ClassVar[str] = "@renovosolutions/cdk-library-aws-sso.IPermissionSet"
@builtins.property
@jsii.member(jsii_name="permissionSetArn")
def permission_set_arn(self) -> builtins.str:
'''The permission set ARN of the permission set.
Such as
``arn:aws:sso:::permissionSet/ins-instanceid/ps-permissionsetid``.
:attribute: true
'''
return typing.cast(builtins.str, jsii.get(self, "permissionSetArn"))
@builtins.property
@jsii.member(jsii_name="ssoInstanceArn")
def sso_instance_arn(self) -> builtins.str:
'''The SSO instance ARN of the permission set.'''
return typing.cast(builtins.str, jsii.get(self, "ssoInstanceArn"))
@jsii.member(jsii_name="grant")
def grant(
self,
id: builtins.str,
*,
principal: typing.Union["PrincipalProperty", typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional["TargetTypes"] = None,
) -> "Assignment":
'''Grant this permission set to a given principal for a given targetId (AWS account identifier) on a given SSO instance.
:param id: -
:param principal: The principal to assign the permission set to.
:param target_id: The target id the permission set will be assigned to.
:param target_type: The entity type for which the assignment will be created. Default: TargetTypes.AWS_ACCOUNT
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__63d08775b809b0e566cdb84b9c89847ac6e6605d16d5345595d249a8f665bbbe)
check_type(argname="argument id", value=id, expected_type=type_hints["id"])
assignment_options = AssignmentOptions(
principal=principal, target_id=target_id, target_type=target_type
)
return typing.cast("Assignment", jsii.invoke(self, "grant", [id, assignment_options]))
# Adding a "__jsii_proxy_class__(): typing.Type" function to the interface
typing.cast(typing.Any, IPermissionSet).__jsii_proxy_class__ = lambda : _IPermissionSetProxy
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.PermissionBoundary",
jsii_struct_bases=[
_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.PermissionsBoundaryProperty
],
name_mapping={
"customer_managed_policy_reference": "customerManagedPolicyReference",
"managed_policy_arn": "managedPolicyArn",
},
)
class PermissionBoundary(
_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.PermissionsBoundaryProperty,
):
def __init__(
self,
*,
customer_managed_policy_reference: typing.Optional[typing.Union[_aws_cdk_ceddda9d.IResolvable, typing.Union[_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.CustomerManagedPolicyReferenceProperty, typing.Dict[builtins.str, typing.Any]]]] = None,
managed_policy_arn: typing.Optional[builtins.str] = None,
) -> None:
'''
:param customer_managed_policy_reference: Specifies the name and path of a customer managed policy. You must have an IAM policy that matches the name and path in each AWS account where you want to deploy your permission set.
:param managed_policy_arn: The AWS managed policy ARN that you want to attach to a permission set as a permissions boundary.
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__4b19ce1fb3e89b904939ce873e787c40e49146fc1e9defc30a9c9e3df0a22c16)
check_type(argname="argument customer_managed_policy_reference", value=customer_managed_policy_reference, expected_type=type_hints["customer_managed_policy_reference"])
check_type(argname="argument managed_policy_arn", value=managed_policy_arn, expected_type=type_hints["managed_policy_arn"])
self._values: typing.Dict[builtins.str, typing.Any] = {}
if customer_managed_policy_reference is not None:
self._values["customer_managed_policy_reference"] = customer_managed_policy_reference
if managed_policy_arn is not None:
self._values["managed_policy_arn"] = managed_policy_arn
@builtins.property
def customer_managed_policy_reference(
self,
) -> typing.Optional[typing.Union[_aws_cdk_ceddda9d.IResolvable, _aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.CustomerManagedPolicyReferenceProperty]]:
'''Specifies the name and path of a customer managed policy.
You must have an IAM policy that matches the name and path in each AWS account where you want to deploy your permission set.
:link: http://docs.aws.amazon.com/AWSCloudFormation/latest/UserGuide/aws-properties-sso-permissionset-permissionsboundary.html#cfn-sso-permissionset-permissionsboundary-customermanagedpolicyreference
'''
result = self._values.get("customer_managed_policy_reference")
return typing.cast(typing.Optional[typing.Union[_aws_cdk_ceddda9d.IResolvable, _aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.CustomerManagedPolicyReferenceProperty]], result)
@builtins.property
def managed_policy_arn(self) -> typing.Optional[builtins.str]:
'''The AWS managed policy ARN that you want to attach to a permission set as a permissions boundary.
:link: http://docs.aws.amazon.com/AWSCloudFormation/latest/UserGuide/aws-properties-sso-permissionset-permissionsboundary.html#cfn-sso-permissionset-permissionsboundary-managedpolicyarn
'''
result = self._values.get("managed_policy_arn")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "PermissionBoundary(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.implements(IPermissionSet)
class PermissionSet(
_aws_cdk_ceddda9d.Resource,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-sso.PermissionSet",
):
def __init__(
self,
scope: _constructs_77d1e7e8.Construct,
id: builtins.str,
*,
name: builtins.str,
sso_instance_arn: builtins.str,
aws_managed_policies: typing.Optional[typing.Sequence[_aws_cdk_aws_iam_ceddda9d.IManagedPolicy]] = None,
customer_managed_policy_references: typing.Optional[typing.Sequence[typing.Union[CustomerManagedPolicyReference, typing.Dict[builtins.str, typing.Any]]]] = None,
description: typing.Optional[builtins.str] = None,
inline_policy: typing.Optional[_aws_cdk_aws_iam_ceddda9d.PolicyDocument] = None,
permissions_boundary: typing.Optional[typing.Union[PermissionBoundary, typing.Dict[builtins.str, typing.Any]]] = None,
relay_state_type: typing.Optional[builtins.str] = None,
session_duration: typing.Optional[_aws_cdk_ceddda9d.Duration] = None,
) -> None:
'''
:param scope: -
:param id: -
:param name: The name of the permission set.
:param sso_instance_arn: The ARN of the SSO instance under which the operation will be executed.
:param aws_managed_policies: The AWS managed policies to attach to the ``PermissionSet``. Default: - No AWS managed policies
:param customer_managed_policy_references: Specifies the names and paths of a customer managed policy. You must have an IAM policy that matches the name and path in each AWS account where you want to deploy your permission set. Default: - No customer managed policies
:param description: The description of the ``PermissionSet``. Default: - No description
:param inline_policy: The IAM inline policy that is attached to the permission set. Default: - No inline policy
:param permissions_boundary: Specifies the configuration of the AWS managed or customer managed policy that you want to set as a permissions boundary. Specify either customerManagedPolicyReference to use the name and path of a customer managed policy, or managedPolicy to use the ARN of an AWS managed policy. A permissions boundary represents the maximum permissions that any policy can grant your role. For more information, see Permissions boundaries for IAM entities in the AWS Identity and Access Management User Guide. Default: - No permissions boundary
:param relay_state_type: Used to redirect users within the application during the federation authentication process. By default, when a user signs into the AWS access portal, chooses an account, and then chooses the role that AWS creates from the assigned permission set, IAM Identity Center redirects the user’s browser to the AWS Management Console. You can change this behavior by setting the relay state to a different console URL. Setting the relay state enables you to provide the user with quick access to the console that is most appropriate for their role. For example, you can set the relay state to the Amazon EC2 console URL (https://console.aws.amazon.com/ec2/) to redirect the user to that console when they choose the Amazon EC2 administrator role. Default: - No redirection
:param session_duration: The length of time that the application user sessions are valid for.
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__9e1c8a3a6bbb48874bcc41baf50aebc1d5b3c196abd7611f29b515aa6e0683d9)
check_type(argname="argument scope", value=scope, expected_type=type_hints["scope"])
check_type(argname="argument id", value=id, expected_type=type_hints["id"])
props = PermissionSetProps(
name=name,
sso_instance_arn=sso_instance_arn,
aws_managed_policies=aws_managed_policies,
customer_managed_policy_references=customer_managed_policy_references,
description=description,
inline_policy=inline_policy,
permissions_boundary=permissions_boundary,
relay_state_type=relay_state_type,
session_duration=session_duration,
)
jsii.create(self.__class__, self, [scope, id, props])
@jsii.member(jsii_name="fromPermissionSetArn")
@builtins.classmethod
def from_permission_set_arn(
cls,
scope: _constructs_77d1e7e8.Construct,
id: builtins.str,
permission_set_arn: builtins.str,
) -> IPermissionSet:
'''Reference an existing permission set by ARN.
:param scope: -
:param id: -
:param permission_set_arn: -
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__1534cc83982ea2274a4ef806b10a9b22e16bd0a2ac53cb7e2d7d7e7e7f04522a)
check_type(argname="argument scope", value=scope, expected_type=type_hints["scope"])
check_type(argname="argument id", value=id, expected_type=type_hints["id"])
check_type(argname="argument permission_set_arn", value=permission_set_arn, expected_type=type_hints["permission_set_arn"])
return typing.cast(IPermissionSet, jsii.sinvoke(cls, "fromPermissionSetArn", [scope, id, permission_set_arn]))
@jsii.member(jsii_name="grant")
def grant(
self,
id: builtins.str,
*,
principal: typing.Union["PrincipalProperty", typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional["TargetTypes"] = None,
) -> "Assignment":
'''Grant this permission set to a given principal for a given targetId (AWS account identifier) on a given SSO instance.
:param id: -
:param principal: The principal to assign the permission set to.
:param target_id: The target id the permission set will be assigned to.
:param target_type: The entity type for which the assignment will be created. Default: TargetTypes.AWS_ACCOUNT
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__4cdc85e58620129e7f246a5cfde6e7c38745a843ac7234984317b311e146af34)
check_type(argname="argument id", value=id, expected_type=type_hints["id"])
assignment_options = AssignmentOptions(
principal=principal, target_id=target_id, target_type=target_type
)
return typing.cast("Assignment", jsii.invoke(self, "grant", [id, assignment_options]))
@builtins.property
@jsii.member(jsii_name="cfnPermissionSet")
def cfn_permission_set(self) -> _aws_cdk_aws_sso_ceddda9d.CfnPermissionSet:
'''The underlying CfnPermissionSet resource.'''
return typing.cast(_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet, jsii.get(self, "cfnPermissionSet"))
@builtins.property
@jsii.member(jsii_name="permissionSetArn")
def permission_set_arn(self) -> builtins.str:
'''The permission set ARN of the permission set.'''
return typing.cast(builtins.str, jsii.get(self, "permissionSetArn"))
@builtins.property
@jsii.member(jsii_name="ssoInstanceArn")
def sso_instance_arn(self) -> builtins.str:
'''The SSO instance the permission set belongs to.'''
return typing.cast(builtins.str, jsii.get(self, "ssoInstanceArn"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.PermissionSetAttributes",
jsii_struct_bases=[],
name_mapping={
"permission_set_arn": "permissionSetArn",
"sso_instance_arn": "ssoInstanceArn",
},
)
class PermissionSetAttributes:
def __init__(
self,
*,
permission_set_arn: builtins.str,
sso_instance_arn: builtins.str,
) -> None:
'''Attributes for a permission set.
:param permission_set_arn: The permission set ARN of the permission set. Such as ``arn:aws:sso:::permissionSet/ins-instanceid/ps-permissionsetid``.
:param sso_instance_arn: The SSO instance ARN of the permission set.
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__9f41d64f11736d4f2d569f1d67bc6cf42babd45687b8f3b4e4f35e92c7ff0fe3)
check_type(argname="argument permission_set_arn", value=permission_set_arn, expected_type=type_hints["permission_set_arn"])
check_type(argname="argument sso_instance_arn", value=sso_instance_arn, expected_type=type_hints["sso_instance_arn"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"permission_set_arn": permission_set_arn,
"sso_instance_arn": sso_instance_arn,
}
@builtins.property
def permission_set_arn(self) -> builtins.str:
'''The permission set ARN of the permission set.
Such as
``arn:aws:sso:::permissionSet/ins-instanceid/ps-permissionsetid``.
'''
result = self._values.get("permission_set_arn")
assert result is not None, "Required property 'permission_set_arn' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def sso_instance_arn(self) -> builtins.str:
'''The SSO instance ARN of the permission set.'''
result = self._values.get("sso_instance_arn")
assert result is not None, "Required property 'sso_instance_arn' is missing"
return typing.cast(builtins.str, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "PermissionSetAttributes(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.PermissionSetProps",
jsii_struct_bases=[],
name_mapping={
"name": "name",
"sso_instance_arn": "ssoInstanceArn",
"aws_managed_policies": "awsManagedPolicies",
"customer_managed_policy_references": "customerManagedPolicyReferences",
"description": "description",
"inline_policy": "inlinePolicy",
"permissions_boundary": "permissionsBoundary",
"relay_state_type": "relayStateType",
"session_duration": "sessionDuration",
},
)
class PermissionSetProps:
def __init__(
self,
*,
name: builtins.str,
sso_instance_arn: builtins.str,
aws_managed_policies: typing.Optional[typing.Sequence[_aws_cdk_aws_iam_ceddda9d.IManagedPolicy]] = None,
customer_managed_policy_references: typing.Optional[typing.Sequence[typing.Union[CustomerManagedPolicyReference, typing.Dict[builtins.str, typing.Any]]]] = None,
description: typing.Optional[builtins.str] = None,
inline_policy: typing.Optional[_aws_cdk_aws_iam_ceddda9d.PolicyDocument] = None,
permissions_boundary: typing.Optional[typing.Union[PermissionBoundary, typing.Dict[builtins.str, typing.Any]]] = None,
relay_state_type: typing.Optional[builtins.str] = None,
session_duration: typing.Optional[_aws_cdk_ceddda9d.Duration] = None,
) -> None:
'''The properties of a new permission set.
:param name: The name of the permission set.
:param sso_instance_arn: The ARN of the SSO instance under which the operation will be executed.
:param aws_managed_policies: The AWS managed policies to attach to the ``PermissionSet``. Default: - No AWS managed policies
:param customer_managed_policy_references: Specifies the names and paths of a customer managed policy. You must have an IAM policy that matches the name and path in each AWS account where you want to deploy your permission set. Default: - No customer managed policies
:param description: The description of the ``PermissionSet``. Default: - No description
:param inline_policy: The IAM inline policy that is attached to the permission set. Default: - No inline policy
:param permissions_boundary: Specifies the configuration of the AWS managed or customer managed policy that you want to set as a permissions boundary. Specify either customerManagedPolicyReference to use the name and path of a customer managed policy, or managedPolicy to use the ARN of an AWS managed policy. A permissions boundary represents the maximum permissions that any policy can grant your role. For more information, see Permissions boundaries for IAM entities in the AWS Identity and Access Management User Guide. Default: - No permissions boundary
:param relay_state_type: Used to redirect users within the application during the federation authentication process. By default, when a user signs into the AWS access portal, chooses an account, and then chooses the role that AWS creates from the assigned permission set, IAM Identity Center redirects the user’s browser to the AWS Management Console. You can change this behavior by setting the relay state to a different console URL. Setting the relay state enables you to provide the user with quick access to the console that is most appropriate for their role. For example, you can set the relay state to the Amazon EC2 console URL (https://console.aws.amazon.com/ec2/) to redirect the user to that console when they choose the Amazon EC2 administrator role. Default: - No redirection
:param session_duration: The length of time that the application user sessions are valid for.
'''
if isinstance(permissions_boundary, dict):
permissions_boundary = PermissionBoundary(**permissions_boundary)
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__81a500faaedc99c37669b04cc4cb3ded8f1161916dab975452e5e55c1cb669f0)
check_type(argname="argument name", value=name, expected_type=type_hints["name"])
check_type(argname="argument sso_instance_arn", value=sso_instance_arn, expected_type=type_hints["sso_instance_arn"])
check_type(argname="argument aws_managed_policies", value=aws_managed_policies, expected_type=type_hints["aws_managed_policies"])
check_type(argname="argument customer_managed_policy_references", value=customer_managed_policy_references, expected_type=type_hints["customer_managed_policy_references"])
check_type(argname="argument description", value=description, expected_type=type_hints["description"])
check_type(argname="argument inline_policy", value=inline_policy, expected_type=type_hints["inline_policy"])
check_type(argname="argument permissions_boundary", value=permissions_boundary, expected_type=type_hints["permissions_boundary"])
check_type(argname="argument relay_state_type", value=relay_state_type, expected_type=type_hints["relay_state_type"])
check_type(argname="argument session_duration", value=session_duration, expected_type=type_hints["session_duration"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"name": name,
"sso_instance_arn": sso_instance_arn,
}
if aws_managed_policies is not None:
self._values["aws_managed_policies"] = aws_managed_policies
if customer_managed_policy_references is not None:
self._values["customer_managed_policy_references"] = customer_managed_policy_references
if description is not None:
self._values["description"] = description
if inline_policy is not None:
self._values["inline_policy"] = inline_policy
if permissions_boundary is not None:
self._values["permissions_boundary"] = permissions_boundary
if relay_state_type is not None:
self._values["relay_state_type"] = relay_state_type
if session_duration is not None:
self._values["session_duration"] = session_duration
@builtins.property
def name(self) -> builtins.str:
'''The name of the permission set.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def sso_instance_arn(self) -> builtins.str:
'''The ARN of the SSO instance under which the operation will be executed.'''
result = self._values.get("sso_instance_arn")
assert result is not None, "Required property 'sso_instance_arn' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def aws_managed_policies(
self,
) -> typing.Optional[typing.List[_aws_cdk_aws_iam_ceddda9d.IManagedPolicy]]:
'''The AWS managed policies to attach to the ``PermissionSet``.
:default: - No AWS managed policies
'''
result = self._values.get("aws_managed_policies")
return typing.cast(typing.Optional[typing.List[_aws_cdk_aws_iam_ceddda9d.IManagedPolicy]], result)
@builtins.property
def customer_managed_policy_references(
self,
) -> typing.Optional[typing.List[CustomerManagedPolicyReference]]:
'''Specifies the names and paths of a customer managed policy.
You must have an IAM policy that matches the name and path in each
AWS account where you want to deploy your permission set.
:default: - No customer managed policies
'''
result = self._values.get("customer_managed_policy_references")
return typing.cast(typing.Optional[typing.List[CustomerManagedPolicyReference]], result)
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''The description of the ``PermissionSet``.
:default: - No description
'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def inline_policy(
self,
) -> typing.Optional[_aws_cdk_aws_iam_ceddda9d.PolicyDocument]:
'''The IAM inline policy that is attached to the permission set.
:default: - No inline policy
'''
result = self._values.get("inline_policy")
return typing.cast(typing.Optional[_aws_cdk_aws_iam_ceddda9d.PolicyDocument], result)
@builtins.property
def permissions_boundary(self) -> typing.Optional[PermissionBoundary]:
'''Specifies the configuration of the AWS managed or customer managed policy that you want to set as a permissions boundary.
Specify either
customerManagedPolicyReference to use the name and path of a customer
managed policy, or managedPolicy to use the ARN of an AWS managed
policy.
A permissions boundary represents the maximum permissions that any
policy can grant your role. For more information, see Permissions boundaries
for IAM entities in the AWS Identity and Access Management User Guide.
:default: - No permissions boundary
:see: https://docs.aws.amazon.com/IAM/latest/UserGuide/access_policies_boundaries.html
'''
result = self._values.get("permissions_boundary")
return typing.cast(typing.Optional[PermissionBoundary], result)
@builtins.property
def relay_state_type(self) -> typing.Optional[builtins.str]:
'''Used to redirect users within the application during the federation authentication process.
By default, when a user signs into the AWS access portal, chooses an account,
and then chooses the role that AWS creates from the assigned permission set,
IAM Identity Center redirects the user’s browser to the AWS Management Console.
You can change this behavior by setting the relay state to a different console
URL. Setting the relay state enables you to provide the user with quick access
to the console that is most appropriate for their role. For example, you can
set the relay state to the Amazon EC2 console URL (https://console.aws.amazon.com/ec2/)
to redirect the user to that console when they choose the Amazon EC2
administrator role.
:default: - No redirection
:see: https://docs.aws.amazon.com/singlesignon/latest/userguide/howtopermrelaystate.html
'''
result = self._values.get("relay_state_type")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def session_duration(self) -> typing.Optional[_aws_cdk_ceddda9d.Duration]:
'''The length of time that the application user sessions are valid for.'''
result = self._values.get("session_duration")
return typing.cast(typing.Optional[_aws_cdk_ceddda9d.Duration], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "PermissionSetProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-aws-sso.PrincipalProperty",
jsii_struct_bases=[],
name_mapping={"principal_id": "principalId", "principal_type": "principalType"},
)
class PrincipalProperty:
def __init__(
self,
*,
principal_id: builtins.str,
principal_type: "PrincipalTypes",
) -> None:
'''
:param principal_id: The id of the principal.
:param principal_type: The type of the principal.
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__ae5a8b88d8d2941e16b21bcde3962a352b3c0bcdac3ecd7b005ecb7859a601d5)
check_type(argname="argument principal_id", value=principal_id, expected_type=type_hints["principal_id"])
check_type(argname="argument principal_type", value=principal_type, expected_type=type_hints["principal_type"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"principal_id": principal_id,
"principal_type": principal_type,
}
@builtins.property
def principal_id(self) -> builtins.str:
'''The id of the principal.'''
result = self._values.get("principal_id")
assert result is not None, "Required property 'principal_id' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def principal_type(self) -> "PrincipalTypes":
'''The type of the principal.'''
result = self._values.get("principal_type")
assert result is not None, "Required property 'principal_type' is missing"
return typing.cast("PrincipalTypes", result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "PrincipalProperty(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.enum(jsii_type="@renovosolutions/cdk-library-aws-sso.PrincipalTypes")
class PrincipalTypes(enum.Enum):
USER = "USER"
GROUP = "GROUP"
@jsii.enum(jsii_type="@renovosolutions/cdk-library-aws-sso.TargetTypes")
class TargetTypes(enum.Enum):
AWS_ACCOUNT = "AWS_ACCOUNT"
@jsii.implements(IAssignment)
class Assignment(
_aws_cdk_ceddda9d.Resource,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-aws-sso.Assignment",
):
'''The assignment construct.
Has no import method because there is no attributes to import.
'''
def __init__(
self,
scope: _constructs_77d1e7e8.Construct,
id: builtins.str,
*,
permission_set: IPermissionSet,
principal: typing.Union[PrincipalProperty, typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional[TargetTypes] = None,
) -> None:
'''
:param scope: -
:param id: -
:param permission_set: The permission set to assign to the principal.
:param principal: The principal to assign the permission set to.
:param target_id: The target id the permission set will be assigned to.
:param target_type: The entity type for which the assignment will be created. Default: TargetTypes.AWS_ACCOUNT
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__5eca7b2cf47acacd3579840411d5aee15f9a9bf2f86aefe69c49c627750efbf7)
check_type(argname="argument scope", value=scope, expected_type=type_hints["scope"])
check_type(argname="argument id", value=id, expected_type=type_hints["id"])
props = AssignmentProps(
permission_set=permission_set,
principal=principal,
target_id=target_id,
target_type=target_type,
)
jsii.create(self.__class__, self, [scope, id, props])
__all__ = [
"Assignment",
"AssignmentAttributes",
"AssignmentOptions",
"AssignmentProps",
"CustomerManagedPolicyReference",
"IAssignment",
"IPermissionSet",
"PermissionBoundary",
"PermissionSet",
"PermissionSetAttributes",
"PermissionSetProps",
"PrincipalProperty",
"PrincipalTypes",
"TargetTypes",
]
publication.publish()
def _typecheckingstub__7ae439f39faaa114233bc0b1a16b1904736952c8fbf1932dc6475eed1f3d817a(
*,
principal: typing.Union[PrincipalProperty, typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional[TargetTypes] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__3677dd84d66e98c8bbf6b4b630ea271a32c589d1f12d4383871d2614207cad7b(
*,
principal: typing.Union[PrincipalProperty, typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional[TargetTypes] = None,
permission_set: IPermissionSet,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__d76e58f544c6bf87eb4ed6045f0b6948c3216099d824187a7cc57942e89eb9fe(
*,
name: builtins.str,
path: typing.Optional[builtins.str] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__63d08775b809b0e566cdb84b9c89847ac6e6605d16d5345595d249a8f665bbbe(
id: builtins.str,
*,
principal: typing.Union[PrincipalProperty, typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional[TargetTypes] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__4b19ce1fb3e89b904939ce873e787c40e49146fc1e9defc30a9c9e3df0a22c16(
*,
customer_managed_policy_reference: typing.Optional[typing.Union[_aws_cdk_ceddda9d.IResolvable, typing.Union[_aws_cdk_aws_sso_ceddda9d.CfnPermissionSet.CustomerManagedPolicyReferenceProperty, typing.Dict[builtins.str, typing.Any]]]] = None,
managed_policy_arn: typing.Optional[builtins.str] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__9e1c8a3a6bbb48874bcc41baf50aebc1d5b3c196abd7611f29b515aa6e0683d9(
scope: _constructs_77d1e7e8.Construct,
id: builtins.str,
*,
name: builtins.str,
sso_instance_arn: builtins.str,
aws_managed_policies: typing.Optional[typing.Sequence[_aws_cdk_aws_iam_ceddda9d.IManagedPolicy]] = None,
customer_managed_policy_references: typing.Optional[typing.Sequence[typing.Union[CustomerManagedPolicyReference, typing.Dict[builtins.str, typing.Any]]]] = None,
description: typing.Optional[builtins.str] = None,
inline_policy: typing.Optional[_aws_cdk_aws_iam_ceddda9d.PolicyDocument] = None,
permissions_boundary: typing.Optional[typing.Union[PermissionBoundary, typing.Dict[builtins.str, typing.Any]]] = None,
relay_state_type: typing.Optional[builtins.str] = None,
session_duration: typing.Optional[_aws_cdk_ceddda9d.Duration] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__1534cc83982ea2274a4ef806b10a9b22e16bd0a2ac53cb7e2d7d7e7e7f04522a(
scope: _constructs_77d1e7e8.Construct,
id: builtins.str,
permission_set_arn: builtins.str,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__4cdc85e58620129e7f246a5cfde6e7c38745a843ac7234984317b311e146af34(
id: builtins.str,
*,
principal: typing.Union[PrincipalProperty, typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional[TargetTypes] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__9f41d64f11736d4f2d569f1d67bc6cf42babd45687b8f3b4e4f35e92c7ff0fe3(
*,
permission_set_arn: builtins.str,
sso_instance_arn: builtins.str,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__81a500faaedc99c37669b04cc4cb3ded8f1161916dab975452e5e55c1cb669f0(
*,
name: builtins.str,
sso_instance_arn: builtins.str,
aws_managed_policies: typing.Optional[typing.Sequence[_aws_cdk_aws_iam_ceddda9d.IManagedPolicy]] = None,
customer_managed_policy_references: typing.Optional[typing.Sequence[typing.Union[CustomerManagedPolicyReference, typing.Dict[builtins.str, typing.Any]]]] = None,
description: typing.Optional[builtins.str] = None,
inline_policy: typing.Optional[_aws_cdk_aws_iam_ceddda9d.PolicyDocument] = None,
permissions_boundary: typing.Optional[typing.Union[PermissionBoundary, typing.Dict[builtins.str, typing.Any]]] = None,
relay_state_type: typing.Optional[builtins.str] = None,
session_duration: typing.Optional[_aws_cdk_ceddda9d.Duration] = None,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__ae5a8b88d8d2941e16b21bcde3962a352b3c0bcdac3ecd7b005ecb7859a601d5(
*,
principal_id: builtins.str,
principal_type: PrincipalTypes,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__5eca7b2cf47acacd3579840411d5aee15f9a9bf2f86aefe69c49c627750efbf7(
scope: _constructs_77d1e7e8.Construct,
id: builtins.str,
*,
permission_set: IPermissionSet,
principal: typing.Union[PrincipalProperty, typing.Dict[builtins.str, typing.Any]],
target_id: builtins.str,
target_type: typing.Optional[TargetTypes] = None,
) -> None:
"""Type checking stubs"""
pass | /renovosolutions.aws-cdk-aws-sso-0.1.149.tar.gz/renovosolutions.aws-cdk-aws-sso-0.1.149/src/sso/__init__.py | 0.661486 | 0.335324 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk
import aws_cdk.aws_events
import aws_cdk.aws_lambda
import aws_cdk.aws_s3
import aws_cdk.aws_sns
import constructs
class Certbot(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-certbot.Certbot",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
hosted_zone_names: typing.Sequence[builtins.str],
letsencrypt_domains: builtins.str,
letsencrypt_email: builtins.str,
bucket: typing.Optional[aws_cdk.aws_s3.Bucket] = None,
enable_insights: typing.Optional[builtins.bool] = None,
enable_object_deletion: typing.Optional[builtins.bool] = None,
function_description: typing.Optional[builtins.str] = None,
function_name: typing.Optional[builtins.str] = None,
insights_arn: typing.Optional[builtins.str] = None,
layers: typing.Optional[typing.Sequence[aws_cdk.aws_lambda.ILayerVersion]] = None,
object_prefix: typing.Optional[builtins.str] = None,
preferred_chain: typing.Optional[builtins.str] = None,
re_issue_days: typing.Optional[jsii.Number] = None,
removal_policy: typing.Optional[aws_cdk.RemovalPolicy] = None,
run_on_deploy: typing.Optional[builtins.bool] = None,
run_on_deploy_wait_minutes: typing.Optional[jsii.Number] = None,
schedule: typing.Optional[aws_cdk.aws_events.Schedule] = None,
sns_topic: typing.Optional[aws_cdk.aws_sns.Topic] = None,
timeout: typing.Optional[aws_cdk.Duration] = None,
) -> None:
'''
:param scope: -
:param id: -
:param hosted_zone_names: Hosted zone names that will be required for DNS verification with certbot.
:param letsencrypt_domains: The comma delimited list of domains for which the Let's Encrypt certificate will be valid. Primary domain should be first.
:param letsencrypt_email: The email to associate with the Let's Encrypt certificate request.
:param bucket: The S3 bucket to place the resulting certificates in. If no bucket is given one will be created automatically.
:param enable_insights: Whether or not to enable Lambda Insights. Default: false
:param enable_object_deletion: Whether or not to enable automatic object deletion if the provided bucket is deleted. Has no effect if a bucket is given as a property Default: false
:param function_description: The description for the resulting Lambda function.
:param function_name: The name of the resulting Lambda function.
:param insights_arn: Insights layer ARN for your region. Defaults to layer for US-EAST-1
:param layers: Any additional Lambda layers to use with the created function. For example Lambda Extensions
:param object_prefix: The prefix to apply to the final S3 key name for the certificates. Default is no prefix.
:param preferred_chain: Set the preferred certificate chain. Default: 'None'
:param re_issue_days: The numbers of days left until the prior cert expires before issuing a new one. Default: 30
:param removal_policy: The removal policy for the S3 bucket that is automatically created. Has no effect if a bucket is given as a property Default: RemovalPolicy.RETAIN
:param run_on_deploy: Whether or not to schedule a trigger to run the function after each deployment. Default: true
:param run_on_deploy_wait_minutes: How many minutes to wait before running the post deployment Lambda trigger. Default: 10
:param schedule: The schedule for the certificate check trigger. Default: events.Schedule.cron({ minute: '0', hour: '0', weekDay: '1' })
:param sns_topic: The SNS topic to notify when a new cert is issued. If no topic is given one will be created automatically.
:param timeout: The timeout duration for Lambda function. Default: Duraction.seconds(180)
'''
props = CertbotProps(
hosted_zone_names=hosted_zone_names,
letsencrypt_domains=letsencrypt_domains,
letsencrypt_email=letsencrypt_email,
bucket=bucket,
enable_insights=enable_insights,
enable_object_deletion=enable_object_deletion,
function_description=function_description,
function_name=function_name,
insights_arn=insights_arn,
layers=layers,
object_prefix=object_prefix,
preferred_chain=preferred_chain,
re_issue_days=re_issue_days,
removal_policy=removal_policy,
run_on_deploy=run_on_deploy,
run_on_deploy_wait_minutes=run_on_deploy_wait_minutes,
schedule=schedule,
sns_topic=sns_topic,
timeout=timeout,
)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="handler")
def handler(self) -> aws_cdk.aws_lambda.Function:
return typing.cast(aws_cdk.aws_lambda.Function, jsii.get(self, "handler"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-certbot.CertbotProps",
jsii_struct_bases=[],
name_mapping={
"hosted_zone_names": "hostedZoneNames",
"letsencrypt_domains": "letsencryptDomains",
"letsencrypt_email": "letsencryptEmail",
"bucket": "bucket",
"enable_insights": "enableInsights",
"enable_object_deletion": "enableObjectDeletion",
"function_description": "functionDescription",
"function_name": "functionName",
"insights_arn": "insightsARN",
"layers": "layers",
"object_prefix": "objectPrefix",
"preferred_chain": "preferredChain",
"re_issue_days": "reIssueDays",
"removal_policy": "removalPolicy",
"run_on_deploy": "runOnDeploy",
"run_on_deploy_wait_minutes": "runOnDeployWaitMinutes",
"schedule": "schedule",
"sns_topic": "snsTopic",
"timeout": "timeout",
},
)
class CertbotProps:
def __init__(
self,
*,
hosted_zone_names: typing.Sequence[builtins.str],
letsencrypt_domains: builtins.str,
letsencrypt_email: builtins.str,
bucket: typing.Optional[aws_cdk.aws_s3.Bucket] = None,
enable_insights: typing.Optional[builtins.bool] = None,
enable_object_deletion: typing.Optional[builtins.bool] = None,
function_description: typing.Optional[builtins.str] = None,
function_name: typing.Optional[builtins.str] = None,
insights_arn: typing.Optional[builtins.str] = None,
layers: typing.Optional[typing.Sequence[aws_cdk.aws_lambda.ILayerVersion]] = None,
object_prefix: typing.Optional[builtins.str] = None,
preferred_chain: typing.Optional[builtins.str] = None,
re_issue_days: typing.Optional[jsii.Number] = None,
removal_policy: typing.Optional[aws_cdk.RemovalPolicy] = None,
run_on_deploy: typing.Optional[builtins.bool] = None,
run_on_deploy_wait_minutes: typing.Optional[jsii.Number] = None,
schedule: typing.Optional[aws_cdk.aws_events.Schedule] = None,
sns_topic: typing.Optional[aws_cdk.aws_sns.Topic] = None,
timeout: typing.Optional[aws_cdk.Duration] = None,
) -> None:
'''
:param hosted_zone_names: Hosted zone names that will be required for DNS verification with certbot.
:param letsencrypt_domains: The comma delimited list of domains for which the Let's Encrypt certificate will be valid. Primary domain should be first.
:param letsencrypt_email: The email to associate with the Let's Encrypt certificate request.
:param bucket: The S3 bucket to place the resulting certificates in. If no bucket is given one will be created automatically.
:param enable_insights: Whether or not to enable Lambda Insights. Default: false
:param enable_object_deletion: Whether or not to enable automatic object deletion if the provided bucket is deleted. Has no effect if a bucket is given as a property Default: false
:param function_description: The description for the resulting Lambda function.
:param function_name: The name of the resulting Lambda function.
:param insights_arn: Insights layer ARN for your region. Defaults to layer for US-EAST-1
:param layers: Any additional Lambda layers to use with the created function. For example Lambda Extensions
:param object_prefix: The prefix to apply to the final S3 key name for the certificates. Default is no prefix.
:param preferred_chain: Set the preferred certificate chain. Default: 'None'
:param re_issue_days: The numbers of days left until the prior cert expires before issuing a new one. Default: 30
:param removal_policy: The removal policy for the S3 bucket that is automatically created. Has no effect if a bucket is given as a property Default: RemovalPolicy.RETAIN
:param run_on_deploy: Whether or not to schedule a trigger to run the function after each deployment. Default: true
:param run_on_deploy_wait_minutes: How many minutes to wait before running the post deployment Lambda trigger. Default: 10
:param schedule: The schedule for the certificate check trigger. Default: events.Schedule.cron({ minute: '0', hour: '0', weekDay: '1' })
:param sns_topic: The SNS topic to notify when a new cert is issued. If no topic is given one will be created automatically.
:param timeout: The timeout duration for Lambda function. Default: Duraction.seconds(180)
'''
self._values: typing.Dict[str, typing.Any] = {
"hosted_zone_names": hosted_zone_names,
"letsencrypt_domains": letsencrypt_domains,
"letsencrypt_email": letsencrypt_email,
}
if bucket is not None:
self._values["bucket"] = bucket
if enable_insights is not None:
self._values["enable_insights"] = enable_insights
if enable_object_deletion is not None:
self._values["enable_object_deletion"] = enable_object_deletion
if function_description is not None:
self._values["function_description"] = function_description
if function_name is not None:
self._values["function_name"] = function_name
if insights_arn is not None:
self._values["insights_arn"] = insights_arn
if layers is not None:
self._values["layers"] = layers
if object_prefix is not None:
self._values["object_prefix"] = object_prefix
if preferred_chain is not None:
self._values["preferred_chain"] = preferred_chain
if re_issue_days is not None:
self._values["re_issue_days"] = re_issue_days
if removal_policy is not None:
self._values["removal_policy"] = removal_policy
if run_on_deploy is not None:
self._values["run_on_deploy"] = run_on_deploy
if run_on_deploy_wait_minutes is not None:
self._values["run_on_deploy_wait_minutes"] = run_on_deploy_wait_minutes
if schedule is not None:
self._values["schedule"] = schedule
if sns_topic is not None:
self._values["sns_topic"] = sns_topic
if timeout is not None:
self._values["timeout"] = timeout
@builtins.property
def hosted_zone_names(self) -> typing.List[builtins.str]:
'''Hosted zone names that will be required for DNS verification with certbot.'''
result = self._values.get("hosted_zone_names")
assert result is not None, "Required property 'hosted_zone_names' is missing"
return typing.cast(typing.List[builtins.str], result)
@builtins.property
def letsencrypt_domains(self) -> builtins.str:
'''The comma delimited list of domains for which the Let's Encrypt certificate will be valid.
Primary domain should be first.
'''
result = self._values.get("letsencrypt_domains")
assert result is not None, "Required property 'letsencrypt_domains' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def letsencrypt_email(self) -> builtins.str:
'''The email to associate with the Let's Encrypt certificate request.'''
result = self._values.get("letsencrypt_email")
assert result is not None, "Required property 'letsencrypt_email' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def bucket(self) -> typing.Optional[aws_cdk.aws_s3.Bucket]:
'''The S3 bucket to place the resulting certificates in.
If no bucket is given one will be created automatically.
'''
result = self._values.get("bucket")
return typing.cast(typing.Optional[aws_cdk.aws_s3.Bucket], result)
@builtins.property
def enable_insights(self) -> typing.Optional[builtins.bool]:
'''Whether or not to enable Lambda Insights.
:default: false
'''
result = self._values.get("enable_insights")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def enable_object_deletion(self) -> typing.Optional[builtins.bool]:
'''Whether or not to enable automatic object deletion if the provided bucket is deleted.
Has no effect if a bucket is given as a property
:default: false
'''
result = self._values.get("enable_object_deletion")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def function_description(self) -> typing.Optional[builtins.str]:
'''The description for the resulting Lambda function.'''
result = self._values.get("function_description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def function_name(self) -> typing.Optional[builtins.str]:
'''The name of the resulting Lambda function.'''
result = self._values.get("function_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def insights_arn(self) -> typing.Optional[builtins.str]:
'''Insights layer ARN for your region.
Defaults to layer for US-EAST-1
'''
result = self._values.get("insights_arn")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def layers(self) -> typing.Optional[typing.List[aws_cdk.aws_lambda.ILayerVersion]]:
'''Any additional Lambda layers to use with the created function.
For example Lambda Extensions
'''
result = self._values.get("layers")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_lambda.ILayerVersion]], result)
@builtins.property
def object_prefix(self) -> typing.Optional[builtins.str]:
'''The prefix to apply to the final S3 key name for the certificates.
Default is no prefix.
'''
result = self._values.get("object_prefix")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def preferred_chain(self) -> typing.Optional[builtins.str]:
'''Set the preferred certificate chain.
:default: 'None'
'''
result = self._values.get("preferred_chain")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def re_issue_days(self) -> typing.Optional[jsii.Number]:
'''The numbers of days left until the prior cert expires before issuing a new one.
:default: 30
'''
result = self._values.get("re_issue_days")
return typing.cast(typing.Optional[jsii.Number], result)
@builtins.property
def removal_policy(self) -> typing.Optional[aws_cdk.RemovalPolicy]:
'''The removal policy for the S3 bucket that is automatically created.
Has no effect if a bucket is given as a property
:default: RemovalPolicy.RETAIN
'''
result = self._values.get("removal_policy")
return typing.cast(typing.Optional[aws_cdk.RemovalPolicy], result)
@builtins.property
def run_on_deploy(self) -> typing.Optional[builtins.bool]:
'''Whether or not to schedule a trigger to run the function after each deployment.
:default: true
'''
result = self._values.get("run_on_deploy")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def run_on_deploy_wait_minutes(self) -> typing.Optional[jsii.Number]:
'''How many minutes to wait before running the post deployment Lambda trigger.
:default: 10
'''
result = self._values.get("run_on_deploy_wait_minutes")
return typing.cast(typing.Optional[jsii.Number], result)
@builtins.property
def schedule(self) -> typing.Optional[aws_cdk.aws_events.Schedule]:
'''The schedule for the certificate check trigger.
:default: events.Schedule.cron({ minute: '0', hour: '0', weekDay: '1' })
'''
result = self._values.get("schedule")
return typing.cast(typing.Optional[aws_cdk.aws_events.Schedule], result)
@builtins.property
def sns_topic(self) -> typing.Optional[aws_cdk.aws_sns.Topic]:
'''The SNS topic to notify when a new cert is issued.
If no topic is given one will be created automatically.
'''
result = self._values.get("sns_topic")
return typing.cast(typing.Optional[aws_cdk.aws_sns.Topic], result)
@builtins.property
def timeout(self) -> typing.Optional[aws_cdk.Duration]:
'''The timeout duration for Lambda function.
:default: Duraction.seconds(180)
'''
result = self._values.get("timeout")
return typing.cast(typing.Optional[aws_cdk.Duration], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "CertbotProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"Certbot",
"CertbotProps",
]
publication.publish() | /renovosolutions.aws-cdk-certbot-2.2.162.tar.gz/renovosolutions.aws-cdk-certbot-2.2.162/src/certbot/__init__.py | 0.675229 | 0.171824 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk.aws_events
import constructs
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.BaseRuleProps",
jsii_struct_bases=[],
name_mapping={
"description": "description",
"enabled": "enabled",
"event_bus": "eventBus",
"event_state": "eventState",
"rule_name": "ruleName",
"targets": "targets",
},
)
class BaseRuleProps:
def __init__(
self,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional["EventStates"] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
self._values: typing.Dict[str, typing.Any] = {}
if description is not None:
self._values["description"] = description
if enabled is not None:
self._values["enabled"] = enabled
if event_bus is not None:
self._values["event_bus"] = event_bus
if event_state is not None:
self._values["event_state"] = event_state
if rule_name is not None:
self._values["rule_name"] = rule_name
if targets is not None:
self._values["targets"] = targets
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''A description of the rule's purpose.
:default: - A rule for new account creation in Organizations
'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def enabled(self) -> typing.Optional[builtins.bool]:
'''Indicates whether the rule is enabled.
:default: true
'''
result = self._values.get("enabled")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def event_bus(self) -> typing.Optional[aws_cdk.aws_events.IEventBus]:
'''The event bus to associate with this rule.
:default: - The default event bus.
'''
result = self._values.get("event_bus")
return typing.cast(typing.Optional[aws_cdk.aws_events.IEventBus], result)
@builtins.property
def event_state(self) -> typing.Optional["EventStates"]:
'''Which event state should this rule trigger for.
:default: - EventStates.SUCCEEDED
'''
result = self._values.get("event_state")
return typing.cast(typing.Optional["EventStates"], result)
@builtins.property
def rule_name(self) -> typing.Optional[builtins.str]:
'''A name for the rule.
:default:
- AWS CloudFormation generates a unique physical ID and uses that ID
for the rule name. For more information, see Name Type.
'''
result = self._values.get("rule_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def targets(self) -> typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]]:
'''Targets to invoke when this rule matches an event.
:default: - No targets.
'''
result = self._values.get("targets")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "BaseRuleProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class CreatedAccountByOrganizationsRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.CreatedAccountByOrganizationsRule",
):
'''A rule for matching events from CloudTrail where Organizations created a new account.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional["EventStates"] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = BaseRuleProps(
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class CreatedAccountRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.CreatedAccountRule",
):
'''A rule for matching events from CloudTrail where Control Tower created a new account.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional["EventStates"] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = OuRuleProps(
ou_id=ou_id,
ou_name=ou_name,
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class DeregisteredOrganizationalUnitRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.DeregisteredOrganizationalUnitRule",
):
'''A rule for matching events from CloudTrail where Control Tower deregistered an Organizational Unit.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional["EventStates"] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = OuRuleProps(
ou_id=ou_id,
ou_name=ou_name,
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class DisabledGuardrailRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.DisabledGuardrailRule",
):
'''A rule for matching events from CloudTrail where a guard rail was disabled via Control Tower for an Organizational Unit.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
guardrail_behavior: typing.Optional["GuardrailBehaviors"] = None,
guardrail_id: typing.Optional[builtins.str] = None,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional["EventStates"] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param guardrail_behavior: The guardrail behavior to match.
:param guardrail_id: The guardrail ID to match.
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = GuardrailRuleProps(
guardrail_behavior=guardrail_behavior,
guardrail_id=guardrail_id,
ou_id=ou_id,
ou_name=ou_name,
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class EnabledGuardrailRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.EnabledGuardrailRule",
):
'''A rule for matching events from CloudTrail where a guardrail was enabled via Control Tower for an Organizational Unit.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
guardrail_behavior: typing.Optional["GuardrailBehaviors"] = None,
guardrail_id: typing.Optional[builtins.str] = None,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional["EventStates"] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param guardrail_behavior: The guardrail behavior to match.
:param guardrail_id: The guardrail ID to match.
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = GuardrailRuleProps(
guardrail_behavior=guardrail_behavior,
guardrail_id=guardrail_id,
ou_id=ou_id,
ou_name=ou_name,
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
@jsii.enum(
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.EventStates"
)
class EventStates(enum.Enum):
SUCCEEDED = "SUCCEEDED"
FAILED = "FAILED"
@jsii.enum(
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.GuardrailBehaviors"
)
class GuardrailBehaviors(enum.Enum):
DETECTIVE = "DETECTIVE"
PREVENTATIVE = "PREVENTATIVE"
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.GuardrailRuleProps",
jsii_struct_bases=[BaseRuleProps],
name_mapping={
"description": "description",
"enabled": "enabled",
"event_bus": "eventBus",
"event_state": "eventState",
"rule_name": "ruleName",
"targets": "targets",
"guardrail_behavior": "guardrailBehavior",
"guardrail_id": "guardrailId",
"ou_id": "ouId",
"ou_name": "ouName",
},
)
class GuardrailRuleProps(BaseRuleProps):
def __init__(
self,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
guardrail_behavior: typing.Optional[GuardrailBehaviors] = None,
guardrail_id: typing.Optional[builtins.str] = None,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
) -> None:
'''
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
:param guardrail_behavior: The guardrail behavior to match.
:param guardrail_id: The guardrail ID to match.
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
'''
self._values: typing.Dict[str, typing.Any] = {}
if description is not None:
self._values["description"] = description
if enabled is not None:
self._values["enabled"] = enabled
if event_bus is not None:
self._values["event_bus"] = event_bus
if event_state is not None:
self._values["event_state"] = event_state
if rule_name is not None:
self._values["rule_name"] = rule_name
if targets is not None:
self._values["targets"] = targets
if guardrail_behavior is not None:
self._values["guardrail_behavior"] = guardrail_behavior
if guardrail_id is not None:
self._values["guardrail_id"] = guardrail_id
if ou_id is not None:
self._values["ou_id"] = ou_id
if ou_name is not None:
self._values["ou_name"] = ou_name
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''A description of the rule's purpose.
:default: - A rule for new account creation in Organizations
'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def enabled(self) -> typing.Optional[builtins.bool]:
'''Indicates whether the rule is enabled.
:default: true
'''
result = self._values.get("enabled")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def event_bus(self) -> typing.Optional[aws_cdk.aws_events.IEventBus]:
'''The event bus to associate with this rule.
:default: - The default event bus.
'''
result = self._values.get("event_bus")
return typing.cast(typing.Optional[aws_cdk.aws_events.IEventBus], result)
@builtins.property
def event_state(self) -> typing.Optional[EventStates]:
'''Which event state should this rule trigger for.
:default: - EventStates.SUCCEEDED
'''
result = self._values.get("event_state")
return typing.cast(typing.Optional[EventStates], result)
@builtins.property
def rule_name(self) -> typing.Optional[builtins.str]:
'''A name for the rule.
:default:
- AWS CloudFormation generates a unique physical ID and uses that ID
for the rule name. For more information, see Name Type.
'''
result = self._values.get("rule_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def targets(self) -> typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]]:
'''Targets to invoke when this rule matches an event.
:default: - No targets.
'''
result = self._values.get("targets")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]], result)
@builtins.property
def guardrail_behavior(self) -> typing.Optional[GuardrailBehaviors]:
'''The guardrail behavior to match.'''
result = self._values.get("guardrail_behavior")
return typing.cast(typing.Optional[GuardrailBehaviors], result)
@builtins.property
def guardrail_id(self) -> typing.Optional[builtins.str]:
'''The guardrail ID to match.'''
result = self._values.get("guardrail_id")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def ou_id(self) -> typing.Optional[builtins.str]:
'''The OU ID to match.'''
result = self._values.get("ou_id")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def ou_name(self) -> typing.Optional[builtins.str]:
'''The OU name to match.'''
result = self._values.get("ou_name")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "GuardrailRuleProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.OuRuleProps",
jsii_struct_bases=[BaseRuleProps],
name_mapping={
"description": "description",
"enabled": "enabled",
"event_bus": "eventBus",
"event_state": "eventState",
"rule_name": "ruleName",
"targets": "targets",
"ou_id": "ouId",
"ou_name": "ouName",
},
)
class OuRuleProps(BaseRuleProps):
def __init__(
self,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
) -> None:
'''
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
'''
self._values: typing.Dict[str, typing.Any] = {}
if description is not None:
self._values["description"] = description
if enabled is not None:
self._values["enabled"] = enabled
if event_bus is not None:
self._values["event_bus"] = event_bus
if event_state is not None:
self._values["event_state"] = event_state
if rule_name is not None:
self._values["rule_name"] = rule_name
if targets is not None:
self._values["targets"] = targets
if ou_id is not None:
self._values["ou_id"] = ou_id
if ou_name is not None:
self._values["ou_name"] = ou_name
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''A description of the rule's purpose.
:default: - A rule for new account creation in Organizations
'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def enabled(self) -> typing.Optional[builtins.bool]:
'''Indicates whether the rule is enabled.
:default: true
'''
result = self._values.get("enabled")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def event_bus(self) -> typing.Optional[aws_cdk.aws_events.IEventBus]:
'''The event bus to associate with this rule.
:default: - The default event bus.
'''
result = self._values.get("event_bus")
return typing.cast(typing.Optional[aws_cdk.aws_events.IEventBus], result)
@builtins.property
def event_state(self) -> typing.Optional[EventStates]:
'''Which event state should this rule trigger for.
:default: - EventStates.SUCCEEDED
'''
result = self._values.get("event_state")
return typing.cast(typing.Optional[EventStates], result)
@builtins.property
def rule_name(self) -> typing.Optional[builtins.str]:
'''A name for the rule.
:default:
- AWS CloudFormation generates a unique physical ID and uses that ID
for the rule name. For more information, see Name Type.
'''
result = self._values.get("rule_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def targets(self) -> typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]]:
'''Targets to invoke when this rule matches an event.
:default: - No targets.
'''
result = self._values.get("targets")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]], result)
@builtins.property
def ou_id(self) -> typing.Optional[builtins.str]:
'''The OU ID to match.'''
result = self._values.get("ou_id")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def ou_name(self) -> typing.Optional[builtins.str]:
'''The OU name to match.'''
result = self._values.get("ou_name")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OuRuleProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class RegisteredOrganizationalUnitRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.RegisteredOrganizationalUnitRule",
):
'''A rule for matching events from CloudTrail where Control Tower registered a new Organizational Unit.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = BaseRuleProps(
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class SetupLandingZoneRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.SetupLandingZoneRule",
):
'''A rule for matching events from CloudTrail where a landing zone was setup via Control Tower.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = BaseRuleProps(
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class UpdatedLandingZoneRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.UpdatedLandingZoneRule",
):
'''A rule for matching events from CloudTrail where a landing zone was updated via Control Tower.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = BaseRuleProps(
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
class UpdatedManagedAccountRule(
aws_cdk.aws_events.Rule,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.UpdatedManagedAccountRule",
):
'''A rule for matching events from CloudTrail where Control Tower updated a managed account.'''
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
account_id: typing.Optional[builtins.str] = None,
account_name: typing.Optional[builtins.str] = None,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
) -> None:
'''
:param scope: -
:param id: -
:param account_id: The account ID to match.
:param account_name: The account name to match.
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
'''
props = AccountRuleProps(
account_id=account_id,
account_name=account_name,
ou_id=ou_id,
ou_name=ou_name,
description=description,
enabled=enabled,
event_bus=event_bus,
event_state=event_state,
rule_name=rule_name,
targets=targets,
)
jsii.create(self.__class__, self, [scope, id, props])
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-control-tower-lifecycle-events.AccountRuleProps",
jsii_struct_bases=[BaseRuleProps],
name_mapping={
"description": "description",
"enabled": "enabled",
"event_bus": "eventBus",
"event_state": "eventState",
"rule_name": "ruleName",
"targets": "targets",
"account_id": "accountId",
"account_name": "accountName",
"ou_id": "ouId",
"ou_name": "ouName",
},
)
class AccountRuleProps(BaseRuleProps):
def __init__(
self,
*,
description: typing.Optional[builtins.str] = None,
enabled: typing.Optional[builtins.bool] = None,
event_bus: typing.Optional[aws_cdk.aws_events.IEventBus] = None,
event_state: typing.Optional[EventStates] = None,
rule_name: typing.Optional[builtins.str] = None,
targets: typing.Optional[typing.Sequence[aws_cdk.aws_events.IRuleTarget]] = None,
account_id: typing.Optional[builtins.str] = None,
account_name: typing.Optional[builtins.str] = None,
ou_id: typing.Optional[builtins.str] = None,
ou_name: typing.Optional[builtins.str] = None,
) -> None:
'''
:param description: A description of the rule's purpose. Default: - A rule for new account creation in Organizations
:param enabled: Indicates whether the rule is enabled. Default: true
:param event_bus: The event bus to associate with this rule. Default: - The default event bus.
:param event_state: Which event state should this rule trigger for. Default: - EventStates.SUCCEEDED
:param rule_name: A name for the rule. Default: - AWS CloudFormation generates a unique physical ID and uses that ID for the rule name. For more information, see Name Type.
:param targets: Targets to invoke when this rule matches an event. Default: - No targets.
:param account_id: The account ID to match.
:param account_name: The account name to match.
:param ou_id: The OU ID to match.
:param ou_name: The OU name to match.
'''
self._values: typing.Dict[str, typing.Any] = {}
if description is not None:
self._values["description"] = description
if enabled is not None:
self._values["enabled"] = enabled
if event_bus is not None:
self._values["event_bus"] = event_bus
if event_state is not None:
self._values["event_state"] = event_state
if rule_name is not None:
self._values["rule_name"] = rule_name
if targets is not None:
self._values["targets"] = targets
if account_id is not None:
self._values["account_id"] = account_id
if account_name is not None:
self._values["account_name"] = account_name
if ou_id is not None:
self._values["ou_id"] = ou_id
if ou_name is not None:
self._values["ou_name"] = ou_name
@builtins.property
def description(self) -> typing.Optional[builtins.str]:
'''A description of the rule's purpose.
:default: - A rule for new account creation in Organizations
'''
result = self._values.get("description")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def enabled(self) -> typing.Optional[builtins.bool]:
'''Indicates whether the rule is enabled.
:default: true
'''
result = self._values.get("enabled")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def event_bus(self) -> typing.Optional[aws_cdk.aws_events.IEventBus]:
'''The event bus to associate with this rule.
:default: - The default event bus.
'''
result = self._values.get("event_bus")
return typing.cast(typing.Optional[aws_cdk.aws_events.IEventBus], result)
@builtins.property
def event_state(self) -> typing.Optional[EventStates]:
'''Which event state should this rule trigger for.
:default: - EventStates.SUCCEEDED
'''
result = self._values.get("event_state")
return typing.cast(typing.Optional[EventStates], result)
@builtins.property
def rule_name(self) -> typing.Optional[builtins.str]:
'''A name for the rule.
:default:
- AWS CloudFormation generates a unique physical ID and uses that ID
for the rule name. For more information, see Name Type.
'''
result = self._values.get("rule_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def targets(self) -> typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]]:
'''Targets to invoke when this rule matches an event.
:default: - No targets.
'''
result = self._values.get("targets")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_events.IRuleTarget]], result)
@builtins.property
def account_id(self) -> typing.Optional[builtins.str]:
'''The account ID to match.'''
result = self._values.get("account_id")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def account_name(self) -> typing.Optional[builtins.str]:
'''The account name to match.'''
result = self._values.get("account_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def ou_id(self) -> typing.Optional[builtins.str]:
'''The OU ID to match.'''
result = self._values.get("ou_id")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def ou_name(self) -> typing.Optional[builtins.str]:
'''The OU name to match.'''
result = self._values.get("ou_name")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AccountRuleProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"AccountRuleProps",
"BaseRuleProps",
"CreatedAccountByOrganizationsRule",
"CreatedAccountRule",
"DeregisteredOrganizationalUnitRule",
"DisabledGuardrailRule",
"EnabledGuardrailRule",
"EventStates",
"GuardrailBehaviors",
"GuardrailRuleProps",
"OuRuleProps",
"RegisteredOrganizationalUnitRule",
"SetupLandingZoneRule",
"UpdatedLandingZoneRule",
"UpdatedManagedAccountRule",
]
publication.publish() | /renovosolutions.aws_cdk_control_tower_lifecycle_events-0.1.43-py3-none-any.whl/control-tower-lifecycle-events/__init__.py | 0.672439 | 0.201322 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk.aws_events
import constructs
class At(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-one-time-event.At",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
date: datetime.datetime,
) -> None:
'''
:param scope: -
:param id: -
:param date: The future date to use for one time event.
'''
props = AtProps(date=date)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="schedule")
def schedule(self) -> aws_cdk.aws_events.Schedule:
return typing.cast(aws_cdk.aws_events.Schedule, jsii.get(self, "schedule"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-one-time-event.AtProps",
jsii_struct_bases=[],
name_mapping={"date": "date"},
)
class AtProps:
def __init__(self, *, date: datetime.datetime) -> None:
'''
:param date: The future date to use for one time event.
'''
self._values: typing.Dict[str, typing.Any] = {
"date": date,
}
@builtins.property
def date(self) -> datetime.datetime:
'''The future date to use for one time event.'''
result = self._values.get("date")
assert result is not None, "Required property 'date' is missing"
return typing.cast(datetime.datetime, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AtProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class OnDeploy(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-one-time-event.OnDeploy",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
offset_minutes: typing.Optional[jsii.Number] = None,
) -> None:
'''
:param scope: -
:param id: -
:param offset_minutes: The number of minutes to add to the current time when generating the expression. Should exceed the expected time for the appropriate resources to converge. Default: 10
'''
props = OnDeployProps(offset_minutes=offset_minutes)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="schedule")
def schedule(self) -> aws_cdk.aws_events.Schedule:
return typing.cast(aws_cdk.aws_events.Schedule, jsii.get(self, "schedule"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-one-time-event.OnDeployProps",
jsii_struct_bases=[],
name_mapping={"offset_minutes": "offsetMinutes"},
)
class OnDeployProps:
def __init__(self, *, offset_minutes: typing.Optional[jsii.Number] = None) -> None:
'''
:param offset_minutes: The number of minutes to add to the current time when generating the expression. Should exceed the expected time for the appropriate resources to converge. Default: 10
'''
self._values: typing.Dict[str, typing.Any] = {}
if offset_minutes is not None:
self._values["offset_minutes"] = offset_minutes
@builtins.property
def offset_minutes(self) -> typing.Optional[jsii.Number]:
'''The number of minutes to add to the current time when generating the expression.
Should exceed the expected time for the appropriate resources to converge.
:default: 10
'''
result = self._values.get("offset_minutes")
return typing.cast(typing.Optional[jsii.Number], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "OnDeployProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"At",
"AtProps",
"OnDeploy",
"OnDeployProps",
]
publication.publish() | /renovosolutions.aws_cdk_one_time_event-2.0.200-py3-none-any.whl/one_time_event/__init__.py | 0.739328 | 0.278637 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk.aws_ec2
import aws_cdk.aws_iam
import aws_cdk.aws_route53
import constructs
import managed_instance_role
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-renovo-instance-service.AmiLookup",
jsii_struct_bases=[],
name_mapping={"name": "name", "owners": "owners", "windows": "windows"},
)
class AmiLookup:
def __init__(
self,
*,
name: builtins.str,
owners: typing.Optional[typing.Sequence[builtins.str]] = None,
windows: typing.Optional[builtins.bool] = None,
) -> None:
'''
:param name: The name string to use for AMI lookup.
:param owners: The owners to use for AMI lookup.
:param windows: Is this AMI expected to be windows?
'''
self._values: typing.Dict[str, typing.Any] = {
"name": name,
}
if owners is not None:
self._values["owners"] = owners
if windows is not None:
self._values["windows"] = windows
@builtins.property
def name(self) -> builtins.str:
'''The name string to use for AMI lookup.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def owners(self) -> typing.Optional[typing.List[builtins.str]]:
'''The owners to use for AMI lookup.'''
result = self._values.get("owners")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
@builtins.property
def windows(self) -> typing.Optional[builtins.bool]:
'''Is this AMI expected to be windows?'''
result = self._values.get("windows")
return typing.cast(typing.Optional[builtins.bool], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "AmiLookup(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class InstanceService(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-renovo-instance-service.InstanceService",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
instance_type: aws_cdk.aws_ec2.InstanceType,
machine_image: aws_cdk.aws_ec2.IMachineImage,
name: builtins.str,
vpc: aws_cdk.aws_ec2.IVpc,
allow_all_outbound: typing.Optional[builtins.bool] = None,
availability_zones: typing.Optional[typing.Sequence[builtins.str]] = None,
block_devices: typing.Optional[typing.Sequence[aws_cdk.aws_ec2.BlockDevice]] = None,
disable_inline_rules: typing.Optional[builtins.bool] = None,
enable_cloudwatch_logs: typing.Optional[builtins.bool] = None,
enabled_no_public_ingress_aspect: typing.Optional[builtins.bool] = None,
enable_no_db_ports_aspect: typing.Optional[builtins.bool] = None,
enable_no_remote_management_ports_aspect: typing.Optional[builtins.bool] = None,
instance_role: typing.Optional[managed_instance_role.ManagedInstanceRole] = None,
key_name: typing.Optional[builtins.str] = None,
parent_domain: typing.Optional[builtins.str] = None,
private_ip_address: typing.Optional[builtins.str] = None,
require_imdsv2: typing.Optional[builtins.bool] = None,
security_group: typing.Optional[aws_cdk.aws_ec2.SecurityGroup] = None,
subnet_type: typing.Optional[aws_cdk.aws_ec2.SubnetType] = None,
use_imdsv2_custom_aspect: typing.Optional[builtins.bool] = None,
user_data: typing.Optional[aws_cdk.aws_ec2.UserData] = None,
) -> None:
'''
:param scope: -
:param id: -
:param instance_type: The type of instance to launch.
:param machine_image: AMI to launch.
:param name: The name of the service the instance is for.
:param vpc: The VPC to launch the instance in.
:param allow_all_outbound: Whether the instance could initiate connections to anywhere by default.
:param availability_zones: Select subnets only in the given AZs.
:param block_devices: Specifies how block devices are exposed to the instance. You can specify virtual devices and EBS volumes
:param disable_inline_rules: Whether to disable inline ingress and egress rule optimization for the instances security group. If this is set to true, ingress and egress rules will not be declared under the SecurityGroup in cloudformation, but will be separate elements. Inlining rules is an optimization for producing smaller stack templates. Sometimes this is not desirable, for example when security group access is managed via tags. The default value can be overriden globally by setting the context variable '@aws-cdk/aws-ec2.securityGroupDisableInlineRules'. Default: false
:param enable_cloudwatch_logs: Whether or not to enable logging to Cloudwatch Logs. Default: true
:param enabled_no_public_ingress_aspect: Whether or not to prevent security group from containing rules that allow access from the public internet: Any rule with a source from 0.0.0.0/0 or ::/0. If these sources are used when this is enabled and error will be added to CDK metadata and deployment and synth will fail.
:param enable_no_db_ports_aspect: Whether or not to prevent security group from containing rules that allow access to relational DB ports: MySQL, PostgreSQL, MariaDB, Oracle, SQL Server. If these ports are opened when this is enabled an error will be added to CDK metadata and deployment and synth will fail. Default: true
:param enable_no_remote_management_ports_aspect: Whether or not to prevent security group from containing rules that allow access to remote management ports: SSH, RDP, WinRM, WinRM over HTTPs. If these ports are opened when this is enabled an error will be added to CDK metadata and deployment and synth will fail. Default: true
:param instance_role: The role to use for this instance. Default: - A new ManagedInstanceRole will be created for this instance
:param key_name: Name of the SSH keypair to grant access to the instance.
:param parent_domain: The parent domain of the service.
:param private_ip_address: Defines a private IP address to associate with the instance.
:param require_imdsv2: Whether IMDSv2 should be required on this instance. Default: true
:param security_group: The security group to use for this instance. Default: - A new SecurityGroup will be created for this instance
:param subnet_type: The subnet type to launch this service in. Default: ec2.SubnetType.PRIVATE_WITH_NAT
:param use_imdsv2_custom_aspect: Whether to use th IMDSv2 custom aspect provided by this library or the default one provided by AWS. Turned on by default otherwise we need to apply a feature flag to every project using an instance or apply a breaking change to instance construct ids. Default: true
:param user_data: The user data to apply to the instance.
'''
props = InstanceServiceProps(
instance_type=instance_type,
machine_image=machine_image,
name=name,
vpc=vpc,
allow_all_outbound=allow_all_outbound,
availability_zones=availability_zones,
block_devices=block_devices,
disable_inline_rules=disable_inline_rules,
enable_cloudwatch_logs=enable_cloudwatch_logs,
enabled_no_public_ingress_aspect=enabled_no_public_ingress_aspect,
enable_no_db_ports_aspect=enable_no_db_ports_aspect,
enable_no_remote_management_ports_aspect=enable_no_remote_management_ports_aspect,
instance_role=instance_role,
key_name=key_name,
parent_domain=parent_domain,
private_ip_address=private_ip_address,
require_imdsv2=require_imdsv2,
security_group=security_group,
subnet_type=subnet_type,
use_imdsv2_custom_aspect=use_imdsv2_custom_aspect,
user_data=user_data,
)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instance")
def instance(self) -> aws_cdk.aws_ec2.Instance:
'''The underlying instance resource.'''
return typing.cast(aws_cdk.aws_ec2.Instance, jsii.get(self, "instance"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceAvailabilityZone")
def instance_availability_zone(self) -> builtins.str:
'''The availability zone of the instance.'''
return typing.cast(builtins.str, jsii.get(self, "instanceAvailabilityZone"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceCfn")
def instance_cfn(self) -> aws_cdk.aws_ec2.CfnInstance:
'''The underlying CfnInstance resource.'''
return typing.cast(aws_cdk.aws_ec2.CfnInstance, jsii.get(self, "instanceCfn"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceEc2PrivateDnsName")
def instance_ec2_private_dns_name(self) -> builtins.str:
'''Private DNS name for this instance assigned by EC2.'''
return typing.cast(builtins.str, jsii.get(self, "instanceEc2PrivateDnsName"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceEc2PublicDnsName")
def instance_ec2_public_dns_name(self) -> builtins.str:
'''Public DNS name for this instance assigned by EC2.'''
return typing.cast(builtins.str, jsii.get(self, "instanceEc2PublicDnsName"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceId")
def instance_id(self) -> builtins.str:
'''The instance's ID.'''
return typing.cast(builtins.str, jsii.get(self, "instanceId"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instancePrivateIp")
def instance_private_ip(self) -> builtins.str:
'''Private IP for this instance.'''
return typing.cast(builtins.str, jsii.get(self, "instancePrivateIp"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceProfile")
def instance_profile(self) -> aws_cdk.aws_iam.CfnInstanceProfile:
'''The instance profile associated with this instance.'''
return typing.cast(aws_cdk.aws_iam.CfnInstanceProfile, jsii.get(self, "instanceProfile"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceRole")
def instance_role(self) -> managed_instance_role.ManagedInstanceRole:
'''The instance role associated with this instance.'''
return typing.cast(managed_instance_role.ManagedInstanceRole, jsii.get(self, "instanceRole"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="osType")
def os_type(self) -> aws_cdk.aws_ec2.OperatingSystemType:
'''The type of OS the instance is running.'''
return typing.cast(aws_cdk.aws_ec2.OperatingSystemType, jsii.get(self, "osType"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="securityGroup")
def security_group(self) -> aws_cdk.aws_ec2.SecurityGroup:
'''The security group associated with this instance.'''
return typing.cast(aws_cdk.aws_ec2.SecurityGroup, jsii.get(self, "securityGroup"))
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="instanceDnsName")
def instance_dns_name(self) -> typing.Optional[aws_cdk.aws_route53.ARecord]:
'''DNS record for this instance created in Route53.'''
return typing.cast(typing.Optional[aws_cdk.aws_route53.ARecord], jsii.get(self, "instanceDnsName"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-renovo-instance-service.InstanceServiceProps",
jsii_struct_bases=[],
name_mapping={
"instance_type": "instanceType",
"machine_image": "machineImage",
"name": "name",
"vpc": "vpc",
"allow_all_outbound": "allowAllOutbound",
"availability_zones": "availabilityZones",
"block_devices": "blockDevices",
"disable_inline_rules": "disableInlineRules",
"enable_cloudwatch_logs": "enableCloudwatchLogs",
"enabled_no_public_ingress_aspect": "enabledNoPublicIngressAspect",
"enable_no_db_ports_aspect": "enableNoDBPortsAspect",
"enable_no_remote_management_ports_aspect": "enableNoRemoteManagementPortsAspect",
"instance_role": "instanceRole",
"key_name": "keyName",
"parent_domain": "parentDomain",
"private_ip_address": "privateIpAddress",
"require_imdsv2": "requireImdsv2",
"security_group": "securityGroup",
"subnet_type": "subnetType",
"use_imdsv2_custom_aspect": "useImdsv2CustomAspect",
"user_data": "userData",
},
)
class InstanceServiceProps:
def __init__(
self,
*,
instance_type: aws_cdk.aws_ec2.InstanceType,
machine_image: aws_cdk.aws_ec2.IMachineImage,
name: builtins.str,
vpc: aws_cdk.aws_ec2.IVpc,
allow_all_outbound: typing.Optional[builtins.bool] = None,
availability_zones: typing.Optional[typing.Sequence[builtins.str]] = None,
block_devices: typing.Optional[typing.Sequence[aws_cdk.aws_ec2.BlockDevice]] = None,
disable_inline_rules: typing.Optional[builtins.bool] = None,
enable_cloudwatch_logs: typing.Optional[builtins.bool] = None,
enabled_no_public_ingress_aspect: typing.Optional[builtins.bool] = None,
enable_no_db_ports_aspect: typing.Optional[builtins.bool] = None,
enable_no_remote_management_ports_aspect: typing.Optional[builtins.bool] = None,
instance_role: typing.Optional[managed_instance_role.ManagedInstanceRole] = None,
key_name: typing.Optional[builtins.str] = None,
parent_domain: typing.Optional[builtins.str] = None,
private_ip_address: typing.Optional[builtins.str] = None,
require_imdsv2: typing.Optional[builtins.bool] = None,
security_group: typing.Optional[aws_cdk.aws_ec2.SecurityGroup] = None,
subnet_type: typing.Optional[aws_cdk.aws_ec2.SubnetType] = None,
use_imdsv2_custom_aspect: typing.Optional[builtins.bool] = None,
user_data: typing.Optional[aws_cdk.aws_ec2.UserData] = None,
) -> None:
'''
:param instance_type: The type of instance to launch.
:param machine_image: AMI to launch.
:param name: The name of the service the instance is for.
:param vpc: The VPC to launch the instance in.
:param allow_all_outbound: Whether the instance could initiate connections to anywhere by default.
:param availability_zones: Select subnets only in the given AZs.
:param block_devices: Specifies how block devices are exposed to the instance. You can specify virtual devices and EBS volumes
:param disable_inline_rules: Whether to disable inline ingress and egress rule optimization for the instances security group. If this is set to true, ingress and egress rules will not be declared under the SecurityGroup in cloudformation, but will be separate elements. Inlining rules is an optimization for producing smaller stack templates. Sometimes this is not desirable, for example when security group access is managed via tags. The default value can be overriden globally by setting the context variable '@aws-cdk/aws-ec2.securityGroupDisableInlineRules'. Default: false
:param enable_cloudwatch_logs: Whether or not to enable logging to Cloudwatch Logs. Default: true
:param enabled_no_public_ingress_aspect: Whether or not to prevent security group from containing rules that allow access from the public internet: Any rule with a source from 0.0.0.0/0 or ::/0. If these sources are used when this is enabled and error will be added to CDK metadata and deployment and synth will fail.
:param enable_no_db_ports_aspect: Whether or not to prevent security group from containing rules that allow access to relational DB ports: MySQL, PostgreSQL, MariaDB, Oracle, SQL Server. If these ports are opened when this is enabled an error will be added to CDK metadata and deployment and synth will fail. Default: true
:param enable_no_remote_management_ports_aspect: Whether or not to prevent security group from containing rules that allow access to remote management ports: SSH, RDP, WinRM, WinRM over HTTPs. If these ports are opened when this is enabled an error will be added to CDK metadata and deployment and synth will fail. Default: true
:param instance_role: The role to use for this instance. Default: - A new ManagedInstanceRole will be created for this instance
:param key_name: Name of the SSH keypair to grant access to the instance.
:param parent_domain: The parent domain of the service.
:param private_ip_address: Defines a private IP address to associate with the instance.
:param require_imdsv2: Whether IMDSv2 should be required on this instance. Default: true
:param security_group: The security group to use for this instance. Default: - A new SecurityGroup will be created for this instance
:param subnet_type: The subnet type to launch this service in. Default: ec2.SubnetType.PRIVATE_WITH_NAT
:param use_imdsv2_custom_aspect: Whether to use th IMDSv2 custom aspect provided by this library or the default one provided by AWS. Turned on by default otherwise we need to apply a feature flag to every project using an instance or apply a breaking change to instance construct ids. Default: true
:param user_data: The user data to apply to the instance.
'''
self._values: typing.Dict[str, typing.Any] = {
"instance_type": instance_type,
"machine_image": machine_image,
"name": name,
"vpc": vpc,
}
if allow_all_outbound is not None:
self._values["allow_all_outbound"] = allow_all_outbound
if availability_zones is not None:
self._values["availability_zones"] = availability_zones
if block_devices is not None:
self._values["block_devices"] = block_devices
if disable_inline_rules is not None:
self._values["disable_inline_rules"] = disable_inline_rules
if enable_cloudwatch_logs is not None:
self._values["enable_cloudwatch_logs"] = enable_cloudwatch_logs
if enabled_no_public_ingress_aspect is not None:
self._values["enabled_no_public_ingress_aspect"] = enabled_no_public_ingress_aspect
if enable_no_db_ports_aspect is not None:
self._values["enable_no_db_ports_aspect"] = enable_no_db_ports_aspect
if enable_no_remote_management_ports_aspect is not None:
self._values["enable_no_remote_management_ports_aspect"] = enable_no_remote_management_ports_aspect
if instance_role is not None:
self._values["instance_role"] = instance_role
if key_name is not None:
self._values["key_name"] = key_name
if parent_domain is not None:
self._values["parent_domain"] = parent_domain
if private_ip_address is not None:
self._values["private_ip_address"] = private_ip_address
if require_imdsv2 is not None:
self._values["require_imdsv2"] = require_imdsv2
if security_group is not None:
self._values["security_group"] = security_group
if subnet_type is not None:
self._values["subnet_type"] = subnet_type
if use_imdsv2_custom_aspect is not None:
self._values["use_imdsv2_custom_aspect"] = use_imdsv2_custom_aspect
if user_data is not None:
self._values["user_data"] = user_data
@builtins.property
def instance_type(self) -> aws_cdk.aws_ec2.InstanceType:
'''The type of instance to launch.'''
result = self._values.get("instance_type")
assert result is not None, "Required property 'instance_type' is missing"
return typing.cast(aws_cdk.aws_ec2.InstanceType, result)
@builtins.property
def machine_image(self) -> aws_cdk.aws_ec2.IMachineImage:
'''AMI to launch.'''
result = self._values.get("machine_image")
assert result is not None, "Required property 'machine_image' is missing"
return typing.cast(aws_cdk.aws_ec2.IMachineImage, result)
@builtins.property
def name(self) -> builtins.str:
'''The name of the service the instance is for.'''
result = self._values.get("name")
assert result is not None, "Required property 'name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def vpc(self) -> aws_cdk.aws_ec2.IVpc:
'''The VPC to launch the instance in.'''
result = self._values.get("vpc")
assert result is not None, "Required property 'vpc' is missing"
return typing.cast(aws_cdk.aws_ec2.IVpc, result)
@builtins.property
def allow_all_outbound(self) -> typing.Optional[builtins.bool]:
'''Whether the instance could initiate connections to anywhere by default.'''
result = self._values.get("allow_all_outbound")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def availability_zones(self) -> typing.Optional[typing.List[builtins.str]]:
'''Select subnets only in the given AZs.'''
result = self._values.get("availability_zones")
return typing.cast(typing.Optional[typing.List[builtins.str]], result)
@builtins.property
def block_devices(
self,
) -> typing.Optional[typing.List[aws_cdk.aws_ec2.BlockDevice]]:
'''Specifies how block devices are exposed to the instance.
You can specify virtual devices and EBS volumes
'''
result = self._values.get("block_devices")
return typing.cast(typing.Optional[typing.List[aws_cdk.aws_ec2.BlockDevice]], result)
@builtins.property
def disable_inline_rules(self) -> typing.Optional[builtins.bool]:
'''Whether to disable inline ingress and egress rule optimization for the instances security group.
If this is set to true, ingress and egress rules will not be declared under the SecurityGroup in cloudformation, but will be separate elements.
Inlining rules is an optimization for producing smaller stack templates.
Sometimes this is not desirable, for example when security group access is managed via tags.
The default value can be overriden globally by setting the context variable '@aws-cdk/aws-ec2.securityGroupDisableInlineRules'.
:default: false
'''
result = self._values.get("disable_inline_rules")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def enable_cloudwatch_logs(self) -> typing.Optional[builtins.bool]:
'''Whether or not to enable logging to Cloudwatch Logs.
:default: true
'''
result = self._values.get("enable_cloudwatch_logs")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def enabled_no_public_ingress_aspect(self) -> typing.Optional[builtins.bool]:
'''Whether or not to prevent security group from containing rules that allow access from the public internet: Any rule with a source from 0.0.0.0/0 or ::/0.
If these sources are used when this is enabled and error will be added to CDK metadata and deployment and synth will fail.
'''
result = self._values.get("enabled_no_public_ingress_aspect")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def enable_no_db_ports_aspect(self) -> typing.Optional[builtins.bool]:
'''Whether or not to prevent security group from containing rules that allow access to relational DB ports: MySQL, PostgreSQL, MariaDB, Oracle, SQL Server.
If these ports are opened when this is enabled an error will be added to CDK metadata and deployment and synth will fail.
:default: true
'''
result = self._values.get("enable_no_db_ports_aspect")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def enable_no_remote_management_ports_aspect(
self,
) -> typing.Optional[builtins.bool]:
'''Whether or not to prevent security group from containing rules that allow access to remote management ports: SSH, RDP, WinRM, WinRM over HTTPs.
If these ports are opened when this is enabled an error will be added to CDK metadata and deployment and synth will fail.
:default: true
'''
result = self._values.get("enable_no_remote_management_ports_aspect")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def instance_role(
self,
) -> typing.Optional[managed_instance_role.ManagedInstanceRole]:
'''The role to use for this instance.
:default: - A new ManagedInstanceRole will be created for this instance
'''
result = self._values.get("instance_role")
return typing.cast(typing.Optional[managed_instance_role.ManagedInstanceRole], result)
@builtins.property
def key_name(self) -> typing.Optional[builtins.str]:
'''Name of the SSH keypair to grant access to the instance.'''
result = self._values.get("key_name")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def parent_domain(self) -> typing.Optional[builtins.str]:
'''The parent domain of the service.'''
result = self._values.get("parent_domain")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def private_ip_address(self) -> typing.Optional[builtins.str]:
'''Defines a private IP address to associate with the instance.'''
result = self._values.get("private_ip_address")
return typing.cast(typing.Optional[builtins.str], result)
@builtins.property
def require_imdsv2(self) -> typing.Optional[builtins.bool]:
'''Whether IMDSv2 should be required on this instance.
:default: true
'''
result = self._values.get("require_imdsv2")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def security_group(self) -> typing.Optional[aws_cdk.aws_ec2.SecurityGroup]:
'''The security group to use for this instance.
:default: - A new SecurityGroup will be created for this instance
'''
result = self._values.get("security_group")
return typing.cast(typing.Optional[aws_cdk.aws_ec2.SecurityGroup], result)
@builtins.property
def subnet_type(self) -> typing.Optional[aws_cdk.aws_ec2.SubnetType]:
'''The subnet type to launch this service in.
:default: ec2.SubnetType.PRIVATE_WITH_NAT
'''
result = self._values.get("subnet_type")
return typing.cast(typing.Optional[aws_cdk.aws_ec2.SubnetType], result)
@builtins.property
def use_imdsv2_custom_aspect(self) -> typing.Optional[builtins.bool]:
'''Whether to use th IMDSv2 custom aspect provided by this library or the default one provided by AWS.
Turned on by default otherwise we need to apply a feature flag to every project using an instance or
apply a breaking change to instance construct ids.
:default: true
:see: https://github.com/jericht/aws-cdk/blob/56c01aedc4f745eec79409c99b749f516ffc39e1/packages/%40aws-cdk/aws-ec2/lib/aspects/require-imdsv2-aspect.ts#L95
'''
result = self._values.get("use_imdsv2_custom_aspect")
return typing.cast(typing.Optional[builtins.bool], result)
@builtins.property
def user_data(self) -> typing.Optional[aws_cdk.aws_ec2.UserData]:
'''The user data to apply to the instance.'''
result = self._values.get("user_data")
return typing.cast(typing.Optional[aws_cdk.aws_ec2.UserData], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "InstanceServiceProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
class ManagedLoggingPolicy(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-renovo-instance-service.ManagedLoggingPolicy",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
os: builtins.str,
) -> None:
'''
:param scope: -
:param id: -
:param os: The OS of the instance this policy is for.
'''
props = ManagedLoggingPolicyProps(os=os)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="policy")
def policy(self) -> aws_cdk.aws_iam.ManagedPolicy:
return typing.cast(aws_cdk.aws_iam.ManagedPolicy, jsii.get(self, "policy"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-renovo-instance-service.ManagedLoggingPolicyProps",
jsii_struct_bases=[],
name_mapping={"os": "os"},
)
class ManagedLoggingPolicyProps:
def __init__(self, *, os: builtins.str) -> None:
'''
:param os: The OS of the instance this policy is for.
'''
self._values: typing.Dict[str, typing.Any] = {
"os": os,
}
@builtins.property
def os(self) -> builtins.str:
'''The OS of the instance this policy is for.'''
result = self._values.get("os")
assert result is not None, "Required property 'os' is missing"
return typing.cast(builtins.str, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "ManagedLoggingPolicyProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"AmiLookup",
"InstanceService",
"InstanceServiceProps",
"ManagedLoggingPolicy",
"ManagedLoggingPolicyProps",
]
publication.publish() | /renovosolutions.aws_cdk_renovo_instance_service-2.8.0-py3-none-any.whl/renovo-instance-service/__init__.py | 0.702836 | 0.224874 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from ._jsii import *
import aws_cdk.aws_s3
import constructs
class RenovoS3Bucket(
constructs.Construct,
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-renovo-s3-bucket.RenovoS3Bucket",
):
def __init__(
self,
scope: constructs.Construct,
id: builtins.str,
*,
lifecycle_rules: typing.Sequence[aws_cdk.aws_s3.LifecycleRule],
name: typing.Optional[builtins.str] = None,
) -> None:
'''
:param scope: -
:param id: -
:param lifecycle_rules: Rules that define how Amazon S3 manages objects during their lifetime.
:param name: The name of the bucket.
'''
props = RenovoS3BucketProps(lifecycle_rules=lifecycle_rules, name=name)
jsii.create(self.__class__, self, [scope, id, props])
@builtins.property # type: ignore[misc]
@jsii.member(jsii_name="bucket")
def bucket(self) -> aws_cdk.aws_s3.Bucket:
return typing.cast(aws_cdk.aws_s3.Bucket, jsii.get(self, "bucket"))
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-renovo-s3-bucket.RenovoS3BucketProps",
jsii_struct_bases=[],
name_mapping={"lifecycle_rules": "lifecycleRules", "name": "name"},
)
class RenovoS3BucketProps:
def __init__(
self,
*,
lifecycle_rules: typing.Sequence[aws_cdk.aws_s3.LifecycleRule],
name: typing.Optional[builtins.str] = None,
) -> None:
'''
:param lifecycle_rules: Rules that define how Amazon S3 manages objects during their lifetime.
:param name: The name of the bucket.
'''
self._values: typing.Dict[str, typing.Any] = {
"lifecycle_rules": lifecycle_rules,
}
if name is not None:
self._values["name"] = name
@builtins.property
def lifecycle_rules(self) -> typing.List[aws_cdk.aws_s3.LifecycleRule]:
'''Rules that define how Amazon S3 manages objects during their lifetime.'''
result = self._values.get("lifecycle_rules")
assert result is not None, "Required property 'lifecycle_rules' is missing"
return typing.cast(typing.List[aws_cdk.aws_s3.LifecycleRule], result)
@builtins.property
def name(self) -> typing.Optional[builtins.str]:
'''The name of the bucket.'''
result = self._values.get("name")
return typing.cast(typing.Optional[builtins.str], result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "RenovoS3BucketProps(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
__all__ = [
"RenovoS3Bucket",
"RenovoS3BucketProps",
]
publication.publish() | /renovosolutions.aws-cdk-renovo-s3-bucket-2.1.195.tar.gz/renovosolutions.aws-cdk-renovo-s3-bucket-2.1.195/src/renovo_s3_bucket/__init__.py | 0.716814 | 0.174709 | __init__.py | pypi |
import abc
import builtins
import datetime
import enum
import typing
import jsii
import publication
import typing_extensions
from typeguard import check_type
from ._jsii import *
import aws_cdk.aws_route53 as _aws_cdk_aws_route53_ceddda9d
@jsii.data_type(
jsii_type="@renovosolutions/cdk-library-route53targets.LoadBalancerTargetAttributes",
jsii_struct_bases=[],
name_mapping={"dns_name": "dnsName", "hosted_zone_id": "hostedZoneId"},
)
class LoadBalancerTargetAttributes:
def __init__(self, *, dns_name: builtins.str, hosted_zone_id: builtins.str) -> None:
'''
:param dns_name: The DNS name of the load balancer.
:param hosted_zone_id: The hosted zone ID of the load balancer.
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__d7d0adc99abf87b53514ecd4cfc8ed4245e5f713a5e9663749820c1058bbed7b)
check_type(argname="argument dns_name", value=dns_name, expected_type=type_hints["dns_name"])
check_type(argname="argument hosted_zone_id", value=hosted_zone_id, expected_type=type_hints["hosted_zone_id"])
self._values: typing.Dict[builtins.str, typing.Any] = {
"dns_name": dns_name,
"hosted_zone_id": hosted_zone_id,
}
@builtins.property
def dns_name(self) -> builtins.str:
'''The DNS name of the load balancer.'''
result = self._values.get("dns_name")
assert result is not None, "Required property 'dns_name' is missing"
return typing.cast(builtins.str, result)
@builtins.property
def hosted_zone_id(self) -> builtins.str:
'''The hosted zone ID of the load balancer.'''
result = self._values.get("hosted_zone_id")
assert result is not None, "Required property 'hosted_zone_id' is missing"
return typing.cast(builtins.str, result)
def __eq__(self, rhs: typing.Any) -> builtins.bool:
return isinstance(rhs, self.__class__) and rhs._values == self._values
def __ne__(self, rhs: typing.Any) -> builtins.bool:
return not (rhs == self)
def __repr__(self) -> str:
return "LoadBalancerTargetAttributes(%s)" % ", ".join(
k + "=" + repr(v) for k, v in self._values.items()
)
@jsii.implements(_aws_cdk_aws_route53_ceddda9d.IAliasRecordTarget)
class LoadBalancerTargetFromAttributes(
metaclass=jsii.JSIIMeta,
jsii_type="@renovosolutions/cdk-library-route53targets.LoadBalancerTargetFromAttributes",
):
def __init__(self, *, dns_name: builtins.str, hosted_zone_id: builtins.str) -> None:
'''
:param dns_name: The DNS name of the load balancer.
:param hosted_zone_id: The hosted zone ID of the load balancer.
'''
load_balancer_target_attributes = LoadBalancerTargetAttributes(
dns_name=dns_name, hosted_zone_id=hosted_zone_id
)
jsii.create(self.__class__, self, [load_balancer_target_attributes])
@jsii.member(jsii_name="bind")
def bind(
self,
_record: _aws_cdk_aws_route53_ceddda9d.IRecordSet,
_zone: typing.Optional[_aws_cdk_aws_route53_ceddda9d.IHostedZone] = None,
) -> _aws_cdk_aws_route53_ceddda9d.AliasRecordTargetConfig:
'''Return hosted zone ID and DNS name, usable for Route53 alias targets.
:param _record: -
:param _zone: -
'''
if __debug__:
type_hints = typing.get_type_hints(_typecheckingstub__e73ec795700ccbb5bcf9f6b715372577573289ebb6b702e4703287effb1115d6)
check_type(argname="argument _record", value=_record, expected_type=type_hints["_record"])
check_type(argname="argument _zone", value=_zone, expected_type=type_hints["_zone"])
return typing.cast(_aws_cdk_aws_route53_ceddda9d.AliasRecordTargetConfig, jsii.invoke(self, "bind", [_record, _zone]))
__all__ = [
"LoadBalancerTargetAttributes",
"LoadBalancerTargetFromAttributes",
]
publication.publish()
def _typecheckingstub__d7d0adc99abf87b53514ecd4cfc8ed4245e5f713a5e9663749820c1058bbed7b(
*,
dns_name: builtins.str,
hosted_zone_id: builtins.str,
) -> None:
"""Type checking stubs"""
pass
def _typecheckingstub__e73ec795700ccbb5bcf9f6b715372577573289ebb6b702e4703287effb1115d6(
_record: _aws_cdk_aws_route53_ceddda9d.IRecordSet,
_zone: typing.Optional[_aws_cdk_aws_route53_ceddda9d.IHostedZone] = None,
) -> None:
"""Type checking stubs"""
pass | /renovosolutions.aws_cdk_route53targets-0.0.131-py3-none-any.whl/route53targets/__init__.py | 0.595022 | 0.205117 | __init__.py | pypi |
import os
import warnings
from typing import Optional, Callable
from functools import wraps
def deprecated(message: str = ""):
"""Mark a function as deprecated.
This is used as a decorator to mark some functions as deprecated without needing to import
warnings repeatedly. The function that uses the decorator will be called but will display
a deprecation warning with a supplied message.
Arguments:
message (str): The message or reason for deprecation. Defaults to a generic statement
generated by the function's name.
Returns:
warnable (Callable): The function with a warning wrapper.
"""
def warnable(call: Callable):
@wraps(call)
def do_call(*args, **kwargs):
warnings.warn(message if message else call.__name__ + " is deprecated.",
category=DeprecationWarning)
call(*args, **kwargs)
return do_call
return warnable
@deprecated(message="Please use isfile or is from the os.path module.")
def file_exists(directory: str = os.getcwd(), item: str = '') -> Optional[str]:
"""Determine whether a file exists in a certain directory.
**Note**: This function is being deprecated in favor of the utilities provided in the `os`
module.
Args:
dir (str): The directory to search in.
item (str): The item to search for in the directory.
Returns:
fname (str): File name if it's found or `None` if it doesn't find anything
"""
cwd = os.listdir(directory)
for listed_item in cwd:
if item in listed_item:
return listed_item
return None
@deprecated("This function is no longer supported.")
def verify_built_files(directory: str = os.getcwd()) -> Optional[str]:
"""Determine if the Ren'Py distributions have been built already by looking for the `-dists`
directory.
This function will check if the directory exists in itself.
Args:
dir (str): The directory to search.
Returns:
isdir (bool): Whether the directory exists or not.
"""
return file_exists(directory=directory, item="-dists")
@deprecated("This function is longer supported.")
def find_mac_build(directory: str) -> bool:
"""Determine whether the macOS builds have been created.
Args:
dir (str): The directory to search in
Returns:
isfile (bool): Whether the macOS ZIP file exists.
"""
return file_exists(directory=directory, item="-mac") | /renpy_distribute_tools-0.4.1.post1-py3-none-any.whl/renpy_distribute_tools/util.py | 0.838184 | 0.251205 | util.py | pypi |
import subprocess as proc
import enum
class ButlerPlatformType(enum.Enum):
"""Enumerations for platform types."""
WINDOWS = "win"
DARWIN = "mac"
LINUX = "linux"
OTHER = ""
DEFAULT_TAG_RULES = {
"win": ButlerPlatformType.WINDOWS,
"mac": ButlerPlatformType.DARWIN,
"linux": ButlerPlatformType.LINUX
}
class Butler(object):
"""The handler for publishing content to Itch.io using Butler.
Attributes:
author (str): The itch.io username that is publishing content.
project (str): The project that the author is publishing content for.
bin (str): The path to the Butler executable.
tag_rules (dict): A dictionary containing rules for additional tags.
"""
def __init__(self, author, project, **kwargs):
"""Initialize a Butler class.
Arguments:
author (str): The itch.io user that will submit a project.
project (str): The project that the user will submit.
Kwargs:
exec (str): The path to the Butler executable file. Defaults to "butler".
"""
self.author = author
self.project = project
self.tag_rules = DEFAULT_TAG_RULES.copy()
self.bin = kwargs["exec"] if "exec" in kwargs else "butler"
def push(self, file: str, **kwargs):
"""Push the file to the Itch.io page.
Arguments:
file (str): The path to the file to push.
**kwargs: Arbitrary keyword arguments
Kwargs:
user_version (str): The user version to use, if any.
with_tag_rule (str): The tag rule to use. This is used as a channel.
with_custom_tag (str): The tag to use at the end of the tag rule name.
"""
channel = self.author + "/" + self.project
if "with_tag_rule" in kwargs:
tag: ButlerPlatformType = self.tag_rules.get(
kwargs["with_tag_rule"], ButlerPlatformType.OTHER)
channel += ":" + tag.value
if "with_custom_tag" in kwargs:
channel += "-" + kwargs["with_custom_tag"]
command = [self.bin, "push", file, channel]
if "user_version" in kwargs:
command += ["--userversion", kwargs["user_version"]]
return proc.check_call(command)
def add_tag_rule(self, name: str, platform: ButlerPlatformType):
"""Create a new rule for the project's channel tags.
Arguments:
name (str): The rule that will determine what platform to store it under.
platform (ButlerPlatformType): The platform for that rule.
"""
self.tag_rules[name] = platform | /renpy_distribute_tools-0.4.1.post1-py3-none-any.whl/renpy_distribute_tools/itch.py | 0.663342 | 0.172729 | itch.py | pypi |
import os
import subprocess
from sys import platform
from functools import wraps
def darwin_only(call):
"""A decorator for macOS-specific commands.
This should be used to denote that a function only works on macOS due to reliance on built-in
tools from macOS or Xcode.
"""
@wraps(call)
def darwin_call(*args, **kwargs):
if platform.lower() != "darwin":
raise OSError("Function %s only works on macOS." % (call))
return call(*args, **kwargs)
return darwin_call
@darwin_only
def package_app_zip(app: str):
"""Create a ZIP file of the app.
Args:
app (str): The path to the macOS to make an archive of
"""
if os.path.isdir(app):
zip_commands = ["ditto", "-c", "-k", "--rsrc",
"--keepParent", app, app + ".zip"]
subprocess.check_call(zip_commands)
else:
raise NotADirectoryError(
"The .app file is either missing or not present.")
@darwin_only
def build_pkg(app: str, identity: str, package_name: str):
"""Create an installable package from a macOS app.
By default, it will create an app package that installs to `/Applications/`. This package
installer can also be used to submit an app to the Mac App Store.
If the package name isn't a file path, `.pkg` will automatically be appended at the end of the
name.
Args:
app (str): The path to the app to create a package of.
identity (str): The identity to sign the package with
package_name (str): The name or path of the resulting package.
"""
package_file = package_name
if ".pkg" not in package_name:
package_file = package_name + ".pkg"
commands = ["productbuild", "--component", app,
"/Applications", "--sign", identity, package_file]
return subprocess.check_call(commands)
@darwin_only
def code_sign(identity: str, app_directory: str, **kwargs):
"""Digitally sign a macOS application with a signing identity and any entitlements.
Args:
identity (str): The identity to use during signing, usually a Developer ID.
app_directory (str): The path to the macOS application for signing.
**kwargs: Arbitrary keyword arguments.
Kwargs:
entitlements (str): (Optional) The path to the entitlements the app should be signed with.
enable_hardened_runtime (bool): Whether to sign the app with the hardened runtime on.
"""
commands = ["codesign",
"--timestamp",
"--deep",
"--force",
"--no-strict",
"--sign",
identity,
app_directory]
if "entitlements" in kwargs:
commands.append("--entitlements")
commands.append(kwargs["entitlements"])
if "enable_hardened_runtime" in kwargs and kwargs["enable_hardened_runtime"]:
commands.append("--options=runtime")
return subprocess.check_call(commands)
@darwin_only
def upload_to_notary(app: str,
identifier: str,
username: str,
password: str,
**kwargs) -> str:
"""Upload a macOS application archive to Apple's notary service for notarization.
Args:
app (str): The path to the macOS application to send to Apple.
identifier (str): The bundle identifier of the application.
username (str): The username (email address) of the Apple ID to notarize under.
password (str): The password of the Apple ID to notarize under.
**kwargs: Arbitrary keyword arguments.
Kwargs:
provider (str): The App Store Connect or iTunes Connect provider associated with the Apple
ID used to sign the app.
Returns:
uuid_str (str): The request UUID.
"""
package_app_zip(app)
commands = ["xcrun", "altool", "-t", "osx", "-f", app + ".zip",
"--notarize-app", "--primary-bundle-id", identifier,
"-u", username, "-p", password]
if "provider" in kwargs:
commands += ["-itc_provider", kwargs["provider"]]
result = subprocess.check_output( # pylint:disable=unexpected-keyword-arg
commands, text=True)
os.remove(app + ".zip")
result = result.split("\n")
trimmed = result[1:]
if len(trimmed) < 1:
return ""
return trimmed[0].replace("RequestUUID = ", "")
@darwin_only
def check_notary_status(uuid: str, username: str, password: str) -> int:
"""Get the notarization status of a given UUID.
Arguments:
uuid (str): The UUID of the app to check the status of.
username (str): The user that submitted the notarization request.
password (str): The password to use to sign into Apple.
Returns:
status (int): The status code associated with the UUID notarization request. A code of `-1`
indicates that getting the status code failed, either because the item could not be
found or because no status code has been given yet.
"""
result = subprocess.check_output( # pylint:disable=unexpected-keyword-arg
["xcrun", "altool", "--notarization-info", uuid, "-u", username, "-p", password], text=True)
status = [x for x in result.replace(
" ", "").split("\n") if "Status Code" in x]
if len(status) < 1:
return -1
status = status[0].replace("Status Code", "").replace(" ", "").split(":")
while '' in status:
status.remove('')
if len(status) < 1:
return -1
return int(status[0])
@darwin_only
def staple(app: str):
"""Staple a notarization ticket to a notarized app.
Args:
app (str): The path of the macOS app to staple the ticket to.
"""
commands = ["xcrun", "stapler", "staple", app]
return subprocess.check_call(commands) | /renpy_distribute_tools-0.4.1.post1-py3-none-any.whl/renpy_distribute_tools/apple.py | 0.712932 | 0.223038 | apple.py | pypi |
from random import SystemRandom
from paillier.crypto import encrypt, secure_addition, scalar_multiplication, secure_subtraction
from eqt.util.exceptions import ProtocolError
from eqt.util.log import logger
r = SystemRandom()
class PartyA:
def __init__(self, pk, a, b, length, kappa):
self.xis, self.xors, self.cis, self.delta_b = None, None, None, None
self._pk = pk
self._n, _ = pk
self._a = a
self._b = b
self._l = length
self._k = kappa
self._r = None
self._delta = None
def generate_r(self):
self._r = r.getrandbits((self._l + 1 + self._k))
@property
def x(self):
try:
return secure_addition(secure_subtraction(self._a, self._b, self._n),
encrypt(self._pk, self._r), self._n)
except TypeError:
raise ProtocolError("x was accessed before r was generated.")
def compute_xor(self):
if self.xis is None:
raise ProtocolError("xis was accessed before it was set")
r_bits = f'{self._r:b}'[::-1].rjust(len(self.xis), '0')[:len(self.xis)]
assert len(r_bits) == len(self.xis), f"r_bits and xis should be the same length, are " \
f"{len(r_bits)} and {len(self.xis)}"
self.xors = list(self._get_xor(r_bits, self.xis))
def determine_delta(self):
self._delta = r.randrange(0, 2)
def compute_cis(self):
if self._delta == 0:
logger.debug("Computing hamming distance")
self._compute_cis_hamming()
else:
logger.debug("Computing secure comparison based on DGK")
self._compute_cis_comparison()
def _compute_cis_hamming(self):
c0 = self._compute_c0()
cis = [encrypt(self._pk, r.getrandbits(self._k))
for _ in range(self._l - 1)]
self.cis = [c0] + cis
def _compute_c0(self):
c0 = self._product(self.xors)
p = r.getrandbits(self._k)
return scalar_multiplication(c0, p, self._n)
def _compute_cis_comparison(self):
cis = list(self._build_cis_comparison())
self.cis = [scalar_multiplication(ci, r.getrandbits(self._k), self._n) for ci in cis]
def _build_cis_comparison(self):
for i in range(0, self._l):
if i == self._l - 1:
prod_square = encrypt(self._pk, 0)
else:
prod = self._product(self.xors, i+1)
prod_square = scalar_multiplication(prod, 2, self._n)
neg_1 = encrypt(self._pk, -1)
result = secure_addition(self.xors[i], prod_square, self._n)
result = secure_addition(neg_1, result, self._n)
yield result
def shuffle_cis(self):
if self.cis is None:
raise ProtocolError("[ci] was shuffled before it was set")
r.shuffle(self.cis)
@property
def curly_theta(self):
if self.delta_b is None:
raise ProtocolError("Result was requested before protocol was run")
if self._delta == 0:
return self.delta_b
return self._inverse(self.delta_b)
def _get_xor(self, r_bits, x_bits):
for ri, xi in zip(r_bits, x_bits):
if int(ri) == 0:
yield int(xi)
else:
yield self._inverse(xi)
def _product(self, xors, start=0):
r = xors[start]
for x in xors[start+1:]:
r = secure_addition(r, x, self._n)
return r
def _inverse(self, ciphertext):
inverse_xi = scalar_multiplication(ciphertext, -1, self._n)
e1 = encrypt(self._pk, 1)
return secure_addition(inverse_xi, e1, self._n) | /rens_eqt-1.5.1-py3-none-any.whl/eqt/party_a.py | 0.863536 | 0.325923 | party_a.py | pypi |
from paillier.crypto import decrypt, encrypt
from eqt.util.exceptions import ProtocolError
from eqt.party_a import PartyA
from eqt.party_b import PartyB
from eqt.util.log import logger
class Protocol:
def __init__(self, a, b, kappa, keysize=2048):
self.__a = a
self.__b = b
self._l = max(len(f'{x:b}') for x in (a, b))
self._kappa = kappa
self._keysize = keysize
self._setup()
def _setup(self):
self.party_b = PartyB(length=self._l, keysize=self._keysize)
a = encrypt(self.party_b.pk, self.__a)
b = encrypt(self.party_b.pk, self.__b)
self.party_a = PartyA(self.party_b.pk, a, b, self._l, self._kappa)
def start(self):
logger.debug('Starting protocol')
self._step1()
self._step2()
self._step3()
self._step4()
self._coinsteps()
self._step12()
self._step13()
self._step14()
self._step15()
@property
def result(self):
result = self.party_a.curly_theta
if result is None:
raise ProtocolError("Start hasn't been called or protocol failed")
return result
@property
def decrypted_result(self):
return decrypt(self.party_b.pk, self.party_b.sk, self.result)
def validate_outcome(self):
outcome = bool(self.decrypted_result)
equality = self.__a == self.__b
if outcome != equality:
logger.error(f"Equality was {equality} but protocol said it was {outcome}")
def _step1(self):
self.party_a.generate_r()
self.party_b.x = self.party_a.x
def _step2(self):
self.party_a.xis = self.party_b.xis
def _step3(self):
self.party_a.compute_xor()
def _step4(self):
self.party_a.determine_delta()
def _coinsteps(self):
self.party_a.compute_cis()
def _step12(self):
self.party_a.shuffle_cis()
self.party_b.cis = self.party_a.cis
def _step13(self):
pass
def _step14(self):
self.party_a.delta_b = self.party_b.delta
def _step15(self):
pass | /rens_eqt-1.5.1-py3-none-any.whl/eqt/protocol.py | 0.80905 | 0.223547 | protocol.py | pypi |
from renson_endura_delta.general_enum import DataType
class FieldEnum:
"""Enum of all the possible fields that can be read."""
name: str = None
field_type: DataType = None
def __init__(self, name: str, field_type: DataType):
"""Create enum with values name and field type."""
self.name = name
self.field_type = field_type
FIRMWARE_VERSION = FieldEnum("Firmware version", DataType.STRING)
CO2_QUALITY_FIELD = FieldEnum("CO2", DataType.QUALITY)
AIR_QUALITY_FIELD = FieldEnum("IAQ", DataType.QUALITY)
CO2_FIELD = FieldEnum("CO2", DataType.NUMERIC)
AIR_FIELD = FieldEnum("IAQ", DataType.NUMERIC)
CURRENT_LEVEL_FIELD = FieldEnum("Current ventilation level", DataType.LEVEL)
CURRENT_AIRFLOW_EXTRACT_FIELD = FieldEnum("Current ETA airflow", DataType.NUMERIC)
CURRENT_AIRFLOW_INGOING_FIELD = FieldEnum("Current SUP airflow", DataType.NUMERIC)
OUTDOOR_TEMP_FIELD = FieldEnum("T21", DataType.NUMERIC)
INDOOR_TEMP_FIELD = FieldEnum("T11", DataType.NUMERIC)
FILTER_REMAIN_FIELD = FieldEnum("Filter remaining time", DataType.NUMERIC)
HUMIDITY_FIELD = FieldEnum("RH11", DataType.NUMERIC)
FROST_PROTECTION_FIELD = FieldEnum("Frost protection active", DataType.BOOLEAN)
MANUAL_LEVEL_FIELD = FieldEnum("Manual level", DataType.STRING)
TIME_AND_DATE_FIELD = FieldEnum("Date and time", DataType.STRING)
BREEZE_TEMPERATURE_FIELD = FieldEnum("Breeze activation temperature", DataType.NUMERIC)
BREEZE_ENABLE_FIELD = FieldEnum("Breeze enable", DataType.BOOLEAN)
BREEZE_LEVEL_FIELD = FieldEnum("Breeze level", DataType.STRING)
DAYTIME_FIELD = FieldEnum("Start daytime", DataType.STRING)
NIGHTTIME_FIELD = FieldEnum("Start night-time", DataType.STRING)
DAY_POLLUTION_FIELD = FieldEnum("Day pollution-triggered ventilation level", DataType.STRING)
NIGHT_POLLUTION_FIELD = FieldEnum("Night pollution-triggered ventilation level", DataType.STRING)
HUMIDITY_CONTROL_FIELD = FieldEnum("Trigger internal pollution alert on RH", DataType.BOOLEAN)
AIR_QUALITY_CONTROL_FIELD = FieldEnum("Trigger internal pollution alert on IAQ", DataType.BOOLEAN)
CO2_CONTROL_FIELD = FieldEnum("Trigger internal pollution alert on CO2", DataType.BOOLEAN)
CO2_THRESHOLD_FIELD = FieldEnum("CO2 threshold", DataType.NUMERIC)
CO2_HYSTERESIS_FIELD = FieldEnum("CO2 hysteresis", DataType.NUMERIC)
BREEZE_MET_FIELD = FieldEnum("Breeze conditions met", DataType.BOOLEAN)
PREHEATER_FIELD = FieldEnum("Preheater enabled", DataType.BOOLEAN)
BYPASS_TEMPERATURE_FIELD = FieldEnum("Bypass activation temperature", DataType.NUMERIC)
BYPASS_LEVEL_FIELD = FieldEnum("Bypass level", DataType.STRING)
FILTER_PRESET_FIELD = FieldEnum("Filter preset time", DataType.NUMERIC)
DEVICE_NAME_FIELD = FieldEnum("Device name", DataType.STRING)
FIRMWARE_VERSION_FIELD = FieldEnum("Firmware version", DataType.STRING)
HARDWARE_VERSION_FIELD = FieldEnum("Hardware version", DataType.STRING)
DEVICE_TYPE = FieldEnum("Device type", DataType.STRING)
MAC_ADDRESS = FieldEnum("MAC", DataType.STRING) | /renson_endura_delta-1.5.2-py3-none-any.whl/renson_endura_delta/field_enum.py | 0.744749 | 0.410284 | field_enum.py | pypi |
import json
import logging
from datetime import datetime
import re
import requests
from renson_endura_delta.field_enum import FieldEnum, FIRMWARE_VERSION, FIRMWARE_VERSION_FIELD
from renson_endura_delta.general_enum import (Level, Quality,
ServiceNames, DataType, Level)
_LOGGER = logging.getLogger(__name__)
class ValueData:
"""Class for getting Renson data."""
def __init__(self, value):
"""Construct the class."""
self.Value = value
class RensonVentilation:
"""Main class to get data and post data to the Renson unit."""
data_url = "http://[host]/JSON/ModifiedItems?wsn=150324488709"
service_url = "http://[host]/JSON/Vars/[field]?index0=0&index1=0&index2=0"
firmware_server_url = "http://www.renson-app.com/endura_delta/firmware/check.php"
firmware_dowload_url = "http://www.renson-app.com/endura_delta/firmware/files/"
host = None
def __init__(self, host: str):
"""Initialize Renson Ventilation class by giving the host name or ip address."""
self.host = host
def connect(self) -> bool:
try:
response = requests.get(self.data_url.replace("[host]", self.host))
return response.status_code == 200
except Exception:
return False
def get_all_data(self):
response = requests.get(self.data_url.replace("[host]", self.host))
if response.status_code == 200:
return response.json()
else:
_LOGGER.error(f"Error communicating with API: {response.status_code}")
return ''
def get_field_value(self, all_data, fieldname: str) -> str:
"""Search for the field in the Reson JSON and return the value of it."""
for data in all_data["ModifiedItems"]:
if data["Name"] == fieldname == FIRMWARE_VERSION_FIELD.name:
return data["Value"].split()[-1]
elif data["Name"] == fieldname:
return data["Value"]
return ''
def parse_value(self, value, data_type):
"""Parse value to correct type"""
if data_type == DataType.NUMERIC:
return self.parse_numeric(value)
elif data_type == DataType.STRING:
return value
elif data_type == DataType.LEVEL:
return self.parse_data_level(value).value
elif data_type == DataType.BOOLEAN:
return self.parse_boolean(value)
elif data_type == DataType.QUALITY:
return self.parse_quality(value).value
def __get_service_url(self, field: ServiceNames):
"""Make the full url of the Renson API and return it."""
return self.service_url.replace("[host]", self.host).replace(
"[field]", field.value.replace(" ", "%20")
)
def __get_base_url(self, path: str):
"""Make the base url of the Renson API and return it."""
return "http://" + self.host + path
def parse_numeric(self, value: str) -> float:
"""Get the value of the field and convert it to a numeric type."""
return round(float(value))
def parse_data_level(self, value: str) -> Level:
"""Get the value of the field and convert it to a Level type."""
return Level[value.split()[-1].upper()]
def parse_boolean(self, value: str) -> bool:
"""Get the value of the field and convert it to a boolean type."""
return bool(int(value))
def parse_quality(self, value: str) -> Quality:
"""Get the value of the field and convert it to a Quality type."""
value = round(float(value))
if value < 950:
return Quality.GOOD
elif value < 1500:
return Quality.POOR
else:
return Quality.BAD
def set_manual_level(self, level: Level):
"""Set the manual level of the Renson unit. When set to 'Off' the unit will go back to auto program."""
data = ValueData(level.value)
response = requests.post(
self.__get_service_url(ServiceNames.SET_MANUAL_LEVEL_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
def restart_device(self):
"""Restart device"""
response = requests.post(
self.__get_base_url("/Reset")
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
def sync_time(self):
"""Sync time of the Renson unit to current date and time."""
response = requests.get(self.__get_service_url(ServiceNames.TIME_AND_DATE_FIELD))
if response.status_code == 200:
json_result = response.json()
device_time = datetime.strptime(
json_result["Value"], "%d %b %Y %H:%M"
)
current_time = datetime.now()
if current_time != device_time:
data = ValueData(current_time.strftime("%d %b %Y %H:%M").lower())
requests.post(
self.__get_service_url(ServiceNames.TIME_AND_DATE_FIELD), data=json.dumps(data.__dict__)
)
else:
_LOGGER.error("Ventilation unit did not return 200")
def set_timer_level(self, level: Level, time: int):
"""Set a level for a specific time (in minutes)."""
if level == Level.OFF:
raise Exception("Off is not a valid type for setting manual level")
data = ValueData(str(time) + " min " + level.value)
response = requests.post(self.__get_service_url(ServiceNames.TIMER_FIELD), data=json.dumps(data.__dict__))
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
def set_breeze(self, level: Level, temperature: int, activated: bool):
"""Activate/deactivate breeze feature and give breeze parameters to the function."""
if level == Level.HOLIDAY or level == Level.OFF or level == Level.BREEZE:
raise Exception("Holiday, Off, Breeze are not a valid types for setting breeze level")
data = ValueData(level)
response = requests.post(
self.__get_service_url(ServiceNames.BREEZE_LEVEL_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(temperature))
response = requests.post(
self.__get_service_url(ServiceNames.BREEZE_TEMPERATURE_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(int(activated)))
response = requests.post(
self.__get_service_url(ServiceNames.BREEZE_ENABLE_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
def set_time(self, day: str, night: str):
"""Set day and night time for the device."""
data = ValueData(day)
response = requests.post(self.__get_service_url(ServiceNames.DAYTIME_FIELD), data=json.dumps(data.__dict__))
if response.status_code != 200:
_LOGGER.error("Start daytime cannot be set")
data = ValueData(night)
response = requests.post(self.__get_service_url(ServiceNames.NIGTHTIME_FIELD), data=json.dumps(data.__dict__))
if response.status_code != 200:
_LOGGER.error("Start nighttime cannot be set")
def set_pollution(self, day: Level, night: Level, humidity_control: bool,
airquality_control: bool,
co2_control: bool, co2_threshold: bool, co2_hysteresis: bool):
"""Enable/disable special auto features of the Renson unit."""
if day == Level.HOLIDAY or day == Level.OFF or day == Level.BREEZE:
raise Exception("Holiday, Off, Breeze are not a valid types for setting day level")
if night == Level.HOLIDAY or night == Level.OFF or night == Level.BREEZE:
raise Exception("Holiday, Off, Breeze are not a valid types for setting night level")
data = ValueData(day.value)
response = requests.post(
self.__get_service_url(ServiceNames.DAY_POLLUTION_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(night.value)
response = requests.post(
self.__get_service_url(ServiceNames.NIGHT_POLLUTION_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(int(humidity_control)))
response = requests.post(
self.__get_service_url(ServiceNames.HUMIDITY_CONTROL_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(int(airquality_control)))
response = requests.post(
self.__get_service_url(ServiceNames.AIR_QUALITY_CONTROL_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(int(co2_control)))
response = requests.post(
self.__get_service_url(ServiceNames.CO2_CONTROL_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(int(co2_threshold)))
response = requests.post(
self.__get_service_url(ServiceNames.CO2_THRESHOLD_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
data = ValueData(str(int(co2_hysteresis)))
response = requests.post(
self.__get_service_url(ServiceNames.CO2_HYSTERESIS_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
def set_filter_days(self, days: int):
"""Set the filter days."""
data = ValueData(str(int(days)))
response = requests.post(
self.__get_service_url(ServiceNames.FILTER_DAYS_FIELD), data=json.dumps(data.__dict__)
)
if response.status_code != 200:
_LOGGER.error("Ventilation unit did not return 200")
def is_firmware_up_to_date(self, current_version) -> bool:
"""Check if the Renson firmware is up to date."""
# version = self.get_field_value(FIRMWARE_VERSION).split()[-1]
version = current_version.split()[-1]
json_string = '{"a":"check", "name":"D_' + version + '.fuf"}'
response_server = requests.post(self.firmware_server_url, data=json_string)
if response_server.status_code == 200:
return bool((response_server.json())["latest"])
return False
def get_latest_firmware_version(self) -> str:
"""Get the latest Renson firmware version."""
json_string = '{"a":"check", "name":"D_0.fuf"}'
response_server = requests.post(self.firmware_server_url, data=json_string)
if response_server.status_code == 200:
return re.sub(r"D_(.*)\.fuf", r"\1", response_server.json()["url"])
return "" | /renson_endura_delta-1.5.2-py3-none-any.whl/renson_endura_delta/renson.py | 0.682679 | 0.303035 | renson.py | pypi |
# Rent-A-Bot
[](https://travis-ci.org/cpoisson/rent-a-bot)
[](https://codecov.io/gh/cpoisson/rent-a-bot)
[](https://gitlab.com/cpoisson/rent-a-bot/commits/master)
[](https://gitlab.com/cpoisson/rent-a-bot/commits/master)
---
Rent-a-bot, your automation resource provider.
Exclusive access to a static resource is a common problem in automation, rent-a-bot allows you to abstract your resources
and lock them to prevent any concurrent access.
## Purpose
Rent-a-bot pursue the same objective as Jenkins [Lockable Resource Plugin](https://wiki.jenkins.io/display/JENKINS/Lockable+Resources+Plugin).
This latter works quite well, but only if you use... well... Jenkins.
Rent-A-Bot purpose is to fill the same needs in an environment where multiple automation applications exist.
e.g.
- Multiple Jenkins application servers
- Mixed automation application, gitlab CI + Jenkins
- Shared resources between humans and automates.
## What is a resource?
A resource is defined by a **name** and the existence of a **lock token** indicating if the resource is locked.
Optional available fields help you customize you resources with additional information:
- Resource description
- Lock description
- Endpoint
- Tags
## How to install and run
Clone the repository from GitLab or GitHub
```commandline
git clone git@gitlab.com:cpoisson/rent-a-bot.git
```
```commandline
git clone git@github.com:cpoisson/rent-a-bot.git
```
Create a virtual env (here using virtualenv wrapper)
```commandline
mkvirtualenv rent-a-bot
workon rent-a-bot
```
Install the package
```commandline
pip install . # pip install -e . if you want to install it in editable mode
```
Add Flask environment variables
```commandline
export FLASK_APP=rentabot
export FLASK_DEBUG=true # If you need the debug mode
```
And... run!
```commandline
flask run
```
## How to use it
Alright, rent-a-bot is up and running.
At this stage you can connect to the front end at http://127.0.0.1:5000/ (assuming your flask app listen to the port 500)
You will notice that the resource list is empty (dang...), let's populate it
### Populate the database
You will need a resource descriptor file to populate the database at startup.
```commandline
RENTABOT_RESOURCE_DESCRIPTOR="/absolute/path/to/your/resource/descriptor.yml"
```
### Resource descriptor
The resource descriptor is a YAML file. It's purpose is to declare the resources you want to make available on rent-a-bot
```yaml
# Resources Description
# This file describes resources to populate in the database at rent-a-bot startup
coffee-machine:
description: "Kitchen coffee machine"
endpoint: "tcp://192.168.1.50"
tags: "coffee kitchen food"
3d-printer-1:
description: "Basement 3d printer 1"
endpoint: "tcp://192.168.1.60"
tags: "3d-printer basement tool"
another-resource:
description: "yet another resource"
endpoint: ""
tags: ""
```
Once set, (re)start the flask application. The web view should be populated with your resources.
### RestFul API
#### List resources
GET /api/v1.0/resources
e.g.
```commandline
curl -X GET -i http://localhost:5000/rentabot/api/v1.0/resources
```
#### Access to a given resource
GET /api/v1.0/resources/{resource_id}
e.g.
```commandline
curl -X GET -i http://localhost:5000/rentabot/api/v1.0/resources/2
```
#### Lock a resource
POST /api/v1.0/resources/{resource_id}/lock
e.g.
```commandline
curl -X POST -i http://localhost:5000/rentabot/api/v1.0/resources/6/lock
```
**Note:** If the resource is available, a lock-token will be returned. Otherwise an error code is returned.
### Lock a resource using it's resource id (rid), name (name) or tag (tag).
POST /api/v1.0/resources/lock
e.g.
```commandline
curl -X POST -i http://localhost:5000/rentabot/api/v1.0/resources/lock\?rid\=6
curl -X POST -i http://localhost:5000/rentabot/api/v1.0/resources/lock\?name\=coffee-maker
curl -X POST -i http://localhost:5000/rentabot/api/v1.0/resources/lock\?tag\=coffee\&tag\=kitchen
```
**Notes:**
- If multiple available resources it the criteria, the first available will be returned.
- If criteria types are exclusive, resource id is prioritize over the name and tags, name is prioritize over tags.
#### Unlock a resource
POST /api/v1.0/resources/{resource_id}/unlock?lock-token={resource/lock/token}
```commandline
curl -X POST -i http://localhost:5000/rentabot/api/v1.0/resources/6/unlock\?lock-token\={resource/lock/token}
```
**Note:** If the resource is already unlocked or the lock-token is not valid, an error code is returned.
## How to tests
### Tests implementation
Unit tests are done using py.test and coverage
### How to run unit tests
```commandline
python setup.py test
```
---
## Helpful documentation used to design this application
- [Designing a RESTful API with Python and Flask](https://blog.miguelgrinberg.com/post/designing-a-restful-api-with-python-and-flask)
- [Testing Flask Applications](http://flask.pocoo.org/docs/0.12/testing/#testing)
- [Flask Project Template](https://github.com/xen/flask-project-template)
- [Flask SQLAlchemy](http://flask-sqlalchemy.pocoo.org/2.1/quickstart/)
- [Put versus Post](https://knpuniversity.com/screencast/rest/put-versus-post)
- [Best practice for a pragmatic restful API](http://www.vinaysahni.com/best-practices-for-a-pragmatic-restful-api#ssl)
- [Implementing a RESTful Web API with Python & Flask](http://blog.luisrei.com/articles/flaskrest.html)
- [HTTP status code](https://restpatterns.mindtouch.us/HTTP_Status_Codes)
- [Implementing API Exceptions](http://flask.pocoo.org/docs/0.12/patterns/apierrors/)
- [The Hitchhiker's Guide To Python](http://docs.python-guide.org/en/latest/) | /rent-a-bot-0.1.0.tar.gz/rent-a-bot-0.1.0/README.md | 0.47658 | 0.911377 | README.md | pypi |
Rentswatch Scraper Framework
============================
This package provides an easy and maintenable way to build a
Rentswatch scraper. Rentswatch is a cross-borders investigation that collects data on flat rents in Europe. Its scrapers mainly focus on classified ads.
How to install
--------------
Install using ``pip``...
::
pip install rentswatch-scraper
How to use
----------
Let's take a look at a quick example of using Rentswatch Scraper to
build a simple model-backed scraper to collect data from a website.
First, import the package components to build your scraper:
.. code:: python
#!/usr/bin/env python
from rentswatch_scraper.scraper import Scraper
from rentswatch_scraper.browser import geocode, convert
from rentswatch_scraper.fields import RegexField, ComputedField
from rentswatch_scraper import reporting
To factorize as much code as possible we created an abstract class that
every scraper will implement. For the sake of simplicity we'll use a
*dummy website* as follow:
.. code:: python
class DummyScraper(Scraper):
# Those are the basic meta-properties that define the scraper behavior
class Meta:
country = 'FR'
site = "dummy"
baseUrl = 'http://dummy.io'
listUrl = baseUrl + '/rent/city/paris/list.php'
adBlockSelector = '.ad-page-link'
Without any further configuration, this scraper will start to collect
ads from the list page of ``dummy.io``. To find links to the ads, it
will use the CSS selector ``.ad-page-link`` to get ``<a>`` markups and
follow their ``href`` attributes.
We have now to teach the scraper how to extract key figures from the ad
page.
.. code:: python
class DummyScraper(Scraper):
# HEADS UP: Meta declarations are hidden here
# ...
# ...
# Extract data using a CSS Selector.
realtorName = RegexField('.realtor-title')
# Extract data using a CSS Selector and a Regex.
serviceCharge = RegexField('.description-list', 'charges : (.*)\s€')
# Extract data using a CSS Selector and a Regex.
# This will throw a custom exception if the field is missing.
livingSpace = RegexField('.description-list', 'surface :(\d*)', required=True, exception=reporting.SpaceMissingError)
# Extract the value directly, without using a Regex
totalRent = RegexField('.description-price', required=True, exception=reporting.RentMissingError)
# Store this value as a private property (begining with a underscore).
# It won't be saved in the database but it can be helpful as you we'll see.
_address = RegexField('.description-address')
Every attribute will be saved as an Ad's property, according to the Ad
model.
Some properties may not be extractable from the HTML. You may need to
use a custom function that received existing properties. For this reason
we created a second field type named ``ComputedField``. Since the
properties order of declaration is recorded, we can use previously
declared (and extracted) values to compute new ones.
.. code:: python
class DummyScraper(Scraper):
# ...
# ...
# Use existing properties `totalRent` and `livingSpace` as they were
# extracted before this one.
pricePerSqm = ComputedField(fn=lambda s, values: values["totalRent"] / values["livingSpace"])
# This full exemple uses private properties to find latitude and longitude.
# To do so we use a buid-in function named `convert` that transforms an
# address into a dictionary of coordinates.
_latLng = ComputedField(fn=lambda s, values: geocode(values['_address'], 'FRA') )
# Gets a the dictionary field we want.
latitude = ComputedField(fn=lambda s, values: values['_latLng']['lat'])
longitude = ComputedField(fn=lambda s, values: values['_latLng']['lng'])
All you need to do now is to create an instance of your class and run
the scraper.
.. code:: python
# When you script is executed directly
if __name__ == "__main__":
dummyScraper = DummyScraper()
dummyScraper.run()
API Doc
-------
``class`` Ad
~~~~~~~~~~~~
Attributes
^^^^^^^^^^
As seen above, every Ad attribute might be used as a Scraper attribute to declare which attribute extract.
+----------------------+--------------------------+---------------------------------------------------------------------------+
| Name | Type | Description |
+======================+==========================+===========================================================================+
| ``status`` | *String* | "listed" if needs more scraping, "scraped" if it's done |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``site`` | *String* | Name of the website |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``createdAt`` | *DateTime* | Date the ad was first scraped |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``siteId`` | *String* | The unique ID from the site where it's scrapped from |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``serviceCharge`` | *Float* | Extra costs (heating mostly) |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``baseRent`` | *Float* | Base costs (without heating) |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``totalRent`` | *Float* | Total cost |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``livingSpace`` | *Float* | Surface in square meters |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``pricePerSqm`` | *Float* | Price per square meter |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``furnished`` | *Bool* | True if the flat or house is furnished |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``realtor`` | *Bool* | True if realtor, n if rented by a physical person |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``realtorName`` | *Unicode* | The name of the realtor or person offering the flat |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``latitude`` | *Float* | Latitude |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``longitude`` | *Float* | Longitude |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``balcony`` | *Bool* | True if there is a balcony/terrasse |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``yearConstructed`` | *String* | The year the building was built |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``cellar`` | *Bool* | True if the flat comes with a cellar |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``parking`` | *Bool* | True if the flat comes with a parking or a garage |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``houseNumber`` | *String* | House Number in the street |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``street`` | *String* | Street name (incl. "street") |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``zipCode`` | *String* | ZIP code |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``city`` | *Unicode* | City |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``lift`` | *Bool* | True if a lift is present |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``typeOfFlat`` | *String* | Type of flat (no typology) |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``noRooms`` | *String* | Number of rooms |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``floor`` | *String* | Floor the flat is at |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``garden`` | *Bool* | True if there is a garden |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``barrierFree`` | *Bool* | True if the flat is wheelchair accessible |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``country`` | *String* | Country, 2 letter code |
+----------------------+--------------------------+---------------------------------------------------------------------------+
| ``sourceUrl`` | *String* | URL of the page |
+----------------------+--------------------------+---------------------------------------------------------------------------+
``class`` Scraper
~~~~~~~~~~~~~~~~~
Methods
^^^^^^^
The Scraper class defines a lot of method that we encourage you to
redefine in order to have the full control of your scraper behavior.
+----------------------+------------------------------------------------------------------------------------------------------+
| Name | Description |
+======================+======================================================================================================+
| ``extract_ad`` | Extract ads list from a page's soup. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``fail`` | Print out an error message. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``fetch_ad`` | Fetch a single ad page from the target website then create Ad instances by calling ``èxtract_ad``. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``fetch_series`` | Fetch a single list page from the target website then fetch an ad by calling ``fetch_ad``. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``find_ad_blocks`` | Extract ad block from a page list. Called within ``fetch_series``. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``get_ad_href`` | Extract a href attribute from an ad block. Called within ``fetch_series``. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``get_ad_id`` | Extract a siteId from an ad block. Called within ``fetch_series``. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``get_fields`` | Used internally to generate a list of property to extract from the ad. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``get_series`` | Fetch a list page from the target website. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``has_issue`` | True if we met issues with this ad before. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``is_scraped`` | True if we already scraped this ad before. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``ok`` | Print out an success message. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``prepare`` | Just before saving the values. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``run`` | Run the scrapper. |
+----------------------+------------------------------------------------------------------------------------------------------+
| ``transform_page`` | Transform HTML content of the series page before parsing it. |
+----------------------+------------------------------------------------------------------------------------------------------+
Start a migration
-----------------
Use Yoyo_:
::
yoyo new ./migrations -m "Your migration's description"
And apply it:
::
yoyo apply --database mysql://user:password@host/db ./migrations
.. _Yoyo: https://pypi.python.org/pypi/yoyo-migrations
| /rentswatch-scraper-1.0.1.tar.gz/rentswatch-scraper-1.0.1/README.rst | 0.844345 | 0.789498 | README.rst | pypi |
from random import randint
from .agents import agents
import requests
import re
import os
import socket
import time
import urllib2
import urllib
import json
# Download SocksiPy - A Python SOCKS client module. ( http://code.google.com/p/socksipy-branch/downloads/list )
import socks
ENV = os.environ.get('ENV', 'production')
# On wich port Tor's proxy is avalaible
TOR_PROXY_PORT = int( os.environ.get('TOR_PROXY_PORT', 9050) )
# Maximum throttle delay
THROTTLE_MAX = int( os.environ.get('THROTTLE_MAX', 3) )
def setup():
# Equal 0 if the TOR_PROXY_PORT is open
if socket.socket(socket.AF_INET, socket.SOCK_STREAM).connect_ex(('127.0.0.1', TOR_PROXY_PORT)) == 0:
# Tor must listen for SOCKS request on 9050
socks.setdefaultproxy(socks.PROXY_TYPE_SOCKS5, "127.0.0.1", TOR_PROXY_PORT)
socket.socket = socks.socksocket
# Notify the user
print 'Your requests are routed throught Tor'
# Ensure the timeout is limited to 20 seconds
socket.setdefaulttimeout(20)
# fetch_page sets up the scraper and the VPN, returns the contents of the page
def fetch_page(url, throttle=(ENV == 'production'), postdata = None):
if throttle: time.sleep(randint(0, THROTTLE_MAX))
request = urllib2.Request(url, postdata)
request.add_header('Cache-Control','max-age=0')
request.add_header('User-Agent', agents[randint(0,len(agents)-1)], )
request.add_header('Accept', 'application/json,text/plain,text/html,application/xhtml+xml,application/xml, */*')
response = urllib2.urlopen(request)
return response.read()
def regex(needle, haystk, transform=lambda a:a):
haystk = str(haystk) if type(haystk) is not str else haystk
match = re.search(r''+needle+'', haystk, re.MULTILINE|re.DOTALL|re.IGNORECASE)
res = match.group(1) if match != None else None
return transform(res)
def booleanize(val, null=None):
if type(val) is bool:
return val
elif type(val) is int:
return bool(val)
elif val == 'n':
return False
elif val == 'y':
return True
else:
return null
def convert(amount, from_currency):
currencies = {
"DKK": 0.13,
"CZK": 0.037,
"PLN": 0.24,
"GBP": 1.40,
"SEK": 0.11,
"CHF": 0.920587,
"HUF": 0.0032,
"BRL": 0.25,
"VND": 0.000039,
"EUR": 1
}
try:
amount = float(amount)
except (ValueError, TypeError):
print "Invalid amount"
return False
return amount * currencies[from_currency]
def clean_amounts(totalRent=0, baseRent=0, serviceCharge=0, livingSpace=None, currency='EUR'):
baseRent = float(baseRent)
totalRent = float(totalRent)
livingSpace = float(livingSpace)
try:
serviceCharge = float(serviceCharge)
except (ValueError, TypeError):
serviceCharge = 0
# Computes totalRent and baseRent if they're missing
if totalRent == 0: totalRent = baseRent + serviceCharge
if baseRent == 0: baseRent = totalRent - serviceCharge
# Checks for bogus ads
if totalRent < 30 or totalRent > 25000 or livingSpace < 3 or livingSpace > 1000:
return False
if serviceCharge == 0:
serviceCharge = None
return {
"baseRent": convert(baseRent, currency),
"baseRentOriginalCurrency": baseRent,
"serviceCharge": convert(serviceCharge, currency),
"serviceChargeOriginalCurrency": serviceCharge,
"totalRent": convert(totalRent, currency),
"totalRentOriginalCurrency": totalRent,
"livingSpace": livingSpace,
"pricePerSqm": convert(baseRent, currency) / livingSpace,
"currency": currency
}
def geocode(address, country):
geocoder = "http://nominatim.openstreetmap.org/search?"
geocoder += "format=json&"
geocoder += "limit=10&"
geocoder += "bounded=1&"
geocoder += "osm_type=N&"
geocoder += "countrycodes=%s&" % country
geocoder += "q=%s&" % urllib.quote_plus(address)
page = fetch_page(geocoder)
data = json.loads(page)
if len(data) > 0:
return {
"lat" : float(data[0]["lat"]),
"lng" : float(data[0]["lon"])
}
else:
return {
"lat" : None,
"lng" : None
}
def convert_in_meters(string, separator):
if (separator in string):
feet = string.split(separator)[0]
inches = string.split(separator)[1]
inches = inches.replace(" ", "")
if inches != "" and inches is not None:
inches = float(inches) * 0.025
else:
inches = 0
return float(feet) * 0.305 + inches
else:
feet = string
return float(feet)
def convert_in_sqm(string):
return float(string) * 0.092903
def find_living_space(desc, no_rooms):
patterns = [
{
# Looks for rooms in feet under the format 5'10" x 3'1"
"format_finder_expression": "\d{1,2}'[ ]*\d{1,2}\" x \d{1,2}'[ ]*\d{1,2}\"",
"expression": "(\d{1,2}'[ ]*\d{0,2})(?:\")*(?: max)* x (\d{1,2}'[ ]*\d{0,2})(?:\")*",
"expect" : "several" # several rooms to add up
},
{
# Looks for rooms in feet under the format 10'2 x 9'3
"format_finder_expression": "\d{1,2}'\d{1,2} x \d{1,2}'\d{1,2}(?!\")",
"expression": "(\d{1,2}'\d{0,2})(?: max)* x (\d{1,2}'\d{0,2})",
"expect" : "several"
},
{
# Looks for "646 sq. Ft."
"format_finder_expression": "sq\. ft\.",
"expression": "(\d{3,4})[ ]*sq\. ft\.",
"expect": "one"
}
]
# Removes conversions in meters of the format " (3.81m)"
expression_to_remove = "( \(\d{1,2}.\d{1,2}m\))"
q = re.compile(expression_to_remove)
for match in q.finditer(desc):
desc = desc.replace(match.group(1), "")
# lower cases
desc = desc.lower()
for pattern in patterns:
format_finder_expression = pattern["format_finder_expression"]
if (re.search(format_finder_expression, desc) is not None):
expression = pattern["expression"]
p = re.compile(expression)
living_space = 0
if pattern["expect"] == "one":
match = re.search(expression, desc)
living_space = convert_in_sqm(match.group(1))
elif pattern["expect"] == "several":
for match in p.finditer(desc):
living_space += convert_in_meters(match.group(1), "'") * convert_in_meters(match.group(2), "'")
if (no_rooms != "NULL" and no_rooms != "None"):
if living_space/float(no_rooms) < 12 or living_space/float(no_rooms) > 40:
return None
else:
return living_space | /rentswatch-scraper-1.0.1.tar.gz/rentswatch-scraper-1.0.1/rentswatch_scraper/browser.py | 0.462959 | 0.159185 | browser.py | pypi |
from datetime import datetime
from functools import lru_cache
from typing import Any, Dict, List, Optional, Union, Type, cast
import numpy as np
import pandas as pd
import trimesh
from renumics.spotlight.dtypes import (
Audio,
Category,
Embedding,
Image,
Mesh,
Sequence1D,
Video,
Window,
)
from renumics.spotlight.dtypes.exceptions import InvalidFile, NotADType
from renumics.spotlight.dtypes.typing import (
ColumnType,
ColumnTypeMapping,
is_array_based_column_type,
is_file_based_column_type,
is_scalar_column_type,
)
from renumics.spotlight.io.pandas import (
infer_dtype,
is_empty,
prepare_column,
prepare_hugging_face_dict,
stringify_columns,
to_categorical,
try_literal_eval,
)
from renumics.spotlight.backend import datasource
from renumics.spotlight.backend.data_source import (
Column,
DataSource,
)
from renumics.spotlight.backend.exceptions import (
ConversionFailed,
DatasetColumnsNotUnique,
)
from renumics.spotlight.typing import PathType, is_pathtype
from renumics.spotlight.dataset.exceptions import ColumnNotExistsError
from renumics.spotlight.dtypes.conversion import read_external_value
@datasource(pd.DataFrame)
@datasource(".csv")
class PandasDataSource(DataSource):
"""
access pandas DataFrame table data
"""
_generation_id: int
_uid: str
_df: pd.DataFrame
def __init__(self, source: Union[PathType, pd.DataFrame]):
if is_pathtype(source):
df = pd.read_csv(source)
else:
df = cast(pd.DataFrame, source)
if not df.columns.is_unique:
raise DatasetColumnsNotUnique()
self._generation_id = 0
self._uid = str(id(df))
self._df = df.copy()
@property
def column_names(self) -> List[str]:
return stringify_columns(self._df)
@property
def df(self) -> pd.DataFrame:
"""
Get **a copy** of the served `DataFrame`.
"""
return self._df.copy()
def __len__(self) -> int:
return len(self._df)
def guess_dtypes(self) -> ColumnTypeMapping:
dtype_map = {
str(column_name): infer_dtype(self.df[column_name])
for column_name in self.df
}
return dtype_map
def _parse_column_index(self, column_name: str) -> Any:
column_names = self.column_names
try:
loc = self._df.columns.get_loc(column_name)
except KeyError:
...
else:
if isinstance(self._df.columns, pd.MultiIndex):
return self._df.columns[loc][0]
return self._df.columns[loc]
try:
index = column_names.index(column_name)
except ValueError as e:
raise ColumnNotExistsError(
f"Column '{column_name}' doesn't exist in the dataset."
) from e
return self._df.columns[index]
def get_generation_id(self) -> int:
return self._generation_id
def get_uid(self) -> str:
return self._uid
def get_name(self) -> str:
return "pd.DataFrame"
def get_column(
self,
column_name: str,
dtype: Type[ColumnType],
indices: Optional[List[int]] = None,
simple: bool = False,
) -> Column:
column_index = self._parse_column_index(column_name)
column = self._df[column_index]
if indices is not None:
column = column.iloc[indices]
column = prepare_column(column, dtype)
categories = None
embedding_length = None
if dtype is Category:
# `NaN` category is listed neither in `pandas`, not in our format.
categories = {
category: i for i, category in enumerate(column.cat.categories)
}
column = column.cat.codes
values = column.to_numpy()
elif dtype is datetime:
# We expect datetimes as ISO strings; empty strings instead of `NaT`s.
column = column.dt.strftime("%Y-%m-%dT%H:%M:%S.%f%z")
column = column.mask(column.isna(), "")
values = column.to_numpy()
elif dtype is str:
# Replace `NA`s with empty strings.
column = column.mask(column.isna(), "")
values = column.to_numpy()
if simple:
values = np.array(
[
value[:97] + "..." if len(value) > 100 else value
for value in values
]
)
elif is_scalar_column_type(dtype):
values = column.to_numpy()
elif dtype is Window:
# Replace all `NA` values with `[NaN, NaN]`.
na_mask = column.isna()
column.iloc[na_mask] = pd.Series(
[[float("nan"), float("nan")]] * na_mask.sum()
)
# This will fail, if arrays in column cells aren't aligned.
try:
values = np.asarray(column.to_list(), dtype=float)
except ValueError as e:
raise ValueError(
f"For the window column '{column_name}', column cells "
f"should be sequences of shape (2,) or `NA`s, but "
f"unaligned sequences received."
) from e
if values.ndim != 2 or values.shape[1] != 2:
raise ValueError(
f"For the window column '{column_name}', column cells "
f"should be sequences of shape (2,) or `NA`s, but "
f"sequences of shape {values.shape[1:]} received."
)
elif dtype is Embedding:
na_mask = column.isna()
# This will fail, if arrays in column cells aren't aligned.
try:
embeddings = np.asarray(column[~na_mask].to_list(), dtype=float)
except ValueError as e:
raise ValueError(
f"For the embedding column '{column_name}', column cells "
f"should be sequences of the same shape (n,) or `NA`s, but "
f"unaligned sequences received."
) from e
if embeddings.ndim != 2 or embeddings.shape[1] == 0:
raise ValueError(
f"For the embedding column '{column_name}', column cells "
f"should be sequences of the same shape (n,) or `NA`s, but "
f"sequences of shape {embeddings.shape[1:]} received."
)
values = np.empty(len(column), dtype=object)
if simple:
values[~na_mask] = "[...]"
else:
values[np.where(~na_mask)[0]] = list(embeddings)
embedding_length = embeddings.shape[1]
elif dtype is Sequence1D:
na_mask = column.isna()
values = np.empty(len(column), dtype=object)
values[~na_mask] = "[...]"
elif dtype is np.ndarray:
na_mask = column.isna()
values = np.empty(len(column), dtype=object)
values[~na_mask] = "[...]"
elif is_file_based_column_type(dtype):
# Strings are paths or URLs, let them inplace. Replace
# non-strings with "<in-memory>".
na_mask = column.isna()
column = column.mask(~(column.map(type) == str), "<in-memory>")
values = column.to_numpy(dtype=object)
values[na_mask] = None
else:
raise NotADType()
return Column(
type=dtype,
order=None,
hidden=column_name.startswith("_"),
optional=True,
description=None,
tags=[],
editable=dtype in (bool, int, float, str, Category, Window),
categories=categories,
x_label=None,
y_label=None,
embedding_length=embedding_length,
name=column_name,
values=values,
)
def get_cell_data(
self, column_name: str, row_index: int, dtype: Type[ColumnType]
) -> Any:
"""
Return the value of a single cell, warn if not possible.
"""
self._assert_index_exists(row_index)
column_index = self._parse_column_index(column_name)
raw_value = self._df.iloc[row_index, self._df.columns.get_loc(column_index)]
if dtype is Category:
if pd.isna(raw_value):
return -1
categories = self._get_column_categories(column_index, as_string=True)
return categories.get(raw_value, -1)
if dtype is datetime:
value = pd.to_datetime(raw_value).to_numpy("datetime64[us]").tolist()
# `tolist()` returns `None` for all `NaT`s, `datetime` otherwise.
if value is None:
return ""
return value.isoformat()
if dtype is str:
if pd.isna(raw_value):
return ""
return str(raw_value)
if is_scalar_column_type(dtype):
# `dtype` is `bool`, `int` or `float`.
dtype = cast(Type[Union[bool, int, float]], dtype)
try:
return dtype(raw_value)
except (TypeError, ValueError) as e:
raise ConversionFailed(dtype, raw_value) from e
if dtype is np.ndarray:
if isinstance(raw_value, str):
raw_value = try_literal_eval(raw_value)
if is_empty(raw_value):
return None
return np.asarray(raw_value)
if dtype is Window:
if isinstance(raw_value, str):
raw_value = try_literal_eval(raw_value)
if is_empty(raw_value):
return np.full(2, np.nan)
try:
value = np.asarray(raw_value, dtype=float)
except (TypeError, ValueError) as e:
raise ConversionFailed(dtype, raw_value) from e
if value.ndim != 1 or len(value) != 2:
raise ValueError(
f"Window column cells should be sequences of shape (2,), "
f"but a sequence of shape {value.shape} received for the "
f"column '{column_name}'."
)
if is_empty(raw_value):
return None
if isinstance(raw_value, str):
raw_value = try_literal_eval(raw_value)
if is_file_based_column_type(dtype) and isinstance(raw_value, dict):
raw_value = prepare_hugging_face_dict(raw_value)
if isinstance(raw_value, trimesh.Trimesh) and dtype is Mesh:
value = Mesh.from_trimesh(raw_value)
return value.encode()
if isinstance(raw_value, str) and is_file_based_column_type(dtype):
try:
return read_external_value(str(raw_value), dtype)
except Exception as e:
raise ConversionFailed(dtype, raw_value) from e
if isinstance(raw_value, bytes) and dtype in (Audio, Image, Video):
try:
value = dtype.from_bytes(raw_value) # type: ignore
except Exception as e:
raise ConversionFailed(dtype, raw_value) from e
return value.encode()
if not isinstance(raw_value, dtype) and is_array_based_column_type(dtype):
try:
value = dtype(raw_value)
except (InvalidFile, TypeError, ValueError) as e:
raise ConversionFailed(dtype, raw_value) from e
return value.encode()
if isinstance(raw_value, dtype):
return raw_value.encode()
raise ConversionFailed(dtype, raw_value)
def _get_default_value(self, dtype: Type[ColumnType]) -> Any:
if dtype is int:
return 0
if dtype is bool:
return False
if dtype is str:
return ""
if dtype is Window:
return [float("nan"), float("nan")]
# `dtype` is `float`, `datetime`, `np.ndarray`, or a custom data type.
return np.nan
@lru_cache(maxsize=128)
def _get_column_categories(
self, column_index: Any, as_string: bool = False
) -> Dict[str, int]:
"""
Get categories of a categorical column.
If `as_string` is True, convert the categories to their string
representation.
At the moment, there is no way to add a new category in Spotlight, so we
rely on the previously cached ones.
"""
column = self._df[column_index]
column = to_categorical(column, str_categories=as_string)
return {category: i for i, category in enumerate(column.cat.categories)} | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight_plugins/core/pandas_data_source.py | 0.851104 | 0.328489 | pandas_data_source.py | pypi |
from functools import lru_cache
from hashlib import sha1
from pathlib import Path
from typing import Any, Dict, List, Optional, cast, Union, Type, Tuple
from dataclasses import asdict
import h5py
import numpy as np
from renumics.spotlight.dtypes import Category, Embedding
from renumics.spotlight.dtypes.typing import (
ColumnType,
ColumnTypeMapping,
get_column_type,
)
from renumics.spotlight.typing import PathType, IndexType
from renumics.spotlight.dataset import (
Dataset,
INTERNAL_COLUMN_NAMES,
unescape_dataset_name,
)
from renumics.spotlight.backend.data_source import DataSource, Attrs, Column
from renumics.spotlight.backend.exceptions import (
NoTableFileFound,
CouldNotOpenTableFile,
NoRowFound,
)
from renumics.spotlight.backend import datasource
from renumics.spotlight.dtypes.conversion import (
DTypeOptions,
NormalizedValue,
convert_to_dtype,
)
def unescape_dataset_names(refs: np.ndarray) -> np.ndarray:
"""
Unescape multiple dataset names.
"""
return np.array([unescape_dataset_name(value) for value in refs])
def _decode_attrs(raw_attrs: h5py.AttributeManager) -> Tuple[Attrs, bool, bool]:
"""
Get relevant subset of column attributes.
"""
column_type_name = raw_attrs.get("type", "unknown")
column_type = get_column_type(column_type_name)
categories: Optional[Dict[str, int]] = None
embedding_length: Optional[int] = None
if column_type is Category:
# If one of the attributes does not exist or is empty, an empty dict
# will be created.
categories = dict(
zip(
raw_attrs.get("category_keys", []),
raw_attrs.get("category_values", []),
)
)
elif column_type is Embedding:
embedding_length = raw_attrs.get("value_shape", [0])[0]
tags = raw_attrs.get("tags", np.array([])).tolist()
has_lookup = "lookup_keys" in raw_attrs
is_external = raw_attrs.get("external", False)
return (
Attrs(
type=column_type,
order=raw_attrs.get("order", None),
hidden=raw_attrs.get("hidden", False),
optional=raw_attrs.get("optional", False),
description=raw_attrs.get("description", None),
tags=tags,
editable=raw_attrs.get("editable", False),
categories=categories,
x_label=raw_attrs.get("x_label", None),
y_label=raw_attrs.get("y_label", None),
embedding_length=embedding_length,
),
has_lookup,
is_external,
)
def ref_placeholder_names(mask: np.ndarray) -> np.ndarray:
"""
Generate placeholder names for a ref column based of the given mask.
"""
return np.array(["..." if x else None for x in mask], dtype=object)
class H5Dataset(Dataset):
"""
A `spotlight.Dataset` class extension for better usage in Spotlight backend.
"""
def __enter__(self) -> "H5Dataset":
self.open()
return self
def get_generation_id(self) -> int:
"""
Get the dataset's generation if set.
"""
return int(self._h5_file.attrs.get("spotlight_generation_id", 0))
def read_value(self, column_name: str, index: IndexType) -> NormalizedValue:
"""
Get a dataset value as it is stored in the H5 dataset, resolve references.
"""
self._assert_column_exists(column_name, internal=True)
self._assert_index_exists(index)
column = cast(h5py.Dataset, self._h5_file[column_name])
value = column[index]
if isinstance(value, bytes):
value = value.decode("utf-8")
if self._is_ref_column(column):
if value:
value = self._resolve_ref(value, column_name)[()]
return value.tolist() if isinstance(value, np.void) else value
return None
if self._get_column_type(column.attrs) is Embedding and len(value) == 0:
return None
return value
def read_column(
self, column_name: str, indices: Optional[List[int]] = None
) -> Union[list, np.ndarray]:
"""
Get a decoded dataset column.
"""
self._assert_column_exists(column_name, internal=True)
column = cast(h5py.Dataset, self._h5_file[column_name])
is_string_dtype = h5py.check_string_dtype(column.dtype)
raw_values: np.ndarray
if indices is None:
raw_values = column[:]
else:
raw_values = column[indices]
if is_string_dtype:
raw_values = np.array([x.decode("utf-8") for x in raw_values])
if self._is_ref_column(column):
assert is_string_dtype, "Only new-style string h5 references supported."
return [
value.tolist() if isinstance(value, np.void) else value
for value in self._resolve_refs(raw_values, column_name)
]
if self._get_column_type(column.attrs) is Embedding:
return [None if len(x) == 0 else x for x in raw_values]
return raw_values
def duplicate_row(self, from_index: IndexType, to_index: IndexType) -> None:
"""
Duplicate a dataset's row. Increases the dataset's length by 1.
"""
self._assert_is_writable()
self._assert_index_exists(from_index)
length = self._length
if from_index < 0:
from_index += length
if to_index < 0:
to_index += length
if to_index != length:
self._assert_index_exists(to_index)
for column_name in self.keys() + INTERNAL_COLUMN_NAMES:
column = cast(h5py.Dataset, self._h5_file[column_name])
column.resize(length + 1, axis=0)
if to_index != length:
# Shift all values after the insertion position by one.
raw_values = column[int(to_index) : -1]
if self._get_column_type(column) is Embedding:
raw_values = list(raw_values)
column[int(to_index) + 1 :] = raw_values
column[int(to_index)] = column[from_index]
self._length += 1
self._update_generation_id()
def min_order(self) -> int:
"""
Get minimum order over all columns, return 0 if no column has an order.
One can use `dataset.min_order() - 1` as order for a new column.
"""
return int(
min(
(
self._h5_file[name].attrs.get("order", 0)
for name in self._column_names
),
default=0,
)
)
def _resolve_refs(self, refs: np.ndarray, column_name: str) -> np.ndarray:
raw_values = np.empty(len(refs), dtype=object)
raw_values[:] = [
self._resolve_ref(ref, column_name)[()] if ref else None for ref in refs
]
return raw_values
@datasource(".h5")
class Hdf5DataSource(DataSource):
"""
access h5 table data
"""
def __init__(self, source: PathType):
self._table_file = Path(source)
@property
def column_names(self) -> List[str]:
with self._open_table() as dataset:
return dataset.keys()
def __len__(self) -> int:
with self._open_table() as dataset:
return len(dataset)
def guess_dtypes(self) -> ColumnTypeMapping:
with self._open_table() as dataset:
return {
column_name: dataset.get_column_type(column_name)
for column_name in self.column_names
}
def get_generation_id(self) -> int:
with self._open_table() as dataset:
return dataset.get_generation_id()
def get_uid(self) -> str:
return sha1(str(self._table_file.absolute()).encode("utf-8")).hexdigest()
def get_name(self) -> str:
return str(self._table_file.name)
def get_internal_columns(self) -> List[Column]:
with self._open_table() as dataset:
return [
self.get_column(column_name, dataset.get_column_type(column_name))
for column_name in INTERNAL_COLUMN_NAMES
]
def get_column(
self,
column_name: str,
dtype: Type[ColumnType],
indices: Optional[List[int]] = None,
simple: bool = False,
) -> Column:
with self._open_table() as dataset:
normalized_values = dataset.read_column(column_name, indices=indices)
if dtype is Category:
categories = self._get_column_categories(column_name)
values = [
convert_to_dtype(
value, dtype, DTypeOptions(categories=categories), simple
)
for value in normalized_values
]
else:
values = [
convert_to_dtype(value, dtype, simple=simple)
for value in normalized_values
]
attrs, _, _ = _decode_attrs(dataset._h5_file[column_name].attrs)
attrs.type = dtype
return Column(name=column_name, values=values, **asdict(attrs))
def get_cell_data(
self, column_name: str, row_index: int, dtype: Type[ColumnType]
) -> Any:
"""
return the value of a single cell
"""
# read raw value from h5 table
with self._open_table() as dataset:
try:
normalized_value = dataset.read_value(column_name, row_index)
except IndexError as e:
raise NoRowFound(row_index) from e
# convert normalized value to requested dtype
if dtype is Category:
categories = self._get_column_categories(column_name)
return convert_to_dtype(
normalized_value, dtype, DTypeOptions(categories=categories)
)
return convert_to_dtype(normalized_value, dtype)
@lru_cache(maxsize=128)
def _get_column_categories(self, column_name: str) -> Dict[str, int]:
with self._open_table() as dataset:
attrs = dataset.get_column_attributes(column_name)
try:
return cast(Dict[str, int], attrs["categories"])
except KeyError:
normalized_values = cast(
List[str],
[
convert_to_dtype(value, str, simple=True)
for value in dataset.read_column(column_name)
],
)
category_names = sorted(set(normalized_values))
return {
category_name: i for i, category_name in enumerate(category_names)
}
def _open_table(self, mode: str = "r") -> H5Dataset:
try:
return H5Dataset(self._table_file, mode)
except FileNotFoundError as e:
raise NoTableFileFound(self._table_file) from e
except OSError as e:
raise CouldNotOpenTableFile(self._table_file) from e | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight_plugins/core/hdf5_data_source.py | 0.860486 | 0.47171 | hdf5_data_source.py | pypi |
import importlib
import pkgutil
from dataclasses import dataclass
from types import ModuleType
from pathlib import Path
from typing import Callable, List, Optional
from fastapi import FastAPI
from renumics.spotlight.settings import settings
from renumics.spotlight.develop.project import get_project_info
from renumics.spotlight.io.path import is_path_relative_to
import renumics.spotlight_plugins as plugins_namespace
@dataclass
class Plugin:
"""
Information about an installed and loaded Spotlight Plugin
"""
name: str
priority: int
module: ModuleType
init: Callable[[], None]
activate: Callable[[FastAPI], None]
dev: bool
frontend_entrypoint: Optional[Path]
_plugins: Optional[List[Plugin]] = None
def load_plugins() -> List[Plugin]:
"""
Automatically load, register and initialize plugins
inside the renumics.spotlight.plugins namespace package.
"""
global _plugins
if _plugins is not None:
return _plugins
def noinit() -> None:
"""
noop impl for __init__
"""
def noactivate(_: FastAPI) -> None:
"""
noop impl for __activate__
"""
plugins = {}
for _, name, _ in pkgutil.iter_modules(plugins_namespace.__path__):
module = importlib.import_module(plugins_namespace.__name__ + "." + name)
project = get_project_info()
dev = bool(
settings.dev
and project.root
and is_path_relative_to(module.__path__[0], project.root)
)
main_js = Path(module.__path__[0]) / "frontend" / "main.js"
plugins[name] = Plugin(
name=name,
priority=getattr(module, "__priority__", 1000),
init=getattr(module, "__register__", noinit),
activate=getattr(module, "__activate__", noactivate),
module=module,
dev=dev,
frontend_entrypoint=main_js if main_js.exists() else None,
)
_plugins = sorted(plugins.values(), key=lambda p: p.priority)
for plugin in _plugins:
plugin.init()
return _plugins | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/plugin_loader.py | 0.780412 | 0.186521 | plugin_loader.py | pypi |
import os
import platform
import signal
import sys
from typing import Optional, Tuple, Union
from pathlib import Path
import click
from renumics import spotlight
from renumics.spotlight.dtypes.typing import COLUMN_TYPES_BY_NAME, ColumnTypeMapping
from renumics.spotlight import logging
def cli_dtype_callback(
_ctx: click.Context, _param: click.Option, value: Tuple[str, ...]
) -> Optional[ColumnTypeMapping]:
"""
Parse column types from multiple strings in format
`COLUMN_NAME=DTYPE` to a dict.
"""
if not value:
return None
dtype = {}
for mapping in value:
try:
column_name, dtype_name = mapping.split("=")
except ValueError as e:
raise click.BadParameter(
"Column type setting separator '=' not specified or specified "
"more than once."
) from e
try:
column_type = COLUMN_TYPES_BY_NAME[dtype_name]
except KeyError as e:
raise click.BadParameter(
f"Column types from {list(COLUMN_TYPES_BY_NAME.keys())} "
f"expected, but value '{dtype_name}' recived."
) from e
dtype[column_name] = column_type
return dtype
@click.command() # type: ignore
@click.argument(
"table-or-folder",
type=str,
required=False,
default=os.environ.get("SPOTLIGHT_TABLE_FILE", str(Path.cwd())),
)
@click.option(
"--host",
"-h",
default="localhost",
help="The host that Spotlight should listen on.",
show_default=True,
)
@click.option(
"--port",
"-p",
type=str,
default="auto",
help="The port that Spotlight should listen on (use 'auto' to use a random free port)",
show_default=True,
)
@click.option(
"--layout",
default=None,
help="Preconfigured layout to use as default.",
)
@click.option(
"--dtype",
type=click.UNPROCESSED,
callback=cli_dtype_callback,
multiple=True,
help="Custom column types setting (use COLUMN_NAME={"
+ "|".join(sorted(COLUMN_TYPES_BY_NAME.keys()))
+ "} notation). Multiple settings allowed.",
)
@click.option(
"--no-browser",
is_flag=True,
default=False,
help="Do not automatically show Spotlight in browser.",
)
@click.option(
"--filebrowsing/--no-filebrowsing",
is_flag=True,
default=True,
help="Whether to allow users to browse and open datasets.",
)
@click.option(
"--analyze",
is_flag=True,
default=False,
help="Automatically analyze common dataset errors.",
)
@click.option("-v", "--verbose", is_flag=True)
@click.version_option(spotlight.__version__)
def main(
table_or_folder: str,
host: str,
port: Union[int, str],
layout: Optional[str],
dtype: Optional[ColumnTypeMapping],
no_browser: bool,
filebrowsing: bool,
analyze: bool,
verbose: bool,
) -> None:
"""
Parse CLI arguments and launch Renumics Spotlight.
"""
if verbose:
logging.enable()
signal.signal(signal.SIGINT, lambda *_: sys.exit())
signal.signal(signal.SIGTERM, lambda *_: sys.exit())
if platform.system() != "Windows":
signal.signal(signal.SIGHUP, lambda *_: sys.exit())
spotlight.show(
table_or_folder,
dtype=dtype,
host=host,
port="auto" if port == "auto" else int(port),
layout=layout,
no_browser=no_browser,
allow_filebrowsing=filebrowsing,
wait="forever",
analyze=analyze,
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/cli.py | 0.499756 | 0.232495 | cli.py | pypi |
from datetime import datetime
from typing import (
List,
Optional,
Sequence,
Tuple,
Union,
)
import numpy as np
import trimesh
from typing_extensions import get_args
from renumics.spotlight.typing import BoolType, IntType, NumberType, PathOrUrlType
from renumics.spotlight.dtypes import (
Array1dLike,
Embedding,
Mesh,
Sequence1D,
Image,
ImageLike,
Audio,
Category,
Video,
Window,
)
from renumics.spotlight.dtypes.typing import FileBasedColumnType, NAME_BY_COLUMN_TYPE
# Only pure types.
SimpleColumnType = Union[bool, int, float, str, datetime, Category, Window, Embedding]
RefColumnType = Union[np.ndarray, Embedding, Mesh, Sequence1D, Image, Audio, Video]
ExternalColumnType = FileBasedColumnType
# Pure types, compatible types and `None`.
BoolColumnInputType = Optional[BoolType]
IntColumnInputType = Optional[IntType]
FloatColumnInputType = Optional[Union[float, np.floating]]
StringColumnInputType = Optional[str]
DatetimeColumnInputType = Optional[Union[datetime, np.datetime64]]
CategoricalColumnInputType = Optional[str]
WindowColumnInputType = Optional[
Union[List[NumberType], Tuple[NumberType, NumberType], np.ndarray]
]
ArrayColumnInputType = Optional[Union[np.ndarray, Sequence]]
EmbeddingColumnInputType = Optional[Union[Embedding, Array1dLike]]
AudioColumnInputType = Optional[Union[Audio, PathOrUrlType, bytes]]
ImageColumnInputType = Optional[Union[Image, ImageLike, PathOrUrlType, bytes]]
MeshColumnInputType = Optional[Union[Mesh, trimesh.Trimesh, PathOrUrlType]]
Sequence1DColumnInputType = Optional[Union[Sequence1D, Array1dLike]]
VideoColumnInputType = Optional[Union[Video, PathOrUrlType, bytes]]
# Aggregated input types.
SimpleColumnInputType = Union[
BoolColumnInputType,
IntColumnInputType,
FloatColumnInputType,
StringColumnInputType,
DatetimeColumnInputType,
CategoricalColumnInputType,
WindowColumnInputType,
EmbeddingColumnInputType,
]
RefColumnInputType = Union[
ArrayColumnInputType,
EmbeddingColumnInputType,
AudioColumnInputType,
ImageColumnInputType,
MeshColumnInputType,
Sequence1DColumnInputType,
VideoColumnInputType,
]
ColumnInputType = Union[SimpleColumnInputType, RefColumnInputType]
ExternalColumnInputType = Optional[PathOrUrlType]
REF_COLUMN_TYPE_NAMES = [
NAME_BY_COLUMN_TYPE[column_type] for column_type in get_args(RefColumnType)
]
SIMPLE_COLUMN_TYPE_NAMES = [
NAME_BY_COLUMN_TYPE[column_type] for column_type in get_args(SimpleColumnType)
] | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dataset/typing.py | 0.846546 | 0.387082 | typing.py | pypi |
import os
import shutil
import uuid
from datetime import datetime
from tempfile import TemporaryDirectory
from typing import (
Any,
Callable,
Dict,
Iterable,
List,
Optional,
overload,
Set,
Tuple,
Type,
Union,
cast,
)
import h5py
import numpy as np
import pandas as pd
import prettytable
import trimesh
import validators
from loguru import logger
from typing_extensions import Literal, TypeGuard
from renumics.spotlight.__version__ import __version__
from renumics.spotlight.io.pandas import (
infer_dtypes,
prepare_column,
is_string_mask,
stringify_columns,
)
from renumics.spotlight.typing import (
BoolType,
IndexType,
Indices1dType,
PathOrUrlType,
PathType,
is_integer,
is_iterable,
)
from renumics.spotlight.dtypes import (
Embedding,
Mesh,
Sequence1D,
Image,
Audio,
Category,
Video,
Window,
)
from renumics.spotlight.dtypes.base import DType, FileBasedDType
from renumics.spotlight.dtypes.typing import (
ColumnType,
ColumnTypeMapping,
FileBasedColumnType,
get_column_type,
get_column_type_name,
is_file_based_column_type,
)
from renumics.spotlight.dtypes.conversion import prepare_path_or_url
from . import exceptions
from .typing import (
REF_COLUMN_TYPE_NAMES,
SimpleColumnType,
RefColumnType,
ExternalColumnType,
BoolColumnInputType,
IntColumnInputType,
FloatColumnInputType,
StringColumnInputType,
DatetimeColumnInputType,
CategoricalColumnInputType,
WindowColumnInputType,
ArrayColumnInputType,
EmbeddingColumnInputType,
AudioColumnInputType,
ImageColumnInputType,
MeshColumnInputType,
Sequence1DColumnInputType,
VideoColumnInputType,
SimpleColumnInputType,
RefColumnInputType,
ColumnInputType,
)
INTERNAL_COLUMN_NAMES = ["__last_edited_by__", "__last_edited_at__"]
_EncodedColumnType = Optional[Union[bool, int, float, str, np.ndarray, h5py.Reference]]
def get_current_datetime() -> datetime:
"""
Get current datetime with timezone.
"""
return datetime.now().astimezone()
def escape_dataset_name(name: str) -> str:
r"""
Replace "\" with "\\" and "/" with "\s".
"""
return name.replace("\\", "\\\\").replace("/", r"\s")
def unescape_dataset_name(escaped_name: str) -> str:
r"""
Replace "\\" with "\" and "\s" with "/".
"""
name = ""
i = 0
while i < len(escaped_name):
char = escaped_name[i]
i += 1
if char == "\\":
next_char = escaped_name[i]
i += 1
if next_char == "\\":
name += "\\"
elif next_char == "s":
name += "/"
else:
raise RuntimeError
else:
name += char
return name
_ALLOWED_COLUMN_TYPES: Dict[Type[ColumnType], Tuple[Type, ...]] = {
bool: (np.bool_,),
int: (np.integer,),
float: (np.floating,),
datetime: (np.datetime64,),
}
_ALLOWED_COLUMN_DTYPES: Dict[Type[ColumnType], Tuple[Type, ...]] = {
bool: (np.bool_,),
int: (np.integer,),
float: (np.floating,),
datetime: (np.datetime64,),
Window: (np.floating,),
Embedding: (np.floating,),
}
def _check_valid_value_type(value: Any, column_type: Type[ColumnType]) -> bool:
"""
Check if a value is suitable for the given column type. Instances of the
given type are always suitable for its type, but extra types from
`_ALLOWED_COLUMN_TYPES` are also checked.
"""
allowed_types = (column_type,) + _ALLOWED_COLUMN_TYPES.get(column_type, ())
return isinstance(value, allowed_types)
def _check_valid_value_dtype(dtype: np.dtype, column_type: Type[ColumnType]) -> bool:
"""
Check if an array with the given dtype is suitable for the given column type.
Only types from `_ALLOWED_COLUMN_DTYPES` are checked. All other column types
are assumed to have no dtype equivalent.
"""
allowed_dtypes = _ALLOWED_COLUMN_DTYPES.get(column_type, ())
return any(np.issubdtype(dtype, allowed_dtype) for allowed_dtype in allowed_dtypes)
def _check_valid_array(
value: Any, column_type: Type[ColumnType]
) -> TypeGuard[np.ndarray]:
"""
Check if a value is an array and its type is suitable for the given column type.
"""
return isinstance(value, np.ndarray) and _check_valid_value_dtype(
value.dtype, column_type
)
class Dataset:
"""
Spotlight dataset.
"""
_filepath: str
_mode: str
_h5_file: h5py.File
_closed: bool
_column_names: Set[str]
_length: int
@staticmethod
def _user_column_attributes(column_type: Type[ColumnType]) -> Dict[str, Type]:
attribute_names = {
"order": int,
"hidden": bool,
"optional": bool,
"default": object,
"description": str,
"tags": list,
}
if column_type in {
bool,
int,
float,
str,
Category,
Window,
}:
attribute_names["editable"] = bool
if column_type is Category:
attribute_names["categories"] = dict
if column_type is Sequence1D:
attribute_names["x_label"] = str
attribute_names["y_label"] = str
if issubclass(column_type, FileBasedDType):
attribute_names["lookup"] = dict
attribute_names["external"] = bool
if column_type is Audio:
attribute_names["lossy"] = bool
return attribute_names
@classmethod
def _default_default(cls, column_type: Type[ColumnType]) -> Any:
if column_type is datetime:
return np.datetime64("NaT")
if column_type in (str, Category):
return ""
if column_type is float:
return float("nan")
if column_type is Window:
return [np.nan, np.nan]
if column_type is np.ndarray or issubclass(column_type, DType):
return None
raise exceptions.InvalidAttributeError(
f"`default` argument for optional column of type "
f"{get_column_type_name(column_type)} should be set, but `None` received."
)
def __init__(self, filepath: PathType, mode: str):
self._filepath = os.path.abspath(filepath)
self._check_mode(mode)
self._mode = mode
dirpath = os.path.dirname(self._filepath)
if self._mode in ("w", "w-", "x", "a"):
os.makedirs(dirpath, exist_ok=True)
elif not os.path.isdir(dirpath):
raise FileNotFoundError(f"File {filepath} does not exist.")
self._closed = True
self._column_names = set()
self._length = 0
@property
def filepath(self) -> str:
"""
Dataset file name.
"""
return self._filepath
@property
def mode(self) -> str:
"""
Dataset file open mode.
"""
return self._mode
def __enter__(self) -> "Dataset":
self.open()
return self
def __exit__(self, *args: Any) -> None:
self.close()
def __delitem__(self, item: Union[str, IndexType, Indices1dType]) -> None:
"""
Delete a dataset column or row.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bools", [True, False, False, True])
... dataset.append_int_column("ints", [-1, 0, 1, 2])
... dataset.append_float_column("floats", [-1.0, 0.0, 1.0, float("nan")])
... print(len(dataset))
... print(sorted(dataset.keys()))
4
['bools', 'floats', 'ints']
>>> with Dataset("docs/example.h5", "a") as dataset:
... del dataset[-1]
... print(len(dataset))
... print(sorted(dataset.keys()))
3
['bools', 'floats', 'ints']
>>> with Dataset("docs/example.h5", "a") as dataset:
... del dataset["bools"]
... del dataset["floats"]
... print(len(dataset))
... print(sorted(dataset.keys()))
3
['ints']
"""
self._assert_is_writable()
if isinstance(item, str):
self._assert_column_exists(item)
del self._h5_file[item]
try:
del self._h5_file[f"__group__/{item}"]
except KeyError:
pass
self._column_names.discard(item)
if not self._column_names:
self._length = 0
self._update_internal_columns()
elif isinstance(item, (slice, list, np.ndarray)):
mask = np.full(self._length, True)
try:
mask[item] = False
except Exception as e:
raise exceptions.InvalidIndexError(
f"Indices {item} of type `{type(item)}` do not match "
f"to the dataset with the length {self._length}."
) from e
keep_indices = np.nonzero(mask)[0]
length = len(keep_indices)
if length == self._length:
logger.warning(
"No rows removed because the given indices reference no elements."
)
return
start = (~mask).argmax() if length else 0
start = cast(int, start)
keep_indices = keep_indices[start:]
for column_name in self.keys() + INTERNAL_COLUMN_NAMES:
column = self._h5_file[column_name]
column[start:length] = column[keep_indices]
column.resize(length, axis=0)
self._length = length
elif is_integer(item):
self._assert_index_exists(item)
item = cast(int, item)
if item < 0:
item += self._length
for column_name in self.keys() + INTERNAL_COLUMN_NAMES:
column = self._h5_file[column_name]
raw_values = column[item + 1 :]
if self._get_column_type(column) is Embedding:
raw_values = list(raw_values)
column[item:-1] = raw_values
column.resize(self._length - 1, axis=0)
self._length -= 1
else:
raise exceptions.InvalidIndexError(
f"`item` argument should be a string or an index/indices, but"
f"value {item} of type `{type(item)}` received.`"
)
self._update_generation_id()
@overload
def __getitem__(
self, item: Union[str, Tuple[str, Indices1dType], Tuple[Indices1dType, str]]
) -> np.ndarray:
...
@overload
def __getitem__(self, item: IndexType) -> Dict[str, Optional[ColumnType]]:
...
@overload
def __getitem__(
self, item: Union[Tuple[str, IndexType], Tuple[IndexType, str]]
) -> Optional[ColumnType]:
...
def __getitem__(
self,
item: Union[
str,
IndexType,
Tuple[str, Union[IndexType, Indices1dType]],
Tuple[Union[IndexType, Indices1dType], str],
],
) -> Union[np.ndarray, Dict[str, Optional[ColumnType]], Optional[ColumnType],]:
"""
Get a dataset column, row or value.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bools", [True, False, False])
... dataset.append_int_column("ints", [-1, 0, 1])
... dataset.append_float_column("floats", [-1.0, 0.0, 1.0])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print("bools:", dataset["bools"])
... print("ints:", dataset["ints"])
... row = dataset[0]
... print("0th row:", [(key, row[key]) for key in sorted(row.keys())])
... row = dataset[1]
... print("1th row:", [(key, row[key]) for key in sorted(row.keys())])
... print(dataset["ints", 2])
... print(dataset[2, "floats"])
bools: [ True False False]
ints: [-1 0 1]
0th row: [('bools', True), ('floats', -1.0), ('ints', -1)]
1th row: [('bools', False), ('floats', 0.0), ('ints', 0)]
1
1.0
"""
self._assert_is_opened()
if is_integer(item):
self._assert_index_exists(item)
return {
column_name: self._get_value(self._h5_file[column_name], item)
for column_name in self._column_names
}
column_name, index = self._prepare_item(item) # type: ignore
self._assert_column_exists(column_name, internal=True)
column = self._h5_file[column_name]
if is_integer(index):
return self._get_value(column, index, check_index=True)
if index is None or isinstance(index, (slice, list, np.ndarray)):
return self._get_column(column, index)
raise exceptions.InvalidIndexError(
f"Invalid index {index} of type `{type(index)}` received."
)
@overload
def __setitem__(
self,
item: Union[str, Tuple[str, Indices1dType], Tuple[Indices1dType, str]],
value: Union[ColumnInputType, Iterable[ColumnInputType]],
) -> None:
...
@overload
def __setitem__(self, item: IndexType, value: Dict[str, ColumnInputType]) -> None:
...
@overload
def __setitem__(
self,
item: Union[Tuple[str, IndexType], Tuple[IndexType, str]],
value: ColumnInputType,
) -> None:
...
def __setitem__(
self,
item: Union[
str,
IndexType,
Tuple[str, Union[IndexType, Indices1dType]],
Tuple[Union[IndexType, Indices1dType], str],
],
value: Union[
ColumnInputType, Iterable[ColumnInputType], Dict[str, ColumnInputType]
],
) -> None:
"""
Set a dataset column, row or value.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bools", [True, False, False])
... dataset.append_int_column("ints", [-1, 0, 1])
... dataset.append_float_column("floats", [-1.0, 0.0, 1.0])
>>> with Dataset("docs/example.h5", "a") as dataset:
... row = dataset[0]
... print("0th row before:", [(key, row[key]) for key in sorted(row.keys())])
... dataset[0] = {"bools": True, "ints": 5, "floats": float("inf")}
... row = dataset[0]
... print("0th row after:", [(key, row[key]) for key in sorted(row.keys())])
0th row before: [('bools', True), ('floats', -1.0), ('ints', -1)]
0th row after: [('bools', True), ('floats', inf), ('ints', 5)]
>>> with Dataset("docs/example.h5", "a") as dataset:
... print(dataset["bools", 2])
... dataset["bools", 2] = True
... print(dataset["bools", 2])
False
True
>>> with Dataset("docs/example.h5", "a") as dataset:
... print(dataset["floats", 0])
... dataset[0, "floats"] = -5.0
... print(dataset["floats", 0])
inf
-5.0
"""
self._assert_is_writable()
if is_integer(item):
self._assert_index_exists(item)
if not isinstance(value, dict):
raise exceptions.InvalidDTypeError(
f"Dataset row should be a dict, but value {value} of type "
f"`{type(value)}` received.`"
)
self._set_row(item, value)
else:
column_name, index = self._prepare_item(item) # type: ignore
self._assert_column_exists(column_name)
column = self._h5_file[column_name]
if is_integer(index):
self._set_value(column, index, value, check_index=True) # type: ignore
elif index is None or isinstance(index, (slice, list, np.ndarray)):
self._set_column(column, value, index, preserve_values=True) # type: ignore
else:
raise exceptions.InvalidIndexError(
f"Invalid index {index} of type `{type(index)}` received."
)
self._update_generation_id()
@staticmethod
def _prepare_item(
item: Union[
str,
Tuple[str, Union[IndexType, Indices1dType]],
Tuple[Union[IndexType, Indices1dType], str],
],
) -> Tuple[str, Optional[Union[IndexType, Indices1dType]]]:
if isinstance(item, str):
return item, None
if isinstance(item, tuple) and len(item) == 2:
if isinstance(item[0], str):
return item
if isinstance(item[1], str):
return item[1], item[0]
raise exceptions.InvalidIndexError(
f"dataset index should be a string, an index or a pair of a string "
f"and an index/indices, but value {item} of type `{type(item)}` received."
)
def __iadd__(
self, other: Union[Dict[str, ColumnInputType], "Dataset"]
) -> "Dataset":
if isinstance(other, dict):
self.append_row(**other)
return self
if isinstance(other, Dataset):
self.append_dataset(other)
return self
raise TypeError(
f"`other` argument should be a dict or an `Dataset` instance, "
f"but value {other} of type `{type(other)}` received.`"
)
def __len__(self) -> int:
return self._length
def __str__(self) -> str:
return self._pretty_table().get_string()
def open(self, mode: Optional[str] = None) -> None:
"""
Open previously closed file or reopen file with another mode.
Args:
mode: Optional open mode. If not given, use `self.mode`.
"""
if mode is not None and mode != self._mode:
self._check_mode(mode)
self.close()
self._mode = mode
if self._closed:
self._h5_file = h5py.File(self._filepath, self._mode)
self._closed = False
self._column_names, self._length = self._get_column_names_and_length()
if self._is_writable():
if "spotlight_generation_id" not in self._h5_file.attrs:
self._h5_file.attrs["spotlight_generation_id"] = np.uint64(0)
self._append_internal_columns()
self._column_names.difference_update(set(INTERNAL_COLUMN_NAMES))
def close(self) -> None:
"""
Close file.
"""
if not self._closed:
if self._is_writable():
current_time = get_current_datetime().isoformat()
raw_attrs = self._h5_file.attrs
# Version could be `None`, but *shouldn't* be.
raw_attrs["version"] = __version__
raw_attrs["last_edited_by"] = self._get_username()
raw_attrs["last_edited_at"] = current_time
if "created" not in raw_attrs:
raw_attrs["created"] = __version__
if "created_by" not in raw_attrs:
raw_attrs["created_by"] = self._get_username()
if "created_at" not in raw_attrs:
raw_attrs["created_at"] = current_time
self._h5_file.close()
self._closed = True
self._column_names = set()
self._length = 0
def keys(self) -> List[str]:
"""
Get dataset column names.
"""
return list(self._column_names)
@overload
def iterrows(self) -> Iterable[Dict[str, Optional[ColumnType]]]:
...
@overload
def iterrows(
self, column_names: Union[str, Iterable[str]]
) -> Union[
Iterable[Dict[str, Optional[ColumnType]]], Iterable[Optional[ColumnType]]
]:
...
def iterrows(
self, column_names: Optional[Union[str, Iterable[str]]] = None
) -> Union[
Iterable[Dict[str, Optional[ColumnType]]], Iterable[Optional[ColumnType]]
]:
"""
Iterate through dataset rows.
"""
self._assert_is_opened()
if isinstance(column_names, str):
self._assert_column_exists(column_names)
column = self._h5_file[column_names]
column_type = self._get_column_type(column)
if column.attrs.get("external", False):
for value in column:
column_type = cast(Type[ExternalColumnType], column_type)
yield self._decode_external_value(value, column_type)
elif self._is_ref_column(column):
for ref in column:
column_type = cast(Type[RefColumnType], column_type)
yield self._decode_ref_value(ref, column_type, column_names)
else:
for value in column:
column_type = cast(Type[SimpleColumnType], column_type)
yield self._decode_simple_value(value, column, column_type)
else:
if column_names is None:
column_names = self._column_names
else:
column_names = set(column_names)
if column_names.difference(self._column_names):
raise exceptions.ColumnNotExistsError(
'Columns "'
+ '", "'.join(column_names.difference(self._column_names))
+ '" do not exist.'
)
columns = {
column_name: self._h5_file[column_name] for column_name in column_names
}
for i in range(self._length):
yield {
column_name: self._get_value(column, i)
for column_name, column in columns.items()
}
def from_pandas(
self,
df: pd.DataFrame,
index: bool = False,
dtype: Optional[ColumnTypeMapping] = None,
workdir: Optional[PathType] = None,
) -> None:
"""
Import a pandas dataframe to the dataset.
Only scalar types supported by the Spotlight dataset are imported, the
other are printed in a warning message.
Args:
df: `pandas.DataFrame` to import.
index: Whether to import index of the dataframe as regular dataset
column.
dtype: Optional dict with mapping `column name -> column type` with
column types allowed by Spotlight.
workdir: Optional folder where audio/images/meshes are stored. If
`None`, current folder is used.
Example:
>>> from datetime import datetime
>>> import pandas as pd
>>> from renumics.spotlight import Dataset
>>> df = pd.DataFrame(
... {
... "bools": [True, False, False],
... "ints": [-1, 0, 1],
... "floats": [-1.0, 0.0, 1.0],
... "strings": ["a", "b", "c"],
... "datetimes": datetime.now().astimezone(),
... }
... )
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.from_pandas(df, index=False)
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(len(dataset))
... print(sorted(dataset.keys()))
3
['bools', 'datetimes', 'floats', 'ints', 'strings']
"""
self._assert_is_writable()
if not df.columns.is_unique:
raise exceptions.DatasetColumnsNotUnique(
"DataFrame's columns are not unique"
)
if index:
df = df.reset_index(level=df.index.names) # type: ignore
else:
df = df.copy()
df.columns = pd.Index(stringify_columns(df))
inferred_dtype = infer_dtypes(df, dtype)
for column_name in df.columns:
try:
column = df[column_name]
column_type = inferred_dtype[column_name]
column = prepare_column(column, column_type)
if workdir is not None and is_file_based_column_type(dtype):
# For file-based data types, relative paths should be resolved.
str_mask = is_string_mask(column)
column[str_mask] = column[str_mask].apply(
lambda x: prepare_path_or_url(x, workdir)
)
attrs = {}
if column_type is Category:
attrs["categories"] = column.cat.categories.to_list()
values = column.to_numpy()
# `pandas` uses `NaN`s for unknown values, we use `None`.
values = np.where(pd.isna(values), np.array(None), values)
elif column_type is datetime:
values = column.to_numpy("datetime64[us]")
else:
values = column.to_numpy()
if is_file_based_column_type(column_type):
attrs["external"] = False
attrs["lookup"] = False
self.append_column(
column_name,
column_type,
values,
hidden=column_name.startswith("_"),
optional=column_type not in (bool, int),
**attrs,
)
except Exception as e:
if column_name in (dtype or {}):
raise e
logger.warning(
f"Column '{column_name}' not imported from "
f"`pandas.DataFrame` because of the following error:\n{e}"
)
def from_csv(
self,
filepath: PathType,
dtype: Optional[ColumnTypeMapping] = None,
columns: Optional[Iterable[str]] = None,
workdir: Optional[PathType] = None,
) -> None:
"""
Args:
filepath: Path of csv file to read.
dtype: Optional dict with mapping `column name -> column type` with
column types allowed by Spotlight.
columns: Optional columns to read from csv. If not set, read all
columns.
workdir: Optional folder where audio/images/meshes are stored. If
`None`, csv folder is used.
"""
if columns is not None:
columns = list(set(columns))
df: pd.DataFrame = pd.read_csv(filepath, usecols=columns or None)
if workdir is None:
workdir = os.path.dirname(filepath)
self.from_pandas(df, index=False, dtype=dtype, workdir=workdir)
def to_pandas(self) -> pd.DataFrame:
"""
Export the dataset to pandas dataframe.
Only scalar types of the Spotlight dataset are exported, the others are
printed in a warning message.
Returns:
`pandas.DataFrame` filled with the data of the Spotlight dataset.
Example:
>>> import pandas as pd
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bools", [True, False, False])
... dataset.append_int_column("ints", [-1, 0, 1])
... dataset.append_float_column("floats", [-1.0, 0.0, 1.0])
... dataset.append_string_column("strings", ["a", "b", "c"])
... dataset.append_datetime_column("datetimes", optional=True)
>>> with Dataset("docs/example.h5", "r") as dataset:
... df = dataset.to_pandas()
>>> print(len(df))
3
>>> print(df.columns.sort_values())
Index(['bools', 'datetimes', 'floats', 'ints', 'strings'], dtype='object')
"""
self._assert_is_opened()
df = pd.DataFrame()
for column_name in self._column_names:
column_type = self.get_column_type(column_name)
if column_type in (bool, int, float, str, datetime):
df[column_name] = self[column_name]
elif column_type is Category:
df[column_name] = pd.Categorical.from_codes(
self._h5_file[column_name],
self._h5_file[column_name].attrs["category_keys"], # type: ignore
)
not_exported_columns = self._column_names.difference(df.columns)
if len(not_exported_columns) > 0:
logger.warning(
'Columns "'
+ '", "'.join(not_exported_columns)
+ '" not appended to the dataframe. Please export them manually.'
)
return df
def append_bool_column(
self,
name: str,
values: Union[BoolColumnInputType, Iterable[BoolColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: BoolColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
editable: bool = True,
) -> None:
"""
Create and optionally fill a boolean column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional.
default: Value to use by default if column is optional and no value
or `None` is given. If `optional` is `True`, should be
explicitly set to `True` or `False`.
description: Optional column description.
tags: Optional tags for the column.
editable: Whether column is editable in Spotlight.
Example:
>>> from renumics.spotlight import Dataset
>>> value = False
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bool_values", 5*[value])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["bool_values", 2])
False
"""
self._append_column(
name,
bool,
values,
np.dtype(bool),
order,
hidden,
optional,
default,
description,
tags,
editable=editable,
)
def append_int_column(
self,
name: str,
values: Union[IntColumnInputType, Iterable[IntColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: IntColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
editable: bool = True,
) -> None:
"""
Create and optionally fill an integer column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given. If `optional` is `True`, should be
explicitly set.
description: Optional column description.
tags: Optional tags for the column.
editable: Whether column is editable in Spotlight.
Example:
Find a similar example usage in
:func:`renumics.spotlight.dataset.Dataset.append_bool_column`.
"""
self._append_column(
name,
int,
values,
np.dtype(int),
order,
hidden,
optional,
default,
description,
tags,
editable=editable,
)
def append_float_column(
self,
name: str,
values: Union[FloatColumnInputType, Iterable[FloatColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: FloatColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
editable: bool = True,
) -> None:
"""
Create and optionally fill a float column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than NaN is
specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
editable: Whether column is editable in Spotlight.
Example:
Find a similar example usage in
:func:`renumics.spotlight.dataset.Dataset.append_bool_column`.
"""
self._append_column(
name,
float,
values,
np.dtype(float),
order,
hidden,
optional,
default,
description,
tags,
editable=editable,
)
def append_string_column(
self,
name: str,
values: Union[StringColumnInputType, Iterable[StringColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: Optional[str] = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
editable: bool = True,
) -> None:
"""
Create and optionally fill a float column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than empty
string is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
editable: Whether column is editable in Spotlight.
Example:
Find a similar example usage in
:func:`renumics.spotlight.dataset.Dataset.append_bool_column`.
"""
self._append_column(
name,
str,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
editable=editable,
)
def append_datetime_column(
self,
name: str,
values: Union[
DatetimeColumnInputType, Iterable[DatetimeColumnInputType]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: DatetimeColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
) -> None:
"""
Create and optionally fill a datetime column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
Example:
>>> import numpy as np
>>> import datetime
>>> from renumics.spotlight import Dataset
>>> date = datetime.datetime.now()
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_datetime_column("dates", 5*[date])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["dates", 2] < datetime.datetime.now())
True
"""
self._append_column(
name,
datetime,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
)
def append_array_column(
self,
name: str,
values: Union[ArrayColumnInputType, Iterable[ArrayColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: ArrayColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
) -> None:
"""
Create and optionally fill a numpy array column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset
>>> array_data = np.random.rand(5,3)
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_array_column("arrays", 5*[array_data])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["arrays", 2].shape)
(5, 3)
"""
self._append_column(
name,
np.ndarray,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
)
def append_categorical_column(
self,
name: str,
values: Union[
CategoricalColumnInputType, Iterable[CategoricalColumnInputType]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: Optional[str] = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
editable: bool = True,
categories: Optional[Union[Iterable[str], Dict[str, int]]] = None,
) -> None:
"""
Create and optionally fill a categorical column.
Args:
name: Column name.
categories: The allowed categories for this column ("" is not allowed)
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than empty
string is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
editable: Whether column is editable in Spotlight.
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Category`.
"""
self._append_column(
name,
Category,
values,
np.dtype("int32"),
order,
hidden,
optional,
default,
description,
tags,
editable=editable,
categories=categories,
)
def append_embedding_column(
self,
name: str,
values: Union[
EmbeddingColumnInputType, Iterable[EmbeddingColumnInputType]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: EmbeddingColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
dtype: Union[str, np.dtype] = "float32",
) -> None:
"""
Create and optionally fill a mesh column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
dtype: A valid float numpy dtype. Default is "float32".
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Embedding`.
"""
np_dtype = np.dtype(dtype)
if np_dtype.str[1] != "f":
raise ValueError(
f'A float `dtype` expected, but dtype "{np_dtype.name}" received.'
)
self._append_column(
name,
Embedding,
values,
h5py.vlen_dtype(np_dtype),
order,
hidden,
optional,
default,
description,
tags,
)
def append_sequence_1d_column(
self,
name: str,
values: Union[
Sequence1DColumnInputType, Iterable[Sequence1DColumnInputType]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: Sequence1DColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
x_label: Optional[str] = None,
y_label: Optional[str] = None,
) -> None:
"""
Create and optionally fill a 1d-sequence column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
x_label: Optional x-axis label.
y_label: Optional y-axis label. If `None`, column name is taken.
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Sequence1D`.
"""
if y_label is None:
y_label = name
self._append_column(
name,
Sequence1D,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
x_label=x_label,
y_label=y_label,
)
def append_mesh_column(
self,
name: str,
values: Optional[
Union[MeshColumnInputType, Iterable[Optional[MeshColumnInputType]]]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: Optional[MeshColumnInputType] = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
lookup: Optional[
Union[
BoolType, Iterable[MeshColumnInputType], Dict[str, MeshColumnInputType]
]
] = None,
external: bool = False,
) -> None:
"""
Create and optionally fill a mesh column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
lookup: Optional data lookup/flag for automatic lookup creation.
If `False` (default if `external` is `True`), never add data to
lookup.
If `True` (default if `external` is `False`), add all given
files to the lookup, do nothing for explicitly given data.
If lookup is given, store it explicit, further behaviour is as
for `True`. If lookup is not a dict, keys are created automatically.
external: Whether column should only contain paths/URLs to data and
load it on demand.
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Mesh`.
"""
self._append_column(
name,
Mesh,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
lookup=not external if lookup is None else lookup,
external=external,
)
def append_image_column(
self,
name: str,
values: Union[ImageColumnInputType, Iterable[ImageColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: ImageColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
lookup: Optional[
Union[
BoolType, Iterable[MeshColumnInputType], Dict[str, MeshColumnInputType]
]
] = None,
external: bool = False,
) -> None:
"""
Create and optionally fill an image column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
lookup: Optional data lookup/flag for automatic lookup creation.
If `False` (default if `external` is `True`), never add data to
lookup.
If `True` (default if `external` is `False`), add all given
files to the lookup, do nothing for explicitly given data.
If lookup is given, store it explicit, further behaviour is as
for `True`. If lookup is not a dict, keys are created automatically.
external: Whether column should only contain paths/URLs to data and
load it on demand.
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Image`.
"""
self._append_column(
name,
Image,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
lookup=not external if lookup is None else lookup,
external=external,
)
def append_audio_column(
self,
name: str,
values: Optional[
Union[AudioColumnInputType, Iterable[AudioColumnInputType]]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: ImageColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
lookup: Optional[
Union[
BoolType, Iterable[MeshColumnInputType], Dict[str, MeshColumnInputType]
]
] = None,
external: bool = False,
lossy: Optional[bool] = None,
) -> None:
"""
Create and optionally fill an audio column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
lookup: Optional data lookup/flag for automatic lookup creation.
If `False` (default if `external` is `True`), never add data to
lookup.
If `True` (default if `external` is `False`), add all given
files to the lookup, do nothing for explicitly given data.
If lookup is given, store it explicit, further behaviour is as
for `True`. If lookup is not a dict, keys are created automatically.
external: Whether column should only contain paths/URLs to data and
load it on demand.
lossy: Whether to store data lossy or lossless (default if
`external` is `False`). Not recomended to use with
`external=True` since it requires on demand transcoding which
slows down the execution.
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Audio`.
"""
attrs = {}
if lossy is None and external is False:
lossy = False
if lossy is not None:
attrs["lossy"] = lossy
self._append_column(
name,
Audio,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
lookup=not external if lookup is None else lookup,
external=external,
**attrs,
)
def append_video_column(
self,
name: str,
values: Optional[
Union[VideoColumnInputType, Iterable[VideoColumnInputType]]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: VideoColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
lookup: Optional[
Union[
BoolType, Iterable[MeshColumnInputType], Dict[str, MeshColumnInputType]
]
] = None,
external: bool = False,
) -> None:
"""
Create and optionally fill an video column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
lookup: Optional data lookup/flag for automatic lookup creation.
If `False` (default if `external` is `True`), never add data to
lookup.
If `True` (default if `external` is `False`), add all given
files to the lookup, do nothing for explicitly given data.
If lookup is given, store it explicit, further behaviour is as
for `True`. If lookup is not a dict, keys are created automatically.
external: Whether column should only contain paths/URLs to data and
load it on demand.
"""
self._append_column(
name,
Video,
values,
h5py.string_dtype(),
order,
hidden,
optional,
default,
description,
tags,
lookup=not external if lookup is None else lookup,
external=external,
)
def append_window_column(
self,
name: str,
values: Optional[
Union[WindowColumnInputType, Iterable[WindowColumnInputType]]
] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: WindowColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
editable: bool = True,
) -> None:
"""
Create and optionally fill window column.
Args:
name: Column name.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
editable: Whether column is editable in Spotlight.
Example:
Find an example usage in :class:`renumics.spotlight.dtypes'.Window`.
"""
self._append_column(
name,
Window,
values,
np.dtype("float32"),
order,
hidden,
optional,
default,
description,
tags,
editable=editable,
)
def append_column(
self,
name: str,
column_type: Type[ColumnType],
values: Union[ColumnInputType, Iterable[ColumnInputType]] = None,
order: Optional[int] = None,
hidden: bool = False,
optional: bool = False,
default: ColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
**attrs: Optional[Union[str, bool]],
) -> None:
"""
Create and optionally fill a dataset column of the given type.
Args:
name: Column name.
column_type: Column type.
values: Optional column values. If a single value, the whole column
filled with this value.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
attrs: Optional arguments for the respective append column method.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_column("int", int, range(5))
... dataset.append_column("float", float, 1.0)
... dataset.append_column("bool", bool, True)
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(len(dataset))
... print(sorted(dataset.keys()))
5
['bool', 'float', 'int']
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["int"])
... print(dataset["bool"])
... print(dataset["float"])
[0 1 2 3 4]
[ True True True True True]
[1. 1. 1. 1. 1.]
"""
if column_type is bool:
append_column_fn: Callable = self.append_bool_column
elif column_type is int:
append_column_fn = self.append_int_column
elif column_type is float:
append_column_fn = self.append_float_column
elif column_type is str:
append_column_fn = self.append_string_column
elif column_type is datetime:
append_column_fn = self.append_datetime_column
elif column_type is np.ndarray:
append_column_fn = self.append_array_column
elif column_type is Embedding:
append_column_fn = self.append_embedding_column
elif column_type is Image:
append_column_fn = self.append_image_column
elif column_type is Mesh:
append_column_fn = self.append_mesh_column
elif column_type is Sequence1D:
append_column_fn = self.append_sequence_1d_column
elif column_type is Audio:
append_column_fn = self.append_audio_column
elif column_type is Category:
append_column_fn = self.append_categorical_column
elif column_type is Video:
append_column_fn = self.append_video_column
elif column_type is Window:
append_column_fn = self.append_window_column
else:
raise exceptions.InvalidDTypeError(f"Unknown column type: {column_type}.")
append_column_fn(
name=name,
values=values,
order=order,
hidden=hidden,
optional=optional,
default=default,
description=description,
tags=tags,
**attrs,
)
def append_row(self, **values: ColumnInputType) -> None:
"""
Append a row to the dataset.
Args:
values: A mapping column name -> value. Keys of `values` should
match dataset column names exactly except for optional columns.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bool_values")
... dataset.append_float_column("float_values")
>>> data = {"bool_values":True, "float_values":0.2}
>>> with Dataset("docs/example.h5", "a") as dataset:
... dataset.append_row(**data)
... dataset.append_row(**data)
... print(dataset["float_values", 1])
0.2
"""
self._assert_is_writable()
if not self._column_names:
raise exceptions.InvalidRowError(
"Cannot write a row, dataset has no columns."
)
values = self._encode_row(values)
try:
for column_name, value in values.items():
column = self._h5_file[column_name]
column.resize(self._length + 1, axis=0)
column[-1] = value
except Exception as e:
self._rollback(self._length)
raise e
self._length += 1
self._update_internal_columns(index=-1)
self._update_generation_id()
def append_dataset(self, dataset: "Dataset") -> None:
"""
Append a dataset to the current dataset row-wise.
"""
length = self._length
try:
for row in dataset.iterrows():
self.append_row(**row)
except Exception as e:
if self._length > length:
self._rollback(length)
self._length = length
self._update_internal_columns()
raise e
def insert_row(self, index: IndexType, values: Dict[str, ColumnInputType]) -> None:
"""
Insert a row into the dataset at the given index.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("example.h5", "w") as dataset:
... dataset.append_float_column("floats", [-1.0, 0.0, 1.0])
... dataset.append_int_column("ints", [-1, 0, 2])
... print(len(dataset))
... print(dataset["floats"])
... print(dataset["ints"])
3
[-1. 0. 1.]
[-1 0 2]
>>> with Dataset("example.h5", "a") as dataset:
... dataset.insert_row(2, {"floats": float("nan"), "ints": 1000})
... dataset.insert_row(-3, {"floats": 3.14, "ints": -1000})
... print(len(dataset))
... print(dataset["floats"])
... print(dataset["ints"])
5
[-1. 3.14 0. nan 1. ]
[ -1 -1000 0 1000 2]
"""
self._assert_is_writable()
self._assert_index_exists(index, check_type=True)
index = cast(int, index)
length = len(self)
if index < 0:
index += length
for column_name in self.keys() + INTERNAL_COLUMN_NAMES:
column = self._h5_file[column_name]
column.resize(length + 1, axis=0)
raw_values = column[index:-1]
if self._get_column_type(column) is Embedding:
raw_values = list(raw_values)
column[index + 1 :] = raw_values
self._length += 1
try:
self._set_row(index, values)
except Exception as e:
del self[index]
raise e
self._update_generation_id()
@overload
def pop(self, item: str) -> np.ndarray:
...
@overload
def pop(self, item: IndexType) -> Dict[str, Optional[ColumnType]]:
...
def pop(
self, item: Union[str, IndexType]
) -> Union[np.ndarray, Dict[str, Optional[ColumnType]]]:
"""
Delete a dataset column or row and return it.
"""
x = self[item]
del self[item]
return x
def isnull(self, column_name: str) -> np.ndarray:
"""
Get missing values mask for the given column.
`None`, `NaN` and category "" values are mapped to `True`. So null-mask
for columns of type `bool`, `int` and `string` always has only `False` values.
A `Window` is mapped on `True` only if both start and end are `NaN`.
"""
self._assert_is_opened()
self._assert_column_exists(column_name, internal=True)
column = self._h5_file[column_name]
raw_values = column[()]
column_type = self._get_column_type(column)
if self._is_ref_column(column):
return ~raw_values.astype(bool)
if column_type is datetime:
return np.array([raw_value in ["", b""] for raw_value in raw_values])
if column_type is float:
return np.isnan(raw_values)
if column_type is Category:
return raw_values == -1
if column_type is Window:
return np.isnan(raw_values).all(axis=1)
if column_type is Embedding:
return np.array([len(x) == 0 for x in raw_values])
return np.full(len(self), False)
def notnull(self, column_name: str) -> np.ndarray:
"""
Get non-missing values mask for the given column.
`None`, `NaN` and category "" values are mapped to `False`. So non-null-mask
for columns of type `bool`, `int` and `string` always has only `True` values.
A `Window` is mapped on `True` if at least one of its values is not `NaN`.
"""
return ~self.isnull(column_name)
def rename_column(self, old_name: str, new_name: str) -> None:
"""
Rename a dataset column.
"""
self._assert_is_writable()
self._assert_column_exists(old_name)
self.check_column_name(new_name)
self._assert_column_not_exists(new_name)
self._h5_file[new_name] = self._h5_file[old_name]
if f"__group__/{old_name}" in self._h5_file:
self._h5_file["__group__"].move(old_name, new_name)
del self._h5_file[old_name]
self._column_names.discard(old_name)
self._column_names.add(new_name)
self._update_generation_id()
def prune(self) -> None:
"""
Rebuild the whole dataset with the same content.
This method can be useful after column deletions, in order to decrease
the dataset file size.
"""
self._assert_is_opened()
column_names = self._column_names
# Internal columns could be not appended yet, then do not copy them.
for column_name in INTERNAL_COLUMN_NAMES:
if column_name in self._h5_file and isinstance(
self._h5_file[column_name], h5py.Dataset
):
column_names.add(column_name)
with TemporaryDirectory() as temp_dir:
new_dataset = os.path.join(temp_dir, "dataset.h5")
with h5py.File(new_dataset, "w") as h5_file:
# Copy top-level meta-information, if presented.
for attr_name, attr in self._h5_file.attrs.items():
h5_file.attrs[attr_name] = attr
for column_name in column_names:
column = self._h5_file[column_name]
new_column = h5_file.create_dataset(
column.name,
column.shape,
column.dtype,
maxshape=column.maxshape,
)
for attr_name, attr in column.attrs.items():
new_column.attrs[attr_name] = attr
raw_values = column[()]
if self._is_ref_column(column):
if h5py.check_string_dtype(column.dtype):
# New-style string refs.
for ref in raw_values:
if ref:
h5_dataset = self._resolve_ref(ref, column_name)
if h5_dataset.name not in h5_file:
h5_file.create_dataset(
h5_dataset.name, data=h5_dataset[()]
)
else:
# Old-style refs.
refs = []
for ref in raw_values:
if ref:
h5_dataset = self._resolve_ref(ref, column_name)
if h5_dataset.name in h5_file:
new_h5_dataset = h5_file[h5_dataset.name]
else:
new_h5_dataset = h5_file.create_dataset(
h5_dataset.name, data=h5_dataset[()]
)
refs.append(new_h5_dataset.ref)
else:
refs.append(None)
raw_values = refs
if self._get_column_type(column) is Embedding:
raw_values = list(raw_values)
new_column[:] = raw_values
self.close()
shutil.move(new_dataset, os.path.realpath(self._filepath))
self.open()
@overload
def get_column_type(
self, name: str, as_string: Literal[False] = False
) -> Type[ColumnType]:
...
@overload
def get_column_type(self, name: str, as_string: Literal[True]) -> str:
...
def get_column_type(
self, name: str, as_string: bool = False
) -> Union[Type[ColumnType], str]:
"""
Get type of dataset column.
Args:
name: Column name.
as_string: Get internal name of the column type.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_bool_column("bool")
... dataset.append_datetime_column("datetime")
... dataset.append_array_column("array")
... dataset.append_mesh_column("mesh")
>>> with Dataset("docs/example.h5", "r") as dataset:
... for column_name in sorted(dataset.keys()):
... print(column_name, dataset.get_column_type(column_name))
array <class 'numpy.ndarray'>
bool <class 'bool'>
datetime <class 'datetime.datetime'>
mesh <class 'renumics.spotlight.dtypes.Mesh'>
>>> with Dataset("docs/example.h5", "r") as dataset:
... for column_name in sorted(dataset.keys()):
... print(column_name, dataset.get_column_type(column_name, True))
array array
bool bool
datetime datetime
mesh Mesh
"""
self._assert_is_opened()
if not isinstance(name, str):
raise TypeError(
f"`item` argument should be a string, but value {name} of type "
f"`{type(name)}` received.`"
)
self._assert_column_exists(name, internal=True)
type_name = self._h5_file[name].attrs["type"]
if as_string:
return type_name
return get_column_type(type_name)
def get_column_attributes(
self, name: str
) -> Dict[
str,
Optional[
Union[
bool,
int,
str,
ColumnType,
Dict[str, int],
Dict[str, FileBasedColumnType],
]
],
]:
"""
Get attributes of a column. Available but unset attributes contain None.
Args:
name: Column name.
Example:
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_int_column("int", range(5))
... dataset.append_int_column(
... "int1",
... hidden=True,
... default=10,
... description="integer column",
... tags=["important"],
... editable=False,
... )
>>> with Dataset("docs/example.h5", "r") as dataset:
... attributes = dataset.get_column_attributes("int")
... for key in sorted(attributes.keys()):
... print(key, attributes[key])
default None
description None
editable True
hidden False
optional False
order None
tags None
>>> with Dataset("docs/example.h5", "r") as dataset:
... attributes = dataset.get_column_attributes("int1")
... for key in sorted(attributes.keys()):
... print(key, attributes[key])
default 10
description integer column
editable False
hidden True
optional True
order None
tags ['important']
"""
self._assert_is_opened()
if not isinstance(name, str):
raise TypeError(
f"`item` argument should be a string, but value {name} of type "
f"`{type(name)}` received.`"
)
self._assert_column_exists(name, internal=True)
column = self._h5_file[name]
column_attrs = column.attrs
column_type = self._get_column_type(column_attrs)
allowed_attributes = self._user_column_attributes(column_type)
attrs: Dict[
str,
Optional[
Union[
bool,
int,
str,
ColumnType,
Dict[str, int],
Dict[str, FileBasedColumnType],
]
],
] = {attribute_name: None for attribute_name in allowed_attributes}
attrs.update(
{
attribute_name: attribute_type(column_attrs[attribute_name])
if not attribute_type == object
else column_attrs[attribute_name]
for attribute_name, attribute_type in allowed_attributes.items()
if attribute_name in column_attrs
}
)
if "categories" in attrs:
if column_attrs.get("category_keys") is not None:
attrs["categories"] = dict(
zip(
column_attrs.get("category_keys"),
column_attrs.get("category_values"),
)
)
elif "lookup" in attrs:
if column_attrs.get("lookup_keys") is not None:
attrs["lookup"] = { # type: ignore
key: self._decode_value(ref, column)
for key, ref in zip(
column_attrs.get("lookup_keys"),
column_attrs.get("lookup_values"),
)
}
else:
attrs["lookup"] = False
default: _EncodedColumnType = attrs.get("default")
if default is not None:
attrs["default"] = self._decode_value(default, column)
return attrs
def _assert_valid_attribute(
self, attribute_name: str, attribute_value: ColumnInputType, column_name: str
) -> None:
column = self._h5_file.get(column_name)
column_type = self._get_column_type(column)
allowed_attributes = self._user_column_attributes(column_type)
if attribute_name not in allowed_attributes:
raise exceptions.InvalidAttributeError(
f'Setting an attribute with the name "{attribute_name}" for column '
f'"{column_name}" is not allowed. '
f'Allowed attribute names for "{column_type}" '
f'are: "{list(allowed_attributes.keys())}"'
)
if not isinstance(attribute_value, allowed_attributes[attribute_name]):
raise exceptions.InvalidAttributeError(
f'Attribute "{attribute_name}" for column "{column_name}" '
f"should be an {allowed_attributes[attribute_name]} or `None`, but "
f"value {attribute_value} of type `{type(attribute_value)}` received."
)
if (
attribute_name == "optional"
and not attribute_value
and column.attrs.get("optional")
):
raise exceptions.InvalidAttributeError(
f'Invalid `optional` argument for column "{column_name}" of '
f"type {column_type}. Columns can not be changed from "
f"`optional=False` to `optional=True`."
)
if attribute_name == "tags" and not all(
isinstance(tag, str) for tag in attribute_value
):
raise exceptions.InvalidAttributeError(
f'Invalid `tags` argument for column "{column_name}" of type '
f"{column_type}. Tags should be a `list of str`."
)
@staticmethod
def _write_lookup(
attrs: h5py.AttributeManager,
keys: Union[List, np.ndarray],
values: Union[List, np.ndarray],
column_name: str,
) -> None:
try:
attrs["lookup_keys"] = keys
attrs["lookup_values"] = values
except RuntimeError as e:
raise exceptions.InvalidAttributeError(
f"It seems that you have too many ({len(keys)}) unique values to "
f"store them in a lookup (~4000). To further write new data into "
f'column "{column_name}", you have to first disable the lookup:'
f'\n\t`dataset.set_column_attributes("{column_name}", lookup=False)`'
f"\nAlternatively, you can disable the lookup at column creation, e.g.:"
f"\n\t`dataset.append_<type>_column(<name>, lookup=False)`"
) from e
def set_column_attributes(
self,
name: str,
order: Optional[int] = None,
hidden: Optional[bool] = None,
optional: Optional[bool] = None,
default: ColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
**attrs: Any,
) -> None:
"""
Set attributes of a column.
Args:
name: Column name.
order: Optional Spotlight priority order value. `None` means the
lowest priority.
hidden: Whether column is hidden in Spotlight.
optional: Whether column is optional. If `default` other than `None`
is specified, `optional` is automatically set to `True`.
default: Value to use by default if column is optional and no value
or `None` is given.
description: Optional column description.
tags: Optional tags for the column.
attrs: Optional more ColumnType specific attributes .
"""
self._assert_is_writable()
if not isinstance(name, str):
raise TypeError(
f"`name` argument should be a string, but value {name} of type "
f"`{type(name)}` received.`"
)
self._assert_column_exists(name)
if default is not None:
optional = True
attrs["order"] = order
attrs["hidden"] = hidden
attrs["optional"] = optional
attrs["description"] = description
attrs["tags"] = tags
attrs = {k: v for k, v in attrs.items() if v is not None}
column = self._h5_file[name]
column_type = self._get_column_type(column)
if "lookup" in attrs:
lookup = attrs["lookup"]
if lookup in (True, np.bool_(True)):
attrs["lookup"] = {}
elif lookup in (False, np.bool_(False)):
if "lookup_values" in column.attrs:
for ref in column.attrs["lookup_values"]:
try:
del self._resolve_ref(ref, name).attrs["key"]
except (KeyError, ValueError):
...
del column.attrs["lookup_keys"], column.attrs["lookup_values"]
del attrs["lookup"]
for attribute_name, attribute_value in attrs.items():
self._assert_valid_attribute(attribute_name, attribute_value, name)
if "categories" in attrs:
if any(not v == np.int32(v) for v in attrs["categories"].values()):
raise exceptions.InvalidAttributeError(
f'Attribute "categories" for column "{name}" contains '
"invalid dict - values must be convertible to np.int32."
)
if any(not isinstance(v, str) for v in attrs["categories"].keys()):
raise exceptions.InvalidAttributeError(
f'Attribute "categories" for column "{name}" contains '
"invalid dict - keys must be of type str."
)
if any(v == "" for v in attrs["categories"].keys()):
raise exceptions.InvalidAttributeError(
f'Attribute "categories" for column "{name}" contains '
'invalid dict - "" (empty string) is no allowed as category key.'
)
if len(attrs["categories"].values()) > len(
set(attrs["categories"].values())
):
raise exceptions.InvalidAttributeError(
f'Attribute "categories" for column "{name}" contains '
"invalid dict - keys and values must be unique"
)
if column.attrs.get("category_keys") is not None:
values_must_include = column[:]
if "default" in column.attrs:
values_must_include = np.append(
values_must_include, column.attrs["default"]
)
missing_values = [
encoded_value
for encoded_value in values_must_include
if encoded_value not in attrs["categories"].values()
]
if any(missing_values):
raise exceptions.InvalidAttributeError(
f'Attribute "categories" for column "{name}" '
f"should include an entry for all values (and the default value) "
f"of the column ({set(column[:])}), but "
f"entries(s) having values {set(missing_values)} are missing."
)
attrs["category_keys"] = list(map(str, attrs["categories"].keys()))
attrs["category_values"] = np.array(
list(attrs["categories"].values()), dtype=np.int32
)
del attrs["categories"]
if "lookup" in attrs:
lookup = attrs.pop("lookup")
if "lookup_values" in column.attrs:
raise exceptions.InvalidAttributeError(
f'Lookup for the column "{name}" already set, cannot reset it.'
)
lookup_keys = []
lookup_values = []
for key, value in lookup.items():
ref = self._encode_value(value, column)
self._resolve_ref(ref, name).attrs["key"] = key
lookup_keys.append(key)
lookup_values.append(ref)
self._write_lookup(
column.attrs,
np.array(lookup_keys, dtype=h5py.string_dtype()),
np.array(lookup_values, dtype=h5py.string_dtype()),
name,
)
if "lossy" in attrs:
lossy = attrs["lossy"]
attrs["format"] = "mp3" if lossy else "flac"
if len(column) > 0 and (
column.attrs.get("lossy") != lossy
or column.attrs.get("format") != attrs.get("format")
):
raise exceptions.InvalidAttributeError(
"Cannot change `lossy` attribute after column creation."
)
column.attrs.update(attrs)
if attrs.get("optional"):
old_default = column.attrs.pop("default", None)
# Set new default value.
try:
if default is None and old_default is None:
default = self._default_default(column_type)
if (
default is None
and column_type is Embedding
and not self._is_ref_column(column)
):
# For a non-ref `Embedding` column, replace `None` with an empty array.
default = np.empty(0, column.dtype.metadata["vlen"])
if column_type is Category and default != "":
if default not in column.attrs["category_keys"]:
column.attrs["category_values"] = np.append(
column.attrs["category_values"],
max(column.attrs["category_values"] + 1),
).astype(dtype=np.int32)
column.attrs["category_keys"] = np.append(
column.attrs["category_keys"], np.array(default)
)
if default is not None:
encoded_value = self._encode_value(default, column)
if column_type is datetime and encoded_value is None:
encoded_value = ""
column.attrs["default"] = encoded_value
except Exception as e:
# Rollback
if old_default is not None:
column.attrs["default"] = old_default
raise e
self._update_generation_id()
def _repr_html_(self) -> str:
return self._pretty_table().get_html_string()
def _pretty_table(self) -> prettytable.PrettyTable:
"""
Get `PrettyTable` representation of the dataset.
"""
def _format(value: _EncodedColumnType, ref_type_name: str) -> str:
"""
Get string representation of a dataset value.
"""
if value is None:
return ""
if isinstance(value, h5py.Reference):
return f"<{ref_type_name}>" if value else ""
return str(value)
self._assert_is_opened()
required_keys = (
"type",
"order",
"hidden",
"optional",
"default",
"description",
"tags",
)
table = prettytable.PrettyTable()
column_names = sorted(self._column_names)
columns = [self._h5_file[column_name] for column_name in column_names]
column_reprs = []
for column in columns:
attrs = column.attrs
type_name = attrs["type"]
column_type = get_column_type(type_name)
optional_keys = set(
self._user_column_attributes(column_type).keys()
).difference(required_keys)
column_reprs.append(
[
key + ": " + _format(attrs.get(key), type_name)
for key in required_keys + tuple(sorted(optional_keys))
]
)
column_repr_length = max((len(x) for x in column_reprs), default=0)
for column_repr in column_reprs:
column_repr.extend([""] * (column_repr_length - len(column_repr)))
for column_name, column_repr, column in zip(
column_names, column_reprs, columns
):
type_name = column.attrs["type"]
column_repr.extend(_format(value, type_name) for value in column)
table.add_column(column_name, column_repr)
return table
def _append_column(
self,
name: str,
column_type: Type[ColumnType],
values: Union[ColumnInputType, Iterable[ColumnInputType]],
dtype: np.dtype,
order: Optional[int] = None,
hidden: bool = True,
optional: bool = False,
default: ColumnInputType = None,
description: Optional[str] = None,
tags: Optional[List[str]] = None,
**attrs: Any,
) -> None:
self._assert_is_writable()
self.check_column_name(name)
self._assert_column_not_exists(name)
# Here, only do expensive logic which *should not* be applied in the
# `set_column_attributes` method.
shape: Tuple[int, ...] = (0,)
maxshape: Tuple[Optional[int], ...] = (None,)
if column_type is Category:
categories = attrs.get("categories", None)
if categories is None:
# Values are given, but no categories.
if is_iterable(values):
values = list(values)
categories = set(values)
else:
categories = {values}
categories.difference_update({"", None})
if is_iterable(categories) and not isinstance(categories, dict):
# dict is forced to preserve the order.
categories = list(dict.fromkeys(categories, None).keys())
attrs["categories"] = dict(zip(categories, range(len(categories))))
# Otherwise, exception about type will be raised later in the
# `set_column_attributes` method.
elif column_type is Window:
shape = (0, 2)
maxshape = (None, 2)
elif issubclass(column_type, FileBasedDType):
lookup = attrs.get("lookup", None)
if is_iterable(lookup) and not isinstance(lookup, dict):
# Assume that we can keep all the lookup values in memory.
attrs["lookup"] = {str(i): v for i, v in enumerate(lookup)}
try:
column = self._h5_file.create_dataset(name, shape, dtype, maxshape=maxshape)
self._column_names.add(name)
column.attrs["type"] = get_column_type_name(column_type)
self.set_column_attributes(
name,
order,
hidden,
optional,
default,
description,
tags,
**attrs,
)
self._set_column(column, values)
except Exception as e:
if name in self._h5_file:
del self._h5_file[name]
try:
del self._h5_file[f"__group__/{name}"]
except KeyError:
pass
self._column_names.discard(name)
raise e
self._update_generation_id()
def _set_column(
self,
column: h5py.Dataset,
values: Union[ColumnInputType, Iterable[ColumnInputType]],
indices: Union[None, slice, List[Union[int, bool]], np.ndarray] = None,
preserve_values: bool = False,
) -> None:
column_name = self._get_column_name(column)
row_wise_filling_message = (
f"Dataset has initialized, but unfilled columns and should be "
f"filled row-wise, but values received for column "
f'"{column_name}".'
)
attrs = column.attrs
# Prepare indices.
if indices is None:
column_indices = values_indices = slice(None) # equivalent to `[:]`.
indices_length = self._length
else:
# We can only write unique sorted indices to `h5py` column, so
# prepare such indices.
try:
column_indices = np.arange(self._length, dtype=int)[indices] # type: ignore
except Exception as e:
raise exceptions.InvalidIndexError(
f"Indices {indices} of type `{type(indices)}` do not match "
f"to the dataset with the length {self._length}."
) from e
indices_length = len(column_indices) # type: ignore
if indices_length == 0:
# e.g.: `dataset[column_name, []] = values`.
logger.warning(
"No values set because the given indices reference no elements."
)
return
column_indices, values_indices = np.unique( # type: ignore
column_indices, return_index=True
)
if len(cast(np.ndarray, column_indices)) != indices_length:
# For now, forbid non-unique indices.
raise exceptions.InvalidIndexError(
"When setting multiple values in a column, indices should be unique."
)
# Fix for non-unique indices.
# indices, values_indices = np.unique(indices[::-1], return_index=True)
# values_indices = indices_length - values_indices
target_column_length = self._length
encoded_values: Union[np.ndarray, List[_EncodedColumnType]]
if is_iterable(values):
# Single windows and embeddings also come here.
encoded_values = self._encode_values(values, column)
if len(encoded_values) != indices_length:
if indices_length == 0:
# That is, `self._length` is 0 (otherwise already returned).
if len(self._column_names) == 1:
target_column_length = indices_length = len(encoded_values)
else:
raise exceptions.InvalidShapeError(row_wise_filling_message)
elif len(encoded_values) == 1:
# Stretch the values for all indices.
encoded_values = np.broadcast_to(
encoded_values, (indices_length, *encoded_values.shape[1:])
)
else:
raise exceptions.InvalidShapeError(
f"{indices_length} values or a single value expected "
f'for column "{column_name}", but '
f"{len(encoded_values)} values received."
)
elif indices_length == 0:
# Set an empty column with 0 values, i.e. do nothing.
return
else:
# Reorder values according to the given indices.
encoded_values = encoded_values[values_indices]
if self._get_column_type(column) is Embedding:
encoded_values = list(encoded_values)
elif values is not None:
# A single value is given. `Window` and `Embedding` values should
# never go here, because they are always iterable, even a single value.
if self._length == 0:
if len(self._column_names) == 1:
target_column_length = indices_length = 1
else:
raise exceptions.InvalidShapeError(row_wise_filling_message)
encoded_values = [
self._encode_value(cast(ColumnInputType, values), column)
] * indices_length
elif self._length == 0:
return
elif attrs.get("optional", False):
encoded_values = column.attrs.get(
"default", "" if h5py.check_string_dtype(column.dtype) else None
)
else:
raise exceptions.InvalidDTypeError(
f'Dataset has been initialized and values for non-optional column "{column_name}" '
f"must be provided on column creation. But no values were provided."
)
old_values = column[:] if preserve_values else None
try:
column.resize(target_column_length, axis=0)
column[column_indices] = encoded_values
except Exception as e:
if preserve_values:
column.resize(self._length, axis=0)
column[:] = old_values
raise e
self._length = target_column_length
self._update_internal_columns(column_indices)
def _set_row(self, index: IndexType, row: Dict[str, ColumnInputType]) -> None:
old_row = {
column_name: self._h5_file[column_name][index]
for column_name in self._column_names
}
try:
row = self._encode_row(row)
for column_name, value in row.items():
self._h5_file[column_name][index] = value
except Exception as e:
# Rollback changed row.
for column_name, item_value in old_row.items():
self._h5_file[column_name][index] = item_value
raise e
self._update_internal_columns(index)
def _set_value(
self,
column: h5py.Dataset,
index: IndexType,
value: ColumnInputType,
check_index: bool = False,
) -> None:
if check_index:
self._assert_index_exists(index)
old_value = column[index]
try:
value = self._encode_value(value, column)
if value is None:
attrs = column.attrs
if attrs.get("optional", False):
value = attrs.get("default", None)
column[index] = value
except Exception as e:
column[index] = old_value
raise e
self._update_internal_columns(index)
def _get_column(
self,
column: h5py.Dataset,
indices: Optional[Indices1dType] = None,
) -> np.ndarray:
"""
Read and decode values of the given existing column.
If `indices` is `None`, get the whole column. Otherwise, all possible
indices supported by one-dimensional numpy arrays should work.
"""
if indices is None:
values = column[()]
else:
# We can only read unique increasing indices from h5py,
# so prepare such indices, take values and remap them back.
try:
indices = np.arange(len(self), dtype=int)[indices]
except Exception as e:
raise exceptions.InvalidIndexError(
f"Indices {indices} of type `{type(indices)}` do not match "
f"to the dataset with the length {self._length}."
) from e
indices, mapping = np.unique(indices, return_inverse=True)
values = column[indices][mapping]
return self._decode_values(values, column)
def _decode_values(self, values: np.ndarray, column: h5py.Dataset) -> np.ndarray:
column_type = self._get_column_type(column)
if column.attrs.get("external", False):
column_type = cast(Type[ExternalColumnType], column_type)
return self._decode_external_values(values, column_type)
if self._is_ref_column(column):
column_type = cast(Type[RefColumnType], column_type)
return self._decode_ref_values(values, column, column_type)
column_type = cast(Type[SimpleColumnType], column_type)
return self._decode_simple_values(values, column, column_type)
@staticmethod
def _decode_simple_values(
values: np.ndarray, column: h5py.Dataset, column_type: Type[SimpleColumnType]
) -> np.ndarray:
if column_type is Category:
mapping = dict(
zip(column.attrs["category_values"], column.attrs["category_keys"])
)
mapping[-1] = ""
return np.array([mapping[x] for x in values], dtype=str)
if h5py.check_string_dtype(column.dtype):
# `column_type` is `str` or `datetime`.
values = np.array([x.decode("utf-8") for x in values])
if column_type is str:
return values
# Decode datetimes.
return np.array(
[None if x == "" else datetime.fromisoformat(x) for x in values],
dtype=object,
)
if column_type is Embedding:
null_mask = [len(x) == 0 for x in values]
values[null_mask] = None
# For column types `bool`, `int`, `float` or `Window`, return the array as-is.
return values
def _decode_ref_values(
self, values: np.ndarray, column: h5py.Dataset, column_type: Type[RefColumnType]
) -> np.ndarray:
column_name = self._get_column_name(column)
if column_type in (np.ndarray, Embedding):
# `np.array([<...>], dtype=object)` creation does not work for
# some cases and erases dtypes of sub-arrays, so we use assignment.
decoded_values = np.empty(len(values), dtype=object)
decoded_values[:] = [
self._decode_ref_value(ref, column_type, column_name) for ref in values
]
return decoded_values
return np.array(
[self._decode_ref_value(ref, column_type, column_name) for ref in values],
dtype=object,
)
def _decode_external_values(
self, values: np.ndarray, column_type: Type[ExternalColumnType]
) -> np.ndarray:
return np.array(
[self._decode_external_value(value, column_type) for value in values],
dtype=object,
)
def _get_value(
self, column: h5py.Dataset, index: IndexType, check_index: bool = False
) -> Optional[ColumnType]:
if check_index:
self._assert_index_exists(index)
value = column[index]
return self._decode_value(value, column)
def _get_column_names_and_length(self) -> Tuple[Set[str], int]:
"""
Parse valid columns of the same length from a H5 file. Valid columns
should have a known type stored in column attributes and have the same
length.
If valid columns of different length found, take the columns of the most
frequent (mode) length only. If multiple modes found, take the columns
of the greatest length.
Returns:
column_names: Names of the chosen columns.
length: Length of the chosen columns.
"""
names = []
lengths = []
for name in self._h5_file:
h5_dataset = self._h5_file[name]
if isinstance(h5_dataset, h5py.Dataset):
try:
self._get_column_type(h5_dataset)
except (KeyError, exceptions.InvalidDTypeError):
continue
else:
names.append(name)
shape = h5_dataset.shape
lengths.append(shape[0] if shape else 0)
max_count = 0
length_modes = []
for length in set(lengths):
count = lengths.count(length)
if count == max_count:
length_modes.append(length)
elif count > max_count:
length_modes = [length]
max_count = count
length = max(length_modes, default=0)
column_names = {
column_name
for column_name, column_length in zip(names, lengths)
if column_length == length
}
if len(column_names) < len(names):
logger.info(
f"Columns with different length found. The greatest of the "
f"most frequent length ({length}) chosen and only columns with "
f"this length taken as the dataset's columns."
)
return column_names, length
def _encode_values(
self, values: Iterable[ColumnInputType], column: h5py.Dataset
) -> np.ndarray:
if self._is_ref_column(column):
values = cast(Iterable[RefColumnInputType], values)
return self._encode_ref_values(values, column)
values = cast(Iterable[SimpleColumnInputType], values)
return self._encode_simple_values(values, column)
def _encode_simple_values(
self, values: Iterable[SimpleColumnInputType], column: h5py.Dataset
) -> np.ndarray:
column_type = cast(Type[SimpleColumnType], self._get_column_type(column))
if column_type is Category:
mapping = dict(
zip(column.attrs["category_keys"], column.attrs["category_values"])
)
if column.attrs.get("optional", False):
default = column.attrs.get("default", -1)
mapping[None] = default
if default == -1:
mapping[""] = -1
try:
# Map values and save as the right int type.
return np.array([mapping[x] for x in values], dtype=column.dtype)
except KeyError as e:
column_name = self._get_column_name(column)
raise exceptions.InvalidValueError(
f'Values for the categorical column "{column_name}" '
f"contain unknown categories."
) from e
if column_type is datetime:
if _check_valid_array(values, column_type):
encoded_values = np.array(
[None if x is None else x.isoformat() for x in values.tolist()]
)
else:
encoded_values = np.array(
[self._encode_value(value, column) for value in values]
)
if np.issubdtype(encoded_values.dtype, str):
# That means, we have all strings in array, no `None`s.
return encoded_values
return self._replace_none(encoded_values, column)
if column_type is Window:
encoded_values = self._asarray(values, column, column_type)
if encoded_values.ndim == 1:
if len(encoded_values) == 2:
# A single window, reshape it to an array.
return np.broadcast_to(values, (1, 2)) # type: ignore
if len(encoded_values) == 0:
# An empty array, reshape for compatibility.
return np.broadcast_to(values, (0, 2)) # type: ignore
elif encoded_values.ndim == 2 and encoded_values.shape[1] == 2:
# An array with valid windows.
return encoded_values
column_name = self._get_column_name(column)
raise exceptions.InvalidShapeError(
f'Input values to `Window` column "{column_name}" should have '
f"one of shapes (2,) (a single window) or (n, 2) (multiple "
f"windows), but values with shape {encoded_values.shape} received."
)
if column_type is Embedding:
if _check_valid_array(values, column_type):
# This is the only case we can handle fast and easily, otherwise
# embedding should go through `_encode_value` element-wise.
if values.ndim == 1:
# Handle 1-dimensional input as a single embedding.
self._assert_valid_or_set_embedding_shape(values.shape, column)
values_list = list(np.broadcast_to(values, (1, len(values))))
elif values.ndim == 2:
self._assert_valid_or_set_embedding_shape(values.shape[1:], column)
values_list = list(values)
else:
raise exceptions.InvalidShapeError(
f"Input values for an `Embedding` column should have 1 "
f"or 2 dimensions, but values with shape "
f"{values.shape} received."
)
else:
values_list = [self._encode_value(value, column) for value in values]
encoded_values = np.empty(len(values_list), dtype=object)
encoded_values[:] = values_list
encoded_values = self._replace_none(encoded_values, column)
return encoded_values
# column type is `bool`, `int`, `float` or `str`.
encoded_values = self._asarray(values, column, column_type)
if encoded_values.ndim == 1:
return encoded_values
column_name = self._get_column_name(column)
raise exceptions.InvalidShapeError(
f'Input values to `{column_type}` column "{column_name}" should '
f"be 1-dimensional, but values with shape {encoded_values.shape} "
f"received."
)
def _encode_ref_values(
self, values: Iterable[RefColumnInputType], column: h5py.Dataset
) -> np.ndarray:
encoded_values = np.array(
[self._encode_value(value, column) for value in values]
)
encoded_values = self._replace_none(encoded_values, column)
if h5py.check_string_dtype(column.dtype):
encoded_values[encoded_values == np.array(None)] = ""
return encoded_values
def _asarray(
self,
values: Iterable[SimpleColumnInputType],
column: h5py.Dataset,
column_type: Type[SimpleColumnType],
) -> np.ndarray:
if isinstance(values, np.ndarray):
if _check_valid_value_dtype(values.dtype, column_type):
return values
elif not isinstance(values, (list, tuple, range)):
# Make iterables, dicts etc. convertible to an array.
values = list(values)
# Array can contain `None`s, so do not infer dtype.
encoded_values = np.array(values, dtype=object)
encoded_values = self._replace_none(encoded_values, column)
try:
# At the moment, `None`s are already replaced, so try optimistic
# dtype conversion.
return np.array(encoded_values.tolist(), dtype=column.dtype)
except TypeError as e:
column_name = self._get_column_name(column)
raise exceptions.InvalidValueError(
f'Values for the column "{column_name}" of type {column_type} '
f"are not convertible to the dtype {column.dtype}."
) from e
@staticmethod
def _replace_none(values: np.ndarray, column: h5py.Dataset) -> np.ndarray:
"""replace all None entries with the default value for the column"""
# none_mask = values == np.array(None)
none_mask = [x is None for x in values]
if not any(none_mask):
# no nones present -> just return all the values
return values
if not column.attrs.get("optional", False):
raise exceptions.InvalidDTypeError(
f"`values` argument for non-optional column "
f'"{column.name.lstrip("/")}" contains `None` values.'
)
try:
default = column.attrs["default"]
except KeyError:
# no default value -> keep None as None
return values
if not isinstance(default, str):
default = default.tolist()
# Replace `None`s with the default value.
default_array = np.empty(1, dtype=object)
default_array[0] = default
values[none_mask] = default_array
return values
def _encode_row(
self, values: Dict[str, ColumnInputType]
) -> Dict[str, _EncodedColumnType]:
"""
Encode a single row for writing to dataset.
This method also replaces missed and `None` values with default values
if possible and checks row consistency.
"""
# Encode row.
values = values.copy()
for column_name in self._column_names:
values[column_name] = self._encode_value(
values.get(column_name), self._h5_file[column_name]
)
if values[column_name] is None:
column = self._h5_file[column_name]
attrs = column.attrs
if attrs.get("optional", False):
values[column_name] = attrs.get(
"default", "" if h5py.check_string_dtype(column.dtype) else None
)
# Check row consistency.
if values.keys() == self._column_names:
return values
error_message = (
"Keys of `values` mismatch column names, even with updated "
"default values."
)
missing_keys = self._column_names - set(values.keys())
if missing_keys:
error_message += (
'\n\tKeys "' + '", "'.join(missing_keys) + '" missing in ' "`values`."
)
excessive_keys = set(values.keys()) - self._column_names
if excessive_keys:
error_message += (
'\n\tColumns "'
+ '", "'.join(excessive_keys)
+ '" should be appended to the dataset.'
)
raise exceptions.InvalidRowError(error_message)
def _encode_value(
self, value: ColumnInputType, column: h5py.Dataset
) -> _EncodedColumnType:
"""
Encode a value for writing into a column, *but* do not replace `None`s
with default value (only check that this exists), since batch replace
should be faster.
"""
column_name = self._get_column_name(column)
attrs = column.attrs
if value is None:
if attrs.get("optional", False):
return value
raise exceptions.InvalidDTypeError(
f'No value given for the non-optional column "{column_name}".'
)
if attrs.get("external", False):
value = cast(PathOrUrlType, value)
return self._encode_external_value(value, column)
column_type = self._get_column_type(attrs)
if self._is_ref_column(column):
value = cast(RefColumnInputType, value)
return self._encode_ref_value(value, column, column_type, column_name)
value = cast(SimpleColumnInputType, value)
return self._encode_simple_value(value, column, column_type, column_name)
def _encode_simple_value(
self,
value: SimpleColumnInputType,
column: h5py.Dataset,
column_type: Type[ColumnType],
column_name: str,
) -> _EncodedColumnType:
"""
Encode a non-ref value, e.g. bool, int, float, str, datetime, Category,
Window and Embedding (in last versions).
Value *cannot* be `None` already.
"""
attrs = column.attrs
if column_type is Category:
categories = dict(
zip(attrs.get("category_keys"), attrs.get("category_values"))
)
if attrs.get("optional", False) and attrs.get("default", -1) == -1:
categories[""] = -1
if value not in categories.keys():
raise exceptions.InvalidValueError(
f"Values for {column_type} column "
f'"{column.name.lstrip("/")}" should be one of '
f"{list(categories.keys())} "
f"but value '{value}' received."
)
return categories[value]
if column_type is Window:
value = np.asarray(value, dtype=column.dtype)
if value.shape == (2,):
return value
raise exceptions.InvalidDTypeError(
f"Windows should consist of 2 values, but window of shape "
f"{value.shape} received for column {column_name}."
)
if column_type is Embedding:
# `Embedding` column is not a ref column.
if isinstance(value, Embedding):
value = value.encode(attrs.get("format", None))
value = np.asarray(value, dtype=column.dtype.metadata["vlen"])
self._assert_valid_or_set_embedding_shape(value.shape, column)
return value
self._assert_valid_value_type(value, column_type, column_name)
if isinstance(value, np.str_):
return value.tolist()
if isinstance(value, np.datetime64):
value = value.astype(datetime)
if isinstance(value, datetime):
return value.isoformat()
return value
def _encode_ref_value(
self,
value: RefColumnInputType,
column: h5py.Dataset,
column_type: Type[ColumnType],
column_name: str,
) -> _EncodedColumnType:
"""
Encode a ref value, e.g. np.ndarray, Sequence1D, Image, Mesh, Audio,
Video, and Embedding (in old versions).
Value *cannot* be `None` already.
"""
attrs = column.attrs
key: Optional[str] = None
lookup_keys: List[str] = []
if column_type is Mesh and isinstance(value, trimesh.Trimesh):
value = Mesh.from_trimesh(value)
elif issubclass(column_type, (Audio, Image, Video)) and isinstance(
value, bytes
):
value = column_type.from_bytes(value)
elif is_file_based_column_type(column_type) and isinstance(
value, (str, os.PathLike)
):
try:
lookup_keys = attrs["lookup_keys"].tolist()
except KeyError:
pass # Don't need to search/update, so encode value as usual.
else:
key = str(value)
try:
index = lookup_keys.index(key)
except ValueError:
pass # Index not found, so encode value as usual.
else:
# Return stored ref, do not process data again.
return attrs["lookup_values"][index]
try:
value = column_type.from_file(value)
except Exception:
return None
if issubclass(column_type, (Embedding, Image, Sequence1D)):
if not isinstance(value, column_type):
value = column_type(value) # type: ignore
value = value.encode(attrs.get("format", None)) # type: ignore
elif issubclass(column_type, (Mesh, Audio, Video)):
self._assert_valid_value_type(value, column_type, column_name)
value = value.encode(attrs.get("format", None)) # type: ignore
else:
value = np.asarray(value)
# `value` can be a `np.ndarray` or a `np.void`.
if isinstance(value, np.ndarray):
# Check dtype.
self._assert_valid_or_set_value_dtype(value.dtype, column)
if column_type is Embedding:
self._assert_valid_or_set_embedding_shape(value.shape, column)
dataset_name = str(uuid.uuid4()) if key is None else escape_dataset_name(key)
h5_dataset = self._h5_file.create_dataset(
f"__group__/{column_name}/{dataset_name}", data=value
)
if h5py.check_ref_dtype(column.dtype):
ref = h5_dataset.ref # Legacy handling.
else:
ref = dataset_name
if key is not None:
# `lookup_keys` is not `None`, so `lookup_values` too.
self._write_lookup(
attrs,
lookup_keys + [key],
np.concatenate(
(attrs["lookup_values"], [ref]),
dtype=column.dtype,
),
column_name,
)
h5_dataset.attrs["key"] = key
return ref
def _encode_external_value(self, value: PathOrUrlType, column: h5py.Dataset) -> str:
"""
Encode an external value, i.e. an URL or a path.
Value *should not* be a `None`.
Column *must* be an external column (H5 dataset with string dtype).
"""
if not isinstance(value, (str, os.PathLike)):
column_name = self._get_column_name(column)
raise exceptions.InvalidDTypeError(
f'For the external column "{column_name}" values should '
f"contain only URLs and/or paths (`str` or `os.PathLike`), but "
f"value {value} of type {type(value)} received."
)
value = str(value)
attrs = column.attrs
lookup_keys: Optional[List[str]] = None
# We still can have a lookup.
try:
lookup_keys = attrs["lookup_keys"].tolist()
except KeyError:
pass # Don't need to search/update, so encode value as usual.
else:
lookup_keys = cast(List[str], lookup_keys)
try:
index = lookup_keys.index(value)
except ValueError:
pass # Index not found, so encode value as usual.
else:
# Return stored value, do not process data again.
return attrs["lookup_values"][index]
if not (validators.url(value) or os.path.isfile(value)):
logger.warning(
f'File "{value}" not found, but still written into '
f"the dataset. If it does not appear at the reading "
f"time, a `None` will be returned."
)
if lookup_keys is not None:
# `lookup_keys` is not `None`, so `lookup_values` too.
self._write_lookup(
attrs,
lookup_keys + [value],
np.concatenate(
(attrs["lookup_values"], [value]),
dtype=column.dtype,
),
self._get_column_name(column),
)
return value
@staticmethod
def _assert_valid_value_type(
value: ColumnInputType, column_type: Type[ColumnType], column_name: str
) -> None:
if not _check_valid_value_type(value, column_type):
allowed_types = (column_type,) + _ALLOWED_COLUMN_TYPES.get(column_type, ())
raise exceptions.InvalidDTypeError(
f'Values for non-optional {column_type} column "{column_name}" '
f"should be one of {allowed_types} instances, but value "
f"{value} of type `{type(value)}` received."
)
def _decode_value(
self,
value: Union[
np.bool_, np.integer, np.floating, bytes, str, np.ndarray, h5py.Reference
],
column: h5py.Dataset,
) -> Optional[ColumnType]:
column_type = self._get_column_type(column)
if column.attrs.get("external", False):
value = cast(bytes, value)
column_type = cast(Type[ExternalColumnType], column_type)
return self._decode_external_value(value, column_type)
if self._is_ref_column(column):
value = cast(Union[bytes, h5py.Reference], value)
column_type = cast(Type[RefColumnType], column_type)
column_name = self._get_column_name(column)
return self._decode_ref_value(value, column_type, column_name)
value = cast(Union[np.bool_, np.integer, np.floating, bytes, np.ndarray], value)
column_type = cast(Type[SimpleColumnType], column_type)
return self._decode_simple_value(value, column, column_type)
@staticmethod
def _decode_simple_value(
value: Union[np.bool_, np.integer, np.floating, bytes, str, np.ndarray],
column: h5py.Dataset,
column_type: Type[SimpleColumnType],
) -> Optional[Union[bool, int, float, str, datetime, np.ndarray]]:
if column_type is Window:
value = cast(np.ndarray, value)
return value
if column_type is Embedding:
value = cast(np.ndarray, value)
if len(value) == 0:
return None
return value
if column_type is Category:
mapping = dict(
zip(column.attrs["category_values"], column.attrs["category_keys"])
)
if column.attrs.get("optional", False) and column.attrs.get(
"default", None
) in (-1, None):
mapping[-1] = ""
return mapping[value]
if isinstance(value, bytes):
value = value.decode("utf-8")
if column_type is datetime:
value = cast(str, value)
if value == "":
return None
return datetime.fromisoformat(value)
value = cast(Union[np.bool_, np.integer, np.floating, str], value)
column_type = cast(Type[Union[bool, int, float, str]], column_type)
return column_type(value)
def _decode_ref_value(
self,
ref: Union[bytes, str, h5py.Reference],
column_type: Type[RefColumnType],
column_name: str,
) -> Optional[Union[np.ndarray, Audio, Image, Mesh, Sequence1D, Video]]:
# Value can be a H5 reference or a string reference.
if not ref:
return None
value = self._resolve_ref(ref, column_name)[()]
value = cast(Union[np.ndarray, np.void], value)
if column_type in (np.ndarray, Embedding):
return value
column_type = cast(
Type[Union[Audio, Image, Mesh, Sequence1D, Video]], column_type
)
return column_type.decode(value)
def _decode_external_value(
self,
value: Union[str, bytes],
column_type: Type[ExternalColumnType],
) -> Optional[ExternalColumnType]:
if not value:
return None
if isinstance(value, bytes):
value = value.decode("utf-8")
file = prepare_path_or_url(value, os.path.dirname(self._filepath))
try:
return column_type.from_file(file)
except Exception:
# No matter what happens, we should not crash, but warn instead.
logger.warning(
f"File or URL {value} either does not exist or could not be "
f"loaded by the class `spotlight.{column_type.__name__}`."
f"Instead of script failure the value will be replaced with "
f"`None`."
)
return None
def _append_internal_columns(self) -> None:
"""
Append internal columns to the first created or imported dataset.
"""
internal_column_values = [self._get_username(), get_current_datetime()]
for column_name, value in zip(INTERNAL_COLUMN_NAMES, internal_column_values):
try:
column = self._h5_file[column_name]
except KeyError:
# Internal column does not exist, create.
value = cast(Union[str, datetime], value)
self.append_column(
column_name, type(value), value if self._length > 0 else None
)
else:
# Internal column exists, check type.
try:
type_name = column.attrs["type"]
except KeyError as e:
raise exceptions.InconsistentDatasetError(
f'Internal column "{column_name}" already exists, but '
f"has no type stored in attributes. Remove or rename "
f"the respective h5 dataset."
) from e
column_type = get_column_type(type_name)
if column_type is not type(value):
raise exceptions.InconsistentDatasetError(
f'Internal column "{column_name}" already exists, '
f"but has invalid type `{column_type}` "
f"(`{type(value)}` expected). Remove or rename "
f"the respective h5 dataset."
)
def _update_internal_columns(
self, index: Optional[Union[IndexType, Indices1dType]] = None
) -> None:
"""
Update internal columns.
Indices should be prepared (slice with positive step or unique sorted sequence).
"""
internal_column_values = [
self._get_username(),
get_current_datetime().isoformat(),
]
for column_name, value in zip(INTERNAL_COLUMN_NAMES, internal_column_values):
if column_name not in self._h5_file:
continue
column = self._h5_file[column_name]
column_length = len(column)
if column_length != self._length:
column.resize(self._length, axis=0)
if column_length < self._length:
# A row/rows appended, append values.
column[column_length - self._length - 1 :] = value
elif column_length == self._length:
if index is None:
# A column appended, update all values.
column[:] = value
else:
# A row/rows chenged, update values according to `index`.
column[index] = value
# Otherwise, all columns deleted. All values removed through resize.
def _update_generation_id(self) -> None:
self._h5_file.attrs["spotlight_generation_id"] += 1
def _rollback(self, length: int) -> None:
"""
Rollback dataset after a failed row/dataset append.
Args:
length: Target length of dataset after rollback.
"""
for column_name in self._column_names:
column = self._h5_file[column_name]
column_length = column.shape[0] if column.shape else 0
if column_length <= length:
continue
column.resize(length, axis=0)
def _resolve_ref(
self, ref: Union[h5py.Reference, str, bytes], column_name: str
) -> h5py.Dataset:
if isinstance(ref, bytes):
ref = ref.decode("utf-8")
if isinstance(ref, str):
return self._h5_file[f"__group__/{column_name}/{ref}"]
return self._h5_file[ref]
@staticmethod
def _get_username() -> str:
return ""
@staticmethod
def _get_column_type(
x: Union[str, h5py.Dataset, h5py.AttributeManager]
) -> Type[ColumnType]:
"""
Get column type by its name, or extract it from `h5py` entities.
"""
if isinstance(x, str):
return get_column_type(x)
if isinstance(x, h5py.Dataset):
return get_column_type(x.attrs["type"])
if isinstance(x, h5py.AttributeManager):
return get_column_type(x["type"])
raise TypeError(
f"Argument is expected to ba an instance of type `str`, `h5py.Dataset` "
f"or `h5py.AttributeManager`, but `x` of type {type(x)} received."
)
@staticmethod
def _get_column_name(column: h5py.Dataset) -> str:
"""
Get name of a column.
"""
return column.name.split("/")[-1]
@staticmethod
def _is_ref_column(column: h5py.Dataset) -> bool:
"""
Check if a column is ref column.
"""
return column.attrs["type"] in REF_COLUMN_TYPE_NAMES and (
h5py.check_string_dtype(column.dtype) or h5py.check_ref_dtype(column.dtype)
)
@staticmethod
def _check_mode(mode: str) -> None:
"""
Check an open mode.
"""
if mode not in ("r", "r+", "w", "w-", "x", "a"):
raise exceptions.InvalidModeError(
f'Open mode should be one of "r", "r+", "w", "w-"/"x" or "a" '
f"but {mode} received."
)
@staticmethod
def check_column_name(name: str) -> None:
"""
Check a column name.
"""
if not isinstance(name, str):
raise exceptions.InvalidColumnNameError(
f"Column name should be a string, but value {name} of type "
f"`{type(name)}` received."
)
if "/" in name:
raise exceptions.InvalidColumnNameError(
f'Column name should not contain "/", but "{name}" received.'
)
if name == "__group__":
raise exceptions.InvalidColumnNameError(
'Column name "__group__" is reserved for internal use.'
)
def _is_writable(self) -> bool:
"""
Check whether dataset is writable.
"""
return self._mode != "r"
def _assert_is_opened(self) -> None:
if self._closed:
raise exceptions.ClosedDatasetError("Dataset is closed.")
def _assert_is_writable(self) -> None:
self._assert_is_opened()
if not self._is_writable():
raise exceptions.ReadOnlyDatasetError("Dataset is read-only.")
def _assert_column_not_exists(self, name: str) -> None:
if name in self._column_names:
raise exceptions.ColumnExistsError(f'Column "{name}" already exists.')
if name in self._h5_file:
raise exceptions.ColumnExistsError(
f'"{name}" name is already used in the H5 file '
f'"{self._filepath}" (but not as a Spotlight dataset column).'
)
def _assert_column_exists(
self, name: str, check_type: bool = False, internal: bool = False
) -> None:
if check_type and not isinstance(name, str):
raise TypeError(
f"Column name should be a string, but value {name} of type "
f"`{type(name)}` received.`"
)
column_names = (
self._column_names
if not internal
else self._column_names.union(INTERNAL_COLUMN_NAMES)
)
if name not in column_names:
raise exceptions.ColumnNotExistsError(f'Column "{name}" does not exist.')
def _assert_index_exists(self, index: IndexType, check_type: bool = False) -> None:
if check_type and not is_integer(index):
raise TypeError(
f"Dataset index should be an integer, but value {index} of "
f"type `{type(index)}` received."
)
if index < -self._length or index >= self._length:
raise exceptions.InvalidIndexError(
f"Row {index} does not exist, dataset has length {self._length}."
)
def _assert_valid_or_set_value_dtype(
self, dtype: np.dtype, column: h5py.Dataset
) -> None:
attrs = column.attrs
if "value_dtype" in attrs:
if dtype.str != attrs["value_dtype"]:
column_name = self._get_column_name(column)
raise exceptions.InvalidDTypeError(
f'Values for {attrs["type"]} column "{column_name}" '
f"should have dtype `{np.dtype(attrs['value_dtype'])}`, "
f"but value with dtype `{dtype}` received."
)
elif issubclass(dtype.type, (np.bool_, np.number)):
attrs["value_dtype"] = dtype.str
else:
column_name = self._get_column_name(column)
raise exceptions.InvalidDTypeError(
f'Values for {attrs["type"]} column "{column_name}" should '
f"have numeric or boolean dtype, but value with dtype {dtype} "
f"received."
)
def _assert_valid_or_set_embedding_shape(
self, shape: Tuple[int, ...], column: h5py.Dataset
) -> None:
attrs = column.attrs
if shape == (0,) and attrs.get("optional", False):
# Do not check shape if an empty array given for an optional column.
return
if "value_shape" in attrs:
if shape != attrs["value_shape"]:
column_name = self._get_column_name(column)
raise exceptions.InvalidShapeError(
f'Values for `Embedding` column "{column_name}" '
f'should have shape {attrs["value_shape"]}, but '
f"value with shape {shape} received."
)
elif len(shape) == 1 and shape[0] > 0:
attrs["value_shape"] = shape
else:
column_name = self._get_column_name(column)
raise exceptions.InvalidShapeError(
f'Values for `Embedding` column "{column_name}" should '
f"have shape `(num_features,)`, `num_features > 0`, "
f"but value with shape {shape} received."
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dataset/__init__.py | 0.709221 | 0.289981 | __init__.py | pypi |
from typing import Tuple
import numpy as np
from scipy import interpolate, signal
from skimage.color import rgba2rgb, rgb2gray
from skimage.transform import resize_local_mean
from renumics.spotlight import (
Audio,
Dataset,
Embedding,
Image,
Sequence1D,
Window,
)
from renumics.spotlight.dataset import exceptions
def align_audio_data(dataset: Dataset, column: str) -> Tuple[np.ndarray, np.ndarray]:
"""
Align data from an audio column.
"""
column_type = dataset.get_column_type(column)
if column_type is not Audio:
raise exceptions.InvalidDTypeError(
f'An audio column expected, but column "{column}" of type {column_type} received.'
)
notnull_mask = dataset.notnull(column)
if notnull_mask.sum() == 0:
# No data to interpolate, so either dataset is empty, or no samples are valid.
return np.empty((0, 0), dtype=np.float64), notnull_mask
raw_data = dataset[column, notnull_mask]
sampling_rates = []
steps = []
for sample in raw_data:
sampling_rates.append(sample.sampling_rate)
steps.append(len(sample.data))
# Resample to the least frequent audio sample.
target_sampling_rate = min(sampling_rates)
lengths = np.array(steps) / np.array(sampling_rates)
target_length = lengths.max()
target_steps = int(np.ceil(target_length * target_sampling_rate))
data = []
for sample, sample_steps in zip(raw_data, steps):
y = sample.data
if np.issubdtype(y.dtype, np.integer):
max_value = 2 ** (np.iinfo(y.dtype).bits - 1)
y = y.astype(np.float64) / max_value
if y.ndim > 1:
y = y.mean(axis=-1)
sampling_rate = sample.sampling_rate
if sampling_rate != target_sampling_rate:
ratio = target_sampling_rate / sampling_rate
n_samples = int(np.ceil(sample_steps * ratio))
y = signal.resample(y, n_samples, axis=-1)
padding = np.zeros(target_steps - sample_steps, dtype=y.dtype)
y = np.append(y, padding)
data.append(y)
return np.stack(data), notnull_mask
def align_embedding_data(
dataset: Dataset, column: str
) -> Tuple[np.ndarray, np.ndarray]:
"""
Align data from an embedding column.
"""
column_type = dataset.get_column_type(column)
if column_type is not Embedding:
raise exceptions.InvalidDTypeError(
f'An embedding column expected, but column "{column}" of type {column_type} received.'
)
notnull_mask = dataset.notnull(column)
if notnull_mask.sum() == 0:
# No data to interpolate, so either dataset is empty, or no samples are valid.
return np.empty((0, 0), dtype=np.float64), notnull_mask
return np.array(list(dataset[column, notnull_mask])), notnull_mask
def align_image_data(dataset: Dataset, column: str) -> Tuple[np.ndarray, np.ndarray]:
"""
Align data from an image column.
"""
column_type = dataset.get_column_type(column)
if column_type is not Image:
raise exceptions.InvalidDTypeError(
f'An image column expected, but column "{column}" of type {column_type} received.'
)
notnull_mask = dataset.notnull(column)
if notnull_mask.sum() == 0:
# No data to interpolate, so either dataset is empty, or no samples are valid.
return np.empty(0, dtype=np.float64), notnull_mask
raw_data = dataset[column, notnull_mask]
min_height, min_width = np.inf, np.inf
for sample in raw_data:
height, width = sample.data.shape[:2]
min_height = min(min_height, height)
min_width = min(min_width, width)
data = []
for sample in raw_data:
y = sample.data
height, width = y.shape[:2]
if np.issubdtype(y.dtype, np.integer):
y = y.astype(np.float64) / 255
if y.ndim > 2:
channels = y.shape[2]
if channels == 1:
y = y[:, :]
elif channels == 3:
y = rgb2gray(y)
elif channels == 4:
y = rgb2gray(rgba2rgb(y))
if height != min_height or width != min_width:
y = resize_local_mean(y, (min_height, min_width), preserve_range=True)
y = y.flatten()
data.append(y)
return np.array(data), notnull_mask
def align_sequence_1d_data(
dataset: Dataset, column: str
) -> Tuple[np.ndarray, np.ndarray]:
"""
Align data from an sequence 1D column.
"""
column_type = dataset.get_column_type(column)
if column_type is not Sequence1D:
raise exceptions.InvalidDTypeError(
f'A sequence 1D column expected, but column "{column}" of type {column_type} received.'
)
notnull_mask = dataset.notnull(column)
if notnull_mask.sum() == 0:
# No data to interpolate, so either dataset is empty, or no samples are valid.
return np.empty((0, 0), dtype=np.float64), notnull_mask
raw_data = dataset[column, notnull_mask]
timesteps = np.empty(0, dtype=raw_data[0].index.dtype)
for sample in raw_data:
timesteps = np.union1d(timesteps, sample.index)
# Space the overall count of unique timesteps evenly over the maximal time range.
x = np.linspace(timesteps.min(), timesteps.max(), len(timesteps))
data = []
for sample in raw_data:
f = interpolate.interp1d(
sample.index,
sample.value,
kind="linear",
copy=False,
bounds_error=False,
fill_value=0,
assume_sorted=False,
)
y = f(x)
data.append(y)
return np.stack(data), notnull_mask
def align_column_data(
dataset: Dataset, column: str, allow_nan: bool = False
) -> Tuple[np.ndarray, np.ndarray]:
"""
Align data from an Spotlight dataset column if possible.
"""
column_type = dataset.get_column_type(column)
if column_type is Audio:
data, mask = align_audio_data(dataset, column)
elif column_type is Embedding:
data, mask = align_embedding_data(dataset, column)
elif column_type is Image:
data, mask = align_image_data(dataset, column)
elif column_type is Sequence1D:
data, mask = align_sequence_1d_data(dataset, column)
elif column_type in (bool, int, float, Window):
data = dataset[column].astype(np.float64).reshape((len(dataset), -1))
mask = np.full(len(dataset), True)
else:
raise NotImplementedError(f"{column_type} column currently not supported.")
if not allow_nan:
# Remove "rows" with `NaN`s.
finite_mask = np.isfinite(data).all(axis=1)
if not finite_mask.all():
indices = np.arange(len(dataset))
indices = indices[mask]
indices = indices[finite_mask]
mask = np.full(len(dataset), False)
mask[indices] = True
data = data[finite_mask]
return data, mask | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dataset/descriptors/data_alignment.py | 0.878184 | 0.676166 | data_alignment.py | pypi |
from typing import List, Optional, Tuple
import numpy as np
import pycatch22
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from renumics.spotlight.dataset import Dataset
from renumics.spotlight.dataset.exceptions import ColumnExistsError, InvalidDTypeError
from renumics.spotlight.dtypes import Audio, Sequence1D
from .data_alignment import align_column_data
def pca(
dataset: Dataset,
column: str,
n_components: int = 8,
inplace: bool = False,
suffix: str = "pca",
overwrite: bool = False,
) -> Optional[Tuple[np.ndarray, np.ndarray]]:
"""
Generate PCA embeddings for the given column of a dataset and
optionally write them back into dataset.
"""
embedding_column_name = f"{column}-{suffix}"
if inplace and not overwrite and embedding_column_name in dataset.keys():
raise ColumnExistsError(
f'Column "{embedding_column_name}" already exists. Either set '
f"another `suffix` argument or set `overwrite` argument to `True`."
)
data, mask = align_column_data(dataset, column, allow_nan=False)
if inplace:
if overwrite and embedding_column_name in dataset.keys():
del dataset[embedding_column_name]
dataset.append_embedding_column(embedding_column_name, optional=True)
if len(data) == 0:
if inplace:
return None
return np.empty((0, n_components), dtype=data.dtype), np.full(
len(dataset), False
)
data = StandardScaler().fit_transform(data)
embeddings = PCA(n_components=n_components).fit_transform(data)
if inplace:
dataset[embedding_column_name, mask] = embeddings
return None
return embeddings, mask
def get_catch22_feature_names(catch24: bool = False) -> List[str]:
"""
Get Catch22 feature names in the same order as returned by :func:`catch22`.
"""
# Run Catch22 with dummy data to get feature names.
return pycatch22.catch22_all([0], catch24)["names"]
def catch22(
dataset: Dataset,
column: str,
catch24: bool = False,
inplace: bool = False,
suffix: Optional[str] = None,
overwrite: bool = False,
as_float_columns: bool = False,
) -> Optional[Tuple[np.ndarray, np.ndarray]]:
"""
Generate Catch22 embeddings for the given column of a dataset and
optionally write them back into dataset.
"""
if suffix is None:
suffix = "catch24" if catch24 else "catch22"
column_type = dataset.get_column_type(column)
if column_type not in (Audio, Sequence1D):
raise InvalidDTypeError(
f"catch22 is only applicable to columns of type `Audio` and "
f'`Sequence1D`, but column "{column}" of type {column_type} received.'
)
column_names = []
if as_float_columns:
for name in get_catch22_feature_names(catch24):
column_names.append("-".join((column, suffix, name)))
else:
column_names.append(f"{column}-{suffix}")
if inplace and not overwrite:
for name in column_names:
if name in dataset.keys():
raise ColumnExistsError(
f'Column "{name}" already exists. Either set another '
f"`suffix` argument or set `overwrite` argument to `True`."
)
data, mask = align_column_data(dataset, column, allow_nan=False)
if inplace:
if overwrite:
for name in column_names:
if name in dataset.keys():
del dataset[name]
for name in column_names:
if as_float_columns:
dataset.append_float_column(name, optional=True)
else:
dataset.append_embedding_column(name, optional=True)
if len(data) == 0:
if inplace:
return None
return np.empty((0, 24 if catch24 else 22), dtype=data.dtype), np.full(
len(dataset), False
)
embeddings = np.array(
[pycatch22.catch22_all(sample, catch24)["values"] for sample in data],
dtype=float,
)
if inplace:
if as_float_columns:
for name, values in zip(column_names, embeddings.T):
dataset[name, mask] = values
else:
dataset[column_names[0], mask] = embeddings
return None
return embeddings, mask | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dataset/descriptors/__init__.py | 0.927394 | 0.602939 | __init__.py | pypi |
import inspect
from typing import Iterable
import numpy as np
import cleanlab.outlier
from renumics.spotlight.dtypes import Embedding
from renumics.spotlight.backend.data_source import DataSource
from renumics.spotlight.dtypes.typing import ColumnTypeMapping
from ..decorator import data_analyzer
from ..typing import DataIssue
@data_analyzer
def analyze_with_cleanlab(
data_source: DataSource, dtypes: ColumnTypeMapping
) -> Iterable[DataIssue]:
"""
Find (embedding) outliers with cleanlab
"""
embedding_columns = (col for col, dtype in dtypes.items() if dtype == Embedding)
for column_name in embedding_columns:
col_values = data_source.get_column(column_name, dtypes[column_name]).values
embeddings = np.array(col_values, dtype=object)
mask = _detect_outliers(embeddings)
rows = np.where(mask)[0].tolist()
if len(rows):
yield DataIssue(
severity="medium",
title="Outliers in embeddings",
rows=rows,
columns=[column_name],
description=inspect.cleandoc(
"""
There are outliers in one of your embedding columns.
Here are a few issues that outliers might indicate:
1. Data entry or measurement errors
2. Feature engineering issues, like missing normalization
3. Sampling bias in your dataset
"""
),
)
def _calculate_outlier_scores(embeddings: np.ndarray) -> np.ndarray:
"""
calculate outlier scores for an embedding column
"""
if embeddings.ndim > 1:
mask = np.ones(len(embeddings), dtype=bool)
else:
mask = np.array([value is not None for value in embeddings])
features = np.stack(embeddings[mask]) # type: ignore
scores = np.full(shape=(len(embeddings),), fill_value=np.nan)
# cleanlab's ood needs at least 10 features
# Don't calculate any scores if we have less than 10 embeddings.
if len(features) < 10:
return scores
scores[mask] = cleanlab.outlier.OutOfDistribution().fit_score(
features=features, verbose=False
)
return scores
def _detect_outliers(embeddings: np.ndarray) -> np.ndarray:
"""
detect outliers in an embedding column
"""
scores = _calculate_outlier_scores(embeddings)
return scores < 0.50 | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/analysis/analyzers/cleanlab.py | 0.810516 | 0.552479 | cleanlab.py | pypi |
import os
import inspect
from typing import Iterable, List, Optional, Type
from pathlib import Path
from tempfile import TemporaryDirectory
from contextlib import redirect_stderr, redirect_stdout
import numpy as np
import cleanvision
from renumics.spotlight.backend.data_source import DataSource
from renumics.spotlight.backend.exceptions import ConversionFailed
from renumics.spotlight.dtypes.typing import ColumnTypeMapping, ColumnType
from renumics.spotlight.dtypes import Image
from ..decorator import data_analyzer
from ..typing import DataIssue
_issue_types = {
"is_light_issue": (
"Bright images",
"medium",
"""
Some images in your dataset are unusually bright.
""",
),
"is_dark_issue": (
"Dark images",
"medium",
"""
Some images in your dataset are unusually dark.
""",
),
"is_blurry_issue": (
"Blurry images",
"medium",
"""
Some images in your dataset are blurry.
""",
),
"is_exact_duplicates_issue": (
"Exact duplicates",
"high",
"""
Some images in your dataset are exact duplicates.
""",
),
"is_near_duplicates_issue": (
"Near duplicates",
"medium",
"""
Some images in your dataset are near duplicates.
""",
),
"is_odd_aspect_ratio_issue": (
"Odd aspect ratio",
"medium",
"""
Some images in your dataset have an odd aspect ratio.
""",
),
"is_low_information_issue": (
"Low information",
"medium",
"""
Some images in your dataset have low information content.
""",
),
"is_grayscale_issue": (
"Grayscale images",
"medium",
"""
Some images in your dataset are grayscale.
""",
),
}
def _make_issue(cleanvision_key: str, column: str, rows: List[int]) -> DataIssue:
title, severity, description = _issue_types[cleanvision_key]
return DataIssue(
severity=severity,
title=title,
rows=rows,
columns=[column],
description=inspect.cleandoc(description),
)
def _get_cell_data_safe(
data_source: DataSource, column_name: str, row: int, dtype: Type[ColumnType]
) -> Optional[bytes]:
try:
return data_source.get_cell_data(column_name, row, dtype)
except ConversionFailed:
return None
@data_analyzer
def analyze_with_cleanvision(
data_source: DataSource, dtypes: ColumnTypeMapping
) -> Iterable[DataIssue]:
"""
find image issues using cleanvision
"""
image_columns = [col for col, dtype in dtypes.items() if dtype == Image]
for column_name in image_columns:
# load image data from data source
images = (
_get_cell_data_safe(data_source, column_name, row, dtypes[column_name])
for row in range(len(data_source))
)
# Write images to temporary directory for cleanvision.
# They alsow support huggingface's image format.
# Maybe use that in the future where applicable.
indices_list = []
image_paths = []
with TemporaryDirectory() as tmp:
for i, image_data in enumerate(images):
if not image_data:
continue
path = Path(tmp) / f"{i}.png"
path.write_bytes(image_data)
image_paths.append(str(path))
indices_list.append(i)
if len(image_paths) == 0:
continue
with open(os.devnull, "w", encoding="utf-8") as devnull:
with redirect_stdout(devnull), redirect_stderr(devnull):
lab = cleanvision.Imagelab(filepaths=image_paths)
lab.find_issues()
analysis = lab.issues
indices = np.array(indices_list)
# Iterate over all the different issue types
# and convert them to our internal DataIssue format.
for cleanvision_key in _issue_types:
rows = indices[analysis[cleanvision_key]].tolist()
if rows:
yield _make_issue(cleanvision_key, column_name, rows) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/analysis/analyzers/cleanvision.py | 0.696681 | 0.415314 | cleanvision.py | pypi |
import asyncio
import dataclasses
import functools
import json
from typing import Any, List, Optional, Set, Callable
from fastapi import WebSocket, WebSocketDisconnect
from loguru import logger
from pydantic.dataclasses import dataclass
from typing_extensions import Literal
from renumics.spotlight.dtypes.typing import ColumnTypeMapping
from .data_source import DataSource, sanitize_values
from .tasks import TaskManager, TaskCancelled
from .tasks.reduction import compute_umap, compute_pca
from .exceptions import GenerationIDMismatch
@dataclass
class Message:
"""
Common websocket message model.
"""
type: str
data: Any
@dataclass
class RefreshMessage(Message):
"""
Refresh message model
"""
type: Literal["refresh"] = "refresh"
data: Any = None
@dataclass
class ResetLayoutMessage(Message):
"""
Reset layout message model
"""
type: Literal["resetLayout"] = "resetLayout"
data: Any = None
@dataclass
class ReductionMessage(Message):
"""
Common data reduction message model.
"""
widget_id: str
uid: str
generation_id: int
@dataclass
class ReductionRequestData:
"""
Base data reduction request payload.
"""
indices: List[int]
columns: List[str]
@dataclass
class UMapRequestData(ReductionRequestData):
"""
U-Map request payload.
"""
n_neighbors: int
metric: str
min_dist: float
@dataclass
class PCARequestData(ReductionRequestData):
"""
PCA request payload.
"""
normalization: str
@dataclass
class UMapRequest(ReductionMessage):
"""
U-Map request model.
"""
data: UMapRequestData
@dataclass
class PCARequest(ReductionMessage):
"""
PCA request model.
"""
data: PCARequestData
@dataclass
class ReductionResponseData:
"""
Data reduction response payload.
"""
indices: List[int]
points: List[List[float]]
@dataclass
class ReductionResponse(ReductionMessage):
"""
Data reduction response model.
"""
data: ReductionResponseData
MESSAGE_BY_TYPE = {
"umap": UMapRequest,
"umap_result": ReductionResponse,
"pca": PCARequest,
"pca_result": ReductionResponse,
"refresh": RefreshMessage,
}
class UnknownMessageType(Exception):
"""
Websocket message type is unknown.
"""
def parse_message(raw_message: str) -> Message:
"""
Parse a websocket message from a raw text.
"""
json_message = json.loads(raw_message)
message_type = json_message["type"]
message_class = MESSAGE_BY_TYPE.get(message_type)
if message_class is None:
raise UnknownMessageType(f"Message type {message_type} is unknown.")
return message_class(**json_message)
@functools.singledispatch
async def handle_message(request: Message, connection: "WebsocketConnection") -> None:
"""
Handle incoming messages.
New message types should be registered by decorating with `@handle_message.register`.
"""
raise NotImplementedError
class WebsocketConnection:
"""
Wraps websocket and dispatches messages to message handlers.
"""
websocket: WebSocket
manager: "WebsocketManager"
def __init__(self, websocket: WebSocket, manager: "WebsocketManager") -> None:
self.websocket = websocket
self.manager = manager
async def send_async(self, message: Message) -> None:
"""
Send a message async.
"""
try:
await self.websocket.send_json(dataclasses.asdict(message))
except WebSocketDisconnect:
self._on_disconnect()
except RuntimeError:
# connection already disconnected
pass
def send(self, message: Message) -> None:
"""
Send a message without async.
"""
self.manager.loop.create_task(self.send_async(message))
def _on_disconnect(self) -> None:
self.manager.on_disconnect(self)
self.task_manager.cancel(tag=id(self))
async def listen(self) -> None:
"""
Wait for websocket connection and handle incoming messages.
"""
await self.websocket.accept()
self.manager.on_connect(self)
try:
while True:
try:
message = parse_message(await self.websocket.receive_text())
except UnknownMessageType as e:
logger.warning(str(e))
else:
logger.info(f"WS message with type {message.type} received.")
asyncio.create_task(handle_message(message, self))
except WebSocketDisconnect:
self._on_disconnect()
@property
def task_manager(self) -> TaskManager:
"""
The app's task manager
"""
return self.websocket.app.task_manager
ConnectCallback = Callable[[int], None]
class WebsocketManager:
"""
Manages websocket connections.
"""
loop: asyncio.AbstractEventLoop
connections: Set[WebsocketConnection]
_disconnect_callbacks: Set[ConnectCallback]
_connect_callbacks: Set[ConnectCallback]
def __init__(self, loop: asyncio.AbstractEventLoop) -> None:
self.loop = loop
self.connections = set()
self._disconnect_callbacks = set()
self._connect_callbacks = set()
def create_connection(self, websocket: WebSocket) -> WebsocketConnection:
"""
Create a new websocket connection.
"""
return WebsocketConnection(websocket, self)
def broadcast(self, message: Message) -> None:
"""
Send message to all connected clients.
"""
for connection in self.connections:
connection.send(message)
def add_disconnect_callback(self, callback: ConnectCallback) -> None:
"""
Add a listener that is notified after a connection terminates
"""
self._disconnect_callbacks.add(callback)
def remove_disconnect_callback(self, callback: ConnectCallback) -> None:
"""
Remove the provided listener if possible
"""
try:
self._disconnect_callbacks.remove(callback)
except KeyError:
pass
def add_connect_callback(self, callback: ConnectCallback) -> None:
"""
Add a listener that is notified after a connection is initialized
"""
self._connect_callbacks.add(callback)
def remove_connect_callback(self, callback: ConnectCallback) -> None:
"""
Remove the provided listener if possible
"""
try:
self._connect_callbacks.remove(callback)
except KeyError:
pass
def on_connect(self, connection: WebsocketConnection) -> None:
"""
Handle a new conection
"""
self.connections.add(connection)
for callback in self._connect_callbacks:
callback(len(self.connections))
def on_disconnect(self, connection: WebsocketConnection) -> None:
"""
Handle the termination of an existing connection
"""
self.connections.remove(connection)
for callback in self._disconnect_callbacks:
callback(len(self.connections))
@handle_message.register
async def _(request: UMapRequest, connection: "WebsocketConnection") -> None:
table: Optional[DataSource] = connection.websocket.app.data_source
dtypes: ColumnTypeMapping = connection.websocket.app.dtypes
if table is None:
return None
try:
table.check_generation_id(request.generation_id)
except GenerationIDMismatch:
return
try:
points, valid_indices = await connection.task_manager.run_async(
compute_umap,
(
table,
dtypes,
request.data.columns,
request.data.indices,
request.data.n_neighbors,
request.data.metric,
request.data.min_dist,
),
name=request.widget_id,
tag=id(connection),
)
except TaskCancelled:
...
else:
response = ReductionResponse(
type="umap_result",
widget_id=request.widget_id,
uid=request.uid,
generation_id=request.generation_id,
data=ReductionResponseData(
indices=valid_indices, points=sanitize_values(points)
),
)
await connection.send_async(response)
@handle_message.register
async def _(request: PCARequest, connection: "WebsocketConnection") -> None:
table: Optional[DataSource] = connection.websocket.app.data_source
dtypes: ColumnTypeMapping = connection.websocket.app.dtypes
if table is None:
return None
try:
table.check_generation_id(request.generation_id)
except GenerationIDMismatch:
return
try:
points, valid_indices = await connection.task_manager.run_async(
compute_pca,
(
table,
dtypes,
request.data.columns,
request.data.indices,
request.data.normalization,
),
name=request.widget_id,
tag=id(connection),
)
except TaskCancelled:
...
else:
response = ReductionResponse(
type="pca_result",
widget_id=request.widget_id,
uid=request.uid,
generation_id=request.generation_id,
data=ReductionResponseData(
indices=valid_indices, points=sanitize_values(points)
),
)
await connection.send_async(response) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/websockets.py | 0.831177 | 0.20834 | websockets.py | pypi |
import json
from typing import Dict, Union, cast, Optional
from databases import Database
from renumics.spotlight import appdirs
ConfigValue = Union[Dict, str, int, float, bool]
class Config:
"""
Configuration object that stores key value pairs in an sqlite db on disk.
"""
def __init__(self) -> None:
appdirs.config_dir.mkdir(parents=True, exist_ok=True)
db_path = appdirs.config_dir / "config.db"
db_url = f"sqlite:///{db_path}"
self.database = Database(db_url)
self.connected = False
async def _lazy_init(self) -> None:
"""
lazy initialization function
"""
if not self.connected:
await self.database.connect()
query = """
CREATE TABLE IF NOT EXISTS entries (
name TEXT NOT NULL,
value JSON NOT NULL,
user TEXT,
dataset TEXT,
PRIMARY KEY (name, user, dataset)
)
"""
await self.database.execute(query)
self.connected = True
async def get(
self, name: str, *, dataset: str = "", user: str = ""
) -> Optional[ConfigValue]:
"""
get a stored value by name
"""
await self._lazy_init()
query = """
SELECT value FROM entries
WHERE name = :name AND user = :user AND dataset = :dataset
"""
row = await self.database.fetch_one(
query, {"name": name, "user": user, "dataset": dataset}
)
if not row:
return None
value = json.loads(row[0])
return cast(ConfigValue, value)
async def set(
self, name: str, value: ConfigValue, *, dataset: str = "", user: str = ""
) -> None:
"""
set a config value by name
"""
await self._lazy_init()
values = {
"name": name,
"user": user,
"dataset": dataset,
"value": json.dumps(value),
}
query = """
REPLACE INTO entries (name, value, user, dataset)
VALUES (:name, :value, :user, :dataset)
"""
await self.database.execute(query, values)
async def remove(self, name: str, *, dataset: str = "", user: str = "") -> None:
"""
remove a config value by name
"""
await self._lazy_init()
query = """
DELETE FROM entries
WHERE name = :name AND user = :user AND dataset = :dataset
"""
await self.database.execute(
query, values={"name": name, "user": user, "dataset": dataset}
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/config.py | 0.762733 | 0.16807 | config.py | pypi |
import dataclasses
import hashlib
import io
from datetime import datetime
from abc import ABC, abstractmethod
from typing import Optional, List, Dict, Type, Any, Union, cast
import pandas as pd
import numpy as np
from pydantic.dataclasses import dataclass
from renumics.spotlight.io import audio
from renumics.spotlight.typing import is_iterable
from renumics.spotlight.dataset.exceptions import (
ColumnExistsError,
ColumnNotExistsError,
)
from renumics.spotlight.dtypes import Audio
from renumics.spotlight.dtypes.typing import (
ColumnType,
ColumnTypeMapping,
)
from renumics.spotlight.dtypes.conversion import ConvertedValue
from renumics.spotlight.cache import external_data_cache
from .exceptions import DatasetNotEditable, GenerationIDMismatch, NoRowFound
@dataclasses.dataclass
class Attrs:
"""
Column attributes relevant for Spotlight.
"""
type: Type[ColumnType]
order: Optional[int]
hidden: bool
optional: bool
description: Optional[str]
tags: List[str]
editable: bool
# data type specific fields
categories: Optional[Dict[str, int]]
x_label: Optional[str]
y_label: Optional[str]
embedding_length: Optional[int]
@dataclasses.dataclass
class Column(Attrs):
"""
Column with raw values.
"""
name: str
values: Union[np.ndarray, List[ConvertedValue]]
@dataclass
class CellsUpdate:
"""
A dataset's cell update.
"""
value: Any
author: str
edited_at: str
class DataSource(ABC):
"""abstract base class for different data sources"""
@abstractmethod
def __init__(self, source: Any):
"""
Create Data Source from matching source and dtype mapping.
"""
@property
@abstractmethod
def column_names(self) -> List[str]:
"""
Dataset's available column names.
"""
@property
def df(self) -> Optional[pd.DataFrame]:
"""
Get the source data as a pandas dataframe if possible
"""
return None
@abstractmethod
def __len__(self) -> int:
"""
Get the table's length.
"""
@abstractmethod
def get_generation_id(self) -> int:
"""
Get the table's generation ID.
"""
def check_generation_id(self, generation_id: int) -> None:
"""
Check if table's generation ID matches to the given one.
"""
if self.get_generation_id() != generation_id:
raise GenerationIDMismatch()
@abstractmethod
def guess_dtypes(self) -> ColumnTypeMapping:
"""
Guess data source's dtypes.
"""
@abstractmethod
def get_uid(self) -> str:
"""
Get the table's unique ID.
"""
@abstractmethod
def get_name(self) -> str:
"""
Get the table's human-readable name.
"""
def get_internal_columns(self) -> List[Column]:
"""
Get internal columns if there are any.
"""
return []
@abstractmethod
def get_column(
self,
column_name: str,
dtype: Type[ColumnType],
indices: Optional[List[int]] = None,
simple: bool = False,
) -> Column:
"""
Get column metadata + values
"""
@abstractmethod
def get_cell_data(
self, column_name: str, row_index: int, dtype: Type[ColumnType]
) -> Any:
"""
return the value of a single cell
"""
def get_waveform(self, column_name: str, row_index: int) -> Optional[np.ndarray]:
"""
return the waveform of an audio cell
"""
blob = self.get_cell_data(column_name, row_index, Audio)
if blob is None:
return None
if isinstance(blob, np.void):
blob = blob.tolist()
value_hash = hashlib.blake2b(blob).hexdigest()
cache_key = f"waveform-v2:{value_hash}"
try:
waveform = external_data_cache[cache_key]
return waveform
except KeyError:
...
waveform = audio.get_waveform(io.BytesIO(blob))
external_data_cache[cache_key] = waveform
return waveform
def replace_cells(
self, column_name: str, indices: List[int], value: Any, dtype: Type[ColumnType]
) -> CellsUpdate:
"""
replace multiple cell's value
"""
raise DatasetNotEditable()
def delete_column(self, name: str) -> None:
"""
remove a column from the table
"""
raise DatasetNotEditable()
def delete_row(self, index: int) -> None:
"""
remove a row from the table
"""
raise DatasetNotEditable()
def duplicate_row(self, index: int) -> int:
"""
duplicate a row in the table
"""
raise DatasetNotEditable()
def append_column(self, name: str, dtype_name: str) -> Column:
"""
add a column to the table
"""
raise DatasetNotEditable()
def _assert_index_exists(self, index: int) -> None:
if index < -len(self) or index >= len(self):
raise NoRowFound(index)
def _assert_indices_exist(self, indices: List[int]) -> None:
indices_array = np.array(indices)
if ((indices_array < -len(self)) | (indices_array >= len(self))).any():
raise NoRowFound()
def _assert_column_exists(self, column_name: str) -> None:
if column_name not in self.column_names:
raise ColumnNotExistsError(
f"Column '{column_name}' doesn't exist in the dataset."
)
def _assert_column_not_exists(self, column_name: str) -> None:
if column_name in self.column_names:
raise ColumnExistsError(
f"Column '{column_name}' already exists in the dataset."
)
def _sanitize_value(value: Any) -> Any:
if pd.isna(value):
return None
if isinstance(value, (bool, int, float, str, bytes)):
return value
try:
# Assume `value` is a `numpy` object.
return value.tolist()
except AttributeError:
# Try to send `value` as is.
return value
def sanitize_values(values: Any) -> Any:
"""
sanitize values for serialization
e.g. replace inf, -inf and NaN in float data
"""
if not is_iterable(values):
return _sanitize_value(values)
if isinstance(values, list):
return [sanitize_values(x) for x in values]
# At the moment, `values` should be a `numpy` array.
values = cast(np.ndarray, values)
if issubclass(values.dtype.type, np.inexact):
return np.where(np.isfinite(values), values, np.array(None)).tolist()
return values.tolist()
def idx_column(row_count: int) -> Column:
"""create a column containing the index"""
return Column(
type=int,
order=None,
description=None,
tags=[],
categories=None,
x_label=None,
y_label=None,
embedding_length=None,
name="__idx__",
hidden=True,
editable=False,
optional=False,
values=np.array(range(row_count)),
)
def last_edited_at_column(row_count: int, value: Optional[datetime]) -> Column:
"""create a column containing a constant datetime"""
return Column(
type=datetime,
order=None,
description=None,
tags=[],
categories=None,
x_label=None,
y_label=None,
embedding_length=None,
name="__last_edited_at__",
hidden=True,
editable=False,
optional=False,
values=np.array([value] * row_count, dtype=object),
)
def last_edited_by_column(row_count: int, value: str) -> Column:
"""create a column containing a constant username"""
return Column(
type=str,
order=None,
description=None,
tags=[],
categories=None,
x_label=None,
y_label=None,
embedding_length=None,
name="__last_edited_by__",
hidden=True,
editable=False,
optional=False,
values=np.array(row_count * [value]),
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/data_source.py | 0.878796 | 0.516169 | data_source.py | pypi |
from typing import Any, Optional, Type
from fastapi import status
from renumics.spotlight.dtypes.typing import ColumnType
from renumics.spotlight.typing import IndexType, PathOrUrlType, PathType
class Problem(Exception):
"""
Base API exception.
"""
def __init__(self, title: str, detail: str, status_code: int = 500) -> None:
self.title = title
self.detail = detail
self.status_code = status_code
def __str__(self) -> str:
return f"{self.title}: {self.detail}"
class InvalidPath(Problem):
"""The supported path is outside the project root or points to an incompatible file"""
def __init__(self, path: PathType) -> None:
super().__init__(
"Invalid path", f"Path {path} is not valid.", status.HTTP_403_FORBIDDEN
)
class InvalidDataSource(Problem):
"""The data source can't be opened"""
def __init__(self) -> None:
super().__init__(
"Invalid data source",
"Can't open supplied data source.",
status.HTTP_403_FORBIDDEN,
)
class NoTableFileFound(Problem):
"""raised when the table file could not be found"""
def __init__(self, path: PathType) -> None:
super().__init__(
"Dataset file not found",
f"File {path} does not exist.",
status.HTTP_404_NOT_FOUND,
)
class CouldNotOpenTableFile(Problem):
"""raised when the table file could not be found"""
def __init__(self, path: PathType) -> None:
super().__init__(
"Could not open dataset",
f"File {path} could not be opened.",
status.HTTP_403_FORBIDDEN,
)
class NoRowFound(Problem):
"""raised when a row can't be found in the dataset"""
def __init__(self, index: Optional[IndexType] = None) -> None:
if index is None:
detail = "One of the requested rows does not exist."
else:
detail = f"Row {index} does not exist in the dataset."
super().__init__("Row not found", detail, status.HTTP_404_NOT_FOUND)
class InvalidCategory(Problem):
"""An invalid Category was passed."""
def __init__(self) -> None:
super().__init__(
"Invalid category",
"Given category is not defined.",
status.HTTP_403_FORBIDDEN,
)
class ColumnNotEditable(Problem):
"""Column is not editable"""
def __init__(self, name: str) -> None:
super().__init__(
"Column not editable",
f"Column '{name}' is not editable.",
status.HTTP_403_FORBIDDEN,
)
class InvalidExternalData(Problem):
"""External data is not readable"""
def __init__(self, value: PathOrUrlType) -> None:
super().__init__(
"Invalid external data",
f"Failed to read external data '{value}'.",
status.HTTP_422_UNPROCESSABLE_ENTITY,
)
class GenerationIDMismatch(Problem):
"""
Generation ID does not match to the expected.
"""
def __init__(self) -> None:
super().__init__(
"Dataset out of sync",
"Dataset is out of sync. Refresh the browser.",
status.HTTP_409_CONFLICT,
)
class ConversionFailed(Problem):
"""
Value cannot be converted to the desired dtype.
"""
def __init__(self, dtype: Type[ColumnType], value: Any) -> None:
self.dtype = dtype
self.value = value
super().__init__(
"Type conversion failed",
f"Value of type {type(value)} cannot be converted to type {dtype}.",
status.HTTP_422_UNPROCESSABLE_ENTITY,
)
class DatasetNotEditable(Problem):
"""The dataset is not editable"""
def __init__(self) -> None:
super().__init__(
"Dataset not editable",
"The dataset is not editable.",
status.HTTP_403_FORBIDDEN,
)
class DatasetColumnsNotUnique(Problem):
"""Dataset's columns are not unique"""
def __init__(self) -> None:
super().__init__(
"Dataset columns not unique",
"Dataset's columns are not unique.",
status.HTTP_403_FORBIDDEN,
)
class InvalidLayout(Problem):
"""The layout could not be parsed from the given source"""
def __init__(self) -> None:
super().__init__(
"Invalid layout",
"The layout could not be loaded from given source.",
status.HTTP_500_INTERNAL_SERVER_ERROR,
)
class FilebrowsingNotAllowed(Problem):
"""Filebrowsing is not allowed"""
def __init__(self) -> None:
super().__init__(
"Filebrowsing not allowed",
"Filebrowsing is not allowed.",
status.HTTP_403_FORBIDDEN,
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/exceptions.py | 0.910062 | 0.342819 | exceptions.py | pypi |
import asyncio
import multiprocessing
from concurrent.futures import Future, ProcessPoolExecutor
from concurrent.futures.process import BrokenProcessPool
from typing import Any, Callable, List, Optional, Sequence, TypeVar, Union
from .exceptions import TaskCancelled
from .task import Task
T = TypeVar("T")
class TaskManager:
"""
Handles task creation, deletion and cleanup
"""
tasks: List[Task]
pool: ProcessPoolExecutor
def __init__(self) -> None:
self.tasks = []
self.pool = ProcessPoolExecutor(8, multiprocessing.get_context("spawn"))
def create_task(
self,
func: Callable,
args: Sequence[Any],
name: Optional[str] = None,
tag: Optional[Union[str, int]] = None,
) -> Task:
"""
create and launch a new task
"""
# cancel running task with same name
self.cancel(name=name)
future = self.pool.submit(func, *args)
task = Task(name, tag, future)
self.tasks.append(task)
def _cleanup(_: Future) -> None:
try:
self.tasks.remove(task)
except ValueError:
# task won't be in list when already cancelled. this is fine
pass
future.add_done_callback(_cleanup)
return task
async def run_async(
self,
func: Callable[..., T],
args: Sequence[Any],
name: Optional[str] = None,
tag: Optional[Union[str, int]] = None,
) -> T:
"""
Launch a new task. Await and return result.
"""
task = self.create_task(func, args, name, tag)
try:
return await asyncio.wrap_future(task.future)
except BrokenProcessPool as e:
raise TaskCancelled from e
def cancel(
self, name: Optional[str] = None, tag: Optional[Union[str, int]] = None
) -> None:
"""
Cancel running and queued tasks.
"""
tasks_to_remove = []
if name is not None and tag is not None:
for task in self.tasks:
if task.name == name and task.tag == tag:
tasks_to_remove.append(task)
elif name is not None:
for task in self.tasks:
if task.name == name:
tasks_to_remove.append(task)
elif tag is not None:
for task in self.tasks:
if task.tag == tag:
tasks_to_remove.append(task)
for task in tasks_to_remove:
task.future.cancel()
try:
self.tasks.remove(task)
except ValueError:
# task won't be in list when already cancelled. this is fine
pass
def cancel_all(self) -> None:
"""
Cancel all running and queued tasks.
"""
for task in self.tasks:
task.future.cancel()
self.tasks.clear()
def shutdown(self) -> None:
"""
Shutdown and cleanup tasks and internal process pool.
This task manager instance won't work after this operation!
"""
self.cancel_all()
self.pool.shutdown(wait=True) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/tasks/task_manager.py | 0.756627 | 0.214013 | task_manager.py | pypi |
from typing import List, Tuple, cast
import numpy as np
import pandas as pd
from renumics.spotlight.dataset.exceptions import ColumnNotExistsError
from renumics.spotlight.dtypes import Category, Embedding
from renumics.spotlight.dtypes.typing import ColumnTypeMapping
from ..data_source import DataSource
SEED = 42
class ColumnNotEmbeddable(Exception):
"""
The column is not embeddable
"""
def get_aligned_data(
table: DataSource,
dtypes: ColumnTypeMapping,
column_names: List[str],
indices: List[int],
) -> Tuple[np.ndarray, List[int]]:
"""
Align data from table's columns, remove `NaN`'s.
"""
if not column_names or not indices:
return np.empty(0, np.float64), []
from sklearn import preprocessing
values = []
for column_name in column_names:
column = table.get_column(column_name, dtypes[column_name], indices)
if column.type is Embedding:
if column.embedding_length:
none_replacement = np.full(column.embedding_length, np.nan)
values.append(
np.array(
[
value if value is not None else none_replacement
for value in column.values
]
)
)
elif column.type is Category:
if column.categories:
classes = sorted(column.categories.values())
na_mask = ~np.isin(np.array(column.values), classes)
one_hot_values = preprocessing.label_binarize(
column.values, classes=sorted(column.categories.values())
).astype(float)
one_hot_values[na_mask] = np.nan
values.append(one_hot_values)
else:
values.append(np.full(len(column.values), np.nan))
elif column.type in (int, bool, float):
values.append(np.array(column.values))
else:
raise ColumnNotEmbeddable
data = np.hstack([col.reshape((len(indices), -1)) for col in values])
mask = ~pd.isna(data).any(axis=1)
return data[mask], (np.array(indices)[mask]).tolist()
def compute_umap(
table: DataSource,
dtypes: ColumnTypeMapping,
column_names: List[str],
indices: List[int],
n_neighbors: int,
metric: str,
min_dist: float,
) -> Tuple[np.ndarray, List[int]]:
"""
Prepare data from table and compute U-Map on them.
"""
try:
data, indices = get_aligned_data(table, dtypes, column_names, indices)
except (ColumnNotExistsError, ColumnNotEmbeddable):
return np.empty(0, np.float64), []
if data.size == 0:
return np.empty(0, np.float64), []
from sklearn import preprocessing
if metric == "standardized euclidean":
data = preprocessing.StandardScaler(copy=False).fit_transform(data)
metric = "euclidean"
elif metric == "robust euclidean":
data = preprocessing.RobustScaler(copy=False).fit_transform(data)
metric = "euclidean"
if data.shape[1] == 2:
return data, indices
import umap
embeddings = umap.UMAP(
n_neighbors=n_neighbors, metric=metric, min_dist=min_dist, random_state=SEED
).fit_transform(data)
return cast(np.ndarray, embeddings), indices
def compute_pca(
table: DataSource,
dtypes: ColumnTypeMapping,
column_names: List[str],
indices: List[int],
normalization: str,
) -> Tuple[np.ndarray, List[int]]:
"""
Prepare data from table and compute PCA on them.
"""
from sklearn import preprocessing, decomposition
try:
data, indices = get_aligned_data(table, dtypes, column_names, indices)
except (ColumnNotExistsError, ValueError):
return np.empty(0, np.float64), []
if data.size == 0:
return np.empty(0, np.float64), []
if data.shape[1] == 1:
return np.hstack((data, np.zeros_like(data))), indices
if normalization == "standardize":
data = preprocessing.StandardScaler(copy=False).fit_transform(data)
elif normalization == "robust standardize":
data = preprocessing.RobustScaler(copy=False).fit_transform(data)
reducer = decomposition.PCA(n_components=2, copy=False, random_state=SEED)
embeddings = reducer.fit_transform(data)
return embeddings, indices | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/tasks/reduction.py | 0.838382 | 0.633148 | reduction.py | pypi |
export declare const datakinds: readonly ["int", "float", "bool", "str", "array", "datetime", "Mesh", "Sequence1D", "Embedding", "Image", "Audio", "Video", "Category", "Window", "Unknown"];
export type DataKind = typeof datakinds[number];
export interface BaseDataType<K extends DataKind, B extends boolean = false> {
kind: K;
binary: B;
optional: boolean;
}
export type UnknownDataType = BaseDataType<'Unknown'>;
export type IntegerDataType = BaseDataType<'int'>;
export type FloatDataType = BaseDataType<'float'>;
export type BooleanDataType = BaseDataType<'bool'>;
export type StringDataType = BaseDataType<'str'>;
export type ArrayDataType = BaseDataType<'array'>;
export type DateTimeDataType = BaseDataType<'datetime'>;
export type WindowDataType = BaseDataType<'Window'>;
export type SequenceDataType = BaseDataType<'Sequence1D', true>;
export type MeshDataType = BaseDataType<'Mesh', true>;
export type ImageDataType = BaseDataType<'Image', true>;
export type AudioDataType = BaseDataType<'Audio', true>;
export type VideoDataType = BaseDataType<'Video', true>;
export interface CategoricalDataType extends BaseDataType<'Category'> {
kind: 'Category';
categories: Record<string, number>;
invertedCategories: Record<number, string>;
}
export interface EmbeddingDataType extends BaseDataType<'Embedding'> {
kind: 'Embedding';
embeddingLength: number;
}
export type DataType = UnknownDataType | IntegerDataType | FloatDataType | BooleanDataType | StringDataType | ArrayDataType | DateTimeDataType | MeshDataType | SequenceDataType | EmbeddingDataType | ImageDataType | AudioDataType | VideoDataType | WindowDataType | CategoricalDataType;
export declare const isInteger: (type: DataType) => type is IntegerDataType;
export declare const isFloat: (type: DataType) => type is FloatDataType;
export declare const isBoolean: (type: DataType) => type is BooleanDataType;
export declare const isString: (type: DataType) => type is StringDataType;
export declare const isArray: (type: DataType) => type is ArrayDataType;
export declare const isDateTime: (type: DataType) => type is DateTimeDataType;
export declare const isMesh: (type: DataType) => type is MeshDataType;
export declare const isSequence: (type: DataType) => type is SequenceDataType;
export declare const isEmbedding: (type: DataType) => type is EmbeddingDataType;
export declare const isImage: (type: DataType) => type is ImageDataType;
export declare const isAudio: (type: DataType) => type is AudioDataType;
export declare const isVideo: (type: DataType) => type is VideoDataType;
export declare const isWindow: (type: DataType) => type is WindowDataType;
export declare const isCategorical: (type: DataType) => type is CategoricalDataType;
export declare const isUnknown: (type: DataType) => type is UnknownDataType;
export type NumericalDataType = IntegerDataType | FloatDataType;
export declare const isNumerical: (type: DataType) => type is NumericalDataType;
export type ScalarDataType = NumericalDataType | StringDataType | BooleanDataType;
export declare const isScalar: (type: DataType) => type is ScalarDataType;
export declare function getNullValue(kind: DataKind): number | boolean | string | null; | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/statics/src/datatypes.d.ts | 0.790894 | 0.692063 | datatypes.d.ts | pypi |
import { DataType } from '../../datatypes';
import { TransferFunction } from '../../hooks/useColorTransferFunction';
import { ColumnsStats, DataColumn, DataRow, DataIssue as DataIssue, Filter, IndexArray, TableData } from '../../types';
export type CallbackOrData<T> = ((data: T) => T) | T;
export type Sorting = 'DESC' | 'ASC';
export type DataSelector = 'full' | 'filtered' | 'selected';
export interface Dataset {
uid?: string;
generationID: number;
filename?: string;
loading: boolean;
columnStats: {
full: ColumnsStats;
selected: ColumnsStats;
filtered: ColumnsStats;
};
columns: DataColumn[];
columnsByKey: Record<string, DataColumn>;
columnData: TableData;
isAnalysisRunning: boolean;
issues: DataIssue[];
rowsWithIssues: IndexArray;
colorTransferFunctions: Record<string, {
full: TransferFunction[];
filtered: TransferFunction[];
}>;
recomputeColorTransferFunctions: () => void;
length: number;
indices: Int32Array;
getRow: (index: number) => DataRow;
isIndexSelected: boolean[];
selectedIndices: Int32Array;
isIndexHighlighted: boolean[];
highlightedIndices: Int32Array;
isIndexFiltered: boolean[];
filteredIndices: Int32Array;
sortColumns: Map<DataColumn, Sorting>;
sortBy: (column?: DataColumn, sorting?: Sorting) => void;
columnRelevance: Map<string, number>;
columnRelevanceGeneration: number;
filters: Filter[];
tags: string[];
lastFocusedRow?: number;
openTable: (path: string) => void;
fetch: () => void;
fetchIssues: () => void;
refresh: () => void;
addFilter: (filter: Filter) => void;
removeFilter: (filter: Filter) => void;
toggleFilterEnabled: (filter: Filter) => void;
replaceFilter: (filter: Filter, newFilter: Filter) => void;
selectRows: (rows: CallbackOrData<IndexArray>) => void;
setHighlightedRows: (mask: boolean[]) => void;
highlightRowAt: (rowIndex: number, only?: boolean) => void;
highlightRows: (rows: CallbackOrData<IndexArray>) => void;
dehighlightRowAt: (rowIndex: number) => void;
dehighlightAll: () => void;
relevanceWorker: any;
isComputingRelevance: boolean;
recomputeColumnRelevance: () => void;
focusRow: (row?: number) => void;
}
export declare function convertValue(value: any, type: DataType): any;
export declare function compareColumnOrder(a: DataColumn, b: DataColumn): number;
export declare const useDataset: import("zustand").UseBoundStore<Omit<import("zustand").StoreApi<Dataset>, "subscribe"> & {
subscribe: {
(listener: (selectedState: Dataset, previousSelectedState: Dataset) => void): () => void;
<U>(selector: (state: Dataset) => U, listener: (selectedState: U, previousSelectedState: U) => void, options?: {
equalityFn?: ((a: U, b: U) => boolean) | undefined;
fireImmediately?: boolean | undefined;
} | undefined): () => void;
};
}>; | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/backend/statics/src/stores/dataset/dataset.d.ts | 0.567457 | 0.507385 | dataset.d.ts | pypi |
from pathlib import Path
from typing import Optional
import site
import dataclasses
import toml
from ..settings import settings
def _find_upwards(filename: str, folder: Path) -> Optional[Path]:
"""
Find first matching file upwards in parent folders.
"""
if folder == folder.parent:
return None
path = folder / filename
if path.exists():
return path
return _find_upwards(filename, folder.parent)
@dataclasses.dataclass
class ProjectInfo:
"""
Info about the current dev project
"""
name: str
type: Optional[str]
root: Optional[Path]
def get_project_info() -> ProjectInfo:
"""
Determine location for dev mode
"core": Inside the main spotlight project.
"plugin": Inside a spotlight plugin project.
None: neither.
"""
if not settings.dev:
return ProjectInfo(name="", type=None, root=None)
pyproject_toml = _find_upwards("pyproject.toml", Path.cwd())
if not pyproject_toml:
return ProjectInfo(name="", type=None, root=None)
pyproject_content = toml.load(pyproject_toml)
project_name = pyproject_content["tool"]["poetry"]["name"]
if project_name == "renumics-spotlight":
project_type = "core"
else:
project_type = "plugin"
return ProjectInfo(name=project_name, type=project_type, root=pyproject_toml.parent)
def find_spotlight_repository() -> Optional[Path]:
"""
Find the cloned spotlight repository.
Returns the path to the repo or None, if it could not be located.
"""
project = get_project_info()
if project.type == "core":
# already in the spotlight repo!
return project.root
if project.type == "plugin":
# find .pth file of the editable install, read it and return repo path
for site_packages_folder in site.getsitepackages():
try:
pth = next(Path(site_packages_folder).glob("**/renumics_spotlight.pth"))
return Path(pth.read_text().strip())
except StopIteration:
return None
return None | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/develop/project.py | 0.767951 | 0.225278 | project.py | pypi |
import io
import os
from typing import Dict, IO, Tuple, Union
import av
import numpy as np
import requests
import validators
from renumics.spotlight.requests import headers
from renumics.spotlight.typing import FileType
# Some AV warning messages have ERROR level and can be confusing.
av.logging.set_level(av.logging.CRITICAL)
_PACKET_AV_DATA_FORMATS: Dict[str, str] = {
value: key
for key, value in av.audio.frame.format_dtypes.items()
if key.endswith("p")
}
def prepare_input_file(
file: FileType, timeout: Union[int, float] = 30, reusable: bool = False
) -> Union[str, IO]:
"""
Prepare an input file depending on its type and value:
- download URL (`str`) and open it for file-like reading;
- check that a filepath (`str` or `os.PathLike`) exists and convert it to `str`;
- assume all other types as file-like and return as-is.
Args:
file: A filepath, an URL or a file-like object.
timeout: For URLs only. Download timeout.
reusable: For URLs only. If `True`, file-like return value is reusable
(use `file.seek(0)`). If `False` (default), return value can be used
only once.
Raises:
ValueError: if `file` is `str` or `os.PathLike`, but isn't a
valid/existing URL or an existing file.
"""
if isinstance(file, os.PathLike):
file = str(file)
if not isinstance(file, str):
# Assume that `file` is file-like.
return file
if validators.url(file):
response = requests.get(file, headers=headers, stream=True, timeout=timeout)
if response.ok:
if not reusable:
return response.raw
return io.BytesIO(response.content)
raise ValueError(f"URL {file} not found.")
if not os.path.isfile(file):
raise ValueError(f"File {file} is neither an URL nor an existing file.")
return file
def read_audio(file: FileType) -> Tuple[np.ndarray, int]:
"""
Read an audio file or file-like object using AV.
Args:
file: An audio filepath, file or file-like object. An input audio file.
Returns:
y: Array of shape `(num_samples, num_channels)`. PCM audio data, dtype
left as-is.
sampling_rate: Sampling rate of the audio data.
"""
file = prepare_input_file(file)
with av.open(file, "r") as container:
stream = container.streams.audio[0]
num_channels = stream.codec_context.channels
sampling_rate = stream.codec_context.rate
data = []
for frame in container.decode(audio=0):
frame_array = frame.to_ndarray()
if len(frame_array) == 1:
frame_array = frame_array.reshape((-1, num_channels))
else:
frame_array = frame_array.T
data.append(frame_array)
y = np.concatenate(data, axis=0)
return y, sampling_rate
def write_audio(
file: FileType, data: np.ndarray, sampling_rate: int, format_: str, codec: str
) -> None:
"""
Write audio data to a file or file-like object using AV.
Args:
file: A filepath, file or file-like object. An output audio file.
data: Array of shape `(num_samples, )` or `(num_samples, num_channels)`.
PCM audio data with any dtype supported by AV.
sampling_rate: Sampling rate of the audio data.
format_: An audio format supported by AV.
codec: An audio codec for the given audio format supported by AV
(s. `av.codec.codecs_available`).
"""
if isinstance(file, os.PathLike):
file = str(file)
data_format = _PACKET_AV_DATA_FORMATS[data.dtype.str[1:]]
if data.ndim == 1:
data = data[np.newaxis, :]
elif data.ndim == 2:
data = data.T
else:
raise ValueError(
f"`data` argument is expected to be an array with 1 or 2 dimensions, "
f"but array with {data.ndim} dimensions received."
)
# `AudioFrame.from_ndarray` expects an C-contiguous array as input.
data = np.ascontiguousarray(data)
num_channels = len(data)
frame = av.audio.AudioFrame.from_ndarray(data, data_format, num_channels)
frame.rate = sampling_rate
with av.open(file, "w", format_) as container:
stream = container.add_stream(codec, sampling_rate)
stream.channels = num_channels
container.mux(stream.encode(frame))
container.mux(stream.encode(None))
def transcode_audio(
input_file: FileType, output_file: FileType, output_format: str, output_codec: str
) -> None:
"""
Transcode an input audio file to an output audio file using AV.
Args:
input_file: A filepath, file or file-like object. An input audio file.
output_file: A filepath, file or file-like object. An output audio file.
output_format: An audio format supported by AV.
output_codec: An audio codec for the given audio format supported by AV
(s. `av.codec.codecs_available`).
"""
input_file = prepare_input_file(input_file)
if isinstance(output_file, os.PathLike):
output_file = str(output_file)
with av.open(input_file, "r") as input_container:
input_stream = input_container.streams.audio[0]
with av.open(output_file, "w", output_format) as output_container:
output_stream = output_container.add_stream(output_codec)
for frame in input_container.decode(input_stream):
frame.pts = None
for packet in output_stream.encode(frame):
output_container.mux(packet)
for packet in output_stream.encode(None):
output_container.mux(packet)
def get_format_codec(file: FileType) -> Tuple[str, str]:
"""
Get audio format and audio codec of an audio file.
"""
file = prepare_input_file(file)
with av.open(file, "r") as input_container:
stream = input_container.streams.audio[0]
return input_container.format.name, stream.name
def get_waveform(file: FileType) -> np.ndarray:
"""
Calculate waveform of an audio file or file-like object.
"""
y, input_sampling_rate = read_audio(file)
if y.dtype.str[1] == "i":
y = y.astype(np.float32) / np.iinfo(y.dtype).max
elif y.dtype.str[1] == "u":
y = 2 * y.astype(np.float32) / np.iinfo(y.dtype).max - 1
length = len(y) / input_sampling_rate
sampling_rate = max(50, min(1000, 10000 / length))
step = max(1.0, input_sampling_rate / sampling_rate)
num_windows = round(len(y) / step)
waveform = []
for window in np.array_split(y, num_windows):
waveform.append(window.max())
waveform.append(window.min())
return np.array(waveform) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/io/audio.py | 0.860208 | 0.436442 | audio.py | pypi |
from typing import List, Tuple
import numpy as np
import pygltflib
GLTF_DTYPES_CONVERSION = {
"i1": "<i1",
"i2": "<i2",
"i4": "<i2",
"i8": "<i2",
"i16": "<i2",
"u1": "<u1",
"u2": "<u2",
"u4": "<u4",
"u8": "<u4",
"u16": "<u4",
"f2": "<f4",
"f4": "<f4",
"f8": "<f4",
"f16": "<f4",
}
GLTF_DTYPES = {
pygltflib.BYTE: "<i1",
pygltflib.SHORT: "<i2",
pygltflib.UNSIGNED_BYTE: "<u1",
pygltflib.UNSIGNED_SHORT: "<u2",
pygltflib.UNSIGNED_INT: "<u4",
pygltflib.FLOAT: "<f4",
}
GLTF_DTYPES_LOOKUP = {v[1:]: k for k, v in GLTF_DTYPES.items()}
GLTF_SHAPES = {
pygltflib.SCALAR: (),
pygltflib.VEC2: (2,),
pygltflib.VEC3: (3,),
pygltflib.VEC4: (4,),
} # MAT2, MAT3 and MAT4 are currently not supported.
GLTF_SHAPES_LOOKUP = {v: k for k, v in GLTF_SHAPES.items()}
def check_gltf(gltf: pygltflib.GLTF2) -> None:
"""
Check whether a glTF mesh can be parsed.
glTF mesh is required to have exactly one scene, one node, one mesh, one
primitive with mode "TRIANGLES" (4) and one buffer.
"""
if (
len(gltf.scenes) != 1
or len(gltf.nodes) != 1
or len(gltf.meshes) != 1
or len(gltf.meshes[0].primitives) != 1
):
raise ValueError(
f"glTF file with exactly one scene, one node, one mesh and one "
f"primitive expected, but {len(gltf.scenes)} scenes, "
f"{len(gltf.nodes)} nodes, {len(gltf.meshes)} meshes and "
f"{len(gltf.meshes[0].primitives)} received."
)
if gltf.meshes[0].primitives[0].mode != pygltflib.TRIANGLES:
raise ValueError(
f"glTF mesh with primitive of mode TRIANGLES "
f"({pygltflib.TRIANGLES}) expected, but mode "
f"{gltf.meshes[0].primitives[0].mode} received."
)
if gltf.scene != 0 or gltf.scenes[0].nodes != [0] or gltf.nodes[0].mesh != 0:
raise ValueError("Invalid glTF hierarchy.")
if (
len(gltf.buffers) != 1
or gltf.buffers[0].uri not in [None, ""]
or gltf.buffers[0].byteLength != len(gltf.binary_blob())
):
raise ValueError("Invalid glTF buffer structure.")
def pad_bin_data(bin_data: bytes, bound: int = 4) -> bytes:
"""
Pad binary data to the given bound.
"""
bound = int(bound)
if bound < 2:
return bin_data
data_length = len(bin_data)
if data_length % bound == 0:
return bin_data
return bin_data + b"\0" * (bound - data_length % bound)
def encode_gltf_array(
array: np.ndarray,
bin_data: bytes,
buffer_views: List[pygltflib.BufferView],
accessors: List[pygltflib.Accessor],
buffer_view_target: int = pygltflib.ARRAY_BUFFER,
) -> Tuple[bytes, List[pygltflib.BufferView], List[pygltflib.Accessor]]:
"""
Encode a single array as expected by pygltflib, update binary blob, lists
with buffer views and accessors.
"""
array = array.squeeze().astype(GLTF_DTYPES_CONVERSION[array.dtype.str[1:]])
array_bin_data = array.tobytes()
if array.size > 0:
if array.dtype.str[1] == "f":
# For float arrays, do not count non-finite values.
finite_mask = np.isfinite(array)
max_list = array.max(axis=0, initial=-np.inf, where=finite_mask)
min_list = array.min(axis=0, initial=np.inf, where=finite_mask)
max_list = np.where(
~np.isfinite(max_list), np.array(None), max_list
).tolist()
min_list = np.where(
~np.isfinite(min_list), np.array(None), min_list
).tolist()
else:
max_list = array.max(axis=0).tolist()
min_list = array.min(axis=0).tolist()
if len(array.shape) == 1:
max_list = [max_list]
min_list = [min_list]
else:
max_list = []
min_list = []
accessors.append(
pygltflib.Accessor(
bufferView=len(buffer_views),
byteOffset=0,
componentType=GLTF_DTYPES_LOOKUP[array.dtype.str[1:]],
count=len(array),
type=GLTF_SHAPES_LOOKUP[array.shape[1:]],
max=max_list,
min=min_list,
)
)
buffer_views.append(
pygltflib.BufferView(
buffer=0,
byteOffset=len(bin_data),
byteLength=len(array_bin_data),
target=buffer_view_target,
)
)
return bin_data + pad_bin_data(array_bin_data), buffer_views, accessors
def decode_gltf_arrays(gltf: pygltflib.GLTF2) -> List[np.ndarray]:
"""
Decode all arrays from glTF instance accessors.
It is assumed that only one buffer is used (as it should be for glb format).
"""
bin_data = gltf.binary_blob()
buffer_views = []
for buffer_view in gltf.bufferViews:
buffer_views.append(
bin_data[
buffer_view.byteOffset : buffer_view.byteOffset + buffer_view.byteLength
]
)
arrays = []
for accessor in gltf.accessors:
accessor_bin_data = buffer_views[accessor.bufferView]
shape = (accessor.count, *GLTF_SHAPES[accessor.type])
array_flatten = np.frombuffer(
accessor_bin_data[accessor.byteOffset :],
dtype=GLTF_DTYPES[accessor.componentType],
count=int(np.prod(shape)),
)
array = array_flatten.reshape(shape)
arrays.append(array)
return arrays | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/io/gltf.py | 0.743261 | 0.557123 | gltf.py | pypi |
import uuid
from typing import List, Optional, Union
from pydantic import BaseModel, Field
class Lens(BaseModel, allow_population_by_field_name=True):
"""
Inspector lens configuration model.
Following combinations of lens types and column types are supported by
default (but can be further extended):
"ScalarView": single column of type `bool`, `int`, `float`, `str`,
`datetime.datetime` or `spotlight.Category`
"TextLens": single column of type `str`
"HtmlLens": single column of type `str`
"SafeHtmlLens": single column of type `str`
"MarkdownLens": single column of type `str`
"ArrayLens": single column of type `np.ndarray`,
`spotlight.Embedding` or `spotlight.Window`
"SequenceView": single or multiple columns of type `spotlight.Sequence1D`
"MeshView": single column of type `spotlight.Mesh`
"ImageView": single column of type `spotlight.Image`
"VideoView": single column of type `spotlight.Video`
"AudioView": single column of type `spotlight.Audio`, optional
single column of type `spotlight.Window`
"SpectrogramView": single column of type `spotlight.Audio`, optional
single column of type `spotlight.Window`
"""
type: str = Field(..., alias="view")
columns: List[str] = Field(..., alias="columns")
name: Optional[str] = Field(None, alias="name")
id: Optional[str] = Field(default_factory=lambda: str(uuid.uuid4()), alias="key")
def lens(
internal_type: str, columns: Union[str, List[str]], name: Optional[str] = None
) -> Lens:
"""
Add a viewer to Spotlight inspector widget.
`internal_type` should be set exactly as expected if the frontend. For
supported lens types and respective column types, see `Lens` class.
Prefer to use explicit lens functions defined below.
"""
return Lens(
type=internal_type, # type: ignore
columns=[columns] if isinstance(columns, str) else columns,
name=name,
)
def scalar(column: str, name: Optional[str] = None) -> Lens:
"""
Add scalar value viewer to Spotlight inspector widget.
Supports a single column of type `bool`, `int`, `float`, `str`,
`datetime.datetime` or `spotlight.Category`.
"""
return Lens(type="ScalarView", columns=[column], name=name) # type: ignore
def text(column: str, name: Optional[str] = None) -> Lens:
"""
Add text viewer to Spotlight inspector widget.
Supports a single column of type `str`.
"""
return Lens(type="TextLens", columns=[column], name=name) # type: ignore
def html(column: str, name: Optional[str] = None, unsafe: bool = False) -> Lens:
"""
Add HTML viewer to Spotlight inspector widget.
Supports a single column of type `str`.
"""
if unsafe:
return Lens(type="HtmlLens", columns=[column], name=name) # type: ignore
return Lens(type="SafeHtmlLens", columns=[column], name=name) # type: ignore
def markdown(column: str, name: Optional[str] = None) -> Lens:
"""
Add markdown viewer to Spotlight inspector widget.
Supports a single column of type `str`.
"""
return Lens(type="MarkdownLens", columns=[column], name=name) # type: ignore
def array(column: str, name: Optional[str] = None) -> Lens:
"""
Add array viewer to Spotlight inspector widget.
Supports a single column of type `np.ndarray`, `spotlight.Embedding` or
`spotlight.Window`.
"""
return Lens(type="ArrayLens", columns=[column], name=name) # type: ignore
def sequences(columns: Union[str, List[str]], name: Optional[str] = None) -> Lens:
"""
Add sequence viewer to Spotlight inspector widget.
Supports one or multiple columns of type `spotlight.Sequence1D`.
"""
return Lens(
type="SequenceView", # type: ignore
columns=[columns] if isinstance(columns, str) else columns,
name=name,
)
def mesh(column: str, name: Optional[str] = None) -> Lens:
"""
Add mesh viewer to Spotlight inspector widget.
Supports a single column of type `spotlight.Mesh`.
"""
return Lens(type="MeshView", columns=[column], name=name) # type: ignore
def image(column: str, name: Optional[str] = None) -> Lens:
"""
Add image viewer to Spotlight inspector widget.
Supports a single column of type `spotlight.Image`.
"""
return Lens(type="ImageView", columns=[column], name=name) # type: ignore
def video(column: str, name: Optional[str] = None) -> Lens:
"""
Add video viewer to Spotlight inspector widget.
Supports a single column of type `spotlight.Video`.
"""
return Lens(type="VideoView", columns=[column], name=name) # type: ignore
def audio(
column: str, window_column: Optional[str] = None, name: Optional[str] = None
) -> Lens:
"""
Add audio viewer to Spotlight inspector widget.
Supports a single column of type `spotlight.Audio` with optional second
column of type `spotlight.Window`.
"""
return Lens(
type="AudioView", # type: ignore
columns=[column] if window_column is None else [column, window_column],
name=name,
)
def spectrogram(
column: str, window_column: Optional[str] = None, name: Optional[str] = None
) -> Lens:
"""
Add audio spectrogram viewer to Spotlight inspector widget.
Supports a single column of type `spotlight.Audio` with optional second
column of type `spotlight.Window`.
"""
return Lens(
type="SpectrogramView", # type: ignore
columns=[column] if window_column is None else [column, window_column],
name=name,
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/layout/lenses.py | 0.947186 | 0.699652 | lenses.py | pypi |
from typing import List, Optional
from pydantic import BaseModel, Extra, Field
from typing_extensions import Literal
from .lenses import Lens
class WidgetConfig(BaseModel, allow_population_by_field_name=True):
"""
Base Spotlight widget configuration model.
"""
class Widget(BaseModel, extra=Extra.forbid):
"""
Spotlight widget model.
"""
type: str
name: Optional[str] = None
config: Optional[WidgetConfig] = None
kind: Literal["widget"] = "widget"
class HistogramConfig(WidgetConfig):
"""
Histogram configuration model.
"""
column: Optional[str] = Field(None, alias="columnKey")
stack_by_column: Optional[str] = Field(None, alias="stackByColumnKey")
filter: bool = Field(False, alias="filter")
class Histogram(Widget):
"""
Spotlight histogram model.
"""
type: Literal["histogram"] = "histogram"
config: Optional[HistogramConfig] = None
class ScatterplotConfig(WidgetConfig):
"""
Scatter plot configuration model.
"""
x_column: Optional[str] = Field(None, alias="xAxisColumn")
y_column: Optional[str] = Field(None, alias="yAxisColumn")
color_by_column: Optional[str] = Field(None, alias="colorBy")
size_by_column: Optional[str] = Field(None, alias="sizeBy")
filter: bool = Field(False, alias="filter")
class Scatterplot(Widget):
"""
Spotlight scatter plot model.
"""
type: Literal["scatterplot"] = "scatterplot"
config: Optional[ScatterplotConfig] = None
TableView = Literal["full", "filtered", "selected"]
class TableConfig(WidgetConfig):
"""
Table configuration model.
"""
active_view: TableView = Field("full", alias="tableView")
visible_columns: Optional[List[str]] = Field(None, alias="visibleColumns")
sort_by_columns: Optional[List[List[str]]] = Field(None, alias="sorting")
order_by_relevance: bool = Field(False, alias="orderByRelevance")
class Table(Widget):
"""
Spotlight table model.
"""
type: Literal["table"] = "table"
config: Optional[TableConfig] = None
ReductionMethod = Literal["umap", "pca"]
UmapMetric = Literal[
"euclidean", "standardized euclidean", "robust euclidean", "cosine", "mahalanobis"
]
PCANormalization = Literal["none", "standardize", "robust standardize"]
class SimilaritymapConfig(WidgetConfig):
"""
Similarity map configuration model.
"""
columns: Optional[List[str]] = Field(None, alias="placeBy")
reduction_method: Optional[ReductionMethod] = Field(None, alias="reductionMethod")
color_by_column: Optional[str] = Field(None, alias="colorBy")
size_by_column: Optional[str] = Field(None, alias="sizeBy")
filter: bool = Field(False, alias="filter")
umap_nn: Optional[int] = Field(None, alias="umapNNeighbors")
umap_metric: Optional[UmapMetric] = Field(None, alias="umapMetric")
umap_min_dist: Optional[float] = Field(None, alias="umapMinDist")
pca_normalization: Optional[PCANormalization] = Field(
None, alias="pcaNormalization"
)
umap_balance: Optional[float] = Field(None, alias="umapMenuLocalGlobalBalance")
umap_is_advanced: bool = Field(False, alias="umapMenuIsAdvanced")
class Similaritymap(Widget):
"""
Spotlight similarity map model.
"""
type: Literal["similaritymap"] = "similaritymap"
config: Optional[SimilaritymapConfig] = None
NumInspectorColumns = Literal[1, 2, 4, 6, 8]
class InspectorConfig(WidgetConfig):
"""
Inspector configuration model.
"""
lenses: Optional[List[Lens]] = Field(default_factory=None, alias="views")
num_columns: NumInspectorColumns = Field(4, alias="visibleColumns")
class Inspector(Widget):
"""
Spotlight inspector model.
"""
type: Literal["inspector"] = "inspector"
config: Optional[InspectorConfig] = None
class Issues(Widget):
"""
Spotlight issues widget
"""
type: Literal["IssuesWidget"] = "IssuesWidget" | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/layout/widgets.py | 0.880938 | 0.414129 | widgets.py | pypi |
import os
from pathlib import Path
from typing import Dict, Iterable, List, Optional, Tuple, Union, cast, overload
from pydantic import (
HttpUrl,
ValidationError,
parse_obj_as,
)
import requests
from typing_extensions import Literal
from renumics.spotlight.backend.exceptions import InvalidLayout
from .nodes import (
Layout,
Orientation as _Orientation,
Split,
Tab,
)
from .lenses import Lens
from .widgets import (
Histogram,
HistogramConfig,
Inspector,
InspectorConfig,
Issues,
NumInspectorColumns as _NumInspectorColumns,
PCANormalization as _PCANormalization,
ReductionMethod as _ReductionMethod,
Scatterplot,
ScatterplotConfig,
Similaritymap,
SimilaritymapConfig,
Table,
TableConfig,
TableView as _TableView,
UmapMetric as _UmapMetric,
Widget as _Widget,
)
_WidgetLike = Union[_Widget, str]
_NodeLike = Union[Split, Tab, _WidgetLike, List]
_LayoutLike = Union[str, os.PathLike, Layout, _NodeLike]
def _clean_nodes(nodes: Iterable[_NodeLike]) -> List[Union[Split, Tab]]:
"""
Wrap standalone widgets into tabs with a single widget inside and
lists of widgets into common tabs. Pass splits and tabs as is.
"""
if all(isinstance(node, (_Widget, str)) for node in nodes):
nodes = cast(Iterable[_WidgetLike], nodes)
return [tab(*nodes)]
cleaned_nodes = []
for node in nodes:
if isinstance(node, (Split, Tab)):
cleaned_nodes.append(node)
elif isinstance(node, (_Widget, str)):
cleaned_nodes.append(tab(node))
elif isinstance(node, list):
if all(isinstance(subnode, (_Widget, str)) for subnode in node):
# All widgets inside, group them into one tab.
cleaned_nodes.append(tab(*node))
else:
# Non-homogeneous content, wrap into a split.
cleaned_nodes.append(split(*node))
else:
raise TypeError(
f"Cannot parse layout content. Unexpected node of type {type(node)} received."
)
return cleaned_nodes
def layout(*nodes: _NodeLike, orientation: _Orientation = None) -> Layout:
"""
Create a new layout with the given orientation and given nodes inside.
"""
return Layout(children=_clean_nodes(nodes), orientation=orientation)
def split(
*nodes: _NodeLike, weight: Union[float, int] = 1, orientation: _Orientation = None
) -> Split:
"""
Create a new split with the given weight, orientation and given nodes inside.
"""
return Split(children=_clean_nodes(nodes), weight=weight, orientation=orientation)
def tab(*widgets: _WidgetLike, weight: Union[float, int] = 1) -> Tab:
"""
Create a new tab with the given weight and given widgets inside.
"""
return Tab(
children=[_Widget(type=x) if isinstance(x, str) else x for x in widgets],
weight=weight,
)
def parse(layout_: _LayoutLike) -> Layout:
"""
Parse layout from a file, url, layout or given nodes.
"""
if isinstance(layout_, Layout):
return layout_
try:
parse_obj_as(HttpUrl, layout_)
try:
resp = requests.get(str(layout_), timeout=20)
return Layout(**resp.json())
except (ValidationError, requests.JSONDecodeError) as e:
raise InvalidLayout() from e
except ValidationError:
pass
if (isinstance(layout_, (os.PathLike, str))) and os.path.isfile(layout_):
return Layout.parse_file(Path(layout_))
layout_ = cast(_NodeLike, layout_)
return layout(layout_)
def histogram(
name: Optional[str] = None,
column: Optional[str] = None,
stack_by_column: Optional[str] = None,
filter: bool = False,
) -> Histogram:
"""
Add configured histogram to Spotlight layout.
"""
return Histogram(
name=name,
config=HistogramConfig(
column=column,
stack_by_column=stack_by_column,
filter=filter,
),
)
def inspector(
name: Optional[str] = None,
lenses: Optional[Iterable[Lens]] = None,
num_columns: _NumInspectorColumns = 4,
) -> Inspector:
"""
Add an inspector widget with optionally preconfigured viewers (lenses).
Example:
>>> from renumics.spotlight import layout
>>> from renumics.spotlight.layout import lenses
>>> spotlight_layout = layout.layout(
... layout.inspector(
... "My Inspector",
... [
... lenses.scalar("bool"),
... lenses.scalar("float"),
... lenses.scalar("str"),
... lenses.scalar("datetime"),
... lenses.scalar("category"),
... lenses.scalar("int"),
... lenses.text("str", name="text"),
... lenses.html("str", name="HTML (safe)"),
... lenses.html("str", name="HTML", unsafe=True),
... lenses.markdown("str", name="MD"),
... lenses.array("embedding"),
... lenses.array("window"),
... lenses.array("array"),
... lenses.sequences("sequence"),
... lenses.sequences(["sequence1", "sequence2"], name="sequences"),
... lenses.mesh("mesh"),
... lenses.image("image"),
... lenses.video("video"),
... lenses.audio("audio"),
... lenses.audio("audio", window_column="window", name="windowed audio"),
... lenses.spectrogram("audio"),
... lenses.spectrogram(
... "audio",
... window_column="window",
... name="windowed spectrogram",
... ),
... ],
... num_columns=2,
... )
... )
"""
return Inspector(
name=name,
config=InspectorConfig(
lenses=lenses if lenses is None else list(lenses), num_columns=num_columns
),
)
def scatterplot(
name: Optional[str] = None,
x_column: Optional[str] = None,
y_column: Optional[str] = None,
color_by_column: Optional[str] = None,
size_by_column: Optional[str] = None,
filter: bool = False,
) -> Scatterplot:
"""
Add configured scatter plot to Spotlight layout.
"""
return Scatterplot(
name=name,
config=ScatterplotConfig(
x_column=x_column,
y_column=y_column,
color_by_column=color_by_column,
size_by_column=size_by_column,
filter=filter,
),
)
_UmapBalance = Literal["local", "normal", "global"]
_UMAP_BALANCE_TO_FLOAT: Dict[_UmapBalance, float] = {
"local": 0.0,
"normal": 0.5,
"global": 1.0,
}
@overload
def similaritymap(
name: Optional[str] = None,
columns: Optional[List[str]] = None,
reduction_method: Literal[None] = None,
color_by_column: Optional[str] = None,
size_by_column: Optional[str] = None,
filter: bool = False,
) -> Similaritymap:
...
@overload
def similaritymap(
name: Optional[str] = None,
columns: Optional[List[str]] = None,
reduction_method: Literal["umap"] = "umap",
color_by_column: Optional[str] = None,
size_by_column: Optional[str] = None,
filter: bool = False,
*,
umap_metric: Optional[_UmapMetric] = None,
umap_balance: Optional[_UmapBalance] = None,
) -> Similaritymap:
...
@overload
def similaritymap(
name: Optional[str] = None,
columns: Optional[List[str]] = None,
reduction_method: Literal["pca"] = "pca",
color_by_column: Optional[str] = None,
size_by_column: Optional[str] = None,
filter: bool = False,
*,
pca_normalization: Optional[_PCANormalization] = None,
) -> Similaritymap:
...
def similaritymap(
name: Optional[str] = None,
columns: Optional[List[str]] = None,
reduction_method: Optional[_ReductionMethod] = None,
color_by_column: Optional[str] = None,
size_by_column: Optional[str] = None,
filter: bool = False,
*,
umap_metric: Optional[_UmapMetric] = None,
umap_balance: Optional[_UmapBalance] = None,
pca_normalization: Optional[_PCANormalization] = None,
) -> Similaritymap:
"""
Add configured similarity map to Spotlight layout.
"""
umap_balance_float = None
if reduction_method == "umap":
pca_normalization = None
if umap_balance is not None:
umap_balance_float = _UMAP_BALANCE_TO_FLOAT[umap_balance]
elif reduction_method == "pca":
umap_metric = None
umap_balance = None
return Similaritymap(
name=name,
config=SimilaritymapConfig(
columns=columns,
reduction_method=reduction_method,
color_by_column=color_by_column,
size_by_column=size_by_column,
filter=filter,
umap_metric=umap_metric,
umap_balance=umap_balance_float,
pca_normalization=pca_normalization,
),
)
_TableTab = Literal["all", "filtered", "selected"]
_TABLE_TAB_TO_TABLE_VIEW: Dict[_TableTab, _TableView] = {
"all": "full",
"filtered": "filtered",
"selected": "selected",
}
_SortOrder = Literal["ascending", "descending"]
_SORT_ORDER_MAPPING: Dict[_SortOrder, str] = {
"ascending": "ASC",
"descending": "DESC",
}
def table(
name: Optional[str] = None,
active_view: _TableTab = "all",
visible_columns: Optional[List[str]] = None,
sort_by_columns: Optional[List[Tuple[str, _SortOrder]]] = None,
order_by_relevance: bool = False,
) -> Table:
"""
Add configured table to Spotlight layout.
"""
return Table(
name=name,
config=TableConfig(
active_view=_TABLE_TAB_TO_TABLE_VIEW[active_view],
visible_columns=visible_columns,
sort_by_columns=None
if sort_by_columns is None
else [
[column, _SORT_ORDER_MAPPING[order]]
for column, order in sort_by_columns
],
order_by_relevance=order_by_relevance,
),
)
def issues(name: Optional[str] = None) -> Issues:
"""
Add a widget displaying data issues.
"""
return Issues(name=name) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/layout/__init__.py | 0.870817 | 0.256157 | __init__.py | pypi |
from datetime import datetime
from typing import Any, Dict, Union, Type
import numpy as np
from typing_extensions import TypeGuard, get_args
from . import Audio, Category, Embedding, Image, Mesh, Sequence1D, Video, Window, DType
from .exceptions import NotADType
ColumnType = Union[bool, int, float, str, datetime, Category, Window, np.ndarray, DType]
ScalarColumnType = Union[bool, int, float, str, datetime, Category]
FileBasedColumnType = Union[Audio, Image, Mesh, Video]
ArrayBasedColumnType = Union[Embedding, Image, Sequence1D]
ColumnTypeMapping = Dict[str, Type[ColumnType]]
COLUMN_TYPES_BY_NAME: Dict[str, Type[ColumnType]] = {
"bool": bool,
"int": int,
"float": float,
"str": str,
"datetime": datetime,
"Category": Category,
"Window": Window,
"array": np.ndarray,
"Image": Image,
"Audio": Audio,
"Video": Video,
"Mesh": Mesh,
"Embedding": Embedding,
"Sequence1D": Sequence1D,
}
NAME_BY_COLUMN_TYPE: Dict[Type[ColumnType], str] = {
v: k for k, v in COLUMN_TYPES_BY_NAME.items()
}
def get_column_type_name(column_type: Type[ColumnType]) -> str:
"""
Get name of a column type as string.
"""
try:
return NAME_BY_COLUMN_TYPE[column_type]
except KeyError as e:
raise NotADType(f"Unknown column type: {column_type}.") from e
def get_column_type(x: str) -> Type[ColumnType]:
"""
Get column type by its name.
"""
try:
return COLUMN_TYPES_BY_NAME[x]
except KeyError as e:
raise NotADType(f"Unknown column type: {x}.") from e
def is_column_type(x: Any) -> TypeGuard[Type[ColumnType]]:
"""
Check whether `x` is a Spotlight data type class.
"""
return x in COLUMN_TYPES_BY_NAME.values()
def is_scalar_column_type(x: Any) -> TypeGuard[Type[ScalarColumnType]]:
"""
Check whether `x` is a scalar Spotlight data type class.
"""
return x in get_args(ScalarColumnType)
def is_file_based_column_type(x: Any) -> TypeGuard[Type[FileBasedColumnType]]:
"""
Check whether `x` is a Spotlight column type class whose instances
can be read from/saved into a file.
"""
return x in get_args(FileBasedColumnType)
def is_array_based_column_type(x: Any) -> TypeGuard[Type[ArrayBasedColumnType]]:
"""
Check whether `x` is a Spotlight column type class which can be instantiated
from a single array-like argument.
"""
return x in get_args(ArrayBasedColumnType) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dtypes/typing.py | 0.860852 | 0.397237 | typing.py | pypi |
from typing import Dict, List, Optional, Tuple, Union
import numpy as np
def reindex(point_ids: np.ndarray, *elements: np.ndarray) -> Tuple[np.ndarray, ...]:
"""
Reindex elements which refer to the non-negative unique point ids so that
they refer to the indices of point ids (`np.arange(len(point_ids))`) in the
same way.
"""
if elements:
inverse_point_ids = np.full(
max((x.max(initial=0) for x in (point_ids, *elements))) + 1,
-1,
np.int64,
)
inverse_point_ids[point_ids] = np.arange(len(point_ids))
return tuple(inverse_point_ids[x].astype(x.dtype) for x in elements)
return ()
def attribute_to_array(attribute: Union[np.ndarray, List[np.ndarray]]) -> np.ndarray:
"""
Encode a single attribute or a list of attribute steps as array with shape
`(num_steps, n, ...)` and squeeze all dimensions except for 0 and 1.
"""
if not isinstance(attribute, list):
attribute = [attribute]
attribute = np.asarray(attribute)
attribute = attribute.reshape(
(*attribute.shape[:2], *(x for x in attribute.shape[2:] if x != 1))
)
return attribute
def triangulate(
triangles: Optional[np.ndarray] = None,
triangle_attributes: Optional[
Dict[str, Union[np.ndarray, List[np.ndarray]]]
] = None,
quadrangles: Optional[np.ndarray] = None,
quadrangle_attributes: Optional[
Dict[str, Union[np.ndarray, List[np.ndarray]]]
] = None,
) -> Tuple[np.ndarray, Dict[str, Union[np.ndarray, List[np.ndarray]]]]:
"""
Triangulate quadrangles and respective attributes and append them to the
given triangles/attributes.
"""
attrs = {}
if triangles is None:
trias = np.empty((0, 3), np.uint32)
if triangle_attributes:
raise ValueError(
f"`triangles` not given, but `triangle_attributes` have "
f"{len(triangle_attributes)} items."
)
else:
trias = triangles
if triangle_attributes is not None:
for attr_name, triangle_attr in triangle_attributes.items():
attrs[attr_name] = attribute_to_array(triangle_attr)
if quadrangles is None:
if quadrangle_attributes is not None and len(quadrangle_attributes) != 0:
raise ValueError(
f"`quadrangles` not given, but `quadrangle_attributes` have "
f"{len(quadrangle_attributes)} items."
)
else:
trias = np.concatenate(
(trias, quadrangles[:, [0, 1, 2]], quadrangles[:, [0, 2, 3]])
)
for attr_name, quadrangle_attr in (quadrangle_attributes or {}).items():
quadrangle_attr = attribute_to_array(quadrangle_attr)
try:
attr = attrs[attr_name]
except KeyError:
attrs[attr_name] = np.concatenate(
(quadrangle_attr, quadrangle_attr), axis=1
)
else:
attrs[attr_name] = np.concatenate(
(attr, quadrangle_attr, quadrangle_attr), axis=1
)
for attr_name, attr in attrs.items():
if attr.shape[1] != len(trias):
raise ValueError(
f"Values of attributes should have the same length as "
f"triangles ({len(trias)}), but length {attr.shape[1]} "
f"received."
)
return (
trias,
{
attr_name: attr[0] if len(attr) == 1 else list(attr)
for attr_name, attr in attrs.items()
},
)
def clean(
points: np.ndarray,
triangles: np.ndarray,
point_attributes: Optional[Dict[str, np.ndarray]] = None,
triangle_attributes: Optional[Dict[str, np.ndarray]] = None,
point_displacements: Optional[List[np.ndarray]] = None,
) -> Tuple[
np.ndarray,
np.ndarray,
Dict[str, np.ndarray],
Dict[str, np.ndarray],
List[np.ndarray],
]:
"""
Remove:
degenerated triangles and respective attributes;
invalid triangles and respective attributes;
invalid points and respective attributes;
empty attributes and point displacements.
"""
point_ids = np.arange(len(points))
valid_triangles_mask = (
(triangles[:, 0] != triangles[:, 1])
& (triangles[:, 0] != triangles[:, 2])
& (triangles[:, 1] != triangles[:, 2])
& np.isin(triangles, point_ids).all(axis=1)
)
triangles = triangles[valid_triangles_mask]
triangle_attributes = {
k: [x[valid_triangles_mask] for x in v]
if isinstance(v, list)
else v[valid_triangles_mask]
for k, v in (triangle_attributes or {}).items()
}
valid_points_mask = np.isin(point_ids, triangles)
points = points[valid_points_mask]
point_attributes = {
k: [x[valid_points_mask] for x in v]
if isinstance(v, list)
else v[valid_points_mask]
for k, v in (point_attributes or {}).items()
}
point_displacements = [x[valid_points_mask] for x in point_displacements or []]
point_ids = point_ids[valid_points_mask]
triangles, *_ = reindex(point_ids, triangles)
return (
points,
triangles,
{k: v for k, v in point_attributes.items() if len(v) > 0},
{k: v for k, v in triangle_attributes.items() if len(v) > 0},
[x for x in point_displacements if len(x) > 0],
) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dtypes/triangulation.py | 0.914477 | 0.705354 | triangulation.py | pypi |
from collections import defaultdict
from inspect import signature
from dataclasses import dataclass
import io
import os
from typing import Any, Callable, List, TypeVar, Union, Type, Optional, Dict
import datetime
from filetype import filetype
import numpy as np
import trimesh
import validators
from renumics.spotlight.typing import PathOrUrlType, PathType
from renumics.spotlight.cache import external_data_cache
from renumics.spotlight.io import audio
from renumics.spotlight.io.file import as_file
from .typing import (
ColumnType,
Category,
FileBasedColumnType,
Window,
Sequence1D,
Embedding,
Image,
Audio,
Video,
Mesh,
get_column_type_name,
)
NormalizedValue = Union[
int,
float,
bool,
str,
bytes,
datetime.datetime,
np.ndarray,
None,
list,
]
ConvertedValue = Union[
int, float, bool, str, bytes, datetime.datetime, np.ndarray, None
]
@dataclass(frozen=True)
class DTypeOptions:
"""
All possible dtype options
"""
categories: Optional[Dict[str, int]] = None
class ConversionError(Exception):
"""
Type Conversion failed
"""
class NoConverterAvailable(Exception):
"""
No matching converter could be applied
"""
def __init__(self, value: NormalizedValue, dtype: Type[ColumnType]) -> None:
msg = f"No Converter for {type(value)} -> {dtype}"
super().__init__(msg)
N = TypeVar("N", bound=NormalizedValue)
Converter = Callable[[N, DTypeOptions], ConvertedValue]
ConverterWithoutOptions = Callable[[N], ConvertedValue]
_converters_table: Dict[
Type[NormalizedValue], Dict[Type[ColumnType], List[Converter]]
] = defaultdict(lambda: defaultdict(list))
_simple_converters_table: Dict[
Type[NormalizedValue], Dict[Type[ColumnType], List[Converter]]
] = defaultdict(lambda: defaultdict(list))
def register_converter(
from_type: Type[N],
to_type: Type[ColumnType],
converter: Converter[N],
simple: Optional[bool] = None,
) -> None:
"""
register a converter from NormalizedType to ColumnType
"""
if simple is None:
_simple_converters_table[from_type][to_type].append(converter) # type: ignore
_converters_table[from_type][to_type].append(converter) # type: ignore
elif simple:
_simple_converters_table[from_type][to_type].append(converter) # type: ignore
else:
_converters_table[from_type][to_type].append(converter) # type: ignore
def convert(
from_type: Type[N], to_type: Type[ColumnType], simple: Optional[bool] = None
) -> Callable:
"""
Decorator for simplified registration of converters
"""
def _decorate(
func: Union[Converter[N], ConverterWithoutOptions[N]]
) -> Converter[N]:
if len(signature(func).parameters) == 1:
def _converter(value: Any, _: DTypeOptions) -> ConvertedValue:
return func(value) # type: ignore
else:
_converter = func # type: ignore
register_converter(from_type, to_type, _converter, simple)
return _converter
return _decorate
def convert_to_dtype(
value: NormalizedValue,
dtype: Type[ColumnType],
dtype_options: DTypeOptions = DTypeOptions(),
simple: bool = False,
) -> ConvertedValue:
"""
Convert normalized type from data source to internal Spotlight DType
"""
registered_converters = (
_simple_converters_table[type(value)][dtype]
if simple
else _converters_table[type(value)][dtype]
)
for converter in registered_converters:
try:
return converter(value, dtype_options)
except ConversionError:
pass
try:
if dtype is bool:
return bool(value) # type: ignore
if dtype is int:
return int(value) # type: ignore
if dtype is float:
return float(value) # type: ignore
if dtype is str:
str_value = str(value)
if simple and len(str_value) > 100:
return str_value[:97] + "..."
return str_value
if value is None:
return None
if dtype is np.ndarray:
if simple:
return "[...]"
if isinstance(value, list):
return np.array(value)
if isinstance(value, np.ndarray):
return value
if dtype is Category and np.issubdtype(np.dtype(type(value)), np.integer):
assert dtype_options.categories is not None
value_int = int(value) # type: ignore
if value_int != -1 and value_int not in dtype_options.categories.values():
raise ConversionError()
return value_int
except (TypeError, ValueError) as e:
raise ConversionError() from e
raise NoConverterAvailable(value, dtype)
@convert(datetime.datetime, datetime.datetime)
def _(value: datetime.datetime) -> datetime.datetime:
return value
@convert(str, datetime.datetime)
@convert(np.str_, datetime.datetime)
def _(value: Union[str, np.str_]) -> Optional[datetime.datetime]:
if value == "":
return None
return datetime.datetime.fromisoformat(value)
@convert(np.datetime64, datetime.datetime) # type: ignore
def _(value: np.datetime64) -> datetime.datetime:
return value.tolist()
@convert(str, Category)
@convert(np.str_, Category)
def _(value: Union[str, np.str_], options: DTypeOptions) -> int:
if not options.categories:
return -1
return options.categories[value]
@convert(type(None), Category)
def _(_: None) -> int:
return -1
@convert(int, Category)
def _(value: int) -> int:
return value
@convert(type(None), Window)
def _(_: None) -> np.ndarray:
return np.full((2,), np.nan, dtype=np.float64)
@convert(list, Window)
def _(value: list) -> np.ndarray:
return np.array(value, dtype=np.float64)
@convert(np.ndarray, Window)
def _(value: np.ndarray) -> np.ndarray:
return value.astype(np.float64)
@convert(list, Embedding, simple=False)
def _(value: list) -> np.ndarray:
return np.array(value, dtype=np.float64)
@convert(np.ndarray, Embedding, simple=False)
def _(value: np.ndarray) -> np.ndarray:
return value.astype(np.float64)
@convert(list, Sequence1D, simple=False)
@convert(np.ndarray, Sequence1D, simple=False)
def _(value: Union[np.ndarray, list], _: DTypeOptions) -> np.ndarray:
return Sequence1D(value).encode()
@convert(str, Image, simple=False)
@convert(np.str_, Image, simple=False)
def _(value: Union[str, np.str_]) -> bytes:
data = read_external_value(value, Image)
if data is None:
raise ConversionError()
return data.tolist()
@convert(bytes, Image, simple=False)
@convert(np.bytes_, Image, simple=False)
def _(value: Union[bytes, np.bytes_]) -> bytes:
return Image.from_bytes(value).encode().tolist()
@convert(np.ndarray, Image, simple=False)
def _(value: np.ndarray) -> bytes:
return Image(value).encode().tolist()
@convert(str, Audio, simple=False)
@convert(np.str_, Audio, simple=False)
def _(value: Union[str, np.str_]) -> bytes:
if data := read_external_value(value, Audio):
return data.tolist()
raise ConversionError()
@convert(bytes, Audio, simple=False)
@convert(np.bytes_, Audio, simple=False)
def _(value: Union[bytes, np.bytes_]) -> bytes:
return Audio.from_bytes(value).encode().tolist()
@convert(str, Video, simple=False)
@convert(np.str_, Video, simple=False)
def _(value: Union[str, np.str_]) -> bytes:
if data := read_external_value(value, Video):
return data.tolist()
raise ConversionError()
@convert(bytes, Video, simple=False)
@convert(np.bytes_, Video, simple=False)
def _(value: Union[bytes, np.bytes_]) -> bytes:
return Video.from_bytes(value).encode().tolist()
@convert(str, Mesh, simple=False)
@convert(np.str_, Mesh, simple=False)
def _(value: Union[str, np.str_]) -> bytes:
if data := read_external_value(value, Mesh):
return data.tolist()
raise ConversionError()
@convert(bytes, Mesh, simple=False)
@convert(np.bytes_, Mesh, simple=False)
def _(value: Union[bytes, np.bytes_]) -> bytes:
return value
# this should not be necessary
@convert(trimesh.Trimesh, Mesh, simple=False) # type: ignore
def _(value: trimesh.Trimesh) -> bytes:
return Mesh.from_trimesh(value).encode().tolist()
@convert(list, Embedding, simple=True)
@convert(np.ndarray, Embedding, simple=True)
@convert(list, Sequence1D, simple=True)
@convert(np.ndarray, Sequence1D, simple=True)
@convert(np.ndarray, Image, simple=True)
def _(_: Union[np.ndarray, list]) -> str:
return "[...]"
@convert(str, Image, simple=True)
@convert(np.str_, Image, simple=True)
@convert(str, Audio, simple=True)
@convert(np.str_, Audio, simple=True)
@convert(str, Video, simple=True)
@convert(np.str_, Video, simple=True)
@convert(str, Mesh, simple=True)
@convert(np.str_, Mesh, simple=True)
def _(value: Union[str, np.str_]) -> str:
return str(value)
@convert(bytes, Image, simple=True)
@convert(np.bytes_, Image, simple=True)
@convert(bytes, Audio, simple=True)
@convert(np.bytes_, Audio, simple=True)
@convert(bytes, Video, simple=True)
@convert(np.bytes_, Video, simple=True)
@convert(bytes, Mesh, simple=True)
@convert(np.bytes_, Mesh, simple=True)
def _(_: Union[bytes, np.bytes_]) -> str:
return "<bytes>"
# this should not be necessary
@convert(trimesh.Trimesh, Mesh, simple=True) # type: ignore
def _(_: trimesh.Trimesh) -> str:
return "<object>"
def read_external_value(
path_or_url: Optional[str],
column_type: Type[FileBasedColumnType],
target_format: Optional[str] = None,
workdir: PathType = ".",
) -> Optional[np.void]:
"""
Read a new external value and cache it or get it from the cache if already
cached.
"""
if not path_or_url:
return None
cache_key = f"external:{path_or_url},{get_column_type_name(column_type)}"
if target_format is not None:
cache_key += f"/{target_format}"
try:
value = np.void(external_data_cache[cache_key])
return value
except KeyError:
...
value = _decode_external_value(path_or_url, column_type, target_format, workdir)
external_data_cache[cache_key] = value.tolist()
return value
def prepare_path_or_url(path_or_url: PathOrUrlType, workdir: PathType) -> str:
"""
For a relative path, prefix it with the `workdir`.
For an absolute path or an URL, do nothing.
"""
path_or_url_str = str(path_or_url)
if validators.url(path_or_url_str): # type: ignore
return path_or_url_str
return os.path.join(workdir, path_or_url_str)
def _decode_external_value(
path_or_url: PathOrUrlType,
column_type: Type[FileBasedColumnType],
target_format: Optional[str] = None,
workdir: PathType = ".",
) -> np.void:
"""
Decode an external value as expected by the rest of the backend.
"""
path_or_url = prepare_path_or_url(path_or_url, workdir)
if column_type is Audio:
file = audio.prepare_input_file(path_or_url, reusable=True)
# `file` is a filepath of type `str` or an URL downloaded as `io.BytesIO`.
input_format, input_codec = audio.get_format_codec(file)
if not isinstance(file, str):
file.seek(0)
if target_format is None:
# Try to send data as is.
if input_format in ("flac", "mp3", "wav") or input_codec in (
"aac",
"libvorbis",
"vorbis",
):
# Format is directly supported by the browser.
if isinstance(file, str):
with open(file, "rb") as f:
return np.void(f.read())
return np.void(file.read())
# Convert all other formats/codecs to flac.
output_format, output_codec = "flac", "flac"
else:
output_format, output_codec = Audio.get_format_codec(target_format)
if output_format == input_format and output_codec == input_codec:
# Nothing to transcode
if isinstance(file, str):
with open(file, "rb") as f:
return np.void(f.read())
return np.void(file.read())
buffer = io.BytesIO()
audio.transcode_audio(file, buffer, output_format, output_codec)
return np.void(buffer.getvalue())
if column_type is Image:
with as_file(path_or_url) as file:
kind = filetype.guess(file)
if kind is not None and kind.mime.split("/")[1] in (
"apng",
"avif",
"gif",
"jpeg",
"png",
"webp",
"bmp",
"x-icon",
):
return np.void(file.read())
# `image/tiff`s become blank in frontend, so convert them too.
return Image.from_file(file).encode(target_format)
data_obj = column_type.from_file(path_or_url)
return data_obj.encode(target_format) | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dtypes/conversion.py | 0.851999 | 0.439868 | conversion.py | pypi |
import io
import math
import os
from typing import Dict, IO, List, Optional, Sequence, Tuple, Union
from urllib.parse import urlparse
import imageio.v3 as iio
import numpy as np
import pygltflib
import requests
import trimesh
import validators
from loguru import logger
from renumics.spotlight.requests import headers
from renumics.spotlight.typing import FileType, NumberType, PathType
from . import exceptions, triangulation
from .base import DType, FileBasedDType
from ..io import audio, gltf, file as file_io
Array1dLike = Union[Sequence[NumberType], np.ndarray]
Array2dLike = Union[Sequence[Sequence[NumberType]], np.ndarray]
ImageLike = Union[
Sequence[Sequence[Union[NumberType, Sequence[NumberType]]]], np.ndarray
]
class Embedding(DType):
"""
Data sample projected onto a new space.
Attributes:
data: 1-dimensional array-like. Sample embedding.
dtype: Optional data type of embedding. If `None`, data type inferred
from data.
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset, Embedding
>>> value = np.array(np.random.rand(2))
>>> embedding = Embedding(value)
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_embedding_column("embeddings", 5*[embedding])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(len(dataset["embeddings", 3].data))
2
"""
data: np.ndarray
def __init__(
self, data: Array1dLike, dtype: Optional[Union[str, np.dtype]] = None
) -> None:
data_array = np.asarray(data, dtype)
if data_array.ndim != 1 or data_array.size == 0:
raise ValueError(
f"`data` argument should an array-like with shape "
f"`(num_features,)` with `num_features > 0`, but shape "
f"{data_array.shape} received."
)
if data_array.dtype.str[1] not in "fiu":
raise ValueError(
f"`data` argument should be an array-like with integer or "
f"float dtypes, but dtype {data_array.dtype.name} received."
)
self.data = data_array
@classmethod
def decode(cls, value: Union[np.ndarray, np.void]) -> "Embedding":
if not isinstance(value, np.ndarray):
raise TypeError(
f"`value` argument should be a numpy array, but {type(value)} "
f"received."
)
return cls(value)
def encode(self, _target: Optional[str] = None) -> np.ndarray:
return self.data
class Sequence1D(DType):
"""
One-dimensional ndarray with optional index values.
Attributes:
index: 1-dimensional array-like of length `num_steps`. Index values (x-axis).
value: 1-dimensional array-like of length `num_steps`. Respective values (y-axis).
dtype: Optional data type of sequence. If `None`, data type inferred
from data.
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset, Sequence1D
>>> index = np.arange(100)
>>> value = np.array(np.random.rand(100))
>>> sequence = Sequence1D(index, value)
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_sequence_1d_column("sequences", 5*[sequence])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(len(dataset["sequences", 2].value))
100
"""
index: np.ndarray
value: np.ndarray
def __init__(
self,
index: Optional[Array1dLike],
value: Optional[Array1dLike] = None,
dtype: Optional[Union[str, np.dtype]] = None,
) -> None:
if value is None:
if index is None:
raise ValueError(
"At least one of arguments `index` or `value` should be "
"set, but both `None` values received."
)
value = index
index = None
value_array = np.asarray(value, dtype)
if value_array.dtype.str[1] not in "fiu":
raise ValueError(
f"Input values should be array-likes with integer or float "
f"dtype, but dtype {value_array.dtype.name} received."
)
if index is None:
if value_array.ndim == 2:
if value_array.shape[0] == 2:
self.index = value_array[0]
self.value = value_array[1]
elif value_array.shape[1] == 2:
self.index = value_array[:, 0]
self.value = value_array[:, 1]
else:
raise ValueError(
f"A single 2-dimensional input value should have one "
f"dimension of length 2, but shape {value_array.shape} received."
)
elif value_array.ndim == 1:
self.value = value_array
if dtype is None:
dtype = self.value.dtype
self.index = np.arange(len(self.value), dtype=dtype)
else:
raise ValueError(
f"A single input value should be 1- or 2-dimensional, but "
f"shape {value_array.shape} received."
)
else:
if value_array.ndim != 1:
raise ValueError(
f"Value should be 1-dimensional, but shape {value_array.shape} received."
)
index_array = np.asarray(index, dtype)
if index_array.ndim != 1:
raise ValueError(
f"INdex should be 1-dimensional array-like, but shape "
f"{index_array.shape} received."
)
if index_array.dtype.str[1] not in "fiu":
raise ValueError(
f"Index should be array-like with integer or float "
f"dtype, but dtype {index_array.dtype.name} received."
)
self.value = value_array
self.index = index_array
if len(self.value) != len(self.index):
raise ValueError(
f"Lengths of `index` and `value` should match, but lengths "
f"{len(self.index)} and {len(self.value)} received."
)
@classmethod
def decode(cls, value: Union[np.ndarray, np.void]) -> "Sequence1D":
if not isinstance(value, np.ndarray):
raise TypeError(
f"`value` argument should be a numpy array, but {type(value)} "
f"received."
)
if value.ndim != 2 or value.shape[1] != 2:
raise ValueError(
f"`value` argument should be a numpy array with shape "
f"`(num_steps, 2)`, but shape {value.shape} received."
)
return cls(value[:, 0], value[:, 1])
def encode(self, _target: Optional[str] = None) -> np.ndarray:
return np.stack((self.index, self.value), axis=1)
@classmethod
def empty(cls) -> "Sequence1D":
"""
Create an empty sequence.
"""
return cls(np.empty(0), np.empty(0))
class Mesh(FileBasedDType):
"""
Triangular 3D mesh with optional per-point and per-triangle attributes and
optional per-point displacements over time.
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset, Mesh
>>> points = np.array([[0,0,0],[1,1,1],[0,1,0],[-1,0,1]])
>>> triangles = np.array([[0,1,2],[2,3,0]])
>>> mesh = Mesh(points, triangles)
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_mesh_column("meshes", 5*[mesh])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["meshes", 2].triangles)
[[0 1 2]
[2 3 0]]
"""
_points: np.ndarray
_triangles: np.ndarray
_point_attributes: Dict[str, np.ndarray]
_point_displacements: List[np.ndarray]
_point_indices: np.ndarray
_triangle_indices: np.ndarray
_triangle_attribute_indices: np.ndarray
def __init__(
self,
points: Array2dLike,
triangles: Array2dLike,
point_attributes: Optional[Dict[str, np.ndarray]] = None,
triangle_attributes: Optional[Dict[str, np.ndarray]] = None,
point_displacements: Optional[Union[np.ndarray, List[np.ndarray]]] = None,
):
self._point_attributes = {}
self._point_displacements = []
self._set_points_triangles(points, triangles)
if point_displacements is None:
point_displacements = []
self.point_displacements = point_displacements # type: ignore
self.update_attributes(point_attributes, triangle_attributes)
@property
def points(self) -> np.ndarray:
"""
:code:`np.array` with shape `(num_points, 3)`. Mesh points.
"""
return self._points
@property
def triangles(self) -> np.ndarray:
"""
:code:`np.array` with shape `(num_triangles, 3)`. Mesh triangles stored as their
CCW nodes referring to the `points` indices.
"""
return self._triangles
@property
def point_attributes(self) -> Dict[str, np.ndarray]:
"""
Mapping str -> :code:`np.array` with shape `(num_points, ...)`. Point-wise
attributes corresponding to `points`. All possible shapes of a single
attribute can be found in
`renumics.spotlight.mesh_proc.gltf.GLTF_SHAPES`.
"""
return self._point_attributes
@property
def point_displacements(self) -> List[np.ndarray]:
"""
List of arrays with shape `(num_points, 3)`. Point-wise relative
displacements (offsets) over the time corresponding to `points`.
Timestep 0 is omitted since it is explicit stored as absolute values in
`points`.
"""
return self._point_displacements
@point_displacements.setter
def point_displacements(self, value: Union[np.ndarray, List[np.ndarray]]) -> None:
array = triangulation.attribute_to_array(value)
if array.size == 0:
self._point_displacements = []
else:
array = array.astype(np.float32)
if array.shape[1] != len(self._points):
array = array[:, self._point_indices]
if array.shape[1:] != (len(self._points), 3):
raise ValueError(
f"Point displacements should have the same shape as points "
f"({self._points.shape}), but shape {array.shape[1:]} "
f"received."
)
self._point_displacements = list(array)
@classmethod
def from_trimesh(cls, mesh: trimesh.Trimesh) -> "Mesh":
"""
Import a `trimesh.Trimesh` mesh.
"""
return cls(
mesh.vertices, mesh.faces, mesh.vertex_attributes, mesh.face_attributes
)
@classmethod
def from_file(cls, filepath: PathType) -> "Mesh":
"""
Read mesh from a filepath or an URL.
`trimesh` is used inside, so only supported formats are allowed.
"""
file: Union[str, IO] = (
str(filepath) if isinstance(filepath, os.PathLike) else filepath
)
extension = None
if isinstance(file, str):
if validators.url(file):
response = requests.get(file, headers=headers, timeout=30)
if not response.ok:
raise exceptions.InvalidFile(f"URL {file} does not exist.")
extension = os.path.splitext(urlparse(file).path)[1]
if extension == "":
raise exceptions.InvalidFile(f"URL {file} has no file extension.")
file = io.BytesIO(response.content)
elif not os.path.isfile(file):
raise exceptions.InvalidFile(
f"File {file} is neither an existing file nor an existing URL."
)
try:
mesh = trimesh.load(file, file_type=extension, force="mesh")
except Exception as e:
raise exceptions.InvalidFile(
f"Mesh {filepath} does not exist or could not be read."
) from e
return cls.from_trimesh(mesh)
@classmethod
def empty(cls) -> "Mesh":
"""
Create an empty mesh.
"""
return cls(np.empty((0, 3)), np.empty((0, 3), np.int64))
@classmethod
def decode(cls, value: Union[np.ndarray, np.void]) -> "Mesh":
gltf_mesh = pygltflib.GLTF2.load_from_bytes(value.tobytes())
gltf.check_gltf(gltf_mesh)
arrays = gltf.decode_gltf_arrays(gltf_mesh)
primitive = gltf_mesh.meshes[0].primitives[0]
points = arrays[primitive.attributes.POSITION]
triangles = arrays[primitive.indices].reshape((-1, 3))
point_attributes = {
k[1:]: arrays[v]
for k, v in primitive.attributes.__dict__.items()
if k.startswith("_")
}
point_displacements = [
arrays[target["POSITION"]] for target in primitive.targets
]
return cls(
points, triangles, point_attributes, point_displacements=point_displacements
)
def encode(self, _target: Optional[str] = None) -> np.void:
bin_data, buffer_views, accessors = gltf.encode_gltf_array(
self._triangles.flatten(), b"", [], [], pygltflib.ELEMENT_ARRAY_BUFFER
)
mesh_primitive_attributes_kwargs = {"POSITION": 1}
bin_data, buffer_views, accessors = gltf.encode_gltf_array(
self._points, bin_data, buffer_views, accessors
)
for attr_name, point_attr in self._point_attributes.items():
mesh_primitive_attributes_kwargs["_" + attr_name] = len(buffer_views)
bin_data, buffer_views, accessors = gltf.encode_gltf_array(
point_attr, bin_data, buffer_views, accessors
)
morph_targets = []
for point_displacement in self._point_displacements:
morph_targets.append(pygltflib.Attributes(POSITION=len(buffer_views)))
bin_data, buffer_views, accessors = gltf.encode_gltf_array(
point_displacement, bin_data, buffer_views, accessors
)
gltf_mesh = pygltflib.GLTF2(
asset=pygltflib.Asset(),
scene=0,
scenes=[pygltflib.Scene(nodes=[0])],
nodes=[pygltflib.Node(mesh=0)],
meshes=[
pygltflib.Mesh(
primitives=[
pygltflib.Primitive(
attributes=pygltflib.Attributes(
**mesh_primitive_attributes_kwargs
),
indices=0,
mode=pygltflib.TRIANGLES,
targets=morph_targets,
)
],
)
],
accessors=accessors,
bufferViews=buffer_views,
buffers=[pygltflib.Buffer(byteLength=len(bin_data))],
)
gltf_mesh.set_binary_blob(bin_data)
return np.void(b"".join(gltf_mesh.save_to_bytes()))
def update_attributes(
self,
point_attributes: Optional[Dict[str, np.ndarray]] = None,
triangle_attributes: Optional[Dict[str, np.ndarray]] = None,
) -> None:
"""
Update point and/or triangle attributes dict-like.
"""
if point_attributes:
point_attributes = self._sanitize_point_attributes(point_attributes)
self._point_attributes.update(point_attributes)
if triangle_attributes:
triangle_attributes = self._sanitize_triangle_attributes(
triangle_attributes
)
logger.info("Triangle attributes will be converted to point attributes.")
self._point_attributes.update(
self._triangle_attributes_to_point_attributes(triangle_attributes)
)
def interpolate_point_displacements(self, num_timesteps: int) -> None:
"""subsample time dependent attributes with new time step count"""
if num_timesteps < 1:
raise ValueError(
f"`num_timesteps` argument should be non-negative, but "
f"{num_timesteps} received."
)
current_num_timesteps = len(self._point_displacements)
if current_num_timesteps == 0:
logger.info("No displacements found, so cannot interpolate.")
return
if current_num_timesteps == num_timesteps:
return
def _interpolated_list_access(
arrays: List[np.ndarray], index_float: float
) -> np.ndarray:
"""access a list equally sized numpy arrays with interpolation between two neighbors"""
array_left = arrays[math.floor(index_float)]
array_right = arrays[math.ceil(index_float)]
weight_right = index_float - math.floor(index_float)
return (array_left * (1 - weight_right)) + (array_right * weight_right)
# simplification assumption : timesteps are equally sized
timesteps = np.linspace(0, current_num_timesteps, num_timesteps + 1)[1:]
# add implicit 0 displacement for t=0
displacements = [
np.zeros_like(self._point_displacements[0])
] + self._point_displacements
self._point_displacements = [
_interpolated_list_access(displacements, t) for t in timesteps
]
def _set_points_triangles(
self, points: Array2dLike, triangles: Array2dLike
) -> None:
# Check points.
points_array = np.asarray(points, np.float32)
if points_array.ndim != 2 or points_array.shape[1] != 3:
raise ValueError(
f"`points` argument should be a numpy array with shape "
f"`(num_points, 3)`, but shape {points_array.shape} received."
)
# Check triangles.
triangles_array = np.asarray(triangles, np.uint32)
if triangles_array.ndim != 2 or triangles_array.shape[1] != 3:
raise ValueError(
f"`triangles` argument should be a numpy array with shape "
f"`(num_triangles, 3)`, but shape {triangles_array.shape} received."
)
# Subsample only valid points and triangles.
point_ids = np.arange(len(points_array))
valid_triangles_mask = (
(triangles_array[:, 0] != triangles_array[:, 1])
& (triangles_array[:, 0] != triangles_array[:, 2])
& (triangles_array[:, 1] != triangles_array[:, 2])
& np.isin(triangles_array, point_ids).all(axis=1)
)
self._triangle_indices = np.nonzero(valid_triangles_mask)[0]
self._triangles = triangles_array[self._triangle_indices]
valid_points_mask = np.isin(point_ids, self._triangles)
self._point_indices = np.nonzero(valid_points_mask)[0]
self._points = points_array[self._point_indices]
# Reindex triangles since there can be fewer points than before.
point_ids = point_ids[self._point_indices]
self._triangles, *_ = triangulation.reindex(point_ids, self._triangles)
# Set indices for conversion of triangle attributes to point attributes.
self._triangle_attribute_indices = np.full((len(self._points)), 0, np.uint32)
triangle_indices = np.arange(len(self._triangles), dtype=np.uint32)
self._triangle_attribute_indices[self._triangles[:, 0]] = triangle_indices
self._triangle_attribute_indices[self._triangles[:, 1]] = triangle_indices
self._triangle_attribute_indices[self._triangles[:, 2]] = triangle_indices
def _triangle_attributes_to_point_attributes(
self, triangle_attributes: Dict[str, np.ndarray]
) -> Dict[str, np.ndarray]:
return {
f"element_{attr_name}": triangle_attr[self._triangle_attribute_indices]
for attr_name, triangle_attr in triangle_attributes.items()
}
def _sanitize_point_attributes(
self, point_attributes: Dict[str, np.ndarray]
) -> Dict[str, np.ndarray]:
if not isinstance(point_attributes, dict):
raise TypeError(
f"`point_attributes` argument should be a dict, but "
f"{type(point_attributes)} received."
)
valid_point_attributes = {}
for attr_name, point_attr in point_attributes.items():
point_attr = np.asarray(point_attr)
if len(point_attr) != len(self._points):
point_attr = point_attr[self._point_indices]
if point_attr.dtype.str[1] not in "fiu":
raise ValueError(
f"Point attributes should have one of integer or float "
f'dtypes, but attribute "{attr_name}" of dtype '
f"{point_attr.dtype.name} received."
)
point_attr = point_attr.squeeze()
if point_attr.shape[1:] not in gltf.GLTF_SHAPES_LOOKUP.keys():
logger.warning(
f"Element shape {point_attr.shape[1:]} of the point "
f'attribute "{attr_name}" not supported, attribute will be '
f"removed."
)
continue
valid_point_attributes[attr_name] = point_attr.astype(
gltf.GLTF_DTYPES_CONVERSION[point_attr.dtype.str[1:]]
)
return valid_point_attributes
def _sanitize_triangle_attributes(
self, triangle_attributes: Dict[str, np.ndarray]
) -> Dict[str, np.ndarray]:
if not isinstance(triangle_attributes, dict):
raise TypeError(
f"`triangle_attributes` argument should be a dict, but "
f"{type(triangle_attributes)} received."
)
valid_triangle_attributes = {}
for attr_name, triangle_attr in triangle_attributes.items():
triangle_attr = np.asarray(triangle_attr)
if len(triangle_attr) != len(self._triangles):
triangle_attr = triangle_attr[self._triangle_indices]
if triangle_attr.dtype.str[1] not in "fiu":
raise ValueError(
f"Triangle attributes should have one of integer or float "
f'dtypes, but attribute "{attr_name}" of dtype '
f"{triangle_attr.dtype.name} received."
)
triangle_attr = triangle_attr.squeeze()
if triangle_attr.shape[1:] not in gltf.GLTF_SHAPES_LOOKUP.keys():
logger.warning(
f"Element shape {triangle_attr.shape[1:]} of the triangle "
f'attribute "{attr_name}" not supported, attribute will be '
f"removed."
)
continue
valid_triangle_attributes[attr_name] = triangle_attr.astype(
gltf.GLTF_DTYPES_CONVERSION[triangle_attr.dtype.str[1:]]
)
return valid_triangle_attributes
class Image(FileBasedDType):
"""
An RGB(A) or grayscale image that will be saved in encoded form.
Attributes:
data: Array-like with shape `(num_rows, num_columns)` or
`(num_rows, num_columns, num_channels)` with `num_channels` equal to
3, or 4; with dtype "uint8".
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset, Image
>>> data = np.full([100,100,3], 255, dtype=np.uint8) # white uint8 image
>>> image = Image(data)
>>> float_data = np.random.uniform(0, 1, (100, 100)) # random grayscale float image
>>> float_image = Image(float_data)
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_image_column("images", [image, float_image, data, float_data])
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["images", 0].data[50][50])
... print(dataset["images", 3].data.dtype)
[255 255 255]
uint8
"""
data: np.ndarray
def __init__(self, data: ImageLike) -> None:
data_array = np.asarray(data)
if (
data_array.size == 0
or data_array.ndim != 2
and (data_array.ndim != 3 or data_array.shape[-1] not in (1, 3, 4))
):
raise ValueError(
f"`data` argument should be a numpy array with shape "
f"`(num_rows, num_columns, num_channels)` or "
f"`(num_rows, num_columns)` or with `num_rows > 0`, "
f"`num_cols > 0` and `num_channels` equal to 1, 3, or 4, but "
f"shape {data_array.shape} received."
)
if data_array.dtype.str[1] not in "fiu":
raise ValueError(
f"`data` argument should be a numpy array with integer or "
f"float dtypes, but dtype {data_array.dtype.name} received."
)
if data_array.ndim == 3 and data_array.shape[2] == 1:
data_array = data_array.squeeze(axis=2)
if data_array.dtype.str[1] == "f":
logger.info(
'Image data converted to "uint8" dtype by multiplication with '
"255 and rounding."
)
data_array = (255 * data_array).round()
self.data = data_array.astype("uint8")
@classmethod
def from_file(cls, filepath: FileType) -> "Image":
"""
Read image from a filepath, an URL, or a file-like object.
`imageio` is used inside, so only supported formats are allowed.
"""
with file_io.as_file(filepath) as file:
try:
image_array = iio.imread(file, index=False) # type: ignore
except Exception as e:
raise exceptions.InvalidFile(
f"Image {filepath} does not exist or could not be read."
) from e
return cls(image_array)
@classmethod
def from_bytes(cls, blob: bytes) -> "Image":
"""
Read image from raw bytes.
`imageio` is used inside, so only supported formats are allowed.
"""
try:
image_array = iio.imread(blob, index=False) # type: ignore
except Exception as e:
raise exceptions.InvalidFile(
"Image could not be read from the given bytes."
) from e
return cls(image_array)
@classmethod
def empty(cls) -> "Image":
"""
Create a transparent 1 x 1 image.
"""
return cls(np.zeros((1, 1, 4), np.uint8))
@classmethod
def decode(cls, value: Union[np.ndarray, np.void]) -> "Image":
if isinstance(value, np.void):
buffer = io.BytesIO(value.tolist())
return cls(iio.imread(buffer, extension=".png", index=False))
raise TypeError(
f"`value` should be a `numpy.void` instance, but {type(value)} "
f"received."
)
def encode(self, _target: Optional[str] = None) -> np.void:
buf = io.BytesIO()
iio.imwrite(buf, self.data, extension=".png")
return np.void(buf.getvalue())
class Audio(FileBasedDType):
"""
An Audio Signal that will be saved in encoded form.
All formats and codecs supported by AV are supported for read.
Attributes:
data: Array-like with shape `(num_samples, num_channels)`
with `num_channels` <= 5.
If `data` has a float dtype, its values should be between -1 and 1.
If `data` has an int dtype, its values should be between minimum and
maximum possible values for the particular int dtype.
If `data` has an unsigned int dtype, ist values should be between 0
and maximum possible values for the particular unsigned int dtype.
sampling_rate: Sampling rate (samples per seconds)
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset, Audio
>>> samplerate = 44100
>>> fs = 100 # 100 Hz audio signal
>>> time = np.linspace(0.0, 1.0, samplerate)
>>> amplitude = np.iinfo(np.int16).max * 0.4
>>> data = np.array(amplitude * np.sin(2.0 * np.pi * fs * time), dtype=np.int16)
>>> audio = Audio(samplerate, np.array([data, data]).T) # int16 stereo signal
>>> float_data = 0.5 * np.cos(2.0 * np.pi * fs * time).astype(np.float32)
>>> float_audio = Audio(samplerate, float_data) # float32 mono signal
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_audio_column("audio", [audio, float_audio])
... dataset.append_audio_column("lossy_audio", [audio, float_audio], lossy=True)
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["audio", 0].data[100])
... print(f"{dataset['lossy_audio', 1].data[0, 0]:.5g}")
[12967 12967]
0.4596
"""
data: np.ndarray
sampling_rate: int
def __init__(self, sampling_rate: int, data: Array2dLike) -> None:
data_array = np.asarray(data)
is_valid_multi_channel = (
data_array.size > 0 and data_array.ndim == 2 and data_array.shape[1] <= 5
)
is_valid_mono = data_array.size > 0 and data_array.ndim == 1
if not (is_valid_multi_channel or is_valid_mono):
raise ValueError(
f"`data` argument should be a 1D array for mono data"
f" or a 2D numpy array with shape "
f"`(num_samples, num_channels)` and with num_channels <= 5, "
f"but shape {data_array.shape} received."
)
if data_array.dtype not in [np.float32, np.int32, np.int16, np.uint8]:
raise ValueError(
f"`data` argument should be a numpy array with "
f"dtype np.float32, np.int32, np.int16 or np.uint8, "
f"but dtype {data_array.dtype.name} received."
)
self.data = data_array
self.sampling_rate = sampling_rate
@classmethod
def from_file(cls, filepath: FileType) -> "Audio":
"""
Read audio file from a filepath, an URL, or a file-like object.
`pyav` is used inside, so only supported formats are allowed.
"""
try:
data, sampling_rate = audio.read_audio(filepath)
except Exception as e:
raise exceptions.InvalidFile(
f"Audio file {filepath} does not exist or could not be read."
) from e
return cls(sampling_rate, data)
@classmethod
def from_bytes(cls, blob: bytes) -> "Audio":
"""
Read audio from raw bytes.
`pyav` is used inside, so only supported formats are allowed.
"""
try:
data, sampling_rate = audio.read_audio(io.BytesIO(blob))
except Exception as e:
raise exceptions.InvalidFile(
"Audio could not be read from the given bytes."
) from e
return cls(sampling_rate, data)
@classmethod
def empty(cls) -> "Audio":
"""
Create a single zero-value sample stereo audio signal.
"""
return cls(1, np.zeros((1, 2), np.int16))
@classmethod
def decode(cls, value: Union[np.ndarray, np.void]) -> "Audio":
if isinstance(value, np.void):
buffer = io.BytesIO(value.tolist())
data, sampling_rate = audio.read_audio(buffer)
return cls(sampling_rate, data)
raise TypeError(
f"`value` should be a `numpy.void` instance, but {type(value)} "
f"received."
)
def encode(self, target: Optional[str] = None) -> np.void:
format_, codec = self.get_format_codec(target)
buffer = io.BytesIO()
audio.write_audio(buffer, self.data, self.sampling_rate, format_, codec)
return np.void(buffer.getvalue())
@staticmethod
def get_format_codec(target: Optional[str] = None) -> Tuple[str, str]:
"""
Get an audio format and an audio codec by an `target`.
"""
format_ = "wav" if target is None else target.lstrip(".").lower()
codec = {"wav": "pcm_s16le", "ogg": "libvorbis", "mp3": "libmp3lame"}.get(
format_, format_
)
return format_, codec
class Category(str):
"""
A string value that takes only a limited number of possible values (categories).
The corresponding categories can be got and set with get/set_column_attributes['categories'].
Dummy class for window column creation, should not be explicitly used as
input data.
Example:
>>> import numpy as np
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_categorical_column("my_new_cat",
... categories=["red", "green", "blue"],)
... dataset.append_row(my_new_cat="blue")
... dataset.append_row(my_new_cat="green")
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["my_new_cat", 1])
green
Example:
>>> import numpy as np
>>> import datetime
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_categorical_column("my_new_cat",
... categories=["red", "green", "blue"],)
... current_categories = dataset.get_column_attributes("my_new_cat")["categories"]
... dataset.set_column_attributes("my_new_cat", categories={**current_categories,
... "black":100})
... dataset.append_row(my_new_cat="black")
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["my_new_cat", 0])
black
"""
class Video(FileBasedDType):
"""
A video object. No encoding or decoding is currently performed on the python
side, so all formats will be saved into dataset without compatibility check,
but only the formats supported by your browser (apparently .mp4, .ogg,
.webm, .mov etc.) can be played in Spotlight.
"""
data: bytes
def __init__(self, data: bytes) -> None:
if not isinstance(data, bytes):
raise TypeError(
f"`data` argument should be video bytes, but type {type(data)} "
f"received."
)
self.data = data
@classmethod
def from_file(cls, filepath: PathType) -> "Video":
"""
Read video from a filepath or an URL.
"""
prepared_file = str(filepath) if isinstance(filepath, os.PathLike) else filepath
if not isinstance(prepared_file, str):
raise TypeError(
"`filepath` should be a string or an `os.PathLike` instance, "
f"but value {prepared_file} or type {type(prepared_file)} "
f"received."
)
if validators.url(prepared_file):
response = requests.get(
prepared_file, headers=headers, stream=True, timeout=10
)
if not response.ok:
raise exceptions.InvalidFile(f"URL {prepared_file} does not exist.")
return cls(response.raw.data)
if os.path.isfile(prepared_file):
with open(filepath, "rb") as f:
return cls(f.read())
raise exceptions.InvalidFile(
f"File {prepared_file} is neither an existing file nor an existing URL."
)
@classmethod
def from_bytes(cls, blob: bytes) -> "Video":
"""
Read video from raw bytes.
"""
return cls(blob)
@classmethod
def empty(cls) -> "Video":
"""
Create an empty video instance.
"""
return cls(b"\x00")
@classmethod
def decode(cls, value: Union[np.ndarray, np.void]) -> "Video":
if isinstance(value, np.void):
return cls(value.tolist())
raise TypeError(
f"`value` should be a `numpy.void` instance, but {type(value)} "
f"received."
)
def encode(self, _target: Optional[str] = None) -> np.void:
return np.void(self.data)
class Window:
"""
A pair of two timestamps in seconds which can be later projected onto
continuous data (only :class:`Audio <renumics.spotlight.dtypes.Audio>`
is currently supported).
Dummy class for window column creation
(see :func:`Dataset.append_column <renumics.spotlight.dataset.Dataset.append_column>`),
should not be explicitly used as input data.
To create a window column, use
:func:`Dataset.append_window_column <renumics.spotlight.dataset.Dataset.append_window_column>`
method.
Examples:
>>> import numpy as np
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_window_column("window", [[1, 2]] * 4)
... dataset.append_row(window=(0, 1))
... dataset.append_row(window=np.array([-1, 0]))
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["window"])
[[ 1. 2.]
[ 1. 2.]
[ 1. 2.]
[ 1. 2.]
[ 0. 1.]
[-1. 0.]]
>>> import numpy as np
>>> from renumics.spotlight import Dataset
>>> with Dataset("docs/example.h5", "w") as dataset:
... dataset.append_int_column("start", range(5))
... dataset.append_float_column("end", dataset["start"] + 2)
... print(dataset["start"])
... print(dataset["end"])
[0 1 2 3 4]
[2. 3. 4. 5. 6.]
>>> with Dataset("docs/example.h5", "a") as dataset:
... dataset.append_window_column("window", zip(dataset["start"], dataset["end"]))
>>> with Dataset("docs/example.h5", "r") as dataset:
... print(dataset["window"])
[[0. 2.]
[1. 3.]
[2. 4.]
[3. 5.]
[4. 6.]]
""" | /renumics_spotlight-1.3.0-py3-none-any.whl/renumics/spotlight/dtypes/__init__.py | 0.861013 | 0.52342 | __init__.py | pypi |
class PtzAPIMixin:
"""
API for PTZ functions.
"""
def _send_operation(self, operation, speed, index=None):
if index is None:
data = [{"cmd": "PtzCtrl", "action": 0, "param": {"channel": 0, "op": operation, "speed": speed}}]
else:
data = [{"cmd": "PtzCtrl", "action": 0, "param": {
"channel": 0, "op": operation, "speed": speed, "id": index}}]
return self._execute_command('PtzCtrl', data)
def _send_noparm_operation(self, operation):
data = [{"cmd": "PtzCtrl", "action": 0, "param": {"channel": 0, "op": operation}}]
return self._execute_command('PtzCtrl', data)
def _send_set_preset(self, operation, enable, preset=1, name='pos1'):
data = [{"cmd": "SetPtzPreset", "action": 0, "param": {
"channel": 0, "enable": enable, "id": preset, "name": name}}]
return self._execute_command('PtzCtrl', data)
def go_to_preset(self, speed=60, index=1):
"""
Move the camera to a preset location
:return: response json
"""
return self._send_operation('ToPos', speed=speed, index=index)
def add_preset(self, preset=1, name='pos1'):
"""
Adds the current camera position to the specified preset.
:return: response json
"""
return self._send_set_preset('PtzPreset', enable=1, preset=preset, name=name)
def remove_preset(self, preset=1, name='pos1'):
"""
Removes the specified preset
:return: response json
"""
return self._send_set_preset('PtzPreset', enable=0, preset=preset, name=name)
def move_right(self, speed=25):
"""
Move the camera to the right
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('Right', speed=speed)
def move_right_up(self, speed=25):
"""
Move the camera to the right and up
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('RightUp', speed=speed)
def move_right_down(self, speed=25):
"""
Move the camera to the right and down
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('RightDown', speed=speed)
def move_left(self, speed=25):
"""
Move the camera to the left
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('Left', speed=speed)
def move_left_up(self, speed=25):
"""
Move the camera to the left and up
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('LeftUp', speed=speed)
def move_left_down(self, speed=25):
"""
Move the camera to the left and down
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('LeftDown', speed=speed)
def move_up(self, speed=25):
"""
Move the camera up.
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('Up', speed=speed)
def move_down(self, speed=25):
"""
Move the camera down.
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('Down', speed=speed)
def stop_ptz(self):
"""
Stops the cameras current action.
:return: response json
"""
return self._send_noparm_operation('Stop')
def auto_movement(self, speed=25):
"""
Move the camera in a clockwise rotation.
The camera moves self.stop_ptz() is called.
:return: response json
"""
return self._send_operation('Auto', speed=speed) | /reolink_api-0.0.5-py3-none-any.whl/api/ptz.py | 0.722918 | 0.381853 | ptz.py | pypi |
class UserAPIMixin:
"""User-related API calls."""
def get_online_user(self) -> object:
"""
Return a list of current logged-in users in json format
See examples/response/GetOnline.json for example response data.
:return: response json
"""
body = [{"cmd": "GetOnline", "action": 1, "param": {}}]
return self._execute_command('GetOnline', body)
def get_users(self) -> object:
"""
Return a list of user accounts from the camera in json format.
See examples/response/GetUser.json for example response data.
:return: response json
"""
body = [{"cmd": "GetUser", "action": 1, "param": {}}]
return self._execute_command('GetUser', body)
def add_user(self, username: str, password: str, level: str = "guest") -> bool:
"""
Add a new user account to the camera
:param username: The user's username
:param password: The user's password
:param level: The privilege level 'guest' or 'admin'. Default is 'guest'
:return: whether the user was added successfully
"""
body = [{"cmd": "AddUser", "action": 0,
"param": {"User": {"userName": username, "password": password, "level": level}}}]
r_data = self._execute_command('AddUser', body)[0]
if r_data["value"]["rspCode"] == 200:
return True
print("Could not add user. Camera responded with:", r_data["value"])
return False
def modify_user(self, username: str, password: str) -> bool:
"""
Modify the user's password by specifying their username
:param username: The user which would want to be modified
:param password: The new password
:return: whether the user was modified successfully
"""
body = [{"cmd": "ModifyUser", "action": 0, "param": {"User": {"userName": username, "password": password}}}]
r_data = self._execute_command('ModifyUser', body)[0]
if r_data["value"]["rspCode"] == 200:
return True
print("Could not modify user:", username, "\nCamera responded with:", r_data["value"])
return False
def delete_user(self, username: str) -> bool:
"""
Delete a user by specifying their username
:param username: The user which would want to be deleted
:return: whether the user was deleted successfully
"""
body = [{"cmd": "DelUser", "action": 0, "param": {"User": {"userName": username}}}]
r_data = self._execute_command('DelUser', body)[0]
if r_data["value"]["rspCode"] == 200:
return True
print("Could not delete user:", username, "\nCamera responded with:", r_data["value"])
return False | /reolink_api-0.0.5-py3-none-any.whl/api/user.py | 0.884912 | 0.328826 | user.py | pypi |
class DisplayAPIMixin:
"""API calls related to the current image (osd, on screen display)."""
def get_osd(self) -> object:
"""
Get OSD information.
See examples/response/GetOsd.json for example response data.
:return: response json
"""
body = [{"cmd": "GetOsd", "action": 1, "param": {"channel": 0}}]
return self._execute_command('GetOsd', body)
def get_mask(self) -> object:
"""
Get the camera mask information.
See examples/response/GetMask.json for example response data.
:return: response json
"""
body = [{"cmd": "GetMask", "action": 1, "param": {"channel": 0}}]
return self._execute_command('GetMask', body)
def set_osd(self, bg_color: bool = 0, channel: int = 0, osd_channel_enabled: bool = 0, osd_channel_name: str = "",
osd_channel_pos: str = "Lower Right", osd_time_enabled: bool = 0,
osd_time_pos: str = "Lower Right") -> bool:
"""
Set OSD
:param bg_color: bool
:param channel: int channel id
:param osd_channel_enabled: bool
:param osd_channel_name: string channel name
:param osd_channel_pos: string channel position ["Upper Left","Top Center","Upper Right","Lower Left","Bottom Center","Lower Right"]
:param osd_time_enabled: bool
:param osd_time_pos: string time position ["Upper Left","Top Center","Upper Right","Lower Left","Bottom Center","Lower Right"]
:return: whether the action was successful
"""
body = [{"cmd": "SetOsd", "action": 1, "param": {
"Osd": {"bgcolor": bg_color, "channel": channel,
"osdChannel": {"enable": osd_channel_enabled, "name": osd_channel_name,
"pos": osd_channel_pos},
"osdTime": {"enable": osd_time_enabled, "pos": osd_time_pos}
}
}}]
r_data = self._execute_command('SetOsd', body)[0]
if r_data["value"]["rspCode"] == 200:
return True
print("Could not set OSD. Camera responded with status:", r_data["value"])
return False | /reolink_api-0.0.5-py3-none-any.whl/api/display.py | 0.808559 | 0.455017 | display.py | pypi |
class NetworkAPIMixin:
"""API calls for network settings."""
def set_net_port(self, http_port=80, https_port=443, media_port=9000, onvif_port=8000, rtmp_port=1935,
rtsp_port=554) -> bool:
"""
Set network ports
If nothing is specified, the default values will be used
:param rtsp_port: int
:param rtmp_port: int
:param onvif_port: int
:param media_port: int
:param https_port: int
:type http_port: int
:return: bool
"""
body = [{"cmd": "SetNetPort", "action": 0, "param": {"NetPort": {
"httpPort": http_port,
"httpsPort": https_port,
"mediaPort": media_port,
"onvifPort": onvif_port,
"rtmpPort": rtmp_port,
"rtspPort": rtsp_port
}}}]
self._execute_command('SetNetPort', body, multi=True)
print("Successfully Set Network Ports")
return True
def set_wifi(self, ssid, password) -> object:
body = [{"cmd": "SetWifi", "action": 0, "param": {
"Wifi": {
"ssid": ssid,
"password": password
}}}]
return self._execute_command('SetWifi', body)
def get_net_ports(self) -> object:
"""
Get network ports
See examples/response/GetNetworkAdvanced.json for example response data.
:return: response json
"""
body = [{"cmd": "GetNetPort", "action": 1, "param": {}},
{"cmd": "GetUpnp", "action": 0, "param": {}},
{"cmd": "GetP2p", "action": 0, "param": {}}]
return self._execute_command('GetNetPort', body, multi=True)
def get_wifi(self):
body = [{"cmd": "GetWifi", "action": 1, "param": {}}]
return self._execute_command('GetWifi', body)
def scan_wifi(self):
body = [{"cmd": "ScanWifi", "action": 1, "param": {}}]
return self._execute_command('ScanWifi', body)
def get_network_general(self) -> object:
"""
Get the camera information
See examples/response/GetNetworkGeneral.json for example response data.
:return: response json
"""
body = [{"cmd": "GetLocalLink", "action": 0, "param": {}}]
return self._execute_command('GetLocalLink', body)
def get_network_ddns(self) -> object:
"""
Get the camera DDNS network information
See examples/response/GetNetworkDDNS.json for example response data.
:return: response json
"""
body = [{"cmd": "GetDdns", "action": 0, "param": {}}]
return self._execute_command('GetDdns', body)
def get_network_ntp(self) -> object:
"""
Get the camera NTP network information
See examples/response/GetNetworkNTP.json for example response data.
:return: response json
"""
body = [{"cmd": "GetNtp", "action": 0, "param": {}}]
return self._execute_command('GetNtp', body)
def get_network_email(self) -> object:
"""
Get the camera email network information
See examples/response/GetNetworkEmail.json for example response data.
:return: response json
"""
body = [{"cmd": "GetEmail", "action": 0, "param": {}}]
return self._execute_command('GetEmail', body)
def get_network_ftp(self) -> object:
"""
Get the camera FTP network information
See examples/response/GetNetworkFtp.json for example response data.
:return: response json
"""
body = [{"cmd": "GetFtp", "action": 0, "param": {}}]
return self._execute_command('GetFtp', body)
def get_network_push(self) -> object:
"""
Get the camera push network information
See examples/response/GetNetworkPush.json for example response data.
:return: response json
"""
body = [{"cmd": "GetPush", "action": 0, "param": {}}]
return self._execute_command('GetPush', body)
def get_network_status(self) -> object:
"""
Get the camera status network information
See examples/response/GetNetworkGeneral.json for example response data.
:return: response json
"""
return self.get_network_general() | /reolink_api-0.0.5-py3-none-any.whl/api/network.py | 0.710929 | 0.410284 | network.py | pypi |
class ImageAPIMixin:
"""API calls for image settings."""
def set_adv_image_settings(self,
anti_flicker='Outdoor',
exposure='Auto',
gain_min=1,
gain_max=62,
shutter_min=1,
shutter_max=125,
blue_gain=128,
red_gain=128,
white_balance='Auto',
day_night='Auto',
back_light='DynamicRangeControl',
blc=128,
drc=128,
rotation=0,
mirroring=0,
nr3d=1) -> object:
"""
Sets the advanced camera settings.
:param anti_flicker: string
:param exposure: string
:param gain_min: int
:param gain_max: string
:param shutter_min: int
:param shutter_max: int
:param blue_gain: int
:param red_gain: int
:param white_balance: string
:param day_night: string
:param back_light: string
:param blc: int
:param drc: int
:param rotation: int
:param mirroring: int
:param nr3d: int
:return: response
"""
body = [{
"cmd": "SetIsp",
"action": 0,
"param": {
"Isp": {
"channel": 0,
"antiFlicker": anti_flicker,
"exposure": exposure,
"gain": {"min": gain_min, "max": gain_max},
"shutter": {"min": shutter_min, "max": shutter_max},
"blueGain": blue_gain,
"redGain": red_gain,
"whiteBalance": white_balance,
"dayNight": day_night,
"backLight": back_light,
"blc": blc,
"drc": drc,
"rotation": rotation,
"mirroring": mirroring,
"nr3d": nr3d
}
}
}]
return self._execute_command('SetIsp', body)
def set_image_settings(self,
brightness=128,
contrast=62,
hue=1,
saturation=125,
sharpness=128) -> object:
"""
Sets the camera image settings.
:param brightness: int
:param contrast: string
:param hue: int
:param saturation: int
:param sharpness: int
:return: response
"""
body = [
{
"cmd": "SetImage",
"action": 0,
"param": {
"Image": {
"bright": brightness,
"channel": 0,
"contrast": contrast,
"hue": hue,
"saturation": saturation,
"sharpen": sharpness
}
}
}
]
return self._execute_command('SetImage', body) | /reolink_api-0.0.5-py3-none-any.whl/api/image.py | 0.933157 | 0.266295 | image.py | pypi |
import re
version_regex = re.compile(r"^v(?P<major>[0-9]+)\.(?P<middle>[0-9]+)\.(?P<minor>[0-9]+).(?P<build>[0-9]+)_([0-9]+)")
class SoftwareVersion:
def __init__(self, version_string: str):
self.version_string = version_string
self.is_unknown = False
self.major = 0
self.middle = 0
self.minor = 0
self.build = 0
if version_string.lower() == 'unknown':
self.is_unknown = True
return
match = version_regex.match(version_string)
if match is None:
raise Exception("version_string has invalid version format: {}".format(version_string))
self.major = int(match.group("major"))
self.middle = int(match.group("middle"))
self.minor = int(match.group("minor"))
build = match.group("build")
if build is None:
self.build = 0
else:
self.build = int(match.group("build"))
def is_greater_than(self, target_version: 'SoftwareVersion'):
if self.major > target_version.major:
return True
if target_version.major == self.major:
if self.middle > target_version.middle:
return True
if target_version.middle == self.middle:
if self.minor > target_version.minor:
return True
if target_version.minor == self.minor:
if self.build > target_version.build:
return True
return False
def is_greater_or_equal_than(self, target_version: 'SoftwareVersion'):
if self.major > target_version.major:
return True
if target_version.major == self.major:
if self.middle > target_version.middle:
return True
if target_version.middle == self.middle:
if self.minor > target_version.minor:
return True
if target_version.minor == self.minor:
if self.build >= target_version.build:
return True
return False
def is_lower_than(self, target_version: 'SoftwareVersion'):
if self.major < target_version.major:
return True
if target_version.major == self.major:
if self.middle < target_version.middle:
return True
if target_version.middle == self.middle:
if self.minor < target_version.minor:
return True
if target_version.minor == self.minor:
if self.build < target_version.build:
return True
return False
def is_lower_or_equal_than(self, target_version: 'SoftwareVersion'):
if self.major < target_version.major:
return True
if target_version.major == self.major:
if self.middle < target_version.middle:
return True
if target_version.middle == self.middle:
if self.minor < target_version.minor:
return True
if target_version.minor == self.minor:
if self.build <= target_version.build:
return True
return False
def equals(self, target_version: 'SoftwareVersion'):
if target_version.major == self.major and target_version.middle == self.middle and \
target_version.minor == self.minor and target_version.build == self.build:
return True
return False
def __lt__(self, other):
return self.is_lower_than(other)
def __le__(self, other):
return self.is_lower_or_equal_than(other)
def __gt__(self, other):
return self.is_greater_than(other)
def __ge__(self, other):
return self.is_greater_or_equal_than(other)
def __eq__(self, target_version):
if target_version.major == self.major and target_version.middle == self.middle and \
target_version.minor == self.minor and target_version.build == self.build:
return True
return False
def generate_str_from_numbers(self):
return "{}.{}.{}-{}".format(self.major, self.middle, self.minor, self.build)
#endof class SoftwareVersion | /reolink_ip-0.0.49-py3-none-any.whl/reolink_ip/software_version.py | 0.440469 | 0.218784 | software_version.py | pypi |
from itertools import combinations, chain
from math import ceil
from typing import Set, List
from reoptimization_algorithms.utils.graph.edge import Edge
from reoptimization_algorithms.utils.graph.pvc import PVCUtils
from reoptimization_algorithms.utils.graph.undirected_graph import UndirectedGraph
class UnweightedPVCP:
"""
Class containing reoptimization algorithms of unweighted path vertex cover
"""
@staticmethod
def reoptimize_ptas(
old_graph: "UndirectedGraph",
attach_graph: "UndirectedGraph",
attach_edges: List[Edge],
old_solution: Set[str],
k: int,
epsilon: float = 0.25,
) -> Set[str]:
"""
:math:`(1+\\epsilon)` PTAS approximation for reoptimization of unweighted k path vertex cover under constant size graph insertion
For formalisms and algorithm details refer [1]_
:param old_graph: Old graph
:type old_graph: List[Edge]
:param attach_graph: Constant size graph which is to be inserted
:type attach_graph: UndirectedGraph
:param attach_edges: Edges connecting the old graph and attach graph
:type attach_edges: List[Edge]
:param old_solution: Vertices denoting k-PVCP Solution to old graph
:type old_solution: Set[str]
:param k: length of paths to cover
:type k: int
:param epsilon: epsilon in :math:`(1+\\epsilon)` PTAS approximation
:type epsilon: float, optional (default = 0.25)
:return: Set of vertices
Example
~~~~~~~
.. code-block:: python
import reoptimization_algorithms as ra
old_graph = (ra.UndirectedGraph().add_vertex("4").add_edge("4", "5").add_edge("40", "50")
.add_vertex("6").add_edge("4", "8").add_vertex("99")
.delete_vertex("6"))
attached_graph = ra.UndirectedGraph().add_edge("90", "95")
attach_edges = [ra.Edge("4", "90")]
old_solution = {"8"}
solution = ra.UnweightedPVCP.reoptimize_ptas(old_graph, attached_graph, attach_edges, old_solution, k = 3)
print(solution) # {"4"}
References
~~~~~~~~~~
.. [1] Kumar M., Kumar A., Pandu Rangan C. (2019) Reoptimization of Path Vertex Cover Problem.
In: Du DZ., Duan Z., Tian C. (eds) Computing and Combinatorics. COCOON 2019.
Lecture Notes in Computer Science, vol 11653. Springer, Cham
<https://link.springer.com/chapter/10.1007/978-3-030-26176-4_30>
"""
v_o = set(old_graph.get_vertices())
new_graph = old_graph.disjoint_graph_union(attach_graph, attach_edges)
v_n = set(new_graph.get_vertices())
v_a = v_n.difference(v_o)
if len(v_n) < k:
return set()
m = ceil(len(v_a) / epsilon)
sol_1 = v_n
for c in set(
chain.from_iterable(combinations(v_n, r) for r in range(0, m + 1))
):
candidate_vertices = set(c)
if len(candidate_vertices) < len(sol_1) and PVCUtils.is_k_pvc(
new_graph, candidate_vertices, k
):
sol_1 = candidate_vertices
sol_2 = v_a.union(old_solution)
result = sol_1 if len(sol_1) < len(sol_2) else sol_2
return result | /reoptimization-algorithms-0.1.3.tar.gz/reoptimization-algorithms-0.1.3/src/reoptimization_algorithms/algorithms/unwtd_pvcp.py | 0.938808 | 0.469459 | unwtd_pvcp.py | pypi |
from typing import Dict
from reoptimization_algorithms.utils.graph.edge import Edge
class Vertex:
"""
Vertex class having key, weight and adjacency dictionary of neighbours
Default weight as :py:attr:`Vertex.DEFAULT_VERTEX_WEIGHT`
:param key: Key
:type key: str
:param weight: Weight of the vertex
:type weight: int
:param neighbours: Neighbours of the vertex
:type neighbours: Dict[str, Edge]
"""
def __init__(
self, key: str, weight: float = None, neighbours: Dict[str, "Edge"] = None
):
"""
Vertex class having key, weight and adjacency dictionary of neighbours
Default weight as :py:attr:`Vertex.DEFAULT_VERTEX_WEIGHT`
:param key: Key
:type key: str
:param weight: Weight of the vertex
:type weight: float, optional (default = None)
:param neighbours: Neighbours of the vertex
:type neighbours: Dict[str, Edge], optional (default = None)
"""
self.__key = key
if weight is None:
weight = self.DEFAULT_VERTEX_WEIGHT
self._weight = weight
if neighbours is None:
neighbours = {}
self._neighbours = neighbours
@property
def DEFAULT_VERTEX_WEIGHT(self) -> float:
"""
Default Vertex weight
:return: 1
"""
return 1
@property
def key(self) -> str:
"""
Vertex key
:return: Vertex key
"""
return self.__key
@property
def weight(self) -> float:
"""
Vertex weight
:return: Vertex weight
"""
return self._weight
@weight.setter
def weight(self, weight: float) -> None:
"""
Vertex weight setter
:param weight: Vertex weight
:type weight: float
:return: None
"""
self._weight = weight
@property
def neighbours(self) -> Dict[str, "Edge"]:
"""
Neighbouring vertices
:return: Dictionary of neighbouring vertices as keys and values as edges
"""
return self._neighbours
@neighbours.setter
def neighbours(self, neighbours: Dict[str, "Edge"]) -> None:
"""
Neighbouring vertices
:param neighbours: Dictionary of neighbouring vertices as keys and values as edges
:type neighbours: Dict[str, Edge]
:return: None
"""
self._neighbours = neighbours
def is_neighbour_exists(self, neighbour: str) -> bool:
"""
Checks if neighbour exists
:param neighbour: Neighbour vertex key
:type neighbour: str
:return: Boolean
"""
return neighbour in self._neighbours
def add_neighbour(self, neighbour: str, weight: float = None) -> "Vertex":
"""
Adds a neighbour, default edge weight as :py:attr:`Edge.DEFAULT_EDGE_WEIGHT`
:param neighbour: Neighbour vertex key
:type neighbour: str
:param weight: Edge weight
:type weight: float, optional (default = None)
:return: Self
"""
if self.is_neighbour_exists(neighbour):
raise Exception(
f"Neighbour {neighbour} already exists for {self.__key}, delete it first"
)
self.neighbours[neighbour] = Edge(self.key, neighbour, weight)
return self
def get_neighbour(self, neighbour: str) -> "Edge":
"""
Gets a neighbour
:param neighbour: Neighbour vertex key
:type neighbour: str
:return: Edge representing the neighbour
"""
if not self.is_neighbour_exists(neighbour):
raise Exception(
f"Neighbour {neighbour} does not exists for {self.__key}, create it first"
)
return self.neighbours.get(neighbour)
def update_neighbour(self, neighbour: str, weight: float = None) -> "Vertex":
"""
Updates a neighbour, default edge weight as :py:attr:`Edge.DEFAULT_EDGE_WEIGHT`
:param neighbour: Neighbour vertex key
:type neighbour: str
:param weight: Edge weight to update
:type weight: float, optional (default = None)
:return: Self
"""
self.neighbours[neighbour] = Edge(self.key, neighbour, weight)
return self
def update_weight(self, weight: float) -> "Vertex":
"""
Updates vertex weight
:param weight: Vertex weight
:type weight: float, optional (default = None)
:return: Self
"""
self._weight = weight
return self
def delete_neighbour(self, neighbour: str) -> Edge:
"""
Deletes a neighbour
:param neighbour: Neighbour vertex key
:type neighbour: str
:return: Deleted Edge
"""
if not self.is_neighbour_exists(neighbour):
raise Exception(
f"Neighbour {neighbour} does not exists for {self.__key}, create it first"
)
return self.neighbours.pop(neighbour)
def degree(self) -> int:
"""
Gets degree of the vertex
:return: Degree of the vertex
"""
return len(self.neighbours) | /reoptimization-algorithms-0.1.3.tar.gz/reoptimization-algorithms-0.1.3/src/reoptimization_algorithms/utils/graph/vertex.py | 0.965706 | 0.713007 | vertex.py | pypi |
from reoptimization_algorithms.utils.graph.base_edge import BaseEdge
class Edge(BaseEdge):
"""
Edge class represented as source, destination and weight attributes
Default weight as :py:attr:`Edge.DEFAULT_EDGE_WEIGHT`
:param source: Source vertex key
:type source: str
:param destination: Destination vertex key
:type destination: str
:param weight: Weight
:type weight: float, optional (default = None)
"""
def __init__(self, source: str, destination: str, weight: float = None):
"""
Edge class represented as source, destination and weight attributes
Default weight as :py:attr:`Edge.DEFAULT_EDGE_WEIGHT`
:param source: Source vertex key
:type source: str
:param destination: Destination vertex key
:type destination: str
:param weight: Weight
:type weight: float, optional (default = None)
"""
self._source = source
self._destination = destination
if weight is None:
weight = self.DEFAULT_EDGE_WEIGHT
self._weight = weight
@property
def DEFAULT_EDGE_WEIGHT(self) -> float:
"""
Default Edge Weight
:return: 1
"""
return 1
@property
def source(self) -> str:
"""
Source vertex key
:return: Vertex key
"""
return self._source
@source.setter
def source(self, source: str) -> None:
"""
Source vertex key setter
:param source: Source vertex key
:type source: str
:return: None
"""
self._source = source
@property
def destination(self) -> str:
"""
Destination vertex key
:return: Destination vertex key
"""
return self._destination
@destination.setter
def destination(self, destination: str) -> None:
"""
Destination vertex key setter
:param destination: Destination vertex key
:type destination: str
:return: None
"""
self._destination = destination
@property
def weight(self) -> float:
"""
Edge weight
:return: Edge weight
"""
return self._weight
@weight.setter
def weight(self, weight: float) -> None:
"""
Edge weight setter
:param weight: Edge weight
:type weight: float
:return: None
"""
self._weight = weight | /reoptimization-algorithms-0.1.3.tar.gz/reoptimization-algorithms-0.1.3/src/reoptimization_algorithms/utils/graph/edge.py | 0.947938 | 0.494019 | edge.py | pypi |
import copy
from abc import ABC
from typing import Dict, List, TypeVar
from reoptimization_algorithms.errors.Error import InputError
from reoptimization_algorithms.utils.graph.base_graph import BaseGraph
from reoptimization_algorithms.utils.graph.edge import Edge
from reoptimization_algorithms.utils.graph.vertex import Vertex
T = TypeVar("T", bound="Graph")
class Graph(BaseGraph, ABC):
"""
Graph data structure class, represented as dictionary with vertices as key mapped to neighbouring vertices
T is a type var, bounded to this class, will refer to child class if inherited
:param graph: Graph definition, if None then empty graph is instantiated
:type graph: Dict[str, Vertex]
Example
~~~~~~~
.. code-block:: python
import reoptimization_algorithms as ra
graph = ra.Graph()
graph = graph.add_vertex("4", 14) # Adding vertex
graph.get_vertex("4") # Getting vertex
graph = graph.update_vertex("4", 15) # Update vertex weight
graph = graph.delete_vertex("4") # Deleting vertex
graph = graph.add_edge("4", "5", 11) # Adding Edge
graph.get_edge("4", "5") # Getting Edge
graph = graph.update_edge("4", "5", 10) # Update Edge weight
graph = graph.delete_edge("4", "5") # Deleting Edge
# Add, Update and Deletes can be chained as follows
graph = (ra.Graph().add_vertex("4").add_edge("4", "5").add_edge("40", "50")
.add_vertex("6").add_edge("4", "8").delete_edge("4", "5").add_vertex("99")
.delete_vertex("6"))
"""
def __init__(self, graph: Dict[str, "Vertex"] = None):
"""
Graph data structure class, represented as dictionary with vertices as key mapped to neighbouring vertices
:param graph: Graph definition, if None then empty graph is instantiated
:type graph: Dict[str, Vertex]
"""
if graph is None:
graph = {}
self._graph = graph
@property
def graph(self) -> Dict[str, "Vertex"]:
"""
Graph represented as dictionary with vertices as key mapped to neighbouring vertices
"""
return self._graph
@graph.setter
def graph(self, graph: Dict[str, "Vertex"]) -> None:
"""
Graph setter
:param graph: Graph as dictionary of vertices to set
:type graph: Dict[str, Vertex]
:return: None
"""
self._graph = graph
def is_vertex_exists(self, vertex: str) -> bool:
"""
Checks if the vertex exists in the graph
:param vertex: Vertex key
:type vertex: str
:return: Boolean
"""
return vertex in self._graph
def get_vertex(self, vertex: str) -> "Vertex":
"""
Gets the vertex
:param vertex: Vertex key
:type vertex: str
:return: Vertex
"""
if not self.is_vertex_exists(vertex):
raise Exception(f"Vertex {vertex} does not exists, create it first")
return self._graph.get(vertex)
def add_vertex(self: T, vertex: str, weight: float = None) -> T:
"""
Adds a vertex to the graph, default weight as :py:attr:`Vertex.DEFAULT_VERTEX_WEIGHT <reoptimization_algorithms.utils.graph.vertex.Vertex.DEFAULT_VERTEX_WEIGHT>`
:param vertex: Vertex key
:type vertex: str
:param weight: Vertex weight
:type weight: float, optional (default = None)
:return: Self
"""
if self.is_vertex_exists(vertex):
raise Exception(f"Vertex {vertex} already exists, delete it first")
self._graph[vertex] = Vertex(vertex, weight)
return self
def delete_vertex(self: T, vertex: str) -> T:
"""
Deletes a vertex
:param vertex: Vertex key
:type vertex: str
:return: Self
"""
for v in self.get_vertices():
vertex_obj = self.get_vertex(v)
vertex_obj.is_neighbour_exists(vertex) and vertex_obj.delete_neighbour(
vertex
)
self._graph.pop(vertex)
return self
def update_vertex(self: T, vertex: str, weight: float) -> T:
"""
Updates the vertex weight
:param vertex: Vertex key
:type vertex: str
:param weight: Vertex weight
:type weight: float
:return: Self
"""
self.get_vertex(vertex).weight = weight
return self
def get_vertices(self) -> List[str]:
"""
Gets the list of vertices in the graph
:return: List of vertices keys
"""
return list(self._graph.keys())
def get_isolated_vertices(self) -> List["Vertex"]:
"""
Gets isolated vertices in the graph
:return: List of vertices
"""
vertices = []
for vertex in self.get_vertices():
if not bool(self.get_vertex(vertex).neighbours):
vertices.append(self.get_vertex(vertex))
self.delete_vertex(vertex)
return vertices
def delete_isolated_vertices(self: T) -> T:
"""
Deletes isolated vertices in the graph
:return: Self
"""
for vertex in self.get_isolated_vertices():
self.delete_vertex(vertex.key)
return self
def is_edge_exists(self, source: str, destination: str) -> bool:
"""
Checks if the edge exists in the graph
:param source: Edge source
:type source: str
:param destination: Edge destination
:type destination: str
:return: Boolean
"""
return self.is_vertex_exists(source) and self.get_vertex(
source
).is_neighbour_exists(destination)
def get_edge(self, source: str, destination: str) -> "Edge":
"""
Gets edge from the graph
:param source: Edge source
:type source: str
:param destination: Edge destination
:type destination: str
:return: Edge
"""
return self.get_vertex(source).get_neighbour(destination)
def add_edge(self: T, source: str, destination: str, weight: float = None) -> T:
"""
Adds edge in the graph, default weight as :py:attr:`Edge.DEFAULT_EDGE_WEIGHT <reoptimization_algorithms.utils.graph.edge.Edge.DEFAULT_EDGE_WEIGHT>`
:param source: Edge source
:type source: str
:param destination: Edge destination
:type destination: str
:param weight: Weight
:type weight: float
:return: Self
"""
if not self.is_vertex_exists(source):
self.add_vertex(source)
if not self.is_vertex_exists(destination):
self.add_vertex(destination)
if self.is_edge_exists(source, destination):
raise Exception(
f"Edge stc: {source} dest: {destination} already exists, delete it first"
)
self.get_vertex(source).add_neighbour(destination, weight)
return self
def delete_edge(self: T, source: str, destination: str) -> T:
"""
Deletes the edge in the graph
:param source: Edge source
:type source: str
:param destination: Edge destination
:type destination: str
:return: Self
"""
self.get_vertex(source).delete_neighbour(destination)
return self
def update_edge(self: T, source: str, destination: str, weight: float) -> T:
"""
Updates the edge in the graph
:param source: Edge source
:type source: str
:param destination: Edge destination
:type destination: str
:param weight: Weight
:type weight: float
:return: Self
"""
self.get_vertex(source).update_neighbour(destination, weight)
return self
def get_edges(self) -> List[Dict]:
"""
Gets list of Edges
:return: List of dictionary having Edge source, destination and weight as keys
"""
edges = []
for vertex in self.get_vertices():
vertex_obj = self.get_vertex(vertex)
for destination in vertex_obj.neighbours:
edges.append(vars(vertex_obj.neighbours[destination]))
return edges
def copy(self: T) -> T:
"""
Return a deep copy of the graph
:return: Copied Graph
"""
return copy.deepcopy(self)
def disjoint_graph_union(self: T, attach_graph: T, attach_edges: List["Edge"]) -> T:
"""
Attaches the caller graph with attach graph and attachment edges, make sure the vertices are disjoint
:param attach_graph: Graph to attach
:type attach_graph: T
:param attach_edges: Edges to connect with attach_graph
:type attach_edges: List[Edge]
:raise InputError: If Vertices of calling graph and attach graph are not disjoint
:return: Self
"""
if not set(self.get_vertices()).isdisjoint(set(attach_graph.get_vertices())):
raise InputError(
{
"calling_graph_vertices": self.get_vertices(),
"attach_graph_edges": attach_graph.get_vertices(),
},
"Vertices of calling graph and attach graph must be disjoint",
)
graph = Graph(copy.deepcopy({**self.graph, **attach_graph.graph}))
for edge in attach_edges:
graph.add_edge(edge.source, edge.destination, edge.weight)
return graph | /reoptimization-algorithms-0.1.3.tar.gz/reoptimization-algorithms-0.1.3/src/reoptimization_algorithms/utils/graph/graph.py | 0.895514 | 0.469095 | graph.py | pypi |
from copy import deepcopy
from typing import Dict, List, Union, Any
from reoptimization_algorithms.utils.graph.graph import Graph
from reoptimization_algorithms.utils.graph.vertex import Vertex
class UndirectedGraph(Graph):
"""
Undirected Graph data structure class, inheriting Graph class, represented as dictionary with vertices
as key mapped to neighbouring vertices in a symmetric manner
:param graph: Undirected graph definition, if None then empty graph is instantiated
:type graph: Dict[str, Vertex]
Example
~~~~~~~
.. code-block:: python
import reoptimization_algorithms as ra
graph = ra.UndirectedGraph()
graph = graph.add_vertex("4", 14) # Adding vertex
graph.get_vertex("4") # Getting vertex
graph = graph.update_vertex("4", 15) # Update vertex weight
graph = graph.delete_vertex("4") # Deleting vertex
graph = graph.add_edge("4", "5", 11) # Adding Edge
graph.get_edge("4", "5") # Getting Edge
graph = graph.update_edge("4", "5", 10) # Update Edge weight
graph = graph.delete_edge("4", "5") # Deleting Edge
# Add, Update and Deletes can be chained as follows
graph = (ra.UndirectedGraph().add_vertex("4").add_edge("4", "5").add_edge("40", "50")
.add_vertex("6").add_edge("4", "8").delete_edge("4", "5").add_vertex("99")
.delete_vertex("6"))
"""
def __init__(self, graph: Dict[str, "Vertex"] = None):
"""
Undirected Graph data structure class, inheriting Graph class, represented as dictionary with vertices
as key mapped to neighbouring vertices in a symmetric manner
:param graph: Undirected graph definition, if None then empty graph is instantiated
:type graph: Dict[str, Vertex], optional (default = None)
"""
super().__init__(graph)
if graph is None:
graph = {}
self._graph = graph
def is_edge_exists(self, vertex_1: str, vertex_2: str) -> bool:
"""
Checks if edge exists in the graph
:param vertex_1: vertex 1 of the edge
:type vertex_1: str
:param vertex_2: vertex 2 of the edge
:type vertex_2: str
:return: Boolean
"""
return super().is_edge_exists(vertex_1, vertex_2) and super().is_edge_exists(
vertex_2, vertex_1
)
def update_edge(
self, vertex_1: str, vertex_2: str, weight: float
) -> "UndirectedGraph":
"""
Updates edge weight in the graph
:param vertex_1: vertex 1 of the edge
:type vertex_1: str
:param vertex_2: vertex 2 of the edge
:type vertex_2: str
:param weight: Weight to change with
:type weight: float
:return: Self
"""
if not self.is_edge_exists(vertex_1, vertex_2):
raise Exception(
f"Edge stc: {vertex_1} dest: {vertex_2} does not exists, create it first"
)
super().update_edge(vertex_1, vertex_2, weight)
super().update_edge(vertex_2, vertex_1, weight)
return self
def delete_edge(self, vertex_1: str, vertex_2: str) -> "UndirectedGraph":
"""
Deletes an edge in the graph
:param vertex_1: vertex 1 of the edge
:type vertex_1: str
:param vertex_2: vertex 2 of the edge
:type vertex_2: str
:return: Self
"""
if not self.is_edge_exists(vertex_1, vertex_2):
raise Exception(
f"Edge stc: {vertex_1} dest: {vertex_2} does not exists, create it first"
)
super().delete_edge(vertex_1, vertex_2)
super().delete_edge(vertex_2, vertex_1)
return self
def add_edge(
self, vertex_1: str, vertex_2: str, weight: float = None
) -> "UndirectedGraph":
"""
Adds an edge in the graph
:param vertex_1: vertex 1 of the edge
:type vertex_1: str
:param vertex_2: vertex 2 of the edge
:type vertex_2: str
:param weight: Weight of the edge
:type weight: float, optional (default = None)
:return: Self
"""
if self.is_edge_exists(vertex_1, vertex_2):
raise Exception(
f"Symmetric Edge ({vertex_1}, {vertex_2}) already exists, delete it first"
)
super().add_edge(vertex_1, vertex_2, weight)
super().add_edge(vertex_2, vertex_1, weight)
return self
def get_edges(self) -> List[Dict]:
"""
Gets edges from the graph
:return: List of dictionary having Edge source, destination and weight as keys
"""
edges = super().get_edges()
edge_key_set = set()
symmetric_edges = []
for edge in edges:
key = "::".join(sorted([edge["_source"], edge["_destination"]]))
if key not in edge_key_set:
edge_key_set.add(key)
symmetric_edges.append(edge)
return symmetric_edges
def graph_pretty(self) -> Dict[str, Union[List[Dict[str, Any]], List[dict]]]:
"""
Returns the graph in the pretty format
:return: Vertices as list of {'_Vertex__key': '4', '_weight': 1, '_neighbours': ..} and edges as list of
dictionary {_source, _destination and _weight}
"""
vertices = self.get_vertices()
formatted_vertices = []
for vertex_key in vertices:
formatted_vertex = vars(deepcopy(self.get_vertex(vertex_key)))
for key in formatted_vertex:
if isinstance(formatted_vertex[key], dict):
for k in formatted_vertex[key]:
formatted_vertex[key][k] = vars(formatted_vertex[key][k])
formatted_vertices.append(formatted_vertex)
edges = self.get_edges()
return {"vertices": formatted_vertices, "edges": edges} | /reoptimization-algorithms-0.1.3.tar.gz/reoptimization-algorithms-0.1.3/src/reoptimization_algorithms/utils/graph/undirected_graph.py | 0.953784 | 0.617426 | undirected_graph.py | pypi |
from itertools import combinations, permutations
from typing import Set, List, Tuple, Union, Iterable
from reoptimization_algorithms.utils.graph.undirected_graph import UndirectedGraph
class PVCUtils:
"""
Utility class for Path vertex cover problem
"""
@staticmethod
def is_k_pvc(graph: "UndirectedGraph", vertices: Set["str"], k: int) -> bool:
"""
Checks if the given candidate vertices are k path vertex cover for the graph
:param graph: Undirected graph
:type graph: UndirectedGraph
:param vertices: Vertices to check if they form k path cover
:type vertices: Set['str']
:param k: length of paths which should be covered
:type k: int
:return: Boolean
Example
~~~~~~~
.. code-block:: python
import reoptimization_algorithms as ra
graph = (ra.UndirectedGraph().add_edge("4", "5").add_edge("40", "50")
.add_vertex("6").add_edge("4", "8").add_vertex("99"))
print(ra.PVCUtils.is_k_pvc(graph, {"4"}, 3)) # True
"""
is_k_pvc = True
for k_path in list(combinations(graph.get_vertices(), k)):
k_path_set = set(k_path)
if len(
k_path_set.intersection(vertices)
) == 0 and PVCUtils.is_vertex_set_path(graph, k_path_set):
is_k_pvc = False
break
return is_k_pvc
@staticmethod
def is_vertex_set_path(graph: "UndirectedGraph", vertices: Iterable["str"]) -> bool:
"""
Checks if the vertices form a path of length :math:`k` in graph
:param graph: Undirected graph
:type graph: UndirectedGraph
:param vertices: Vertices to check if they form k path cover
:type vertices: Set['str']
:return: Boolean
Example
~~~~~~~
.. code-block:: python
from reoptimization_algorithms import PVCUtils, UndirectedGraph
graph = (UndirectedGraph().add_edge("4", "5").add_edge("40", "50")
.add_vertex("6").add_edge("4", "8").add_vertex("99"))
print(PVCUtils.is_vertex_set_path(graph, {"4", "5", "8"})) # True
"""
is_k_path = False
for path_perm in list(permutations(vertices)):
if PVCUtils.is_path(graph, path_perm):
is_k_path = True
break
return is_k_path
@staticmethod
def is_path(
graph: "UndirectedGraph", path: Union[List["str"], Tuple["str"]]
) -> bool:
"""
Checks if the path exists in the graph
:param graph: Undirected graph
:type graph: UndirectedGraph
:param path: Path as a list of vertices
:type path: Union[List['str'], Tuple['str']]
:return: Boolean
Example
~~~~~~~
.. code-block:: python
from reoptimization_algorithms import PVCUtils, UndirectedGraph
graph = (UndirectedGraph().add_edge("4", "5").add_edge("40", "50")
.add_vertex("6").add_edge("4", "8").add_vertex("99"))
print(PVCUtils.is_path(graph, ["4"])) # True
"""
is_k_path = True
for v in range(0, len(path) - 1):
if not graph.is_edge_exists(path[v], path[v + 1]):
is_k_path = False
break
return is_k_path | /reoptimization-algorithms-0.1.3.tar.gz/reoptimization-algorithms-0.1.3/src/reoptimization_algorithms/utils/graph/pvc.py | 0.926066 | 0.509215 | pvc.py | pypi |
import os
import sys
from typing import Sequence, List, Callable, Optional
import click
from .core import Editable, ReorderEditableError
def absdirs(positionals: Sequence[str]) -> List[str]:
"""
Convert all paths to abolsute paths, and make sure they all exist
"""
res = []
for pos in positionals:
absfile = os.path.abspath(os.path.expanduser(pos))
if not os.path.exists(absfile):
click.echo(f"{absfile} does not exist", err=True)
sys.exit(1)
res.append(absfile)
return res
@click.group()
def main() -> None:
"""
Manage your editable packages - your easy-install.pth file
"""
def _resolve_editable() -> str:
"""
Find the default easy-install.pth. Exits if a file couldn't be found
"""
editable_pth: Optional[str] = Editable.locate_editable()
if editable_pth is None:
raise ReorderEditableError("Could not locate easy-install.pth")
return editable_pth
def _print_editable_contents(
stderr: bool = False, chosen_editable: Optional[str] = None
) -> None:
"""
Opens the editable file directly and prints its contents
"""
editable_pth: str
if chosen_editable is not None:
editable_pth = chosen_editable
else:
editable_pth = _resolve_editable()
with open(editable_pth, "r") as src:
click.echo(src.read(), nl=False, err=stderr)
@main.command(short_help="print easy-install.pth contents")
def cat() -> None:
"""
Locate and print the contents of your easy-install.pth
"""
try:
_print_editable_contents()
except ReorderEditableError as err:
click.echo(str(err), err=True)
sys.exit(1)
@main.command(short_help="print easy-install.pth file location")
def locate() -> None:
"""
Try to find the easy-install.pth file, and print the location
"""
try:
click.echo(_resolve_editable())
except ReorderEditableError as err:
click.echo(str(err), err=True)
sys.exit(1)
# shared click options/args between check/reorder
SHARED = [
click.option(
"-e",
"--easy-install-location",
"editable_pth",
default=None,
help="Manually provide path to easy-install.pth",
),
click.argument("DIRECTORY", nargs=-1, required=True),
]
def shared(func: Callable[..., None]) -> Callable[..., None]:
"""
Decorator to apply shared arguments to reorder/check
"""
for decorator in SHARED:
func = decorator(func)
return func
@main.command(short_help="check easy-install.pth")
@shared
def check(editable_pth: Optional[str], directory: Sequence[str]) -> None:
"""
If the order specified in your easy-install.pth doesn't match
the order of the directories specified as positional arguments,
exit with a non-zero exit code
Also fails if one of the paths you provide doesn't exist
\b
e.g.
reorder_editable check ./path/to/repo /another/path/to/repo
In this case, ./path/to/repo should be above ./another/path/to/repo
in your easy-install.pth file
"""
dirs = absdirs(directory)
try:
Editable(location=editable_pth).assert_ordered(dirs)
except ReorderEditableError as exc:
click.echo("Error: " + str(exc))
_print_editable_contents(stderr=True, chosen_editable=editable_pth)
sys.exit(1)
@main.command(short_help="reorder easy-install.pth")
@shared
def reorder(editable_pth: Optional[str], directory: Sequence[str]) -> None:
"""
If the order specified in your easy-install.pth doesn't match
the order of the directories specified as positional arguments,
reorder them so that it does. This always places items
you're reordering at the end of your easy-install.pth so
make sure to include all items you care about the order of
Also fails if one of the paths you provide doesn't exist, or
it isn't already in you easy-install.pth
\b
e.g.
reorder_editable reorder ./path/to/repo /another/path/to/repo
If ./path/to/repo was below /another/path/to/repo, this would
reorder items in your config file to fix it so that ./path/to/repo
is above /another/path/to/repo
"""
dirs = absdirs(directory)
try:
Editable(location=editable_pth).reorder(dirs)
except ReorderEditableError as exc:
click.echo("Error: " + str(exc))
_print_editable_contents(stderr=True, chosen_editable=editable_pth)
sys.exit(1)
if __name__ == "__main__":
main(prog_name="reorder_editable") | /reorder_editable-0.1.0-py3-none-any.whl/reorder_editable/__main__.py | 0.485844 | 0.239727 | __main__.py | pypi |
import os
import itertools
import pandas, math, gzip, numpy, argparse, time
from keras import optimizers
from keras.layers import Dense, Dropout, Activation
from keras.models import Sequential, load_model
from keras.callbacks import ModelCheckpoint
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import normalize
from sklearn.utils import shuffle
from sklearn.metrics import classification_report, roc_curve, precision_recall_curve
import matplotlib.pyplot as plt
from scipy import stats
if __name__ == '__main__':
parser = argparse.ArgumentParser(description = 'Builds, test and uses models for the orientation of cDNA reads.')
parser.add_argument('-train', default = False, action = 'store_true',
help = 'Set true to train a model.')
parser.add_argument('-test', default = False, action = 'store_true',
help = 'Set true to test a model.')
parser.add_argument('-predict', default = False, action = 'store_true',
help = 'Set true to use a model to make predictions')
parser.add_argument('-data','--d', action = 'store', type = str, required = True, default = False,
help = 'The path to the input data. Must be either fasta or fastq. Can be compressed in gz.')
parser.add_argument('-source', '--s', action = 'store', type = str, required = True, choices = ['annotation','experimental','mapped', 'csv'],
help = 'The source of the data. Must be either \'experimental\', \' annotation\' or \'mapped\'. Choose experimental for experiments like RNA-direct, annotation for transcriptomes and mapped for mapped cDNA reads. Mapped reads require a paf file to know the orientation.')
parser.add_argument('-format', '--f', action = 'store', type = str, choices = ['fasta', 'fastq', 'auto'], default = 'auto',
help = 'The format of the input data. Auto by deafult. Change only if inconsistencies in the name.')
parser.add_argument('-annotation', '--a', action = 'store', type = str, default = False,
help = 'Path to the paf file if a mapped training set which requires a paf reference is being used.')
parser.add_argument('-use_all_annotation', '-aa', action = 'store_true', default = False,
help = 'Uses all the reads, instead of only keeping antisense,lincRNA,processed_transcript, protein_coding, and retained_intron. Use it also if the fasta has unconventional format and gives errors.')
parser.add_argument('-kmers', '--k', action = 'store', type = int, required = False, default = 5,
help = 'The maximum length of the kmers used for training, testing and using the models.')
parser.add_argument('-fixedkmer', '--fk', action = 'store_true', required = False, default = False,
help = 'Only use the last kmer length')
parser.add_argument('-reads', '--r', action = 'store', type = int, default = 10e10,
help = 'Number of reads to read from the dataset.')
parser.add_argument('-trimming', '--t', action = 'store', type = int, default = False,
help = 'Number of nucleotides to trimm at each side. 0 by default.')
parser.add_argument('-verbose', '--v', action = 'store_true', default = False,
help = 'Whether to print detailed information about the training process.')
parser.add_argument('-epochs', '--e', action = 'store', default = 20, type = int,
help = 'Number of epochs to train the model.')
parser.add_argument('-output', '--o', action = 'store', default = 'output',
help = 'Where to store the outputs. using "--train" outputs a model, while using "-predict" outputs a csv. Corresponding extensions will be added.')
parser.add_argument('-model', '--m', action = 'store',
help = 'The model to test or to predict with.')
parser.add_argument('-reverse_all', '--ra', action = 'store_true', default = False,
help = 'All the sequences will be reversed, instead of half of them')
parser.add_argument('-reads_to_model', '--rm', action = 'store', type = int, default = int(10e10),
help = 'Number of reads to use from the read ones')
parser.add_argument('-one_hot', '--oh', action = 'store_true', default = False,
help = 'Use one hot encoding instead of kmer counting')
options = parser.parse_args()
# Helper functions ------
def reverse_complement(dna):
"""Takes a RNA or DNA sequence string and returns the reverse complement"""
complement = {'A': 'T', 'C': 'G', 'G': 'C', 'T': 'A', 'U':'A', 'N':'N'}
return ''.join([complement[base] for base in dna[::-1]])
def sequences_to_kmers(seq, ks, only_last_kmer = False, full_counting = False, one_hot = False):
"""Converts a sequence to kmers counting. Returns a pandas Series object for easier processing.
- seq: a string containing only nucleotides.
- ks: maximum lenght of the k-mer counting.
- only_last_kmer: calculate only the biggest k-mer, but not the others.
- full_counting: ensures that all possible lectures windows are used to find the kmers. It makes the process
slower but more accurate.
- one_hot: if true, instead of counting kmers, it performs one-hot encoding.
"""
kmers = {}
length = len(seq)
if only_last_kmer:
starting = ks
else:
starting = 1
if full_counting:
windows = ks
else:
windows = 1
for k in range(starting, 1+ks):
for window in range(min(windows, k)):
for i in range(len(seq)//k):
subseq = seq[i*k+window: i*k+k+window]
if 'N' in subseq: # Ensures we discard ambigous nucleotide sequences.
continue
if len(subseq) < k:
continue
if subseq in kmers:
if one_hot:
kmers[subseq] = 1
else:
kmers[subseq] += 1/(length-k+1)
else:
if one_hot:
kmers[subseq] = 1
else:
kmers[subseq] = 1/(length-k+1)
return pandas.Series(kmers)
def generate_sets(data, labels, norm = False, do_not_split = False, no_test = False, mn_reads = int(10e10)):
"""
Generate sets for the training, validating and testing. The return depends on the parameters.
- data: train data. A matrix with columns being normalized counter kmers ordered alphabetically and rows as reads.
- do_not_split: if you want all the data in the same set, but shuffled.
- labels: an array of 0 and 1 for each row in data. 1 means reverse and 0 means forward.
- norm: if True normalizes the data. As the counting kmers are already normalized it's usually not necessary. If
the results are not good enought, set True to normalize across samples, which might help.
- no_test: True if the data provided is not going to be used as test, only as training and validation. Increases the model
performance.
"""
print('generating sets')
if norm:
data = normalize(data)
if do_not_split:
data = shuffle(data)
labels = labels.loc[data.index]
print('sets generated')
return data, labels
X_train, X_cvt, y_train, y_cvt = train_test_split(data[:int(mn_reads)], labels[:int(mn_reads)], train_size = 0.75, random_state = 0)
X_CV, X_test, y_CV, y_test = train_test_split(X_cvt, y_cvt, train_size = 0.50, random_state = 0)
print('sets generated')
if no_test:
return X_train, y_train, X_cvt, y_cvt
else:
return X_train, y_train, X_CV, y_CV, X_test,y_test
def prepare_data(sequences, order = 'forwarded', full_counting = True, ks = 5, drop_duplicates = False,
paf_path = False, ensure_all_kmers = False, only_last_kmer = False, reverse_all = False,
one_hot = False):
"""
Prepares a pandas Series containing nucleotide sequences into a pandas dataframe with kmers counting. Returns a pandas
data frame with the normalized kmer counts as columns and the reads as rows and a pandas Series with the labels (0 for
forward and 1 for reverse).
- drop_duplicates: drops sequences that are very similar at the end or beggining.
- order: can take different values and process data accordingly.
* order = forwarded: It can process any kind of format if all the sequences are in forward orientation. For example,
for RNA-direct or for the transcriptome.
* order = mixed: Doesn't assume everything is forward. Expects a paf file that must be provided by
paf_path argument to know the orientation.
* order = unknown: assumes the order is unknown. Used to predict.
- full_counting: full_counting: ensures that all possible lectures windows are used to find the kmers. It makes the process
slower but more accurate.
- ks: maximum lenght of the k-mer counting.
- ensure_all_kmers: if True, it makes sure all the mers are calculated. Use only for small files if some the prediction fails.
"""
print('Preparing the data')
if drop_duplicates:
sequences = sequences[~sequences.str[:30].duplicated()]
sequences = sequences[~sequences.str[-30:].duplicated()]
sequences = sequences[~sequences.str.contains('N')]
if order == 'forwarded':
print('Assuming the data provided is all in forward')
if reverse_all:
sequences_reverse = sequences.sample(sequences.shape[0])
sequences_reverse.index = sequences_reverse.index + sequences.shape[0]
else:
sequences_reverse = sequences.sample(sequences.shape[0]//2)
sequences = sequences.drop(sequences_reverse.index)
sequences_reverse = sequences_reverse.apply(reverse_complement)
sequences = sequences.apply(sequences_to_kmers, ks = ks, full_counting = full_counting, only_last_kmer=only_last_kmer, one_hot = one_hot)
sequences_reverse = sequences_reverse.apply(sequences_to_kmers, ks = ks, full_counting = full_counting, only_last_kmer=only_last_kmer, one_hot = one_hot)
sequences = pandas.DataFrame(sequences)
sequences_reverse = pandas.DataFrame(sequences_reverse)
sequences['s'] = 0
sequences_reverse['s'] = 1
sequences = pandas.concat([sequences, sequences_reverse])
sequences = sequences.sample(frac = 1)
labels = sequences['s']
data = sequences.drop('s', axis = 1)
data = data.fillna(0)
elif order == 'mixed':
print('Using a paf file to infer the orientation of reads')
ids = pandas.read_table(paf_path, usecols = [0,4], index_col = 0, header = None)
ids = ids[~ids.index.duplicated()]
ids.index.name = 'ID'
ids.columns = ['strand']
sequences = pandas.DataFrame(sequences, columns = ['seq'])
sequences['strand'] = ids['strand']
sequences['strand'] = sequences['strand'].replace(['+','-'], [0,1])
sequences = sequences.dropna()
labels = sequences['strand']
data = sequences.drop('strand', axis = 1)
data = sequences['seq'].apply(sequences_to_kmers, ks = ks, full_counting=full_counting, only_last_kmer=only_last_kmer, one_hot=one_hot)
data = data.fillna(0)
elif order == 'unknown':
labels = sequences
sequences = sequences.apply(sequences_to_kmers, ks = ks, full_counting = full_counting, only_last_kmer=only_last_kmer, one_hot=one_hot)
sequences = pandas.DataFrame(sequences)
data = sequences.fillna(0)
else:
raise NameError('Invalid source format')
if ensure_all_kmers:
bases=['A','T','G','C']
for k in range(ks):
kmers = [''.join(p) for p in itertools.product(bases, repeat=k+1)]
for kmer in kmers:
if kmer not in data.columns:
data[kmer] = 0
data = data.sort_index(axis = 1)
print('Data processed successfully')
return data, labels
# Reading functions ------
def read_experimental_data(path, format_file = 'auto' ,trimming = False, gzip_encoded = 'auto',
n_reads = 50000):
"""Takes a fasta or fastq file and reads it. The fasta can be compressed in gzip format.
- path: the fasta file path.
- trimming: allows to trimming while reading the file, so it's faster than doing it afterwards. False for no trimming.
Use an integer to trim the sequence both sides for that length.
- format: can be 'fasta', 'fastq' or 'auto' to autodetect it.
- gzip_encoded: if True it reads a gzip compressed fasta file. Use False if the fasta is in plain text. 'auto' tries to infer it from the filename.
- n_reads: number of reads to reads. Tune according to available memory.
"""
sequences = []
if gzip_encoded == 'auto':
if path[-2:] == 'gz':
gzip_encoded = True
else:
gzip_encoded = False
if gzip_encoded:
file = gzip.open(path, 'rb')
else:
file = open(path, 'r')
if format_file == 'auto':
if gzip_encoded:
marker = file.readline().decode()[0]
else:
marker = file.readline()[0]
if marker == '@':
format_file = 'fastq'
elif marker == '>':
format_file = 'fasta'
else:
raise NameError('Incorrect format')
if format_file == 'fastq':
n = 4
elif format_file == 'fasta':
n = 2
print('Detected file format: ', format_file)
i = -1
kept = 0
for line in file:
if gzip_encoded:
line = line.decode()
line = line.strip()
i += 1
if i%n == 0:
if kept >= n_reads:
break
if line.startswith('>'):
continue
else:
kept += 1
if trimming:
sequences.append(line.replace('U', 'T')[trimming:-trimming])
else:
sequences.append(line.replace('U', 'T'))
sequences = pandas.Series(sequences)
return sequences
def read_annotation_data(path, format_file = 'auto', n_reads = 50000, trimming = False, gzip_encoded = 'auto', use_all_annotation = False):
"""
This function reads data that doesn't come from an experiment but rather from the reference transcriptome.
- path: path to the transcriptome file in fasta format.
- n_reads: number of aproximate reads to process.
- trimming: allows to trimming while reading the file, so it's faster than doing it afterwards. False for no trimming.
Use an integer to trim the sequence both sides for that length.
- gzip_encoded: if True it reads a gzip compressed fasta file. Use False if the fasta is in plain text. 'auto' tries to infer it from the filename.
"""
sequences = []
if gzip_encoded == 'auto':
if path[-2:] == 'gz':
gzip_encoded = True
else:
gzip_encoded = False
if gzip_encoded:
file_c = gzip.open(path, 'rb')
file = gzip.open(path, 'rb')
else:
file_c = open(path, 'r')
file = open(path, 'r')
if format_file == 'auto':
if gzip_encoded:
line = file_c.readline()
line = line.decode()
marker = line[0]
else:
line = file_c.readline()
marker = line[0]
if marker == '@':
format_file = 'fastq'
elif marker == '>':
if len(line.split('|')) > 2:
separator = '|'
elif len(line.split(' ')) > 2:
separator = ' '
else:
print(line)
print('The file has not the correct format. All the sequence will be kept to avoid errors.')
use_all_annotation = True
format_file = 'fasta'
else:
raise NameError('Incorrect format')
print(separator)
if format_file == 'fastq':
n = 4
elif format_file == 'fasta':
n = 2
kept = 0
keep_next = False
for line in file:
print(line, keep_next)
if kept >= n_reads:
break
if gzip_encoded:
line = line.decode()
if line.startswith('>'):
line = line.strip()
if not use_all_annotation:
if separator == '|':
sline = line.split('|')
read_type = sline[-2]
elif separator == ' ':
sline = line.split(' ')
if ':' in sline[-1]:
read_type = sline[-1].split(':')[1]
else:
use_all_annotation = True
if use_all_annotation or read_type in ['antisense','lincRNA','processed_transcript', 'protein_coding', 'retained_intron']:
if keep_next:
kept += 1
if trimming:
sequences.append(sequence[trimming:-trimming])
else:
sequences.append(sequence)
keep_next = True
sequence = ''
else:
keep_next = False
else:
if keep_next:
sequence += line.strip()
if trimming:
sequences.append(sequence[trimming:-trimming])
else:
sequences.append(sequence)
sequences = pandas.Series(sequences)
return sequences
def read_mapped_data(path, n_reads = 50000, trimming = False, gzip_encoded = 'auto', format_file = 'auto'):
"""
Reads RNA that has been generated by mapping reads into a reference. We only consider antisense, lincRNA,
processed transcripts, protein coding transcripts and retained introns.
- path: path to the transcriptome file in fasta format.
- n_reads: number of aproximate reads to process.
- trimming: allows to trimming while reading the file, so it's faster than doing it afterwards. False for no trimming.
Use an integer to trim the sequence both sides for that length.
- gzip_encoded: if True it reads a gzip compressed fasta file. Use False if the fasta is in plain text. 'auto' tries to infer it from the filename.
"""
sequences = {}
if gzip_encoded == 'auto':
if path[-2:] == 'gz':
gzip_encoded = True
else:
gzip_encoded = False
if gzip_encoded:
file = gzip.open(path, 'rb')
else:
file = open(path, 'r')
file.readline()
file.readline()
i = 0
kept = 0
for line in file:
if gzip_encoded:
line = line.decode()
line = line.strip()
i += 1
if i%4 == 0:
if kept >= n_reads:
break
else:
kept += 1
if trimming:
sequences[indentifier] = line[trimming: -trimming]
else:
sequences[indentifier] = line
elif line.startswith('@'):
indentifier = line.split('\t')[0].split(' ')[0][1:]
file.close()
return pandas.Series(sequences).replace('U', 'T')
def read_cluster_data(path,trimming = False, n_reads = 50000):
"""
"""
sequences = []
ids = []
kept = 0
q = False
file = open(path)
for line in file:
line = line.strip()
if kept >= n_reads:
break
if line == '+':
q = True
continue
elif q==True:
q = False
continue
if line.startswith('@'):
ids.append(line[1:37])
else:
kept += 1
if trimming:
sequences.append(line.replace('U', 'T')[trimming:-trimming])
else:
sequences.append(line.replace('U', 'T'))
sequences = pandas.Series(sequences, index = ids)
return sequences
# Model functions ------
def plain_NN(input_shape, output_shape, n_layers = 5, n_nodes = 5, step_activation = 'relu',
final_activation = 'sigmoid',optimizer = False, kind_of_model = 'classification',
halve_each_layer = False,dropout = False, learning_rate = 0.0001):
"""
Creates a simple neural network model and returns the model object.
-input_shape = integer that represents the number of features used by the model.
-output_shape = integer that represents the number of features the model tries to predict.
-n_layers = the number of layers in the model.
-n_nodes = the number of nodes in each layer.
-step_activation = activation function at each step, can be any that keras uses.
-final_activation = activation function at the final step, can be any that keras uses.
-optimizers = if provided, it uses the optimizers delivered.
-halve each layer = if true, each layer has half the nodes as the previous one.
-dropout = use drouput layers.
-learning_rate = the learning rate for the model to learn.
"""
print('Creating model architecture')
model = Sequential()
model.add(Dense(n_nodes, activation='relu', input_dim=input_shape))
if halve_each_layer:
halver = 2
else:
halver = 1
if dropout:
model.add(Dropout(0.3))
for i in range(n_layers-1):
n_nodes = n_nodes // halver
model.add(Dense(n_nodes,activation = step_activation))
if dropout:
model.add(Dropout(0.3))
model.add(Dense(output_shape, activation = final_activation))
if optimizer:
optimizer = optimizer
else:
optimizer = optimizers.RMSprop(lr = learning_rate)
if kind_of_model == 'classification':
model.compile(optimizer = optimizer, loss = 'binary_crossentropy', metrics = ['accuracy'])
if kind_of_model == 'regression':
model.compile(optimizer = optimizer, loss = 'mse', metrics = ['mae'])
print(model.summary())
return model
def fit_network(model, data, labels, epochs = 10, batch_size = 32, verbose = 1 ,checkpointer = False, no_test = True, mn_reads = int(10e10)):
"""
Fits a neural network into a model and returns the history to easily analyze the performance.
Returns the trained model and the training history, for evaluation purposes.
checkpointer: if given a name, creates a checkpointer with that name.
- model: model to train the data with. Must have the same input shape as number of variables in the train set.
- data: train data. A matrix with columns being normalized counter kmers ordered alphabetically and rows as reads.
- labels: an array of 0 and 1 for each row in data. 1 means reverse and 0 means forward.
- epochs: number of iterations to train the model. Recomended from 10 to 100. The more data the less epochs are necessary.
- verbose: whether to print several information related to the training process.
- batch_size: number of reads to train the model at once during each epochs.
- checkpointer: if given a name, creates a checkpointer with that name.
- no_test: True if the data provided is not going to be used as test, only as training and validation. Increases the model
performance.
"""
if no_test:
X_train, y_train, X_CV, y_CV = generate_sets(data, labels, no_test = no_test, mn_reads = mn_reads)
else:
X_train, y_train, X_CV, y_CV, X_test,y_test = generate_sets(data, labels, mn_reads = mn_reads)
if checkpointer:
print('Using Checkpointer')
model_file = checkpointer+'.model'
checkpointer = ModelCheckpoint(filepath= model_file,
verbose=verbose, save_best_only=True)
train_time0 = time.time()
history = model.fit(X_train.values, y_train.values, batch_size=batch_size,
epochs=epochs,validation_data=(X_CV.values, y_CV.values), verbose=verbose,
callbacks = [checkpointer])
train_time1 = time.time()
print('Elapsed time while training: ', train_time1 - train_time0, ' seconds')
model.load_weights(model_file)
model.save(model_file)
print('Best model train accuracy: ', model.evaluate(X_train.values, y_train.values))
print('Best model validation accuracy: ', model.evaluate(X_CV.values, y_CV.values))
else:
history = model.fit(X_train.values, y_train.values, batch_size=batch_size,
epochs=epochs,validation_data=(X_CV.values, y_CV.values), verbose=verbose)
if not no_test:
print(model.evaluate(X_test.values, y_test.values))
return model, history
def build_kmer_model(kind_of_data, path_data, n_reads, path_paf, trimming, full_counting, ks, verbose = 1,
epochs = 10, checkpointer = 'cDNAOrderPrediction', use_all_annotation = False, only_last_kmer = False, reverse_all = False,
mn_reads = int(10e10), one_hot = False):
"""
Function that automatically reads and processes the data and builds a model with it. Returns the trained model
and the generated dataset and labelset.
- kind_of_data: the kind of data used to train the model. Can be:
* 'experimental' if it comes from RNA direct or similars.
* 'annotation' if it is the transcriptome reference.
* 'mapped' if its a mapped cDNA dataset. It requires a paf file to be provided.
- path_data: path to the data that is going to train the model.
- n_reads: number of approximate reads to process from the train data.
- path_paf: path to the paf file if we are using mapped data.
- trimming: allows to trimming while reading the file, so it's faster than doing it afterwards. False for no trimming.
Use an integer to trim the sequence both sides for that length.
- ks: maximum lenght of the k-mer counting.
- full_counting: ensures that all possible lectures windows are used to find the kmers. It makes the process
slower but more accurate.
- verbose: can be 0 or 1. 1 means ploting several information related to the training process.
- epochs: the number of training iterations.
- checkpointer: if False, the best model is not saved into a file for easy retrieve. If given a name, it saves the model into
a file with that name.
"""
if kind_of_data == 'experimental':
sequences = read_experimental_data(path = path_data, trimming = trimming, n_reads = n_reads)
elif kind_of_data == 'annotation':
sequences = read_annotation_data(path = path_data, trimming = trimming, n_reads = n_reads, use_all_annotation = use_all_annotation)
elif kind_of_data == 'mapped':
sequences = read_mapped_data(path = path_data, trimming = trimming, n_reads = n_reads)
if path_paf:
order = 'mixed'
else:
order = 'forwarded'
data, labels = prepare_data(sequences, order, full_counting, ks, False, path_paf, only_last_kmer=only_last_kmer, reverse_all = reverse_all, one_hot = one_hot)
model = plain_NN(data.shape[1],1, 5, 500, step_activation = 'relu', final_activation = 'sigmoid',
optimizer = False, kind_of_model = 'classification', halve_each_layer = True,dropout = True,
learning_rate = 0.00001)
model, history = fit_network(model, data, labels, epochs = epochs, verbose = verbose, checkpointer = checkpointer, batch_size = 64, mn_reads = mn_reads)
return model, history ,data, labels
def test_model(model, kind_of_data, path_data, n_reads, path_paf, trimming, full_counting, ks, one_hot, return_predictions = False, mn_reads = int(10e10)):
"""
Function that automatically reads and processes the data and test a model with it. Prints several
metrics about the model performance. !!Use the same parameters as used to train the model!!.
- model: trained model.
- kind_of_data: the kind of data used to train the model. Can be:
* 'experimental' if it comes from RNA direct or similars.
* 'annotation' if it is the transcriptome reference.
* 'mapped' if its a mapped cDNA dataset. It requires a paf file to be provided.
- path_data: path to the data that is going to train the model.
- n_reads: number of approximate reads to process from the train data.
- path_paf: path to the paf file if we are using mapped data.
- trimming: allows to trimming while reading the file, so it's faster than doing it afterwards. False for no trimming.
Use an integer to trim the sequence both sides for that length.
- ks: maximum lenght of the k-mer counting.
- full_counting: ensures that all possible lectures windows are used to find the kmers. It makes the process
slower but more accurate.
- return_predictions: if True, the predictions and labels are returned with the metrics.
"""
if kind_of_data == 'experimental':
sequences = read_experimental_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = options.f)
elif kind_of_data == 'annotation':
sequences = read_annotation_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = options.f)
elif kind_of_data == 'mapped':
sequences = read_mapped_data(path = path_data, trimming = trimming, n_reads = n_reads)
if path_paf:
order = 'mixed'
else:
order = 'forwarded'
data, labels = prepare_data(sequences, order, full_counting, ks, False, path_paf, one_hot=one_hot)
data, labels = (data[:int(mn_reads)], labels[:int(mn_reads)])
predictions = model.predict(data.values)
print('----------------------Test Results-----------------------\n')
print(classification_report(labels,predictions.round()))
print('---------------------------------------------------------\n')
if return_predictions:
return predictions, labels
def make_predictions(model, kind_of_data, path_data, n_reads, path_paf, trimming, full_counting, ks, one_hot):
if kind_of_data == 'experimental':
sequences = read_experimental_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = options.f)
elif kind_of_data == 'annotation':
sequences = read_annotation_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = options.f)
elif kind_of_data == 'mapped':
sequences = read_mapped_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = options.f)
elif kind_of_data == 'csv':
sequences = pandas.read_csv(path_data, index_col = 0, squeeze = True, nrows = n_reads)
data, labels = prepare_data(sequences, 'unknown', full_counting, ks, False, path_paf, one_hot = one_hot, ensure_all_kmers = True)
predictions = model.predict(data.values)
data = pandas.DataFrame(labels)
data['predictions'] = predictions
data['orientation'] = 0
data.loc[data['predictions'] > 0.5, 'orientation'] = 1
data.loc[data['predictions'] > 0.5, 0] = data[0].apply(reverse_complement)
data.loc[data['predictions'] < 0.5, 'predictions'] = 1 - data['predictions']
data.columns = ['ForwardSequence', 'Score', 'Orientation']
data.to_csv(options.o+'.csv')
# Plot functions ------
def plot_roc_and_precision_recall_curves(models, kind_of_data, path_data, n_reads, path_paf, trimming,
full_counting, ks, format_file, species):
"""
Plots the ROC and the precision-recall-curve for a set of models and a specific data input.
- models: a list of trained model.
- kind_of_data: the kind of data used to train the model. Can be:
* 'experimental' if it comes from RNA direct or similars.
* 'annotation' if it is the transcriptome reference.
* 'mapped' if its a mapped cDNA dataset. It requires a paf file to be provided.
- path_data: path to the data that is going to train the model.
- n_reads: number of approximate reads to process from the train data.
- path_paf: path to the paf file if we are using mapped data.
- trimming: allows to trimming while reading the file, so it's faster than doing it afterwards. False for no trimming.
Use an integer to trim the sequence both sides for that length.
- ks: maximum lenght of the k-mer counting.
- full_counting: ensures that all possible lectures windows are used to find the kmers. It makes the process
slower but more accurate.
- format_file: The format of the input data: can be auto, fasta or fastq. 'auto' should be used by default.
- species: the name of the species to be plotted in the plots.
"""
if kind_of_data == 'experimental':
sequences = read_experimental_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = format_file)
elif kind_of_data == 'annotation':
sequences = read_annotation_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = format_file)
elif kind_of_data == 'mapped':
sequences = read_mapped_data(path = path_data, trimming = trimming, n_reads = n_reads, format_file = format_file)
if path_paf:
data, labels = prepare_data(sequences, 'mixed', full_counting, ks, False, path_paf, True)
else:
data, labels = prepare_data(sequences, 'forwarded', full_counting, ks, False, path_paf, True)
for model_name in models:
model = load_model(model_name)
prediction = model.predict(data.values)
fpr_grd, tpr_grd, _ = roc_curve(labels, prediction)
plt.plot(fpr_grd, tpr_grd, label = model_name.split('/')[-1])
plt.xlabel('False positive rate', fontsize = 17)
plt.ylabel('True positive rate', fontsize = 17)
plt.tick_params(axis='both', which='major', labelsize=14)
plt.title(species + ' cDNA orientation prediction', fontsize = 18)
plt.legend(fontsize = 15)
plt.tight_layout()
plt.savefig('plots/ROC_' + species + ' cDNA orientation prediction.png', dpi = 200)
plt.close('all')
for model_name in models:
model = load_model(model_name)
prediction = model.predict(data.values)
precision, recall, _ = precision_recall_curve(labels, prediction)
plt.plot(recall[:-2], precision[:-2], label = model_name.split('/')[-1])
plt.xlabel('precision', fontsize = 17)
plt.ylabel('Recall', fontsize = 17)
plt.tick_params(axis='both', which='major', labelsize=14)
plt.title(species + ' cDNA orientation prediction', fontsize = 15)
plt.legend(fontsize = 15)
plt.tight_layout()
plt.savefig('plots/PRC_'+species + ' cDNA orientation prediction.png', dpi = 200)
plt.close('all')
return precision, recall, _
def analyze_clusters(path, model=False, csv=False):
if model:
model = load_model(model)
else:
prediction_csv = pandas.read_csv(csv, index_col = 0, usecols=['id', 'strand', 'predictions'])
results = []
lengths = []
for file in os.listdir(path):
if not file.startswith('.') and file.endswith('.fq'):
full_path = path+'/'+file
sequences = read_experimental_data(full_path, format_file = 'auto' ,trimming = False, gzip_encoded = 'auto', n_reads = int(10e10))
data, labels = prepare_data(sequences, 'unknown', True, 5, False, False,True)
try:
predictons = model.predict(data.values).round()
except:
data, labels = prepare_data(sequences, 'unknown', True, 5, False, False,True)
predictons = model.predict(data.values).round()
agreement = stats.mode(predictons)[1][0][0]/len(predictons)
results.append(agreement)
lengths.append(len(predictons))
return results, lengths
def analyze_clustersv2(path_clusters = '/Users/angelruiz/Desktop/clusters/clusters/hs/final_clusters.csv',
path_prediction = '/Users/angelruiz/Downloads/predictions_cDNA_human_Hs_transcriptome_cnn.csv',
paf_file = '/Users/angelruiz/Desktop/RE_data/cdna_human_no_secondary_mapq_60_unique.paf',
species = 'hs', prediction_id_col = 0, prediction_prediction_col = 4):
clusters = pandas.read_table(path_clusters, names = ['cluster', 'id'], index_col=1)
clusters.index = clusters.index.str[0:36]
clusters.index = clusters.index.str[0:36] # change this to capture the identifier
#clusters.index = clusters.index.str.split('_').str[0]
predictions = pandas.read_csv(path_prediction, usecols=[prediction_id_col, prediction_prediction_col], names = ['id', 'prediction'], index_col=[0], skiprows=1)
predictions = predictions.prediction.round().replace([1.0, 0], ['-', '+'])
labels = pandas.read_table(paf_file, usecols = [0,4], index_col = 0, header = None, names = ['id', 'strand'])
data = clusters.join(predictions).join(labels)
data['ncorrect'] = data['prediction']==data['strand']
cluster_analysis = pandas.DataFrame(data.groupby('cluster')['ncorrect'].sum())
cluster_analysis['size'] = data.groupby('cluster').size()
cluster_analysis['per_correct'] = cluster_analysis['ncorrect']/cluster_analysis['size']
cluster_analysis['correct_cluster'] = cluster_analysis['ncorrect']>=cluster_analysis['size']//2
cluster_analysis[cluster_analysis['size'] > 2]['per_correct'].hist()
plt.title('Accuracy per cluster (Size > 2)')
plt.savefig(species+'_acc_clusters_gt_2.png')
plt.close('all')
cluster_analysis[cluster_analysis['size'] > 1]['per_correct'].hist()
plt.title('Accuracy per cluster (Size > 1)')
plt.savefig(species+'_acc_clusters_gt_1.png')
plt.close('all')
RE_lazy = labels[labels['strand'] == '+'].size/labels.size
RE_correct = sum(cluster_analysis['ncorrect'])/sum(cluster_analysis['size'])
RE_correct_voting = sum(cluster_analysis['size'] * cluster_analysis.correct_cluster)/sum(cluster_analysis['size'])
print('Lazy model accuracy: ', RE_lazy)
print('RE accuracy:', RE_correct)
print('RE plus clustering accuracy:', RE_correct_voting)
return(0)
if __name__ == '__main__':
time0 = time.time()
if options.train:
print('\n----Starting Training Pipeline----\n')
model, history ,data, labels = build_kmer_model(options.s, options.d, options.r, options.a, options.t,
True, options.k, options.v, options.e ,options.o, options.use_all_annotation, options.fk,options.ra, options.rm,
options.oh)
elif options.test:
print('\n----Starting Testing Pipeline----\n')
model = load_model(options.m)
test_model(model, options.s, options.d, options.r, options.a, options.t, True, options.k, options.oh, False,options.rm)
elif options.predict:
print('\n----Starting Prediction Pipeline----\n')
model = load_model(options.m)
print('Model successfully loaded')
print(model.summary())
make_predictions(model, options.s, options.d, options.r, options.a, options.t, True, options.k, options.oh)
print('Predictions saved to:', options.o+'.csv')
time1 = time.time()
delta = time1 - time0
print('Elapsed time\t', delta, ' seconds') | /reorientexpress-0.400.tar.gz/reorientexpress-0.400/reorientexpress.py | 0.623721 | 0.222172 | reorientexpress.py | pypi |
# ReorientExpress
ReorientExpress is a program to create, test and apply models to predict the 5'-to-3' orientation of long-reads from cDNA sequencing with Nanopore or PacBio using deep neural networks for samples without a genome or a transcriptome reference.
----------------------------
# Table of Contents
----------------------------
* [Overview](#overview)
* [Installation](#installation)
* [Commands and options](#commands-and-options)
* [Inputs and Outputs](#inputs-and-outputs)
* [Usage example](#usage-example)
----------------------------
# Overview
----------------------------
ReorientExpress is a tool to predict the orientation of cDNA reads from error-prone long-read sequencing technologies. It was developed with the aim to orientate nanopore long-reads from unstranded cDNA libraries without the need of a genome or transcriptome reference, but it is applicable to any set of long-reads. ReorientExpress implements two Deep Neural Network models: a Multi-Layer Perceptron (MLP) and a Convolutional Neural Network (CNN), and it uses as training input a transcriptome annotation from any species or any other fasta/fasq file of RNA/cDNA sequences for which the orientation is known.
Training or testing data can thus be experimental data, annotation data or also mapped reads (providing the corresponding PAF file).
ReorientExpress has three main modes:
- Training a model.
- Testing a model.
- Using a model to orientate input sequences.
These are implemented in three options: train, test and predict. In train mode, the input data is randomly split into three subsets: training, validation and test, with relative proportions of 0.75, 0.125 and 0.125, respectively. The training set is used to train the weights of the DNN model, the validation set is used to optimize the weights during the training process, and the test set has never been seen for training and is only used at the end to evaluate the accuracy of the model.
----------------------------
# Installation
----------------------------
ReorientExpress has been developed in Python 3.6. It can be directly cloned and used or installed for an easier reuse and dependency management.
Currently, you can use pip to do an authomatic installation:
```
pip3 install reorientexpress
```
If some dependencies are not correctly downloaded and installed, using the following can fix it:
```
pip3 install -r requirements.txt
pip3 install reorientexpress
```
Once the package is installed, ReorientExpress can be used from the command line as any other program.
----------------------------
# Commands and options
----------------------------
Once the package is installed it can be used as an independent program. ReorientExpress has three main functions, one of them must be provided when calling the program:
* -train: takes an input an uses it to train a model.
* -test: takes a model and a labeled input and estimate the accuracy of the model using the labeled input data.
* -predict: takes a model and an input and outputs all the sequences in the predicted 5'-to3' orientation. It also gives a certainty score per input sequence.
The different options available for MLP (reorientexpress.py) are:
* **-h, --help**: Shows a help message with all the options.
* **-train**: Set true to train a model.
* **-test**: Set true to test a model.
* **-predict**: Set true to use a model to make predictions
* **-data D, --d D**: The path to the input data. Must be either fasta or
fastq. Can be compressed in gz format. Mandatory.
* **-source {annotation,experimental,mapped}, --s {annotation,experimental,mapped}**:
The source of the data. Must be either 'experimental',
'annotation' or 'mapped'. Choose experimental for
experiments like RNA-direct, annotation for
transcriptomes or other references and mapped for reads mapped
to a reference transcriptome.
Mapped reads must be in PAF format to extract the orientation.
Mandatory.
* **-format {fasta,fastq,auto}, --f {fasta,fastq,auto}**:
The format of the input data. Auto by deafult. Change
only if inconsistencies in the name.
* **-annotation A, --a A**: Path to the PAF file if a mapped training set is used.
* **-use_all_annotation, -aa**:
Uses all the reads from the annotation, instead of only keeping
protein_coding, lincRNA, processed_transcript, antisense, and retained_intron.
Use it also if the fasta has unconventional format and gives errors.
* **-kmers K, --k K**: The maximum length of the kmers used for training,
testing and using the models. It will use from k=1 up to this number.
* **-reads R, --r R**: Number of reads to use from the dataset.
* **-trimming T, --t T**: Number of nucleotides to trimm at each side. 0 by default.
* **-reverse_all**: Reverse-complement all input sequences to double up the training input,
instead of reverse-complementing just a random half of the input sequences,
which is the default.
* **-verbose, --v**: Flag to print detailed information about the
training process.
* **-epochs E, --e E**: Number of epochs to train the model.
* **-output O, --o O**: Where to store the outputs. using "--train" outputs a
model, while using "-predict" outputs a csv.
Corresponding extensions will be added.
* **-model M, --m M**: The model to test or to predict with.
The different option available for CNN (reoreintexpress-cnn.py) are:
* **-h, --help**: Shows a help message with all the options.
* **-train**: Set true to train a model.
* **-test**: Set true to test a model.
* **-predict**: Set true to use a model to make predictions
* **-data D, --d D**: The path to the input data. Must be either fasta or
fastq. Can be compressed in gz format. Mandatory.
* **-source {annotation,experimental,mapped}, --s {annotation,experimental,mapped}**:
The source of the data. Must be either 'experimental',
'annotation' or 'mapped'. Choose experimental for
experiments like RNA-direct, annotation for
transcriptomes or other references and mapped for reads mapped
to a reference transcriptome.
Mapped reads must be in PAF format to extract the orientation.
Mandatory.
* **-format {fasta,fastq,auto}, --f {fasta,fastq,auto}**:
The format of the input data. Auto by deafult. Change
only if inconsistencies in the name.
* **-annotation A, --a A**: Path to the PAF file if a mapped training set is used.
* **-use_all_annotation, -aa**:
Uses all the reads from the annotation, instead of only keeping
protein_coding, lincRNA, processed_transcript, antisense, and retained_intron.
Use it also if the fasta has unconventional format and gives errors.
* **-win_size W, --w W**: Window size for spliting the sequence.
* **-step_size, --step**: Overlapping size on the the sliding window.
* **-reads R, --r R**: Number of reads to use from the dataset.
* **-trimming T, --t T**: Number of nucleotides to trimm at each side. 0 by default.
* **-reverse_all**: Reverse-complement all input sequences to double up the training input,
instead of reverse-complementing just a random half of the input sequences,
which is the default.
* **-verbose, --v**: Flag to print detailed information about the
training process.
* **-epochs E, --e E**: Number of epochs to train the model.
* **-output O, --o O**: Where to store the outputs. using "--train" outputs a
model, while using "-predict" outputs a csv.
Corresponding extensions will be added.
* **-model M, --m M**: The model to test or to predict with.
----------------------------
# Inputs and Outputs
----------------------------
All the input sequence files can be in fasta or fastq format. They can also be compressed in gz format.
Input sequences can be of three different types, which we call experimental, annotation or mapped, which can be in FASTA or FASTQ formats, either compressed (in .gz format) or uncompressed.
* Experimental data refers to any kind of long-read data for which the orientation is known, such as direct RNA-seq, and reads are considered to be given in the 5’-to-3’ orientation.
* Annotation data refers to the transcript sequences from a reference annotation, such as the human transcriptome reference. It also considers all the sequences to be in the right 5’-to-3’ orientation. Annotation data can also include the transcript type, such as protein coding, processed transcript, etc.
* Mapped data refers to sequencing data, usually cDNA, whose orientation has been annotated by an independent method, e.g. by mapping the reads to a reference. In this case, a PAF file for the mapping, together with the FASTA/FASTQ file, is required.
### Examples of possible inputs:
#### Experimental
<pre>@0e403438-313b-4497-b1c2-2fd3cc685c1d runid=46930771ed1cff73b50bf5c153000aa904eb5c9c read=100 ch=493 sta
rt_time=2017-10-09T18:11:16Z
CCCGGAAAAUGGUGAAGAAAAUUGAAAUCAGCCAGCACGUCCGUUAAGUCACUUGCUUUACCGCGGCAAACCAAGAUGAAGACGAGCUGUGGGAUCUGGCACUA
CUGUGGUUCCAUUGCAUGAACGGGAAGACAGUGGCUGGCGGGUGCCCUGGACGUACAAAUACCACUCCAAUUGUCACGGUAAAGUCCGCCAUCAGAAGACUGAA
GGAGUUGUAGACCAGUAGACGUUCCAUACACAUUGAGACACUACUGGCCUAUAAUAAUUAAAUGGGUUAUUAAUUUAUUUAUGGCUAACAAAUUGUUCCGAGCU
CGUAUUAAACAGAUAUCGAUGUUGUAUUGUUGUAGUAGUAUUGAAGAGCAAAUCCCACCCAUCCUUCCAUCAACAACCUCCCGUUAUUAUACCGUUAUCCCACC
GCCUACCAUCUUCCCAUAAAAUCCAUC
+
$)/*+7B:314:3.,/.6C;4.*'+69-.14:221'%&#"+)'$$%*)'$%&&)*''(+"$&$%)1*.:/0:7522222/--**--*++*/9>/0-&*('%%%)
,+&031=12+(**)#$#$$'&%((-.-4524,,4*+-:.-./(('@7-)5$'%)))3.,)**-),--/*(/0)(%+1.7*+6)+*7:32&'&*,,(/(('.-1/
3.+../)$-/29:66,*-,&.+.8,(#'&&&')1-//.--((%)(111+''&11,2(%&*./,)5..*'*%.0011%$%%#%'-&(-5+,@6>9;'-)5)**%$
#+*,,,15.''%(*)++,,4,---/064'))()($%#%''*-%&'$'##$$)&'+.%+4,(%'*&$/(&''(0(%/',$,.(&)'#,-$$$'-"$$$$&.+%($
"*+$$$$$%$$#0:*'&%&'+#$&$$"</pre>
#### Annotation
<pre>
>ENSMUST00000193812.1|ENSMUSG00000102693.1|OTTMUSG00000049935.1|OTTMUST00000127109.1|4933401J01Rik-201|4933401J01Rik|1070|TEC|
AAGGAAAGAGGATAACACTTGAAATGTAAATAAAGAAAATACCTAATAAAAATAAATAAA
AACATGCTTTCAAAGGAAATAAAAAGTTGGATTCAAAAATTTAACTTTTGCTCATTTGGT
ATAATCAAGGAAAAGACCTTTGCATATAAAATATATTTTGAATAAAATTCAGTGGAAGAA
TGGAATAGAAATATAAGTTTAATGCTAAGTATAAGTACCAGTAAAAGAATAATAAAAAGA
AATATAAGTTGGGTATACAGTTATTTGCCAGCACAAAGCCTTGGGTATGGTTCTTAGCAC
TAAGGAACCAGCCAAATCACCAACAAACAGAGGCATAAGGTTTTAGTGTTTACTATTTGT
ACTTTTGTGGATCATCTTGCCAGCCTGTAGTGCAACCATCTCTAATCCACCACCATGAAG
GGAACTGTGATAATTCACTGGGCTTTTTCTGTGCAAGATGAAAAAAAGCCAGGTGAGGCT
GATTTATGAGTAAGGGATGTGCATTCCTAACTCAAAAATCTGAAATTTGAAATGCCGCCC
</pre>
#### Mapped
Takes a file with the same format as experimental and also a PAF file with the following format:
<pre>
0M1I3M2D4M3D1M1D10M4I11M1D25M1D6M1D10M1D10M
0e04dd74-26bd-47e3-91bf-0e6e97310067 795 2 410 - ENST00000584828.5|ENSG0000018406
0.10|OTTHUMG00000132868.4|OTTHUMT00000444515.1|ADAP2-209|ADAP2|907|protein_coding| 907 398
798 344 432 1 NM:i:88 ms:i:336 AS:i:336 nn:i:0 tp:A:P cm:i:7 s1:i:82
s2:i:67 dv:f:0.1443 cg:Z:4M1I19M2D15M1I8M4I1M1D6M1I29M1D1M2D13M1D5M2I4M1D21M3I28M2I11M3I8M1I13M2I16M
3D12M1I2M3D5M2I16M2I14M4D12M1I9M4I47M2D1M3D24M2I7M1D25M
0e04dd74-26bd-47e3-91bf-0e6e97310067 795 2 405 - ENST00000585130.5|ENSG0000018406
0.10|OTTHUMG00000132868.4|OTTHUMT00000444510.1|ADAP2-211|ADAP2|2271|nonsense_mediated_decay| 2271
1366 1762 340 426 0 NM:i:86 ms:i:334 AS:i:334 nn:i:0 tp:A:S cm:i:6
s1:i:67 dv:f:0.1427 cg:Z:19M2D15M1I7M1I3M2I6M1I29M1D1M2D13M1D5M2I4M1D21M3I28M2I11M3I8M1I13M2I16M3D12
M1I2M3D5M2I16M2I14M4D12M1I9M4I47M2D1M3D24M2I7M1D25M
0e04dd74-26bd-47e3-91bf-0e6e97310067 795 2 405 - ENST00000330889.7|ENSG0000018406
0.10|OTTHUMG00000132868.4|OTTHUMT00000256346.1|ADAP2-201|ADAP2|2934|protein_coding| 2934 1446
1842 340 426 0 NM:i:86 ms:i:334 AS:i:334 nn:i:0 tp:A:S cm:i:6 s1:i:67
</pre>
You can read more about the paf file format [here](https://github.com/lh3/miniasm/blob/master/PAF.md).
### Examples of possible outputs:
Depending on the chosen pipeline, the output can be:
* Training: a keras model object, from the class keras.engine.sequential.Sequential (https://keras.io). It is saved as a binary file that be loaded later.
* Testing: there is no file output. Only the results of the accuracy evaluation displayed on the terminal.
* Predicting: outputs a csv file with all the reads in the predicted 5'-to-3' orientation. It contains three columns: the predicted 5'-to-3' sequence (ForwardedSequence) and the model Score and the read orientation. See below an example:
| Index | ForwardSequence | Score | orientation |
|---|---|---|---|
| 0 | ATGTTGAATAGTTCAAGAAAATATGCTTGTCGTTCCCTATTCAGACAAGCGAACGTCTCA | 0.8915960788726807 | 0 |
| 1 | TTGAGGAGTGATAACAAGGAAAGCCCAAGTGCAAGACAACCACTAGATAGGCTACAACTA | 0.9746999740600586 | 1 |
| 2 | AAGGCCACCATTGCTCTATTGTTGCTAAGTGGTGGGACGTATGCCTATTTATCAAGAAAA | 0.9779879450798035 | 0 |
*Note*: '0' orientation represents '+' and '1' orientation represents '-'. However, the '-' reads are resevse complemented and provided in the 'ForwardSequence' column.
----------------------------
# Usage example
----------------------------
**Note:** The below commands are for MLP model. Similar commands can be used for CNN model will the replacement of *reorientexpress.py* with *reoreintexpress-cnn.py*
To train a model:
```
reorientexpress.py -train -data path_to_data -source annotation --v -output my_model
```
This trains a model with the data stored in path_to_data, which is an annotation file, suchs as a transcriptome and outputs a file called my_model.model which can be later used to make predictions. Prints relevant information.
Example on test_case provided in the repo:
```
reorientexpress.py -train -data ./test_case/annotation/gencode.vM19.transcripts_50k.fa -source annotation --v -output my_model
```
or
```
reorientexpress-cnn.py -train -data ./test_case/annotation/gencode.vM19.transcripts_50k.fa -source annotation --v -output my_model
```
To make predictions:
```
reorientexpress.py -predict -data path_to_data -source experimental -model path_to_model -output my_predictions
```
This takes the experimental data stored in path_to_data and the model stored in path_to_model and predicts the 5'-to-3' orientation of reads, i.e. converts to forward reads the reads that the model predicts are reverse complemented, printing the results in my_predictions.csv. The output format is same as provided in the 'Examples of possible outputs section above'
In the saved_models/ folder we provide a model trained with the human transcriptome annotation and a model trained with the Saccharomyces cerevisiae transcriptome annoation. They can be directly used with the "-model" flag.
Example on test_case provided in the repo:
```
reorientexpress.py -predict -data ./test_case/experimental/Hopkins_Run1_20171011_1D.pass.dedup_60_unique_50k.fastq -model ./saved_models/Hs_transcriptome_mlp.model -source experimental -output my_predictions
```
or
```
reorientexpress-cnn.py -predict -data ./test_case/experimental/Hopkins_Run1_20171011_1D.pass.dedup_60_unique_50k.fastq -model ./saved_models/Hs_transcriptome_mlp.model -source experimental -output my_predictions
```
To test the accuracy of the model:
```
reorientexpress.py -test -data path_to_data -annotation path_of_paf_file -source mapped -model path_to_model
```
Example on test_case provided in the repo:
```
reorientexpress.py -test -data ./test_case/mapped/Hopkins_Run1_20171011_1D.pass.dedup_60_unique_2000.fastq -annotation ./test_case/mapped/cdna_human_no_secondary_mapq_60_unique_2000.paf -model ./saved_models/Hs_transcriptome_mlp.model -source mapped
```
or
```
reorientexpress-cnn.py -test -data ./test_case/mapped/Hopkins_Run1_20171011_1D.pass.dedup_60_unique_2000.fastq -annotation ./test_case/mapped/cdna_human_no_secondary_mapq_60_unique_2000.paf -model ./saved_models/Hs_transcriptome_mlp.model -source mapped
```
The ouput accuracy (precision, recall, F1-score, support) will be displayed on the screen.
| /reorientexpress-0.400.tar.gz/reorientexpress-0.400/README.md | 0.453262 | 0.986205 | README.md | pypi |
import pandas as pd
import knock_knock.target_info
from hits import utilities
def convert_to_sgRNA_coords(target_info, a):
''' given a in coordinates relative to anchor on + strand, convert to relative to sgRNA beginning and sgRNA strand'''
if target_info.sgRNA_feature.strand == '+':
return (a + target_info.anchor) - target_info.sgRNA_feature.start
else:
return target_info.sgRNA_feature.end - (a + target_info.anchor)
def convert_to_anchor_coords(target_info, s):
''' given s in coordinates relative to sgRNA beginning and sgRNA strand, convert to anchor coords + coords'''
if target_info.sgRNA_feature.strand == '+':
return (s + target_info.sgRNA_feature.start) - target_info.anchor
else:
return (target_info.sgRNA_feature.end - s) - target_info.anchor
def convert_deletion(d, source_target_info, dest_target_info):
''' deletions are defined by starts_ats and length.
When switching between anchor/+ and sgRNA/sgRNA strand coordinate, starts_ats may become ends_ats.
'''
start_end_pairs = list(zip(d.starts_ats, d.ends_ats))
sgRNA_coords = [(convert_to_sgRNA_coords(source_target_info, s), convert_to_sgRNA_coords(source_target_info, e)) for s, e in start_end_pairs]
anchor_coords = [(convert_to_anchor_coords(dest_target_info, s), convert_to_anchor_coords(dest_target_info, e)) for s, e in sgRNA_coords]
anchor_coords = [sorted(pair) for pair in anchor_coords]
starts_ats = sorted([s for s, e in anchor_coords])
return knock_knock.target_info.DegenerateDeletion(starts_ats, d.length)
def convert_insertion(ins, source_target_info, dest_target_info):
''' insertion are defined by starts_afters and seqs
When switching between anchor/+ and sgRNA/sgRNA strand coordinate, starts_afters may become starts_before,
and seqs maybe be reverse complemented.
'''
before_after_pairs = [(s, s + 1) for s in ins.starts_afters]
sgRNA_coords = [(convert_to_sgRNA_coords(source_target_info, b), convert_to_sgRNA_coords(source_target_info, a)) for a, b in before_after_pairs]
anchor_coords = [(convert_to_anchor_coords(dest_target_info, b), convert_to_anchor_coords(dest_target_info, a)) for a, b in sgRNA_coords]
anchor_coords = [sorted(pair) for pair in anchor_coords]
starts_afters = sorted([s for s, e in anchor_coords])
if source_target_info.sgRNA_feature.strand != dest_target_info.sgRNA_feature.strand:
seqs = [utilities.reverse_complement(seq) for seq in ins.seqs][::-1]
else:
seqs = ins.seqs
return knock_knock.target_info.DegenerateInsertion(starts_afters, seqs)
def convert_outcomes(outcomes, source_target_info, dest_target_info):
converted = []
for c, s, d in outcomes:
if c == 'deletion':
d = str(convert_deletion(knock_knock.target_info.DegenerateDeletion.from_string(d), source_target_info, dest_target_info))
elif c == 'insertion':
d = str(convert_insertion(knock_knock.target_info.DegenerateInsertion.from_string(d), source_target_info, dest_target_info))
elif c == 'wild type':
if s == 'indels':
d = 'indels'
s = 'clean'
converted.append((c, s, d))
return converted
def compare_pool_to_endogenous(pools, groups, destination_target_info=None, guide=None):
''' '''
if destination_target_info is None:
destination_target_info = pools[0].target_info
all_outcomes = set()
all_fs = []
all_l2fcs = {}
baseline_column_names = []
if not any(isinstance(group, tuple) for group in groups):
groups = [(group, None) for group in groups]
for group, condition in groups:
endogenous_outcomes = [(c, s, d) for c, s, d in group.outcomes_by_baseline_frequency if c != 'uncategorized' and s != 'far from cut' and s != 'mismatches']
endogenous_outcomes_converted = convert_outcomes(endogenous_outcomes, destination_target_info, group.target_info)
group_fs = group.outcome_fractions.loc[endogenous_outcomes]
group_fs.index = pd.MultiIndex.from_tuples(endogenous_outcomes_converted)
means = group.outcome_fraction_condition_means.loc[endogenous_outcomes]
means.index = pd.MultiIndex.from_tuples(endogenous_outcomes_converted)
means = pd.concat({'mean': means}, axis=1)
means.columns.names = ['replicate'] + group_fs.columns.names[:-1]
means.columns = means.columns.reorder_levels(group_fs.columns.names[:-1] + ['replicate'])
group_fs_with_means = pd.concat([group_fs, means], axis=1).sort_index(axis=1)
group_fs_with_means.columns = [f'{group.batch} {group.group} ' + ' '.join(map(str, v)) for v in group_fs_with_means.columns.values]
baseline_column_names.append(f'{group.batch} {group.group} ' + ' '.join(map(str, group.baseline_condition)) + ' mean')
all_fs.append(group_fs_with_means)
all_outcomes |= set(endogenous_outcomes_converted)
if condition is not None:
group_l2fcs = group.log2_fold_change_condition_means.loc[endogenous_outcomes, condition]
group_l2fcs.index = pd.MultiIndex.from_tuples(endogenous_outcomes_converted)
all_l2fcs[f'{group.batch} {group.group}'] = group_l2fcs
for pool in pools:
all_outcomes |= set(pool.non_targeting_fractions.index.values)
df = pd.DataFrame(pool.non_targeting_fractions)
df.columns = [pool.group]
all_fs.append(df)
baseline_column_names.append(pool.group)
if guide is not None:
all_l2fcs[pool.group] = pool.log2_fold_changes[guide]
all_outcomes = [(c, s, d) for c, s, d in all_outcomes if c != 'uncategorized']
all_fs = [v.reindex(all_outcomes).fillna(0) for v in all_fs]
for k, v in all_l2fcs.items():
all_l2fcs[k] = v.reindex(all_outcomes).fillna(0)
fs_df = pd.concat(all_fs, axis=1)
fs_df.index.names = ('category', 'subcategory', 'details')
l2fcs_df = pd.DataFrame(all_l2fcs)
if guide is not None or condition is not None:
l2fcs_df.index.names = ('category', 'subcategory', 'details')
# Collapse genomic insertions to one row.
genomic_insertion_collapsed = fs_df.loc[['genomic insertion']].groupby('subcategory').sum()
fs_df.drop('genomic insertion', inplace=True)
for subcategory, row in genomic_insertion_collapsed.iterrows():
fs_df.loc['genomic insertion', subcategory, 'collapsed'] = row
return fs_df, l2fcs_df, baseline_column_names
def compare_cut_sites(groups):
''' '''
all_fs = []
for group in groups:
group_fs = group.outcome_fractions
means = group.outcome_fraction_condition_means
means = pd.concat({'mean': means}, axis=1)
means.columns.names = ['replicate'] + group_fs.columns.names[:-1]
means.columns = means.columns.reorder_levels(group_fs.columns.names[:-1] + ['replicate'])
group_fs_with_means = pd.concat([group_fs, means], axis=1).sort_index(axis=1)
group_fs_with_means.columns = [f'{group.batch} {group.group} ' + ' '.join(map(str, v)) for v in group_fs_with_means.columns.values]
all_fs.append(group_fs_with_means)
fs_df = pd.concat(all_fs, axis=1).fillna(0)
fs_df.index.names = ('category', 'subcategory', 'details')
# Collapse genomic insertions to one row.
genomic_insertion_collapsed = fs_df.loc[['genomic insertion']].groupby('subcategory').sum()
fs_df.drop('genomic insertion', inplace=True)
fs_df.drop('uncategorized', inplace=True)
for subcategory, row in genomic_insertion_collapsed.iterrows():
fs_df.loc['genomic insertion', subcategory, 'collapsed'] = row
return fs_df | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/convert_outcomes.py | 0.435181 | 0.364862 | convert_outcomes.py | pypi |
from collections import Counter
from knock_knock import outcome_record
from hits.utilities import group_by
from .collapse_cython import hamming_distance_matrix, register_corrections
Pooled_UMI_Outcome = outcome_record.OutcomeRecord_factory(
columns_arg=[
'UMI',
'guide_mismatch',
'cluster_id',
'num_reads',
'inferred_amplicon_length',
'category',
'subcategory',
'details',
'query_name',
'common_sequence_name',
],
converters_arg={
'num_reads': int,
'guide_mismatch': int,
'inferred_amplicon_length': int,
},
)
gDNA_Outcome = outcome_record.OutcomeRecord_factory(
columns_arg=[
'query_name',
'guide_mismatches',
'inferred_amplicon_length',
'category',
'subcategory',
'details',
'common_sequence_name',
],
converters_arg={'inferred_amplicon_length': int},
)
def collapse_pooled_UMI_outcomes(outcome_iter):
def is_relevant(outcome):
return (outcome.category != 'bad sequence' and
outcome.outcome != ('no indel', 'other', 'ambiguous')
)
all_outcomes = [o for o in outcome_iter if is_relevant(o)]
all_outcomes = sorted(all_outcomes, key=lambda u: (u.UMI, u.cluster_id))
all_collapsed_outcomes = []
most_abundant_outcomes = []
for UMI, UMI_outcomes in group_by(all_outcomes, lambda u: u.UMI):
observed = set(u.outcome for u in UMI_outcomes)
collapsed_outcomes = []
for outcome in observed:
relevant = [u for u in UMI_outcomes if u.outcome == outcome]
representative = max(relevant, key=lambda u: u.num_reads)
representative.num_reads = sum(u.num_reads for u in relevant)
collapsed_outcomes.append(representative)
all_collapsed_outcomes.append(representative)
max_count = max(u.num_reads for u in collapsed_outcomes)
has_max_count = [u for u in collapsed_outcomes if u.num_reads == max_count]
if len(has_max_count) == 1:
most_abundant_outcomes.append(has_max_count[0])
all_collapsed_outcomes = sorted(all_collapsed_outcomes, key=lambda u: (u.UMI, u.cluster_id))
return all_collapsed_outcomes, most_abundant_outcomes
def error_correct_outcome_UMIs(outcome_group, max_UMI_distance=1):
# sort UMIs in descending order by number of occurrences.
UMI_read_counts = Counter()
for outcome in outcome_group:
UMI_read_counts[outcome.UMI] += outcome.num_reads
UMIs = [UMI for UMI, read_count in UMI_read_counts.most_common()]
ds = hamming_distance_matrix(UMIs)
corrections = register_corrections(ds, max_UMI_distance, UMIs)
for outcome in outcome_group:
correct_to = corrections.get(outcome.UMI)
if correct_to:
outcome.UMI = correct_to
return outcome_group | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/coherence.py | 0.597608 | 0.172033 | coherence.py | pypi |
from pathlib import Path
from collections import defaultdict
import pandas as pd
from hits import utilities, fasta, mapping_tools
memoized_property = utilities.memoized_property
memoized_with_args = utilities.memoized_with_args
class GuideLibrary:
def __init__(self, base_dir, name):
self.base_dir = Path(base_dir)
self.name = name
self.full_dir = self.base_dir / 'guides' / name
self.reference_STAR_index = '/nvme/indices/refdata-cellranger-GRCh38-1.2.0/star'
self.fns = {
'guides': self.full_dir / 'guides.txt',
'guides_fasta': self.full_dir / 'guides.fasta',
'best_promoters': self.full_dir / 'best_promoters.txt',
'updated_gene_names': self.full_dir / 'updated_gene_names.txt',
'non_targeting_guide_sets': self.full_dir / 'non_targeting_guide_sets.txt',
'protospacers': self.full_dir / 'protospacers.fasta',
'perturbseq_STAR_index': self.full_dir / 'perturbseq_STAR_index',
'cell_cycle_phase_fractions': self.full_dir / 'cell_cycle_phase_fractions.txt',
'cell_cycle_log2_fold_changes': self.full_dir / 'cell_cycle_effects.txt',
'K562_knockdown': self.full_dir / 'K562_knockdown.txt',
}
@memoized_property
def guides_df(self):
guides_df = pd.read_csv(self.fns['guides'], index_col='short_name', sep='\t')
if 'promoter' in guides_df.columns:
guides_df.loc[guides_df['promoter'].isnull(), 'promoter'] = 'P1P2'
else:
guides_df['promoter'] = 'P1P2'
guides_df['best_promoter'] = True
for gene, promoter in self.best_promoters.items():
not_best = guides_df.query('gene == @gene and promoter != @promoter').index
guides_df.loc[not_best, 'best_promoter'] = False
guides_df = guides_df.sort_values(['gene', 'promoter', 'rank'])
return guides_df
@memoized_property
def best_promoters(self):
if self.fns['best_promoters'].exists():
best_promoters = pd.read_csv(self.fns['best_promoters'], index_col='gene', squeeze=True, sep='\t')
else:
best_promoters = {}
return best_promoters
@memoized_property
def old_gene_to_new_gene(self):
updated_gene_names = pd.read_csv(self.fns['updated_gene_names'], index_col=0, squeeze=True, seq='\t')
return updated_gene_names
@memoized_property
def new_gene_to_old_gene(self):
new_to_old_dict = utilities.reverse_dictionary(self.old_gene_to_new_gene)
def new_gene_to_old_gene(new_gene):
return new_to_old_dict.get(new_gene, new_gene)
return new_gene_to_old_gene
def make_protospacer_fasta(self):
with open(self.fns['protospacers'], 'w') as fh:
for name, seq in self.guides_df['protospacer'].items():
# Remove initial G from seq.
record = fasta.Record(name, seq[1:])
fh.write(str(record))
def make_guides_fasta(self):
with open(self.fns['guides_fasta'], 'w') as fh:
for name, seq in self.guides_df['full_seq'].items():
record = fasta.Record(name, seq)
fh.write(str(record))
@memoized_property
def guides(self):
guides = self.guides_df.index.values
return guides
@memoized_property
def non_targeting_guides(self):
return [g for g in self.guides if 'non-targeting' in g]
@memoized_property
def targeting_guides(self):
return [g for g in self.guides if 'non-targeting' not in g and 'eGFP' not in g]
@memoized_property
def genes(self):
return sorted(set(self.guides_df['gene']))
@memoized_property
def genes_with_non_targeting_guide_sets(self):
genes = self.genes + sorted(self.non_targeting_guide_sets)
return genes
def gene_guides(self, gene, only_best_promoter=False):
if isinstance(gene, str):
genes = [gene]
else:
genes = gene
query = 'gene in @genes'
if only_best_promoter:
query += ' and best_promoter'
gene_guides = self.guides_df.query(query).sort_values(['gene', 'promoter', 'rank'])
nt_guides = []
for gene in genes:
nt_guides.extend(self.non_targeting_guide_sets.get(gene, []))
all_guides = list(gene_guides.index) + nt_guides
return all_guides
@memoized_property
def guide_to_gene(self):
guide_to_gene = self.guides_df['gene'].copy()
return guide_to_gene
@memoized_property
def guide_to_gene_with_non_targeting_guide_sets(self):
guide_to_gene = self.guides_df['gene'].copy()
for nt_guide_set, guides in self.non_targeting_guide_sets.items():
for guide in guides:
guide_to_gene[guide] = nt_guide_set
return guide_to_gene
@memoized_property
def guide_barcodes(self):
return self.guides_df['guide_barcode']
@memoized_with_args
def gene_indices(self, gene):
idxs = [i for i, g in enumerate(self.guides_df['gene']) if g == gene]
return min(idxs), max(idxs)
@memoized_property
def cell_cycle_log2_fold_changes(self):
return pd.read_csv(self.fns['cell_cycle_log2_fold_changes'], index_col=0)
@memoized_property
def cell_cycle_phase_fractions(self):
return pd.read_csv(self.fns['cell_cycle_phase_fractions'], index_col=0)
@memoized_property
def K562_knockdown(self):
return pd.read_csv(self.fns['K562_knockdown'], index_col=0)
@memoized_property
def non_targeting_guide_sets(self):
non_targeting_guide_sets = {}
with open(self.fns['non_targeting_guide_sets']) as fh:
for i, line in enumerate(fh):
guides = line.strip().split(',')
set_name = f'non-targeting_set_{i:05d}'
non_targeting_guide_sets[set_name] = list(guides)
return non_targeting_guide_sets
class DummyGuideLibrary:
def __init__(self):
self.guides = ['none']
self.non_targeting_guides = ['none']
self.genes = ['negative_control']
self.name = None
@memoized_property
def guide_to_gene(self):
return defaultdict(lambda: 'none')
def gene_guides(self, gene, **kwargs):
if gene == 'negative_control':
return ['none']
else:
return []
dummy_guide_library = DummyGuideLibrary() | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/guide_library.py | 0.642432 | 0.201715 | guide_library.py | pypi |
import copy
from collections import defaultdict, Counter
import hdbscan
import matplotlib
import matplotlib.cm
import matplotlib.pyplot as plt
import numpy as np
import sklearn.metrics.pairwise
import scipy.cluster.hierarchy as sch
import scipy.spatial.distance as ssd
import seaborn as sns
import pandas as pd
import umap
import hits.utilities
import hits.visualize
import knock_knock.outcome
from . import visualize
memoized_property = hits.utilities.memoized_property
def get_outcomes_and_guides(pool, outcomes, guides, fixed_guide='none', min_UMIs=None, only_best_promoter=True):
if isinstance(outcomes, int):
outcomes = pool.most_frequent_outcomes(fixed_guide)[:outcomes]
if isinstance(guides, int):
phenotype_strengths = pool.chi_squared_per_guide(outcomes, fixed_guide=fixed_guide, only_best_promoter=only_best_promoter)
if min_UMIs is not None:
UMIs = pool.UMI_counts('perfect').loc[fixed_guide]
enough_UMIs = UMIs[UMIs > min_UMIs].index
phenotype_strengths = phenotype_strengths[phenotype_strengths.index.isin(enough_UMIs)]
guides = phenotype_strengths.index[:guides]
return outcomes, guides
class Clusterer:
def __init__(self, **options):
options.setdefault('outcomes_selection_method', 'above_frequency_threshold')
options.setdefault('outcomes_selection_kwargs', {})
options['outcomes_selection_kwargs'].setdefault('threshold', 2e-3)
options.setdefault('guides_selection_method', 'chi_squared_multiple')
options.setdefault('guides_selection_kwargs', {})
options['guides_selection_kwargs'].setdefault('multiple', 2)
options['guides_selection_kwargs'].setdefault('n', 100)
options.setdefault('guides_method', 'HDBSCAN')
options.setdefault('guides_kwargs', {})
options.setdefault('guides_seed', 0)
options.setdefault('outcomes_seed', 0)
options.setdefault('outcomes_min_dist', 0.2)
options.setdefault('guides_min_dist', 0.2)
options.setdefault('outcomes_method', 'hierarchical')
options.setdefault('outcomes_kwargs', {})
options.setdefault('use_high_frequency_counts', False)
options['guides_kwargs'].setdefault('metric', 'cosine')
options['outcomes_kwargs'].setdefault('metric', 'correlation')
self.options = options
def perform_clustering(self, axis):
method = self.options[f'{axis}_method']
if method == 'hierarchical':
func = hierarchical
elif method == 'HDBSCAN':
func = HDBSCAN
elif method == 'precomputed':
func = process_precomputed
else:
raise ValueError(method)
results = func(self.log2_fold_changes, axis, **self.options[f'{axis}_kwargs'])
assign_palette(results, axis)
return results
@memoized_property
def outcome_clustering(self):
return self.perform_clustering('outcomes')
@memoized_property
def guide_clustering(self):
return self.perform_clustering('guides')
@memoized_property
def clustered_log2_fold_changes(self):
return self.log2_fold_changes.loc[self.clustered_outcomes, self.clustered_guides]
@memoized_property
def clustered_guides(self):
return self.guide_clustering['clustered_order']
@memoized_property
def clustered_outcomes(self):
return self.outcome_clustering['clustered_order']
@memoized_property
def outcomes_with_pool(self):
return self.log2_fold_changes.index.values
@memoized_property
def all_log2_fold_changes(self):
return self.guide_log2_fold_changes(self.guide_library.guides)
@memoized_property
def all_guide_correlations(self):
all_l2fcs = self.all_log2_fold_changes
all_corrs = all_l2fcs.corr().stack()
all_corrs.index.names = ['guide_1', 'guide_2']
all_corrs = pd.DataFrame(all_corrs)
all_corrs.columns = ['r']
# Make columns out of index levels.
for k in all_corrs.index.names:
all_corrs[k] = all_corrs.index.get_level_values(k)
guide_to_gene = self.guide_library.guide_to_gene
all_corrs['gene_1'] = guide_to_gene.loc[all_corrs['guide_1']].values
all_corrs['gene_2'] = guide_to_gene.loc[all_corrs['guide_2']].values
for guide in ['guide_1', 'guide_2']:
all_corrs[f'{guide}_is_active'] = all_corrs[guide].isin(self.guides)
all_corrs['both_active'] = all_corrs[['guide_1_is_active', 'guide_2_is_active']].all(axis=1)
distinct_guides = all_corrs.query('guide_1 < guide_2')
return distinct_guides.sort_values('r', ascending=False)
@memoized_property
def guide_embedding(self):
reducer = umap.UMAP(random_state=self.options['guides_seed'],
metric=self.options['guides_kwargs']['metric'],
n_neighbors=10,
min_dist=self.options['guides_min_dist'],
)
embedding = reducer.fit_transform(self.log2_fold_changes.T)
embedding = pd.DataFrame(embedding,
columns=['x', 'y'],
index=self.log2_fold_changes.columns,
)
embedding['color'] = pd.Series(self.guide_clustering['colors'])
embedding['gene'] = [self.guide_to_gene[guide] for guide in embedding.index]
embedding['cluster_assignments'] = pd.Series(self.guide_clustering['cluster_assignments'], index=self.guide_clustering['clustered_order'])
return embedding
@memoized_property
def outcome_embedding(self):
reducer = umap.UMAP(random_state=self.options['outcomes_seed'],
metric=self.options['outcomes_kwargs']['metric'],
n_neighbors=10,
min_dist=self.options['outcomes_min_dist'],
)
embedding = reducer.fit_transform(self.log2_fold_changes)
embedding = pd.DataFrame(embedding,
columns=['x', 'y'],
index=self.log2_fold_changes.index,
)
embedding['color'] = pd.Series(self.outcome_clustering['colors'])
embedding['cluster_assignment'] = pd.Series(self.outcome_clustering['cluster_assignments'], index=self.outcome_clustering['clustered_order'])
MH_lengths = []
deletion_lengths = []
directionalities = []
fractions = []
for pn, c, s, d in embedding.index.values:
pool = self.pn_to_pool[pn]
ti = self.pn_to_target_info[pn]
if c == 'deletion':
deletion = knock_knock.outcome.DeletionOutcome.from_string(d).undo_anchor_shift(ti.anchor)
MH_length = len(deletion.deletion.starts_ats) - 1
deletion_length = deletion.deletion.length
directionality = deletion.classify_directionality(ti)
else:
MH_length = -1
deletion_length = -1
directionality = 'n/a'
MH_lengths.append(MH_length)
deletion_lengths.append(deletion_length)
directionalities.append(directionality)
if self.options['use_high_frequency_counts']:
fraction = pool.high_frequency_outcome_fractions.loc[(c, s, d), 'all_non_targeting']
else:
fraction = pool.outcome_fractions().loc[(c, s, d), ('none', 'all_non_targeting')]
fractions.append(fraction)
embedding['MH length'] = MH_lengths
embedding['deletion length'] = deletion_lengths
embedding['directionality'] = directionalities
embedding['fraction'] = fractions
embedding['log10_fraction'] = np.log10(embedding['fraction'])
effector = self.target_info.effector.name
categories = embedding.index.get_level_values('category')
combined_categories = []
for category, directionality in zip(categories, embedding['directionality']):
if category != 'deletion':
combined_category = category
else:
combined_category = f'{category}, {directionality}'
combined_categories.append(combined_category)
combined_categories = pd.Series(combined_categories, index=embedding.index)
categories_aliased = combined_categories.map(visualize.category_aliases[effector])
value_to_color = visualize.category_alias_colors[effector]
colors = categories_aliased.map(value_to_color)
embedding['combined_categories'] = combined_categories
embedding['categories_aliased'] = categories_aliased
embedding['category_colors'] = colors
embedding['sgRNA'] = embedding.index.get_level_values('pool_name').map(self.pn_to_sgRNA).values
return embedding
def plot_guide_embedding(self,
figsize=(12, 12),
color_by='cluster',
label_genes=True,
label_alpha=1,
legend_location='lower right',
label_size=8,
marker_size=None,
circle_edge_width=0,
gene_line_width=1,
):
fig, ax = plt.subplots(figsize=figsize)
data = self.guide_embedding.copy()
if color_by == 'cluster':
# sort so that guides assigned to clusters are drawn on top
data['sort_by'] = data['cluster_assignments']
data = data.sort_values(by='sort_by')
elif color_by == 'gamma':
min_gamma = -0.3
values = self.guide_library.guides_df['gamma'].loc[data.index]
cmap = visualize.gamma_cmap
norm = matplotlib.colors.Normalize(vmin=min_gamma, vmax=0)
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
colors = [tuple(row) for row in sm.to_rgba(values)]
data['color'] = colors
# sort so that strongest (negative) gammas are drawn on top
data['sort_by'] = -values
data = data.sort_values(by='sort_by')
ax_p = ax.get_position()
if legend_location == 'lower right':
x0 = ax_p.x0 + 0.65 * ax_p.width
y0 = ax_p.y0 + 0.25 * ax_p.height
elif legend_location == 'upper right':
x0 = ax_p.x0 + 0.70 * ax_p.width
y0 = ax_p.y0 + 0.95 * ax_p.height
else:
raise NotImplementedError
cax = fig.add_axes([x0,
y0,
0.25 * ax_p.width,
0.03 * ax_p.height,
])
colorbar = plt.colorbar(mappable=sm, cax=cax, orientation='horizontal')
ticks = [min_gamma, 0]
tick_labels = [str(t) for t in ticks]
tick_labels[0] = '$\leq$' + tick_labels[0]
colorbar.set_ticks(ticks)
colorbar.set_ticklabels(tick_labels)
colorbar.outline.set_alpha(0)
colorbar.set_label(f'gamma', size=6)
cax.tick_params(labelsize=6, width=0.5)
elif color_by in self.guide_library.cell_cycle_log2_fold_changes.index:
phase = color_by
values = self.guide_library.cell_cycle_log2_fold_changes.loc[phase, data.index]
cmap = visualize.cell_cycle_cmap
norm = matplotlib.colors.Normalize(vmin=-1, vmax=1)
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
colors = [tuple(row) for row in sm.to_rgba(values)]
data['color'] = colors
# sort by absolute value of fold changes so that most extreme
# points are drawn on top
data['sort_by'] = np.abs(values)
data = data.sort_values(by='sort_by')
ax_p = ax.get_position()
if legend_location == 'lower right':
x0 = ax_p.x0 + 0.65 * ax_p.width
y0 = ax_p.y0 + 0.25 * ax_p.height
elif legend_location == 'upper right':
x0 = ax_p.x0 + 0.70 * ax_p.width
y0 = ax_p.y0 + 0.95 * ax_p.height
else:
raise NotImplementedError
cax = fig.add_axes([x0,
y0,
0.25 * ax_p.width,
0.03 * ax_p.height,
])
colorbar = plt.colorbar(mappable=sm, cax=cax, orientation='horizontal')
ticks = [-1, 0, 1]
tick_labels = [str(t) for t in ticks]
tick_labels[0] = '$\leq$' + tick_labels[0]
tick_labels[-1] = '$\geq$' + tick_labels[-1]
colorbar.set_ticks(ticks)
colorbar.set_ticklabels(tick_labels)
colorbar.outline.set_alpha(0)
colorbar.set_label(f'log$_2$ fold change\nin {phase} occupancy', size=6)
cax.tick_params(labelsize=6, width=0.5)
if marker_size is None:
marker_size = figsize[0] * 100 / 12
ax.scatter(x='x',
y='y',
color='color',
data=data,
s=marker_size,
alpha=0.8,
edgecolors='black',
linewidths=(circle_edge_width,),
#marker='s',
)
if label_genes == 'all':
for guide, row in self.guide_embedding.iterrows():
ax.annotate(row['gene'],
xy=(row['x'], row['y']),
xytext=(2, 0),
textcoords='offset points',
ha='left',
va='center',
size=label_size,
alpha=label_alpha,
)
elif label_genes:
for gene, rows in self.guide_embedding.groupby('gene', sort=False):
n_rows, _ = rows.shape
for first_index in range(n_rows):
first_row = rows.iloc[first_index]
for second_index in range(first_index + 1, n_rows):
second_row = rows.iloc[second_index]
xs = [first_row['x'], second_row['x']]
ys = [first_row['y'], second_row['y']]
if gene_line_width > 0:
ax.plot(xs, ys, linewidth=gene_line_width, color='black', alpha=0.1)
centroid = (rows['x'].mean(), rows['y'].mean())
if n_rows == 1:
num_guides_string = ''
else:
num_guides_string = f' ({len(rows)})'
ax.annotate(f'{gene}{num_guides_string}',
xy=centroid,
xytext=(0, 0),
textcoords='offset points',
ha='center',
va='center',
size=label_size,
alpha=label_alpha,
)
x_min = data['x'].min()
x_max = data['x'].max()
x_range = x_max - x_min
buffer = 0.05 * x_range
ax.set_xlim(x_min - buffer, x_max + buffer)
y_min = data['y'].min()
y_max = data['y'].max()
y_range = y_max - y_min
buffer = 0.05 * y_range
ax.set_ylim(y_min - buffer, y_max + buffer)
ax.set_xticks([])
ax.set_yticks([])
plt.setp(fig.axes[0].spines.values(), alpha=0.5)
return fig, data
def plot_outcome_embedding(self,
marker_size=35,
alpha=0.8,
color_by='cluster',
ax=None,
figsize=(6, 6),
draw_legend=True,
legend_location='upper left',
legend_text_size=12,
marker='o',
):
if ax is None:
fig, ax = plt.subplots(figsize=figsize)
else:
fig = ax.get_figure()
data = self.outcome_embedding.copy()
common_kwargs = dict(
x='x',
y='y',
s=marker_size,
alpha=alpha,
linewidths=(0,),
clip_on=False,
color='color',
marker=marker,
)
needs_categorical_legend = False
value_to_color = None
if color_by is None:
data['color'] = 'grey'
ax.scatter(data=data,
**common_kwargs,
)
elif color_by == 'cluster':
# To prevent privileging specific clusters, don't
# sort by cluster assignment, but do plot assigned
# points after unasigned ones.
ax.scatter(data=data.query('cluster_assignment == -1'),
**common_kwargs,
)
ax.scatter(data=data.query('cluster_assignment != -1'),
**common_kwargs,
)
else:
data_query = None
if color_by == 'sgRNA':
colors, value_to_color = self.sgRNA_colors()
data['color'] = colors
# Shuffle to randomize z-order.
data = data.sample(frac=1)
needs_categorical_legend = True
elif color_by == 'category' or color_by in data.index.levels[1]:
effector = self.target_info.effector.name
value_to_color = visualize.category_alias_colors[effector]
data['color'] = data['category_colors']
if color_by != 'category':
data.loc[data.index.get_level_values('category') != color_by, 'color'] = 'grey'
# Force insertions to be drawn on top.
data = data.sort_values(by='categories_aliased', key=lambda s: s == 'insertion')
needs_categorical_legend = True
elif color_by in ['MH length', 'deletion length', 'fraction', 'log10_fraction']:
if color_by == 'MH length':
min_length = 0
max_length = 3
cmap = copy.copy(plt.get_cmap('viridis_r'))
tick_step = 1
label = 'nts of flanking\nmicrohomology'
data_query = 'category == "deletion"'
elif color_by == 'deletion length':
min_length = 0
max_length = 30
cmap = copy.copy(plt.get_cmap('Purples'))
tick_step = 10
label = 'deletion length'
data_query = 'category == "deletion"'
elif color_by == 'fraction':
min_length = 0
max_length = 0.1
cmap = copy.copy(plt.get_cmap('YlOrBr'))
tick_step = 0.25
label = 'fraction of outcomes'
data_query = None
elif color_by == 'log10_fraction':
min_length = -3
max_length = -1
cmap = copy.copy(plt.get_cmap('YlOrBr'))
tick_step = 1
label = 'log$_{10}$ baseline\nfraction\nof outcomes\nwithin screen'
data_query = None
else:
raise ValueError(color_by)
if color_by == 'MH length':
norm = matplotlib.colors.Normalize(vmin=min_length, vmax=max_length)
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
colors = sm.to_rgba(np.arange(min_length, max_length + 1))
cmap = matplotlib.colors.ListedColormap(colors)
norm = matplotlib.colors.BoundaryNorm(np.arange(-0.5, max_length + 1), cmap.N)
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
else:
cmap.set_under('white')
norm = matplotlib.colors.Normalize(vmin=min_length, vmax=max_length)
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
vs = self.outcome_embedding[color_by]
colors = [tuple(row) for row in sm.to_rgba(vs)]
data['color'] = colors
data['sort_by'] = vs
data = data.sort_values(by='sort_by')
ax_p = ax.get_position()
if 'left' in legend_location:
x0 = ax_p.x0 + 0.45 * ax_p.width
elif 'middle' in legend_location:
x0 = ax_p.x0 + 0.5 * ax_p.width
else:
x0 = ax_p.x0 + 0.75 * ax_p.width
if 'upper' in legend_location:
y0 = ax_p.y0 + 0.65 * ax_p.height
elif 'middle' in legend_location:
y0 = ax_p.y0 + 0.5 * ax_p.height - 0.125 * ax_p.height
elif 'lower' in legend_location or 'bottom' in legend_location:
y0 = ax_p.y0 + 0.2 * ax_p.height - 0.125 * ax_p.height
cax = fig.add_axes([x0,
y0,
0.03 * ax_p.width,
0.25 * ax_p.height,
])
colorbar = plt.colorbar(mappable=sm, cax=cax)
ticks = np.arange(min_length, max_length + 1, tick_step)
tick_labels = [str(t) for t in ticks]
tick_labels[-1] = '$\geq$' + tick_labels[-1]
if min_length != 0:
tick_labels[0] = '$\leq$' + tick_labels[0]
colorbar.set_ticks(ticks)
colorbar.set_ticklabels(tick_labels)
colorbar.outline.set_alpha(0)
cax.tick_params(labelsize=6, width=0.5, length=2)
cax.annotate(label,
xy=(0, 0.5),
xycoords='axes fraction',
xytext=(-5, 0),
textcoords='offset points',
va='center',
ha='right',
size=legend_text_size,
)
else:
# color_by is be a guide name. This might be a guide that was among
# those used for clustering or might not.
guide = color_by
if guide in self.log2_fold_changes:
values = self.log2_fold_changes[guide]
else:
values = self.guide_log2_fold_changes(guide)
values = values.loc[data.index]
norm = matplotlib.colors.Normalize(vmin=-2, vmax=2)
cmap = visualize.fold_changes_cmap
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
colors = [tuple(row) for row in sm.to_rgba(values)]
data['color'] = colors
# sort by absolute value of fold changes so that most extreme
# points are drawn on top
data['sort_by'] = np.abs(values)
data = data.sort_values(by='sort_by')
ax_p = ax.get_position()
if 'left' in legend_location:
x0 = ax_p.x0 + 0.1 * ax_p.width
elif 'middle' in legend_location:
x0 = ax_p.x0 + 0.4 * ax_p.width
else:
x0 = ax_p.x0 + 0.6 * ax_p.width
if 'upper' in legend_location:
y0 = ax_p.y0 + 0.75 * ax_p.height
elif 'middle' in legend_location:
y0 = ax_p.y0 + 0.5 * ax_p.height - 0.125 * ax_p.height
elif 'lower' in legend_location or 'bottom' in legend_location:
y0 = ax_p.y0 + 0.3 * ax_p.height - 0.125 * ax_p.height
cax = fig.add_axes([x0,
y0,
0.3 * ax_p.width,
0.03 * ax_p.height,
])
colorbar = plt.colorbar(mappable=sm, cax=cax, orientation='horizontal')
ticks = [-2, 0, 2]
tick_labels = [str(t) for t in ticks]
tick_labels[0] = '$\leq$' + tick_labels[0]
tick_labels[-1] = '$\geq$' + tick_labels[-1]
colorbar.set_ticks(ticks)
colorbar.set_ticklabels(tick_labels)
colorbar.outline.set_alpha(0)
colorbar.set_label('log$_2$ fold change\nfrom non-targeting', size=legend_text_size)
cax.tick_params(labelsize=6, width=0.5, length=2)
cax.annotate(guide,
xy=(0.5, 1),
xycoords='axes fraction',
xytext=(0, legend_text_size / 2),
textcoords='offset points',
va='bottom',
ha='center',
size=legend_text_size,
color=self.guide_embedding['color'].get(guide, 'black')
)
if data_query is not None:
to_plot = data.query(data_query)
else:
to_plot = data
ax.scatter(data=to_plot,
**common_kwargs,
)
x_min = data['x'].min()
x_max = data['x'].max()
x_range = x_max - x_min
buffer = 0.05 * x_range
ax.set_xlim(x_min - buffer, x_max + buffer)
y_min = data['y'].min()
y_max = data['y'].max()
y_range = y_max - y_min
buffer = 0.05 * y_range
ax.set_ylim(y_min - buffer, y_max + buffer)
ax.set_xticks([])
ax.set_yticks([])
if draw_legend and needs_categorical_legend:
hits.visualize.draw_categorical_legend(value_to_color, ax, font_size=legend_text_size, legend_location=legend_location)
plt.setp(fig.axes[0].spines.values(), alpha=0.5)
return fig, data, value_to_color
def hierarchical(l2fcs,
axis,
metric='correlation',
method='single',
**fcluster_kwargs,
):
fcluster_kwargs.setdefault('criterion', 'maxclust')
fcluster_kwargs.setdefault('t', 5)
if axis == 'guides':
to_cluster = l2fcs.T
elif axis == 'outcomes':
to_cluster = l2fcs
else:
raise ValueError(axis)
labels = list(to_cluster.index.values)
linkage = sch.linkage(to_cluster,
optimal_ordering=True,
metric=metric,
method=method,
)
dendro = sch.dendrogram(linkage,
no_plot=True,
labels=labels,
)
clustered_order = dendro['ivl']
cluster_ids = sch.fcluster(linkage, **fcluster_kwargs)
# Transform from original order into the order produced by dendrogram.
# Convert from 1-based indexing to 0-based indexing for consistency with other methods.
cluster_assignments = [cluster_ids[labels.index(label)] - 1 for label in clustered_order]
if axis == 'guides':
l2fcs_reordered = l2fcs.loc[:, clustered_order]
elif axis == 'outcomes':
l2fcs_reordered = l2fcs.loc[clustered_order, :].T
else:
raise ValueError(axis)
if metric == 'correlation':
similarities = l2fcs_reordered.corr()
elif metric == 'cosine':
similarities = sklearn.metrics.pairwise.cosine_similarity(l2fcs_reordered.T)
elif metric == 'euclidean':
similarities = 1 / (1 + ssd.squareform(ssd.pdist(l2fcs_reordered.T)))
else:
similarities = None
results = {
'linkage': linkage,
'dendro': dendro,
'clustered_order': clustered_order,
'cluster_assignments': cluster_assignments,
'similarities': similarities,
}
return results
def HDBSCAN(l2fcs,
axis,
min_cluster_size=2,
min_samples=1,
cluster_selection_epsilon=0.2,
metric='cosine',
cluster_selection_method='eom',
):
# cosine_distances wants samples to be rows and features to be columns
if axis == 'guides':
to_cluster = l2fcs.T
elif axis == 'outcomes':
to_cluster = l2fcs
else:
raise ValueError(axis)
if metric == 'cosine':
distances = sklearn.metrics.pairwise.cosine_distances(to_cluster)
elif metric == 'correlation':
distances = 1 - to_cluster.T.corr()
elif metric == 'euclidean':
distances = ssd.squareform(ssd.pdist(to_cluster))
else:
distances = None
labels = list(to_cluster.index.values)
distances = pd.DataFrame(distances, index=labels, columns=labels)
clusterer = hdbscan.HDBSCAN(metric='precomputed',
min_cluster_size=min_cluster_size,
min_samples=min_samples,
cluster_selection_epsilon=cluster_selection_epsilon,
cluster_selection_method=cluster_selection_method,
)
clusterer.fit(distances)
linkage = clusterer.single_linkage_tree_.to_numpy()
linkage = sch.optimal_leaf_ordering(linkage, ssd.squareform(distances))
dendro = sch.dendrogram(linkage,
no_plot=True,
labels=labels,
)
clustered_order = dendro['ivl']
cluster_ids = clusterer.labels_
# Transform from original order into the order produced by dendrogram.
cluster_assignments = [cluster_ids[labels.index(l)] for l in clustered_order]
if axis == 'guides':
l2fcs_reordered = l2fcs.loc[:, clustered_order]
elif axis == 'outcomes':
l2fcs_reordered = l2fcs.loc[clustered_order, :].T
else:
raise ValueError(axis)
if metric == 'correlation':
similarities = l2fcs_reordered.corr()
elif metric == 'cosine':
similarities = sklearn.metrics.pairwise.cosine_similarity(l2fcs_reordered.T)
elif metric == 'euclidean':
similarities = 1 / (1 + ssd.squareform(ssd.pdist(l2fcs_reordered.T)))
else:
similarities = None
results = {
'clustered_order': clustered_order,
'cluster_assignments': cluster_assignments,
'distances': distances.loc[clustered_order, clustered_order],
'similarities': similarities,
'linkage': linkage,
'original_order': labels,
'clusterer': clusterer,
}
return results
def process_precomputed(l2fcs,
axis,
metric='cosine',
**fcluster_kwargs,
):
if axis == 'guides':
to_cluster = l2fcs.T
elif axis == 'outcomes':
to_cluster = l2fcs
else:
raise ValueError(axis)
if axis == 'guides':
l2fcs_reordered = l2fcs
elif axis == 'outcomes':
l2fcs_reordered = l2fcs.T
labels = list(to_cluster.index.values)
if metric == 'correlation':
similarities = l2fcs_reordered.corr()
elif metric == 'cosine':
similarities = sklearn.metrics.pairwise.cosine_similarity(l2fcs_reordered.T)
elif metric == 'euclidean':
similarities = 1 / (1 + ssd.squareform(ssd.pdist(l2fcs_reordered.T)))
else:
similarities = None
results = {
'clustered_order': labels,
'similarities': similarities,
}
return results
def get_cluster_blocks(cluster_assignments):
''' tuples of inclusive index boundaries of connected blocks of cluster ids '''
cluster_ids = set(cluster_assignments)
cluster_blocks = {}
for cluster_id in cluster_ids:
idxs = [idx for idx, c_id in enumerate(cluster_assignments) if c_id == cluster_id]
if len(idxs) == 1:
blocks = [(idxs[0], idxs[0])]
else:
blocks = []
current_value = idxs[0]
block_start = current_value
for current_index in range(1, len(idxs)):
previous_value = current_value
current_value = idxs[current_index]
if current_value - previous_value == 1:
continue
else:
block_end = idxs[current_index - 1]
blocks.append((block_start, block_end))
block_start = current_value
block_end = idxs[current_index]
blocks.append((block_start, block_end))
cluster_blocks[cluster_id] = blocks
return cluster_blocks
class SinglePoolClusterer(Clusterer):
def __init__(self, pool, **options):
self.pool = pool
self.guide_library = self.pool.variable_guide_library
self.pn_to_target_info = {self.pool.short_name: self.pool.target_info}
self.pn_to_sgRNA = {self.pool.short_name: self.pool.target_info.sgRNA}
self.pn_to_pool = {self.pool.short_name: self.pool}
super().__init__(**options)
@memoized_property
def guide_to_gene(self):
return self.pool.variable_guide_library.guide_to_gene
@property
def target_info(self):
return self.pool.target_info
@memoized_property
def outcomes(self):
selection_method = self.options['outcomes_selection_method']
if pd.api.types.is_list_like(selection_method):
outcomes = selection_method
elif selection_method == 'category':
category = self.options['outcomes_selection_kwargs']['category']
min_f = self.options['outcomes_selection_kwargs']['threshold']
outcomes = [(c, s, d) for (c, s, d), f in self.pool.non_targeting_fractions.items() if c == category and f >= min_f]
elif selection_method == 'above_frequency_threshold':
threshold = self.options['outcomes_selection_kwargs']['threshold']
outcomes = self.pool.outcomes_above_simple_threshold(frequency_threshold=threshold,
use_high_frequency_counts=self.options['use_high_frequency_counts'],
)
elif selection_method == 'top_n':
num_outcomes = self.options['outcomes_selection_kwargs']['n']
outcomes = self.pool.most_frequent_outcomes(use_high_frequency_counts=self.options['use_high_frequency_counts'])[:num_outcomes]
else:
outcomes = self.pool.canonical_outcomes
return outcomes
@property
def original_outcome_order_with_pool(self):
return [(self.pool.short_name, *vs) for vs in self.outcomes]
@memoized_property
def guides(self):
if pd.api.types.is_list_like(self.options['guides_selection_method']):
guides = self.options['guides_selection_method']
elif self.options['guides_selection_method'] == 'chi_squared_multiple':
multiple = self.options['guides_selection_kwargs']['multiple']
guides = self.pool.active_guides_above_multiple_of_max_nt(self.outcomes, multiple)
elif self.options['guides_selection_method'] == 'chi_squared_top_n':
n = self.options['guides_selection_kwargs']['n']
guides = self.pool.top_n_active_guides(self.outcomes, n,
use_high_frequency_counts=self.options['use_high_frequency_counts'],
)
else:
guides = self.pool.canonical_active_guides
return guides
@memoized_property
def log2_fold_changes(self):
if self.options['use_high_frequency_counts']:
l2fcs = self.pool.high_frequency_log2_fold_changes.loc[self.outcomes, self.guides]
else:
l2fcs = self.pool.log2_fold_changes().loc[self.outcomes, self.guides]
index = pd.MultiIndex.from_tuples([(self.pool.short_name, *vs) for vs in l2fcs.index])
index.names = ['pool_name'] + l2fcs.index.names
l2fcs.index = index
return l2fcs
def guide_log2_fold_changes(self, guide):
if self.options['use_high_frequency_counts']:
l2fcs = self.pool.high_frequency_log2_fold_changes.loc[self.outcomes, guide]
else:
l2fcs = self.pool.log2_fold_changes().loc[self.outcomes, guide]
index = pd.MultiIndex.from_tuples([(self.pool.short_name, *vs) for vs in l2fcs.index])
index.names = ['pool_name'] + l2fcs.index.names
l2fcs.index = index
return l2fcs
class MultiplePoolClusterer(Clusterer):
def __init__(self, pools, **options):
self.pools = pools
# Note: assumes all pools have the same guide library.
self.guide_library = self.pools[0].variable_guide_library
self.pn_to_pool = {pool.short_name: pool for pool in self.pools}
self.pn_to_target_info = {pool.short_name: pool.target_info for pool in self.pools}
self.pn_to_sgRNA = {pool.short_name: pool.target_info.sgRNA for pool in self.pools}
super().__init__(**options)
def sgRNA_colors(self):
outcome_colors, sgRNA_to_color = hits.visualize.assign_categorical_colors(self.outcome_embedding['sgRNA'])
return outcome_colors, sgRNA_to_color
@memoized_property
def common_guides(self):
return set.intersection(*[set(pool.variable_guide_library.guides) for pool in self.pools])
@property
def target_info(self):
return self.pools[0].target_info
@memoized_property
def guide_to_gene(self):
guide_to_gene = {}
for pool in self.pools:
guide_to_gene.update(pool.variable_guide_library.guide_to_gene)
return guide_to_gene
@memoized_property
def guides(self):
if pd.api.types.is_list_like(self.options['guides_selection_method']):
guides = self.options['guides_selection_method']
else:
active_guide_lists = defaultdict(list)
for pool in self.pools:
for i, guide in enumerate(pool.canonical_active_guides(use_high_frequency_counts=self.options['use_high_frequency_counts'])):
active_guide_lists[guide].append((pool.short_name, i))
# Include guides that were active in at least two screens.
# Only include up to 3 guides per gene, prioritized by average activity rank.
gene_guides = defaultdict(list)
for guide, active_pns in active_guide_lists.items():
if len(active_pns) >= min(len(self.pools), 2):
average_rank = np.mean([rank for pn, rank in active_pns])
gene_guides[self.guide_to_gene[guide]].append((average_rank, guide))
filtered_guides = []
for gene, guides in gene_guides.items():
guides = [g for r, g in sorted(guides)]
filtered_guides.extend(guides[:3])
guides = filtered_guides
return guides
@memoized_property
def pool_specific_outcomes(self):
selection_method = self.options['outcomes_selection_method']
pool_specific_outcomes = {}
for pool in self.pools:
if selection_method == 'category':
category = self.options['outcomes_selection_kwargs']['category']
min_f = self.options['outcomes_selection_kwargs']['threshold']
outcomes = [(c, s, d) for (c, s, d), f in pool.non_targeting_fractions.items() if c == category and f >= min_f]
elif selection_method == 'above_frequency_threshold':
threshold = self.options['outcomes_selection_kwargs']['threshold']
outcomes = pool.outcomes_above_simple_threshold(frequency_threshold=threshold,
use_high_frequency_counts=self.options['use_high_frequency_counts'],
)
elif selection_method == 'top_n':
num_outcomes = self.options['outcomes_selection_kwargs']['n']
outcomes = pool.most_frequent_outcomes('none')[:num_outcomes]
else:
outcomes = pool.canonical_outcomes
pool_specific_outcomes[pool.short_name] = outcomes
return pool_specific_outcomes
@memoized_property
def outcomes(self):
return [(c, s, d) for pn, c, s, d in self.outcomes_with_pool]
@memoized_property
def log2_fold_changes(self):
all_fcs = {}
for pool in self.pools:
if self.options['use_high_frequency_counts']:
fcs = pool.high_frequency_log2_fold_changes
else:
fcs = pool.log2_fold_changes()
all_fcs[pool.short_name] = fcs.loc[self.pool_specific_outcomes[pool.short_name], self.guides]
all_fcs = pd.concat(all_fcs)
all_fcs.index.names = ['pool_name'] + all_fcs.index.names[1:]
return all_fcs
def guide_log2_fold_changes(self, guide):
''' Look up log2 fold changes for a guide that may or may not be one of the guides
used for clustering.
'''
all_fcs = {}
for pool in self.pools:
if self.options['use_high_frequency_counts']:
fcs = pool.high_frequency_log2_fold_changes
else:
fcs = pool.log2_fold_changes()
all_fcs[pool.short_name] = fcs.loc[self.pool_specific_outcomes[pool.short_name], guide]
all_fcs = pd.concat(all_fcs)
all_fcs.index.names = ['pool_name'] + all_fcs.index.names[1:]
return all_fcs
def get_cluster_genes(results, guide_to_gene):
cluster_genes = defaultdict(Counter)
for cluster_id, guide in zip(results['cluster_assignments'], results['clustered_order']):
gene = guide_to_gene[guide]
cluster_genes[cluster_id][gene] += 1
return cluster_genes
def assign_palette(results, axis):
if 'cluster_assignments' in results:
num_clusters = len(set(results['cluster_assignments']))
if axis == 'guides':
palette = sns.husl_palette(num_clusters)
elif axis == 'outcomes':
palette = sns.color_palette('muted', n_colors=num_clusters)
else:
raise ValueError(axis)
results['palette'] = palette
grey = matplotlib.colors.to_rgb('silver')
cluster_colors = {i: palette[i] for i in range(num_clusters)}
results['cluster_colors'] = cluster_colors
results['colors'] = {key: cluster_colors.get(i, grey) for key, i in zip(results['clustered_order'], results['cluster_assignments'])} | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/cluster.py | 0.603465 | 0.208723 | cluster.py | pypi |
from collections import defaultdict
import numpy as np
import mpmath
import pandas as pd
import scipy.stats
import scipy.optimize
import scipy.special
n_choose_k = scipy.special.comb
def fit_beta_binomial(numerator_counts, denominator_counts):
p = numerator_counts.sum() / denominator_counts.sum()
def negative_log_likelihood(a):
b = a / p - a
count_pairs = zip(numerator_counts, denominator_counts)
return -sum(scipy.stats.betabinom.logpmf(n, d, a, b) for n, d in count_pairs)
results = scipy.optimize.minimize_scalar(negative_log_likelihood, bounds=(0, None))
if not results.success:
raise ValueError(results)
else:
alpha = results.x
beta = alpha / p - alpha
return alpha, beta
def beta_binomial_pvals(numerator, denominator, alpha, beta):
# Note: sf required to maintain precision.
# Since sf is P(> n), outcome_count - 1 required to produce P(>= n)
pvals = {
'down': scipy.stats.betabinom.cdf(numerator, denominator, alpha, beta),
'up': scipy.stats.betabinom.sf(numerator - 1, denominator, alpha, beta),
}
return pvals
def p_k_of_n_less(n, k, sorted_ps):
if k > n:
return 1
else:
a = sorted_ps[k - 1]
total = 0
for free_dimensions in range(0, n - k + 1):
total += n_choose_k(n, free_dimensions) * (1 - a)**free_dimensions * a**(n - free_dimensions)
return total
def compute_outcome_guide_statistics(pool, numerator_outcomes, denominator_outcomes=None):
numerator_counts, denominator_counts = extract_numerator_and_denominator_counts(pool, numerator_outcomes, denominator_outcomes)
frequencies = numerator_counts / denominator_counts
nt_guides = pool.variable_guide_library.non_targeting_guides
nt_numerator_counts = numerator_counts.loc[nt_guides]
nt_denominator_counts = denominator_counts.loc[nt_guides]
nt_fraction = nt_numerator_counts.sum() / nt_denominator_counts.sum()
alpha, beta = fit_beta_binomial(nt_numerator_counts, nt_denominator_counts)
ps = beta_binomial_pvals(numerator_counts, denominator_counts, alpha, beta)
genes = pool.variable_guide_library.guides_df['gene']
capped_fc = np.minimum(2**5, np.maximum(2**-5, frequencies / nt_fraction))
guides_df = pd.DataFrame({'denominator_count': denominator_counts,
'numerator_count': numerator_counts,
'frequency': frequencies,
'log2_fold_change': np.log2(capped_fc),
'p_down': ps['down'],
'p_up': ps['up'],
'gene': genes,
})
guides_df.index.name = 'guide'
recalculate_low_precision_pvals(guides_df, alpha, beta)
# Bonferonni factor of 2
guides_df['p_relevant'] = guides_df[['p_down', 'p_up']].min(axis=1) * 2
guides_df['-log10_p_relevant'] = -np.log10(np.maximum(np.finfo(np.float64).tiny, guides_df['p_relevant']))
return guides_df, nt_fraction
def recalculate_low_precision_pvals(guides_df, alpha, beta):
for guide, row in guides_df.iterrows():
if row['p_down'] < 1e-8:
B = scipy.stats.betabinom(row['denominator_count'], alpha, beta)
p_down = float(mpmath.fsum([mpmath.exp(B.logpmf(v)) for v in range(row['numerator_count'] + 1)]))
guides_df.loc[guide, 'p_down'] = p_down
if row['p_up'] < 1e-8:
B = scipy.stats.betabinom(row['denominator_count'], alpha, beta)
p_up = float(mpmath.fsum([mpmath.exp(B.logpmf(v)) for v in range(row['numerator_count'], row['denominator_count'])]))
guides_df.loc[guide, 'p_up'] = p_up
def compute_outcome_gene_statistics(guides_df):
ps = defaultdict(list)
max_k = 9
gene_order = []
for gene, rows in guides_df.groupby('gene'):
if gene == 'negative_control':
continue
gene_order.append(gene)
for direction in ('down', 'up'):
sorted_ps = sorted(rows[f'p_{direction}'].values)
n = len(sorted_ps)
for k in range(1, max_k + 1):
ps[direction, k].append(p_k_of_n_less(n, k, sorted_ps))
uncorrected_ps_df = pd.DataFrame(ps, index=gene_order).min(axis=1, level=0)
guides_per_gene = guides_df.groupby('gene').size()
bonferonni_factor = np.minimum(max_k, guides_per_gene)
corrected_ps_df = np.minimum(1, uncorrected_ps_df.multiply(bonferonni_factor, axis=0))
up_genes = corrected_ps_df.query('up < down')['up'].sort_values(ascending=False).index
grouped_fcs = guides_df.groupby('gene')['log2_fold_change']
gene_log2_fold_changes = pd.DataFrame({
'up': grouped_fcs.nlargest(2).mean(level=0),
'down': grouped_fcs.nsmallest(2).mean(level=0),
})
gene_log2_fold_changes['relevant'] = gene_log2_fold_changes['down']
gene_log2_fold_changes.loc[up_genes, 'relevant'] = gene_log2_fold_changes.loc[up_genes, 'up']
corrected_ps_df['relevant'] = corrected_ps_df['down']
corrected_ps_df.loc[up_genes, 'relevant'] = corrected_ps_df.loc[up_genes, 'up']
genes_df = pd.concat({'log2_fold_change': gene_log2_fold_changes, 'p': corrected_ps_df}, axis=1)
negative_log10_p = -np.log10(np.maximum(np.finfo(np.float64).tiny, genes_df['p']))
negative_log10_p = pd.concat({'-log10_p': negative_log10_p}, axis=1)
genes_df = pd.concat([genes_df, negative_log10_p], axis=1)
genes_df.index.name = 'gene'
return genes_df | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/beta_binomial.py | 0.766905 | 0.333775 | beta_binomial.py | pypi |
from collections import defaultdict
import numpy as np
import pandas as pd
import hits.utilities
memoized_with_key = hits.utilities.memoized_with_key
memoized_property = hits.utilities.memoized_property
class Bootstrapper():
def __init__(self, all_cells, guides):
self.all_cells = all_cells.copy()
self.guides = guides
self._guide_cell_bcs = {}
self.guide_UMI_fraction_bootstraps = defaultdict(list)
def compute_guide_UMI_fractions(self, guide, bootstrap=False):
cell_bcs = self.guide_cell_bcs(guide)
if bootstrap:
cell_bcs = np.random.choice(cell_bcs, size=len(cell_bcs), replace=True)
cells = self.all_cells[cell_bcs]
UMI_counts_per_gene = cells.X.A.sum(axis=0)
total_UMIs = cells.obs['num_UMIs'].sum()
UMI_fractions_per_gene = UMI_counts_per_gene / total_UMIs
return UMI_fractions_per_gene
@memoized_with_key
def guide_UMI_fractions(self, guide):
return self.compute_guide_UMI_fractions(guide)
@memoized_with_key
def guide_cell_bcs(self, guide):
if guide == 'non-targeting':
nt_guides = [g for g in guides if g.startswith('non-')]
cell_bcs = self.all_cells.obs[self.all_cells.obs['sgRNA_name'].isin(nt_guides)].index
else:
cell_bcs = self.all_cells.obs.query('sgRNA_name == @guide').index
return cell_bcs
@memoized_property
def guide_UMI_fractions_df(self):
fs = [self.guide_UMI_fractions(guide) for guide in self.guides]
return pd.DataFrame(fs, index=self.guides, columns=[ENSG_to_name[ensg] for ensg in ENSGs])
@memoized_property
def guide_knockdown_df(self):
return self.guide_UMI_fractions_df / self.guide_UMI_fractions('non-targeting')
def quantile_UMI_fractions_df(self, q):
fs = [self.quantile_guide_UMI_fractions(guide, q) for guide in self.guides]
return pd.DataFrame(fs, index=self.guides, columns=[ENSG_to_name[ensg] for ensg in ENSGs])
def quantile_knockdown_df(self, q):
return self.quantile_UMI_fractions_df(q) / self.guide_UMI_fractions('non-targeting')
def add_bootstrap(self, guide):
guide_fs = self.compute_guide_UMI_fractions(guide, bootstrap=True)
self.guide_UMI_fraction_bootstraps[guide].append(guide_fs)
def add_all_bootstraps(self, n):
for guide in progress(self.guides):
for i in range(n):
self.add_bootstrap(guide)
def quantile_guide_UMI_fractions(self, guide, q):
bootstraps = np.array(self.guide_UMI_fraction_bootstraps[guide])
return np.quantile(bootstraps, q, axis=0)
@memoized_property
def guide_knockdown(self):
kds = []
targeting_guides = []
for guide in self.guides:
gene = guides_df.loc[guide, 'gene']
if gene != 'negative_control':
kd = self.guide_knockdown_df.loc[guide, gene]
kds.append(kd)
targeting_guides.append(guide)
return pd.Series(kds, index=targeting_guides) | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/bootstrapper.py | 0.577019 | 0.295001 | bootstrapper.py | pypi |
import bisect
import multiprocessing
import pandas as pd
import pysam
import scipy.sparse
from hits import utilities
memoized_property = utilities.memoized_property
memoized_with_args = utilities.memoized_with_args
import knock_knock.outcome_record
from repair_seq.common_sequences import CommonSequenceSplitter
def run_stage(GroupClass, group_args, sample_name, stage):
group = GroupClass(*group_args)
if sample_name in group.sample_names:
# Important to do this branch first, since preprocssing happens before common sequence collection.
exp = group.sample_name_to_experiment(sample_name, no_progress=True)
elif sample_name in group.common_sequence_chunk_exp_names:
exp = group.common_sequence_chunk_exp_from_name(sample_name)
else:
raise ValueError(sample_name)
exp.process(stage=stage)
class ExperimentGroup:
def __init__(self):
self.fns = {
'common_sequences_dir': self.results_dir / 'common_sequences',
'common_sequence_outcomes': self.results_dir / 'common_sequences' / 'common_sequence_outcomes.txt',
'common_sequence_special_alignments': self.results_dir / 'common_sequences' / 'all_special_alignments.bam',
'total_outcome_counts': self.results_dir / 'total_outcome_counts.txt',
'outcome_counts': self.results_dir / 'outcome_counts.npz',
'genomic_insertion_length_distributions': self.results_dir / 'genomic_insertion_length_distribution.txt',
}
def process(self, num_processes):
with multiprocessing.Pool(num_processes) as pool:
print('preprocessing')
args = [(type(self), self.group_args, sample_name, 'preprocess') for sample_name in self.sample_names]
pool.starmap(run_stage, args)
self.make_common_sequences()
for stage in [
'align',
'categorize',
]:
print('common sequences', stage)
args = [(type(self), self.group_args, chunk_exp_name, stage) for chunk_exp_name in self.common_sequence_chunk_exp_names]
pool.starmap(run_stage, args)
self.merge_common_sequence_outcomes()
for stage in [
'align',
'categorize',
'visualize',
]:
print(stage)
args = [(type(self), self.group_args, sample_name, stage) for sample_name in self.sample_names]
pool.starmap(run_stage, args)
self.make_outcome_counts()
def make_common_sequences(self):
''' Identify all sequences that occur more than once across preprocessed
reads for all experiments in the group and write them into common sequences
experiments to be categorized.
'''
splitter = CommonSequenceSplitter(self)
description = 'Collecting common sequences'
exps = self.experiments(no_progress=True)
for exp in self.progress(exps, desc=description, total=self.num_experiments):
reads = exp.reads_by_type(self.preprocessed_read_type)
splitter.update_counts((read.seq for read in reads))
splitter.write_files()
@memoized_property
def common_sequence_chunk_exp_names(self):
''' Names of all common sequence chunk experiments. '''
return sorted([d.name for d in self.fns['common_sequences_dir'].iterdir() if d.is_dir()])
def common_sequence_chunk_exps(self):
''' Iterator over common sequence chunk experiments. '''
for chunk_name in self.common_sequence_chunk_exp_names:
yield self.common_sequence_chunk_exp_from_name(chunk_name)
@memoized_property
def common_names(self):
''' List of all names assigned to common sequence artificial reads. '''
return [outcome.query_name for outcome in self.common_sequence_outcomes]
@memoized_property
def common_sequence_outcomes(self):
outcomes = []
for exp in self.common_sequence_chunk_exps():
for outcome in exp.outcome_iter():
outcomes.append(outcome)
return outcomes
@memoized_property
def common_name_to_common_sequence(self):
name_to_seq = {}
for outcome in self.common_sequence_outcomes:
name_to_seq[outcome.query_name] = outcome.seq
return name_to_seq
@memoized_property
def common_sequence_to_common_name(self):
return utilities.reverse_dictionary(self.common_name_to_common_sequence)
@memoized_property
def common_name_to_special_alignment(self):
name_to_al = {}
if self.fns['common_sequence_special_alignments'].exists():
for al in pysam.AlignmentFile(self.fns['common_sequence_special_alignments']):
name_to_al[al.query_name] = al
return name_to_al
@memoized_property
def common_sequence_to_outcome(self):
common_sequence_to_outcome = {}
for outcome in self.common_sequence_outcomes:
common_sequence_to_outcome[outcome.seq] = outcome
return common_sequence_to_outcome
def merge_common_sequence_outcomes(self):
with self.fns['common_sequence_outcomes'].open('w') as fh:
for outcome in self.common_sequence_outcomes:
fh.write(f'{outcome}\n')
@memoized_property
def name_to_chunk(self):
chunks = list(self.common_sequence_chunk_exps())
starts = [int(chunk.sample_name.split('-')[0]) for chunk in chunks]
def name_to_chunk(name):
number = int(name.split('_')[0])
start_index = bisect.bisect(starts, number) - 1
chunk = chunks[start_index]
return chunk
return name_to_chunk
def get_common_seq_alignments(self, seq):
name = self.common_sequence_to_common_name[seq]
als = self.get_read_alignments(name)
return als
@memoized_property
def common_sequence_to_alignments(self):
common_sequence_to_alignments = {}
for chunk_exp in self.common_sequence_chunk_exps():
for common_name, als in chunk_exp.alignment_groups(read_type='nonredundant'):
seq = self.common_name_to_common_sequence[common_name]
common_sequence_to_alignments[seq] = als
return common_sequence_to_alignments
def get_read_alignments(self, name):
if isinstance(name, int):
name = self.common_names[name]
chunk = self.name_to_chunk(name)
als = chunk.get_read_alignments(name, read_type='nonredundant')
return als
def get_read_layout(self, name, **kwargs):
als = self.get_read_alignments(name)
l = self.categorizer(als, self.target_info, mode=self.layout_mode, error_corrected=False, **kwargs)
return l
def get_read_diagram(self, read_id, only_relevant=True, **diagram_kwargs):
layout = self.get_read_layout(read_id)
if only_relevant:
layout.categorize()
to_plot = layout.relevant_alignments
else:
to_plot = layout.alignments
for k, v in self.diagram_kwargs.items():
diagram_kwargs.setdefault(k, v)
diagram = knock_knock.visualize.ReadDiagram(to_plot, self.target_info, **diagram_kwargs)
return diagram
def make_outcome_counts(self):
all_counts = {}
description = 'Loading outcome counts'
items = self.progress(self.full_condition_to_experiment.items(), desc=description)
for condition, exp in items:
try:
all_counts[condition] = exp.outcome_counts
except (FileNotFoundError, pd.errors.EmptyDataError):
pass
all_outcomes = set()
for counts in all_counts.values():
all_outcomes.update(counts.index.values)
outcome_order = sorted(all_outcomes)
outcome_to_index = {outcome: i for i, outcome in enumerate(outcome_order)}
counts = scipy.sparse.dok_matrix((len(outcome_order), len(self.full_conditions)), dtype=int)
description = 'Combining outcome counts'
full_conditions = self.progress(self.full_conditions, desc=description)
for c_i, condition in enumerate(full_conditions):
if condition in all_counts:
for outcome, count in all_counts[condition].items():
o_i = outcome_to_index[outcome]
counts[o_i, c_i] = count
scipy.sparse.save_npz(self.fns['outcome_counts'], counts.tocoo())
df = pd.DataFrame(counts.toarray(),
columns=self.full_conditions,
index=pd.MultiIndex.from_tuples(outcome_order),
)
df.sum(axis=1).to_csv(self.fns['total_outcome_counts'], header=False)
## Collapse potentially equivalent outcomes together.
#collapsed = pd.concat({pg: collapse_categories(df.loc[pg]) for pg in [True, False] if pg in df.index.levels[0]})
#coo = scipy.sparse.coo_matrix(np.array(collapsed))
#scipy.sparse.save_npz(self.fns['collapsed_outcome_counts'], coo)
#collapsed.sum(axis=1).to_csv(self.fns['collapsed_total_outcome_counts'], header=False)
@memoized_with_args
def outcome_counts_df(self, collapsed):
if collapsed:
prefix = 'collapsed_'
else:
prefix = ''
key = prefix + 'outcome_counts'
sparse_counts = scipy.sparse.load_npz(self.fns[key])
df = pd.DataFrame(sparse_counts.toarray(),
index=self.total_outcome_counts(collapsed).index,
columns=pd.MultiIndex.from_tuples(self.full_conditions),
)
df.index.names = self.outcome_index_levels
df.columns.names = self.outcome_column_levels
return df
@memoized_with_args
def total_outcome_counts(self, collapsed):
if collapsed:
prefix = 'collapsed_'
else:
prefix = ''
key = prefix + 'total_outcome_counts'
return pd.read_csv(self.fns[key], header=None, index_col=list(range(len(self.outcome_index_levels))), na_filter=False)
@memoized_property
def genomic_insertion_length_distributions(self):
df = pd.read_csv(self.fns['genomic_insertion_length_distributions'], index_col=[0, 1, 2])
df.columns = [int(c) for c in df.columns]
return df
class CommonSequencesExperiment:
@property
def final_Outcome(self):
return knock_knock.outcome_record.CommonSequenceOutcomeRecord
@memoized_property
def results_dir(self):
return self.experiment_group.fns['common_sequences_dir'] / self.sample_name
@memoized_property
def seq_to_outcome(self):
return {}
@memoized_property
def seq_to_alignments(self):
return {}
@memoized_property
def names_with_common_seq(self):
return {}
@property
def preprocessed_read_type(self):
return 'nonredundant'
@property
def read_types_to_align(self):
return ['nonredundant']
def make_nonredundant_sequence_fastq(self):
pass | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/experiment_group.py | 0.536313 | 0.18743 | experiment_group.py | pypi |
import argparse
import gzip
import multiprocessing
import subprocess
import tempfile
from pathlib import Path
from collections import Counter
import anndata as ad
import numpy as np
import pandas as pd
import pysam
import scanpy as sc
import scipy.sparse
import yaml
from pomegranate import GeneralMixtureModel, PoissonDistribution, NormalDistribution
from hits import utilities, mapping_tools, fasta, fastq, bus, sam
from hits.utilities import memoized_property
import repair_seq.guide_library
def build_guide_index(guides_fn, index_dir):
''' index entries are in same orientation as R2 '''
index_dir = Path(index_dir)
index_dir.mkdir(exist_ok=True)
fasta_fn = index_dir / 'expected_R2s.fasta'
guides_df = pd.read_csv(guides_fn, sep='\t', index_col=0)
before_ps = 'AGTACCAAGTTGATAACGGACTAGCCTTATTTAAACTTGCTATGCTGTTTCCAGCTTAGCTCTTAAAC'
# Note: Cs here are from untemplated addition and are not deterministically 3.
after_ps = 'CCCATATAAGAAA'
with fasta_fn.open('w') as fh:
for name, protospacer in guides_df['protospacer'].items():
expected_R2 = before_ps + utilities.reverse_complement(protospacer) + after_ps
fh.write(str(fasta.Record(name, expected_R2)))
pysam.faidx(str(fasta_fn))
mapping_tools.build_STAR_index([fasta_fn], index_dir)
bustools_dir = index_dir / 'bustools_annotations'
bustools_dir.mkdir(exist_ok=True)
matrix_fn = bustools_dir / 'matrix.ec'
with matrix_fn.open('w') as fh:
for i, name in enumerate(guides_df.index):
fh.write(f'{i}\t{i}\n')
transcript_to_gene_fn = bustools_dir / 'transcripts_to_genes.txt'
with transcript_to_gene_fn.open('w') as fh:
for i, name in enumerate(guides_df.index):
fh.write(f'{name}\t{name}\t{name}\n')
transcripts_fn = bustools_dir / 'transcripts.txt'
with transcripts_fn.open('w') as fh:
for i, name in enumerate(guides_df.index):
fh.write(f'{name}\n')
def load_bustools_counts(prefix):
prefix = str(prefix)
data = sc.read_mtx(str(prefix) + '.mtx')
data.obs.index = pd.read_csv(prefix + '.barcodes.txt', header=None)[0].values
data.var.index = pd.read_csv(prefix + '.genes.txt', header=None)[0].values
return data
def fit_mixture_model(counts):
''' Code adapted from https://github.com/josephreplogle/guide_calling '''
data = np.log2(counts + 1)
reshaped_data = data.reshape(-1, 1)
xs = np.linspace(-2, max(data) + 2, 1000)
# Re-fit the model until it has converged with both components given non-zero weight
# and the Poisson component in the first position with lower mean.
while True:
model = GeneralMixtureModel.from_samples([PoissonDistribution, NormalDistribution], 2, reshaped_data)
if 0 in model.weights:
# One component was eliminated
continue
elif np.isnan(model.probability(xs)).any():
continue
elif model.distributions[0].parameters[0] > model.distributions[1].parameters[0]:
continue
elif model.distributions[0].name != 'PoissonDistribution':
continue
else:
break
labels = model.predict(reshaped_data)
xs = np.linspace(0, max(data) + 2, 1000)
p_second_component = model.predict_proba(xs.reshape(-1, 1))[:, 1]
threshold = 2**xs[np.argmax(p_second_component >= 0.5)]
return labels, threshold
class PerturbseqLane:
def __init__(self, base_dir, group, name):
self.base_dir = Path(base_dir)
self.group = group
self.name = name
self.data_dir = self.base_dir / 'data' / self.group
self.barcode_length = 16
self.UMI_length = 10
full_sample_sheet = load_sample_sheet(self.data_dir / 'sample_sheet.yaml')
sample_sheet = full_sample_sheet['lanes'][name]
self.output_dir = self.base_dir / 'results' / self.group / self.name
self.sgRNA_dir = self.output_dir / 'sgRNA'
self.GEX_dir = self.output_dir / 'GEX'
self.cellranger_dir = self.output_dir / 'cellranger_output'
self.guide_library = repair_seq.guide_library.GuideLibrary(self.base_dir, full_sample_sheet['guide_libary'])
self.guide_index = self.guide_library.fns['perturbseq_STAR_index']
self.whitelist_fn = Path(full_sample_sheet['whitelist_fn'])
self.sgRNA_fns = {
'dir': self.sgRNA_dir,
'R1_fns': [self.data_dir / fn for fn in sample_sheet['sgRNA_R1_fns']],
'R2_fns': [self.data_dir / fn for fn in sample_sheet['sgRNA_R2_fns']],
'STAR_output_prefix': self.sgRNA_dir / 'STAR' / 'sgRNA.',
'bam': self.sgRNA_dir / 'sgRNA.bam',
'bus': self.sgRNA_dir / 'sgRNA.bus',
'counts': self.sgRNA_dir / 'counts',
'genemap': self.guide_index / 'bustools_annotations' / 'transcripts_to_genes.txt',
'ecmap': self.guide_index / 'bustools_annotations' / 'matrix.ec',
'txnames': self.guide_index / 'bustools_annotations' / 'transcripts.txt',
}
self.GEX_fns = {
'dir': self.GEX_dir,
'R1_fns': [self.data_dir / fn for fn in sample_sheet['GEX_R1_fns']],
'R2_fns': [self.data_dir / fn for fn in sample_sheet['GEX_R2_fns']],
'bus': self.GEX_dir / 'output.bus',
'counts': self.GEX_dir / 'counts',
'kallisto_index': Path(full_sample_sheet['kallisto_index']),
'genemap': Path(full_sample_sheet['kallisto_genemap']),
'ecmap': self.GEX_dir / 'matrix.ec',
'txnames': self.GEX_dir / 'transcripts.txt',
'cellranger_filtered_feature_bc_matrix_dir': self.cellranger_dir / 'filtered_feature_bc_matrix',
'cellranger_barcodes': self.cellranger_dir / 'filtered_feature_bc_matrix' / 'barcodes.tsv.gz',
'sgRNA_counts_list': self.cellranger_dir / 'sgRNA_counts_list.csv.gz',
'sgRNA_counts_csv': self.cellranger_dir / 'sgRNA_counts.csv.gz',
'sgRNA_counts_h5ad': self.cellranger_dir / 'sgRNA_counts.h5ad',
}
self.fns = {
'annotated_counts': self.output_dir / 'counts.h5ad',
}
missing_files = []
files_to_check = [
self.sgRNA_fns['R1_fns'],
self.sgRNA_fns['R2_fns'],
self.GEX_fns['R1_fns'],
self.GEX_fns['R2_fns'],
]
for fns in files_to_check:
for fn in fns:
if not fn.exists():
missing_files.append(fn)
if missing_files:
print(f'{self.name} specifies non-existent files: {[str(fn) for fn in missing_files]}')
def map_sgRNA_reads(self):
output_prefix = self.sgRNA_fns['STAR_output_prefix']
output_prefix.parent.mkdir(parents=True, exist_ok=True)
mapping_tools.map_STAR(self.sgRNA_fns['R2_fns'],
self.guide_index,
output_prefix,
mode='guide_alignment',
sort=False,
include_unmapped=True,
num_threads=1,
bam_fn=self.sgRNA_fns['bam'],
)
def convert_sgRNA_bam_to_bus(self):
barcode_length = self.barcode_length
UMI_length = self.UMI_length
R1s = fastq.reads(self.sgRNA_fns['R1_fns'], up_to_space=True)
R2_alignment_groups = sam.grouped_by_name(self.sgRNA_fns['bam'])
with self.sgRNA_fns['bus'].open('wb') as fh:
bus.write_header(fh, self.barcode_length, self.UMI_length)
for (qname, als), R1 in zip(R2_alignment_groups, R1s):
if qname != R1.name:
raise ValueError(qname, R1.name)
ref_ids = {al.reference_id for al in als if not al.is_unmapped}
if len(ref_ids) == 1:
barcode = R1.seq[:self.barcode_length]
UMI = R1.seq[barcode_length:barcode_length + UMI_length]
if 'N' in barcode or 'N' in UMI:
continue
record = bus.Record(barcode, UMI, ref_ids.pop(), 1, 0)
fh.write(record.pack())
def convert_bus_to_counts(self, fns):
with tempfile.TemporaryDirectory(prefix=str(fns['dir'] / 'tmp')) as temp_dir_name:
correct_command = [
'bustools', 'correct',
'--whitelist', str(self.whitelist_fn),
'--pipe',
str(fns['bus']),
]
correct_process = subprocess.Popen(correct_command, stdout=subprocess.PIPE, stderr=subprocess.DEVNULL)
sort_command = [
'bustools', 'sort',
'--temp', temp_dir_name,
'--pipe',
'-',
]
sort_process = subprocess.Popen(sort_command, stdin=correct_process.stdout, stdout=subprocess.PIPE, stderr=subprocess.DEVNULL)
correct_process.stdout.close()
count_command = [
'bustools', 'count',
'--output', str(fns['counts']),
'--genemap', str(fns['genemap']),
'--ecmap', str(fns['ecmap']),
'--txnames', str(fns['txnames']),
'--genecounts',
'-',
]
count_process = subprocess.Popen(count_command, stdin=sort_process.stdout, stdout=subprocess.PIPE, stderr=subprocess.DEVNULL)
sort_process.stdout.close()
count_process.communicate()
def pseudoalign_GEX_reads(self):
self.GEX_dir.mkdir(parents=True, exist_ok=True)
fastq_fns = []
for R1_fn, R2_fn in zip(self.GEX_fns['R1_fns'], self.GEX_fns['R2_fns']):
fastq_fns.extend([str(R1_fn), str(R2_fn)])
kallisto_command = [
'kallisto', 'bus',
'-i', str(self.GEX_fns['kallisto_index']),
'-x', '10xv2',
'-o', str(self.GEX_dir),
]
kallisto_command.extend(fastq_fns)
try:
subprocess.run(kallisto_command, check=True, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
except subprocess.CalledProcessError as e:
for line in e.stdout.splitlines():
print(line.decode())
for line in e.stderr.splitlines():
print(line.decode())
raise
@memoized_property
def sgRNA_data(self):
return load_bustools_counts(self.sgRNA_fns['counts'])
@memoized_property
def GEX_data(self):
return load_bustools_counts(self.GEX_fns['counts'])
@memoized_property
def ENSG_to_name(self):
names_fn = '/lab/solexa_weissman/indices/refdata-cellranger-hg19-1.2.0/kallisto/transcripts_to_genes_hg19.txt'
updated_names_fn = self.base_dir / 'guides' / 'DDR_library' / 'updated_gene_names.txt'
updated_names = pd.read_csv(updated_names_fn, sep='\t', index_col='old_name', squeeze=True)
ENSG_to_name = {}
names_seen = Counter()
for line in open(names_fn):
ENST, ENSG, name = line.strip().split()
if ENSG in ENSG_to_name:
continue
names_seen[name] += 1
if name in updated_names:
name = updated_names[name]
if names_seen[name] > 1:
name_to_use = f'{name}_{names_seen[name]}'
else:
name_to_use = name
ENSG_to_name[ENSG] = name_to_use
ENSG_to_name['negative_control'] = 'negative_control'
return pd.Series(ENSG_to_name)
@memoized_property
def name_to_ENSG(self):
return pd.Series(utilities.reverse_dictionary(self.ENSG_to_name))
def combine_sgRNA_and_GEX_counts(self):
gex_data = self.GEX_data
gex_data.var['name'] = [self.ENSG_to_name[g] for g in gex_data.var.index.values]
sgRNA_data = self.sgRNA_data
gex_data.obs['num_UMIs'] = np.sum(gex_data.X, axis=1).A1
sgRNA_data.obs['num_UMIs'] = np.sum(sgRNA_data.X, axis=1).A1
sgRNA_data.obs['highest_count'] = sgRNA_data.X.max(axis=1).todense().A1
sgRNA_data.obs['highest_index'] = sgRNA_data.X.argmax(axis=1).A1
sgRNA_data.obs['fraction_highest'] = sgRNA_data.obs['highest_count'] / sgRNA_data.obs['num_UMIs']
gex_cellBCs = gex_data.obs_names
gex_data.obs['sgRNA_highest_index'] = sgRNA_data.obs['highest_index'].reindex(gex_cellBCs, fill_value=-1).astype(int)
gex_data.obs['sgRNA_highest_count'] = sgRNA_data.obs['highest_count'].reindex(gex_cellBCs, fill_value=0).astype(int)
gex_data.obs['sgRNA_fraction_highest'] = sgRNA_data.obs['fraction_highest'].reindex(gex_cellBCs, fill_value=0)
gex_data.obs['sgRNA_num_UMIs'] = sgRNA_data.obs['num_UMIs'].reindex(gex_cellBCs, fill_value=0).astype(int)
gex_data.obs['sgRNA_name'] = [sgRNA_data.var_names[i] if i != -1 else 'none' for i in gex_data.obs['sgRNA_highest_index']]
# For performance reasons, go ahead and discard any BCs with < 1000 UMIs.
gex_data = gex_data[gex_data.obs.query('num_UMIs >= 1000').index]
valid_cellBCs = gex_data.obs.query('num_UMIs > 5e3').index
guide_calls, num_guides = self.fit_guide_count_mixture_models(valid_cellBCs)
gex_data.obs['MM_guide_call'] = guide_calls.reindex(gex_cellBCs, fill_value='none')
gex_data.obs['MM_num_guides'] = num_guides.reindex(gex_cellBCs, fill_value=-1).astype(int)
gex_data.write(self.fns['annotated_counts'])
def fit_guide_count_mixture_models(self, valid_cellBCs):
sgRNA_data = self.sgRNA_data[valid_cellBCs]
guides_present = np.zeros(sgRNA_data.X.shape)
for g, guide in enumerate(sgRNA_data.var.index):
labels, _ = fit_mixture_model(sgRNA_data.obs_vector(guide))
guides_present[:, g] = labels
guide_calls = sgRNA_data.var_names[guides_present.argmax(axis=1)].values
num_guides = guides_present.sum(axis=1)
guide_calls = pd.Series(guide_calls, index=sgRNA_data.obs_names)
num_guides = pd.Series(num_guides, index=sgRNA_data.obs_names)
return guide_calls, num_guides
@memoized_property
def cellranger_barcodes(self):
bcs = []
with gzip.open(self.GEX_fns['cellranger_barcodes'], 'rt') as fh:
for line in fh:
bcs.append(line.strip())
return pd.Index(bcs)
def make_guide_count_tables(self):
cells_with_dummy_lane = [f'{cell_bc}-1' for cell_bc in self.sgRNA_data.obs_names]
self.sgRNA_data.obs.index = cells_with_dummy_lane
cells_in_both = self.cellranger_barcodes.intersection(self.sgRNA_data.obs_names)
sgRNA_data = self.sgRNA_data[cells_in_both]
sgRNA_data.write(self.GEX_fns['sgRNA_counts_h5ad'])
df = sgRNA_data.to_df().astype(int)
df.index.name = 'cell_barcode'
df.columns.name = 'guide_identity'
df.to_csv(self.GEX_fns['sgRNA_counts_csv'])
stacked = df.stack()
stacked.name = 'UMI_count'
stacked.index.names = ('cell_barcode', 'guide_identity')
stacked.to_csv(self.GEX_fns['sgRNA_counts_list'])
@memoized_property
def annotated_counts(self):
return sc.read_h5ad(self.fns['annotated_counts'])
def process(self):
#self.map_sgRNA_reads()
#self.convert_sgRNA_bam_to_bus()
#self.convert_bus_to_counts(self.sgRNA_fns)
#self.pseudoalign_GEX_reads()
#self.convert_bus_to_counts(self.GEX_fns)
#self.combine_sgRNA_and_GEX_counts()
self.make_guide_count_tables()
def load_sample_sheet(sample_sheet_fn):
sample_sheet = yaml.safe_load(Path(sample_sheet_fn).read_text())
return sample_sheet
class MultipleLanes:
def __init__(self, base_dir, group):
self.base_dir = Path(base_dir)
self.group = group
sample_sheet_fn = self.base_dir / 'data' / group / 'sample_sheet.yaml'
full_sample_sheet = load_sample_sheet(sample_sheet_fn)
self.guide_library = repair_seq.guide_library.GuideLibrary(self.base_dir, full_sample_sheet['guide_libary'])
self.lanes = [PerturbseqLane(self.base_dir, self.group, name) for name in full_sample_sheet['lanes']]
self.results_dir = self.base_dir / 'results' / self.group
self.cellranger_dir = self.results_dir / 'cellranger_aggregated' / 'outs'
self.fns = {
'all_cells': self.results_dir / 'all_cells.h5ad',
}
self.GEX_fns = {
'cellranger_filtered_feature_bc_matrix_dir': self.cellranger_dir / 'filtered_feature_bc_matrix',
'cellranger_barcodes': self.cellranger_dir / 'filtered_feature_bc_matrix' / 'barcodes.tsv.gz',
'sgRNA_counts_list': self.cellranger_dir / 'sgRNA_counts_list.csv.gz',
'sgRNA_counts_csv': self.cellranger_dir / 'sgRNA_counts.csv.gz',
'sgRNA_counts_h5ad': self.cellranger_dir / 'sgRNA_counts.h5ad',
'guide_assignments': self.cellranger_dir / 'guide_assignments.csv.gz',
}
@memoized_property
def ENSG_to_name(self):
return self.lanes[0].ENSG_to_name
@memoized_property
def name_to_ENSG(self):
return self.lanes[0].name_to_ENSG
def combine_counts(self):
all_cells = {}
for lane in self.lanes:
gex_data = lane.annotated_counts
#good_cell_query = 'num_UMIs > 0.5e4 and sgRNA_num_UMIs > 1e2 and sgRNA_highest_count / sgRNA_num_UMIs > 0.9'
cells = gex_data.obs.index
cells_with_lane = [f'{cell_bc}-{lane.name[-1]}' for cell_bc in cells]
all_cells[lane.name] = gex_data
all_cells[lane.name].obs.index = cells_with_lane
all_Xs = scipy.sparse.vstack([all_cells[name].X for name in sorted(all_cells)])
all_obs = pd.concat([all_cells[name].obs for name in sorted(all_cells)])
all_var = all_cells[sorted(all_cells)[0]].var
all_cells = sc.AnnData(all_Xs, all_obs, all_var)
all_cells.write(self.fns['all_cells'])
def make_guide_count_tables(self):
all_sgRNA_counts = []
for lane in self.lanes:
sgRNA_counts = sc.read_h5ad(lane.GEX_fns['sgRNA_counts_h5ad'])
lane_num = lane.name[-1]
sgRNA_counts.obs.index = [f'{cell_bc.rsplit("-", 1)[0]}-{lane_num}' for cell_bc in sgRNA_counts.obs_names]
all_sgRNA_counts.append(sgRNA_counts)
sgRNA_data = ad.concat(all_sgRNA_counts)
sgRNA_data.write(self.GEX_fns['sgRNA_counts_h5ad'])
df = sgRNA_data.to_df().astype(int)
df.index.name = 'cell_barcode'
df.columns.name = 'guide_identity'
df.to_csv(self.GEX_fns['sgRNA_counts_csv'])
stacked = df.stack()
stacked.name = 'UMI_count'
stacked.index.names = ('cell_barcode', 'guide_identity')
stacked.to_csv(self.GEX_fns['sgRNA_counts_list'])
@memoized_property
def cells(self):
return sc.read_h5ad(self.fns['all_cells'])
def process_in_pool(lane):
lane.process()
def parallel(lanes, max_procs):
pool = multiprocessing.Pool(processes=max_procs)
pool.map(process_in_pool, lanes.lanes)
#lanes.combine_counts()
if __name__ == '__main__':
parser = argparse.ArgumentParser()
parser.add_argument('--base_dir', type=Path, required=True)
parser.add_argument('--group', type=Path, required=True)
parser.add_argument('--max_procs', type=int, required=True)
args = parser.parse_args()
lanes = MultipleLanes(args.base_dir, args.group)
parallel(lanes, args.max_procs) | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/perturb_seq.py | 0.517083 | 0.290503 | perturb_seq.py | pypi |
import numpy as np
import pandas as pd
def extract_numerator_and_denominator_counts(pool, numerator_outcomes, denominator_outcomes=None):
if all(outcome in pool.category_counts.index for outcome in list(numerator_outcomes)):
count_source = pool.category_counts
elif all(outcome in pool.subcategory_counts.index for outcome in list(numerator_outcomes)):
count_source = pool.subcategory_counts
else:
count_source = pool.outcome_counts('perfect')['none']
numerator_counts = count_source.loc[numerator_outcomes].sum(axis='index')
if denominator_outcomes is None:
denominator_counts = pool.UMI_counts
else:
if not all(outcome in count_source.index for outcome in denominator_outcomes):
raise ValueError(denominator_outcomes)
denominator_counts = count_source.loc[denominator_outcomes].sum(axis='index')
# Make sure neither counts has 'all_non_targeting' in index.
for counts in [numerator_counts, denominator_counts]:
counts.drop(['all_non_targeting', 'eGFP_NT2'], errors='ignore', inplace=True)
return numerator_counts, denominator_counts
def compute_outcome_guide_statistics(pool, numerator_outcomes, denominator_outcomes=None):
numerator_counts, denominator_counts = extract_numerator_and_denominator_counts(pool, numerator_outcomes, denominator_outcomes)
frequencies = numerator_counts / denominator_counts
nt_guides = pool.variable_guide_library.non_targeting_guides
nt_numerator_counts = numerator_counts.loc[nt_guides]
nt_denominator_counts = denominator_counts.loc[nt_guides]
nt_fraction = nt_numerator_counts.sum() / nt_denominator_counts.sum()
capped_fc = np.minimum(2**5, np.maximum(2**-5, frequencies / nt_fraction))
guides_df = pd.DataFrame({
'denominator_count': denominator_counts,
'numerator_count': numerator_counts,
'frequency': frequencies,
'log2_fold_change': np.log2(capped_fc),
'gene': pool.variable_guide_library.guides_df['gene'],
'best_promoter': pool.variable_guide_library.guides_df['best_promoter'],
})
guides_df['non-targeting'] = guides_df.index.isin(pool.variable_guide_library.non_targeting_guides)
guides_df.index.name = 'guide'
for set_name, guides in pool.variable_guide_library.non_targeting_guide_sets.items():
guides_df.loc[guides, 'gene'] = set_name
guides_df.drop('eGFP_NT2', errors='ignore', inplace=True)
return guides_df, nt_fraction
def convert_to_gene_statistics(guides_df, only_best_promoters=True):
gene_order = []
for gene, rows in guides_df.groupby('gene'):
gene_order.append(gene)
all_means = {}
for gene, rows in guides_df.query('best_promoter').groupby('gene')['log2_fold_change']:
means = {}
ordered = sorted(rows)
ordered_by_abs = sorted(rows, key=np.abs, reverse=True)
for n in [1, 2, 3]:
means[f'lowest_{n}'] = np.mean(ordered[:n])
means[f'highest_{n}'] = np.mean(ordered[-n:])
means[f'extreme_{n}'] = np.mean(ordered_by_abs[:n])
all_means[gene] = means
genes_df = pd.DataFrame(all_means).T
genes_df.index.name = 'gene'
return genes_df | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/statistics.py | 0.5 | 0.294836 | statistics.py | pypi |
import bisect
from collections import defaultdict
import matplotlib.pyplot as plt
import matplotlib.colors
import numpy as np
import pandas as pd
import scipy.stats
import seaborn as sns
from scipy.stats.stats import pearsonr
import hits.utilities
memoized_property = hits.utilities.memoized_property
import knock_knock.outcome
from . import visualize
from .visualize import outcome_diagrams, clustermap
from . import pooled_screen
ALL_NON_TARGETING = pooled_screen.ALL_NON_TARGETING
class ReplicatePair:
def __init__(self, pools, pn_pair, use_high_frequency_counts=False):
self.pools = pools
self.pn_pair = pn_pair
self.pn0 = pn_pair[0]
self.pn1 = pn_pair[1]
self.pool0 = self.pools[pn_pair[0]]
self.pool1 = self.pools[pn_pair[1]]
self.target_info = self.pool0.target_info
self.use_high_frequency_counts = use_high_frequency_counts
@memoized_property
def average_nt_fractions(self):
if self.use_high_frequency_counts:
nt_fracs = {pn: self.pools[pn].high_frequency_outcome_fractions[ALL_NON_TARGETING] for pn in self.pn_pair}
else:
nt_fracs = {pn: self.pools[pn].non_targeting_fractions() for pn in self.pn_pair}
return pd.concat(nt_fracs, axis=1).fillna(0).mean(axis=1).sort_values(ascending=False)
@memoized_property
def common_guides(self):
return sorted(set(self.pool0.variable_guide_library.guides) & set(self.pool1.variable_guide_library.guides))
@memoized_property
def common_guides_df(self):
return self.pool0.variable_guide_library.guides_df.loc[self.common_guides]
@memoized_property
def common_non_targeting_guides(self):
return self.common_guides_df.query('gene == "negative_control"').index
def outcomes_above_simple_threshold(self, threshold):
return [(c, s, d) for (c, s, d), f in self.average_nt_fractions.items() if f >= threshold
and c not in ['uncategorized', 'genomic insertion']
]
def union_of_top_n_guides(self, outcomes, n):
all_guides = set()
for pn in self.pn_pair:
pool = self.pools[pn]
top_guides = pool.top_n_active_guides(outcomes, n, use_high_frequency_counts=self.use_high_frequency_counts)
all_guides.update(top_guides)
# Only consider guides that were present in both screens.
all_guides = all_guides & set(self.common_guides)
all_guides = sorted(all_guides)
return all_guides
def log2_fold_changes(self, outcomes, guides):
if self.use_high_frequency_counts:
data = {pn: self.pools[pn].high_frequency_log2_fold_changes.loc[outcomes, guides] for pn in self.pn_pair}
else:
data = {pn: self.pools[pn].log2_fold_changes().loc[outcomes, guides] for pn in self.pn_pair}
return pd.concat(data, axis=1)
def outcome_r_matrix(self, outcomes, guides):
log2_fold_changes = self.log2_fold_changes(outcomes, guides)
r_matrix = np.zeros((len(outcomes), len(outcomes)))
for i in range(len(outcomes)):
row = log2_fold_changes.iloc[i]
r, p = scipy.stats.pearsonr(row[self.pn0], row[self.pn1])
r_matrix[i, i] = r
highest_rs = []
for i in range(len(outcomes)):
row_i = log2_fold_changes.iloc[i]
for j in range(i + 1, len(outcomes)):
row_j = log2_fold_changes.iloc[j]
r, p = scipy.stats.pearsonr(row_i[self.pn0], row_j[self.pn0])
r_matrix[i, j] = r
highest_rs.append((r, i, j, log2_fold_changes.index.values[i], log2_fold_changes.index.values[j]))
r, p = scipy.stats.pearsonr(row_i[self.pn1], row_j[self.pn1])
r_matrix[j, i] = r
highest_rs = sorted(highest_rs, reverse=True)
return r_matrix, highest_rs
def outcome_r_series(self, guides=None):
if guides is None:
pool = self.pools[self.pn0]
guides = pool.variable_guide_library.guides
outcomes = self.average_nt_fractions.iloc[:100].index.values
log2_fold_changes = self.log2_fold_changes(outcomes, guides)
rs = []
ps = []
for outcome in outcomes:
row = log2_fold_changes.loc[outcome]
r, p = scipy.stats.pearsonr(row[self.pn0], row[self.pn1])
rs.append(r)
ps.append(p)
return pd.Series(rs, index=pd.MultiIndex.from_tuples(outcomes))
def guide_r_matrix(self, outcomes, guides):
log2_fold_changes = self.log2_fold_changes(outcomes, guides)
r_matrix = np.zeros((len(guides), len(guides)))
for i, guide in enumerate(guides):
cols = log2_fold_changes.xs(guide, axis=1, level=1)
r, p = scipy.stats.pearsonr(cols[self.pn0], cols[self.pn1])
r_matrix[i, i] = r
highest_rs = []
for i, guide_i in enumerate(guides):
cols_i = log2_fold_changes.xs(guide_i, axis=1, level=1)
for j in range(i + 1, len(guides)):
guide_j = guides[j]
cols_j = log2_fold_changes.xs(guide_j, axis=1, level=1)
r, p = scipy.stats.pearsonr(cols_i[self.pn0], cols_j[self.pn0])
r_matrix[i, j] = r
highest_rs.append((r, guides[i], guides[j]))
r, p = scipy.stats.pearsonr(cols_i[self.pn1], cols_j[self.pn1])
r_matrix[j, i] = r
highest_rs = sorted(highest_rs, reverse=True)
return pd.DataFrame(r_matrix, index=guides, columns=guides), highest_rs
def guide_r_series(self, outcomes):
guides = self.common_guides
log2_fold_changes = self.log2_fold_changes(outcomes, guides)
rs = []
ps = []
for guide in guides:
cols = log2_fold_changes.xs(guide, axis=1, level=1)
r, p = scipy.stats.pearsonr(cols[self.pn0], cols[self.pn1])
rs.append(r)
ps.append(p)
return pd.Series(rs, index=guides, name='r').sort_values(ascending=False)
def plot_outcome_diagrams_with_correlations(self,
threshold=5e-3,
n_guides=100,
window=(-20, 20),
draw_heatmap=False,
):
outcomes = self.outcomes_above_simple_threshold(threshold)
guides = self.union_of_top_n_guides(outcomes, n_guides)
r_series = self.outcome_r_series(guides=guides)
inches_per_nt = 0.1
text_size = 7
inches_per_outcome = 0.2
scale = 0.5
g = outcome_diagrams.DiagramGrid(outcomes,
self.target_info,
window=window,
cut_color='PAM',
draw_wild_type_on_top=True,
flip_if_reverse=False,
inches_per_nt=inches_per_nt * scale,
text_size=text_size * scale,
inches_per_outcome=inches_per_outcome * scale,
line_widths=0.75,
title=None,
block_alpha=0.2,
)
g.add_ax('frequency',
width_multiple=10,
title='Baseline % of\noutcomes in cells\nwith non-targeting\nCRISPRi sgRNAs',
title_size=7,
gap_multiple=2,
)
g.plot_on_ax('frequency', self.average_nt_fractions,
transform=lambda f: f * 100,
marker='o',
color='black',
clip_on=False,
markersize=1,
line_alpha=0.5,
)
plt.setp(g.axs_by_name['frequency'].get_xticklabels(), size=6)
g.axs_by_name['frequency'].tick_params(axis='x', which='both', pad=2)
g.add_ax('correlation',
gap_multiple=3,
width_multiple=10,
title='Correlation between\nreplicates in log$_2$ fold\nchanges in frequencies\nof each outcome\nacross active\nCRISPRi sgRNAs',
title_size=7,
)
norm = matplotlib.colors.Normalize(vmin=-1, vmax=1)
sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=visualize.correlation_cmap)
g.plot_on_ax('correlation',
r_series,
marker=None,
color=sm.to_rgba(0.5),
clip_on=False,
markersize=3,
line_alpha=0.5,
)
rs = r_series.loc[outcomes][::-1]
colors = [tuple(row) for row in sm.to_rgba(rs)]
g.axs_by_name['correlation'].scatter(rs, np.arange(len(rs)),
color=colors,
linewidths=(0,),
s=6,
clip_on=False,
zorder=10,
)
g.axs_by_name['correlation'].set_xlim(-1, 1)
g.axs_by_name['correlation'].set_xticks([-1, -0.5, 0, 0.5, 1])
g.axs_by_name['correlation'].set_xticks([-0.75, -0.25, 0.25, 0.75], minor=True)
g.axs_by_name['correlation'].tick_params(axis='x', which='both', pad=2)
g.style_fold_change_ax('correlation')
g.axs_by_name['correlation'].axvline(0, color='black', alpha=0.5)
plt.setp(g.axs_by_name['correlation'].get_xticklabels(), size=6)
if draw_heatmap:
r_matrix, _ = self.outcome_r_matrix(outcomes, guides)
# Note reversing of row order.
g.add_heatmap(r_matrix[::-1], 'r',
cmap=visualize.correlation_cmap,
draw_tick_labels=False,
vmin=-1, vmax=1,
gap_multiple=2,
)
g.draw_outcome_categories()
return g.fig
def plot_guide_r_matrix(self, threshold=5e-3, n_guides=100, only_genes_with_multiple=True):
outcomes = self.outcomes_above_simple_threshold(threshold)
guides = self.union_of_top_n_guides(outcomes, n_guides)
if only_genes_with_multiple:
gene_counts = self.common_guides_df.loc[guides, 'gene'].value_counts()
genes_with_multiple = set(gene_counts[gene_counts > 1].index)
guides = [guide for guide in guides if self.common_guides_df.loc[guide, 'gene'] in genes_with_multiple]
guide_r_matrix, _ = self.guide_r_matrix(outcomes, guides)
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(guide_r_matrix, cmap=clustermap.correlation_cmap, vmin=-1, vmax=1)
ax.set_xticks([])
ax.set_yticks([])
plt.setp(ax.spines.values(), visible=False)
title = '''\
correlations between guides
diagonal: reproducibility between replicates (correlation of guide i in rep 1 with guide i in rep 2)
above diagonal: correlations between distinct guides in rep 1
below diagonal: correlations between distinct guides in rep 2
only genes with more than one active guide are included so that blocks on diagonal show within-gene consistency
'''
ax.set_title(title)
return fig, guides
def guide_guide_correlation_reproducibility(self, threshold=5e-3, n_guides=100):
outcomes = self.outcomes_above_simple_threshold(threshold)
guides = self.union_of_top_n_guides(outcomes, n_guides)
guide_r_matrix, _ = self.guide_r_matrix(outcomes, guides)
upper_t = np.triu_indices_from(guide_r_matrix, 1)
fig, ax = plt.subplots(figsize=(6, 6))
ax.scatter(guide_r_matrix.values[upper_t], guide_r_matrix.T.values[upper_t], s=20, linewidths=(0,), alpha=0.5)
kwargs = dict(color='black', alpha=0.3)
ax.axhline(0, **kwargs)
ax.axvline(0, **kwargs)
ax.plot([-1, 1], [-1, 1], **kwargs)
ax.set_ylabel('guide-guide correlation in replicate 2')
ax.set_xlabel('guide-guide correlation in replicate 1')
return fig
def outcome_outcome_correlation_scatter(self, threshold=5e-3, n_guides=100, rasterize=False):
threshold = 5e-3
outcomes = self.outcomes_above_simple_threshold(threshold)
guides = self.union_of_top_n_guides(outcomes, n_guides)
log2_fold_changes = self.log2_fold_changes(outcomes, guides)
rs = {}
for pn in self.pn_pair:
all_corrs = log2_fold_changes[pn].T.corr()
all_corrs.index = np.arange(len(all_corrs))
all_corrs.columns = np.arange(len(all_corrs))
rs[pn] = all_corrs.stack()
rs = pd.DataFrame(rs)
rs.index.names = ['outcome_1', 'outcome_2']
for i in [1, 2]:
k = f'outcome_{i}'
rs[k] = rs.index.get_level_values(k)
full_categories = self.full_categories(outcomes)
cat_to_highlight = 'deletion, bidirectional'
relevant_indices = [outcomes.index(outcome) for outcome in full_categories[cat_to_highlight]]
df = rs.query('outcome_1 < outcome_2').copy()
df['color'] = 'grey'
to_highlight = df['outcome_1'].isin(relevant_indices) & df['outcome_2'].isin(relevant_indices)
df.loc[to_highlight, 'color'] = visualize.Cas9_category_colors[cat_to_highlight]
x = self.pn0
y = self.pn1
fig, ax = plt.subplots(figsize=(1.25, 1.25))
kwargs = dict(x=x, y=y, color='color', linewidths=(0,), rasterized=rasterize)
ax.scatter(data=df.query('color == "grey"'), label='', alpha=0.4, s=12, **kwargs)
ax.scatter(data=df.query('color != "grey"'), label='bidirectional deletions', s=15, **kwargs)
ax.annotate('pairs of\nbidirectional\ndeletions',
xy=(0, 1),
xycoords='axes fraction',
xytext=(2, -1),
textcoords='offset points',
ha='left',
va='top',
color=visualize.Cas9_category_colors[cat_to_highlight],
size=6,
)
ax.annotate('all other\npairs',
xy=(0, 1),
xycoords='axes fraction',
xytext=(2, -21),
textcoords='offset points',
ha='left',
va='top',
color='grey',
size=6,
)
kwargs = dict(color='black', alpha=0.3, linewidth=0.75)
ax.axhline(0, **kwargs)
ax.axvline(0, **kwargs)
ax.plot([-1, 1], [-1, 1], **kwargs)
ticks = [-1, -0.5, 0, 0.5, 1]
ax.set_xticks(ticks)
ax.set_yticks(ticks)
plt.setp(ax.spines.values(), linewidth=0.5)
ax.tick_params(labelsize=6, width=0.5, length=2)
ax.set_ylabel('Replicate 2', size=6)
ax.set_xlabel('Replicate 1', size=6)
ax.set_title('Correlations between\ndistinct outcomes', size=7)
return fig
def plot_outcome_r_matrix(self, threshold=5e-3, n_guides=100):
outcomes = self.outcomes_above_simple_threshold(threshold)
guides = self.union_of_top_n_guides(outcomes, n_guides)
outcome_r_matrix, _ = self.outcome_r_matrix(outcomes, guides)
fig, ax = plt.subplots(figsize=(6, 6))
ax.imshow(outcome_r_matrix, cmap=visualize.correlation_cmap, vmin=-1, vmax=1)
ax.set_xticks([])
ax.set_yticks([])
plt.setp(ax.spines.values(), visible=False)
title = '''\
correlations between outcomes
diagonal: reproducibility between replicates (correlation of outcome i in rep 1 with outcome i in rep 2)
above diagonal: correlations between distinct outcomes in rep 1
below diagonal: correlations between distinct outcome in rep 2
'''
ax.set_title(title)
return fig
def plot_guide_reproducibility(self, outcomes):
rs = self.guide_r_series(outcomes)
fig, ax = plt.subplots(figsize=(12, 6))
sorted_rs = rs.sort_values(ascending=False)
xs = np.arange(len(sorted_rs))
df = pd.DataFrame(sorted_rs)
df['x'] = xs
df['color'] = 'black'
df.loc[self.common_non_targeting_guides, 'color'] = 'tab:orange'
kwargs = dict(x='x', y='r', linewidths=(0,), color='color')
ax.scatter(data=df.query('color == "black"'), s=10, label='', **kwargs)
ax.scatter(data=df.query('color != "black"'), s=25, label='non-targeting guides', **kwargs)
ax.legend()
ax.set_ylim(-0.75, 1)
ax.set_xlim(-0.01 * len(rs), 1.01 * len(rs))
ax.axhline(0, color='black')
ax.set_xlabel('guides ranked by between-replicate correlation', size=12)
ax.set_ylabel('correlation between replicates in outcome redistribution profile', size=12)
return fig
def compare_single_outcomes(self,
outcome_ranks=None,
top_n_guides=None,
manual_data_lims=None,
manual_ticks=None,
correlation_label=None,
scale=2,
):
x = self.pn0
y = self.pn1
outcomes = self.outcomes_above_simple_threshold(1e-3)
if top_n_guides is not None:
relevant_guides = list(self.union_of_top_n_guides(outcomes, top_n_guides))
guides = relevant_guides + list(self.common_non_targeting_guides)
else:
guides = self.common_guides
relevant_guides = guides
log2_fold_changes = self.log2_fold_changes(outcomes, guides).stack().sort_index()
if isinstance(outcome_ranks, int):
outcome_ranks = np.arange(outcome_ranks)
num_outcomes = len(outcome_ranks)
fig, axs = plt.subplots(1, num_outcomes, figsize=(scale * num_outcomes, scale * 0.5), gridspec_kw=dict(wspace=1))
for outcome_i, ax in zip(outcome_ranks, axs):
outcome = outcomes[outcome_i]
df = log2_fold_changes.loc[outcome]
if manual_data_lims is not None:
data_lims = manual_data_lims
else:
data_lims = (np.floor(df.min().min() - 0.1), np.ceil(df.max().max() + 0.1))
df['color'] = visualize.targeting_guide_color
df.loc[self.common_non_targeting_guides, 'color'] = visualize.nontargeting_guide_color
df = df.dropna()
r, p = scipy.stats.pearsonr(df.loc[relevant_guides, x], df.loc[relevant_guides, y])
ax.scatter(x=x, y=y, data=df, color='color', linewidth=(0,), s=10, alpha=0.9, clip_on=False)
ax.set_xlim(*data_lims)
ax.set_ylim(*data_lims)
ax.set_xlabel(f'Replicate 1', size=6)
ax.set_ylabel(f'Replicate 2', size=6)
outcome_string = '\n'.join(outcome)
ax.set_title(f'Rank: {outcome_i}\n{outcome_string}', size=6)
line_kwargs = dict(alpha=0.2, color='black', linewidth=0.5)
hits.visualize.draw_diagonal(ax, **line_kwargs)
ax.axhline(0, **line_kwargs)
ax.axvline(0, **line_kwargs)
if correlation_label == 'r_squared':
label = f'r$^2$ = {r**2:0.2f}'
elif correlation_label == 'r':
label = f'r = {r:0.2f}'
else:
label = None
if label is not None:
ax.annotate(label,
xy=(0, 1),
xycoords='axes fraction',
xytext=(3, -3),
textcoords='offset points',
ha='left',
va='top',
size=6,
)
ax.set_aspect('equal')
ax.tick_params(labelsize=6)
if manual_ticks is not None:
ax.set_xticks(manual_ticks)
ax.set_yticks(manual_ticks)
plt.setp(ax.spines.values(), linewidth=0.5)
ax.tick_params(width=0.5)
return fig
def compare_single_guides(self, guide_ranks=5, threshold=2e-3, manual_data_lims=None, manual_ticks=None):
x = self.pn0
y = self.pn1
outcomes = self.outcomes_above_simple_threshold(threshold)
log2_fold_changes = self.log2_fold_changes(outcomes, self.common_guides)
r_series = self.guide_r_series(outcomes).sort_values(ascending=False)
if isinstance(guide_ranks, int):
guide_ranks = np.arange(guide_ranks)
num_guides = len(guide_ranks)
fig, axs = plt.subplots(1, num_guides, figsize=(2 * num_guides, 1), gridspec_kw=dict(wspace=1))
for guide_i, ax in zip(guide_ranks, axs):
guide = r_series.index[guide_i]
df = log2_fold_changes.xs(guide, axis=1, level=1).copy()
if manual_data_lims is not None:
data_lims = manual_data_lims
else:
data_lims = (np.floor(df.min().min() - 0.1), np.ceil(df.max().max() + 0.1))
df['color'] = 'black'
r, p = scipy.stats.pearsonr(df[x], df[y])
ax.scatter(x=x, y=y, data=df, color='color', linewidth=(0,), s=10, alpha=0.5, marker='D', clip_on=False)
ax.set_xlim(*data_lims)
ax.set_ylim(*data_lims)
ax.set_xlabel(f'Replicate 1', size=6)
ax.set_ylabel(f'Replicate 2', size=6)
ax.set_title(f'{guide}', size=6)
line_kwargs = dict(alpha=0.2, color='black', linewidth=0.5)
hits.visualize.draw_diagonal(ax, **line_kwargs)
ax.axhline(0, **line_kwargs)
ax.axvline(0, **line_kwargs)
ax.tick_params(labelsize=6)
ax.annotate(f'r$^2$ = {r**2:0.2f}',
xy=(0, 1),
xycoords='axes fraction',
xytext=(2, -2),
textcoords='offset points',
ha='left',
va='top',
size=6,
)
ax.set_aspect('equal')
if manual_ticks is not None:
ax.set_xticks(manual_ticks)
ax.set_yticks(manual_ticks)
plt.setp(ax.spines.values(), linewidth=0.5)
ax.tick_params(width=0.5)
return fig
def active_guides_for_same_gene(self, threshold=5e-3, num_guides=100):
outcomes = self.outcomes_above_simple_threshold(threshold)
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
rs = self.guide_guide_correlations(threshold)
for ax, pn in zip(axs, self.pn_pair):
pool = self.pools[pn]
active_guides = pool.top_n_active_guides(outcomes, num_guides)
active_pairs = rs.query('guide_1 in @active_guides and guide_2 in @active_guides')
same_gene = active_pairs.query('gene_1 == gene_2')
bins = np.linspace(-1, 1, 30)
common_kwargs = dict(bins=bins, linewidth=2, histtype='step', density=True)
ax.hist(active_pairs[pn], **common_kwargs)
ax.hist(same_gene[pn], **common_kwargs)
ax.set_xlim(-1.02, 1.02)
return fig
def guide_guide_correlations(self, threshold=5e-3):
outcomes = self.outcomes_above_simple_threshold(threshold)
log2_fold_changes = self.log2_fold_changes(outcomes, self.common_guides)
rs = {}
for pn in self.pn_pair:
all_corrs = log2_fold_changes[pn].corr().stack()
rs[pn] = all_corrs
rs = pd.DataFrame(rs)
rs.index.names = ['guide_1', 'guide_2']
# Make columns out of index levels.
for i in [1, 2]:
k = f'guide_{i}'
rs[k] = rs.index.get_level_values(k)
rs[f'gene_{i}'] = self.common_guides_df.loc[rs[k], 'gene'].values
upper_triangle = rs.query('guide_1 < guide_2')
return upper_triangle
def guide_guide_correlation_scatter(self, threshold=5e-3, num_guides=100, bad_guides=None, rasterize=False):
outcomes = self.outcomes_above_simple_threshold(5e-3)
guides = self.union_of_top_n_guides(outcomes, num_guides)
if bad_guides is not None:
guides = [g for g in guides if g not in bad_guides]
rs = self.guide_guide_correlations(threshold)
active_rs = rs.query('guide_1 in @guides and guide_2 in @guides').copy()
active_rs['color'] = 'grey'
color_for_same = 'tab:cyan'
active_rs.loc[active_rs['gene_1'] == active_rs['gene_2'], 'color'] = color_for_same
g = sns.JointGrid(height=1.75, space=0.5, xlim=(-1.02, 1.02), ylim=(-1.02, 1.02))
ax = g.ax_joint
distinct = active_rs.query('color == "grey"')
same = active_rs.query('color == @color_for_same')
x = self.pn0
y = self.pn1
kwargs = dict(x=x, y=y, color='color', linewidths=(0,), clip_on=False, rasterized=rasterize)
ax.scatter(data=distinct, s=5, alpha=0.2, **kwargs)
ax.scatter(data=same, s=10, alpha=0.95, **kwargs)
plt.setp(ax.spines.values(), linewidth=0.5)
ax.annotate('same\ngene',
xy=(0, 1),
xycoords='axes fraction',
xytext=(3, 0),
textcoords='offset points',
ha='left',
va='top',
color=color_for_same,
size=6,
)
ax.annotate('different\ngenes',
xy=(0, 1),
xycoords='axes fraction',
xytext=(3, -16),
textcoords='offset points',
ha='left',
va='top',
color='grey',
size=6,
)
hits.visualize.draw_diagonal(ax, alpha=0.2)
ax.axhline(0, color='black', alpha=0.2)
ax.axvline(0, color='black', alpha=0.2)
ax.set_xlabel('Replicate 1', size=6)
ax.set_ylabel('Replicate 2', size=6)
ax.set_title('Correlations between\ndistinct CIRSPRi sgRNAs', size=7, y=1.2)
ax.tick_params(labelsize=6, width=0.5, length=2)
ticks = [-1, -0.5, 0, 0.5, 1]
ax.set_xticks(ticks)
ax.set_yticks(ticks)
for ax, pn, orientation in [(g.ax_marg_x, x, 'vertical'), (g.ax_marg_y, y, 'horizontal')]:
bins = np.linspace(-1, 1, 30)
kwargs = dict(histtype='step', density=True, orientation=orientation)
max_n = 0
n, *rest = ax.hist(distinct[pn], color='grey', bins=bins, **kwargs)
max_n = max(max_n, max(n))
focused_bins = bins[bisect.bisect_right(bins, min(same[pn])) - 1:]
n, *rest = ax.hist(same[pn], color=color_for_same, bins=focused_bins, **kwargs)
max_n = max(max_n, max(n))
if orientation == 'vertical':
set_lim = ax.set_ylim
else:
set_lim = ax.set_xlim
set_lim(0, max_n * 1.05)
plt.setp(ax.spines.values(), linewidth=0.5)
ax.tick_params(labelsize=6, width=0.5, length=2)
return g.fig, active_rs
def full_categories(self, outcomes):
full_categories = defaultdict(list)
for c, s, d in outcomes:
ti = self.target_info
if c == 'deletion':
deletion = knock_knock.outcome.DeletionOutcome.from_string(d).undo_anchor_shift(ti.anchor)
directionality = deletion.classify_directionality(ti)
full_category = f'{c}, {directionality}'
else:
full_category = c
full_categories[full_category].append((c, s, d))
return full_categories
def get_bidirectional_deletions(self, threshold=5e-3):
ti = self.pool0.target_info
def get_min_removed(d):
deletion = knock_knock.outcome.DeletionOutcome.from_string(d).undo_anchor_shift(ti.anchor)
min_removed_before = max(0, ti.cut_after - max(deletion.deletion.starts_ats) + 1)
min_removed_after = max(0, min(deletion.deletion.ends_ats) - ti.cut_after)
return min_removed_before, min_removed_after
relevant_outcomes = []
for c, s, d in self.outcomes_above_simple_threshold(threshold):
if c == 'deletion':
min_before, min_after = get_min_removed(d)
if min_before > 0 and min_after > 0:
relevant_outcomes.append((c, s, d))
return relevant_outcomes
class PoolReplicates:
def __init__(self, pools, short_name):
self.pools = pools
self.short_name = short_name
self.target_info = self.pools[0].target_info
self.variable_guide_library = self.pools[0].variable_guide_library
@memoized_property
def outcome_fractions(self):
return pd.concat({pool.name: pool.outcome_fractions('perfect')['none'] for pool in self.pools}, axis=1).fillna(0)
@memoized_property
def outcome_fraction_means(self):
return self.outcome_fractions.mean(axis=1, level=1)
@memoized_property
def outcome_fraction_stds(self):
return self.outcome_fractions.std(axis=1, level=1)
@memoized_property
def non_targeting_fraction_means(self):
return self.outcome_fraction_means[ALL_NON_TARGETING].sort_values(ascending=False)
@memoized_property
def non_targeting_fraction_stds(self):
return self.outcome_fraction_stds[ALL_NON_TARGETING].loc[self.non_targeting_fraction_means.index]
@memoized_property
def log2_fold_changes(self):
fs = self.outcome_fraction_means
fold_changes = fs.div(fs[ALL_NON_TARGETING], axis=0)
return np.log2(fold_changes)
@memoized_property
def log2_fold_change_intervals(self):
fs = self.outcome_fraction_means
fold_changes = fs.div(fs[ALL_NON_TARGETING], axis=0)
return np.log2(fold_changes)
@memoized_property
def category_fractions(self):
all_fs = {pool.name: pool.category_fractions for pool in self.pools}
return pd.concat(all_fs, axis=1).fillna(0)
@memoized_property
def category_fraction_means(self):
return self.category_fractions.groupby(axis=1, level=1).mean()
@memoized_property
def category_fraction_stds(self):
return self.category_fractions.std(axis=1, level=1)
@memoized_property
def categories_by_baseline_frequency(self):
return self.category_fraction_means[ALL_NON_TARGETING].sort_values(ascending=False).index.values
@memoized_property
def category_fraction_differences(self):
return pd.concat({pool.name: pool.category_fraction_differences for pool in self.pools}, axis=1).fillna(0)
@memoized_property
def category_fraction_difference_means(self):
return self.category_fraction_differences.groupby(axis=1, level=1).mean()
@memoized_property
def category_fraction_difference_stds(self):
return self.category_fraction_differences.std(axis=1, level=1)
@memoized_property
def category_log2_fold_changes(self):
return pd.concat({pool.name: pool.category_log2_fold_changes for pool in self.pools}, axis=1).fillna(0)
@memoized_property
def category_log2_fold_change_means(self):
return self.category_log2_fold_changes.groupby(axis=1, level=1).mean()
@memoized_property
def cateogry_log2_fold_change_stds(self):
return self.category_log2_fold_changes.std(axis=1, level=1)
@memoized_property
def gene_level_category_statistics(self):
return pd.concat({pool.name: pool.gene_level_category_statistics for pool in self.pools}, axis=1)
@memoized_property
def gene_level_category_statistic_means(self):
return self.gene_level_category_statistics.groupby(axis=1, level=[1, 2]).mean()
def plot_ranked_category_statistics(self, category, stat='extreme_2', top_n=3, bottom_n=3, y_lim=(-2, 2)):
df = self.gene_level_category_statistics.xs([category, stat], level=[1, 2], axis=1).copy()
first, second = [pool.name for pool in self.pools]
df['color'] = 'black'
df.loc[['MSH2', 'MSH6'], 'color'] = 'tab:green'
df.loc[['PMS2', 'MLH1'], 'color'] = 'tab:green'
df.loc['HLTF', 'color'] = 'tab:orange'
#df.loc[[f'POLD{i}' for i in range(1, 5)], 'color'] = 'tab:red'
#df.loc[[f'RFC{i}' for i in range(1, 6)], 'color'] = 'tab:purple'
#df.loc[['MRE11', 'RAD50', 'NBN'], 'color'] = 'tab:pink'
df['mean'] = df.mean(axis=1).sort_values()
df = df.sort_values('mean')
df['x'] = np.arange(len(df))
df = df.drop('negative_control', errors='ignore')
fig, ax = plt.subplots(figsize=(6, 4))
for to_plot, alpha in [
(df.query('color == "black"'), 0.5),
(df.query('color != "black"'), 0.9),
]:
ax.scatter(x='x', y='mean', data=to_plot, linewidths=(0,), s=10, c='color', alpha=alpha, clip_on=False)
for _, row in df.iterrows():
xs = [row['x'], row['x']]
ys = sorted(row[[first, second]])
ax.plot(xs, ys, alpha=0.2, color=row['color'], clip_on=False)
ax.axhline(0, color='black')
ax.set_ylim(*y_lim)
ax.set_xlim(-0.02 * len(df), 1.02 * len(df))
label_kwargs = dict(
ax=ax,
xs='x',
ys='mean',
labels='gene',
initial_distance=10,
arrow_alpha=0.15,
color='color',
avoid=True,
)
hits.visualize.label_scatter_plot(data=df.iloc[::-1][:top_n], vector='left', **label_kwargs)
hits.visualize.label_scatter_plot(data=df.iloc[:bottom_n], vector='right', **label_kwargs)
ax.set_xticks([])
ax.set_ylabel('log2 fold-change from non-targeting\n(average of 2 most extreme guides,\naverage of 2 replicates)')
for side in ['bottom', 'top', 'right']:
ax.spines[side].set_visible(False)
ax.grid(axis='y', alpha=0.2)
ax.set_title(category, size=16)
return fig | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/replicates.py | 0.480479 | 0.30832 | replicates.py | pypi |
import random
from collections import Counter, defaultdict
from pathlib import Path
from multiprocessing import Pool
import pandas as pd
import numpy as np
import h5py
import tqdm
from knock_knock import experiment
from hits import utilities
progress = tqdm.tqdm
def get_quantile(leq, negative_geq, q):
if q <= 0.5:
# largest value such that no more than q * total are less than or equal to value '''
total = leq[-1]
quantile = np.searchsorted(leq, total * q)
else:
# smallest value such that no more than (1 - q) * total are greater than or equal to value '''
total = negative_geq[0]
quantile = np.searchsorted(negative_geq, total * (1 - q), side='right')
return quantile
def quantiles_to_record(num_samples):
to_record = {
'median': 0.5,
}
for exponent in range(1, int(np.log10(num_samples)) + 1):
to_record['down_{}'.format(exponent)] = 10**-exponent
to_record['up_{}'.format(exponent)] = 1 - 10**-exponent
return to_record
def sample_negative_controls(guide_outcomes, num_cells, num_samples):
progress = utilities.identity
print(num_cells)
counts_by_sample = []
outcome_names = set()
qs = quantiles_to_record(num_samples)
guides = list(guide_outcomes)
for _ in range(num_samples):
cells = []
while len(cells) < num_cells:
guide = random.choice(guides)
cells.extend(guide_outcomes[guide])
cells = random.sample(cells, num_cells)
outcomes = ((outcome.category, outcome.subcategory, outcome.details) for outcome in cells)
counts = Counter(outcomes)
counts_by_sample.append(counts)
outcome_names.update(counts)
outcome_groups = {
'deletion': {outcome for outcome in outcome_names if outcome[:2] == ('indel', 'deletion')},
'insertion': {outcome for outcome in outcome_names if outcome[:2] == ('indel', 'insertion')},
'partial_donor': {outcome for outcome in outcome_names if outcome[:2] == ('no indel', 'other')},
}
outcome_to_groups = defaultdict(set)
for outcome_group_name, outcome_group in outcome_groups.items():
for outcome in outcome_group:
outcome_to_groups[outcome].add(outcome_group_name)
filled_counts = {k: np.zeros(num_samples, int) for k in list(outcome_groups) + ['not_observed']}
results = {}
def process_frequencies(frequencies):
full_frequencies = np.zeros(num_cells + 1, int)
full_frequencies[:len(frequencies)] = frequencies
leq = np.cumsum(full_frequencies)
geq = leq[-1] - leq + full_frequencies
negative_geq = -geq
results = {
'frequencies': frequencies,
'quantiles': {}
}
for key, q in qs.items():
results['quantiles'][key] = get_quantile(leq, negative_geq, q)
return results
for outcome in progress(outcome_names):
filled = np.array([counts[outcome] for counts in counts_by_sample])
frequencies = utilities.counts_to_array(Counter(filled))
results[outcome] = process_frequencies(frequencies)
for outcome_group_name in outcome_to_groups[outcome]:
filled_counts[outcome_group_name] += filled
for outcome, filled in filled_counts.items():
frequencies = utilities.counts_to_array(Counter(filled))
results[outcome] = process_frequencies(frequencies)
return results
if __name__ == '__main__':
base_dir = Path('/home/jah/projects/britt')
group_name = '2018_09_07_rep1'
pool = experiment.PooledExperiment(base_dir, group_name)
num_cells_list = np.concatenate([np.arange(10, 100, 5),
np.arange(100, 1000, 50),
np.arange(1000, 5000, 100),
np.arange(5000, 20000, 1000),
np.arange(20000, 40000, 5000),
],
)
num_samples = 100000
guide_outcomes = pool.non_targeting_outcomes
args_list = [(guide_outcomes, num_cells, num_samples) for num_cells in num_cells_list]
with Pool(processes=20) as process_pool:
all_results = process_pool.starmap(sample_negative_controls, args_list, chunksize=1)
#all_results = [sample_negative_controls(*args) for args in progress(args_list)]
by_num_cells = dict(zip(num_cells_list, all_results))
by_outcome = {}
all_outcome_names = set()
for num_cells in num_cells_list:
all_outcome_names.update(by_num_cells[num_cells])
all_outcome_names.remove('not_observed')
for outcome_name in all_outcome_names:
by_outcome[outcome_name] = {
'quantiles': defaultdict(list),
'frequencies': {},
}
for num_cells in num_cells_list:
results = by_num_cells[num_cells]
for outcome_name in all_outcome_names:
outcome_results = results.get(outcome_name, results['not_observed'])
for q, v in outcome_results['quantiles'].items():
by_outcome[outcome_name]['quantiles'][q].append(v)
by_outcome[outcome_name]['frequencies'][num_cells] = outcome_results['frequencies']
hdf5_fn = pool.fns['quantiles']
with h5py.File(hdf5_fn, 'w') as f:
f.create_dataset('num_cells', data=num_cells_list)
f.attrs['num_samples'] = num_samples
for outcome, results in progress(by_outcome.items()):
if isinstance(outcome, tuple):
name = '_'.join(outcome)
else:
name = outcome
group = f.create_group(name)
frequencies = group.create_group('frequencies')
for num_cells, vs in results['frequencies'].items():
frequencies.create_dataset(str(num_cells), data=vs)
quantiles = group.create_group('quantiles')
for q, vs in results['quantiles'].items():
quantiles.create_dataset(str(q), data=vs) | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/quantiles.py | 0.53048 | 0.359814 | quantiles.py | pypi |
import bokeh.palettes
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
idx = pd.IndexSlice
def plot_length_distributions(pool,
guide_combinations_to_plot,
species='hg19',
smoothing_window=10,
draw_legend=True,
):
def get_length_distributions_for_multiple_guides(pool, species, guide_pairs):
# counts for each specified guide
length_counts = pool.genomic_insertion_length_counts.loc[species].loc[guide_pairs]
# total counts at each length across all specified guides
all_guide_length_counts = length_counts.sum(axis='rows')
# total UMIs across all specified guides
total_UMIs = pool.UMI_counts_for_all_fixed_guides().loc[guide_pairs].sum()
all_guide_length_fractions = all_guide_length_counts / total_UMIs
smoothed_fractions = all_guide_length_fractions.rolling(2 * smoothing_window + 1, center=True, min_periods=1).mean()
return all_guide_length_fractions, smoothed_fractions
relevant_lengths = idx[25:pool.blunt_insertion_length_detection_limit]
upper_limit = pool.blunt_insertion_length_detection_limit
x_for_above_limit = upper_limit + 75
x_max = x_for_above_limit + 25
fig_width = 0.0075 * x_max
axs = {}
fig, (axs['frequency'], axs['log2_fc']) = plt.subplots(2, 1,
figsize=(fig_width, 3),
gridspec_kw=dict(hspace=0.4,
height_ratios=[3, 2],
),
)
axs['frequency_high'] = axs['frequency'].twinx()
nt_fs, smoothed_nt_fs = get_length_distributions_for_multiple_guides(pool, species, pool.non_targeting_guide_pairs)
for label, guides, color, line_width, alpha in guide_combinations_to_plot:
fs, smoothed_fs = get_length_distributions_for_multiple_guides(pool, species, guides)
axs['frequency'].plot(smoothed_fs.loc[relevant_lengths] * 100, color=color, alpha=alpha, linewidth=line_width, label=label)
axs['frequency_high'].plot(x_for_above_limit, fs.iloc[-1] * 100, 'o', markersize=2, color=color)
l2fcs = np.log2(smoothed_fs / smoothed_nt_fs)
axs['log2_fc'].plot(np.maximum(-5, l2fcs.loc[relevant_lengths]), color=color, alpha=alpha, linewidth=line_width)
fc = fs.iloc[-1] / nt_fs.iloc[-1]
log2_fc = np.log2(fc)
axs['log2_fc'].plot(x_for_above_limit, log2_fc, 'o', markersize=2, color=color)
for ax in (axs['log2_fc'],):
min_fc = -4
max_fc = 4
ax.set_ylim(min_fc, max_fc)
for fc in np.arange(min_fc, max_fc + 1):
ax.axhline(fc, color='black', alpha=0.1)
for ax in axs.values():
ax.set_xlim(0, x_max)
axs['frequency'].set_ylabel('Percentage of\nrepair outcomes', size=7)
axs['log2_fc'].set_ylabel('Log$_2$ fold change\nfrom non-targeting', size=7)
axs['log2_fc'].set_yticks(np.arange(-4, 6, 2))
for ax in axs.values():
main_ticks = list(range(0, upper_limit, 50))
main_tick_labels = [f'{x:,}' for x in main_ticks]
extra_ticks = [x_for_above_limit] + list(range(25, upper_limit, 50))
extra_tick_labels = ['longer than\ndetection limit'] + ['']*len(list(range(25, upper_limit, 50)))
ax.set_xticks(main_ticks + extra_ticks)
ax.set_xticklabels(main_tick_labels + extra_tick_labels, size=6)
ax.axvline(25, color='black', alpha=0.2)
ax.axvline(pool.blunt_insertion_length_detection_limit, color='black', alpha=0.2)
axs['frequency_high'].annotate(
'upper\ndetection limit',
xy=(pool.blunt_insertion_length_detection_limit, 1),
xycoords=('data', 'axes fraction'),
xytext=(0, 3),
textcoords='offset points',
ha='center',
va='bottom',
size=6,
)
axs['frequency_high'].annotate(
'lower\ndetection limit',
xy=(25, 1),
xycoords=('data', 'axes fraction'),
xytext=(0, 3),
textcoords='offset points',
ha='center',
va='bottom',
size=6,
)
axs['log2_fc'].set_xlabel('Length of captured\ngenomic sequence', size=7)
axs['log2_fc'].axhline(0, color=bokeh.palettes.Greys9[2])
axs['frequency_high'].spines['bottom'].set_visible(False)
axs['frequency'].spines['top'].set_visible(False)
axs['frequency'].set_ylim(0, 0.072)
axs['frequency_high'].set_ylim(0, 6)
if draw_legend:
axs['frequency'].legend(bbox_to_anchor=(0.5, 1.25),
loc='lower center',
fontsize=6,
)
for ax in axs.values():
ax.tick_params(labelsize=6, width=0.5)
plt.setp(ax.spines.values(), linewidth=0.5)
return fig | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/visualize/templated_insertions.py | 0.679285 | 0.457803 | templated_insertions.py | pypi |
from collections import defaultdict
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import scipy.cluster.hierarchy as sch
import pandas as pd
import hits.visualize
import hits.utilities
import knock_knock.outcome
import repair_seq.cluster
import repair_seq.visualize
import repair_seq.visualize.outcome_diagrams
import repair_seq.visualize.heatmap
memoized_property = hits.utilities.memoized_property
class Clustermap:
def __init__(self,
clusterer,
fig=None,
diagram_ax_rectangle=None,
**options,
):
'''
diagram_ax_rectangle: rectangle (in figure fraction coords) for diagrams to occupy in existing figure
'''
options.setdefault('upside_down', False)
options.setdefault('draw_fold_changes', True)
options.setdefault('draw_outcome_clusters', False)
options.setdefault('draw_guide_clusters', True)
options.setdefault('draw_outcome_similarities', True)
options.setdefault('draw_guide_similarities', True)
options.setdefault('draw_colorbars', True)
options.setdefault('guide_library', None)
options.setdefault('diagram_kwargs', {})
options.setdefault('gene_text_size', 14)
options.setdefault('guide_to_color', {})
options.setdefault('guide_to_alias', {})
options.setdefault('guides_on_top', False)
#options.setdefault('guide_to_color', {})
options.setdefault('alphabetical_guides', False)
options.setdefault('original_outcome_order', False)
options.setdefault('colorbar_text_size', 12)
options['diagram_kwargs'].setdefault('window', (-30, 30))
self.options = options
self.clusterer = clusterer
self.inches_per_guide = 14 / 100
self.num_guides = len(self.clusterer.guides)
self.num_outcomes = len(self.clusterer.outcomes)
self.axs = {}
self.width_inches = {}
self.height_inches = {}
self.x0 = {}
self.x1 = {}
self.y0 = {}
self.y1 = {}
self.ims = {}
self.fig = fig
if self.fig is None:
# want 1 nt of diagram to be 3/4ths as wide as 1 guide of heatmap
window_start, window_end = self.options['diagram_kwargs']['window']
window_size = window_end - window_start + 1
self.width_inches['diagrams'] = self.inches_per_guide * window_size * 0.75
self.height_inches['diagrams'] = self.inches_per_guide * self.num_outcomes
self.fig = plt.figure(figsize=(self.width_inches['diagrams'], self.height_inches['diagrams']))
diagram_ax_rectangle = (0, 0, 1, 1)
self.diagram_ax_rectangle = diagram_ax_rectangle
self.fig_width_inches, self.fig_height_inches = self.fig.get_size_inches()
self.height_per_guide = self.inches_per_guide / self.fig_height_inches
self.width_per_guide = self.inches_per_guide / self.fig_width_inches
self.draw()
def draw(self):
self.draw_diagrams()
if self.options['draw_fold_changes']:
self.draw_fold_changes()
if self.options['draw_outcome_similarities']:
self.draw_outcome_similarities()
if self.options['draw_guide_similarities']:
self.draw_guide_similarities()
if self.options['draw_guide_clusters']:
self.draw_guide_clusters()
self.annotate_guide_clusters()
if self.options['draw_outcome_clusters']:
self.draw_outcome_clusters()
if self.options['draw_colorbars']:
self.draw_colorbars()
def width(self, ax_name):
return self.width_inches[ax_name] / self.fig_width_inches
def height(self, ax_name):
return self.height_inches[ax_name] / self.fig_height_inches
def rectangle(self, ax_name):
width = self.width(ax_name)
height = self.height(ax_name)
if ax_name in self.x0:
x0 = self.x0[ax_name]
elif ax_name in self.x1:
x0 = self.x1[ax_name] - width
else:
raise ValueError(ax_name)
if ax_name in self.y0:
y0 = self.y0[ax_name]
elif ax_name in self.y1:
y0 = self.y1[ax_name] - height
else:
raise ValueError(ax_name)
return (x0, y0, width, height)
def add_axes(self, ax_name, sharex=None, sharey=None):
ax = self.fig.add_axes(self.rectangle(ax_name), sharex=self.axs.get(sharex), sharey=self.axs.get(sharey))
self.axs[ax_name] = ax
return ax
def get_position(self, ax_name):
return self.axs[ax_name].get_position()
def draw_diagrams(self):
self.x0['diagrams'], self.y0['diagrams'], width, height = self.diagram_ax_rectangle
self.width_inches['diagrams'] = width * self.fig_width_inches
self.height_inches['diagrams'] = height * self.fig_height_inches
ax = self.add_axes('diagrams')
# See TODO comment in fold changes on reversing here.
if self.options['original_outcome_order']:
outcomes = self.clusterer.original_outcome_order_with_pool[::-1]
else:
outcomes = self.clusterer.clustered_outcomes[::-1]
repair_seq.visualize.outcome_diagrams.plot(outcomes,
self.clusterer.pn_to_target_info,
ax=ax,
replacement_text_for_complex={
'genomic insertion, hg19, <=75 nts': 'capture of genomic sequence ≤75 nts',
'genomic insertion, hg19, >75 nts': 'capture of genomic sequence >75 nts',
},
**self.options['diagram_kwargs'],
)
ax.set_position(self.rectangle('diagrams'))
def draw_frequencies(self, frequencies,
x_lims=(np.log10(1.99e-3), np.log10(2.01e-1)),
manual_ticks=None,
include_percentage=False,
label_size=6,
**plot_kwargs,
):
diagrams_position = self.get_position('diagrams')
self.x0['frequencies'] = diagrams_position.x1 + diagrams_position.width * 0.02
self.y0['frequencies'] = diagrams_position.y0
self.width_inches['frequencies'] = self.inches_per_guide * 15
self.height_inches['frequencies'] = self.inches_per_guide * self.num_outcomes
ax = self.add_axes('frequencies', sharey='diagrams')
ax.set_yticks([])
ax.xaxis.tick_top()
ax.spines['left'].set_alpha(0.3)
ax.spines['right'].set_alpha(0.3)
ax.tick_params(labelsize=6)
ax.grid(axis='x', alpha=0.3, clip_on=False)
ax.spines['bottom'].set_visible(False)
x_min, x_max = x_lims
ax.set_xlim(x_min, x_max)
x_ticks = []
for exponent in [6, 5, 4, 3, 2, 1, 0]:
xs = np.log10(np.arange(1, 10) * 10**-exponent)
for x in xs:
if x_min < x < x_max:
ax.axvline(x, color='black', alpha=0.05, clip_on=False)
if exponent <= 3:
multiples = [1, 5]
else:
multiples = [1]
for multiple in multiples:
x = multiple * 10**-exponent
if x_min <= np.log10(x) <= x_max:
x_ticks.append(x)
if manual_ticks is not None:
x_ticks = manual_ticks
ax.set_xticks(np.log10(x_ticks))
ax.set_xticklabels([f'{100 * x:g}' + ('%' if include_percentage else '') for x in x_ticks], size=label_size)
for side in ['left', 'right']:
ax.spines[side].set_visible(False)
ax.plot(frequencies, np.arange(len(frequencies)), clip_on=False, **plot_kwargs)
def draw_fold_changes(self):
diagrams_position = self.get_position('diagrams')
self.x0['fold changes'] = diagrams_position.x1 + diagrams_position.width * 0.02
self.y0['fold changes'] = diagrams_position.y0
self.width_inches['fold changes'] = self.inches_per_guide * self.num_guides
self.height_inches['fold changes'] = self.inches_per_guide * self.num_outcomes
# TODO: fix weirdness in y axis here. Lining up diagrams, fold changes, and
# outcome similarities requires sharey and reversing outcome order in diagrams.
ax = self.add_axes('fold changes', sharey='diagrams')
heatmap_to_plot = self.clusterer.clustered_log2_fold_changes
if self.options['alphabetical_guides']:
heatmap_to_plot = heatmap_to_plot.rename(columns=self.options['guide_to_alias'])
heatmap_to_plot = heatmap_to_plot.sort_index(axis=1)
if self.options['original_outcome_order']:
heatmap_to_plot = heatmap_to_plot.loc[self.clusterer.original_outcome_order_with_pool]
im = ax.imshow(heatmap_to_plot,
cmap=repair_seq.visualize.fold_changes_cmap,
vmin=-2, vmax=2,
interpolation='none',
)
self.ims['fold changes'] = im
ax.axis('off')
on_top = self.options['upside_down'] or self.options['guides_on_top']
for x, guide in enumerate(heatmap_to_plot.columns):
color = self.options['guide_to_color'].get(guide, 'black')
alias = self.options['guide_to_alias'].get(guide, guide)
ax.annotate(alias,
xy=(x, 1 if on_top else 0),
xycoords=('data', 'axes fraction'),
xytext=(0, 3 if on_top else -3),
textcoords='offset points',
rotation=90,
ha='center',
va='bottom' if on_top else 'top',
size=7 if color == 'black' else 9,
color=color,
weight='normal' if color == 'black' else 'bold',
)
def draw_outcome_similarities(self):
fold_changes_position = self.get_position('fold changes')
gap = 0.5 * self.width_per_guide
self.x0['outcome similarity'] = fold_changes_position.x1 + gap
self.y0['outcome similarity'] = fold_changes_position.y0
self.width_inches['outcome similarity'] = self.height_inches['fold changes'] / 2
self.height_inches['outcome similarity'] = self.height_inches['fold changes']
ax = self.add_axes('outcome similarity')
vs = self.clusterer.outcome_clustering['similarities']
for side in ['left', 'top', 'bottom']:
ax.spines[side].set_visible(False)
ax.spines['right'].set_color('white')
im = ax.imshow(vs,
cmap=repair_seq.visualize.correlation_cmap,
vmin=-1,
vmax=1,
interpolation='none',
)
self.ims['outcome similarities'] = im
transform = matplotlib.transforms.Affine2D().rotate_deg(45) + ax.transData
im.set_transform(transform)
ax.set_xticks([])
ax.set_yticks([])
diagonal_length = np.sqrt(2 * len(vs)**2)
ax.set_xlim(0, diagonal_length / 2)
ax.set_ylim(-np.sqrt(2) / 2, -np.sqrt(2) / 2 + diagonal_length)
def draw_guide_similarities(self):
fold_changes_position = self.get_position('fold changes')
self.x0['guide similarity'] = fold_changes_position.x0
gap = 0.5 * self.height_per_guide
if self.options['upside_down']:
self.y1['guide similarity'] = fold_changes_position.y0 - gap
else:
y = fold_changes_position.y1 + gap
if self.options['guides_on_top']:
y += 5 * self.height_per_guide
self.y0['guide similarity'] = y
self.width_inches['guide similarity'] = self.width_inches['fold changes']
self.height_inches['guide similarity'] = self.width_inches['fold changes'] / 2
ax = self.add_axes('guide similarity')
vs = self.clusterer.guide_clustering['similarities']
im = ax.imshow(vs,
cmap=repair_seq.visualize.correlation_cmap,
vmin=-1,
vmax=1,
interpolation='none',
)
self.ims['guide similarities'] = im
ax.set_xticks([])
ax.set_yticks([])
plt.setp(ax.spines.values(), visible=False)
transform = matplotlib.transforms.Affine2D().rotate_deg(-45) + ax.transData
im.set_transform(transform)
diag_length = np.sqrt(2 * len(vs)**2)
if self.options['upside_down']:
ax.set_ylim(diag_length / 2, 0)
else:
ax.set_ylim(0, -diag_length / 2)
ax.set_xlim(-np.sqrt(2) / 2, -np.sqrt(2) / 2 + diag_length)
def draw_guide_clusters(self):
guide_clustering = self.clusterer.guide_clustering
gap_before_clusters = self.height_per_guide * 10
fold_changes_position = self.get_position('fold changes')
self.x0['guide clusters'] = fold_changes_position.x0
if self.options['upside_down']:
self.y0['guide clusters'] = fold_changes_position.y1 + gap_before_clusters
else:
self.y1['guide clusters'] = fold_changes_position.y0 - gap_before_clusters
self.width_inches['guide clusters'] = self.width_inches['fold changes']
self.height_inches['guide clusters'] = self.inches_per_guide * 1
ax = self.add_axes('guide clusters')
assignments = guide_clustering['cluster_assignments']
cluster_blocks = repair_seq.cluster.get_cluster_blocks(assignments)
for cluster_id, blocks in cluster_blocks.items():
if cluster_id == -1:
continue
for block_start, block_end in blocks:
y = 1
x_start = block_start + 0.1
x_end = block_end + 0.9
ax.plot([x_start, x_end],
[y, y],
color=guide_clustering['cluster_colors'][cluster_id],
linewidth=6,
solid_capstyle='butt',
clip_on=False,
)
ax.set_ylim(0.5, 1.5)
ax.set_xlim(0, len(self.clusterer.guides))
ax.axis('off')
def draw_guide_cell_cycle_effects(self):
ax_key = 'guide cell cycle'
guide_order = self.clusterer.guide_clustering['clustered_order']
cell_cycle_fold_changes = self.clusterer.guide_library.cell_cycle_log2_fold_changes[guide_order]
gap_between = self.height_per_guide * 6
fold_changes_position = self.get_position('fold changes')
self.x0[ax_key] = fold_changes_position.x0
if self.options['upside_down']:
self.y0[ax_key] = fold_changes_position.y1 + gap_between
else:
self.y1[ax_key] = fold_changes_position.y0 - gap_between
self.width_inches[ax_key] = self.width_inches['fold changes']
self.height_inches[ax_key] = self.inches_per_guide * 3
ax = self.add_axes(ax_key)
im = ax.imshow(cell_cycle_fold_changes,
cmap=repair_seq.visualize.cell_cycle_cmap,
vmin=-1, vmax=1,
interpolation='none',
)
for y, phase in enumerate(cell_cycle_fold_changes.index):
ax.annotate(phase,
xy=(0, y),
xycoords=('axes fraction', 'data'),
xytext=(-5, 0),
textcoords='offset points',
ha='right',
va='center',
)
ax.axis('off')
colorbar_key = f'{ax_key} colorbar'
heatmap_position = self.get_position(ax_key)
self.x0[colorbar_key] = heatmap_position.x1 + self.width_per_guide * 5
self.y0[colorbar_key] = heatmap_position.y0 + self.height_per_guide * 0.75
self.width_inches[colorbar_key] = self.inches_per_guide * 5
self.height_inches[colorbar_key] = self.inches_per_guide * 1.5
colorbar_ax = self.add_axes(colorbar_key)
colorbar = plt.colorbar(mappable=im, cax=colorbar_ax, orientation='horizontal')
colorbar.set_label(f'log$_2$ fold change\nin cell-cycle phase\noccupancy')
def draw_guide_dendrogram(self):
clusters_position = self.get_position('guide clusters')
gap_after_clusters = self.height_per_guide * 0.5
self.x0['guide dendrogram'] = self.x0['fold changes']
self.y1['guide dendrogram'] = clusters_position.y0 - gap_after_clusters
self.height_inches['guide dendrogram'] = self.inches_per_guide * 10
self.width_inches['guide dendrogram'] = self.width_inches['fold changes']
ax = self.add_axes('guide dendrogram')
sch.dendrogram(self.clusterer.guide_clustering['linkage'],
ax=ax,
color_threshold=-1,
above_threshold_color='black',
orientation='bottom',
)
ax.axis('off')
def draw_outcome_clusters(self):
outcome_clustering = self.clusterer.outcome_clustering
diagrams_position = self.get_position('diagrams')
gap_before_clusters = self.width_per_guide * 0.5
self.x1['outcome clusters'] = diagrams_position.x0 - gap_before_clusters
self.y0['outcome clusters'] = diagrams_position.y0
self.height_inches['outcome clusters'] = self.height_inches['fold changes']
self.width_inches['outcome clusters'] = self.inches_per_guide * 0.5
ax = self.add_axes('outcome clusters')
cluster_blocks = repair_seq.cluster.get_cluster_blocks(outcome_clustering['cluster_assignments'])
for cluster_id, blocks in cluster_blocks.items():
if cluster_id == -1:
continue
for block_start, block_end in blocks:
y_start = block_start + 0.1
y_end = block_end + 0.9
x = 1
ax.plot([x, x],
[y_start, y_end],
color=outcome_clustering['cluster_colors'][cluster_id],
linewidth=6,
solid_capstyle='butt',
clip_on=False,
)
ax.set_xlim(0.5, 1.5)
ax.set_ylim(0, len(self.clusterer.outcomes))
ax.axis('off')
def draw_outcome_categories(self):
diagrams_position = self.get_position('diagrams')
gap_before_categories = self.width_per_guide * 0.5
self.x1['outcome categories'] = diagrams_position.x0 - gap_before_categories
self.y0['outcome categories'] = diagrams_position.y0
self.height_inches['outcome categories'] = self.height_inches['fold changes']
self.width_inches['outcome categories'] = self.inches_per_guide * 2
ax = self.add_axes('outcome categories')
full_categories = defaultdict(list)
outcomes = self.clusterer.clustered_outcomes[::-1]
for pn, c, s, d in outcomes:
ti = self.clusterer.pn_to_target_info[pn]
if c == 'deletion':
deletion = knock_knock.outcome.DeletionOutcome.from_string(d).undo_anchor_shift(ti.anchor)
directionality = deletion.classify_directionality(ti)
full_category = f'{c}, {directionality}'
else:
full_category = c
full_categories[full_category].append((pn, c, s, d))
x = 0
effector = self.clusterer.target_info.effector.name
category_display_order = repair_seq.visualize.category_display_order[effector]
category_colors = repair_seq.visualize.category_colors[effector]
for cat in category_display_order:
cat_outcomes = full_categories[cat]
if len(cat_outcomes) > 0:
indices = sorted([len(outcomes) - 1 - outcomes.index(outcome) for outcome in cat_outcomes])
connected_blocks = []
current_block_start = indices[0]
current_idx = indices[0]
for next_idx in indices[1:]:
if next_idx - current_idx > 1:
block = (current_block_start, current_idx)
connected_blocks.append(block)
current_block_start = next_idx
current_idx = next_idx
# Close off last block
block = (current_block_start, current_idx)
connected_blocks.append(block)
for first, last in connected_blocks:
ax.plot([x, x], [first - 0.4, last + 0.4],
linewidth=6,
color=category_colors[cat],
clip_on=False,
solid_capstyle='butt',
)
x -= 1
ax.set_xlim(x - 1, 1)
ax.set_ylim(-0.5, len(outcomes) - 0.5)
ax.axis('off')
def draw_outcome_dendrogram(self):
clusters_position = self.get_position('outcome clusters')
gap_after_clusters = self.width_per_guide * 0.5
self.height_inches['outcome dendro'] = self.height_inches['outcome clusters']
self.width_inches['outcome dendro'] = self.inches_per_guide * 10
self.x1['outcome dendro'] = clusters_position.x0 - gap_after_clusters
self.y0['outcome dendro'] = clusters_position.y0
ax = self.add_axes('outcome dendro')
sch.dendrogram(self.clusterer.outcome_clustering['linkage'],
ax=ax,
color_threshold=-1,
above_threshold_color='black',
orientation='left',
)
ax.axis('off')
def annotate_guide_clusters(self):
guide_clustering = self.clusterer.guide_clustering
assignments = guide_clustering['cluster_assignments']
cluster_blocks = repair_seq.cluster.get_cluster_blocks(assignments)
cluster_genes = repair_seq.cluster.get_cluster_genes(guide_clustering, self.clusterer.guide_to_gene)
cluster_colors = guide_clustering['cluster_colors']
ax = self.axs['guide clusters']
if self.options['upside_down']:
initial_offset = 7
step_sign = 1
va = 'bottom'
else:
initial_offset = -5
step_sign = -1
va = 'top'
for cluster_id in cluster_blocks:
if cluster_id == -1:
continue
blocks = cluster_blocks[cluster_id]
for block in blocks:
if block[1] - block[0] == 0:
continue
genes = cluster_genes[cluster_id]
gene_and_counts = sorted(genes.most_common(), key=lambda gene_and_count: (-gene_and_count[1], gene_and_count[0]))
for i, (gene, count) in enumerate(gene_and_counts):
count_string = f' x{count}' if count > 1 else ''
ax.annotate(f'{gene}{count_string}',
xy=(np.mean(blocks[0]) + 0.5, 0),
xycoords=('data', 'axes fraction'),
xytext=(0, initial_offset + step_sign * (self.options['gene_text_size'] + 1) * i),
textcoords='offset points',
ha='center',
va=va,
color=cluster_colors[cluster_id],
size=self.options['gene_text_size'],
)
def draw_colorbars(self):
text_size = self.options['colorbar_text_size']
# Similarity colorbar.
self.x0['similarity colorbar'] = self.x0['guide similarity']
self.y0['similarity colorbar'] = self.y0['guide similarity'] + 20 * self.height_per_guide
self.width_inches['similarity colorbar'] = self.inches_per_guide * 1.5
self.height_inches['similarity colorbar'] = self.inches_per_guide * 18
ax = self.add_axes('similarity colorbar')
cbar = plt.colorbar(self.ims['guide similarities'], cax=ax, orientation='vertical', ticks=[-1, 0, 1])
cbar.outline.set_alpha(0.1)
cbar.ax.tick_params(labelsize=text_size)
ax.annotate('correlation\nbetween\nrepair outcome\nredistribution\nprofiles',
xy=(1, 0.5),
xycoords='axes fraction',
xytext=(text_size * 1.5, 0),
textcoords='offset points',
ha='left',
va='center',
size=text_size,
)
# Fold changes colorbar.
diagrams_position = self.get_position('diagrams')
self.x0['fold changes colorbar'] = diagrams_position.x0 + 10 * self.width_per_guide
self.y0['fold changes colorbar'] = diagrams_position.y1 + 10 * self.height_per_guide
self.width_inches['fold changes colorbar'] = self.inches_per_guide * 15
self.height_inches['fold changes colorbar'] = self.inches_per_guide * 1.5
ax = self.add_axes('fold changes colorbar')
repair_seq.visualize.heatmap.add_fold_change_colorbar(self.fig, self.ims['fold changes'], cbar_ax=ax, text_size=text_size)
return cbar
class SinglePoolClustermap(Clustermap):
def __init__(self, pool, **kwargs):
self.pool = pool
self.target_info = pool.target_info
super().__init__(**kwargs)
class MultiplePoolClustermap(Clustermap):
def __init__(self, pools, **kwargs):
self.pools = pools
self.target_info = self.pools[0].target_info
super().__init__(**kwargs)
@memoized_property
def guides(self):
if self.manual_guides is not None:
return self.manual_guides
else:
active_guide_lists = defaultdict(list)
for pool in self.pools:
for i, guide in enumerate(pool.canonical_active_guides):
active_guide_lists[guide].append((pool.short_name, i))
# Include guides that were active in at least two screens.
# Only include up to 3 guides per gene, prioritized by average activity rank.
gene_guides = defaultdict(list)
for guide, active_pns in active_guide_lists.items():
if len(active_pns) >= min(len(self.pools), 2):
average_rank = np.mean([rank for pn, rank in active_pns])
gene_guides[self.options['guide_library'].guide_to_gene[guide]].append((average_rank, guide))
filtered_guides = []
for gene, guides in gene_guides.items():
guides = [g for r, g in sorted(guides)]
filtered_guides.extend(guides[:3])
return filtered_guides
@memoized_property
def fold_changes(self):
all_fcs = {}
for pool in self.pools:
fcs = pool.log2_fold_changes.loc[pool.canonical_outcomes, self.guides]
all_fcs[pool.short_name] = fcs
all_fcs = pd.concat(all_fcs)
return all_fcs
@memoized_property
def outcomes(self):
return [(c, s, d) for pn, c, s, d in self.fold_changes.index.values]
@memoized_property
def clustered_outcomes(self):
return [(c, s, d) for pn, c, s, d in self.clustered_fold_changes.index.values] | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/visualize/clustermap.py | 0.466359 | 0.224034 | clustermap.py | pypi |
import bokeh.palettes
import numpy as np
import matplotlib.pyplot as plt
import hits.utilities
import repair_seq.visualize
import repair_seq.visualize.outcome_diagrams
def draw_ssODN_configurations(pools=None,
tis=None,
draw_SNVs_on_target=True,
flip_target=False,
):
if pools is None:
common_ti = tis[0]
else:
common_ti = pools[0].target_info
tis = [pool.target_info for pool in pools]
rect_height = 0.25
fig, ax = plt.subplots(figsize=(25, 4))
def draw_rect(x0, x1, y0, y1, alpha, color='black', fill=True):
path = [
[x0, y0],
[x0, y1],
[x1, y1],
[x1, y0],
]
patch = plt.Polygon(path,
fill=fill,
closed=True,
alpha=alpha,
color=color,
linewidth=0 if fill else 1.5,
clip_on=False,
)
ax.add_patch(patch)
def mark_5_and_3(y, x_start, x_end, color):
if y > 0:
left_string = '5\''
right_string = '3\''
else:
right_string = '5\''
left_string = '3\''
ax.annotate(left_string, (x_start, y),
xytext=(-5 if not flip_target else 5, 0),
textcoords='offset points',
ha='right' if not flip_target else 'left',
va='center',
color=color,
)
ax.annotate(right_string, (x_end, y),
xytext=(5 if not flip_target else -5, 0),
textcoords='offset points',
ha='left' if not flip_target else 'right',
va='center',
color=color,
)
kwargs = dict(ha='center', va='center', fontfamily='monospace',)
offset_at_cut = 0
ys = {
'target_+': 0.15,
'target_-': -0.15,
'donor_+': 0.75,
'donor_-': -0.75,
}
colors = bokeh.palettes.Set2[8]
donor_x_min = -1
donor_x_max = 1
for ti, color in zip(tis, colors):
_, offset, is_reverse_complement = ti.best_donor_target_alignment
# For ss donors, ti.donor_sequence is the actual stranded sequence
# supplied. There is no need to complement anything, since it is drawn
# on the bottom with the - strand if it aligned to the reverse complement
# of the target strandedness.
if is_reverse_complement:
aligned_donor_seq = ti.donor_sequence[::-1]
donor_y = ys['donor_-']
else:
aligned_donor_seq = ti.donor_sequence
donor_y = ys['donor_+']
donor_cut_after = ti.cut_after - offset
donor_before_cut = aligned_donor_seq[:donor_cut_after + 1]
donor_after_cut = aligned_donor_seq[donor_cut_after + 1:]
offset_at_this_cut = ti.cut_after - common_ti.cut_after + offset_at_cut
for b, x in zip(donor_before_cut[::-1], np.arange(len(donor_before_cut))):
final_x = -x + offset_at_this_cut - 0.5
donor_x_min = min(donor_x_min, final_x)
ax.annotate(b,
(final_x, donor_y),
**kwargs,
)
for b, x in zip(donor_after_cut, np.arange(len(donor_after_cut))):
final_x = x + offset_at_this_cut + 0.5
donor_x_max = max(donor_x_max, final_x)
ax.annotate(b,
(final_x, donor_y),
**kwargs,
)
if is_reverse_complement:
y = ys['donor_-']
ys['donor_-'] -= 0.3
else:
y = ys['donor_+']
ys['donor_+'] += 0.3
draw_rect(-len(donor_before_cut) + offset_at_this_cut,
len(donor_after_cut) + offset_at_this_cut,
donor_y + rect_height / 2,
donor_y - rect_height / 2,
alpha=0.5,
fill=True,
color=color,
)
mark_5_and_3(donor_y, -len(donor_before_cut), len(donor_after_cut), color)
for name, info in ti.donor_SNVs['target'].items():
x = info['position'] - ti.cut_after
if x >= 0:
x = -0.5 + offset_at_this_cut + x
else:
x = -0.5 + offset_at_this_cut + x
draw_rect(x - 0.5, x + 0.5, donor_y - rect_height / 2, donor_y + rect_height / 2, 0.2)
# Draw resected target.
resect_before = int(np.abs(np.floor(donor_x_min))) + 1
resect_after = int(np.abs(np.ceil(donor_x_max))) + 1
x_min = -resect_before - 5
x_max = resect_after + 5
before_cut = common_ti.target_sequence[:common_ti.cut_after + 1][x_min:]
after_cut = common_ti.target_sequence[common_ti.cut_after + 1:][:x_max]
for b, x in zip(before_cut[::-1], np.arange(len(before_cut))):
final_x = -x - offset_at_cut - 0.5
ax.annotate(b,
(final_x, ys['target_+']),
**kwargs,
)
if x < resect_before:
alpha = 0.3
else:
alpha = 1
ax.annotate(hits.utilities.complement(b),
(final_x, ys['target_-']),
alpha=alpha,
**kwargs,
)
for b, x in zip(after_cut, np.arange(len(after_cut))):
final_x = x + offset_at_cut + 0.5
if x < resect_after:
alpha = 0.3
else:
alpha = 1
ax.annotate(b,
(final_x, ys['target_+']),
alpha=alpha,
**kwargs,
)
ax.annotate(hits.utilities.complement(b),
(final_x, ys['target_-']),
**kwargs,
)
alpha = 0.1
draw_rect(offset_at_cut + resect_after, x_max, ys['target_+'] - rect_height / 2, ys['target_+'] + rect_height / 2, alpha)
draw_rect(0, x_max, ys['target_-'] - rect_height / 2, ys['target_-'] + rect_height / 2, alpha)
draw_rect(0, x_min, ys['target_+'] - rect_height / 2, ys['target_+'] + rect_height / 2, alpha)
draw_rect(-offset_at_cut - resect_before, x_min, ys['target_-'] - rect_height / 2, ys['target_-'] + rect_height / 2, alpha)
ax.set_xlim(x_min, x_max)
ax.set_ylim(-2, 2)
if flip_target:
ax.invert_xaxis()
ax.invert_yaxis()
ax.plot([0, 0], [ys['target_-'] - rect_height, ys['target_+'] + rect_height], color='black', linestyle='--', alpha=0.5)
mark_5_and_3(ys['target_+'], x_min, x_max, 'black')
mark_5_and_3(ys['target_-'], x_min, x_max, 'black')
ax.set_xticks([])
ax.set_yticks([])
ax.set_frame_on(False)
if draw_SNVs_on_target:
for name, info in common_ti.donor_SNVs['target'].items():
x = info['position'] - ti.cut_after
if x >= 0:
x = -0.5 + offset_at_cut + x
else:
x = -0.5 - offset_at_cut + x
for y in [ys['target_+'], ys['target_-']]:
draw_rect(x - 0.5, x + 0.5, y - rect_height / 2, y + rect_height / 2, 0.2)
for name, PAM_slice in common_ti.PAM_slices.items():
sgRNA = common_ti.sgRNA_features[name]
if sgRNA.strand == '+':
y_start = ys['target_+'] + rect_height / 4
y_end = y_start + rect_height / 3
else:
y_start = ys['target_-'] - rect_height / 4
y_end = y_start - rect_height / 3
x_start = PAM_slice.start - common_ti.cut_after - 1 + 0.2 # offset empirically determined, confused by it
x_end = PAM_slice.stop - common_ti.cut_after - 1 - 0.2
PAM_feature = common_ti.PAM_features[common_ti.target, f'{name}_PAM']
PAM_color = PAM_feature.attribute['color']
#draw_rect(x_start, x_end, ys['target_-'] - rect_height / 2, ys['target_+'] + rect_height / 2, 0.9, color=PAM_color, fill=False)
ax.plot([x_start, x_start, x_end, x_end], [y_start, y_end, y_end, y_start], color=PAM_color, linewidth=2)
x_start = sgRNA.start - common_ti.cut_after - 1 + 0.1
x_end = sgRNA.end - common_ti.cut_after - 0.1
ax.plot([x_start, x_start, x_end, x_end], [y_start, y_end, y_end, y_start], color=sgRNA.attribute['color'], linewidth=2)
#draw_rect(x_start, x_end, ys['target_-'] - rect_height / 2, ys['target_+'] + rect_height / 2, 0.5, color=sgRNA.attribute['color'], fill=False)
return fig, ax
def conversion_tracts(pool,
heatmap_genes=None,
plot_genes=None,
guides=None,
fc_ylims=None,
fc_xlims=(-4, 2),
frequency_threshold=0.002,
outcomes=None,
gene_to_sort_by='MLH1',
x_lims=None,
just_heatmaps=False,
draw_labels=True,
draw_conversion_plots=True,
gene_to_color=None,
diagram_kwargs=None,
pools_for_diagram=None,
flip_target=False,
ax_on_bottom=False,
**kwargs,
):
fracs = pool.non_targeting_fractions()
if outcomes is not None:
fracs = fracs.reindex(outcomes, fill_value=0)
if pools_for_diagram is None:
pools_for_diagram = [pool]
outcomes = [(c, s, d) for (c, s, d), f in fracs.items() if c == 'donor' and f > frequency_threshold]
fig, configuration_ax = draw_ssODN_configurations(pools_for_diagram, flip_target=flip_target, draw_SNVs_on_target=False)
xs = pool.SNV_name_to_position - pool.target_info.cut_after - 0.5
# overall incorporation frequency plot
c_ax_x_min, c_ax_x_max = configuration_ax.get_xlim()
data_width = c_ax_x_max - c_ax_x_min
c_ax_p = configuration_ax.get_position()
if x_lims is not None:
x_min, x_max = x_lims
else:
x_min = int(np.floor(min(xs))) - 2
x_max = int(np.ceil(max(xs))) + 2
left = c_ax_p.x0 + ((x_min - c_ax_x_min) / data_width * c_ax_p.width)
width = abs((x_max - x_min) / data_width * c_ax_p.width)
height = 0.5 * c_ax_p.height
gene_guides_by_activity = pool.gene_guides_by_activity()
if gene_to_color is None:
gene_to_color = {gene: repair_seq.visualize.good_colors[i] for i, gene in enumerate(heatmap_genes)}
if draw_conversion_plots:
frequency_ax = fig.add_axes([left, c_ax_p.y0 - height, width, height])
ys = pool.conversion_fractions['all_non_targeting']
frequency_ax.plot(xs, ys * 100, 'o-', color='black', linewidth=2)
frequency_ax.axvline(0, linestyle='--', color='black', alpha=0.5)
frequency_ax.set_ylim(0, max(ys * 100) * 1.1)
frequency_ax.set_ylabel('overall\nconversion\npercentage', size=12)
frequency_ax.set_xlim(x_min, x_max)
plt.setp(frequency_ax.get_xticklabels(), visible=False)
# log2 fold changes plot
f_ax_p = frequency_ax.get_position()
height = 0.75 * c_ax_p.height
gap = height * 0.1
fold_change_ax = fig.add_axes([f_ax_p.x0, f_ax_p.y0 - gap - height, f_ax_p.width, height], sharex=frequency_ax)
guide_sets = [
('negative_control',
pool.variable_guide_library.non_targeting_guides,
'non-targeting',
dict(color='black', alpha=0.2),
),
]
for gene_i, gene in enumerate(plot_genes):
guide_sets.append((gene,
gene_guides_by_activity[gene][:1],
None,
dict(color=gene_to_color[gene], alpha=0.8, linewidth=2.5, markersize=10),
),
)
max_y = 1
min_y = -2
for gene_i, (gene, gene_guides, label, gene_kwargs) in enumerate(guide_sets):
fold_change_ax.annotate(gene,
xy=(1, 1),
xycoords='axes fraction',
xytext=(5, -10 - 16 * gene_i),
textcoords='offset points',
color=gene_kwargs['color'],
size=14,
)
for i, guide in enumerate(gene_guides):
ys = pool.conversion_log2_fold_changes[guide]
max_y = np.ceil(max(max_y, max(ys)))
min_y = np.floor(min(min_y, min(ys)))
label_to_use = None
if i == 0:
if label is None:
label_to_use = gene
else:
label_to_use = label
else:
label_to_use = ''
fold_change_ax.plot(xs, ys, '.-', label=label_to_use, **gene_kwargs)
plt.setp(fold_change_ax.get_xticklabels(), visible=False)
if fc_ylims is None:
fc_ylims = (max(-6, min_y), min(5, max_y))
fold_change_ax.set_ylim(*fc_ylims)
#fold_change_ax.set_yticks(np.arange(-3, 2))
fold_change_ax.grid(alpha=0.5, axis='y')
fold_change_ax.axhline(0, color='black', alpha=0.2)
fold_change_ax.axvline(0, linestyle='--', color='black', alpha=0.5)
fold_change_ax.set_ylabel('log2 fold-change\nin conversion\nfrom non-targeting', size=12)
fc_ax_p = fold_change_ax.get_position()
# Make height such that nts are roughly square, with a slop factor for spacing between rows.
fig_width_inches, fig_height_inches = fig.get_size_inches()
#width_inches = fc_ax_p.width * fig_width_inches
width_inches = width * fig_width_inches
height_inches = width_inches * len(outcomes) / abs(x_max - x_min)
#diagram_width = fc_ax_p.width
diagram_width = width
diagram_height = height_inches / fig_height_inches * 2
#diagram_left = fc_ax_p.x0
diagram_left = left
if draw_conversion_plots:
diagram_bottom = fc_ax_p.y0 - 0.4 * fc_ax_p.height - diagram_height
else:
diagram_bottom = 0
#diagram_bottom = c_ax_p.y0 - 0.4 * c_ax_p.height - diagram_height
diagram_rect = [diagram_left, diagram_bottom, diagram_width, diagram_height]
diagram_ax = fig.add_axes(diagram_rect, sharex=frequency_ax if draw_conversion_plots else None)
if gene_to_sort_by is None:
sorted_outcomes = outcomes[::-1]
else:
sorted_outcomes = pool.sort_outcomes_by_gene_phenotype(outcomes, gene_to_sort_by)[::-1]
if diagram_kwargs is None:
diagram_kwargs = {
'flip_if_reverse': False,
}
diagram_kwargs.update(dict(
window=(x_min, x_max),
preserve_x_lims=True,
shift_x=-0 if just_heatmaps else 0.5,
draw_donor_on_top=True,
draw_wild_type_on_top=True,
))
# Note: plot does a weird flip of outcomes
diagram_grid = repair_seq.visualize.outcome_diagrams.DiagramGrid(sorted_outcomes[::-1],
pool.target_info,
diagram_ax=diagram_ax,
ax_on_bottom=ax_on_bottom,
**diagram_kwargs,
)
diagram_grid.add_ax('log10 frequency',
side=kwargs.get('frequency_side', 'left'),
width_multiple=4,
gap_multiple=2.5,
title='Percentage of\noutcomes for\n non-targeting\nsgRNAs' if draw_labels else '',
title_size=12,
)
diagram_grid.add_ax('log2 fold change',
side='right',
width_multiple=7,
gap_multiple=1.5,
title='Log$_2$ fold change\nfrom non-targeting' if draw_labels else '',
title_size=12,
)
log10_frequencies = np.log10(pool.non_targeting_fractions().loc[sorted_outcomes])
diagram_grid.plot_on_ax('log10 frequency', log10_frequencies, marker='o', markersize=2.5, linewidth=1, line_alpha=0.9, marker_alpha=0.9, color='black', clip_on=False)
x_min, x_max = np.log10(0.95 * frequency_threshold), np.log10(0.21)
diagram_grid.axs_by_name['log10 frequency'].set_xlim(x_min, x_max)
diagram_grid.style_log10_frequency_ax('log10 frequency')
if kwargs.get('frequency_side', 'left') == 'left':
diagram_grid.axs_by_name['log10 frequency'].invert_xaxis()
fcs = pool.log2_fold_changes().loc[sorted_outcomes]
for gene in plot_genes:
if gene == 'DNA2':
# override since growth phenotype leads to low UMI counts for strong guides
guide = 'DNA2_1'
elif gene == 'MLH1':
guide = 'MLH1_1'
elif gene == 'PMS2':
guide = 'PMS2_1'
elif gene == 'MSH6':
guide = 'MSH6_1'
elif gene == 'RBBP8':
guide = 'RBBP8_1'
elif gene == 'NBN':
guide = 'NBN_1',
elif gene == 'MRE11':
guide = 'MRE11_1'
else:
guide = gene_guides_by_activity[gene][0]
print(guide)
diagram_grid.plot_on_ax('log2 fold change', fcs[guide], marker='o', markersize=2.5, marker_alpha=0.9, line_alpha=0.9, linewidth=1, color=gene_to_color[gene], clip_on=False)
diagram_grid.style_fold_change_ax('log2 fold change')
diagram_grid.axs_by_name['log2 fold change'].set_xlim(*fc_xlims)
for gene_i, gene in enumerate(heatmap_genes):
if gene == 'DNA2':
# override since growth phenotype leads to low UMI counts for strong guides
guides = ['DNA2_1', 'DNA2_3']
elif gene == 'MLH1':
guides = ['MLH1_1', 'MLH1_2']
elif gene == 'PMS2':
guides = ['PMS2_1', 'PMS2_2']
elif gene == 'MSH6':
guides = ['MSH6_1', 'MSH6_2']
elif gene == 'RBBP8':
guides = ['RBBP8_1', 'RBBP8_2']
elif gene == 'NBN':
guides = ['NBN_1', 'NBN_2']
elif gene == 'MRE11':
guides = ['MRE11_1', 'MRE11_3']
else:
guides = gene_guides_by_activity[gene][:2]
vals = fcs[guides]
if gene_i == 0:
gap_multiple = 1
else:
gap_multiple = 0.25
heatmap_ax = diagram_grid.add_heatmap(vals, f'heatmap {gene}', gap_multiple=gap_multiple, color=gene_to_color[gene])
if not draw_labels:
heatmap_ax.set_xticklabels([])
if just_heatmaps:
fig.delaxes(configuration_ax)
if draw_conversion_plots:
fig.delaxes(frequency_ax)
fig.delaxes(fold_change_ax)
return fig, diagram_grid | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/visualize/conversion_tracts.py | 0.422981 | 0.496521 | conversion_tracts.py | pypi |
import copy
import pickle
import string
import bokeh.io
import bokeh.models
import bokeh.plotting
import numpy as np
from hits.visualize.callback import build_js_callback
def scatter(data_source,
outcome_names=None,
plot_width=2000,
plot_height=800,
initial_guides=None,
initial_genes=None,
save_as=None,
):
if save_as is not None and save_as != 'layout':
bokeh.io.output_file(save_as)
if initial_guides is not None and initial_genes is not None:
raise ValueError('can only specify one of initial_guides or initial_genes')
if initial_guides is None:
initial_guides = []
if isinstance(data_source, str):
with open(data_source, 'rb') as fh:
data = pickle.load(fh)
else:
data = copy.deepcopy(data_source)
pool_names = data['pool_names']
if outcome_names is None:
outcome_names = data['outcome_names']
guides_df = data['guides_df']
nt_fractions = data['nt_percentages']
initial_dataset = data['initial_dataset']
initial_outcome = data['initial_outcome']
guides_df.columns = ['_'.join(t) if t[1] != '' else t[0] for t in data['guides_df'].columns.values]
table_keys = [
'frequency',
'percentage',
'ys',
'total_UMIs',
'gene_p_up',
'gene_p_down',
'log2_fold_change',
]
for key in table_keys:
guides_df[key] = guides_df[f'{initial_dataset}_{initial_outcome}_{key}']
scatter_source = bokeh.models.ColumnDataSource(data=guides_df, name='scatter_source')
scatter_source.data[guides_df.index.name] = guides_df.index
if initial_genes is not None:
initial_indices = np.array(guides_df.query('gene in @initial_genes')['x'])
else:
initial_indices = np.array(guides_df.loc[initial_guides]['x'])
scatter_source.selected.indices = initial_indices
filtered_data = {k: [scatter_source.data[k][i] for i in initial_indices] for k in scatter_source.data}
filtered_source = bokeh.models.ColumnDataSource(data=filtered_data, name='filtered_source')
x_min = -1
x_max = len(guides_df)
y_min = 0
y_max = guides_df['percentage'].max() * 1.2
tools = [
'reset',
'undo',
'pan',
'box_zoom',
'box_select',
'tap',
'wheel_zoom',
'save',
]
fig = bokeh.plotting.figure(plot_width=plot_width, plot_height=plot_height,
tools=tools, active_drag='box_select', active_scroll='wheel_zoom',
)
fig.toolbar.logo = None
fig.x_range = bokeh.models.Range1d(x_min, x_max, name='x_range')
fig.y_range = bokeh.models.Range1d(y_min, y_max, name='y_range')
x_range_callback = build_js_callback(__file__, 'screen_range', format_kwargs={'lower_bound': x_min, 'upper_bound': x_max})
y_range_callback = build_js_callback(__file__, 'screen_range', format_kwargs={'lower_bound': 0, 'upper_bound': 100})
for prop in ['start', 'end']:
fig.x_range.js_on_change(prop, x_range_callback)
fig.y_range.js_on_change(prop, y_range_callback)
circles = fig.circle(x='x',
y='percentage',
source=scatter_source,
color='color', selection_color='color', nonselection_color='color',
alpha=0.8,
selection_line_alpha=0, nonselection_line_alpha=0, line_alpha=0,
size=5,
)
tooltips = [
('CRISPRi sgRNA', '@guide'),
('Frequency of outcome', '@frequency'),
('Log2 fold change from nt', '@log2_fold_change'),
('Total UMIs', '@total_UMIs'),
]
hover = bokeh.models.HoverTool(renderers=[circles])
hover.tooltips = tooltips
fig.add_tools(hover)
confidence_intervals = fig.multi_line(xs='xs', ys='ys',
source=scatter_source,
color='color',
selection_color='color',
nonselection_color='color',
alpha=0.4,
)
interval_button = bokeh.models.widgets.Toggle(label='Show confidence intervals', active=True)
# Slightly counterintuitive - directionality of link matters here.
# Add the link to the emitter of the signal, with the listener as the arg.
interval_button.js_link('active', confidence_intervals, 'visible')
labels = bokeh.models.LabelSet(x='x', y='percentage', text='guide',
source=filtered_source,
level='glyph',
x_offset=4, y_offset=0,
text_font_size='6pt',
text_color='color',
text_baseline='middle',
name='labels',
)
fig.add_layout(labels)
fig.xgrid.visible = False
fig.ygrid.visible = False
dataset_menu = bokeh.models.widgets.MultiSelect(options=pool_names, value=[initial_dataset], name='dataset_menu', title='Screen condition:', size=len(pool_names) + 2, width=400)
outcome_menu = bokeh.models.widgets.MultiSelect(options=outcome_names, value=[initial_outcome], name='outcome_menu', title='Outcome category:', size=len(outcome_names) + 2, width=400)
nt_fraction = bokeh.models.Span(location=nt_fractions[f'{initial_dataset}_{initial_outcome}'], dimension='width', line_alpha=0.5)
fig.add_layout(nt_fraction)
cutoff_slider = bokeh.models.Slider(start=-10, end=-2, value=-5, step=1, name='cutoff_slider', title='log10 p-value significance threshold')
filter_buttons = {}
filter_buttons['down'] = bokeh.models.widgets.Button(label='Filter to genes that significantly decrease')
filter_buttons['up'] = bokeh.models.widgets.Button(label='Filter to genes that significantly increase')
text_input = bokeh.models.TextInput(title='Search sgRNAs:', name='search')
fig.outline_line_color = 'black'
first_letters = [g[0] for g in guides_df.index]
x_tick_labels = {first_letters.index(c): c for c in string.ascii_uppercase if c in first_letters}
x_tick_labels[first_letters.index('n')] = 'nt'
fig.xaxis.ticker = sorted(x_tick_labels)
fig.xaxis.major_label_overrides = x_tick_labels
table_col_names = [
('guide', 'CRISPRi sgRNA', 50),
('gene', 'Gene', 50),
('frequency', 'Frequency of outcome', 50),
('log2_fold_change', 'Log2 fold change', 50),
('total_UMIs', 'Total UMIs', 50),
]
columns = []
for col_name, col_label, width in table_col_names:
width = 50
if col_name == 'gene':
formatter = bokeh.models.widgets.HTMLTemplateFormatter(template='<a href="https://www.genecards.org/cgi-bin/carddisp.pl?gene=<%= value %>" target="_blank"><%= value %></a>')
elif col_name == 'log2_fold_change':
formatter = bokeh.models.widgets.NumberFormatter(format='0.00')
elif col_name == 'total_UMIs':
formatter = bokeh.models.widgets.NumberFormatter(format='0,0')
elif col_name == 'frequency':
formatter = bokeh.models.widgets.NumberFormatter(format='0.00%')
else:
formatter = bokeh.models.widgets.StringFormatter()
column = bokeh.models.widgets.TableColumn(field=col_name,
title=col_label,
formatter=formatter,
width=width,
)
columns.append(column)
save_button = bokeh.models.widgets.Button(label='Save table', name='save_button')
table = bokeh.models.widgets.DataTable(source=filtered_source,
columns=columns,
width=600,
height=300,
sortable=False,
reorderable=False,
name='table',
index_position=None,
)
fig.xaxis.axis_label = 'CRISPRi sgRNAs (ordered alphabetically)'
fig.yaxis.axis_label = 'Frequency of outcome'
for axis in (fig.xaxis, fig.yaxis):
axis.axis_label_text_font_size = '14pt'
axis.axis_label_text_font_style = 'bold'
fig.yaxis.formatter = bokeh.models.PrintfTickFormatter(format="%6.2f%%")
fig.yaxis.major_label_text_font = 'courier'
fig.yaxis.major_label_text_font_style = 'bold'
title = bokeh.models.Title(text=f'Outcome category: {initial_outcome}', text_font_size='14pt', name='subtitle')
subtitle = bokeh.models.Title(text=f'Screen condition: {initial_dataset}', text_font_size='14pt', name='title')
fig.add_layout(title, 'above')
fig.add_layout(subtitle, 'above')
top_widgets = bokeh.layouts.column([bokeh.layouts.Spacer(height=70),
dataset_menu,
outcome_menu,
text_input,
])
bottom_widgets = bokeh.layouts.column([interval_button,
cutoff_slider,
filter_buttons['up'],
filter_buttons['down'],
save_button,
])
first_row = bokeh.layouts.row([fig, top_widgets])
second_row = bokeh.layouts.row([bokeh.layouts.Spacer(width=90), table, bottom_widgets])
final_layout = bokeh.layouts.column([first_row, bokeh.layouts.Spacer(height=50), second_row])
menu_js = build_js_callback(__file__, 'screen_menu',
args=dict(
dataset_menu=dataset_menu,
outcome_menu=outcome_menu,
scatter_source=scatter_source,
filtered_source=filtered_source,
y_range=fig.y_range,
title=title,
subtitle=subtitle,
nt_fraction=nt_fraction,
),
format_kwargs={'nt_fractions': str(nt_fractions)},
)
for menu in [dataset_menu, outcome_menu]:
menu.js_on_change('value', menu_js)
selection_callback = build_js_callback(__file__, 'screen_scatter_selection',
args=dict(
scatter_source=scatter_source,
filtered_source=filtered_source,
),
)
scatter_source.selected.js_on_change('indices', selection_callback)
search_callback = build_js_callback(__file__, 'screen_search',
args=dict(
scatter_source=scatter_source,
),
)
text_input.js_on_change('value', search_callback)
for direction in filter_buttons:
callback = build_js_callback(__file__, 'screen_significance_filter',
args=dict(
scatter_source=scatter_source,
cutoff_slider=cutoff_slider,
),
format_kwargs=dict(
direction=direction,
),
)
filter_buttons[direction].js_on_click(callback)
save_callback = build_js_callback(__file__, 'scatter_save_button',
args=dict(
filtered_source=filtered_source,
),
format_kwargs={'column_names': table_col_names},
)
save_button.js_on_click(save_callback)
if save_as == 'layout':
return final_layout
elif save_as is not None:
bokeh.io.save(final_layout)
else:
bokeh.io.show(final_layout) | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/visualize/interactive/alphabetical.py | 0.475605 | 0.297773 | alphabetical.py | pypi |
import bokeh.plotting
import matplotlib.colors
import matplotlib.cm
import numpy as np
import pandas as pd
from hits.visualize import callback
from repair_seq import visualize
def plot(clusterer):
all_l2fcs = clusterer.all_log2_fold_changes
corrs = all_l2fcs.corr()
initial_guide = 'POLQ_1'
corrs['x'] = np.arange(len(corrs))
corrs['y'] = corrs[initial_guide]
sorted_pairs = sorted(enumerate(corrs['y']), key=lambda pair: pair[1], reverse=True)
corrs['ranked_indices'] = [pair[0] for pair in sorted_pairs]
corrs['color'] = visualize.targeting_guide_color
corrs.loc[clusterer.guide_library.non_targeting_guides, 'color'] = visualize.nontargeting_guide_color
corrs.index.name = 'sgRNA'
corrs_source = bokeh.models.ColumnDataSource(data=corrs)
data = clusterer.outcome_embedding.copy()
def bind_hexer(scalar_mappable):
def to_hex(v):
return matplotlib.colors.to_hex(scalar_mappable.to_rgba(v))
return to_hex
norm = matplotlib.colors.Normalize(vmin=-2, vmax=2)
cmap = visualize.fold_changes_cmap
l2fcs_sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
l2fcs_to_hex = bind_hexer(l2fcs_sm)
norm = matplotlib.colors.Normalize(vmin=0, vmax=30)
cmap = matplotlib.cm.get_cmap('Purples')
deletion_length_sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
deletion_length_to_hex = bind_hexer(deletion_length_sm)
norm = matplotlib.colors.Normalize(vmin=-3, vmax=-1)
cmap = matplotlib.cm.get_cmap('YlOrBr')
log10_fraction_sm = matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
log10_fraction_to_hex = bind_hexer(log10_fraction_sm)
colors = {}
for guide in clusterer.guide_library.guides:
values = all_l2fcs[guide]
hex_values = [matplotlib.colors.to_hex(row) for row in l2fcs_sm.to_rgba(values)]
colors[f'color_{guide}'] = hex_values
target_colors, target_to_color = clusterer.sgRNA_colors()
target_to_hex_color = {target: matplotlib.colors.to_hex(color) for target, color in target_to_color.items()}
colors['target'] = [matplotlib.colors.to_hex(row) for row in target_colors]
colors = pd.DataFrame(colors, index=data.index)
relevant_data = clusterer.outcome_embedding[['x', 'y', 'category_colors', 'MH length', 'deletion length', 'log10_fraction']]
all_colors = pd.concat([relevant_data, colors], axis=1)
all_colors['first_color'] = all_colors[f'color_{initial_guide}']
all_colors['blank'] = ['#FFFFFF' for _ in all_colors.index]
all_colors['second_color'] = all_colors['blank']
color_source = bokeh.models.ColumnDataSource(data=all_colors)
color_mappers = {
'none': None,
'MH length': bokeh.models.LinearColorMapper(low=-0.5, high=3.5, palette=bokeh.palettes.viridis(4)[::-1], low_color='white'),
'l2fcs': bokeh.models.LinearColorMapper(low=-2, high=2, palette=[l2fcs_to_hex(v) for v in np.linspace(-2, 2, 500)]),
'deletion length': bokeh.models.LinearColorMapper(low=0, high=30, palette=[deletion_length_to_hex(v) for v in np.linspace(0, 30, 500)]),
'log10_fraction': bokeh.models.LinearColorMapper(low=-3, high=-1, palette=[log10_fraction_to_hex(v) for v in np.linspace(-3, -1, 500)]),
}
figs = {}
big_frame_size = 600
small_frame_size = 450
for name, color_key, transform_key in [('l2fcs', 'first_color', 'none'),
('category', 'category_colors', 'none'),
('target', 'target', 'none'),
('MH length', 'MH length', 'MH length'),
('deletion length', 'deletion length', 'deletion length'),
('log10_fraction', 'log10_fraction', 'log10_fraction'),
]:
fig = bokeh.plotting.figure(frame_width=big_frame_size, frame_height=big_frame_size, min_border=0)
fig.scatter('x', 'y',
source=color_source,
fill_color={'field': color_key, 'transform': color_mappers[transform_key]},
line_color=None,
size=7,
)
fig.grid.visible = False
fig.xaxis.visible = False
fig.yaxis.visible = False
fig.toolbar_location = None
fig.toolbar.active_drag = None
fig.outline_line_color = 'black'
fig.outline_line_alpha = 0.5
figs[name] = fig
color_menu = bokeh.models.widgets.Select(title='CRISPRi sgRNA',
options=list(clusterer.guide_library.guides),
value=initial_guide,
width=150,
)
corrs_fig = bokeh.plotting.figure(plot_width=big_frame_size + small_frame_size + 50 + 20,
plot_height=320,
toolbar_location=None,
active_drag=None,
min_border=0,
)
corrs_scatter = corrs_fig.scatter(x='x', y='y', source=corrs_source,
fill_color='color',
line_color=None,
size=6,
)
top_indices = corrs['ranked_indices'].iloc[:10]
top_guides = corrs.iloc[top_indices].index.values
filtered_corrs = corrs.loc[top_guides, ['x', 'y', 'color']].copy()
filtered_corrs_source = bokeh.models.ColumnDataSource(data=filtered_corrs)
labels = bokeh.models.LabelSet(x='x',
y='y',
text='sgRNA',
level='glyph',
x_offset=7,
y_offset=0,
source=filtered_corrs_source,
text_font_size='8pt',
text_baseline='middle',
)
corrs_fig.add_layout(labels)
corrs_fig.xaxis.axis_label = 'CRISPRi sgRNAs (ordered alphabetically)'
corrs_fig.xaxis.axis_label_text_font_style = 'normal'
corrs_fig.xaxis.axis_label_text_font_size = '14pt'
corrs_fig.yaxis.axis_label = f'Correlation with {initial_guide}'
corrs_fig.yaxis.axis_label_text_font_style = 'normal'
corrs_fig.yaxis.axis_label_text_font_size = '14pt'
line_kwargs = dict(
color='black',
nonselection_color='black',
line_width=1,
)
x_bounds = [-5, max(corrs['x']) + 5]
corrs_fig.line(x=x_bounds, y=[0, 0], **line_kwargs)
xs = [
corrs.loc[initial_guide, 'x'],
corrs.loc[initial_guide, 'x'],
100,
100,
]
ys = [
corrs.loc[initial_guide, 'y'] + 0.05,
1.12,
1.12,
1.2,
]
left_line_source = bokeh.models.ColumnDataSource(pd.DataFrame({'x': xs, 'y': ys}))
corrs_fig.line(x='x', y='y', source=left_line_source, line_alpha=0.5, **line_kwargs)
xs = [
corrs.loc[initial_guide, 'x'],
corrs.loc[initial_guide, 'x'],
300,
300,
]
ys = [
corrs.loc[initial_guide, 'y'] + 0.05,
1.08,
1.08,
1.2,
]
right_line_source = bokeh.models.ColumnDataSource(pd.DataFrame({'x': xs, 'y': ys}))
right_line = corrs_fig.line(x='x', y='y', source=right_line_source, visible=False, line_alpha=0.5, **line_kwargs)
corrs_fig.y_range = bokeh.models.Range1d(-0.7, 1.2)
corrs_fig.yaxis.bounds = (-0.6, 1)
corrs_fig.x_range = bokeh.models.Range1d(*x_bounds)
corrs_fig.xgrid.visible = False
corrs_fig.xaxis.ticker = []
corrs_fig.yaxis.ticker = np.linspace(-0.6, 1, 9)
corrs_fig.outline_line_color = None
corrs_fig.xaxis.ticker = []
corrs_fig.xaxis.axis_line_color = None
nt_text = pd.DataFrame([
{'x': max(corrs['x']) - len(clusterer.guide_library.non_targeting_guides) / 2,
'y': -0.4,
'text': 'non-\ntargeting',
'color': visualize.nontargeting_guide_color,
}
])
corrs_fig.text(x='x', y='y', text='text', source=nt_text, color='color', text_align='center', text_font_style='bold', text_font_size={'value': '12px'})
second_scatter = bokeh.plotting.figure(frame_width=small_frame_size,
frame_height=small_frame_size,
toolbar_location=None,
min_border=0,
outline_line_color=None,
)
second_scatter.scatter('x', 'y', source=color_source, fill_color='second_color', line_color=None, size=6)
second_scatter.grid.visible = False
second_scatter.xaxis.visible = False
second_scatter.yaxis.visible = False
common_colorbar_kwargs = dict(
orientation='horizontal',
width=200,
location=(0.1 * big_frame_size, 0.6 * big_frame_size),
major_tick_out=5,
major_tick_in=0,
margin=100,
padding=10,
title_text_font_style='normal',
background_fill_color=None,
)
color_bars = {
'l2fcs': bokeh.models.ColorBar(
color_mapper=color_mappers['l2fcs'],
major_tick_line_color='black',
ticker=bokeh.models.FixedTicker(ticks=[-2, -1, 0, 1, 2]),
major_label_overrides={2: '≥2', -2: '≤-2'},
title=f'Log₂ fold change in outcome frequency\nfor the selected CRISPRi sgRNA',
**common_colorbar_kwargs,
),
'MH length': bokeh.models.ColorBar(
color_mapper=color_mappers['MH length'],
major_tick_line_color=None,
ticker=bokeh.models.FixedTicker(ticks=[0, 1, 2, 3]),
major_label_overrides={3: '≥3'},
title='Flanking microhomology (nts)',
**common_colorbar_kwargs,
),
'deletion length': bokeh.models.ColorBar(
color_mapper=color_mappers['deletion length'],
major_tick_line_color='black',
ticker=bokeh.models.FixedTicker(ticks=[0, 10, 20, 30]),
major_label_overrides={30: '≥30'},
title='Deletion length (nts)',
**common_colorbar_kwargs,
),
'log10_fraction': bokeh.models.ColorBar(
color_mapper=color_mappers['log10_fraction'],
major_tick_line_color='black',
ticker=bokeh.models.FixedTicker(ticks=[-3, -2, -1]),
major_label_overrides={-3: '≤0.1%', -2: '1%', -1: '≥10%'},
title='Baseline frequency of outcome',
**common_colorbar_kwargs,
),
}
for key in color_bars:
figs[key].add_layout(color_bars[key])
# Draw title and various text labels on big scatter plots.
common_text_kwargs = dict(
x='x',
y='y',
text='text',
text_align='center',
color='color',
)
category_text = pd.DataFrame([
{'x': -4, 'y': -3.5, 'text': 'bidirectional\ndeletions', 'color': visualize.category_colors['SpCas9']['deletion, bidirectional']},
{'x': 3, 'y': 5.5, 'text': 'insertions\nI', 'color': visualize.category_colors['SpCas9']['insertion']},
{'x': 4.5, 'y': 3.6, 'text': 'insertions\nII', 'color': visualize.category_colors['SpCas9']['insertion']},
{'x': -7, 'y': 1.7, 'text': 'insertions\nIII', 'color': visualize.category_colors['SpCas9']['insertion']},
{'x': -7, 'y': -2.2, 'text': 'insertions\nwith\ndeletions', 'color': visualize.category_colors['SpCas9']['insertion with deletion']},
{'x': 6.4, 'y': 2, 'text': 'unedited', 'color': visualize.category_colors['SpCas9']['wild type']},
{'x': 9.9, 'y': 5.2, 'text': 'capture of\ngenomic\nsequence\nat break', 'color': visualize.category_colors['SpCas9']['genomic insertion']},
{'x': 9.9, 'y': 3.8, 'text': '≤75 nts', 'color': visualize.category_colors['SpCas9']['genomic insertion']},
{'x': 10.4, 'y': 1.9, 'text': '>75 nts', 'color': visualize.category_colors['SpCas9']['genomic insertion']},
{'x': 10, 'y': -0.5, 'text': 'deletions\nconsistent\nwith either\nside', 'color': visualize.category_colors['SpCas9']['deletion, ambiguous']},
{'x': 2.5, 'y': -4, 'text': 'deletions\non only\nPAM-distal\nside', 'color': visualize.category_colors['SpCas9']['deletion, PAM-distal']},
{'x': 1, 'y': 1.2, 'text': 'deletions\non only\nPAM-proximal\nside', 'color': visualize.category_colors['SpCas9']['deletion, PAM-proximal']},
])
figs['category'].text(source=category_text, text_font_size={'value': '14px'}, **common_text_kwargs)
title_text = pd.DataFrame([{'x': -3, 'y': 5, 'text': 'Cas9 outcome\nembedding', 'color': 'black'}])
for fig in figs.values():
fig.text(source=title_text, text_font_size={'value': '26px'}, **common_text_kwargs)
target_text = pd.DataFrame([
{'x': -3, 'y': 4.5 - 0.5 * i, 'text': f'Cas9 target {i}', 'color': target_to_hex_color[f'SpCas9 target {i}']}
for i in [1, 2, 3, 4]
])
figs['target'].text(source=target_text, text_font_size={'value': '18px'}, **common_text_kwargs)
# Layout the components of the dashboard.
def row(*children):
return bokeh.models.layouts.Row(children=list(children))
def col(*children):
return bokeh.models.layouts.Column(children=list(children))
tabs = [
bokeh.models.Panel(child=figs['category'], title='Category'),
bokeh.models.Panel(child=figs['target'], title='Target site'),
bokeh.models.Panel(child=figs['MH length'], title='MH length'),
bokeh.models.Panel(child=figs['deletion length'], title='Deletion length'),
bokeh.models.Panel(child=figs['log10_fraction'], title='Baseline frequency'),
bokeh.models.Panel(child=figs['l2fcs'], title='Log₂ fold changes'),
]
tabs = bokeh.models.Tabs(tabs=tabs, active=5)
color_by = bokeh.models.Div(text='Color by:')
def v_gap(height):
return bokeh.layouts.Spacer(height=height)
def h_gap(width):
return bokeh.layouts.Spacer(width=width)
#final_layout = row([color_by, tabs, col([row([color_menu, h_gap(20), second_scatter]), v_gap(20), corrs_fig])])
gap_above = 40
menu_height = 59 # Empirically determined
tab_menu_height = 29 # Empirically determined
gap_between = big_frame_size + tab_menu_height - small_frame_size - gap_above - menu_height
final_layout = col(row(col(v_gap(2), color_by), tabs, h_gap(20), col(v_gap(gap_above), color_menu, v_gap(gap_between), second_scatter)),
row(corrs_fig),
)
# Build and attach callbacks at the end, so all necessary args are available.
menu_callback = callback.build_js_callback(__file__, 'UMAP_menu',
args=dict(color_source=color_source,
corrs_source=corrs_source,
filtered_corrs_source=filtered_corrs_source,
left_line_source=left_line_source,
color_menu=color_menu,
corrs_y_axis=corrs_fig.yaxis,
tabs=tabs,
),
)
color_menu.js_on_change('value', menu_callback)
hover_callback = callback.build_js_callback(__file__, 'UMAP_hover',
args=dict(color_source=color_source,
corrs_source=corrs_source,
filtered_corrs_source=filtered_corrs_source,
right_line_source=right_line_source,
second_scatter=second_scatter,
right_line=right_line,
),
)
hover = bokeh.models.HoverTool(renderers=[corrs_scatter],
callback=hover_callback,
)
#hover.tooltips = [
# (corrs.index.name, '@{{{0}}}'.format(corrs.index.name)),
#]
hover.tooltips = None
corrs_fig.add_tools(hover)
return final_layout | /repair_seq-1.0.3.tar.gz/repair_seq-1.0.3/repair_seq/visualize/interactive/UMAP.py | 0.536799 | 0.416144 | UMAP.py | pypi |
# repassh
SSH wrapper to load private keys via `ssh-add` when they are first needed, with port knocking support.
Heavily based on [ssh-ident](https://github.com/ccontavalli/ssh-ident).
## Usage
Use this script to start ssh-agents and load ssh keys on demand,
when they are first needed.
All you have to do is modify your .bashrc to have:
```
alias ssh='/path/to/repassh'
```
or add a link to `repassh` from a directory in your PATH, for example:
```
ln -s /path/to/repassh ~/bin/ssh
```
If you use scp or rsync regularly, you should add a few more lines described
below.
In any case, `repassh`:
- will start ssh-agent and load the keys you need the first time you
actually need them, once. No matter how many terminals, ssh or login
sessions you have, no matter if your home is shared via NFS.
- can prepare and use a different agent and different set of keys depending
on the host you are connecting to, or the directory you are using ssh
from.
This allows for isolating keys when using agent forwarding with different
sites (eg, university, work, home, secret evil internet identity, ...).
It also allows to use multiple accounts on sites like github, unfuddle
and gitorious easily.
- allows to specify different options for each set of keys. For example, you
can provide a -t 60 to keep keys loaded for at most 60 seconds. Or -c to
always ask for confirmation before using a key.
- performs port knocking before attempting to connect via `ssh` if configured.
## Installation
All you need to run `repassh` is a standard installation of python >= 3.6.
To install it, run:
```
pip install repassh
```
Then you can use the `repassh` command just as you'd use `ssh`.
## Alternatives
In `.bashrc` you can define an alias:
```
alias ssh=/path/to/repassh
```
then all you have to do is:
```
ssh somewhere
```
`repassh` will be called instead of `ssh`, and it will:
- check if `ssh-agent` is running. If not, it will start one.
- try to load all the keys in `~/.ssh`, if not loaded.
If you use `ssh` again, `repassh` will reuse the same agent
and the same keys.
## About scp, rsync, and friends
`scp`, `rsync`, and most similar tools internally invoke `ssh`. If you don't tell
them to use `repassh` instead, key loading won't work. There are a few ways
to solve the problem:
### Rename or link
Rename `repassh` to `ssh` or create a symlink `ssh` pointing to
`repassh` in a directory in your PATH before `/usr/bin` or `/bin`.
For example:
```
ln -s /path/to/repassh ~/bin/ssh
export PATH="~/bin:$PATH"
```
Make sure `echo $PATH` shows `~/bin` *before* `/usr/bin` or `/bin`. You
can verify this is working as expected with `which ssh`, which should
show `~/bin/ssh`.
This works for `rsync` and `git`, among others, but not for `scp` and `sftp`, as
these do not look for `ssh` in your `PATH` but use a hard-coded path to the
binary.
If you want to use `repassh` with `scp` or `sftp`, you can simply create
symlinks for them as well:
```
ln -s /path/to/repassh ~/bin/scp
ln -s /path/to/repassh ~/bin/sftp
```
### More aliases
Add a few more aliases in your .bashrc file, for example:
```
alias scp='BINARY_SSH=scp /path/to/repassh'
alias rsync='BINARY_SSH=rsync /path/to/repassh'
...
```
The first alias will make the `scp` command invoke `repassh` instead,
but tell `repassh` to invoke `scp` instead of the plain `ssh` command
after loading the necessary agents and keys.
Note that aliases don't work from scripts - if you have any script that
you expect to use with `repassh`, you may prefer the first method, or you will
need to update the script accordingly.
### Tell other programs to use `repassh` instead of `ssh`
Use command specific methods to force them to use `repassh` instead of
`ssh`, for example:
```
rsync -e '/path/to/repassh' ...
scp -S '/path/to/repassh' ...
```
## Config file with multiple identities
To have multiple identities:
1. create a `$XDG_CONFIG_HOME/repassh/config.json` file. In this file, you need to tell `repassh`
which identities to use and when. The file should be a valid JSON (ignore/remove the lines
starting with #, they are comments, but JSON does not have comments):
```
{
# Specifies which identity to use depending on the path I'm running ssh
# from.
# For example: ("mod-xslt", "personal") means that for any path that
# contains the word "mod-xslt", the "personal" identity should be used.
# This is optional - don't include any MATCH_PATH if you don't need it.
"MATCH_PATH": [
["mod-xslt", "personal"],
["repassh", "personal"],
["opt/work", "work"],
["opt/private", "secret"]
],
# If any of the ssh arguments have 'cweb' in it, the 'personal' identity
# has to be used. For example: "ssh myhost.cweb.com" will have cweb in
# argv, and the "personal" identity will be used.
# This is optional - don't include any MATCH_ARGV if you don't
# need it.
"MATCH_ARGV": [
("cweb", "personal"),
("corp", "work")
],
# KNOCK_PATH and KNOCK_ARGV work the same way as MATCH_PATH and MATCH_ARGV,
# but instead of identity name you can provide port knocking configuration
# executed before the ssh connection attempt.
# Parameters:
# - host: hostname
# - ports: sequence of ports
# - delay: time to wait between knocks (ms) [0]
# - timeout: timeout for knocks (ms) [100]
# - use_udp: set for UDP knocks, TCP is used by default
"KNOCK_ARGV": [
[
"remotehost",
{
"host": "remotehost.domain.org",
"ports": [123, 234, 345],
"delay": 50
}
]
],
# Note that if no match is found, the DEFAULT_IDENTITY is used. This is
# generally your loginname, no need to change it.
# This is optional - don't include any DEFAULT_IDENTITY if you don't
# need it.
# "DEFAULT_IDENTITY": "foo",
# Use running `ssh-agent`, true by default
# If `SSH_AUTH_SOCK` and `SSH_AGENT_PID` environment variables are set
# and the agent responds then it will be used instead of executing a new
# one based on identity matching.
# If the agent does not respond, a new one is started just like
# `USE_RUNNING_AGENT` would be false.
# "USE_RUNNING_AGENT": true,
# This is optional - don't include any SSH_ADD_OPTIONS if you don't
# need it.
"SSH_ADD_OPTIONS": {
# Regardless, ask for confirmation before using any of the
# work keys.
"work": "-c",
# Forget about secret keys after ten minutes. repassh will
# automatically ask you your passphrase again if they are needed.
"secret": "-t 600"
},
# This is optional - don't include any SSH_OPTIONS if you don't
# need it.
# Otherwise, provides options to be passed to 'ssh' for specific
# identities.
"SSH_OPTIONS": {
# Disable forwarding of the agent, but enable X forwarding,
# when using the work profile.
"work": "-Xa",
# Always forward the agent when using the secret identity.
"secret": "-A"
},
# Options to pass to ssh by default.
# If you don't specify anything, UserRoaming=no is passed, due
# to CVE-2016-0777. Leave it empty to disable this.
"SSH_DEFAULT_OPTIONS": "-oUseRoaming=no",
# Which options to use by default if no match with SSH_ADD_OPTIONS
# was found. Note that repassh hard codes -t 7200 to prevent your
# keys from remaining in memory for too long.
"SSH_ADD_DEFAULT_OPTIONS": "-t 7200",
# Output verbosity
# valid values are:
# LOG_ERROR = 1, LOG_WARN = 2, LOG_INFO = 3, LOG_DEBUG = 4
"VERBOSITY": 3
}
```
2. Create the directory where all the identities and agents
will be kept:
```
mkdir -p ~/.ssh/identities; chmod u=rwX,go= -R ~/.ssh
```
3. Create a directory for each identity, for example:
```
mkdir -p ~/.ssh/identities/personal
mkdir -p ~/.ssh/identities/work
mkdir -p ~/.ssh/identities/secret
```
4. Generate (or copy) keys for those identities:
```
# Default keys are for my personal account
$ cp ~/.ssh/id_rsa* ~/.ssh/identities/personal
# Generate keys to be used for work only, rsa
$ ssh-keygen -t rsa -b 4096 -f ~/.ssh/identities/work/id_rsa
...
```
Now if you run:
```
$ ssh corp.mywemployer.com
```
`repassh` will be invoked and:
1. checks `ssh` argv, determine that the *work* identity has to be used.
2. checks `~/.ssh/agents` for a *work* agent loaded. If there is no
agent, it will prepare one.
3. checks `~/.ssh/identities/work/` for a list of keys to load for this
identity. It will try to load any key that is not already loaded in
the agent.
4. finally run `ssh` with the environment setup such that it will have
access only to the agent for the identity work, and the corresponding
keys.
Note that `repassh` needs to access both your private and public keys. Note
also that it identifies public keys by the .pub extension. All files in your
identities subdirectories will be considered keys.
If you want to only load keys that have "key" in the name, you can add
to your `config.json`:
```
PATTERN_KEYS = "key"
```
The default is:
```
PATTERN_KEYS = r"/(id_.*|identity.*|ssh[0-9]-.*)"
```
You can also redefine:
```
DIR_IDENTITIES = "$HOME/.ssh/identities"
DIR_AGENTS = "$HOME/.ssh/agents"
```
To point somewhere else if you so desire.
| /repassh-1.2.0.tar.gz/repassh-1.2.0/README.md | 0.521715 | 0.710226 | README.md | pypi |
import io
from dataclasses import dataclass
from typing import List
from repetita_parser.errors import ParseError
from repetita_parser.types import PathLike
try:
import networkx as nx
except ImportError:
_has_networkx = False
else:
_has_networkx = True
NODES_ID = "NODES"
EDGES_ID = "EDGES"
NODES_MEMO_LINE = "label x y\n"
EDGES_MEMO_LINE = "label src dest weight bw delay\n"
@dataclass
class Node:
label: str
x: float
y: float
@dataclass
class Edge:
label: str
src: int
dest: int
weight: float
bandwidth: float
delay: float
class Topology:
def __init__(self, nodes: List[Node], edges: List[Edge], source_file: PathLike) -> None:
self.nodes: List[Node] = nodes
self.edges: List[Edge] = edges
self.source_file = source_file
def __eq__(self, other) -> bool:
"""
Comparison for equality is only defined in terms of the topology
structure, i.e., two instances can be equal although their source files
differ.
"""
return all(
[
self.nodes == other.nodes,
self.edges == other.edges,
]
)
def __ne__(self, other) -> bool:
"""
Comparison for equality is only defined in terms of the topology
structure, i.e., two instances can be equal although their source files
differ.
"""
return not (self == other)
def as_nx_graph(self):
"""
Convert the topology to a `networkx.MultiDiGraph`. In the graph, nodes
are represented by their index into `self.nodes`. Node and edge objects
carry their respective `Node` and `Edge` objects in their attributes
under the `obj` key.
This function requires NetworkX to be installed. If you call this
function without NetworkX available, it will raise an ImportError.
"""
if not _has_networkx:
msg = "NetworkX is required to call this function"
raise ImportError(msg)
else:
graph = nx.MultiDiGraph()
for node_idx, node in enumerate(self.nodes):
graph.add_node(node_idx, obj=node)
for edge in self.edges:
graph.add_edge(edge.src, edge.dest, obj=edge)
return graph
def export(self, target: io.TextIOBase) -> None:
# Write node info
target.writelines(
[
f"{NODES_ID} {len(self.nodes)}\n",
NODES_MEMO_LINE,
]
)
target.writelines([f"{n.label} {n.x} {n.y}\n" for n in self.nodes])
target.write("\n")
# Write edge info
target.writelines(
[
f"{EDGES_ID} {len(self.edges)}\n",
EDGES_MEMO_LINE,
]
)
target.writelines([f"{e.label} {e.src} {e.dest} {e.weight} {e.bandwidth} {e.delay}\n" for e in self.edges])
@dataclass
class _ParserState:
stream: io.TextIOWrapper
file_path: PathLike
line_idx: int
@property
def line_num(self) -> int:
return self.line_idx + 1
def _parse_nodes(state: _ParserState) -> List[Node]:
num_node_fields = 3
# If this changes, we have to touch the impl
assert len(NODES_MEMO_LINE.strip().split(" ")) == num_node_fields
nodes: List[Node] = []
# Nodes and edges are separated by a blank line
while (line := state.stream.readline()) != "\n":
fields = line.strip("\n").split()
if state.line_idx == 1:
if line != NODES_MEMO_LINE:
msg = "expected nodes memo line"
raise ParseError(msg, state.file_path, state.line_num)
elif len(fields) != num_node_fields:
msg = "not all node fields present"
raise ParseError(msg, state.file_path, state.line_num)
else:
label = fields[0]
x = float(fields[1])
y = float(fields[2])
nodes.append(Node(label, x, y))
state.line_idx += 1
state.line_idx += 1
return nodes
def _parse_edges(state: _ParserState) -> List[Edge]:
num_edge_fields = 6
# If this changes, we have to touch the impl
assert num_edge_fields == len(EDGES_MEMO_LINE.strip().split(" "))
edges: List[Edge] = []
start_line_idx = state.line_idx
# At EOF, we read an empty string which is falsey
while line := state.stream.readline():
fields = line.strip("\n").split()
if state.line_idx == start_line_idx:
if line != EDGES_MEMO_LINE:
msg = "expected edges memo line"
raise ParseError(msg, state.file_path, state.line_num)
else:
if len(fields) != num_edge_fields:
msg = "not all edge fields present"
raise ParseError(msg, state.file_path, state.line_num)
label = fields[0]
src = int(fields[1])
dest = int(fields[2])
weight = float(fields[3])
bw = float(fields[4])
delay = float(fields[5])
edges.append(Edge(label, src, dest, weight, bw, delay))
state.line_idx += 1
state.line_idx += 1
return edges
def parse(file_path: PathLike) -> Topology:
with open(file_path) as f:
cur_line_idx = 0
line = f.readline()
fields = line.strip("\n").split()
if fields[0] != NODES_ID:
msg = "expected nodes header line"
raise ParseError(msg, file_path, cur_line_idx + 1)
cur_line_idx += 1
state = _ParserState(f, file_path, cur_line_idx)
nodes = _parse_nodes(state)
line = f.readline()
state.line_idx += 1
fields = line.strip("\n").split()
if fields[0] != EDGES_ID:
msg = "expected edges header line"
raise ParseError(msg, file_path, cur_line_idx + 1)
edges = _parse_edges(state)
return Topology(nodes, edges, file_path) | /repetita_parser-4.0.0.tar.gz/repetita_parser-4.0.0/src/repetita_parser/topology.py | 0.842734 | 0.435121 | topology.py | pypi |
from dataclasses import dataclass
from io import TextIOBase
from typing import List
from repetita_parser.errors import ParseError
from repetita_parser.types import PathLike
DEMANDS_ID = "DEMANDS"
DEMANDS_MEMO_LINE = "label src dest bw\n"
@dataclass
class Demand:
label: str
src: int
dest: int
bandwidth: float
class Demands:
"""
Wrapper object for demands. Use `Demands.list` to get access to the
actual `Demand` objects.
"""
def __init__(self, demands: List[Demand], source_file: PathLike) -> None:
self.list = demands
self.source_file = source_file
def export(self, target: TextIOBase) -> None:
target.writelines(
[
f"{DEMANDS_ID} {len(self.list)}\n",
DEMANDS_MEMO_LINE,
]
)
target.writelines([f"{d.label} {d.src} {d.dest} {d.bandwidth}\n" for d in self.list])
def __eq__(self, other) -> bool:
"""
Comparison for equality is only defined in terms of the demands
themselves , i.e., two instances can be equal although their source
files differ.
"""
return self.list == other.list
def __ne__(self, other) -> bool:
"""
Comparison for equality is only defined in terms of the demands
themselves , i.e., two instances can be equal although their source
files differ.
"""
return not (self.list == other.list)
def parse(file_path: PathLike) -> Demands:
num_demand_fields = 4
# If this changes, we have to touch the impl
assert num_demand_fields == len(DEMANDS_MEMO_LINE.strip().split(" "))
demands: List[Demand] = []
with open(file_path) as f:
for line_idx, line in enumerate(f.readlines()):
fields = line.strip("\n").split()
if line_idx == 0:
# Two fields: `DEMANDS_ID` and number of demands
num_header_fields = 2
if len(fields) != num_header_fields or fields[0] != DEMANDS_ID:
msg = "expected demands header line"
raise ParseError(msg, file_path, line_idx + 1)
elif line_idx == 1:
if line != DEMANDS_MEMO_LINE:
msg = "expected demands memo line"
raise ParseError(msg, file_path, line_idx + 1)
else:
if len(fields) != num_demand_fields:
msg = "not all demand fields present"
raise ParseError(msg, file_path, line_idx + 1)
label = fields[0]
src = int(fields[1])
dest = int(fields[2])
bw = float(fields[3])
demands.append(Demand(label, src, dest, bw))
return Demands(demands, file_path) | /repetita_parser-4.0.0.tar.gz/repetita_parser-4.0.0/src/repetita_parser/demands.py | 0.874988 | 0.300868 | demands.py | pypi |
from io import TextIOBase
from os import PathLike
from string import Template
import numpy as np
from repetita_parser import demands, errors, topology
def _build_tm(topology: topology.Topology, demands: demands.Demands) -> np.ndarray:
"""
Per the format specification, demands between the same node pair can occur
multiple times. This function collapses the list of demands into a
two-dimensional traffic matrix that sums all demands between any given pair
into a single value.
"""
num_nodes = len(topology.nodes)
tm = np.zeros(shape=(num_nodes, num_nodes))
for d in demands.list:
tm[d.src, d.dest] += d.bandwidth
return tm
class Instance:
def __init__(self, topology_file: PathLike, demands_file: PathLike) -> None:
self.topology: topology.Topology = topology.parse(topology_file)
self.demands: demands.Demands = demands.parse(demands_file)
# Check if all indices in parsed demands are valid for parsed topology
min_node_idx, max_node_idx = 0, len(self.topology.nodes) - 1
for d in self.demands.list:
src_ok = min_node_idx <= d.src <= max_node_idx
dest_ok = min_node_idx <= d.dest <= max_node_idx
if not (src_ok and dest_ok):
msg_template = Template(f"demand {d.label}: node index $index does not exist in topology")
if not src_ok:
msg = msg_template.substitute(index=d.src)
else:
msg = msg_template.substitute(index=d.dest)
# XXX: In theory, both indices could be invalid. In that case,
# we only report the invalid source index.
raise errors.ValidationError(msg, topology_file, demands_file)
self.traffic_matrix = _build_tm(self.topology, self.demands)
"""
Total traffic demand from node `i` to node `j` at `traffic_matrix[i, j]`
"""
def __eq__(self, other) -> bool:
return all(
[
self.topology == other.topology,
self.demands == other.demands,
]
)
def __ne__(self, other) -> bool:
return not (self == other)
def export(self, topology_target: TextIOBase, demands_target: TextIOBase) -> None:
self.topology.export(topology_target)
self.demands.export(demands_target) | /repetita_parser-4.0.0.tar.gz/repetita_parser-4.0.0/src/repetita_parser/instance.py | 0.811825 | 0.351728 | instance.py | pypi |
repex (REPlace (regular) EXpression)
====================================
[](https://travis-ci.org/cloudify-cosmo/repex)
[](https://ci.appveyor.com/project/Cloudify/repex/branch/master)
[](https://pypi.python.org/pypi/repex)
[](https://img.shields.io/pypi/pyversions/repex.svg)
[](https://requires.io/github/cloudify-cosmo/repex/requirements/?branch=master)
[](https://codecov.io/github/cloudify-cosmo/repex?branch=master)
[](https://landscape.io/github/cloudify-cosmo/repex)
[](https://pypi.python.org/pypi/repex)
NOTE: Beginning with `repex 0.4.1`, file attributes are kept when replacing.
NOTE: Beginning with `repex 0.4.3`, Windows is officially supported (and tested via appveyor).
NOTE: Beggining with `repex 1.0.0`, Python 3 is officially supported.
NOTE: `repex 1.1.0` has breaking CLI and API changes. See [CHANGES](CHANGES) for more information.
NOTE: `repex 1.2.0` does not allow to set variables in the config without providing them.
`repex` replaces strings in single/multiple files based on regular expressions.
Why not Jinja you ask? Because sometimes you have existing files which are not templated in which you'd like to replace things.. and even if they're in your control, sometimes templates are just not viable if you need something working OOB.
Why not use sed you ask? Because `repex` provides some layers of protection and an easy to use config yaml in which you easily add new files and folders to iterate through.
The layers are:
* Match and only then replace in the matched regular expression which allows the user to provide context for the replacement instead of just iterating through the entire file.
* Check for existing strings in a file before replacing anything.
* Exclude files and folders so that you don't screw up.
* Validate that the replacement went as expected by allowing to execute a validation function post-replacement.
AND, you can use variables (sorta Jinja2 style). How cool is that? See reference config below.
## Installation
`repex` is supported and tested on Python 2.6, 2.7, 3.3+ and PyPy.
```shell
pip install repex
```
For dev:
```shell
pip install https://github.com/cloudify-cosmo/repex/archive/master.tar.gz
```
## Usage
### CLI
Repex exposes a CLI which can be used to do one of two things:
1. Use repex's power to basically replace sed in the command line.
2. Execute repex using a config file.
NOTE: When passing a config file, repex will ignore any options passed which are not `config-only`.
```
$ rpx -h
...
Usage: rpx [OPTIONS] [REGEX_PATH]
Replace strings in one or multiple files.
You must either provide `REGEX_PATH` or use the `-c` flag to provide a
valid repex configuration.
`REGEX_PATH` can be: a regex of paths under `basedir`, a path to a single
directory under `basedir`, or a path to a single file.
It's important to note that if the `REGEX_PATH` is a path to a directory,
the `-t,--ftype` flag must be provided.
Options:
-r, --replace TEXT A regex string to replace. Mutually
exclusive with: [config]
-w, --replace-with TEXT Non-regex string to replace with. Mutually
exclusive with: [config]
-m, --match TEXT Context regex match for `replace`. If this
is ommited, the context will be the entire
content of the file. Mutually exclusive
with: [config]
-t, --ftype TEXT A regex file name to look for. Defaults to
`None`, which means that `PATH_TO_HANDLE`
must be a path to a single file [non-config
only]. Mutually exclusive with: [to_file,
config]
-b, --basedir TEXT Where to start looking for `path` from.
Defaults to the cwd. Mutually exclusive
with: [config]
-x, --exclude-paths TEXT Paths to exclude when searching for files to
handle. This can be used multiple times.
Mutually exclusive with: [config]
-i, --must-include TEXT Files found must include this string. This
can be used multiple times. Mutually
exclusive with: [config]
--validator TEXT Validator file:function (e.g.
validator.py:valid_func [non-config only].
Mutually exclusive with: [config]
--validator-type [per_file|per_type]
Type of validation to perform. `per_type`
will validate the last file found while
`per_file` will run validation for each file
found. Defaults to `per_type` [non-config
only]. Mutually exclusive with: [config]
--to-file TEXT File path to write the output to. Mutually
exclusive with: [ftype, config]
-c, --config TEXT Path to a repex config file. Mutually
exclusive with: [REGEX_PATH]
--vars-file TEXT Path to YAML based vars file. Mutually
exclusive with: [REGEX_PATH]
--var TEXT A variable to pass to Repex. Can be used
multiple times. Format should be
`'key'='value'`. Mutually exclusive with:
[REGEX_PATH]
--tag TEXT A tag to match with a set of tags in the
config. Can be used multiple times. Mutually
exclusive with: [REGEX_PATH]
--validate / --no-validate Validate the config (defaults to True).
Mutually exclusive with: [validate_only,
REGEX_PATH]
--validate-only Only validate the config, do not run
(defaults to False). Mutually exclusive
with: [validate, REGEX_PATH]
--diff Write the diff to a file under `cwd/.rpx
/diff-TIMESTAMP` (defaults to False)
-v, --verbose Show verbose output
-h, --help Show this message and exit.
...
```
#### Using repex like sed
Just like sed:
```bash
rpx /path/to/my/file --replace 3.3 --rwith 3.4
```
Much, much more than sed:
```bash
rpx 'check_validity/resources/.*'
-t VERSION \
-r '3.3.0-m\d+' \
-w 2.1.1 \
-i blah -i yay! \
-x check_validity/resources/VERSION -x another/VERSION \
--validator check_validity/resources/validator.py:validate \
--diff -v
```
This will look for all files named "VERSION" under all folders named "check_validity/resources/.*" (recursively); replace all strings matching "3.3.0-m\d+" with "2.1.1"; validate using the "validate" function found in "check_validity/resources/validator.py" only if the files found include the strings "blah" and "yay!" excluding specifically the files "check_validity/resources/VERSION" and "another/VERSION". A git style diff file will be generated.
Note that you must either escape special chars or use single quotes where applicable, that is, where regex strings are provided and bash expansion takes place.
#### Notes
* In complex scenarios, while the CLI can execute repex, it will be more likely that you would use the Python API to execute the `iterate` function as you will most probably want to dynamically pass variables according to certain logic provided by your system.
* Variables provided via the `--var` flag will override variables provided within the `--vars-file`.
* Currently, you can't pass variables which contain a `=` within them.
#### Passing a config file to the CLI
Passing a config file to the CLI is done as follows:
```bash
rpx -c config.yaml -t my_tag -v --vars-file vars.yaml --var 'x'='y' --var 'version'='3.3.0-m3'
```
See below for how to use the config file.
### Config file based usage
Using a config file adds some cool features and allows to run repex on multiple paths using a single config file.
Let's say you have files named "VERSION" in different directories which look like this:
```json
{
"date": "",
"commit": "",
"version": "3.3.0-m2",
"version_other": "3.1.2-m1",
"build": "8"
}
```
And you'd like to replace 3.3.0-m2 with 3.3.0-m3 in all of those files
You would create a repex config.yaml with the following:
```yaml
variables:
base_dir: .
valstr: 'date'
regex: '\d+(\.\d+){1,2}(-(m|rc)(\d+)?)?'
paths:
- type: VERSION
path: resources
tags:
- my_tag
- my_other_tag
excluded:
- x/y/VERSION
base_directory: "{{ .base_dir }}"
match: '"version": "{{ .regex }}'
replace: "{{ .regex }}"
with: "{{ .version }}"
validate_before: true
diff: true
must_include:
- "{{ .valstr }}"
- commit
- version
validator:
type: per_file
path: '{{ .basedir }}/validator/script/path.py'
function: my_validation_function
```
and do the following
```python
import os
import repex
CONFIG_YAML_FILE = "config.yaml"
VERSION = os.environ['VERSION'] # '3.1.0-m3'
variables = {
'version': VERSION,
}
repex.iterate(
config_file_path=CONFIG_YAML_FILE,
config=None, # config is simply the dict form of the contents of `CONFIG_YAML_FILE`.
tags=['my_tag1', 'my_tag2'] # tags to match
variables=variables,
validate=True, # validate config schema
validate_only=False, # only validate config schema without running
with_diff=True # write the diff to a file
)
```
and even add a validator file:
```python
def my_validation_function(version_file_path, logger):
logger.debug('Validating my thing...')
result = verify_replacement()
return result == 'yay! it passed!'
```
## Config YAML Explained
IMPORTANT NOTE: variables MUST be enclosed within single or double quotes or they will not expand! Might fix that in future versions...
ANOTHER IMPORTANT NOTE: variables must be structured EXACTLY like this: {{ .VER_NAME }}
Don't forget the spaces!
- `variables` is a dict of variables you can use throughout the config. See below for more info.
- `type` is a regex string representing the file name you're looking for.
- `path` is a regex string representing the path in which you'd like to search for files (so, for instance, if you only want to replace files in directory names starting with "my-", you would write "my-.*"). If `path` is a path to a single file, the `type` attribute must not be configured.
- `tags` is a list of tags to apply to the path. Tags are used for Repex's triggering mechanism to allow you to choose which paths you want to address in every single execution. More on that below.
- `excluded` is a list of excluded paths. The paths must be relative to the working directory, NOT to the `path` variable.
- `base_directory` is the directory from which you'd like to start the recursive search for files. If `path` is a path to a file, this property can be omitted. Alternatively, you can set the `base_directory` and a `path` relative to it.
- `match` is the initial regex based string you'd like to match before replacing the expression. This provides a more robust way of replacing strings where you first match the exact area in which you'd like to replace the expression and only then match the expression you want to replace within it. It also provides a way to replace only specific instances of an expression, and not all.
- `replace` - which regex would you like to replace?
- `with` - what you replace with.
- `must_include` - as an additional layer of security, you can specify a set of regex based strings to look for to make sure that the files you're dealing with are the actual files you'd like to replace the expressions in.
- `validator` - validator allows you to run a validation function after replacing expressions. It receives `type` which can be either `per_file` or `per_type` where `per_file` runs the validation on every file while `per_type` runs once for every `type` of file; it receives a `path` to the script and a `function` within the script to call. Note that each validation function must return `True` if successful while any other return value will fail the validation. The validating function receives the file's path as and a logger as arguments.
- `diff` - if `true`, will write a git-like unified diff to a file under `cwd/.rpx/diff-TIMESTAMP`. Note that `PATH_REGEX` can be anything which means that the names of the files will look somewhat weird. The diff will be written for each replacement. See below for an example.
In case you're providing a path to a file rather than a directory:
- `type` and `base_directory` are depracated
- you can provide a `to_file` key with the path to the file you'd like to create after replacing.
## Tags
Tags allow a user to choose a set of paths on each execution.
A user could apply a list of tags to a path and then, executing repex will address these paths according to the following logic:
* If a user supplied a list of tags and the path was applied a list of tags, the path will be addressed only if matching tags were found.
* If a user supplied a list of tags and the path contains no tags, the path will be ignored.
* If a user did not supply tags and the path contains tags, the path will be ignored.
* If a user did not supply tags and the path does not contain tags, the path will be addressed.
* If a user proivded `any` as a tag, all paths, regardless of whether they have or haven't tags will be addressed.
## Variables
Variables are one of the strongest features of repex. They provide a way of injecting dynamic info to the config.
Variables can be declared in 4 ways:
- Provided via the CLI
- Provided via the API.
- Harcoded in the config under a top level `variables` section.
- Set as Environment Variables.
Note that variables can also be used within variables in the config.
See the example above for a variable definition reference.
Some important facts about variables:
- Variables with the same name sent via the API will override the hardcoded ones.
- API provided or hardcoded variables can be overriden if env vars exist with the same name but in upper case and prefixed with `REPEX_VAR_` (so the variable "version" can be overriden by an env var called "REPEX_VAR_VERSION".) This can help with, for example, using the $BUILD_NUMBER env var in Jenkins to update a file with the new build number.
Note that if any variables are required but not provided, repex will fail stating that they must be provided.
## Diff
NOTE: THIS IS WIP! Use sparingly.
Repex has the ability to write a git-like unified diff for every replacement that occurs. The diff is written to a file under `cwd/.rpx/` and will contain something that looks like the following:
```text
$ cat .rpx/diff-20170119T115322
...
2017-01-19 11:53:22 tests/resources/multiple/mock_VERSION
0 ---
1 +++
2 @@ -1,7 +1,7 @@
3 {
4 "date": "",
5 "commit": "",
6 - "version": "3.1.0-m2",
7 + "version": "xxx",
8 "versiond": "3.1.0-m2",
9 "build": "8"
10 }
2017-01-19 11:53:22 tests/resources/multiple/folders/mock_VERSION
0 ---
1 +++
2 @@ -1,7 +1,7 @@
3 {
4 "date": "",
5 "commit": "",
6 - "version": "3.1.0-m2",
7 + "version": "xxx",
8 "versiond": "3.1.0-m2",
9 "build": "8"
10 }
...
```
There is currently no way to ask repex to not generate the diff for every file, so take that into consideration when working with a large amount of files.
Diff generation is off by default. Note that other than providing the overriding `--diff` (or `with_diff` in `iterate`) flag, you can set `diff` for each path in the config.
## Testing
```shell
git clone git@github.com:cloudify-cosmo/repex.git
cd repex
pip install tox
tox
```
## Contributions..
Pull requests are always welcome..
| /repex-1.3.1.tar.gz/repex-1.3.1/README.md | 0.659624 | 0.933461 | README.md | pypi |
import sys
from copy import deepcopy
from datetime import datetime, timedelta
from typing import Any, ClassVar, Type, Union
from uuid import uuid4
from msgspec import Meta, Struct, field, json
from repid.data import PrioritiesT
from repid.data.protocols import (
DelayPropertiesT,
ResultPropertiesT,
RetriesPropertiesT,
)
from repid.utils import VALID_ID, VALID_NAME
if sys.version_info >= (3, 9):
from typing import Annotated
else:
from typing_extensions import Annotated
try:
from croniter import croniter
CRON_SUPPORT = True
except ImportError: # pragma: no cover
CRON_SUPPORT = False
def enc_hook(obj: Any) -> Any:
if isinstance(obj, timedelta):
return obj.total_seconds()
raise TypeError(f"Objects of type {type(obj)} are not supported")
def dec_hook(type: Type, obj: Any) -> Any: # noqa: A002
if type is timedelta:
return timedelta(seconds=obj)
raise TypeError(f"Objects of type {type} are not supported")
class RoutingKey(Struct, array_like=True, omit_defaults=True, frozen=True):
topic: Annotated[str, Meta(pattern=str(VALID_NAME))]
queue: Annotated[str, Meta(pattern=str(VALID_NAME))] = "default"
priority: Annotated[int, Meta(ge=1)] = PrioritiesT.MEDIUM.value
id_: Annotated[str, Meta(pattern=str(VALID_ID))] = field(default_factory=lambda: uuid4().hex)
class ArgsBucket(Struct, array_like=True, omit_defaults=True, frozen=True):
data: str
timestamp: datetime = field(default_factory=datetime.now)
ttl: Union[timedelta, None] = None
def encode(self) -> str:
return encoder.encode(self).decode("cp1252")
@classmethod
def decode(cls, data: str) -> "ArgsBucket":
return args_bucket_decoder.decode(data.encode())
@property
def is_overdue(self) -> bool:
if self.ttl is None:
return False
return datetime.now(tz=self.timestamp.tzinfo) > self.timestamp + self.ttl
class ResultBucket(Struct, array_like=True, omit_defaults=True, frozen=True):
data: str
# perf_counter_ns
started_when: int
finished_when: int
success: bool = True
exception: Union[str, None] = None
timestamp: datetime = field(default_factory=datetime.now)
ttl: Union[timedelta, None] = None
def encode(self) -> str:
return encoder.encode(self).decode("cp1252")
@classmethod
def decode(cls, data: str) -> "ResultBucket":
return result_bucket_decoder.decode(data.encode())
@property
def is_overdue(self) -> bool:
if self.ttl is None:
return False
return datetime.now(tz=self.timestamp.tzinfo) > self.timestamp + self.ttl
class RetriesProperties(Struct, array_like=True, omit_defaults=True, frozen=True):
max_amount: int = 0
already_tried: int = 0
def encode(self) -> str:
return encoder.encode(self).decode("cp1252")
@classmethod
def decode(cls, data: str) -> "RetriesProperties":
return retries_decoder.decode(data.encode())
class ResultProperties(Struct, array_like=True, omit_defaults=True, frozen=True):
id_: str = field(default_factory=lambda: uuid4().hex)
ttl: Union[timedelta, None] = None
def encode(self) -> str:
return encoder.encode(self).decode("cp1252")
@classmethod
def decode(cls, data: str) -> "ResultProperties":
return result_decoder.decode(data.encode())
class DelayProperties(Struct, array_like=True, omit_defaults=True, frozen=True):
delay_until: Union[datetime, None] = None
defer_by: Union[timedelta, None] = None
cron: Union[str, None] = None
next_execution_time: Union[datetime, None] = None
def encode(self) -> str:
return encoder.encode(self).decode("cp1252")
@classmethod
def decode(cls, data: str) -> "DelayProperties":
return delay_decoder.decode(data.encode())
class Parameters(Struct, array_like=True, omit_defaults=True, frozen=True):
RETRIES_CLASS: ClassVar[Type[RetriesPropertiesT]] = RetriesProperties
RESULT_CLASS: ClassVar[Type[ResultPropertiesT]] = ResultProperties
DELAY_CLASS: ClassVar[Type[DelayPropertiesT]] = DelayProperties
execution_timeout: timedelta = field(default_factory=lambda: timedelta(minutes=10))
result: Union[ResultProperties, None] = None
retries: RetriesProperties = field(default_factory=RetriesProperties)
delay: DelayProperties = field(default_factory=DelayProperties)
timestamp: datetime = field(default_factory=datetime.now)
ttl: Union[timedelta, None] = None
def encode(self) -> str:
return encoder.encode(self).decode("cp1252")
@classmethod
def decode(cls, data: str) -> "Parameters":
return parameters_decoder.decode(data.encode())
@property
def is_overdue(self) -> bool:
if self.ttl is None:
return False
return datetime.now(tz=self.timestamp.tzinfo) > self.timestamp + self.ttl
@property
def compute_next_execution_time(self) -> Union[datetime, None]:
now = datetime.now()
if self.delay.delay_until is not None and self.delay.delay_until > now:
return self.delay.delay_until
if self.delay.defer_by is not None:
defer_by_times = (now - self.timestamp) // self.delay.defer_by + 1
time_offset = self.delay.defer_by * defer_by_times
return self.timestamp + time_offset
if self.delay.cron is not None:
if not CRON_SUPPORT:
raise ImportError("Croniter is not installed.") # pragma: no cover
return croniter(self.delay.cron, now).get_next(ret_type=datetime) # type: ignore[no-any-return]
return None
def _prepare_reschedule(self) -> "Parameters":
copy = deepcopy(self)
object.__setattr__(copy.retries, "already_tried", 0)
object.__setattr__(copy.delay, "next_execution_time", self.compute_next_execution_time)
object.__setattr__(copy, "timestamp", datetime.now())
return copy
def _prepare_retry(self, next_retry: timedelta) -> "Parameters":
copy = deepcopy(self)
object.__setattr__(copy.retries, "already_tried", copy.retries.already_tried + 1)
object.__setattr__(
copy.delay,
"next_execution_time",
datetime.now() + next_retry,
)
return copy
encoder = json.Encoder(enc_hook=enc_hook)
args_bucket_decoder = json.Decoder(ArgsBucket, dec_hook=dec_hook)
result_bucket_decoder = json.Decoder(ResultBucket, dec_hook=dec_hook)
retries_decoder = json.Decoder(RetriesProperties, dec_hook=dec_hook)
result_decoder = json.Decoder(ResultProperties, dec_hook=dec_hook)
delay_decoder = json.Decoder(DelayProperties, dec_hook=dec_hook)
parameters_decoder = json.Decoder(Parameters, dec_hook=dec_hook) | /repid_msgspec-0.0.1.tar.gz/repid_msgspec-0.0.1/repid_msgspec/plugin.py | 0.586049 | 0.270169 | plugin.py | pypi |
import inspect
from dataclasses import dataclass
from functools import wraps
from typing import Any, Callable, Dict, List
from pydantic import BaseModel, Field, create_model
from repilot.utils.schema_utils import minify_docstring
@dataclass
class Tool:
name: str
pydoc: str | None
signature: str
basemodel: BaseModel
return_type: str
func: Callable
is_async: bool = False
@property
def minified_doc(self):
if self.pydoc is None:
return ""
return minify_docstring(self.pydoc.split(":param")[0])
def __repr__(self):
return f"{self.name}:{self.minified_doc} {self.json_schema}"
@property
def json_schema(self):
return self.basemodel.schema()["properties"]
@property
def details(self):
return {
"name": self.name,
"doc": self.minified_doc,
"signature": self.signature,
"return_type": self.return_type,
"json_schema": self.json_schema,
"is_async": self.is_async,
}
class ToolKit:
def __init__(self, init_func: Callable | None = None):
self.tools: List[Tool] = []
self.tools_dict: Dict[str, Tool] = {}
if init_func:
init_func(self)
def register(self, func: Callable) -> Callable:
# Dynamically generate Pydantic request model based on function arguments
args_annotations = getattr(func, "__annotations__", {})
inputs_annotations = {
k: v for k, v in args_annotations.items() if k != "return"
}
# Get function signature for default values
sig = inspect.signature(func)
params = sig.parameters
# Define a dictionary to store the field definitions for the Pydantic model
field_definitions: Dict[str, Any] = {}
for arg_name, arg_type in inputs_annotations.items():
# Use the argument name as the field name
field_name = arg_name
# Use the argument type and any additional validation constraints from
# annotations to define the field type
default_value = (
params[arg_name].default
if params[arg_name].default != inspect.Parameter.empty
else ...
)
field_definitions[field_name] = (arg_type, Field(default_value))
# Dynamically create a Pydantic model from the field definitions
Request = create_model("InputModel", **field_definitions)
is_async = inspect.iscoroutinefunction(func)
@wraps(func)
def wrapper(*args, **kwargs):
# Get parameter names from function signature
params = inspect.signature(func).parameters.keys()
# Match parameter values with names and convert them to a dictionary
kwargs.update(zip(params, args))
# Create request object from validated kwargs
request = Request(**kwargs)
# Call original function with validated and converted kwargs
return func(**request.dict())
# Create tool object and append it to the tools list
tool = Tool(
name=func.__name__,
pydoc=func.__doc__,
basemodel=Request,
signature=str(inspect.signature(func)),
return_type=func.__annotations__["return"].__name__
if "return" in func.__annotations__
else None,
func=wrapper,
is_async=is_async,
)
self.tools.append(tool)
self.tools_dict[func.__name__] = tool
return wrapper
def tool_names(self):
return [tool.name for tool in self.tools]
def __getitem__(self, item):
return self.tools_dict[item]
def __iter__(self):
return iter(self.tools)
def __len__(self):
return len(self.tools)
def __repr__(self):
return f"ToolKit({self.tools})" | /repilot_client-0.1.2.tar.gz/repilot_client-0.1.2/repilot/tool.py | 0.875508 | 0.237797 | tool.py | pypi |
# repka
[](https://pypi.org/project/repka/)
[](https://travis-ci.org/github/potykion/repka)
Repository pattern implementation - isolate db manipulation from domain models
## Installation
Via pip:
```
pip install repka
```
Via poetry:
```
poetry add repka
```
## Usage
### repka.api.BaseRepository
BaseRepository used to execute sql-queries (via [aiopg & sqlalchemy](https://github.com/aio-libs/aiopg)) and convert sql-rows to/from [pydantic](https://github.com/samuelcolvin/pydantic) models
```python
import sqlalchemy as sa
from repka.api import AiopgRepository, IdModel
from repka.utils import create_async_db_connection
# Define pydantic model
# It should inherit repka.api.IdModel
# to set id on entity insert, to update entity with id and more
# IdModel inherits pydantic.BaseModel and defines int id field
class Task(IdModel):
title: str
# Define sqlachemy table with same model columns
metadata = sa.MetaData()
tasks_table = sa.Table(
"tasks", metadata,
sa.Column("id", sa.Integer, primary_key=True, autoincrement=True),
sa.Column("title", sa.String)
)
# Define repository
# You should inherit repka.api.BaseRepository and
# set sqlalchemy-table via table property
# Kwargs is sql-row data returned by sqlalchemy
class TaskRepo(AiopgRepository[Task]):
table = tasks_table
# To use the repository you should instantiate it with async sqlalchemy-connection
db_url = "postgresql://postgres@localhost/test"
async with create_async_db_connection(db_url) as conn:
repo = TaskRepo(conn)
# Now you can use the repo
# Here we select first task with matching title
task = await repo.first(tasks_table.c.title == "My first task")
```
#### BaseRepository methods
>`T` means generic type passed to BaseRepository (e.g. `BaseRepository[Task]` means that type of `T` is `Task`)
##### Select methods
- `repo.first(*filters: BinaryExpression, orders: Optional[Columns])` - get first entity matching sqlalchemy {filters} and {orders}; if no entity matches {filters} then `None` is returned
> Example of {filters}: `table.c.title == 'test task'` - equals to sql where clause: `where title = 'test task'`
> Example of {orders}: `table.c.title` - equals to sql order by clause: `order by title`
- `repo.get_by_ids(entity_ids: List[int])` - get all entities whose id in {entity_ids} (same as sql `where id in ({entity_ids})`)
- `repo.get_by_id(entity_id: int)` - get entity with id = {entity_id}
- `repo.get_or_create(filters: Optional[List[BinaryExpression]], defaults: Optional[Dict])` - get entity that matches {filters} if no entity found create new entity with {defaults}; return tuple of entity and entity existence flag
- `repo.get_all(filters: Optional[List[BinaryExpression]], orders: Optional[Columns])` - return all entities matching {filters} and {orders}
- `repo.get_all_ids(filters: Optional[List[BinaryExpression]], orders: Optional[Columns])` - return ids of entites matching {filters} and {orders}
- `repo.exists(*filters: BinaryExpression)` - check that entity matching {filters} exists using sql `count` statement
##### Insert methods
- `repo.insert(entity: T)` - insert entity to db table and set id field to the entity
- `repo.insert_many(entities: List[T])` - insert multiple entities and set ids to them in single transaction
##### Update methods
- `repo.update(entity: T)` - updates entity in db
- `repo.update_partial(entity: T, **updated_values)` - update entity fields via kwargs and update entity fields in db
- `repo.update_many(entities: List[T])` - update multiple entities in single transaction
##### Delete methods
- `repo.delete(*filters: BinaryExpression)` - delete entities matching {filters} via sql `delete` statement
> To delete all entities pass `None` as an arg: `repo.delete(None)`
- `repo.delete_by_id(entity_id: int)` - delete entity with {entity_id}
- `repo.delete_by_ids(entity_ids: List[int])` - delete entities whose id in {entity_ids}
##### Other methods & properties
- `repo.serialize(entity: T)` - convert {entity} to dict (e.g. in `insert` and `update` methods)
- `repo.deserialize(**kwargs)` - convert {kwargs} to entity (e.g. in `first` and `get_all` methods)
- `repo.execute_in_transaction()` - context manager that allows execute multiple queries in transaction
Example: delete all old entities and insert new one in single transaction:
```python
async with repo.execute_in_transaction():
await repo.delete()
await repo.insert(Task(title="New task"))
```
- `repo.ignore_default` - list of entity fields that will be ignored on insert and set after insert if they equal to default field value.
Useful for auto incrementing / default fields like dates or sequence numbers
#### ContextVar support
You can create lazy-connection repositories with context vars
```python
from contextvars import ContextVar
from repka.utils import create_async_db_connection
# Create context var and instantiate repository
db_connection = ContextVar("db_connection")
repo = TaskRepo(db_connection)
# Now you should set the context var somewhere (e.g. in middleware)
# And start using the repository
async with create_async_db_connection(db_url) as conn:
db_connection.set(conn)
await repo.insert(Task(title="New task"))
```
#### Other sqlalchemy repositories
Following repositories have same api as `AiopgRepository` (select methods, insert methods, etc.)
- `repka.api.FakeRepo` - repository that uses lists instead of database tables, can be used as mock
- This repository is implemented partially, because implementing sqlalchemy features (like filters or orders) is hard and pointless for python lists
### repka.json_.DictJsonRepo
This kind of repository used to save/load json objects from file:
```python
from repka.json_ import DictJsonRepo
songs = [
{"artist": "Pig Destroyer", "title": "Thumbsucker"},
{"artist": "Da Menace", "title": "Bag of Funk"}
]
repo = DictJsonRepo()
repo.write(songs, "songs.json")
assert repo.read("songs.json") == songs
```
#### DictJsonRepo methods
- `repo.read(filename: str)` - read json file with {filename}, return its content as json primitive (list, dict, str, etc.)
- `repo.write(data: T, filename: str)` - serialize json primitive {data} and save it to file with {filename}
- `repo.read_or_write_default(filename: str, default_factory: Callable[[], T])` - check file with {filename} exists, read its content if exists, execute {default_factory} and write it to file with {filename} otherwise
- Example: read data from `test.json` or create `test.json` with `[{"field": "value"}]` if no such file:
```python
repo = DictJsonRepo()
repo.read_or_write_default("test.json", lambda: [{"field": "value"}])
```
#### DictJsonRepo constructor
- `DictJsonRepo(directory: str)` - set directory where files will be read / written; if not set current working directory will be used
- Example: read files from `data/` dir:
```python
repo = DictJsonRepo("data")
repo.read("test.json") # will read "./data/test.json"
```
## Development and contribution
### Dependencies
Install production and development dependencies via poetry:
```
poetry install
```
### Tests
To run tests:
1. Setup [database url](https://docs.sqlalchemy.org/en/13/core/engines.html#database-urls) via `DB_URL` environment variable (e.g. via .env file)
**WARNING:** Every test run will drop all tables from the db
2. Run tests via `pytest`
### Contribution
1. Create fork/branch for new feature/fix/whatever
2. [Optional] Install pre-commit hooks: `pre-commit install` (for manual pre-commit run use`pre-commit run -a`)
3. When you done create pull request and wait for approval
### Deploy
To deploy new version you need to increment version via bump2version and publish it to PyPI via poetry:
```
bump2version major/minor/patch
poetry publish --build
```
Don't forget to fill the CHANGELOG.md before release
| /repka-3.2.0.tar.gz/repka-3.2.0/README.md | 0.664867 | 0.889816 | README.md | pypi |
from typing import Callable, Any, Literal
import os
from rich.console import Console
console = Console()
menus = Literal["", "featflags", "settings"]
def handle_error(name: str, default: Any, menu: menus = ""):
"""Handles the error by returning the default value and printing an
informative error message.
Parameters
----------
name: str
The name of the environment variable
default: Any
The value to return if the converter fails
menu: menus
If provided, will tell the user where to fix the setting
Returns
----------
Any
The default value
"""
base = f"[red]Invalid variable provided for variable '{name}'."
if menu:
base += f" Please change the setting in the `{menu}` menu."
base += "[/red]\n"
console.print(base)
return default
def load_env_vars(
name: str, converter: Callable, default: Any, menu: menus = ""
) -> Any:
"""Loads an environment variable and attempts to convert it to the correct data type.
Will return the provided default if it fails
Parameters
----------
name: str
The name of the environment variable
converter: Callable
The function to convert the env variable to the desired format
default: Any
The value to return if the converter fails
menu: menus
If provided, will tell the user where to fix the setting
Returns
----------
Any
The value or the default
"""
raw_var = os.getenv(name, str(default))
try:
return converter(raw_var)
except ValueError:
return handle_error(name, default, menu)
except AttributeError:
return handle_error(name, default, menu)
except TypeError:
return handle_error(name, default, menu)
def strtobool(val):
"""Convert a string representation of truth to true (1) or false (0).
True values are 'y', 'yes', 't', 'true', 'on', and '1'; false values
are 'n', 'no', 'f', 'false', 'off', and '0'. Raises ValueError if
'val' is anything else.
"""
val = str(val).lower()
if val in ("y", "yes", "t", "true", "on", "1"):
output = 1
elif val in ("n", "no", "f", "false", "off", "0"):
output = 0
else:
raise ValueError(f"invalid truth value {val}")
return output | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/base_helpers.py | 0.862265 | 0.279828 | base_helpers.py | pypi |
__docformat__ = "numpy"
import functools
import logging
import os
from ssl import SSLError
import pandas as pd
from requests.exceptions import RequestException
from openbb_terminal import feature_flags as obbff
from openbb_terminal.rich_config import console # pragma: allowlist secret
logger = logging.getLogger(__name__)
def log_start_end(func=None, log=None):
"""Wrap function to add a log entry at execution start and end.
Parameters
----------
func : optional
Function, by default None
log : optional
Logger, by default None
Returns
-------
Wrapped function
"""
assert callable(func) or func is None # nosec
def decorator(func):
@functools.wraps(func)
def wrapper(*args, **kwargs):
logging_name = ""
args_passed_in_function = [
repr(a) for a in args if isinstance(a, (pd.DataFrame, pd.Series)) or a
]
if (
len(args) == 2
and args_passed_in_function
and (
"__main__.TerminalController" in args_passed_in_function[0]
or (
"openbb_terminal." in args_passed_in_function[0]
and "_controller" in args_passed_in_function[0]
)
)
):
logging_name = args_passed_in_function[0].split()[0][1:]
args_passed_in_function = args_passed_in_function[1:]
logger_used = logging.getLogger(logging_name) if logging_name else log
logger_used.info(
"START",
extra={"func_name_override": func.__name__},
)
if os.environ.get("DEBUG_MODE") == "true":
value = func(*args, **kwargs)
log.info("END", extra={"func_name_override": func.__name__})
return value
try:
value = func(*args, **kwargs)
logger_used.info("END", extra={"func_name_override": func.__name__})
return value
except RequestException as e:
console.print(
"[red]There was an error connecting to the API."
" Please try again later.\n[/red]"
)
logger_used.exception(
"Exception: %s",
str(e),
extra={"func_name_override": func.__name__},
)
return []
except SSLError as e:
console.print(
"[red]There was an error connecting to the API."
" Please check whether your wifi is blocking this site.\n[/red]"
)
logger_used.exception(
"Exception: %s",
str(e),
extra={"func_name_override": func.__name__},
)
return []
except Exception as e:
console.print(f"[red]Error: {e}\n[/red]")
logger_used.exception(
"Exception: %s",
str(e),
extra={"func_name_override": func.__name__},
)
return []
return wrapper
return decorator(func) if callable(func) else decorator
# pylint: disable=import-outside-toplevel
def check_api_key(api_keys):
"""
Wrapper around the view or controller function and
print message statement to the console if API keys are not yet defined.
"""
def decorator(func):
@functools.wraps(func)
def wrapper_decorator(*args, **kwargs):
if obbff.ENABLE_CHECK_API:
import openbb_terminal.config_terminal as cfg
undefined_apis = []
for key in api_keys:
# Get value of the API Keys
if getattr(cfg, key) == "REPLACE_ME" and key not in [
"API_KEY_ALPHAVANTAGE"
]:
undefined_apis.append(key)
if undefined_apis:
undefined_apis_name = ", ".join(undefined_apis)
console.print(
f"[red]{undefined_apis_name} not defined. "
"Set API Keys in ~/.openbb_terminal/.env or under keys menu.[/red]\n"
) # pragma: allowlist secret
return None
return func(*args, **kwargs)
return wrapper_decorator
return decorator
def disable_check_api():
obbff.ENABLE_CHECK_API = False | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/decorators.py | 0.591959 | 0.153803 | decorators.py | pypi |
<!-- markdownlint-disable MD033 -->
# OpenBB SDK
OpenBB SDK gives you direct and programmatic access to all capabilities of the OpenBB Terminal.
You will have the necessary building blocks to create your own financial tools and applications,
whether that be a visualization dashboard or a custom report in a Jupyter Notebook.
With OpenBB SDK, you can access normalized financial data from dozens of data providers,
without having to develop your own integrations from scratch.
On top of financial data feeds, OpenBB SDK also provides you with a toolbox to perform financial analysis
on a variety of asset classes, including stocks, crypto, ETFs, funds; the economy as well as your portfolios.
OpenBB SDK is created and maintained by OpenBB team together with the contributions from hundreds of community members.
This gives us an unrivaled speed of development and the ability to maintain stable integrations with numerous third-party data providers.
Developing and maintaining an full-blown investment research infrastructure from the ground up takes a lot of time and effort.
However, it does not have to be. Take advantage of OpenBB SDK with its out-of-the-box data connectors and financial analysis toolkit.
So that you can focus on designing and building your financial reports and applications.
## SDK structure
The OpenBB SDK consists of the core package and extension toolkits.
The core package includes all necessary functionality for you to start researching or developing your dashboards and applications.
The toolkits that you can extend the OpenBB SDK with are:
- Portfolio Optimization Toolkit.
- Forecasting Toolkit.
## System and Platform Requirements
The SDK core package is expected to work in any officially supported python version 3.8 and higher (3.9 recommended).
Optimization and Forecasting toolkits installation requires specific settings on computers powered by Apple Silicon, the newer Windows ARM and Raspberry Pi.
### Minimal and Recommended System Requirements
- A computer with a modern CPU (released in the past 5 years)
- At least 8GB of RAM, 16+ recommended
- SSD drive with at least 12GB of storage space available
- Internet connection
**NOTES ON THE INTERNET CONNECTIVITY:** Installation of the SDK with all the toolkits would require downloading around 4GB of data.
Querying data does not require a lot of bandwidth but you will certainly have a more pleasant experience if you will be on a fast internet line. 4G networks provide a good enough experience so if you're traveling your personal hot-spot will do.
While it's technically possible to use a subset of the functionality in off-line mode, you will not be able to use any data that is queried from the APIs of data providers and services.
### Platform Specific Requirements
**Portfolio Optimization Toolkit and Forecasting Toolkit on Apple Silicon:** To install the Forecasting toolkit on M1/M2 macs you need to use the x86_64 version of conda and install certain dependencies from conda-forge. Follow the [instructions in this section](https://github.com/OpenBB-finance/OpenBBTerminal/blob/main/openbb_terminal/README.md#1-install-miniconda)
**Base Linux Docker containers:** To have the package work in base linux containers like python's `slim-buster` you need to install a C anc C++ compiler that's not bundled with the distribution.
Run `sudo apt update && sudo apt install gcc cmake`
## Installation
We provide a simple installation method in order to utilize the OpenBB SDK. You must first create an environment,
which allows you to isolate the SDK from the rest of your system. It is our recommendation that you utilize a
`conda` environment because there are optional features, such as `forecast`, that utilize libraries that are
specifically sourced from `conda-forge`. Due to this, if you do not use a conda environment, you will not be
able to use some of these features. As such, the installation steps will be written under the assumption that
you are using conda.
### Steps
#### 1. **Install [Miniconda](https://docs.conda.io/en/latest/miniconda.html)**
Download the `x86_64` Miniconda for your respective system and follow along with it's installation instructions. The Miniconda architecture MUST be `x86_64` in order to use the forecasting toolkit. Follow the [instructions in this section](https://github.com/OpenBB-finance/OpenBBTerminal/blob/main/openbb_terminal/README.md#1-install-miniconda)
#### 2. **Create the virtual environment**
```bash
conda create -n obb python=3.9.6 -y
```
#### 3. **Activate the virtual environment**
```bash
conda activate obb
```
#### 4. **Install OpenBB SDK Core package**
```bash
pip install openbb
```
#### 5. **(Optional) Install the Toolkits**
##### 5.1 **If you would like to use the Portfolio Optimization features**
On Apple Silicon Macs (M1/M2) install dependency from conda-forge
```bash
conda install -c conda-forge cvxpy=1.2.2 -y
```
And install the Portfolio Optimization Toolkit
```bash
pip install "openbbterminal[optimization]"
```
##### 5.2 **If you would like ML Forecasting features**
On Apple Silicon Macs (M1/M2) install dependency from conda-forge
```bash
conda install -c conda-forge lightgbm=3.3.3 -y
```
And install the Forecasting Toolkit
```bash
pip install "openbbterminal[prediction]"
```
##### 5.2 **If you would like to use both Portfolio Optimization and ML forecast features**
On Apple Silicon Macs (M1/M2) install dependencies from conda-forge
```bash
conda install -c conda-forge lightgbm=3.3.3 cvxpy=1.2.2 -y
```
And install the Both Toolkits
```bash
pip install "openbbterminal[all]"
```
Congratulations! You have successfully installed `openbbterminal` on an environment and are now able to begin using it. However, it is important to note that if you close out of your CLI you must re-activate your environment in order begin using it again.
## Setup
### 1. Import OpenBB SDK
First off, import OpenBB SDK into your python script or Jupyter Notebook with:
```python
from openbb_terminal.sdk import openbb
```
This imports all Terminal commands at once. To see all the available commands, you can press `tab` in jupyter notebook.
Another approach is to check out [OpenBB SDK Documentation](https://openbb-finance.github.io/OpenBBTerminal/sdk/), where you can explore its capabilities
### 2. Customize chart style
With OpenBB SDK, you can customize your chart style. You can switch between `dark` and `light` easily using this block of code:
```python
from openbb_terminal.sdk import TerminalStyle
theme = TerminalStyle("light", "light", "light")
```
<img width="813" alt="Screenshot 2022-10-03 at 23 56 52" src="https://user-images.githubusercontent.com/40023817/193700307-cbb12edc-0a5d-4804-9f3c-a798efd9e69d.png">
OR
```python
from openbb_terminal.sdk import TerminalStyle
theme = TerminalStyle("dark", "dark", "dark")
```
<img width="791" alt="Screenshot 2022-10-03 at 23 46 33" src="https://user-images.githubusercontent.com/40023817/193699221-e154995b-653c-40fd-8fc6-a3f8d39638db.png">
### 3. Access Documentation
Each and every command of OpenBB SDK has detailed documentation about input parameters and returned outputs. You can access them in multiple ways:
**Approach 1: Press `shift + tab`.**
This will work out of the box if you're using Jupyter Notebook. In case your IDE is VSCode, you will need to install the [Jupyter PowerToys
extension](https://marketplace.visualstudio.com/items?itemName=ms-toolsai.vscode-jupyter-powertoys).
<img width="788" alt="Screenshot 2022-10-03 at 23 31 55" src="https://user-images.githubusercontent.com/40023817/193697567-e7143252-c560-441e-84fd-cbe38aeaf0ea.png">
**Approach 2: Type `help(command)`.**
You can also type `help(command)`, see example below, to see the command' docstring.
<img width="871" alt="Screenshot 2022-10-03 at 23 33 05" src="https://user-images.githubusercontent.com/40023817/193697676-39351008-386d-4c4c-89f2-3de7d8d4e89d.png">
**Approach 3: Use OpenBB SDK Documentation page.**
Finally, if you prefer to check documentation on a web browser, [OpenBB SDK Documentation](https://openbb-finance.github.io/OpenBBTerminal/sdk/) will be your best friend. You can browse available commands and search for any specific one that you need.
<img width="1200" alt="Screenshot 2022-10-03 at 18 41 48" src="https://user-images.githubusercontent.com/40023817/193643316-c063df03-4172-487f-ba47-ee60f36a3fef.png">
### 4. Set API Keys
You can set your external API keys through OpenBB SDK.
- Single API setup
```python
openbb.keys.fmp(key="example")
openbb.keys.reddit(
client_id="example",
client_secret="example",
password="example",
username="example",
useragent="example")
```
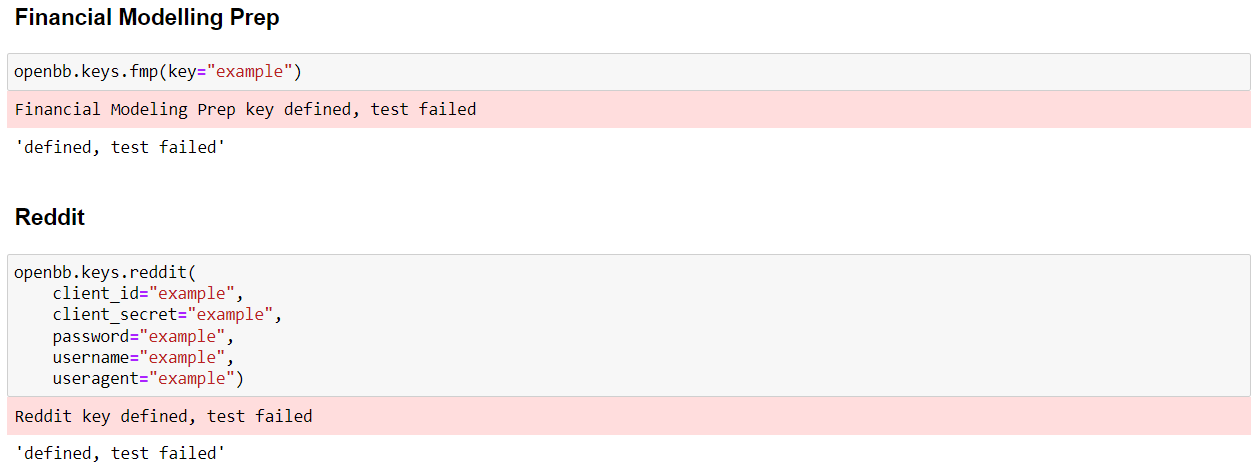
- API key setup with persistence: `persist=True` means that your key will be saved and can be reused after, otherwise it will be lost when you restart the kernel.
```python
openbb.keys.fmp(key="example", persist=True)
```
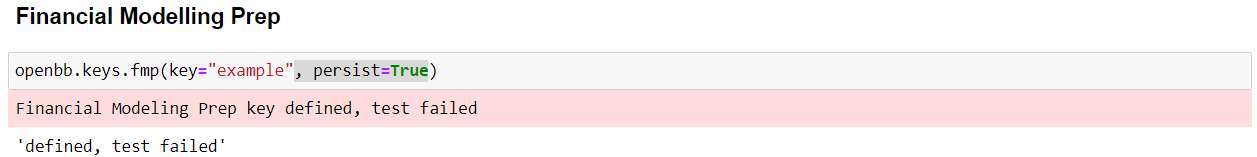
- Set multiple keys from dictionary
```python
d = {
"fed": {
"key":"XXXXX"
},
"binance": {
"key":"YYYYY",
"secret":"example"
},
}
openbb.keys.set_keys(d)
```
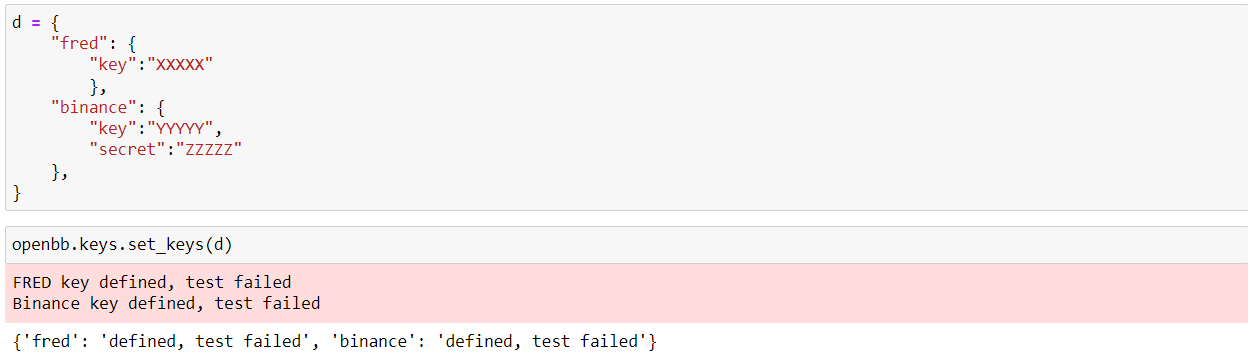
- Get info about API setup arguments
```python
openbb.keys.get_keys_info()
```
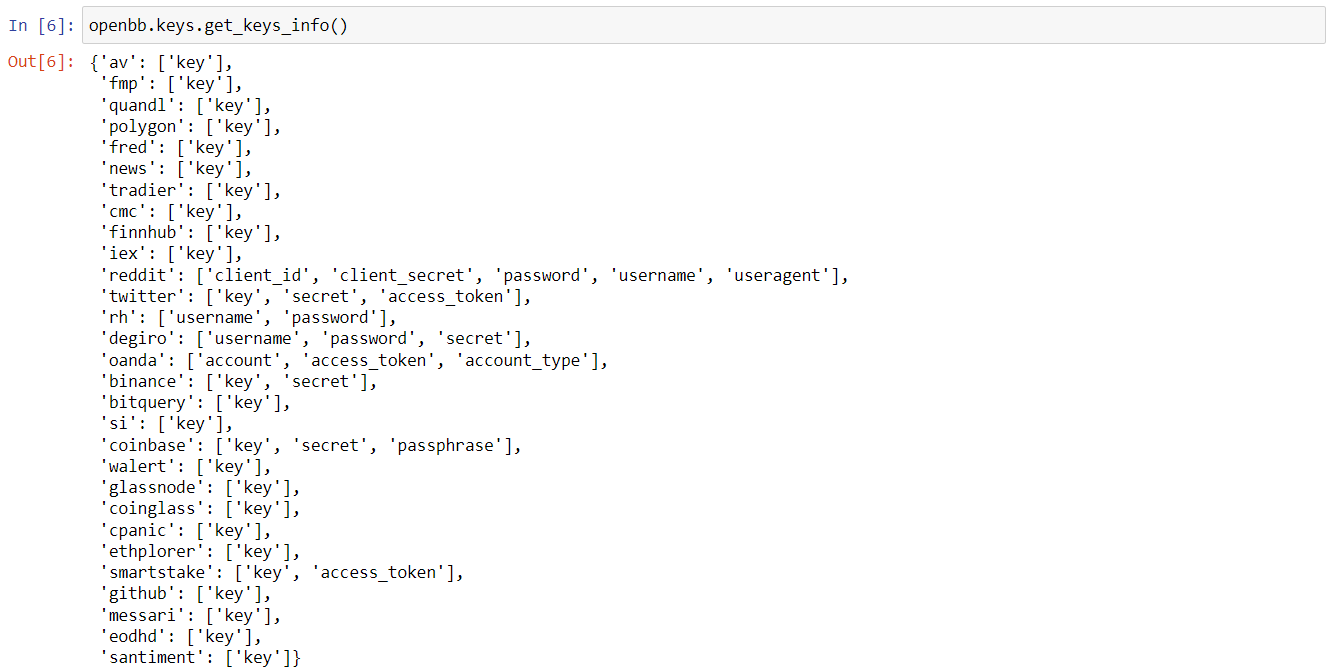
- Get your defined keys
```python
openbb.keys.mykeys()
openbb.keys.mykeys(show=True)
```
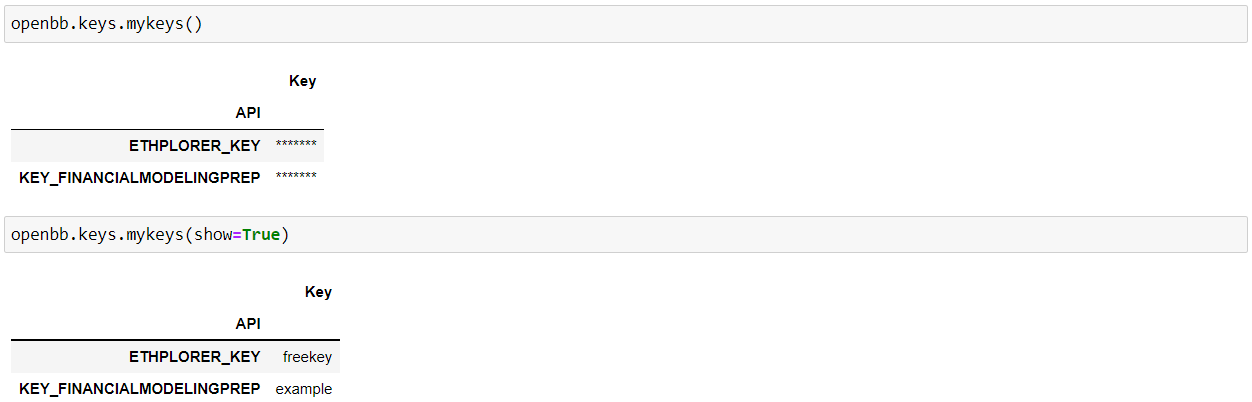
## Usage
Now, let's explore what OpenBB SDK can do. At a high level, you can break down OpenBB SDK's functionalities into two main buckets: (1) Data layer and (2) Toolbox layer.
### 1. Data Layer
### **Getting financial data from multiple data sources using one single SDK**
OpenBB SDK provides you access to normalized financial data from dozens of data sources, without having to built your own integration or relying on multiple third-party packages. Let's explore how we can do that.
First, you will need to load in the desired ticker. If it's not on the top of your mind, make use of our search functionality.
```python
openbb.stocks.search("apple")
```
<img width="652" alt="Screenshot 2022-10-04 at 00 00 14" src="https://user-images.githubusercontent.com/40023817/193700663-b91d57a9-4581-4f7e-a6da-764c0c9de092.png">
We want to load `Apple Inc.` listed on US exchange, so our ticker should be `AAPL`. If you want to load `Apple Inc.` from Brazilian exchange, you should load in `AAPL34.SA`.
```python
df = openbb.stocks.load("AAPL")
```
What's extremely powerful about OpenBB SDK is that you can specify the data source. Depending on the asset class, we have a list of available data sources and it's only getting bigger with contributions from our open-source community.
```python
## From YahooFinance
df_yf = openbb.stocks.load("AAPL", source='YahooFinance')
## From AlphaVantage
df_av = openbb.stocks.load("AAPL", source='AlphaVantage')
## From IEXCloud
df_iex = openbb.stocks.load("AAPL", source='IEXCloud')
## From Polygon
df_pol = openbb.stocks.load("AAPL", source='Polygon')
```
### **Easy option to switch between obtaining underlying data and charts**
Depending on your needs, you can get the outputs in form of data (e.g. `pandas dataframe`) or charts. If the latter is what you want, simple add `chart=True` as the last parameter.
### 1. Getting underlying data
```python
openbb.economy.index(indices = ['sp500', 'nyse_ny', 'russell1000'], start_date = '2010-01-01')
```
<img width="575" alt="Screenshot 2022-10-04 at 00 02 23" src="https://user-images.githubusercontent.com/40023817/193700891-f4d93440-31e3-411e-9931-3a38782f68e3.png">
You might be wondering how to find all the available indices. This type of information should be available in the docstring. Let's give it a try.
<img width="906" alt="Screenshot 2022-10-04 at 13 20 58" src="https://user-images.githubusercontent.com/40023817/193817866-b05cacee-a11b-4c44-b8c3-efb51bb9c892.png">
As mentioned in the docstring, you can access it with the following helper function.
```python
openbb.economy.available_indices()
```
<img width="1078" alt="Screenshot 2022-10-04 at 00 16 36" src="https://user-images.githubusercontent.com/40023817/193702595-ecbfc84d-3ed1-4f89-9086-e975b01c4b12.png">
### 2. Getting charts
```python
openbb.economy.index(indices = ['sp500', 'nyse_ny', 'russell1000'], start_date = '2010-01-01', chart=True)
```
<img width="741" alt="Screenshot 2022-10-04 at 00 03 57" src="https://user-images.githubusercontent.com/40023817/193701075-796ffabe-3266-4d71-9a81-3042e8ca5fc8.png">
## 2. Toolbox Layer
In addition to financial data, you can also get access to a robust and powerful toolbox to perform analysis on different asset classes and on your portfolio.
Imagine that you would like to leverage existing financial calculations from OpenBB and apply them on your own data. This can be done easily - OpenBB SDK's commands usually accept a `dataframe` as an input. Here you can load it your data, either via a `csv`, `excel` file, or connecting directly with an `API` or a `database`. The possibilities are endless.
Let's go through an example to see how we can do it in a few simple steps. Here we shall see how to use `portfolio optimization` functionalities from OpenBB SDK.
### Step 1. Loading order book
Here we will use an example orderbook for illustration purposes. You can choose to upload your own orderbook instead.
```python
order_book_path = "portfolio/allocation/60_40_Portfolio.xlsx"
tickers, categories = openbb.portfolio.po.load(excel_file = order_book_path)
```
### Step 2. Optimizing portfolio
We provide multiple portfolio optimization techniques. You can utilize basic mean-variance techniques, such as optimizing for the maximum Sharpe ratio, or minimum variance, as well as advanced optimization techniques including Hierarchical Risk Parity and Nested Clustered Optimization.
```python
## Max Sharpe optimization
weights_max_sharpe, data_returns_max_sharpe = openbb.portfolio.po.maxsharpe(tickers)
print("Max Sharpe")
weights_max_sharpe
```
<img width="734" alt="Screenshot 2022-10-04 at 13 23 45" src="https://user-images.githubusercontent.com/40023817/193818381-e3e75455-ea91-4bdd-a903-0874ac8700dc.png">
```python
## Minimum risk optimization
weights_min_risk, data_returns_min_risk = openbb.portfolio.po.minrisk(tickers)
print("Min Risk")
weights_min_risk
```
<img width="742" alt="Screenshot 2022-10-04 at 13 24 45" src="https://user-images.githubusercontent.com/40023817/193818556-89380c7c-94c3-4e5c-8848-28058c9cf056.png">
```python
## Hierarchical Risk Parity optimization
weights_hrp, data_returns_hrp = openbb.portfolio.po.hrp(tickers)
print("Hierarchical Risk Parity")
weights_hrp
```
<img width="736" alt="Screenshot 2022-10-04 at 13 34 39" src="https://user-images.githubusercontent.com/40023817/193820500-1bcde650-f517-4aed-b989-b2bd92bebbb8.png">
After having obtained the asset allocation outcomes, you can plot a correlation heatmap across tickers, as well as their individual risk contribution.
```python
openbb.portfolio.po.plot(data=data_returns_hrp,weights=weights_hrp,heat=True)
```
<img width="734" alt="Screenshot 2022-10-04 at 13 35 14" src="https://user-images.githubusercontent.com/40023817/193820624-3e6da926-aea9-4963-bd54-fd1a6df0fda3.png">
```python
openbb.portfolio.po.plot(data=data_returns_hrp,weights=weights_hrp,rc_chart=True)
```
<img width="737" alt="Screenshot 2022-10-04 at 13 36 10" src="https://user-images.githubusercontent.com/40023817/193820817-82f8727f-0e12-4794-b128-d6ebe20b2c4f.png">
These techniques have an extensive list of parameters and thus the optimization outcome is highly dependent on the chosen parameters. For instance, you can refer to the documentation below.
<img width="747" alt="Screenshot 2022-10-04 at 00 35 00" src="https://user-images.githubusercontent.com/40023817/193704210-b75ddee3-1da3-432b-90f8-6966e85bb345.png">
This allows us to alter certain assumption which also modify the asset allocation.
```python
weights_hrp_2, data_returns_hrp_2 = openbb.portfolio.po.hrp(
tickers,
interval="5y",
risk_measure="cVaR",
risk_aversion=0.8
)
pd.DataFrame([weights_hrp, weights_hrp_2], index=["Basic", "Extended"]).T
```
<img width="401" alt="Screenshot 2022-10-04 at 00 37 18" src="https://user-images.githubusercontent.com/40023817/193704462-d006deee-f009-4330-9918-0e0d661636d8.png">
The basic method was optimized for *variance*. The extended method increases the period of historical data, optimizes for conditional Value at Risk and has a lower risk aversion.
```python
openbb.portfolio.po.plot(data=data_returns_hrp,weights=weights_hrp,pie=True)
```
<img width="735" alt="Screenshot 2022-10-04 at 13 38 12" src="https://user-images.githubusercontent.com/40023817/193821181-0cb8cc51-3532-4542-b098-b23222330142.png">
```python
openbb.portfolio.po.plot(data=data_returns_hrp_2,weights=weights_hrp_2,pie=True)
```
<img width="735" alt="Screenshot 2022-10-04 at 13 38 30" src="https://user-images.githubusercontent.com/40023817/193821231-e92839b5-47d1-4a1a-81c2-61244bb6d925.png">
## Useful tips
### 1. Display matplotlib charts in Jupyter Notebook
To display matplotlib charts inside the Jupyter notebook output cells, you can use the block of code below, and initialize it at the top of the Notebook.
```python
import matplotlib.pyplot as plt
import matplotlib_inline.backend_inline
from openbb_terminal.sdk import openbb
%matplotlib inline
matplotlib_inline.backend_inline.set_matplotlib_formats("svg")
```
### 2. Take advantage of `external_axes`
The code below utilizes the `external_axes` parameter to get two axis in one chart.
```python
import matplotlib.pyplot as plt
from openbb_terminal.sdk import openbb
fig, (ax1, ax2) = plt.subplots(nrows=2, ncols=1, figsize=(11, 5), dpi=150)
openbb.stocks.dps.dpotc(
"aapl",
external_axes=[ax1, ax2],
chart=True,
)
fig.tight_layout()
```
You can also do this to save output charts in a variable for later uses.
### For more examples, we'd recommend checking out our [curated Jupyter Notebook reports](https://github.com/OpenBB-finance/OpenBBTerminal/tree/main/openbb_terminal/reports). They are excellent demonstration on how to use the SDK to its fullest extent
| /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/SDK_README.md | 0.566738 | 0.85741 | SDK_README.md | pypi |
__docformat__ = "numpy"
import os
from pathlib import Path
import argparse
import json
from importlib import machinery, util
from typing import Union, List, Dict, Optional
import matplotlib.pyplot as plt
from matplotlib import font_manager, ticker
from openbb_terminal.core.config.paths import (
MISCELLANEOUS_DIRECTORY,
USER_DATA_DIRECTORY,
)
# pylint: disable=too-few-public-methods
class ModelsNamespace:
"""A namespace placeholder for the menu models.
This class is used in all api wrappers to create a `models` namespace and import
all the model functions.
"""
def __init__(self, folders: Union[str, List[str]]) -> None:
"""Import all menu models into the models namespace.
Instantiation of the namespace requires either a path to the folder that
contains model files or a list of such folders.
Parameters
----------
folders : Union[str, List[str]]
a folder or a list of folders to import models from
"""
if isinstance(folders, str):
folders = [folders]
for folder in folders:
menu_models = [
(
f.replace("_model.py", ""),
os.path.abspath(os.path.join(folder, f)),
)
for f in os.listdir(folder)
if f.endswith("_model.py")
]
for model_name, model_file in menu_models:
loader = machinery.SourceFileLoader(model_name, model_file)
spec = util.spec_from_loader(model_name, loader)
if spec is not None:
setattr(self, model_name, util.module_from_spec(spec))
loader.exec_module(getattr(self, model_name))
else:
pass
# pylint: disable=R0902
class TerminalStyle:
"""The class that helps with handling of style configurations.
It serves styles for 3 libraries. For `Matplotlib` this class serves absolute paths
to the .mplstyle files. For `Matplotlib Finance` and `Rich` this class serves custom
styles as python dictionaries.
"""
DEFAULT_STYLES_LOCATION = MISCELLANEOUS_DIRECTORY / "styles" / "default"
USER_STYLES_LOCATION = USER_DATA_DIRECTORY / "styles" / "user"
mpl_styles_available: Dict[str, str] = {}
mpl_style: str = ""
mpl_rcparams_available: Dict[str, str] = {}
mpl_rcparams: Dict = {}
mpf_styles_available: Dict[str, str] = {}
mpf_style: Dict = {}
console_styles_available: Dict[str, str] = {}
console_style: Dict[str, str] = {}
down_color: str = ""
up_color: str = ""
xticks_rotation: str = ""
tight_layout_padding: int = 0
pie_wedgeprops: Dict = {}
pie_startangle: int = 0
line_width: float = 1.5
volume_bar_width: float = 0.8
def __init__(
self,
mpl_style: Optional[str] = "",
mpf_style: Optional[str] = "",
console_style: Optional[str] = "",
) -> None:
"""Instantiate a terminal style class
The stylesheet files should be placed to the `styles/default` or `styles/user`
folders. The parameters required for class instantiation are stylesheet names
without extensions (following matplotlib convention).
Ex. `styles/default/boring.mplstyle` should be passed as `boring`.
Parameters
----------
mpl_style : str, optional
Style name without extension, by default ""
mpf_style : str, optional
Style name without extension, by default ""
console_style : str, optional
Style name without extension, by default ""
"""
# To import all styles from terminal repo folder to user data
for folder in [self.DEFAULT_STYLES_LOCATION, self.USER_STYLES_LOCATION]:
self.load_available_styles_from_folder(folder)
self.load_custom_fonts_from_folder(folder)
if mpl_style in self.mpl_styles_available:
self.mpl_style = self.mpl_styles_available[mpl_style]
else:
self.mpl_style = self.mpl_styles_available["dark"]
if mpl_style in self.mpl_rcparams_available:
with open(self.mpl_rcparams_available[mpl_style]) as stylesheet:
self.mpl_rcparams = json.load(stylesheet)
else:
with open(self.mpl_rcparams_available["dark"]) as stylesheet:
self.mpl_rcparams = json.load(stylesheet)
if mpf_style in self.mpf_styles_available:
with open(self.mpf_styles_available[mpf_style]) as stylesheet:
self.mpf_style = json.load(stylesheet)
self.mpf_style["base_mpl_style"] = self.mpl_style
else:
with open(self.mpf_styles_available["dark"]) as stylesheet:
self.mpf_style = json.load(stylesheet)
self.mpf_style["base_mpl_style"] = self.mpl_style
if "openbb_config" in self.console_styles_available:
with open(self.console_styles_available["openbb_config"]) as stylesheet:
self.console_style = json.load(stylesheet)
elif console_style in self.console_styles_available:
with open(self.console_styles_available[console_style]) as stylesheet:
self.console_style = json.load(stylesheet)
else:
with open(self.console_styles_available["dark"]) as stylesheet:
self.console_style = json.load(stylesheet)
self.applyMPLstyle()
def load_custom_fonts_from_folder(self, folder: Path) -> None:
"""Load custom fonts form folder.
TTF and OTF fonts are loaded into the mpl font manager and are available for
selection in mpl by their name (for example "Consolas" or "Hack").
Parameters
----------
folder : str
Path to the folder containing the fonts
"""
if not folder.exists():
return
for font_file in folder.iterdir():
if not font_file.is_file():
continue
if font_file.name.endswith(".otf") or font_file.name.endswith(".ttf"):
font_path = os.path.abspath(os.path.join(folder, font_file))
font_manager.fontManager.addfont(font_path)
def load_available_styles_from_folder(self, folder: Path) -> None:
"""Load custom styles from folder.
Parses the styles/default and styles/user folders and loads style files.
To be recognized files need to follow a naming convention:
*.mplstyle - matplotlib stylesheets
*.mplrc.json - matplotlib rc stylesheets that are not handled by mplstyle
*.mpfstyle.json - matplotlib finance stylesheets
*.richstyle.json - rich stylesheets
Parameters
----------
folder : str
Path to the folder containing the stylesheets
"""
if not folder.exists():
return
for stf in folder.iterdir():
if not stf.is_file():
continue
if stf.name.endswith(".mplstyle"):
self.mpl_styles_available[
stf.name.replace(".mplstyle", "")
] = os.path.join(folder, stf)
elif stf.name.endswith(".mplrc.json"):
self.mpl_rcparams_available[
stf.name.replace(".mplrc.json", "")
] = os.path.join(folder, stf)
elif stf.name.endswith(".mpfstyle.json"):
self.mpf_styles_available[
stf.name.replace(".mpfstyle.json", "")
] = os.path.join(folder, stf)
elif stf.name.endswith(".richstyle.json"):
self.console_styles_available[
stf.name.replace(".richstyle.json", "")
] = os.path.join(folder, stf)
def applyMPLstyle(self):
"""Apply style to the current matplotlib context."""
plt.style.use(self.mpl_style)
self.xticks_rotation = self.mpl_rcparams["xticks_rotation"]
self.tight_layout_padding = self.mpl_rcparams["tight_layout_padding"]
self.pie_wedgeprops = self.mpl_rcparams["pie_wedgeprops"]
self.pie_startangle = self.mpl_rcparams["pie_startangle"]
self.mpf_style["mavcolors"] = plt.rcParams["axes.prop_cycle"].by_key()["color"]
self.down_color = self.mpf_style["marketcolors"]["volume"]["down"]
self.up_color = self.mpf_style["marketcolors"]["volume"]["up"]
self.line_width = plt.rcParams["lines.linewidth"]
try:
self.volume_bar_width = self.mpl_rcparams["volume_bar_width"]
except Exception():
pass
def get_colors(self, reverse: bool = False) -> List:
"""Get hex color sequence from the stylesheet."""
plt.style.use(self.mpl_style)
colors = plt.rcParams["axes.prop_cycle"].by_key()["color"]
if reverse:
colors.reverse()
return colors
def style_primary_axis(
self,
ax: plt.Axes,
data_index: Optional[List[int]] = None,
tick_labels: Optional[List[str]] = None,
):
"""Apply styling to a primary axis.
Parameters
----------
ax : plt.Axes
A matplolib axis
"""
ax.yaxis.set_label_position("right")
ax.grid(axis="both", visible=True, zorder=0)
if (
all([data_index, tick_labels])
and isinstance(data_index, list)
and isinstance(tick_labels, list)
):
ax.xaxis.set_major_formatter(
ticker.FuncFormatter(
lambda value, _: tick_labels[int(value)]
if int(value) in data_index
else ""
)
)
ax.xaxis.set_major_locator(ticker.MaxNLocator(6, integer=True))
ax.tick_params(axis="x", labelrotation=self.xticks_rotation)
def style_twin_axis(self, ax: plt.Axes):
"""Apply styling to a twin axis.
Parameters
----------
ax : plt.Axes
A matplolib axis
"""
ax.yaxis.set_label_position("left")
def style_twin_axes(self, ax1: plt.Axes, ax2: plt.Axes):
"""Apply styling to a twin axes
Parameters
----------
ax1 : plt.Axes
Primary matplolib axis
ax2 : plt.Axes
Twinx matplolib axis
"""
ax1.tick_params(axis="x", labelrotation=self.xticks_rotation)
ax1.grid(axis="both", visible=True, zorder=0)
ax2.grid(visible=False)
def add_label(
self,
fig: plt.figure,
):
"""Add a text label to a figure in a funny position.
Parameters
----------
fig : plt.figure
A matplotlib figure
"""
label = "OpenBB Terminal"
fig.text(
0.99,
0.0420,
label,
fontsize=12,
color="gray",
alpha=0.5,
horizontalalignment="right",
)
# pylint: disable=import-outside-toplevel
def add_cmd_source(
self,
fig: plt.figure,
):
"""Add a text label to a figure in a funny position.
Parameters
----------
fig : plt.figure
A matplotlib figure
"""
from openbb_terminal.helper_funcs import command_location
if command_location:
fig.text(
0.01,
0.5,
command_location,
rotation=90,
fontsize=12,
color="gray",
alpha=0.5,
verticalalignment="center",
)
# pylint: disable=import-outside-toplevel
def visualize_output(self, force_tight_layout: bool = True):
"""Show chart in an interactive widget."""
import openbb_terminal.feature_flags as obbff
if obbff.USE_CMD_LOCATION_FIGURE:
self.add_cmd_source(plt.gcf())
if obbff.USE_WATERMARK:
self.add_label(plt.gcf())
if force_tight_layout:
plt.tight_layout(pad=self.tight_layout_padding)
if obbff.USE_ION:
plt.ion()
plt.show()
class AllowArgsWithWhiteSpace(argparse.Action):
def __call__(self, parser, namespace, values, option_string=None):
setattr(namespace, self.dest, " ".join(values)) | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/helper_classes.py | 0.751739 | 0.259201 | helper_classes.py | pypi |
from typing import Literal, Tuple, Callable, Dict
import pandas as pd
from pandas._typing import Axis
DENOMINATION = Literal[
"Trillions", "Billions", "Millions", "Tens of thousands", "Thousands", "Units", ""
]
def transform(
df: pd.DataFrame,
sourceDenomination: DENOMINATION = "Units",
targetDenomination: DENOMINATION = None,
maxValue: float = None,
axis: Axis = 0,
skipPredicate: Callable[[pd.Series], bool] = None,
) -> Tuple[pd.DataFrame, DENOMINATION]:
"""Transforms data frame by denomination.
Args:
df (pd.DataFrame): Source data frame
sourceDenomination (DENOMINATION, optional): Current denomination. Defaults to Units.
targetDenomination (DENOMINATION, optional): Desired denomination. Defaults to None, meaning we will find it.
maxValue (float, optional): Max value of the data frame. Defaults to None, meaning df.max().max() will be used.
axis (Axis, optional): Axis to apply to skip predicate. Defaults to 0.
skipPredicate (Callable[[pd.Series], bool], optional): Predicate for skipping a transform.
Returns:
pd.DataFrame
Tuple[pd.DataFrame, DENOMINATION]: Pair of transformed data frame and applied denomination.
"""
def apply(
df: pd.DataFrame, source: DENOMINATION, target: DENOMINATION
) -> pd.DataFrame:
multiplier = get_denominations()[source] / get_denominations()[target]
df = df.astype(float)
return df.apply(
lambda series: series
if skipPredicate is not None and skipPredicate(series)
else series * multiplier,
axis,
)
if targetDenomination is not None:
return (
apply(df, sourceDenomination, targetDenomination),
targetDenomination,
)
if maxValue is None:
maxValue = df.max(numeric_only=True).max(numeric_only=True)
foundTargetDenomination = get_denomination(maxValue)
return (
apply(df, sourceDenomination, foundTargetDenomination[0]),
foundTargetDenomination[0],
)
def get_denominations() -> Dict[DENOMINATION, float]:
"""Gets all supported denominations and their lower bound value
Returns:
Dict[DENOMINATION, int]: All supported denominations and their lower bound value
"""
return {
"Trillions": 1_000_000_000_000,
"Billions": 1_000_000_000,
"Millions": 1_000_000,
"Tens of thousands": 10_000,
"Thousands": 1_000,
"Units": 1,
}
def get_denomination(value: float) -> Tuple[DENOMINATION, float]:
"""Gets denomination that fits the supplied value.
If no denomination found, 'Units' is returned.
Args:
value (int): Value
Returns:
Tuple[DENOMINATION, int]:Denomination that fits the supplied value
"""
return next((x for x in get_denominations().items() if value >= x[1]), ("Units", 1)) | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/helpers_denomination.py | 0.935971 | 0.512022 | helpers_denomination.py | pypi |
from typing import (
Any,
Dict,
List,
Set,
Iterable,
Mapping,
Optional,
Union,
Pattern,
Callable,
)
from prompt_toolkit.completion import CompleteEvent, Completer, Completion
from prompt_toolkit.document import Document
from prompt_toolkit.formatted_text import AnyFormattedText
NestedDict = Mapping[str, Union[Any, Set[str], None, Completer]]
# pylint: disable=too-many-arguments,global-statement,too-many-branches,global-variable-not-assigned
class WordCompleter(Completer):
"""
Simple autocompletion on a list of words.
:param words: List of words or callable that returns a list of words.
:param ignore_case: If True, case-insensitive completion.
:param meta_dict: Optional dict mapping words to their meta-text. (This
should map strings to strings or formatted text.)
:param WORD: When True, use WORD characters.
:param sentence: When True, don't complete by comparing the word before the
cursor, but by comparing all the text before the cursor. In this case,
the list of words is just a list of strings, where each string can
contain spaces. (Can not be used together with the WORD option.)
:param match_middle: When True, match not only the start, but also in the
middle of the word.
:param pattern: Optional compiled regex for finding the word before
the cursor to complete. When given, use this regex pattern instead of
default one (see document._FIND_WORD_RE)
"""
def __init__(
self,
words: Union[List[str], Callable[[], List[str]]],
ignore_case: bool = False,
display_dict: Optional[Mapping[str, AnyFormattedText]] = None,
meta_dict: Optional[Mapping[str, AnyFormattedText]] = None,
WORD: bool = False,
sentence: bool = False,
match_middle: bool = False,
pattern: Optional[Pattern[str]] = None,
) -> None:
assert not (WORD and sentence)
self.words = words
self.ignore_case = ignore_case
self.display_dict = display_dict or {}
self.meta_dict = meta_dict or {}
self.WORD = WORD
self.sentence = sentence
self.match_middle = match_middle
self.pattern = pattern
def get_completions(
self,
document: Document,
_complete_event: CompleteEvent,
) -> Iterable[Completion]:
# Get list of words.
words = self.words
if callable(words):
words = words()
# Get word/text before cursor.
if self.sentence:
word_before_cursor = document.text_before_cursor
else:
word_before_cursor = document.get_word_before_cursor(
WORD=self.WORD, pattern=self.pattern
)
if (
"--" in document.text_before_cursor
and document.text_before_cursor.rfind(" --")
>= document.text_before_cursor.rfind(" -")
):
word_before_cursor = f'--{document.text_before_cursor.split("--")[-1]}'
elif f"--{word_before_cursor}" == document.text_before_cursor:
word_before_cursor = document.text_before_cursor
if self.ignore_case:
word_before_cursor = word_before_cursor.lower()
def word_matches(word: str) -> bool:
"""True when the word before the cursor matches."""
if self.ignore_case:
word = word.lower()
if self.match_middle:
return word_before_cursor in word
return word.startswith(word_before_cursor)
for a in words:
if word_matches(a):
display = self.display_dict.get(a, a)
display_meta = self.meta_dict.get(a, "")
yield Completion(
text=a,
start_position=-len(word_before_cursor),
display=display,
display_meta=display_meta,
)
class NestedCompleter(Completer):
"""
Completer which wraps around several other completers, and calls any the
one that corresponds with the first word of the input.
By combining multiple `NestedCompleter` instances, we can achieve multiple
hierarchical levels of autocompletion. This is useful when `WordCompleter`
is not sufficient.
If you need multiple levels, check out the `from_nested_dict` classmethod.
"""
complementary: List = list()
def __init__(
self, options: Dict[str, Optional[Completer]], ignore_case: bool = True
) -> None:
self.flags_processed: List = list()
self.original_options = options
self.options = options
self.ignore_case = ignore_case
self.complementary = list()
def __repr__(self) -> str:
return f"NestedCompleter({self.options!r}, ignore_case={self.ignore_case!r})"
@classmethod
def from_nested_dict(cls, data: dict) -> "NestedCompleter":
"""
Create a `NestedCompleter`, starting from a nested dictionary data
structure, like this:
.. code::
data = {
'show': {
'version': None,
'interfaces': None,
'clock': None,
'ip': {'interface': {'brief'}}
},
'exit': None
'enable': None
}
The value should be `None` if there is no further completion at some
point. If all values in the dictionary are None, it is also possible to
use a set instead.
Values in this data structure can be a completers as well.
"""
options: Dict[str, Any] = {}
for key, value in data.items():
if isinstance(value, Completer):
options[key] = value
elif isinstance(value, dict):
options[key] = cls.from_nested_dict(value)
elif isinstance(value, set):
options[key] = cls.from_nested_dict({item: None for item in value})
elif isinstance(key, str) and isinstance(value, str):
options[key] = options[value]
else:
assert value is None
options[key] = None
for items in cls.complementary:
if items[0] in options:
options[items[1]] = options[items[0]]
elif items[1] in options:
options[items[0]] = options[items[1]]
return cls(options)
def get_completions(
self, document: Document, complete_event: CompleteEvent
) -> Iterable[Completion]:
# Split document.
cmd = ""
text = document.text_before_cursor.lstrip()
if " " in text:
cmd = text.split(" ")[0]
if "-" in text:
if text.rfind("--") == -1:
unprocessed_text = "-" + text.split("-")[-1]
elif text.rfind("-") - 1 > text.rfind("--"):
unprocessed_text = "-" + text.split("-")[-1]
else:
unprocessed_text = "--" + text.split("--")[-1]
else:
unprocessed_text = text
stripped_len = len(document.text_before_cursor) - len(text)
# Check if there are multiple flags for the same command
if self.complementary:
for same_flags in self.complementary:
if (
same_flags[0] in self.flags_processed
and same_flags[1] not in self.flags_processed
) or (
same_flags[1] in self.flags_processed
and same_flags[0] not in self.flags_processed
):
if same_flags[0] in self.flags_processed:
self.flags_processed.append(same_flags[1])
elif same_flags[1] in self.flags_processed:
self.flags_processed.append(same_flags[0])
if cmd:
self.options = {
k: self.original_options.get(cmd).options[k] # type: ignore
for k in self.original_options.get(cmd).options.keys() # type: ignore
if k not in self.flags_processed
}
else:
self.options = {
k: self.original_options[k]
for k in self.original_options.keys()
if k not in self.flags_processed
}
# If there is a space, check for the first term, and use a subcompleter.
if " " in unprocessed_text:
first_term = unprocessed_text.split()[0]
# user is updating one of the values
if unprocessed_text[-1] != " ":
self.flags_processed = [
flag for flag in self.flags_processed if flag != first_term
]
if self.complementary:
for same_flags in self.complementary:
if (
same_flags[0] in self.flags_processed
and same_flags[1] not in self.flags_processed
) or (
same_flags[1] in self.flags_processed
and same_flags[0] not in self.flags_processed
):
if same_flags[0] in self.flags_processed:
self.flags_processed.remove(same_flags[0])
elif same_flags[1] in self.flags_processed:
self.flags_processed.remove(same_flags[1])
if cmd and self.original_options.get(cmd):
self.options = self.original_options
else:
self.options = {
k: self.original_options[k]
for k in self.original_options.keys()
if k not in self.flags_processed
}
if "-" not in text:
completer = self.options.get(first_term)
else:
if cmd in self.options and self.options.get(cmd):
completer = self.options.get(cmd).options.get(first_term) # type: ignore
else:
completer = self.options.get(first_term)
# If we have a sub completer, use this for the completions.
if completer is not None:
remaining_text = unprocessed_text[len(first_term) :].lstrip()
move_cursor = len(text) - len(remaining_text) + stripped_len
new_document = Document(
remaining_text,
cursor_position=document.cursor_position - move_cursor,
)
# Provides auto-completion but if user doesn't take it still keep going
if " " in new_document.text:
if (
new_document.text in [f"{opt} " for opt in self.options]
or unprocessed_text[-1] == " "
):
self.flags_processed.append(first_term)
if cmd:
self.options = {
k: self.original_options.get(cmd).options[k] # type: ignore
for k in self.original_options.get(cmd).options.keys() # type: ignore
if k not in self.flags_processed
}
else:
self.options = {
k: self.original_options[k]
for k in self.original_options.keys()
if k not in self.flags_processed
}
# In case the users inputs a single boolean flag
elif not completer.options: # type: ignore
self.flags_processed.append(first_term)
if self.complementary:
for same_flags in self.complementary:
if (
same_flags[0] in self.flags_processed
and same_flags[1] not in self.flags_processed
) or (
same_flags[1] in self.flags_processed
and same_flags[0] not in self.flags_processed
):
if same_flags[0] in self.flags_processed:
self.flags_processed.append(same_flags[1])
elif same_flags[1] in self.flags_processed:
self.flags_processed.append(same_flags[0])
if cmd:
self.options = {
k: self.original_options.get(cmd).options[k] # type: ignore
for k in self.original_options.get(cmd).options.keys() # type: ignore
if k not in self.flags_processed
}
else:
self.options = {
k: self.original_options[k]
for k in self.original_options.keys()
if k not in self.flags_processed
}
else:
# This is a NestedCompleter
yield from completer.get_completions(new_document, complete_event)
# No space in the input: behave exactly like `WordCompleter`.
else:
# check if the prompt has been updated in the meantime
if " " in text or "-" in text:
actual_flags_processed = [
flag for flag in self.flags_processed if flag in text
]
if self.complementary:
for same_flags in self.complementary:
if (
same_flags[0] in actual_flags_processed
and same_flags[1] not in actual_flags_processed
) or (
same_flags[1] in actual_flags_processed
and same_flags[0] not in actual_flags_processed
):
if same_flags[0] in actual_flags_processed:
actual_flags_processed.append(same_flags[1])
elif same_flags[1] in actual_flags_processed:
actual_flags_processed.append(same_flags[0])
if len(actual_flags_processed) < len(self.flags_processed):
self.flags_processed = actual_flags_processed
if cmd:
self.options = {
k: self.original_options.get(cmd).options[k] # type: ignore
for k in self.original_options.get(cmd).options.keys() # type: ignore
if k not in self.flags_processed
}
else:
self.options = {
k: self.original_options[k]
for k in self.original_options.keys()
if k not in self.flags_processed
}
command = self.options.get(cmd)
if command:
options = command.options # type: ignore
else:
options = {}
command_options = [f"{cmd} {opt}" for opt in options.keys()]
text_list = [text in val for val in command_options]
if cmd and cmd in self.options.keys() and text_list:
completer = WordCompleter(
list(self.options.get(cmd).options.keys()), # type: ignore
ignore_case=self.ignore_case,
)
elif bool([val for val in self.options.keys() if text in val]):
completer = WordCompleter(
list(self.options.keys()), ignore_case=self.ignore_case
)
else:
# The user has delete part of the first command and we need to reset options
if bool([val for val in self.original_options.keys() if text in val]):
self.options = self.original_options
self.flags_processed = list()
completer = WordCompleter(
list(self.options.keys()), ignore_case=self.ignore_case
)
# This is a WordCompleter
yield from completer.get_completions(document, complete_event) | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/custom_prompt_toolkit.py | 0.873983 | 0.338405 | custom_prompt_toolkit.py | pypi |
from __future__ import annotations
import random
import re
import requests
from bs4 import BeautifulSoup
from openbb_terminal.helper_funcs import get_user_agent
from openbb_terminal.rich_config import console
__docformat__ = "numpy"
class ThoughtOfTheDay:
"""ThoughtOfTheDay class"""
def __init__(self, urls: dict[str, str] = None):
"""Constructor"""
self.metadata: dict = {}
if urls is None:
self.urls = {
"Marcus_Aurelius": "https://www.goodreads.com/author/quotes/17212.Marcus_Aurelius",
"Epictetus": "https://www.goodreads.com/author/quotes/13852.Epictetus",
"Seneca": "https://www.goodreads.com/author/quotes/4918776.Seneca",
"Marcus_Tullius_Cicero": "https://www.goodreads.com/author/quotes/13755.Marcus_Tullius_Cicero",
"Aristotle": "https://www.goodreads.com/author/quotes/2192.Aristotle",
"Plato": "https://www.goodreads.com/author/quotes/879.Plato",
"Pythagoras": "https://www.goodreads.com/author/quotes/203707.Pythagoras",
}
return
self.urls = urls
def get_urls(self) -> dict:
"""Getter method for URLs"""
return self.urls
def get_metadata(self, author: str) -> dict:
"""Loads metadata for a given author
Parameters
----------
author : str
Author key - Marcus_Aurelius, Epictetus, Seneca, Marcus_Tullius_Cicero, Aristotle, Plato, Pythagoras
Returns
----------
dict
Metadata dictionary that includes number of quotes, number of pages and first 30 quotes
"""
quotes_page = BeautifulSoup(
requests.get(
self.urls[author],
headers={"User-Agent": get_user_agent()},
).text,
"lxml",
)
find_navigation = quotes_page.find("em", {"class": "current"}).find_parent(
"div"
)
page_count = [
a_page_ref.text.strip("\n")
for a_page_ref in find_navigation.find_all("a", href=True)
]
ret = {}
ret["pages"] = page_count[-2]
find_count = quotes_page.find(string=re.compile("Showing 1-30 of"))
quote_count = re.search(r"Showing 1-30 of (?P<number>[\d,]+)", find_count)
if quote_count:
ret["quoutes"] = quote_count.group("number")
all_quotes = quotes_page.find_all("div", {"class": "quote"})
ret["quotes"] = []
for a_quote in all_quotes:
parsed_quote = {}
parsed_quote = a_quote.find("div", {"class": "quoteText"}).text
ret["quotes"].append(parsed_quote)
return ret
def quote_to_str(self, a_quote: str) -> str:
"""Format a quote parsed from Goodreads into a string
Parameters
----------
a_quote : str
A quote formatted by Goodreads
Returns
----------
str
A string version of the quote
"""
ready = []
prev = None
for a_line in a_quote:
if not prev:
ready.append(a_line)
prev = a_line
continue
if a_line != "\n":
ready.append(a_line)
prev = a_line
continue
if prev == " \n":
ready.pop()
prev = None
return "".join(map(str, ready))
def get_thought_of_the_day():
"""Pick a thought of the day"""
totd = ThoughtOfTheDay()
quotes = []
for an_author in totd.get_urls():
metadata = totd.get_metadata(an_author)
quotes = quotes + metadata["quotes"]
console.print("Thought of the day:")
console.print(
totd.quote_to_str(quotes[random.randint(0, len(quotes) - 1)]) # nosec
)
console.print("\n") | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/thought_of_the_day.py | 0.611498 | 0.341555 | thought_of_the_day.py | pypi |
__docformat__ = "numpy"
# pylint:disable=too-many-lines
import argparse
import logging
from datetime import datetime, timedelta
from typing import List
from openbb_terminal.custom_prompt_toolkit import NestedCompleter
from openbb_terminal import feature_flags as obbff
from openbb_terminal.decorators import log_start_end
from openbb_terminal.helper_funcs import (
EXPORT_BOTH_RAW_DATA_AND_FIGURES,
EXPORT_ONLY_RAW_DATA_ALLOWED,
valid_date,
parse_and_split_input,
)
from openbb_terminal.parent_classes import BaseController
from openbb_terminal.rich_config import console, MenuText
from openbb_terminal.menu import session
from openbb_terminal.futures import yfinance_model, yfinance_view
logger = logging.getLogger(__name__)
def valid_expiry_date(s: str) -> str:
"""Argparse type to check date is in valid format"""
try:
if not s:
return s
return datetime.strptime(s, "%Y-%m").strftime("%Y-%m")
except ValueError as value_error:
logging.exception(str(value_error))
raise argparse.ArgumentTypeError(f"Not a valid date: {s}") from value_error
class FuturesController(BaseController):
"""Futures Controller class"""
CHOICES_COMMANDS = [
"search",
"curve",
"historical",
]
curve_type = "futures"
PATH = "/futures/"
CHOICES_GENERATION = True
def __init__(self, queue: List[str] = None):
"""Constructor"""
super().__init__(queue)
self.all_tickers = yfinance_model.FUTURES_DATA["Ticker"].unique().tolist()
self.all_exchanges = yfinance_model.FUTURES_DATA["Exchange"].unique().tolist()
self.all_categories = yfinance_model.FUTURES_DATA["Category"].unique().tolist()
if session and obbff.USE_PROMPT_TOOLKIT:
self.choices: dict = self.choices_default
self.choices["historical"].update({c: None for c in self.all_tickers})
self.choices["historical"]["--ticker"] = {c: None for c in self.all_tickers}
self.choices["curve"].update({c: None for c in self.all_tickers})
self.completer = NestedCompleter.from_nested_dict(self.choices) # type: ignore
def parse_input(self, an_input: str) -> List:
"""Parse controller input
Overrides the parent class function to handle github org/repo path convention.
See `BaseController.parse_input()` for details.
"""
# Filtering out sorting parameters with forward slashes like P/E
sort_filter = r"((\ -s |\ --sortby ).*?(P\/E|Fwd P\/E|P\/S|P\/B|P\/C|P\/FCF)*)"
custom_filters = [sort_filter]
commands = parse_and_split_input(
an_input=an_input, custom_filters=custom_filters
)
return commands
def print_help(self):
"""Print help"""
mt = MenuText("futures/")
mt.add_cmd("search")
mt.add_raw("\n")
mt.add_cmd("historical")
mt.add_cmd("curve")
console.print(text=mt.menu_text, menu="Futures")
@log_start_end(log=logger)
def call_search(self, other_args: List[str]):
"""Process search command"""
parser = argparse.ArgumentParser(
add_help=False,
formatter_class=argparse.ArgumentDefaultsHelpFormatter,
prog="search",
description="""Search futures. [Source: YahooFinance]""",
)
parser.add_argument(
"-e",
"--exchange",
dest="exchange",
type=str,
choices=self.all_exchanges,
default="",
help="Select the exchange where the future exists",
)
parser.add_argument(
"-c",
"--category",
dest="category",
type=str,
choices=self.all_categories,
default="",
help="Select the category where the future exists",
)
parser.add_argument(
"-d",
"--description",
dest="description",
type=str,
nargs="+",
default="",
help="Select the description future you are interested in",
)
ns_parser = self.parse_known_args_and_warn(
parser, other_args, export_allowed=EXPORT_ONLY_RAW_DATA_ALLOWED
)
if ns_parser:
yfinance_view.display_search(
category=ns_parser.category,
exchange=ns_parser.exchange,
description=" ".join(ns_parser.description),
export=ns_parser.export,
)
@log_start_end(log=logger)
def call_historical(self, other_args: List[str]):
"""Process historical command"""
parser = argparse.ArgumentParser(
add_help=False,
formatter_class=argparse.ArgumentDefaultsHelpFormatter,
prog="historical",
description="""Display futures historical. [Source: YahooFinance]""",
)
parser.add_argument(
"-t",
"--ticker",
dest="ticker",
type=str,
default="",
help="Future ticker to display timeseries separated by comma when multiple, e.g.: BLK,QI",
required="-h" not in other_args,
)
parser.add_argument(
"-s",
"--start",
dest="start",
type=valid_date,
help="Initial date. Default: 3 years ago",
default=(datetime.now() - timedelta(days=3 * 365)),
)
parser.add_argument(
"-e",
"--expiry",
dest="expiry",
type=valid_expiry_date,
help="Select future expiry date with format YYYY-MM",
default="",
)
if other_args and "-" not in other_args[0][0]:
other_args.insert(0, "-t")
ns_parser = self.parse_known_args_and_warn(
parser,
other_args,
export_allowed=EXPORT_BOTH_RAW_DATA_AND_FIGURES,
raw=True,
)
if ns_parser:
yfinance_view.display_historical(
symbols=ns_parser.ticker.upper().split(","),
expiry=ns_parser.expiry,
start_date=ns_parser.start.strftime("%Y-%m-%d"),
raw=ns_parser.raw,
export=ns_parser.export,
)
@log_start_end(log=logger)
def call_curve(self, other_args: List[str]):
"""Process curve command"""
parser = argparse.ArgumentParser(
add_help=False,
formatter_class=argparse.ArgumentDefaultsHelpFormatter,
prog="curve",
description="""Display futures curve. [Source: YahooFinance]""",
)
parser.add_argument(
"-t",
"--ticker",
dest="ticker",
type=lambda x: str(x).upper(),
default="",
help="Future curve to be selected",
required="-h" not in other_args,
metavar="TICKER",
choices=self.all_tickers,
)
if other_args and "-" not in other_args[0][0]:
other_args.insert(0, "-t")
ns_parser = self.parse_known_args_and_warn(
parser,
other_args,
export_allowed=EXPORT_ONLY_RAW_DATA_ALLOWED,
raw=True,
)
if ns_parser:
yfinance_view.display_curve(
symbol=ns_parser.ticker.upper(),
raw=ns_parser.raw,
export=ns_parser.export,
) | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/futures/futures_controller.py | 0.746878 | 0.15876 | futures_controller.py | pypi |
__docformat__ = "numpy"
from typing import Optional, List
from itertools import cycle
import logging
import os
from datetime import datetime, timedelta
from matplotlib import pyplot as plt
from openbb_terminal.config_terminal import theme
from openbb_terminal.config_plot import PLOT_DPI
from openbb_terminal.decorators import log_start_end
from openbb_terminal.futures import yfinance_model
from openbb_terminal.helper_funcs import (
export_data,
plot_autoscale,
print_rich_table,
is_valid_axes_count,
)
from openbb_terminal.rich_config import console
from openbb_terminal.futures.futures_helper import make_white
logger = logging.getLogger(__name__)
@log_start_end(log=logger)
def display_search(
category: str = "",
exchange: str = "",
description: str = "",
export: str = "",
):
"""Display search futures [Source: Yahoo Finance]
Parameters
----------
category: str
Select the category where the future exists
exchange: str
Select the exchange where the future exists
description: str
Select the description of the future
export: str
Type of format to export data
"""
df = yfinance_model.get_search_futures(category, exchange, description)
if df.empty:
console.print("[red]No futures data found.\n[/red]")
return
print_rich_table(df)
console.print()
export_data(
export,
os.path.dirname(os.path.abspath(__file__)),
"search",
df,
)
@log_start_end(log=logger)
def display_historical(
symbols: List[str],
expiry: str = "",
start_date: Optional[str] = None,
raw: bool = False,
export: str = "",
external_axes: Optional[List[plt.Axes]] = None,
):
"""Display historical futures [Source: Yahoo Finance]
Parameters
----------
symbols: List[str]
List of future timeseries symbols to display
expiry: str
Future expiry date with format YYYY-MM
start_date : Optional[str]
Initial date like string (e.g., 2021-10-01)
raw: bool
Display futures timeseries in raw format
export: str
Type of format to export data
external_axes : Optional[List[plt.Axes]], optional
External axes (1 axis is expected in the list), by default None
"""
if start_date is None:
start_date = (datetime.now() - timedelta(days=3 * 365)).strftime("%Y-%m-%d")
symbols_validated = list()
for symbol in symbols:
if symbol in yfinance_model.FUTURES_DATA["Ticker"].unique().tolist():
symbols_validated.append(symbol)
else:
console.print(f"[red]{symbol} is not a valid symbol[/red]")
symbols = symbols_validated
if not symbols:
console.print("No symbol was provided.\n")
return
historicals = yfinance_model.get_historical_futures(symbols, expiry)
if historicals.empty:
console.print(f"No data was found for the symbols: {', '.join(symbols)}\n")
return
if raw or len(historicals) == 1:
if not raw and len(historicals) == 1:
console.print(
"\nA single datapoint is not enough to depict a chart, data is presented below."
)
print_rich_table(
historicals[historicals.index > datetime.strptime(start_date, "%Y-%m-%d")],
headers=list(historicals.columns),
show_index=True,
title="Futures timeseries",
)
console.print()
else:
# This plot has 1 axis
if not external_axes:
_, ax = plt.subplots(figsize=plot_autoscale(), dpi=PLOT_DPI)
elif is_valid_axes_count(external_axes, 1):
(ax,) = external_axes
else:
return
colors = cycle(theme.get_colors())
if len(symbols) > 1:
name = list()
for tick in historicals["Adj Close"].columns.tolist():
if len(historicals["Adj Close"][tick].dropna()) == 1:
console.print(
f"\nA single datapoint on {tick} is not enough to depict a chart, data shown below."
)
naming = yfinance_model.FUTURES_DATA[
yfinance_model.FUTURES_DATA["Ticker"] == tick
]["Description"].values[0]
print_rich_table(
historicals[
historicals["Adj Close"][tick].index
> datetime.strptime(start_date, "%Y-%m-%d")
]["Adj Close"][tick]
.dropna()
.to_frame(),
headers=[naming],
show_index=True,
title="Futures timeseries",
)
continue
name.append(
yfinance_model.FUTURES_DATA[
yfinance_model.FUTURES_DATA["Ticker"] == tick
]["Description"].values[0]
)
ax.plot(
historicals["Adj Close"][tick].dropna().index,
historicals["Adj Close"][tick].dropna().values,
color=next(colors, "#FCED00"),
)
ax.legend(name)
first = datetime.strptime(start_date, "%Y-%m-%d")
if historicals["Adj Close"].index[0] > first:
first = historicals["Adj Close"].index[0]
ax.set_xlim(first, historicals["Adj Close"].index[-1])
theme.style_primary_axis(ax)
make_white(ax)
if external_axes is None:
theme.visualize_output()
else:
if len(historicals["Adj Close"]) == 1:
console.print(
f"\nA single datapoint on {symbols[0]} is not enough to depict a chart, data shown below."
)
print_rich_table(
historicals[
historicals["Adj Close"].index
> datetime.strptime(start_date, "%Y-%m-%d")
],
headers=list(historicals["Adj Close"].columns),
show_index=True,
title="Futures timeseries",
)
else:
name = yfinance_model.FUTURES_DATA[
yfinance_model.FUTURES_DATA["Ticker"] == symbols[0]
]["Description"].values[0]
ax.plot(
historicals["Adj Close"].dropna().index,
historicals["Adj Close"].dropna().values,
color=next(colors, "#FCED00"),
)
if expiry:
ax.set_title(f"{name} with expiry {expiry}")
else:
ax.set_title(name)
first = datetime.strptime(start_date, "%Y-%m-%d")
if historicals["Adj Close"].index[0] > first:
first = historicals["Adj Close"].index[0]
ax.set_xlim(first, historicals["Adj Close"].index[-1])
theme.style_primary_axis(ax)
make_white(ax)
if external_axes is None:
theme.visualize_output()
export_data(
export,
os.path.dirname(os.path.abspath(__file__)),
"historical",
historicals[historicals.index > datetime.strptime(start_date, "%Y-%m-%d")],
)
@log_start_end(log=logger)
def display_curve(
symbol: str,
raw: bool = False,
export: str = "",
external_axes: Optional[List[plt.Axes]] = None,
):
"""Display curve futures [Source: Yahoo Finance]
Parameters
----------
symbol: str
Curve future symbol to display
raw: bool
Display futures timeseries in raw format
export: str
Type of format to export data
external_axes : Optional[List[plt.Axes]], optional
External axes (1 axis is expected in the list), by default None
"""
if symbol not in yfinance_model.FUTURES_DATA["Ticker"].unique().tolist():
console.print(f"[red]'{symbol}' is not a valid symbol[/red]")
return
df = yfinance_model.get_curve_futures(symbol)
if df.empty:
console.print("[red]No future data found to generate curve.[/red]\n")
return
if raw:
print_rich_table(
df,
headers=list(df.columns),
show_index=True,
title="Futures curve",
)
console.print()
else:
# This plot has 1 axis
if not external_axes:
_, ax = plt.subplots(figsize=plot_autoscale(), dpi=PLOT_DPI)
elif is_valid_axes_count(external_axes, 1):
(ax,) = external_axes
else:
return
name = yfinance_model.FUTURES_DATA[
yfinance_model.FUTURES_DATA["Ticker"] == symbol
]["Description"].values[0]
colors = cycle(theme.get_colors())
ax.plot(
df.index,
df.values,
marker="o",
linestyle="dashed",
linewidth=2,
markersize=8,
color=next(colors, "#FCED00"),
)
make_white(ax)
ax.set_title(name)
theme.style_primary_axis(ax)
if external_axes is None:
theme.visualize_output()
export_data(
export,
os.path.dirname(os.path.abspath(__file__)),
"curve",
df,
) | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/futures/yfinance_view.py | 0.776665 | 0.280924 | yfinance_view.py | pypi |
__docformat__ = "numpy"
import os
import sys
import logging
from typing import List
from datetime import datetime
import yfinance as yf
import pandas as pd
from dateutil.relativedelta import relativedelta
from openbb_terminal.decorators import log_start_end
from openbb_terminal.core.config.paths import MISCELLANEOUS_DIRECTORY
# pylint: disable=attribute-defined-outside-init
logger = logging.getLogger(__name__)
FUTURES_DATA = pd.read_csv(MISCELLANEOUS_DIRECTORY / "futures" / "futures.csv")
MONTHS = {
1: "F",
2: "G",
3: "H",
4: "J",
5: "K",
6: "M",
7: "N",
8: "Q",
9: "U",
10: "V",
11: "X",
12: "Z",
}
class HiddenPrints:
def __enter__(self):
self._original_stdout = sys.stdout
sys.stdout = open(os.devnull, "w")
def __exit__(self, exc_type, exc_val, exc_tb):
sys.stdout.close()
sys.stdout = self._original_stdout
@log_start_end(log=logger)
def get_search_futures(
category: str = "",
exchange: str = "",
description: str = "",
):
"""Get search futures [Source: Yahoo Finance]
Parameters
----------
category: str
Select the category where the future exists
exchange: str
Select the exchange where the future exists
description: str
Select the description where the future exists
"""
df = FUTURES_DATA
if category:
df = df[df["Category"] == category]
if exchange:
df = df[df["Exchange"] == exchange]
if description:
df = df[
[description.lower() in desc.lower() for desc in df["Description"].values]
]
return df
@log_start_end(log=logger)
def get_historical_futures(symbols: List[str], expiry: str = "") -> pd.DataFrame:
"""Get historical futures [Source: Yahoo Finance]
Parameters
----------
symbols: List[str]
List of future timeseries symbols to display
expiry: str
Future expiry date with format YYYY-MM
Returns
-------
pd.DataFrame
Dictionary with sector weightings allocation
"""
if expiry:
symbols_with_expiry = list()
for symbol in symbols:
expiry_date = datetime.strptime(expiry, "%Y-%m")
exchange = FUTURES_DATA[FUTURES_DATA["Ticker"] == symbol][
"Exchange"
].values[0]
symbols_with_expiry.append(
f"{symbol}{MONTHS[expiry_date.month]}{str(expiry_date.year)[-2:]}.{exchange}"
)
return yf.download(symbols_with_expiry, progress=False, period="max")
df = yf.download([t + "=F" for t in symbols], progress=False, period="max")
if len(symbols) > 1:
df.columns = pd.MultiIndex.from_tuples(
[(tup[0], tup[1].replace("=F", "")) for tup in df.columns]
)
return df
@log_start_end(log=logger)
def get_curve_futures(
symbol: str = "",
) -> pd.DataFrame:
"""Get curve futures [Source: Yahoo Finance]
Parameters
----------
symbol: str
symbol to get forward curve
Returns
-------
pd.DataFrame
Dictionary with sector weightings allocation
"""
if symbol not in FUTURES_DATA["Ticker"].unique().tolist():
return pd.DataFrame()
exchange = FUTURES_DATA[FUTURES_DATA["Ticker"] == symbol]["Exchange"].values[0]
today = datetime.today()
futures_index = list()
futures_curve = list()
for i in range(36):
future = today + relativedelta(months=i)
future_symbol = (
f"{symbol}{MONTHS[future.month]}{str(future.year)[-2:]}.{exchange}"
)
with HiddenPrints():
data = yf.download(future_symbol, progress=False)
if not data.empty:
futures_index.append(future.strftime("%Y-%b"))
futures_curve.append(data["Adj Close"].values[-1])
if not futures_index:
return pd.DataFrame()
futures_index = pd.to_datetime(futures_index)
return pd.DataFrame(index=futures_index, data=futures_curve, columns=["Futures"]) | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/futures/yfinance_model.py | 0.665845 | 0.245582 | yfinance_model.py | pypi |
# Help translating terminal to other languages
```
# Install libraries if they are not installed locally
# !pip install pyyaml
# !pip install txtai[pipeline]
import os
import yaml
from txtai.pipeline import Translation
translate = Translation()
# Select language to translate
language_to_translate = "pt"
# Read default english dictionary
english_dictionary = {}
with open("en.yml", "r") as stream:
try:
english_dictionary = yaml.safe_load(stream)["en"]
except yaml.YAMLError as exc:
print(exc)
# Translated file
translated_file = f"{language_to_translate}.yml"
```
## Update other language dictionary or create from scratch
```
language_dictionary = {}
if os.path.exists(translated_file):
# Read already translated dictionary
with open(translated_file, "r") as stream:
try:
language_dictionary = yaml.safe_load(stream)[language_to_translate]
except yaml.YAMLError as exc:
print(exc)
# Iterate through english dictionary
for k in english_dictionary:
# Check if that variable doesn't existed on the already translated dictionary
if k not in language_dictionary:
# If the variable doesn't exist, let's update it to it's default
language_dictionary[k] = translate(
english_dictionary[k], language_to_translate
)
else:
# Iterate through english dictionary
for k in english_dictionary:
try:
# Replace english value by its translation
language_dictionary[k] = translate(
english_dictionary[k], language_to_translate
)
except:
# Keep english language because there was an issue with the convertion
language_dictionary[k] = english_dictionary[k]
# Save dictionary to another language
with open(f"{language_to_translate}.yml", "w") as file:
yaml.dump(
{language_to_translate: language_dictionary}, stream=file, allow_unicode=True
)
```
## Update all other languages based on english dictionary
```
for language_to_translate in [
val.split(".")[0] for val in os.listdir() if val.endswith("yml") and val != "en.yml"
]:
# Read already translated dictionary
with open(translated_file, "r") as stream:
try:
language_dictionary = yaml.safe_load(stream)[language_to_translate]
except yaml.YAMLError as exc:
print(exc)
# Iterate through english dictionary
for k in english_dictionary:
# Check if that variable doesn't existed on the already translated dictionary
if k not in language_dictionary:
# If the variable doesn't exist, let's update it to it's default
language_dictionary[k] = translate(
english_dictionary[k], language_to_translate
)
# Save dictionary to another language
with open(f"{language_to_translate}.yml", "w") as file:
yaml.dump(
{language_to_translate: language_dictionary},
stream=file,
allow_unicode=True,
)
```
| /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/miscellaneous/i18n/help_translation.ipynb | 0.438785 | 0.557725 | help_translation.ipynb | pypi |
# OpenBB Terminal : `Integration Testing`
This document is part of the `OpenBB Terminal` library documentation.
It aims to provide necessary information in order to:
- Build `integration tests`
- Run `integration tests`
- Identify errors noted from `integration tests`
## 1.1. Why have integration tests ?
The purpose of integration tests is to provide standard usage examples that can be programmatically run
to make sure that a specific functionality of the terminal and the process to utilize that functionality
is working properly.
## 2. How to build integration tests ?
Integration tests themselves are the same as manually running a certain series of steps. As a result,
writing tests is easy because the integration test script is just the total list of steps necessary to
use a command, command with specific argument, or series of commands.
When you contribute a new feature to the Terminal, it's important that integration tests are added for
this particular feature. It is a part of the Checklist for the PR to be approved.
All the `integration tests` should be insides the `scripts` folder. The naming convention for scripts
should be `test_<menu>_<command>.openbb` if you are testing a specific command or `test_<menu>.openbb`
if you are testing the entire menu. However, it is encouraged to create as specific of integration tests
as possible to identify errors more precisely. Additionally, all tests must end with the `exit` command.
These files can be given dynamic output with the following syntax `${key=default}`. Please note that
both key and default can only contain letters and numbers with NO special characters. Each dynamic
argument MUST contain a key and a default value.
### Examples
Testing a specific command and it's arguments:
```zsh
script: test_alt_covid.openbb
- - - - - - - - - -
alternative
covid
country Australia
reset
slopes
country US
ov
country Russia
deaths
country United Kingdom
cases
country Canada
rates
exit
```
Testing an entire menu
```zsh
test_stocks_options.openbb
- - - - - - - - - -
stocks
options
screen
view
view high_IV
set high_IV
scr
q
unu
calc
load aapl
exp 0
pcr
info
chains
oi
vol
voi
hist 100
grhist 100
plot -x ltd -y iv
parity
binom
load spy
vsurf
exit
```
## 3. How to run integration tests ?
### Conda Terminal
After navigating to the location of the OpenBBTerminal repo, one can run integration tests in a
few different ways using the wildcard expression. Please include a `-t` with `terminal.py` to run
the tests.
- Run all integration tests:
```zsh
python terminal.py -t
```
- Run some integration tests:
```zsh
python terminal.py -f stocks cryptocurrency -t
```
*This specific example runs all of the stocks integration tests. One can use this same format for different tests.*
- Run one integration tests:
```zsh
python terminal.py -f alternative/test_alt_covid.openbb -t
```
*Note that the base path is `OpenBBTerminal/openbb_terminal/miscellaneous/scripts`.*
- Run integration tests with arguments by adding --key=value
```zsh
python terminal.py --ticker=aapl -t
```
- To see a lot of possible keys, run the following:
```zsh
python terminal.py -h -t
```
If there are any test failures a csv will be generated with detailed information on the failures.
### Installer Terminal
Integration tests can also be used on installers, which is a packaged version of the conda terminal.
More information on how to build an installer can be found [here](/build/README.md).
- Run all integration tests:
```zsh
/Full/Path/To/OpenBB\ Terminal/.OpenBB/OpenBBTerminal /Full/Path/To/OpenBBTerminal/OpenBBTerminal/scripts/*.openbb -t
```
- Run some integration tests:
```zsh
/Full/Path/To/OpenBB\ Terminal/.OpenBB/OpenBBTerminal /Full/Path/To/OpenBBTerminal/OpenBBTerminal/scripts/test_stocks_*.openbb -t
```
- Run one integration tests:
```zsh
/Full/Path/To/OpenBB\ Terminal/.OpenBB/OpenBBTerminal /Full/Path/To/OpenBBTerminal/OpenBBTerminal/scripts/test_alt_covid.openbb -t
```
The `-t` argument runs the integration tests in 'test' mode. It is effectively a 'quiet' mode where one
doesn't see the terminal in action. If one were to remove this argument, then the terminal and subsequent
steps will be seen running in real time.
## 4. Errors
If the `-t` argument is given, errors that occur during an integration test are shown after all the
selected tests have run. The specific script that has some type of failure occurring, the reason why
it failed, and a summary of all the integration tests that were run is printed. An example is as followed:
```zsh
/scripts/test_stocks_ba.openbb: Implement enable_gui in a subclass
/scripts/test_stocks_disc.openbb: No tables found
/scripts/test_stocks_dps.openbb: No tables found
/scripts/test_stocks_scr.openbb: 429 Client Error: Too Many Requests for url: https://finviz.com/screener.ashx?v=111&s=ta_toplosers&ft=4&r=101
Summary: Successes: 55 Failures: 4
```
If the `-t` argument is not given, then the reason why a specific failure occurs within an integration
test is printed inline while the test is being run.
If there is an error, one can identify the command and or series of steps that causes it fairly easily.
Output from the integration tests can also be viewed in the `integration_test_output` folder .
| /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/miscellaneous/scripts/README.md | 0.789153 | 0.97567 | README.md | pypi |
__docformat__ = "numpy"
from datetime import datetime
from typing import Optional
import requests
import pandas as pd
from openbb_terminal.config_terminal import API_POLYGON_KEY as api_key
from openbb_terminal.helper_funcs import get_user_agent
from openbb_terminal.decorators import check_api_key
from openbb_terminal.rich_config import console
# pylint: disable=unsupported-assignment-operation
@check_api_key(["API_POLYGON_KEY"])
def get_historical(
fx_pair: str,
multiplier: int = 1,
timespan: str = "day",
start_date: str = "2000-01-01",
end_date: Optional[str] = None,
) -> pd.DataFrame:
"""Load historical fx data from polygon
Parameters
----------
fx_pair: str
Forex pair to download
multiplier: int
Multiplier for timespan. 5 with minute timespan indicated 5min windows
timespan: str
Window to aggregate data.
start_date : str
Initial date, format YYYY-MM-DD
end_date : Optional[str]
Final date, format YYYY-MM-DD
Returns
-------
pd.DataFrame
Dataframe of historical forex prices
"""
if end_date is None:
end_date = datetime.now().date().strftime("%Y-%m-%d")
request_url = (
f"https://api.polygon.io/v2/aggs/ticker/C:{fx_pair}/range"
f"/{multiplier}/{timespan}/{start_date}/{end_date}?adjusted=true&sort=desc&limit=50000&apiKey={api_key}"
)
json_response = requests.get(
request_url, headers={"User-Agent": get_user_agent()}
).json()
if json_response["status"] == "ERROR":
console.print(f"[red]{json_response['error']}[/red]\n")
return pd.DataFrame()
if "results" not in json_response.keys():
console.print("[red]Error in polygon request[/red]\n")
return pd.DataFrame()
historical = pd.DataFrame(json_response["results"]).rename(
columns={
"o": "Open",
"c": "Close",
"h": "High",
"l": "Low",
"t": "date",
"v": "Volume",
"n": "Transactions",
}
)
historical["date"] = pd.to_datetime(historical.date, unit="ms")
historical = historical.sort_values(by="date", ascending=True)
historical = historical.set_index("date")
return historical | /repl_openbb-2.1.7-py3-none-any.whl/openbb_terminal/forex/polygon_model.py | 0.859664 | 0.208743 | polygon_model.py | pypi |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.