code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
Lambda School Data Science
*Unit 2, Sprint 3, Module 3*
---
# Permutation & Boosting
You will use your portfolio project dataset for all assignments this sprint.
## Assignment
Complete these tasks for your project, and document your work.
- [ ] If you haven't completed assignment #1, please do so first.
- [ ] Continue to clean and explore your data. Make exploratory visualizations.
- [ ] Fit a model. Does it beat your baseline?
- [ ] Try xgboost.
- [ ] Get your model's permutation importances.
You should try to complete an initial model today, because the rest of the week, we're making model interpretation visualizations.
But, if you aren't ready to try xgboost and permutation importances with your dataset today, that's okay. You can practice with another dataset instead. You may choose any dataset you've worked with previously.
The data subdirectory includes the Titanic dataset for classification and the NYC apartments dataset for regression. You may want to choose one of these datasets, because example solutions will be available for each.
## Reading
Top recommendations in _**bold italic:**_
#### Permutation Importances
- _**[Kaggle / Dan Becker: Machine Learning Explainability](https://www.kaggle.com/dansbecker/permutation-importance)**_
- [Christoph Molnar: Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/feature-importance.html)
#### (Default) Feature Importances
- [Ando Saabas: Selecting good features, Part 3, Random Forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/)
- [Terence Parr, et al: Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)
#### Gradient Boosting
- [A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning](https://machinelearningmastery.com/gentle-introduction-gradient-boosting-algorithm-machine-learning/)
- [An Introduction to Statistical Learning](http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 8
- _**[Gradient Boosting Explained](https://www.gormanalysis.com/blog/gradient-boosting-explained/)**_ — Ben Gorman
- [Gradient Boosting Explained](http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html) — Alex Rogozhnikov
- [How to explain gradient boosting](https://explained.ai/gradient-boosting/) — Terence Parr & Jeremy Howard
```
!pip install category_encoders==2.*
import pandas as pd
import numpy as np
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
source_file1 = '/content/redacted_sales_data.csv'
df = pd.read_csv(source_file1)
df.tail(3)
df.head()
# the column names have a trailing space, this removes it
df = df.rename(columns={'Name ':'Name', 'Price ':'Price', 'Tax ':'Tax',
'Total Price ':'Total Price', 'Total Paid ':'Total Paid',
'Terminal ':'Terminal', 'User ':'User', 'Date ':'Date'})
### drop name column
df = df.drop(['Name'], axis=1)
## drop the last 12 rows
df.tail(13)
df = df.dropna(axis=0)
df.tail()
## drop total paid because it's redundant
## and drop terminal because it's not informative
## rename "User" column for clarity
df = df.drop(['Total Paid'], axis=1)
df = df.drop(['Terminal'], axis=1)
df = df.rename(columns={'Total Price':'Total'})
df = df.rename(columns={'User':'Employee'})
df.head()
#TIME to choose a target!!!
df['Total'].describe()
df.isna().sum()
df["Above ATP"] = df["Total"] >= df.Total.mean()
df['Date'] = pd.to_datetime(df['Date'])
df['Week'] = df['Date'].dt.week
df['Day'] = df['Date'].dt.day
df['Hour'] = df['Date'].dt.hour
df['Minute'] = df['Date'].dt.minute
df['Second'] = df['Date'].dt.second
df['Month'] = df['Date'].dt.month
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['Date']
df = df.drop(columns=unusable_variance)
df.dtypes
df.nunique().value_counts()
df['Month'].value_counts()
train = df[df['Month'] <= 3]
val = df[df['Month'] == 4]
test = df[df['Month'] >= 5]
'''
!pip install category_encoders==2.*
'''
# The status_group column is the target
target = 'Above ATP'
# Get a dataframe with all train columns except the target & Date
features = train.columns.drop([target])
print(features)
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test
y_val = test[target]
transformers = make_pipeline(
ce.ordinal.OrdinalEncoder(),
SimpleImputer()
)
X_train_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.transform(X_val_permuted)
model = RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
model.fit(X_train_transformed, y_train)
#column = train.columns.drop([target])
#X_train_t = X_train.drop(columns=column)
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
score_without = pipeline.score(X_val.drop, y_val)
print(f'Validation Accuracy without {column}: {score_without}')
# Fit with column
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
score_with = pipeline.score(X_val, y_val)
print(f'Validation Accuracy with {column}: {score_with}')
# Compare the error with & without column
print(f'Drop-Column Importance for {column}: {score_with - score_without}')
'''import eli5
from eli5.sklearn import PermutationImportance
permuter = PermutationImportance(
model,
scoring='accuracy',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed, y_val)'''
```
| github_jupyter |
# Multi-Fidelity
<div class="btn btn-notebook" role="button">
<img src="../_static/images/colab_logo_32px.png"> [Run in Google Colab](https://colab.research.google.com/drive/1Cc9TVY_Tl_boVzZDNisQnqe6Qx78svqe?usp=sharing)
</div>
<div class="btn btn-notebook" role="button">
<img src="../_static/images/github_logo_32px.png"> [View on GitHub](https://github.com/adapt-python/notebooks/blob/d0364973c642ea4880756cef4e9f2ee8bb5e8495/Multi_fidelity.ipynb)
</div>
The following example is a 1D regression multi-fidelity issue. Blue points are low fidelity observations and orange points are high fidelity observations. The goal is to use both datasets to learn the task on the [0, 1] interval.
To tackle this challenge, we use here the parameter-based method: [RegularTransferNN](#RegularTransferNN)
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from sklearn.metrics import mean_absolute_error, mean_squared_error
import tensorflow as tf
from tensorflow.keras import Model, Sequential
from tensorflow.keras.optimizers import Adam, SGD, RMSprop, Adagrad
from tensorflow.keras.layers import Dense, Input, Dropout, Conv2D, MaxPooling2D, Flatten, Reshape, GaussianNoise, BatchNormalization
from tensorflow.keras.constraints import MinMaxNorm
from tensorflow.keras.regularizers import l2
from tensorflow.keras.callbacks import Callback
from tensorflow.keras.models import clone_model
from adapt.parameter_based import RegularTransferNN
```
## Setup
```
np.random.seed(0)
Xs = np.linspace(0, 1, 200)
ys = (1 - Xs**2) * np.sin(2 * 2 * np.pi * Xs) - Xs + 0.1 * np.random.randn(len(Xs))
Xt = Xs[:100]
yt = (1 - Xt**2) * np.sin(2 * 2 * np.pi * Xt) - Xt - 1.5
gt = (1 - Xs**2) * np.sin(2 * 2 * np.pi * Xs) - Xs - 1.5
plt.figure(figsize=(10,6))
plt.plot(Xs, ys, '.', label="low fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xt, yt, '.', label="high fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xs, gt, c="black", alpha=0.7, ls="--", label="Ground truth")
plt.legend(fontsize=14)
plt.xlabel("X", fontsize=16)
plt.ylabel("y = f(X)", fontsize=16)
plt.show()
```
## Network
```
np.random.seed(0)
tf.random.set_seed(0)
model = Sequential()
model.add(Dense(100, activation='relu', input_shape=(1,)))
model.add(Dense(100, activation='relu'))
model.add(Dense(1))
model.compile(optimizer=Adam(0.001), loss='mean_squared_error')
```
## Low fidelity only
```
np.random.seed(0)
tf.random.set_seed(0)
model_low = clone_model(model)
model_low.compile(optimizer=Adam(0.001), loss='mean_squared_error')
model_low.fit(Xs, ys, epochs=800, batch_size=34, verbose=0);
yp = model_low.predict(Xs.reshape(-1,1))
score = mean_absolute_error(gt.ravel(), yp.ravel())
plt.figure(figsize=(10,6))
plt.plot(Xs, ys, '.', label="low fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xt, yt, '.', label="high fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xs, gt, c="black", alpha=0.7, ls="--", label="Ground truth")
plt.plot(Xs, yp, c="red", alpha=0.9, lw=3, label="Predictions")
plt.legend(fontsize=14)
plt.xlabel("X", fontsize=16)
plt.ylabel("y = f(X)", fontsize=16)
plt.title("Low Fidelity Only -- MAE = %.3f"%score, fontsize=18)
plt.show()
```
## High fidelity only
```
np.random.seed(0)
tf.random.set_seed(0)
model_high = clone_model(model)
model_high.compile(optimizer=Adam(0.001), loss='mean_squared_error')
model_high.fit(Xt, yt, epochs=800, batch_size=34, verbose=0);
yp = model_high.predict(Xs.reshape(-1,1))
score = mean_absolute_error(gt.ravel(), yp.ravel())
plt.figure(figsize=(10,6))
plt.plot(Xs, ys, '.', label="low fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xt, yt, '.', label="high fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xs, gt, c="black", alpha=0.7, ls="--", label="Ground truth")
plt.plot(Xs, yp, c="red", alpha=0.9, lw=3, label="Predictions")
plt.legend(fontsize=14)
plt.xlabel("X", fontsize=16)
plt.ylabel("y = f(X)", fontsize=16)
plt.title("Low Fidelity Only -- MAE = %.3f"%score, fontsize=18)
plt.show()
```
## [RegularTransferNN](https://adapt-python.github.io/adapt/generated/adapt.parameter_based.RegularTransferNN.html)
```
model_reg = RegularTransferNN(model_low, lambdas=1000., random_state=1, optimizer=Adam(0.0001))
model_reg.fit(Xt.reshape(-1,1), yt, epochs=1200, batch_size=34, verbose=0);
yp = model_reg.predict(Xs.reshape(-1,1))
score = mean_absolute_error(gt.ravel(), yp.ravel())
plt.figure(figsize=(10,6))
plt.plot(Xs, ys, '.', label="low fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xt, yt, '.', label="high fidelity", ms=15, alpha=0.9, markeredgecolor="black")
plt.plot(Xs, gt, c="black", alpha=0.7, ls="--", label="Ground truth")
plt.plot(Xs, yp, c="red", alpha=0.9, lw=3, label="Predictions")
plt.legend(fontsize=14)
plt.xlabel("X", fontsize=16)
plt.ylabel("y = f(X)", fontsize=16)
plt.title("Low Fidelity Only -- MAE = %.3f"%score, fontsize=18)
plt.show()
```
| github_jupyter |
## **Semana de Data Science**
- Minerando Dados
## Aula 01
### Conhecendo a base de dados
Monta o drive
```
from google.colab import drive
drive.mount('/content/drive')
```
Importando as bibliotecas básicas
```
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Carregando a Base de Dados
```
# carrega o dataset de london
from sklearn.datasets import load_boston
boston = load_boston()
# descrição do dataset
print (boston.DESCR)
# cria um dataframe pandas
data = pd.DataFrame(boston.data, columns=boston.feature_names)
# imprime as 5 primeiras linhas do dataset
data.head()
```
Conhecendo as colunas da base de dados
**`CRIM`**: Taxa de criminalidade per capita por cidade.
**`ZN`**: Proporção de terrenos residenciais divididos por lotes com mais de 25.000 pés quadrados.
**`INDUS`**: Essa é a proporção de hectares de negócios não comerciais por cidade.
**`CHAS`**: variável fictícia Charles River (= 1 se o trecho limita o rio; 0 caso contrário)
**`NOX`**: concentração de óxido nítrico (partes por 10 milhões)
**`RM`**: Número médio de quartos entre as casas do bairro
**`IDADE`**: proporção de unidades ocupadas pelos proprietários construídas antes de 1940
**`DIS`**: distâncias ponderadas para cinco centros de emprego em Boston
**`RAD`**: Índice de acessibilidade às rodovias radiais
**`IMPOSTO`**: taxa do imposto sobre a propriedade de valor total por US $ 10.000
**`B`**: 1000 (Bk - 0,63) ², onde Bk é a proporção de pessoas de descendência afro-americana por cidade
**`PTRATIO`**: Bairros com maior proporção de alunos para professores (maior valor de 'PTRATIO')
**`LSTAT`**: porcentagem de status mais baixo da população
**`MEDV`**: valor médio de casas ocupadas pelos proprietários em US $ 1000
Adicionando a coluna que será nossa variável alvo
```
# adiciona a variável MEDV
data['MEDV'] = boston.target
# imprime as 5 primeiras linhas do dataframe
data.head()
data.describe()
```
### Análise e Exploração dos Dados
Nesta etapa nosso objetivo é conhecer os dados que estamos trabalhando.
Podemos a ferramenta **Pandas Profiling** para essa etapa:
```
# Instalando o pandas profiling
pip install https://github.com/pandas-profiling/pandas-profiling/archive/master.zip
# import o ProfileReport
from pandas_profiling import ProfileReport
# executando o profile
profile = ProfileReport(data, title='Relatório - Pandas Profiling', html={'style':{'full_width':True}})
profile
# salvando o relatório no disco
profile.to_file(output_file="Relatorio01.html")
```
**Observações**
* *O coeficiente de correlação varia de `-1` a `1`.
Se valor é próximo de 1, isto significa que existe uma forte correlação positiva entre as variáveis. Quando esse número é próximo de -1, as variáveis tem uma forte correlação negativa.*
* *A relatório que executamos acima nos mostra que a nossa variável alvo (**MEDV**) é fortemente correlacionada com as variáveis `LSTAT` e `RM`*
* *`RAD` e `TAX` são fortemente correlacionadas, podemos remove-las do nosso modelo para evitar a multi-colinearidade.*
* *O mesmo acontece com as colunas `DIS` and `AGE` a qual tem a correlação de -0.75*
* *A coluna `ZN` possui 73% de valores zero.*
## Aula 02
Obtendo informações da base de dados manualmente
```
# Check missing values
data.isnull().sum()
# um pouco de estatística descritiva
data.describe()
```
Analisando a Correlação das colunas da base de dados
```
# Calcule a correlaçao
correlacoes = data.corr()
# Usando o método heatmap do seaborn
%matplotlib inline
plt.figure(figsize=(16, 6))
sns.heatmap(data=correlacoes, annot=True)
```
Visualizando a relação entre algumas features e variável alvo
```
# Importando o Plot.ly
import plotly.express as px
# RM vs MEDV (Número de quartos e valor médio do imóvel)
fig = px.scatter(data, x=data.RM, y=data.MEDV)
fig.show()
# LSTAT vs MEDV (índice de status mais baixo da população e preço do imóvel)
fig = px.scatter(data, x=data.LSTAT, y=data.MEDV)
fig.show()
# PTRATIO vs MEDV (percentual de proporção de alunos para professores e o valor médio de imóveis)
fig = px.scatter(data, x=data.PTRATIO, y=data.MEDV)
fig.show()
```
#### Analisando Outliers
```
# estatística descritiva da variável RM
data.RM.describe()
# visualizando a distribuição da variável RM
import plotly.figure_factory as ff
labels = ['Distribuição da variável RM (número de quartos)']
fig = ff.create_distplot([data.RM], labels, bin_size=.2)
fig.show()
# Visualizando outliers na variável RM
import plotly.express as px
fig = px.box(data, y='RM')
fig.update_layout(width=800,height=800)
fig.show()
```
Visualizando a distribuição da variável MEDV
```
# estatística descritiva da variável MEDV
data.MEDV.describe()
# visualizando a distribuição da variável MEDV
import plotly.figure_factory as ff
labels = ['Distribuição da variável MEDV (preço médio do imóvel)']
fig = ff.create_distplot([data.MEDV], labels, bin_size=.2)
fig.show()
```
Analisando a simetria do dado
```
# carrega o método stats da scipy
from scipy import stats
# imprime o coeficiente de pearson
stats.skew(data.MEDV)
```
Coeficiente de Pearson
* Valor entre -1 e 1 - distribuição simétrica.
* Valor maior que 1 - distribuição assimétrica positiva.
* Valor maior que -1 - distribuição assimétrica negativa.
```
# Histogram da variável MEDV (variável alvo)
fig = px.histogram(data, x="MEDV", nbins=50, opacity=0.50)
fig.show()
# Visualizando outliers na variável MEDV
import plotly.express as px
fig = px.box(data, y='MEDV')
fig.update_layout( width=800,height=800)
fig.show()
# imprimindo os 16 maiores valores de MEDV
data[['RM','LSTAT','PTRATIO','MEDV']].nlargest(16, 'MEDV')
# filtra os top 16 maiores registro da coluna MEDV
top16 = data.nlargest(16, 'MEDV').index
# remove os valores listados em top16
data.drop(top16, inplace=True)
# visualizando a distribuição da variável MEDV
import plotly.figure_factory as ff
labels = ['Distribuição da variável MEDV (número de quartos)']
fig = ff.create_distplot([data.MEDV], labels, bin_size=.2)
fig.show()
# Histogram da variável MEDV (variável alvo)
fig = px.histogram(data, x="MEDV", nbins=50, opacity=0.50)
fig.show()
# imprime o coeficiente de pearson
# o valor de inclinação..
stats.skew(data.MEDV)
```
**Definindo um Baseline**
- `Uma baseline é importante para ter marcos no projeto`.
- `Permite uma explicação fácil para todos os envolvidos`.
- `É algo que sempre tentaremos ganhar na medida do possível`.
```
# converte os dados
data.RM = data.RM.astype(int)
data.info()
# definindo a regra para categorizar os dados
categorias = []
# Se número de quartos for menor igual a 4 este será pequeno, senão se for menor que 7 será médio, senão será grande.
# alimenta a lista categorias
for i in data.RM.iteritems():
valor = (i[1])
if valor <= 4:
categorias.append('Pequeno')
elif valor < 7:
categorias.append('Medio')
else:
categorias.append('Grande')
# imprimindo categorias
categorias
# cria a coluna categorias no dataframe data
data['categorias'] = categorias
# imprime 5 linhas do dataframe
data.head()
# imprime a contagem de categorias
data.categorias.value_counts()
# agrupa as categorias e calcula as médias
medias_categorias = data.groupby(by='categorias')['MEDV'].mean()
# imprime a variável medias_categorias
medias_categorias
# criando o dicionario com chaves medio, grande e pequeno e seus valores
dic_baseline = {'Grande': medias_categorias[0], 'Medio': medias_categorias[1], 'Pequeno': medias_categorias[2]}
# imprime dicionario
dic_baseline
# cria a função retorna baseline
def retorna_baseline(num_quartos):
if num_quartos <= 4:
return dic_baseline.get('Pequeno')
elif num_quartos < 7:
return dic_baseline.get('Medio')
else:
return dic_baseline.get('Grande')
# chama a função retorna baseline
retorna_baseline(10)
# itera sobre os imoveis e imprime o valor médio pelo número de quartos.
for i in data.RM.iteritems():
n_quartos = i[1]
print('Número de quartos é: {} , Valor médio: {}'.format(n_quartos,retorna_baseline(n_quartos)))
# imprime as 5 primeiras linhas do dataframe
data.head()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/readikus/code-samples/blob/main/google-foo-bar/Challenge_2_1_Lovely_Lucky_LAMBs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Lovely Lucky LAMBs
## Problem Definition:
Being a henchman isn't all drudgery. Occasionally, when Commander Lambda is feeling generous, she'll hand out Lucky LAMBs (Lambda's All-purpose Money Bucks). Henchmen can use Lucky LAMBs to buy things like a second
pair of socks, a pillow for their bunks, or even a third daily meal!
However, actually passing out LAMBs isn't easy. Each henchman squad has a strict seniority ranking which must be respected - or else the henchmen will revolt and you'll all get demoted back to minions again!
There are 4 key rules which you must follow in order to avoid a revolt:
1. The most junior henchman (with the least seniority) gets exactly 1 LAMB. (There will always be at least 1 henchman on a team.)
2. A henchman will revolt if the person who ranks immediately above them gets more than double the number of LAMBs they do.
3. A henchman will revolt if the amount of LAMBs given to their next two subordinates combined is more than the number of LAMBs they get. (Note that the two most junior henchmen won't have two subordinates, so this rule doesn't apply to them. The 2nd most junior henchman would require at least as many LAMBs as the most junior henchman.)
4. You can always find more henchmen to pay - the Commander has plenty of employees. If there are enough LAMBs left over such that another henchman could be added as the most senior while obeying the other rules, you must always add and pay that henchman.
Note that you may not be able to hand out all the LAMBs. A single LAMB cannot be subdivided. That is, all henchmen must get a positive integer number of LAMBs.
Write a function called solution(total_lambs), where total_lambs is the integer number of LAMBs in the handout you are trying to divide. It should return an integer which represents the difference between the minimum and maximum number of henchmen who can share the LAMBs (that is, being as generous as possible to those you pay and as stingy as possible, respectively) while still obeying all of the above rules to avoid a revolt. For instance, if you had 10 LAMBs and were as generous as possible, you could only pay 3 henchmen (1, 2, and 4 LAMBs, in order of ascending seniority), whereas if you were as stingy as possible, you could pay 4 henchmen (1, 1, 2, and 3 LAMBs). Therefore, solution(10) should return 4-3 = 1.
To keep things interesting, Commander Lambda varies the sizes of the Lucky LAMB payouts. You can expect total_lambs to always be a positive integer less than 1 billion (10 ^ 9).
```
import math
def solution(total_lambs):
return abs(int(stingy(total_lambs) - generous(total_lambs)))
def generous(lambs):
return int(math.log(lambs + 1, 2))
fib = [1, 1]
def stingy(max):
# handle base cases
if (max == 1):
return 1
elif max == 2:
return 2
# keep track of the running total and the sequence
total = 2
i = 2
while total < max:
# go up following the fib. sequence - i.e. sum last 2 numbers
# ensure we have computed the ith fib. number
if len(fib) <= i:
fib.append(fib[i - 1] + fib[i - 2])
# will it put us above the max?
if ((total + fib[i]) > max):
return i
total += fib[i]
i += 1
return i
import unittest
class TestSolution(unittest.TestCase):
def test_solution(self):
self.assertEqual(solution(10), 1)
def test_generous(self):
self.assertEqual(generous(1), 1)
self.assertEqual(generous(2), 1)
self.assertEqual(generous(10), 3)
def test_stingy(self):
self.assertEqual(stingy(143), 10)
self.assertEqual(stingy(2), 2)
unittest.main(argv=[''], verbosity=2, exit=False)
```
**Notes on solution:** the number series for the generous algorithm follows the Fibonacci sequence, so we can make use of the various properities of that (i.e. dynamic programming/memonization).
| github_jupyter |
```
from __future__ import print_function
```
## Neural synthesis, feature visualization, and DeepDream notes
This notebook introduces what we'll call here "neural synthesis," the technique of synthesizing images using an iterative process which optimizes the pixels of the image to achieve some desired state of activations in a convolutional neural network.
The technique in its modern form dates back to around 2009 and has its origins in early attempts to visualize what features were being learned by the different layers in the network (see [Erhan et al](https://pdfs.semanticscholar.org/65d9/94fb778a8d9e0f632659fb33a082949a50d3.pdf), [Simonyan et al](https://arxiv.org/pdf/1312.6034v2.pdf), and [Mahendran & Vedaldi](https://arxiv.org/pdf/1412.0035v1.pdf)) as well as in trying to identify flaws or vulnerabilities in networks by synthesizing and feeding them adversarial examples (see [Nguyen et al](https://arxiv.org/pdf/1412.1897v4.pdf), and [Dosovitskiy & Brox](https://arxiv.org/pdf/1506.02753.pdf)). The following is an example from Simonyan et al on visualizing image classification models.
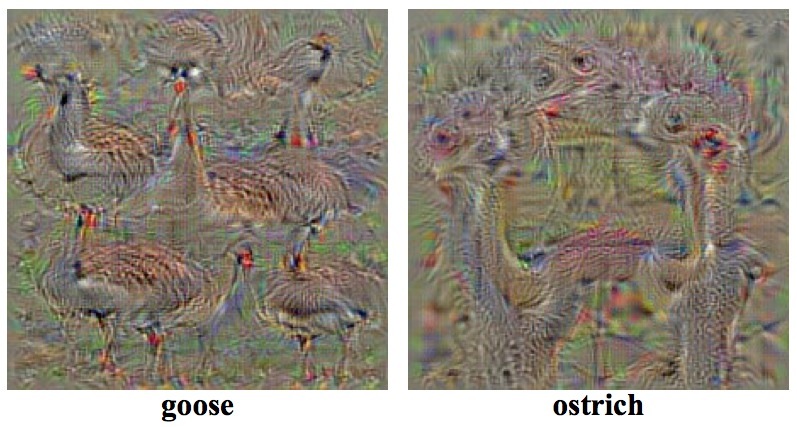

In 2012, the technique became widely known after [Le et al](https://googleblog.blogspot.in/2012/06/using-large-scale-brain-simulations-for.html) published results of an experiment in which a deep neural network was fed millions of images, predominantly from YouTube, and unexpectedly learned a cat face detector. At that time, the network was trained for three days on 16,000 CPU cores spread over 1,000 machines!

In 2015, following the rapid proliferation of cheap GPUs, Google software engineers [Mordvintsev, Olah, and Tyka](https://research.googleblog.com/2015/06/inceptionism-going-deeper-into-neural.html) first used it for ostensibly artistic purposes and introduced several innovations, including optimizing pixels over multiple scales (octaves), improved regularization, and most famously, using real images (photographs, paintings, etc) as input and optimizing their pixels so as to enhance whatever activations the network already detected (hence "hallucinating" or "dreaming"). They nicknamed their work "Deepdream" and released the first publicly available code for running it [in Caffe](https://github.com/google/deepdream/), which led to the technique being widely disseminated on social media, [puppyslugs](https://www.google.de/search?q=puppyslug&safe=off&tbm=isch&tbo=u&source=univ&sa=X&ved=0ahUKEwiT3aOwvtnXAhUHKFAKHXqdCBwQsAQIKQ&biw=960&bih=979) and all. Some highlights of their original work follow, with more found in [this gallery](https://photos.google.com/share/AF1QipPX0SCl7OzWilt9LnuQliattX4OUCj_8EP65_cTVnBmS1jnYgsGQAieQUc1VQWdgQ?key=aVBxWjhwSzg2RjJWLWRuVFBBZEN1d205bUdEMnhB).


A number of creative innovations were further introduced by [Mike Tyka](http://www.miketyka.com) including optimizing several channels along pre-arranged masks, and using feedback loops to generate video. Some examples of his work follow.

This notebook builds upon the code found in [tensorflow's deepdream example](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/examples/tutorials/deepdream). The first part of this notebook will summarize that one, including naive optimization, multiscale generation, and Laplacian normalization. The code from that notebook is lightly modified and is mostly found in the the [lapnorm.py](../notebooks/lapnorm.py) script, which is imported into this notebook. The second part of this notebook builds upon that example by showing how to combine channels and mask their gradients, warp the canvas, and generate video using a feedback loop. Here is a [gallery of examples](http://www.genekogan.com/works/neural-synth/) and a [video work](https://vimeo.com/246047871).
Before we get started, we need to make sure we have downloaded and placed the Inceptionism network in the data folder. Run the next cell if you haven't already downloaded it.
```
#Grab inception model from online and unzip it (you can skip this step if you've already downloaded the model.
!wget -P . https://storage.googleapis.com/download.tensorflow.org/models/inception5h.zip
!unzip inception5h.zip -d inception5h/
!rm inception5h.zip
```
To get started, make sure all of the folloing import statements work without error. You should get a message telling you there are 59 layers in the network and 7548 channels.
```
from io import BytesIO
import math, time, copy, json, os
import glob
from os import listdir
from os.path import isfile, join
from random import random
from io import BytesIO
from enum import Enum
from functools import partial
import PIL.Image
from IPython.display import clear_output, Image, display, HTML
import numpy as np
import scipy.misc
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
# import everything from lapnorm.py
from lapnorm import *
```
Let's inspect the network now. The following will give us the name of all the layers in the network, as well as the number of channels they contain. We can use this as a lookup table when selecting channels.
```
for l, layer in enumerate(layers):
layer = layer.split("/")[1]
num_channels = T(layer).shape[3]
print(layer, num_channels)
```
The basic idea is to take any image as input, then iteratively optimize its pixels so as to maximally activate a particular channel (feature extractor) in a trained convolutional network. We reproduce tensorflow's recipe here to read the code in detail. In `render_naive`, we take `img0` as input, then for `iter_n` steps, we calculate the gradient of the pixels with respect to our optimization objective, or in other words, the diff for all of the pixels we must add in order to make the image activate the objective. The objective we pass is a channel in one of the layers of the network, or an entire layer. Declare the function below.
```
def render_naive(t_obj, img0, iter_n=20, step=1.0):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
img = img0.copy()
for i in range(iter_n):
g, score = sess.run([t_grad, t_score], {t_input:img})
# normalizing the gradient, so the same step size should work
g /= g.std()+1e-8 # for different layers and networks
img += g*step
return img
```
Now let's try running it. First, we initialize a 200x200 block of colored noise. We then select the layer `mixed4d_5x5_bottleneck_pre_relu` and the 20th channel in that layer as the objective, and run it through `render_naive` for 40 iterations. You can try to optimize at different layers or different channels to get a feel for how it looks.
```
img0 = np.random.uniform(size=(200, 200, 3)) + 100.0
layer = 'mixed4d_3x3_bottleneck_pre_relu'
channel = 140
img1 = render_naive(T(layer)[:,:,:,channel], img0, 40, 1.0)
display_image(img1)
```
The above isn't so interesting yet. One improvement is to use repeated upsampling to effectively detect features at multiple scales (what we call "octaves") of the image. What we do is we start with a smaller image and calculate the gradients for that, going through the procedure like before. Then we upsample it by a particular ratio and calculate the gradients and modify the pixels of the result. We do this several times.
You can see that `render_multiscale` is similar to `render_naive` except now the addition of the outer "octave" loop which repeatedly upsamples the image using the `resize` function.
```
def render_multiscale(t_obj, img0, iter_n=10, step=1.0, octave_n=3, octave_scale=1.4):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
img = img0.copy()
for octave in range(octave_n):
if octave>0:
hw = np.float32(img.shape[:2])*octave_scale
img = resize(img, np.int32(hw))
for i in range(iter_n):
g = calc_grad_tiled(img, t_grad)
# normalizing the gradient, so the same step size should work
g /= g.std()+1e-8 # for different layers and networks
img += g*step
print("octave %d/%d"%(octave+1, octave_n))
clear_output()
return img
```
Let's try this on noise first. Note the new variables `octave_n` and `octave_scale` which control the parameters of the scaling. Thanks to tensorflow's patch to do the process on overlapping subrectangles, we don't have to worry about running out of memory. However, making the overall size large will mean the process takes longer to complete.
```
h, w = 200, 200
octave_n = 3
octave_scale = 1.4
iter_n = 50
img0 = np.random.uniform(size=(h, w, 3)) + 100.0
layer = 'mixed4c_5x5_bottleneck_pre_relu'
channel = 20
img1 = render_multiscale(T(layer)[:,:,:,channel], img0, iter_n, 1.0, octave_n, octave_scale)
display_image(img1)
```
Now load a real image and use that as the starting point. We'll use the kitty image in the assets folder. Here is the original.
<img src="../assets/kitty.jpg" alt="kitty" style="width: 280px;"/>
```
h, w = 320, 480
octave_n = 3
octave_scale = 1.4
iter_n = 60
img0 = load_image('../assets/kitty.jpg', h, w)
layer = 'mixed4d_5x5_bottleneck_pre_relu'
channel = 21
img1 = render_multiscale(T(layer)[:,:,:,channel], img0, iter_n, 1.0, octave_n, octave_scale)
display_image(img1)
```
Now we introduce Laplacian normalization. The problem is that although we are finding features at multiple scales, it seems to have a lot of unnatural high-frequency noise. We apply a [Laplacian pyramid decomposition](https://en.wikipedia.org/wiki/Pyramid_%28image_processing%29#Laplacian_pyramid) to the image as a regularization technique and calculate the pixel gradient at each scale, as before.
```
def render_lapnorm(t_obj, img0, iter_n=10, step=1.0, oct_n=3, oct_s=1.4, lap_n=4):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
# build the laplacian normalization graph
lap_norm_func = tffunc(np.float32)(partial(lap_normalize, scale_n=lap_n))
img = img0.copy()
for octave in range(oct_n):
if octave>0:
hw = np.float32(img.shape[:2])*oct_s
img = resize(img, np.int32(hw))
for i in range(iter_n):
g = calc_grad_tiled(img, t_grad)
g = lap_norm_func(g)
img += g*step
print('.', end='')
print("octave %d/%d"%(octave+1, oct_n))
clear_output()
return img
```
With Laplacian normalization and multiple octaves, we have the core technique finished and are level with the Tensorflow example. Try running the example below and modifying some of the numbers to see how it affects the result. Remember you can use the layer lookup table at the top of this notebook to recall the different layers that are available to you. Note the differences between early (low-level) layers and later (high-level) layers.
```
h, w = 300, 400
octave_n = 3
octave_scale = 1.4
iter_n = 20
img0 = np.random.uniform(size=(h, w, 3)) + 100.0
layer = 'mixed5b_pool_reduce_pre_relu'
channel = 99
img1 = render_lapnorm(T(layer)[:,:,:,channel], img0, iter_n, 1.0, octave_n, octave_scale)
display_image(img1)
```
Now we are going to modify the `render_lapnorm` function in three ways.
1) Instead of passing just a single channel or layer to be optimized (the objective, `t_obj`), we can pass several in an array, letting us optimize several channels simultaneously (it must be an array even if it contains just one element).
2) We now also pass in `mask`, which is a numpy array of dimensions (`h`,`w`,`n`) where `h` and `w` are the height and width of the source image `img0` and `n` is equal to the number of objectives in `t_obj`. The mask is like a gate or multiplier of the gradient for each channel. mask[:,:,0] gets multiplied by the gradient of the first objective, mask[:,:,1] by the second and so on. It should contain a float between 0 and 1 (0 to kill the gradient, 1 to let all of it pass). Another way to think of `mask` is it's like `step` for every individual pixel for each objective.
3) Internally, we use a convenience function `get_mask_sizes` which figures out for us the size of the image and mask at every octave, so we don't have to worry about calculating this ourselves, and can just pass in an img and mask of the same size.
```
def lapnorm_multi(t_obj, img0, mask, iter_n=10, step=1.0, oct_n=3, oct_s=1.4, lap_n=4, clear=True):
mask_sizes = get_mask_sizes(mask.shape[0:2], oct_n, oct_s)
img0 = resize(img0, np.int32(mask_sizes[0]))
t_score = [tf.reduce_mean(t) for t in t_obj] # defining the optimization objective
t_grad = [tf.gradients(t, t_input)[0] for t in t_score] # behold the power of automatic differentiation!
# build the laplacian normalization graph
lap_norm_func = tffunc(np.float32)(partial(lap_normalize, scale_n=lap_n))
img = img0.copy()
for octave in range(oct_n):
if octave>0:
hw = mask_sizes[octave] #np.float32(img.shape[:2])*oct_s
img = resize(img, np.int32(hw))
oct_mask = resize(mask, np.int32(mask_sizes[octave]))
for i in range(iter_n):
g_tiled = [lap_norm_func(calc_grad_tiled(img, t)) for t in t_grad]
for g, gt in enumerate(g_tiled):
img += gt * step * oct_mask[:,:,g].reshape((oct_mask.shape[0],oct_mask.shape[1],1))
print('.', end='')
print("octave %d/%d"%(octave+1, oct_n))
if clear:
clear_output()
return img
```
Try first on noise, as before. This time, we pass in two objectives from different layers and we create a mask where the top half only lets in the first channel, and the bottom half only lets in the second.
```
h, w = 300, 400
octave_n = 3
octave_scale = 1.4
iter_n = 15
img0 = np.random.uniform(size=(h, w, 3)) + 100.0
objectives = [T('mixed3a_3x3_pre_relu')[:,:,:,79],
T('mixed5a_1x1_pre_relu')[:,:,:,200],
T('mixed4b_5x5_bottleneck_pre_relu')[:,:,:,22]]
# mask
mask = np.zeros((h, w, 3))
mask[0:100,:,0] = 1.0
mask[100:200,:,1] = 1.0
mask[200:,:,2] = 1.0
img1 = lapnorm_multi(objectives, img0, mask, iter_n, 1.0, octave_n, octave_scale)
display_image(img1)
```
Now the same thing, but we optimize over the kitty instead and pick new channels.
```
h, w = 400, 400
octave_n = 3
octave_scale = 1.4
iter_n = 30
img0 = load_image('../assets/kitty.jpg', h, w)
objectives = [T('mixed4d_3x3_bottleneck_pre_relu')[:,:,:,99],
T('mixed5a_5x5_bottleneck_pre_relu')[:,:,:,40]]
# mask
mask = np.zeros((h, w, 2))
mask[:,:200,0] = 1.0
mask[:,200:,1] = 1.0
img1 = lapnorm_multi(objectives, img0, mask, iter_n, 1.0, octave_n, octave_scale)
display_image(img1)
```
Let's make a more complicated mask. Here we use numpy's `linspace` function to linearly interpolate the mask between 0 and 1, going from left to right, in the first channel's mask, and the opposite for the second channel. Thus on the far left of the image, we let in only the second channel, on the far right only the first channel, and in the middle exactly 50% of each. We'll make a long one to show the smooth transition. We'll also visualize the first channel's mask right afterwards.
```
h, w = 256, 1024
img0 = np.random.uniform(size=(h, w, 3)) + 100.0
octave_n = 3
octave_scale = 1.4
objectives = [T('mixed4c_3x3_pre_relu')[:,:,:,50],
T('mixed4d_5x5_bottleneck_pre_relu')[:,:,:,29]]
mask = np.zeros((h, w, 2))
mask[:,:,0] = np.linspace(0,1,w)
mask[:,:,1] = np.linspace(1,0,w)
img1 = lapnorm_multi(objectives, img0, mask, iter_n=40, step=1.0, oct_n=3, oct_s=1.4, lap_n=4)
print("image")
display_image(img1)
print("masks")
display_image(255*mask[:,:,0])
display_image(255*mask[:,:,1])
```
One can think up many clever ways to make masks. Maybe they are arranged as overlapping concentric circles, or along diagonal lines, or even using [Perlin noise](https://github.com/caseman/noise) to get smooth organic-looking variation.
Here is one example making a circular mask.
```
h, w = 500, 500
cy, cx = 0.5, 0.5
# circle masks
pts = np.array([[[i/(h-1.0),j/(w-1.0)] for j in range(w)] for i in range(h)])
ctr = np.array([[[cy, cx] for j in range(w)] for i in range(h)])
pts -= ctr
dist = (pts[:,:,0]**2 + pts[:,:,1]**2)**0.5
dist = dist / np.max(dist)
mask = np.ones((h, w, 2))
mask[:, :, 0] = dist
mask[:, :, 1] = 1.0-dist
img0 = np.random.uniform(size=(h, w, 3)) + 100.0
octave_n = 3
octave_scale = 1.4
objectives = [T('mixed3b_5x5_bottleneck_pre_relu')[:,:,:,9],
T('mixed4d_5x5_bottleneck_pre_relu')[:,:,:,17]]
img1 = lapnorm_multi(objectives, img0, mask, iter_n=20, step=1.0, oct_n=3, oct_s=1.4, lap_n=4)
display_image(img1)
```
Now we show how to use an existing image as a set of masks, using k-means clustering to segment it into several sections which become masks.
```
import sklearn.cluster
k = 3
h, w = 320, 480
img0 = load_image('../assets/kitty.jpg', h, w)
imgp = np.array(list(img0)).reshape((h*w, 3))
clusters, assign, _ = sklearn.cluster.k_means(imgp, k)
assign = assign.reshape((h, w))
mask = np.zeros((h, w, k))
for i in range(k):
mask[:,:,i] = np.multiply(np.ones((h, w)), (assign==i))
for i in range(k):
display_image(mask[:,:,i]*255.)
img0 = np.random.uniform(size=(h, w, 3)) + 100.0
octave_n = 3
octave_scale = 1.4
objectives = [T('mixed4b_3x3_bottleneck_pre_relu')[:,:,:,111],
T('mixed5b_pool_reduce_pre_relu')[:,:,:,12],
T('mixed4b_5x5_bottleneck_pre_relu')[:,:,:,11]]
img1 = lapnorm_multi(objectives, img0, mask, iter_n=20, step=1.0, oct_n=3, oct_s=1.4, lap_n=4)
display_image(img1)
```
Now, we move on to generating video. The most straightforward way to do this is using feedback; generate one image in the conventional way, and then use it as the input to the next generation, rather than starting with noise again. By itself, this would simply repeat or intensify the features found in the first image, but we can get interesting results by perturbing the input to the second generation slightly before passing it in. For example, we can crop it slightly to remove the outer rim, then resize it to the original size and run it through again. If we do this repeatedly, we will get what looks like a constant zooming-in motion.
The next block of code demonstrates this. We'll make a small square with a single feature, then crop the outer rim by around 5% before making the next one. We'll repeat this 20 times and look at the resulting frames. For simplicity, we'll just set the mask to 1 everywhere. Note, we've also set the `clear` variable in `lapnorm_multi` to false so we can see all the images in sequence.
```
h, w = 200, 200
# start with random noise
img = np.random.uniform(size=(h, w, 3)) + 100.0
octave_n = 3
octave_scale = 1.4
objectives = [T('mixed4d_5x5_bottleneck_pre_relu')[:,:,:,11]]
mask = np.ones((h, w, 1))
# repeat the generation loop 20 times. notice the feedback -- we make img and then use it the initial input
for f in range(20):
img = lapnorm_multi(objectives, img, mask, iter_n=20, step=1.0, oct_n=3, oct_s=1.4, lap_n=4, clear=False)
display_image(img) # let's see it
scipy.misc.imsave('frame%05d.png'%f, img) # ffmpeg to save the frames
img = resize(img[10:-10,10:-10,:], (h, w)) # before looping back, crop the border by 10 pixels, resize, repeat
```
If you look at all the frames, you can see the zoom-in effect. Zooming is just one of the things we can do to get interesting dynamics. Another cropping technique might be to shift the canvas in one direction, or maybe we can slightly rotate the canvas around a pivot point, or perhaps distort it with perlin noise. There are many things that can be done to get interesting and compelling results. Try also combining these with different ways of aking and modifying masks, and the combinatorial space of possibilities grows immensely. Most ambitiously, you can try training your own convolutional network from scratch and using it instead of Inception to get more custom effects. Thus as we see, the technique of feature visualization provides a wealth of possibilities to generate interesting video art.
| github_jupyter |
<a href="https://colab.research.google.com/github/ik-okoro/DS-Unit-2-Linear-Models/blob/master/module3-ridge-regression/LS_DS_213_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Lambda School Data Science
*Unit 2, Sprint 1, Module 3*
---
# Ridge Regression
## Assignment
We're going back to our other **New York City** real estate dataset. Instead of predicting apartment rents, you'll predict property sales prices.
But not just for condos in Tribeca...
- [ ] Use a subset of the data where `BUILDING_CLASS_CATEGORY` == `'01 ONE FAMILY DWELLINGS'` and the sale price was more than 100 thousand and less than 2 million.
- [ ] Do train/test split. Use data from January — March 2019 to train. Use data from April 2019 to test.
- [ ] Do one-hot encoding of categorical features.
- [ ] Do feature selection with `SelectKBest`.
- [ ] Fit a ridge regression model with multiple features. Use the `normalize=True` parameter (or do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html) beforehand — use the scaler's `fit_transform` method with the train set, and the scaler's `transform` method with the test set)
- [ ] Get mean absolute error for the test set.
- [ ] As always, commit your notebook to your fork of the GitHub repo.
The [NYC Department of Finance](https://www1.nyc.gov/site/finance/taxes/property-rolling-sales-data.page) has a glossary of property sales terms and NYC Building Class Code Descriptions. The data comes from the [NYC OpenData](https://data.cityofnewyork.us/browse?q=NYC%20calendar%20sales) portal.
## Stretch Goals
Don't worry, you aren't expected to do all these stretch goals! These are just ideas to consider and choose from.
- [ ] Add your own stretch goal(s) !
- [ ] Instead of `Ridge`, try `LinearRegression`. Depending on how many features you select, your errors will probably blow up! 💥
- [ ] Instead of `Ridge`, try [`RidgeCV`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeCV.html).
- [ ] Learn more about feature selection:
- ["Permutation importance"](https://www.kaggle.com/dansbecker/permutation-importance)
- [scikit-learn's User Guide for Feature Selection](https://scikit-learn.org/stable/modules/feature_selection.html)
- [mlxtend](http://rasbt.github.io/mlxtend/) library
- scikit-learn-contrib libraries: [boruta_py](https://github.com/scikit-learn-contrib/boruta_py) & [stability-selection](https://github.com/scikit-learn-contrib/stability-selection)
- [_Feature Engineering and Selection_](http://www.feat.engineering/) by Kuhn & Johnson.
- [ ] Try [statsmodels](https://www.statsmodels.org/stable/index.html) if you’re interested in more inferential statistical approach to linear regression and feature selection, looking at p values and 95% confidence intervals for the coefficients.
- [ ] Read [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapters 1-3, for more math & theory, but in an accessible, readable way.
- [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
```
%%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
!pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
import numpy as np
import pandas as pd
import pandas_profiling
# Read New York City property sales data
df = pd.read_csv(DATA_PATH+'condos/NYC_Citywide_Rolling_Calendar_Sales.csv', parse_dates=[-1], index_col=[-1])
# Change column names: replace spaces with underscores
df.columns = [col.replace(' ', '_') for col in df]
# SALE_PRICE was read as strings.
# Remove symbols, convert to integer
df['SALE_PRICE'] = (
df['SALE_PRICE']
.str.replace('$','')
.str.replace('-','')
.str.replace(',','')
.astype(int)
)
# BOROUGH is a numeric column, but arguably should be a categorical feature,
# so convert it from a number to a string
df['BOROUGH'] = df['BOROUGH'].astype(str)
# Reduce cardinality for NEIGHBORHOOD feature
# Get a list of the top 10 neighborhoods
top10 = df['NEIGHBORHOOD'].value_counts()[:10].index
# At locations where the neighborhood is NOT in the top 10,
# replace the neighborhood with 'OTHER'
df.loc[~df['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
df.head()
df.columns
df["BUILDING_CLASS_CATEGORY"].describe()
len(df)
# Subset data
df = df[(df["BUILDING_CLASS_CATEGORY"] == "01 ONE FAMILY DWELLINGS") & ((df["SALE_PRICE"] > 100000) | (df["SALE_PRICE"] < 2000000))]
len(df)
df.dtypes
df['ZIP_CODE'].value_counts()
```
Zip code should be an object but it's going to be dropped anyways so don't bother converting
```
df['EASE-MENT'].value_counts(dropna=False)
```
Drop easement as well
```
df['BLOCK'].value_counts(dropna=False)
df['LOT'].value_counts(dropna=False)
```
Don't bother changing lot and block variables before dropping
```
df.select_dtypes("object").head()
df.select_dtypes("object").nunique()
df["NEIGHBORHOOD"].value_counts()
df["APARTMENT_NUMBER"].value_counts(dropna=False)
```
Going to drop apartment number
```
df['BUILDING_CLASS_AT_PRESENT'].value_counts()
# Drop building class columns and convert land square feet
df['LAND_SQUARE_FEET'] = df['LAND_SQUARE_FEET'].str.replace(",", "").astype(int)
df = df.drop(["BUILDING_CLASS_AT_PRESENT", "BUILDING_CLASS_AT_TIME_OF_SALE"], axis=1)
df.select_dtypes("object").nunique()
# Drop address and apartments as well
df = df.drop(["ADDRESS", "APARTMENT_NUMBER"], axis=1)
df.select_dtypes("object").nunique()
```
Good with these categorical variables
```
# Drop the other wrong format columns
df = df.drop(['BLOCK', 'LOT', 'EASE-MENT', 'ZIP_CODE'], axis=1)
target = "SALE_PRICE"
y = df[target]
X = df.drop(target, axis=1)
cutoff = "2019-04-01"
mask = X.index < cutoff
X_train, y_train = X.loc[mask], y.loc[mask]
X_test, y_test = X.loc[~mask], y.loc[~mask]
assert len(X) == len(X_test) + len(X_train)
y_train.mean()
from sklearn.metrics import mean_absolute_error
mean_absolute_error(y_train, ([y_train.mean()]*len(y_train)))
len(X_train.columns)
X_train.columns
# Encoding
from category_encoders import OneHotEncoder
encode = OneHotEncoder(use_cat_names=True)
XT_train = encode.fit_transform(X_train)
len(XT_train.columns)
XT_train.columns
XT_test = encode.transform(X_test)
len(XT_test.columns)
# Not advisable but perform selectkbest iteratively simple linear regression on train data then compare with test
from sklearn.linear_model import LinearRegression
from sklearn.feature_selection import SelectKBest, f_regression
for i in range(1, len(XT_train.columns) + 1):
print(f"{i} Features")
selector = SelectKBest(score_func = f_regression, k=i)
XT_train_selected = selector.fit_transform(XT_train, y_train)
XT_test_selected = selector.transform(XT_test)
lin_reg = LinearRegression()
lin_reg.fit(XT_train_selected, y_train)
print(f"Test Mean Absolute Error: {mean_absolute_error(y_test, lin_reg.predict(XT_test_selected)).round(2)}")
```
Guess I'm using k=18?
```
selector = SelectKBest(k = 18)
XTT_train = selector.fit_transform(XT_train, y_train)
XTT_test = selector.transform(XT_test)
# Use RidgeCV to find best lambda before using Ridge
from sklearn.linear_model import Ridge, RidgeCV
alphas = [0.0001, 56, 79, 0.001, 0.01, 0.05, 0.1, 0.5, 1, 5, 10, 50, 100, 500]
ridge_cv = RidgeCV(alphas = alphas, normalize = True)
ridge_cv.fit(XTT_train, y_train)
ridge_cv.alpha_
# Use alpha = 1 then
ridge = Ridge(normalize=True)
ridge.fit(XTT_train, y_train)
print("RIDGE train MAE:", mean_absolute_error(y_train, ridge.predict(XTT_train)))
print("RIDGE test MAE:", mean_absolute_error(y_test, ridge.predict(XTT_test)))
from sklearn.metrics import r2_score
print("Training R^2:", r2_score(y_train, ridge.predict(XTT_train)))
print("Testing R^2:", r2_score(y_test, ridge.predict(XTT_test)))
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.backends.backend_pdf as pdf
import matplotlib.patches as pch
import eleanor_constants as EL
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
%matplotlib inline
### PLOT STARVED TRAJECTORIES
savename = "./figures/S0_starved.pdf"
df = pd.read_csv('./data/experiment_IDs/cleaned_static_data.csv')
df = df[df['starved'] == '1day']
df = df[df['dead'] == 'no']
color = EL.c_starve
odors = ["F", "FE", "Y",
"W", "I", "O",
"A", "G", "Q", "I2"]
odorkeys = ["Food", "Food extract", "Yeast RNA", "Water",
"Indole 100uM", "O-cresol",
"Amino acids", "Glucose",
"Quinine", "Indole 10mM"]
fig = plt.figure(figsize=(14, 12*4.125/3))
for i, (odor, odorkey) in enumerate(zip(odors, odorkeys)):
if odor != "":
col = 4
ax = fig.add_subplot(8, col, np.floor(i/col)*col+(i+1)+col, aspect="equal")
histax = fig.add_subplot(8, col, np.floor(i/col)*col+(i+1))
ax.set_xlim(0, 80-1)
ax.set_ylim(30-1, 0)
ax.set_xticks([])
ax.set_yticks([])
ax.spines['bottom'].set_color(EL.c_greyax)
ax.spines['top'].set_color(EL.c_greyax)
ax.spines['right'].set_color(EL.c_greyax)
ax.spines['left'].set_color(EL.c_greyax)
xlist = []
temp = df[df['treatment_odor'] == EL.treatments.get(odor)].copy()
temp['fname'] = './data/trajectories/video_calculations/' + temp['animal_ID'] + '-experiment.csv'
for n in temp["fname"].values:
temp2 = pd.read_csv(n)
x = temp2["pos_x_mm"].values
y = temp2["pos_y_mm"].values
xlist += x.tolist()
ax.plot(x, y, lw=0.75, color=EL.c_greyax, alpha=0.2)
ax.scatter([x[-1]], [y[-1]], color="k", alpha=0.5, lw=0, s=25, zorder=20)
histax.set_xlim(0, 80)
histax.set_ylim(0, 0.07)
if odor in ['Q', "I2"]:
histax.set_ylim(0, 0.18)
histax.text(40, 0.18, odorkey+', starved, n='+str(len(temp)),
ha='center', va='bottom', clip_on=False)
else:
histax.text(40, 0.07, odorkey+', starved, n='+str(len(temp)),
ha='center', va='bottom', clip_on=False)
histax.hist(xlist, bins=80, color=color, density=True, lw=0, clip_on=False)
histax.set_xlabel("Arena location (mm, 0-80)")
histax.set_ylabel("Probability Density (%)")
histax.spines['top'].set_visible(False)
histax.spines['right'].set_visible(False)
histax.spines['left'].set_color(EL.c_greyax)
histax.spines['bottom'].set_color(EL.c_greyax)
# SET BOUNDARIES AND SAVE FIGURE -----------------------------------------------
plt.tight_layout()
fig.subplots_adjust(wspace=0.4)
pp = pdf.PdfPages(savename, keep_empty=False)
pp.savefig(fig)
pp.close()
plt.show()
### PLOT FED TRAJECTORIES
savename = "./figures/S0_fed.pdf"
df = pd.read_csv('./data/experiment_IDs/cleaned_static_data.csv')
df = df[df['starved'] == 'no']
df = df[df['dead'] == 'no']
color = EL.c_fed
odors = ["F", "FE", "Y",
"W", "I", "O",
"A", "G", "Q"]
odorkeys = ["Food", "Food extract", "Yeast RNA", "Water",
"Indole", "O-cresol",
"Amino acids", "Glucose", "Quinine"]
fig = plt.figure(figsize=(14, 12))
for i, (odor, odorkey) in enumerate(zip(odors, odorkeys)):
ax = fig.add_subplot(6, 3, np.floor(i/3)*3+(i+1)+3, aspect="equal")
histax = fig.add_subplot(6, 3, np.floor(i/3)*3+(i+1))
ax.set_xlim(0, 80-1)
ax.set_ylim(30-1, 0)
ax.set_xticks([])
ax.set_yticks([])
ax.spines['bottom'].set_color(EL.c_greyax)
ax.spines['top'].set_color(EL.c_greyax)
ax.spines['right'].set_color(EL.c_greyax)
ax.spines['left'].set_color(EL.c_greyax)
xlist = []
temp = df[df['treatment_odor'] == EL.treatments.get(odor)].copy()
temp['fname'] = './data/trajectories/video_calculations/' + temp['animal_ID'] + '-experiment.csv'
for n in temp["fname"].values:
temp2 = pd.read_csv(n)
x = temp2["pos_x_mm"].values
y = temp2["pos_y_mm"].values
xlist += x.tolist()
ax.plot(x, y, lw=0.75, color=EL.c_greyax, alpha=0.2)
ax.scatter([x[-1]], [y[-1]], color="k", alpha=0.5, lw=0, s=25, zorder=20)
histax.set_xlim(0, 80)
histax.set_ylim(0, 0.1)
if odor == "Q":
histax.set_ylim(0, 0.14)
histax.set_yticks([0, 0.07, 0.14])
histax.text(40, 0.14, odorkey+', fed, n='+str(len(temp)),
ha='center', va='bottom', clip_on=False)
else:
histax.text(40, 0.1, odorkey+', fed, n='+str(len(temp)),
ha='center', va='bottom', clip_on=False)
histax.hist(xlist, bins=80, color=color, density=True, lw=0)
histax.set_xlabel("Arena location (mm, 0-80)")
histax.set_ylabel("Probability Density (%)")
histax.spines['top'].set_visible(False)
histax.spines['right'].set_visible(False)
histax.spines['left'].set_color(EL.c_greyax)
histax.spines['bottom'].set_color(EL.c_greyax)
# SET BOUNDARIES AND SAVE FIGURE -----------------------------------------------
plt.tight_layout()
fig.subplots_adjust(wspace=0.2)
pp = pdf.PdfPages(savename, keep_empty=False)
pp.savefig(fig)
pp.close()
plt.show()
```
| github_jupyter |
# Intro to scikit-learn, SVMs and decision trees
<hr style="clear:both">
This notebook is part of a series of exercises for the CIVIL-226 Introduction to Machine Learning for Engineers course at EPFL. Copyright (c) 2021 [VITA](https://www.epfl.ch/labs/vita/) lab at EPFL
Use of this source code is governed by an MIT-style license that can be found in the LICENSE file or at https://www.opensource.org/licenses/MIT
**Author(s):** [David Mizrahi](mailto:david.mizrahi@epfl.ch)
<hr style="clear:both">
This is the final exercise of this course. In this exercise, we'll introduce the scikit-learn package, and use it to train SVMs and decision trees. We'll end with a small note on how to use scikit-learn for unsupervised learning.
## 1. Intro to scikit-learn
[scikit-learn](https://scikit-learn.org/stable/index.html) is a very popular Python package, built on top of NumPy, which provides efficient implementations of many popular machine learning algorithms.
It can be used for:
- Generating and loading popular datasets
- Preprocessing (feature extraction and expansion, normalization)
- Supervised learning (classification and regression)
- Unsupervised learning (clustering and dimensionality reduction)
- Model selection (grid search, train/test split, cross-validation)
- Evaluation (with many metrics for all kinds of tasks)
### 1.1. Data representation in scikit-learn
In scikit-learn, data is represented in the same way it was in the previous exercises. That is:
- The features are represented as a 2D features matrix (usually named `X`), most often contained in a NumPy array or Pandas DataFrame.
- The label (or target) array is often called `y`, and is usually contained in a NumPy array or Pandas Series.
In mathematical notation, this is:
- features: $\boldsymbol{X} \in \mathbb{R}^{N \times D}$, $\forall \ \boldsymbol{x}^{(i)} \in \boldsymbol{X}: \boldsymbol{x}^{(i)} \in \mathbb{R}^{D}$
- label (or target): $\boldsymbol{y} \in \mathbb{R}^{N}$
where $N$ is the number of examples in our dataset, and $D$ is the number of features per example
scikit-learn offers many utilities for splitting and preprocessing data.
- For splitting data, there are functions such as [`model_selection.train_test_split()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html#sklearn.model_selection.train_test_split) which splits arrays or matrices into random train and test subsets, or [`model_selection.KFold()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html#sklearn.model_selection.KFold) and similar functions which provides train/test indices for cross-validation. These functions are extremely handy, and are often used to split NumPy or Pandas arrays even when the training and models come from a library other than scikit-learn.
- For preprocessing data, scikit-learn offers many utility functions which can standardize data (e.g. [`preprocessing.StandardScaler()`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler)), impute, discretize and perform feature expansion. For more information, refer to the [official preprocessing tutorial](https://scikit-learn.org/stable/modules/preprocessing.html#).
### 1.2. Estimator API
For **supervised learning**, scikit-learn implements many algorithms we've seen in this class such as:
- Nearest neighbors
- Linear regression
- Logistic regression
- Support vector machines
- Decision trees
- Ensembles (such as random forests)
In scikit-learn, these algorithms are called **estimators**, and they use a clean, uniform and streamlined API, which makes it very easy to switch to a new model or algorithm.
Here is an example of many of the estimators available with scikit-learn. [Source](https://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html)
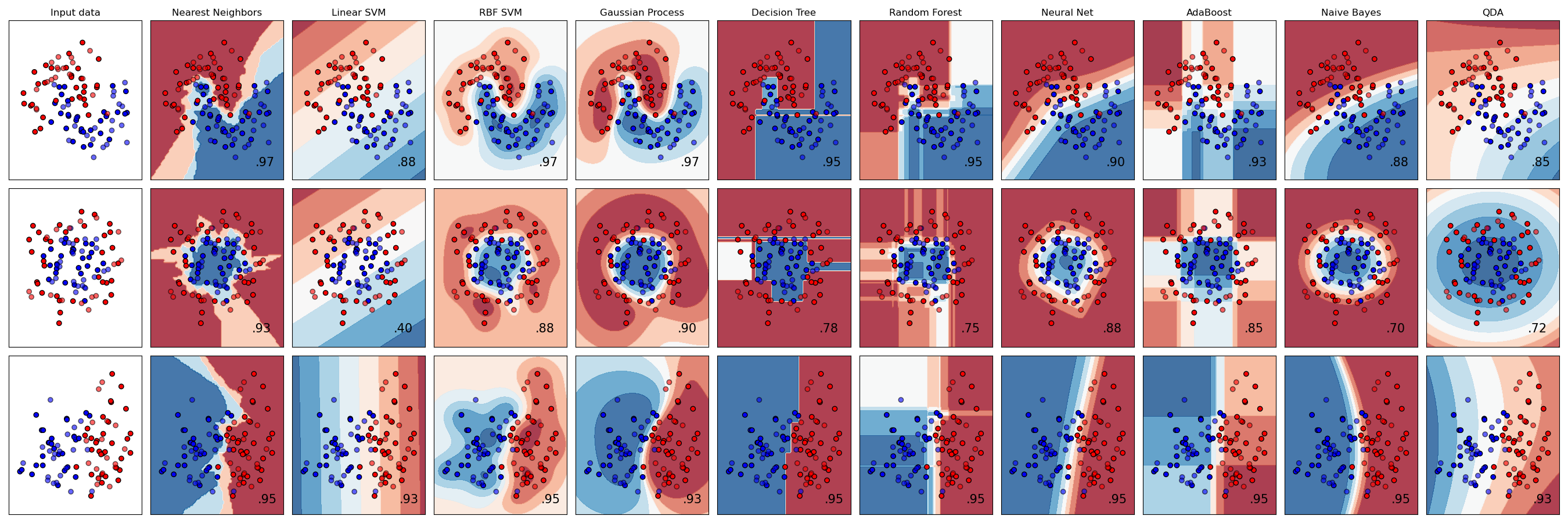
Here are the steps to follow when using the scikit-learn estimator API:
1. Arrange data into a features matrix (`X`) and target vector (`y`).
2. Choose a class of model by importing the appropriate estimator class (e.g. `linear_model.LogisticRegression()`, `svm.SVC()`, etc...)
3. Choose model hyperparameters by instantiating this class with desired values.
4. Fit the model to your data by calling the `fit()` method of the model instance.
5. Apply the model to new data: for supervised learning, we predict labels for unknown data using the `predict()` method.
The steps to follow when using scikit-learn estimators for unsupervised learning are almost identical.
### 1.3. Example: Logistic regression on the Iris dataset
As an example, we'll walk through how to use scikit-learn to train a logistic regression model for multi-class classification on the Iris dataset.
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_theme(style="white", context="notebook", palette="dark")
# !!! sklearn is how the scikit-learn package is called in Python
import sklearn
```
#### 1.3.1. Loading the dataset
```
from sklearn import datasets
# Iris is a toy dataset , which is directly available in sklearn.datasets
iris = datasets.load_iris()
X = iris.data[:, :2] # we only take the first two features for simpler visualizations
y = iris.target
print(f"Type of X: {type(X)} | Shape of X: {X.shape}")
print(f"Type of y: {type(y)} | Shape of y: {y.shape}")
```
#### 1.3.2. Splitting and scaling
```
from sklearn.model_selection import train_test_split
# Split data using train_test_split, use 30% of the data as a test set and set a random state for reproducibility
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
print(f"Shape of X_train: {X_train.shape} | Shape of y_train: {y_train.shape}")
print(f"Shape of X_test: {X_test.shape} | Shape of y_test: {y_test.shape}")
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# Fit with the mean / std of the training data
scaler.fit(X_train)
# Scale both the training / test data
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
print(f"Mean of X_train: {X_train.mean():.3f}| Std of X_train: {X_train.std():.3f}")
print(f"Mean of X_test: {X_test.mean():.3f}| Std of X_test: {X_test.std():.3f}")
```
#### 1.3.3. Training
```
from sklearn.linear_model import LogisticRegression
# Initialize a logistic regression model with L2 regularization
# and regularization strength 1e-4 (as C is inverse of regularization strength)
logreg = LogisticRegression(penalty="l2", C=1e4)
# Train the model
logreg.fit(X_train, y_train)
# Get train accuracy
train_acc = logreg.score(X_train, y_train)
print(f"Train accuracy: {train_acc * 100:.2f}%")
```
#### 1.3.4. Decision boundaries
We can use matplotlib to view the decision boundaries of our trained model.
```
# This code is beyond the scope of this class, no need to understand what it does.
# Source: https://scikit-learn.org/stable/auto_examples/linear_model/plot_iris_logistic.html
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
x_min, x_max = X_train[:, 0].min() - .5, X_train[:, 0].max() + .5
y_min, y_max = X_train[:, 1].min() - .5, X_train[:, 1].max() + .5
h = .02 # step size in the mesh
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
Z = logreg.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Paired, shading='auto', alpha=0.1, antialiased=True)
# Plot also the training points
scatter = plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train, edgecolors='k', cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.legend(handles=scatter.legend_elements()[0], labels=list(iris.target_names))
plt.show()
```
#### 1.3.5. Test accuracy
```
# Get test accuracy
test_acc = logreg.score(X_test, y_test)
print(f"Test accuracy: {test_acc * 100:.2f}%")
```
#### 1.3.6. Other metrics
```
# We can easily use other metrics using sklearn.metrics
from sklearn.metrics import balanced_accuracy_score
# First we'll use the balanced accuracy
y_pred_train = logreg.predict(X_train)
train_balanced_acc = balanced_accuracy_score(y_train, y_pred_train)
y_pred_test = logreg.predict(X_test)
test_balanced_acc = balanced_accuracy_score(y_test, y_pred_test)
print(f"Train balanced acc: {train_balanced_acc*100:.2f}%")
print(f"Test balanced acc: {test_balanced_acc*100:.2f}%")
from sklearn.metrics import plot_confusion_matrix
# Now we'll plot the confusion matrix of the testing data
plot_confusion_matrix(logreg, X_test, y_test, display_labels=iris.target_names, cmap=plt.cm.Blues)
plt.show()
```
### 1.4. Additional scikit-learn resources
This tutorial very briefly covers the scikit-learn package, and how it can be used to train a simple classifier. This package is capable of a lot more than what was shown here, as you will see in the rest of this exercise. If you want a more in-depth look at scikit-learn, take a look at these resources:
- scikit-learn Getting Started tutorial: https://scikit-learn.org/stable/getting_started.html
- scikit-learn User Guide: https://scikit-learn.org/stable/user_guide.html
- scikit-learn cheatsheet by Datacamp: https://s3.amazonaws.com/assets.datacamp.com/blog_assets/Scikit_Learn_Cheat_Sheet_Python.pdf
- scikit-learn tutorial from the Python Data Science Handbook: https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html
## 2. Support Vector Machines
In class, we have covered the theory behind SVMs, and how they can be used to perform non-linear classification using the "kernel trick". In this exercise, you'll see how SVMs can easily be trained with scikit-learn, and how the choice of kernel can impact the performance on a non-linearly separable dataset.
### 2.1. Linear SVM
First we'll show how to train a simple SVM classifier.
In scikit-learn, the corresponding estimator is called `SVC` (Support Vector Classifier).
In this part, we'll use a toy dataset which is linearly separable, generated using the `datasets.make_blobs()` function.
```
from helpers import plot_svc_decision_function
from sklearn.datasets import make_blobs
# Generate a linearly separable dataset
X, y = make_blobs(n_samples=150, centers=2, random_state=0, cluster_std=0.70)
# Split into train / test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Plot training and test data (color is for classes, shape is for train / test)
plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train, s=50, marker='o', cmap="viridis", label="train")
plt.scatter(X_test[:, 0], X_test[:, 1], c=y_test, s=50, marker='^', cmap="viridis", label="test")
plt.legend()
plt.show()
```
For this part, we'll train a SVM with a linear kernel. This corresponds to the basic SVM model that you've seen in class.
When initializing an instance of the SVC class, you can specify a regularization parameter C, and the strength of regularization is inversely proportional to C. That is, a high value of C leads to low regularization and a low C leads to high regularization.
Try changing the value of C. How does it affect the support vectors?
**Answer:**
YOUR ANSWER HERE
```
from sklearn.svm import SVC # SVC = Support vector classifier
# C is the regularization parameter. The strength of regularization is inversely proportional to C.
# Try very large and very small values of C
model = SVC(kernel='linear', C=1)
model.fit(X_train, y_train)
# Print training accuracy
train_acc = model.score(X_train, y_train)
print(f"Train accuracy: {train_acc * 100:.2f}%")
# Show decision function and support vectors
plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train, s=50, cmap="viridis")
plt.title(f"Kernel = {model.kernel} | C = {model.C}")
plot_svc_decision_function(model, plot_support=True)
# Print test accuracy
test_acc = model.score(X_test, y_test)
print(f"Test accuracy: {test_acc * 100:.2f}%")
```
### 2.2. Kernel SVM
Let's now use a non-linearly separable dataset, to observe the effect of the kernel function in SVMs.
```
from sklearn.datasets import make_circles
# Generate a circular dataset
X, y = make_circles(n_samples=400, noise=0.25, factor=0, random_state=0)
# Split into train / test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# Plot training and test data (color is for classes, shape is for train / test)
plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train, marker='o', cmap="viridis", label="train")
plt.scatter(X_test[:, 0], X_test[:, 1], c=y_test, marker='^', cmap="viridis", label="test")
plt.legend()
plt.show()
```
As you've seen in class, we can use kernel functions to allow SVMs to operate in high-dimensional, implicit feature spaces, without needing to compute the coordinates of the data in that space. We have seen a variety of kernel functions, such as the polynomial kernel and the RBF kernel.
In this exercise, experiment with the different kernels, such as:
- the linear kernel (`linear`): $\langle x, x'\rangle$
- the polynomial kernel (`poly`): $(\gamma \langle x, x'\rangle + r)^d$ (try out different degrees)
- the radial basis function kernel (`rbf`): $\exp(-\gamma \|x-x'\|^2)$
Your task is to experiment with these kernels to see which one does the best on this dataset.
How does the kernel affect the decision boundary? Which kernel and value of C would you pick to maximize your model's performance?
**Note:** Use the the helper function `plot_svc_decision_function()` to view the decision boundaries for each model.
```
# Use as many code cells as needed to try out different kernels and values of C
### YOUR CODE HERE ###
```
**Answer:**
YOUR ANSWER HERE
**To go further**: To learn more about SVMs in scikit-learn, and how to use them for multi-class classification and regression, check out the documentation page: https://scikit-learn.org/stable/modules/svm.html
## 3. Trees
Decision trees are a very intuitive way to classify objects: they ask a series of questions to infer the target variable.
A decision tree is a set of nested decision rules. At each node $i$, the $d_i$-th feature of the input vector $ \boldsymbol{x}$ is compared to a treshold value $t$. The vector $\boldsymbol{x}$ is passed down to the left or right branch depending on whether $d_i$ is less than or greater than $t$. This process is repeated for each node encountered until a reaching leaf node, which specifies the predicted output.
<img src="images/simple_tree.png" width=400></img>
*Example of a simple decision tree on the Palmer Penguins dataset*
Decision trees are usually constructed from the top-down, by choosing a feature at each step that best splits the set of items. There are different metrics for measuring the "best" feature to pick, such as the Gini impurity and the entropy / information gain. We won't dive into them here, but we recommend reading Chapter 18 of ["Probabilistic Machine Learning: An Introduction"](https://probml.github.io/pml-book/) by K.P. Murphy if you want to learn more about them.
Decision trees are popular for several reasons:
- They are **easy to interpret**.
- They can handle mixed discrete and continuous inputs.
- They are insensitive to monotone transformations of the inputs, so there is no need to standardize the data.
- They perform automatic feature selection.
- They are fast to fit, and scale well to large data sets.
Unfortunately, trees usually do not predict as accurately as other models we have seen previously, such as neural networks and SVMs.
It is however possible to significantly improve their performance through an ensemble learning method called **random forests**, which consists of constructing a multitude of decision trees at training time and averaging their outputs at test time. While random forests usually perform better than a single decision tree, they are much less interpretable. We won't cover random forests in this exercise, but keep in mind that they can be easily implemented in scikit-learn using the [`ensemble` module](https://scikit-learn.org/stable/modules/ensemble.html).
### 3.1. Training decision trees
In this part, we will work on the Titanic dataset obtained at the end of the `05-pandas` tutorial. Our goal is to train a model to predict whether or not a passenger survived the shipwreck and to find out which features are the most useful for predicting this.
```
import pandas as pd
titanic = pd.read_csv("data/titanic.csv")
titanic.head(5)
# Split into X and y
X = titanic.drop(columns="survived")
y = titanic["survived"]
# Convert to NumPy (needed for interpretability function later on)
X_numpy, y_numpy = X.to_numpy(), y.to_numpy()
# Use 80% of data for train/val, 20% for test
X_trainval, X_test, y_trainval, y_test = train_test_split(X_numpy, y_numpy, test_size=0.2, random_state=42)
# Use 80% of trainval for train, 20% for val
X_train, X_val, y_train, y_val = train_test_split(X_trainval, y_trainval, test_size=0.2, random_state=42)
```
It is now your turn to train decision trees in scikit-learn. They follow the same estimator API as all other supervised learning models, so the implementation is very straightforward. For more information, check out the [`tree` module](https://scikit-learn.org/stable/modules/tree.html#tree).
**Your task:** Initialize a `DecisionTreeClassifier` and train it on `X_train` and `y_train`.
- Use "entropy" as the `criterion`
- Try out different values for the max tree depth. How does it affect the train and validation accuracy?
```
import sklearn.tree as tree
from sklearn.tree import DecisionTreeClassifier
# Use the entropy (information gain) as the criterion
# Try varying the max depth
### YOUR CODE HERE ###
model = ...
### END CODE HERE ###
train_acc = model.score(X_train, y_train)
print(f"Train accuracy: {train_acc * 100:.2f}%")
val_acc = model.score(X_val, y_val)
print(f"Validation accuracy: {val_acc * 100:.2f}%")
test_acc = model.score(X_test, y_test)
print(f"Test accuracy: {test_acc * 100:.2f}%")
```
**Answer:** YOUR ANSWER HERE
### 3.2. Interpretability of trees
In this section, we'll show you how to visualize decision trees and interpret the decision made for some examples of our test set.
**Your task:** Run the next few cells to better understand the structure of the tree you just built. Can you identify which features are the most important for predicting whether or not a passenger survived?
```
# Plots the decision tree
# Try out a max plot depth of 2 or 3, tree will be hard to read otherwise
plt.figure(figsize=(30, 10))
tree.plot_tree(model, max_depth=2, filled=True, feature_names=X.columns, class_names=["Perished", "Survived"],
impurity=False, proportion=True, rounded=True)
plt.show()
# Prints the decision tree as text
# Will be very long if max depth is high
# Class 0 = Perished, Class 1 = Survived
print(tree.export_text(model, feature_names=list(X.columns)))
# Explaining the decisions (complicated code, no need to understand what it does exactly)
def explain_decision(sample_id: int = 0):
"""Prints rules followed to obtain prediction for a sample of the test set
Code adapted from:
https://scikit-learn.org/stable/auto_examples/tree/plot_unveil_tree_structure.html
"""
sample_id = sample_id
class_names=["Perished", "Survived"]
n_nodes = model.tree_.node_count
children_left = model.tree_.children_left
children_right = model.tree_.children_right
feature = model.tree_.feature
threshold = model.tree_.threshold
node_indicator = model.decision_path(X_test)
leaf_id = model.apply(X_test)
# obtain ids of the nodes `sample_id` goes through, i.e., row `sample_id`
node_index = node_indicator.indices[node_indicator.indptr[sample_id]:
node_indicator.indptr[sample_id + 1]]
prediction = class_names[model.predict(X_test[sample_id:sample_id+1])[0]]
print(f"Prediction for sample {sample_id}: {prediction}\n")
print("Rules used:")
for node_id in node_index:
# continue to the next node if it is a leaf node
if leaf_id[sample_id] == node_id:
continue
# check if value of the split feature for sample 0 is below threshold
if (X_test[sample_id, feature[node_id]] <= threshold[node_id]):
threshold_sign = "<="
else:
threshold_sign = ">"
print("- node {node}: ({feature} = {value}) "
"{inequality} {threshold}".format(
node=node_id,
feature=X.columns[feature[node_id]],
value=X_test[sample_id, feature[node_id]],
inequality=threshold_sign,
threshold=threshold[node_id]))
# For binary variables, 1 = True, 0 = False
# e.g. sex_male = 1 -> male, sex_male = 0 -> female
# Many of the features are redundant (e.g. sex_male and sex_female)
# so the tree doesn't always choose the same features
explain_decision(sample_id=0)
explain_decision(sample_id=1)
```
**Answer:**
YOUR ANSWER HERE
**To go further:** Decision trees and random forests can also be used for regression, check out the scikit-learn pages on [trees](https://scikit-learn.org/stable/modules/tree.html#tree) and [ensemble methods](https://scikit-learn.org/stable/modules/ensemble.html#ensemble) for more info.
## 4. A small note on unsupervised learning
While we won't cover them in this exercise, most of the unsupervised learning techniques seen in class can be easily implemented with scikit-learn.
As an example, here is how to use the k-means clustering algorithm on a toy dataset consisting of 7 unlabeled blobs of points.
When choosing $k=7$, k-means manages to almost perfectly recover the original blobs.
```
from sklearn.cluster import KMeans
# Generate unlabeled data
X_blobs, _ = make_blobs(n_samples=200, centers=7, random_state=0, cluster_std=0.60)
plt.scatter(X_blobs[:, 0], X_blobs[:, 1], c="grey", alpha=0.5)
plt.title("Unlabeled data")
plt.show()
# Run k-means on data to find the blobs
# Try changing the value of k
k = 7
kmeans = KMeans(n_clusters=k)
labels = kmeans.fit_predict(X_blobs)
# Display clusters and their centers
plt.scatter(X_blobs[:,0], X_blobs[:,1], c=labels, cmap="viridis", alpha=0.5)
for c in kmeans.cluster_centers_:
plt.scatter(c[0], c[1], marker="*", s=80, color="blue")
plt.title(f"K-Means with {k} clusters")
plt.show()
```
To learn more about how to practically implement these techniques, check out these resources:
**For dimensionality reduction:**
- [PCA from the Python Data Science handbook](https://jakevdp.github.io/PythonDataScienceHandbook/05.09-principal-component-analysis.html)
- [Manifold learning from the Python Data Science handbook](https://jakevdp.github.io/PythonDataScienceHandbook/05.10-manifold-learning.html)
- [Decomposition page on scikit-learn's website](https://scikit-learn.org/stable/modules/decomposition.html)
- [Manifold learning page on scikit-learn's website](https://scikit-learn.org/stable/modules/manifold.html)
**For clustering:**
- [k-means from the Python Data Science handbook](https://jakevdp.github.io/PythonDataScienceHandbook/05.11-k-means.html)
- [Gaussian mixtures from the Python Data Science handbook](https://jakevdp.github.io/PythonDataScienceHandbook/05.12-gaussian-mixtures.html)
- [Clustering page on scikit-learn's website](https://scikit-learn.org/stable/modules/clustering.html)
## Congratulations!
Congratulations on completing this final exercise!
Throughout this series of exercises, you learned about the fundamental tools and libraries used in machine learning, and worked on practical implementations of many of the most commonly used techniques in this field.
As long as these exercises have been, they are still too short to cover several other interesting and important machine learning topics, but we believe you now have all the tools at your disposal to learn about them on your own, if you desire to do so.
Thank you for sticking with us through the end, we really hope you enjoyed the exercises in this course!
<img src="images/thats_all_folks.png" width=400></img>
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import pickle
%matplotlib inline
thirteen_genre_df = pd.read_pickle('/content/13_genre.pkl')
thirteen_genre_df.head()
thirteen_genre_df = thirteen_genre_df.set_index('Genre_first')
thirteen_genre_df
thirteen_genre_list = sorted(list(set([x for x in thirteen_genre_df.index.to_numpy()])))
thirteen_genre_list
```
## Collecting all the description of specific genre
```
thr_g = thirteen_genre_df.groupby('Genre_first')['Description']
Action_gf = thr_g.get_group('Action')
Action_gf = Action_gf.str.cat(sep='.')
Action_gf
# thr_g = thirteen_genre_df.groupby('Genre_first')['Description']
Adventure_gf = thr_g.get_group('Adventure')
Adventure_gf = Adventure_gf.str.cat(sep='.')
Adventure_gf
Animation_gf = thr_g.get_group('Animation')
Animation_gf = Animation_gf.str.cat(sep='.')
Animation_gf
Biography_gf = thr_g.get_group('Biography')
Biography_gf = Biography_gf.str.cat(sep='.')
Biography_gf
Comedy_gf = thr_g.get_group('Comedy')
Comedy_gf = Comedy_gf.str.cat(sep='.')
Comedy_gf
Crime_gf = thr_g.get_group('Crime')
Crime_gf = Crime_gf.str.cat(sep='.')
Crime_gf
Drama_gf = thr_g.get_group('Drama')
Drama_gf = Drama_gf.str.cat(sep='.')
Drama_gf
Fantasy_gf = thr_g.get_group('Fantasy')
Fantasy_gf = Fantasy_gf.str.cat(sep='.')
Fantasy_gf
Horror_gf = thr_g.get_group('Horror')
Horror_gf = Horror_gf.str.cat(sep='.')
Horror_gf
Mystery_gf = thr_g.get_group('Mystery')
Mystery_gf = Mystery_gf.str.cat(sep='.')
Mystery_gf
Romance_gf = thr_g.get_group('Romance')
Romance_gf = Romance_gf.str.cat(sep='.')
Romance_gf
Sci_Fi_gf = thr_g.get_group('Sci-Fi')
Sci_Fi_gf = Sci_Fi_gf.str.cat(sep='.')
Sci_Fi_gf
Thriller_gf = thr_g.get_group('Thriller')
Thriller_gf = Thriller_gf.str.cat(sep='.')
Thriller_gf
thirteen_genre_list
```
## Creating dictionary out of the series generated
```
thirteen_genre_dict = dict(zip(thirteen_genre_list,[Action_gf,Adventure_gf,Animation_gf,Biography_gf,
Comedy_gf,Crime_gf,Drama_gf,Fantasy_gf,Horror_gf,
Mystery_gf,Romance_gf,Sci_Fi_gf,Thriller_gf]))
(thirteen_genre_dict)
```
## Generating dataframe out of the dictionary
```
only_thirteen_genre = pd.DataFrame.from_dict(thirteen_genre_dict,orient='index',columns=['Description'])
only_thirteen_genre.head()
```
## Finding top common words from all genres.
```
from collections import Counter
# genre_list
# common_words_list = []
for i in only_thirteen_genre['Description']:
common_words_list = [word for word, count in Counter(i.split()).most_common() if count >1]
common_words_list
from textblob import TextBlob
only_thirteen_genre['polarity'] = only_thirteen_genre['Description'].apply(lambda x : TextBlob(x).sentiment.polarity)
only_thirteen_genre['subjectivity'] = only_thirteen_genre['Description'].apply(lambda x : TextBlob(x).sentiment.subjectivity)
only_thirteen_genre
plt.figure(figsize=(12,9))
for indexes , genre in enumerate(thirteen_genre_list):
x = only_thirteen_genre.polarity[indexes]
y = only_thirteen_genre.subjectivity[indexes]
plt.scatter(x,y,color='blue')
plt.text(x+0.001,y+0.001,genre)
plt.title("Sentiment Anlaysis")
plt.xlabel("<-- Negative -------- Positive -->")
plt.ylabel('<-- Facts -------- Opinions -->')
from wordcloud import WordCloud
from sklearn.feature_extraction import text
stop_words = text.ENGLISH_STOP_WORDS.union(common_words_list)
wc = WordCloud(stopwords=stop_words, background_color='white',random_state=42,max_font_size=150,colormap='Dark2')
plt.figure(figsize=(12,9))
for indexes , genre in enumerate(only_thirteen_genre['Description']):
wc.generate(genre)
plt.subplot(5,3,indexes+1)
plt.imshow(wc,interpolation='bilinear')
plt.axis("off")
plt.title(thirteen_genre_list[indexes])
```
| github_jupyter |
# Generating thousands of images
These landscapes will be used to train generative ML models. Therefore, we need a few thousand images for training and testing. This notebook will have similar commands as the `Example` notebook, but will make many images with the same format and save them all.
I am going to update this to randomize a few things:
1. Randomize where the horizon appears. As it stands, every image had $R(\theta)=[-15,30]$, which means the models can learn the horizon boundary too easily.
2. Randomize the number of trees. Previously all landscapes had 25 trees unless they randomly appeared directly in front of one another. Now I'll draw from a Poisson distribution to figure out how many trees.
3. Randomize the "scene" between a sunset with darkening skies and a bright fall day where the trees are more reddish.
```
import numpy as np
import numpy.random as npr
import matplotlib.pyplot as plt
%matplotlib inline
import LandscapeGenerator as LG
np.random.seed(1234)
#Create some objects that will be reused to avoid calling constructors many times
field_of_view = np.zeros((2, 2))
field_of_view[1, 0:2] = 0, 360
M, N = 128, 128*8
dimensions = [M, N]
L = LG.LGenerator(dimensions)
def get_random_FOV():
dtheta = 0*npr.randn()
field_of_view[0, 0:2] = -15 + dtheta, 30 + dtheta
return field_of_view
def make_tree(leaf_rgb_means = [80, 160, 110],
trunk_rgb_means = [105, 75, 50]):
return LG.TreeFeature(height = 6 + 2*npr.rand(),
branch_radius = 1.5 + npr.rand(),
distance = 6 + 40*npr.rand(),
phi = 360 * npr.rand(),
leaf_rgb_means = leaf_rgb_means + 20*npr.randn(3),
trunk_rgb_means = trunk_rgb_means + 5*npr.randn(3))
def make_sunset_landscape():
L.clear_features()
L.set_field_of_view(get_random_FOV())
L.add_feature(LG.SkyFeature(rgb_means = [20, 100, 200], rgb_SDs = [5, 5, 5]))
L.add_feature(LG.SkyFeature(theta_boundary = 10, rgb_means = [20, 80, 160], rgb_SDs = [5, 5, 5]))
L.add_feature(LG.SkyFeature(theta_boundary = 20, rgb_means = [5, 40, 120], rgb_SDs = [5, 5, 5]))
L.add_feature(LG.GrassFeature(theta_boundary = 0))
L.add_feature(LG.GrassFeature(theta_boundary = -5, rgb_means=[50, 200, 70]))
L.add_feature(LG.SunFeature(theta = 10*npr.rand(), phi = 360 * npr.rand()))
L.add_feature(LG.SkyGradientFeature(rgb_peaks = [150, 0, 50]))
Ntrees = npr.poisson(25)
trees = [make_tree() for _ in range(Ntrees)]
trees.sort(key = lambda x: x.distance, reverse = True)
for t in trees:
L.add_feature(t)
return L.generate()
def make_fall_landscape():
L.clear_features()
L.set_field_of_view(get_random_FOV())
L.add_feature(LG.SkyFeature(rgb_means = [60, 210, 240], rgb_SDs = [5, 5, 5]))
L.add_feature(LG.GrassFeature(theta_boundary = 0, rgb_means=[50, 230, 70]))
L.add_feature(LG.GrassFeature(theta_boundary = -5, rgb_means=[50, 230, 120]))
L.add_feature(LG.SunFeature(theta = L.field_of_view[0, 1] * npr.rand(), phi = 360 * npr.rand()))
#L.add_feature(LG.SkyGradientFeature(rgb_peaks = [150, 0, 50]))
Ntrees = npr.poisson(25)
trees = [make_tree(leaf_rgb_means = [190, 70, 45], trunk_rgb_means = [105, 80, 65]) for _ in range(Ntrees)]
trees.sort(key = lambda x: x.distance, reverse = True)
for t in trees:
L.add_feature(t)
return L.generate()
N_pics = 10
data = np.zeros((N_pics, M, N, 3))
for i in range(N_pics):
if npr.randint(2):
data[i] = make_sunset_landscape()
else:
data[i] = make_fall_landscape()
#np.save(f"panoramas_{M}x{N}_Npics{N_pics}_rgb", data)
#Visualize
nr = 10
fig, ax = plt.subplots(nr, 1, figsize=(20, 5))
plt.subplots_adjust(hspace=0)
#ax[2].set_xticks([])
#ax[2].set_xticklabels([])
for i in range(nr):
ax[i].axis("off")
#ax[i].set_yticks([])
#ax[i].set_yticklabels([])
ax[i].imshow(data[np.random.randint(low = 0, high = N_pics)])
fig, ax = plt.subplots(1, 1, figsize=(10,10))
ax.imshow(data[3])
ax.axis("off")
fig.savefig("example2.png", dpi=400, bbox_inches="tight", transparent=True)
```
| github_jupyter |
# NIRCam F444W Coronagraphic Observations of Vega
---
Here we create the basics for a MIRI simulation to observe the Fomalhaut system with the FQPM 1550. This includes simulating the Fomalhaut stellar source behind the center of the phase mask, some fake off-axis companions, and a debris disk model that crosses the mask's quadrant boundaries.
From JWST PID 1193, the pointing file provides some the relevant information. Here's a truncated version of the pointing file for the first roll position. In this example, we skip the target acquisition observations and only simulate the science exposure image.
```
* Fomalhaut-1550C-Rot1 (Obs 6)
** Visit 6:1
Aperture Name Target RA Dec V2 V3 IdlX IdlY Level Type
MIRIM_TABLOCK 1 VEGA +279.23474 +38.78369 -407.464 -387.100 +0.000 +0.000 TARGET T_ACQ
MIRIM_TA1550_UR 1 VEGA +279.23474 +38.78369 -395.471 -365.842 +0.000 +0.000 TARGET T_ACQ
MIRIM_TA1550_CUR 1 VEGA +279.23474 +38.78369 -391.430 -370.519 +0.122 +0.134 TARGET T_ACQ
MIRIM_MASK1550 1 VEGA +279.23474 +38.78369 -389.892 -372.181 +0.000 +0.000 TARGET SCIENCE
```
Final outputs will be detector-sampled slope images (counts/sec).
```
# Import the usual libraries
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
#import matplotlib.patches as mpatches
# Enable inline plotting
%matplotlib inline
# Progress bar
from tqdm.auto import trange, tqdm
import webbpsf_ext, pysiaf
from astropy.io import fits
from webbpsf_ext import image_manip, setup_logging, spectra, coords
```
# NIRCam Observation with MASK430R+F444W
## 1. Create PSF structure
```
# Mask information
filt = 'F444W'
mask = 'MASK430R'
pupil = 'CIRCLYOT'
# Initiate instrument class with selected filters, pupil mask, and image mask
inst = webbpsf_ext.NIRCam_ext(filter=filt, pupil_mask=pupil, image_mask=mask)
# Set desired PSF size and oversampling
inst.fov_pix = 256
inst.oversample = 2
# Calculate PSF coefficients
inst.npsf = 10
inst.gen_psf_coeff()
# Calculate position-dependent PSFs due to FQPM
# Equivalent to generating a giant library to interpolate over
inst.gen_wfemask_coeff()
```
## 1.1 Observation setup
__Configuring observation settings__
Observations consist of nested visit, mosaic tiles, exposures, and dithers. In this section, we configure a pointing class that houses information for a single observation defined in the APT .pointing file. The primary information includes a pointing reference SIAF aperturne name, RA and Dec of the ref aperture, Base X/Y offset relative to the ref aperture position, and Dith X/Y offsets. From this information, along with the V2/V3 position angle, we can determine the orientation and location of objects on the detector focal plane.
**Note**: The reference aperture is not necessarily the same as the observed aperture. For instance, you may observe simultaneously with four of NIRCam's SWA detectors, so the reference aperture would be the entire SWA channel, while the observed apertures are A1, A2, A3, and A4.
```
# Import class to setup pointing info
from webbpsf_ext.coords import jwst_point
# Observed and reference apertures
ap_obs = 'NRCA5_MASK430R'
ap_ref = 'NRCA5_MASK430R'
# Define the RA/Dec of reference aperture and telescope position angle
# Position angle is angle of V3 axis rotated towards East
ra_ref, dec_ref = (+279.23474, +38.78369)
pos_ang = 0
# Set any baseline pointing offsets (e.g., specified in APT's Special Requirements)
base_offset=(0,0)
# Define a list of nominal dither offsets
dith_offsets = [(0,0)]
# Telescope pointing information
tel_point = jwst_point(ap_obs, ap_ref, ra_ref, dec_ref, pos_ang=pos_ang,
base_offset=base_offset, dith_offsets=dith_offsets,
base_std=0, dith_std=0)
print(f"Reference aperture: {tel_point.siaf_ap_ref.AperName}")
print(f" Nominal RA, Dec = ({tel_point.ra_ref:.6f}, {tel_point.dec_ref:.6f})")
print(f"Observed aperture: {tel_point.siaf_ap_obs.AperName}")
print(f" Nominal RA, Dec = ({tel_point.ra_obs:.6f}, {tel_point.dec_obs:.6f})")
print(f"Relative offsets in 'idl' for each dither position (incl. pointing errors)")
for i, offset in enumerate(tel_point.position_offsets_act):
print(f" Position {i}: ({offset[0]:.4f}, {offset[1]:.4f}) arcsec")
```
## 1.2 Add central source
Here we define the stellar atmosphere parameters for Fomalhaut, including spectral type, optional values for (Teff, log_g, metallicity), normalization flux and bandpass, as well as RA and Dec.
```
from webbpsf_ext import miri_filter, nircam_filter, bp_2mass
from webbpsf_ext.image_manip import pad_or_cut_to_size
def make_spec(name=None, sptype=None, flux=None, flux_units=None, bp_ref=None, **kwargs):
"""
Create pysynphot stellar spectrum from input dictionary properties.
"""
from webbpsf_ext import stellar_spectrum
# Renormalization arguments
renorm_args = (flux, flux_units, bp_ref)
# Create spectrum
sp = stellar_spectrum(sptype, *renorm_args, **kwargs)
if name is not None:
sp.name = name
return sp
# Information necessary to create pysynphot spectrum of star
obj_params = {
'name': 'Vega',
'sptype': 'A0V',
'Teff': 9602, 'log_g': 4.1, 'metallicity': -0.5,
'dist': 7.7,
'flux': 16.09, 'flux_units': 'Jy', 'bp_ref': miri_filter('F1550C'),
'RA_obj' : +279.23474, # RA (decimal deg) of source
'Dec_obj' : +38.78369, # Dec (decimal deg) of source
}
# Create stellar spectrum and add to dictionary
sp_star = make_spec(**obj_params)
obj_params['sp'] = sp_star
# Get `sci` coord positions
coord_obj = (obj_params['RA_obj'], obj_params['Dec_obj'])
xsci, ysci = tel_point.radec_to_frame(coord_obj, frame_out='sci')
# Get sci position shifts from center in units of detector pixels
siaf_ap = tel_point.siaf_ap_obs
xsci_cen, ysci_cen = siaf_ap.reference_point('sci')
xsci_off, ysci_off = (xsci-xsci_cen, ysci-ysci_cen)
# and now oversampled pixel shifts
osamp = inst.oversample
xsci_off_over = xsci_off * osamp
ysci_off_over = ysci_off * osamp
print("Image shifts (oversampled pixels):", xsci_off_over, ysci_off_over)
# Create PSF
# PSFs already includes geometric distortions based on SIAF info
sp = obj_params['sp']
xtel, ytel = siaf_ap.convert(xsci, ysci, 'sci', 'tel')
hdul = inst.calc_psf_from_coeff(sp=sp, coord_vals=(xtel,ytel), coord_frame='tel')
# Expand PSF to full frame and offset to proper position
ny_pix, nx_pix = (siaf_ap.YSciSize, siaf_ap.XSciSize)
ny_pix_over, nx_pix_over = np.array([ny_pix, nx_pix]) * osamp
shape_new = (ny_pix*osamp, nx_pix*osamp)
delyx = (ysci_off_over, xsci_off_over)
image_full = pad_or_cut_to_size(hdul[0].data, shape_new, offset_vals=delyx)
# Make new HDUList of target (just central source so far)
hdul_full = fits.HDUList(fits.PrimaryHDU(data=image_full, header=hdul[0].header))
fig, ax = plt.subplots(1,1)
extent = 0.5 * np.array([-1,1,-1,1]) * inst.fov_pix * inst.pixelscale
ax.imshow(hdul_full[0].data, extent=extent)
ax.set_xlabel('Arcsec')
ax.set_ylabel('Arcsec')
ax.tick_params(axis='both', color='white', which='both')
for k in ax.spines.keys():
ax.spines[k].set_color('white')
ax.xaxis.get_major_locator().set_params(nbins=9, steps=[1, 2, 5, 10])
ax.yaxis.get_major_locator().set_params(nbins=9, steps=[1, 2, 5, 10])
fig.tight_layout()
```
## 1.3 Convolve extended disk image
Properly including extended objects is a little more complicated than for point sources. First, we need properly format the input model to a pixel binning and flux units appropriate for the simulations (ie., pixels should be equal to oversampled PSFs with flux units of counts/sec). Then, the image needs to be rotated relative to the 'idl' coordinate plane and subsequently shifted for any pointing offsets. Once in the appropriate 'idl' system
### 1.3.1 PSF Grid
```
# Create grid locations for array of PSFs to generate
field_rot = 0 if inst._rotation is None else inst._rotation
rvals = 10**(np.linspace(-2,1,7))
thvals = np.linspace(0, 360, 4, endpoint=False)
rvals_all = [0]
thvals_all = [0]
for r in rvals:
for th in thvals:
rvals_all.append(r)
thvals_all.append(th)
rvals_all = np.array(rvals_all)
thvals_all = np.array(thvals_all)
xgrid_off, ygrid_off = coords.rtheta_to_xy(rvals_all, thvals_all)
# xyoff_half = 10**(np.linspace(-2,1,5))
# xoff = yoff = np.concatenate([-1*xyoff_half[::-1],[0],xyoff_half])
# # Mask Offset grid positions in arcsec
# xgrid_off, ygrid_off = np.meshgrid(xoff, yoff)
# xgrid_off, ygrid_off = xgrid_off.flatten(), ygrid_off.flatten()
# Science positions in detector pixels
xoff_sci_asec, yoff_sci_asec = coords.xy_rot(-1*xgrid_off, -1*ygrid_off, -1*field_rot)
xsci = xoff_sci_asec / siaf_ap.XSciScale + siaf_ap.XSciRef
ysci = yoff_sci_asec / siaf_ap.YSciScale + siaf_ap.YSciRef
xtel, ytel = siaf_ap.convert(xsci, ysci, 'sci', 'tel')
plt.plot(xtel, ytel, marker='o', ls='none', alpha=0.5)
%%time
# Now, create all PSFs, one for each (xsci, ysci) location
# Only need to do this once. Can be used for multiple dither positions.
hdul_psfs = inst.calc_psf_from_coeff(coord_vals=(xtel, ytel), coord_frame='tel', return_oversample=True)
```
### 1.3.2 Disk Model Image
```
# Disk model information
disk_params = {
'file': "Vega/Vega_F444Wsccomb.fits",
'pixscale': inst.pixelscale,
'wavelength': 4.4,
'units': 'Jy/pixel',
'dist' : 7.7,
'cen_star' : False,
}
# Open model and rebin to PSF sampling
# Scale to instrument wavelength assuming grey scattering function
# Converts to phot/sec/lambda
hdul_disk_model = image_manip.make_disk_image(inst, disk_params, sp_star=obj_params['sp'])
# Rotation necessary to go from sky coordinates to 'idl' frame
rotate_to_idl = -1*(tel_point.siaf_ap_obs.V3IdlYAngle + tel_point.pos_ang)
```
### 1.3.3 Dither Position
```
# Select the first dither location offset
delx, dely = tel_point.position_offsets_act[0]
hdul_out = image_manip.rotate_shift_image(hdul_disk_model, PA_offset=rotate_to_idl,
delx_asec=delx, dely_asec=dely)
sci_cen = (siaf_ap.XSciRef, siaf_ap.YSciRef)
# Distort image on 'sci' coordinate grid
im_sci, xsci_im, ysci_im = image_manip.distort_image(hdul_out, ext=0, to_frame='sci', return_coords=True,
aper=siaf_ap, sci_cen=sci_cen)
# Distort image onto 'tel' (V2, V3) coordinate grid for plot illustration
im_tel, v2_im, v3_im = image_manip.distort_image(hdul_out, ext=0, to_frame='tel', return_coords=True,
aper=siaf_ap, sci_cen=sci_cen)
# Plot locations for PSFs that we will generate
fig, ax = plt.subplots(1,1)
# Show image in V2/V3 plane
extent = [v2_im.min(), v2_im.max(), v3_im.min(), v3_im.max()]
ax.imshow(im_tel**0.1, extent=extent)
# Add on SIAF aperture boundaries
tel_point.plot_inst_apertures(ax=ax, clear=False, label=True)
tel_point.plot_ref_aperture(ax=ax)
tel_point.plot_obs_aperture(ax=ax, color='C4')
# Add PSF location points
v2, v3 = siaf_ap.convert(xsci, ysci, 'sci', 'tel')
ax.scatter(v2, v3, marker='.', alpha=0.5, color='C3', edgecolors='none', linewidths=0)
ax.set_title('Model disk image and PSF Locations in SIAF FoV')
fig.tight_layout()
```
This particular disk image is oversized, so we will need to crop the image after convolving PSFs. We may want to consider trimming some of this image prior to convolution, depending on how some of the FoV is blocked before reaching the coronagraphic optics.
```
# If the image is too large, then this process will eat up much of your computer's RAM
# So, crop image to more reasonable size (20% oversized)
osamp = inst.oversample
xysize = int(1.2 * np.max([siaf_ap.XSciSize,siaf_ap.YSciSize]) * osamp)
xy_add = osamp - np.mod(xysize, osamp)
xysize += xy_add
im_sci = pad_or_cut_to_size(im_sci, xysize)
hdul_disk_model_sci = fits.HDUList(fits.PrimaryHDU(data=im_sci, header=hdul_out[0].header))
# Convolve image
im_conv = image_manip.convolve_image(hdul_disk_model_sci, hdul_psfs, aper=siaf_ap)
import scipy
from scipy import fftpack
from astropy.convolution import convolve, convolve_fft
psf = hdul_psfs[-1].data
im_temp = im_sci.copy()
norm = psf.sum()
psf = psf / norm
res = convolve_fft(im_temp, psf, fftn=fftpack.fftn, ifftn=fftpack.ifftn, allow_huge=True)
res *= norm
im_conv = res
# Add cropped image to final oversampled image
im_conv = pad_or_cut_to_size(im_conv, hdul_full[0].data.shape)
hdul_full[0].data = im_conv
def quick_ref_psf(idl_coord, inst, tel_point, out_shape, sp=None):
"""
Create a quick reference PSF for subtraction of the science target.
"""
# Observed SIAF aperture
siaf_ap = tel_point.siaf_ap_obs
# Location of observation
xidl, yidl = idl_coord
# Get offset in SCI pixels
xsci_off, ysci_off = np.array(siaf_ap.convert(xidl, yidl, 'idl', 'sci')) - \
np.array(siaf_ap.reference_point('sci'))
# Get oversampled pixels offests
osamp = inst.oversample
xsci_off_over, ysci_off_over = np.array([xsci_off, ysci_off]) * osamp
yx_offset = (ysci_off_over, xsci_off_over)
# Create PSF
prev_log = webbpsf_ext.conf.logging_level
setup_logging('WARN', verbose=False)
xtel, ytel = siaf_ap.convert(xidl, yidl, 'idl', 'tel')
hdul_psf_ref = inst.calc_psf_from_coeff(sp=sp, coord_vals=(xtel, ytel), coord_frame='tel')
setup_logging(prev_log, verbose=False)
im_psf = pad_or_cut_to_size(hdul_psf_ref[0].data, out_shape, offset_vals=yx_offset)
return im_psf
# Subtract a reference PSF from the science data
coord_vals = tel_point.position_offsets_act[0]
im_psf = quick_ref_psf(coord_vals, inst, tel_point, hdul_full[0].data.shape, sp=sp_star)
im_ref = image_manip.frebin(im_psf, scale=1/osamp)
# Rebin science data to detector pixels
im_sci = image_manip.frebin(hdul_full[0].data, scale=1/osamp) + im_ref
imdiff = im_sci - im_ref
# De-rotate to sky orientation
imrot = image_manip.rotate_offset(imdiff, rotate_to_idl, reshape=False, cval=np.nan)
from matplotlib.colors import LogNorm
from webbpsf_ext.coords import plotAxes
fig, axes = plt.subplots(1,3, figsize=(12,4.5))
############################
# Plot raw image
ax = axes[0]
im = im_sci
mn = np.median(im)
std = np.std(im)
vmin = 0
vmax = mn+10*std
xsize_asec = siaf_ap.XSciSize * siaf_ap.XSciScale
ysize_asec = siaf_ap.YSciSize * siaf_ap.YSciScale
extent = [-1*xsize_asec/2, xsize_asec/2, -1*ysize_asec/2, ysize_asec/2]
norm = LogNorm(vmin=im.max()/1e5, vmax=im.max())
ax.imshow(im, extent=extent, norm=norm)
ax.set_title("Raw Image (log scale)")
ax.set_xlabel('XSci (arcsec)')
ax.set_ylabel('YSci (arcsec)')
plotAxes(ax, angle=-1*siaf_ap.V3SciYAngle)
############################
# Basic PSF subtraction
# Subtract a near-perfect reference PSF
ax = axes[1]
norm = LogNorm(vmin=imdiff.max()/1e5, vmax=imdiff.max())
ax.imshow(imdiff, extent=extent, norm=norm)
ax.set_title("PSF Subtracted (log scale)")
ax.set_xlabel('XSci (arcsec)')
ax.set_ylabel('YSci (arcsec)')
plotAxes(ax, angle=-1*siaf_ap.V3SciYAngle)
############################
# De-rotate to sky orientation
ax = axes[2]
ax.imshow(imrot, extent=extent, norm=norm)
ax.set_title("De-Rotated (log scale)")
ax.set_xlabel('RA offset (arcsec)')
ax.set_ylabel('Dec offset (arcsec)')
plotAxes(ax, position=(0.95,0.35), label1='E', label2='N')
for i, ax in enumerate(axes.flatten()):
ax.xaxis.get_major_locator().set_params(nbins=7, steps=[1, 2, 5, 10])
ax.yaxis.get_major_locator().set_params(nbins=7, steps=[1, 2, 5, 10])
fig.suptitle(f"Fomalhaut ({siaf_ap.AperName})", fontsize=14)
fig.tight_layout()
# Save image to FITS file
hdu_diff = fits.PrimaryHDU(imdiff)
copy_keys = [
'PIXELSCL', 'DISTANCE',
'INSTRUME', 'FILTER', 'PUPIL', 'CORONMSK',
'APERNAME', 'MODULE', 'CHANNEL',
'DET_NAME', 'DET_X', 'DET_Y', 'DET_V2', 'DET_V3'
]
hdr = hdu_diff.header
for head_temp in (inst.psf_coeff_header, hdul_out[0].header):
for key in copy_keys:
try:
hdr[key] = (head_temp[key], head_temp.comments[key])
except (AttributeError, KeyError):
pass
hdr['PIXELSCL'] = inst.pixelscale
name = obj_params['name']
outfile = f'Vega/{name}_{inst.aperturename}_.fits'
hdu_diff.writeto(outfile, overwrite=True)
```
| github_jupyter |
### Insert Spaces Between Words
Given a sentence of words which do not contain spaces between them, list all possible sentences with
spaces in the appropriate places.
For example, given s = "catsanddog", dict = ["cat", "cats", "and", "sand", "dog"], the solution is ["cats and dog", "cat sand dog"].
See [The example problem](https://www.programcreek.com/2014/03/leetcode-word-break-ii-java/)
```
debugging = False
#debugging = True
logging = True
def dbg(f, *args):
if debugging:
print((' DBG:' + f).format(*args))
def log(f, *args):
if logging:
print((f).format(*args))
def logError(f, *args):
if logging:
print(('*** ERROR:' + f).format(*args))
def className(instance):
return type(instance).__name__
class TrieDict(object):
def insert(node, word):
for ch in word:
if ch in node.children:
node = node.children[ch]
else:
newn = TrieDict()
node.children[ch] = newn
node = newn
node.endWord = True
def __init__(node, words=[]):
node.children = {}
node.endWord = False
for word in words:
node.insert(word)
def nextNode(node, ch):
return None if ch not in node.children else node.children[ch]
def toList(node):
words = []
def addWords(node, word, words):
for c in node.children:
n = node.children[c]
w = word + c
if n.endWord:
words.append(w)
addWords(n, w, words)
addWords(node, '', words)
return words
def __str__(node):
return str(node.toList())
dictRoot = TrieDict(
['tit', 'bell', 'cat', 'cats', 'cattle',
'and', 'at', 'a', 'an', 'bet', 'bat', 'dog', 'dogs', 'sand',
'battle', 'cab', 'jot', 'jab', 'belt', 'let', 'tell',
'boat', 'tab', 'oat', 'toad', 'tot', 'trot', 'eat',
'east', 'eel', 'seal', 'lead', 'lee', 'tat', 'the', 'lab',
'lot', 'motor', 'debt', 'jolt', 'bomb', 'rot', 'very',
'quick', 'lazy', 'jump', 'jumped', 'over', 'ugly'
])
catdogDict = TrieDict(["cat", "cats", "and", "sand", "dog"])
print(catdogDict)
str(dictRoot)
def splitWords(s, rootDict):
results = set()
def findSplits(currentWord, partialResult, partialSentence, tdict):
if len(partialSentence):
c = partialSentence[0]
s = partialSentence[1:]
r = partialResult + c
n = tdict.nextNode(c)
if n:
if n.endWord:
if len(s) == 0:
results.add(r)
return # Done with sentence, don't add extra space.
findSplits('', r + ' ', s, rootDict)
findSplits(c, r, s, n)
# If this letter doesn't match a dictionary path then this is not
# a valid word, and thus the sentence is dead to us.
findSplits('', '', s, rootDict)
return results
print(splitWords('', dictRoot))
print(splitWords("catsanddog", dictRoot))
print(splitWords("anuglyveryquickdogjumpedoverthelazycat", dictRoot))
```
| github_jupyter |
## Naive Bayes Classification
```
#Author-Vishal Burman
import matplotlib.pyplot as plt
from mxnet import nd, gluon
import math
%matplotlib inline
#We download the mnist dataset through mxnet's Gluon high-level API
#We normalize the image and remove the last dimension
def transform(data, label):
return nd.floor(data/128).astype('float32').squeeze(axis=-1), label
mnist_train=gluon.data.vision.MNIST(train=True, transform=transform)
mnist_test=gluon.data.vision.MNIST(train=False, transform=transform)
#We can access a particular example
image, label=mnist_train[2]
image.shape, label
image.shape, label.dtype
#We can also access multiple examples at the same time
images, labels=mnist_train[10:38]
images.shape, label.dtype
#Creating a function to visualise these samples
def show_images(imgs, num_rows, num_cols, titles=None, scale=1.5):
"""Plot a list of images."""
figsize = (num_cols * scale, num_rows * scale)
axes = plt.subplots(num_rows, num_cols, figsize=figsize)[1].flatten()
for i, (ax, img) in enumerate(zip(axes, imgs)):
ax.imshow(img.asnumpy())
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
if titles:
ax.set_title(titles[i])
return axes
show_images(images, 2, 9);
##Using the naive bayes classifier on mnist dataset
#Using the chain rule of probablities
#p(x|y)=p(x1|y).p(x2|x1, y)....p(xd|x1...xd-1, y)
#The above calculation is still like 2^d parameters
#If we make an assumption that each feature are independent
#The equation reduces to argmax{Product(p(xi|y)*p(y)) i=i->d}
#First we calculate the p(y) which is the probablity of each number appearing in the dataset
X, Y=mnist_train[:]
n_y=nd.zeros((10))
for y in range(10):
n_y[y]=(Y==y).sum()
P_y=n_y/n_y.sum()
P_y
n_x=nd.zeros((10, 28, 28))
for y in range(10):
n_x[y]=nd.array(X.asnumpy()[Y==y].sum(axis=0))
P_xy = (n_x+1) / (n_y+1).reshape((10,1,1))
show_images(P_xy, 2, 5);
def bayes_pred(x):
x=x.expand_dims(axis=0)
p_xy=P_xy*x+(1-P_xy)*(1-x)
p_xy=p_xy.reshape((10, -1)).prod(axis=1)
return p_xy*P_y
image, label=mnist_test[0]
bayes_pred(image)
#Well the above went horribly wrong...its numerical underflow ie multiplying all the small numbers
#leads to something smaller until it is rounded down to zero
#To fix the above problem we use the following property:
#log ab=log a+ log b
a=0.1
print("underflow", a**784)
print("logarithm is normal", 784*math.log(a))
log_P_xy=nd.log(P_xy)
log_P_xy_neg=nd.log(1-P_xy)
log_P_y=nd.log(P_y)
def bayes_pred_stable(x):
x = x.expand_dims(axis=0) # (28, 28) -> (1, 28, 28)
p_xy = log_P_xy * x + log_P_xy_neg * (1-x)
p_xy = p_xy.reshape((10,-1)).sum(axis=1) # p(x|y)
return p_xy + log_P_y
py = bayes_pred_stable(image)
py
#Checking if the prediction is correct
py.argmax(axis=0).asscalar()==label
def predict(X):
return [bayes_pred_stable(x).argmax(axis=0).asscalar() for x in X]
X, y = mnist_test[:18]
show_images(X, 2, 9, titles=predict(X));
#Computing the overall accuracy of the classifier
X, y = mnist_test[:]
py = predict(X)
'Validation accuracy', (nd.array(py).asnumpy() == y).sum() / len(y)
#The poor performance is due to incorrect statistical assumptions that we made in our model:
#We assumed each and every pixel is independently generated depending only on the label
```
| github_jupyter |
# Linear Regression Example
Let's walk through the steps of the official documentation example. Doing this will help your ability to read from the documentation, understand it, and then apply it to your own problems.
```
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('Linear Regression').getOrCreate()
from pyspark.ml.regression import LinearRegression
# Load training data
training = spark.read.format("libsvm").load("data/sample_linear_regression_data.txt")
```
Interesting! We haven't seen libsvm formats before. In fact the aren't very popular when working with datasets in Python, but the Spark Documentation makes use of them a lot because of their formatting. Let's see what the training data looks like:
```
training.show()
```
This is the format that Spark expects. Two columns with the names "label" and "features".
The "label" column then needs to have the numerical label, either a regression numerical value, or a numerical value that matches to a classification grouping. Later on we will talk about unsupervised learning algorithms that by their nature do not use or require a label.
The feature column has inside of it a vector of all the features that belong to that row. Usually what we end up doing is combining the various feature columns we have into a single 'features' column using the data transformations we've learned about.
Let's continue working through this simple example!
```
# These are the default values for the featuresCol, labelCol, predictionCol
lr = LinearRegression(featuresCol='features', labelCol='label', predictionCol='prediction')
# You could also pass in additional parameters for regularization, do the reading
# in ISLR to fully understand that, after that its just some simple parameter calls.
# Check the documentation with Shift+Tab for more info!
# Fit the model
lrModel = lr.fit(training)
# Print the coefficients and intercept for linear regression
print("Coefficients: {}".format(str(lrModel.coefficients))) # For each feature...
print('\n')
print("Intercept:{}".format(str(lrModel.intercept)))
```
There is a summary attribute that contains even more info!
```
# Summarize the model over the training set and print out some metrics
trainingSummary = lrModel.summary
```
Lots of info, here are a few examples:
```
trainingSummary.residuals.show()
print("RMSE: {}".format(trainingSummary.rootMeanSquaredError))
print("r2: {}".format(trainingSummary.r2))
```
## Train/Test Splits
But wait! We've commited a big mistake, we never separated our data set into a training and test set. Instead we trained on ALL of the data, something we generally want to avoid doing. Read ISLR and check out the theory lecture for more info on this, but remember we won't get a fair evaluation of our model by judging how well it does again on the same data it was trained on!
Luckily Spark DataFrames have an almost too convienent method of splitting the data! Let's see it:
```
all_data = spark.read.format("libsvm").load("sample_linear_regression_data.txt")
# Pass in the split between training/test as a list.
# No correct, but generally 70/30 or 60/40 splits are used.
# Depending on how much data you have and how unbalanced it is.
train_data,test_data = all_data.randomSplit([0.7,0.3])
train_data.show()
test_data.show()
unlabeled_data = test_data.select('features')
unlabeled_data.show()
```
Now we only train on the train_data
```
correct_model = lr.fit(train_data)
```
Now we can directly get a .summary object using the evaluate method:
```
test_results = correct_model.evaluate(test_data)
test_results.residuals.show()
print("RMSE: {}".format(test_results.rootMeanSquaredError))
```
Well that is nice, but realistically we will eventually want to test this model against unlabeled data, after all, that is the whole point of building the model in the first place. We can again do this with a convenient method call, in this case, transform(). Which was actually being called within the evaluate() method. Let's see it in action:
```
predictions = correct_model.transform(unlabeled_data)
predictions.show()
```
Okay, so this data is a bit meaningless, so let's explore this same process with some data that actually makes a little more intuitive sense!
# Linear Regression Code Along
This notebook is the reference for the video lecture on the Linear Regression Code Along. Basically what we do here is examine a dataset with Ecommerce Customer Data for a company's website and mobile app. Then we want to see if we can build a regression model that will predict the customer's yearly spend on the company's product.
First thing to do is start a Spark Session
```
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('lr_example').getOrCreate()
from pyspark.ml.regression import LinearRegression
# Use Spark to read in the Ecommerce Customers csv file.
data = spark.read.csv("Ecommerce_Customers.csv",inferSchema=True,header=True)
# Print the Schema of the DataFrame
data.printSchema()
data.show()
data.head()
for item in data.head():
print(item)
```
## Setting Up DataFrame for Machine Learning
```
# A few things we need to do before Spark can accept the data!
# It needs to be in the form of two columns
# ("label","features")
# Import VectorAssembler and Vectors
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
data.columns
assembler = VectorAssembler(
inputCols=["Avg Session Length", "Time on App",
"Time on Website",'Length of Membership'],
outputCol="features")
output = assembler.transform(data)
output.select("features").show()
output.show()
final_data = output.select("features",'Yearly Amount Spent')
train_data,test_data = final_data.randomSplit([0.7,0.3])
train_data.describe().show()
test_data.describe().show()
# Create a Linear Regression Model object
lr = LinearRegression(labelCol='Yearly Amount Spent')
# Fit the model to the data and call this model lrModel
lrModel = lr.fit(train_data,)
# Print the coefficients and intercept for linear regression
print("Coefficients: {} Intercept: {}".format(lrModel.coefficients,lrModel.intercept))
test_results = lrModel.evaluate(test_data)
# Interesting results....
test_results.residuals.show()
unlabeled_data = test_data.select('features')
predictions = lrModel.transform(unlabeled_data)
predictions.show()
print("RMSE: {}".format(test_results.rootMeanSquaredError))
print("MSE: {}".format(test_results.meanSquaredError))
```
Excellent results! Let's see how you handle some more realistically modeled data in the Consulting Project!
# Data Transformations
You won't always get data in a convienent format, often you will have to deal with data that is non-numerical, such as customer names, or zipcodes, country names, etc...
A big part of working with data is using your own domain knowledge to build an intuition of how to deal with the data, sometimes the best course of action is to drop the data, other times feature-engineering is a good way to go, or you could try to transform the data into something the Machine Learning Algorithms will understand.
Spark has several built in methods of dealing with thse transformations, check them all out here: http://spark.apache.org/docs/latest/ml-features.html
Let's see some examples of all of this!
```
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('data').getOrCreate()
df = spark.read.csv('fake_customers.csv',inferSchema=True,header=True)
df.show()
```
## Data Features
### StringIndexer
We often have to convert string information into numerical information as a categorical feature. This is easily done with the StringIndexer Method:
```
from pyspark.ml.feature import StringIndexer
df = spark.createDataFrame(
[(0, "a"), (1, "b"), (2, "c"), (3, "a"), (4, "a"), (5, "c")],
["user_id", "category"])
indexer = StringIndexer(inputCol="category", outputCol="categoryIndex")
indexed = indexer.fit(df).transform(df)
indexed.show()
```
The next step would be to encode these categories into "dummy" variables.
### VectorIndexer
VectorAssembler is a transformer that combines a given list of columns into a single vector column. It is useful for combining raw features and features generated by different feature transformers into a single feature vector, in order to train ML models like logistic regression and decision trees. VectorAssembler accepts the following input column types: all numeric types, boolean type, and vector type. In each row, the values of the input columns will be concatenated into a vector in the specified order.
Assume that we have a DataFrame with the columns id, hour, mobile, userFeatures, and clicked:
id | hour | mobile | userFeatures | clicked
----|------|--------|------------------|---------
0 | 18 | 1.0 | [0.0, 10.0, 0.5] | 1.0
userFeatures is a vector column that contains three user features. We want to combine hour, mobile, and userFeatures into a single feature vector called features and use it to predict clicked or not. If we set VectorAssembler’s input columns to hour, mobile, and userFeatures and output column to features, after transformation we should get the following DataFrame:
id | hour | mobile | userFeatures | clicked | features
----|------|--------|------------------|---------|-----------------------------
0 | 18 | 1.0 | [0.0, 10.0, 0.5] | 1.0 | [18.0, 1.0, 0.0, 10.0, 0.5]
```
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
dataset = spark.createDataFrame(
[(0, 18, 1.0, Vectors.dense([0.0, 10.0, 0.5]), 1.0)],
["id", "hour", "mobile", "userFeatures", "clicked"])
dataset.show()
assembler = VectorAssembler(
inputCols=["hour", "mobile", "userFeatures"],
outputCol="features")
output = assembler.transform(dataset)
print("Assembled columns 'hour', 'mobile', 'userFeatures' to vector column 'features'")
output.select("features", "clicked").show()
```
There ar emany more data transformations available, we will cover them once we encounter a need for them, for now these were the most important ones.
Let's continue on to Linear Regression!
# Linear Regression Project
Congratulations! You've been contracted by Hyundai Heavy Industries to help them build a predictive model for some ships. [Hyundai Heavy Industries](http://www.hyundai.eu/en) is one of the world's largest ship manufacturing companies and builds cruise liners.
You've been flown to their headquarters in Ulsan, South Korea to help them give accurate estimates of how many crew members a ship will require.
They are currently building new ships for some customers and want you to create a model and use it to predict how many crew members the ships will need.
Here is what the data looks like so far:
Description: Measurements of ship size, capacity, crew, and age for 158 cruise
ships.
Variables/Columns
Ship Name 1-20
Cruise Line 21-40
Age (as of 2013) 46-48
Tonnage (1000s of tons) 50-56
passengers (100s) 58-64
Length (100s of feet) 66-72
Cabins (100s) 74-80
Passenger Density 82-88
Crew (100s) 90-96
It is saved in a csv file for you called "cruise_ship_info.csv". Your job is to create a regression model that will help predict how many crew members will be needed for future ships. The client also mentioned that they have found that particular cruise lines will differ in acceptable crew counts, so it is most likely an important feature to include in your analysis!
```
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('cruise').getOrCreate()
df = spark.read.csv('cruise_ship_info.csv',inferSchema=True,header=True)
df.printSchema()
df.show()
df.describe().show()
```
## Dealing with the Cruise_line categorical variable
Ship Name is a useless arbitrary string, but the cruise_line itself may be useful. Let's make it into a categorical variable!
```
df.groupBy('Cruise_line').count().show()
from pyspark.ml.feature import StringIndexer
indexer = StringIndexer(inputCol="Cruise_line", outputCol="cruise_cat")
indexed = indexer.fit(df).transform(df)
indexed.head(5)
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
indexed.columns
assembler = VectorAssembler(
inputCols=['Age',
'Tonnage',
'passengers',
'length',
'cabins',
'passenger_density',
'cruise_cat'],
outputCol="features")
output = assembler.transform(indexed)
output.select("features", "crew").show()
final_data = output.select("features", "crew")
train_data,test_data = final_data.randomSplit([0.7,0.3])
from pyspark.ml.regression import LinearRegression
# Create a Linear Regression Model object
lr = LinearRegression(labelCol='crew')
# Fit the model to the data and call this model lrModel
lrModel = lr.fit(train_data)
# Print the coefficients and intercept for linear regression
print("Coefficients: {} Intercept: {}".format(lrModel.coefficients,lrModel.intercept))
test_results = lrModel.evaluate(test_data)
print("RMSE: {}".format(test_results.rootMeanSquaredError))
print("MSE: {}".format(test_results.meanSquaredError))
print("R2: {}".format(test_results.r2))
# R2 of 0.86 is pretty good, let's check the data a little closer
from pyspark.sql.functions import corr
df.select(corr('crew','passengers')).show()
df.select(corr('crew','cabins')).show()
```
Okay, so maybe it does make sense! Well that is good news for us, this is information we can bring to the company!
| github_jupyter |
# Covid-19 Analysis using python language.
> I will do some analysis on the Death rate of the pandemic Covid-19 using python.
- badges: true
- comments: true
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime
worldometer_df = pd.read_csv('worldometer_snapshots_April18_to_May18.csv')
worldometer_df = pd.read_csv('worldometers_snapshots_April18_to_September20.csv')
worldometer_df = pd.read_csv('worldometers_snapshots_April18_to_August1.csv')
worldometer_df = pd.read_csv('worldometers_snapshots_April18_to_July30.csv')
worldometer_df = pd.read_csv('worldometers_snapshots_April18_to_July3.csv')
worldometer_df = pd.read_csv('worldometers_snapshots_October11_to_October12.csv')
worldometer_df
country_name = 'Bangladesh'
country_df = worldometer_df.loc[worldometer_df['Country'] == country_name, :].reset_index(drop=True)
country_df
selected_date = datetime.strptime('10/11/2020', '%d/%m/%Y')
selected_date_df = worldometer_df.loc[worldometer_df['Date'] == selected_date.strftime('%Y-%m-%d'), :].reset_index(drop=True)
selected_date_df
last_date = datetime.strptime('12/10/2020', '%d/%m/%Y')
last_date_df = worldometer_df.loc[worldometer_df['Date'] == last_date.strftime('%Y-%m-%d'), :].reset_index(drop=True)
last_date_df
last_date_df['Case Fatality Ratio'] = last_date_df['Total Deaths'] / last_date_df['Total Cases']
plt.figure(figsize=(12,8))
plt.hist(100 * np.array(last_date_df['Case Fatality Ratio']), bins=np.arange(35))
plt.xlabel('Death Rate (%)', fontsize=16)
plt.ylabel('Number of Countries', fontsize=16)
plt.title('Histogram of Death Rates for various Countries', fontsize=18)
plt.show()
min_number_of_cases = 1000
greatly_affected_df = last_date_df.loc[last_date_df['Total Cases'] > min_number_of_cases,:]
plt.figure(figsize=(12,8))
plt.hist(100 * np.array(greatly_affected_df['Case Fatality Ratio']), bins=np.arange(35))
plt.xlabel('Death Rate (%)', fontsize=16)
plt.ylabel('Number of Countries', fontsize=16)
plt.title('Histogram of Death Rates for various Countries', fontsize=18)
plt.show()
last_date_df['Num Tests per Positive Case'] = last_date_df['Total Tests'] / last_date_df['Total Cases']
min_number_of_cases = 1000
greatly_affected_df = last_date_df.loc[last_date_df['Total Cases'] > min_number_of_cases,:]
x_axis_limit = 80
death_rate_percent = 100 * np.array(greatly_affected_df['Case Fatality Ratio'])
num_test_per_positive = np.array(greatly_affected_df['Num Tests per Positive Case'])
num_test_per_positive[num_test_per_positive > x_axis_limit] = x_axis_limit
total_num_deaths = np.array(greatly_affected_df['Total Deaths'])
population = np.array(greatly_affected_df['Population'])
plt.figure(figsize=(16,12))
plt.scatter(x=num_test_per_positive, y=death_rate_percent,
s=0.5*np.power(np.log(1+population),2),
c=np.log10(1+total_num_deaths))
plt.colorbar()
plt.ylabel('Death Rate (%)', fontsize=16)
plt.xlabel('Number of Tests per Positive Case', fontsize=16)
plt.title('Death Rate as function of Testing Quality', fontsize=18)
plt.xlim(-1, x_axis_limit + 12)
plt.ylim(-0.2,17)
# plot on top of the figure the names of the
#countries_to_display = greatly_affected_df['Country'].unique().tolist()
countries_to_display = ['USA', 'Russia', 'Spain', 'Bangladesh', 'Brazil', 'UK', 'Italy', 'France',
'Germany', 'India', 'Canada', 'Belgium', 'Mexico', 'Netherlands',
'Sweden', 'Portugal', 'UAE', 'Poland', 'Indonesia', 'Romania',
'Israel','Thailand','Kyrgyzstan','El Salvador', 'S. Korea',
'Denmark', 'Serbia', 'Norway', 'Algeria', 'Bahrain','Slovenia',
'Greece','Cuba','Hong Kong','Lithuania', 'Australia', 'Morocco',
'Malaysia', 'Nigeria', 'Moldova', 'Ghana', 'Armenia', 'Bolivia',
'Iraq', 'Hungary', 'Cameroon', 'Azerbaijan']
for country_name in countries_to_display:
country_index = greatly_affected_df.index[greatly_affected_df['Country'] == country_name]
plt.text(x=num_test_per_positive[country_index] + 0.5,
y=death_rate_percent[country_index] + 0.2,
s=country_name, fontsize=10)
plt.show()
good_testing_threshold = 300
good_testing_df = greatly_affected_df.loc[greatly_affected_df['Num Tests per Positive Case'] > good_testing_threshold,:]
good_testing_df
estimated_death_rate_percent = 100 * good_testing_df['Total Deaths'].sum() / good_testing_df['Total Cases'].sum()
print('Death Rate only for "good testing countries" is %.2f%s' %(estimated_death_rate_percent,'%'))
```
| github_jupyter |
# Oakland 311 calls
This notebook is part of Assignment #6.
```
import pandas as pd
import altair as alt
import requests
```
## Download data
Download [311 call data](https://data.oaklandca.gov/Infrastructure/Service-requests-received-by-the-Oakland-Call-Cent/quth-gb8e) from the City of Oakland's Open Data Portal.
```
url = 'https://data.oaklandca.gov/api/views/quth-gb8e/rows.csv?accessType=DOWNLOAD'
r = requests.get(url, allow_redirects=False)
# write the content of the request into a file called `oakland_311.csv`
open('oakland_311.csv', 'wb').write(r.content)
```
## Import the csv into a pandas dataframe
Assign the dataframe into a variable called oakland_311_original
```
oakland_311_original = pd.read_csv('oakland_311.csv')
```
## View data
View the first 5 rows and the last 5 rows of the dataframe.
```
oakland_311_original.head()
oakland_311_original.tail()
```
Try to call the dataframe on its own.
```
oakland_311_original
```
See more info on the dataframe
```
oakland_311_original.info()
```
## Check the types of data
```
oakland_311_original.head()
```
Need to convert two columns: DATETIMEINIT and DATETIMECLOSED
```
oakland_311_original['DATETIMEINIT'].min()
oakland_311_original['DATETIMEINIT'].max()
```
The operations might be wrong.
## Clean data part 1: Properly type the data
### Copy the original dataframe
```
oakland_311 = oakland_311_original.copy()
```
### Covert columns to datetime
```
oakland_311.head()
oakland_311['DATETIMEINIT'] = pd.to_datetime(oakland_311['DATETIMEINIT'])
```
Compare DATETIMEINIT and DATETIMECLOSED
```
oakland_311.head()
oakland_311.info()
oakland_311['DATETIMECLOSED'] = pd.to_datetime(oakland_311['DATETIMECLOSED'])
oakland_311.head()
```
### Now get the earliest and latest date of both columns
```
oakland_311['DATETIMEINIT'].min()
oakland_311['DATETIMEINIT'].max()
```
It shows that the data of year 2009 and 2021 is not complete. May need to exclude these two years later.
```
oakland_311['DATETIMECLOSED'].min()
oakland_311['DATETIMECLOSED'].max()
```
### Get the difference of the two date columns
Create a new column called Close_Time that shows how long it took for a case to be closed.
```
oakland_311['Close_Time'] = oakland_311['DATETIMECLOSED'] - oakland_311['DATETIMEINIT']
```
Check the result.
```
oakland_311.head()
```
## Explore some operations on the new column
### Mean
```
oakland_311['Close_Time'].mean()
```
### Median
```
oakland_311['Close_Time'].median()
```
### Min
```
oakland_311['Close_Time'].min()
```
-2981 days?? It doesn't make sense. Check if this is an error.
```
oakland_311[oakland_311['Close_Time'] < '0 days']
```
All these all errors? Need to ask the city for more information.
### Max
```
oakland_311['Close_Time'].max()
```
This seems like way too long. There might still be an error.
```
oakland_311[oakland_311['Close_Time'] >= '4293 days']
```
May need to ask the city for more info as well.
### Sort data
Check the first 10 rows of sorted data.
```
oakland_311.sort_values(by=['Close_Time'], ascending=False).head(10)
```
## Clean data part 2: Check and vet the data
### Check if there's unique identifier for every row
```
oakland_311['REQUESTID'].nunique()
len(oakland_311)
```
There numbers of unique case IDs and rows in the dataframe are both 852673! There might not be duplicates.
### Subsetting
Still check for any existing duplicates.
```
oakland_311[oakland_311.duplicated()]
```
No exact duplicates found.
```
oakland_311[oakland_311['REQUESTID'].duplicated()]
```
Still no duplicates found.
### Assert
Double-check if the length of the dataframe now matches the number of unique IDs.
```
assert len(oakland_311) == oakland_311['REQUESTID'].nunique()
```
## Export a clean version of the data to a csv
create a new notebook and exclude the column of row ids
```
oakland_311.to_csv('oakland_311_clean.csv', index = False)
```
| github_jupyter |
## 6-2. Qulacs を用いた variational quantum eigensolver (VQE) の実装
この節では、OpenFermion・PySCF を用いて求めた量子化学ハミルトニアンについて、Qulacs を用いてシミュレータ上で variational quantum eigensolver (VQE) を実行し、基底状態を探索する例を示す。
必要なもの
- qulacs
- openfermion
- openfermion-pyscf
- pyscf
- scipy
- numpy
### 必要なパッケージのインストール・インポート
```
## Google Colaboratory上で実行する場合バグを回避するためscipyをダウングレード
!pip install scipy==1.2.1
## 各種ライブラリがインストールされていない場合は実行してください
## Google Colaboratory上で実行する場合'You must restart the runtime in order to use newly installed versions.'と出ますが無視してください。
## runtimeを再開するとクラッシュします。
!pip install qulacs pyscf openfermion openfermionpyscf
import qulacs
from openfermion.transforms import get_fermion_operator, jordan_wigner
from openfermion.transforms import get_sparse_operator
from openfermion.hamiltonians import MolecularData
from openfermionpyscf import run_pyscf
from scipy.optimize import minimize
from pyscf import fci
import numpy as np
import matplotlib.pyplot as plt
```
### ハミルトニアンを作る
前節と同様の手順で、ハミルトニアンを PySCF によって計算する。
```
basis = "sto-3g"
multiplicity = 1
charge = 0
distance = 0.977
geometry = [["H", [0,0,0]],["H", [0,0,distance]]]
description = "tmp"
molecule = MolecularData(geometry, basis, multiplicity, charge, description)
molecule = run_pyscf(molecule,run_scf=1,run_fci=1)
n_qubit = molecule.n_qubits
n_electron = molecule.n_electrons
fermionic_hamiltonian = get_fermion_operator(molecule.get_molecular_hamiltonian())
jw_hamiltonian = jordan_wigner(fermionic_hamiltonian)
```
### ハミルトニアンを qulacs ハミルトニアンに変換する
Qulacs では、ハミルトニアンのようなオブザーバブルは `Observable` クラスによって扱われる。OpenFermion のハミルトニアンを Qulacs の `Observable` に変換する関数 `create_observable_from_openfermion_text` が用意されているので、これを使えば良い。
```
from qulacs import Observable
from qulacs.observable import create_observable_from_openfermion_text
qulacs_hamiltonian = create_observable_from_openfermion_text(str(jw_hamiltonian))
```
### ansatz を構成する
Qulacs 上で量子回路を構成する。ここでは、量子回路は超伝導量子ビットによる実験 (A. Kandala _et. al._ , “Hardware-efficient variational quantum eigensolver for small molecules and quantum magnets“, Nature **549**, 242–246) で用いられたものを模して作った。
```
from qulacs import QuantumState, QuantumCircuit
from qulacs.gate import CZ, RY, RZ, merge
depth = n_qubit
def he_ansatz_circuit(n_qubit, depth, theta_list):
"""he_ansatz_circuit
Returns hardware efficient ansatz circuit.
Args:
n_qubit (:class:`int`):
the number of qubit used (equivalent to the number of fermionic modes)
depth (:class:`int`):
depth of the circuit.
theta_list (:class:`numpy.ndarray`):
rotation angles.
Returns:
:class:`qulacs.QuantumCircuit`
"""
circuit = QuantumCircuit(n_qubit)
for d in range(depth):
for i in range(n_qubit):
circuit.add_gate(merge(RY(i, theta_list[2*i+2*n_qubit*d]), RZ(i, theta_list[2*i+1+2*n_qubit*d])))
for i in range(n_qubit//2):
circuit.add_gate(CZ(2*i, 2*i+1))
for i in range(n_qubit//2-1):
circuit.add_gate(CZ(2*i+1, 2*i+2))
for i in range(n_qubit):
circuit.add_gate(merge(RY(i, theta_list[2*i+2*n_qubit*depth]), RZ(i, theta_list[2*i+1+2*n_qubit*depth])))
return circuit
```
### VQE のコスト関数を定義する
[5-1節](5.1_variational_quantum_eigensolver.ipynb)で説明した通り、VQE はパラメータ付きの量子回路 $U(\theta)$ から出力される状態 $|\psi(\theta)\rangle = U(\theta)|0\rangle$ に関するハミルトニアンの期待値
$$\langle H (\theta)\rangle = \langle\psi(\theta)|H|\psi(\theta)\rangle$$
を最小化することで、近似的な基底状態を得る。以下にこのハミルトニアンの期待値を返す関数を定義する。
```
def cost(theta_list):
state = QuantumState(n_qubit) #|00000> を準備
circuit = he_ansatz_circuit(n_qubit, depth, theta_list) #量子回路を構成
circuit.update_quantum_state(state) #量子回路を状態に作用
return qulacs_hamiltonian.get_expectation_value(state) #ハミルトニアンの期待値を計算
```
### VQE を実行する
準備ができたので、VQE を実行する。最適化には scipy に実装されている BFGS 法を用い、初期パラメータはランダムに選ぶ。数十秒で終わるはずである。
```
cost_history = []
init_theta_list = np.random.random(2*n_qubit*(depth+1))*1e-1
cost_history.append(cost(init_theta_list))
method = "BFGS"
options = {"disp": True, "maxiter": 50, "gtol": 1e-6}
opt = minimize(cost, init_theta_list,
method=method,
callback=lambda x: cost_history.append(cost(x)))
```
実行結果をプロットしてみると、正しい解に収束していることが見て取れる。
```
plt.rcParams["font.size"] = 18
plt.plot(cost_history, color="red", label="VQE")
plt.plot(range(len(cost_history)), [molecule.fci_energy]*len(cost_history), linestyle="dashed", color="black", label="Exact Solution")
plt.xlabel("Iteration")
plt.ylabel("Energy expectation value")
plt.legend()
plt.show()
```
興味のある読者は、水素原子間の距離 `distance` を様々に変えて基底状態を計算し、水素分子が最も安定になる原子間距離を探してみてほしい。(ansatzの性能にもよるが、およそ0.74オングストロームになるはずである)
| github_jupyter |
# Let's try some DS on Thanksgiving data!!
Using a SurveyMonkey poll, we asked 1,058 respondents on Nov. 17, 2015 the following questions about their Thanksgiving:
* Do you celebrate Thanksgiving?
* What is typically the main dish at your Thanksgiving dinner?
* Other (please specify)
* How is the main dish typically cooked?
* Other (please specify)
* What kind of stuffing/dressing do you typically have?
* Other (please specify)
* What type of cranberry sauce do you typically have?
* Other (please specify)
* Do you typically have gravy?
* Which of these side dishes are typically served at your Thanksgiving dinner? Please select all that apply.
* Brussel sprouts
* Carrots
* Cauliflower
* Corn
* Cornbread
* Fruit salad
* Green beans/green bean casserole
* Macaroni and cheese
* Mashed potatoes
* Rolls/biscuits
* Vegetable salad
* Yams/sweet potato casserole
* Other (please specify)
* Which type of pie is typically served at your Thanksgiving dinner? Please select all that apply.
* Apple
* Buttermilk
* Cherry
* Chocolate
* Coconut cream
* Key lime
* Peach
* Pecan
* Pumpkin
* Sweet Potato
* None
* Other (please specify)
* Which of these desserts do you typically have at Thanksgiving dinner? Please select all that apply.
* Apple cobbler
* Blondies
* Brownies
* Carrot cake
* Cheesecake
* Cookies
* Fudge
* Ice cream
* Peach cobbler
* None
* Other (please specify)
* Do you typically pray before or after the Thanksgiving meal?
* How far will you travel for Thanksgiving?
* Will you watch any of the following programs on Thanksgiving? Please select all that apply.
* Macy's Parade
* What's the age cutoff at your "kids' table" at Thanksgiving?
* Have you ever tried to meet up with hometown friends on Thanksgiving night?
* Have you ever attended a "Friendsgiving?"
* Will you shop any Black Friday sales on Thanksgiving Day?
* Do you work in retail?
* Will you employer make you work on Black Friday?
* How would you describe where you live?
* Age
* What is your gender?
* How much total combined money did all members of your HOUSEHOLD earn last year?
* US Region
Download data here: https://github.com/opencubelabs/notebooks/blob/master/data_science/data/thanksgiving.csv
```
import pandas as pd
data = pd.read_csv("data/thanksgiving.csv", encoding="Latin-1")
data.head()
data["Do you celebrate Thanksgiving?"].unique()
data.columns[59:]
```
Using this Thanksgiving survey data, we can answer quite a few interesting questions, like:
* Do people in Suburban areas eat more Tofurkey than people in Rural areas?
* Where do people go to Black Friday sales most often?
* Is there a correlation between praying on Thanksgiving and income?
* What income groups are most likely to have homemade cranberry sauce?
Let's get familier with applying, grouping and aggregation in Pandas
### Applying functions to Series in pandas
There are times when we're using pandas that we want to apply a function to every row or every column in the data. A good example is getting from the values in our What is your gender? column to numeric values. We'll assign 0 to Male, and 1 to Female.
**pandas.Series.apply**
```
data["What is your gender?"].value_counts(dropna=False)
import math
def gender_code(gender_string):
if isinstance(gender_string, float) and math.isnan(gender_string):
return gender_string
return int(gender_string == "Female")
data["gender"] = data["What is your gender?"].apply(gender_code)
data["gender"].value_counts(dropna=False)
```
### Applying functions to DataFrames in pandas
We can use the apply method on DataFrames as well as Series. When we use the **pandas.DataFrame.apply** method, an entire row or column will be passed into the function we specify. By default, apply will work across each column in the DataFrame. If we pass the axis=1 keyword argument, it will work across each row.
```
data.apply(lambda x: x.dtype).head() # find data type of each column
# Let's clean up the income using apply method
data["How much total combined money did all members of your HOUSEHOLD earn last year?"].value_counts(dropna=False)
```
We have 4 different patterns for the values in the column:
* X to Y — an example is 25,000 USD to 49,999 USD --> Average value
* NaN --> No change
* X and up — an example is 200,000 USD and up. --> Value
* Prefer not to answer --> NaN
```
import numpy as np
def clean_income(value):
if value == "$200,000 and up":
return 200000
elif value == "Prefer not to answer":
return np.nan
elif isinstance(value, float) and math.isnan(value):
return np.nan
value = value.replace(",", "").replace("$", "")
income_high, income_low = value.split(" to ")
return (int(income_high) + int(income_low)) / 2
data["income"] = data["How much total combined money did all members of your HOUSEHOLD earn last year?"].apply(clean_income)
data["income"].head()
```
### Grouping data with pandas
When performing data analysis, it's often useful to explore only a subset of the data. For example, what if we want to compare income between people who tend to eat homemade cranberry sauce for Thanksgiving vs people who eat canned cranberry sauce?
**pandas.DataFrame.groupby**
```
# let's check the unique values in the columns
data["What type of cranberry saucedo you typically have?"].value_counts()
homemade = data[data["What type of cranberry saucedo you typically have?"] == "Homemade"]
canned = data[data["What type of cranberry saucedo you typically have?"] == "Canned"]
print(homemade["income"].mean())
print(canned["income"].mean())
grouped = data.groupby("What type of cranberry saucedo you typically have?")
grouped
grouped.groups
grouped.size()
for name, group in grouped:
print(name)
print(group.shape)
print(type(group))
grouped["income"]
```
### Aggregating values in groups
If all we could do was split a DataFrame into groups, it wouldn't be of much use. The real power of groups is in the computations we can do after creating groups. We do these computations through the **pandas.GroupBy.aggregate** method, which we can abbreviate as agg. This method allows us to perform the same computation on every group.
For example, we could find the average income for people who served each type of cranberry sauce for Thanksgiving (Canned, Homemade, None, etc).
```
grouped["income"].agg(np.mean)
grouped.agg(np.mean)
%matplotlib inline
sauce = grouped.agg(np.mean)
sauce["income"].plot(kind="bar")
```
### Aggregating with multiple columns
We can call groupby with multiple columns as input to get more granular groups. If we use the *What type of cranberry saucedo you typically have?* and *What is typically the main dish at your Thanksgiving dinner?* columns as input, we'll be able to find the average income of people who eat Homemade cranberry sauce and Tofurkey, for example:
```
grouped = data.groupby(["What type of cranberry saucedo you typically have?", "What is typically the main dish at your Thanksgiving dinner?"])
grouped.agg(np.mean)
```
Above output gives us some interesting patterns:
* People who have Turducken and Homemade cranberry sauce seem to have high household incomes.
* People who eat Canned cranberry sauce tend to have lower incomes, but those who also have Roast Beef have the lowest incomes.
* It looks like there's one person who has Canned cranberry sauce and doesn't know what type of main dish he's having.
```
grouped["income"].agg([np.mean, np.sum, np.std]).head(10) # Aggregating with multiple functions
```
### Using apply on groups
One of the limitations of aggregation is that each function has to return a single number. While we can perform computations like finding the mean, we can't for example, call value_counts to get the exact count of a category. We can do this using the **pandas.GroupBy.apply** method. This method will apply a function to each group, then combine the results.
In the below code, we'll apply value_counts to find the number of people who live in each area type (Rural, Suburban, etc) who eat different kinds of main dishes for Thanksgiving:
```
grouped = data.groupby("How would you describe where you live?")["What is typically the main dish at your Thanksgiving dinner?"]
grouped.apply(lambda x:x.value_counts())
```
The above table shows us that people who live in different types of areas eat different Thanksgiving main dishes at about the same rate.

| github_jupyter |
<a href="https://colab.research.google.com/github/juliocnsouzadev/notebooks/blob/develop/Bert_tokenizer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Stage 1: Importing dependencies
```
!pip install bert-for-tf2
!pip install sentencepiece
import numpy as np
import math
import re
import pandas as pd
from bs4 import BeautifulSoup
import random
import tensorflow as tf
import tensorflow_hub as hub
from tensorflow.keras import layers
import bert
```
# Stage 2: Data preprocessing
## Loading files
Download and unzip the files
```
!wget http://cs.stanford.edu/people/alecmgo/trainingandtestdata.zip
!unzip trainingandtestdata.zip
!ls
cols = ["sentiment", "id", "date", "query", "user", "text"]
data = pd.read_csv(
"training.1600000.processed.noemoticon.csv",
header=None,
names=cols,
engine="python",
encoding="latin1"
)
data.head()
data.drop(["id", "date", "query", "user"],
axis=1,
inplace=True)
```
## Preprocessing
### Cleaning
```
def clean_tweet(tweet):
tweet = BeautifulSoup(tweet, "lxml").get_text()
# Removing the @
tweet = re.sub(r"@[A-Za-z0-9]+", ' ', tweet)
# Removing the URL links
tweet = re.sub(r"https?://[A-Za-z0-9./]+", ' ', tweet)
# Keeping only letters
tweet = re.sub(r"[^a-zA-Z.!?']", ' ', tweet)
# Removing additional whitespaces
tweet = re.sub(r" +", ' ', tweet)
return tweet
data_clean = [clean_tweet(tweet) for tweet in data.text]
data_labels = data.sentiment.values
data_labels[data_labels == 4] = 1
```
### Tokenization
We need to create a BERT layer to have access to meta data for the tokenizer (like vocab size).
```
FullTokenizer = bert.bert_tokenization.FullTokenizer
bert_layer = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/1",
trainable=False)
vocab_file = bert_layer.resolved_object.vocab_file.asset_path.numpy()
do_lower_case = bert_layer.resolved_object.do_lower_case.numpy()
tokenizer = FullTokenizer(vocab_file, do_lower_case)
def encode_sentence(sent):
return tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sent))
data_inputs = [encode_sentence(sentence) for sentence in data_clean]
```
### Dataset creation
We will create padded batches (so we pad sentences for each batch independently), this way we add the minimum of padding tokens possible. For that, we sort sentences by length, apply padded_batches and then shuffle.
```
data_with_len = [[sent, data_labels[i], len(sent)]
for i, sent in enumerate(data_inputs)]
random.shuffle(data_with_len)
data_with_len.sort(key=lambda x: x[2])
sorted_all = [(sent_lab[0], sent_lab[1])
for sent_lab in data_with_len if sent_lab[2] > 7]
all_dataset = tf.data.Dataset.from_generator(lambda: sorted_all,
output_types=(tf.int32, tf.int32))
next(iter(all_dataset))
BATCH_SIZE = 32
all_batched = all_dataset.padded_batch(BATCH_SIZE, padded_shapes=((None, ), ()))
NB_BATCHES = math.ceil(len(sorted_all) / BATCH_SIZE)
NB_BATCHES_TEST = NB_BATCHES // 10
all_batched.shuffle(NB_BATCHES)
test_dataset = all_batched.take(NB_BATCHES_TEST)
train_dataset = all_batched.skip(NB_BATCHES_TEST)
```
# Stage 3: Model building
```
class DCNN(tf.keras.Model):
def __init__(self,
vocab_size,
emb_dim=128,
nb_filters=50,
FFN_units=512,
nb_classes=2,
dropout_rate=0.1,
training=False,
name="dcnn"):
super(DCNN, self).__init__(name=name)
self.embedding = layers.Embedding(vocab_size,
emb_dim)
self.bigram = layers.Conv1D(filters=nb_filters,
kernel_size=2,
padding="valid",
activation="relu")
self.trigram = layers.Conv1D(filters=nb_filters,
kernel_size=3,
padding="valid",
activation="relu")
self.fourgram = layers.Conv1D(filters=nb_filters,
kernel_size=4,
padding="valid",
activation="relu")
self.pool = layers.GlobalMaxPool1D()
self.dense_1 = layers.Dense(units=FFN_units, activation="relu")
self.dropout = layers.Dropout(rate=dropout_rate)
if nb_classes == 2:
self.last_dense = layers.Dense(units=1,
activation="sigmoid")
else:
self.last_dense = layers.Dense(units=nb_classes,
activation="softmax")
def call(self, inputs, training):
x = self.embedding(inputs)
x_1 = self.bigram(x) # batch_size, nb_filters, seq_len-1)
x_1 = self.pool(x_1) # (batch_size, nb_filters)
x_2 = self.trigram(x) # batch_size, nb_filters, seq_len-2)
x_2 = self.pool(x_2) # (batch_size, nb_filters)
x_3 = self.fourgram(x) # batch_size, nb_filters, seq_len-3)
x_3 = self.pool(x_3) # (batch_size, nb_filters)
merged = tf.concat([x_1, x_2, x_3], axis=-1) # (batch_size, 3 * nb_filters)
merged = self.dense_1(merged)
merged = self.dropout(merged, training)
output = self.last_dense(merged)
return output
```
# Stage 4: Training
```
VOCAB_SIZE = len(tokenizer.vocab)
EMB_DIM = 200
NB_FILTERS = 100
FFN_UNITS = 256
NB_CLASSES = 2
DROPOUT_RATE = 0.2
NB_EPOCHS = 5
Dcnn = DCNN(vocab_size=VOCAB_SIZE,
emb_dim=EMB_DIM,
nb_filters=NB_FILTERS,
FFN_units=FFN_UNITS,
nb_classes=NB_CLASSES,
dropout_rate=DROPOUT_RATE)
if NB_CLASSES == 2:
Dcnn.compile(loss="binary_crossentropy",
optimizer="adam",
metrics=["accuracy"])
else:
Dcnn.compile(loss="sparse_categorical_crossentropy",
optimizer="adam",
metrics=["sparse_categorical_accuracy"])
from google.colab import drive
drive.mount("/content/drive")
checkpoint_path = "/content/drive/MyDrive/Colab Notebooks/Bert/ckpt_bert_tok"
ckpt = tf.train.Checkpoint(Dcnn=Dcnn)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=1)
if ckpt_manager.latest_checkpoint:
ckpt.restore(ckpt_manager.latest_checkpoint)
print("Latest checkpoint restored!!")
class MyCustomCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs=None):
ckpt_manager.save()
print("Checkpoint saved at {}.".format(checkpoint_path))
Dcnn.fit(train_dataset,
epochs=NB_EPOCHS,
callbacks=[MyCustomCallback()])
```
# Stage 5: Evaluation
```
results = Dcnn.evaluate(test_dataset)
print(results)
def get_prediction(sentence):
tokens = encode_sentence(sentence)
inputs = tf.expand_dims(tokens, 0)
output = Dcnn(inputs, training=False)
sentiment = math.floor(output*2)
if sentiment == 0:
print("Ouput of the model: {}\nPredicted sentiment: negative.".format(
output))
elif sentiment == 1:
print("Ouput of the model: {}\nPredicted sentiment: positive.".format(
output))
get_prediction("This movie was pretty interesting.")
get_prediction("Life is wonderful.")
get_prediction("Feeling like crap.")
get_prediction("You are smart like a donkey.")
```
| github_jupyter |
# Deep Neural Networks (DNN) Model Development
## Preparing Packages
```
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
import tensorflow as tf
from sklearn import metrics
from numpy import genfromtxt
from scipy import stats
from sklearn import preprocessing
from keras.callbacks import ModelCheckpoint
from keras.callbacks import Callback
from keras.models import load_model
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score, f1_score, precision_score, recall_score
import keras
from keras.layers import Dense, Flatten, Reshape,Dropout
from keras.layers import Conv2D, MaxPooling2D, LSTM
from keras.models import Sequential
from sklearn.model_selection import train_test_split
import timeit #package for recording the model running time
import time
from keras.callbacks import EarlyStopping
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential, load_model
from keras.layers import Activation, Dropout, Flatten, Dense, Conv2D, Conv3D, MaxPooling3D, Reshape, BatchNormalization, MaxPooling2D
from keras.applications.inception_resnet_v2 import InceptionResNetV2
from keras.callbacks import ModelCheckpoint
from keras import metrics
from keras.optimizers import Adam
from keras import backend as K
from sklearn.metrics import fbeta_score
from sklearn.model_selection import KFold,StratifiedKFold,ShuffleSplit,StratifiedShuffleSplit
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import classification_report,f1_score,accuracy_score
```
## Preparing Functions
```
def win_seg(data,windowsize,overlap):#function for overlap segmentation
length=int((data.shape[0]*data.shape[1]-windowsize)/(windowsize*overlap)+1)
newdata=np.empty((length,windowsize, data.shape[2],1))
data_dim=data.shape[2]
layers=data.shape[3]
data=data.reshape(-1,data_dim,layers)
for i in range(0,length) :
start=int(i*windowsize*overlap)
end=int(start+windowsize)
newdata[i]=data[start:end]
return newdata
def lab_vote(data,windowsize):
y_data=data.reshape(-1,windowsize,1,1)
y_data=win_seg(y_data,windowsize,0.5)
y_data=y_data.reshape(y_data.shape[0],y_data.shape[1],y_data.shape[2])
y_data=stats.mode(y_data,axis=1)
y_data=y_data.mode
y_data=y_data.reshape(-1,1)
y_data=np.float64(keras.utils.to_categorical(y_data))
return y_data
def lab_vote_cat(data,windowsize): # non one-hot coding
y_data=data.reshape(-1,windowsize,1,1)
y_data=win_seg(y_data,windowsize,0.5)
y_data=y_data.reshape(y_data.shape[0],y_data.shape[1],y_data.shape[2])
y_data=stats.mode(y_data,axis=1)
y_data=y_data.mode
y_data=y_data.reshape(-1,1)
return y_data
def write_csv(data):
a = np.asarray(data)
a.tofile('check.csv',sep=',',format='%10.5f')
def average(lst):
a = np.array(lst)
return np.mean(a)
class TimeHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.times = []
def on_epoch_begin(self, batch, logs={}):
self.epoch_time_start = time.time()
def on_epoch_end(self, batch, logs={}):
self.times.append(time.time() - self.epoch_time_start)
def f1(y_true, y_pred):
y_pred = K.round(y_pred)
tp = K.sum(K.cast(y_true*y_pred, 'float'), axis=0)
# tn = K.sum(K.cast((1-y_true)*(1-y_pred), 'float'), axis=0)
fp = K.sum(K.cast((1-y_true)*y_pred, 'float'), axis=0)
fn = K.sum(K.cast(y_true*(1-y_pred), 'float'), axis=0)
p = tp / (tp + fp + K.epsilon())
r = tp / (tp + fn + K.epsilon())
f1 = 2*p*r / (p+r+K.epsilon())
f1 = tf.where(tf.is_nan(f1), tf.zeros_like(f1), f1)
return K.mean(f1)
```
## Convolutional LSTM Model Development
```
#loading the training and testing data
os.chdir("...") #changing working directory
buffer = np.float64(preprocessing.scale(genfromtxt('S3_X.csv', delimiter=','))) # using S3 as an example
x_data=buffer.reshape(-1,40,30,1)
x_data=win_seg(x_data,40,0.5) # data segmentation with 0.5 overlap
#majority vote on training label
buffer = np.float64(genfromtxt('S3_Y.csv', delimiter=','))-1 #0 based index
y_data=lab_vote(buffer,40)
y_data2=lab_vote_cat(buffer,40) # for stratification purposes
#five round Stratified Random Shuffle
SRS=StratifiedShuffleSplit(n_splits=5, test_size=0.1, random_state=42) #split the train and test by 9:1
#model evaluation metrics
acc_score=list()
f_score=list()
eopch_time_record=list()
oper_time_record=list()
i=0
for train_index, test_index in SRS.split(x_data,y_data):
X_train, X_test = x_data[train_index], x_data[test_index]
y_train, y_test = y_data[train_index], y_data[test_index]
#split the train data into training (training the model) and validation (tuning hypeparameters) by 8:2
X_training, X_validation, y_training, y_validation = train_test_split(X_train, y_train, test_size=0.20)
#setup model parameters
data_dim = X_train.shape[2] #y of 2D Motion Image
timesteps = X_train.shape[1] #x of 2D Motion Image
num_classes = y_train.shape[1]
batchsize=300
epcoh=300
#build model
model = Sequential()
#five convolutional layers as an exmaple, adjust the convolutional layer depth if needed
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh',input_shape=(timesteps, data_dim,1)))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
#turn the multilayer tensor into single layer tensor
model.add(Reshape((40, -1),input_shape=(40,30,64)))
model.add(Dropout(0.5)) #add dropout layers for controlling overfitting
model.add(LSTM(128, return_sequences=True, input_shape=(40, 1920))) # returns a sequence of vectors
model.add(Dropout(0.5)) #add dropout layers for controlling overfitting
model.add(LSTM(128)) # return a single vector
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),metrics=['accuracy',f1])
checkpointer = ModelCheckpoint(filepath="2D_CNN5_LSTM_checkpoint(F1)_sss_%s.h5" % i, monitor='val_f1',verbose=1, mode='max', save_best_only=True)
time_callback = TimeHistory() #record the model training time for each epoch
callbacks_list = [checkpointer,time_callback]
train_history=model.fit(X_training, y_training,
batch_size=batchsize, epochs=epcoh,callbacks=callbacks_list,
validation_data=(X_validation, y_validation))
eopch_time=time_callback.times
eopch_time_record.append(eopch_time) #record the traing time of each epoch
CNN_LSTM_model=load_model("2D_CNN5_LSTM_checkpoint(F1)_sss_%s.h5" % i, custom_objects={'f1': f1})
#model operation and timing
start=timeit.default_timer()
y_pred=CNN_LSTM_model.predict(X_test)
stop=timeit.default_timer()
oper_time=stop-start
oper_time_record.append(oper_time)
#check the model test result
y_pred=CNN_LSTM_model.predict(X_test)
y_pred = np.argmax(y_pred, axis=1)
Y_test=np.argmax(y_test, axis=1)
acc_score.append(accuracy_score(Y_test, y_pred)) # Evaluation of accuracy
f_score.append(f1_score(Y_test, y_pred,average='macro')) # Evaluation of F1 score
print("This is the", i+1, "out of ",5, "Shuffle")
i+=1
del model #delete the model for retrain the neural network from scrach, instead of starting from trained model
# record performance
performance=pd.DataFrame(columns=['Acc_score','Macro_Fscore','Average_Epoch','Average_Run'])
performance['Acc_score']=acc_score
performance['Macro_Fscore']=f_score
performance['Average_Epoch']=average(eopch_time_record)
performance['Average_Run']=average(oper_time_record)
performance.to_csv("2DConv5LSTM_Performance_sss_test.csv")
```
## Baseline LSTM Model Development
```
acc_score=list()
f_score=list()
eopch_time_record=list()
oper_time_record=list()
#loading data
buffer = np.float64(preprocessing.scale(genfromtxt('S3_X.csv', delimiter=',')))
x_data=buffer.reshape(-1,40,30,1)
x_data=win_seg(x_data,40,0.5) # data segmentation with 0.5 overlap
x_data=x_data.reshape(x_data.shape[0],x_data.shape[1],x_data.shape[2]) #reshape the dataset as LSTM input shape
#majority vote on training label
buffer = np.float64(genfromtxt('S3_Y.csv', delimiter=','))-1 #0 based index
y_data=lab_vote(buffer,40)
i=0
for train_index, test_index in SRS.split(x_data,y_data):
X_train, X_test = x_data[train_index], x_data[test_index]
y_train, y_test = y_data[train_index], y_data[test_index]
#split the train data into training (training the model) and validation (tuning hypeparameters) by 8:2
X_training, X_validation, y_training, y_validation = train_test_split(X_train, y_train, test_size=0.20)
#setup model parameters
data_dim = X_train.shape[2] #y of figure
timesteps = X_train.shape[1] #x of figure
num_classes = y_train.shape[1]
batchsize=300
epcoh=300
#Build Model
model = Sequential()
model.add(LSTM(128, return_sequences=True, input_shape=(timesteps, data_dim))) # returns a sequence of vectors of dimension 64
model.add(Dropout(0.5)) #add dropout layers for controlling overfitting
model.add(LSTM(128)) # return a single vector of dimension 64
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),metrics=['accuracy',f1])
checkpointer = ModelCheckpoint(filepath='LSTM_checkpoint(F1)_sss_%s.h5' % i, monitor='val_f1',verbose=1,mode='max', save_best_only=True)
time_callback = TimeHistory() #record the model training time for each epoch
callbacks_list = [checkpointer,time_callback]
model.fit(X_training, y_training,
batch_size=batchsize, epochs=epcoh,callbacks=callbacks_list,
validation_data=(X_validation, y_validation))
eopch_time=time_callback.times
eopch_time_record.append(eopch_time) #record the traing time of each epoch
LSTM_model=load_model('LSTM_checkpoint(F1)_sss_%s.h5' % i,custom_objects={'f1': f1})
#model operation and timing
start=timeit.default_timer()
y_pred=LSTM_model.predict(X_test)
stop=timeit.default_timer()
oper_time=stop-start
oper_time_record.append(oper_time)
#check the model test result
y_pred = np.argmax(y_pred, axis=1)
Y_test=np.argmax(y_test, axis=1)
acc_score.append(accuracy_score(Y_test, y_pred))
f_score.append(f1_score(Y_test, y_pred,average='macro'))
print("This is the", i+1, "out of ",5, "Shuffle")
del model #delete the model for retrain the neural network from scrach, instead of starting from trained model
i+=1
# record performance
performance=pd.DataFrame(columns=['Acc_score','Macro_Fscore','Average_Epoch','Average_Run'])
performance['Acc_score']=acc_score
performance['Macro_Fscore']=f_score
performance['Average_Epoch']=average(eopch_time_record)
performance['Average_Run']=average(oper_time_record)
performance.to_csv("LSTM_Performance_sss_test.csv")
```
## Baseline CNN Model
```
acc_score=list()
f_score=list()
eopch_time_record=list()
oper_time_record=list()
i=0
for train_index, test_index in SRS.split(x_data,y_data):
X_train, X_test = x_data[train_index], x_data[test_index]
y_train, y_test = y_data[train_index], y_data[test_index]
#split the train data into training (training the model) and validation (tuning hypeparameters) by 8:2
X_training, X_validation, y_training, y_validation = train_test_split(X_train, y_train, test_size=0.20)
#setup model parameters
data_dim = X_train.shape[2] #y of figure
timesteps = X_train.shape[1] #x of figure
num_classes = y_train.shape[1]
batchsize=300
epcoh=300
#Build Model
model = Sequential()
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh',input_shape=(timesteps, data_dim,1)))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Conv2D(64, kernel_size=(5, 30), strides=(1, 1),padding='same',
activation='tanh'))
model.add(Flatten())
model.add(Dropout(0.5)) #add dropout layers for controlling overfitting
model.add(Dense(128, activation='tanh'))
model.add(Dropout(0.5)) #add dropout layers for controlling overfitting
model.add(Dense(128, activation='tanh'))
model.add(Dense(num_classes, activation='softmax'))#second flat fully connected layer for softmatrix (classification)
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),metrics=['accuracy',f1])
checkpointer = ModelCheckpoint(filepath='2D_CNN_checkpoint(F1)_sss_%s.h5' % i, monitor='val_f1',mode='max',verbose=1, save_best_only=True)
time_callback = TimeHistory() #record the model training time for each epoch
callbacks_list = [checkpointer,time_callback]
model.fit(X_training, y_training,
batch_size=batchsize, epochs=epcoh,callbacks=callbacks_list,
validation_data=(X_validation, y_validation))
eopch_time=time_callback.times
eopch_time_record.append(eopch_time) #record the traingtime of each epoch
CNN_model=load_model('2D_CNN_checkpoint(F1)_sss_%s.h5' % i, custom_objects={'f1': f1})
#model operation and timing
start=timeit.default_timer()
y_pred=CNN_model.predict(X_test)
stop=timeit.default_timer()
oper_time=stop-start
oper_time_record.append(oper_time)
#check the model test result
y_pred = np.argmax(y_pred, axis=1)
Y_test=np.argmax(y_test, axis=1)
acc_score.append(accuracy_score(Y_test, y_pred))
f_score.append(f1_score(Y_test, y_pred,average='macro'))
print("This is the", i+1, "out of ",5, "Shuffle")
del model #delete the model for retrain the neural network from scrach, instead of starting from trained model
i+=1
# record performance
import pandas as pd
performance=pd.DataFrame(columns=['Acc_score','Macro_Fscore','Average_Epoch','Average_Run'])
performance['Acc_score']=acc_score
performance['Macro_Fscore']=f_score
performance['Average_Epoch']=average(eopch_time_record)
performance['Average_Run']=average(oper_time_record)
performance.to_csv("2DConv_Performance_sss_test.csv")
```
# Benchmark Machine Learing-based Model Development
## Packages Preparation
```
import os
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import classification_report,f1_score,accuracy_score
import timeit
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score, f1_score, precision_score, recall_score
import matplotlib.pyplot as plt
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import RFECV
from sklearn.datasets import make_classification
```
## Functions Preparation
```
def win_seg(data,windowsize,overlap):#function for overlap segmentation
length=int((data.shape[0]*data.shape[1]-windowsize)/(windowsize*overlap)+1)
newdata=np.empty((length,windowsize, data.shape[2],1))
data_dim=data.shape[2]
layers=data.shape[3]
data=data.reshape(-1,data_dim,layers)
for i in range(0,length) :
start=int(i*windowsize*overlap)
end=int(start+windowsize)
newdata[i]=data[start:end]
return newdata
def lab_vote(data,windowsize):
y_data=data.reshape(-1,windowsize,1,1)
y_data=win_seg(y_data,windowsize,0.5)
y_data=y_data.reshape(y_data.shape[0],y_data.shape[1],y_data.shape[2])
y_data=stats.mode(y_data,axis=1)
y_data=y_data.mode
y_data=y_data.reshape(-1,1)
y_data=np.float64(keras.utils.to_categorical(y_data))
return y_data
def lab_vote_cat(data,windowsize): # non one-hot coding
y_data=data.reshape(-1,windowsize,1,1)
y_data=win_seg(y_data,windowsize,0.5)
y_data=y_data.reshape(y_data.shape[0],y_data.shape[1],y_data.shape[2])
y_data=stats.mode(y_data,axis=1)
y_data=y_data.mode
y_data=y_data.reshape(-1,1)
return y_data
def preparation(dataset):
x_data=preprocessing.scale(pd.read_csv(dataset).iloc[:,1:]) #Column-wise normalization
y_data=pd.read_csv(dataset).iloc[:,0]
X_train, X_test, y_train, y_test = train_test_split(x_data, y_data, test_size=0.20, random_state=42)#split the data into train and test by 8:2
return X_train, X_test, x_data,y_train, y_test, y_data
def TrainModels(X_train, X_test, y_train, y_test):
# Time cost
train_time=[]
run_time=[]
#SVM
svm=SVC(gamma='auto',random_state=42)
start = timeit.default_timer()
svm.fit(X_train,y_train)
stop = timeit.default_timer()
train_time.append(stop - start)
start = timeit.default_timer()
svm_pre=pd.DataFrame(data=svm.predict(X_test))
stop = timeit.default_timer()
run_time.append(stop - start)
#Naive Bayes
nb=GaussianNB()
start = timeit.default_timer()
nb.fit(X_train,y_train)
stop = timeit.default_timer()
train_time.append(stop - start)
start = timeit.default_timer()
nb_pre=pd.DataFrame(data=nb.predict(X_test))
stop = timeit.default_timer()
run_time.append(stop - start)
#KNN
knn=KNeighborsClassifier(n_neighbors=7) # based on a simple grid search
start = timeit.default_timer()
knn.fit(X_train,y_train)
stop = timeit.default_timer()
train_time.append(stop - start)
start = timeit.default_timer()
knn_pre=pd.DataFrame(data=knn.predict(X_test))
stop = timeit.default_timer()
run_time.append(stop - start)
#Decision Tree
dt=DecisionTreeClassifier(random_state=42)
start = timeit.default_timer()
dt.fit(X_train,y_train)
stop = timeit.default_timer()
train_time.append(stop - start)
start = timeit.default_timer()
dt_pre= pd.DataFrame(data=dt.predict(X_test))
stop = timeit.default_timer()
run_time.append(stop - start)
#Random Forest
rf=RandomForestClassifier(n_estimators=100)
start = timeit.default_timer()
rf.fit(X_train,y_train)
stop = timeit.default_timer()
train_time.append(stop - start)
start = timeit.default_timer()
rf_pre=pd.DataFrame(data=rf.predict(X_test))
stop = timeit.default_timer()
run_time.append(stop - start)
report = pd.DataFrame(columns=['Models','Accuracy','Macro F1','Micro F1','Train Time','Run Time'])
report['Models']=modelnames
for i in range(len(result.columns)):
report.iloc[i,1]=accuracy_score(y_test, result.iloc[:,i])
report.iloc[i,2]=f1_score(y_test, result.iloc[:,i],average='macro')
report.iloc[i,3]=f1_score(y_test, result.iloc[:,i],average='micro')
if i<len(train_time):
report.iloc[i,4]=train_time[i]
report.iloc[i,5]=run_time[i]
return report
```
## Sliding Window Segmentation
```
#loading the training and testing data
os.chdir("...") #changing working directory
buffer = np.float64(genfromtxt('S3_X.csv', delimiter=','))
x_data=buffer.reshape(-1,40,30,1)
x_data=win_seg(x_data,40,0.5) # data segmentation with 0.5 overlap
x_data=x_data.reshape(-1,40,30)
x_data_pd=x_data.reshape(-1,30)
x_data_pd = pd.DataFrame(data=x_data_pd)
adj_win=[i//40+1 for i in range(len(x_data_pd.iloc[:,0]))]
x_data_pd["adjwin"]=adj_win
x_data_pd.to_csv("S3_X_ML.csv")
#majority vote on training label
buffer = np.float64(genfromtxt('S3_Y.csv', delimiter=',')) #0 based index
y_data=lab_vote(buffer,40)
y_data2=lab_vote_cat(buffer,40) # for stratification purposes
y_data_pd = pd.DataFrame(data=y_data2)
y_data_pd.to_csv("S3_Y_ML.csv")
```
## Feature Selection Using Recursive Feature Elimination
```
X, y = X_train, y_train
svc = SVC(kernel="linear")
rfecv = RFECV(estimator=svc, step=1, cv=StratifiedKFold(10),scoring='f1_macro')
rfecv.fit(X, y)
print("Optimal number of features : %d" % rfecv.n_features_)
#plot number of features VS. cross-validation scores
plt.figure()
plt.xlabel("Number of features selected")
plt.ylabel("Cross validation score (nb of correct classifications)")
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_)
plt.show()
# Export the best features
sel_features=pd.DataFrame()
sel_features["label"]=y_test
fullfeatures=pd.read_csv("fullfeatures.csv")
names=list(fullfeatures.columns.values)[1:]
for index, val in enumerate(list(rfecv.support_)):
if val:
sel_features=pd.concat([sel_features,fullfeatures.iloc[:,index+1]],axis=1)
sel_features.to_csv("S3_Dataset_ML_SelectetedFeatures.csv")
```
## Test on Selected Features
```
X_train, X_test, X_data,y_train, y_test, y_data=preparation("S3_Dataset_ML_SelectetedFeatures.csv")
sf = ShuffleSplit(n_splits=5, test_size=0.1, random_state=42) # Random Shuffle
SRS = StratifiedShuffleSplit(n_splits=5, test_size=0.1, random_state=42) # Stratified Shuffle
finalreport = pd.DataFrame(columns=['Models','Accuracy','Macro F1','Micro F1','Train Time','Run Time'])
for train_index, test_index in SRS.split(X_data, y_data):
X_train, X_test = X_data[train_index], X_data[test_index]
y_train, y_test = y_data[train_index], y_data[test_index]
finalreport=finalreport.append(TrainModels(X_train, X_test, y_train, y_test))
finalreport.to_csv("S3_Dataset_ML_SelectetedFeatures_Evalucation.csv")
```
| github_jupyter |
<h2>Cheat sheet for numpy/scipy factorizations and operations on sparse matrices</h2>
Python's API for manipulating sparse matrices is not as well designed as Matlab's.
In Matlab, you can do (almost) anything to a sparse matrix with the same syntax
as a dense matrix, or any mixture of dense and sparse. In numpy/scipy, you often
have to use different syntax for sparse matrices. Here is my own cheat sheet for
how to operations that involve sparse matrices in numpy/scipy.
This is also a cheat sheet for some of the dense matrix factorizations in scipy, namely LU, Cholesky, and QR.
```
# These are the standard imports for CS 111.
# This list may change as the quarter goes on.
import os
import math
import numpy as np
import numpy.linalg as npla
import scipy as sp
import scipy.sparse.linalg as spla
from scipy import sparse
from scipy import linalg
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits.mplot3d import axes3d
%matplotlib tk
# create sparse from list of triples
triples = [
(0, 0, 4.0),
(0, 1, -1.0),
(0, 2, -1.0),
(1, 0, -1.0),
(1, 1, 4.0),
(1, 3, -1.0),
(2, 0, -1.0),
(2, 2, 4.0),
(2, 3, -1.0),
(3, 1, -1.0),
(3, 2,-1.0),
(3, 3, 4.0)
]
rownum = [t[0] for t in triples]
colnum = [t[1] for t in triples]
values = [t[2] for t in triples]
nrows = 4
ncols = 4
A = sparse.csr_matrix((values, (rownum, colnum)), shape = (nrows, ncols))
print('\nA:'); print(A)
# sparse to dense
Ad = A.todense()
print('\nAd:'); print(Ad)
# dense to sparse
As = sparse.csr_matrix(Ad)
print('\nAs:'); print(As)
# sparse matrix times dense vector
v = np.array(range(4))
print('\nv:', v)
w = As.dot(v)
print('\nw with As.dot:', w)
w = A @ v
print('\nw with A @ v :', w)
w = Ad @ v
print('\nw with Ad @ v:', w)
# sparse matrix times sparse matrix
Bs = As @ As
print('\nAs @ As:'); print(Bs)
Bd = Ad @ Ad
print('\nAd @ Ad:'); print(Bd)
Bdiff = Bs - Bd
print('\ndiff:'); print(Bdiff)
norm_diff = npla.norm(Bdiff)
print('\nnorm_diff:', norm_diff)
# sparse transpose
Ast = As.T
print('\nAs.T:'); print(Ast)
Adt = Ad.T
print('\nAd.T:'); print(Adt)
norm_diff = npla.norm(Adt - Ast)
print('\nnorm_diff:', norm_diff)
# indexing sparse matrix
print('\nAs[2,3]:', As[2,3])
print('\nAs[2,:]:'); print(As[2,:])
print('\nAs[:2,1:]:'); print(As[:2,1:])
# dense Ax = b solver
x = npla.solve(Ad,v)
print('\nrhs :', v)
print('\nx :', x)
print('\nA @ x:', Ad @ x)
print('\nrelative residual norm:', npla.norm(v - Ad @ x) / npla.norm(v))
# sparse Ax = b solver
x = spla.spsolve(As,v)
print('\nrhs :', v)
print('\nx:', x)
print('\nA @ x:', As @ x)
print('\nrelative residual norm:', npla.norm(v - As @ x) / npla.norm(v))
# dense least squares solver
B = np.round(10*np.random.rand(6,4))
print('\nB:'); print(B)
b = np.random.rand(6)
solution = npla.lstsq(B, b, rcond = None)
x = solution[0]
print('\nrhs :', b)
print('\nx :', x)
print('\nB @ x:', B @ x)
print('\nrelative residual norm:', npla.norm(b - B @ x) / npla.norm(b))
# dense LU factorization
B = np.round(10*np.random.rand(4,4))
print('\nB:'); print(B)
P, L, U = linalg.lu(B)
print('\nP:'); print(P)
print('\nL:'); print(L)
print('\nU:'); print(U)
norm_diff = npla.norm(B - P @ L @ U)
print('\nnorm_diff:', norm_diff)
# dense Cholesky factorization
L = linalg.cholesky(Ad, lower = True) # omit second parameter to get upper triangular factor
print('\nL:'); print(L)
print('\nL @ L.T:'); print(L @ L.T)
print('\nAd:'); print(Ad)
print('\nnorm_diff:', npla.norm(L @ L.T - Ad))
# dense QR factorization
print('\nB:'); print(B)
Q,R = linalg.qr(B)
print('\nQ:'); print(Q)
print('\nQ @ Q.t:'); print(Q @ Q.T)
print('\nR:'); print(R)
print('\nQ @ R:'); print(Q @ R)
print('\nnorm_diff:', npla.norm(Q @ R - B))
# sparse LU factorization
print('\nB:'); print(B)
Bs = sparse.csc_matrix(B)
print('\nBs:'); print(Bs)
lu = spla.splu(Bs)
print('\nL:'); print(lu.L)
print('\nU:'); print(lu.U)
print('\nperm_r:', lu.perm_r)
print('\nperm_c:', lu.perm_c)
# sparse LU factorization of large temperature matrix
AA = cs111.make_A(100)
print('\nA dimensions, nonzeros:', AA.shape, AA.size)
AA = sparse.csc_matrix(AA)
lu = spla.splu(AA)
print('\nL dimensions, nonzeros:', lu.L.shape, lu.L.size)
#plt.spy(lu.L)
# sparse Cholesky factorization (hard to do, there's a python wrapper for cholmod somewhere)
```
| github_jupyter |
## Classification - Before and After MMLSpark
### 1. Introduction
<p><img src="https://images-na.ssl-images-amazon.com/images/G/01/img16/books/bookstore/landing-page/1000638_books_landing-page_bookstore-photo-01.jpg" style="width: 500px;" title="Image from https://images-na.ssl-images-amazon.com/images/G/01/img16/books/bookstore/landing-page/1000638_books_landing-page_bookstore-photo-01.jpg" /><br /></p>
In this tutorial, we perform the same classification task in two
different ways: once using plain **`pyspark`** and once using the
**`mmlspark`** library. The two methods yield the same performance,
but one of the two libraries is drastically simpler to use and iterate
on (can you guess which one?).
The task is simple: Predict whether a user's review of a book sold on
Amazon is good (rating > 3) or bad based on the text of the review. We
accomplish this by training LogisticRegression learners with different
hyperparameters and choosing the best model.
### 2. Read the data
We download and read in the data. We show a sample below:
```
rawData = spark.read.parquet("wasbs://publicwasb@mmlspark.blob.core.windows.net/BookReviewsFromAmazon10K.parquet")
rawData.show(5)
```
### 3. Extract more features and process data
Real data however is more complex than the above dataset. It is common
for a dataset to have features of multiple types: text, numeric,
categorical. To illustrate how difficult it is to work with these
datasets, we add two numerical features to the dataset: the **word
count** of the review and the **mean word length**.
```
from pyspark.sql.functions import udf
from pyspark.sql.types import *
def wordCount(s):
return len(s.split())
def wordLength(s):
import numpy as np
ss = [len(w) for w in s.split()]
return round(float(np.mean(ss)), 2)
wordLengthUDF = udf(wordLength, DoubleType())
wordCountUDF = udf(wordCount, IntegerType())
from mmlspark.stages import UDFTransformer
wordLength = "wordLength"
wordCount = "wordCount"
wordLengthTransformer = UDFTransformer(inputCol="text", outputCol=wordLength, udf=wordLengthUDF)
wordCountTransformer = UDFTransformer(inputCol="text", outputCol=wordCount, udf=wordCountUDF)
from pyspark.ml import Pipeline
data = Pipeline(stages=[wordLengthTransformer, wordCountTransformer]) \
.fit(rawData).transform(rawData) \
.withColumn("label", rawData["rating"] > 3).drop("rating")
data.show(5)
```
### 4a. Classify using pyspark
To choose the best LogisticRegression classifier using the `pyspark`
library, need to *explictly* perform the following steps:
1. Process the features:
* Tokenize the text column
* Hash the tokenized column into a vector using hashing
* Merge the numeric features with the vector in the step above
2. Process the label column: cast it into the proper type.
3. Train multiple LogisticRegression algorithms on the `train` dataset
with different hyperparameters
4. Compute the area under the ROC curve for each of the trained models
and select the model with the highest metric as computed on the
`test` dataset
5. Evaluate the best model on the `validation` set
As you can see below, there is a lot of work involved and a lot of
steps where something can go wrong!
```
from pyspark.ml.feature import Tokenizer, HashingTF
from pyspark.ml.feature import VectorAssembler
# Featurize text column
tokenizer = Tokenizer(inputCol="text", outputCol="tokenizedText")
numFeatures = 10000
hashingScheme = HashingTF(inputCol="tokenizedText",
outputCol="TextFeatures",
numFeatures=numFeatures)
tokenizedData = tokenizer.transform(data)
featurizedData = hashingScheme.transform(tokenizedData)
# Merge text and numeric features in one feature column
featureColumnsArray = ["TextFeatures", "wordCount", "wordLength"]
assembler = VectorAssembler(
inputCols = featureColumnsArray,
outputCol="features")
assembledData = assembler.transform(featurizedData)
# Select only columns of interest
# Convert rating column from boolean to int
processedData = assembledData \
.select("label", "features") \
.withColumn("label", assembledData.label.cast(IntegerType()))
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.ml.classification import LogisticRegression
# Prepare data for learning
train, test, validation = processedData.randomSplit([0.60, 0.20, 0.20], seed=123)
# Train the models on the 'train' data
lrHyperParams = [0.05, 0.1, 0.2, 0.4]
logisticRegressions = [LogisticRegression(regParam = hyperParam)
for hyperParam in lrHyperParams]
evaluator = BinaryClassificationEvaluator(rawPredictionCol="rawPrediction",
metricName="areaUnderROC")
metrics = []
models = []
# Select the best model
for learner in logisticRegressions:
model = learner.fit(train)
models.append(model)
scoredData = model.transform(test)
metrics.append(evaluator.evaluate(scoredData))
bestMetric = max(metrics)
bestModel = models[metrics.index(bestMetric)]
# Get AUC on the validation dataset
scoredVal = bestModel.transform(validation)
print(evaluator.evaluate(scoredVal))
```
### 4b. Classify using mmlspark
Life is a lot simpler when using `mmlspark`!
1. The **`TrainClassifier`** Estimator featurizes the data internally,
as long as the columns selected in the `train`, `test`, `validation`
dataset represent the features
2. The **`FindBestModel`** Estimator find the best model from a pool of
trained models by find the model which performs best on the `test`
dataset given the specified metric
3. The **`CompueModelStatistics`** Transformer computes the different
metrics on a scored dataset (in our case, the `validation` dataset)
at the same time
```
from mmlspark.train import TrainClassifier, ComputeModelStatistics
from mmlspark.automl import FindBestModel
# Prepare data for learning
train, test, validation = data.randomSplit([0.60, 0.20, 0.20], seed=123)
# Train the models on the 'train' data
lrHyperParams = [0.05, 0.1, 0.2, 0.4]
logisticRegressions = [LogisticRegression(regParam = hyperParam)
for hyperParam in lrHyperParams]
lrmodels = [TrainClassifier(model=lrm, labelCol="label", numFeatures=10000).fit(train)
for lrm in logisticRegressions]
# Select the best model
bestModel = FindBestModel(evaluationMetric="AUC", models=lrmodels).fit(test)
# Get AUC on the validation dataset
predictions = bestModel.transform(validation)
metrics = ComputeModelStatistics().transform(predictions)
print("Best model's AUC on validation set = "
+ "{0:.2f}%".format(metrics.first()["AUC"] * 100))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/HknyYtbz/cng-562/blob/master/YATBAZ_malaria_cell_images.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Data Loading
```
import numpy as np
import pandas as pd
```
Please upload your Kaggle API JSON named as kaggle.json to download the data
```
from google.colab import files
#Kaggle api key upload
files.upload()
#Essential downloads for the project, especially the kaggle dataset download
!pip install -U -q kaggle
!mkdir -p ~/.kaggle
!cp kaggle.json ~/.kaggle/
!chmod 600 /root/.kaggle/kaggle.json
!kaggle datasets download -d iarunava/cell-images-for-detecting-malaria
!ls
import zipfile
zip_ref = zipfile.ZipFile("/content/cell-images-for-detecting-malaria.zip", 'r')
zip_ref.extractall("/content/")
zip_ref.close()
import torch
import numpy as np
from PIL import Image
# check if CUDA is available
train_on_gpu = torch.cuda.is_available()
device = None
if not train_on_gpu:
print('CUDA is not available. Training on CPU ...')
device = "cpu"
else:
print('CUDA is available! Training on GPU ...')
device = "cuda"
```
#Preprocessing
##Image to Tensor Conversion & Train-Test Split
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
test_size = 0.2
train_size= 0.8
# convert data to a normalized torch.FloatTensor
transform = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
#transforms.RandomRotation(10),
transforms.Resize((108,108)),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
# choose the training and test datasets
dataset = datasets.ImageFolder("/content/cell_images", transform=transform)
dataloader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, shuffle=True)
# obtain training indices that will be used for validation
num_data = len(dataset)
indices = list(range(num_data))
np.random.shuffle(indices)
split = int(np.floor(test_size * num_data))
train_idx1, test_idx = indices[split:], indices[:split]
num_train_data = len(dataset) - split
split2 = int(np.floor(valid_size * num_train_data))
train_idx, valid_idx = train_idx1[split2:], train_idx1[:split2]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
test_sampler = SubsetRandomSampler(test_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders (combine dataset and sampler)
train_loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, sampler=train_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, sampler=test_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, sampler=valid_sampler, num_workers=num_workers)
# specify the image classes
classes = ['Parasitized','Uninfected']
```
## Data Visualization
```
import matplotlib.pyplot as plt
%matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0))) # convert from Tensor image
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
# display 20 images
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title(classes[labels[idx]])
```
#CNN Model
```
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# convolutional layer (sees 360x360x3 image tensor)
self.conv1 = nn.Conv2d(3, 32, 3, padding=1)
# convolutional layer (sees 16x16x16 tensor)
self.conv2 = nn.Conv2d(32, 32, 3, padding=1)
# convolutional layer (sees 8x8x32 tensor)
self.conv3 = nn.Conv2d(32, 32, 3, padding=1)
# max pooling layer
self.pool = nn.MaxPool2d(2, 2)
# linear layer (64 * 12 * 12 -> 4096)
self.fc1 = nn.Linear(32*13*13, 512)
# linear layer (500 -> 10)
self.fc2 = nn.Linear(512,256)
self.fc3 = nn.Linear(256,2)
# dropout layer (p=0.25)
self.dropout = nn.Dropout(0.25)
def forward(self, x):
# add sequence of convolutional and max pooling layers
#print(x.shape)
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
#print(x.shape)
# flatten image input
x = x.view(-1, 32 * 13 * 13)
#print(x.shape)
# add dropout layer
# add 1st hidden layer, with relu activation function
x = self.dropout(F.relu(self.fc1(x)))
# add dropout layer
# add 2nd hidden layer, with relu activation function
x = self.dropout(F.relu(self.fc2(x)))
x = self.dropout(self.fc3(x))
return x
# create a complete CNN
model = Net()
print(model)
# move tensors to GPU if CUDA is available
if train_on_gpu:
model.cuda()
import torch.optim as optim
# specify loss function (categorical cross-entropy)
criterion = nn.CrossEntropyLoss()
# specify optimizer
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
##Training Part
```
n_epochs = 30
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval()
for batch_idx, (data, target) in enumerate(valid_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), "/content/malaria_model.pt")
valid_loss_min = valid_loss
```
##Testing Part
```
model.load_state_dict(torch.load("/content/malaria_model.pt"))
test_loss = 0.0
class_correct = list(0. for i in range(len(classes)))
class_total = list(0. for i in range(len(classes)))
model.eval()
# iterate over test data
for batch_idx, (data, target) in enumerate(test_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
ct = 0
for i in range(len(list(target.data)) ):
try:
label = target.data[i]
#print(correct)
class_correct[label] += correct[i].item()
class_total[label] += 1
except IndexError:
ct+=1
continue
#print("Problematic Count: %d"%ct)
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(len(classes)):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
#Transfer Learning
##DenseNet
```
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
model2 =models.densenet121(pretrained=True)
print(model2)
for param in model2.parameters():
param.requires_grad = False
from collections import OrderedDict
classifier = nn.Sequential(
OrderedDict([ ('fc1', nn.Linear(1024, 500)),
('relu', nn.ReLU()),
('fc2', nn.Linear(500, 2)),
('output', nn.LogSoftmax(dim=1))
])
)
model2.classifier = classifier
print(model2)
optimizer = optim.Adam(model2.classifier.parameters(), lr=0.003)
criterion = nn.NLLLoss()
model2.to(device)
```
###Training
```
n_epochs = 30
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model2.train()
for batch_idx, (data, target) in enumerate(train_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model2(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model2.eval()
for batch_idx, (data, target) in enumerate(valid_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model2(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model2.state_dict(), "/content/malaria_modelDense.pt")
valid_loss_min = valid_loss
```
### Testing
```
model2.load_state_dict(torch.load("/content/malaria_modelDense.pt"))
test_loss = 0.0
class_correct = list(0. for i in range(len(classes)))
class_total = list(0. for i in range(len(classes)))
model2.eval()
# iterate over test data
for batch_idx, (data, target) in enumerate(test_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model2(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
ct = 0
for i in range(len(list(target.data)) ):
try:
label = target.data[i]
#print(correct)
class_correct[label] += correct[i].item()
class_total[label] += 1
except IndexError:
ct+=1
continue
#print("Problematic Count: %d"%ct)
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(len(classes)):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
##ResNet50
```
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
model3 =models.resnet50(pretrained=True)
print(model3)
from collections import OrderedDict
classifier = nn.Sequential(OrderedDict([
("fc",nn.Linear(2048, 1000)),
("relu",nn.ReLU()),
("dropout",nn.Dropout(0.2)),
("a1",nn.Linear(1000, 512)),
("relu",nn.ReLU()),
("dropout",nn.Dropout(0.2)),
("a2",nn.Linear(512, 256)),
("relu2",nn.ReLU()),
("dropout2",nn.Dropout(0.2)),
("a3",nn.Linear(256, 2)),
("output", nn.LogSoftmax(dim=1))]))
model3.fc = classifier
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model3.fc.parameters(), lr=0.003)
model3.to(device);
print(model3)
```
### Training
```
n_epochs = 30
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model3.train()
for batch_idx, (data, target) in enumerate(train_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model3(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model3.eval()
for batch_idx, (data, target) in enumerate(valid_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model3(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model3.state_dict(), "/content/malaria_modelResNet50.pt")
valid_loss_min = valid_loss
```
###Testing
```
model3.load_state_dict(torch.load("/content/malaria_modelResNet50.pt"))
test_loss = 0.0
class_correct = list(0. for i in range(len(classes)))
class_total = list(0. for i in range(len(classes)))
model3.eval()
# iterate over test data
for batch_idx, (data, target) in enumerate(test_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model3(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
#print("Pred: %s"+str(pred))
#print("Real: %s"+str(target.data))
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
ct = 0
for i in range(len(list(target.data)) ):
try:
label = target.data[i]
#print(correct)
class_correct[label] += correct[i].item()
class_total[label] += 1
except IndexError:
ct+=1
continue
#print("Problematic Count: %d"%ct)
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(len(classes)):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
#Conclusion
In this dataset, we had many cell images labeled as infected or not infected. I simply load the data using PyTorch (with their respective label) and generated a train and test sample ( In the process I coverted images to sensors and appied some transformations to make the model more robust). Apart from that no processing done on the data.
I used my own CNN network with 3 convolutional layer and 3 fully connected layer. Convolutional layers have 32 3x3 kernels and max pooling layer between them. For training I have also generated a validation set from the training set to avoid overfitting and saved the model when the validation loss is decreased. The accuracy obtained is 95%.
Having tried the model I implemented, I have also tried pretrained DenseNet and ResNet by modifying their fully connected layers and obtained 93% and 91% respectively.
As a future work, other pretrained models can be used or different CNN architectures can be applied.
All in all, in this dataset, I had a chance to explore CNN and well-known architectures better.
| github_jupyter |
# Coordinate Descent
### Lower Bound, Take 4
Ensure feasibility of "Lower Bound, Take 2" by adjusting alpha as necessary.
### Lower Bound, Take 3
Ensure feasibility by allowing a stochastic mixture with the MLE. Doesn't work (not DCP).
Assume $r_{\min} = 0$ for simplicity. Idea for online solving $$
\begin{aligned}
&\!\min_{Q \succeq 0} &\qquad& \sum_{(w,r)} w r Q_{w, r}, \\
&\text{subject to} & & \sum_{(w,r)} w Q_{w,r} = 1, \\
& & & \sum_{(w,r)} Q_{w,r} = 1, \\
& & & \sum_n \log(Q_{w_n, r_n}) \geq \phi
\end{aligned}
$$ where $\phi = -\frac{1}{2} \chi^{2,\alpha}_{(1)} + \sum_n \log(Q^{\text{(mle)}}_{w_n, r_n})$. Because the support of $Q$ is at most the empirical support plus $(w_{\min}, 0)$ and $(w_{\max}, 0)$ we will maintain two variables $q_{\min}$ and $q_{\max}$ corresponding to $w_{\min}$ and $w_{\max}$ respectively. Otherwise we need one primal variable for each data point. However we will use two primal variables corresponding to $r = 0$ and $r = 1$ respectively. We will split each datapoint into two points whose fractional counts are proportional to $r$ and $(1 - r)$ respectively.
At time $t$ we receive $(w_t, r_t)$ and we want to determine $q_t$. We are allowed to choose the new $q_{\min}$ and $q_{\max}$ arbitrarily. For points $q_{<t}$ we are allowed to scale them by $\psi_0$ and $\psi_1$, corresponding to points with $r = 0$ and $r = 1$ respectively. We are also allowed to stochastically mix in the maximum likelihood solution according to $\psi_{\text{mle}}$. We assume $(q_{0,<t}, q_{1,<t}, q_{<t,\min}, q_{<t,\max})$ is feasible before receiving $(w_t, r_t)$. Then
$$
\begin{aligned}
&\!\min_{q_{0,t}, q_{1,t}, q_{\min}, q_{\max}, \psi_0, \psi_1 \succeq 0, \psi_{\text{mle}} \in [0, 1]} &\qquad& q_{1,t} w_t + \psi_1 v_{<t} + \psi_{\text{mle}} v_{\text{mle}} \\
&\text{subject to} & & w_t q_{0,t} + w_t q_{1,t} + w_{\min} q_{\min} + w_{\max} q_{\max} + \psi_0 w_{0,<t} q_{0,<t} + \psi_1 w_{1,<t} q_{1,<t} = 1, \\
& & & q_{0,t} + q_{1,t} + q_{\min} + q_{\max} + \psi_0 q_{0,<t} + \psi_1 q_{1,<t} = 1, \\
& & & (1 - r_t) \log(q_{0,t}) + r_t \log(q_{1,t}) + (t - r_{<t}) \log(\psi_0) + r_{<t} \log(\psi_1) \geq \phi - \mathcal{L}_{<t} = -\frac{1}{2} \chi^{2,\alpha}_{(1)} + \mathcal{L}^{\text{(mle)}}_t - \mathcal{L}_{<t}
\end{aligned}
$$ where $v_{<t}$ is the previous lower bound, $\mathcal{L}^{\text{(mle)}}_t$ is the mle likelihood of the observed data including point $t$, and $\mathcal{L}_{<t}$ is the previously obtained likelihood of the observed data for the lower bound.
### Lower Bound, Take 2
Better, but still has feasibility issues as the stream progresses.
Assume $r_{\min} = 0$ for simplicity. Idea for online solving $$
\begin{aligned}
&\!\min_{Q \succeq 0} &\qquad& \sum_{(w,r)} w r Q_{w, r}, \\
&\text{subject to} & & \sum_{(w,r)} w Q_{w,r} = 1, \\
& & & \sum_{(w,r)} Q_{w,r} = 1, \\
& & & \sum_n \log(Q_{w_n, r_n}) \geq \phi
\end{aligned}
$$ where $\phi = -\frac{1}{2} \chi^{2,\alpha}_{(1)} + \sum_n \log(Q^{\text{(mle)}}_{w_n, r_n})$. Because the support of $Q$ is at most the empirical support plus $(w_{\min}, 0)$ and $(w_{\max}, 0)$ we will maintain two variables $q_{\min}$ and $q_{\max}$ corresponding to $w_{\min}$ and $w_{\max}$ respectively. Otherwise we need one primal variable for each data point.
At time $t$ we receive $(w_t, r_t)$ and we want to determine $q_t$. We are allowed to choose the new $q_{\min}$ and $q_{\max}$ arbitrarily. For points $q_{<t}$ we are allowed to scale them by $\psi_0$ and $\psi_1$, corresponding to points with $r = 0$ and $r = 1$ respectively. We assume $(q_{0,<t}, q_{1,<t}, q_{<t,\min}, q_{<t,\max})$ is feasible before receiving $(w_t, r_t)$. Then
$$
\begin{aligned}
&\!\min_{q_t, q_{\min}, q_{\max},\psi_0, \psi_1 \succeq 0} &\qquad& q_t w_t r_t + \psi_1 v_{<t} \\
&\text{subject to} & & w_t q_t + w_{\min} q_{\min} + w_{\max} q_{\max} + \psi_0 w_{0,<t} q_{0,<t} + \psi_1 w_{1,<t} q_{1,<t} = 1, \\
& & & q_t + q_{\min} + q_{\max} + \psi_0 q_{0,<t} + \psi_1 q_{1,<t} = 1, \\
& & & \log(q_t) + (t - r_{<t}) \log(\psi_0) + r_{<t} \log(\psi_1) \geq \phi - \mathcal{L}_{<t} = -\frac{1}{2} \chi^{2,\alpha}_{(1)} + \mathcal{L}^{\text{(mle)}}_t - \mathcal{L}_{<t}
\end{aligned}
$$ where $v_{<t}$ is the previous lower bound, $\mathcal{L}^{\text{(mle)}}_t$ is the mle likelihood of the observed data including point $t$, and $\mathcal{L}_{<t}$ is the previously obtained likelihood of the observed data for the lower bound.
After computing $q_t$ we increment $q_{0,<t}$ and $q_{1,<t}$ proportional to $1 - r_t$ and $r_t$ respectively; and $w_{0,<t}$ and $w_{1,<t}$ by $w_t (1 - r_t)$ and $w_t r_t$ respectively.
### Lower Bound
Starts out good, but then runs into infeasibility.
Assume $r_{\min} = 0$ for simplicity. Idea for online solving $$
\begin{aligned}
&\!\min_{Q \succeq 0} &\qquad& \sum_{(w,r)} w r Q_{w, r}, \\
&\text{subject to} & & \sum_{(w,r)} w Q_{w,r} = 1, \\
& & & \sum_{(w,r)} Q_{w,r} = 1, \\
& & & \sum_n \log(Q_{w_n, r_n}) \geq \phi
\end{aligned}
$$ where $\phi = -\frac{1}{2} \chi^{2,\alpha}_{(1)} + \sum_n \log(Q^{\text{(mle)}}_{w_n, r_n})$. Because the support of $Q$ is at most the empirical support plus $(w_{\min}, 0)$ and $(w_{\max}, 0)$ we will maintain two variables $q_{\min}$ and $q_{\max}$ corresponding to $w_{\min}$ and $w_{\max}$ respectively. Otherwise we need one primal variable for each data point.
At time $t$ we receive $(w_t, r_t)$ and we want to determine $q_t$. We are allowed to choose the new $q_{\min}$ and $q_{\max}$ arbitrarily. For points $q_{<t}$ we only allowed to scale them by $\psi$. We assume $(q_{<t}, q_{<t,\min}, q_{<t,\max})$ is feasible before receiving $(w_t, r_t)$. Then
$$
\begin{aligned}
&\!\min_{q_t, q_{\min}, q_{\max}, \psi \succeq 0} &\qquad& q_t w_t r_t + \psi v_{<t} \\
&\text{subject to} & & w_t q_t + w_{\min} q_{\min} + w_{\max} q_{\max} + \psi w_{<t} q_{<t} = 1, \\
& & & q_t + q_{\min} + q_{\max} + \psi q_{<t} = 1, \\
& & & \log(q_t) + t \log(\psi) \geq \phi - \mathcal{L}_{<t} = -\frac{1}{2} \chi^{2,\alpha}_{(1)} + \mathcal{L}^{\text{(mle)}}_t - \mathcal{L}_{<t}
\end{aligned}
$$ where $v_{<t}$ is the previous lower bound, $\mathcal{L}^{\text{(mle)}}_t$ is the mle likelihood of the observed data including point $t$, and $\mathcal{L}_{<t}$ is the previously obtained likelihood of the observed data for the lower bound. Substituting $q_{<t} + q_{<t,\min} + q_{<t,\max} = 1$ and $w_{<t} q_{<t} + w_{\min} q_{<t,\min} + w_{\max} q_{<t,\max} = 1$ yields
$$
\begin{aligned}
&\!\min_{q_t, q_{\min}, q_{\max} \succeq 0, \psi \in [0, 1]} &\qquad& q_t w_t r_t + \psi v_{<t} \\
&\text{subject to} & & w_t q_t + w_{\min} q_{\min} + w_{\max} q_{\max} + \psi (1 - w_{\min} q_{<t,\min} - w_{\max} q_{<t,\max}) = 1, \\
& & & q_t + q_{\min} + q_{\max} + \psi (1 - q_{<t,\max} - q_{<t,\min}) = 1 \\
& & & -\log(q_t) - t \log(\psi) - \frac{1}{2} \chi^{2,\alpha}_{(1)} + \mathcal{L}^{\text{(mle)}}_t - \mathcal{L}_{<t} \leq 0
\end{aligned}
$$
### MLE
Idea for online solving $$
\begin{aligned}
&\!\max_{Q \succeq 0} &\qquad& \sum_n \log(Q_{w_n, r_n}), \\
&\text{subject to} & & \sum_{(w,r)} w Q_{w,r} = 1, \\
& & & \sum_{(w,r)} Q_{w,r} = 1.
\end{aligned}
$$
Because the support of $Q$ is at most the empirical support plus $w_{\min}$ and $w_{\max}$ we will maintain two variables $q_{\min}$ and $q_{\max}$ corresponding to $w_{\min}$ and $w_{\max}$ respectively. Otherwise we need one primal variable for each data point.
At time $t$ we receive $(w_t, r_t)$ and we want to determine $q_t$. We are allowed to choose the new $q_{\min}$ and $q_{\max}$ arbitrarily. For points $q_{<t}$ we only allowed to scale them by $\psi$. We assume $(q_{<t}, q_{<t,\min}, q_{<t,\max})$ is feasible before receiving $(w_t, r_t)$. Then
$$
\begin{aligned}
&\!\max_{q_t, q_{\min}, q_{\max}, \psi > 0} &\qquad& t \log(\psi) + \log(q_t), \\
&\text{subject to} & & w_t q_t + w_{\min} q_{\min} + w_{\max} q_{\max} + \psi w_{<t} q_{<t} = 1, \\
& & & q_t + q_{\min} + q_{\max} + \psi q_{<t} = 1
\end{aligned}
$$
Substituting $q_{<t} + q_{<t,\min} + q_{<t,\max} = 1$ and $w_{<t} q_{<t} + w_{\min} q_{<t,\min} + w_{\max} q_{<t,\max} = 1$ yields
$$
\begin{aligned}
&\!\max_{q_t, q_{\min}, q_{\max}, \psi > 0} &\qquad& t \log(\psi) + \log(q_t), \\
&\text{subject to} & & w_t q_t + w_{\min} q_{\min} + w_{\max} q_{\max} + \psi (1 - w_{\min} q_{<t,\min} - w_{\max} q_{<t,\max}) = 1, \\
& & & q_t + q_{\min} + q_{\max} + \psi (1 - q_{<t,\max} - q_{<t,\min}) = 1
\end{aligned}
$$
At the beginning of time we can initialize with $$
\begin{aligned}
q_{0,\min} &= \frac{1 - w_{\min}}{w_{\max} - w_{\min}} \\
q_{0,\max} &= \frac{w_{\max} - 1}{w_{\max} - w_{\min}}
\end{aligned}
$$
### Code
```
class OnlineCoordinateDescentMLE:
def __init__(self, wmin, wmax):
from cvxopt import matrix
assert wmax > 1
assert wmin >= 0
assert wmin < wmax
self.wmin = wmin
self.wmax = wmax
self.qmin = (wmax - 1) / (wmax - wmin)
self.qmax = (1 - wmin) / (wmax - wmin)
self.obj = 0
self.vmin = 0
self.lastphi = 0
self.G = matrix([ [ -1, 0, 0, 0 ],
[ 0, -1, 0, 0 ],
[ 0, 0, -1, 0 ],
[ 0, 0, 0, -1 ],
],
tc='d').T
self.h = matrix([ 0, 0, 0, 0 ], tc='d')
self.b = matrix([ 1 / wmax, 1 ], tc='d')
self.t = 0
def update(self, c, w, r):
from cvxopt import matrix, solvers
assert c > 0
safet = max(self.t, 1)
x0 = matrix([ c / (c + safet),
self.qmin * safet / (c + safet),
self.qmax * safet / (c + safet),
safet / (c + safet) ],
tc='d')
def F(x=None, z=None):
import math
if x is None: return 0, x0
if x[0] <= 0 or x[3] <= 0:
return None
f = -c * math.log(x[0]) / safet - self.t * math.log(x[3]) / safet
jf = matrix([ -c / (safet * x[0]), 0, 0, -self.t / (safet * x[3]) ], tc='d').T
if z is None: return f, jf
hf = z[0] * matrix([ [ (c / safet) * 1/x[0]**2, 0, 0, 0 ],
[ 0, 0, 0, 0 ],
[ 0, 0, 0, 0 ],
[ 0, 0, 0, (self.t / safet) * 1/x[3]**2 ]
], tc='d')
return f, jf, hf
A = matrix([
[ float(w) / self.wmax,
self.wmin / self.wmax,
1,
(1 / self.wmax - (self.wmin / self.wmax) * self.qmin - self.qmax) ],
[ 1, 1, 1, (1 - self.qmin - self.qmax) ]
],
tc='d')
soln = solvers.cp(F=F, G=self.G, h=self.h, A=A.T, b=self.b, options={'show_progress': False})
from pprint import pformat
assert soln['status'] == 'optimal', pformat([ soln, self.t ])
self.obj -= safet * soln['primal objective']
self.lastq = soln['x'][0]
self.qmin = soln['x'][1]
self.qmax = soln['x'][2]
self.lastphi = soln['x'][3]
self.vmin = soln['x'][0] * w * r + soln['x'][3] * self.vmin
self.t += c
return self.lastq
class OnlineCoordinateDescentLB:
class Flass:
def __init__(self):
pass
def __init__(self, wmin, wmax, alpha):
assert wmax > 1
assert wmin >= 0
assert wmin < wmax
self.wmin = wmin
self.wmax = wmax
self.qmin = (wmax - 1) / (wmax - wmin)
self.qmax = (1 - wmin) / (wmax - wmin)
self.vlb = 0
self.wq0 = 0
self.wq1 = 0
self.q0t = 0
self.q1t = 0
self.t = 0
self.rt = 0
self.llb = 0
self.alpha = alpha
self.mle = OnlineCoordinateDescentMLE(wmin=wmin, wmax=wmax)
from scipy.stats import chi2
import cvxpy as cp
self.vars = OnlineCoordinateDescentLB.Flass()
self.vars.qt = cp.Variable(nonneg=True)
self.vars.qmin = cp.Variable(nonneg=True)
self.vars.qmax = cp.Variable(nonneg=True)
self.vars.psi0 = cp.Variable(nonneg=True)
self.vars.psi1 = cp.Variable(nonneg=True)
self.params = OnlineCoordinateDescentLB.Flass()
self.params.w = cp.Parameter(nonneg=True)
self.params.wr = cp.Parameter(nonneg=True)
self.params.vlb = cp.Parameter(nonneg=True)
self.params.wq0 = cp.Parameter(nonneg=True)
self.params.wq1 = cp.Parameter(nonneg=True)
self.params.q0t = cp.Parameter(nonneg=True)
self.params.q1t = cp.Parameter(nonneg=True)
self.params.c = cp.Parameter(nonneg=True)
self.params.tminusrt = cp.Parameter(nonneg=True)
self.params.rt = cp.Parameter(nonneg=True)
self.params.constraintrhs = cp.Parameter()
self.prob = cp.Problem(cp.Minimize(self.params.wr * self.vars.qt + self.params.vlb * self.vars.psi1), [
self.params.w * self.vars.qt
+ (self.wmin / self.wmax) * self.vars.qmin
+ self.vars.qmax
+ self.params.wq0 * self.vars.psi0
+ self.params.wq1 * self.vars.psi1
== 1 / self.wmax,
self.vars.qt
+ self.vars.qmin
+ self.vars.qmax
+ self.params.q0t * self.vars.psi0
+ self.params.q1t * self.vars.psi1 == 1,
self.params.c * cp.log(self.vars.qt)
+ self.params.tminusrt * cp.log(self.vars.psi0)
+ self.params.rt * cp.log(self.vars.psi1)
>= self.params.constraintrhs
])
def innersolve(self, c, w, r, alpha):
from scipy.stats import chi2
safet = max(self.t, 1)
halfchisq = 0.5 * chi2.isf(q=alpha, df=1)
self.params.w.value = w / self.wmax
self.params.wr.value = w * r
self.params.vlb.value = self.vlb
self.params.wq0.value = self.wq0 / self.wmax
self.params.wq1.value = self.wq1 / self.wmax
self.params.q0t.value = self.q0t
self.params.q1t.value = self.q1t
self.params.c.value = c / safet
self.params.tminusrt.value = (self.t - self.rt) / safet
self.params.rt.value = self.rt / safet
self.params.constraintrhs.value = (-halfchisq + self.mle.obj - self.llb) / safet
self.prob.solve(verbose=False)
return (self.prob.value,
self.prob.status,
(1 - r) * self.vars.qt.value if self.vars.qt.value is not None else None,
r * self.vars.qt.value if self.vars.qt.value is not None else None,
self.vars.qmin.value,
self.vars.qmax.value,
self.vars.psi0.value,
self.vars.psi1.value
)
def updatev3(self, c, w, r):
import cvxpy as cp
import math
from scipy.special import xlogy
qmle = self.mle.update(c, w, r)
q0t = cp.Variable(nonneg=True)
q1t = cp.Variable(nonneg=True)
qmin = cp.Variable(nonneg=True)
qmax = cp.Variable(nonneg=True)
psi0 = cp.Variable(nonneg=True)
psi1 = cp.Variable(nonneg=True)
psimle = cp.Variable(nonneg=True)
safet = max(self.t, 1)
# prob = cp.Problem(cp.Minimize((float(w) * q1t + self.vlb * psi1) * (1 - psimle) + self.mle.vhat * psimle), [
prob = cp.Problem(cp.Minimize(float(w) * q1t + self.vlb * psi1 + self.mle.vhat * psimle), [
float(w / self.wmax) * q0t
+ float(w / self.wmax) * q1t
+ (self.wmin / self.wmax) * qmin
+ qmax
+ psi0 * (self.wq0 / self.wmax)
+ psi1 * (self.wq1 / self.wmax)
== 1 / self.wmax,
q0t + q1t + qmin + qmax + psi0 * self.q0t + psi1 * self.q1t == 1,
float(c * (1 - r) / safet) * cp.log(q0t)
+ float(c * r / safet) * cp.log(q1t)
+ float((self.t - self.rt) / safet) * cp.log(psi0)
+ float(self.rt / safet) * cp.log(psi1)
# >= ((-self.halfchisq + self.mle.obj) / safet) * cp.inv_pos(1 - psimle) - (self.llb / safet),
>= ((-self.halfchisq + self.mle.obj) / safet) * (1 + psimle) - (self.llb / safet),
psimle <= 1,
psimle >= 0.9
])
prob.solve(verbose=False)
assert prob.status[:7] == 'optimal', prob.solve(verbose=True)
self.vlb = (w * q1t.value + self.vlb * psi1.value) * (1 - psimle.value) + psimle.value * self.mle.vhat
self.qmin = qmin.value
self.qmax = qmax.value
self.llb += ( xlogy(c * (1 - r), q0t.value)
+ xlogy(c * r, q1t.value)
+ xlogy(self.t - self.rt, psi0.value)
+ xlogy(self.rt, psi1.value)
)
self.llb = (1 - psimle.value) * self.llb + psimle.value * self.mle.obj
self.t += c
self.rt += c * r
self.q0t = q0t.value + psi0.value * self.q0t
self.wq0 = w * q0t.value + psi0.value * self.wq0
self.q1t = q1t.value + psi1.value * self.q1t
self.wq1 = w * q1t.value + psi1.value * self.wq1
myq = (1 - r) * q0t.value + r * q1t.value
myq = (1 - psimle.value) * myq + psimle.value * qmle
return myq, { 'self.q0t': self.q0t,
'self.q1t': self.q1t,
'qmin': qmin.value,
'qmax': qmax.value,
'vlb': self.vlb,
'llb': self.llb,
'vmle': self.mle.vhat,
'self.rt': self.rt,
'psimle': psimle.value,
}
def __initturg__(self, wmin, wmax, alpha):
assert wmax > 1
assert wmin >= 0
assert wmin < wmax
self.wmin = wmin
self.wmax = wmax
self.qmin = (wmax - 1) / (wmax - wmin)
self.qmax = (1 - wmin) / (wmax - wmin)
self.vlb = 0
self.wq0 = 0
self.wq1 = 0
self.q0t = 0
self.q1t = 0
self.t = 0
self.rt = 0
self.llb = 0
self.alpha = alpha
self.mle = OnlineCoordinateDescentMLE(wmin=wmin, wmax=wmax)
from scipy.stats import chi2
import cvxpy as cp
self.vars = OnlineCoordinateDescentLB.Flass()
self.vars.q0t = cp.Variable(nonneg=True)
self.vars.q1t = cp.Variable(nonneg=True)
self.vars.qmin = cp.Variable(nonneg=True)
self.vars.qmax = cp.Variable(nonneg=True)
self.vars.psi0 = cp.Variable(nonneg=True)
self.vars.psi1 = cp.Variable(nonneg=True)
self.params = OnlineCoordinateDescentLB.Flass()
self.params.w = cp.Parameter(nonneg=True)
self.params.wcost = cp.Parameter(nonneg=True)
self.params.vlb = cp.Parameter(nonneg=True)
self.params.wq0 = cp.Parameter(nonneg=True)
self.params.wq1 = cp.Parameter(nonneg=True)
self.params.q0t = cp.Parameter(nonneg=True)
self.params.q1t = cp.Parameter(nonneg=True)
self.params.coneminusr = cp.Parameter(nonneg=True)
self.params.cr = cp.Parameter(nonneg=True)
self.params.tminusrt = cp.Parameter(nonneg=True)
self.params.rt = cp.Parameter(nonneg=True)
self.params.constraintrhs = cp.Parameter()
self.prob = cp.Problem(cp.Minimize(self.params.wcost * self.vars.q1t + self.params.vlb * self.vars.psi1), [
self.params.w * self.vars.q0t
+ self.params.w * self.vars.q1t
+ (self.wmin / self.wmax) * self.vars.qmin
+ self.vars.qmax
+ self.params.wq0 * self.vars.psi0
+ self.params.wq1 * self.vars.psi1
== 1 / self.wmax,
self.vars.q0t
+ self.vars.q1t
+ self.vars.qmin
+ self.vars.qmax
+ self.params.q0t * self.vars.psi0
+ self.params.q1t * self.vars.psi1 == 1,
self.params.coneminusr * cp.log(self.vars.q0t)
+ self.params.cr * cp.log(self.vars.q1t)
+ self.params.tminusrt * cp.log(self.vars.psi0)
+ self.params.rt * cp.log(self.vars.psi1)
>= self.params.constraintrhs
])
def innersolveflass(self, c, w, r, alpha):
# doesn't work, not sure why (?)
from cvxopt import matrix, spdiag, solvers
import numpy as np
from scipy.stats import chi2
assert 0 < c
assert 0 <= r
assert r <= 1
safet = max(self.t, 1)
halfchisq = 0.5 * chi2.isf(q=alpha, df=1)
print([
w / self.wmax,
w,
self.vlb,
self.wq0 / self.wmax,
self.wq1 / self.wmax,
self.q0t,
self.q1t,
c * (1 - r) / safet,
c * r / safet,
(self.t - self.rt) / safet,
self.rt / safet,
(-halfchisq + self.mle.obj - self.llb) / safet
])
G = matrix(-np.eye(6), tc='d')
h = matrix(0, size=(6,1), tc='d')
A = matrix([ [ w / self.wmax,
w / self.wmax,
self.wmin / self.wmax,
1,
self.wq0 / self.wmax,
self.wq1 / self.wmax
],
[ 1, 1, 1, 1, self.q0t, self.q1t ]
],
tc='d')
b = matrix([ 1 / self.wmax, 1 ], tc='d')
cost = matrix(0, size=(6,1), tc='d')
cost[1] = float(w)
cost[5] = self.vlb
x0 = matrix([ c * (1 - r) / (c + safet),
c * r / (c + safet),
self.qmin * safet / (c + safet),
self.qmax * safet / (c + safet),
safet / (c + safet),
safet / (c + safet)
], tc='d')
def F(x=None, z=None):
from scipy.special import xlogy
if x is None: return 1, x0
if any(z < 0 for z in x):
return None
f = ( xlogy(c * (1 - r), x[0])
+ xlogy(c * r, x[1])
+ xlogy(self.t - self.rt, x[3])
+ xlogy(self.rt, x[4])
+ halfchisq
- self.mle.obj
+ self.llb)
f *= -1 / safet
jf = matrix(0, size=(1, 6), tc='d')
jf[0] = c * (1 - r) / x[0] if c * (1 - r) > 0 else 0
jf[1] = c * r / x[1] if c * r > 0 else 0
jf[4] = (self.t - self.rt) / x[4] if self.t > self.rt else 0
jf[5] = self.rt / x[5] if self.rt > 0 else 0
jf *= -1 / safet
if z is None: return f, jf
hf = spdiag([
-c * (1 - r) / x[0]**2 if c * (1 - r) > 0 else 0,
-c * r / x[1]**2 if c * r > 0 else 0,
0,
0,
-(self.t - self.rt) / x[4]**2 if self.t > self.rt else 0,
-self.rt / x[5]**2 if self.rt > 0 else 0
])
hf *= -z[0] / safet
return f, jf, hf
soln = solvers.cpl(c=cost, F=F, G=G, h=h, A=A.T, b=b)
from pprint import pformat
import numpy
assert soln['status'][:7] == 'optimal', pformat({ 'soln': soln,
'solnx': [ z for z in soln['x'] ],
'datum': (c, w, r),
'F(x=x0)': F(x=x0),
'A': numpy.matrix(A),
'b': [ z for z in b ],
'A.x0 - b': [ z for z in A.T*x0 - b ],
'G.x0 - h': [ z for z in G*x0 - h ],
'F(x=soln)': F(x=soln['x']),
'A.x - b': [ z for z in A.T*soln['x'] - b ],
'G.x - h': [ z for z in G*soln['x'] - h ],
})
return (soln['primal objective'], soln['status'],
(1 - r) * soln['x'][0],
r * soln['x'][0]
) + tuple(soln['x'][1:])
def innersolveturg(self, c, w, r, alpha):
from scipy.stats import chi2
safet = max(self.t, 1)
halfchisq = 0.5 * chi2.isf(q=alpha, df=1)
self.params.w.value = w / self.wmax
self.params.wcost.value = w
self.params.vlb.value = self.vlb
self.params.wq0.value = self.wq0 / self.wmax
self.params.wq1.value = self.wq1 / self.wmax
self.params.q0t.value = self.q0t
self.params.q1t.value = self.q1t
self.params.coneminusr.value = c * (1 - r) / safet
self.params.cr.value = c * r / safet
self.params.tminusrt.value = (self.t - self.rt) / safet
self.params.rt.value = self.rt / safet
self.params.constraintrhs.value = (-halfchisq + self.mle.obj - self.llb) / safet
self.prob.solve(verbose=False)
return (self.prob.value,
self.prob.status,
self.vars.q0t.value,
self.vars.q1t.value,
self.vars.qmin.value,
self.vars.qmax.value,
self.vars.psi0.value,
self.vars.psi1.value
)
def update(self, c, w, r):
import math
from scipy.special import xlogy
self.mle.update(c, w, r)
alpha = self.alpha
(pvalue, pstatus, q0t, q1t, qmin, qmax, psi0, psi1) = self.innersolve(c, w, r, alpha)
if pstatus[:7] != 'optimal':
alphalb = 0
alphaub = alpha
while alphaub - alphalb >= 1e-3:
alphatest = 0.5 * (alphalb + alphaub)
(pvalue, pstatus, q0t, q1t, qmin, qmax, psi0, psi1) = self.innersolve(c, w, r, alphatest)
if pstatus[:7] == 'optimal':
alphalb = alphatest
else:
alphaub = alphatest
alpha = alphalb
(pvalue, pstatus, q0t, q1t, qmin, qmax, psi0, psi1) = self.innersolve(c, w, r, alpha)
assert pstatus[:7] == 'optimal', { 'alpha': alpha, 'pstatus': pstatus }
self.vlb = pvalue
self.qmin = qmin
self.qmax = qmax
self.llb += ( xlogy(c * (1 - r), q0t)
+ xlogy(c * r, q1t)
+ xlogy(self.t - self.rt, psi0)
+ xlogy(self.rt, psi1)
)
self.t += c
self.rt += c * r
self.q0t = q0t + psi0 * self.q0t
self.wq0 = w * q0t + psi0 * self.wq0
self.q1t = q1t + psi1 * self.q1t
self.wq1 = w * q1t + psi1 * self.wq1
return (1 - r) * q0t + r * q1t, { # 'self.q0t': self.q0t,
# 'self.q1t': self.q1t,
# 'qmin': qmin.value,
# 'qmax': qmax.value,
'vlb': self.vlb,
'vmle': self.mle.vmin,
# 'self.rt': self.rt,
'alpha': alpha,
}
def __initv1__(self, wmin, wmax, alpha):
from scipy.stats import chi2
from cvxopt import matrix
assert wmax > 1
assert wmin >= 0
assert wmin < wmax
self.wmin = wmin
self.wmax = wmax
self.qmin = (wmax - 1) / (wmax - wmin)
self.qmax = (1 - wmin) / (wmax - wmin)
self.G = matrix([ [ -1, 0, 0, 0 ],
[ 0, -1, 0, 0 ],
[ 0, 0, -1, 0 ],
[ 0, 0, 0, -1 ],
],
tc='d').T
self.h = matrix([ 0, 0, 0, 0 ], tc='d')
self.b = matrix([ 1 / wmax, 1 ], tc='d')
self.t = 0
self.halfchisq = 0.5 * chi2.isf(q=alpha, df=1)
self.llb = 0
self.vlb = 0
self.mle = OnlineCoordinateDescentMLE(wmin=wmin, wmax=wmax)
def updatev1(self, c, w, r):
import cvxpy as cp
import math
self.mle.update(c, w, r)
qt = cp.Variable(nonneg=True)
qmin = cp.Variable(nonneg=True)
qmax = cp.Variable(nonneg=True)
psi = cp.Variable(nonneg=True)
safet = max(self.t, 1)
prob = cp.Problem(cp.Minimize(float(w * r) * qt + self.vlb * psi), [
float(w / self.wmax) * qt
+ (self.wmin / self.wmax) * qmin
+ qmax
+ (1 / self.wmax - (self.wmin / self.wmax) * self.qmin - self.qmax) * psi == 1 / self.wmax,
qt + qmin + qmax + (1 - self.qmin - self.qmax) * psi == 1,
float(c / safet) * cp.log(qt) + (self.t / safet) * cp.log(psi) >= (-self.halfchisq + self.mle.obj - self.llb) / safet
])
prob.solve(verbose=False)
if prob.status[:7] != 'optimal':
# just maximize likelihood to recover (?)
pass
from pprint import pformat
assert prob.status[:7] == 'optimal', pformat({ 'datum': [ c, w, r ],
'x': [ z.value for z in [ qt, qmin, qmax, psi ] ],
'prob': prob.status,
't': self.t,
'lmle': self.mle.obj,
'llb': self.llb,# + c * math.log(qt.value) + self.t * math.log(psi.value),
'halfchisq': self.halfchisq,
})
self.vlb = prob.value
self.qmin = qmin.value
self.qmax = qmax.value
self.lastphi = psi.value
self.llb += c * math.log(qt.value) + self.t * math.log(psi.value)
self.t += c
return { 'soln': [ z.value for z in [ qt, qmin, qmax, psi ] ], 'lmle': self.mle.obj,
'llb': self.llb, 'halfchisq': self.halfchisq, 'vlb': self.vlb, 'vhat': self.mle.vhat }
if False:
from cvxopt import matrix, solvers
import math
assert c > 0
self.mle.update(c, w, r)
lmle = self.mle.obj
safet = max(self.t, 1)
x0 = matrix([ c / (c + safet),
self.qmin * safet / (c + safet),
self.qmax * safet / (c + safet),
safet / (c + safet) ], tc='d')
def F(x=None, z=None):
if x is None: return 1, x0
if x[0] <= 0 or x[3] <= 0:
return None
f = -c * math.log(x[0]) - self.t * math.log(x[3]) - self.halfchisq + lmle - self.llb
f /= safet
jf = matrix([ -c / (safet * x[0]), 0, 0, -self.t / (safet * x[3]) ], tc='d').T
if z is None: return f, jf
hf = z[0] * matrix([ [ (c / safet) * 1/x[0]**2, 0, 0, 0 ],
[ 0, 0, 0, 0 ],
[ 0, 0, 0, 0 ],
[ 0, 0, 0, (self.t / safet) * 1/x[3]**2 ]
], tc='d')
return f, jf, hf
cost = matrix([ float(w * r), 0, 0, self.vlb ], tc='d')
A = matrix([
[ float(w) / self.wmax,
self.wmin / self.wmax,
1,
(1 / self.wmax - (self.wmin / self.wmax) * self.qmin - self.qmax) ],
[ 1, 1, 1, (1 - self.qmin - self.qmax) ]
],
tc='d')
soln = solvers.cpl(c=cost, F=F, G=self.G, h=self.h, A=A.T, b=self.b, options={'show_progress': True,
'maxiters': 100 })
from pprint import pformat
assert soln['status'] == 'optimal', pformat({ 'datum': [ c, w, r ],
'x': [ z for z in soln['x'] ],
'soln': soln,
't': self.t,
'lmle': lmle,
'llb': self.llb,
'F(x0)': F(x=x0),
'F(x)': F(soln['x'])
})
print(pformat((F(x=x0), F(x=soln['x']))))
self.vlb = soln['primal objective']
self.qmin = soln['x'][1]
self.qmax = soln['x'][2]
self.lastphi = soln['x'][3]
self.llb += c * math.log(soln['x'][0]) + self.t * math.log(soln['x'][3])
self.t += c
return { 'soln': [ z for z in soln['x'] ], 'lmle': lmle,
'llb': self.llb, 'halfchisq': self.halfchisq, 'vlb': self.vlb, 'vhat': self.mle.vhat }
class Test:
def flass():
import environments.ControlledRangeVariance
import MLE.MLE
def batchtoonline(samples, seed=45):
import numpy as np
state = np.random.RandomState(seed)
n = sum(c for c, w, r in samples)
while n > 0:
p = np.array([ c for c, w, r in samples ], dtype='float64') / n
what = state.choice(len(samples), p=p)
c = min(samples[what][0], 1)
yield (c, samples[what][1], samples[what][2])
samples[what] = (samples[what][0] - c, samples[what][1], samples[what][2])
n -= c
env = environments.ControlledRangeVariance.ControlledRangeVariance(seed=45, wsupport=[0,2,1000], expwsq=100)
_, samples = env.sample(1 + (1 << 16))
ocd = OnlineCoordinateDescentLB(wmin=0, wmax=1000, alpha=0.05)
from pprint import pformat
t = 0
for n, (c, w, r) in enumerate(batchtoonline(samples)):
qt, qex = ocd.update(c, w, r)
t += c
if (n & (n - 1) == 0 and n & 0xAAAAAAAA == 0) or w == 1000:
vlb = '{:.5}'.format(qex['vlb'])
vmle = '{:.5}'.format(qex['vmle'])
print([ c, w, r, t, '{:.3g}'.format(qt), '{:.3f}'.format(t * qt), { 'vlb': vlb, 'vmle': vmle, 'alpha': qex['alpha'] } ],
flush=True)
Test.flass()
```
# Histogram Based
```
from importlib import reload
import environments.ControlledRangeVariance
import MLE.MLE
reload(environments.ControlledRangeVariance)
reload(MLE.MLE)
def batchtoonline(samples, seed=45):
import numpy as np
state = np.random.RandomState(seed)
n = sum(c for c, w, r in samples)
while n > 0:
p = np.array([ c for c, w, r in samples ], dtype='float64') / n
what = state.choice(len(samples), p=p)
c = min(samples[what][0], 1)
yield (c, samples[what][1], samples[what][2])
samples[what] = (samples[what][0] - c, samples[what][1], samples[what][2])
n -= c
env = environments.ControlledRangeVariance.ControlledRangeVariance(seed=45, wsupport=[0,2,1000], expwsq=100)
happrox = MLE.MLE.Online.HistApprox(wmin=0, wmax=1000, numbuckets=10)
onlineci = MLE.MLE.Online.CI(wmin=0, wmax=1000, rmin=0, rmax=1, alpha=0.05)
onlinemle = MLE.MLE.Online.MLE(wmin=0, wmax=1000, rmin=0, rmax=1)
_, samples = env.sample(1 + (1 << 16))
from pprint import pformat
print(pformat(samples), flush=True)
t = 0
for n, (c, w, r) in enumerate(batchtoonline(samples)):
happrox.update(c, w, r)
onlineci.update(happrox.iterator)
onlinemle.update(happrox.iterator)
t += c
if (n & (n - 1) == 0 and n & 0xAAAAAAAA == 0) or w == 1000:
vmle = MLE.MLE.estimate(happrox.iterator, wmin=0, wmax=1000)[1]['vmin']
vlb = MLE.MLE.asymptoticconfidenceinterval(happrox.iterator, wmin=0, wmax=1000)[0][0]
print([ c, w, r, t, 'n/a', 'n/a', { 'vlb': '{:.3f}'.format(vlb), 'vmle': '{:.3f}'.format(vmle), 'alpha': 0.05 } ],
flush=True)
# from pprint import pformat
# print(pformat(
# {
# 'n': n,
# 'onlineci': onlineci.getqfunc(),
# 'onlinemle': onlinemle.getqfunc(),
# 'batchmle': MLE.MLE.estimate(happrox.iterator, wmin=0, wmax=1000),
# 'batchci': MLE.MLE.asymptoticconfidenceinterval(happrox.iterator, wmin=0, wmax=1000)
# }),
# flush=True)
```
# Forest Covertype
### Full Information Online Learning
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
def flass():
from sklearn.datasets import fetch_covtype
from sklearn.decomposition import PCA
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from math import ceil
import numpy as np
cov = fetch_covtype()
cov.data = PCA(whiten=True).fit_transform(cov.data)
classes = np.unique(cov.target - 1)
ndata = len(cov.target)
order = np.random.RandomState(seed=42).permutation(ndata)
ntrain = ceil(0.2 * ndata)
Object = lambda **kwargs: type("Object", (), kwargs)()
train = Object(data = cov.data[order[:ntrain]], target = cov.target[order[:ntrain]] - 1)
test = Object(data = cov.data[order[ntrain:]], target = cov.target[order[ntrain:]] - 1)
for lr in (1, ):
print("**** lr = {} ****".format(lr))
print('{:8.8s}\t{:8.8s}\t{:10.10s}'.format('n', 'emp loss', 'since last'))
classweights = { k: lr for k, _ in Counter(train.target).items() }
cls = SGDClassifier(loss='log', class_weight=classweights, shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
blocksize = 32
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = np.array([ np.outer(t, np.append(t, [1])).ravel() for z in ind for t in ( train.data[z], ) ])
actual = [ train.target[z] for z in ind ]
if n > 0:
pred = cls.predict(v)
for p, a in zip(pred, actual):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f}\t{:<10.3f}'.format(loss.n, loss.mean(), sincelast.mean()), flush=True)
sincelast = OnlineDRO.EasyAcc()
cls.partial_fit(v, actual, classes=classes)
print('{:<8d}\t{:<8.3f}\t{:<10.3f}'.format(loss.n, loss.mean(), sincelast.mean()), flush=True)
sincelast = OnlineDRO.EasyAcc()
preds = cls.predict(np.array([np.outer(d, np.append(d, [1])).ravel() for d in test.data]))
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
OnlineDRO.flass()
```
### Partial Information Online Learning, Softmax Logging Policy
Uniform $(\tau = 0)$ and softmax $(\tau = 4)$ are pretty similar for off-policy learning but uniform has larger regret.
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
def flass():
from scipy.special import softmax
from sklearn.datasets import fetch_covtype
from sklearn.decomposition import PCA
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from math import ceil
import numpy as np
cov = fetch_covtype()
cov.data = PCA(whiten=True).fit_transform(cov.data)
classes = np.unique(cov.target - 1)
ndata = len(cov.target)
order = np.random.RandomState(seed=42).permutation(ndata)
ntrain = ceil(0.2 * ndata)
Object = lambda **kwargs: type("Object", (), kwargs)()
train = Object(data = cov.data[order[:ntrain]], target = cov.target[order[:ntrain]] - 1)
test = Object(data = cov.data[order[ntrain:]], target = cov.target[order[ntrain:]] - 1)
blocksize = 32
for lr, tau in ( (x, y) for x in np.logspace(-2.5, -2, 1) for y in (0, 4, ) ):
print("*** lr = {} tau = {} ***".format(lr, tau), flush=True)
print('{:8.8s}\t{:8.8s}\t{:10.10s}\t{:8.8s}\t{:10.10s}'.format(
'n', 'emp loss', 'since last', 'log pv', 'since last')
)
cls = SGDClassifier(loss='log', shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
logpv = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
loggerrand = np.random.RandomState(seed=2112)
logchoices = [None]*len(train.data)
pchoices = [None]*len(train.data)
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = np.array([ np.outer(t, np.append(t, [1])).ravel() for z in ind for t in ( train.data[z], ) ])
if n == 0 and pno == 0:
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, size=1)
logchoices[z] = choice[0]
pchoices[z] = 1.0 / len(classes)
else:
predlogp = cls.predict_proba(v)
soft = softmax(tau * predlogp, axis=1)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, p=soft[i,:], size=1)
logchoices[z] = choice[0]
pchoices[z] = soft[i, choice[0]]
pred = cls.predict(v)
actual = [ train.target[z] for z in ind ]
for i, (p, a) in enumerate(zip(pred, actual)):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
logpv += soft[i, a]
logpvsl += soft[i, a]
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
x = np.array([ v[i] for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
y = np.array([ logchoices[z] for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
w = np.array([ (lr / len(classes)) * (1 / pchoices[z])
for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
if np.any(x):
cls.partial_fit(x, y, classes=classes, sample_weight=w)
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
preds = cls.predict(np.array([np.outer(d, np.append(d, [1])).ravel() for d in test.data]))
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
OnlineDRO.flass()
```
### Bound Online Learning
Either MLE, lower bound, or upper bound. Some count decay seems better than no count decay. Upper bound seems to want less count decay than the MLE or lower bound. All forms of bound learning have lower regret than IPS learning.
TODO: "delayed batch" online learning.
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
class OnlineCressieReadLB:
from math import inf
def __init__(self, alpha, gamma=1, wmin=0, wmax=inf):
import numpy as np
self.alpha = alpha
self.gamma = gamma
self.n = 0
self.sumw = 0
self.sumwsq = 0
self.sumwr = 0
self.sumwsqr = 0
self.sumwsqrsq = 0
self.wmin = wmin
self.wmax = wmax
self.duals = None
self.mleduals = None
def update(self, c, w, r):
if c > 0:
assert w + 1e-6 >= self.wmin and w <= self.wmax + 1e-6, 'w = {} < {} < {}'.format(self.wmin, w, self.wmax)
assert r >= 0 and r <= 1, 'r = {}'.format(r)
decay = self.gamma ** c
self.n = decay * self.n + c
self.sumw = decay * self.sumw + c * w
self.sumwsq = decay * self.sumwsq + c * w**2
self.sumwr = decay * self.sumwr + c * w * r
self.sumwsqr = decay * self.sumwsqr + c * (w**2) * r
self.sumwsqrsq = decay * self.sumwsqrsq + c * (w**2) * (r**2)
self.duals = None
self.mleduals = None
return self
def recomputeduals(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.duals = CrMinusTwo.intervalimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, self.sumwsqrsq,
self.wmin, self.wmax, self.alpha, raiseonerr=True)
def recomputedualsmle(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.mleduals = CrMinusTwo.estimateimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, None, None,
self.wmin, self.wmax, raiseonerr=True)
def qlb(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][0]['qfunc'](c, w, r) if self.duals[1][0] is not None else 1
def qub(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][1]['qfunc'](c, w, r) if self.duals[1][1] is not None else 1
def qmle(self, c, w, r):
if self.mleduals is None:
self.recomputedualsmle()
assert self.mleduals is not None
return self.mleduals[1]['qfunc'](c, w, r) if self.mleduals[1] is not None else 1
def flass():
from scipy.special import softmax
from sklearn.datasets import fetch_covtype
from sklearn.decomposition import PCA
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from math import ceil, exp
import numpy as np
cov = fetch_covtype()
cov.data = PCA(whiten=True).fit_transform(cov.data)
classes = np.unique(cov.target - 1)
ndata = len(cov.target)
order = np.random.RandomState(seed=42).permutation(ndata)
ntrain = ceil(0.2 * ndata)
Object = lambda **kwargs: type("Object", (), kwargs)()
train = Object(data = cov.data[order[:ntrain]], target = cov.target[order[:ntrain]] - 1)
test = Object(data = cov.data[order[ntrain:]], target = cov.target[order[ntrain:]] - 1)
blocksize = 32
for lr, tau, what, gamma in ( (x, y, z, g) for x in np.logspace(-2.5, -2, 1) for y in (4, )
for z in ('mle', 'ub', 'lb') for g in (1, 0.9999, 0.999, 0.99, ) ):
print("*** lr = {} tau = {} what = {} gamma = {} ***".format(lr, tau, what, gamma), flush=True)
print('{:8.8s}\t{:8.8s}\t{:10.10s}\t{:8.8s}\t{:10.10s}\t{:8.8s}\t{:10.10s}'.format(
'n', 'eff n', 'since last', 'emp loss', 'since last', 'log pv', 'since last')
)
cls = SGDClassifier(loss='log', shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
logpv = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effn = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
loggerrand = np.random.RandomState(seed=2112)
logchoices = [None]*len(train.data)
pchoices = [None]*len(train.data)
ocrl = OnlineDRO.OnlineCressieReadLB(alpha=0.05,
gamma=gamma,
wmin=0,
wmax=exp(tau) + len(classes) - 1
)
qfunc = ocrl.qmle if what == 'mle' else ocrl.qlb if what == 'lb' else ocrl.qub
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = np.array([ np.outer(t, np.append(t, [1])).ravel() for z in ind for t in ( train.data[z], ) ])
if n == 0 and pno == 0:
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, size=1)
logchoices[z] = choice[0]
pchoices[z] = 1.0 / len(classes)
else:
predlogp = cls.predict_proba(v)
soft = softmax(tau * predlogp, axis=1)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, p=soft[i,:], size=1)
logchoices[z] = choice[0]
pchoices[z] = soft[i, choice[0]]
pred = cls.predict(v)
actual = [ train.target[z] for z in ind ]
for i, (p, a) in enumerate(zip(pred, actual)):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
logpv += soft[i, a]
logpvsl += soft[i, a]
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, effn.mean(), effnsl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
for i, z in enumerate(ind):
r = 1 if logchoices[z] == train.target[z] else 0
w = 1 / pchoices[z]
ocrl.update(1, w, r)
if n == 0 and pno == 0:
sampweight = np.array([ lr for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
else:
sampweight = np.array([ lr * w * ocrl.n
* max(0, qfunc(1, w, 1))
for i, z in enumerate(ind)
if logchoices[z] == train.target[z]
for w in (1 / pchoices[z],)# if logchoices[z] == pred[i] else 0,)
])
effn += sampweight.sum() / (lr * blocksize)
effnsl += sampweight.sum() / (lr * blocksize)
x = np.array([ v[i] for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
y = np.array([ logchoices[z] for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
if np.any(x):
cls.partial_fit(x, y, classes=classes, sample_weight=sampweight)
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, effn.mean(), effnsl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
preds = cls.predict(np.array([np.outer(d, np.append(d, [1])).ravel() for d in test.data]))
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
OnlineDRO.flass()
```
### Delayed Batch Bound Online Learning
Process larger batches as sets of smaller batches to emulate time delay in policy updates without changing optimization properties (i.e., SGD batch size).
Everything still seems to work.
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
class OnlineCressieReadLB:
from math import inf
def __init__(self, alpha, gamma=1, wmin=0, wmax=inf):
import numpy as np
self.alpha = alpha
self.gamma = gamma
self.n = 0
self.sumw = 0
self.sumwsq = 0
self.sumwr = 0
self.sumwsqr = 0
self.sumwsqrsq = 0
self.wmin = wmin
self.wmax = wmax
self.duals = None
self.mleduals = None
def update(self, c, w, r):
if c > 0:
assert w + 1e-6 >= self.wmin and w <= self.wmax + 1e-6, 'w = {} < {} < {}'.format(self.wmin, w, self.wmax)
assert r >= 0 and r <= 1, 'r = {}'.format(r)
decay = self.gamma ** c
self.n = decay * self.n + c
self.sumw = decay * self.sumw + c * w
self.sumwsq = decay * self.sumwsq + c * w**2
self.sumwr = decay * self.sumwr + c * w * r
self.sumwsqr = decay * self.sumwsqr + c * (w**2) * r
self.sumwsqrsq = decay * self.sumwsqrsq + c * (w**2) * (r**2)
self.duals = None
self.mleduals = None
return self
def recomputeduals(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.duals = CrMinusTwo.intervalimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, self.sumwsqrsq,
self.wmin, self.wmax, self.alpha, raiseonerr=True)
def recomputedualsmle(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.mleduals = CrMinusTwo.estimateimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, None, None,
self.wmin, self.wmax, raiseonerr=True)
def qlb(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][0]['qfunc'](c, w, r) if self.duals[1][0] is not None else 1
def qub(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][1]['qfunc'](c, w, r) if self.duals[1][1] is not None else 1
def qmle(self, c, w, r):
if self.mleduals is None:
self.recomputedualsmle()
assert self.mleduals is not None
return self.mleduals[1]['qfunc'](c, w, r) if self.mleduals[1] is not None else 1
def flass():
from scipy.special import softmax
from sklearn.datasets import fetch_covtype
from sklearn.decomposition import PCA
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from math import ceil, exp
import numpy as np
cov = fetch_covtype()
cov.data = PCA(whiten=True).fit_transform(cov.data)
classes = np.unique(cov.target - 1)
ndata = len(cov.target)
order = np.random.RandomState(seed=42).permutation(ndata)
ntrain = ceil(0.2 * ndata)
Object = lambda **kwargs: type("Object", (), kwargs)()
train = Object(data = cov.data[order[:ntrain]], target = cov.target[order[:ntrain]] - 1)
test = Object(data = cov.data[order[ntrain:]], target = cov.target[order[ntrain:]] - 1)
subblocksize = 32
delay = 8
blocksize = delay * subblocksize
for lr, tau, what, gamma in ( (x, y, z, g) for x in np.logspace(-2.5, -2, 1) for y in (4, )
for z in ('mle', 'ub', 'lb') for g in (1, 0.9999, 0.999, 0.99,) ):
print("*** lr = {} tau = {} what = {} gamma = {} ***".format(lr, tau, what, gamma), flush=True)
print('{:8.8s}\t{:8.8s}\t{:10.10s}\t{:8.8s}\t{:10.10s}\t{:8.8s}\t{:10.10s}'.format(
'n', 'eff n', 'since last', 'emp loss', 'since last', 'log pv', 'since last')
)
cls = SGDClassifier(loss='log', shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
logpv = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effn = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
loggerrand = np.random.RandomState(seed=2112)
logchoices = [None]*len(train.data)
pchoices = [None]*len(train.data)
ocrl = OnlineDRO.OnlineCressieReadLB(alpha=0.05,
gamma=gamma,
wmin=0,
wmax=exp(tau) + len(classes) - 1
)
qfunc = ocrl.qmle if what == 'mle' else ocrl.qlb if what == 'lb' else ocrl.qub
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = np.array([ np.outer(t, np.append(t, [1])).ravel() for z in ind for t in ( train.data[z], ) ])
if n == 0 and pno == 0:
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, size=1)
logchoices[z] = choice[0]
pchoices[z] = 1.0 / len(classes)
else:
predlogp = cls.predict_proba(v)
soft = softmax(tau * predlogp, axis=1)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, p=soft[i,:], size=1)
logchoices[z] = choice[0]
pchoices[z] = soft[i, choice[0]]
pred = cls.predict(v)
actual = [ train.target[z] for z in ind ]
for i, (p, a) in enumerate(zip(pred, actual)):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
logpv += soft[i, a]
logpvsl += soft[i, a]
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, effn.mean(), effnsl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
for i, z in enumerate(ind):
r = 1 if logchoices[z] == train.target[z] else 0
w = 1 / pchoices[z]
ocrl.update(1, w, r)
for d in range(delay):
x = np.array([ v[i]
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
y = np.array([ logchoices[z]
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
if n == 0 and pno == 0:
sampweight = np.array([ lr
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
else:
sampweight = np.array([ lr * w * ocrl.n
* max(0, qfunc(1, w, 1))
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z]
for w in (1 / pchoices[z],)
])
effn += sampweight.sum() / (lr * subblocksize)
effnsl += sampweight.sum() / (lr * subblocksize)
if np.any(x):
cls.partial_fit(x, y, classes=classes, sample_weight=sampweight)
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, effn.mean(), effnsl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
# from pprint import pformat
# print(pformat(ocrl.__dict__))
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
preds = cls.predict(np.array([np.outer(d, np.append(d, [1])).ravel() for d in test.data]))
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
OnlineDRO.flass()
```
### Auto-Temperature + Delayed Batch Bound Online Learning
Best regret achieved with lower bound optimization.
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
class OnlineCressieReadLB:
from math import inf
def __init__(self, alpha, gamma=1, wmin=0, wmax=inf):
import numpy as np
self.alpha = alpha
self.gamma = gamma
self.n = 0
self.sumw = 0
self.sumwsq = 0
self.sumwr = 0
self.sumwsqr = 0
self.sumwsqrsq = 0
self.wmin = wmin
self.wmax = wmax
self.duals = None
self.mleduals = None
def update(self, c, w, r):
if c > 0:
assert w + 1e-6 >= self.wmin and w <= self.wmax + 1e-6, 'w = {} < {} < {}'.format(self.wmin, w, self.wmax)
assert r >= 0 and r <= 1, 'r = {}'.format(r)
decay = self.gamma ** c
self.n = decay * self.n + c
self.sumw = decay * self.sumw + c * w
self.sumwsq = decay * self.sumwsq + c * w**2
self.sumwr = decay * self.sumwr + c * w * r
self.sumwsqr = decay * self.sumwsqr + c * (w**2) * r
self.sumwsqrsq = decay * self.sumwsqrsq + c * (w**2) * (r**2)
self.duals = None
self.mleduals = None
return self
def recomputeduals(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.duals = CrMinusTwo.intervalimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, self.sumwsqrsq,
self.wmin, self.wmax, self.alpha, raiseonerr=True)
def recomputedualsmle(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.mleduals = CrMinusTwo.estimateimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, None, None,
self.wmin, self.wmax, raiseonerr=True)
def qlb(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][0]['qfunc'](c, w, r) if self.duals[1][0] is not None else 1
def qub(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][1]['qfunc'](c, w, r) if self.duals[1][1] is not None else 1
def qmle(self, c, w, r):
if self.mleduals is None:
self.recomputedualsmle()
assert self.mleduals is not None
return self.mleduals[1]['qfunc'](c, w, r) if self.mleduals[1] is not None else 1
def autotune(pre, target, taumax):
from scipy.optimize import root_scalar
def f(tau):
from scipy.special import softmax
import numpy as np
soft = softmax(tau * pre, axis=1)
minsoft = np.min(soft, axis=1)
return np.mean(minsoft) - target
fmax = f(taumax)
if fmax >= 0:
return taumax
taumin = 0
fmin = f(taumin)
assert fmin > 0, { 'fmin': fmin, 'target': target }
root = root_scalar(f, bracket=(taumin, taumax))
assert root.converged, root
return root.root
def flass():
from scipy.special import softmax
from sklearn.datasets import fetch_covtype
from sklearn.decomposition import PCA
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from math import ceil, exp
import numpy as np
cov = fetch_covtype()
cov.data = PCA(whiten=True).fit_transform(cov.data)
classes = np.unique(cov.target - 1)
ndata = len(cov.target)
order = np.random.RandomState(seed=42).permutation(ndata)
ntrain = ceil(0.2 * ndata)
Object = lambda **kwargs: type("Object", (), kwargs)()
train = Object(data = cov.data[order[:ntrain]], target = cov.target[order[:ntrain]] - 1)
test = Object(data = cov.data[order[ntrain:]], target = cov.target[order[ntrain:]] - 1)
subblocksize = 32
delay = 8
blocksize = delay * subblocksize
for lr, taumax, target, what, gamma in ( (x, 8, y, z, g) for x in np.logspace(-2.5, -2, 1)
for z in ('mle', 'ub', 'lb') for g in (0.9999, 0.999,) for y in (0.025, 0.05, 0.1, 0.2, ) ):
print("*** lr = {} taumax = {} target = {} what = {} gamma = {} ***".format(lr, taumax, target, what, gamma), flush=True)
print('{:8.8s}\t{:8.8s} [{:10.10s}]\t{:8.8s} [{:10.10s}]\t{:8.8s} [{:10.10s}]\t{:8.8s} [{:10.10s}]'.format(
'n', 'eff n', 'since last', 'av tau', 'since last', 'emp loss', 'since last', 'log pv', 'since last')
)
try:
target /= len(classes)
cls = SGDClassifier(loss='log', shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
logpv = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effn = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
avtau = OnlineDRO.EasyAcc()
avtausl = OnlineDRO.EasyAcc()
loggerrand = np.random.RandomState(seed=2112)
logchoices = [None]*len(train.data)
pchoices = [None]*len(train.data)
ocrl = OnlineDRO.OnlineCressieReadLB(alpha=0.05,
gamma=gamma,
wmin=0,
# wmax=exp(taumax) + len(classes) - 1
)
qfunc = ocrl.qmle if what == 'mle' else ocrl.qlb if what == 'lb' else ocrl.qub
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = np.array([ np.outer(t, np.append(t, [1])).ravel() for z in ind for t in ( train.data[z], ) ])
if n == 0 and pno == 0:
pred = np.zeros(blocksize)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, size=1)
logchoices[z] = choice[0]
pchoices[z] = 1.0 / len(classes)
else:
predlogp = cls.predict_proba(v)
tau = OnlineDRO.autotune(predlogp, target, taumax)
avtau += tau
avtausl += tau
soft = softmax(tau * predlogp, axis=1)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, p=soft[i,:], size=1)
logchoices[z] = choice[0]
pchoices[z] = soft[i, choice[0]]
pred = cls.predict(v)
actual = [ train.target[z] for z in ind ]
for i, (p, a) in enumerate(zip(pred, actual)):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
logpv += soft[i, a]
logpvsl += soft[i, a]
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]'.format(
loss.n, effn.mean(), effnsl.mean(), avtau.mean(), avtausl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
avtausl = OnlineDRO.EasyAcc()
for i, z in enumerate(ind):
r = 1 if logchoices[z] == train.target[z] else 0
w = 1 / pchoices[z] if pred[i] == logchoices[z] else 0
ocrl.update(1, w, r)
for d in range(delay):
x = np.array([ v[i]
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
y = np.array([ logchoices[z]
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
if n == 0 and pno == 0:
sampweight = np.array([ lr
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
else:
sampweight = np.array([ lr * w * ocrl.n
* max(0, qfunc(1, w, 1))
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z]
for w in (1 / pchoices[z],)
])
effn += sampweight.sum() / (lr * subblocksize)
effnsl += sampweight.sum() / (lr * subblocksize)
if np.any(x):
cls.partial_fit(x, y, classes=classes, sample_weight=sampweight)
print('{:<8d}\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]'.format(
loss.n, effn.mean(), effnsl.mean(), avtau.mean(), avtausl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
avtausl = OnlineDRO.EasyAcc()
preds = cls.predict(np.array([np.outer(d, np.append(d, [1])).ravel() for d in test.data]))
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
except KeyboardInterrupt:
raise
except:
pass
OnlineDRO.flass()
```
# 20 Newsgroups
Really hard ... 20 actions and only 22K examples.
### Full Information Online Learning
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
def flass():
from scipy.sparse import vstack
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
import numpy as np
train = fetch_20newsgroups(subset='train', remove=('headers', 'footers', 'quotes'))
classes = np.unique(train.target)
test = fetch_20newsgroups(subset='test', remove=('headers', 'footers', 'quotes'))
for ngram, norm, lr, nfeat in ( (2, 'l2', 1, 20), ):
print("**** ngram = {} norm = {} lr = {} nfeat = {} ****".format(ngram, norm, lr, nfeat))
print('{:8.8s}\t{:8.8s}\t{:10.10s}'.format('n', 'emp loss', 'since last'))
vectorizer = HashingVectorizer(n_features = 1 << nfeat, norm=norm, ngram_range=(1, ngram), alternate_sign=True)
docs = vectorizer.transform(train.data)
testdocs = vectorizer.transform(test.data)
classweights = { k: lr for k in classes }
cls = SGDClassifier(loss='log', class_weight=classweights, shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
blocksize = 32
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = vstack([ docs[z] for z in ind ])
actual = [ train.target[z] for z in ind ]
if n > 0:
pred = cls.predict(v)
for p, a in zip(pred, actual):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f}\t{:<10.3f}'.format(loss.n, loss.mean(), sincelast.mean()), flush=True)
sincelast = OnlineDRO.EasyAcc()
cls.partial_fit(v, actual, classes=classes)
print('{:<8d}\t{:<8.3f}\t{:<10.3f}'.format(loss.n, loss.mean(), sincelast.mean()), flush=True)
sincelast = OnlineDRO.EasyAcc()
preds = cls.predict(testdocs)
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
OnlineDRO.flass()
```
### Partial Information Online Learning, Softmax Logging Policy
Uniform $(\tau = 0)$ and softmax $(\tau = 4)$ are pretty similar for off-policy learning but uniform has larger regret.
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
def flass():
from scipy.sparse import vstack
from scipy.special import softmax
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
import numpy as np
train = fetch_20newsgroups(subset='train', remove=('headers', 'footers', 'quotes'))
classes = np.unique(train.target)
test = fetch_20newsgroups(subset='test', remove=('headers', 'footers', 'quotes'))
blocksize = 32
for ngram, norm, lr, nfeat, tau in ( (2, 'l2', z, 20, x) for x in (0, 4,) for z in (0.5, ) ):
print("**** ngram = {} norm = {} lr = {} nfeat = {} tau = {} ****".format(ngram, norm, lr, nfeat, tau))
print('{:8.8s}\t{:8.8s}\t{:10.10s}\t{:8.8s}\t{:10.10s}'.format(
'n', 'emp loss', 'since last', 'log pv', 'since last')
)
vectorizer = HashingVectorizer(n_features = 1 << nfeat, norm=norm, ngram_range=(1, ngram), alternate_sign=True)
docs = vectorizer.transform(train.data)
testdocs = vectorizer.transform(test.data)
classweights = { k: lr for k in classes }
cls = SGDClassifier(loss='log', class_weight=classweights, shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
logpv = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
loggerrand = np.random.RandomState(seed=2112)
logchoices = [None]*len(train.data)
pchoices = [None]*len(train.data)
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = vstack([ docs[z] for z in ind ])
if n == 0 and pno == 0:
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, size=1)
logchoices[z] = choice[0]
pchoices[z] = 1.0 / len(classes)
else:
predlogp = cls.predict_proba(v)
soft = softmax(tau * predlogp, axis=1)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, p=soft[i,:], size=1)
logchoices[z] = choice[0]
pchoices[z] = soft[i, choice[0]]
pred = cls.predict(v)
actual = [ train.target[z] for z in ind ]
for i, (p, a) in enumerate(zip(pred, actual)):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
logpv += soft[i, a]
logpvsl += soft[i, a]
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
# y = np.array([ logchoices[z] for i, z in enumerate(ind) ])
# w = np.array([ (lr / len(classes)) * (1 / pchoices[z])
# if logchoices[z] == train.target[z]
# else -(lr / len(classes)**2) * (1/pchoices[z])
# for i, z in enumerate(ind) ])
y = np.array([ logchoices[z] for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
w = np.array([ (lr / len(classes)) * (1 / pchoices[z])
for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
if np.any(y):
x = vstack([ v[i] for i, z in enumerate(ind) if logchoices[z] == train.target[z] ])
# x = vstack([ v[i] for i, z in enumerate(ind) ])
cls.partial_fit(x, y, classes=classes, sample_weight=w)
print('{:<8d}\t{:<8.3f}\t{:<10.3f}\t{:<8.3f}\t{:<10.3f}'.format(
loss.n, loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
preds = cls.predict(testdocs)
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
OnlineDRO.flass()
```
### Auto-Temperature and Bound Optimization
Nothing working great, but regret is better than other approaches.
```
class OnlineDRO:
class EasyAcc:
def __init__(self):
self.n = 0
self.sum = 0
def __iadd__(self, other):
self.n += 1
self.sum += other
return self
def mean(self):
return self.sum / max(self.n, 1)
class OnlineCressieReadLB:
from math import inf
def __init__(self, alpha, gamma=1, wmin=0, wmax=inf):
import numpy as np
self.alpha = alpha
self.gamma = gamma
self.n = 0
self.sumw = 0
self.sumwsq = 0
self.sumwr = 0
self.sumwsqr = 0
self.sumwsqrsq = 0
self.wmin = wmin
self.wmax = wmax
self.duals = None
self.mleduals = None
def update(self, c, w, r):
if c > 0:
assert w + 1e-6 >= self.wmin and w <= self.wmax + 1e-6, 'w = {} < {} < {}'.format(self.wmin, w, self.wmax)
assert r >= 0 and r <= 1, 'r = {}'.format(r)
decay = self.gamma ** c
self.n = decay * self.n + c
self.sumw = decay * self.sumw + c * w
self.sumwsq = decay * self.sumwsq + c * w**2
self.sumwr = decay * self.sumwr + c * w * r
self.sumwsqr = decay * self.sumwsqr + c * (w**2) * r
self.sumwsqrsq = decay * self.sumwsqrsq + c * (w**2) * (r**2)
self.duals = None
self.mleduals = None
return self
def recomputeduals(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.duals = CrMinusTwo.intervalimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, self.sumwsqrsq,
self.wmin, self.wmax, self.alpha, raiseonerr=True)
def recomputedualsmle(self):
from MLE.MLE import CrMinusTwo as CrMinusTwo
self.mleduals = CrMinusTwo.estimateimpl(self.n, self.sumw, self.sumwsq,
self.sumwr, self.sumwsqr, None, None,
self.wmin, self.wmax, raiseonerr=True)
def qlb(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][0]['qfunc'](c, w, r) if self.duals[1][0] is not None else 1
def qub(self, c, w, r):
if self.duals is None:
self.recomputeduals()
assert self.duals is not None
return self.duals[1][1]['qfunc'](c, w, r) if self.duals[1][1] is not None else 1
def qmle(self, c, w, r):
if self.mleduals is None:
self.recomputedualsmle()
assert self.mleduals is not None
return self.mleduals[1]['qfunc'](c, w, r) if self.mleduals[1] is not None else 1
def autotune(pre, target, taumax):
from scipy.optimize import root_scalar
def f(tau):
from scipy.special import softmax
import numpy as np
soft = softmax(tau * pre, axis=1)
minsoft = np.min(soft, axis=1)
return np.mean(minsoft) - target
fmax = f(taumax)
if fmax >= 0:
return taumax
taumin = 0
fmin = f(taumin)
assert fmin > 0, { 'fmin': fmin, 'target': target }
root = root_scalar(f, bracket=(taumin, taumax))
assert root.converged, root
return root.root
def flass():
from scipy.sparse import vstack
from scipy.special import softmax
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from math import exp
import numpy as np
train = fetch_20newsgroups(subset='train', remove=('headers', 'footers', 'quotes'))
classes = np.unique(train.target)
test = fetch_20newsgroups(subset='test', remove=('headers', 'footers', 'quotes'))
subblocksize = 32
delay = 8
blocksize = delay * subblocksize
for lr, taumax, target, what, gamma in ( (x, 8, y, z, g) for x in (0.5,)
for z in ('mle', 'ub', 'lb')
for g in (0.9999, 0.999,)
for y in (0.025, 0.05, 0.1, 0.2, ) ):
ngram = 2
norm = 'l2'
nfeat = 20
print("*** lr = {} taumax = {} target = {} what = {} gamma = {} ***".format(lr, taumax, target, what, gamma), flush=True)
print('{:8.8s}\t{:8.8s} [{:10.10s}]\t{:8.8s} [{:10.10s}]\t{:8.8s} [{:10.10s}]\t{:8.8s} [{:10.10s}]'.format(
'n', 'eff n', 'since last', 'av tau', 'since last', 'emp loss', 'since last', 'log pv', 'since last')
)
try:
target /= len(classes)
vectorizer = HashingVectorizer(n_features = 1 << nfeat, norm=norm, ngram_range=(1, ngram), alternate_sign=True)
docs = vectorizer.transform(train.data)
testdocs = vectorizer.transform(test.data)
classweights = { k: lr for k in classes }
cls = SGDClassifier(loss='log', class_weight=classweights, shuffle=False)
loss = OnlineDRO.EasyAcc()
sincelast = OnlineDRO.EasyAcc()
logpv = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effn = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
avtau = OnlineDRO.EasyAcc()
avtausl = OnlineDRO.EasyAcc()
loggerrand = np.random.RandomState(seed=2112)
logchoices = [None]*len(train.data)
pchoices = [None]*len(train.data)
ocrl = OnlineDRO.OnlineCressieReadLB(alpha=0.05,
gamma=gamma,
wmin=0,
wmax=exp(taumax) + len(classes) - 1
)
qfunc = ocrl.qmle if what == 'mle' else ocrl.qlb if what == 'lb' else ocrl.qub
for pno in range(1):
order = np.random.RandomState(seed=42+pno).permutation(len(train.data))
for n, ind in enumerate(zip(*(iter(order),)*blocksize)):
v = vstack([ docs[z] for z in ind ])
if n == 0 and pno == 0:
pred = np.zeros(blocksize)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, size=1)
logchoices[z] = choice[0]
pchoices[z] = 1.0 / len(classes)
else:
predlogp = cls.predict_proba(v)
tau = OnlineDRO.autotune(predlogp, target, taumax)
avtau += tau
avtausl += tau
soft = softmax(tau * predlogp, axis=1)
for i, z in enumerate(ind):
if logchoices[z] is None:
choice = loggerrand.choice(a=classes, p=soft[i,:], size=1)
logchoices[z] = choice[0]
pchoices[z] = soft[i, choice[0]]
pred = cls.predict(v)
actual = [ train.target[z] for z in ind ]
for i, (p, a) in enumerate(zip(pred, actual)):
loss += 0 if p == a else 1
sincelast += 0 if p == a else 1
logpv += soft[i, a]
logpvsl += soft[i, a]
if (n & (n - 1) == 0): # and n & 0xAAAAAAAA == 0):
print('{:<8d}\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]'.format(
loss.n, effn.mean(), effnsl.mean(), avtau.mean(), avtausl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
avtausl = OnlineDRO.EasyAcc()
for i, z in enumerate(ind):
r = 1 if logchoices[z] == train.target[z] else 0
w = 1 / pchoices[z] if pred[i] == logchoices[z] else 0
ocrl.update(1, w, r)
for d in range(delay):
y = np.array([ logchoices[z]
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
if n == 0 and pno == 0:
sampweight = np.array([ lr
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
else:
sampweight = np.array([ lr * w * ocrl.n
* max(0, qfunc(1, w, 1))
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z]
for w in (1 / pchoices[z],)
])
effn += sampweight.sum() / (lr * subblocksize)
effnsl += sampweight.sum() / (lr * subblocksize)
if np.any(y):
x = vstack([ v[i]
for i, z in enumerate(ind)
if (d-1)*subblocksize <= i and i < d*subblocksize
if logchoices[z] == train.target[z] ])
cls.partial_fit(x, y, classes=classes, sample_weight=sampweight)
print('{:<8d}\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]\t{:<8.3f} [{:<10.3f}]'.format(
loss.n, effn.mean(), effnsl.mean(), avtau.mean(), avtausl.mean(), loss.mean(), sincelast.mean(), logpv.mean(), logpvsl.mean()),
flush=True)
sincelast = OnlineDRO.EasyAcc()
logpvsl = OnlineDRO.EasyAcc()
effnsl = OnlineDRO.EasyAcc()
avtausl = OnlineDRO.EasyAcc()
preds = cls.predict(testdocs)
ascores = []
for b in range(16):
bootie = np.random.RandomState(90210+b).choice(len(test.target), replace=True, size=len(test.target))
ascores.append(accuracy_score(y_true=test.target[bootie], y_pred=preds[bootie]))
print("test accuracy: {}".format(np.quantile(ascores, [0.05, 0.5, 0.95])))
except KeyboardInterrupt:
raise
except:
pass
OnlineDRO.flass()
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Classify Flowers with Transfer Learning
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/hub/tutorials/image_feature_vector"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/image_feature_vector.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/hub/blob/master/examples/colab/image_feature_vector.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/hub/examples/colab/image_feature_vector.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
<td>
<a href="https://tfhub.dev/google/imagenet/mobilenet_v2_035_128/feature_vector/2"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
</td>
</table>
Have you ever seen a beautiful flower and wondered what kind of flower it is? Well, you're not the first, so let's build a way to identify the type of flower from a photo!
For classifying images, a particular type of *deep neural network*, called a *convolutional neural network* has proved to be particularly powerful. However, modern convolutional neural networks have millions of parameters. Training them from scratch requires a lot of labeled training data and a lot of computing power (hundreds of GPU-hours or more). We only have about three thousand labeled photos and want to spend much less time, so we need to be more clever.
We will use a technique called *transfer learning* where we take a pre-trained network (trained on about a million general images), use it to extract features, and train a new layer on top for our own task of classifying images of flowers.
## Setup
```
import collections
import io
import math
import os
import random
from six.moves import urllib
from IPython.display import clear_output, Image, display, HTML
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
import tensorflow_hub as hub
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn.metrics as sk_metrics
import time
```
## The flowers dataset
The flowers dataset consists of images of flowers with 5 possible class labels.
When training a machine learning model, we split our data into training and test datasets. We will train the model on our training data and then evaluate how well the model performs on data it has never seen - the test set.
Let's download our training and test examples (it may take a while) and split them into train and test sets.
Run the following two cells:
```
FLOWERS_DIR = './flower_photos'
TRAIN_FRACTION = 0.8
RANDOM_SEED = 2018
def download_images():
"""If the images aren't already downloaded, save them to FLOWERS_DIR."""
if not os.path.exists(FLOWERS_DIR):
DOWNLOAD_URL = 'http://download.tensorflow.org/example_images/flower_photos.tgz'
print('Downloading flower images from %s...' % DOWNLOAD_URL)
urllib.request.urlretrieve(DOWNLOAD_URL, 'flower_photos.tgz')
!tar xfz flower_photos.tgz
print('Flower photos are located in %s' % FLOWERS_DIR)
def make_train_and_test_sets():
"""Split the data into train and test sets and get the label classes."""
train_examples, test_examples = [], []
shuffler = random.Random(RANDOM_SEED)
is_root = True
for (dirname, subdirs, filenames) in tf.gfile.Walk(FLOWERS_DIR):
# The root directory gives us the classes
if is_root:
subdirs = sorted(subdirs)
classes = collections.OrderedDict(enumerate(subdirs))
label_to_class = dict([(x, i) for i, x in enumerate(subdirs)])
is_root = False
# The sub directories give us the image files for training.
else:
filenames.sort()
shuffler.shuffle(filenames)
full_filenames = [os.path.join(dirname, f) for f in filenames]
label = dirname.split('/')[-1]
label_class = label_to_class[label]
# An example is the image file and it's label class.
examples = list(zip(full_filenames, [label_class] * len(filenames)))
num_train = int(len(filenames) * TRAIN_FRACTION)
train_examples.extend(examples[:num_train])
test_examples.extend(examples[num_train:])
shuffler.shuffle(train_examples)
shuffler.shuffle(test_examples)
return train_examples, test_examples, classes
# Download the images and split the images into train and test sets.
download_images()
TRAIN_EXAMPLES, TEST_EXAMPLES, CLASSES = make_train_and_test_sets()
NUM_CLASSES = len(CLASSES)
print('\nThe dataset has %d label classes: %s' % (NUM_CLASSES, CLASSES.values()))
print('There are %d training images' % len(TRAIN_EXAMPLES))
print('there are %d test images' % len(TEST_EXAMPLES))
```
## Explore the data
The flowers dataset consists of examples which are labeled images of flowers. Each example contains a JPEG flower image and the class label: what type of flower it is. Let's display a few images together with their labels.
```
#@title Show some labeled images
def get_label(example):
"""Get the label (number) for given example."""
return example[1]
def get_class(example):
"""Get the class (string) of given example."""
return CLASSES[get_label(example)]
def get_encoded_image(example):
"""Get the image data (encoded jpg) of given example."""
image_path = example[0]
return tf.gfile.GFile(image_path, 'rb').read()
def get_image(example):
"""Get image as np.array of pixels for given example."""
return plt.imread(io.BytesIO(get_encoded_image(example)), format='jpg')
def display_images(images_and_classes, cols=5):
"""Display given images and their labels in a grid."""
rows = int(math.ceil(len(images_and_classes) / cols))
fig = plt.figure()
fig.set_size_inches(cols * 3, rows * 3)
for i, (image, flower_class) in enumerate(images_and_classes):
plt.subplot(rows, cols, i + 1)
plt.axis('off')
plt.imshow(image)
plt.title(flower_class)
NUM_IMAGES = 15 #@param {type: 'integer'}
display_images([(get_image(example), get_class(example))
for example in TRAIN_EXAMPLES[:NUM_IMAGES]])
```
## Build the model
We will load a [TF-Hub](https://tensorflow.org/hub) image feature vector module, stack a linear classifier on it, and add training and evaluation ops. The following cell builds a TF graph describing the model and its training, but it doesn't run the training (that will be the next step).
```
LEARNING_RATE = 0.01
tf.reset_default_graph()
# Load a pre-trained TF-Hub module for extracting features from images. We've
# chosen this particular module for speed, but many other choices are available.
image_module = hub.Module('https://tfhub.dev/google/imagenet/mobilenet_v2_035_128/feature_vector/2')
# Preprocessing images into tensors with size expected by the image module.
encoded_images = tf.placeholder(tf.string, shape=[None])
image_size = hub.get_expected_image_size(image_module)
def decode_and_resize_image(encoded):
decoded = tf.image.decode_jpeg(encoded, channels=3)
decoded = tf.image.convert_image_dtype(decoded, tf.float32)
return tf.image.resize_images(decoded, image_size)
batch_images = tf.map_fn(decode_and_resize_image, encoded_images, dtype=tf.float32)
# The image module can be applied as a function to extract feature vectors for a
# batch of images.
features = image_module(batch_images)
def create_model(features):
"""Build a model for classification from extracted features."""
# Currently, the model is just a single linear layer. You can try to add
# another layer, but be careful... two linear layers (when activation=None)
# are equivalent to a single linear layer. You can create a nonlinear layer
# like this:
# layer = tf.layers.dense(inputs=..., units=..., activation=tf.nn.relu)
layer = tf.layers.dense(inputs=features, units=NUM_CLASSES, activation=None)
return layer
# For each class (kind of flower), the model outputs some real number as a score
# how much the input resembles this class. This vector of numbers is often
# called the "logits".
logits = create_model(features)
labels = tf.placeholder(tf.float32, [None, NUM_CLASSES])
# Mathematically, a good way to measure how much the predicted probabilities
# diverge from the truth is the "cross-entropy" between the two probability
# distributions. For numerical stability, this is best done directly from the
# logits, not the probabilities extracted from them.
cross_entropy = tf.nn.softmax_cross_entropy_with_logits_v2(logits=logits, labels=labels)
cross_entropy_mean = tf.reduce_mean(cross_entropy)
# Let's add an optimizer so we can train the network.
optimizer = tf.train.GradientDescentOptimizer(learning_rate=LEARNING_RATE)
train_op = optimizer.minimize(loss=cross_entropy_mean)
# The "softmax" function transforms the logits vector into a vector of
# probabilities: non-negative numbers that sum up to one, and the i-th number
# says how likely the input comes from class i.
probabilities = tf.nn.softmax(logits)
# We choose the highest one as the predicted class.
prediction = tf.argmax(probabilities, 1)
correct_prediction = tf.equal(prediction, tf.argmax(labels, 1))
# The accuracy will allow us to eval on our test set.
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
```
## Train the network
Now that our model is built, let's train it and see how it perfoms on our test set.
```
# How long will we train the network (number of batches).
NUM_TRAIN_STEPS = 100 #@param {type: 'integer'}
# How many training examples we use in each step.
TRAIN_BATCH_SIZE = 10 #@param {type: 'integer'}
# How often to evaluate the model performance.
EVAL_EVERY = 10 #@param {type: 'integer'}
def get_batch(batch_size=None, test=False):
"""Get a random batch of examples."""
examples = TEST_EXAMPLES if test else TRAIN_EXAMPLES
batch_examples = random.sample(examples, batch_size) if batch_size else examples
return batch_examples
def get_images_and_labels(batch_examples):
images = [get_encoded_image(e) for e in batch_examples]
one_hot_labels = [get_label_one_hot(e) for e in batch_examples]
return images, one_hot_labels
def get_label_one_hot(example):
"""Get the one hot encoding vector for the example."""
one_hot_vector = np.zeros(NUM_CLASSES)
np.put(one_hot_vector, get_label(example), 1)
return one_hot_vector
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for i in range(NUM_TRAIN_STEPS):
# Get a random batch of training examples.
train_batch = get_batch(batch_size=TRAIN_BATCH_SIZE)
batch_images, batch_labels = get_images_and_labels(train_batch)
# Run the train_op to train the model.
train_loss, _, train_accuracy = sess.run(
[cross_entropy_mean, train_op, accuracy],
feed_dict={encoded_images: batch_images, labels: batch_labels})
is_final_step = (i == (NUM_TRAIN_STEPS - 1))
if i % EVAL_EVERY == 0 or is_final_step:
# Get a batch of test examples.
test_batch = get_batch(batch_size=None, test=True)
batch_images, batch_labels = get_images_and_labels(test_batch)
# Evaluate how well our model performs on the test set.
test_loss, test_accuracy, test_prediction, correct_predicate = sess.run(
[cross_entropy_mean, accuracy, prediction, correct_prediction],
feed_dict={encoded_images: batch_images, labels: batch_labels})
print('Test accuracy at step %s: %.2f%%' % (i, (test_accuracy * 100)))
def show_confusion_matrix(test_labels, predictions):
"""Compute confusion matrix and normalize."""
confusion = sk_metrics.confusion_matrix(
np.argmax(test_labels, axis=1), predictions)
confusion_normalized = confusion.astype("float") / confusion.sum(axis=1)
axis_labels = list(CLASSES.values())
ax = sns.heatmap(
confusion_normalized, xticklabels=axis_labels, yticklabels=axis_labels,
cmap='Blues', annot=True, fmt='.2f', square=True)
plt.title("Confusion matrix")
plt.ylabel("True label")
plt.xlabel("Predicted label")
show_confusion_matrix(batch_labels, test_prediction)
```
## Incorrect predictions
Let's a take a closer look at the test examples that our model got wrong.
- Are there any mislabeled examples in our test set?
- Is there any bad data in the test set - images that aren't actually pictures of flowers?
- Are there images where you can understand why the model made a mistake?
```
incorrect = [
(example, CLASSES[prediction])
for example, prediction, is_correct in zip(test_batch, test_prediction, correct_predicate)
if not is_correct
]
display_images(
[(get_image(example), "prediction: {0}\nlabel:{1}".format(incorrect_prediction, get_class(example)))
for (example, incorrect_prediction) in incorrect[:20]])
```
## Exercises: Improve the model!
We've trained a baseline model, now let's try to improve it to achieve better accuracy. (Remember that you'll need to re-run the cells when you make a change.)
### Exercise 1: Try a different image model.
With TF-Hub, trying a few different image models is simple. Just replace the `"https://tfhub.dev/google/imagenet/mobilenet_v2_050_128/feature_vector/2"` handle in the `hub.Module()` call with a handle of different module and rerun all the code. You can see all available image modules at [tfhub.dev](https://tfhub.dev/s?module-type=image-feature-vector).
A good choice might be one of the other [MobileNet V2 modules](https://tfhub.dev/s?module-type=image-feature-vector&network-architecture=mobilenet-v2). Many of the modules -- including the MobileNet modules -- were trained on the [ImageNet dataset](http://image-net.org/challenges/LSVRC/2012/index#task) which contains over 1 million images and 1000 classes. Choosing a network architecture provides a tradeoff between speed and classification accuracy: models like MobileNet or NASNet Mobile are fast and small, more traditional architectures like Inception and ResNet were designed for accuracy.
For the larger Inception V3 architecture, you can also explore the benefits of pre-training on a domain closer to your own task: it is also available as a [module trained on the iNaturalist dataset](https://tfhub.dev/google/inaturalist/inception_v3/feature_vector/1) of plants and animals.
### Exercise 2: Add a hidden layer.
Stack a hidden layer between extracted image features and the linear classifier (in function `create_model()` above). To create a non-linear hidden layer with e.g. 100 nodes, use [tf.layers.dense](https://www.tensorflow.org/api_docs/python/tf/compat/v1/layers/dense) with units set to 100 and activation set to `tf.nn.relu`. Does changing the size of the hidden layer affect the test accuracy? Does adding second hidden layer improve the accuracy?
### Exercise 3: Change hyperparameters.
Does increasing *number of training steps* improves final accuracy? Can you *change the learning rate* to make your model converge more quickly? Does the training *batch size* affect your model's performance?
### Exercise 4: Try a different optimizer.
Replace the basic GradientDescentOptimizer with a more sophisticate optimizer, e.g. [AdagradOptimizer](https://www.tensorflow.org/api_docs/python/tf/compat/v1/train/AdagradOptimizer). Does it make a difference to your model training? If you want to learn more about the benefits of different optimization algorithms, check out [this post](http://ruder.io/optimizing-gradient-descent/).
## Want to learn more?
If you are interested in a more advanced version of this tutorial, check out the [TensorFlow image retraining tutorial](https://www.tensorflow.org/hub/tutorials/image_retraining) which walks you through visualizing the training using TensorBoard, advanced techniques like dataset augmentation by distorting images, and replacing the flowers dataset to learn an image classifier on your own dataset.
You can learn more about TensorFlow at [tensorflow.org](http://tensorflow.org) and see the TF-Hub API documentation is available at [tensorflow.org/hub](https://www.tensorflow.org/hub/). Find available TensorFlow Hub modules at [tfhub.dev](http://tfhub.dev) including more image feature vector modules and text embedding modules.
Also check out the [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/) which is Google's fast-paced, practical introduction to machine learning.
| github_jupyter |
# DRF of CNS-data
```
%load_ext autoreload
%autoreload 2
%matplotlib notebook
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import networkx as nx
from matplotlib.colors import LogNorm
from sklearn.utils import shuffle
from itertools import groupby
from matplotlib.figure import figaspect
# package developed for the analysis
from world_viewer.cns_world import CNSWorld
from world_viewer.glasses import Glasses
# DTU Data Wrapper
from sensible_raw.loaders import loader
```
## 1) Load and Prepare Data
```
# load data for analysis
cns = CNSWorld()
cns.load_world(opinions = ['fitness'], read_cached = False, stop=False, write_pickle = False, continous_op=False)
# load analysis tools
cns_glasses = Glasses(cns)
# remove not needed data in order to save mermory
cns.d_ij = None
# set analysis parameters
analysis = 'expo_frac'
opinion_type = "op_fitness"
binning = True
n_bins = 10
save_plots = True
show_plot = True
4.1_CopenhagenDataRelativeExposure.ipynb# load previously calculated exposure instead of recalculate it
exposure = pd.read_pickle("tmp/fitness_exposure_tx7.pkl")
# alternative: recalculate exposure value
# exposure = cns_glasses.calc_exposure("expo_frac", "op_fitness", exposure_time = 7)
# filter by degre
degree = exposure.groupby("node_id").n_nbs.mean().to_frame("avg").reset_index()
exposure = exposure.loc[degree.loc[degree.avg >= 4,"node_id"]]
exposure = exposure.loc[exposure.n_nbs_mean > 1/7]
# cut time series in time slices: spring + summer
exposure.reset_index(inplace=True)
start_spring = "2014-02-01"
end_spring = "2014-04-30"
exposure_spring = exposure.loc[(exposure.time >= pd.to_datetime(start_spring)) & (exposure.time <= pd.to_datetime(end_spring))].copy()
start_summer = "2014-07-01"
end_summer = "2014-09-30"
exposure_summer = exposure.loc[(exposure.time >= pd.to_datetime(start_summer)) & (exposure.time <= pd.to_datetime(end_summer))].copy()
exposure_spring.set_index(['node_id','time'],inplace=True)
exposure_summer.set_index(['node_id','time'],inplace=True)
exposure.set_index(['node_id','time'],inplace=True)
# column "exposure" equals relative exposure
# column "n_influencer_summed" equals absolute exposure
# use absolute exposure for further calculations
exposure.rename(columns={"exposure":"exposure_old", "n_influencer_summed":"exposure"},inplace=True)
exposure_spring.rename(columns={"exposure":"exposure_old", "n_influencer_summed":"exposure"},inplace=True)
exposure_summer.rename(columns={"exposure":"exposure_old", "n_influencer_summed":"exposure"},inplace=True)
# calculate if nodes changed trait after experiencing a certain exposure
# save value as column "op_change" (bool)
data_spring, expo_agg_spring = cns_glasses.opinion_change_per_exposure(exposure_spring, opinion_type, opinion_change_time = 1)
data_summer, expo_agg_summer = cns_glasses.opinion_change_per_exposure(exposure_summer, opinion_type, opinion_change_time = 1)
data_full, expo_agg_full = cns_glasses.opinion_change_per_exposure(exposure, opinion_type, opinion_change_time = 1)
# save calculated values on hard drive
expo_agg_spring.to_pickle("tmp/final/exposure_filtered_spring.pkl")
```
## 2) Plot Dose Response Functions (FIG.: 4.9)
```
# plot drf for full timeseries
fig, ax = plt.subplots(1,2,subplot_kw = {"adjustable":'box', "aspect":200/0.25})
cns_glasses.output_folder = "final/"
suffix = "_full"
data = data_full[data_full.exposure <= 200]
bin_width=1
q_binning=True
bin_width=5
n_bins=15
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == True], "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=False, show_plot=show_plot, y_lower_lim=0, y_upper_lim = 0.2, fig=fig, ax=ax[0], label="become active", q_binning = q_binning, \
loglog=False, step_plot=True, color="forestgreen", suffix=suffix,x_lim=200)
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == False], "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=True, show_plot=show_plot, y_lower_lim=0, y_upper_lim = 0.2, fig=fig, ax=ax[1], label="become passive", loglog=False, \
q_binning=q_binning, step_plot=True, color="darkorange", suffix=suffix,x_lim=200)
# plot drf for summer timeseries
fig, ax = plt.subplots(1,2,subplot_kw = {"adjustable":'box', "aspect":200/0.25})
cns_glasses.output_folder = "final/"
suffix = "_summer"
data = data_summer[data_summer.exposure <= 200]
q_binning=False
bin_width=15
n_bins=20
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == True].dropna(), "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=False, show_plot=show_plot, y_lower_lim=-0.01, y_upper_lim = 0.25, fig=fig, ax=ax[0], label="become active", q_binning = q_binning, loglog=False, step_plot=True, color="forestgreen", suffix=suffix,x_lim=200)
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == False].dropna(), "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=True, show_plot=show_plot, y_lower_lim=-0.01, y_upper_lim = 0.25, fig=fig, ax=ax[1], label="become passive", loglog=False, q_binning=q_binning, step_plot=True, color="darkorange", suffix=suffix,x_lim=200)
# plot drf for spring timeseries
x_max = 330
w, h = figaspect(0.5)
fig, ax = plt.subplots(1,2,figsize=(w,h))
cns_glasses.output_folder = "final/"
suffix = "_spring"
data = data_spring[data_spring.exposure <= x_max]
q_binning=False
bin_width=15
n_bins=15
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == True], "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=False, show_plot=show_plot, y_lower_lim=-0.01, y_upper_lim = 0.2, fig=fig, ax=ax[0], label="become active", \
q_binning = q_binning, loglog=False, step_plot=True, color="forestgreen", suffix=suffix, x_lim=x_max)
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == False], "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=True, show_plot=show_plot, y_lower_lim=-0.01, y_upper_lim = 0.2, fig=fig, ax=ax[1], label="become passive", loglog=False, \
q_binning=q_binning, step_plot=True, color="darkorange", suffix=suffix, x_lim=x_max)
fig.savefig("tmp/final/empirical_drfs.pdf" , bbox_inches='tight')
x_max = 330
fig, ax = plt.subplots()
cns_glasses.output_folder = "final/"
suffix = "_spring"
data = data_spring[data_spring.exposure <= x_max]
q_binning=False
bin_width=15
n_bins=15
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == True], "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=False, show_plot=show_plot, y_lower_lim=-0.01, y_upper_lim = 0.2, fig=fig, ax=ax, label="become active", \
q_binning = q_binning, loglog=False, step_plot=True, color="forestgreen", suffix=suffix, x_lim=x_max,marker="^", markersize=5)
fig.savefig("tmp/final/empirical_drf_1.pdf" , bbox_inches='tight')
cns_glasses.plot_opinion_change_per_exposure_number(data[data.op_fitness == False], "expo_nmb", binning, n_bins=n_bins, bin_width=bin_width, \
save_plots=True, show_plot=show_plot, y_lower_lim=-0.01, y_upper_lim = 0.2, fig=fig, ax=ax, label="become passive", loglog=False, \
q_binning=q_binning, step_plot=True, color="darkorange", suffix=suffix, x_lim=x_max, marker=".")
fig.savefig("tmp/final/empirical_drf_2.pdf" , bbox_inches='tight')
```
## 3) Plot Distribution of the Absolute Exposure (FIG.: 4.10)
```
expo = expo_agg_spring[expo_agg_spring.op_fitness==True].reset_index()
#expo = expo.loc[(expo.time > "2013-09-01") & (expo.time < "2014-09-01")]
#expo.time = expo.time.dt.dayofyear
expo.time = expo.time.astype("int")
mean_expo = expo.groupby("time").exposure.mean().to_frame("mean exposure").reset_index()
mean_expo.set_index("time",inplace=True)
fig,ax = plt.subplots()
expo.dropna(inplace=True)
#expo = expo[expo.exposure < 250]
plot = ax.hist2d(expo.time,expo.exposure,norm=LogNorm(), bins = [len(expo.time.unique())
,120])#, vmin=1, vmax=100)
expo.groupby("time").exposure.mean().plot(label=r"mean exposure $<K>$",color="red",linestyle="--")
ax.legend(loc="upper left")
ax.set_xlabel("time")
ax.set_ylabel(r"absolute exposure $K$")
ax.set_xticklabels(pd.to_datetime(ax.get_xticks()).strftime('%d. %B %Y'), rotation=40, ha="right")
cbar = fig.colorbar(plot[3])
cbar.set_label('# number of occurrences')
fig.savefig("tmp/final/abs_expo_distrib_spring.pdf",bbox_inches='tight')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/broker-workshop/tutorials/blob/main/Pitt-Google/Pitt-Google-Tutorial-Code-Samples.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Pitt-Google Broker Tutorial: Code Samples
| | Learning Objective | Section | Notes |
|---|---|---|---|
| 1. | Understand What, Where, How to access | 1) Introduction | ZTF data. Google Cloud SDK (Python, command-line). `pgb-utils` Python package. |
| 2. | Access the data | 3) BigQuery Database | Alert data (no cutouts), lightcurves, cone search. Standard SQL queries (Python, command-line). `pgb-utils` helper functions. |
| | | 4) Files in Cloud Storage | Alert packets in Avro file format (includes cutouts). Direct downloads. |
| 3. | Process the data | 5) Apache Beam data pipelines | End-to-end, working examples with templates for user-defined functions. |
---
This "Code Samples" notebook is a condensed version of the full [Pitt-Google-Tutorial.ipynb](https://colab.research.google.com/github/broker-workshop/tutorials/blob/main/Pitt-Google/Pitt-Google-Tutorial.ipynb), which you can refer to for further information.
---
# 1) Introduction
## 1a) Data overview
See full tutorial.
## 1b) `pgb_utils` overview
See full tutorial.
## 1c) Note on costs
Everything we do in this tutorial falls well within Google's [Free Tier](https://cloud.google.com/free). See full tutorial for more information.
---
# 2) Setup
1. Use the [Cloud Resource Manager](https://console.cloud.google.com/cloud-resource-manager) to create a GCP project.
It is a simple process, and you do not need to enable billing of any kind. Take note of the auto-generated "Project ID", you will need it below in order to make API calls.
2. Installs, imports, etc.:
```
pgb_project_name = 'pitt-google-broker-prototype'
pgb_project_id = 'ardent-cycling-243415'
# ENTER YOUR GCP PROJECT ID HERE
my_project_id =
!{'pip install pgb-utils'}
import apache_beam as beam
import aplpy
from astropy import coordinates as coord
from astropy import units as u
from astropy.io import fits
import fastavro
from google.colab import auth, drive
import gzip
import io
from matplotlib import pyplot as plt
import pandas as pd
from pathlib import Path
from google.cloud import bigquery, storage
import pgb_utils as pgb
auth.authenticate_user()
# follow the instructions to authorize Google Cloud SDK
# For demonstration of command-line tools, create a fnc for running shell commands
def run(cmd: str):
print('>> {}'.format(cmd))
!{cmd}
print('')
# Connect your Google Drive file system
# to be used in the sections on File Storage and Apache Beam pipelines
drive.mount('/content/drive')
# follow the instructions to authorize access
# create a path for later
colabpath = '/content/drive/MyDrive/Colab\ Notebooks/PGB_tutorial'
colabpath_noesc = '/content/drive/MyDrive/Colab Notebooks/PGB_tutorial'
run(f'mkdir -p {colabpath}')
# Colab Hint: Click the "Files" icon on the left to view a file browser.
# Colab Hint: Click the "Code snippets" icon (<>) on the left and search for
# "access drive" to learn how to interact with Drive.
```
---
# 3) BigQuery Database
## 3a) Python
### Table names and schemas
```
# Create a Client for the BigQuery connections below
pgb.bigquery.create_client(my_project_id)
# this is just a convenience wrapper, as are many pgb functions. look at
# its source code for guidance on using `google.cloud` libraries directly.
# Colab Hint: Mouse over the function name to see its definition and
# source code (make sure the cell is selected).
pgb.bigquery.get_dataset_table_names()
pgb.bigquery.get_table_info('DIASource')
# Colab Hint: Right-click this cell and select "Copy to scratch cell"
# so you can use this as a reference in later queries.
```
---
### Query lightcurves and other history
```
# Choose the history data you want returned
columns = ['jd', 'fid', 'magpsf', 'sigmapsf']
# 'objectId' and 'candid' will be included automatically
# options are in the 'DIASource' table
# pgb.bigquery.get_table_info('DIASource')
# Choose specific objects (optional. we'll choose some to minimize load time)
objectIds = ['ZTF18aczuwfe', 'ZTF18aczvqcr', 'ZTF20acqgklx', 'ZTF18acexdlh']
# you can also restrict sample size using the `limit` keyword below
```
To retrieve lightcurves and other history, we must query for objects' "DIASource" observations and aggregate the results by `objectId`.
`pgb.bigquery.query_objects()` is a convenience wrapper that let's you grab all the results at once, or step through them using a generator.
It's options are demonstrated below.
```
# Option 1: Get a single DataFrame of all results
lcs_df = pgb.bigquery.query_objects(columns, objectIds=objectIds)
# This will execute a dry run and tell you how much data will be processed.
# You will be asked to confirm before proceeding.
# In the future we'll skip this using
dry_run = False
lcs_df.sample(10)
# cleaned of duplicates
```
Congratulations! You've now retrieved your first data from the transient table.
It is a DataFrame containing the candidate observations for every object we requested, indexed by `objectId` and `candid` (candidate ID). It includes the columns we requested in the query.
Queries can return large datasets. You may want to use a generator to step through objects individually, and avoid loading the entire dataset into memory at once.
`query_objects()` can return one for you:
```
# Option 2: Get a generator that yields a DataFrame for each objectId
iterator = True
objects = pgb.bigquery.query_objects(columns,
objectIds=objectIds,
iterator=iterator,
dry_run=dry_run
)
# cleaned of duplicates
for lc_df in objects:
print(f'\nobjectId: {lc_df.objectId}') # objectId in metadata
print(lc_df.sample(5))
```
Each DataFrame contains data on a single object, and is indexed by `candid`. The `objectId` is in the metadata.
`query_objects()` can also return a json formatted strings of the query results, or the raw `query_job` object that is returned by the Google Cloud SDK. See the full tutorial for details.
---
#### Plot a lightcurve
```
# Get an object's lightcurve DataFrame with the minimum required columns
columns = ['jd','fid','magpsf','sigmapsf','diffmaglim']
objectId = 'ZTF20acqgklx'
lc_df = pgb.bigquery.query_objects(columns, objectIds=[objectId], dry_run=False)
# make the plot
pgb.figures.plot_lightcurve(lc_df, objectId=objectId)
# this function was adapted from:
# https://github.com/ZwickyTransientFacility/ztf-avro-alert/blob/master/notebooks/Filtering_alerts.ipynb
```
---
### Cone Search
See full tutorial.
---
### Direct access: Google Cloud SDK
See full tutorial.
---
## 3b) Command-line tool `bq`
All commands in this section are executed using the `run` function we created earlier to run commands on the command-line.
```
# Get help
run('bq help query')
# view the schema of a table
run('bq show --schema --format=prettyjson ardent-cycling-243415:ztf_alerts.DIASource')
# run('bq show --schema --format=prettyjson ardent-cycling-243415:ztf_alerts.alerts')
# Note: The first time you make a call with `bq` you will ask you to
# initialize a .bigqueryrc configuration file. Follow the directions.
# Query
bq_query = """bq query \
--use_legacy_sql=false \
'SELECT
objectId, candid, t0, x0, x1, c, chisq, ndof
FROM
`ardent-cycling-243415.ztf_alerts.salt2`
WHERE
ndof>0 and chisq/ndof<2
LIMIT
10'
"""
run(bq_query)
```
---
# 4) Files in Cloud Storage
## 4a) Python
See full tutorial.
### Download files
### Plot cutouts and lightcurves
## 4b) Command-line tool `gsutil`
See full tutorial.
---
# 5) Apache Beam data pipelines
[Apache Beam](https://beam.apache.org/) is an SDK that facilitates writing and executing data pipelines. The [Apache Beam Programming Guide](https://beam.apache.org/documentation/programming-guide/) is very useful!
In this "Code Samples" notebook, we demonstrate working examples that retrieve and process ZTF data and write out the results. The full tutorial has much more information on basic concepts and some of the many available options.
---
## 5a) A demo example
See full tutorial.
---
## 5b) Descriptions
See full tutorial.
---
## 5c) Pitt-Google working examples
In these examples, we will query the database for object histories (lightcurves) and cast them to DataFrames, apply a filter and a processing function, and write the results to a text file.
Note that Beam _overloads_ some operators:
- `|` means `apply`
- `>>` allows you to name the step with the preceeding string. It is optional. We use it here to improve readability. Various UIs like Dataflow use it in their displays.
Here's a preview of the pipeline we will create and run.
Pipeline:
```python
with beam.Pipeline() as pipeline:
(
pipeline
| 'Read from BigQuery' >> beam.io.ReadFromBigQuery(**read_args)
| 'Type cast to DataFrame' >> beam.ParDo(pgb.beam.ExtractHistoryDf())
| 'Is nearby known SS object' >> beam.Filter(nearby_ssobject)
| 'Calculate mean magnitudes' >> beam.ParDo(calc_mean_mags())
| 'Write results' >> beam.io.WriteToText(beam_outputs_prefix)
)
```
With comments:
```python
# 0. Instantiate a pipeline object
with beam.Pipeline() as pipeline:
(
# 1. Start the pipeline by piping it to a "read" function. Then format the incoming data as a DataFrame.
pipeline
| 'Read from BigQuery' >> beam.io.ReadFromBigQuery(**read_args)
| 'Type cast to DataFrame' >> beam.ParDo(pgb.beam.ExtractHistoryDf())
# 2. PROCESS the data with USER-DEFINED FUNCTIONS
# Apply a Filter
| 'Is nearby known SS object' >> beam.Filter(nearby_ssobject)
# Apply a processing function
| 'Calculate mean magnitudes' >> beam.ParDo(calc_mean_mags())
# 3. Output the results
| 'Write results' >> beam.io.WriteToText(beam_outputs_prefix)
)
```
First, let's define our user-defined functions: a filter and a processing function.
Both will take as input `lc_df`, a single object's lightcurve (or history) DataFrame.
_The functions will work as-is, or you can use them as templates to create your own._
_Colab Hint_: Right-click on one of the code cells with a function definition and select "Copy to scratch cell". Use the new scratch cell to change the function and experiment with the pipeline.
```
# Filter for likely solar system objects
def nearby_ssobject(lc_df):
"""Keep only objects that are within 5" of a known solar system object.
To be called with `beam.Filter(nearby_ssobject)`.
"""
ssdistnr = lc_df['ssdistnr'].mean()
ssobject_is_near = (ssdistnr > 0) and (ssdistnr < 5)
return ssobject_is_near
# generally: return a bool where `True` means we keep this df, else drop it
# Processing: Calculate and return the mean magnitude per passband
class calc_mean_mags(beam.DoFn):
"""Class that wraps our `process()` function to calculate mean magnitudes.
To be called with `beam.ParDo(calc_mean_mags())`.
"""
def process(self, lc_df):
"""Calculate mean magnitudes per passband."""
meanmags = lc_df[['fid','magpsf']].groupby('fid').mean()
# we will write this to a file, so let's format it nicely
output = []
for fid, row in meanmags.iterrows():
output.append(f"{lc_df.objectId},{fid},{row['magpsf']}")
return output
# generally: return a list containing 0 or more elements, each of which
# becomes an element in the `ParDo`'s output collection.
```
Now let's configure and run specific pipelines.
### Lightcurve pipeline
Set configs
See the "BigQuery Database" section for options to generate the query statement, or write your own SQL.
Use `pgb.bigquery.get_table_info('DIASource')` in a scratch cell to view options for column names.
```
# keyword args for `ReadFromBigQuery()`
columns = ['jd', 'fid', 'magpsf', 'sigmapsf', 'ssdistnr']
limit = 2000 # just to reduce runtime
query = pgb.bigquery.object_history_sql_statement(columns, limit=limit) # str
read_args = {
'query': query,
'project': my_project_id,
'use_standard_sql': True,
'gcs_location': f'gs://{pgb_project_id}-workshop_beam_test',
# courtesy location for temp files, available for the workshop duration
}
# path to write the results
outputs_prefix = f'{colabpath}/outputs/meanmags'
beam_outputs_prefix = f'{colabpath_noesc}/outputs/meanmags'
```
Define and run the pipeline
```
with beam.Pipeline() as pipeline:
(
pipeline
| 'Read BigQuery' >> beam.io.ReadFromBigQuery(**read_args)
| 'Extract lightcurve df' >> beam.ParDo(pgb.beam.ExtractHistoryDf())
| 'Nearby SS object' >> beam.Filter(nearby_ssobject)
| 'Calc mean mags' >> beam.ParDo(calc_mean_mags())
| 'Write results' >> beam.io.WriteToText(beam_outputs_prefix)
)
```
Congratulations! You just ran an Apache Beam pipeline that fetches ZTF data, filters and processes it, and writes out the results!
Let's look at the first 10 results. Remember, there are no ordering guarantees.
```
run('head -n 10 {}-00000-of-*'.format(outputs_prefix))
```
### Cone Search pipeline
See full tutorial.
| github_jupyter |
# Example data analysis notebook
This notebook downloads and analyses some surface air temperature anomaly data from [Berkeley Earth](http://berkeleyearth.org/).
Import the required libraries.
```
import matplotlib.pyplot as plt
import pandas as pd
import requests
```
Use the [requests](http://docs.python-requests.org/) library to download the data file for Australia.
```
# URL to the data
url = 'http://berkeleyearth.lbl.gov/auto/Regional/TAVG/Text/australia-TAVG-Trend.txt'
```
## Open & Clean the Data
To begin there are some data cleaning steps that you need to implement here.
Often when you are building a workflow you build it out in it's entirety first
to get the pieces working. And then you turn those pieces into functions to
modularize and scale your workflow
```
temp_df = pd.read_csv(url,
skiprows=69,
delim_whitespace=True)
all_cols = temp_df.columns[1:]
# Remove the last row
temp_df = temp_df.iloc[:, :-1]
# CLEANUP: Drop the commas from the column names & Add a day column
temp_df.columns = [acol.replace(',', '') for acol in all_cols]
temp_df = temp_df.assign(Day=1)
# Finally create a date time column
temp_df["date"] = pd.to_datetime(temp_df[['Year', 'Month', 'Day']])
temp_df.set_index("date", inplace=True)
temp_df
```
# Initial Visualization
Plot the data to explore it!
```
f, ax = plt.subplots(figsize=(10, 6))
temp_df.plot(y="Anomaly",
ax=ax,
legend=False,
color="Purple")
ax.set(title="Temperature anomaly for Australia",
xlabel='Date',
ylabel='Temperature anomaly (C)')
plt.grid()
plt.show()
```
## Calculate Moving Average
Next you calculate a 12-month moving average for a smoother time series.
```
temp_moving_avg = temp_df["Anomaly"].rolling(window=12).mean()
temp_moving_avg
```
## Plot the Data
```
f, ax = plt.subplots(figsize=(10, 6))
temp_df.plot(y="Anomaly",
ax=ax,
legend=True,
color="DarkGrey",
label="Anomaly")
temp_moving_avg.plot(ax=ax,
legend=True,
color="Purple",
linewidth=2,
label="Moving average")
ax.set(title="Temperature Anomaly for Australia",
xlabel='Date',
ylabel='Temperature anomaly (C)')
plt.grid()
plt.show()
```
## Modularize Your Workflow
The function below may be bigger than you want. It is a
starting place.
```
from agu_oss import open_and_clean
help(open_and_clean)
url = 'http://berkeleyearth.lbl.gov/auto/Regional/TAVG/Text/australia-TAVG-Trend.txt'
temp_australia = open_and_clean(url)
temp_australia.head()
f, ax = plt.subplots(figsize=(10, 6))
temp_australia.plot(y="Anomaly",
color="purple",
ax=ax)
plt.show()
```
## Try this on another location
http://berkeleyearth.lbl.gov/country-list/
http://berkeleyearth.lbl.gov/auto/Regional/TAVG/Text/belize-TAVG-Trend.txt
```
url_belize = "http://berkeleyearth.lbl.gov/auto/Regional/TAVG/Text/belize-TAVG-Trend.txt"
temp_belize = open_and_clean(url_belize)
f, ax = plt.subplots(figsize=(10, 6))
temp_belize.plot(y="Anomaly",
color="purple",
ax=ax)
plt.show()
```
| github_jupyter |
```
# 그래프, 수학 기능 추가
# Add graph and math features
import pylab as py
```
# 순차법<br>Sequential Method
10의 제곱근을 구한다고 생각해 보자.<br>Let's try to find the square root of 10.
$$
x=\sqrt{10} = 10 ^ \frac{1}{2} = 10 ^ {0.5}
$$
계산기라면 이런 식으로 구할 수 있을 것이다.<br>With a calcuator, an engineer can find it as follows.
```
print('sqrt(10) =', 10 ** 0.5)
```
조금 다른 방식으로 해 보자.<br>Let's try a different way.
양변을 제곱해 보자.<br>Let's square both sides.
$$
\begin{align}
x^2 &= \left(10 ^ {0.5}\right)^2 \\
x^2 &= 10
\end{align}
$$
이 관계를 그래프로 표현해 보면 다음과 같을 것이다.<br>An engineer can visualize this relationship as follows.
```
# x 의 범위와 간격을 지정
# Specify range and interval of x
x = py.arange(-5, 5, 0.2)
# y = x^2
py.plot(x, x**2, 'k.')
# y = 10
py.plot(x, 10*py.ones_like(x), 'r.')
```
양변에서 10을 빼 보자.<br>Let's subtract 10 from both sides.
$$
\begin{align}
x^2-10 &= 10-10\\
x^2-10 &= 0
\end{align}
$$
이 관계도 그려보자.<br>Let's plot this, too.
```
# x 의 범위와 간격을 지정
# Specify range and interval of x
x = py.arange(-5, 5, 0.2)
# y = x^2
py.plot(x, x**2 - 10, 'k.')
# y = 0
py.plot(x, py.zeros_like(x), 'r.')
# 모눈 표시
# Indicate grid
py.grid()
```
위 방정식을 만족하는 $x$ 는 10의 제곱근일 것이다.<br>$x$ satisfying the equation above would be the square root of 10.
이러한 $x$를 위 방정식의 **근** 이라고 부른다.<br>We call such $x$ a **root** of the equation above.
그리고 그러한 x는 아래와 같은 함수를 0으로 만들 것이다.<br>And such $x$ would make following function zero.
$$
f(x) = x^2 - 10=0
$$
```
# 이 함수를 0으로 만드는 x 를 찾고자 함
# Intend to find x making the function below zero
def f(x):
return x ** 2 - 10
```
컴퓨터의 연산능력을 이용하여 근을 찾아 보도록 하자.<br>Let's try to find the root using the computer's computation capability.
여러 가능한 방법 가운데 하나는 $x$를 어떤 *초기값*으로부터 시작하여 일정 *간격* 으로 증가시키면서 $f(x)=x^2-10$을 계산해 보는 것이다.<br>
One of the possible ways is to evaluate $f(x)=x^2-10$ starting from an *initial value* of $x$, increasing by a constant *interval*.
그렇게 반복하다가 $\left|f(x)\right|=\left|x^2-10\right|$ 이 어떤 *만족스러운 수준* 이하가 되면, 그 때의 $x$ 값이 $\sqrt{10}$의 *근사값*이 될 것이다.<br>
During the iteration, the $x$ making $\left|f(x)\right|=\left|x^2-10\right|$ below a certain *satisfactory level* would be the *approximation* of $\sqrt{10}.$
$$
\left(\left|x^2-10\right|<\epsilon \right) \equiv \left(\left|x^2-10\right|\approx 0 \right)
$$
$$
\left(\left|f(x)\right|<\epsilon \right) \equiv \left(\left|f(x)\right|\approx 0 \right)
$$
여기서 $\epsilon$ 값의 의미는, 예를 들어 $\left|y\right| < \epsilon$ 이면 $y \approx 0$ 이라고 보는 것이다.<br>
Here, $\epsilon$ means that if $\left|y\right| < \epsilon$ then $y \approx 0$.
## 왜 $\epsilon$ 이 필요한가?<br>Why do we need $\epsilon$?
위의 그래프의 일부를 확대해보자.<br>Let's zoom into the plot above.
```
# x 의 범위와 간격을 지정
# Specify range and interval of x
x = py.arange(2.8, 3.5, 0.2)
# y = f(x) = x * x - 10
py.plot(x, f(x), 'ko')
# y = 0
py.plot(x, py.zeros_like(x), 'ro')
# x 축 이름표
# x axis label
py.xlabel('x')
# 모눈 표시
# Indicate grid
py.grid()
```
위 그림에서 $|x^2-10|$이 $x=3.0$과 $x=3.2$ 사이 어딘가에서 0이 될 것임을 짐작할 수 있다.<br>
We can see that $|x^2-10|$ will be zero somewhere between $x=3.0$ and $x=3.2$.
그 값을 더 정확히 알고 싶다면 점을 그리는 간격을 줄이면 될 것이다.<br>
To find more precise value, we can reduce the interval of the dots.
```
# x 의 범위와 간격을 지정
# Specify range and interval of x
x = py.arange(2.8, 3.5, 0.05)
# y = f(x) = x * x - 10
py.plot(x, f(x), 'ko')
# y = 0
py.plot(x, py.zeros_like(x), 'ro')
# x 축 이름표
# x axis label
py.xlabel('x')
# 모눈 표시
# Indicate grid
py.grid()
```
사람의 경우는 그 간격을 무한히 줄여서 점이 **연속적**이 되는 것을 생각할 수 있지만, 컴퓨터는 **이산적**인 그래프까지만 그릴 수 있다.<br>
A person may think infinitely small interval so that the dots become **continuous**, however, computer plots can only be **discrete**.
컴퓨터는 **정밀도**라는 어떤 수준 이하로 점을 그리는 간격을 줄일 수 없기 때문이다.<br>Because computers are unable to make the interval of the dots smaller than a certain level called **precision**.
사람은 $x^2-10$ 을 0으로 만들 수 있는 $x$ 를 생각할 수 있지만, 컴퓨터가 찾아 낼 수 있는 최선의 $\sqrt{10}$의 근사값으로도 $x^2-10$ 이 정확히 0이 되지 않을 수 있는 것이다.<br>
While a person may think about $x$ making $x^2-10$ exact zero, even computer's best approximation of $\sqrt{10}$ may not exactly make $x^2-10$ zero.
그래서 컴퓨터의 계산 결과 절대값이 **$\epsilon$** 값보다 작으면, 그 결과는 0과 같은 것으로 생각할 것이다.<br>
Hence, if a computation result has absolute value smaller then **$\epsilon$**, we would regard that the result is the same as zero.
```
import plot_root_finding
x_start = 2.8
x_end = 3.5
x_interval = 0.05
plot_root_finding.plot(f, x_start, x_end, x_interval);
```
### 순차법 구현<br>Implementing Sequantial Method
아래 python 프로그램은 순차법으로 $\sqrt{10}$을 계산한다<br>Following python script calculates $\sqrt{10}$ using sequential method.
```
%%time
# y_i 의 절대값이 이 값 보다 작으면 y_i = 0으로 본다
# If absolute value of y_i is smaller than this, we would think y_i = 0.
epsilon = 1e-3
# sqrt_10 의 초기값
# Initial value for sqrt_10
sqrt_10 = 'Not Found'
# 순차법의 매개변수
# Parameters of the Sequantial Method
# x_i 의 초기값
# Initial value of x_i
x_init = 0
# x_i 의 마지막 값
# Final value of x_i
x_final = 4
# i 번째 x 값과 i+1 번째 x 값 사이의 간격
# The interval between i'th and (i+1)'th x's
x_interval = 1e-5
# 일련의 x_i 값을 미리 준비한다
# Prepare a series of x_i values in advance
x_array = py.arange(x_init, x_final+x_interval*0.5, x_interval)
# 몇번 반복했는지 측정해 보자
# Let's count the number of iterations
counter = 0
# x_i 에 관한 반복문
# x_i loop
for x_i in x_array:
# Evaluate the function
y_i = x_i ** 2 - 10
counter += 1
# Check if absolute value is smaller than epsilon
if abs(y_i) < epsilon:
sqrt_10 = x_i
# found
break
# 반복 횟수
# Number of iterations
print('counter =', counter)
# 순차법으로 찾은 10의 제곱근
# Square root of 10 that we just found using the sequential method
print('sqrt_10 =', sqrt_10)
# 아래 연산의 결과는 0인가?
# Is the result of the following calculation zero?
print('sqrt_10 ** 2 - 10 =', sqrt_10 ** 2 - 10)
```
`epsilon`, `x_init`, `x_interval` 등 매개 변수를 바꾸어 보면서 결과가 어떻게 달라지는지 확인 해 보라.<br>
See how the results change as you change parameters such as `epsilon`, `x_init`, and `x_interval`.
### 순차법을 함수로 구현<br>Implementing Sequantial Method in a Function
위 프로그램으로 어떤 수의 제곱근을 구할 수 있었다. 다른 함수의 근을 구하기에 더 편리한 형태로 바꾸면 더 좋을 것이다.<br>
We could the sqare root of a number. It would be even better modify so that we can easily find roots of other functions.
```
def sequential(f, x_init, x_interval, epsilon, x_final):
# result 의 초기값
# Initial value for sqrt_10
result = 'Not Found'
# 일련의 x_i 값을 미리 준비한다
# Prepare a series of x_i values in advance
x_array = py.arange(x_init, x_final+x_interval*0.5, x_interval)
# 몇번 반복했는지 측정해 보자
# Let's count the number of iterations
counter = 0
# x_i 에 관한 반복문
# x_i loop
for x_i in x_array:
# Evaluate the function
y_i = f(x_i)
counter += 1
# Check if absolute value is smaller than epsilon
if abs(y_i) < epsilon:
result = x_i
# found
break
# 반복 횟수
# Number of iterations
print('counter =', counter)
return result
```
이번에는 5의 제곱근을 구해보자.<br>This time, let's find the square root of five.
```
def find_sqrt_5(x):
return (x ** 2) - 5
```
매개변수는 다음과 같다.<br>
Parameters are as follows.
```
# y_i 의 절대값이 이 값 보다 작으면 y_i = 0으로 본다
# If absolute value of y_i is smaller than this, we would think y_i = 0.
epsilon = 1e-3
# 순차법의 매개변수
# Parameters of the Sequantial Method
# x_i 의 초기값
# Initial value of x_i
x_init = 0
# x_i 의 마지막 값
# Final value of x_i
x_final = 4
# i 번째 x 값과 i+1 번째 x 값 사이의 간격
# The interval between i'th and (i+1)'th x's
x_interval = 1.0 / (2**12)
```
제곱근을 구하기 위해 순차법 함수는 다음과 같이 호출할 수 있다.<br>To find the square root of five, we can call the function of the sequential method as follows.
```
sqrt_5 = sequential(find_sqrt_5, x_init, x_interval, epsilon, x_final)
```
순차법으로 찾은 5의 제곱근<br>
Square root of 5 that we just found using the sequential method
```
print('sqrt_5 =', sqrt_5)
```
아래 연산의 결과는 0인가?<br>
Is the result of the following calculation zero?
```
print('sqrt_5 ** 2 - 5 =', find_sqrt_5(sqrt_5))
```
$\epsilon$ 값과 비교<br>Check with the $\epsilon$
```
assert abs(find_sqrt_5(sqrt_5)) < epsilon
```
도전 과제 1: 다음 매개 변수값을 하나씩 바꾸어 보고 그 영향에 대한 의견을 적으시오.<br>Try this 1: Change one parameter value at a time and write down your opinion on its influence.
|매개변수<br>Parameter | 현재값<br>Current value | 바꾸어 본 값<br>New value | 영향<br>Influence |
|:------:|:------:|:----------:|:----------:|
|`epsilon` | `1e-3` | ? | ? |
|`x_init` | `0` | ? | ? |
|`x_interval` | `1.0 / (2**12)` | ? | ? |
도전 과제 2: $sin^2(\theta)=0.5$ 인 $\theta$(도)를 구해 보시오.<br>Try this 2: Find $\theta$(degree) satisfying $sin^2(\theta)=0.5$.
## Final Bell<br>마지막 종
메뉴의 Cell/Run All 명령을 실행시켰을 때 문서의 모든 셀이 실행되었으면 종소리를 냄<br>When all cells executed by Cell/Run All command of the menu, generate the bell sound.
```
# stackoverfow.com/a/24634221
import os
os.system("printf '\a'")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score, silhouette_samples
from yellowbrick.cluster import SilhouetteVisualizer
%matplotlib inline
df = pd.read_csv('customers.csv')
df.shape
df.head()
# considering only annual income and spending score for clustering
X = df.iloc[:,[3,4]].values # numpy array
X[:10]
# plotting the data points before running clustering algorithm
plt.scatter(X[:,0],X[:,1])
```
#### Elbow method for finding number of clusters (K value)
```
SSE_cluster = []
for i in range(2,11): # choosing 10 clusters
kmeans = KMeans(n_clusters=i,init='k-means++',random_state=0)
kmeans.fit(X) # Compute k-means clustering
SSE_cluster.append(kmeans.inertia_)
# Documentation
# n_clusters : The number of clusters to form as well as the number of centroids to generate.
# init: can be {‘k-means++’, ‘random’}.
# ‘k-means++’ : selects initial cluster centers for k-mean clustering in a smart way to speed up convergence.
#It initializes the centroids very very far.
# ‘random’: choose n_clusters observations (rows) at random from data for the initial centroids.
# random_stateint: similar to set_seed(). Determines random number generation for centroid initialization.
# inertia_ : Sum of squared distances of samples to their closest cluster center,weighted by the sample weights if provided.
plt.plot(range(2,11),SSE_cluster) # plotting numbers of clusers vs SSE
plt.title('The Elbow Method')
plt.xlabel('Number of clusters')
plt.ylabel('SSE')
plt.show()
```
#### Number of clusters will be 5
```
# Fitting K means to the dataset
kmeans = KMeans(n_clusters=5,init='k-means++',random_state=0)
y_kmeans = kmeans.fit_predict(X) # Compute cluster centers and predict cluster index for each sample
y_kmeans # contains index of the cluster each sample belongs to
kmeans.cluster_centers_
kmeans.labels_
plt.scatter(X[y_kmeans==0,0], X[y_kmeans==0,1], c='red', label='Cluser1')
plt.scatter(X[y_kmeans==1,0], X[y_kmeans==1,1], c='blue', label='Cluser2')
plt.scatter(X[y_kmeans==2,0], X[y_kmeans==2,1], c='green', label='Cluser3')
plt.scatter(X[y_kmeans==3,0], X[y_kmeans==3,1], c='cyan', label='Cluser4')
plt.scatter(X[y_kmeans==4,0], X[y_kmeans==4,1], c='magenta', label='Cluser5')
# displaying the centroids
plt.scatter(kmeans.cluster_centers_[:,0],kmeans.cluster_centers_[:,1],s = 50, c='yellow', label='Centroids') # s: pixel value of the points
plt.legend()
plt.title('Clusters of customers')
plt.xlabel('Annual Income (k$)')
plt.xlabel('Spending Score (1-100)')
```
#### VALIDATING THE CLUSTER MODEL WITH SILHOUETTE SCORE
```
score = silhouette_score(X, y_kmeans)
score
#Using Silhouette Plot
visualizer = SilhouetteVisualizer(kmeans,colors = 'yellowbrick')
#Fit the data to the visualizer
visualizer.fit(X)
#Render the figure
visualizer.show()
range_n_clusters = [2,3,4,5,6]
for n_clusters in range_n_clusters:
# Create a subplot with 1 row and 2 columns
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.set_size_inches(18, 7)
# The 1st subplot is the silhouette plot
# The silhouette coefficient can range from -1, 1 but in this example all
# lie within [-0.1, 1]
ax1.set_xlim([-0.1, 1])
# The (n_clusters+1)*10 is for inserting blank space between silhouette
# plots of individual clusters, to demarcate them clearly.
ax1.set_ylim([0, len(X) + (n_clusters + 1) * 10])
# Initialize the clusterer with n_clusters value and a random generator
# seed of 10 for reproducibility.
clusterer = KMeans(n_clusters=n_clusters, random_state=10)
cluster_labels = clusterer.fit_predict(X)
# The silhouette_score gives the average value for all the samples.
# This gives a perspective into the density and separation of the formed
# clusters
silhouette_avg = silhouette_score(X, cluster_labels)
print("For n_clusters =", n_clusters,
"The average silhouette_score is :", silhouette_avg)
# Compute the silhouette scores for each sample
sample_silhouette_values = silhouette_samples(X, cluster_labels)
y_lower = 10
for i in range(n_clusters):
# Aggregate the silhouette scores for samples belonging to
# cluster i, and sort them
ith_cluster_silhouette_values = \
sample_silhouette_values[cluster_labels == i]
ith_cluster_silhouette_values.sort()
size_cluster_i = ith_cluster_silhouette_values.shape[0]
y_upper = y_lower + size_cluster_i
color = cm.nipy_spectral(float(i) / n_clusters)
ax1.fill_betweenx(np.arange(y_lower, y_upper),
0, ith_cluster_silhouette_values,
facecolor=color, edgecolor=color, alpha=0.7)
# Label the silhouette plots with their cluster numbers at the middle
ax1.text(-0.05, y_lower + 0.5 * size_cluster_i, str(i))
# Compute the new y_lower for next plot
y_lower = y_upper + 10 # 10 for the 0 samples
ax1.set_title("The silhouette plot for the various clusters.")
ax1.set_xlabel("The silhouette coefficient values")
ax1.set_ylabel("Cluster label")
# The vertical line for average silhouette score of all the values
ax1.axvline(x=silhouette_avg, color="red", linestyle="--")
ax1.set_yticks([]) # Clear the yaxis labels / ticks
ax1.set_xticks([-0.1, 0, 0.2, 0.4, 0.6, 0.8, 1])
# 2nd Plot showing the actual clusters formed
colors = cm.nipy_spectral(cluster_labels.astype(float) / n_clusters)
ax2.scatter(X[:, 0], X[:, 1], marker='.', s=30, lw=0, alpha=0.7,
c=colors, edgecolor='k')
# Labeling the clusters
centers = clusterer.cluster_centers_
# Draw white circles at cluster centers
ax2.scatter(centers[:, 0], centers[:, 1], marker='o',
c="white", alpha=1, s=200, edgecolor='k')
for i, c in enumerate(centers):
ax2.scatter(c[0], c[1], marker='$%d$' % i, alpha=1,
s=50, edgecolor='k')
ax2.set_title("The visualization of the clustered data.")
ax2.set_xlabel("Feature space for the 1st feature")
ax2.set_ylabel("Feature space for the 2nd feature")
plt.suptitle(("Silhouette analysis for KMeans clustering on sample data "
"with n_clusters = %d" % n_clusters),
fontsize=14, fontweight='bold')
plt.show()
```
# Silhouette clusterinng
https://www.youtube.com/watch?v=DpRPd274-0E&list=RDCMUCNU_lfiiWBdtULKOw6X0Dig&start_radio=1&rv=DpRPd274-0E&t=4
https://scikit-learn.org/stable/auto_examples/cluster/plot_kmeans_silhouette_analysis.html#sphx-glr-auto-examples-cluster-plot-kmeans-silhouette-analysis-py
https://github.com/krishnaik06/Silhouette-clustering-/blob/master/Untitled.ipynb
| github_jupyter |
# Policy Evaluation in Contextual Bandits
** *
This IPython notebook illustrates the usage of the [contextualbandits](https://www.github.com/david-cortes/contextualbandits) package's `evaluation` module through a simulation with public datasets.
** Small note: if the TOC here is not clickable or the math symbols don't show properly, try visualizing this same notebook from nbviewer following [this link](http://nbviewer.jupyter.org/github/david-cortes/contextualbandits/blob/master/example/policy_evaluation.ipynb). **
** *
### Sections
[1. Problem description](#p1)
[2. Methods](#p2)
[3. Experiments](#p3)
[4. References](#p4)
** *
<a id="p1"></a>
## 1. Problem description
For a general description of the contextual bandits problem, see the first part of the package's guide [Online Contextual Bandits](http://nbviewer.jupyter.org/github/david-cortes/contextualbandits/blob/master/example/online_contextual_bandits.ipynb).
The previous two guides [Online Contextual Bandits](http://nbviewer.jupyter.org/github/david-cortes/contextualbandits/blob/master/example/online_contextual_bandits.ipynb) and [Off-policy Learning in Contextual Bandits](http://nbviewer.jupyter.org/github/david-cortes/contextualbandits/blob/master/example/offpolicy_learning.ipynb) evaluated the performance of different policies by looking at the actions they would have chosen in a fully-labeled dataset for multi-label classification.
However, in contextual bandits settings one doesn't have access to fully-labeled data, and the data that one has is usually very biased, as it is collected through some policy that aims to maximize rewards. In this situation, it is a lot more difficult to evaluate the performance of a new policy. This module deals with such problem.
** *
<a id="p2"></a>
## 2. Methods
This module implements two policy evaluation methods:
* `evaluateRejectionSampling` (see _"A contextual-bandit approach to personalized news article recommendation"_), for both online and offline policies.
* `evaluateDoublyRobust` (see _"Doubly Robust Policy Evaluation and Learning"_).
Both of these are based on a train-test split - that is, the policy is trained with some data and evaluated on different data.
The best way to obtain a good estimate of the performance of a policy is to collect some data on which actions are chosen at random. When such data is available, one can iterate through it, let the policy choose an action for each observation, and if it matches with what was chosen, take it along with its rewards for evaluation purposes, skip it if not. This simple rejection sampling method is unbiased and let's you evaluate both online and offline algorithms. **It must be stressed that evaluating data like this only works when the actions of this test sample are chosen at random, otherwise the estimates will be biased (and likely very wrong)**.
When such data is not available and there is reasonable variety of actions chosen, another option is doubly-robust estimates. These are meant for the case of continuous rewards, and don't work as well with discrete rewards though, especially when there are many labels, but they can still be tried.
The doubly-robust estimate requires, as it names suggests, two estimates: one of the reward that each arm will give, and another of the probability or score that the policy that collected the data gave to each arm it chose for each observation.
In a scenario such as online advertising, we don't need the second estimate if we record the scores that the models output along with the covariates-action-reward history. When using the functions from this package's `online` module, you can get such estimates for some of the policies by using their `predict_proba_separate` function.
For the first estimate, there are different options to obtain it. One option is to fit a (non-online) model to both the train and test sets to make reward estimates on the test set, or fit it only on the test set (while the policy to be evaluated is fitted to the training set); or perhaps even use the score estimates from the old policy (which chose the actions on the training and test data) or from the new policy. The function `evaluateDoublyRobust` provides an API that can accomodate all these methods.
** *
<a id="p3"></a>
## 3. Experiments
Just like in the previous guide [Off-policy Learning in Contextual Bandits](http://nbviewer.jupyter.org/github/david-cortes/contextualbandits/blob/master/example/offpolicy_learning.ipynb), I will simualate data generated from a policy by fitting a logistic regression model with a sample of the **fully-labeled** data, then let it choose actions for some more data, and take those actions and rewards as input for a new policy, along with the estimated reward probabilities for the actions that were chosen.
The new policy will then be evaluated on a test sample with actions already pre-selected, and the estimates from the methods here will be compared with the real rewards, which we can know because the data is fully labeled.
The data are again the Bibtext and Mediamill datasets.
** *
Loading the Bibtex dataset again:
```
import pandas as pd, numpy as np, re
from sklearn.preprocessing import MultiLabelBinarizer
def parse_data(file_name):
features = list()
labels = list()
with open(file_name, 'rt') as f:
f.readline()
for l in f:
if bool(re.search("^[0-9]", l)):
g = re.search("^(([0-9]{1,2},?)+)\s(.*)$", l)
labels.append([int(i) for i in g.group(1).split(",")])
features.append(eval("{" + re.sub("\s", ",", g.group(3)) + "}"))
else:
l = l.strip()
labels.append([])
features.append(eval("{" + re.sub("\s", ",", l) + "}"))
features = pd.DataFrame.from_dict(features).fillna(0).as_matrix()
mlb = MultiLabelBinarizer()
y = mlb.fit_transform(labels)
return features, y
features, y = parse_data("Bibtex_data.txt")
print(features.shape)
print(y.shape)
```
Simulating a stationary exploration policy and a test set:
```
from sklearn.linear_model import LogisticRegression
# the 'explorer' polcy will be fit with this small sample of the rows
st_seed = 0
end_seed = 2000
# then it will choose actions for this larger sample, which will be the input for the new policy
st_exploration = 0
end_exploration = 3000
# the new policy will be evaluated with a separate test set
st_test = 3000
end_test = 7395
# separating the covariates data for each case
Xseed = features[st_seed:end_seed, :]
Xexplore_sample = features[st_exploration:end_exploration, :]
Xtest = features[st_test:end_test, :]
nchoices = y.shape[1]
# now constructing an exploration policy as explained above, with fully-labeled data
explorer = LogisticRegression()
np.random.seed(100)
explorer.fit(Xseed, np.argmax(y[st_seed:end_seed], axis=1))
# letting the exploration policy choose actions for the new policy input
np.random.seed(100)
actions_explore_sample=explorer.predict(Xexplore_sample)
rewards_explore_sample=y[st_exploration:end_exploration, :]\
[np.arange(end_exploration - st_exploration), actions_explore_sample]
# extracting the probabilities it estimated
ix_internal_actions = {j:i for i,j in enumerate(explorer.classes_)}
ix_internal_actions = [ix_internal_actions[i] for i in actions_explore_sample]
ix_internal_actions = np.array(ix_internal_actions)
prob_actions_explore = explorer.predict_proba(Xexplore_sample)[np.arange(Xexplore_sample.shape[0]),
ix_internal_actions]
# generating a test set with random actions
actions_test = np.random.randint(nchoices, size=end_test - st_test)
rewards_test = y[st_test:end_test, :][np.arange(end_test - st_test), actions_test]
```
Rejection sampling estimate:
```
from contextualbandits.online import SeparateClassifiers
from contextualbandits.evaluation import evaluateRejectionSampling
new_policy = SeparateClassifiers(LogisticRegression(C=0.1), y.shape[1])
np.random.seed(100)
new_policy.fit(Xexplore_sample, actions_explore_sample, rewards_explore_sample)
np.random.seed(100)
est_r, ncases = evaluateRejectionSampling(new_policy, X=Xtest, a=actions_test, r=rewards_test, online=False)
np.random.seed(100)
real_r = np.mean(y[st_test:end_test,:][np.arange(end_test - st_test), new_policy.predict(Xtest)])
print('Test set Rejection Sampling mean reward estimate (new policy)')
print('Estimated mean reward: ',est_r)
print('Sample size: ', ncases)
print('----------------')
print('Real mean reward: ', real_r)
```
We can also evaluate the exploration policy with the same method:
```
np.random.seed(100)
est_r, ncases = evaluateRejectionSampling(explorer, X=Xtest, a=actions_test, r=rewards_test, online=False)
real_r = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), explorer.predict(Xtest)])
print('Test set Rejection Sampling mean reward estimate (old policy)')
print('Estimated mean reward: ', est_r)
print('Sample size: ', ncases)
print('----------------')
print('Real mean reward: ', real_r)
```
_(Remember that the exploration policy was fit with a smaller set of fully-labeled data, thus it's no surprise it performs a lot better)_
The estimates are not exact, but they are somewhat close to the real values as expected. They get better the more cases are successfully sampled, and their estimate should follow the central limit theorem.
** *
To be stressed again, such an evaluation method only works when the data was collected by choosing actions at random. **If we evaluate it with the actions chosen by the exploration policy, the results will be totally biased as demonstrated here:**
```
actions_test_biased = explorer.predict(Xtest)
rewards_test_biased = y[st_test:end_test, :][np.arange(end_test - st_test), actions_test_biased]
est_r, ncases = evaluateRejectionSampling(new_policy, X=Xtest, a=actions_test_biased,\
r=rewards_test_biased, online=False)
real_r = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), new_policy.predict(Xtest)])
print('Biased Test set Rejection Sampling mean reward estimate (new policy)')
print('Estimated mean reward: ', est_r)
print('Sample size: ', ncases)
print('----------------')
print('Real mean reward: ', real_r)
print("(Don't try rejection sampling on a biased test set)")
```
We can also try Doubly-Robust estimates, but these work poorly for a dataset like this:
```
from contextualbandits.evaluation import evaluateDoublyRobust
# getting estimated probabilities for the biased test sample chosen by the old policy
ix_internal_actions = {j:i for i,j in enumerate(explorer.classes_)}
ix_internal_actions = [ix_internal_actions[i] for i in actions_test_biased]
ix_internal_actions = np.array(ix_internal_actions)
prob_actions_test_biased = explorer.predict_proba(Xtest)[np.arange(Xtest.shape[0]), ix_internal_actions]
# actions that the new policy will choose
np.random.seed(1)
pred = new_policy.predict(Xtest)
# method 1: estimating rewards by fitting another model to the whole data (train + test)
model_fit_on_all_data = SeparateClassifiers(LogisticRegression(), y.shape[1])
np.random.seed(1)
model_fit_on_all_data.fit(np.r_[Xexplore_sample, Xtest],
np.r_[actions_explore_sample, actions_test_biased],
np.r_[rewards_explore_sample, rewards_test_biased])
np.random.seed(1)
est_r_dr_whole = evaluateDoublyRobust(pred, X=Xtest, a=actions_test_biased, r=rewards_test_biased,\
p=prob_actions_test_biased, reward_estimator = model_fit_on_all_data)
# method 2: estimating rewards by fitting another model to the test data only
np.random.seed(1)
est_r_dr_test_only = evaluateDoublyRobust(pred, X=Xtest, a=actions_test_biased, r=rewards_test_biased,\
p=prob_actions_test_biased, reward_estimator = LogisticRegression(), nchoices=y.shape[1])
print('Biased Test set mean reward estimates (new policy)')
print('DR estimate (reward estimator fit on train+test): ', est_r_dr_whole)
print('DR estimate (reward estimator fit on test only): ', est_r_dr_test_only)
print('----------------')
print('Real mean reward: ', real_r)
```
Both estimates are very wrong, but they are still less wrong than the wrongly-conducted rejection sampling from before.
** *
Finally, rejection sampling can also be used to evaluate online policies - in this case though, be aware that the estimate will only be considered up to a certain number of rounds (as many as it accepts, but it will end up rejecting the majority), but online policies keep improving with time.
Here I will use the Mediamill dataset instead, as it has a lot more data:
```
from contextualbandits.online import BootstrappedUCB
features, y = parse_data("Mediamill_data.txt")
nchoices = y.shape[1]
Xall=features
actions_random = np.random.randint(nchoices, size = Xall.shape[0])
rewards_actions = y[np.arange(y.shape[0]), actions_random]
online_policy = BootstrappedUCB(LogisticRegression(), y.shape[1])
evaluateRejectionSampling(online_policy,
X = Xall,
a = actions_random,
r = rewards_actions,
online = True,
start_point_online = 'random',
batch_size = 5)
```
** *
<a id="p4"></a>
## 4. References
* Li, L., Chu, W., Langford, J., & Schapire, R. E. (2010, April). A contextual-bandit approach to personalized news article recommendation. In Proceedings of the 19th international conference on World wide web (pp. 661-670). ACM.
* Dudík, M., Langford, J., & Li, L. (2011). Doubly robust policy evaluation and learning. arXiv preprint arXiv:1103.4601.
| github_jupyter |
```
!pip install scikit-learn==1.0
!pip install xgboost==1.4.2
!pip install catboost==0.26.1
!pip install pandas==1.3.3
!pip install radiant-mlhub==0.3.0
!pip install rasterio==1.2.8
!pip install numpy==1.21.2
!pip install pathlib==1.0.1
!pip install tqdm==4.62.3
!pip install joblib==1.0.1
!pip install matplotlib==3.4.3
!pip install Pillow==8.3.2
!pip install torch==1.9.1
!pip install plotly==5.3.1
gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
if gpu_info.find('failed') >= 0:
print('Select the Runtime > "Change runtime type" menu to enable a GPU accelerator, ')
print('and then re-execute this cell.')
else:
print(gpu_info)
import pandas as pd
import numpy as np
import random
import torch
def seed_all(seed_value):
random.seed(seed_value) # Python
np.random.seed(seed_value) # cpu vars
torch.manual_seed(seed_value) # cpu vars
if torch.cuda.is_available():
torch.cuda.manual_seed(seed_value)
torch.cuda.manual_seed_all(seed_value) # gpu vars
torch.backends.cudnn.deterministic = True #needed
torch.backends.cudnn.benchmark = False
seed_all(13)
# from google.colab import drive
# drive.mount('/content/drive')
import warnings
warnings.filterwarnings("ignore")
import gc
import pandas as pd
import numpy as np
from sklearn.metrics import *
from xgboost import XGBClassifier
from catboost import CatBoostClassifier
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import roc_auc_score
from sklearn.ensemble import VotingClassifier
from sklearn.linear_model import LogisticRegression
from indices_creation import *
```
## Data Load Step
1. We load the mean aggregations for both train and test. The mean aggregations contain the labels and field IDs.
2. The quantile aggregations contain the field IDs.
```
import os
os.getcwd()
train_df_mean = pd.read_csv('train_mean.csv')
#### we need to drop 'label' and 'field_id' later in the code
test_df_mean = pd.read_csv('test_mean.csv')
#### we need to drop 'field_id' later in the code
train_df_median = pd.read_csv('train_median.csv')
#### we need to drop 'field_id' later in the code
test_df_median = pd.read_csv('test_median.csv')
#### we need to drop 'field_id' later in the code
train_size = pd.read_csv('size_of_field_train.csv')
test_size = pd.read_csv('size_of_field_test.csv')
train_size = train_size.rename({'Field_id':'field_id'},axis=1)
test_size = test_size.rename({'Field_id':'field_id'},axis=1)
train_df_median = train_df_median.merge(train_size, on =['field_id'],how='left')
test_df_median = test_df_median.merge(test_size, on =['field_id'],how='left')
cluster_df = pd.read_csv('seven_cluster.csv')
cluster_df = cluster_df.rename({'cluster_label':'cluster_label_7'},axis=1)
train_df_median = train_df_median.merge(cluster_df,on=['field_id'],how='left')
test_df_median = test_df_median.merge(cluster_df,on=['field_id'],how='left')
gc.collect()
full_nearest=pd.read_csv('full_nearest_radius_0.25.csv')
full_nearest
train_df_median = train_df_median.merge(full_nearest,on=['field_id'],how='left')
print(train_df_median.shape)
test_df_median = test_df_median.merge(full_nearest,on=['field_id'],how='left')
```
## Removing Erroneous data points
We observed some data points for which the labels were floats, we will remove them (they are few in number) to make sure our model is learning from correctly labelled data points
```
print(f'The shape of train data before outlier removal - {train_df_mean.shape}')
train_df_mean = train_df_mean[train_df_mean.label.isin(list(range(1,10)))]
print(f'The shape of train data after outlier removal - {train_df_mean.shape}')
relevant_fids = train_df_mean['field_id'].values.tolist()
train_df_median = train_df_median[train_df_median['field_id'].isin(relevant_fids)]
print(f'The shape of median train data - {train_df_median.shape} and mean train data {train_df_mean.shape}' )
### two extra columns in train_df_mean being 'label' and 'size_of_field'
```
### Extract date list
We extract the list of all dates where observations were seen for index generation
```
cols = ['B01_','B02_','B03_','B04_','B05_','B06_','B07_','B08_','B09_','B8A_','B11_','B12_']
columns_available = train_df_mean.columns.tolist()
cols2consider = []
for col in cols:
cols2consider.extend( [c for c in columns_available if col in c])
bands_with_dates = [c for c in columns_available if 'B01_' in c]
dates = [c.replace('B01_','') for c in bands_with_dates]
print(f'The sample showing the commencement dates where observations were seen is {dates[:10]}')
print(f'The sample showing the ending dates where observations were seen is {dates[-10:]}')
```
### Removal of field ID column
We consider only the relevant columns to be considered for the next step
```
train_df_mean = train_df_mean[cols2consider+['label']]
test_df_mean = test_df_mean[cols2consider]
train_df_median = train_df_median[cols2consider+['size_of_field']+['cluster_label_7']+full_nearest.columns.tolist()]
test_df_median = test_df_median[cols2consider+['size_of_field']+['cluster_label_7']+full_nearest.columns.tolist()]
```
### Indices Creation
We will create the indices for train and test data for mean aggregates using the indices coded in indices_creation.py module
```
# train_df_mean = get_band_ndvi_red(train_df_mean,dates)
# train_df_mean = get_band_afri(train_df_mean,dates)
# train_df_mean = get_band_evi2(train_df_mean,dates)
# train_df_mean = get_band_ndmi(train_df_mean,dates)
# train_df_mean = get_band_ndvi(train_df_mean,dates)
# train_df_mean = get_band_evi(train_df_mean,dates)
# train_df_mean = get_band_bndvi(train_df_mean,dates)
# train_df_mean = get_band_nli(train_df_mean,dates)
# train_df_mean = get_band_lci(train_df_mean,dates)
# test_df_mean = get_band_ndvi_red(test_df_mean,dates)
# test_df_mean = get_band_afri(test_df_mean,dates)
# test_df_mean = get_band_evi2(test_df_mean,dates)
# test_df_mean = get_band_ndmi(test_df_mean,dates)
# test_df_mean = get_band_ndvi(test_df_mean,dates)
# test_df_mean = get_band_evi(test_df_mean,dates)
# test_df_mean = get_band_bndvi(test_df_mean,dates)
# test_df_mean = get_band_nli(test_df_mean,dates)
# test_df_mean = get_band_lci(test_df_mean,dates)
```
We will create the indices for train and test data for median aggregates using the indices coded in indices_creation.py module
```
train_df_median = get_band_ndvi_red(train_df_median,dates)
train_df_median = get_band_afri(train_df_median,dates)
train_df_median = get_band_evi2(train_df_median,dates)
train_df_median = get_band_ndmi(train_df_median,dates)
train_df_median = get_band_ndvi(train_df_median,dates)
train_df_median = get_band_evi(train_df_median,dates)
train_df_median = get_band_bndvi(train_df_median,dates)
train_df_median = get_band_nli(train_df_median,dates)
# train_df_median = get_band_lci(train_df_median,dates)
test_df_median = get_band_ndvi_red(test_df_median,dates)
test_df_median = get_band_afri(test_df_median,dates)
test_df_median = get_band_evi2(test_df_median,dates)
test_df_median = get_band_ndmi(test_df_median,dates)
test_df_median = get_band_ndvi(test_df_median,dates)
test_df_median = get_band_evi(test_df_median,dates)
test_df_median = get_band_bndvi(test_df_median,dates)
test_df_median = get_band_nli(test_df_median,dates)
# test_df_median = get_band_lci(test_df_median,dates)
# train_df_median = train_df_median.drop(cols2consider,axis=1)
# test_df_median = test_df_median.drop(cols2consider,axis=1)
train_df_mean.shape,train_df_median.shape,test_df_mean.shape,test_df_median.shape
######### Saving the label variable and dropping it from the data
train_y = train_df_mean['label'].values
train_df_mean = train_df_mean.drop(['label'],axis=1)
train_df_mean.replace([np.inf, -np.inf], np.nan, inplace=True)
test_df_mean.replace([np.inf, -np.inf], np.nan, inplace=True)
train_df_median.replace([np.inf, -np.inf], np.nan, inplace=True)
test_df_median.replace([np.inf, -np.inf], np.nan, inplace=True)
# train_df_slope.replace([np.inf, -np.inf], np.nan, inplace=True)
# test_df_slope.replace([np.inf, -np.inf], np.nan, inplace=True)
train = train_df_median.values
test = test_df_median.values
# train = pd.concat([train_df_median,train_df_slope],axis=1).values
# test = pd.concat([test_df_median,test_df_slope],axis=1).values
print(f'The shape of model ready train data is {train.shape} and model ready test data is {test.shape}')
print(f'The shape of target is {train_y.shape}')
train1 = pd.read_csv('train_with_slopes.csv')
test1 = pd.read_csv('test_with_slopes.csv')
train1.replace([np.inf, -np.inf], np.nan, inplace=True)
test1.replace([np.inf, -np.inf], np.nan, inplace=True)
train2=pd.concat([pd.DataFrame(train1.values,columns=train1.columns),train_df_median[['size_of_field','cluster_label_7']+full_nearest.columns.tolist()].reset_index(drop=True)],axis=1)
test2=pd.concat([pd.DataFrame(test1.values,columns=test1.columns),test_df_median[['size_of_field','cluster_label_7']+full_nearest.columns.tolist()].reset_index(drop=True)],axis=1)
train2.head()
del train2['field_id']
del test2['field_id']
pivot=pd.read_csv('pivottable.csv')
pivot
train2=train2.merge(pivot,how='left',on='cluster_label_7')
test2=test2.merge(pivot,how='left',on='cluster_label_7')
train2
del train_df_mean,train_df_median,train1,train_size,test_df_mean,test_df_median,test1,test_size
import gc
gc.collect()
train = train2.values
test = test2.values
train.shape,test.shape
# (1616-1520)/8
oof_pred = np.zeros((len(train), 9))
y_pred_final = np.zeros((len(test),9 ))
num_models = 3
temperature = 50
n_splits = 15
error = []
kf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=13)
for fold, (tr_ind, val_ind) in enumerate(kf.split(train, train_y)):
wghts = [0]*num_models
logloss = []
X_train, X_val = train[tr_ind], train[val_ind]
# X_train1, X_val1 = train_max[tr_ind], train_max[val_ind]
y_train, y_val = train_y[tr_ind], train_y[val_ind]
model1 = XGBClassifier(n_estimators=2000,random_state=13,learning_rate=0.04,colsample_bytree=0.95,reg_lambda=11,
tree_method='gpu_hist',eval_metric='mlogloss')
model2 = CatBoostClassifier(task_type='GPU',verbose=False,n_estimators=5000,random_state=13,auto_class_weights='SqrtBalanced',max_depth=9,learning_rate=0.06)
model3 = CatBoostClassifier(task_type='GPU',verbose=False,n_estimators=5000,random_state=13,auto_class_weights='SqrtBalanced',max_depth=10,learning_rate=0.04)
# model4 = CatBoostClassifier(task_type='GPU',verbose=False,n_estimators=5000,random_state=13,auto_class_weights='SqrtBalanced',max_depth=11)
model1.fit(X_train,y_train)
val_pred1 = model1.predict_proba(X_val)
logloss.append(log_loss(y_val,val_pred1))
print('validation logloss model1 fold-',fold+1,': ',log_loss(y_val,val_pred1))
model2.fit(X_train,y_train)
val_pred2 = model2.predict_proba(X_val)
logloss.append(log_loss(y_val,val_pred2))
print('validation logloss model2 fold-',fold+1,': ',log_loss(y_val,val_pred2))
model3.fit(X_train,y_train)
val_pred3 = model3.predict_proba(X_val)
logloss.append(log_loss(y_val,val_pred3))
print('validation logloss model3 fold-',fold+1,': ',log_loss(y_val,val_pred3))
# model4.fit(X_train,y_train)
# val_pred4 = model4.predict_proba(X_val)
# logloss.append(log_loss(y_val,val_pred4))
# print('validation logloss model4 fold-',fold+1,': ',log_loss(y_val,val_pred4))
wghts = np.exp(-temperature*np.array(logloss/sum(logloss)))
wghts = wghts/sum(wghts)
print(wghts)
val_pred = wghts[0]*val_pred1+wghts[1]*val_pred2+wghts[2]*val_pred3 #+wghts[3]*val_pred4
print('Validation logloss for fold- ',fold+1,': ',log_loss(y_val,val_pred))
oof_pred[val_ind] = val_pred
y_pred_final += (wghts[0]*model1.predict_proba(test)+
wghts[1]*model2.predict_proba(test)+wghts[2]*model3.predict_proba(test)
)/(n_splits)
print('OOF LogLoss :- ',(log_loss(train_y,oof_pred)))
outputs = y_pred_final.copy()
test_df = pd.read_csv('test_mean.csv')
field_ids_test = test_df['field_id'].values.tolist()
data_test = pd.DataFrame(outputs)
data_test['field_id'] = field_ids_test
data_test = data_test[data_test.field_id != 0]
data_test
data_test = data_test.rename(columns={
0:'Lucerne/Medics',
1:'Planted pastures (perennial)',
2:'Fallow',
3:'Wine grapes',
4:'Weeds',
5:'Small grain grazing',
6:'Wheat',
7:'Canola',
8:'Rooibos'
})
pred_df = data_test[['field_id', 'Lucerne/Medics', 'Planted pastures (perennial)', 'Fallow', 'Wine grapes', 'Weeds', 'Small grain grazing', 'Wheat', 'Canola', 'Rooibos']]
pred_df['field_id'] = pred_df['field_id'].astype(int)
pred_df = pred_df.sort_values(by=['field_id'],ascending=True)
pred_df
pred_df.to_csv('trial1_sep_salim.csv',index=False)
```
| github_jupyter |
## 1 卷积神经网络
在之前的神经网络学习过程中,使用的都是全连接神经网络,全连接神经网络对识别和预测都有非常好的效果。在之前使用 MNIST 数据集的实践过程中,输入神经网络的是是一幅 28 行 28 列的 784 个像素点的灰度值,但是仅两层神经网络就有十多万个待训练参数(第一层$784\times128$个$\omega+128个b$,第二层$128\times10$个$z\omega+10个b$,共 101770 个参数)。
在实际项目中,输入神经网络的是具有更高分辨率的彩色图片,使得送入全连接网络的输入特征数特别多,随着隐藏层数的增加,网络规模过大,待优化参数过多,很容易造成过拟合。**为了减少待训练参数,在实际应用时会先对原始图片进行特征提取,再把提取到的特征送给全连接网络**。而卷积计算就是一种有效的提取图像特征的方法。
### 1.1 卷积层
#### 1.1.1 卷积过程
卷积(Convolutional)的计算过程:
(1)一般会用一个正方形的卷积核,按指定步长,在输入特征图上滑动,遍历输入特征图中的每个像素点。滑动过程如下所示:

(2)每移动一个步长,卷积核会与输入特征图出现重合区域,重合区域对应元素相乘、求和再加上偏置项得到输出特征的每一个像素点。如果输入特征是单通道灰度图,那么使用的就是深度为 1 的单通道卷积核。那么计算过程如下:
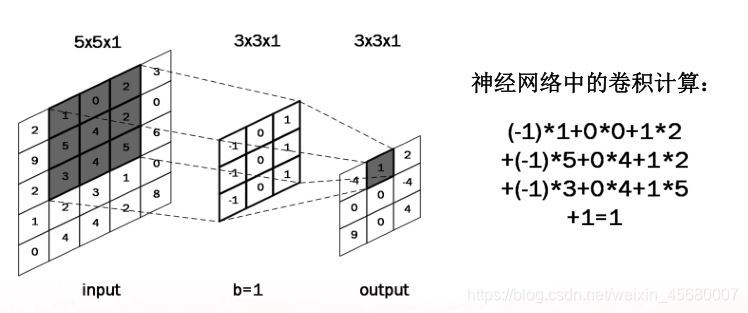
**单通道与卷积核深度为 1 有什么关系呢**?首先要知道图片除去长度和宽度,还有一个表示图片色彩的通道(channel)数,灰度图就是单通道,RGB 彩色图就是三通道。要想让卷积核与输入特征图对应点匹配上,就必须让卷积核的深度与输入特征图的通道数保持一致,所以**输入特征图的通道数(深度)决定了当前层卷积核的深度**。那么对于单通道的特征图而言,通道数为 1 自然卷积核的深度也为 1。
综上所述,如果输入特征是三通道彩色图,则需要使用一个 $3\times3\times3$ 的卷积核,结合上面两幅图片,去理解下面的卷积计算过程:
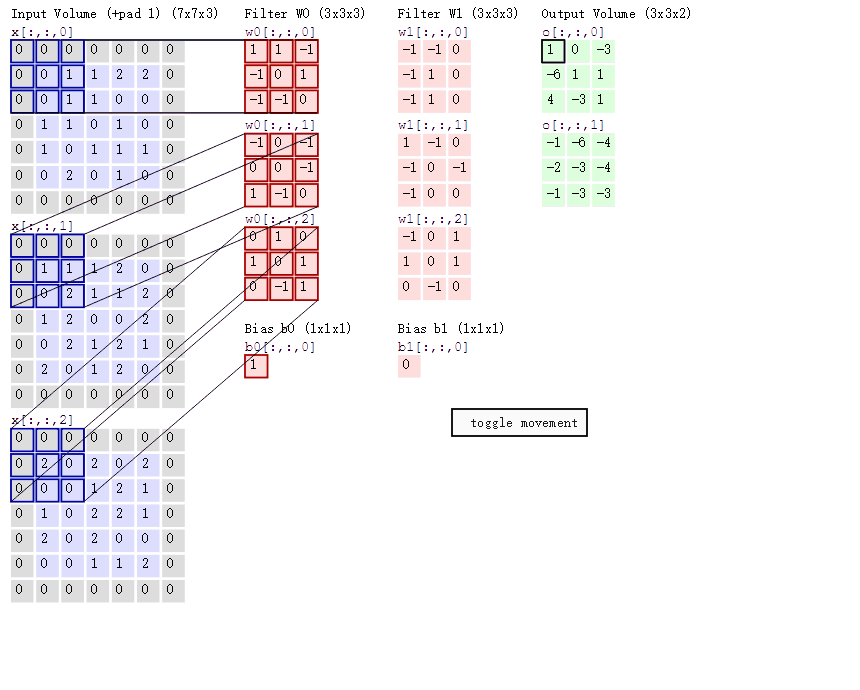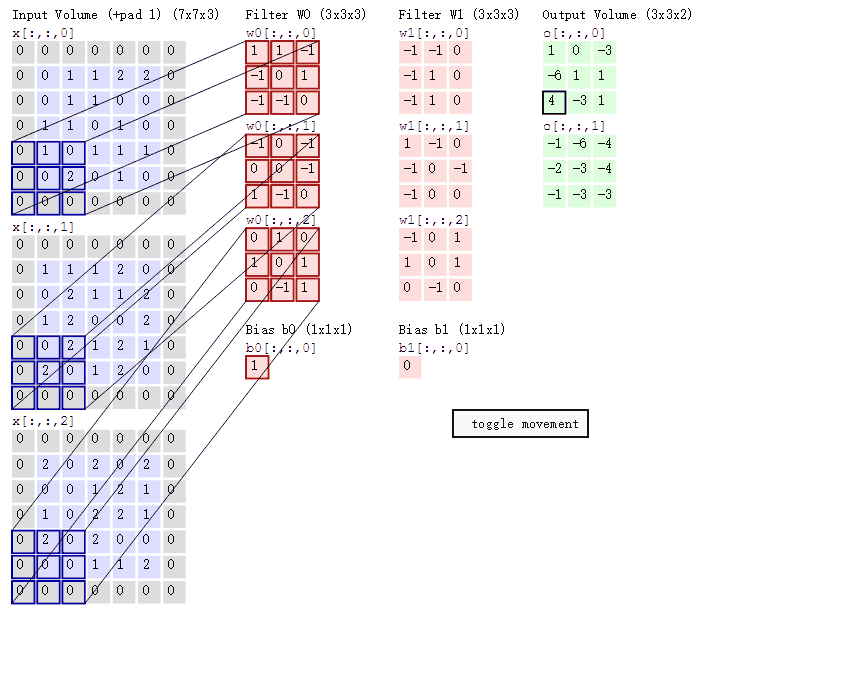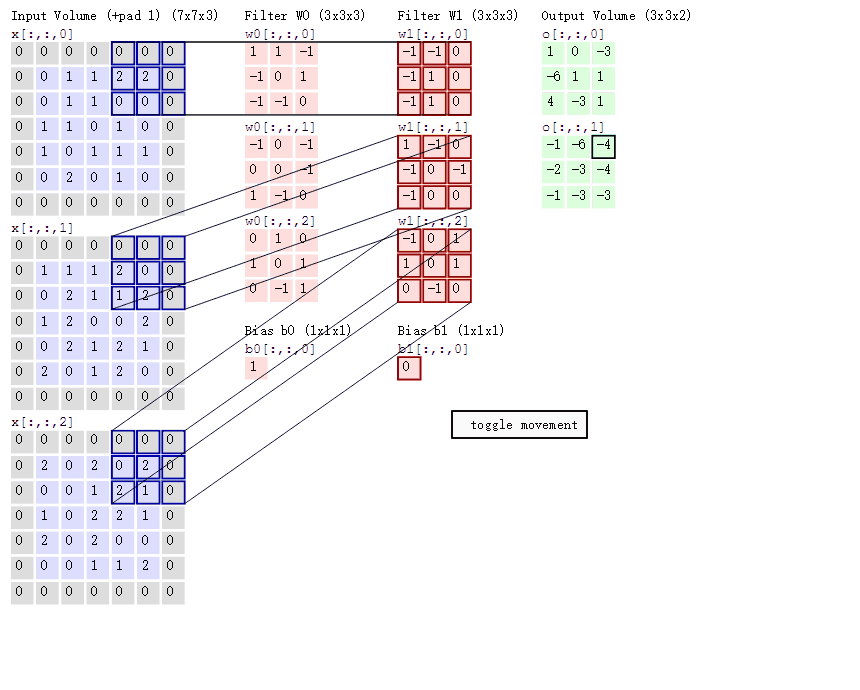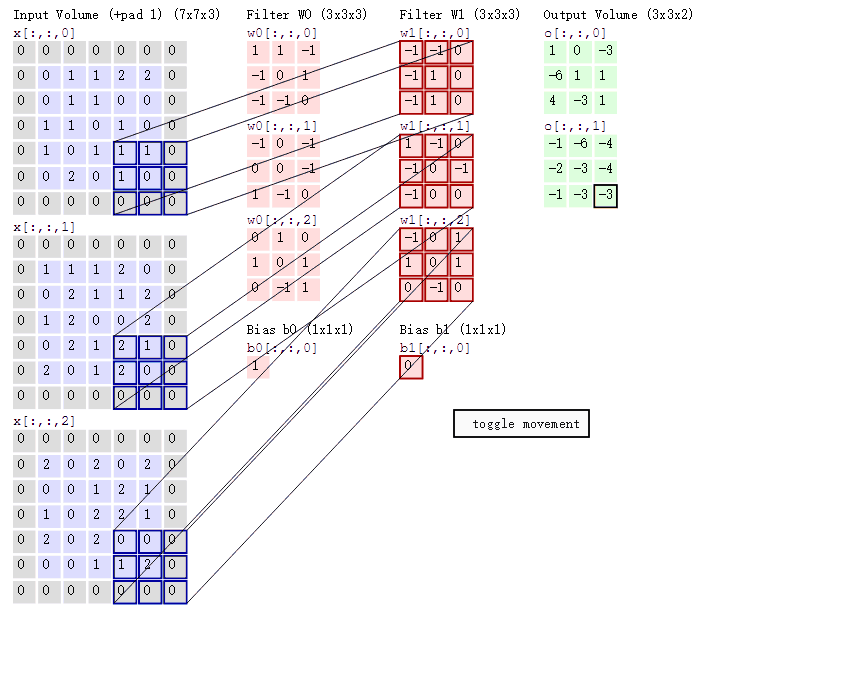
在上面的动图中,使用了两个卷积核 $\text{filter } \omega_0$ 和 $\text{filter } \omega_1$,分别得到了最右边 output 下的两个输出特征图,所以当前层使用了几个卷积核,就有几个输出特征图,即**当前卷积核的个数决定了当前层输出特征图的深度**。如果觉得某层模型的特征提取能力不足,可以在这一层多用几个卷积核提高这一层的特征提取能力。卷积核的深度是固定的,但卷积核的个数是任意的。
#### 1.1.2感受野
感受野(Receptive Field):卷积神经网络各输出特征图中的每个像素点,在原始输入图片上映射区域的大小。简单点说就是,输出特征图上的一个点对应输入图上的区域。有一张图来理解就是:
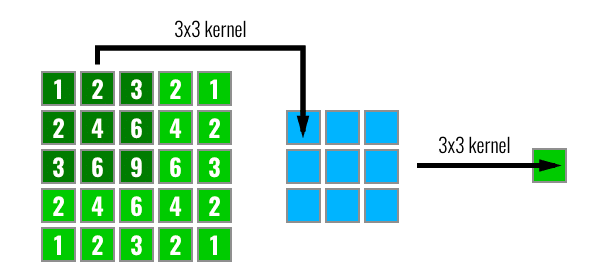
根据上面这个图可以发现,对于一个 $5\times5$ 的输入特征图,经过两个 $3\times3$ 的卷积核的运算,可以得到一个像素点,那么对于这一个像素点来说,映射到输入特征图的感受野就是 5, 正好与输入特征图的大小一致。那么进一步再思考一下,如果对一个 $5\times5$ 的输入特征图,使用一个 $5\times5$ 的卷积核进行运算,得到也会是一个像素点,而对于这一个像素点来说它映射到输入特征图的感受野也是 5。
那么问题来了,既然对于一个相同的输入特征图,应用两个 $3\times3$ 的卷积核和应用一个 $5\times5$ 卷积核的特征提取能力是一样的,那么应该使用那种方案比较好?
这个时候就要考虑,两个方案所承载的待训练参数量和计算量了:
+ 对于两个 $3\times3$ 卷积核,参数量为 $3\times3 + 3\times3 = 18$ 个
+ 对于一个 $5\times5$ 卷积核,参数量为 $5 \times 5 = 25$ 个
在计算量上,假设输入特征图宽、高为 $x$,卷积核宽、高为 $k$,卷积计算步长为 1,输出图边长计算公式为 `输出图边长=(输入图边长-卷积核长+1)/步长`,那么输出特征图的像素个数就是输出图边长的平方,每个像素点都需要进行 $k^2$ 次乘加运算。因此总的计算量就是像素个数乘以乘加运算的次数:
+ 对于两个 $3\times3$ 卷积核,计算量为 $3\times3\times(x-3+1)^2 + 3\times3\times(x-3+1-3+1)^2=18x^2-108x+180$ 个
+ 对于一个 $5\times5$ 卷积核,计算量为 $5\times5(x-5+1)^2=25x^2-200x+400$ 个
直接给出结论,当 $x>10$ 时,两层 $3\times3$ 卷积核会比一个 $5\times5$ 卷积核的计算量少,也就是说前者会比后者的性能要好。这也是为什么现在的神经网络在卷积计算中常使用两层 $3\times3$ 卷积核替换一层 $5\times5$ 卷积核。
#### 1.1.3 填充
填充(padding):为了保持输出图像尺寸与输入图像一致,经常会在输入图像周围进行全零填充。参考下面的动图:

可以发现,输入特征图经过 $3\times$ 卷积核的运算后,得到的输出特征图的大小与输入特征图的大小一致,其原因就在于,在进行卷积运算之前,在输入特征图周围进行了全 0 填充。此外,是否使用全 0 填充,对输出特征图尺寸的大小也有影响:
$$
L_{o}=\begin{cases}
\lceil \frac{L_{i}}{Step} \rceil & \text{ if } padding=same \\
\lceil \frac{L_{i} - L_{k}+1}{Step} \rceil & \text{ if } padding=valid
\end{cases}
$$
+ $L_{o}$:输出特征图边长
+ $L_{i}$:输入特征图边长
+ $L_{k}$:卷积核边长
+ $Step$:步长
+ $padding=same$:全 0 填充
+ $padding=valid$:不使用全 0 填充
如果计算结果为小数,需要向上取整,比如计算结果若为 2.2,则结果应该变为 3。后面的条件 `padding=same` 和 `padding=valid` 就是与 TensorFlow 的 API 相对应,在使用 `tf.keras.layers.Conv2D()` 来设置卷积层时,对于参数 `padding` 的取值就是 `same` 或者 `valid`。
在 TensorFlow 中,使用 `tf.keras.layers.Conv2D` 来定义卷积层
```python
tf.keras.layers.Conv2D (
filters=卷积核个数,
kernel_size=卷积核尺寸, #正方形写核长整数,或(核高h,核宽w)
strides=滑动步长, #横纵向相同写步长整数,或(纵向步长h,横向步长w),默认1
padding='same'或'valid', #使用全零填充是“same”,不使用是“valid”(默认)
activation='relu'或'sigmoid'或'tanh'或'softmax'等 , #如卷积层之后还有批标准化操作,则不在这里使用激活函数
input_shape=(高,宽,通道数) #输入特征图维度,可省略
)
```
### 1.2 批标准化层
神经网络对 0 附近的数据更敏感,但随着网络层数的增加,特征数据会出现偏离 0 均值的情况。标准化可以使数据符合以 0 为均值 1 为标准差的标准正态分布,把偏移的特征数据重新拉回到 0 附近。
批标准化(Batch Normalization,BN)就是对一小批(batch)数据进行标准化处理,使数据回归标准正态分布。常用在卷积操作和激活操作之间。
批标准化后,第 k 个卷积核的输出特征图中第 i 个像素点的值:
$$H_i^{'k}=\frac{H_i^k-\mu_{batch}^k}{\sigma_{batch}^k}$$
+ $H_i^k$:批标准化前,第 k 个卷积核,输出特征图中第 i 个像素点的值
+ $\mu_{batch}^k$:批标准化前,第 k 个卷积核,batch 张输出特征图中所有像素点平均值 $\mu_{batch}^k=\frac{1}{m}\sum_{i=1}^mH_i^k$
+ $\sigma_{batch}^k$:批标准化前,第 k 个卷积核,batch 张输出特征图中所有像素点标准差 $\sigma_{batch}^k=\sqrt{\delta+\frac{1}{m}\sum_{i=1}^m(H_i^k-\mu_{batch}^k)^2}$
BN 操作将原本偏移的特征数据重新拉回到 0 均值,使进入激活函数的数据分布在激活函数线性区,使得输入数据的微小变化,更明显的体现到激活函数的输出,提升了激活函数对输入数据的区分力。但是这种简单的特征数据标准化,使特征数据完全满足标准正态分布,集中在激活函数中心的线性区域。使激活函数丧失了非线性特性。因此在 BN 操作中为每个卷积核引入了两个可训练参数:缩放因子$\gamma$ 和 偏移因子$\beta$,在反向传播时会与其他参数一同被训练优化,使标准正态分布后的特征数据通过缩放因子和偏移因子优化了特征数据分布的宽窄和偏移量,保证了网络的非线性表达力。特征数据分布 $X_i^k$ 表示如下
$$X_i^k=\gamma H_i^{'k}+\beta_k$$
在 TensorFlow 中,使用 `tf.keras.layers.BatchNormalization()` 来表示批标准化层
### 1.3 池化层
池化(Pooling)操作用于减少卷积神经网络中的特征数量。主要法有最大池化和均值池化。最大值池化可以提取图片纹理,均值池化可以保留背景特征。如果用 $2\times2$ 的池化核对输入图片以 2 为步长进行池化,输出图片将变为输入图片的四分之一。

最大值池化就是把与池化核覆盖区域的最大值输出,如下图(a);均值池化就是把与池化核覆盖区域的均值输出,如下图(b)。

在 Tensorflow 中分别使用 的是`tf.keras.layers.MaxPool2D` 函数和 `tf.keras.layers.AveragePooling2D` 函数,具体的使用方法如下:
最大值池化
```python
tf.keras.layers.MaxPool2D(
pool_size=池化核尺寸, #正方形写核长整数,或(核高h,核宽w)
strides=池化步长, #步长整数, 或(纵向步长h,横向步长w),默认为pool_size
padding='same'或'valid')#使用全零填充是“same”,不使用是“valid”(默认)
```
均值池化
```python
tf.keras.layers.AveragePooling2D(
pool_size=池化核尺寸, #正方形写核长整数,或(核高h,核宽w)
strides=池化步长, #步长整数, 或(纵向步长h,横向步长w),默认为pool_size
padding='same'或'valid')#使用全零填充是“same”,不使用是“valid”(默认)
```
### 1.4 Dropout 层
为了缓解神经网络过拟合,在训练过程中,将一部分隐藏层的神经元按照一定比例从神经网络中临时舍弃,在使用时,再把舍弃的神经元恢复链接。如下图所示,图(a)为标准的全连接神经网络,图(b)为使用 dropout 后的网络:
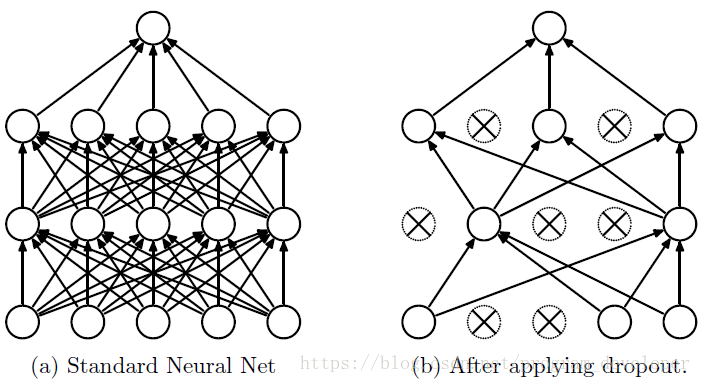
在 TensorFlow 中,使用 `tf.keras.layers.Dropout(舍弃的概率)` 来表示 dropout 层。如果要临时舍弃 20% 的神经元,可以把舍弃的概率设为 0.2。
### 1.5 总结
以上简单的介绍了卷积神经网络基本操作步骤。卷积神经网络在完成特征提取后,就会把数据送入到全连接网络中,因此对于卷积神经网络的整个基本流程可以概括如下:
> 卷积 —》 批标准化 -》 激活 -》 池化 -》 Dropout -》 全连接
>
> **C**onvolutional —》 **B**atch Normal -》 **A**ctivation -》 **P**ooling -》 **D**ropout -》 全连接
说完了卷积神经网络,那如何用一句话总结“卷积”到底是什么呢?
说白了,**卷积就是特征提取器**,就是将数据送入全连接网络之前,对图像进行特征提取的操作。依据上面的执行流程,就是大写加粗的 **CBAPD**。
体现到用 TensorFlow 来搭建网络结构模型中,就如下所示:
```python
model = tf.keras.models.Sequential([
Conv2D(filters=6, kernel_size=(5, 5), padding='same'), # 卷积层 C
BatchNormalization(), # BN 层 B
Activation('relu'), # 激活层 A
MaxPool2D(pool_size=(2, 2), strides=2, padding='same'),# 池化层 P
Dropout(0.2), # Dropout层 D
])
```
| github_jupyter |
```
#hide
#skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
#default_exp data.load
#export
from fastai.torch_basics import *
from torch.utils.data.dataloader import _MultiProcessingDataLoaderIter,_SingleProcessDataLoaderIter,_DatasetKind
_loaders = (_MultiProcessingDataLoaderIter,_SingleProcessDataLoaderIter)
#hide
from nbdev.showdoc import *
bs = 4
letters = list(string.ascii_lowercase)
```
## DataLoader helpers
fastai includes a replacement for Pytorch's *DataLoader* which is largely API-compatible, and adds a lot of useful functionality and flexibility. Before we look at the class, there are a couple of helpers we'll need to define.
```
#export
def _wif(worker_id):
set_num_threads(1)
info = get_worker_info()
ds = info.dataset.d
ds.num_workers,ds.offs = info.num_workers,info.id
set_seed(info.seed)
ds.wif()
class _FakeLoader:
def _fn_noops(self, x=None, *args, **kwargs): return x
_IterableDataset_len_called,_auto_collation,collate_fn,drop_last = None,False,_fn_noops,False
_index_sampler,generator,prefetch_factor = Inf.count,None,2
dataset_kind = _dataset_kind = _DatasetKind.Iterable
def __init__(self, d, pin_memory, num_workers, timeout, persistent_workers):
self.dataset,self.default,self.worker_init_fn = self,d,_wif
store_attr('d,pin_memory,num_workers,timeout,persistent_workers')
def __iter__(self): return iter(self.d.create_batches(self.d.sample()))
@property
def multiprocessing_context(self): return (None,multiprocessing)[self.num_workers>0]
@contextmanager
def no_multiproc(self):
old_num_workers = self.num_workers
try:
self.num_workers = 0
yield self.d
finally: self.num_workers = old_num_workers
_collate_types = (ndarray, Tensor, typing.Mapping, str)
#export
def fa_collate(t):
"A replacement for PyTorch `default_collate` which maintains types and handles `Sequence`s"
b = t[0]
return (default_collate(t) if isinstance(b, _collate_types)
else type(t[0])([fa_collate(s) for s in zip(*t)]) if isinstance(b, Sequence)
else default_collate(t))
#e.g. x is int, y is tuple
t = [(1,(2,3)),(1,(2,3))]
test_eq(fa_collate(t), default_collate(t))
test_eq(L(fa_collate(t)).map(type), [Tensor,tuple])
t = [(1,(2,(3,4))),(1,(2,(3,4)))]
test_eq(fa_collate(t), default_collate(t))
test_eq(L(fa_collate(t)).map(type), [Tensor,tuple])
test_eq(L(fa_collate(t)[1]).map(type), [Tensor,tuple])
#export
def fa_convert(t):
"A replacement for PyTorch `default_convert` which maintains types and handles `Sequence`s"
return (default_convert(t) if isinstance(t, _collate_types)
else type(t)([fa_convert(s) for s in t]) if isinstance(t, Sequence)
else default_convert(t))
t0 = array([1,2])
t = [t0,(t0,t0)]
test_eq(fa_convert(t), default_convert(t))
test_eq(L(fa_convert(t)).map(type), [Tensor,tuple])
#export
class SkipItemException(Exception):
"Raised to notify `DataLoader` to skip an item"
pass
show_doc(SkipItemException, title_level=3)
```
## DataLoader -
```
#export
@funcs_kwargs
class DataLoader(GetAttr):
_noop_methods = 'wif before_iter after_item before_batch after_batch after_iter'.split()
for o in _noop_methods: exec(f"def {o}(self, x=None, *args, **kwargs): return x")
_methods = _noop_methods + 'create_batches create_item create_batch retain \
get_idxs sample shuffle_fn do_batch create_batch'.split()
_default = 'dataset'
def __init__(self, dataset=None, bs=None, num_workers=0, pin_memory=False, timeout=0, batch_size=None,
shuffle=False, drop_last=False, indexed=None, n=None, device=None, persistent_workers=False, **kwargs):
if batch_size is not None: bs = batch_size # PyTorch compatibility
assert not (bs is None and drop_last)
if indexed is None: indexed = (hasattr(dataset,'__getitem__')
and not isinstance(dataset, IterableDataset))
if not indexed and shuffle: raise ValueError("Can only shuffle an indexed dataset (not an iterable one).")
if n is None:
try: n = len(dataset)
except TypeError: pass
store_attr('dataset,bs,shuffle,drop_last,indexed,n,pin_memory,timeout,device')
self.rng,self.num_workers,self.offs = random.Random(random.randint(0,2**32-1)),1,0
if sys.platform == "win32" and IN_NOTEBOOK and num_workers > 0:
print("Due to IPython and Windows limitation, python multiprocessing isn't available now.")
print("So `number_workers` is changed to 0 to avoid getting stuck")
num_workers = 0
self.fake_l = _FakeLoader(self, pin_memory, num_workers, timeout, persistent_workers=persistent_workers)
def __len__(self):
if self.n is None: raise TypeError
if self.bs is None: return self.n
return self.n//self.bs + (0 if self.drop_last or self.n%self.bs==0 else 1)
def get_idxs(self):
idxs = Inf.count if self.indexed else Inf.nones
if self.n is not None: idxs = list(itertools.islice(idxs, self.n))
if self.shuffle: idxs = self.shuffle_fn(idxs)
return idxs
def sample(self):
return (b for i,b in enumerate(self.__idxs) if i//(self.bs or 1)%self.num_workers==self.offs)
def __iter__(self):
self.randomize()
self.before_iter()
self.__idxs=self.get_idxs() # called in context of main process (not workers/subprocesses)
for b in _loaders[self.fake_l.num_workers==0](self.fake_l):
# pin_memory causes tuples to be converted to lists, so convert them back to tuples
if self.pin_memory and type(b) == list: b = tuple(b)
if self.device is not None: b = to_device(b, self.device)
yield self.after_batch(b)
self.after_iter()
if hasattr(self, 'it'): del(self.it)
def create_batches(self, samps):
if self.dataset is not None: self.it = iter(self.dataset)
res = filter(lambda o:o is not None, map(self.do_item, samps))
yield from map(self.do_batch, self.chunkify(res))
def new(self, dataset=None, cls=None, **kwargs):
if dataset is None: dataset = self.dataset
if cls is None: cls = type(self)
cur_kwargs = dict(dataset=dataset, num_workers=self.fake_l.num_workers, pin_memory=self.pin_memory, timeout=self.timeout,
bs=self.bs, shuffle=self.shuffle, drop_last=self.drop_last, indexed=self.indexed, device=self.device)
for n in self._methods:
o = getattr(self, n)
if not isinstance(o, MethodType): cur_kwargs[n] = o
return cls(**merge(cur_kwargs, kwargs))
@property
def prebatched(self): return self.bs is None
def do_item(self, s):
try: return self.after_item(self.create_item(s))
except SkipItemException: return None
def chunkify(self, b): return b if self.prebatched else chunked(b, self.bs, self.drop_last)
def shuffle_fn(self, idxs): return self.rng.sample(idxs, len(idxs))
def randomize(self): self.rng = random.Random(self.rng.randint(0,2**32-1))
def retain(self, res, b): return retain_types(res, b[0] if is_listy(b) else b)
def create_item(self, s):
if self.indexed: return self.dataset[s or 0]
elif s is None: return next(self.it)
else: raise IndexError("Cannot index an iterable dataset numerically - must use `None`.")
def create_batch(self, b): return (fa_collate,fa_convert)[self.prebatched](b)
def do_batch(self, b): return self.retain(self.create_batch(self.before_batch(b)), b)
def to(self, device): self.device = device
def one_batch(self):
if self.n is not None and len(self)==0: raise ValueError(f'This DataLoader does not contain any batches')
with self.fake_l.no_multiproc(): res = first(self)
if hasattr(self, 'it'): delattr(self, 'it')
return res
#export
add_docs(DataLoader, "API compatible with PyTorch DataLoader, with a lot more callbacks and flexibility",
get_idxs = "Return a list of indices to reference the dataset. Calls `shuffle_fn` internally if `shuffle=True`.",
sample = "Same as `get_idxs` but returns a generator of indices to reference the dataset.",
create_batches = "Takes output of `sample` as input, and returns batches of data. Does not apply `after_batch`.",
new = "Create a new `DataLoader` with given arguments keeping remaining arguments same as original `DataLoader`.",
prebatched = "Check if `bs` is None.",
do_item = "Combines `after_item` and `create_item` to get an item from dataset by providing index as input.",
chunkify = "Used by `create_batches` to turn generator of items (`b`) into batches.",
shuffle_fn = "Returns a random permutation of `idxs`.",
randomize = "Set's `DataLoader` random number generator state.",
retain = "Cast each item of `res` to type of matching item in `b` if its a superclass.",
create_item = "Subset of the dataset containing the index values of sample if exists, else next iterator.",
create_batch = "Collate a list of items into a batch.",
do_batch = "Combines `create_batch` and `before_batch` to get a batch of items. Input is a list of items to collate.",
to = "Sets `self.device=device`.",
one_batch = "Return one batch from `DataLoader`.",
wif = "See pytorch `worker_init_fn` for details.",
before_iter = "Called before `DataLoader` starts to read/iterate over the dataset.",
after_item = "Takes output of `create_item` as input and applies this function on it.",
before_batch = "It is called before collating a list of items into a batch. Input is a list of items.",
after_batch = "After collating mini-batch of items, the mini-batch is passed through this function.",
after_iter = "Called after `DataLoader` has fully read/iterated over the dataset.")
```
Arguments to `DataLoader`:
* `dataset`: dataset from which to load the data. Can be either map-style or iterable-style dataset.
* `bs` (int): how many samples per batch to load (if `batch_size` is provided then `batch_size` will override `bs`). If `bs=None`, then it is assumed that `dataset.__getitem__` returns a batch.
* `num_workers` (int): how many subprocesses to use for data loading. `0` means that the data will be loaded in the main process.
* `pin_memory` (bool): If `True`, the data loader will copy Tensors into CUDA pinned memory before returning them.
* `timeout` (float>0): the timeout value in seconds for collecting a batch from workers.
* `batch_size` (int): It is only provided for PyTorch compatibility. Use `bs`.
* `shuffle` (bool): If `True`, then data is shuffled every time dataloader is fully read/iterated.
* `drop_last` (bool): If `True`, then the last incomplete batch is dropped.
* `indexed` (bool): The `DataLoader` will make a guess as to whether the dataset can be indexed (or is iterable), but you can override it with this parameter. `True` by default.
* `n` (int): Defaults to `len(dataset)`. If you are using iterable-style dataset, you can specify the size with `n`.
* `device` (torch.device): Defaults to `default_device()` which is CUDA by default. You can specify device as `torch.device('cpu')`.
Override `create_item` and use the default infinite sampler to get a stream of unknown length (`stop()` when you want to stop the stream).
```
class RandDL(DataLoader):
def create_item(self, s):
r = random.random()
return r if r<0.95 else stop()
L(RandDL())
L(RandDL(bs=4, drop_last=True)).map(len)
dl = RandDL(bs=4, num_workers=4, drop_last=True)
L(dl).map(len)
test_num_workers = 0 if sys.platform == "win32" else 4
test_eq(dl.fake_l.num_workers, test_num_workers)
with dl.fake_l.no_multiproc():
test_eq(dl.fake_l.num_workers, 0)
L(dl).map(len)
test_eq(dl.fake_l.num_workers, test_num_workers)
def _rand_item(s):
r = random.random()
return r if r<0.95 else stop()
L(DataLoader(create_item=_rand_item))
```
If you don't set `bs`, then `dataset` is assumed to provide an iterator or a `__getitem__` that returns a batch.
```
ds1 = DataLoader(letters)
test_eq(L(ds1), letters)
test_eq(len(ds1), 26)
test_shuffled(L(DataLoader(letters, shuffle=True)), letters)
ds1 = DataLoader(letters, indexed=False)
test_eq(L(ds1), letters)
test_eq(len(ds1), 26)
t2 = L(tensor([0,1,2]),tensor([3,4,5]))
ds2 = DataLoader(t2)
test_eq_type(L(ds2), t2)
t3 = L(array([0,1,2], dtype=np.int64),array([3,4,5], dtype=np.int64))
ds3 = DataLoader(t3)
test_eq_type(L(ds3), t3.map(tensor))
ds4 = DataLoader(t3, create_batch=noop, after_iter=lambda: setattr(t3, 'f', 1))
test_eq_type(L(ds4), t3)
test_eq(t3.f, 1)
```
If you do set `bs`, then `dataset` is assumed to provide an iterator or a `__getitem__` that returns a single item of a batch.
```
def twoepochs(d): return ' '.join(''.join(list(o)) for _ in range(2) for o in d)
ds1 = DataLoader(letters, bs=4, drop_last=True, num_workers=0)
test_eq(twoepochs(ds1), 'abcd efgh ijkl mnop qrst uvwx abcd efgh ijkl mnop qrst uvwx')
ds1 = DataLoader(letters,4,num_workers=2)
test_eq(twoepochs(ds1), 'abcd efgh ijkl mnop qrst uvwx yz abcd efgh ijkl mnop qrst uvwx yz')
ds1 = DataLoader(range(12), bs=4, num_workers=3)
test_eq_type(L(ds1), L(tensor([0,1,2,3]),tensor([4,5,6,7]),tensor([8,9,10,11])))
ds1 = DataLoader([str(i) for i in range(11)], bs=4, after_iter=lambda: setattr(t3, 'f', 2))
test_eq_type(L(ds1), L(['0','1','2','3'],['4','5','6','7'],['8','9','10']))
test_eq(t3.f, 2)
it = iter(DataLoader(map(noop,range(20)), bs=4, num_workers=1))
test_eq_type([next(it) for _ in range(3)], [tensor([0,1,2,3]),tensor([4,5,6,7]),tensor([8,9,10,11])])
```
Iterable dataloaders require specific tests.
```
class DummyIterableDataset(IterableDataset):
def __iter__(self):
yield from range(11)
ds1 = DataLoader(DummyIterableDataset(), bs=4)
# Check it yields fine, and check we can do multiple passes
for i in range(3):
test_eq_type(L(ds1), L(tensor([0,1,2,3]),tensor([4,5,6,7]),tensor([8,9,10])))
# Check `drop_last` works fine (with multiple passes, since this will prematurely terminate the iterator)
ds1 = DataLoader(DummyIterableDataset(), bs=4, drop_last=True)
for i in range(3):
test_eq_type(L(ds1), L(tensor([0,1,2,3]),tensor([4,5,6,7])))
class SleepyDL(list):
def __getitem__(self,i):
time.sleep(random.random()/50)
return super().__getitem__(i)
t = SleepyDL(letters)
%time test_eq(DataLoader(t, num_workers=0), letters)
%time test_eq(DataLoader(t, num_workers=2), letters)
%time test_eq(DataLoader(t, num_workers=4), letters)
dl = DataLoader(t, shuffle=True, num_workers=1)
test_shuffled(L(dl), letters)
test_shuffled(L(dl), L(dl))
L(dl)
class SleepyQueue():
"Simulate a queue with varying latency"
def __init__(self, q): self.q=q
def __iter__(self):
while True:
time.sleep(random.random()/100)
try: yield self.q.get_nowait()
except queues.Empty: return
q = Queue()
for o in range(30): q.put(o)
it = SleepyQueue(q)
if not (sys.platform == "win32" and IN_NOTEBOOK):
%time test_shuffled(L(DataLoader(it, num_workers=4)), L(range(30)))
class A(TensorBase): pass
for nw in (0,2):
t = A(tensor([1,2]))
dl = DataLoader([t,t,t,t,t,t,t,t], bs=4, num_workers=nw)
b = first(dl)
test_eq(type(b), A)
t = (A(tensor([1,2])),)
dl = DataLoader([t,t,t,t,t,t,t,t], bs=4, num_workers=nw)
b = first(dl)
test_eq(type(b[0]), A)
list(DataLoader(list(range(50)),bs=32,shuffle=True,num_workers=3))
class A(TensorBase): pass
t = A(tensor(1,2))
tdl = DataLoader([t,t,t,t,t,t,t,t], bs=4, num_workers=2, after_batch=to_device)
b = first(tdl)
test_eq(type(b), A)
# Unknown attributes are delegated to `dataset`
test_eq(tdl.pop(), tensor(1,2))
```
Override `get_idxs` to return the same index until consumption of the DL. This is intented to test consistent sampling behavior when `num_workers`>1.
```
class AdamantDL(DataLoader):
def get_idxs(self):
r=random.randint(0,self.n-1)
return [r] * self.n
test_eq(torch.cat(tuple(AdamantDL((list(range(50))),bs=16,num_workers=4))).unique().numel(),1)
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
# from subprocess import Popen, PIPE
# # test num_workers > 0 in scripts works when python process start method is spawn
# process = Popen(["python", "dltest.py"], stdout=PIPE)
# _, err = process.communicate(timeout=15)
# exit_code = process.wait()
# test_eq(exit_code, 0)
```
| github_jupyter |
## バックテスト用コード
```
flg = {
'buy_signal': 0, # False
'sell_signal': 0, # False
'order': {
'exist': False,
'side': '',
'price': 0,
'count': 0,
},
'position':{
'exist': False,
'side': '',
'price': 0,
'count': 0,
},
'records': {
'buy_count': 0,
'buy_winning': 0,
'buy_return': [], # 利益率
'buy_profit': [],
'buy_holding_periods': [],
'sell_count': 0,
'sell_winning': 0,
'sell_return': [], # 利益率
'sell_profit': [],
'sell_holding_periods': [],
'slippage': [],
'log': [], # textに書き出すログ内容
}
}
import requests
from datetime import datetime
import time
import json
import ccxt
import numpy as np
# テスト用の初期設定
chart_sec = 300 # 5分足
lot = 1 # トレードの枚数
slippage = 0.0005 #手数料やスリッページ
close_condition = 0 #n足後経過するまでは決済しない
# データ取得
def get_price(min, before = 0, after = 0):
price = []
params = {'periods', min}
if before:
params['before'] = before
if after:
params['after'] = after
response = requests.get('https://api.cryptowat.ch/markets/bitflyer/btcfxjpy/ohlc', params)
data = response.json()
if data['result'][str(min)]:
for i in data['result'][str(min)]:
if i[1] and i[2] and i[3] and i[4]:
price.append({
'close_time': i[0],
'close_time_dt': datetime.fromtimestamp(i[0]).strftime('%Y/%m/%d %H:%M'),
'open_price': i[1],
'high_price': i[2],
'low_price': i[3],
'close_price': i[4],
})
return price
else:
print('データがありません')
return
# jsonファイルからの読み込み
def get_price_from_file(path):
file = open(path, 'r', encoding='UTF-8')
price = json.load(file)
return price
# データと時間の表示
def print_price(data):
print('時間: ' + datetime.fromtimestamp(data['close_time']).strftime('%Y/%m/%d %H:%M') + '始値: ' + str(data['open_price']) + '終値: ' + str(data['close_price']))
# データと時間をロギング
def log_price(data):
log = '時間: ' + datetime.fromtimestamp(data['close_time']).strftime('%Y/%m/%d %H:%M') + '始値: ' + str(data['open_price']) + '終値: ' + str(data['close_price']) + '\n'
flg['records']['log'].append(log) #ログの内容もflgに追記していいく
return flg # ログが追記された フラグを返す
# candleが陽線、陰線の基準を満たしているかどうか
def check_candle(data, side):
try:
realbody_rate = abs(data['close_price'] - data['open_price'] / data['high_price'] - data['low_price'])
increase_rate = data['close_price'] / data['open_price'] - 1
except ZeroDivisionError as e:
return False
if side == 'buy':
if data['close_price'] < data['open_price']:
return False
# elif increase_rate < 0.0003:
# return False
# elif realbody_rate < 0.5:
# return False
else:
return True
if side == 'sell':
if data['close_price'] > data['open_price']:
return False
elif increase_rate > -0.0003:
return False
elif realbody_rate < 0.5:
return False
else:
return True
# candleが連続上昇してるか
def check_ascend(data, last_data):
if data['open_price'] > last_data['open_price'] and data['close_price'] > last_data['close_price']:
return True
else:
return False
# candleが連続下落してるか
def check_descen(data, last_data):
if data['open_price'] < last_data['open_price'] and data['close_price'] < last_data['close_price']:
return True
else:
return False
# buy_signalで指値を出す
def buy_signal( data,last_data,flg ):
if flg['buy_signal'] == 0 and check_candle( data,'buy' ):
flg['buy_signal'] = 1
elif flg['buy_signal'] == 1 and check_candle( data,'buy' ) and check_ascend( data,last_data ):
flg['buy_signal'] = 2
elif flg['buy_signal'] == 2 and check_candle( data,'buy' ) and check_ascend( data,last_data ):
log = '3本連続で陽線 なので' + str(data['close_price']) + '円で買い指値を入れます\n'
flg['records']['log'].append(log)
flg['buy_signal'] = 3
# ここにapiのbuyのコード
flg['order']['exist'] = True
flg['order']['side'] = 'BUY'
flg['order']['price'] = round(data['close_price'] * lot)
else:
flg['buy_signal'] = 0
return flg
# sell_signalで指値を出す
def sell_signal( data, last_data, flg ):
if flg['sell_signal'] == 0 and check_candle( data,'sell' ):
flg['sell_signal'] = 1
elif flg['sell_signal'] == 1 and check_candle( data,'sell' ) and check_descend( data,last_data ):
flg['sell_signal'] = 2
elif flg['sell_signal'] == 2 and check_candle( data,'sell' ) and check_descend( data,last_data ):
log = '3本連続で陰線 なので' + str(data['close_price']) + '円で売り指値を入れます\n'
flg['records']['log'].append(log)
flg['sell_signal'] = 3
# ここにapiのsellコードを入れる
flg['order']['exist'] = True
flg['order']['side'] = 'SELL'
flg['order']['price'] = round(data['close_price'] * lot)
else:
flg['sell_signal'] = 0
return flg
# 手仕舞いsignalで成行決済
# 足ごとにポジションがあればメインコードから実行される。ポジションがある時の足の本数分だけ実行されることになる
def close_position(data, last_data, flg):
flg['position']['count'] += 1 # ポジションを持っている足のチェックごとにインクリメント。ポジションがない足の時はカウントしない。決済されたら、0にリセット
if flg['position']['side'] == 'BUY':
if data['close_price'] < last_data['close_price'] and flg['position']['count'] > close_condition:
log = '前回の終値を下回ったので' + str(data['close_price']) + '円あたりで成行で決済します\n'
flg['records']['log'].append(log)
# 成行決済のコード
records(flg, data)
flg['position']['exist'] = False
flg['position']['count'] = 0
if flg['position']['side'] == 'SELL':
if data['close_price'] > last_data['close_peice'] and flg['position']['count'] > close_condition:
log = '前回の終値を上回ったので' + str(data['close_price']) + '円あたりで成行で決済します\n'
flg['records']['log'].append(log)
# 成行決済のコード
records( flg,data )
flg['position']['exist'] = False
flg['position']['count'] = 0
return flg
def check_order(flg):
# 注文状況確認、約定済みなら以下を実行
flg['order']['exist'] = False
flg['order']['count'] = 0
flg['position']['exist'] = True
flg['position']['side'] = flg['order']['side']
flg['position']['price'] = flg['order']['price']
# 一定時間約定してなければキャンセル
return flg
# パフォーマンスの記録
def records(data, flg):
entry_price = flg['position']['price']
exit_price = round(data['close_price'] * lot)
trade_cost = round(exit_price * slippage)
log = 'スリッページ・手数料として ' + str(trade_cost) + '円を考慮します\n'
flg['records']['log'].append(log)
flg['records']['slippage'].append(trade_cost)
# 値幅
buy_profit = exit_price - entry_price - trade_cost
sell_profit = -(exit_price - entry_price) - trade_cost
# 保有時間の記録
# 利益の有無
if flg['position']['side'] == 'BUY':
flg['records']['buy_count'] += 1
flg['records']['buy_profit'].append(buy_profit)
flg['records']['buy_return'].append(round(buy_profit / entry_price * 100, 4))
flg['records']['buy_holding_periods'].append(flg['position']['count'])
if buy_profit > 0:
flg['records']['buy_winning'] += 1
log = str(buy_profit) + '円の利益です\n'
flg['records']['log'].append(log)
else:
log = str(buy_profit) + '円の損失です\n'
flg['records']['log'].append(log)
if flg['position']['side'] == 'SELL':
flg['records']['sell_count'] += 1
flg['records']['sell_profit'].append(sell_profit)
flg['records']['sell_return'].append(round(sell_profit / entry_price * 100, 4))
flg['records']['sell_holding_periods'].append(flg['position']['count'])
if sell_profit > 0:
flg['records']['sell_winning'] += 1
log = str(sell_profit) + '円の利益です\n'
flg['records']['log'].append(log)
else:
log = str(sell_profit) + '円の損失です\n'
flg['records']['log'].append(log)
return flg
# 集計用の関数
def backtest(flg):
buy_gross_profit = np.sum(flg['records']['buy_profit'])
sell_gross_profit = np.sum(flg['records']['sell_profit'])
print('バックテストの結果')
print('--------------------------')
print('買いエントリの成績')
print('--------------------------')
print('トレード回数 : {}回'.format(flg['records']['buy-count']))
print('勝率 : {}%'.format(round(flg['records']['buy-winning'] / flg['records']['buy-count'] * 100, 1)))
print('平均リターン : {}%'.format(round(np.average(flg['records']['buy-return']), 4)))
print('総損益 : {}円'.format(np.sum(flg['records']['buy-profit'])))
print('平均保有期間 : {}足分'.format(round(np.average(flg['records']['buy-holding-periods']),1)))
print('--------------------------')
print('売りエントリの成績')
print('--------------------------')
print('トレード回数 : {}回'.format(flg['records']['sell-count']))
print('勝率 : {}%'.format(round(flg['records']['sell-winning'] / flg['records']['sell-count'] * 100, 1)))
print('平均リターン : {}%'.format(round(np.average(flg['records']['sell-return']), 4)))
print('総損益 : {}円'.format(np.sum(flg['records']['sell-profit'])))
print('平均保有期間 : {}足分'.format(round(np.average(flg['records']['sell-holding-periods']),1)))
print('--------------------------')
print('総合の成績')
print('--------------------------')
print('総損益 : {}円'.format(np.sum(flg['records']['sell-profit']) + np.sum(flg['records']['buy-profit'])))
print('手数料合計 : {}円'.format(np.sum(flg['records']['slippage'])))
# ログファイルの出力
info_time = datetime.now().strftime('%Y/%m/%d-%H:%M')
f = open('./{info_time}-log.txt', 'wt', encoding='UTF-8')
f.writelines(flg['records']['log'])
# メインコード
price = get_price_from_file('path')
# price = get_price(chart_sec, after=1514764800)
print('--------------------------')
print('テスト期間:')
print('開始時点: ' + str(price[0]['close_time_dt']))
print('終了時点: ' + str(price[-1]['close_time_dt']))
print(str(len(price)) + '件のローソク足データで検証')
print('--------------------------')
last_data = price[0]
while i < len(price):
# 未約定を確認
if flg['order']['exist']:
flg = check_order(flg)
data = price[i]
flg = log_price(data, flg)
# ポジションがあれば決済条件を確認、条件を満たせば決済
if flg['position']['exist']:
flg = close_position(data, last_data, flg)
else:
flg = buy_signal(data, last_data, flg) # buy条件チェック
flg = sell_signal(data, last_data, flg) # sell条件チェック
last_data['close_time'] = data['close_time']
last_data['open_price'] = data['open_price']
last_data['close_price'] = data['close_price']
i += 1
backtest(flg)
```
| github_jupyter |
# Keras tutorial - Emotion Detection in Images of Faces
Welcome to the first assignment of week 2. In this assignment, you will:
1. Learn to use Keras, a high-level neural networks API (programming framework), written in Python and capable of running on top of several lower-level frameworks including TensorFlow and CNTK.
2. See how you can in a couple of hours build a deep learning algorithm.
#### Why are we using Keras?
* Keras was developed to enable deep learning engineers to build and experiment with different models very quickly.
* Just as TensorFlow is a higher-level framework than Python, Keras is an even higher-level framework and provides additional abstractions.
* Being able to go from idea to result with the least possible delay is key to finding good models.
* However, Keras is more restrictive than the lower-level frameworks, so there are some very complex models that you would still implement in TensorFlow rather than in Keras.
* That being said, Keras will work fine for many common models.
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "v2a".
* You can find your original work saved in the notebook with the previous version name ("v2").
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of updates
* Changed back-story of model to "emotion detection" from "happy house."
* Cleaned/organized wording of instructions and commentary.
* Added instructions on how to set `input_shape`
* Added explanation of "objects as functions" syntax.
* Clarified explanation of variable naming convention.
* Added hints for steps 1,2,3,4
## Load packages
* In this exercise, you'll work on the "Emotion detection" model, which we'll explain below.
* Let's load the required packages.
```
import numpy as np
from keras import layers
from keras.layers import Input, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D
from keras.layers import AveragePooling2D, MaxPooling2D, Dropout, GlobalMaxPooling2D, GlobalAveragePooling2D
from keras.models import Model
from keras.preprocessing import image
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras.applications.imagenet_utils import preprocess_input
import pydot
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
from kt_utils import *
import keras.backend as K
K.set_image_data_format('channels_last')
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
%matplotlib inline
```
**Note**: As you can see, we've imported a lot of functions from Keras. You can use them by calling them directly in your code. Ex: `X = Input(...)` or `X = ZeroPadding2D(...)`.
In other words, unlike TensorFlow, you don't have to create the graph and then make a separate `sess.run()` call to evaluate those variables.
## 1 - Emotion Tracking
* A nearby community health clinic is helping the local residents monitor their mental health.
* As part of their study, they are asking volunteers to record their emotions throughout the day.
* To help the participants more easily track their emotions, you are asked to create an app that will classify their emotions based on some pictures that the volunteers will take of their facial expressions.
* As a proof-of-concept, you first train your model to detect if someone's emotion is classified as "happy" or "not happy."
To build and train this model, you have gathered pictures of some volunteers in a nearby neighborhood. The dataset is labeled.
<img src="images/face_images.png" style="width:550px;height:250px;">
Run the following code to normalize the dataset and learn about its shapes.
```
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# Normalize image vectors
X_train = X_train_orig/255.
X_test = X_test_orig/255.
# Reshape
Y_train = Y_train_orig.T
Y_test = Y_test_orig.T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
```
**Details of the "Face" dataset**:
- Images are of shape (64,64,3)
- Training: 600 pictures
- Test: 150 pictures
## 2 - Building a model in Keras
Keras is very good for rapid prototyping. In just a short time you will be able to build a model that achieves outstanding results.
Here is an example of a model in Keras:
```python
def model(input_shape):
"""
input_shape: The height, width and channels as a tuple.
Note that this does not include the 'batch' as a dimension.
If you have a batch like 'X_train',
then you can provide the input_shape using
X_train.shape[1:]
"""
# Define the input placeholder as a tensor with shape input_shape. Think of this as your input image!
X_input = Input(input_shape)
# Zero-Padding: pads the border of X_input with zeroes
X = ZeroPadding2D((3, 3))(X_input)
# CONV -> BN -> RELU Block applied to X
X = Conv2D(32, (7, 7), strides = (1, 1), name = 'conv0')(X)
X = BatchNormalization(axis = 3, name = 'bn0')(X)
X = Activation('relu')(X)
# MAXPOOL
X = MaxPooling2D((2, 2), name='max_pool')(X)
# FLATTEN X (means convert it to a vector) + FULLYCONNECTED
X = Flatten()(X)
X = Dense(1, activation='sigmoid', name='fc')(X)
# Create model. This creates your Keras model instance, you'll use this instance to train/test the model.
model = Model(inputs = X_input, outputs = X, name='HappyModel')
return model
```
#### Variable naming convention
* Note that Keras uses a different convention with variable names than we've previously used with numpy and TensorFlow.
* Instead of creating unique variable names for each step and each layer, such as
```
X = ...
Z1 = ...
A1 = ...
```
* Keras re-uses and overwrites the same variable at each step:
```
X = ...
X = ...
X = ...
```
* The exception is `X_input`, which we kept separate since it's needed later.
#### Objects as functions
* Notice how there are two pairs of parentheses in each statement. For example:
```
X = ZeroPadding2D((3, 3))(X_input)
```
* The first is a constructor call which creates an object (ZeroPadding2D).
* In Python, objects can be called as functions. Search for 'python object as function and you can read this blog post [Python Pandemonium](https://medium.com/python-pandemonium/function-as-objects-in-python-d5215e6d1b0d). See the section titled "Objects as functions."
* The single line is equivalent to this:
```
ZP = ZeroPadding2D((3, 3)) # ZP is an object that can be called as a function
X = ZP(X_input)
```
**Exercise**: Implement a `HappyModel()`.
* This assignment is more open-ended than most.
* Start by implementing a model using the architecture we suggest, and run through the rest of this assignment using that as your initial model. * Later, come back and try out other model architectures.
* For example, you might take inspiration from the model above, but then vary the network architecture and hyperparameters however you wish.
* You can also use other functions such as `AveragePooling2D()`, `GlobalMaxPooling2D()`, `Dropout()`.
**Note**: Be careful with your data's shapes. Use what you've learned in the videos to make sure your convolutional, pooling and fully-connected layers are adapted to the volumes you're applying it to.
```
# GRADED FUNCTION: HappyModel
def HappyModel(input_shape):
"""
Implementation of the HappyModel.
Arguments:
input_shape -- shape of the images of the dataset
(height, width, channels) as a tuple.
Note that this does not include the 'batch' as a dimension.
If you have a batch like 'X_train',
then you can provide the input_shape using
X_train.shape[1:]
"""
### START CODE HERE ###
# Feel free to use the suggested outline in the text above to get started, and run through the whole
# exercise (including the later portions of this notebook) once. The come back also try out other
# network architectures as well.
# Define the input placeholder as a tensor with shape input_shape. Think of this as your input image!
X_input = Input(input_shape)
# Zero-Padding: pads the border of X_input with zeroes
X = ZeroPadding2D((3, 3))(X_input)
# CONV -> BN -> RELU Block applied to X
X = Conv2D(32, (7, 7), strides = (1, 1), name = 'conv0')(X)
X = BatchNormalization(axis = 3, name = 'bn0')(X)
X = Activation('relu')(X)
# MAXPOOL
X = MaxPooling2D((2, 2), name='max_pool')(X)
# FLATTEN X (means convert it to a vector) + FULLYCONNECTED
X = Flatten()(X)
X = Dense(1, activation='sigmoid', name='fc')(X)
# Create model. This creates your Keras model instance, you'll use this instance to train/test the model.
model = Model(inputs = X_input, outputs = X, name='HappyModel')
### END CODE HERE ###
return model
```
You have now built a function to describe your model. To train and test this model, there are four steps in Keras:
1. Create the model by calling the function above
2. Compile the model by calling `model.compile(optimizer = "...", loss = "...", metrics = ["accuracy"])`
3. Train the model on train data by calling `model.fit(x = ..., y = ..., epochs = ..., batch_size = ...)`
4. Test the model on test data by calling `model.evaluate(x = ..., y = ...)`
If you want to know more about `model.compile()`, `model.fit()`, `model.evaluate()` and their arguments, refer to the official [Keras documentation](https://keras.io/models/model/).
#### Step 1: create the model.
**Hint**:
The `input_shape` parameter is a tuple (height, width, channels). It excludes the batch number.
Try `X_train.shape[1:]` as the `input_shape`.
```
### START CODE HERE ### (1 line)
happyModel = HappyModel(X_train.shape[1:])
### END CODE HERE ###
```
#### Step 2: compile the model
**Hint**:
Optimizers you can try include `'adam'`, `'sgd'` or others. See the documentation for [optimizers](https://keras.io/optimizers/)
The "happiness detection" is a binary classification problem. The loss function that you can use is `'binary_cross_entropy'`. Note that `'categorical_cross_entropy'` won't work with your data set as its formatted, because the data is an array of 0 or 1 rather than two arrays (one for each category). Documentation for [losses](https://keras.io/losses/)
```
### START CODE HERE ### (1 line)
happyModel.compile(loss='binary_crossentropy', optimizer='adam', metrics = ["accuracy"])
### END CODE HERE ###
```
#### Step 3: train the model
**Hint**:
Use the `'X_train'`, `'Y_train'` variables. Use integers for the epochs and batch_size
**Note**: If you run `fit()` again, the `model` will continue to train with the parameters it has already learned instead of reinitializing them.
```
### START CODE HERE ### (1 line)
happyModel.fit(x = X_train, y = Y_train, epochs = 5, batch_size = 10)
### END CODE HERE ###
```
#### Step 4: evaluate model
**Hint**:
Use the `'X_test'` and `'Y_test'` variables to evaluate the model's performance.
```
### START CODE HERE ### (1 line)
preds = happyModel.evaluate(x = X_test, y = Y_test)
### END CODE HERE ###
print()
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
```
#### Expected performance
If your `happyModel()` function worked, its accuracy should be better than random guessing (50% accuracy).
To give you a point of comparison, our model gets around **95% test accuracy in 40 epochs** (and 99% train accuracy) with a mini batch size of 16 and "adam" optimizer.
#### Tips for improving your model
If you have not yet achieved a very good accuracy (>= 80%), here are some things tips:
- Use blocks of CONV->BATCHNORM->RELU such as:
```python
X = Conv2D(32, (3, 3), strides = (1, 1), name = 'conv0')(X)
X = BatchNormalization(axis = 3, name = 'bn0')(X)
X = Activation('relu')(X)
```
until your height and width dimensions are quite low and your number of channels quite large (≈32 for example).
You can then flatten the volume and use a fully-connected layer.
- Use MAXPOOL after such blocks. It will help you lower the dimension in height and width.
- Change your optimizer. We find 'adam' works well.
- If you get memory issues, lower your batch_size (e.g. 12 )
- Run more epochs until you see the train accuracy no longer improves.
**Note**: If you perform hyperparameter tuning on your model, the test set actually becomes a dev set, and your model might end up overfitting to the test (dev) set. Normally, you'll want separate dev and test sets. The dev set is used for parameter tuning, and the test set is used once to estimate the model's performance in production.
## 3 - Conclusion
Congratulations, you have created a proof of concept for "happiness detection"!
## Key Points to remember
- Keras is a tool we recommend for rapid prototyping. It allows you to quickly try out different model architectures.
- Remember The four steps in Keras:
1. Create
2. Compile
3. Fit/Train
4. Evaluate/Test
## 4 - Test with your own image (Optional)
Congratulations on finishing this assignment. You can now take a picture of your face and see if it can classify whether your expression is "happy" or "not happy". To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Write your image's name in the following code
4. Run the code and check if the algorithm is right (0 is not happy, 1 is happy)!
The training/test sets were quite similar; for example, all the pictures were taken against the same background (since a front door camera is always mounted in the same position). This makes the problem easier, but a model trained on this data may or may not work on your own data. But feel free to give it a try!
```
### START CODE HERE ###
img_path = 'images/my_image.jpg'
### END CODE HERE ###
img = image.load_img(img_path, target_size=(64, 64))
imshow(img)
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
print(happyModel.predict(x))
```
## 5 - Other useful functions in Keras (Optional)
Two other basic features of Keras that you'll find useful are:
- `model.summary()`: prints the details of your layers in a table with the sizes of its inputs/outputs
- `plot_model()`: plots your graph in a nice layout. You can even save it as ".png" using SVG() if you'd like to share it on social media ;). It is saved in "File" then "Open..." in the upper bar of the notebook.
Run the following code.
```
happyModel.summary()
plot_model(happyModel, to_file='HappyModel.png')
SVG(model_to_dot(happyModel).create(prog='dot', format='svg'))
```
| github_jupyter |
```
import math
import matplotlib.pyplot as plt
import numpy as np
class RollingWindowPerceptron:
def __init__(self, M=2000, n=0.5, p=4, expected=0, eps=1e-5):
self.M = M
self.n = n
self.p = p
self.eps = eps
self.w = np.zeros((p + 1,))
self.w[0] = expected
def _quad_error(self, diffs):
s = sum([math.pow(x, 2) for x in diffs])
e = math.sqrt(s)
return e
@staticmethod
def mse(sigmas):
return np.sqrt((sigmas**2).sum())
def fit(self, X, y):
epoch = 0
curr_error = None
while epoch < self.M or (epoch > self.M and curr_error > self.eps):
errors = []
for s, target in zip(X,y):
s = [1] + s
net = sum([s[i] * self.w[i] for i in range(len(s))])
sigma = target - net
errors.append(sigma)
if sigma != 0:
for i in range(len(s)):
self.w[i] += self.n * sigma * s[i]
epoch += 1
if epoch == self.M:
break
print(np.clip(errors, -1.,1.))
curr_error = self._quad_error(errors)
def predict(self, x):
X = [1] + x[len(x) - self.p:]
return sum([X[i]*self.w[i] for i in range(len(X))])
```
# График исходной функции на интервале [a, b]
```
def func(t):
return np.cos(t)**2 - 0.05
N = 20
a = -1
b = 0.5
p = 4
t1 = np.linspace(a, b, num=N)
x1 = func(t1)
plt.plot(t1, x1)
plt.plot(t1, x1, 'go')
plt.grid(True)
```
# Прогноз функции при M=2000 на интервале (b, 2b-a]
```
X_train = np.array([x1[shift:shift+p] for shift in range(x1.size-p)])
y_train = [x1[i] for i in range(p, x1.size)]
perceptron = RollingWindowPerceptron()
perceptron.fit(X_train, y_train)
pred_t = list(set(t2) - set(t1))
pred_t.sort()
pred_x = []
work_x = [] + x1
for i in range(len(x2) - len(x1)):
pred = perceptron.predict(work_x)
pred_x.append(pred)
work_x.append(pred)
plt.plot(t2, x2)
plt.plot(pred_t, pred_x, 'ro')
plt.grid(True)
```
# Прогноз функции при M=4000 на интервале (b, 2b-a]
```
perceptron = RollingWindowPerceptron(M=4000)
perceptron.fit(x2)
pred_t = list(set(t2) - set(t1))
pred_t.sort()
pred_x = []
work_x = [] + x1
for i in range(len(x2) - len(x1)):
pred = perceptron.predict(work_x)
pred_x.append(pred)
work_x.append(pred)
plt.plot(t2, x2)
plt.plot(pred_t, pred_x, 'ro')
plt.grid(True)
```
# Зависимость ошибки от различной ширины окна
```
errors = []
for p_i in range(0, 8): #TODO: до 15-20, когда будет исправлена ошибка с оверфлоу (сейчас оверфлоу при 8)
perceptron = RollingWindowPerceptron(p=p_i)
perceptron.fit(x2)
work_x = [] + x1
for i in range(len(x2) - len(x1)):
pred = perceptron.predict(work_x)
work_x.append(pred)
diffs = [work_x[i] - x2[i] for i in range(len(work_x))]
errors.append(perceptron._quad_error(diffs))
plt.plot(errors)
plt.grid(True)
```
# Зависимость ошибки от различной нормы обучения
```
errors = []
n_range = list(np.arange(0.1, 0.91, 0.01))
for n_i in n_range: #TODO: до 1, когда будет исправлена ошибка с оверфлоу (сейчас оверфлоу при 0.9)
perceptron = RollingWindowPerceptron(n=n_i)
perceptron.fit(x2)
work_x = [] + x1
for i in range(len(x2) - len(x1)):
pred = perceptron.predict(work_x)
work_x.append(pred)
diffs = [work_x[i] - x2[i] for i in range(len(work_x))]
errors.append(perceptron._quad_error(diffs))
plt.plot(n_range, errors)
plt.grid(True)
```
| github_jupyter |
```
from __future__ import absolute_import, division, print_function, unicode_literals
import os, sys
from os.path import abspath
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import warnings
warnings.filterwarnings('ignore')
import keras.backend as k
from keras.models import Sequential
from keras.layers import Dense, Flatten, Conv2D, MaxPooling2D, Activation, Dropout
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import tensorflow as tf
tf.compat.v1.disable_eager_execution()
tf.get_logger().setLevel('ERROR')
from art.estimators.classification import KerasClassifier
from art.attacks.poisoning import PoisoningAttackBackdoor, PoisoningAttackCleanLabelBackdoor
from art.attacks.poisoning.perturbations import add_pattern_bd
from art.utils import load_mnist, preprocess, to_categorical
from art.defences.trainer import AdversarialTrainerMadryPGD
from art.estimators.classification.deep_partition_ensemble import DeepPartitionEnsemble
```
# Load the Data
```
(x_raw, y_raw), (x_raw_test, y_raw_test), min_, max_ = load_mnist(raw=True)
# Random Selection:
n_train = np.shape(x_raw)[0]
num_selection = 10000
random_selection_indices = np.random.choice(n_train, num_selection)
x_raw = x_raw[random_selection_indices]
y_raw = y_raw[random_selection_indices]
# Poison training data
percent_poison = .33
x_train, y_train = preprocess(x_raw, y_raw)
x_train = np.expand_dims(x_train, axis=3)
x_test, y_test = preprocess(x_raw_test, y_raw_test)
x_test = np.expand_dims(x_test, axis=3)
# Shuffle training data
n_train = np.shape(y_train)[0]
shuffled_indices = np.arange(n_train)
np.random.shuffle(shuffled_indices)
x_train = x_train[shuffled_indices]
y_train = y_train[shuffled_indices]
```
# Initialize the Model Architecture
```
# Create Keras convolutional neural network - basic architecture from Keras examples
# Source here: https://github.com/keras-team/keras/blob/master/examples/mnist_cnn.py
def create_model():
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=x_train.shape[1:]))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
```
# Set up the Model Backdoor
```
backdoor = PoisoningAttackBackdoor(add_pattern_bd)
example_target = np.array([0, 0, 0, 0, 0, 0, 0, 0, 0, 1])
pdata, plabels = backdoor.poison(x_test, y=example_target)
plt.imshow(pdata[0].squeeze())
```
# Create the poison data
For this example, we will select 9 as the target class. Thus, the adversary's goal is to poison the model so adding a trigger will result in the trained model misclassifying the triggered input as a 9.
First, the adversary will create a proxy classifier (i.e., a classifier that is similar to the target classifier). As the clean label attack generates noise using PGD in order to encourage the trained classifier to rely on the trigger, it is important that the generated noise be transferable. Thus, adversarial training is used.
```
# Poison some percentage of all non-nines to nines
targets = to_categorical([9], 10)[0]
proxy = AdversarialTrainerMadryPGD(KerasClassifier(create_model()), nb_epochs=10, eps=0.15, eps_step=0.001)
proxy.fit(x_train, y_train)
attack = PoisoningAttackCleanLabelBackdoor(backdoor=backdoor, proxy_classifier=proxy.get_classifier(),
target=targets, pp_poison=percent_poison, norm=2, eps=5,
eps_step=0.1, max_iter=200)
pdata, plabels = attack.poison(x_train, y_train)
poisoned = pdata[np.all(plabels == targets, axis=1)]
poisoned_labels = plabels[np.all(plabels == targets, axis=1)]
print(len(poisoned))
for i in range(len(poisoned)):
if poisoned[i][0][0] != 0:
plt.imshow(poisoned[i].squeeze())
plt.show()
print(f"Index: {i} Label: {np.argmax(poisoned_labels[i])}")
break
```
# Initialize the classification models
We will initialize four models. The first is a single model architecture. The other three are DPA models with varying ensemble sizes to demonstrate the tradeoff between clean accuracy and poison accuracy. This make take some time because of the model copying.
```
model = KerasClassifier(create_model())
dpa_model_10 = DeepPartitionEnsemble(model, ensemble_size=10)
dpa_model_20 = DeepPartitionEnsemble(model, ensemble_size=20)
dpa_model_30 = DeepPartitionEnsemble(model, ensemble_size=30)
```
Train the models on the poisoned data
```
model.fit(pdata, plabels, nb_epochs=10)
dpa_model_10.fit(pdata, plabels, nb_epochs=10)
dpa_model_20.fit(pdata, plabels, nb_epochs=10)
dpa_model_30.fit(pdata, plabels, nb_epochs=10)
```
# Evaluate the performance of the trained models on unpoisoned data
The performance of the models appears normal. We see that for the DPA models, the performance drops slightly as the ensemble size increases
```
clean_preds = np.argmax(model.predict(x_test), axis=1)
clean_correct = np.sum(clean_preds == np.argmax(y_test, axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nClean test set accuracy (model): %.2f%%" % (clean_acc * 100))
# Display image, label, and prediction for a clean sample to show how the poisoned model classifies a clean sample
c = 0 # class to display
i = 0 # image of the class to display
c_idx = np.where(np.argmax(y_test, 1) == c)[0][i] # index of the image in clean arrays
plt.imshow(x_test[c_idx].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(clean_preds[c_idx]))
clean_preds = np.argmax(dpa_model_10.predict(x_test), axis=1)
clean_correct = np.sum(clean_preds == np.argmax(y_test, axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nClean test set accuracy (DPA model_10): %.2f%%" % (clean_acc * 100))
# Display image, label, and prediction for a clean sample to show how the poisoned model classifies a clean sample
c = 0 # class to display
i = 0 # image of the class to display
c_idx = np.where(np.argmax(y_test, 1) == c)[0][i] # index of the image in clean arrays
plt.imshow(x_test[c_idx].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(clean_preds[c_idx]))
clean_preds = np.argmax(dpa_model_20.predict(x_test), axis=1)
clean_correct = np.sum(clean_preds == np.argmax(y_test, axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nClean test set accuracy (DPA model_20): %.2f%%" % (clean_acc * 100))
# Display image, label, and prediction for a clean sample to show how the poisoned model classifies a clean sample
c = 0 # class to display
i = 0 # image of the class to display
c_idx = np.where(np.argmax(y_test, 1) == c)[0][i] # index of the image in clean arrays
plt.imshow(x_test[c_idx].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(clean_preds[c_idx]))
clean_preds = np.argmax(dpa_model_30.predict(x_test), axis=1)
clean_correct = np.sum(clean_preds == np.argmax(y_test, axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nClean test set accuracy (DPA model_30): %.2f%%" % (clean_acc * 100))
# Display image, label, and prediction for a clean sample to show how the poisoned model classifies a clean sample
c = 0 # class to display
i = 0 # image of the class to display
c_idx = np.where(np.argmax(y_test, 1) == c)[0][i] # index of the image in clean arrays
plt.imshow(x_test[c_idx].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(clean_preds[c_idx]))
```
# Evaluate the performance of the trained models on poisoned data
When the trigger is added, we see a shift in performance. The single model performs the worst as no defense is in place to mitigate the effect of the poisoned. The DPA models show some robustnesss to the poison as they partition the training data, which spreads the effect of the poison between models in the ensemble.
```
not_target = np.logical_not(np.all(y_test == targets, axis=1))
px_test, py_test = backdoor.poison(x_test[not_target], y_test[not_target])
poison_preds = np.argmax(model.predict(px_test), axis=1)
clean_correct = np.sum(poison_preds == np.argmax(y_test[not_target], axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nPoison test set accuracy (model): %.2f%%" % (clean_acc * 100))
c = 0 # index to display
plt.imshow(px_test[c].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(poison_preds[c]))
poison_preds = np.argmax(dpa_model_10.predict(px_test), axis=1)
clean_correct = np.sum(poison_preds == np.argmax(y_test[not_target], axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nPoison test set accuracy (DPA model_10): %.2f%%" % (clean_acc * 100))
c = 0 # index to display
plt.imshow(px_test[c].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(poison_preds[c]))
poison_preds = np.argmax(dpa_model_20.predict(px_test), axis=1)
clean_correct = np.sum(poison_preds == np.argmax(y_test[not_target], axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nPoison test set accuracy (DPA model_20): %.2f%%" % (clean_acc * 100))
c = 0 # index to display
plt.imshow(px_test[c].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(poison_preds[c]))
poison_preds = np.argmax(dpa_model_30.predict(px_test), axis=1)
clean_correct = np.sum(poison_preds == np.argmax(y_test[not_target], axis=1))
clean_total = y_test.shape[0]
clean_acc = clean_correct / clean_total
print("\nPoison test set accuracy (DPA model_30): %.2f%%" % (clean_acc * 100))
c = 0 # index to display
plt.imshow(px_test[c].squeeze())
plt.show()
clean_label = c
print("Prediction: " + str(poison_preds[c]))
```
| github_jupyter |
```
#export
from fastai2.basics import *
from nbdev.showdoc import *
#default_exp callback.schedule
```
# Hyperparam schedule
> Callback and helper functions to schedule any hyper-parameter
```
from fastai2.test_utils import *
```
## Annealing
```
#export
def annealer(f):
"Decorator to make `f` return itself partially applied."
@functools.wraps(f)
def _inner(start, end): return partial(f, start, end)
return _inner
```
This is the decorator we will use for all of our scheduling functions, as it transforms a function taking `(start, end, pos)` to something taking `(start, end)` and return a function depending of `pos`.
```
#export
@annealer
def SchedLin(start, end, pos): return start + pos*(end-start)
@annealer
def SchedCos(start, end, pos): return start + (1 + math.cos(math.pi*(1-pos))) * (end-start) / 2
@annealer
def SchedNo (start, end, pos): return start
@annealer
def SchedExp(start, end, pos): return start * (end/start) ** pos
SchedLin.__doc__ = "Linear schedule function from `start` to `end`"
SchedCos.__doc__ = "Cosine schedule function from `start` to `end`"
SchedNo .__doc__ = "Constant schedule function with `start` value"
SchedExp.__doc__ = "Exponential schedule function from `start` to `end`"
annealings = "NO LINEAR COS EXP".split()
p = torch.linspace(0.,1,100)
fns = [SchedNo, SchedLin, SchedCos, SchedExp]
#export
def SchedPoly(start, end, power):
"Polynomial schedule (of `power`) function from `start` to `end`"
def _inner(pos): return start + (end - start) * pos ** power
return _inner
for fn, t in zip(fns, annealings):
plt.plot(p, [fn(2, 1e-2)(o) for o in p], label=t)
f = SchedPoly(2,1e-2,0.5)
plt.plot(p, [f(o) for o in p], label="POLY(0.5)")
plt.legend();
show_doc(SchedLin)
sched = SchedLin(0, 2)
test_eq(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.5, 1., 1.5, 2.])
show_doc(SchedCos)
sched = SchedCos(0, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.29289, 1., 1.70711, 2.])
show_doc(SchedNo)
sched = SchedNo(0, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0., 0., 0., 0.])
show_doc(SchedExp)
sched = SchedExp(1, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [1., 1.18921, 1.41421, 1.68179, 2.])
show_doc(SchedPoly)
sched = SchedPoly(0, 2, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.125, 0.5, 1.125, 2.])
p = torch.linspace(0.,1,100)
pows = [0.5,1.,2.]
for e in pows:
f = SchedPoly(2, 0, e)
plt.plot(p, [f(o) for o in p], label=f'power {e}')
plt.legend();
#export
def combine_scheds(pcts, scheds):
"Combine `scheds` according to `pcts` in one function"
assert sum(pcts) == 1.
pcts = tensor([0] + L(pcts))
assert torch.all(pcts >= 0)
pcts = torch.cumsum(pcts, 0)
def _inner(pos):
if pos == 1.: return scheds[-1](1.)
idx = (pos >= pcts).nonzero().max()
actual_pos = (pos-pcts[idx]) / (pcts[idx+1]-pcts[idx])
return scheds[idx](actual_pos.item())
return _inner
```
`pcts` must be a list of positive numbers that add up to 1 and is the same length as `scheds`. The generated function will use `scheds[0]` from 0 to `pcts[0]` then `scheds[1]` from `pcts[0]` to `pcts[0]+pcts[1]` and so forth.
```
p = torch.linspace(0.,1,100)
f = combine_scheds([0.3,0.2,0.5], [SchedLin(0.,1.), SchedNo(1.,1.), SchedCos(1., 0.)])
plt.plot(p, [f(o) for o in p]);
#hide
test_close([f(0.), f(0.15), f(0.3), f(0.4), f(0.5), f(0.7), f(1.)],
[0., 0.5, 1., 1., 1., 0.65451, 0.])
#export
def combined_cos(pct, start, middle, end):
"Return a scheduler with cosine annealing from `start`→`middle` & `middle`→`end`"
return combine_scheds([pct,1-pct], [SchedCos(start, middle), SchedCos(middle, end)])
```
This is a useful helper function for the [1cycle policy](https://sgugger.github.io/the-1cycle-policy.html). `pct` is used for the `start` to `middle` part, `1-pct` for the `middle` to `end`. Handles floats or collection of floats. For example:
```
f = combined_cos(0.25,0.5,1.,0.)
plt.plot(p, [f(o) for o in p]);
#hide
test_close([f(0.), f(0.1), f(0.25), f(0.5), f(1.)], [0.5, 0.67275, 1., 0.75, 0.])
f = combined_cos(0.25, np.array([0.25,0.5]), np.array([0.5,1.]), np.array([0.,0.]))
test_close([f(0.), f(0.1), f(0.25), f(0.5), f(1.)],
[[0.25,0.5], [0.33638,0.67275], [0.5,1.], [0.375,0.75], [0.,0.]])
```
## ParamScheduler -
```
#export
@docs
class ParamScheduler(Callback):
"Schedule hyper-parameters according to `scheds`"
run_after,run_valid = TrainEvalCallback,False
def __init__(self, scheds): self.scheds = scheds
def begin_fit(self): self.hps = {p:[] for p in self.scheds.keys()}
def begin_batch(self): self._update_val(self.pct_train)
def _update_val(self, pct):
for n,f in self.scheds.items(): self.opt.set_hyper(n, f(pct))
def after_batch(self):
for p in self.scheds.keys(): self.hps[p].append(self.opt.hypers[-1][p])
def after_fit(self):
if hasattr(self.learn, 'recorder'): self.recorder.hps = self.hps
_docs = {"begin_fit": "Initialize container for hyper-parameters",
"begin_batch": "Set the proper hyper-parameters in the optimizer",
"after_batch": "Record hyper-parameters of this batch",
"after_fit": "Save the hyper-parameters in the recorder if there is one"}
```
`scheds` is a dictionary with one key for each hyper-parameter you want to schedule, with either a scheduler or a list of schedulers as values (in the second case, the list must have the same length as the the number of parameters groups of the optimizer).
```
learn = synth_learner()
sched = {'lr': SchedLin(1e-3, 1e-2)}
learn.fit(1, cbs=ParamScheduler(sched))
n = len(learn.dbunch.train_dl)
test_close(learn.recorder.hps['lr'], [1e-3 + (1e-2-1e-3) * i/n for i in range(n)])
#hide
#test discriminative lrs
def _splitter(m): return [[m.a], [m.b]]
learn = synth_learner(splitter=_splitter)
sched = {'lr': combined_cos(0.5, np.array([1e-4,1e-3]), np.array([1e-3,1e-2]), np.array([1e-5,1e-4]))}
learn.fit(1, cbs=ParamScheduler(sched))
show_doc(ParamScheduler.begin_fit)
show_doc(ParamScheduler.begin_batch)
show_doc(ParamScheduler.after_batch)
show_doc(ParamScheduler.after_fit)
#export
@patch
def fit_one_cycle(self:Learner, n_epoch, lr_max=None, div=25., div_final=1e5, pct_start=0.25, wd=defaults.wd,
moms=(0.95,0.85,0.95), cbs=None, reset_opt=False):
"Fit `self.model` for `n_epoch` using the 1cycle policy."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr_max is None else lr_max)
lr_max = np.array([h['lr'] for h in self.opt.hypers])
scheds = {'lr': combined_cos(pct_start, lr_max/div, lr_max, lr_max/div_final),
'mom': combined_cos(pct_start, *moms)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
```
The 1cycle policy was introduced by Leslie N. Smith et al. in [Super-Convergence: Very Fast Training of Neural Networks Using Large Learning Rates](https://arxiv.org/abs/1708.07120). It schedules the learning rate with a cosine annealing from `lr_max/div` to `lr_max` then `lr_max/div_final` (pass an array to `lr_max` if you want to use differential learning rates) and the momentum with cosine annealing according to the values in `moms`. The first phase takes `pct_start` of the training. You can optionally pass additional `cbs` and `reset_opt`.
```
#Integration test: training a few epochs should make the model better
learn = synth_learner(lr=1e-2)
xb,yb = learn.dbunch.one_batch()
init_loss = learn.loss_func(learn.model(xb), yb)
learn.fit_one_cycle(2)
assert learn.loss < init_loss
#Scheduler test
lrs,moms = learn.recorder.hps['lr'],learn.recorder.hps['mom']
test_close(lrs, [combined_cos(0.25,1e-2/25,1e-2,1e-7)(i/20) for i in range(20)])
test_close(moms, [combined_cos(0.25,0.95,0.85,0.95)(i/20) for i in range(20)])
#export
@patch
def plot_sched(self:Recorder, figsize=None):
rows,cols = (len(self.hps)+1)//2, min(2, len(self.hps))
figsize = figsize or (6*cols,4*rows)
_, axs = plt.subplots(rows, cols, figsize=figsize)
axs = axs.flatten() if len(self.hps) > 1 else L(axs)
for p,ax in zip(self.hps.keys(), axs):
ax.plot(self.hps[p])
ax.set_ylabel(p)
#hide
#test discriminative lrs
def _splitter(m): return [[m.a], [m.b]]
learn = synth_learner(splitter=_splitter)
learn.fit_one_cycle(1, lr_max=slice(1e-3,1e-2))
#n = len(learn.dbunch.train_dl)
#est_close(learn.recorder.hps['lr'], [1e-3 + (1e-2-1e-3) * i/n for i in range(n)])
learn = synth_learner()
learn.fit_one_cycle(2)
learn.recorder.plot_sched()
#export
@patch
def fit_flat_cos(self:Learner, n_epoch, lr=None, div_final=1e5, pct_start=0.75, wd=defaults.wd,
cbs=None, reset_opt=False):
"Fit `self.model` for `n_epoch` at flat `lr` before a cosine annealing."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr is None else lr)
lr = np.array([h['lr'] for h in self.opt.hypers])
scheds = {'lr': combined_cos(pct_start, lr, lr, lr/div_final)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
learn = synth_learner()
learn.fit_flat_cos(2)
learn.recorder.plot_sched()
#export
@patch
def fit_sgdr(self:Learner, n_cycles, cycle_len, lr_max=None, cycle_mult=2, cbs=None, reset_opt=False, wd=defaults.wd):
"Fit `self.model` for `n_cycles` of `cycle_len` using SGDR."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr_max is None else lr_max)
lr_max = np.array([h['lr'] for h in self.opt.hypers])
n_epoch = cycle_len * (cycle_mult**n_cycles-1)//(cycle_mult-1)
pcts = [cycle_len * cycle_mult**i / n_epoch for i in range(n_cycles)]
scheds = [SchedCos(lr_max, 0) for _ in range(n_cycles)]
scheds = {'lr': combine_scheds(pcts, scheds)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
```
This schedule was introduced by Ilya Loshchilov et al. in [SGDR: Stochastic Gradient Descent with Warm Restarts](https://arxiv.org/abs/1608.03983). It consists of `n_cycles` that are cosine annealings from `lr_max` (defaults to the `Learner` lr) to 0, with a length of `cycle_len * cycle_mult**i` for the `i`-th cycle (first one is `cycle_len`-long, then we multiply the length by `cycle_mult` at each epoch). You can optionally pass additional `cbs` and `reset_opt`.
```
#slow
learn = synth_learner()
with learn.no_logging(): learn.fit_sgdr(3, 1)
test_eq(learn.n_epoch, 7)
iters = [k * len(learn.dbunch.train_dl) for k in [0,1,3,7]]
for i in range(3):
n = iters[i+1]-iters[i]
#The start of a cycle can be mixed with the 0 of the previous cycle with rounding errors, so we test at +1
test_close(learn.recorder.lrs[iters[i]+1:iters[i+1]], [SchedCos(learn.lr, 0)(k/n) for k in range(1,n)])
learn.recorder.plot_sched()
#export
@delegates(Learner.fit_one_cycle)
@patch
def fine_tune(self:Learner, epochs, base_lr=3e-3, freeze_epochs=1, lr_mult=100,
pct_start=0.3, div=5.0, **kwargs):
"Fine tune with `freeze` for `freeze_epochs` then with `unfreeze` from `epochs` using discriminative LR"
self.freeze()
self.fit_one_cycle(freeze_epochs, slice(base_lr), pct_start=0.99, **kwargs)
self.unfreeze()
self.fit_one_cycle(epochs, slice(base_lr/lr_mult, base_lr), pct_start=pct_start, div=div, **kwargs)
```
## LRFind -
```
#export
@docs
class LRFinder(ParamScheduler):
"Training with exponentially growing learning rate"
run_after=Recorder
def __init__(self, start_lr=1e-7, end_lr=10, num_it=100, stop_div=True):
if is_listy(start_lr):
self.scheds = {'lr': [SchedExp(s, e) for (s,e) in zip(start_lr,end_lr)]}
else: self.scheds = {'lr': SchedExp(start_lr, end_lr)}
self.num_it,self.stop_div = num_it,stop_div
def begin_fit(self):
super().begin_fit()
self.learn.save('_tmp')
self.best_loss = float('inf')
def begin_batch(self):
self._update_val(self.train_iter/self.num_it)
def after_batch(self):
super().after_batch()
if self.smooth_loss < self.best_loss: self.best_loss = self.smooth_loss
if self.smooth_loss > 4*self.best_loss and self.stop_div: raise CancelFitException()
if self.train_iter >= self.num_it: raise CancelFitException()
def begin_validate(self): raise CancelValidException()
def after_fit(self):
tmp_f = self.path/self.model_dir/'_tmp.pth'
if tmp_f.exists():
self.learn.load('_tmp')
os.remove(tmp_f)
_docs = {"begin_fit": "Initialize container for hyper-parameters and save the model",
"begin_batch": "Set the proper hyper-parameters in the optimizer",
"after_batch": "Record hyper-parameters of this batch and potentially stop training",
"after_fit": "Save the hyper-parameters in the recorder if there is one and load the original model",
"begin_validate": "Skip the validation part of training"}
#slow
with tempfile.TemporaryDirectory() as d:
learn = synth_learner(path=Path(d))
init_a,init_b = learn.model.a,learn.model.b
with learn.no_logging(): learn.fit(20, cbs=LRFinder(num_it=100))
assert len(learn.recorder.lrs) <= 100
test_eq(len(learn.recorder.lrs), len(learn.recorder.losses))
#Check stop if diverge
if len(learn.recorder.lrs) < 100: assert learn.recorder.losses[-1] > 4 * min(learn.recorder.losses)
#Test schedule
test_eq(learn.recorder.lrs, [SchedExp(1e-7, 10)(i/100) for i in range_of(learn.recorder.lrs)])
#No validation data
test_eq([len(v) for v in learn.recorder.values], [1 for _ in range_of(learn.recorder.values)])
#Model loaded back properly
test_eq(learn.model.a, init_a)
test_eq(learn.model.b, init_b)
test_eq(learn.opt.state_dict()['state'], [{}, {}])
show_doc(LRFinder.begin_fit)
show_doc(LRFinder.begin_batch)
show_doc(LRFinder.after_batch)
show_doc(LRFinder.begin_validate)
#export
@patch
def plot_lr_find(self:Recorder, skip_end=5):
"Plot the result of an LR Finder test (won't work if you didn't do `learn.lr_find()` before)"
lrs = self.lrs if skip_end==0 else self.lrs [:-skip_end]
losses = self.losses if skip_end==0 else self.losses[:-skip_end]
fig, ax = plt.subplots(1,1)
ax.plot(lrs, losses)
ax.set_ylabel("Loss")
ax.set_xlabel("Learning Rate")
ax.set_xscale('log')
#export
@patch
def lr_find(self:Learner, start_lr=1e-7, end_lr=10, num_it=100, stop_div=True, show_plot=True):
"Launch a mock training to find a good learning rate"
n_epoch = num_it//len(self.dbunch.train_dl) + 1
cb=LRFinder(start_lr=start_lr, end_lr=end_lr, num_it=num_it, stop_div=stop_div)
with self.no_logging(): self.fit(n_epoch, cbs=cb)
if show_plot: self.recorder.plot_lr_find()
```
First introduced by Leslie N. Smith in [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/pdf/1506.01186.pdf), the LR Finder trains the model with exponentially growing learning rates from `start_lr` to `end_lr` for `num_it` and stops in case of divergence (unless `stop_div=False`) then plots the losses vs the learning rates with a log scale.
A good value for the learning rates is then either:
- when the slope is the steepest
- one tenth of the minimum before the divergence
```
#slow
with tempfile.TemporaryDirectory() as d:
learn = synth_learner(path=Path(d))
learn.lr_find()
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
# Data Prediction
#### Importing Libraries
```
import tensorflow as tf
from tensorflow.keras import models
import numpy as np
from PIL import Image
import cv2
import imutils
```
#### Global Variables
```
bg = None
temp_image = 'temp.png'
```
### Resize Image
Used to resize the image given as input.
```
def resizeImage(imageName):
basewidth = 100
img = Image.open(imageName)
wpercent = (basewidth/float(img.size[0]))
hsize = int((float(img.size[1])*float(wpercent)))
img = img.resize((basewidth, hsize), Image.ANTIALIAS)
img.save(imageName)
```
## Running Average
```
def run_avg(image, aWeight):
global bg
# initialize the background
if bg is None:
bg = image.copy().astype("float")
return
# compute weighted average, accumulate it and update the background
cv2.accumulateWeighted(image, bg, aWeight)
```
## Segimentation
```
def segment(image, threshold=25):
global bg
# find the absolute difference between background and current frame
diff = cv2.absdiff(bg.astype("uint8"), image)
# threshold the diff image so that we get the foreground
thresholded = cv2.threshold(diff,
threshold,
255,
cv2.THRESH_BINARY)[1]
# get the contours in the thresholded image
(cnts, _) = cv2.findContours(thresholded.copy(),
cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
# return None, if no contours detected
if len(cnts) == 0:
return
else:
# based on contour area, get the maximum contour which is the hand
segmented = max(cnts, key=cv2.contourArea)
return (thresholded, segmented)
```
## Getting Predicted Class
```
def getPredictedClass():
# read the image
image = cv2.imread(temp_image)
# convert to greyscale
gray_image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# change to numpy array
gray_image = gray_image.reshape(89, 100, 1)
# change to numpy array with shape (1, 89, 100, 1) so model can receive it
gray_image = np.array([gray_image])
# predict the image
prediction = model.predict(gray_image)
# return a numpy array with all values for layers
return prediction
```
## Displaying Result
```
def showStatistics(prediction):
gestures = ['fist', 'palm', 'swing', 'ok']
n = len(gestures)
x = 30
y = 30
height = (n+3)*y
width = 500
textImage = np.zeros((height, width, 3), np.uint8)
for i in range(0, len(gestures)):
cv2.putText(textImage,
gestures[i] + ' : ' + f"{prediction[0][i]:.2f}" ,
(x, y),
cv2.FONT_HERSHEY_SIMPLEX,
1,
(255, 255, 255),
2)
y = y + 30
predicted_gesture = gestures[np.argmax(prediction)]
sum = 0.00
for i in prediction[0]:
sum += i
confidence = (np.amax(prediction) / sum) * 100
cv2.putText(textImage,
"Gesture: " + predicted_gesture,
(x, y),
cv2.FONT_HERSHEY_SIMPLEX,
1,
(255, 255, 255),
2)
y += 30
cv2.putText(textImage,
"Confidence: " + str(confidence) + "%",
(x, y),
cv2.FONT_HERSHEY_SIMPLEX,
1,
(255, 255, 255),
2)
cv2.imshow("Statistics ", textImage)
```
#### (BUG) GPU memory overflow
> this is a workaround
```
#
method1 = False
if method1:
import os
os.environ['TF_FORCE_GPU_ALLOW_GROWTH'] = 'true'
tf.compat.v1.reset_default_graph()
gpu_options.allow_growth = True
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
#
method2 = False
if method2:
config = tf.compat.v1.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.7
tf.compat.v1.keras.backend.set_session(
tf.compat.v1.Session(config=config))
#
method3 = False
if method3:
physical_devices = tf.config.list_physical_devices('GPU')
for device in physical_devices:
tf.config.experimental.set_memory_growth(device, True)
```
## Loading Model
```
tf.compat.v1.reset_default_graph()
model = models.load_model('./TrainedModel/')
```
## Predicting
```
# initialize weight for running average
aWeight = 0.5
# region of interest (ROI) coordinates
top, right, bottom, left = 10, 350, 225, 590
# initialize num of frames
num_frames = 0
start_recording = False
# get the reference to the webcam
camera = cv2.VideoCapture(0)
```
Press `s` to start recording, and press `q` to quit.
Please wait for some time till black background is formed.
In case you get camera not found error, intialize the camera again.
```
# keep looping, until interrupted
while(True):
# get the current frame
grabbed, frame = camera.read()
if grabbed:
# resize the frame
frame = imutils.resize(frame, width=700)
# flip the frame so that it is not the mirror view
frame = cv2.flip(frame, 1)
# clone the frame
clone = frame.copy()
# get the ROI
roi = frame[top:bottom, right:left]
# convert the roi to grayscale and blur it
gray = cv2.cvtColor(roi, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (7, 7), 0)
# to get the background, keep looking till a threshold is reached
# so that our running average model gets calibrated
if num_frames < 30:
run_avg(gray, aWeight)
else:
# segment the hand region
hand = segment(gray)
# check whether hand region is segmented
if hand is not None:
# if yes, unpack the thresholded image and
# segmented region
(thresholded, segmented) = hand
# draw the segmented region and display the frame
cv2.drawContours(
clone, [segmented + (right, top)], -1, (0, 0, 255))
if start_recording:
cv2.imwrite(temp_image, thresholded)
resizeImage(temp_image)
# predictedClass, confidence = getPredictedClass()
prediction = getPredictedClass()
showStatistics(prediction)
cv2.imshow("Thesholded", thresholded)
# draw the segmented hand
cv2.rectangle(clone, (left, top), (right, bottom), (0, 255, 0), 2)
# increment the number of frames
num_frames += 1
# display the frame with segmented hand
cv2.imshow("Video Feed", clone)
# observe the keypress by the user
keypress = cv2.waitKey(1) & 0xFF
# if the user pressed "q", then stop looping
if keypress == ord("q"):
break
if keypress == ord("s"):
start_recording = True
else:
print("Error, Please check your camera")
print(camera)
break
# relaease the resources
camera.release()
cv2.destroyAllWindows()
# remove temporary image file
import os
os.remove(temp_image)
```
| github_jupyter |
# Multi-Layer Perceptron, MNIST
MLP를 훈련하여 이미지 분류기를 생성해 본다. 손글씨 데이터[MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database를 사용하여 0~9를 분류하는 모형을 만든다.
다음과 같은 절차로 수행한다.:
>1. Load and visualize the data
2. Define a neural network
3. Train the model
4. Evaluate the performance of our trained model on a test dataset!
시작하기 위해 필요한 패키지를 import한다.
```
# import libraries
import torch
import numpy as np
```
---
## 1. Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
torchvision에서 제공되는 MNIST 데이터 셋을 다운로드 하여 수행한다. `batch_size`는 필요에 따라 선택할 수 있다.
MNIST 데이터 셋을 dataloader를 생성하여 신경망의 입력 데이터로 사용한다.
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='../data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='../data', train=False,
download=True, transform=transform)
print("Number of training data : ", len(train_data))
print("Number of test data : ", len(test_data))
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize a Batch of Training Data
모형을 구성하기 전에 읽어 들인 데이터를 확인하는 차원에서 이미지 데이터를 확인한다.
```
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 10, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### View an Image in More Detail
개별적으로 이미지 하나를 출력해 보면 다음과 같다.
```
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
```
---
## 2. Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
손글씨 인식을 위해 MLP(multi layer perceptron)모형을 구성한다.
기본적인 구조는 784개(28X28)의 pixel정보를 784dim의 텐서로 읽어 들여 0~9 사이의 분류값으로 분류하는 모형을 구현하기 위한 것이다.
- 2개의 hidden layer로 구성한다.
```
import torch.nn as nn
import torch.nn.functional as F
# define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# number of hidden nodes in each layer (512)
hidden_1 = 512
hidden_2 = 512
# linear layer (784 -> hidden_1)
self.fc1 = nn.Linear(28 * 28, hidden_1)
# linear layer (n_hidden -> hidden_2)
self.fc2 = nn.Linear(hidden_1, hidden_2)
# linear layer (n_hidden -> 10)
self.fc3 = nn.Linear(hidden_2, 10)
def forward(self, x):
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
# add hidden layer, with relu activation function
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
# initialize the NN
model = Net()
print(model)
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
classification을 위해서 loss 함수는 cross-entropy를 사용한다.
```
# specify loss function (categorical cross-entropy)
criterion = nn.CrossEntropyLoss()
# specify optimizer (stochastic gradient descent) and learning rate = 0.01
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
```
---
## 3. Train the Network
각각의 batch에 대하여 다음과 같은 순서로 훈련을 진행한다.:
1. Clear the gradients of all optimized variables
2. Forward pass: compute predicted outputs by passing inputs to the model
3. Calculate the loss
4. Backward pass: compute gradient of the loss with respect to model parameters
5. Perform a single optimization step (parameter update)
6. Update average training loss
epoch의 횟수는 훈련시간을 고려하여 정하도록 한다. n_epochs=10으로 하면 훈련시간은 약 6~7분 정도 소요될 것이다.
훈련이 진행되면서 loss의 감소를 확인한다.valid loss가 가장 작은 모형을 저장한다.
```
# number of epochs to train the model
n_epochs = 5
# initialize tracker for minimum validation loss
valid_loss_min = np.Inf # set initial "min" to infinity
print("Training the model.....")
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train() # prep model for training
for data, target in train_loader:
data = data.view(data.shape[0], -1)
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval() # prep model for evaluation
for data, target in valid_loader:
data = data.view(data.shape[0], -1)
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update running validation loss
valid_loss += loss.item()*data.size(0)
# print training/validation statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch+1,
train_loss,
valid_loss
))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'model.pt')
valid_loss_min = valid_loss
```
### Load the Model with the Lowest Validation Loss
훈련중에 저장한 최소 valid loss를 가지는 모형을 불러들인다.
```
model.load_state_dict(torch.load('model.pt'))
```
---
## 4. Test the Trained Network
마지막으로 best model을 이용하여 **test data**를 사용하여 생성된 모형을 테스트한다.
훈련에 사용되지 않은 테스트 데이터를 사용하여 모형을 평가한다.
test loss와 accuracy로 평가할 수 있다.
```
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for evaluation
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize Sample Test Results
테스트 결과를 시각적으로 판단할 수 있다.
해당 테스트 결과에서 `predicted (ground-truth)`형태로 예측값을 확인한다. 모형에 의한 예측값이 실제 데이터의 참값(ground-truth)를 맞추었으면 녹색 틀리면 빨간색으로 표시한다.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 10, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
```
| github_jupyter |
# Import libraries needed to plot data
```
import math
import numpy as np
import pandas as pd
import scipy.special
from bokeh.layouts import gridplot
from bokeh.io import show, output_notebook, save, output_file
from bokeh.plotting import figure
from bokeh.models import BoxAnnotation, HoverTool, ColumnDataSource, NumeralTickFormatter
from scipy.stats import lognorm, norm
```
Set plots to ouput in notebook instead of as a new tab in the browser, comment out or delete if you want the output as a new browser tab
```
# Bokeh output to notebook setting
output_notebook()
```
# Create main functions used to plot the different outputs, CHANGE AT YOUR OWN RISK
```
# Find P10, P50, and P90
def find_nearest(array, value):
array = np.asarray(array)
idx = (np.abs(array - value)).argmin()
return (array[idx], idx)
def make_plot_cdf(title, hist, edges, x, pdf, cdf, x_label):
p = figure(title=title, background_fill_color="#fafafa", x_axis_type='log')
p.quad(top=hist, bottom=0, left=edges[:-1], right=edges[1:],
fill_color="navy", line_color="white", alpha=0.5)
p.line(x, cdf, line_color="orange", line_width=2, alpha=0.7, legend="CDF")
p.x_range.start = 1
p.y_range.start = 0
p.legend.location = "center_right"
p.legend.background_fill_color = "#fefefe"
p.xaxis.axis_label = x_label
p.yaxis.axis_label = 'Pr(x)'
p.grid.grid_line_color = "white"
p.left[0].formatter.use_scientific = False
p.xaxis[0].formatter = NumeralTickFormatter(format="0,0")
return p
def make_plot_probit(title, input_data, x_label):
'''Creates Probit plot for EUR and data that has a log-normal distribution.
'''
# Calculate log-normal distribtion for input data
sigma, floc, scale = lognorm.fit(input_data, floc=0)
mu = math.log(scale)
x = np.linspace(0.001, np.max(input_data) + np.mean(input_data), 1000)
pdf = 1/(x * sigma * np.sqrt(2*np.pi)) * \
np.exp(-(np.log(x)-mu)**2 / (2*sigma**2))
cdf = (1+scipy.special.erf((np.log(x)-mu)/(np.sqrt(2)*sigma)))/2
p = figure(title=title, background_fill_color="#fafafa", x_axis_type='log')
# Prepare input data for plot
input_data_log = np.log(input_data)
# Get percentile of each point by getting rank/len(data)
input_data_log_sorted = np.argsort(input_data_log)
ranks = np.empty_like(input_data_log_sorted)
ranks[input_data_log_sorted] = np.arange(len(input_data_log))
# Add 1 to length of data because norm._ppf(1) is infinite, which will occur for highest ranked value
input_data_log_perc = [(x + 1)/(len(input_data_log_sorted) + 1)
for x in ranks]
input_data_y_values = norm._ppf(input_data_log_perc)
# Prepare fitted line for plot
x_y_values = norm._ppf(cdf)
# Values to display on y axis instead of z values from ppf
y_axis = [1 - x for x in cdf]
# Plot input data values
p.scatter(input_data, input_data_y_values, size=15,
line_color="navy", legend="Input Data", marker='circle_cross')
p.line(x, x_y_values, line_width=3, line_color="red", legend="Best Fit")
# calculate P90, P50, P10
p10_param = find_nearest(cdf, 0.9)
p10 = round(x[p10_param[1]])
p50_param = find_nearest(cdf, 0.5)
p50 = round(x[p50_param[1]])
p90_param = find_nearest(cdf, 0.1)
p90 = round(x[p90_param[1]])
# Add P90, P50, P10 markers
p.scatter(p90, norm._ppf(0.10), size=15, line_color="black",
fill_color='darkred', legend=f"P90 = {int(p90)}", marker='square_x')
p.scatter(p50, norm._ppf(0.50), size=15, line_color="black",
fill_color='blue', legend=f"P50 = {int(p50)}", marker='square_x')
p.scatter(p10, norm._ppf(0.90), size=15, line_color="black",
fill_color='red', legend=f"P10 = {int(p10)}", marker='square_x')
# Add P90, P50, P10 segments
# p.segment(1, norm._ppf(0.10), np.max(x), norm._ppf(0.10), line_dash='dashed', line_width=2, line_color='black', legend="P90")
# p.segment(1, norm._ppf(0.50), np.max(x), norm._ppf(0.50), line_dash='dashed', line_width=2, line_color='black', legend="P50")
# p.segment(1, norm._ppf(0.90), np.max(x), norm._ppf(0.90), line_dash='dashed', line_width=2, line_color='black', legend="P10")
p.segment(p90, -4, p90, np.max(x_y_values), line_dash='dashed',
line_width=2, line_color='darkred', legend=f"P90 = {int(p90)}")
p.segment(p50, -4, p50, np.max(x_y_values), line_dash='dashed',
line_width=2, line_color='blue', legend=f"P50 = {int(p50)}")
p.segment(p10, -4, p10, np.max(x_y_values), line_dash='dashed',
line_width=2, line_color='red', legend=f"P10 = {int(p10)}")
# Find min for x axis
x_min = int(np.log10(np.min(input_data)))
power_of_10 = 10**(x_min)
# Plot Styling
p.x_range.start = power_of_10
p.y_range.start = -3
p.legend.location = "top_left"
p.legend.background_fill_color = "#fefefe"
p.xaxis.axis_label = x_label
p.yaxis.axis_label = 'Z'
p.left[0].formatter.use_scientific = False
p.xaxis[0].formatter = NumeralTickFormatter(format="0,0")
p.yaxis.visible = False
p.title.text = title
p.title.align = 'center'
p.legend.click_policy = "hide"
return p
def make_plot_pdf(title, hist, edges, x, pdf, x_label):
source = ColumnDataSource(data = {
'x' : x,
'pdf': pdf,
})
p = figure(background_fill_color="#fafafa", )
p.quad(top=hist, bottom=0, left=edges[:-1], right=edges[1:],
fill_color="navy", line_color="white", alpha=0.5)
p.line('x', 'pdf', line_color="black", line_width=4, alpha=0.8, legend="PDF",
hover_alpha=0.4, hover_line_color="black", source=source)
# calculate P90, P50, P10
p10_param = find_nearest(cdf, 0.9)
p10 = round(x[p10_param[1]])
p50_param = find_nearest(cdf, 0.5)
p50 = round(x[p50_param[1]])
p90_param = find_nearest(cdf, 0.1)
p90 = round(x[p90_param[1]])
p.line((p90, p90), [0, np.max(pdf)],
line_color='darkred', line_width=3, legend=f"P90 = {int(p90)}")
p.line((p50, p50), [0, np.max(pdf)],
line_color='blue', line_width=3, legend=f"P50 = {int(p50)}")
p.line((p10, p10), [0, np.max(pdf)],
line_color='red', line_width=3, legend=f"P10 = {int(p10)}")
lower = BoxAnnotation(left=p90, right=p50,
fill_alpha=0.1, fill_color='darkred')
middle = BoxAnnotation(left=p50, right=p10,
fill_alpha=0.1, fill_color='blue')
upper = BoxAnnotation(
left=p10, right=x[-1], fill_alpha=0.1, fill_color='darkred')
# Hover Tool
p.add_tools(HoverTool(
tooltips=[
( x_label, '@x{f}' ),
( 'Probability', '@pdf{%0.6Ff}' ), # use @{ } for field names with spaces
]))
# Plot Styling
p.add_layout(lower)
p.add_layout(middle)
p.add_layout(upper)
p.y_range.start = 0
p.x_range.start = 0
p.legend.location = "center_right"
p.legend.background_fill_color = "#fefefe"
p.xaxis.axis_label = x_label
p.yaxis.axis_label = 'Pr(x)'
p.grid.grid_line_color = "white"
p.left[0].formatter.use_scientific = False
p.xaxis[0].formatter = NumeralTickFormatter(format="0,0")
p.title.text = title
p.title.align = 'center'
return p
```
## The data you want to analyze needs to be set equal to the **input_data** variable below. This example uses the dataset supplied in the /Test_Data tab. The input data can be a list, numpy array, pandas series, or DataFrame Column.
```
data = pd.read_csv(
"https://raw.githubusercontent.com/mwentzWW/petrolpy/master/petrolpy/Test_Data/EUR_Data.csv")
data
input_data = data["CUM_MBO"]
```
The **input_data** is fit to a log normal model
```
# lognorm.fit returns (shape, floc, scale)
# shape is sigma or the standard deviation, scale = exp(median)
sigma, floc, scale = lognorm.fit(input_data, floc=0)
mu = math.log(scale)
```
The model parameters are used to construct the histogram, probability density function (pdf) and cumulative density function (cdf)
```
hist, edges = np.histogram(input_data, density=True, bins='auto')
x = np.linspace(0.001, np.max(input_data) + np.mean(input_data), 1000)
pdf = 1/(x * sigma * np.sqrt(2*np.pi)) * \
np.exp(-(np.log(x)-mu)**2 / (2*sigma**2))
cdf = (1+scipy.special.erf((np.log(x)-mu)/(np.sqrt(2)*sigma)))/2
mean = np.exp(mu + 0.5*(sigma**2))
```
Now we create one of each plot, for basic use the only thing you will want to change is the label argument. Replace 'Cum MBO' with whatever label you want for your data.
```
plot_cdf = make_plot_cdf("Log Normal Distribution (n = {}, mean = {}, σ = {})".format(round(len(
input_data), 2), int(mean), round(sigma, 2)), hist, edges, x, pdf, cdf, 'Cum MBO')
plot_pdf = make_plot_pdf("Log Normal Distribution (n = {}, mean = {}, σ = {})".format(round(
len(input_data), 2), int(mean), round(sigma, 2)), hist, edges, x, pdf, 'Cum MBO')
plot_dist = make_plot_probit("Log Normal Distribution (n = {}, mean = {}, σ = {})".format(
round(len(input_data), 2), int(mean), round(sigma, 2)), input_data, 'Cum MBO')
show(plot_cdf)
```
# The show function will return the plot generated. If you want to save the output as an html file, remove the # from the lines below.
```
#output_file("plot_pdf.html")
#save(plot_pdf)
show(plot_pdf)
#output_file("plot_dist.html")
#save(plot_dist)
show(plot_dist)
```
Below are examples of how to calculate the value of each percentile in the cdf. The P50, P10, and P90 are calculated below.
```
# P50 value
p50_param = find_nearest(cdf, 0.5)
p50_value = round(x[p50_param[1]])
p50_value
# P10 value, only 10% of values will have this value or more
p10_param = find_nearest(cdf, 0.9)
p10_value = round(x[p10_param[1]])
p10_value
# P90 value, 90% of values will have this value or more
p90_param = find_nearest(cdf, 0.1)
p90_value = round(x[p90_param[1]])
p90_value
```
| github_jupyter |
## Creating an experiment - UPDATED MAR 29 2022
The first step towards starting the analysis pipeline and creating the `Experiment` object is to create a dictionary (`ExperimentMetainfo`) to collect the metadata fields required for initializing `Experiment`. In `ExperimentMetainfo`, we provide initial information about the experiment.
```
import os
import packerlabimaging as pli
from packerlabimaging.main.paq import PaqData
from packerlabimaging.processing.imagingMetadata import PrairieViewMetadata
LOCAL_DATA_PATH = '/Users/prajayshah/data/oxford-data-to-process/'
REMOTE_DATA_PATH = '/home/pshah/mnt/qnap/Data/'
BASE_PATH = LOCAL_DATA_PATH
ExperimentMetainfo = {
'dataPath': f'{BASE_PATH}/2020-12-19/2020-12-19_t-013/2020-12-19_t-013_Cycle00001_Ch3.tif',
'saveDir': f'{BASE_PATH}/2020-12-19/',
'expID': 'RL109',
'comment': 'two photon imaging + alloptical trials',
}
# create the experiment
# expobj = pli.Experiment(**ExperimentMetainfo)
```
Once created, the new Experiment gets saved to the disk using python's pickle protocol at the location specified in the options while creating the Experiment.
```
# we can now load the new expobj from disk storage using pickle in pli.import_obj():
expobj = pli.import_obj('/home/pshah/Documents/code/packerlabimaging/tests/RL109_analysis.pkl')
# to save the Experiment to disk using pickle from `expobj.save()`:
expobj.save()
# to save the Experiment with a custom path, simply provide a custom .pkl path and use the .save_pkl() method:
expobj.save_pkl(pkl_path='/home/pshah/mnt/qnap/Analysis/2021-01-25/PS12/PS12_new_path.pkl')
```
## Adding trials to an experiment
After creating the initial `expobj` experiment, we move onto loading each trial from the experiment.
There is a built-in `ImagingTrial` workflow for data processing and analysis of an all optical imaging experiment trial called `AllOpticalTrial`. This extends the `TwoPhotonImaging` trial workflow.
The `AllOpticalTrial` is setup in an analogous manner as the `TwoPhotonImaging` workflow. In addition to the fields required for the `TwoPhotonImaging` trial, we supply additional fields that allow for addition of all-optical specific sub-modules. In particular, we supply a `naparm_path` which triggers the workflow to run the `naparm` analysis sub-module for analysis of 2-photon photostimulation protocols setup by NAPARM during the all-optical experiment.
```
initialization_dict = {'naparm_path': f'{BASE_PATH}/2020-12-19/photostim/2020-12-19_RL109_ps_014/',
'dataPath': f'{BASE_PATH}/2020-12-19/2020-12-19_t-013/2020-12-19_t-013_Cycle00001_Ch3.tif',
'saveDir': f'{BASE_PATH}/2020-12-19/',
'date': '2020-12-19',
'trialID': 't-013',
'expID': 'RL109',
'expGroup': 'all optical trial with LFP',
'comment': ''}
from packerlabimaging import AllOpticalTrial
# create the all optical trial
paqs_loc = f'{BASE_PATH}/2020-12-19/2020-12-19_RL109_013.paq' # path to the .paq files for the selected trials
dataPath = initialization_dict['dataPath']
imparams = PrairieViewMetadata(pv_xml_dir=os.path.dirname(dataPath), microscope='Bruker 2pPlus')
tmdata = PaqData.paqProcessingTwoPhotonImaging(paq_path=paqs_loc, frame_channel='frame_clock')
aotrial = AllOpticalTrial(imparams=imparams, tmdata=tmdata, **initialization_dict)
```
Load in the newly created `AllOpticalTrial` object:
```
aotrial = pli.import_obj(pkl_path='')
```
As with all other `ImagingTrial` objects, each `AllOpticalTrial` object is added to the overall experiment.
```
# after adding a trial to the experiment, it can be loaded using the Experiment object as well:
```
| github_jupyter |
<img src="Logo.png" width="100" align="left"/>
# <center> Preparatory Unit project:</center>
Congratulations on finishing the lessons content for this preparatory unit!!
At this stage it's important to test your theoritical concepts from a practical side and that's exactely the goal of this project.
## Some guidelines:
1. To run a cell you can use the shortcut use : Shift + Enter
2. Only sections mentioned as To-Do are the places where you should put in your own code other than that we do not recommend that you change the provided code.
3. You will be graded for the visibility of your code so make sure you respect the correct indentation and your code contains suitable variables names.
4. This notebook is designed in a sequential way so if you solve your project on different days make sure to run the previous cells before you can run the one you want.
5. Teacher assistants in th slack space remain available to answer any questions you might have.
>Best of luck !
## Project Sections:
In this project you will have a chance to practice most of the important aspects we saw throughout The Preparatory Unit.
This project is divided into 5 sections:
1. [Setting the environement](#set_env)
2. [Importing necessary tools](#importing)
3. [SQLite section](#sql)
4. [Data types section](#datatypes)
5. [Linear Algebra section](#algebra)
### 1. Setting the environement: <a id='set_env'></a>
```
# Make sure you have virtualenv installed
!pip install --user virtualenv
# To-Do: create a virtual environement called myenv
!python -m venv myenv
# Activate the environement
! myenv\Scripts\activate.bat
# Add this virtual environement to Jupyter notebook
!pip install --user ipykernel
!python -m ipykernel install --user --name=myenv
# Install the necessary dependencies
!pip install scipy
!pip install numpy
```
> Please check if you have sqlite installed on your device. For more informations head to the sql lesson
### 2. Importing necessary tools:<a id='importing'></a>
```
from data import database_manager as dm
import utils
from matplotlib import pyplot
from linear_algebra import curve_fitting as cf
```
### 3. SQLite section : <a id='sql'></a>
```
# create a connection to the database
connection = dm.create_connection("longley.db")
# To-Do : retrieve rows of the table
rows = dm.select_all(connection)
dm.print_rows(rows)
```
> Since at this stage we already retrieved our data it's more memory efficient to close the connection to our database.
```
#To-Do close connection using the close_connection function from the data_manager file (dm)
dm.close_connection(connection)
```
### 4. Data types section : <a id='datatypes'></a>
Let's check the datatypes of the retrieved rows
```
rows
```
> This is a list containing multiple tuples, each tuple is a row in the Table with each element within this tuple being a string.
We will be executing mathematical operations on these values and hence we need them in numerical format. Each value contains decimal fractions which means the suitable type to convert to is either double or float. In this case we need to convert these values to a float fomat. Head up to the "utils.py" file and set the function convert_to_floats to be able to do so.
```
# To-Do convert to an ndarray of floats by calling the function convert_to_floats from the utils file
# make sure to set some requirements in that function before you call it here
data = utils.convert_to_floats(rows)
# let's check the shape
data.shape
# Let's see the format
data
```
### 5. Linear Algebra section: <a id='algebra'></a>
```
# Let's check if the two variables GNP.deflator and year are correlated
x, y = data[:,5],data[:, 0]
pyplot.scatter(x, y)
pyplot.xlabel("Year")
pyplot.ylabel("GNP.deflactor")
pyplot.show()
```
> You can clearly see that the two variables: GNP.deflator (y axis) and year (x axis). In other words the GNP.deflactor is increasing throughout the years.
Under this trend it makes sense that we can fit a line to these data points, a line that can describe this trend. And this is our task for this section.
#### Explanation:
Curve fitting aims to find the perfect curve equation for a number of correlated variables. In our example we aim to find the equation for the line that can perfectly fit this point . Such a line should be at minimum distance from all points in average.
Because we are dealing with two variables only, the line's equation should be of the form : y = a*x + b . Which is a typical linear equation.
To acheieve this you will have to :
1. Head to the file linear_algebra/curve_fiiting.py file.
2. Set the objective function's code (function set_objective), objective function is the function that returns the typical shape of our wanted linear equation ( a*x+b), Please delete the "pass" statement and write your code.
3. Here in this notebook in the cell below, call the function get_results and pass to it x and y and get back the optimal values of "a" and "b".
```
# To-Do get the values of a and b using the get_result function
a,b = cf.get_results(x,y)
# plotting the result
from numpy import arange
pyplot.scatter(x, y)
# define a sequence of inputs between the smallest and largest known inputs
x_line = arange(min(x), max(x), 1)
# calculate the output for the range
y_line = cf.set_objective(x_line, a, b)
# create a line plot for the mapping function
pyplot.plot(x_line, y_line, '--', color='red')
pyplot.show()
```
> yohooo ! It's indeed working!!!
# Final thoughts :
This curve fitting process can have many use cases within the machine learning workflow.
A curve fitting can be used as a way to fill in missing values. Datasets aren't always clean. In fact in 90% of the cases we need to do some pre-processing and cleaning for the data before using it in any analysis. In many cases, this cleaning can include filling the missing values, in other words you have some data points with some missing values for some features, if we know that we have a "model" a curve that is supposed to model the trend(or correlation between two of our existing features we can use it to infer these missing values. So as a result Curve fitting can be used in the data cleaning step of the workflow.
Another use case, is when the curve fitting is our end goal, Thus we are cleaning and modeling because the end objective is to have such an equation, in this case the curve fitting is the heart of the Machine learning project.
| github_jupyter |
```
import warnings
warnings.filterwarnings('ignore')
import ipywidgets as widgets
from IPython.display import display, clear_output
#!jupyter nbextension enable --py widgetsnbextension --sys-prefix
#!jupyter serverextension enable voila --sys-prefix
# Image Widget
file = open("grandma.jpg", "rb")
image = file.read()
image_headline = widgets.Image(
value=image,
format='jpg',
width='300'
)
label_headline = widgets.Label(
value='Photo by CDC on Unsplash',
style={'description_width': 'initial'}
)
vbox_headline = widgets.VBox([image_headline, label_headline])
# grandson/granddaughter
grand = widgets.ToggleButtons(
options=['grandson', 'granddaughter']
)
# name
name = widgets.Text(placeholder='Your name here')
date = widgets.DatePicker(description='Pick a Date')
# number of friends
friends = widgets.IntSlider(
value=3, # default value
min=0,
max=10,
step=1,
style={'description_width': 'initial', 'handle_color': '#16a085'}
)
# button send
button_send = widgets.Button(
description='Send to grandma',
tooltip='Send',
style={'description_width': 'initial'}
)
output = widgets.Output()
def on_button_clicked(event):
with output:
clear_output()
print("Sent message: ")
print(f"Dear Grandma! This is your favourite {grand.value}, {name.value}.")
print(f"I would love to come over on {date.value} for dinner, if that's okay for you!")
print(f"Also, if you don't mind, I'll bring along {friends.value} hungry ghosts for your delicious food!")
button_send.on_click(on_button_clicked)
vbox_result = widgets.VBox([button_send, output])
# stacked right hand side
text_0 = widgets.HTML(value="<h1>Dear Grandma!</h1>")
text_1 = widgets.HTML(value="<h2>This is your favourite</h2>")
text_2= widgets.HTML(value="<h2>I would love to come over on </h2>")
text_3= widgets.HTML(value="<h2>for dinner, if that's okay for you!</h2>")
text_4= widgets.HTML(value="<h2>Also, if you don't mind, I'll bring along </h2>")
text_5= widgets.HTML(value="<h2>hungry ghosts for dinner, if that's okay for you!</h2>")
vbox_text = widgets.VBox([text_0, text_1, grand, name, text_2, date, text_3, text_4, friends, text_5, vbox_result])
page = widgets.HBox([vbox_headline, vbox_text])
display(page)
!pip freeze > requirements.txt
```
| github_jupyter |
```
import pandas as pd
df = pd.read_csv("./swissvotes-dataset.csv", delimiter=";", na_values=".")
# Exclude data from votes with missing values (very first and of the last)
df = df.iloc[1:-12, :]
# Extract some data rows for comparison and verification
volk = df["volk"].iloc[:]
stand = df["stand"].iloc[:]
kt_ja = df["kt-ja"].iloc[:]
kt_nein = df["kt-nein"].iloc[:]
ktjaproz = df["ktjaproz"].iloc[:]
staende = kt_ja + kt_nein
invalid_results = (staende > 23).sum() + (staende < 22).sum()
print("Number of invalid staende results: {}".format(invalid_results))
volk_wrong = (df["volk"] != (df["volkja-proz"] > 50.0)).sum()
print("Check calculation of volk: {}".format(volk_wrong))
delta_ktjaproz = (df["ktjaproz"] - (100.0 * df["kt-ja"] / staende)).abs().sum()
print("Total absolute sum of deltas for ktjaproz: {}".format(delta_ktjaproz))
delta_volkja_proz = (df["volkja-proz"] - (100.0 * df["volkja"] / df["gultig"])).abs().sum()
print("Total absolute sum of deltas for volkja-proz: {}".format(delta_volkja_proz))
delta_bet = (df["bet"] - (100.0 * df["stimmen"] / df["berecht"])).abs().sum()
print("Total absolute sum of deltas for bet: {}".format(delta_bet))
invalid_berecht = (df["berecht"] < df["stimmen"]).sum()
print("Number of invalid berecht: {}".format(invalid_berecht))
delta_stimmen = (df["stimmen"] - df["leer"] - df["ungultig"] - df["gultig"])
print("Total absolute sum of deltas for stimmen: {}".format(delta_stimmen.abs().sum()))
stimmen_discrepancies = (delta_stimmen != 0).sum()
print("Number of discrepancies in stimmen: {}".format(stimmen_discrepancies))
delta_gultig = (df["gultig"] - df["volkja"] - df["volknein"])
print("Total absolute sum of deltas for gultig: {}".format(delta_gultig.abs().sum()))
gultig_discrepancies = (delta_gultig != 0).sum()
print("Number of discrepancies in gultig: {}".format(gultig_discrepancies))
#df = df.reset_index(drop=True)
c_berecht = df.iloc[:, df.columns.str.endswith("-berecht")]
c_stimmen = df.iloc[:, df.columns.str.endswith("-stimmen")]
c_stimmen.columns = c_stimmen.columns.str.replace("-stimmen", "")
c_bet = df.iloc[:, df.columns.str.endswith("-bet")]
c_gultig = df.iloc[:, df.columns.str.endswith("-gultig")]
c_gultig.columns = c_gultig.columns.str.replace("-gultig", "")
c_ja = df.iloc[:, df.columns.str.endswith("-ja")]
c_ja = c_ja.drop(columns=["inserate-ja", "kt-ja"])
c_ja.columns = c_ja.columns.str.replace("-ja", "")
c_nein = df.iloc[:, df.columns.str.endswith("-nein")]
c_nein = c_nein.drop(columns=["inserate-nein", "kt-nein"])
c_nein.columns = c_nein.columns.str.replace("-nein", "")
c_japroz = df.iloc[:, df.columns.str.endswith("-japroz")]
c_japroz.columns = c_japroz.columns.str.replace("-japroz", "")
c_annahme = df.iloc[:, df.columns.str.endswith("-annahme")]
c_annahme.columns = c_annahme.columns.str.replace("-annahme", "")
c_discrepancies_gultig = (c_gultig - c_ja - c_nein != 0)
print("Discrepancies gultig at cantonal level: {}".format(c_discrepancies_gultig.abs().sum().sum()))
number_cantons = c_annahme.count(axis=1)
c_weight = c_annahme.count(axis=0)
c_weight[:] = 1.0
c_weight['ar','ai','ow','nw','bl','bs'] = 0.5
wrong_result = (c_annahme != (c_japroz >= 50.0))
import numpy as np
coord = np.where(wrong_result)
coordinates = [(x,y) for x, y in zip(coord[0], coord[1])]
discrepancies_staende = ((c_annahme * c_weight).sum(axis=1) != kt_ja)
discrepancies_staende.index[discrepancies_staende == True]
stand_accepted = (c_annahme * c_weight).sum(axis=1) > (number_cantons/2)
(stand_accepted != stand).sum()
(number_cantons - staende - 3.0).sum()
```
| github_jupyter |
```
import json
import random
import scipy
import numpy as np
import matplotlib.pyplot as plt
from skimage import color
from skimage import io
from skimage import filters, feature
from sklearn.utils import shuffle
from sklearn import svm
from keras.models import Sequential
import keras.layers as l
from tensorflow.keras.utils import to_categorical
```
# Loading data
This code is used to load the data from the assets directory
```
images = []
bunnies = []
data = None
with open("../../Boundary_Box/coordinates.json", 'r+') as file:
data = json.loads(file.read())
for jpg in data:
for coords in data[jpg]:
im = io.imread(f"../../Boundary_Box/assets/{jpg}")
x_coords = [coords['begin'][0], coords['end'][0]]
y_coords = [coords['begin'][1], coords['end'][1]]
images.append(im)
bunnies.append(im[ min(y_coords) : max(y_coords) , min(x_coords) : max(x_coords) ])
```
# Utility functions
This code can be used as utility later on
```
def max_pooling(image, n):
shape = image.shape
output = np.zeros((shape[0]//n, shape[1]//n))
for y in range(1, shape[1]-1, n):
for x in range(1, shape[0]-1, n):
region = image[x-(n//2):x+((n//2)+1), y-(n//2):y+((n//2)+1)]
output[(x-1)//n, (y-1)//n] = region.max()
return output
```
# Filters
This code is used to test if there is a different way to make some features more visible to use as patches, instead of just using black and white images.
* **Obeservations:**\
It looks like that using a smooth filter where the selected pixel weighs more than the rest, combined with a laplacian filter from the scipy library seems to really enhance subtle features.
* **Conclusion:**\
I will be using the combination of a smooth filter with a laplacian filter to extract patches and train the neural network.
```
im = images[230]
b = bunnies[230]
smooth_mean_1=[ [1/9,1/9,1/9],
[1/9,1/9,1/9],
[1/9,1/9,1/9]]
smooth_mean_2=[ [1/16,1/16,1/16],
[1/16,1/5,1/16],
[1/16,1/16,1/16]]
laplacian=[ [0.5,1,0.5],
[1,-6,1],
[0.5,1,0.5]]
laplace_im = filters.laplace(b)
smoothIMG=scipy.ndimage.convolve(b, smooth_mean_1)
laplacianIMG=scipy.ndimage.convolve(smoothIMG, laplacian)
laplacianIMG += 127
smoothIMG2=scipy.ndimage.convolve(b, smooth_mean_2)
laplacian2=scipy.ndimage.convolve(smoothIMG2, laplacian)
laplacian2 += 127
smoothIMG3=scipy.ndimage.convolve(b, smooth_mean_2)
laplacian3=filters.laplace(smoothIMG3)
fig, ax = plt.subplots(1,6, figsize=(25, 8))
ax[0].imshow(b, cmap='gray')
ax[0].set_title("original")
ax[1].imshow(laplace_im, cmap='gray')
ax[1].set_title("raw laplacian")
ax[2].imshow(laplacianIMG, cmap='gray')
ax[2].set_title("smooth9 and selfmade laplacian")
ax[3].imshow(laplacian2, cmap='gray')
ax[3].set_title("smooth16 with self made laplacian")
ax[4].imshow(laplacian3, cmap='gray')
ax[4].set_title("smooth16 with raw laplacian")
ax[5].imshow(feature.canny(b, sigma=0.5), cmap='gray')
ax[5].set_title("canny sigma 2. lowT=40, highT=120")
plt.show()
```
# Patch selection
This code is used to test to see if I can select patches from the boundary boxes.
* **Result:**\
I used an edge detection filter on the image and boundary box image, then I managed to create 50 patches from within the boundary box, and 50 outside of it.
```
# pic_num = random.sample(range(0, len(images)), len(images)-1)
patch_size = 30 # (N x N) pixels
patches, labels = [], []
smooth_filter = [
[1/16,1/16,1/16],
[1/16,1/5,1/16],
[1/16,1/16,1/16]
]
for i in range(len(images)):
im = images[i]
bunny = bunnies[i]
# smooth_bunny=scipy.ndimage.convolve(bunny, smooth_filter)
# bunny=filters.laplace(smooth_bunny)
# smooth_image=scipy.ndimage.convolve(im, smooth_filter)
# im=filters.laplace(smooth_image)
for j in range(50):
# boundary box coordinates
x_b = random.randint((patch_size//2)+1, bunny.shape[0]-(patch_size//2))
y_b = random.randint((patch_size//2)+1, bunny.shape[1]-(patch_size//2))
x_coords_b = [x_b-patch_size//2, x_b+patch_size//2]
y_coords_b = [y_b-patch_size//2, y_b+patch_size//2]
patch_b = bunny[ min(x_coords_b) : max(x_coords_b), min(y_coords_b) : max(y_coords_b) ]
patches.append(patch_b)
labels.append(1)
##############################################################################################################
# outside boundary box coordinates range
x_range = [x for x in range((patch_size//2), im.shape[0]-(patch_size//2))]
y_range = [y for y in range((patch_size//2), im.shape[1]-(patch_size//2))]
# remove subsection of the boundary box from the list with all coordinates
name = list(data.keys())[i]
y_coords_data_arrays = [data[name][0]['begin'][0], data[name][0]['end'][0]]
x_coords_data_arrays = [data[name][0]['begin'][1], data[name][0]['end'][1]]
del x_range[min(x_coords_data_arrays)-(patch_size//2)+1 : max(x_coords_data_arrays)+(patch_size//2)]
del y_range[min(y_coords_data_arrays)-(patch_size//2)+1 : max(y_coords_data_arrays)+(patch_size//2)]
#pick random coordinate
x = random.choice(y_range)
y = random.choice(x_range)
x_coords = [x-patch_size//2, x+patch_size//2]
y_coords = [y-patch_size//2, y+patch_size//2]
patch_pic = im[ min(y_coords) : max(y_coords), min(x_coords) : max(x_coords) ]
patches.append(patch_pic)
labels.append(0)
##############################################################################################################
############## VISUALIZATION ###############
# fig, ax = plt.subplots(2, 2, figsize=(12, 13))
# ax[0,0].imshow(patch_pic, cmap='gray')
# ax[0,0].set_title(f"Patch outside boundary box")
# ax[0,1].imshow(images[i], cmap='gray')
# ax[0,1].set_title(f"{list(data.keys())[i]}")
# ax[0,1].scatter(x=[x_coords[0], x_coords[1]], y=[y_coords[0], y_coords[1]], c='b')
# ax[0,1].scatter(x=[y_coords_data_arrays[0], y_coords_data_arrays[1]], y=[x_coords_data_arrays[0], x_coords_data_arrays[1]], c='r')
# ax[1,0].imshow(patch_b, cmap='gray')
# ax[1,0].set_title(f"Patch within boundary box")
# ax[1,1].imshow(bunnies[i], cmap='gray')
# ax[1,1].set_title(f"Boundary box content")
# ax[1,1].scatter(x=[y_coords_b[0], y_coords_b[1]], y=[x_coords_b[0], x_coords_b[1]], c='r')
# plt.show()
############################################
fig, ax = plt.subplots(1)
ax.imshow(patches[110], cmap='gray')
ax.set_title(f"{labels[110]}")
plt.show()
```
# Neural networks
This code is used to train neural networks
### Loading test- and train data/labels
```
data_arr, label = np.array(patches), np.array(labels)
data_arr, label = shuffle(data_arr, label)
len_data = len(data_arr)
train_data = data_arr[:len_data//3 *2]
train_labels = label[:len_data//3 *2]
test_data = data_arr[len_data//3 *2:]
test_labels = label[len_data//3 *2:]
# Normalizeren van de images
train_data = (train_data / 255) - 0.5
test_data = (test_data / 255) - 0.5
# Reshapen van de images zodat ze de juiste dimensies hebben
train_data = np.expand_dims(train_data, axis=4)
test_data = np.expand_dims(test_data, axis=4)
print(train_data.shape)
# plt.imshow(train_data[126])
```
## Creating a SVM model
This code is used to train a SVM
```
x = len(train_data[0])
y = len(train_data[0][0])
train_data2 = np.reshape(train_data, (len(train_data), x*y) )
test_data2 = np.reshape(test_data, (len(test_data), x*y) )
print(train_data2.shape)
clf = svm.SVC(gamma='scale', C=100)
clf.fit(train_data2, train_labels)
correct = 0
for i, data in enumerate(test_data2):
res = clf.predict([data])
if res[0] == test_labels[i]:
correct += 1
print("Accuracy =", round((correct/len(test_data2))*100, 2) )
```
## Creating the CNN model
### Creating the model
This code creates a CNN model with given layers so that it can be used to classify input into 2 classes
```
shape = train_data.shape
num_filters = 20
filter_size = 8
pool_size = 8
model = Sequential([
l.Conv3D(20, (8, 8, 1), input_shape=(shape[1], shape[2], shape[3], shape[4]), strides=(3,3,3)),
l.Dense(100, activation='relu'),
l.MaxPooling3D(pool_size=3, padding='same'),
l.Conv3D(20, (2, 2, 1), strides = (1,1,3)),
# l.Dense(50, activation='sigmoid'), DIT GEEFT MEER ACCURACY
l.Flatten(None),
l.Dense(2, activation='softmax')
])
model.summary()
```
### Compiling the model
```
model.compile('adam', loss='categorical_crossentropy', metrics=['accuracy'])
model.fit(train_data, to_categorical(train_labels), epochs=10)
validation_data = train_data, to_categorical(test_labels)
test_loss, test_acc = model.evaluate(test_data, to_categorical(test_labels), verbose=2)
print(test_acc)
predictions = model.predict([test_data])
im = np.reshape(test_data[100], (30, 30, 3))
print(predictions[100])
plt.imshow(im)
```
# Hit or miss?
This code is used to generate a list to evaluate which pixel belongs to a bunny and which doesnt. And then proceed to calculate a boundary box
```
a = np.array([1,2,3,4])
print(a.shape)
np.reshape(a, (2,2))
print(a.shape)
patch_size = 30
patches_per_image = []
im = images[71]
for x in range(patch_size//2, im.shape[0]-patch_size//2, patch_size):
new_row = []
for y in range(patch_size//2, im.shape[1]-patch_size//2, patch_size):
patch = im[ x-patch_size//2 : x+patch_size//2, y-patch_size//2 : y+patch_size//2 ]
eval_patch = np.reshape(patch, (1,30,30,3,1)) / 255 - 0.5
res = model.predict([eval_patch])
new_row.append(res)
patches_per_image.append(new_row)
# patches.append(patch_b)
# labels.append(1)
plt.imshow(images[71])
dat = np.array(patches_per_image)
lst = []
for x in range(dat.shape[0]):
row = []
for y in range(dat.shape[1]):
# row.append( 0 if round(np.max(dat[x][y]),2) < 0.95 else round(np.max(dat[x][y]),2))
row.append(np.argmax(dat[x][y]))
lst.append(row)
plt.imshow(lst)
plt.axis('off')
plt.savefig("detection.jpg")
size = 8
hit_list = np.array(lst)
blobs = {
}
inc = 5
increase = 5
threshold = 10
for x in range(size//2, hit_list.shape[0], size):
for y in range(size//2, hit_list.shape[1], size):
begin_x, end_x = x-size//2, x+size//2
begin_y, end_y = y-size//2, y+size//2
box = hit_list[ begin_x : end_x , begin_y : end_y]
amount = np.count_nonzero(box == 1)
if amount > threshold:
while np.count_nonzero(box == 1) > threshold:
amount = np.count_nonzero(box == 1)
threshold = amount
new_boxes = {
"top" : hit_list[ begin_x - inc : end_x, begin_y : end_y ],
"bottom" : hit_list[ begin_x : end_x + inc, begin_y : end_y ],
"left" : hit_list[ begin_x : end_x, begin_y - inc : end_y ],
"right" : hit_list[ begin_x : end_x, begin_y : end_y + inc ]
}
best_v = amount + 5
best_b = box
side = None
for key, value in new_boxes.items():
val = np.count_nonzero(value == 1)
if val > best_v:
best_v = val
best_b = value
side = key
if best_v == np.count_nonzero(box == 1):
break
else:
box = best_b
begin_x = begin_x - inc if side == "top" else begin_x
begin_y = begin_y - inc if side == "left" else begin_y
end_x = end_x + inc if side == "bottom" else end_x
end_y = end_y + inc if side == "right" else end_y
# substract one increase from the 'inc' variable because if done with the while loop, the 'inc' value will be one step ahead
blobs[f'{len(blobs) +1}'] = {
"box" : box,
"begin" :
[
(begin_x - (inc - increase)) if side == "top" else begin_x,
(begin_y - (inc - increase)) if side == "left" else begin_y
]
,
"end" :
[
(end_x + (inc - increase)) if side == "bottom" else end_x,
(end_y + (inc - increase)) if side == "right" else end_y
]
}
fig, ax = plt.subplots(2, 2, figsize=(8,6))
ax[0,0].imshow(images[71])
ax[0,0].set_title("original image")
ax[0,1].imshow(hit_list)
ax[0,1].set_title("points matched with a bunny")
ax[1,0].imshow(blobs['1']['box'])
ax[1,0].set_title("Most interesting area")
ax[1,1].imshow(images[71])
ax[1,1].set_title("Boundary box calculated")
ax[1,1].add_patch(plt.Rectangle((
blobs['1']['begin'][1]*30, blobs['1']['begin'][0]*30
),
blobs['1']['end'][0]*30 - blobs['1']['begin'][0]*30,
blobs['1']['end'][1]*30 - blobs['1']['begin'][1]*30,
fill = False,
edgecolor = 'red',
lw = 3
)
)
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Create Points to cluster
Points = pd.DataFrame()
Points.loc[:,0] = [1.91,0.9,1.26,0.61,1.25,1.04,0.53,0.99,1.11,0.1,-0.15,0.83,0.72,0.69,0.74,
0.72,1.09,0.68,0.67,0.82,0.74,0.94,0.64,1.44,0.76,1.06,0.79,0.88,0.76,0.85,
0.88,0.75,0.83,0.85,0.35,0.63,-0.14,-0.04,0.3,-0.52,-0.27,-0.32,-0.08,-0.39,
-0.06,0.09,-0.51,-0.22,-0.03,-0.12,0.01,-0.21,-0.21,0.37,1.18,0,0,-0.66,-0.1,
1.01,1.19,-0.3,-2.2,-1.82,-1.33,-0.84,-2.17,-1.67,-1.38,-1.39,-1.32,-1.49,
-2.16,-1.64,-1.44,-1.58,-1.53,-1.53,-0.27,-1.32,-0.89,-0.33,-1.29]
Points.loc[:,1] = [1.43,0.79,0.52,1.55,0.66,0.62,1.33,1.27,1.04,2.41,1.83,1.02,1.17,0.97,0.91,
0.14,0.53,1.15,0.96,0.87,0.27,-0.15,0.82,0.72,0.84,1.52,0.93,0.91,0.87,0.93,
0.97,1,0.86,0.88,0.55,-1.99,-0.78,-0.32,0.67,-1.75,-0.7,-0.51,-0.37,-0.55,
-0.42,-0.48,0.64,-0.49,-0.51,-0.32,-0.48,-0.57,-0.32,-0.28,-1.51,-0.41,-0.44,
-2.27,-0.67,-0.32,0.43,-1.26,-1.85,-0.16,-0.89,0.05,-0.38,-0.53,-1.75,-0.98,
-0.33,-1.41,-1.33,-0.9,-0.72,-0.77,-0.66,-0.81,-0.87,-0.94,-1.73,0.55,-0.7]
# Create initial cluster centroids
ClusterCentroidGuesses = pd.DataFrame()
ClusterCentroidGuesses.loc[:,0] = [-1, 1, 0]
ClusterCentroidGuesses.loc[:,1] = [2, -2, 0]
def FindLabelOfClosest(Points, ClusterCentroids): # determine Labels from Points and ClusterCentroids
NumberOfClusters, NumberOfDimensions = ClusterCentroids.shape # dimensions of the initial Centroids
Distances = np.array([float('inf')]*NumberOfClusters) # centroid distances
NumberOfPoints, NumberOfDimensions = Points.shape
Labels = np.array([-1]*NumberOfPoints)
for PointNumber in range(NumberOfPoints): # assign labels to all data points
for ClusterNumber in range(NumberOfClusters): # for each cluster
# Get distances for each cluster
# print(" Replace this line with code")
print(f'PointNumber: {PointNumber}, ClusterNumber{ClusterNumber}')
# Distances[ClusterNumber] =
Labels[PointNumber] = np.argmin(Distances) # assign to closest cluster
return Labels # return the a label for each point
def CalculateClusterCentroid(Points, Labels): # determine centroid of Points with the same label
ClusterLabels = np.unique(Labels) # names of labels
NumberOfPoints, NumberOfDimensions = Points.shape
ClusterCentroids = pd.DataFrame(np.array([[float('nan')]*NumberOfDimensions]*len(ClusterLabels)))
for ClusterNumber in ClusterLabels: # for each cluster
# get mean for each label
ClusterCentroids.loc[ClusterNumber, :] = np.mean(Points.loc[ClusterNumber == Labels, :])
return ClusterCentroids # return the a label for each point
def KMeans(Points, ClusterCentroidGuesses):
ClusterCentroids = ClusterCentroidGuesses.copy()
Labels_Previous = None
# Get starting set of labels
Labels = FindLabelOfClosest(Points, ClusterCentroids)
while not np.array_equal(Labels, Labels_Previous):
# Re-calculate cluster centers based on new set of labels
print(" Replace this line with code")
Labels_Previous = Labels.copy() # Must make a deep copy
# Determine new labels based on new cluster centers
print(" Replace this line with code")
return Labels, ClusterCentroids
def Plot2DKMeans(Points, Labels, ClusterCentroids, Title):
for LabelNumber in range(max(Labels)+1):
LabelFlag = Labels == LabelNumber
color = ['c', 'm', 'y', 'b', 'g', 'r', 'c', 'm', 'y', 'b', 'g', 'r', 'c', 'm', 'y'][LabelNumber]
marker = ['s', 'o', 'v', '^', '<', '>', '8', 'p', '*', 'h', 'H', 'D', 'd', 'P', 'X'][LabelNumber]
plt.scatter(Points.loc[LabelFlag,0], Points.loc[LabelFlag,1],
s= 100, c=color, edgecolors="black", alpha=0.3, marker=marker)
plt.scatter(ClusterCentroids.loc[LabelNumber,0], ClusterCentroids.loc[LabelNumber,1], s=200, c="black", marker=marker)
plt.title(Title)
plt.show()
Labels, ClusterCentroids = KMeans(Points, ClusterCentroidGuesses)
Title = 'K-Means Test'
Plot2DKMeans(Points, Labels, ClusterCentroids, Title)
```
| github_jupyter |
# TreeDLib
```
%load_ext autoreload
%autoreload 2
%load_ext sql
#from treedlib import *
# Note: reloading for submodules doesn't work, so we load directly here
from treedlib.util import *
from treedlib.structs import *
from treedlib.templates import *
from treedlib.features import *
import lxml.etree as et
import numpy as np
```
We define three classes of operators:
* _NodeSets:_ $S : 2^T \mapsto 2^T$
* _Indicators:_ $I : 2^T \mapsto \{0,1\}^F$
* _Combinators:_ $C : \{0,1\}^F \times \{0,1\}^F \mapsto \{0,1\}^F$
where $T$ is a given input tree, and $F$ is the dimension of the feature space.
## Binning
```
%sql postgresql://ajratner@localhost:6432/genomics_ajratner2
res_seq = %sql SELECT * FROM genepheno_features WHERE feature LIKE '%SEQ%'
res_dep = %sql SELECT * FROM genepheno_features WHERE feature NOT LIKE '%SEQ%'
%matplotlib inline
import matplotlib.pyplot as plt
seq_lens = [len(rs.feature.split('_')) for rs in res_seq]
n, bins, patches = plt.hist(seq_lens, 50, normed=1, facecolor='green', alpha=0.75)
print([np.percentile(seq_lens, p) for p in [25,50,75]])
n, bins, patches = plt.hist(dep_lens, 50, normed=1, facecolor='green', alpha=0.75)
dep_lens = [len(rs.feature.split('_')) for rs in res_dep]
print([np.percentile(dep_lens, p) for p in [25,50,75]])
```
## Adding new feature types...
```
ds = {'GENE': ['TFB1M']}
gen_feats = compile_relation_feature_generator()
for f in gen_feats(xt.root, gidxs, pidxs):
print(f)
```
# Genomics Debugging Pipeline
* Fix this!
* _Features to add:_
* modifiers of VBs in between
* candidates in between?
* Better way to do siblings, when siblings have children...?
* LeftAll / RightAll
* Also, get unigrams, etc.
* **Use wildcard, e.g. "ABC now causes" --> WORD:LEFT-OF-MENTION[?_causes]**?
* Modifiers before e.g. "We investigated whether..." / NEGATIONS (see Johannes's email / list)
* Handle negation words explicitly?
```
from random import shuffle
RESULTS_ROOT = '/lfs/raiders7/hdd/ajratner/dd-genomics/alex-results/'
def get_exs(header, rel_path, root=RESULTS_ROOT):
rids = []
in_section = False
with open(root + rel_path, 'rb') as f:
for line in f:
if in_section and len(line.strip()) == 0:
break
elif in_section:
rids.append('_'.join(map(lambda x : x[0].upper() + x[1:], line.strip().split('_'))))
elif line.strip() == header:
in_section = True
return rids
false_pos = get_exs('False Positives:', '02-01-16/stats_causation_1500.tsv')
false_negs = get_exs('False Negatives:', '02-01-16/stats_causation_1500.tsv')
#shuffle(false_pos)
#shuffle(false_negs)
#relation_id = false_negs[12]
#print(relation_id)
#relation_id = '20396601_Body.0_287_0_20396601_Body.0_287_25'
relation_id = '18697824_Abstract.0_1_24_18697824_Abstract.0_1_6'
# Connect to correct db
%sql postgresql://ajratner@localhost:6432/genomics_ajratner
# Getting the component IDs
id = relation_id.split('_')
doc_id = id[0]
section_id = id[1][0].upper() + id[1][1:]
sent_id = int(id[2])
gidxs = map(int, relation_id.split(doc_id)[1].strip('_').split('_')[-1].split('-'))
pidxs = map(int, relation_id.split(doc_id)[2].strip('_').split('_')[-1].split('-'))
cids = [gidxs, pidxs]
# Get sentence from db + convert to XMLTree
res = %sql SELECT words, lemmas, poses, ners, dep_paths AS "dep_labels", dep_parents FROM sentences_input WHERE doc_id = :doc_id AND section_id = :section_id AND sent_id = :sent_id;
rows = [dict((k, v.split('|^|')) for k,v in dict(row).items()) for row in res]
xts = map(corenlp_to_xmltree, rows)
xt = xts[0]
# Show XMLTree
xt.render_tree(highlight=[gidxs, pidxs])
# Print TreeDLib features
#print_gen(get_relation_features(xt.root, gidxs, pidxs))
RightNgrams(RightSiblings(Mention(0)), 'lemma').print_apply(xt.root, cids)
seen.add("blah")
"blah" in seen
dict_sub = compile_dict_sub(brown_clusters_path="clusters_VB_NN.lemma.tsv")
Ngrams(Between(Mention(0), Mention(1)), 'word', 2).print_apply(xt.root, cids, dict_sub=dict_sub)
xt.root.xpath("//*[@dep_label='dobj']/@word")
Indicator(Between(Mention(0), Mention(1)), 'dep_label').print_apply(xt.root, cids)
Ngrams(Between(Mention(0), Mention(1)), 'word', 2).print_apply(xt.root, cids)
dict_sub = compile_dict_sub([
('FOUND', set(['found', 'identified', 'discovered'])),
('CAUSES', set(['causes']))
])
Ngrams(Between(Mention(0), Mention(1)), 'word', 2).print_apply(xt.root, cids, dict_sub=dict_sub)
Ngrams(Children(Filter(Between(Mention(0), Mention(1)), 'pos', 'VB')), 'word', 1).print_apply(xt.root, cids)
Ngrams(Children(Filter(Between(Mention(0), Mention(1)), 'pos', 'VB')), 'word', 1).print_apply(xt.root, cids)
```
## Error analysis round 4
### False negatives:
* [0] `24065538_Abstract.0_2_8_24065538_Abstract.0_2_14`:
* **Should this be association instead?**
* "... have been found... however studies of the association between ... and OSA risk have reported inconsistent findings"
* [1] `8844207_Abstract.0_5_6_8844207_Abstract.0_5_1`:
* **"known {{G}} mutations"**
* [2] `24993959_Abstract.1_3_36_24993959_Abstract.1_3_46`:
* `UnicodeDecodeError`!
* [3] `22653594_Abstract.0_1_5_22653594_Abstract.0_1_25-26-27`:
* **Incorrectly labeled**
* [4] `21282350_Abstract.0_1_13_21282350_Abstract.0_1_20`:
* `UnicodeDecodeError`!
* [5] `11461952_Abstract.0_10_8_11461952_Abstract.0_10_15-16`:
* "This study deomstrates that ... can be responsible for ..."
* "{{G}} responsible for {{P}}"
* [6] `25110572_Body.0_103_42_25110572_Body.0_103_18-19`:
* **Incorrectly labeled??**
* [7] `22848613_Body.0_191_7_22848613_Body.0_191_15`:
* **Incorrectly labeled??**
* [8] `19016241_Abstract.0_2_29_19016241_Abstract.0_2_34-35`:
* **Incorrectly labeled??**
* "weakly penetrant"
* [9] `19877056_Abstract.0_2_37_19877056_Abstract.0_2_7`:
* **"{{P}} attributable to {{G}}"**
* [10] `11079449_Abstract.0_5_48_11079449_Abstract.0_5_41`:
* **_Tough example: ref to a list of pairs!_**
* [11] `11667976_Body.0_6_31_11667976_Body.0_6_34-35`:
* **Is this correctly labeled...?**
* [12] `11353725_Abstract.0_7_13_11353725_Abstract.0_7_9`:
* **Is this correctly labeled...?**
* [13] `20499351_Body.0_120_6_20499351_Body.0_120_10-11-12`:
* "Patients homozygous for {{g}} mutation had"
* "had" on path between
* [14] `10511432_Abstract.0_1_12_10511432_Abstract.0_1_23`:
* **Incorrectly labeled...??**
* [15] `17033686_Abstract.0_4_4_17033686_Abstract.0_4_12`:
* "misense mutation in {{G}} was described in a family with {{P}}"
* **_Incorrectly labeled...?_**
* [16] `23288328_Body.0_179_20_23288328_Body.0_179_24-25`:
* **{{G}} - related {{P}}**
* [17] `21203343_Body.0_127_4_21203343_Body.0_127_19`:
* "have been reported in"- **Incorrectly labeled?**
* [18] `9832037_Abstract.0_2_13_9832037_Abstract.0_2_26-27-28`:
* "{{G}} sympotms include {{P}}", "include"
* [19] `18791638_Body.0_8_6_18791638_Body.0_8_0`:
* "{{P}} results from {{G}}"
```
%%sql
-- Get the features + weights for an example
SELECT f.feature, w.weight
FROM
genepheno_features f,
dd_inference_result_variables_mapped_weights w
WHERE
f.relation_id = :relation_id
AND w.description = 'inf_istrue_genepheno_causation_inference--' || f.feature
ORDER BY w.weight DESC;
res = _
sum(r[1] for r in res)
%sql SELECT expectation FROM genepheno_causation_inference_label_inference WHERE relation_id = :relation_id;
```
## Error analysis round 3
### False Positives:
* [0] `18478198_Abstract.0_2_29_18478198_Abstract.0_2_11-12`:
* "our aim was to establish whether"
* [1] `17508172_Abstract.0_4_21_17508172_Abstract.0_4_32`:
* "role"
* "sodium ion channel"
* [2] `19561293_Abstract.0_3_7_19561293_Abstract.0_3_10-11`:
* "are currently unknown"
* [3] `19956409_Abstract.0_1_8_19956409_Abstract.0_1_21`:
* r'^To evaluate'
* "the possible role"
* [4] `19714249_Body.0_130_10_19714249_Body.0_130_18`:
* '^Although"
* "potential role"
* "needs to be replicated"
* "suggests", "possible", "role"
* [5] `16297188_Title.0_1_5_16297188_Title.0_1_14`:
* "role"
* **Incorrectly supervised...?**
* [6] `24412566_Body.0_70_72_24412566_Body.0_70_6`:
* **_Long one with other genes in between..._**
* [7] `16837472_Abstract.3_1_19_16837472_Abstract.3_1_10`:
* "needs to be further studied"
* "associated"
* [8] `14966353_Abstract.0_1_41_14966353_Abstract.0_1_5`:
* `UnicodeError`!
* [9] `15547491_Abstract.0_1_23_15547491_Abstract.0_1_7-8-9-10`:
* r'^To analyze'
## Error analysis round 2
With new DSR code:
### False Positives
* [0] `17183713_Body.0_111_12_17183713_Body.0_111_25`:
* **"unlikely" on path between**
* [1] `19561293_Abstract.0_3_7_19561293_Abstract.0_3_10-11`:
* _"are unknown"- not on dep path between..._
* **Labeling error- doesn't this imply that there is a causal relation??**
* [2] `17167409_Abstract.3_2_5_17167409_Abstract.3_2_13`:
* **"is _not_ a common cause of..." - NEG modifying primary VB on path between!!!**
* [3] `18538017_Body.0_12_5_18538017_Body.0_12_17`:
* **Labeling error!? (marked because only partial P...?)**
* [4] `20437121_Abstract.0_1_30_20437121_Abstract.0_1_15`:
* "to determine" - in phrase between
* [5] `10435725_Abstract.0_1_14_10435725_Abstract.0_1_20`:
* "in mice" - off the main VB
* [6] `23525542_Abstract.0_7_12_23525542_Abstract.0_7_24`:
* **is _not_ due to..."- NEG modifying primary VB on path between!!!**
* [7] `19995275_Abstract.0_1_2_19995275_Abstract.0_1_18`:
* "has been implicated... in various studies with conflicting results"
### False Negatives
* [0] `23874215_Body.0_172_3_23874215_Body.0_172_23-24-25-26`:
* "role", "detected" - dep path between
* [1] `17507029_Abstract.0_2_13_17507029_Abstract.0_2_6-7-8-9-10`:
* "caused by" but also "association"... should do dep path in between...?
* _a tough one..._
* [2] `15219231_Body.0_121_8_15219231_Body.0_121_35`:
* **Incorrect label**
* [3] `25110572_Body.0_103_42_25110572_Body.0_103_18-19`:
* **Incorrect label- should be association?**
* [4] `17909190_Abstract.0_3_16_17909190_Abstract.0_3_25`:
* **Incorrectly labeled...?**
* [5] `22803640_Abstract.0_3_14_22803640_Abstract.0_3_24-25`:
* **Incorrectly labeled- should be association?**
* [6] `11170071_Abstract.0_1_3_11170071_Abstract.0_1_21`:
* **Incorrectly labeled- wrong mention**
* [7] `10511432_Abstract.0_1_12_10511432_Abstract.0_1_23`:
* "A variety of mutations have been detected in patients with..."- should this be association?
* [8] `10797440_Abstract.0_3_16_10797440_Abstract.0_3_3`:
* _This one seems like should be straight-forwards..._
* **{{P}} are due to {{G}}**
* [9] `23275784_Body.0_82_29_23275784_Body.0_82_13`:
* _This one seems like should be straight-forwards..._
* **{{P}} result of / due to mutations in {{G}}**
```
# Filler
```
### To investigate:
1. Correlation with length of sentence? - **_No._**
2. Low-MI words like '\_', 'the', 'gene'?
3. _[tdl] Include sequence patterns too?_
### FNs / recall analysis notes
* `10982191_Title.0_1_8_10982191_Title.0_1_21-22-23`:
* Shorter sentence
* neg. weight from "gene" in between... is this just super common?
* `19353431_Abstract.0_2_12_19353431_Abstract.0_2_1`:
* Shorter sentence
* neg. weight from "gene" in between... is this just super common?
* `23285148_Body.0_4_32_23285148_Body.0_4_3`:
* **Incorrectly labeled: should be false**
* `23316347_Body.0_202_25_23316347_Body.0_202_54`:
* _Longer sentence..._
* **BUG: Missing a left-of-mention (G: "mutation")!**
* neg. weight from "\_" in betweeen
* **BUG: left-of-mention[delay] happens twice!**
* A lot of negative weight from "result"...?
* `21304894_Body.0_110_4_21304894_Body.0_110_9-10-11`:
* Shorter sentence
* A lot of negative weight from "result"...?
* **Is this just from a low-quality DSR?**
* Duplicated features again!
* `21776272_Body.0_60_46_21776272_Body.0_60_39-40`:
* Longer sentence
* A slightly tougher example: an inherited disorder ... with mutations in gene...
* neg. weight from "gene" in between... is this just super common?
* `19220582_Abstract.0_2_20_19220582_Abstract.0_2_5`:
* 'We identified a mutation in a family with...' - should this be a positive example??
* neg. weight from "gene" in between... is this just super common?
* neg. weight from "identify" and "affect"...?
* **'c. mutation' - mutation doesn't get picked up as it's a child off the path...**
* `23456818_Body.0_148_9_23456818_Body.0_148_21-22`:
* `LEMMA:PARENTS-OF-BETWEEN-MENTION-and-MENTION[determine]` has huge negative weight
* gene, patient, distribution, etc. - neg weight
* negative impact from `PARENTS OF`...
* `20429427_Abstract.0_1_2_20429427_Abstract.0_1_14`:
* **Key word like "mutation" is off main path... ("responsible -> mutation -> whose")**
* **STOPWORDS: "the"**
* **BUG: dep_path labels are all None...**, **BUG: left-siblings doubled**
* `21031598_Body.0_24_25_21031598_Body.0_24_9`:
* Need a feature like `direct parent of mention`
* NEG: 'site', 'gene'
* `INV_`
* `22670894_Title.0_1_16_22670894_Title.0_1_7-8`:
* NEG: 'the', 'gene', 'locus'
* **'due to' just dropped from the dep tree!**
* `22887726_Abstract.0_5_33_22887726_Abstract.0_5_54-55`:
* **Incorrectly labeled for causation?**
* `19641605_Abstract.0_3_14_19641605_Abstract.0_3_22`:
* This one has "cause", exp = 0.89, seems like dead match...
* **BUG: doubles of stuff!!!!!**
* `23879989_Abstract.0_1_3_23879989_Abstract.0_1_12-13`:
* This one has "cause", exp = 0.87, seems like dead match...
* **BUG: doubles of stuff!!!!!**
* `LEMMA:FILTER-BY(pos=NN):BETWEEN-MENTION-and-MENTION[_]`
* 'distinct', 'mutation _ cause'...
* **_Why does '\_' have such negative weight??_**
* `21850180_Body.0_62_14_21850180_Body.0_62_26-27`:
* This one again seems like should be a dead match...
* **BUG: Double of word "three"!**
* Key word "responsible" not included...?
* NEG: 'identify', 'i.e.', '_ _ _'
* `20683840_Abstract.0_4_12_20683840_Abstract.0_4_33`:
* UnicodeError!
* `17495019_Title.0_1_5_17495019_Title.0_1_18`:
* **Incorrectly labeled for causation?**
* _Why is '% patients' positive...?_
* `18283249_Abstract.0_3_2_18283249_Abstract.0_3_16-17-18`:
* **'are one of the factors' - is this correctly labeled for causation?**
* `21203343_Body.0_10_3_21203343_Body.0_10_20`:
* **'are described in...' - this at least seems on the border of "causation"**
* expectation 0.85
* **BUG: doubles**
* NEG: `_`
* `24312213_Body.0_110_66_24312213_Body.0_110_73`:
* **Interesting example of isolated subtree which should be direct match!**
* Expectation 0.42???
* NEG: 'mutation result', `_`, 'result', 'influence'
### Final tally:
* 55%: Negative weight from features that seem like they should be stop words
* 25%: Incorrectly labeled or on the border
* 40%: Bug of some sort in TreeDLib
* 30%: Features that seems suprisingly weighted- due to low-quality DSRs?
## TODO:
1. Fix bugs in treedlib - DONE
2. Filter "stopwords" i.e. low-Chi-squared features - DONE
3. Add manual weights to DSRs in `config.py`
## Testing the low-Chi-squared hypothesis
```
%sql SELECT COUNT(*) FROM genepheno_features;
%sql SELECT COUNT(DISTINCT(feature)) FROM genepheno_features;
%%sql
SELECT
gc.is_correct, COUNT(*)
FROM
genepheno_causation gc,
genepheno_features gf
WHERE
gc.relation_id = gf.relation_id
AND gf.feature LIKE '%the%'
GROUP BY
gc.is_correct;
%sql SELECT is_correct, COUNT(*) FROM genepheno_causation GROUP BY is_correct;
P_T = 40022.0/(116608.0+40022.0)
P_F = 116608.0/(116608.0+40022.0)
print(P_T)
print(P_F)
from collections import defaultdict
feats = defaultdict(lambda : [0,0])
with open('/lfs/raiders7/hdd/ajratner/dd-genomics/alex-results/chi-sq/chi-sq-gp.tsv', 'rb') as f:
for line in f:
feat, label, count = line.split('\t')
b = 0 if label == 't' else 1
feats[feat][b] = int(count)
feats['INV_DEP_LABEL:BETWEEN-MENTION-and-MENTION[nsubj_vmod_prepc_by]']
chi_sqs = []
for feat, counts in feats.items():
total = float(counts[0] + counts[1])
chi_sqs.append([
(P_T-(counts[0]/total))**2 + (P_F-(counts[1]/total))**2,
feat
])
chi_sqs.sort()
with open('/lfs/raiders7/hdd/ajratner/dd-genomics/alex-results/chi-sq/chi-sq-gp-computed.tsv', 'wb') as f:
for x in chi_sqs:
f.write('\t'.join(map(str, x[::-1]))+'\n')
len(chi_sqs)
chi_sqs[500000]
thes = filter(lambda x : 'the' in x[1], chi_sqs)
len(thes)
thes[:100]
```
## Testing the length-bias hypothesis
Is their a bias towards longer sentences (because more high-weight keywords?)
```
rows = []
with open('/lfs/raiders7/hdd/ajratner/dd-genomics/alex-results/test-len-corr/all_rel_sents.tsv', 'rb') as f:
for line in f:
r = line.rstrip().split('\t')
rows.append([float(r[1]), len(r[2].split('|^|'))])
print(len(rows))
from scipy.stats import pearsonr
exps, lens = zip(*filter(lambda r : r[0] > 0.7, rows))
pearsonr(exps, lens)
%matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import random
exps, lens = zip(*random.sample(filter(lambda r : r[0] > 0.5, rows), 1000))
plt.scatter(lens, exps)
```
## Debugging pipeline
We'll debug here, also to show the general most current procedure for debugging treedlib on examples in a SQL database (e.g. from DeepDive)
```
%sql postgresql://ajratner@localhost:5432/deepdive_spouse
%%sql
SELECT sentence_text
FROM sentences
WHERE doc_id = '79205745-b593-4b98-8a94-da6b8238fefc' AND sentence_index = 32;
res = %sql SELECT tokens AS "words", lemmas, pos_tags, ner_tags, dep_types AS "dep_labels", dep_tokens AS "dep_parents" FROM sentences WHERE doc_id = '79205745-b593-4b98-8a94-da6b8238fefc' AND sentence_index = 32;
xts = map(corenlp_to_xmltree, res)
xt = xts[0]
xt.render_tree(highlight=[[21,22], [33,34]])
print_gen(get_relation_features(xt.root, [21,22], [33,34]))
```
## Feature focus: Preceding statements which nullify or negate meaning
Example:
> _Ex1:_ To investigate whether mutations in the SURF1 gene are a cause of Charcot-Marie-Tooth -LRB- CMT -RRB- disease
> _Ex2:_ To investigate the genetic effect of a new mutation found in exon 17 of the myophosphorylase -LRB- PYGM -RRB- gene as a cause of McArdle disease -LRB- also known as type 5 glycogenosis -RRB-.
Notes:
* These seem to mostly be **_modifiers of the primary verb_**?
* We are only sampling from a limited set of patterns of sentences (due to narrow DSR set) currently...
* Modifiers in general...?
* _I know how RNNs claim to / do handle this phenomenon..._ *
```
%%sql
SELECT relation_id
FROM genepheno_causation
WHERE doc_id = '15262743' AND section_id = 'Abstract.0' AND sent_id = 1;
ex1_id = ('24027061', 'Abstract.0', 1)
ex1_raw="""
<node dep_parent="0" lemma="investigate" ner="O" pos="VB" word="investigate" word_idx="1"><node dep_parent="2" dep_path="aux" lemma="to" ner="O" pos="TO" word="To" word_idx="0"/><node dep_parent="2" dep_path="ccomp" lemma="cause" ner="O" pos="NN" word="cause" word_idx="10"><node dep_parent="11" dep_path="mark" lemma="whether" ner="O" pos="IN" word="whether" word_idx="2"/><node dep_parent="11" dep_path="nsubj" lemma="mutation" ner="O" pos="NNS" word="mutations" word_idx="3"><node dep_parent="4" dep_path="prep_in" lemma="gene" ner="O" pos="NN" word="gene" word_idx="7"><node dep_parent="8" dep_path="det" lemma="the" ner="O" pos="DT" word="the" word_idx="5"/><node dep_parent="8" dep_path="nn" lemma="surf1" ner="O" pos="NN" word="SURF1" word_idx="6"/></node></node><node dep_parent="11" dep_path="cop" lemma="be" ner="O" pos="VBP" word="are" word_idx="8"/><node dep_parent="11" dep_path="det" lemma="a" ner="O" pos="DT" word="a" word_idx="9"/><node dep_parent="11" dep_path="prep_of" lemma="Charcot-Marie-Tooth" ner="O" pos="NNP" word="Charcot-Marie-Tooth" word_idx="12"/><node dep_parent="11" dep_path="dep" lemma="disease" ner="O" pos="NN" word="disease" word_idx="16"><node dep_parent="17" dep_path="appos" lemma="CMT" ner="O" pos="NNP" word="CMT" word_idx="14"/></node></node></node>
"""
xt1 = XMLTree(et.fromstring(ex1_raw))
ex2_id = ('15262743', 'Abstract.0', 1)
ex2_raw="""
<node dep_parent="0" lemma="investigate" ner="O" pos="VB" word="investigate" word_idx="1"><node dep_parent="2" dep_path="aux" lemma="to" ner="O" pos="TO" word="To" word_idx="0"/><node dep_parent="2" dep_path="dobj" lemma="effect" ner="O" pos="NN" word="effect" word_idx="4"><node dep_parent="5" dep_path="det" lemma="the" ner="O" pos="DT" word="the" word_idx="2"/><node dep_parent="5" dep_path="amod" lemma="genetic" ner="O" pos="JJ" word="genetic" word_idx="3"/><node dep_parent="5" dep_path="prep_of" lemma="mutation" ner="O" pos="NN" word="mutation" word_idx="8"><node dep_parent="9" dep_path="det" lemma="a" ner="O" pos="DT" word="a" word_idx="6"/><node dep_parent="9" dep_path="amod" lemma="new" ner="O" pos="JJ" word="new" word_idx="7"/><node dep_parent="9" dep_path="vmod" lemma="find" ner="O" pos="VBN" word="found" word_idx="9"><node dep_parent="10" dep_path="prep_in" lemma="exon" ner="O" pos="NN" word="exon" word_idx="11"><node dep_parent="12" dep_path="num" lemma="17" ner="NUMBER" pos="CD" word="17" word_idx="12"/><node dep_parent="12" dep_path="prep_of" lemma="gene" ner="O" pos="NN" word="gene" word_idx="19"><node dep_parent="20" dep_path="det" lemma="the" ner="O" pos="DT" word="the" word_idx="14"/><node dep_parent="20" dep_path="nn" lemma="myophosphorylase" ner="O" pos="NN" word="myophosphorylase" word_idx="15"/><node dep_parent="20" dep_path="nn" lemma="pygm" ner="O" pos="NN" word="PYGM" word_idx="17"/></node></node><node dep_parent="10" dep_path="prep_as" lemma="cause" ner="O" pos="NN" word="cause" word_idx="22"><node dep_parent="23" dep_path="det" lemma="a" ner="O" pos="DT" word="a" word_idx="21"/><node dep_parent="23" dep_path="prep_of" lemma="disease" ner="O" pos="NN" word="disease" word_idx="25"><node dep_parent="26" dep_path="nn" lemma="McArdle" ner="PERSON" pos="NNP" word="McArdle" word_idx="24"/><node dep_parent="26" dep_path="vmod" lemma="know" ner="O" pos="VBN" word="known" word_idx="28"><node dep_parent="29" dep_path="advmod" lemma="also" ner="O" pos="RB" word="also" word_idx="27"/><node dep_parent="29" dep_path="prep_as" lemma="glycogenosis" ner="O" pos="NN" word="glycogenosis" word_idx="32"><node dep_parent="33" dep_path="nn" lemma="type" ner="O" pos="NN" word="type" word_idx="30"/><node dep_parent="33" dep_path="num" lemma="5" ner="NUMBER" pos="CD" word="5" word_idx="31"/></node></node></node></node></node></node></node></node>
"""
xt2 = XMLTree(et.fromstring(ex2_raw))
xt1.render_tree()
xt2.render_tree()
```
### Testing XML speeds
How does it compare between:
* parse to XML via this python code, store as string, then parse from string at runtime
* just parse to XML at runtime via this python code?
```
# Map sentence to xmltree
%time xts = map(corenlp_to_xmltree, rows)
# Pre-process to xml string
xmls = [xt.to_str() for xt in map(corenlp_to_xmltree, rows)]
# Parse @ runtime using lxml
%time roots = map(et.fromstring, xmls)
```
### Table example
```
# Some wishful thinking...
table_xml = """
<div class="table-wrapper">
<h3>Causal genomic relationships</h3>
<table>
<tr><th>Gene</th><th>Variant</th><th>Phenotype</th></tr>
<tr><td>ABC</td><td><i>AG34</i></td><td>Headaches during defecation</td></tr>
<tr><td>BDF</td><td><i>CT2</i></td><td>Defecation during headaches</td></tr>
<tr><td>XYG</td><td><i>AT456</i></td><td>Defecasomnia</td></tr>
</table>
</div>
"""
from IPython.core.display import display_html, HTML
display_html(HTML(table_xml))
```
| github_jupyter |
# Week 10 - Create and manage a digital bookstore collection
*© 2021 Colin Conrad*
Welcome to Week 10 of INFO 6270! Last week marked an important milestone, in the sense that you completed the second course unit on core data science skills. Starting this week, we will have three labs on "other skills" that are valuable to data scientists but do not constitute the core practices. This week we will change gears into a topic that is important to most data-related careers: SQL. Many former students have stressed the importance of having basic knowledge of this topic in the workforce.
Many of you are likely to have covered SQL in other courses and this lab assumes that you have seen it before. If you have not explored SQL before, I strongly recommend that you spend a few hours working your way through this week's reading. The Khan Academy [Intro to SQL: Querying and managing data](https://www.khanacademy.org/computing/computer-programming/sql) is an excellent course and may be interesting to you even if you have covered it before. It is important to qualify that they use MySQL, which is ever-so slightly different from the tool that we will use this week: SQLite. The principles are the same regardless.
**This week, we will achieve the following objectives:**
- Create a SQL table with Python
- Conduct simple SQL queries
- Create relations between tables
- Conduct a more complex SQL query
- Create and query a relational table
# Case: Kobo Inc.
[Rakuten Kobo Inc.]( https://www.kobo.com/) is a Canadian company which sells e-books, audiobooks and e-readers and is a competitor to Amazon's bookselling business. Founded as a subsidiary of the Indigo Books and Music Inc., in 2010 Kobo was spun off as an independent company in order to form partnerships with other book retailors which whished to compete with Amazon. The company was later acquired by Japanese e-commerce giant Rakuten and today competes with Amazon for e-book dominance in Canada and Japan.
Like many e-commerce companies, Rakuten Kobo keeps a database of products for download. Though it is unclear whether they use an SQL database specifically, it is likely that the company stores its inventories using a relational database framework. As an e-commerce company, they maintain an inventory of thousands of items, many of which are related to transactions which happen every day. In this lab we will explore hypothetical data structures which may be similar to the technology that Rakuten Kobo uses on the backend.
# Objective 1: Create an SQL table with Python
If you have [kept up with this week's readings](https://www.khanacademy.org/computing/computer-programming/sql), you likely already know that SQL stands for _Structured Query Language_ and is the standard method for communicating with a relational database. Databases are the backbone of virtually every information system used in businesses and organizations. Relational databases are the most common variety of database; it is not difficult to see why using and managing SQL is important. In fact, _**MI, MDI and MBA students who work for companies or governments consistently express to me that this skill is among the most important skills for securing gainful employment*_. If you have not done a module on SQL, I strongly recommend that you at least look through the Khan Academy materials!
It will probably be no surprise to you by now that you can also connect to an SQL database using Python. In this lab we use a simple SQL database called [SQLite](https://www.sqlite.org/index.html). Without going into the details of SQLite, just know that this is a fully functional SQL database that is optimized for small datasets. We will use Python' `sqlite3` library to create, query and retrieve data from an SQLite database. You can read the [documentation for this library here](https://docs.python.org/2/library/sqlite3.html).
As usual, we will start by importing the `sqlite3` library. This library will allow us to connect to an SQLite database.
```
import sqlite3 # import SQLite
```
It is important to remember that `sqlite3` is not the database itself but the means to connect to the database. To make `sqlite3` work we must first create a *connection* to a database. Though it does not yet exist, we will connect to `kobo.db` in your data folder. When `sqlite3` connects to a database which does not yet exist, it will automatically create a new database for you.
If you are interested, you can open the `/data` folder and see the database that was just created!
```
conn = sqlite3.connect('data/kobo.db') # this is your database connection. We will create a database in the data folder
```
Normally we would connect to an SQL database using a command line, which would allow us to execute SQL commands, similarly to the Khan Academy example. Similarly, SQLite must connect to the database and it does so using a series of `cursor()` methods. We can create a cursor object by declaring it as below. This will allow us to use python to execute SQL queries of our database.
```
c = conn.cursor() # this is the tool for interfacing between SQLite and Python
```
Good work! We are now connected to a database. Let's talk about that a bit before proceeding.
### A bit more about relational databases
Relational databases have been around a long time (in computer years) and were originally proposed by E. F. Codd (1970). What makes relational databases different from data tables (which we have explored so far) is that they actually consist of many tables which *relate* to one another in a variety of ways. For example:
- The Government of Canada may maintain a table of citizens and a table of passports. Each passport belongs to exactly one citizen and each citizen may have a passport. This is an example of a one-to-one relationship.
- A library database could have table of patrons and a table of books. This is a one-to-many relationship. Each patron could borrow many books, though each book only lent to one patron at a given time.
- A university database may have table consisting of courses and a table consisting of students. This is a many-to-many relationship. Each course contains many students and each student takes many courses.
Each entity in a relational database is typically represented by a key. For now, we will only focus on the `primary key`, a unique indicator of each entry in a table. Primary keys are the way that you can navigate relationships between tables.
Let's start by creating a table. In SQL you can create a table using the CREATE TABLE command. We will create a table for e-readers called *readers* which will contain the following information:
- id (integer, primary key)
- date_added
- model (unique)
- description
- quantity
- price
### SQLite data types
The following command executes the order to create this table. Note that the command to create table column must also specify the data type. The following are some SQL data types for your reference with their python equivalent in brackets:
- integer (integer ... aka complete numbers)
- real (float ... aka decimal value)
- text (string)
- blob (binary number)
- null (an empty value)
```
# this is the SQL query to create the readers table
c.execute('''CREATE TABLE readers
(id integer primary key, date_added text, model text unique, description text, quantity integer, price real)''')
```
Finally, after executing the table we must *commit* the change to the database and *close* our connection. The following code accomplishes this.
```
conn.commit() # commits the query to the database
conn.close # closes the connection to the database
```
The data will be saved for subsequent sessions.
## *Challenge Question 1 (2 points)*
Modify the code below to create a table for *books* which contains the information below. We will return to this later.
- id (integer, primary key)
- date_added (text)
- author (text)
- title (text)
- publisher (text)
- ISBN (text, unique)
- price (real)
```
# insert your code here
```
# Objective 2: Conduct simple SQL queries
Once we have some tables up and running we can start to add values to the tables. Similarly to other SQL databases, we do this by using the `INSERT INTO` command. Adding values to an SQLite database is a matter of using this command as well as the corresponding values.
The following command will `INSERT INTO` the database information about the [Kobo Forma](https://ca.kobobooks.com/products/kobo-forma?store=ca-en&utm_source=Kobo&utm_medium=TopNav&utm_campaign=Forma). Execute the cell below to add the command to the cursor.
```
conn = sqlite3.connect('data/kobo.db') # connect to the DB
c = conn.cursor() # create the cursor
c.execute("INSERT INTO readers VALUES (1, '2020-02-22', 'Forma', 'To make the reading experience better for ravenous booklovers who read for hours on end, and want a lightweight, portable alternative to heavy print books, we’ve delivered our most comfortable eReader yet with waterproof reliability, the choice of landscape or portrait mode, and the expanded access of book borrowing. Storage size available in 8GB and 32GB.',10000, 299.99)")
```
The `INSERT INTO` command will add an entry to the table as long as the entry exactly matches the requirements of the table columns. However, the way that we inserted the value is not ideal. Typically it is not advisable to specify the primary key value for an entry. Primary keys are designed to auto increment.
Let's delete the value that we just created by using the `DELETE` command. This command will remove a value from a table depending on whether that value meets the specified condition. The simplest way to delete the value is to tell SQL to delete all values from readers where id = 1.
```
c.execute('''DELETE FROM readers WHERE id=1''')
```
To properly add a value to the readers table we would typically specify the values that we wish to add using the INSERT INTO command. By doing this there is less room for error.
```
c.execute('''INSERT INTO readers(date_added, model, description, quantity, price)
VALUES ('2020-02-22', 'Forma', 'To make the reading experience better for ravenous booklovers who read for hours on end, and want a lightweight, portable alternative to heavy print books, we’ve delivered our most comfortable eReader yet with waterproof reliability, the choice of landscape or portrait mode, and the expanded access of book borrowing. Storage size available in 8GB and 32GB.',10000, 299.99)''')
```
So far so good, however manually specifying an INSERT INTO command can become burdensome. In SQL it is often important to enter many entries into the database. In SQLite and Python it is a matter of creating a list of queries. For example, we could add information about Kobo's other reading products by creating one such list.
```
readers_to_add = [
('2020-02-22', 'Libra H20', 'The perfect balance between innovative digital technology, a comfortable reading experience, and modern design is here. Offered in black or white, and with four colourful accessories to pair with, Kobo Libra H2O lets you choose the model that best suits you. Read longer than ever before with its comfortable ergonomic design, the option for landscape orientation, and easy-to-use page-turn buttons. With ComfortLight PRO and full waterproofing, Kobo Libra H2O lets you enjoy your books day or night, in any environment. Kobo Libra H2O is designed for a better reading life.',10000, 199.99),
('2020-02-22', 'Clara HD', 'Kobo Clara HD is the perfect reading companion for any booklover. It always provides the best light to read by with ComfortLight PRO, and a natural, print-like reading experience on its superior 6” HD screen. Easily customizable features help customers new to eReading to read the way they prefer. With 8 GB of on-board memory and the ability to carry up to 6,000 eBooks, Kobo Clara HD always has room for your next escape',10000, 139.99)
]
```
We then want to use the sqlite3 cursor's `executemany` method to execute multiple queries. The following code achieves this.
```
c.executemany('INSERT INTO readers(date_added, model, description, quantity, price) VALUES (?,?,?,?,?)', readers_to_add)
conn.commit() # commits the query to the database
```
## Selecting data
When working as a data scientist or analyst, perhaps the most important SQL skill to have is to query the database. In SQL, queries are typically executed using the `SELECT` command. The command `SELECT * FROM readers`, for instance will retrieve all of the entries from the `readers` table. Try executing the cell below.
```
c.execute("SELECT * FROM readers")
print(c.fetchone())
```
**Gotchya!** This code indeed retrieves all of the entries, but only prints one of them. This is because I used the `fetchone()` method to print only a single entry from the database. This is a surprisingly helpful function in most circumstances because you will not want to print all of the contents of a hundred-thousand entry database!
When the cursor retrieves query entries it saves them in a list behind the scene, similarly to the `csv` library previously explored. If we wanted to print multiple entries we would loop through them using a `for` loop, just like in previous weeks. Try executing the cell below to retrieve all of the readers entered so far.
```
for row in c.execute("SELECT * FROM readers"):
print(row)
```
In addition to executing entire entries, SQL can be used to select only specific columns. To do this, you would replace the `*` with the fields that you desire to retrieve. The following code retrieves the `id` and `model` from the `readers` table.
```
for row in c.execute("SELECT id, model FROM readers"):
print(row)
```
SQL does not order itself the same way as a CSV spreadsheet, so often you need to specify the order that you desire to retrieve the information in. You can use the ORDER BY command to achieve this.
```
for row in c.execute("SELECT id, model FROM readers ORDER BY id"):
print(row)
```
## *Challenge Question 2 (1 point)*
Write a script that [inserts information about this book](https://www.kobo.com/ca/en/ebook/pride-and-prejudice-32) into your database. Consider today to be the `date_added`; you can retrieve the rest of the necessary data from the web page using the link provided.
```
# insert your code here
```
## *Challenge Question 3 (1 point)*
Print a line that retrieves the `author` and `title` of the book that you just entered. Refer back to the examples for more information on how to do this.
```
# insert your code here
```
## *Challenge Question 4 (1 point)*
Using a list, add two more books to add to this table. They can be any books from the Kobo website.
```
# insert your code here
c.executemany('INSERT INTO books VALUES (?,?,?,?,?,?,?)', books_to_add)
conn.commit() # commits the query to the database
```
# Objective 3: Create relations between tables
As mentioned earlier, perhaps the most powerful feature of relational databases are the relationships that tables have to one another. So far, we have not specified relations between the `readers` and `books` tables. We do not need to because these two entities do not interact in meaningful way.
Users with Kobo accounts will contain information about the e-readers that they own. If we wanted to create a table for users, we would probably create a table that looks something like the following.
```
c.execute('''CREATE TABLE users(id integer primary key, date_joined text, email text)''')
```
However, this does not contain any information about other tables! What we need to relate to other tables is a `FOREIGN KEY`, a value from another table. Foreign keys are usually primary keys from another table that can be used to link two tables together. They need to be specified when creating a table that relates to another.
Let's drop that bad table before proceeding.
```
c.execute('''DROP TABLE users''')
```
To specify a foreign key in SQLite, you must first specify the value in your table and then declare it as a foreign key. The code below creates an integer called userreader, and then declares it to be a foreign key and `REFERENCES` it to the id column in the `readers` table.
```
c.execute('''CREATE TABLE users(
id integer primary key,
date_joined text,
email text,
userreader integer,
FOREIGN KEY(userreader) REFERENCES readers(id))''')
```
We can now enter an entry into the users table which identifies the users' primary e-reader. The following line creates an a user entry and connects that user to the e-reader with the ID 2 (aka Libra H20).
```
c.execute('''INSERT INTO users VALUES (1, '2020-02-22','colin.conrad@dal.ca', 2)''')
```
We can also query the user table to check our sanity. Let's do that before proceeding.
```
c.execute('''SELECT * FROM users''')
for row in c:
print(row)
```
## *Challenge Question 5 (1 point)*:
Take a momemnt to generate some more data. Create three more users and add them to the database. At least one of the users should have the Clara HD reader assigned to them. We will return to this later.
```
# insert your code here
```
# Objective 4: Conduct a complex SQL query
Great! It's now time to move on to something slightly more complex. Let's start by adding a few more users. I am sure that these email addresses may be familiar to some of you!
```
users_to_add = [
('2010-01-26','harry@hogwarts.co.uk', 1),
('2010-01-26','hermione@hogwarts.co.uk', 3),
('2010-01-26','ron@hogwarts.co.uk', 2),
('2010-01-26','ginny@hogwarts.co.uk', 2),
('2010-01-26','severus@hogwarts.co.uk', 2),
('2010-01-26','dumbledore@hogwarts.co.uk', 2),
('2010-01-26','luna@hogwarts.co.uk', 3)
]
c.executemany('INSERT INTO users(date_joined, email, userreader ) VALUES (?,?,?)', users_to_add)
c.execute('''select * from users''')
for r in c:
print(r)
```
We are now ready to see relational tables in action. So far we have only executed queries from single table. Now that we have a table with a foreign key we can create a JOIN query. These types of queries draw data from multiple tables.
Let's create a query that shows us the `date_joined`, `email` and the name of the `model` that they own. The query below achieves this.
```
c.execute('''SELECT users.date_joined, users.email, readers.model
FROM users
JOIN readers
ON users.userreader = readers.id''')
for r in c: # print the query results
print(r)
```
_Magic right?_ The beauty of relational databases is that you do not have to duplicate data because you are can retrieve the necessary data from other tables. This is extremely helpful when managing larger databases.
Let's unpack this query a bit. The query contained:
- `SELECT` which specified the data that you wished to retrieve and the table the data belongs to
- `FROM` specifies the main table (we could have chosen the `readers` table here as well in this case)
- `JOIN` specifies the table that you wish to match
- `ON` specifies the relation between the two tables, in this case they are linked by the `userreader` foreign key
Take a while to study this before proceeding. Consider trying different `JOIN` queries.
### Constraining your query with `WHERE`
There is one more thing that we should discuss before wrapping up this objective. Nearly every `SELECT` query can be limited by specifying a `WHERE` condition, which helps you limit the amount of data retrieved. For instance, we might wish to query only those users who own the Clara HD model of reader. To do this we would change the query to something like the following:
```
c.execute('''SELECT users.email, readers.model
FROM users
JOIN readers
ON users.userreader = readers.id
WHERE readers.id=3''') # you could also specify readers.model="Clara HD" in this instance
for r in c:
print(r)
```
`SELECT` queries constrained by `JOIN` and `WHERE` are among the most common type of queries used by business analysts in industry. Using these queries, you can retrieve desired data and generate reports for analysis.
## *Challenge Question 6 (2 points)*:
Create a query which retrieves the following data:
- The users' email address
- The model that they purchased
- The price of the model that they purchased
- Only select users who have purchased the Libra H20 model
Be sure to print your results for easy grading!
```
# insert your code here
```
# Objective 5: Create and query a relational table
The final thing worth mentioning is that SQL databases do not always manage so-called "one-to-many" relationships, such as the relationship between users and e-readers. These types of relationships can be managed with a foreign key and they are much simpler.
Often however, you will be faced with a "many-to-many" relationship. In our Kobo example, we could envision a scenario where each owns many books, but each book is owned by many people. A foreign key alone will not help us here. To illustrate this issue, let's start by adding some more books. Execute the cell below.
```
books_to_add = [
('2020-02-22','J. K. Rowling','Harry Potter and the Philosophers Stone', 'Pottermore Publishing','9781781100219', 10.99),
('2020-02-22','J. K. Rowling','Harry Potter and the Chamber of Secrets', 'Pottermore Publishing','9781781100226', 10.99),
('2020-02-22','J. K. Rowling','Harry Potter and the Prisoner of Azkaban', 'Pottermore Publishing','9781781100233', 10.99),
('2020-02-22','J. K. Rowling','Harry Potter and the Goblet of Fire', 'Pottermore Publishing','9781781105672', 10.99),
('2020-02-22','J. K. Rowling','Harry Potter and the Order of the Phoenix', 'Pottermore Publishing','9781781100240', 10.99),
('2020-02-22','J. K. Rowling','Harry Potter and the Half-Blood Prince', 'Pottermore Publishing','9781781100257', 10.99),
('2020-02-22','J. K. Rowling','Harry Potter and the Deathly Hallows', 'Pottermore Publishing','9781781100264', 10.99)
]
c.executemany('INSERT INTO books(date_added, author, title, publisher, isbn, price ) VALUES (?,?,?,?,?,?)', books_to_add)
c.execute('''select * from books''')
for r in c:
print(r)
```
Many-to-many relationships cannot be expressed with two tables. When faced with these sort of situations, you must create an intemediary table that contains records of the relationships between the entities. Let's create a new table called `userbooks`. This table can consist of a series of `user.id` and `book.id` pairings held as foreign keys. We can keep this one simple because its entire purpose is to hold those relationships.
```
# creates the intermediary table
c.execute('''CREATE TABLE userbooks(
userid integer,
bookid integer,
FOREIGN KEY(userid) REFERENCES users(id),
FOREIGN KEY(bookid) REFERENCES books(id))''')
```
Just like any other table, we can `INSERT INTO` the userbooks table values which correspond to the users' books. The following code should assign a relationship between user 1 (likely `colin.conrad@dal.ca`) and book 4, likely to be Harry Potter, if implemented correctly.
```
c.execute('''INSERT INTO userbooks VALUES (1, 4)''') # Colin owns Harry Potter and the Philosophers Stone
```
Similarly you can express the books which you would like to add using a list.
```
userbooks_to_add = [
('1', '5'), # Colin owns Harry Potter and the Chamber of Secrets
('2', '4'), # Justin Trudeau owns Harry Potter and the Philosophers Stone
('2', '5'), # ... etc
('2', '6'),
('2', '7'),
('2', '8'),
('3', '4'),
('3', '5')
]
c.executemany('INSERT INTO userbooks(userid, bookid ) VALUES (?,?)', userbooks_to_add)
c.execute('''select * from userbooks''')
for r in c:
print(r)
```
Finally, we can retrieve records from a many-to-many relationship by `SELECTING` from the intermediary table and `JOIN`ing on the other two tables. The code below should retrieve records for `colin.conrad@dal.ca`.
```
c.execute('''SELECT users.email, books.title
FROM userbooks
JOIN users
ON users.id = userbooks.userid
JOIN books
ON books.id = userbooks.bookid
WHERE users.id=1''') # you could also specify readers.model="Clara HD" in this instance
for r in c:
print(r)
```
## *Challenge Question 7 (2 points)*
Let's try another complex query. Select the `user.email` and `books.title` for all users who own the Forma reader. Execute the cell below before conducting this query. **Hint:** the Forma reader's ID is 1.
### Execute this cell
```
userbooks_to_add = [
('4', '1'),
('4', '2'),
('4', '3')
]
c.executemany('INSERT INTO userbooks(userid, bookid ) VALUES (?,?)', userbooks_to_add)
```
### Enter your code here
```
# insert your code here
```
## Other Stuff - Connecting to MySQL (not just SQLite)
Finally, a final note on connecting to MySQL databases or other related tools. You can use Python to connect to MySQL environments easily using similar skills to what you have explored here. Python provides a [MySQL connector](https://www.w3schools.com/python/python_mysql_getstarted.asp) to establish a connection to a remote MySQL server. For example, the call below could be used to connect to establish a connection to a remove SQL server.
I will provide a video demonstration of a connection to one of our research servers. You could similarly use this code to connect to your own MySQL server if you have one.
```
# connect to a MySQL db by entering a host name. Be sure to install the MySQL connector before attempting it
import mysql.connector
# establish the connection
mydb = mysql.connector.connect(
host="qsslab.mysql.database.azure.com",
user="conradc@qsslab.mysql.database.azure.com",
password="Zt9!L#lG2fsM",
database="userdb_conradc"
)
print(mydb)
```
With the remote connection established, you could then execute commands as you normally would. The code below was used to select all of the `tweets` from Colin's `tweets` table.
```
mycursor = mydb.cursor()
mycursor.execute("SELECT * FROM tweets")
myresult = mycursor.fetchall()
for x in myresult:
print(x)
```
## References
Codd, E. F. (1970). A Relational Model of Data for Large Shared Data Banks. *Communications of the ACM. 13*(6), 377–387. doi:10.1145/362384.362685.
W3SChools (2021). Python MySQL. Retrieved from: https://www.w3schools.com/python/python_mysql_getstarted.asp
Khan Academy (2019). Intro to SQL: Querying and managing data. Retrieved from: https://www.khanacademy.org/computing/computer-programming/sql
| github_jupyter |
<a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/hbayes_binom_rats_pymc3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
We fit a hierarchical beta-binomial model to some count data derived from rat survival. (In the book, we motivate this in terms of covid incidence rates.)
Based on https://docs.pymc.io/notebooks/GLM-hierarchical-binominal-model.html
```
import sklearn
import scipy.stats as stats
import scipy.optimize
import matplotlib.pyplot as plt
import seaborn as sns
import time
import numpy as np
import os
import pandas as pd
#!pip install pymc3 # colab uses 3.7 by default (as of April 2021)
# arviz needs 3.8+
#!pip install pymc3>=3.8 # fails to update
!pip install pymc3==3.11
import pymc3 as pm
print(pm.__version__)
import arviz as az
print(az.__version__)
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
import pandas as pd
#import seaborn as sns
import pymc3 as pm
import arviz as az
import theano.tensor as tt
np.random.seed(123)
# rat data (BDA3, p. 102)
y = np.array([
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1,
1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 1, 5, 2,
5, 3, 2, 7, 7, 3, 3, 2, 9, 10, 4, 4, 4, 4, 4, 4, 4,
10, 4, 4, 4, 5, 11, 12, 5, 5, 6, 5, 6, 6, 6, 6, 16, 15,
15, 9, 4
])
n = np.array([
20, 20, 20, 20, 20, 20, 20, 19, 19, 19, 19, 18, 18, 17, 20, 20, 20,
20, 19, 19, 18, 18, 25, 24, 23, 20, 20, 20, 20, 20, 20, 10, 49, 19,
46, 27, 17, 49, 47, 20, 20, 13, 48, 50, 20, 20, 20, 20, 20, 20, 20,
48, 19, 19, 19, 22, 46, 49, 20, 20, 23, 19, 22, 20, 20, 20, 52, 46,
47, 24, 14
])
N = len(n)
def logp_ab(value):
''' prior density'''
return tt.log(tt.pow(tt.sum(value), -5/2))
with pm.Model() as model:
# Uninformative prior for alpha and beta
ab = pm.HalfFlat('ab',
shape=2,
testval=np.asarray([1., 1.]))
pm.Potential('p(a, b)', logp_ab(ab))
alpha = pm.Deterministic('alpha', ab[0])
beta = pm.Deterministic('beta', ab[1])
X = pm.Deterministic('X', tt.log(ab[0]/ab[1]))
Z = pm.Deterministic('Z', tt.log(tt.sum(ab)))
theta = pm.Beta('theta', alpha=ab[0], beta=ab[1], shape=N)
p = pm.Binomial('y', p=theta, observed=y, n=n)
#trace = pm.sample(1000, tune=2000, target_accept=0.95)
trace = pm.sample(1000, tune=500)
#az.plot_trace(trace)
#plt.savefig('../figures/hbayes_binom_rats_trace.png', dpi=300)
print(az.summary(trace))
J = len(n)
post_mean = np.zeros(J)
samples = trace[theta]
post_mean = np.mean(samples, axis=0)
print('post mean')
print(post_mean)
alphas = trace['alpha']
betas = trace['beta']
alpha_mean = np.mean(alphas)
beta_mean = np.mean(betas)
hyper_mean = alpha_mean/(alpha_mean + beta_mean)
print('hyper mean')
print(hyper_mean)
mle = y / n
pooled_mle = np.sum(y) / np.sum(n)
print('pooled mle')
print(pooled_mle)
#axes = az.plot_forest(
# trace, var_names='theta', credible_interval=0.95, combined=True, colors='cycle')
axes = az.plot_forest(
trace, var_names='theta', hdi_prob=0.95, combined=True, colors='cycle')
y_lims = axes[0].get_ylim()
axes[0].vlines(hyper_mean, *y_lims)
#plt.savefig('../figures/hbayes_binom_rats_forest95.pdf', dpi=300)
J = len(n)
fig, axs = plt.subplots(4,1, figsize=(10,10))
plt.subplots_adjust(hspace=0.3)
axs = np.reshape(axs, 4)
xs = np.arange(J)
ax = axs[0]
ax.bar(xs, y)
ax.set_title('number of postives')
ax = axs[1]
ax.bar(xs, n)
ax.set_title('popn size')
ax = axs[2]
ax.bar(xs, mle)
ax.set_ylim(0, 0.5)
ax.hlines(pooled_mle, 0, J, 'r', lw=3)
ax.set_title('MLE (red line = pooled)')
ax = axs[3]
ax.bar(xs, post_mean)
ax.hlines(hyper_mean, 0, J, 'r', lw=3)
ax.set_ylim(0, 0.5)
ax.set_title('posterior mean (red line = hparam)')
#plt.savefig('../figures/hbayes_binom_rats_barplot.pdf', dpi=300)
J = len(n)
xs = np.arange(J)
fig, ax = plt.subplots(1,1)
ax.bar(xs, y)
ax.set_title('number of postives')
#plt.savefig('../figures/hbayes_binom_rats_outcomes.pdf', dpi=300)
fig, ax = plt.subplots(1,1)
ax.bar(xs, n)
ax.set_title('popn size')
#plt.savefig('../figures/hbayes_binom_rats_popsize.pdf', dpi=300)
fig, ax = plt.subplots(1,1)
ax.bar(xs, mle)
ax.set_ylim(0, 0.5)
ax.hlines(pooled_mle, 0, J, 'r', lw=3)
ax.set_title('MLE (red line = pooled)')
#plt.savefig('../figures/hbayes_binom_rats_MLE.pdf', dpi=300)
fig, ax = plt.subplots(1,1)
ax.bar(xs, post_mean)
ax.hlines(hyper_mean, 0, J, 'r', lw=3)
ax.set_ylim(0, 0.5)
ax.set_title('posterior mean (red line = hparam)')
#plt.savefig('../figures/hbayes_binom_rats_postmean.pdf', dpi=300)
```
| github_jupyter |
# Basics and Package Structure
If you're just interested in pulling data, you will primarily be using `nba_api.stats.endpoints`.
This submodule contains a class for each API endpoint supported by stats.nba.com.
For example, [the PlayerCareerStats class](https://github.com/swar/nba_api/blob/master/nba_api/stats/endpoints/playercareerstats.py) is initialized with a player ID and returns some career statistics for the player.
```
from nba_api.stats.endpoints import playercareerstats
# Anthony Davis
career = playercareerstats.PlayerCareerStats(player_id='203076')
career.get_data_frames()[0]
```
`career`, above, is a `PlayerCareerStats` object.
This class (and the other endpoint classes) supports several methods of accessing the data: `get_dict()`, `get_json()`, `get_data_frames()`, and more.
`get_data_frames()` returns a list of pandas DataFrames, and when working in notebooks, this is often your best option for viewing data.
In general, the first DataFrame in this list is the primary returned data structure and the one you'll want to look at.
Almost all of the endpoint classes take at least one required argument, along with several optional ones.
In the case of `PlayerCareerStats`, a player ID is required, but the user may also specify a league ID.
At the time of writing this notebook, these are the endpoints available:
<table><tr></tr><tr><td>boxscoreadvancedv2</td><td>boxscorefourfactorsv2</td><td>boxscoremiscv2</td><td>boxscoreplayertrackv2</td></tr><tr><td>boxscorescoringv2</td><td>boxscoresummaryv2</td><td>boxscoretraditionalv2</td><td>boxscoreusagev2</td></tr><tr><td>commonallplayers</td><td>commonplayerinfo</td><td>commonplayoffseries</td><td>commonteamroster</td></tr><tr><td>commonteamyears</td><td>defensehub</td><td>draftcombinedrillresults</td><td>draftcombinenonstationaryshooting</td></tr><tr><td>draftcombineplayeranthro</td><td>draftcombinespotshooting</td><td>draftcombinestats</td><td>drafthistory</td></tr><tr><td>franchisehistory</td><td>homepageleaders</td><td>homepagev2</td><td>infographicfanduelplayer</td></tr><tr><td>leaderstiles</td><td>leaguedashlineups</td><td>leaguedashplayerbiostats</td><td>leaguedashplayerclutch</td></tr><tr><td>leaguedashplayerptshot</td><td>leaguedashplayershotlocations</td><td>leaguedashplayerstats</td><td>leaguedashptdefend</td></tr><tr><td>leaguedashptstats</td><td>leaguedashptteamdefend</td><td>leaguedashteamclutch</td><td>leaguedashteamptshot</td></tr><tr><td>leaguedashteamshotlocations</td><td>leaguedashteamstats</td><td>leaguegamefinder</td><td>leaguegamelog</td></tr><tr><td>leagueleaders</td><td>leaguestandings</td><td>playbyplay</td><td>playbyplayv2</td></tr><tr><td>playerawards</td><td>playercareerstats</td><td>playercompare</td><td>playerdashboardbyclutch</td></tr><tr><td>playerdashboardbygamesplits</td><td>playerdashboardbygeneralsplits</td><td>playerdashboardbylastngames</td><td>playerdashboardbyopponent</td></tr><tr><td>playerdashboardbyshootingsplits</td><td>playerdashboardbyteamperformance</td><td>playerdashboardbyyearoveryear</td><td>playerdashptpass</td></tr><tr><td>playerdashptreb</td><td>playerdashptshotdefend</td><td>playerdashptshots</td><td>playerfantasyprofile</td></tr><tr><td>playerfantasyprofilebargraph</td><td>playergamelog</td><td>playergamestreakfinder</td><td>playernextngames</td></tr><tr><td>playerprofilev2</td><td>playersvsplayers</td><td>playervsplayer</td><td>playoffpicture</td></tr><tr><td>scoreboard</td><td>scoreboardv2</td><td>shotchartdetail</td><td>shotchartlineupdetail</td></tr><tr><td>teamdashboardbyclutch</td><td>teamdashboardbygamesplits</td><td>teamdashboardbygeneralsplits</td><td>teamdashboardbylastngames</td></tr><tr><td>teamdashboardbyopponent</td><td>teamdashboardbyshootingsplits</td><td>teamdashboardbyteamperformance</td><td>teamdashboardbyyearoveryear</td></tr><tr><td>teamdashlineups</td><td>teamdashptpass</td><td>teamdashptreb</td><td>teamdashptshots</td></tr><tr><td>teamdetails</td><td>teamgamelog</td><td>teamgamestreakfinder</td><td>teamhistoricalleaders</td></tr><tr><td>teaminfocommon</td><td>teamplayerdashboard</td><td>teamplayeronoffdetails</td><td>teamplayeronoffsummary</td></tr><tr><td>teamvsplayer</td><td>teamyearbyyearstats</td><td>videodetails</td><td>videoevents</td></tr><tr><td>videostatus</td></tr></table>
### Getting Team and Player IDs
The package also includes utilities for fetching player and team information available under `nba_api.stats.static`.
You can use this to fetch player IDs and team IDs, which are often used as inputs to API endpoints.
```
from nba_api.stats.static import teams
# get_teams returns a list of 30 dictionaries, each an NBA team.
nba_teams = teams.get_teams()
print('Number of teams fetched: {}'.format(len(nba_teams)))
nba_teams[:3]
from nba_api.stats.static import players
# get_players returns a list of dictionaries, each representing a player.
nba_players = players.get_players()
print('Number of players fetched: {}'.format(len(nba_players)))
nba_players[:5]
```
To search for an individual team or player by its name (or other attribute), dictionary comprehensions are your friend.
```
spurs = [team for team in nba_teams
if team['full_name'] == 'San Antonio Spurs'][0]
spurs
big_fundamental = [player for player in nba_players
if player['full_name'] == 'Tim Duncan'][0]
big_fundamental
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import rcParams
rcParams['legend.fontsize'] = 14
rcParams['axes.labelsize'] = 14
rcParams['axes.titlesize'] = 14
%matplotlib inline
```
# Tutorial apply IMU erreur model
## Creating a trajectory and generating inertial readings
First we need to generate a trajectory.
```
from pyins import sim
dt = 0.5
t = 3 * 3600
n_samples = int(t / dt)
lat = np.full(n_samples, 50.0)
lon = np.full(n_samples, 60.0)
alt = np.zeros_like(lat)
h = np.full(n_samples, 10.0)
r = np.full(n_samples, -5.0)
p = np.full(n_samples, 3.0)
traj, gyro_ref, accel_ref = sim.from_position(dt, lat, lon, alt, h, p, r)
step_plot = int(60/dt) # display of one point per min.
```
## IMU model
### gyro model
```
gyro_scale_error = None
gyro_scale_asym = None
gyro_align = None
gyro_bias = np.array([1e-8, -2e-8, 3e-8])
gyro_noise = np.array([1e-6, 1e-6, 1e-6])
```
### accel model
```
accel_scale_error = None
accel_scale_asym = None
accel_align = None
accel_bias = np.array([3e-3, -4e-3, 2e-3])
accel_noise = np.array([3e-4, 3e-4 , 3e-4])
imu = sim.ImuErrors(gyro_scale_error, gyro_scale_asym, gyro_align, gyro_bias, gyro_noise,
accel_scale_error, accel_scale_asym, accel_align, accel_bias, accel_noise)
```
Application of the error model to the reference increment
```
gyro, accel = imu.apply(dt, gyro_ref, accel_ref)
```
## Initial condition errors
```
d_lat = 100
d_lon = -200
d_VE = 1
d_VN = -2
d_h = 0.01
d_p = -0.02
d_r = 0.03
```
Adding initial errors to the trajectory
```
from pyins.coord import perturb_ll
lat0, lon0 = perturb_ll(traj.lat[0], traj.lon[0], d_lat, d_lon)
VE0 = traj.VE[0] + d_VE
VN0 = traj.VN[0] + d_VN
h0 = traj.h[0] + d_h
p0 = traj.p[0] + d_p
r0 = traj.r[0] + d_r
```
## Integration of increments
```
from pyins.integrate import coning_sculling, integrate
theta, dv = coning_sculling(gyro, accel)
traj_c = integrate(dt, lat0, lon0, VE0, VN0, h0, p0, r0, theta, dv)
```
The reference trajectory and the trajectory resulting from integration are drawn below, the initial point is marked with a cross.
```
plt.plot(traj.lon, traj.lat)
plt.plot(traj.lon[0], traj.lat[0], 'kx', markersize=12)
plt.plot(traj_c.lon, traj_c.lat)
plt.xlabel("lon, deg")
plt.ylabel("lat, deg")
```
## Computing Integration Errors
```
from pyins.filt import traj_diff
error_true = traj_diff(traj_c, traj)
from helper import plot_traj
plot_traj(dt, error_true, step_plot, coord_unit="m")
```
## Propagation of IMU and initial errors
Calculation of increment errors, Gyro and accelerometer errors (in SI units) can be constant or specified for each time stamp in `traj`.
```
e_g, e_a = gyro - gyro_ref, accel - accel_ref
e_g = np.vstack((np.zeros(3), e_g))
e_a = np.vstack((np.zeros(3), e_a))
from pyins.filt import propagate_errors
error_linear = propagate_errors(dt, traj, d_lat, d_lon, d_VE, d_VN, d_h,
d_p, d_r, e_g, e_a)
plot_traj(dt, error_linear, step_plot, coord_unit="m")
```
## Comparison of integration and propagation method
```
error_scale = np.mean(np.abs(error_true))
rel_diff = (error_true - error_linear) / error_scale
rel_diff.abs().max()
```
| github_jupyter |
# Think Bayes
This notebook presents example code and exercise solutions for Think Bayes.
Copyright 2018 Allen B. Downey
MIT License: https://opensource.org/licenses/MIT
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import classes from thinkbayes2
from thinkbayes2 import Hist, Pmf, Suite, Beta
import thinkplot
import numpy as np
```
## The dinner party
Suppose you are having a dinner party with 10 guests and 4 of them are allergic to cats. Because you have cats, you expect 50% of the allergic guests to sneeze during dinner. At the same time, you expect 10% of the non-allergic guests to sneeze. What is the distribution of the total number of guests who sneeze?
```
# Solution
n_allergic = 4
n_non = 6
p_allergic = 0.5
p_non = 0.1
pmf = MakeBinomialPmf(n_allergic, p_allergic) + MakeBinomialPmf(n_non, p_non)
thinkplot.Hist(pmf)
# Solution
pmf.Mean()
```
## The Gluten Problem
[This study from 2015](http://onlinelibrary.wiley.com/doi/10.1111/apt.13372/full) showed that many subjects diagnosed with non-celiac gluten sensitivity (NCGS) were not able to distinguish gluten flour from non-gluten flour in a blind challenge.
Here is a description of the study:
>"We studied 35 non-CD subjects (31 females) that were on a gluten-free diet (GFD), in a double-blind challenge study. Participants were randomised to receive either gluten-containing flour or gluten-free flour for 10 days, followed by a 2-week washout period and were then crossed over. The main outcome measure was their ability to identify which flour contained gluten.
>"The gluten-containing flour was correctly identified by 12 participants (34%)..."
Since 12 out of 35 participants were able to identify the gluten flour, the authors conclude "Double-blind gluten challenge induces symptom recurrence in just one-third of patients fulfilling the clinical diagnostic criteria for non-coeliac gluten sensitivity."
This conclusion seems odd to me, because if none of the patients were sensitive to gluten, we would expect some of them to identify the gluten flour by chance. So the results are consistent with the hypothesis that none of the subjects are actually gluten sensitive.
We can use a Bayesian approach to interpret the results more precisely. But first we have to make some modeling decisions.
1. Of the 35 subjects, 12 identified the gluten flour based on resumption of symptoms while they were eating it. Another 17 subjects wrongly identified the gluten-free flour based on their symptoms, and 6 subjects were unable to distinguish. So each subject gave one of three responses. To keep things simple I follow the authors of the study and lump together the second two groups; that is, I consider two groups: those who identified the gluten flour and those who did not.
2. I assume (1) people who are actually gluten sensitive have a 95% chance of correctly identifying gluten flour under the challenge conditions, and (2) subjects who are not gluten sensitive have only a 40% chance of identifying the gluten flour by chance (and a 60% chance of either choosing the other flour or failing to distinguish).
Using this model, estimate the number of study participants who are sensitive to gluten. What is the most likely number? What is the 95% credible interval?
```
# Solution
# Here's a class that models the study
class Gluten(Suite):
def Likelihood(self, data, hypo):
"""Computes the probability of the data under the hypothesis.
data: tuple of (number who identified, number who did not)
hypothesis: number of participants who are gluten sensitive
"""
# compute the number who are gluten sensitive, `gs`, and
# the number who are not, `ngs`
gs = hypo
yes, no = data
n = yes + no
ngs = n - gs
pmf1 = MakeBinomialPmf(gs, 0.95)
pmf2 = MakeBinomialPmf(ngs, 0.4)
pmf = pmf1 + pmf2
return pmf[yes]
# Solution
prior = Gluten(range(0, 35+1))
thinkplot.Pdf(prior)
# Solution
posterior = prior.Copy()
data = 12, 23
posterior.Update(data)
# Solution
thinkplot.Pdf(posterior)
thinkplot.Config(xlabel='# who are gluten sensitive',
ylabel='PMF', legend=False)
# Solution
posterior.CredibleInterval(95)
```
| github_jupyter |
NASA MARS NEWS
```
#import dependencies
from bs4 import BeautifulSoup as bs
from splinter import Browser
import pandas as pd
import time
import os
import requests
from selenium.webdriver.chrome.options import Options
from splinter.exceptions import ElementDoesNotExist
#pointing to the directory where chromedriver exists
executable_path = {"executable_path": "users/anali/bin/chromedriver"}
browser = Browser("chrome", **executable_path, headless = False)
#visiting the page
#url = "https://mars.nasa.gov/news/"
url = "https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest"
# launch browser
browser.visit(url)
#check if the page has been loaded
browser.is_element_present_by_name('list_date', wait_time=10)
#create HTML object
html = browser.html
#parse HTML with beautiful object
soup = bs(html,"html.parser")
#extract title and paragraph
news_date = soup.find('div', class_='list_date').text
news_title = soup.find('div', class_='content_title').text
news_p = soup.find('div', class_='article_teaser_body').text
print(f"Date: {news_date}")
print(f"Title: {news_title}")
print(f"Para: {news_p}")
```
#--------------------------------------------
# JPL Mars Space Images - Featured Image
#--------------------------------------------
```
url_image = "https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars"
browser.visit(url_image)
#Getting the base url
from urllib.parse import urlsplit
base_url = "{0.scheme}://{0.netloc}/".format(urlsplit(url_image))
print(base_url)
#Design an xpath selector to grab the image
xpath = "//*[@id=\"page\"]/section[3]/div/ul/li[1]/a/div/div[2]/img"
#Use splinter to click on the mars featured image
#to bring the full resolution image
results = browser.find_by_xpath(xpath)
img = results[0]
img.click()
#get image url using BeautifulSoup
html_image = browser.html
soup = bs(html_image, "html.parser")
img_url = soup.find("img", class_="fancybox-image")["src"]
featured_image_url = base_url + img_url
print(featured_image_url)
#RESULT: featured_image_url = https://www.jpl.nasa.gov//spaceimages/images/largesize/PIA24053_hires.jpg
```
##Mars Weather
```
# twitter url to visit
url = 'https://twitter.com/marswxreport?lang=en'
# launch browser
browser.visit(url)
# create beautifulsoup object
html = browser.html
soup = bs(html, "html.parser")
print(soup.prettify())
#find tweet and extract text
mars_weather = soup.find_all('span')
for i in range(len(mars_weather)):
if ("InSight" in mars_weather[i].text):
weather = mars_weather[i].text
print(weather)
break
#<span class="css-901oao css-16my406 r-1qd0xha r-ad9z0x r-bcqeeo r-qvutc0">
#InSight sol 597 (2020-08-01) low -91.0ºC (-131.8ºF) high -16.9ºC (1.6ºF)
#winds from the WNW at 8.0 m/s (17.9 mph) gusting to 20.2 m/s (45.1 mph)
#pressure at 7.90 hPa
#</span>
```
##Mars Facts
```
url_facts = "https://space-facts.com/mars/"
table = pd.read_html(url_facts)
table[0]
df_mars_facts = table[0]
df_mars_facts.columns = ["Parameter", "Values"]
df_mars_facts.set_index(["Parameter"])
df_mars_facts.to_html()
```
Mars Hemispheres
```
url4 = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(url4)
html4 = browser.html
soup4 = bs(html4, 'html.parser')
# First, get a list of all of the hemispheres
links = browser.find_by_css("a.product-item h3")
hemisphere_image_urls = []
# First, get a list of all of the hemispheres
links = browser.find_by_css("a.product-item h3")
# Next, loop through those links, click the link, find the sample anchor, return the href
for i in range(len(links)):
hemisphere = {}
# We have to find the elements on each loop to avoid a stale element exception
browser.find_by_css("a.product-item h3")[i].click()
# Next, we find the Sample image anchor tag and extract the href
sample_elem = browser.find_link_by_text('Sample').first
hemisphere['img_url'] = sample_elem['href']
# Get Hemisphere title
hemisphere['title'] = browser.find_by_css("h2.title").text
# Append hemisphere object to list
hemisphere_image_urls.append(hemisphere)
# Finally, we navigate backwards
browser.back()
hemisphere_image_urls
```
| github_jupyter |
# The DataFetcher
The DataFetcher class is by detex to serve seismic data to other functions and classes. It is designed to use data from local directories as well as remote clients (like the [obspy FDSN client](https://docs.obspy.org/packages/obspy.fdsn.html)). In the future I hope to add functionality to the DataFetcher to allow it to check data availability and quality. We will start by looking at the DataFetcher class docs, using the DataFetcher on local data directories, and then setting up a DataFetcher to use a remote client.
## DataFetcher docs
Let's print the current version of detex and the docstring associated with the DataFetcher class in order to get an idea of what it does and what options are available.
```
import detex
print('Current detex version is %s' % (detex.__version__))
print ('-------------------------------')
print (detex.getdata.DataFetcher.__doc__)
```
Some of the more important parameters to pay attention to are the ones controlling the duration of files and the response removal.
* Parameters that control data duration, number of files, and file type:
1. timeBeforeOrigin
2. timeAfterOrigin
3. conDatDuration
4. secBuf
* Parameters that control response removal (more on obspy response removal [here](https://docs.obspy.org/packages/autogen/obspy.core.stream.Stream.remove_response.html#obspy.core.stream.Stream.remove_response))
1. removeResponse (True or False)
2. opType ("DISP" (m), "VEL" (m/s), or "ACC" (m/s^2))
3. prefilt
Also, for less than perfect data, the fillZeros parameter can be very important to avoid discarding data with small gaps. More on this in the [clustering section](../Clustering/clustering.md).
## DataFetcher with local directories
Often it can be faster to download, preform some processing, and save data once rather than using clients each time detex needs seismic data. This is not always the case, however, if a database on the same network as your workstation is the remote client you wish to use. As an example, let's create a local data directory and then set up a DataFetcher instance to pull data from it.
### Create local directories
In order to create the data directories we first need to let Detex know which stations and events it should look for. To do this we use the template key and station key files (more on that in the [required files section](../RequiredFiles/required_files.md).
For this example lets use a subset of the template key and station key used in the intro tutorial.
```
import detex
stakey = detex.util.readKey('StationKey.csv', key_type='station')
stakey
temkey = detex.util.readKey('TemplateKey.csv', key_type='template')
temkey
```
Next we need to call makeDataDirectories (or getAllData which was kept for backward compatibility).
```
%time detex.getdata.makeDataDirectories() # make directories and time how long it takes (the %time magic only works in ipython)
```
Before we look at the downloaded data lets discuss some of the parameters that you should pay attention to when calling the makeDataDirectories function. You should notice that many of the makeDataDirectories function's input arguments are similar to DataFetchers arguments. This is because under the hood the makeDataDirectories function is simply using a DataFetcher attached to a client (IRIS by default). If you wanted to use something besides IRIS you would just need to pass a DataFetcher instance attached to another client as the fetch argument.
One unique argument that makeDataDirectories needs is the formatOut, which is the format to use when saving the data to disk. Any format obspy can read/write should be acceptable. Options are: 'mseed', 'sac', 'GSE2', 'sacxy', 'q', 'sh_asc', 'slist', 'tspair', 'segy', 'su', 'pickle', 'h5' (if obspyh5 is installed). Default is mseed, although the makeDataDirectories call by default will remove instrument response thus necessitating that the data are in a float format and therefore devaluing the mseed compression advantage.
I recommend you look at the entire doc string of the function, but I wont print it here. You should think about what parameters will work best for your data set before just using the defaults.
Now let's take a look at the newly created data directories. This is most easily accomplished by reading the SQLite database that was created to index the directories.
```
import os
index_file = os.path.join('ContinuousWaveForms', '.index.db')
ind = detex.util.loadSQLite(index_file, 'ind')
ind
```
The fields in the database table "ind" are as follows:
| Field | Description |
|:-----:| :---------: |
| Path | A list of indicies to reference values in the indkey table for building absolute paths|
| FileName | The name of the particular file represented by the current row |
| Starttime | time stamp of the start time in the file |
| Endtime | time stamp of the end time in the file |
| Gaps | The total number of gaps in the file |
| Nc | The number of unique channels |
| Nt | The number of traces (without gaps Nc = Nt) |
| Duration | Duration of seismic data in seconds |
| Station | network.station |
When the DataFetcher loads files from a directory it first reads the index to find the paths to load. Because of this, the directory structure not important. For example, if you already have a directory that contains some files in an obspy readable format you can index it with the detex.util.indexDirectory function. Once indexed the directory can be used by the DataFetcher class.
It can be useful to use the index for data quality checks. For example, let's look for files that are shorter than expected, that are missing channels, or that have gaps (even though we can see these TA data don't have any such issues).
```
# look for gaps
ind_gaps = ind[ind.Gaps > 0]
print("There are %d files with gaps" % len(ind_gaps))
# Look for durations at least 2 minutes less than the expected duration
expected_duration = 3720
ind_short = ind[3720 - ind.Duration > 120]
print("There are %d files with shorter than expected durations" % len(ind_short))
# look for missing channels
expected_channels = 3
ind_missing = ind[ind.Nc < expected_channels]
print("There are %d files with less than %d channels" % (len(ind_missing), expected_channels))
```
### Initiate DataFetcher
Now we are ready to create a DataFetcher instance and point it at the newly created directory. We will also explore some of the DataFetcher methods.
```
# Create two fetchers, one pointed at the continuous data and the other at the event data
con_fetcher = detex.getdata.DataFetcher('dir', directoryName='ContinuousWaveForms', removeResponse=False)
eve_fetcher = detex.getdata.DataFetcher('dir', directoryName='EventWaveForms', removeResponse=False)
```
As of version 1.0.4 the DataFetcher has 3 public methods:
1. getStream - fetches an a stream from an arbitrary network, station, channel, location (which the user must define). If no data are fetchable then None is returned.
2. getConData - creates a generator for fetching all data avaliable for the stations, channels, and date ranges found in a station key.
3. getTemData - fetches data related to those described by the template key, but also needs a station key to know which stations to look for.
Let's look at an example use of each method:
```
# getStream example
import obspy
## set variables
utc1 = obspy.UTCDateTime('2009-091T04-13-00') - 5
utc2 = utc1 + 60
net = 'TA'
sta = 'M17A'
chan = 'BH?'
## fetch
st = con_fetcher.getStream(utc1, utc2, net, sta, chan)
## plot the boring data
%pylab inline
st.plot()
# getConData example
## Read station key and use only TA M17A
stakey = detex.util.readKey('StationKey.csv', key_type='station')
stakey = stakey[stakey.STATION=='M17A']
## Create a generator for fetching continuous data
congen = con_fetcher.getConData(stakey) # note if we dont pass a duration the default is used
## loop over generator and calculate sta/lta values to see if we can find an event
from obspy.signal.trigger import classicSTALTA # for simplicity let's use the basic sta/lta
from obspy.signal.trigger import plotTrigger
sta = 0.5 # short term average in seconds
lta = 2 # long term average in seconds
ratio_max = 0 # int variables to keep track of max and time it occurs
time_max = 0
trace_max = None
cft_max = None
for st in congen: # iterate through the generator until it is exhausted
trace = st.select(component = 'z')[0] # select vertical component
trace.filter('bandpass', freqmin=1, freqmax=10, zerophase=True, corners=2) #filter
sr = trace.stats.sampling_rate # get sampling rate
starttime = trace.stats.starttime
cft = classicSTALTA(trace.data, int(sta * sr), int(lta * sr)) # run sta/lta
cft_max = max(cft) # get max value
if cft_max > ratio_max: # if the max is greater than old max
ratio_max = cft_max # set new max
time_max = starttime + cft.argmax()/float(sr) # set time max
trace_max = trace.copy()
cft_max = cft
print("The max sta/lta was %.2f occured at %s" % (ratio_max, time_max))
plotTrigger(trace, cft, ratio_max*.92, ratio_max/1.5)
## Let's get a closer look
st = con_fetcher.getStream(time_max-10, time_max+35, 'TA', 'M17A', 'BHZ')
st.filter('bandpass', freqmin=1, freqmax=5, zerophase=True, corners=2)
st.plot()
# getConData example
## Create stream generator
evegen = eve_fetcher.getTemData("TemplateKey.csv", "StationKey.csv")
# note: the temkey and stakey parameters can either be paths to csv files or DataFrames
## iterate through each of the known events plot a spectrogram of the one with highest amplitude
amp_max = 0
tr_max = None
for st, evename in evegen:
trace = st.select(component = 'z')[0]
trace.detrend('linear')
trace.filter('bandpass', freqmin=1, freqmax=10, zerophase=True, corners=2)
z_max = max(trace.data)
if z_max > amp_max:
amp_max = z_max
tr_max = trace.copy()
tr_max.plot()
tr_max.spectrogram()
```
## DataFetcher with clients
Detex should be able to handle a wide variety of obspy client types, including FDSN, NEIC, EARTHWORM, etc. However, as of version 1.0.4 I have only tested IRIS extensively so using other clients may take a bit of debugging. More tests and bug fixes will follow in future versions.
### IRIS FDSN client
In order to use the DataFetcher we first need to set up a client object. We will create an FDSN client then initiate an instance of the DataFetcher class and use the getStream function to fetch an obspy stream.
```
import detex
import obspy
from obspy.fdsn import Client
#setup client
client = Client("IRIS")
# setup fetcher
fetcher = detex.getdata.DataFetcher(method='client', client=client)
# set info
utc1 = obspy.UTCDateTime('2009-03-19T19-06-07') - 5
utc2 = utc1 + 60
net = 'TA'
sta = 'M17A'
chan = 'BH?'
# fetch a stream
st = fetcher.getStream(utc1, utc2, net, sta, chan)
# plot waveforms
%pylab inline
st.filter('bandpass', freqmin=1, freqmax=10, corners=2, zerophase=True)
st.plot()
```
If the waveforms look strange it is because this event is actually a blast at a surface coal mine.
The other methods demonstrated in previous sections also work with the DataFetcher attached to IRIS, so I wont illustrate them again here.
It should be noted that by default the instrument responses have been removed. This can be controlled with the removeResponse input argument which is either set to True or False.
This should give you all the information you need on how detex gets its data and how to set up a custom DataFetcher to be used by other detex classes.
# Next Section
The [next section](../Clustering/clustering.md) covers how to perform waveform similarity analysis in preparation for subspace detection.
| github_jupyter |
# Deserialisation
Consider a straightforward YAML serialisation for our model:
```
class Element:
def __init__(self, symbol):
self.symbol = symbol
class Molecule:
def __init__(self):
self.elements= {} # Map from element to number of that element in the molecule
def add_element(self, element, number):
self.elements[element] = number
def to_struct(self):
return {x.symbol: self.elements[x] for x in self.elements}
class Reaction:
def __init__(self):
self.reactants = { } # Map from reactants to stoichiometries
self.products = { } # Map from products to stoichiometries
def add_reactant(self, reactant, stoichiometry):
self.reactants[reactant] = stoichiometry
def add_product(self, product, stoichiometry):
self.products[product] = stoichiometry
def to_struct(self):
return {
'reactants' : [x.to_struct() for x in self.reactants],
'products' : [x.to_struct() for x in self.products],
'stoichiometries' : list(self.reactants.values())+
list(self.products.values())
}
class System:
def __init__(self):
self.reactions=[]
def add_reaction(self, reaction):
self.reactions.append(reaction)
def to_struct(self):
return [x.to_struct() for x in self.reactions]
c=Element("C")
o=Element("O")
h=Element("H")
co2 = Molecule()
co2.add_element(c,1)
co2.add_element(o,2)
h2o = Molecule()
h2o.add_element(h,2)
h2o.add_element(o,1)
o2 = Molecule()
o2.add_element(o,2)
h2 = Molecule()
h2.add_element(h,2)
glucose = Molecule()
glucose.add_element(c,6)
glucose.add_element(h,12)
glucose.add_element(o,6)
combustion_glucose = Reaction()
combustion_glucose.add_reactant(glucose, 1)
combustion_glucose.add_reactant(o2, 6)
combustion_glucose.add_product(co2, 6)
combustion_glucose.add_product(h2o, 6)
combustion_hydrogen = Reaction()
combustion_hydrogen.add_reactant(h2,2)
combustion_hydrogen.add_reactant(o2,1)
combustion_hydrogen.add_product(h2o,2)
s=System()
s.add_reaction(combustion_glucose)
s.add_reaction(combustion_hydrogen)
s.to_struct()
import yaml
print(yaml.dump(s.to_struct()))
```
# Deserialising non-normal data structures
We can see that this data structure, although seemingly
sensible, is horribly non-normal.
* The stoichiometries information requires us to align each one to the corresponding molecule in order.
* Each element is described multiple times: we will have to ensure that each mention of `C` comes back to the same constructed element object.
```
class DeSerialiseStructure:
def __init__(self):
self.elements = {}
self.molecules = {}
def add_element(self, candidate):
if candidate not in self.elements:
self.elements[candidate]=Element(candidate)
return self.elements[candidate]
def add_molecule(self, candidate):
if tuple(candidate.items()) not in self.molecules:
m = Molecule()
for symbol, number in candidate.items():
m.add_element(self.add_element(symbol), number)
self.molecules[tuple(candidate.items())]=m
return self.molecules[tuple(candidate.items())]
def parse_system(self, system):
s = System()
for reaction in system:
r = Reaction()
stoichiometries = reaction['stoichiometries']
for molecule in reaction['reactants']:
r.add_reactant(self.add_molecule(molecule),
stoichiometries.pop(0))
for molecule in reaction['products']:
r.add_product(self.add_molecule(molecule),
stoichiometries.pop(0))
s.add_reaction(r)
return s
de_serialiser = DeSerialiseStructure()
round_trip = de_serialiser.parse_system(s.to_struct())
round_trip.to_struct()
de_serialiser.elements
de_serialiser.molecules
list(round_trip.reactions[0].reactants.keys())[1]
list(round_trip.reactions[1].reactants.keys())[1]
```
In making this, we ended up choosing primary keys for our datatypes:
```
list(de_serialiser.molecules.keys())
```
Again, we note that a combination of columns uniquely defining an item
is a valid key - there is a key correspondence between
a candidate key in the database sense and a "hashable" data structure that can be used to a
key in a `dict`.
Note that to make this example even reasonably doable, we didn't add additional data to the objects (mass, rate etc)
# Normalising a YAML structure
To make this structure easier to de-serialise, we can make a normalised file-format, by defining primary keys (hashable types) for each entity on write:
```
class SaveSystem:
def __init__(self):
self.elements = set()
self.molecules = set()
def element_key(self, element):
return element.symbol
def molecule_key(self, molecule):
key=''
for element, number in molecule.elements.items():
key+=element.symbol
key+=str(number)
return key
def save(self, system):
for reaction in system.reactions:
for molecule in reaction.reactants:
self.molecules.add(molecule)
for element in molecule.elements:
self.elements.add(element)
for molecule in reaction.products:
self.molecules.add(molecule)
for element in molecule.elements:
self.elements.add(element)
result = {
'elements' : [self.element_key(element)
for element in self.elements],
'molecules' : {
self.molecule_key(molecule):
{self.element_key(element): number
for element, number
in molecule.elements.items()}
for molecule in self.molecules},
'reactions' : [{
'reactants' : {
self.molecule_key(reactant) : stoich
for reactant, stoich
in reaction.reactants.items()
},
'products' : {
self.molecule_key(product) : stoich
for product, stoich
in reaction.products.items()
}}
for reaction in system.reactions]
}
return result
saver = SaveSystem()
print(yaml.dump(saver.save(s)))
```
We can see that to make an easily parsed file format, without having to
guess-recognise repeated entities based on their names
(which is highly subject to data entry error), we effectively recover
the same tables as found for the database model.
An alternative is to use a simple integer for such a primary key:
```
class SaveSystemI:
def __init__(self):
self.elements = {}
self.molecules = {}
def add_element(self, element):
if element not in self.elements:
self.elements[element]=len(self.elements)
return self.elements[element]
def add_molecule(self, molecule):
if molecule not in self.molecules:
self.molecules[molecule]=len(self.molecules)
return self.molecules[molecule]
def element_key(self, element):
return self.elements[element]
def molecule_key(self, molecule):
return self.molecules[molecule]
def save(self, system):
for reaction in system.reactions:
for molecule in reaction.reactants:
self.add_molecule(molecule)
for element in molecule.elements:
self.add_element(element)
for molecule in reaction.products:
self.add_molecule(molecule)
for element in molecule.elements:
self.add_element(element)
result = {
'elements' : [element.symbol
for element in self.elements],
'molecules' : {
self.molecule_key(molecule):
{self.element_key(element): number
for element, number
in molecule.elements.items()}
for molecule in self.molecules},
'reactions' : [{
'reactants' : {
self.molecule_key(reactant) : stoich
for reactant, stoich
in reaction.reactants.items()
},
'products' : {
self.molecule_key(product) : stoich
for product, stoich
in reaction.products.items()
}}
for reaction in system.reactions]
}
return result
saver = SaveSystemI()
print(yaml.dump(saver.save(s)))
```
## Reference counting
The above approach of using a dictionary to determine the integer keys
for objects is a bit clunky.
Another good approach is to use counted objects either via a static member or by using a factory pattern:
```
class Element:
def __init__(self, symbol, id):
self.symbol = symbol
self.id = id
class Molecule:
def __init__(self, id):
self.elements= {} # Map from element to number of that element in the molecule
self.id=id
def add_element(self, element, number):
self.elements[element] = number
def to_struct(self):
return {x.symbol: self.elements[x] for x in self.elements}
class Reaction:
def __init__(self):
self.reactants = { } # Map from reactants to stoichiometries
self.products = { } # Map from products to stoichiometries
def add_reactant(self, reactant, stoichiometry):
self.reactants[reactant] = stoichiometry
def add_product(self, product, stoichiometry):
self.products[product] = stoichiometry
def to_struct(self):
return {
'reactants' : [x.to_struct() for x in self.reactants],
'products' : [x.to_struct() for x in self.products],
'stoichiometries' : list(self.reactants.values())+
list(self.products.values())
}
class System: # This will be our factory
def __init__(self):
self.reactions=[]
self.elements=[]
self.molecules=[]
def add_element(self, symbol):
new_element = Element(symbol, len(self.elements))
self.elements.append(new_element)
return new_element
def add_molecule(self):
new_molecule = Molecule(len(self.molecules))
self.molecules.append(new_molecule)
return new_molecule
def add_reaction(self):
new_reaction=Reaction()
self.reactions.append(new_reaction)
return new_reaction
def save(self):
result = {
'elements' : [element.symbol
for element in self.elements],
'molecules' : {
molecule.id:
{element.id: number
for element, number
in molecule.elements.items()}
for molecule in self.molecules},
'reactions' : [{
'reactants' : {
reactant.id : stoich
for reactant, stoich
in reaction.reactants.items()
},
'products' : {
product.id : stoich
for product, stoich
in reaction.products.items()
}}
for reaction in self.reactions]
}
return result
s2=System()
c=s2.add_element("C")
o=s2.add_element("O")
h=s2.add_element("H")
co2 = s2.add_molecule()
co2.add_element(c,1)
co2.add_element(o,2)
h2o = s2.add_molecule()
h2o.add_element(h,2)
h2o.add_element(o,1)
o2 = s2.add_molecule()
o2.add_element(o,2)
h2 = s2.add_molecule()
h2.add_element(h,2)
glucose = s2.add_molecule()
glucose.add_element(c,6)
glucose.add_element(h,12)
glucose.add_element(o,6)
combustion_glucose = s2.add_reaction()
combustion_glucose.add_reactant(glucose, 1)
combustion_glucose.add_reactant(o2, 6)
combustion_glucose.add_product(co2, 6)
combustion_glucose.add_product(h2o, 6)
combustion_hydrogen = s2.add_reaction()
combustion_hydrogen.add_reactant(h2,2)
combustion_hydrogen.add_reactant(o2,1)
combustion_hydrogen.add_product(h2o,2)
s2.save()
```
## Binary file formats
Now we're getting toward a numerically-based data structure, using
integers for object keys, we should think about binary serialisation.
Binary file formats are much smaller than human-readable text based formats,
so important when handling really big datasets.
One can compress a textual file format, of course, and with good compression algorithms this will be similar in size to the binary file. (C.f. discussions of Shannon information density!) However,
this has performance implications.
A hand-designed binary format is fast and small, at the loss of human readability.
The problem with binary file formats, is that, lacking complex data structures, one needs to supply the *length* of an item before that item:
```
class FakeSaveBinary: # Pretend binary-style writing to a list
# to make it easier to read at first.
def save(self, system, buffer):
buffer.append(len(system.elements))
for element in system.elements:
buffer.append(element.symbol)
buffer.append(len(system.molecules))
for molecule in system.molecules:
buffer.append(len(molecule.elements))
for element, number in molecule.elements.items():
buffer.append(element.id)
buffer.append(number)
buffer.append(len(system.reactions))
for reaction in system.reactions:
buffer.append(len(reaction.reactants))
for reactant, stoich in reaction.reactants.items():
buffer.append(reactant.id)
buffer.append(stoich)
buffer.append(len(reaction.products))
for product, stoich in reaction.products.items():
buffer.append(product.id)
buffer.append(stoich)
import io
arraybuffer = []
FakeSaveBinary().save(s2, arraybuffer)
arraybuffer
```
Deserialisation is left as an exercise for the reader.
## Endian-robust binary file formats
Having prepared our data as a sequence of data which can be recorded in a single byte,
we might think a binary file format on disk is as simple as saving
each number in one byte:
```
# First, turn symbol characters to equivalent integers (ascii)
intarray = [x.encode('ascii')[0] if type(x)==str else x for x in arraybuffer]
bytearray(intarray)
with open('system.mol','bw') as binfile:
binfile.write(bytearray(intarray))
```
However, this misses out on an unfortunate problem if we end up with large enough numbers to need more than one byte per integer, or we want to represent floats: different computer designs but the most-significant bytes of a multi-byte integer or float at the beginning or
end ('big endian' or 'little endian' data).
To get around this, we need to use a portable standard for making binary files.
One choice is **XDR**:
```
class XDRSavingSystem(System):
def __init__(self, system):
# Shallow Copy constructor
self.elements = system.elements
self.reactions = system.reactions
self.molecules = system.molecules
def save(self):
import xdrlib
buffer = xdrlib.Packer()
el_symbols = list(map(lambda x: x.symbol.encode('utf-8'),
self.elements))
buffer.pack_array(el_symbols,
buffer.pack_string)
#AUTOMATICALLY packs the length of the array first!
def _pack_pair(item):
buffer.pack_int(item[0].id)
buffer.pack_int(item[1])
def _pack_molecule(mol):
buffer.pack_array(mol.elements.items(),
_pack_pair)
buffer.pack_array(self.molecules, _pack_molecule)
def _pack_reaction(reaction):
buffer.pack_array(reaction.reactants.items(),
_pack_pair)
buffer.pack_array(reaction.products.items(),
_pack_pair)
buffer.pack_array(self.reactions, _pack_reaction)
return buffer
xdrsys = XDRSavingSystem(s2)
xdrbuff = xdrsys.save()
xdrbuff.get_buffer()
```
## A higher level approach to binary file formats: HDF5
This was quite painful. We've shown you it because it is very likely
you will encounter this kind of unpleasant binary file format in your work.
However, the recommended approach to building binary file formats is to use HDF5, a much higher level binary file format.
HDF5's approach requires you to represent your system in terms of high-dimensional matrices, like NumPy arrays.
It then saves these, and handles all the tedious number-of-field management for you.
```
import numpy as np
class HDF5SavingSystem(System):
def __init__(self, system):
# Shallow Copy constructor
self.elements = system.elements
self.reactions = system.reactions
self.molecules = system.molecules
def element_symbols(self):
return list(map(lambda x: x.symbol.encode('ascii'),
self.elements))
def molecule_matrix(self):
molecule_matrix = np.zeros((len(self.elements),
len(self.molecules)),dtype=int)
for molecule in self.molecules:
for element, n in molecule.elements.items():
molecule_matrix[element.id,
molecule.id]=n
return molecule_matrix
def reaction_matrix(self):
reaction_matrix = np.zeros((len(self.molecules),
len(self.reactions)),dtype=int)
for i, reaction in enumerate(self.reactions):
for reactant,n in reaction.reactants.items():
reaction_matrix[reactant.id,i]=-1*n
for product, n in reaction.products.items():
reaction_matrix[product.id,i]=n
return reaction_matrix
def write(self, filename):
import h5py
hdf = h5py.File(filename,'w')
string_type = h5py.special_dtype(vlen=bytes)
hdf.create_dataset('symbols', (len(self.elements),1),
string_type, self.element_symbols())
hdf.create_dataset('molecules', data=self.molecule_matrix())
hdf.create_dataset('reactions', data=self.reaction_matrix())
hdf.close()
saver=HDF5SavingSystem(s2)
saver.element_symbols()
saver.molecule_matrix()
saver.reaction_matrix()
saver.write('foo.hdf5')
import h5py
hdf_load=h5py.File('foo.hdf5')
np.array(hdf_load['reactions'])
```
Using a `sparse matrix` storage would be even better here, but we don't have time for that!
| github_jupyter |
```
import datetime
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.cm as cmx
df = pd.read_csv("./data/users/user_survey_raw.csv")
col_names = ["ts","version","application","switch","share","choice","cyclone_confidence","cyclone_nps","cyclone_narrative",
"cyclone_over_fast","fast_confidence","fast_nps","fast_narrative","fast_over_cyclone"]
df.columns = col_names
df.head()
t = df["version"]
out = df.groupby(["choice"])
cyclone = out.get_group("Cyclone DDS")
test = cyclone.groupby("version").count()["ts"].tolist()
cyclone.groupby("version").count()["ts"].keys().tolist()
# Let's start with a pie chart for ROS version, one for All, Cyclone, Fast
fig, ax = plt.subplots(1,3,figsize=(20, 10))
fig.tight_layout()
names = df.groupby("version").count()["ts"].keys().tolist()
colors = ["firebrick","darkorange","gold","olivedrab","dodgerblue","royalblue","purple"]
color_assign = {}
for idx,name in enumerate(names):
color_assign[name] = colors[idx]
################################################
title = "ROS 2 Version Used"
ax[0].set_title(title)
names = df.groupby("version").count()["ts"].keys().tolist()
vals = df.groupby("version").count()["ts"].values
c_list = []
for name in names:
c_list.append(color_assign[name])
patches,text,auto =ax[0].pie(vals,autopct='%1.1f%%', colors=c_list, shadow=True,radius=0.9)
ax[0].legend(names)
ax[0].legend(names,loc='lower right')
################################################
title = "ROS 2 Version Used -- Cyclone DDS Users"
ax[1].set_title(title)
out = df.groupby(["choice"])
cyclone = out.get_group("Cyclone DDS")
names = cyclone.groupby("version").count()["ts"].keys().tolist()
vals = cyclone.groupby("version").count()["ts"].tolist()
c_list = []
for name in names:
c_list.append(color_assign[name])
patches,text,auto =ax[1].pie(vals,colors=c_list, autopct='%1.1f%%', shadow=True,radius=0.9)
ax[1].legend(names,loc='lower right')
################################################
title = "ROS 2 Version Used -- Fast DDS Users"
ax[2].set_title(title)
out = df.groupby(["choice"])
fast = out.get_group("Fast DDS")
names = fast.groupby("version").count()["ts"].keys().tolist()
vals = fast.groupby("version").count()["ts"].tolist()
c_list = []
for name in names:
c_list.append(color_assign[name])
patches,text,auto =ax[2].pie(vals, colors=c_list,autopct='%1.1f%%', shadow=True,radius=0.9)
ax[2].legend(names)
ax[2].legend(names,loc='lower right')
plt.suptitle("ROS Distro Used By Preferred DDS Implementation",fontsize="xx-large")
plt.savefig("./plots/ROS2Version.png")
plt.show()
cm = plt.get_cmap('Reds')
# Now do the user choice
fig, ax = plt.subplots(1,2,figsize=(20, 10))
plt.suptitle("Preferred DDS Implementation and Confidence in Selected Implementation",fontsize="xx-large")
################################################
title = "Preferred DDS Implementation"
ax[0].set_title(title,fontsize="x-large")
names = df.groupby("choice").count()["ts"].keys().tolist()
vals = df.groupby("choice").count()["ts"].values
print(vals)
patches,text,auto =ax[0].pie(vals,autopct='%1.1f%%', colors=["red","blue"], shadow=True,radius=0.9)
ax[0].legend(names)
################################################
title = "ROS 2 Version Used -- Cyclone DDS Users"
ax[1].set_title(title,fontsize="x-large")
out = df.groupby(["choice"])
cyclone = out.get_group("Cyclone DDS")
names = cyclone.groupby("cyclone_confidence").count()["ts"].keys().tolist()
vals = cyclone.groupby("cyclone_confidence").count()["ts"].tolist()
cm = plt.get_cmap('Reds')
cs = []
for v in names:
cs.append(cm((float(v)/10.0)))
out = df.groupby(["choice"])
fast = out.get_group("Fast DDS")
vals2 = fast.groupby("fast_confidence").count()["ts"].tolist()
names2 = fast.groupby("fast_confidence").count()["ts"].keys().tolist()
cm = plt.get_cmap('Blues')
for v in names2:
cs.append(cm((float(v)/10.0)))
vals = vals + vals2
names = names + names2
patches,text,auto =ax[1].pie(vals, autopct='%1.1f%%', colors=cs,shadow=True,radius=0.9)
ax[1].legend(names)
title = "Preferred DDS Implementation by Type and Confidence Score"
ax[1].set_title(title)
plt.savefig("./plots/ROS2Choice.png")
plt.show()
cm = plt.get_cmap('Reds')
# Now do the user choice
fig, ax = plt.subplots(1,2,figsize=(20, 10))
plt.suptitle("Percentage of Respondents that Tried both Implementations and Their Preferences",fontsize="xx-large")
################################################
title = "Did you try more than one DDS / RMW vendor this year?"
ax[0].set_title(title,fontsize="x-large")
names = df.groupby("switch").count()["ts"].keys().tolist()
vals = df.groupby("switch").count()["ts"].values
print(vals)
patches,text,auto =ax[0].pie(vals,autopct='%1.1f%%', colors=["red","blue"], shadow=True,radius=0.9)
ax[0].legend(names)
################################################
out = df.groupby(["switch"])
switch = out.get_group("Yes")
names = switch.groupby("choice").count()["ts"].keys().tolist()
vals = switch.groupby("choice").count()["ts"].tolist()
new_names = []
for name in names:
new_names.append("Tried both implementations, preferred " + name)
names = new_names
out = df.groupby(["switch"])
no_switch = out.get_group("No")
vals2 = no_switch.groupby("choice").count()["ts"].tolist()
names2 = no_switch.groupby("choice").count()["ts"].keys().tolist()
new_names2 = []
for name in names2:
new_names2.append("Has only tried " + name)
names2 = new_names2
vals = vals + vals2
names = names + names2
colors = ["darkred","royalblue","indianred","cornflowerblue"]
patches,text,auto =ax[1].pie(vals, autopct='%1.1f%%',colors=colors, shadow=True,radius=0.9)
ax[1].legend(names)
title = "User Preference and Experimentation"
ax[1].set_title(title,fontsize="x-large")
plt.savefig("./plots/SwitchChoice.png")
plt.show()
fig, ax = plt.subplots(2,1,figsize=(20, 20))
plt.suptitle("Net Promoter Score by Preferred DDS Implementation",fontsize="xx-large")
out = df.groupby(["choice"])
cyclone = out.get_group("Cyclone DDS")
print(np.mean(cyclone["cyclone_nps"]))
print(np.median(cyclone["cyclone_nps"]))
ax[0].hist(cyclone["cyclone_nps"],density=False,align="mid",range=[0,10],bins=10, color="red")
ax[0].grid()
ax[0].set_xlim(0,11)
ticks = np.arange(0,11,1)
tick_names = ["{0}".format(t) for t in ticks]
ax[0].set_xticks(ticks)
ax[0].set_xticklabels(tick_names)
ax[0].set_ylabel("Respondents",fontsize="x-large")
ax[0].set_xlabel("Net Promotoer Score",fontsize="x-large")
ax[0].set_title("Cyclone DDS Net Promoter Score",fontsize="x-large")
out = df.groupby(["choice"])
fast = out.get_group("Fast DDS")
print(np.mean(fast["fast_nps"]))
print(np.median(fast["fast_nps"]))
bottom = np.arange(0,10,1)
ax[1].hist(fast["fast_nps"],density=False,align="mid",range=[0,10],bins=10, color="blue")
ax[1].grid()
ax[1].set_xlim(0,11)
ticks = np.arange(0,11,1)
tick_names = ["{0}".format(t) for t in ticks]
ax[1].set_xticks(ticks)
ax[1].set_xticklabels(tick_names)
ax[1].set_ylabel("Respondents",fontsize="x-large")
ax[1].set_xlabel("Net Promotoer Score",fontsize="x-large")
ax[1].set_title("Fast DDS Net Promoter Score",fontsize="x-large")
plt.savefig("./plots/DDSNPS.png")
plt.show()
```
| github_jupyter |
## Face and Facial Keypoint detection
After you've trained a neural network to detect facial keypoints, you can then apply this network to *any* image that includes faces. The neural network expects a Tensor of a certain size as input and, so, to detect any face, you'll first have to do some pre-processing.
1. Detect all the faces in an image using a face detector (we'll be using a Haar Cascade detector in this notebook).
2. Pre-process those face images so that they are grayscale, and transformed to a Tensor of the input size that your net expects. This step will be similar to the `data_transform` you created and applied in Notebook 2, whose job was tp rescale, normalize, and turn any iimage into a Tensor to be accepted as input to your CNN.
3. Use your trained model to detect facial keypoints on the image.
---
In the next python cell we load in required libraries for this section of the project.
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
```
#### Select an image
Select an image to perform facial keypoint detection on; you can select any image of faces in the `images/` directory.
```
import cv2
# load in color image for face detection
image = cv2.imread('images/obamas.jpg')
# switch red and blue color channels
# --> by default OpenCV assumes BLUE comes first, not RED as in many images
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# plot the image
fig = plt.figure(figsize=(9,9))
plt.imshow(image)
```
## Detect all faces in an image
Next, you'll use one of OpenCV's pre-trained Haar Cascade classifiers, all of which can be found in the `detector_architectures/` directory, to find any faces in your selected image.
In the code below, we loop over each face in the original image and draw a red square on each face (in a copy of the original image, so as not to modify the original). You can even [add eye detections](https://docs.opencv.org/3.4.1/d7/d8b/tutorial_py_face_detection.html) as an *optional* exercise in using Haar detectors.
An example of face detection on a variety of images is shown below.
<img src='images/haar_cascade_ex.png' width=80% height=80%/>
```
# load in a haar cascade classifier for detecting frontal faces
face_cascade = cv2.CascadeClassifier('detector_architectures/haarcascade_frontalface_default.xml')
# run the detector
# the output here is an array of detections; the corners of each detection box
# if necessary, modify these parameters until you successfully identify every face in a given image
faces = face_cascade.detectMultiScale(image, 1.2, 2)
print(faces)
# make a copy of the original image to plot detections on
image_with_detections = image.copy()
# loop over the detected faces, mark the image where each face is found
for (x,y,w,h) in faces:
# draw a rectangle around each detected face
# you may also need to change the width of the rectangle drawn depending on image resolution
cv2.rectangle(image_with_detections,(x,y),(x+w,y+h),(255,0,0),3)
fig = plt.figure(figsize=(9,9))
plt.imshow(image_with_detections)
```
## Loading in a trained model
Once you have an image to work with (and, again, you can select any image of faces in the `images/` directory), the next step is to pre-process that image and feed it into your CNN facial keypoint detector.
First, load your best model by its filename.
```
import torch
from models import Net
net = Net()
## TODO: load the best saved model parameters (by your path name)
## You'll need to un-comment the line below and add the correct name for *your* saved model
net.load_state_dict(torch.load('saved_models/keypoints_model_1.pt'))
## print out your net and prepare it for testing (uncomment the line below)
net.eval()
```
## Keypoint detection
Now, we'll loop over each detected face in an image (again!) only this time, you'll transform those faces in Tensors that your CNN can accept as input images.
### TODO: Transform each detected face into an input Tensor
You'll need to perform the following steps for each detected face:
1. Convert the face from RGB to grayscale
2. Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
3. Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
4. Reshape the numpy image into a torch image.
You may find it useful to consult to transformation code in `data_load.py` to help you perform these processing steps.
### TODO: Detect and display the predicted keypoints
After each face has been appropriately converted into an input Tensor for your network to see as input, you'll wrap that Tensor in a Variable() and can apply your `net` to each face. The ouput should be the predicted the facial keypoints. These keypoints will need to be "un-normalized" for display, and you may find it helpful to write a helper function like `show_keypoints`. You should end up with an image like the following with facial keypoints that closely match the facial features on each individual face:
<img src='images/michelle_detected.png' width=30% height=30%/>
```
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=1):
for i in range(batch_size):
plt.figure(figsize=(10,5))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[0].data # get the image from it's wrapper
image = image.numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[0].data
predicted_key_pts = predicted_key_pts.numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts*50.0+100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
image_copy = np.copy(image)
from data_load import Rescale, RandomCrop, Normalize, ToTensor
import torch
margin = 50
# loop over the detected faces from your haar cascade
for (x,y,w,h) in faces:
# Select the region of interest that is the face in the image
roi = image_copy[y-margin:y+h+margin, x-margin:x+w+margin]
## TODO: Convert the face region from RGB to grayscale
roi = cv2.cvtColor(roi, cv2.COLOR_RGB2GRAY)
## TODO: Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
roi = roi/255.0
## TODO: Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
roi = cv2.resize(roi, (224, 224))
## TODO: Reshape the numpy image shape (H x W x C) into a torch image shape (C x H x W)
roi = roi.reshape(1, roi.shape[0], roi.shape[1], 1)
roi = torch.from_numpy(roi.transpose((0, 3, 1, 2)))
## TODO: Make facial keypoint predictions using your loaded, trained network
## perform a forward pass to get the predicted facial keypoints
roi = roi.type(torch.FloatTensor)
output_pts = net(roi)
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
## TODO: Display each detected face and the corresponding keypoints
visualize_output(roi, output_pts)
```
| github_jupyter |
```
import pandas as pd
in_patient_train = pd.read_csv('Train_Inpatientdata-1542865627584.csv')
```
40,474 rows in the in-patient datafile
```
in_patient_train.shape
```
30 columns, across provider, amount reimbursed, physicians, admission date, all the codes.
```
in_patient_train.head()
```
description of continuous variables. 10,300usd average reimbursed claim. The deductible is 1,068usd and that is always the same, across nearly 40,000 records. That's odd.
```
in_patient_train.describe()
```
There are 112 claims without an attending physician, which seems odd for in-patient admissions. Not sure what that means. Other NAs make sense to me, since there won't necessarily be 2nd and so forth diagnosis codes and it seems possible there wouldn't be any procedure codes since physician might not be able to do anything (and thus submit a claim.) Notice there are 899 NAs in deductible paid.
```
import numpy as np
a = np.sum(in_patient_train.isnull())
a[a>0]
```
Looking at unique counts. It does appear the claimID is the unique identifier since that has 40474 unique values, the number of rows in the dataframe. I note that there are 398 unique values of claim start date, with more than a few dates with only one or two claims starting on those dates, vs 145 on other days. On the other hand, for claim end date, the smallest count is 75 on Christmas 2009. That also seems odd to me. There seem to be physicians with 300 or more claims, and then a number that show up only on 1 claim.
```
for colmn in in_patient_train.columns:
print(in_patient_train[colmn].value_counts())
```
Let's look at the distribution of claims by patient.
```
b = in_patient_train.groupby('BeneID').agg(['count'])
b.columns = [x[0] for x in b.columns]
b
```
most patients have only 1 or 2 claims, with a rightward skew
```
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.hist(b['ClaimID'])
plt.xlabel('patient claims')
plt.ylabel('count of patients with # of claims')
c = in_patient_train.groupby('AttendingPhysician').agg(['count'])
c.columns = [x[0] for x in c.columns]
c
```
Most physicians only have 0-25 claims, but a few are on out to nearly 400 claims
```
plt.hist(c['ClaimID'])
plt.xlabel('number of claims for attending physician')
plt.ylabel('count of physicians with # of claims')
d = in_patient_train.groupby('OtherPhysician').agg(['count'])
d.columns = [x[0] for x in d.columns]
d
```
same pattern for 'other' physician, with most having 0-25 claims, but a few showing up on 400 claims
```
plt.hist(d['ClaimID'])
plt.xlabel('number of claims for other physician')
plt.ylabel('count of physicians with # of claims')
e = in_patient_train.groupby('OperatingPhysician').agg(['count'])
e.columns = [x[0] for x in e.columns]
e
plt.hist(e['ClaimID'])
plt.xlabel('number of claims for operating physician')
plt.ylabel('count of physicians with # of claims')
```
The amount of reimbursement is up to 10,000 USD, but there are some out to 100,000USD
```
plt.hist(in_patient_train['InscClaimAmtReimbursed'])
plt.xlabel('amount of insurance claim reimbursement')
plt.ylabel('count of claims with that reimbursement')
```
count of diagnostic codes and procedure codes per claim, then see what average number is.
```
in_patient_train.loc[in_patient_train['ClaimID']=='CLM46614',['ClmDiagnosisCode_7','ClmDiagnosisCode_9','ClmDiagnosisCode_10']]
np.sum(in_patient_train.loc[in_patient_train['ClaimID']=='CLM46614',['ClmDiagnosisCode_7','ClmDiagnosisCode_9','ClmDiagnosisCode_10']].count(axis=0))
in_patient_train.columns
diag_colmn = ['ClmDiagnosisCode_1', 'ClmDiagnosisCode_2', 'ClmDiagnosisCode_3','ClmDiagnosisCode_4',
'ClmDiagnosisCode_5', 'ClmDiagnosisCode_6','ClmDiagnosisCode_7', 'ClmDiagnosisCode_8',
'ClmDiagnosisCode_9', 'ClmDiagnosisCode_10']
proced_colmn = ['ClmProcedureCode_1', 'ClmProcedureCode_2','ClmProcedureCode_3', 'ClmProcedureCode_4',
'ClmProcedureCode_5','ClmProcedureCode_6']
in_patient_train['Num_diag_codes'] = [np.sum(in_patient_train.loc[in_patient_train['ClaimID']==claim,diag_colmn].\
count(axis=0)) for claim in in_patient_train['ClaimID']]
in_patient_train['Num_diag_codes']
```
9 appears to be a popular number of diagnostic codes for a claim
```
plt.hist(in_patient_train['Num_diag_codes'])
plt.xlabel('number of diagnostic codes on claim')
plt.ylabel('count of claims with # of diag codes')
in_patient_train['Num_proc_codes'] = [np.sum(in_patient_train.loc[in_patient_train['ClaimID']==claim,proced_colmn].\
count(axis=0)) for claim in in_patient_train['ClaimID']]
```
a little under half admissions have no procedures, about the same number have one procedure, and a few claims have up to 4 procedures.
```
plt.hist(in_patient_train['Num_proc_codes'])
plt.xlabel('number of procedure codes on claim')
plt.ylabel('count of claims with # of proc codes')
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import scipy as sp
#Visuallization
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import seaborn as sns
from pandas.plotting import scatter_matrix
sns.set_style('whitegrid')
mpl.style.use('ggplot')
%matplotlib inline
pylab.rcParams['figure.figsize'] = 12,8
import random
import time
import warnings
warnings.filterwarnings('ignore')
# from sklearn.linear_model import LogisticRegression
# from sklearn.svm import SVC, LinearSVC
# from sklearn.ensemble import RandomForestClassifier
# from sklearn.naive_bayes import GaussianNB
from sklearn import svm, tree,linear_model, neighbors, naive_bayes, ensemble, discriminant_analysis, gaussian_process
from xgboost import XGBClassifier
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn import feature_selection
from sklearn import model_selection
from sklearn import metrics
#Import data
path = '~/Documents/Kaggle/Titanic/'
data_raw = pd.read_csv(path+'train.csv')
data_val = pd.read_csv(path+'test.csv')
#make copy and group train and test dataset in list since they are both similar and can be treated with same operations and analysis
data = data_raw.copy(deep=True)
data_group = [data,data_val]
#checking missing values all fields
print('train data missing values: ')
print(data.isnull().sum())
print('-'*20)
print('test data missing values: ')
print(data_val.isnull().sum())
#Imputation missing values with median and mode values
for eachset in data_group:
eachset['Age'].fillna(eachset['Age'].median(),inplace=True)
eachset['Embarked'].fillna(eachset['Embarked'].mode()[0],inplace=True)
eachset['Fare'].fillna(eachset['Fare'].median(),inplace=True)
#list of columns to drop since they dont have useful info
drop_columns = ['PassengerId','Cabin','Ticket']
data.drop(drop_columns,axis=1,inplace=True)
print('train data missing')
print(data.isnull().sum())
print('-'*20)
print('test data missing')
print(data_val.isnull().sum())
### Feature egnineering
## Create FamilySize column by add number of SibSp and Parch
## Create IsAlone column
## Create Title column extracted from Name column
#creating new field called IsAlone, having 1 as True, 0 as False
for eachset in data_group:
eachset['FamilySize'] = eachset['SibSp'] + eachset['Parch'] + 1
eachset['IsAlone'] = 1 #initialize with 1
eachset['IsAlone'].loc[eachset['FamilySize']>1]=0 #change to 0 if family size > 1
#extract Title from Name column
eachset['Title'] = eachset['Name'].str.split(", ",expand=True)[1].str.split(".",expand=True)[0]
eachset['FareBin'] = pd.qcut(eachset['Fare'],4)
eachset['AgeBin'] = pd.cut(eachset['Age'].astype(int),5)
# cut and qcut diff: https://stackoverflow.com/questions/30211923/what-is-the-difference-between-pandas-qcut-and-pandas-cut/30214901
# discretization of continuous variable explained: http://www.uta.fi/sis/tie/tl/index/Datamining6.pdf
#change titles which have count less than 10 to 'Misc'
title_names = (data['Title'].value_counts() < 10)
data['Title'] = data['Title'].apply(lambda x: 'Misc' if title_names.loc[x]==True else x)
label = LabelEncoder()
for eachset in data_group:
eachset['Sex_Code'] = label.fit_transform(eachset['Sex'])
eachset['Embarked_Code'] = label.fit_transform(eachset['Embarked'])
eachset['Title_Code'] = label.fit_transform(eachset['Title'])
eachset['AgeBin_Code'] = label.fit_transform(eachset['AgeBin'])
eachset['FareBin_Code'] = label.fit_transform(eachset['FareBin'])
Target = ['Survived']
data_x = ['Sex','Pclass','Embarked','Title','SibSp','Parch','Age','Fare','FamilySize','IsAlone']
data_x_calc =['Sex_Code','Pclass','Embarked_Code','Title_Code','SibSp','Parch','Age','Fare']
data_xy = Target + data_x
data_x_bin = ['Sex_Code','Pclass','Embarked_Code','Title_Code','FamilySize','AgeBin_Code','FareBin_Code']
data_xy_bin = Target + data_x_bin
data_dummy = pd.get_dummies(data[data_x])
data_x_dummy = data_dummy.columns.tolist()
data_xy_dummy = Target + data_x_dummy
train_x,test_x,train_y,test_y = model_selection.train_test_split(data[data_x_calc],data[Target],random_state=0)
train_x_bin,test_x_bin,train_y_dummy,test_y_dummy = model_selection.train_test_split(data_dummy[data_x_dummy],data[Target],random_state = 0)
train_x_bin.head()
# Explore discrete field with survive rate using group by
for x in data_x:
if data[x].dtype!='float64':
print('Survival Correlation by:',x)
print(data[[x,Target[0]]].groupby(x,as_index=False).mean())
print('-'*20)
# Explore title field
print(pd.crosstab(data['Title'],data[Target[0]]))
### Visuallization for EDA
plt.figure(figsize=[16,12])
plt.subplot(231)
plt.boxplot(x=data['Fare'],showmeans=True,meanline = True)
plt.title('Fare Boxplot')
plt.ylabel('Fare ($)')
plt.subplot(232)
plt.boxplot(data['Age'],showmeans=True,meanline = True)
plt.title('Age Boxplot')
plt.ylabel('Age (Years)')
plt.subplot(233)
plt.boxplot(data['FamilySize'],showmeans=True,meanline=True)
plt.title('Family Size Boxplot')
plt.ylabel('Family Size (#)')
plt.subplot(234)
plt.hist(x=[data[data['Survived']==1]['Fare'], data[data['Survived']==0]['Fare']],stacked=True,color=['b','r'],label=['Survived','Dead'])
plt.title('Fare hist by survival')
plt.ylabel('# of Passengers')
plt.xlabel('Fare ($)')
plt.legend()
plt.subplot(235)
plt.hist(x=[data[data['Survived']==1]['Age'], data[data['Survived']==0]['Age']],stacked=True,color=['b','r'],label=['Survived','Dead'])
plt.title('Age hist by survival')
plt.ylabel('# of Passengers')
plt.xlabel('Age (years)')
plt.legend()
plt.subplot(236)
plt.hist(x=[data[data['Survived']==1]['FamilySize'], data[data['Survived']==0]['FamilySize']],stacked=True,color=['b','r'],label=['Survived','Dead'])
plt.title('Family Size hist by survival')
plt.ylabel('# of Passengers')
plt.xlabel('Family Size')
plt.legend()
fig, saxis = plt.subplots(2, 3,figsize=(16,12))
sns.barplot(x='Embarked',y='Survived',data=data,ax=saxis[0,0])
sns.barplot(x='Pclass',y='Survived',order = [1,2,3],data=data,ax=saxis[0,1])
sns.barplot(x='IsAlone',y='Survived',order = [1,0],data=data,ax=saxis[0,2])
sns.pointplot(x='FareBin',y='Survived',data=data,ax=saxis[1,0])
sns.pointplot(x='AgeBin',y='Survived',data=data,ax=saxis[1,1])
sns.pointplot(x='FamilySize',y='Survived',data=data,ax=saxis[1,2])
fig, (axis1,axis2,axis3) = plt.subplots(1,3,figsize=(14,12))
sns.boxplot(x='Pclass',y='Fare',hue='Survived',data=data,ax=axis1)
axis1.set_title('Pclass vs Fare')
sns.violinplot(x='Pclass',y='Age',hue='Survived',split=True,data=data,ax=axis2)
axis2.set_title('Pclass vs Age')
sns.boxplot(x='Pclass',y='FamilySize',hue='Survived',data=data,ax=axis3)
axis3.set_title('Pclass vs FamilySize')
## Compare sex to other fields with survival rate
fig, qaxis = plt.subplots(1,3,figsize=(14,12))
sns.barplot(x='Sex',y='Survived',hue='Embarked', data =data, ax=qaxis[0])
axis1.set_title('Sex vs Embarked survival comparision')
sns.barplot(x='Sex',y='Survived',hue='Pclass', data =data, ax=qaxis[1])
axis1.set_title('Sex vs Pclass survival comparision')
sns.barplot(x='Sex',y='Survived',hue='IsAlone', data =data, ax=qaxis[2])
axis1.set_title('Sex vs IsAlone survival comparision')
fig,(maxis1,maxis2) = plt.subplots(1,2,figsize=(14,12))
#how family affect sex and survival rate
sns.pointplot(x='FamilySize',y='Survived',hue='Sex',data=data,
palette = {'male': 'blue','female':'pink'},
markers = ['*','o'],linestyles=['-','--'],ax =maxis1)
#how class affect sex and survival rate
sns.pointplot(x='Pclass',y='Survived',hue='Sex',data=data,
palette = {'male': 'blue','female':'pink'},
markers = ['*','o'],linestyles=['-','--'],ax =maxis2)
sns.heatmap(data.corr(),square=True,cbar_kws={'shrink':.9},annot=True)
MLA = [
#ensemble
ensemble.AdaBoostClassifier(),
ensemble.BaggingClassifier(),
ensemble.ExtraTreesClassifier(),
ensemble.GradientBoostingClassifier(),
ensemble.RandomForestClassifier(),
#Gaussian Process
gaussian_process.GaussianProcessClassifier(),
#GLM
linear_model.LogisticRegressionCV(),
linear_model.PassiveAggressiveClassifier(),
linear_model.RidgeClassifierCV(),
linear_model.SGDClassifier(),
linear_model.Perceptron(),
#Naive Bayes
naive_bayes.BernoulliNB(),
naive_bayes.GaussianNB(),
#Nearest Neighbor
neighbors.KNeighborsClassifier(),
#SVM
svm.SVC(probability=True),
svm.NuSVC(probability=True),
svm.LinearSVC(),
#Trees
tree.DecisionTreeClassifier(),
tree.ExtraTreeClassifier(),
#Discriminant Analysis
discriminant_analysis.LinearDiscriminantAnalysis(),
discriminant_analysis.QuadraticDiscriminantAnalysis(),
#XG Boost
XGBClassifier()
]
cv_split = model_selection.ShuffleSplit(n_splits=10,test_size=0.3,train_size=0.6,random_state =0)
MLA_columns = ['MLA Name','MLA Parameters','MLA train accuracy mean','MLA test accuracy mean','MLA test accuracy 3*STD','MLA Time']
MLA_compare = pd.DataFrame(columns=MLA_columns)
MLA_predict = data[Target]
row_index=0
for alg in MLA:
MLA_name = alg.__class__.__name__
MLA_compare.loc[row_index,'MLA Name'] = MLA_name
MLA_compare.loc[row_index,'MLA Parameters'] = str(alg.get_params())
cv_results = model_selection.cross_validate(alg,data[data_x_bin],data[Target],cv=cv_split)
MLA_compare.loc[row_index,'MLA Time'] = cv_results['fit_time'].mean()
MLA_compare.loc[row_index,'MLA train accuracy mean'] = cv_results['test_score'].mean()
MLA_compare.loc[row_index,'MLA test accuracy mean'] = cv_results['test_score'].mean()
MLA_compare.loc[row_index,'MLA test accuracy 3*STD'] = cv_results['test_score'].std()*3
alg.fit(data[data_x_bin],data[Target])
MLA_predict[MLA_name] = alg.predict(data[data_x_bin])
row_index+=1
MLA_compare.sort_values(by=['MLA test accuracy mean'],ascending=False,inplace=True)
MLA_compare
sns.barplot(x='MLA test accuracy mean',y='MLA Name',data= MLA_compare,color='r')
plt.title('ML Alg Accuracy Score')
plt.xlabel('Accuracy Score')
plt.ylabel('Alg')
cv_results
```
| github_jupyter |
<img src='./img/logoline_12000.png' align='right' width='100%'></img>
# Tutorial on creating a climate index for wind chill
In this tutorial we will plot a map of wind chill over Europe using regional climate reanalysis data (UERRA) of wind speed and temperature. From the WEkEO Jupyterhub we will download this data from the Climate Data Store (CDS) of the Copernicus Climate Change Service (C3S). The tutorial comprises the following steps:
1. [Search and download](#search_download) regional climate reanalysis data (UERRA) of 10m wind speed and 2m temperature.
2. [Read data](#read_data): Once downloaded, we will read and understand the data, including its variables and coordinates.
3. [Calculate wind chill index](#wind_chill): We will calculate the wind chill index from the two parameters of wind speed and temperature, and view a map of average wind chill over Europe.
4. [Calculate wind chill with ERA5](#era5): In order to assess the reliability of the results, repeat the process with ERA5 reanalysis data and compare the results with those derived with UERRA.
<img src='./img/climate_indices.png' align='center' width='100%'></img>
## <a id='search_download'></a>1. Search and download data
Before we begin we must prepare our environment. This includes installing the Application Programming Interface (API) of the CDS, and importing the various python libraries that we will need.
#### Install CDS API
To install the CDS API, run the following command. We use an exclamation mark to pass the command to the shell (not to the Python interpreter).
```
!pip install cdsapi
```
#### Import libraries
We will be working with data in NetCDF format. To best handle this data we need a number of libraries for working with multidimensional arrays, in particular Xarray. We will also need libraries for plotting and viewing data, in particular Matplotlib and Cartopy.
```
# CDS API
import cdsapi
# Libraries for working with multidimensional arrays
import numpy as np
import xarray as xr
# Libraries for plotting and visualising data
import matplotlib.path as mpath
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import cartopy.feature as cfeature
```
#### Enter your CDS API key
Please follow the steps at this link to obtain a User ID and a CDS API key:
https://cds.climate.copernicus.eu/api-how-to
Once you have these, please enter them in the fields below by replacing "UID" with your User ID, and "API_KEY" with your API key.
```
CDS_URL = "https://cds.climate.copernicus.eu/api/v2"
# enter your CDS authentication key:
CDS_KEY = "UID:API_KEY"
```
#### Search for climate data to calculate wind chill index
The wind chill index we will calculate takes two parameters as input, these are 2m near-surface air temperature, and 10m wind speed. Data for these parameters are available as part of the UERRA regional reanalysis dataset for Europe for the period 1961 to 2019. We will search for this data on the CDS website: http://cds.climate.copernicus.eu. The specific dataset we will use is the UERRA regional reanalysis for Europe on single levels from 1961 to 2019.
<img src='./img/CDS.jpg' align='left' width='45%'></img> <img src='./img/CDS_UERRA.png' align='right' width='45%'></img>
Having selected the dataset, we now need to specify what product type, variables, temporal and geographic coverage we are interested in. These can all be selected in the **"Download data"** tab. In this tab a form appears in which we will select the following parameters to download:
- Origin: `UERRA-HARMONIE`
- Variable: `10m wind speed` and `2m temperature` (these will need to be selected one at a time)
- Year: `1998 to 2019`
- Month: `December`
- Day: `15`
- Time: `12:00`
- Format: `NetCDF`
<img src='./img/CDS_UERRA_download.png' align='center' width='45%'></img>
At the end of the download form, select **"Show API request"**. This will reveal a block of code, which you can simply copy and paste into a cell of your Jupyter Notebook (see cells below). You will do this twice: once for 10m wind speed and again for 2m temperature.
#### Download data
Having copied the API requests into the cells below, run these to retrieve and download the data you requested into your local directory.
```
c = cdsapi.Client(url=CDS_URL, key=CDS_KEY)
c.retrieve(
'reanalysis-uerra-europe-single-levels',
{
'origin': 'uerra_harmonie',
'variable': '10m_wind_speed',
'year': [
'1998', '1999', '2000',
'2001', '2002', '2003',
'2004', '2005', '2006',
'2007', '2008', '2009',
'2010', '2011', '2012',
'2013', '2014', '2015',
'2016', '2017', '2018',
],
'month': '12',
'day': '15',
'time': '12:00',
'format': 'netcdf',
},
'UERRA_ws10m.nc')
c = cdsapi.Client(url=CDS_URL, key=CDS_KEY)
c.retrieve(
'reanalysis-uerra-europe-single-levels',
{
'origin': 'uerra_harmonie',
'variable': '2m_temperature',
'year': [
'1998', '1999', '2000',
'2001', '2002', '2003',
'2004', '2005', '2006',
'2007', '2008', '2009',
'2010', '2011', '2012',
'2013', '2014', '2015',
'2016', '2017', '2018',
],
'month': '12',
'day': '15',
'time': '12:00',
'format': 'netcdf',
},
'UERRA_t2m.nc')
```
## <a id='read_data'></a>2. Read Data
Now that we have downloaded the data, we can start to play ...
We have requested the data in NetCDF format. This is a commonly used format for array-oriented scientific data.
To read and process this data we will make use of the Xarray library. Xarray is an open source project and Python package that makes working with labelled multi-dimensional arrays simple, efficient, and fun! We will read the data from our NetCDF file into an Xarray **"dataset"**
```
fw = 'UERRA_ws10m.nc'
ft = 'UERRA_t2m.nc'
# Create Xarray Dataset
dw = xr.open_dataset(fw)
dt = xr.open_dataset(ft)
```
Now we can query our newly created Xarray datasets ...
```
dw
dt
```
We see that dw (dataset for wind speed) has one variable called **"si10"**. If you view the documentation for this dataset on the CDS you will see that this is the wind speed valid for a grid cell at the height of 10m above the surface. It is computed from both the zonal (u) and the meridional (v) wind components by $\sqrt{(u^{2} + v^{2})}$. The units are m/s.
The other dataset, dt (2m temperature), has a variable called **"t2m"**. According to the documentation on the CDS this is air temperature valid for a grid cell at the height of 2m above the surface, in units of Kelvin.
While an Xarray **dataset** may contain multiple variables, an Xarray **data array** holds a single multi-dimensional variable and its coordinates. To make the processing of the **si10** and **t2m** data easier, we will convert them into Xarray data arrays.
```
# Create Xarray Data Arrays
aw = dw['si10']
at = dt['t2m']
```
## <a id='wind_chill'></a>3. Calculate wind chill index
There are several indices to calculate wind chill based on air temperature and wind speed. Until recently, a commonly applied index was the following:
$\textit{WCI} = (10 \sqrt{\upsilon}-\upsilon + 10.5) \cdot (33 - \textit{T}_{a})$
where:
- WCI = wind chill index, $kg*cal/m^{2}/h$
- $\upsilon$ = wind velocity, m/s
- $\textit{T}_{a}$ = air temperature, °C
We will use the more recently adopted North American and United Kingdom wind chill index, which is calculated as follows:
$\textit{T}_{WC} = 13.12 + 0.6215\textit{T}_{a} - 11.37\upsilon^{0.16} + 0.3965\textit{T}_{a}\upsilon^{0.16}$
where:
- $\textit{T}_{WC}$ = wind chill index
- $\textit{T}_{a}$ = air temperature in degrees Celsius
- $\upsilon$ = wind speed at 10 m standard anemometer height, in kilometres per hour
To calculate $\textit{T}_{WC}$ we first have to ensure our data is in the right units. For the wind speed we need to convert from m/s to km/h, and for air temperature we need to convert from Kelvin to degrees Celsius:
```
# wind speed, convert from m/s to km/h: si10 * 1000 / (60*60)
w = aw * 3600 / 1000
# air temperature, convert from Kelvin to Celsius: t2m - 273.15
t = at - 273.15
```
Now we can calculate the North American and United Kingdom wind chill index:
$\textit{T}_{WC} = 13.12 + 0.6215\textit{T}_{a} - 11.37\upsilon^{0.16} + 0.3965\textit{T}_{a}\upsilon^{0.16}$
```
twc = 13.12 + (0.6215*t) - (11.37*(w**0.16)) + (0.3965*t*(w**0.16))
```
Let's calculate the average wind chill for 12:00 on 15 December for the 20 year period from 1998 to 2019:
```
twc_mean = twc.mean(dim='time')
```
Now let's plot the average wind chill for this time over Europe:
```
# create the figure panel
fig = plt.figure(figsize=(10,10))
# create the map using the cartopy Orthographic projection
ax = plt.subplot(1,1,1, projection=ccrs.Orthographic(central_longitude=8., central_latitude=42.))
# add coastlines
ax.coastlines()
ax.gridlines(draw_labels=False, linewidth=1, color='gray', alpha=0.5, linestyle='--')
# provide a title
ax.set_title('Wind Chill Index 12:00, 15 Dec, 1998 to 2019')
# plot twc
im = plt.pcolormesh(twc_mean.longitude, twc_mean.latitude,
twc_mean, cmap='viridis', transform=ccrs.PlateCarree())
# add colourbar
cbar = plt.colorbar(im)
cbar.set_label('Wind Chill Index')
```
Can you identify areas where frostbite may occur (see chart below)?
<img src='./img/Windchill_effect_en.svg' align='left' width='60%'></img>
RicHard-59, CC BY-SA 3.0 <https://creativecommons.org/licenses/by-sa/3.0>, via Wikimedia Commons
## <a id='era5'></a>4. Exercise: Repeat process with ERA5 data and compare results
So far you have plotted wind chill using the UERRA regional reanalysis dataset, but how accurate is this plot? One way to assess a dataset is to compare it with an alternative independent one to see what differences there may be. An alternative to UERRA is the ERA5 reanalysis data that you used in the previous tutorials. Repeat the steps above with ERA5 and compare your results with those obtained using UERRA.
<hr>
| github_jupyter |
# Automatic music generation system (AMGS) - Pop genre
An affective rule-based generative music system that generates retro pop music.
```
import numpy as np
import pandas as pd
import mido
import scipy.io
import time
import statistics
from numpy.random import choice
from IPython.display import clear_output
import math
import json
# set up midi ports
print(mido.get_output_names())
percussion = mido.open_output('IAC Driver Bus 1')
piano = mido.open_output('IAC Driver Bus 2')
# read in composed progressions
with open('composed_progressions.txt') as json_file:
data = json.load(json_file)
```
# Scales, progressions and patterns
This section determines the scales, chord progressions, melodic patterns and rhythmic patterns used by the system.
```
import playerContainer
import progressionsContainer as progs
# initialize helper functions
player = playerContainer.PlayerContainer()
# set relative positions of notes in major and parallel minor scales
# MIDI note numbers for C major: 60 (C4), 62 (D), 64 (E), 65 (F), 67 (G), 69 (A), 71 (B)
tonic = 60
majorScale = [tonic, tonic+2, tonic+4, tonic+5, tonic+7, tonic+9, tonic+11]
minorScale = [tonic, tonic+2, tonic+3, tonic+5, tonic+7, tonic+8, tonic+10]
# test sound -> should hear note being played through audio workstation
ichannel = 1
ivelocity = 64
msg = mido.Message('note_on',channel=ichannel,note=tonic,velocity=ivelocity)
piano.send(msg)
time.sleep(0.50)
msg = mido.Message('note_off',channel=ichannel,note=tonic,velocity=ivelocity)
piano.send(msg)
# draft: percussion
# Ableton's drum pads are mapped by default to MIDI notes 36-51
ichannel = 10
ivelocity = 64
inote = 51
msg = mido.Message('note_on',channel=ichannel,note=inote,velocity=ivelocity)
percussion.send(msg)
```
# Player (Main)
This section puts together all the functions and generates music based on the current arousal and valence values.
**Arousal-based params**
1. roughness. Lower roughness -> higher note density.
3. loudness
4. tempo. Minimum = 60bpm, maximum = 160bpm
**Valence-based params**
1. voicing
2. chord progression
```
# artificially determine arousal-valence trajectory
#np.array([0.9, 0.8, 0.7, 0.6, 0.5, 0.4, 0.3, 0.2])
input_arousal = np.array([0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9])
input_arousal = np.repeat(input_arousal, 8)
input_valence = np.array([0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9])
input_valence = np.repeat(input_valence, 8)
# or randomly generate a trajectory
rng = np.random.default_rng()
# low arousal, low valence, 40-bar progression
input_arousal = rng.integers(50, size=40)/100
input_valence = rng.integers(50, size=40)/100
# high arousal, low valence, 40-bar progression
input_arousal = rng.integers(50, high=100, size=40)/100
input_valence = rng.integers(50, size=40)/100
# low arousal, high valence, 40-bar progression
input_arousal = rng.integers(50, size=40)/100
input_valence = rng.integers(50, high=100, size=40)/100
# high arousal, high valence, 40-bar progression
input_arousal = rng.integers(50, high=100, size=40)/100
input_valence = rng.integers(50, high=100, size=40)/100
input_arousal
input_arousal = np.array([0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9])
input_arousal = np.repeat(input_arousal, 4)
input_valence = np.array([0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9])
input_valence = np.repeat(input_valence, 4)
print(input_valence)
print(input_arousal)
```
* melody generator, harmony generator, bass generator
* implement voice leading logic
**POSSIBLE CHANGES**
* maybe we can do moving average tempo instead -> but is that sacrificing accuracy of emotion feedback?
```
# initialize params: next_chord, minimal loudness, velocity, current_motive
next_chord = []
current_motive=0
# initialize memory of previous harmony and melody notes (partially determines current harmony/melody notes)
prev_noteset, melody_note = [], []
# keep track of current bar
for bar in range(len(input_arousal)):
# set arousal and valence, keep track of current bar in 8-bar progressions
arousal = input_arousal[bar]
valence = input_valence[bar]
bar = bar%8
print("arousal: ", arousal, "---valence: ", valence, "---bar: ", bar)
# set simple params: roughness, voicing, loudness, tempo
roughness = 1-arousal
low_loudness = 40 + (arousal*40)
loudness = (round(arousal*10))/10*40+60
bpm = 60 + arousal * 100
volume = int(50 + (arousal*30))
# allocate note densities
n_subdivisions = 8
if arousal >= 0.75:
arousal_cat='high'
elif arousal >= 0.40:
arousal_cat='moderate'
else:
arousal_cat='low'
activate1 = [x for x in data['rhythmic_motives'] if x['bar']==bar if x['arousal']==arousal_cat][0]['motive']
activate2 = player.setRoughness(n_subdivisions, roughness+0.3)
# change volume of instruments
# instruments[0]: piano, instruments[1]: clarinet, instruments[2]: strings
msg = mido.Message('control_change',channel=ichannel,control=7,value=volume)
instruments[0].send(msg), instruments[1].send(msg), instruments[1].send(msg)
# select chord to be sounded
if next_chord==[]:
# if next chord has not already been determined, then select randomly as usual
chord, next_chord = progs.selectChord(data['progressions'], valence, bar)
else:
chord = next_chord.pop(0)
# generate set of all valid notes within range (based on current valence)
noteset = progs.createChord(chord, majorScale)
n_notes = len(noteset)
midi_low = [x for x in data['range'] if x['valence']==math.floor(valence * 10)/10][0]['midi_low']
midi_high = [x for x in data['range'] if x['valence']==math.floor(valence * 10)/10][0]['midi_high']
range_noteset = player.setRange(midi_low, midi_high, noteset)
print("chord: ", chord[3], "---notes in noteset: ", noteset, "----notes in full range: ", range_noteset)
# initialize memory of previous chord
if prev_noteset==[]:
prev_noteset=noteset
# allocate probabilities of register for each note in chord.
bright = player.setPitch(n_notes, valence)
# determine if scale patterns should be drawn from major or minor scale
if valence<0.4:
scale = player.setRange(midi_low, midi_high, minorScale)
else:
scale = player.setRange(midi_low, midi_high, majorScale)
scale.sort()
# do we want to add in a percussion instrument?
# play bass (root note) -> want to try bassoon? instruments = [piano, clarinet, strings]
current_velocity = np.random.randint(low_loudness,loudness)
note = mido.Message('note_on', channel=1, note=min(noteset) - 12, velocity=current_velocity)
instruments[2].send(note)
# play "accompaniment"/harmony chords
chord_voicing = progs.harmonyVL(prev_noteset, noteset, range_noteset)
print("chord voicing: ", chord_voicing)
for i in range(len(chord_voicing)):
note = mido.Message('note_on',
channel=1,
note=int(chord_voicing[i]+bright[i]*12),
velocity=current_velocity)
instruments[0].send(note)
# update value of prev_noteset
prev_noteset=chord_voicing
# plays "foreground" melody
for beat in range(0,n_subdivisions):
# determine which extensions to sound and create tone
if (activate1[beat] == 1):
note1 = int(noteset[0]+bright[0]*12)
msg = mido.Message('note_on',
channel=1,
note=note1,
velocity=current_velocity)
instruments[0].send(msg)
if (activate2[beat] == 1):
# use melodic motives for voice leading logic
current_motive = player.selectMotive(data['melodic_motives'], current_motive, arousal)
melody_note = player.melodyVL_motives(current_motive, melody_note, noteset, scale)
print('melody note is: ',melody_note)
msg = mido.Message('note_on',
channel=1,
note=melody_note,
velocity=current_velocity+10)
instruments[0].send(msg)
# length of pause determined by tempo.
time.sleep((60/bpm)/(n_subdivisions/4))
# shut all down
instruments[0].reset()
instruments[1].reset()
instruments[2].reset()
instruments[0].reset()
instruments[1].reset()
instruments[2].reset()
#clear_output()
```
Three voices: bass, harmony and melody
* Bass - String ensemble, Harmony and melody - Piano
* Bass - String ensemble, Harmony and melody - Piano, melody - Clarinet (doubling)
* Bass - Clarinet, Harmony and melody - Piano
```
[x for x in zip(chord_voicing, bright*12)]
melody_note
```
# Archive
```
if np.random.rand(1)[0] < arousal:
violin.send(msg)
# write control change (cc) message. Controller number 7 maps to volume.
volume = 80
msg = mido.Message('control_change',channel=ichannel,control=7,value=volume)
piano.send(msg)
# initial idea for melody voice leading - pick closest note
# note how this doesn't depend on arousal or valence at all, basically only controls musicality
def melodyVL_closestNote(melody_note, noteset, range_noteset):
"""
Controls voice leading of melodic line by picking the closest available next note based on previous note
the melody tends to stay around the same register with this implementation
"""
rand_idx = np.random.randint(2,n_notes)
# randomly initialize melody
if melody_note==[]:
melody_note = int(noteset[rand_idx]+bright[rand_idx]*12)
else:
melody_note = min(range_noteset, key=lambda x:abs(x-melody_note))
return melody_note
# initialize params: next_chord, minimal loudness, stadard velocity, current_motive
next_chord = []
low_loudness = 50
default_velocity = 80
current_motive=0
# initialize memory of previous harmony and melody notes (partially determines current harmony/melody notes)
prev_chord, melody_note = [], []
# keep track of current bar
for bar in range(len(input_arousal)):
# set arousal and valence, keep track of current bar in 8-bar progressions
arousal = input_arousal[bar]
valence = input_valence[bar]
bar = bar%8
print("arousal: ", arousal, "---valence: ", valence, "---bar: ", bar)
# set simple params: roughness, voicing, loudness, tempo
roughness = 1-arousal
voicing = valence
loudness = (round(arousal*10))/10*40+60
bpm = 60 + arousal * 100
# first vector (activate1) determines density of background chords
# second vector (activate2) determines density of melody played by piano
# TBC: n_subdivisions should eventually be determined by rhythmic pattern
n_subdivisions = 4
activate1 = player.setRoughness(n_subdivisions, roughness+0.4)
activate2 = player.setRoughness(n_subdivisions, roughness+0.2)
# select chord to be sounded
if next_chord==[]:
# if next chord has not already been determined, then select randomly as usual
chord, next_chord = progs.selectChord(data['progressions'], valence, bar)
else:
chord = next_chord.pop(0)
# generate set of all valid notes within range (based on current valence)
noteset = progs.createChord(chord, majorScale)
n_notes = len(noteset)
midi_low = [x for x in data['range'] if x['valence']==valence][0]['midi_low']
midi_high = [x for x in data['range'] if x['valence']==valence][0]['midi_high']
range_noteset = player.setRange(midi_low, midi_high, noteset)
print("chord: ", chord[3], "---notes in noteset: ", noteset, "----notes in full range: ", range_noteset)
# allocate probabilities of register for each note in chord.
bright = player.setPitch(n_notes, voicing)
# determine if scale patterns should be drawn from major or minor scale
if valence<0.4:
scale = player.setRange(midi_low, midi_high, minorScale)
else:
scale = player.setRange(midi_low, midi_high, majorScale)
scale.sort()
# play "accompaniment"/harmony chords
# TO CHANGE: if all notes in noteset above C4 octave, tranpose whole noteset down an octave.
# Create tone for each note in chord. Serves as the harmony of the generated music
for n in noteset:
note = mido.Message('note_on',
channel=1,
#note=int(noteset[i]+bright[i]*12),
note=n,
velocity=np.random.randint(low_loudness,loudness))
piano.send(note)
# NEW: added in bass (taking lowest value in noteset and transpose down 1-2 octaves)
# this should probably be played by cello, not piano
note = mido.Message('note_on', channel=1, note=min(noteset) - 24, velocity=default_velocity)
piano.send(note)
# plays "foreground" melody [0, 0, 0, 0] [0, 1, 1, 0]
for beat in range(0,n_subdivisions):
# determine which extensions to sound and create tone
#activate1 = player.setRoughness(n_subdivisions, roughness) -> moving this here lets us change subdivision every beat
# alternatively: determine downbeat probability separately.
if (activate1[beat] == 1):
note1 = int(noteset[0]+bright[0]*12)
msg = mido.Message('note_on',
channel=1,
note=note1,
velocity=np.random.randint(low_loudness,loudness))
piano.send(msg)
# add note_off message
if (activate2[beat] == 1):
# use "closest note" voice leading logic
#melody_note = melodyVL_closestNote(melody_note)
# use melodic motives for voice leading logic
current_motive = selectMotive(data['melodic_motives'], current_motive, arousal)
melody_note = melodyVL_motives(current_motive, melody_note, noteset, scale)
print('melody note is: ',melody_note)
msg = mido.Message('note_on',
channel=1,
note=melody_note,
velocity=np.random.randint(low_loudness,loudness))
piano.send(msg)
# length of pause determined by tempo. This formula works when smallest subdivision = eighth notes
time.sleep(0.50/(bpm/60))
#piano.send(mido.Message('note_off', channel=1, note=note1, velocity=64))
#piano.send(mido.Message('note_off', channel=1, note=note2, velocity=64))
# shut all down
# see if you can change the release param
piano.reset()
# generate scale for maximum range of player (C1-C6, MIDI note numbers 24-84)
range_majorScale = player.setRange(24, 84, majorScale)
range_majorScale.sort()
range_minorScale = player.setRange(24, 84, minorScale)
range_minorScale.sort()
range_majorScale.index(60)
temp = [1, 2, 3, 4]
temp[-1]
[x for x in data['melodic_motives'] if x['arousal']=='low' if x['current_motive']=='CT'][0]['motive_weights']
motives = [1, -1, 0, 'CT']
motive_weights=[0.15, 0.15, 0.3, 0.4]
choice(len(motives), 1, p=motive_weights)[0]
def inversion(noteset, inversion):
"""
increases the chord (noteset)'s inversion
"""
noteset.sort()
for i in range(inversion):
while noteset[i] < noteset[-1]:
noteset[i]+=12
return noteset
def decrease_inversion(noteset, inversion):
"""
decreases the chord (noteset)'s inversion
"""
noteset.sort()
for i in range(inversion):
while noteset[-1-i] > noteset[0]:
noteset[-1-i]-=12
return noteset
# implement voice leading logic for bass
temp = 61
print(range_noteset)
# this chooses the closest available note
min(range_noteset, key=lambda x:abs(x-temp))
# I think another possibility is to min. total distance moved for the hamony chords (which is more human)
print(noteset)
setRange(data['range'], 0.1, noteset)
```
| github_jupyter |
### Dropout with L2 Weight Regularization
```
import numpy as np
import keras
from keras.models import Sequential
from matplotlib import pyplot as plt
from keras.layers import Dense,Flatten
from keras.layers import Conv2D, MaxPooling2D,BatchNormalization,Dropout
from keras.utils import np_utils
from sklearn.metrics import confusion_matrix, f1_score, precision_score, recall_score, classification_report
from keras import optimizers,regularizers
class AccuracyHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.acc = []
self.loss = []
self.val_f1s = []
self.val_recalls = []
self.val_precisions = []
def on_epoch_end(self, batch, logs={}):
self.acc.append(logs.get('acc'))
self.loss.append(logs.get('loss'))
X_val, y_val = self.validation_data[0], self.validation_data[1]
y_predict = np.asarray(model.predict(X_val))
y_val = np.argmax(y_val, axis=1)
y_predict = np.argmax(y_predict, axis=1)
self.val_recalls.append(recall_score(y_val, y_predict, average=None))
self.val_precisions.append(precision_score(y_val, y_predict, average=None))
self.val_f1s.append(f1_score(y_val,y_predict, average=None))
data = np.load('/home/aj/assignments/assign2/outfile.npz')
X_train=data["X_train.npy"]
X_test=data["X_test.npy"]
y_train=data["y_train.npy"]
y_test=data["y_test.npy"]
# reshape to be [samples][pixels][width][height]
X_train = X_train.reshape(X_train.shape[0],28, 28,3).astype('float32')
X_test = X_test.reshape(X_test.shape[0],28, 28,3).astype('float32')
# normalize inputs from 0-255 to 0-1
X_train = X_train / 255
X_test = X_test / 255
# one hot encode outputs
# y_train = np_utils.to_categorical(y_train)
# y_test = np_utils.to_categorical(y_test)
print(y_train.shape)
num_classes = y_test.shape[1]
print(num_classes)
input_shape=(28,28,3)
epochs=15
batch_size = 512
history = AccuracyHistory()
def create_deep_model(opt,loss):
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),activation='relu',input_shape=input_shape))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.20))
model.add(Conv2D(64, (3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(128, (3, 3), activation='relu',padding='same'))
model.add(Dropout(0.25))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.25))
model.add(Dense(num_classes, activation='softmax',kernel_regularizer=regularizers.l2(0.01)))
model.compile(optimizer=opt,loss=loss,metrics=['accuracy'])
return model
def create_optimizer(opt_name,lr,decay):
if opt_name == "SGD":
opt = optimizers.SGD(lr=lr, decay=decay)
elif opt_name == "Adam":
opt = optimizers.Adam(lr=lr, decay=decay)
elif opt_name == "RMSprop":
opt = optimizers.RMSprop(lr=lr, decay=decay)
elif opt_name == "Adagrad":
opt = optimizers.Adagrad(lr=lr, decay=decay)
return opt
def create_model(filters,filt1_size,conv_stride,pool_size,pool_stride,opt,loss):
model=Sequential()
model.add(Conv2D(filters, kernel_size=(filt1_size, filt1_size), strides=(conv_stride, conv_stride),activation='relu',input_shape=input_shape))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(pool_size, pool_size), strides=(pool_stride,pool_stride), padding='valid'))
model.add(Flatten())
model.add(Dense(1024,activation='relu'))
model.add(Dense(num_classes, activation='softmax'))
model.compile(optimizer=opt,loss=loss,metrics=['accuracy'])
return model
def fit_model(epochs,batch_size):
model.fit(X_train, y_train,batch_size=batch_size,epochs=epochs,validation_split=0.05,callbacks=[history])
score = model.evaluate(X_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
y_pred = model.predict_classes(X_test)
cnf_mat = confusion_matrix(np.argmax(y_test,axis=1), y_pred)
return cnf_mat,score,y_pred
lr = 0.001
decay = 1e-6
#decay = 0.0
epochs=10
batch_size = 1024
opt = create_optimizer('Adam',lr,decay)
loss = "categorical_crossentropy"
# filters,filt1_size,conv_stride,pool_size,pool_stride = 32,7,1,2,2
model = create_deep_model(opt,loss)
print(model.summary())
cnf_mat,score,y_pred = fit_model(epochs,batch_size)
from keras.models import load_model
model.save('Dropout_model_Line.h5')
fscore=f1_score(np.argmax(y_test,axis=1), y_pred,average=None)
recall=recall_score(np.argmax(y_test,axis=1), y_pred,average=None)
prec=precision_score(np.argmax(y_test,axis=1), y_pred,average=None)
def plot(r1,r2,data,Info):
plt.plot(range(r1,r2),data)
plt.xlabel('Epochs')
plt.ylabel(Info)
plt.show()
plot(1,epochs+1,history.acc,'Accuracy')
plot(1,epochs+1,history.loss,'Loss')
plt.plot(recall,label='Recall')
plt.plot(prec,label='Precision')
plt.xlabel('Class')
plt.ylabel('F-score vs Recall vs Precision')
plt.plot(fscore,label='F-score')
plt.legend()
avg_fscore=np.mean(fscore)
print(avg_fscore)
avg_precision=np.mean(prec)
print(avg_precision)
avg_recall=np.mean(recall)
print(avg_recall)
cnf_mat = confusion_matrix(np.argmax(y_test,axis=1), y_pred)
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
conf = cnf_mat
fig, ax = plt.subplots(figsize=(30,30))
im = ax.imshow(conf,alpha=0.5)
# plt.show()
# We want to show all ticks...
ax.set_xticks(np.arange(cnf_mat.shape[0]))
ax.set_yticks(np.arange(cnf_mat.shape[1]))
# ... and label them with the respective list entries
ax.set_xticklabels(np.arange(0,96))
ax.set_yticklabels(np.arange(0,96))
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
for i in range(cnf_mat.shape[0]):
for j in range(cnf_mat.shape[1]):
text = ax.text(j, i, conf[i, j],
ha="center", va="center",color="black",fontsize=10)
ax.set_title("Confusion matrix",fontsize=20)
fig.tight_layout()
# fig.savefig('plot1_cnf.png')
plt.show()
del model
```
| github_jupyter |
```
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Getting started: Training and prediction with Keras in AI Platform
<img src="https://storage.googleapis.com/cloud-samples-data/ai-platform/census/keras-tensorflow-cmle.png" alt="Keras, TensorFlow, and AI Platform logos" width="300px">
<table align="left">
<td>
<a href="https://cloud.google.com/ml-engine/docs/tensorflow/getting-started-keras">
<img src="https://cloud.google.com/_static/images/cloud/icons/favicons/onecloud/super_cloud.png"
alt="Google Cloud logo" width="32px"> Read on cloud.google.com
</a>
</td>
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/cloudml-samples/blob/main/notebooks/tensorflow/getting-started-keras.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/cloudml-samples/blob/main/notebooks/tensorflow/getting-started-keras.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
## Overview
This tutorial shows how to train a neural network on AI Platform
using the Keras sequential API and how to serve predictions from that
model.
Keras is a high-level API for building and training deep learning models.
[tf.keras](https://www.tensorflow.org/guide/keras) is TensorFlow’s
implementation of this API.
The first two parts of the tutorial walk through training a model on Cloud
AI Platform using prewritten Keras code, deploying the trained model to
AI Platform, and serving online predictions from the deployed model.
The last part of the tutorial digs into the training code used for this model and ensuring it's compatible with AI Platform. To learn more about building
machine learning models in Keras more generally, read [TensorFlow's Keras
tutorials](https://www.tensorflow.org/tutorials/keras).
### Dataset
This tutorial uses the [United States Census Income
Dataset](https://archive.ics.uci.edu/ml/datasets/census+income) provided by the
[UC Irvine Machine Learning
Repository](https://archive.ics.uci.edu/ml/index.php). This dataset contains
information about people from a 1994 Census database, including age, education,
marital status, occupation, and whether they make more than $50,000 a year.
### Objective
The goal is to train a deep neural network (DNN) using Keras that predicts
whether a person makes more than $50,000 a year (target label) based on other
Census information about the person (features).
This tutorial focuses more on using this model with AI Platform than on
the design of the model itself. However, it's always important to think about
potential problems and unintended consequences when building machine learning
systems. See the [Machine Learning Crash Course exercise about
fairness](https://developers.google.com/machine-learning/crash-course/fairness/programming-exercise)
to learn about sources of bias in the Census dataset, as well as machine
learning fairness more generally.
### Costs
This tutorial uses billable components of Google Cloud Platform (GCP):
* AI Platform
* Cloud Storage
Learn about [AI Platform
pricing](https://cloud.google.com/ml-engine/docs/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Before you begin
You must do several things before you can train and deploy a model in
AI Platform:
* Set up your local development environment.
* Set up a GCP project with billing and the necessary
APIs enabled.
* Authenticate your GCP account in this notebook.
* Create a Cloud Storage bucket to store your training package and your
trained model.
### Set up your local development environment
**If you are using Colab or AI Platform Notebooks**, your environment already meets
all the requirements to run this notebook. You can skip this step.
**Otherwise**, make sure your environment meets this notebook's requirements.
You need the following:
* The Google Cloud SDK
* Git
* Python 3
* virtualenv
* Jupyter notebook running in a virtual environment with Python 3
The Google Cloud guide to [Setting up a Python development
environment](https://cloud.google.com/python/setup) and the [Jupyter
installation guide](https://jupyter.org/install) provide detailed instructions
for meeting these requirements. The following steps provide a condensed set of
instructions:
1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
2. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
3. [Install
virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
and create a virtual environment that uses Python 3.
4. Activate that environment and run `pip install jupyter` in a shell to install
Jupyter.
5. Run `jupyter notebook` in a shell to launch Jupyter.
6. Open this notebook in the Jupyter Notebook Dashboard.
### Set up your GCP project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a GCP project.](https://console.cloud.google.com/cloud-resource-manager)
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the AI Platform ("Cloud Machine Learning Engine") and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "<your-project-id>" #@param {type:"string"}
! gcloud config set project $PROJECT_ID
```
### Authenticate your GCP account
**If you are using AI Platform Notebooks**, your environment is already
authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions
when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
1. In the GCP Console, go to the [**Create service account key**
page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
2. From the **Service account** drop-down list, select **New service account**.
3. In the **Service account name** field, enter a name.
4. From the **Role** drop-down list, select
**Machine Learning Engine > AI Platform Admin** and
**Storage > Storage Object Admin**.
5. Click *Create*. A JSON file that contains your key downloads to your
local environment.
6. Enter the path to your service account key as the
`GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
```
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
if 'google.colab' in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
else:
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you submit a training job using the Cloud SDK, you upload a Python package
containing your training code to a Cloud Storage bucket. AI Platform runs
the code from this package. In this tutorial, AI Platform also saves the
trained model that results from your job in the same bucket. You can then
create an AI Platform model version based on this output in order to serve
online predictions.
Set the name of your Cloud Storage bucket below. It must be unique across all
Cloud Storage buckets.
You may also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Make sure to [choose a region where Cloud
AI Platform services are
available](https://cloud.google.com/ml-engine/docs/tensorflow/regions).
```
BUCKET_NAME = "<your-bucket-name>" #@param {type:"string"}
REGION = "us-central1" #@param {type:"string"}
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION gs://$BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al gs://$BUCKET_NAME
```
## Part 1. Quickstart for training in AI Platform
This section of the tutorial walks you through submitting a training job to Cloud
AI Platform. This job runs sample code that uses Keras to train a deep neural
network on the United States Census data. It outputs the trained model as a
[TensorFlow SavedModel
directory](https://www.tensorflow.org/guide/saved_model#save_and_restore_models)
in your Cloud Storage bucket.
### Get training code and dependencies
First, download the training code and change the notebook's working directory:
```
# Clone the repository of AI Platform samples
! git clone --depth 1 https://github.com/GoogleCloudPlatform/cloudml-samples
# Set the working directory to the sample code directory
%cd cloudml-samples/census/tf-keras
```
Notice that the training code is structured as a Python package in the
`trainer/` subdirectory:
```
# `ls` shows the working directory's contents. The `p` flag adds trailing
# slashes to subdirectory names. The `R` flag lists subdirectories recursively.
! ls -pR
```
Run the following cell to install Python dependencies needed to train the model locally. When you run the training job in AI Platform,
dependencies are preinstalled based on the [runtime
version](https://cloud.google.com/ml-engine/docs/tensorflow/runtime-version-list)
you choose.
```
! pip install -r requirements.txt
```
### Train your model locally
Before training on AI Platform, train the job locally to verify the file
structure and packaging is correct.
For a complex or resource-intensive job, you
may want to train locally on a small sample of your dataset to verify your code.
Then you can run the job on AI Platform to train on the whole dataset.
This sample runs a relatively quick job on a small dataset, so the local
training and the AI Platform job run the same code on the same data.
Run the following cell to train a model locally:
```
# Explicitly tell `gcloud ai-platform local train` to use Python 3
! gcloud config set ml_engine/local_python $(which python3)
# This is similar to `python -m trainer.task --job-dir local-training-output`
# but it better replicates the AI Platform environment, especially for
# distributed training (not applicable here).
! gcloud ai-platform local train \
--package-path trainer \
--module-name trainer.task \
--job-dir local-training-output
```
### Train your model using AI Platform
Next, submit a training job to AI Platform. This runs the training module
in the cloud and exports the trained model to Cloud Storage.
First, give your training job a name and choose a directory within your Cloud
Storage bucket for saving intermediate and output files:
```
JOB_NAME = 'my_first_keras_job'
JOB_DIR = 'gs://' + BUCKET_NAME + '/keras-job-dir'
```
Run the following command to package the `trainer/` directory, upload it to the
specified `--job-dir`, and instruct AI Platform to run the
`trainer.task` module from that package.
The `--stream-logs` flag lets you view training logs in the cell below. You can
also see logs and other job details in the GCP Console.
### Hyperparameter tuning
You can optionally perform hyperparameter tuning by using the included
`hptuning_config.yaml` configuration file. This file tells AI Platform to tune the batch size and learning rate for training over multiple trials to maximize accuracy.
In this example, the training code uses a [TensorBoard
callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard),
which [creates TensorFlow `Summary`
`Event`s](https://www.tensorflow.org/api_docs/python/tf/summary/FileWriter#add_summary)
during training. AI Platform uses these events to track the metric you want to
optimize. Learn more about [hyperparameter tuning in
AI Platform Training](https://cloud.google.com/ml-engine/docs/tensorflow/hyperparameter-tuning-overview).
```
! gcloud ai-platform jobs submit training $JOB_NAME \
--package-path trainer/ \
--module-name trainer.task \
--region $REGION \
--python-version 3.7 \
--runtime-version 1.15 \
--job-dir $JOB_DIR \
--stream-logs
```
## Part 2. Quickstart for online predictions in AI Platform
This section shows how to use AI Platform and your trained model from Part 1
to predict a person's income bracket from other Census information about them.
### Create model and version resources in AI Platform
To serve online predictions using the model you trained and exported in Part 1,
create a *model* resource in AI Platform and a *version* resource
within it. The version resource is what actually uses your trained model to
serve predictions. This structure lets you adjust and retrain your model many times and
organize all the versions together in AI Platform. Learn more about [models
and
versions](https://cloud.google.com/ai-platform/prediction/docs/projects-models-versions-jobs).
While you specify `--region $REGION` in gcloud commands, you will use regional endpoint. You can also specify `--region global` to use global endpoint. Please note that you must create versions using the same endpoint as the one you use to create the model. Learn more about available [regional endpoints](https://cloud.google.com/ai-platform/prediction/docs/regional-endpoints).
First, name and create the model resource:
```
MODEL_NAME = "my_first_keras_model"
! gcloud ai-platform models create $MODEL_NAME \
--region $REGION
```
Next, create the model version. The training job from Part 1 exported a timestamped
[TensorFlow SavedModel
directory](https://www.tensorflow.org/guide/saved_model#structure_of_a_savedmodel_directory)
to your Cloud Storage bucket. AI Platform uses this directory to create a
model version. Learn more about [SavedModel and
AI Platform](https://cloud.google.com/ml-engine/docs/tensorflow/deploying-models).
You may be able to find the path to this directory in your training job's logs.
Look for a line like:
```
Model exported to: gs://<your-bucket-name>/keras-job-dir/keras_export/1545439782
```
Execute the following command to identify your SavedModel directory and use it to create a model version resource:
```
MODEL_VERSION = "v1"
# Get a list of directories in the `keras_export` parent directory
KERAS_EXPORT_DIRS = ! gsutil ls $JOB_DIR/keras_export/
# Update the directory as needed, in case you've trained
# multiple times
SAVED_MODEL_PATH = keras_export
# Create model version based on that SavedModel directory
! gcloud ai-platform versions create $MODEL_VERSION \
--region $REGION \
--model $MODEL_NAME \
--runtime-version 1.15 \
--python-version 3.7 \
--framework tensorflow \
--origin $SAVED_MODEL_PATH
```
### Prepare input for prediction
To receive valid and useful predictions, you must preprocess input for prediction in the same way that training data was preprocessed. In a production
system, you may want to create a preprocessing pipeline that can be used identically at training time and prediction time.
For this exercise, use the training package's data-loading code to select a random sample from the evaluation data. This data is in the form that was used to evaluate accuracy after each epoch of training, so it can be used to send test predictions without further preprocessing:
```
from trainer import util
_, _, eval_x, eval_y = util.load_data()
prediction_input = eval_x.sample(20)
prediction_targets = eval_y[prediction_input.index]
prediction_input
```
Notice that categorical fields, like `occupation`, have already been converted to integers (with the same mapping that was used for training). Numerical fields, like `age`, have been scaled to a
[z-score](https://developers.google.com/machine-learning/crash-course/representation/cleaning-data). Some fields have been dropped from the original
data. Compare the prediction input with the raw data for the same examples:
```
import pandas as pd
_, eval_file_path = util.download(util.DATA_DIR)
raw_eval_data = pd.read_csv(eval_file_path,
names=util._CSV_COLUMNS,
na_values='?')
raw_eval_data.iloc[prediction_input.index]
```
Export the prediction input to a newline-delimited JSON file:
```
import json
with open('prediction_input.json', 'w') as json_file:
for row in prediction_input.values.tolist():
json.dump(row, json_file)
json_file.write('\n')
! cat prediction_input.json
```
The `gcloud` command-line tool accepts newline-delimited JSON for online
prediction, and this particular Keras model expects a flat list of
numbers for each input example.
AI Platform requires a different format when you make online prediction requests to the REST API without using the `gcloud` tool. The way you structure
your model may also change how you must format data for prediction. Learn more
about [formatting data for online
prediction](https://cloud.google.com/ml-engine/docs/tensorflow/prediction-overview#prediction_input_data).
### Submit the online prediction request
Use `gcloud` to submit your online prediction request.
```
! gcloud ai-platform predict \
--region $REGION \
--model $MODEL_NAME \
--version $MODEL_VERSION \
--json-instances prediction_input.json
```
Since the model's last layer uses a [sigmoid function](https://developers.google.com/machine-learning/glossary/#sigmoid_function) for its activation, outputs between 0 and 0.5 represent negative predictions ("<=50K") and outputs between 0.5 and 1 represent positive ones (">50K").
Do the predicted income brackets match the actual ones? Run the following cell
to see the true labels.
```
prediction_targets
```
## Part 3. Developing the Keras model from scratch
At this point, you have trained a machine learning model on AI Platform, deployed the trained model as a version resource on AI Platform, and received online predictions from the deployment. The next section walks through recreating the Keras code used to train your model. It covers the following parts of developing a machine learning model for use with AI Platform:
* Downloading and preprocessing data
* Designing and training the model
* Visualizing training and exporting the trained model
While this section provides more detailed insight to the tasks completed in previous parts, to learn more about using `tf.keras`, read [TensorFlow's guide to Keras](https://www.tensorflow.org/tutorials/keras). To learn more about structuring code as a training packge for AI Platform, read [Packaging a training application](https://cloud.google.com/ml-engine/docs/tensorflow/packaging-trainer) and reference the [complete training code](https://github.com/GoogleCloudPlatform/cloudml-samples/tree/master/census/tf-keras), which is structured as a Python package.
### Import libraries and define constants
First, import Python libraries required for training:
```
import os
from six.moves import urllib
import tempfile
import numpy as np
import pandas as pd
import tensorflow as tf
# Examine software versions
print(__import__('sys').version)
print(tf.__version__)
print(tf.keras.__version__)
```
Then, define some useful constants:
* Information for downloading training and evaluation data
* Information required for Pandas to interpret the data and convert categorical fields into numeric features
* Hyperparameters for training, such as learning rate and batch size
```
### For downloading data ###
# Storage directory
DATA_DIR = os.path.join(tempfile.gettempdir(), 'census_data')
# Download options.
DATA_URL = 'https://storage.googleapis.com/cloud-samples-data/ai-platform' \
'/census/data'
TRAINING_FILE = 'adult.data.csv'
EVAL_FILE = 'adult.test.csv'
TRAINING_URL = '%s/%s' % (DATA_URL, TRAINING_FILE)
EVAL_URL = '%s/%s' % (DATA_URL, EVAL_FILE)
### For interpreting data ###
# These are the features in the dataset.
# Dataset information: https://archive.ics.uci.edu/ml/datasets/census+income
_CSV_COLUMNS = [
'age', 'workclass', 'fnlwgt', 'education', 'education_num',
'marital_status', 'occupation', 'relationship', 'race', 'gender',
'capital_gain', 'capital_loss', 'hours_per_week', 'native_country',
'income_bracket'
]
_CATEGORICAL_TYPES = {
'workclass': pd.api.types.CategoricalDtype(categories=[
'Federal-gov', 'Local-gov', 'Never-worked', 'Private', 'Self-emp-inc',
'Self-emp-not-inc', 'State-gov', 'Without-pay'
]),
'marital_status': pd.api.types.CategoricalDtype(categories=[
'Divorced', 'Married-AF-spouse', 'Married-civ-spouse',
'Married-spouse-absent', 'Never-married', 'Separated', 'Widowed'
]),
'occupation': pd.api.types.CategoricalDtype([
'Adm-clerical', 'Armed-Forces', 'Craft-repair', 'Exec-managerial',
'Farming-fishing', 'Handlers-cleaners', 'Machine-op-inspct',
'Other-service', 'Priv-house-serv', 'Prof-specialty', 'Protective-serv',
'Sales', 'Tech-support', 'Transport-moving'
]),
'relationship': pd.api.types.CategoricalDtype(categories=[
'Husband', 'Not-in-family', 'Other-relative', 'Own-child', 'Unmarried',
'Wife'
]),
'race': pd.api.types.CategoricalDtype(categories=[
'Amer-Indian-Eskimo', 'Asian-Pac-Islander', 'Black', 'Other', 'White'
]),
'native_country': pd.api.types.CategoricalDtype(categories=[
'Cambodia', 'Canada', 'China', 'Columbia', 'Cuba', 'Dominican-Republic',
'Ecuador', 'El-Salvador', 'England', 'France', 'Germany', 'Greece',
'Guatemala', 'Haiti', 'Holand-Netherlands', 'Honduras', 'Hong', 'Hungary',
'India', 'Iran', 'Ireland', 'Italy', 'Jamaica', 'Japan', 'Laos', 'Mexico',
'Nicaragua', 'Outlying-US(Guam-USVI-etc)', 'Peru', 'Philippines', 'Poland',
'Portugal', 'Puerto-Rico', 'Scotland', 'South', 'Taiwan', 'Thailand',
'Trinadad&Tobago', 'United-States', 'Vietnam', 'Yugoslavia'
]),
'income_bracket': pd.api.types.CategoricalDtype(categories=[
'<=50K', '>50K'
])
}
# This is the label (target) we want to predict.
_LABEL_COLUMN = 'income_bracket'
### Hyperparameters for training ###
# This the training batch size
BATCH_SIZE = 128
# This is the number of epochs (passes over the full training data)
NUM_EPOCHS = 20
# Define learning rate.
LEARNING_RATE = .01
```
### Download and preprocess data
#### Download the data
Next, define functions to download training and evaluation data. These functions also fix minor irregularities in the data's formatting.
```
def _download_and_clean_file(filename, url):
"""Downloads data from url, and makes changes to match the CSV format.
The CSVs may use spaces after the comma delimters (non-standard) or include
rows which do not represent well-formed examples. This function strips out
some of these problems.
Args:
filename: filename to save url to
url: URL of resource to download
"""
temp_file, _ = urllib.request.urlretrieve(url)
with tf.io.gfile.GFile(temp_file, 'r') as temp_file_object:
with tf.io.gfile.GFile(filename, 'w') as file_object:
for line in temp_file_object:
line = line.strip()
line = line.replace(', ', ',')
if not line or ',' not in line:
continue
if line[-1] == '.':
line = line[:-1]
line += '\n'
file_object.write(line)
tf.io.gfile.remove(temp_file)
def download(data_dir):
"""Downloads census data if it is not already present.
Args:
data_dir: directory where we will access/save the census data
"""
tf.io.gfile.makedirs(data_dir)
training_file_path = os.path.join(data_dir, TRAINING_FILE)
if not tf.io.gfile.exists(training_file_path):
_download_and_clean_file(training_file_path, TRAINING_URL)
eval_file_path = os.path.join(data_dir, EVAL_FILE)
if not tf.io.gfile.exists(eval_file_path):
_download_and_clean_file(eval_file_path, EVAL_URL)
return training_file_path, eval_file_path
```
Use those functions to download the data for training and verify that you have CSV files for training and evaluation:
```
training_file_path, eval_file_path = download(DATA_DIR)
# You should see 2 files: adult.data.csv and adult.test.csv
!ls -l $DATA_DIR
```
Next, load these files using Pandas and examine the data:
```
# This census data uses the value '?' for fields (column) that are missing data.
# We use na_values to find ? and set it to NaN values.
# https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html
train_df = pd.read_csv(training_file_path, names=_CSV_COLUMNS, na_values='?')
eval_df = pd.read_csv(eval_file_path, names=_CSV_COLUMNS, na_values='?')
# Here's what the data looks like before we preprocess the data.
train_df.head()
```
#### Preprocess the data
The first preprocessing step removes certain features from the data and
converts categorical features to numerical values for use with Keras.
Learn more about [feature engineering](https://developers.google.com/machine-learning/crash-course/representation/feature-engineering) and [bias in data](https://developers.google.com/machine-learning/crash-course/fairness/types-of-bias).
```
UNUSED_COLUMNS = ['fnlwgt', 'education', 'gender']
def preprocess(dataframe):
"""Converts categorical features to numeric. Removes unused columns.
Args:
dataframe: Pandas dataframe with raw data
Returns:
Dataframe with preprocessed data
"""
dataframe = dataframe.drop(columns=UNUSED_COLUMNS)
# Convert integer valued (numeric) columns to floating point
numeric_columns = dataframe.select_dtypes(['int64']).columns
dataframe[numeric_columns] = dataframe[numeric_columns].astype('float32')
# Convert categorical columns to numeric
cat_columns = dataframe.select_dtypes(['object']).columns
dataframe[cat_columns] = dataframe[cat_columns].apply(lambda x: x.astype(
_CATEGORICAL_TYPES[x.name]))
dataframe[cat_columns] = dataframe[cat_columns].apply(lambda x: x.cat.codes)
return dataframe
prepped_train_df = preprocess(train_df)
prepped_eval_df = preprocess(eval_df)
```
Run the following cell to see how preprocessing changed the data. Notice in particular that `income_bracket`, the label that you're training the model to predict, has changed from `<=50K` and `>50K` to `0` and `1`:
```
prepped_train_df.head()
```
Next, separate the data into features ("x") and labels ("y"), and reshape the label arrays into a format for use with `tf.data.Dataset` later:
```
# Split train and test data with labels.
# The pop() method will extract (copy) and remove the label column from the dataframe
train_x, train_y = prepped_train_df, prepped_train_df.pop(_LABEL_COLUMN)
eval_x, eval_y = prepped_eval_df, prepped_eval_df.pop(_LABEL_COLUMN)
# Reshape label columns for use with tf.data.Dataset
train_y = np.asarray(train_y).astype('float32').reshape((-1, 1))
eval_y = np.asarray(eval_y).astype('float32').reshape((-1, 1))
```
Scaling training data so each numerical feature column has a mean of 0 and a standard deviation of 1 [can improve your model](https://developers.google.com/machine-learning/crash-course/representation/cleaning-data).
In a production system, you may want to save the means and standard deviations from your training set and use them to perform an identical transformation on test data at prediction time. For convenience in this exercise, temporarily combine the training and evaluation data to scale all of them:
```
def standardize(dataframe):
"""Scales numerical columns using their means and standard deviation to get
z-scores: the mean of each numerical column becomes 0, and the standard
deviation becomes 1. This can help the model converge during training.
Args:
dataframe: Pandas dataframe
Returns:
Input dataframe with the numerical columns scaled to z-scores
"""
dtypes = list(zip(dataframe.dtypes.index, map(str, dataframe.dtypes)))
# Normalize numeric columns.
for column, dtype in dtypes:
if dtype == 'float32':
dataframe[column] -= dataframe[column].mean()
dataframe[column] /= dataframe[column].std()
return dataframe
# Join train_x and eval_x to normalize on overall means and standard
# deviations. Then separate them again.
all_x = pd.concat([train_x, eval_x], keys=['train', 'eval'])
all_x = standardize(all_x)
train_x, eval_x = all_x.xs('train'), all_x.xs('eval')
```
Finally, examine some of your fully preprocessed training data:
```
# Verify dataset features
# Note how only the numeric fields (not categorical) have been standardized
train_x.head()
```
### Design and train the model
#### Create training and validation datasets
Create an input function to convert features and labels into a
[`tf.data.Dataset`](https://www.tensorflow.org/guide/datasets) for training or evaluation:
```
def input_fn(features, labels, shuffle, num_epochs, batch_size):
"""Generates an input function to be used for model training.
Args:
features: numpy array of features used for training or inference
labels: numpy array of labels for each example
shuffle: boolean for whether to shuffle the data or not (set True for
training, False for evaluation)
num_epochs: number of epochs to provide the data for
batch_size: batch size for training
Returns:
A tf.data.Dataset that can provide data to the Keras model for training or
evaluation
"""
if labels is None:
inputs = features
else:
inputs = (features, labels)
dataset = tf.data.Dataset.from_tensor_slices(inputs)
if shuffle:
dataset = dataset.shuffle(buffer_size=len(features))
# We call repeat after shuffling, rather than before, to prevent separate
# epochs from blending together.
dataset = dataset.repeat(num_epochs)
dataset = dataset.batch(batch_size)
return dataset
```
Next, create these training and evaluation datasets.Use the `NUM_EPOCHS`
and `BATCH_SIZE` hyperparameters defined previously to define how the training
dataset provides examples to the model during training. Set up the validation
dataset to provide all its examples in one batch, for a single validation step
at the end of each training epoch.
```
# Pass a numpy array by using DataFrame.values
training_dataset = input_fn(features=train_x.values,
labels=train_y,
shuffle=True,
num_epochs=NUM_EPOCHS,
batch_size=BATCH_SIZE)
num_eval_examples = eval_x.shape[0]
# Pass a numpy array by using DataFrame.values
validation_dataset = input_fn(features=eval_x.values,
labels=eval_y,
shuffle=False,
num_epochs=NUM_EPOCHS,
batch_size=num_eval_examples)
```
#### Design a Keras Model
Design your neural network using the [Keras Sequential API](https://www.tensorflow.org/guide/keras#sequential_model).
This deep neural network (DNN) has several hidden layers, and the last layer uses a sigmoid activation function to output a value between 0 and 1:
* The input layer has 100 units using the ReLU activation function.
* The hidden layer has 75 units using the ReLU activation function.
* The hidden layer has 50 units using the ReLU activation function.
* The hidden layer has 25 units using the ReLU activation function.
* The output layer has 1 units using a sigmoid activation function.
* The optimizer uses the binary cross-entropy loss function, which is appropriate for a binary classification problem like this one.
Feel free to change these layers to try to improve the model:
```
def create_keras_model(input_dim, learning_rate):
"""Creates Keras Model for Binary Classification.
Args:
input_dim: How many features the input has
learning_rate: Learning rate for training
Returns:
The compiled Keras model (still needs to be trained)
"""
Dense = tf.keras.layers.Dense
model = tf.keras.Sequential(
[
Dense(100, activation=tf.nn.relu, kernel_initializer='uniform',
input_shape=(input_dim,)),
Dense(75, activation=tf.nn.relu),
Dense(50, activation=tf.nn.relu),
Dense(25, activation=tf.nn.relu),
Dense(1, activation=tf.nn.sigmoid)
])
# Custom Optimizer:
# https://www.tensorflow.org/api_docs/python/tf/train/RMSPropOptimizer
optimizer = tf.keras.optimizers.RMSprop(
lr=learning_rate)
# Compile Keras model
model.compile(
loss='binary_crossentropy', optimizer=optimizer, metrics=['accuracy'])
return model
```
Next, create the Keras model object and examine its structure:
```
num_train_examples, input_dim = train_x.shape
print('Number of features: {}'.format(input_dim))
print('Number of examples: {}'.format(num_train_examples))
keras_model = create_keras_model(
input_dim=input_dim,
learning_rate=LEARNING_RATE)
# Take a detailed look inside the model
keras_model.summary()
```
#### Train and evaluate the model
Define a learning rate decay to encourage model paramaters to make smaller
changes as training goes on:
```
# Setup Learning Rate decay.
lr_decay_cb = tf.keras.callbacks.LearningRateScheduler(
lambda epoch: LEARNING_RATE + 0.02 * (0.5 ** (1 + epoch)),
verbose=True)
# Setup TensorBoard callback.
tensorboard_cb = tf.keras.callbacks.TensorBoard(
os.path.join(JOB_DIR, 'keras_tensorboard'),
histogram_freq=1)
```
Finally, train the model. Provide the appropriate `steps_per_epoch` for the
model to train on the entire training dataset (with `BATCH_SIZE` examples per step) during each epoch. And instruct the model to calculate validation
accuracy with one big validation batch at the end of each epoch.
```
history = keras_model.fit(training_dataset,
epochs=NUM_EPOCHS,
steps_per_epoch=int(num_train_examples/BATCH_SIZE),
validation_data=validation_dataset,
validation_steps=1,
callbacks=[lr_decay_cb, tensorboard_cb],
verbose=1)
```
### Visualize training and export the trained model
#### Visualize training
Import `matplotlib` to visualize how the model learned over the training period.
```
! pip install matplotlib
from matplotlib import pyplot as plt
%matplotlib inline
```
Plot the model's loss (binary cross-entropy) and accuracy, as measured at the
end of each training epoch:
```
# Visualize History for Loss.
plt.title('Keras model loss')
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['training', 'validation'], loc='upper right')
plt.show()
# Visualize History for Accuracy.
plt.title('Keras model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.legend(['training', 'validation'], loc='lower right')
plt.show()
```
Over time, loss decreases and accuracy increases. But do they converge to a
stable level? Are there big differences between the training and validation
metrics (a sign of overfitting)?
Learn about [how to improve your machine learning
model](https://developers.google.com/machine-learning/crash-course/). Then, feel
free to adjust hyperparameters or the model architecture and train again.
#### Export the model for serving
AI Platform requires when you [create a model version
resource](https://cloud.google.com/ml-engine/docs/tensorflow/deploying-models#create_a_model_version).
Since not all optimizers can be exported to the SavedModel format, you may see
warnings during the export process. As long you successfully export a serving
graph, AI Platform can used the SavedModel to serve predictions.
```
# Export the model to a local SavedModel directory
export_path = tf.keras.experimental.export_saved_model(keras_model, 'keras_export')
print("Model exported to: ", export_path)
```
You may export a SavedModel directory to your local filesystem or to Cloud
Storage, as long as you have the necessary permissions. In your current
environment, you granted access to Cloud Storage by authenticating your GCP account and setting the `GOOGLE_APPLICATION_CREDENTIALS` environment variable.
AI Platform training jobs can also export directly to Cloud Storage, because
AI Platform service accounts [have access to Cloud Storage buckets in their own
project](https://cloud.google.com/ml-engine/docs/tensorflow/working-with-cloud-storage).
Try exporting directly to Cloud Storage:
```
# Export the model to a SavedModel directory in Cloud Storage
export_path = tf.keras.experimental.export_saved_model(keras_model, JOB_DIR + '/keras_export')
print("Model exported to: ", export_path)
```
You can now deploy this model to AI Platform and serve predictions by
following the steps from Part 2.
## Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Alternatively, you can clean up individual resources by running the following
commands:
```
# Delete model version resource
! gcloud ai-platform versions delete $MODEL_VERSION --region $REGION --quiet --model $MODEL_NAME
# Delete model resource
! gcloud ai-platform models delete $MODEL_NAME --region $REGION --quiet
# Delete Cloud Storage objects that were created
! gsutil -m rm -r $JOB_DIR
# If the training job is still running, cancel it
! gcloud ai-platform jobs cancel $JOB_NAME --quiet --verbosity critical
```
If your Cloud Storage bucket doesn't contain any other objects and you would like to delete it, run `gsutil rm -r gs://$BUCKET_NAME`.
## What's next?
* View the [complete training
code](https://github.com/GoogleCloudPlatform/cloudml-samples/tree/master/census/tf-keras) used in this guide, which structures the code to accept custom
hyperparameters as command-line flags.
* Read about [packaging
code](https://cloud.google.com/ml-engine/docs/tensorflow/packaging-trainer) for an AI Platform training job.
* Read about [deploying a
model](https://cloud.google.com/ml-engine/docs/tensorflow/deploying-models) to serve predictions.
| github_jupyter |
# 652 Week 4 Tips and Tricks #
Jamie White
Revision: [ F2021 20 September 2021 ]
## Before starting your assignments... ##
... make a clean copy of your notebook in case you have to start over.
In Coursera:
`File > Make a copy...`
## Remember to work through the Week 4 networkX tutorial ##
## <font color='red'>Warning! </font>
<font color='red' size=2>Please **AVOID** using `community` and `modularity` as your variable names. These are imported as preserved names for networkx submodules. Changing their representations would result in autograder failuers.</font>
## <font color='green'>Advice </font>
<font color='Green' size=2>You do not need to run the notebook Girvan-Newman.ipynb</font>
There is a pickle file with the output in assets. Just load it as instructed.
## Load some libraries to get started... ##
```
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
from networkx.drawing.nx_pydot import graphviz_layout
import ndlib.models.ModelConfig as mc
import ndlib.models.epidemics as ep
import operator
import random
import json
# %matplotlib inline
```
## How are communities represented in networkX? ##
In Assignment 4, a community (singular) is a set of node ID's. The communities (plural) are contained in a list. Thus,
```
# each community is a set of node ID's. sample_communities is a list of communities
sample_communities = [
{512, 513, 514, 515, 516, 517},
{96, 97, 92, 93, 94, 95},
{288, 289, 290, 285, 286, 287},
{0, 1, 2, 3, 4, 5}
]
type(sample_communities)
a_community = sample_communities[0]
a_community
type(a_community)
another_community = sample_communities[1]
another_community
type(another_community)
```
## Communities in the Map of Science network ##
```
G = nx.read_gml('assets/MapOfScience.gml', label='id')
```
### Classes and communities ###
*If the different domains of science are communities, what are they?*
Below is a sample of the data for each node (here, nodes 0 and 1). Each node is represented as a tuple with a node label and a dictionary `(0,{...})`.
In the dictionary for each node, the value for the 'Class' key indicates a domain of science. For Nodes 0 and 1, the domain of science is 'Applied', so they are part of the 'Applied' network community. The 'Class' of the community to which Nodes 0 and 1 belong is also 'Applied.'
```
# recall from the assignement
list(G.nodes(data=True))[0:2]
# what are the classes?
list_of_communities = set([d['Class'] for n,d in G.nodes(data=True)]) # set of unique community labels
list_of_communities
# classes are domains of science, let's make a list
domains = list(list_of_communities)
len(domains)
# let's grab 2 domains (communities)
domains = list(list_of_communities)[:2]
len(domains)
domains
# find the nodes in those domains (communities)
communities = [set([n for n,d in list(G.nodes(data=True)) if (d['Class']) == domain]) for domain in domains]
# recall from above that communties are
a_community = communities[0]
# let's have a look
type(communities), len(communities), type(a_community), len(a_community), list(a_community)[:10]
```
## Partitions ###
Hint: Based on the Week 4 networkX tutorial.
```
sizes = [50, 40, 30]
probs = [[0.25, 0.04, 0.03], [0.04, 0.3, 0.05], [0.03, 0.05, 0.40]]
G = nx.stochastic_block_model(sizes, probs, seed=0)
pos = nx.spring_layout(G)
plt.figure(figsize=(5,5))
nx.draw_networkx(G, pos=pos, with_labels=False,
node_color='blue', node_size=10,
edge_color='#eeeeee', width=1)
```
### What is this partition thing? ###
The partitioning algorithms return an iterator over tuples of sets of nodes in `G`. Each set of nodes is a community, each tuple is a sequence of communities at a particular level of the algorithm.
☞ "at a particular level of the algorithm" means the number of communities it finds.
```
# uncomment the line below to have a look
# nx.algorithms.community.centrality.girvan_newman?
# use girvan_newman to generate partitions
partitions = nx.algorithms.community.centrality.girvan_newman(G)
# finding all the ways the algorithm split the graph
# partitions is a generator, so it is consumed when you run this cell.
partition_splits = list(partitions)
# How many possible ways did the algorithm split the graph into communities?
len(partition_splits)
# the partitions generator is consumed, so...
empty_generator = list(partitions)
# the list is not generated
len(empty_generator)
# find out how many communities were in the first split?
first_split = partition_splits[0]
len(first_split)
# recall each community is a set of nodes.
# Convert the set to a list to look at 10 nodes
list(first_split[0])[:10]
# how many nodes were in the first community of the first split?
first_split_community_1 = first_split[0]
len(first_split_community_1)
# how many nodes were in the second community of the second split?
first_split_community_2 = first_split[1]
len(first_split_community_2)
# testing if node any nodes in community_1 are in community_2
first_split_community_1 in first_split_community_2
# let's look at the second split
second_split = partition_splits[1]
# it looks like there are three communities in the second split
len(second_split)
# let's assign the communities to variables
second_split_community_1 = second_split[0]
second_split_community_2 = second_split[1]
second_split_community_3 = second_split[2]
# and see how big they are
len(second_split_community_1),len(second_split_community_2),len(second_split_community_3)
```
### Testing or setting node properties based on community membership ###
The tutorial sets color for a plot by community, but you could set other properites, or you could test if the node meets some condition.
It might be useful to break down the steps for 3 communities
```
# pick some colors
color_list = ['green', 'red', 'blue', 'black']
# list of nodes without colors
node_color_list = [i for i in G.nodes]
# iterate through the nodes. i is a counter, n is the node ID
for i, n in enumerate(node_color_list):
# if node is in the first community from the second split,
# color it green
if n in second_split_community_1:
node_color_list[i] = color_list[0]
# if node is in the second community from the second split
# color it red
elif n in second_split_community_2:
node_color_list[i] = color_list[1]
# if node is in the third community from the second split
# color it red
elif n in second_split_community_3:
node_color_list[i] = color_list[2]
# otherwise color it grey
else:
node_color_list[i] = color_list[3]
#see how we did
plt.figure(figsize=(5,5))
nx.draw_networkx(G, pos=pos, with_labels=False,
node_color=node_color_list, node_size=10,
edge_color='#eeeeee', width=1)
```
#### Conclusion ####
Because there are few black nodes (I don't see any), and the three communities look separated, the second partition probably did a good job.
The different metrics in Assignment 4 will quantitate the separation:
```
mod = nx.community.modularity( G, nodes_by_domain ) # modularity
cov = nx.community.coverage( G, nodes_by_domain ) # coverage
perf = nx.community.performance(G, nodes_by_domain ) # performance
den = avg_measure(G, nodes_by_domain, density_one_community ) # density
sep = avg_measure(G, nodes_by_domain, separability_one_community ) # separability
```
## Good luck on Assignment 4! ##
| github_jupyter |
```
con <- url("http://www2.math.su.se/~esbj/GLMbook/moppe.sas")
data <- readLines(con, n = 200L, warn = FALSE, encoding = "unknown")
close(con)
data.start <- grep("^cards;", data) + 1L
data.end <- grep("^;", data[data.start:999L]) + data.start - 2L
table.1.2 <- read.table(text = data[data.start:data.end],
header = FALSE,
sep = "",
quote = "",
col.names = c("premiekl", "moptva", "zon", "dur",
"medskad", "antskad", "riskpre", "helpre", "cell"),
na.strings = NULL,
colClasses = c(rep("factor", 3), "numeric",
rep("integer", 4), "NULL"),
comment.char = "")
rm(con, data, data.start, data.end)
comment(table.1.2) <-
c("Title: Partial casco moped insurance from Wasa insurance, 1994--1999",
"Source: http://www2.math.su.se/~esbj/GLMbook/moppe.sas",
"Copyright: http://www2.math.su.se/~esbj/GLMbook/")
table.1.2$skadfre = with(table.1.2, antskad / dur)
comment(table.1.2$premiekl) <-
c("Name: Class",
"Code: 1=Weight over 60kg and more than 2 gears",
"Code: 2=Other")
comment(table.1.2$moptva) <-
c("Name: Age",
"Code: 1=At most 1 year",
"Code: 2=2 years or more")
comment(table.1.2$zon) <-
c("Name: Zone",
"Code: 1=Central and semi-central parts of Sweden's three largest cities",
"Code: 2=suburbs and middle-sized towns",
"Code: 3=Lesser towns, except those in 5 or 7",
"Code: 4=Small towns and countryside, except 5--7",
"Code: 5=Northern towns",
"Code: 6=Northern countryside",
"Code: 7=Gotland (Sweden's largest island)")
comment(table.1.2$dur) <-
c("Name: Duration",
"Unit: year")
comment(table.1.2$medskad) <-
c("Name: Claim severity",
"Unit: SEK")
comment(table.1.2$antskad) <- "Name: No. claims"
comment(table.1.2$riskpre) <-
c("Name: Pure premium",
"Unit: SEK")
comment(table.1.2$helpre) <-
c("Name: Actual premium",
"Note: The premium for one year according to the tariff in force 1999",
"Unit: SEK")
comment(table.1.2$skadfre) <-
c("Name: Claim frequency",
"Unit: /year")
save(table.1.2, file = "table.1.2.RData")
print(table.1.2)
install.packages(c("data.table", "foreach", "ggplot2"), dependencies = TRUE, repos = "http://cran.us.r-project.org")
if (!exists("table.1.2"))
load("table.1.2.RData")
library("foreach")
table27 <-
data.frame(rating.factor =
c(rep("Vehicle class", nlevels(table.1.2$premiekl)),
rep("Vehicle age", nlevels(table.1.2$moptva)),
rep("Zone", nlevels(table.1.2$zon))),
class =
c(levels(table.1.2$premiekl),
levels(table.1.2$moptva),
levels(table.1.2$zon)),
stringsAsFactors = FALSE)
new.cols <-
foreach (rating.factor = c("premiekl", "moptva", "zon"),
.combine = rbind) %do%
{
nclaims <- tapply(table.1.2$antskad, table.1.2[[rating.factor]], sum)
sums <- tapply(table.1.2$dur, table.1.2[[rating.factor]], sum)
n.levels <- nlevels(table.1.2[[rating.factor]])
contrasts(table.1.2[[rating.factor]]) <-
contr.treatment(n.levels)[rank(-sums, ties.method = "first"), ]
data.frame(duration = sums, n.claims = nclaims)
}
table27 <- cbind(table27, new.cols)
rm(new.cols)
model.frequency <-
glm(antskad ~ premiekl + moptva + zon + offset(log(dur)),
data = table.1.2, family = poisson)
rels <- coef( model.frequency )
rels <- exp( rels[1] + rels[-1] ) / exp( rels[1] )
table27$rels.frequency <-
c(c(1, rels[1])[rank(-table27$duration[1:2], ties.method = "first")],
c(1, rels[2])[rank(-table27$duration[3:4], ties.method = "first")],
c(1, rels[3:8])[rank(-table27$duration[5:11], ties.method = "first")])
model.severity <-
glm(medskad ~ premiekl + moptva + zon,
data = table.1.2[table.1.2$medskad > 0, ],
family = Gamma("log"), weights = antskad)
rels <- coef( model.severity )
rels <- exp( rels[1] + rels[-1] ) / exp( rels[1] )
table27$rels.severity <-
c(c(1, rels[1])[rank(-table27$duration[1:2], ties.method = "first")],
c(1, rels[2])[rank(-table27$duration[3:4], ties.method = "first")],
c(1, rels[3:8])[rank(-table27$duration[5:11], ties.method = "first")])
table27$rels.pure.premium <- with(table27, rels.frequency * rels.severity)
print(table27, digits = 2)
```
| github_jupyter |
<img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# Remotive - Post daily jobs on slack
<a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Remotive/Remotive_Post_daily_jobs_on_slack.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
**Tags:** #remotive #jobs #slack #gsheet #naas_drivers #automation #opendata #text
**Author:** [Sanjeet Attili](https://www.linkedin.com/in/sanjeet-attili-760bab190/)
## Input
### Import libraries
```
import pandas as pd
from bs4 import BeautifulSoup
import requests
from datetime import datetime
import time
from naas_drivers import gsheet, slack
import naas
```
### Setup slack channel configuration
```
SLACK_TOKEN = "xoxb-1481042297777-3085654341191-xxxxxxxxxxxxxxxxxxxxxxxxx"
SLACK_CHANNEL = "05_work"
```
### Setup sheet log data
```
spreadsheet_id = "1EBefhkbmqaXMZLRCiafabf6xxxxxxxxxxxxxxxxxxx"
sheet_name = "SLACK_CHANNEL_POSTS"
```
### Setup Remotive
#### Get categories from Remotive
```
def get_remotejob_categories():
req_url = f"https://remotive.io/api/remote-jobs/categories"
res = requests.get(req_url)
try:
res.raise_for_status()
except requests.HTTPError as e:
return e
res_json = res.json()
# Get categories
jobs = res_json.get('jobs')
return pd.DataFrame(jobs)
df_categories = get_remotejob_categories()
df_categories
```
#### Enter your parameters
```
categories = ['data'] # Pick the list of categories in columns "slug"
date_from = - 10 # Choose date difference in days from now => must be negative
```
### Set the Scheduler
```
naas.scheduler.add(recurrence="0 9 * * *")
# # naas.scheduler.delete() # Uncomment this line to delete your scheduler if needed
```
## Model
### Get the sheet log of jobs
```
df_jobs_log = gsheet.connect(spreadsheet_id).get(sheet_name=sheet_name)
df_jobs_log
```
### Get all jobs posted after timestamp_date
All jobs posted after the date from will be fetched.<br>
In summary, we can set the value, in seconds, of 'search_data_from' to fetch all jobs posted since this duration
```
REMOTIVE_DATETIME = "%Y-%m-%dT%H:%M:%S"
NAAS_DATETIME = "%Y-%m-%d %H:%M:%S"
def get_remotive_jobs_since(jobs, date):
ret = []
for job in jobs:
publication_date = datetime.strptime(job['publication_date'], REMOTIVE_DATETIME).timestamp()
if publication_date > date:
ret.append({
'URL': job['url'],
'TITLE': job['title'],
'COMPANY': job['company_name'],
'PUBLICATION_DATE': datetime.fromtimestamp(publication_date).strftime(NAAS_DATETIME)
})
return ret
def get_category_jobs_since(category, date, limit):
url = f"https://remotive.io/api/remote-jobs?category={category}&limit={limit}"
res = requests.get(url)
if res.json()['jobs']:
publication_date = datetime.strptime(res.json()['jobs'][-1]['publication_date'], REMOTIVE_DATETIME).timestamp()
if len(res.json()['jobs']) < limit or date > publication_date:
print(f"Jobs from catgory {category} fetched ✅")
return get_remotive_jobs_since(res.json()['jobs'], date)
else:
return get_category_jobs_since(category, date, limit + 5)
return []
def get_jobs_since(categories: list,
date_from: int):
if date_from >= 0:
return("'date_from' must be negative. Please update your parameter.")
# Transform datefrom int to
search_jobs_from = date_from * 24 * 60 * 60 # days in seconds
timestamp_date = time.time() + search_jobs_from
jobs = []
for category in categories:
jobs += get_category_jobs_since(category, timestamp_date, 5)
print(f'- All job since {datetime.fromtimestamp(timestamp_date)} have been fetched -')
return pd.DataFrame(jobs)
df_jobs = get_jobs_since(categories, date_from=date_from)
df_jobs
```
### Remove duplicate jobs
```
def remove_duplicates(df1, df2):
# Get jobs log
jobs_log = df1.URL.unique()
# Exclude jobs already log from jobs
df2 = df2[~df2.URL.isin(jobs_log)]
return df2.sort_values(by="PUBLICATION_DATE")
df_new_jobs = remove_duplicates(df_jobs_log, df_jobs)
df_new_jobs
```
## Output
### Add new jobs on the sheet log
```
gsheet.connect(spreadsheet_id).send(sheet_name=sheet_name,
data=df_new_jobs,
append=True)
```
### Send all jobs link to the slack channel
```
if len(df_new_jobs) > 0:
for _, row in df_new_jobs.iterrows():
url = row.URL
slack.connect(SLACK_TOKEN).send(SLACK_CHANNEL, f"<{url}>")
else:
print("Nothing to published in Slack !")
```
| github_jupyter |
# Unsupervised Classification
A script that uses unsupervised classification to explore the data a little bit and see if we can reconstruct the labels directly without using supervised learning.
```
import common, preprocess, numpy as np
import pandas as pd
zImages = preprocess.getKeyFeatures()
```
We're going to first try using just the features that we previously selected.
```
df = pd.DataFrame()
for k, keepKey in enumerate(zImages):
band = zImages[keepKey]
df[keepKey] = band.flatten()
subset = df.sample(n=100000)
```
While we could run the clustering algorithms directly on the entire images, in practice that takes too much compute time, and typically gains you very little unless you want to overfit your data, so random downsampling is typically a good idea.
```
labelImage = common.loadNumpy('labels')
h,w= labelImage.shape
```
First up is the classic KMeans clustering algorithm. KMeans works well when there is large separation between clusters. I would expect it to not do so well here because of the significant amount of similarity between clusters just looking at the band images visually. Another big problem with K-means is the need to select the number of classes that you want. In this case, we're just going to select 7 because I did some testing and saw that it was really making very small clusters when we increased it beyond this.
```
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=7)
kmeans.fit(subset)
classes = kmeans.predict(df)
kclass = classes.reshape(h,w)
```
We're going to compare the K-means clustering results with the desired classes directly in image format. Note: the colorbars are not directly comparable because the class numbers don't correspond to one another yet, but in general we can see some similar patterns. Here are some correspondences that I think stand out based on the colors in each image:
| K-means | Labeled Image |
|---------|---------------|
| purple | orange |
|pink | green |
|orange | red |
|green | brown |
```
from pylab import *
figure(figsize=(20,20))
subplot(211)
title('K-means')
imshow(kclass,cmap='tab10',vmax=10)
colorbar()
subplot(212)
title('Labeled')
imshow(labelImage,cmap='tab10',vmax=10)
colorbar()
from scipy.stats import mode
classMapping = {}
for k in range(10):
mask = kclass == k
most_likely, _ = mode(labelImage[mask])
if len(most_likely) > 0:
classMapping[k] = most_likely[0]
print('most likely class: ', k,most_likely)
figure(figsize=(20,20))
subplot(211)
title('K-means remapped to match original classes')
kremap = np.zeros_like(kclass)
for kc, mc in classMapping.items():
kremap[kclass == kc] = mc
# kremap[kclass == 4] = 1 # purple -> orange
# kremap[kclass == 6] = 2 # pink -> green
# kremap[kclass == 1] = 3 # orange -> red
# kremap[kclass == 2] = 5 # green -> brown
imshow(kremap,cmap='tab10', vmax=10)
colorbar()
subplot(212)
title('Labeled')
imshow(labelImage,cmap='tab10',vmax=10)
colorbar()
```
Another way to visualize the data is to remap the classes from Kmeans to the classes in the labeled dataset. I estimated which class each Kmeans cluster belongs to by simply looking at the most likely occuring class at the corresponding points in the label image. As the image above shows, there is actually pretty decent correspondence betwen the Kmeans clusters and some of the classes, but it's not a perfect match.
We could try other clustering algorithms to see if we can get better results, but ultimately I think it's not really worth the effort to see something marginally better when we're going to do supervised classification anyways...
| github_jupyter |
<font color='purple'><b><center>~ Proyecto Segundo Sprint ~</center></b></font>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('C:/Users/Usuario/Downloads/DS_Proyecto_01_Datos_Properati.csv')
```
<font color='purple'><b>~ Elección de datos ~</b></font>
Los datos que se utilizarán para predecir el precio de la propiedad son la cantidad de habitaciones, baños y superficie total. El resto de datos se descartan ya que los datos elegidos dan la mayor cantidad de informacón sobre el inmueble y su posible precio.
<font color='purple'><b>~ Análisis exploratorio de datos ~</b></font>
```
data
data.describe()
data.isna().sum()/data.shape[0]*100
data.isnull().sum()
```
Las variables de baños, superficie total, superficie cubierta, latitud y longitud son las únicas en presentar datos nulos. Las variables de superficie son las que presentan la mayor cantidad de datos nulos.
<font color='purple'><b>~ Correlación ~</b></font>
```
corr = data.corr()
plt.figure(figsize = (10,10))
sns.heatmap(corr, square=True, annot=True).set_title('Correlación')
```
Aquí se puede observar que las variables que están más fuertemente correlacionadas con el precio son baños, log_sc, log_st, habitaciones y dormitorios en orden descendente.
<font color='purple'><b>~ Gráficas de distribución ~</b></font>
```
plt.figure(figsize = (20,10))
sns.countplot(data=data, x = "bedrooms", palette='crest').set_title('Número de Habitaciones')
plt.figure(figsize = (20,10))
sns.countplot(data=data, x= "bathrooms", palette='crest').set_title('Número de Baños')
plt.figure(figsize = (20,10))
sns.distplot(data.surface_total, bins=20).set_title('Superficie Total')
plt.figure(figsize = (20,10))
sns.distplot(data.surface_covered, bins=20).set_title('Superficie Cubierta')
plt.figure(figsize = (20,10))
sns.countplot(data=data, x= "l2", order = data['l2'].value_counts().index, palette= 'crest').set_title('Zona')
plt.figure(figsize = (20,10))
sns.countplot(data=data, x= "l3", order = data['l3'].value_counts().index, palette= 'crest').set_title('Barrios')
plt.xticks(rotation=90)
plt.subplots_adjust(bottom=0.15)
```
<font color='purple'><b>~ Presencia de datos atípicos ~</b></font>
```
plt.figure(figsize = (20,10))
sns.boxplot(data.surface_total).set_title('Boxplot ST')
plt.figure(figsize = (20,10))
sns.boxplot(data.surface_covered).set_title('Boxplot SC')
```
Se puede observar la presencia de outliers en las variables `surface_total` y `surface_covered` además de una alta dispersión de los datos.
<font color='purple'><center><b>~ Pre-procesamiento de datos ~</b></center></font>
Para realizar los modelos de regresión se realizará un filtrado de datos de tal forma que se evaluen las propiedades con valores menores a los 4000000 dolares, las propiedades de tipo PH, departamento y casa y aquellas cuya superficie este entre los 15 y 1000 m2. También se hará imputación de los valores nulos de la variable `bathrooms` utilizando su media.
```
df2 = data[(data['price'] <= 4000000)] #filtando propiedades con precios menores a 4000000 dolares
df2 = df2[(df2['l2'] == "Capital Federal") & (df2.property_type.isin(['Departamento','Casa','PH']))] #filtrando por zona y tipo de propiedades, las más publicadas
df2 = df2[(df2['surface_total']>= 15) & (df2['surface_total'] <= 1000)] #filtrando propiedades con superficies entre 15 y 1000 m2
df2
df2['bathrooms'].fillna(df2['bathrooms'].mean(), inplace = True)
df2.isnull().sum()
df2.describe()
```
<font color='purple'><b>Modelo Benchmark</b></font>
```
X = df2[['surface_total', 'bedrooms', 'bathrooms']]
y= df2.price
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.30, random_state=42)
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
from sklearn.tree import DecisionTreeRegressor
tree_regressor = DecisionTreeRegressor(max_depth=12, random_state=42)
tree_regressor.fit(X_train, y_train)
#Arbol de decisión
y_train_pred = tree_regressor.predict(X_train)
y_test_pred = tree_regressor.predict(X_test)
from sklearn.metrics import mean_squared_error
rmse_train = np.sqrt(mean_squared_error(y_train, y_train_pred))
rmse_test = np.sqrt(mean_squared_error(y_test, y_test_pred))
print(f'Raíz del error cuadrático medio en Train: {rmse_train}')
print(f'Raíz del error cuadrático medio en Test: {rmse_test}')
```
<font color='purple'><b>PCA</b></font>
```
encode_b = pd.get_dummies(df2.l3)
df_destino = pd.concat([df2, encode_b], axis=1)
df_destino
X = df_destino[['surface_total', 'bedrooms', 'bathrooms']]
y= df_destino.price
from sklearn.decomposition import PCA
pca = PCA(n_components=3)
X_nuevo = pca.fit_transform(X)
pca.components_
X_train, X_test, y_train, y_test = train_test_split(X_nuevo, y, test_size=0.33, random_state=42)
regresor2 = DecisionTreeRegressor(max_depth=10)
regresor2.fit(X_train,y_train)
y_train_pred = regresor2.predict(X_train)
y_test_pred = regresor2.predict(X_test)
#PCA
print(mean_squared_error(y_train,y_train_pred))
print(mean_squared_error(y_test,y_test_pred))
regresor2.feature_importances_
```
Para evaluar los resultados del modelo se hace uso de RMSE para elegir el modelo que presente los errores más bajos entre los modelos utlizados. Ya que los errores están afectados por el efecto de los valores atípicos de superficie, esto no busca mostrar los errores minimos posibles que harían a este modelo más eficiente.
```
from sklearn.metrics import mean_squared_error
modelos = ['Árbol de Decisión', 'PCA']
for i, model in enumerate([tree_regressor, regresor2]):
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
print(f'Modelo: {modelos[i]}')
rmse_train = np.sqrt(mean_squared_error(y_train, y_train_pred))
rmse_test = np.sqrt(mean_squared_error(y_test, y_test_pred))
print(f'Raíz del error cuadrático medio en Train: {rmse_train}')
print(f'Raíz del error cuadrático medio en Test: {rmse_test}')
plt.figure(figsize = (8,4))
plt.subplot(1,2,1)
sns.histplot(np.array(y_train) - y_train_pred, bins = 30, label = 'train', color='powderblue')
sns.histplot(np.array(y_test) - y_test_pred, bins = 30, label = 'test', color = 'darkorchid')
plt.xlabel('errores')
plt.legend()
ax = plt.subplot(1,2,2)
ax.scatter(y_test,y_test_pred, s =2, c='#e59fed')
lims = [
np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes]
]
ax.plot(lims, lims, 'k-', alpha=0.75, zorder=0)
plt.xlabel('y (test)')
plt.ylabel('y_pred (test)')
plt.tight_layout()
plt.show()
```
<font color='purple'><b>Conclusión</b></font>
De acuerdo a los resultados obtenidos, el modelo elegido entre los dos es PCA al presentar errores menores a los de árbol de decisión.
<font color='purple'><b>Problemas a solucionar</b></font>
Ambos modelos muestran errores menores en `train` que en `test` lo que podría significar cierto sobreajuste aunque en ninguno de los dos la diferencia es muy grande o significativa.
| github_jupyter |
# Ingetsing larger catalogs: PanStarrs DR1 Mean/Thin Objects
This notebook follows up on *insert_example*, be sure to go through that one first.
Here we show a possible way in which the catalog ingestion step can be sped up though the use of mulitprocessing. In particular, we will try to import some of the files from a subset and compilation of PS1 DR1 mean and thin object tables. The entire subsample contains 1.92B sources, selected requiring nDetections>2.
Reference: https://panstarrs.stsci.edu/
The test files we will be using for this test can be downloaded from:
https://desycloud.desy.de/index.php/s/stCkA6uJ8ayKvjI
```
import CatalogPusher
import pandas as pd
import concurrent.futures
from healpy import ang2pix
import importlib
importlib.reload(CatalogPusher)
# build the pusher object and point it to the raw files.
ps1p = CatalogPusher.CatalogPusher(
catalog_name = 'ps1_test', # short name of the catalog
data_source = '../testdata/PS1DR1_test/', # where to find the data (other options are possible)
file_type = '*.csv.gz' # filter files (there is col definition file in data_source)
)
# define the reader for the raw files (import column names from file.)
headfile = '../testdata/PS1DR1_test/column_headings.csv'
with open(headfile, 'r') as header:
catcols=[c.strip() for c in header.readline().split(',')]
ps1p.assign_file_reader(
reader_func = pd.read_csv, # callable to use to read the raw_files.
read_chunks = True, # weather or not the reader process each file into smaller chunks.
names=catcols, # All other arguments are passed directly to this function.
chunksize=50000,
engine='c')
# define modifier. This time the healpix grid is finer (an orer 16 corresponds to 3")
hp_nside16=2**16
def ps1_modifier(srcdict):
ra=srcdict['raMean'] if srcdict['raMean']<180. else srcdict['raMean']-360.
srcdict['pos']={
'type': 'Point',
'coordinates': [ra, srcdict['decMean']]
}
srcdict['hpxid_12']=int(
ang2pix(hp_nside16, srcdict['raMean'], srcdict['decMean'], lonlat = True, nest = True))
return srcdict
ps1p.assign_dict_modifier(ps1_modifier)
# wrap up the file pushing function so that we can
# use multiprocessing to speed up the catalog ingestion
def pushfiles(filerange):
ps1p.push_to_db(
coll_name = 'srcs',
index_on = ['hpxid_12'],
filerange = filerange,
overwrite_coll = False,
dry = False)
# each job will run on a subgroup of all the files
file_groups = ps1p.file_groups(group_size=4)
with concurrent.futures.ProcessPoolExecutor(max_workers = 2) as executor:
executor.map(pushfiles, file_groups)
```
| github_jupyter |
AMUSE tutorial on multiple code in a single bridge
====================
A cascade of bridged codes to address the problem of running multiple planetary systems in, for example, a star cluster. This is just an example of how to initialize such a cascaded bridge without any stellar evolution, background potentials. The forces for one planetary system on the planets in the other systems are ignored to save computer time. This gives rise to some energy errors, and inconsistencies (for example when one star tries to capture planets from another system. The latter will not happen here.
This can be addressed by intorducing some logic in checking what stars are nearby which planets.
```
import numpy
from amuse.units import (units, constants)
from amuse.lab import Particles
from amuse.units import nbody_system
from matplotlib import pyplot
## source https://en.wikipedia.org/wiki/TRAPPIST-1
trappist= {"b": {"m": 1.374 | units.MEarth,
"a": 0.01154 | units.au,
"e": 0.00622,
"i": 89.56},
"c": {"m": 1.308 | units.MEarth,
"a": 0.01580 | units.au,
"e": 0.00654,
"i": 89.70},
"d": {"m": 0.388 | units.MEarth,
"a": 0.02227 | units.au,
"e": 0.00837,
"i": 89.89},
"e": {"m": 0.692 | units.MEarth,
"a": 0.02925 | units.au,
"e": 0.00510,
"i": 89.736},
"f": {"m": 1.039 | units.MEarth,
"a": 0.03849 | units.au,
"e": 0.01007,
"i": 89.719},
"g": {"m": 1.321 | units.MEarth,
"a": 0.04683 | units.au,
"e": 0.00208,
"i": 89.721},
"h": {"m": 0.326 | units.MEarth,
"a": 0.06189 | units.au,
"e": 0.00567,
"i": 89.796}
}
def trappist_system():
from amuse.ext.orbital_elements import new_binary_from_orbital_elements
from numpy.random import uniform
star = Particles(1)
setattr(star, "name", "")
setattr(star, "type", "")
star[0].mass = 0.898 | units.MSun
star[0].position = (0,0,0) | units.au
star[0].velocity = (0,0,0) | units.kms
star[0].name = "trappist"
star[0].type = "star"
bodies = Particles(len(trappist))
setattr(bodies, "name", "")
setattr(bodies, "type", "")
for bi, planet in zip(bodies, trappist):
true_anomaly = uniform(0, 360)
b = new_binary_from_orbital_elements(star.mass,
trappist[planet]['m'],
trappist[planet]["a"],
trappist[planet]["e"],
true_anomaly = true_anomaly,
inclination = trappist[planet]["i"],
G = constants.G)
bi.name = planet
bi.type = "planet"
bi.mass = b[1].mass
bi.position = b[1].position - b[0].position
bi.velocity = b[1].velocity - b[0].velocity
return star | bodies
from amuse.community.ph4.interface import ph4
from amuse.community.hermite.interface import Hermite
from amuse.ic.plummer import new_plummer_model
import numpy.random
numpy.random.seed(1624973942)
converter=nbody_system.nbody_to_si(1 | units.MSun, 0.1|units.parsec)
t1 = trappist_system()
t2 = trappist_system()
t3 = trappist_system()
p = new_plummer_model(3, convert_nbody=converter)
t1.position += p[0].position
t1.velocity += p[0].velocity
t2.position += p[1].position
t2.velocity += p[1].velocity
t3.position += p[2].position
t3.velocity += p[2].velocity
converter=nbody_system.nbody_to_si(t1.mass.sum(), 0.1|units.au)
bodies = Particles(0)
gravity1 = ph4(converter)
t = gravity1.particles.add_particles(t1)
bodies.add_particles(t1)
gravity2 = ph4(converter)
t = gravity2.particles.add_particles(t2)
bodies.add_particles(t2)
gravity3 = Hermite(converter)
t = gravity3.particles.add_particles(t3)
bodies.add_particles(t3)
channel_from_g1 = gravity1.particles.new_channel_to(bodies)
channel_from_g2 = gravity2.particles.new_channel_to(bodies)
channel_from_g3 = gravity3.particles.new_channel_to(bodies)
from amuse.plot import scatter
from matplotlib import pyplot
scatter(bodies.x-bodies[0].x, bodies.z-bodies[0].z)
#pyplot.xlim(-0.1, 0.1)
#pyplot.ylim(-0.1, 0.1)
pyplot.show()
def plot(bodies):
from amuse.plot import scatter
from matplotlib import pyplot
stars = bodies[bodies.type=='star']
planets = bodies-stars
pyplot.scatter((stars.x-bodies[0].x).value_in(units.au),
(stars.z-bodies[0].z).value_in(units.au), c='r', s=100)
pyplot.scatter((planets.x-bodies[0].x).value_in(units.au),
(planets.z-bodies[0].z).value_in(units.au), c='b', s=10)
pyplot.xlim(-0.1, 0.1)
pyplot.ylim(-0.1, 0.1)
pyplot.show()
plot(bodies)
from amuse.couple import bridge
gravity = bridge.Bridge()
gravity.add_system(gravity1, (gravity2,gravity3))
gravity.add_system(gravity2, (gravity1,gravity3))
gravity.add_system(gravity3, (gravity1,gravity2))
from amuse.lab import zero
Etot_init = gravity.kinetic_energy + gravity.potential_energy
Etot_prev = Etot_init
gravity.timestep = 100.0| units.yr
time = zero
dt = 200.0|units.yr
t_end = 1000.0| units.yr
while time < t_end:
time += dt
gravity.evolve_model(time)
Etot_prev_se = gravity.kinetic_energy + gravity.potential_energy
channel_from_g1.copy()
channel_from_g2.copy()
channel_from_g3.copy()
plot(bodies)
print(bodies[1].position.in_(units.au))
Ekin = gravity.kinetic_energy
Epot = gravity.potential_energy
Etot = Ekin + Epot
print("T=", time.in_(units.yr), end=' ')
print("E= ", Etot/Etot_init, "Q= ", Ekin/Epot, end=' ')
print("dE=", (Etot_init-Etot)/Etot, "ddE=", (Etot_prev-Etot)/Etot)
Etot_prev = Etot
gravity.stop()
```
| github_jupyter |
# BERT Fine-Tuning Sentence Classification
> BERT Fine-Tuning Tutorial with PyTorch
>
> ref: https://colab.research.google.com/drive/1Y4o3jh3ZH70tl6mCd76vz_IxX23biCPP
- toc: true
- badges: true
- comments: true
- categories: [bert, jupyter]
# 1. Setup
## 1.1. Using Colab GPU for Training
```
import tensorflow as tf
# Get the GPU device name.
device_name = tf.test.gpu_device_name()
# The device name should look like the following:
if device_name == '/device:GPU:0':
print('Found GPU at: {}'.format(device_name))
else:
raise SystemError('GPU device not found')
import torch
# If there's a GPU available,
if torch.cuda.is_available():
# Tell PyTorch to user the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will user the GPU:', torch.cuda.get_device_name(0))
# If not,
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
```
## 1.2. Installing the Hugging Face Library
```
! pip install transformers -q
! pip install wget -q
import wget
import os
print('Downloading dataset...')
# The URL for the dataset zip file.
url = 'https://nyu-mll.github.io/CoLA/cola_public_1.1.zip'
# Download the file (if we haven't already)
if not os.path.exists('./cola_public_1.1.zip'):
wget.download(url, './cola_public_1.1.zip')
# Unzip the dataset (if we haven't already)
if not os.path.exists('./cola_public/'):
! unzip cola_public_1.1.zip
! head cola_public/*/*
```
## 2.2. Parse
```
import pandas as pd
# Load the dataset into a pandas dataframe.
df = pd.read_csv("./cola_public/raw/in_domain_train.tsv", delimiter='\t', header=None, names=['sentence_source', 'label', 'label_notes', 'sentence'])
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(df.shape[0]))
# Display 10 random rows from the data
df.sample(10)
df[df.label == 0].sample(5)[['sentence', 'label']]
# Get the lists of sentences and their labels.
sentences = df.sentence.values
labels = df.label.values
```
# 3. Tokenization & Input Formatting
## 3.1. BERT Tokenizer
```
from transformers import BertTokenizer
# Load the BERT tokenizer.
print('Loading BERT tokenizer...')
tokenizer = BertTokenizer.from_pretrained('bert-base-multilingual-uncased', do_lower_case=True)
# Print the original sentence.
print(' Original: ', sentences[0])
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(sentences[0]))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sentences[0])))
korean = "안녕하세요. 반갑습니다. 너는 이름이 뭐니? 오늘 날씨가 맑고 좋구나."
# Print the original sentence.
print(' Original: ', korean)
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(korean))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(korean)))
```
When we actually convert all of our sentences, we'll use the `tokenize.encode` functio to handle both steps, rather than calling `tokenize` and `convert_tokens_to_ids` seperately.
## 3.2. Required Formatting
## 3.3. Tokenize Dataset
```
max_len = 0
# For every sentence,
for sent in sentences:
# Tokenize the text and add `[CLS]` and `[SEP]` tokens.
input_ids = tokenizer.encode(sent, add_special_tokens=True)
# Update the maximum sentence langth.
max_len = max(max_len, len(input_ids))
print('Max sentence length: ', max_len)
# Tokenize all of the sentences and map the tokens to their word IDs.
input_ids = []
attention_masks = []
# For every sentence,
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_Tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding.)
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.LongTensor(input_ids)
attention_masks = torch.FloatTensor(attention_masks)
labels = torch.tensor(labels)
# Print sentence 0, now as a list of IDs.
print('Original: ', sentences[0])
print('Token IDs: ', input_ids[0])
```
## 3.4. Training & Validation Split
```
from torch.utils.data import TensorDataset, random_split
# Combine the training inputs into a TensorDataset.
dataset = TensorDataset(input_ids, attention_masks, labels)
# Create a 90-10 train-validation split.
# Calculate the number of samples to include in each set.
train_size = int(0.9 * len(dataset))
val_size = len(dataset) - train_size
# Devide the dataset by randomly selecting samples.
train_dataset, val_dataset = random_split(dataset, [train_size, val_size])
print('{:>5} training samples'.format(train_size))
print('{:>5} validation samples'.format(val_size))
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# The DataLoader needs to know our batch size for training, so we specify it here.
# For fine-tuning BERT on a specific task, the authors recommend a batch size of 16 or 32.
batch_size = 32
# Create the DataLoaders for our training and validation sets.
# We'll take training samples in random order.
train_dataloader = DataLoader(
train_dataset, # The training samples.
sampler = RandomSampler(train_dataset), # Select batches randomly
batch_size = batch_size # Trains with this batch size.
)
# For validation the order doesn't matter, so we'll just read them sequentially.
validation_dataloader = DataLoader(
val_dataset, # The validation samples.
sampler = SequentialSampler(val_dataset), # Pull out batches sequentially.
batch_size = batch_size # Evaluate with this batch size.
)
train_dataloader.batch_size
# 7695/32 = 240.46875
```
# 4. Train Our Classification Model
## 4.1. BertForSequenceClassification
* BertModel
* BertForPreTraining
* BertForMaskedLM
* BertForNextSentencePredicion
* BertForSequenceClassification
* BertForTokenClassification
* BertForQuestionAnswering
```
device
from transformers import BertForSequenceClassification, AdamW, BertConfig
# Load BertForSequenceClassification, the pretrained BERT model with
# a single linear classification layer on top.
model = BertForSequenceClassification.from_pretrained(
"bert-base-uncased", # Use the 12-layer BERT model, with an uncased vocab.
num_labels = 2, # The number of output labels--2 for binary classification.
# You can increase this for multi-class tasks.
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
# Tell pytorch to run this model on the GPU.
# if device == 'cuda':
model.cuda()
# Get all of the model's parameters as a list of tuples.
params = list(model.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
```
## 4.2. Optimizer & Learning Rate Scheduler
```
# Note: AdamW is a class from the huggingface library (as opposed to pytorch)
# I believe the 'W' stands for 'Weight Decay fix'
optimizer = AdamW(model.parameters(),
lr = 2e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.dam_epsilon - default is 1e-8
)
from transformers import get_linear_schedule_with_warmup
# Number of training epochs. The BERT authors recommend between 2 and 4.
# We chose to run for 4, but we'll see later that this my be over-fitting the training data.
epochs = 4
# Total number of training steps is [number of batches] x [number of epochs].
# (Note that this is not the same as the number of training samples).
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps=0, # Default value in run_glue.py
num_training_steps = total_steps)
```
## 4.3. Training Loop
**Training:**
* Unpack our data inputs and labels
* Load data onto the GPU for acceleration
* Clear out the gradients calculated in the previous pass.
- In pytorch the gradients accumulate by default (useful for things like RNNs) unless you explicitly clear them out.
* Forward pass (feed input data through the network)
* Backward pass (backpropagation)
* Tell the network to update parameters with optimizer.step()
* Track variables for monitoring progress
**Evaluation:**
* Unpack our data inputs and labels
* Load data onto the GPU for acceleration
* Forward pass (feed input data through the network)
* Compute loss on our validation data and track variables for monitoring progress
```
import numpy as np
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
import time
import datetime
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
print(" ", elapsed, elapsed_rounded, str(datetime.timedelta(seconds=elapsed_rounded)))
return str(datetime.timedelta(seconds=elapsed_rounded))
import random
import numpy as np
# This training code is based on the `run_glue.py` script here:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L128
# Set the seed value all over the place to make this reproducible.
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# We'll store a number of quantities such as training and validation loss,
# validation accuracy, and timings.
training_stats = []
# Measure the total training time for the whole run.
total_t0 = time.time()
# For each epoch,
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
# Perform one full pass over the training set.
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i+1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_train_loss = 0
# Put the model into training mode. Don't be mislead--the call to
# `train` just changes the *mode*, it doesn't *perform* the training.
# `dropout` and `batchnorm` layers behave differently during training vs test
# (source: https://stackoverflow.com/questions/51433378/what-does-model-train-do-in-pytorch)
model.train()
# For each batch of training data,
for step, batch in enumerate(train_dataloader):
# # of step : 241
# len of batch : 32
# size of batch[0] : 64
# Progress update every 40 batches.
if step % 40 == 0 and not step == 0:
# Calcuate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}'.format(step, len(train_dataloader), elapsed))
# Unpack this training batch from our dataloader
#
# As we unpack the batch, we'll also copy each tensor to the GPU using `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1] : attention masks
# [2] : labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
print(type(batch[0]))
print(batch[0])
print(type(b_input_ids))
print(b_input_ids)
# Always clear any previously calculated gradients before performing backward pass.
# PyTorch doesn't do this automatically because accumulating the gradients is
# "convenient while training RNNs". (source: https://stackoverflow.com/questions/48001598/why-do-we-need-to-call-zero-grad-in-pytorch)
model.zero_grad()
# Perform a forwad pass (evaluate the model on this training batch).
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
# It returns different numbers of parameters depending on what arguments
# are given and what flags are set. For our usage here, it returns the loss (because we provided labels)
# and the "logits"--the model outputs prior to activation.
loss, logits = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the training loss over all of the batches so that we can calculate the average loss at the end.
# `loss` is a Tensor containing a single value; the `.item()` function just returns the Python value from the tensor.
total_train_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are modified
# based on their gradients, the learning rate, etc.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over all of the batches.
avg_train_loss = total_train_loss / len(train_dataloader)
# Measure how long this epoch took.
training_time = format_time(time.time() - t0)
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epoch took: {:}".format(training_time))
# ========================================
# Validation
# ========================================
# After the completion of each training epoch, measure our performance on our validation set.
print("")
print("Running Validation...")
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently during evaluation.
model.eval()
# Tracking variables
total_eval_accuracy = 0
total_eval_loss = 0
nb_eval_steps = 0
# Evaluate data for one epoch
for batch in validation_dataloader:
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using the `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Tell pytorch not to bother with constructing the compute graph during the forward pass,
# since this is only needed for backprop (training).
with torch.no_grad():
# Forward pass, calculate logit predictions.
# token_type_ids is the same as the "segment ids", which differentiates sentence 1 and 2 in 2-sentence tasks.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
# Get the "logits" output byt the model. The "logits" are the output
# values prior to applying an activation function like the softmax.
(loss, logits) = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the validation loss.
total_eval_loss += loss.item()
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentence,
# and accumulate it over all batches.
total_eval_accuracy += flat_accuracy(logits, label_ids)
# Report the final accuracy for this validation run.
avg_val_accuracy = total_eval_accuracy / len(validation_dataloader)
print(" Accuracy: {0:.2f}".format(avg_val_accuracy))
# Calculate the average loss over all of the batches.
avg_val_loss = total_eval_loss / len(validation_dataloader)
# Measure how long the validation run took.
validation_time = format_time(time.time() - t0)
print(" Validation Loss: {0:.2f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
# Record all statistics from this epoch.
training_stats.append(
{
'epoch': epoch_i + 1,
'Training Loss': avg_train_loss,
'Valid. Loss': avg_val_loss,
'Valid. Accur.': avg_val_accuracy,
'Training Time': training_time,
'Validation Time': validation_time
}
)
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
```
| github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex SDK: AutoML training tabular binary classification model for online prediction
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/sdk/sdk_automl_tabular_binary_classification_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/sdk/sdk_automl_tabular_binary_classification_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex SDK to create tabular binary classification models and do online prediction using Google Cloud's [AutoML](https://cloud.google.com/vertex-ai/docs/start/automl-users).
### Dataset
The dataset used for this tutorial is the [Bank Marketing](gs://cloud-ml-tables-data/bank-marketing.csv). This dataset does not require any feature engineering. The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket.
### Objective
In this tutorial, you create an AutoML tabular binary classification model and deploy for online prediction from a Python script using the Vertex SDK. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Google Cloud Console.
The steps performed include:
- Create a Vertex `Dataset` resource.
- Train the model.
- View the model evaluation.
- Deploy the `Model` resource to a serving `Endpoint` resource.
- Make a prediction.
- Undeploy the `Model`.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex SDK.
```
import sys
import os
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = '--user'
else:
USER_FLAG = ''
! pip3 install --upgrade google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex SDK and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" #@param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/ai-platform-unified/docs/general/locations)
```
REGION = 'us-central1' #@param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" #@param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
```
import google.cloud.aiplatform as aip
```
## Initialize Vertex SDK
Initialize the Vertex SDK for your project and corresponding bucket.
```
aip.init(project=PROJECT_ID, staging_bucket=BUCKET_NAME)
```
# Tutorial
Now you are ready to start creating your own AutoML tabular binary classification model.
## Create a Dataset Resource
First, you create an tabular Dataset resource for the Bank Marketing dataset.
### Data preparation
The Vertex `Dataset` resource for tabular has a couple of requirements for your tabular data.
- Must be in a CSV file or a BigQuery query.
#### CSV
For tabular binary classification, the CSV file has a few requirements:
- The first row must be the heading -- note how this is different from Vision, Video and Language where the requirement is no heading.
- All but one column are features.
- One column is the label, which you will specify when you subsequently create the training pipeline.
#### Location of Cloud Storage training data.
Now set the variable `IMPORT_FILE` to the location of the CSV index file in Cloud Storage.
```
IMPORT_FILE = 'gs://cloud-ml-tables-data/bank-marketing.csv'
```
#### Quick peek at your data
You will use a version of the Bank Marketing dataset that is stored in a public Cloud Storage bucket, using a CSV index file.
Start by doing a quick peek at the data. You count the number of examples by counting the number of rows in the CSV index file (`wc -l`) and then peek at the first few rows.
You also need for training to know the heading name of the label column, which is save as `label_column`. For this dataset, it is the last column in the CSV file.
```
count = ! gsutil cat $IMPORT_FILE | wc -l
print("Number of Examples", int(count[0]))
print("First 10 rows")
! gsutil cat $IMPORT_FILE | head
heading = ! gsutil cat $IMPORT_FILE | head -n1
label_column = str(heading).split(',')[-1].split("'")[0]
print("Label Column Name", label_column)
if label_column is None:
raise Exception("label column missing")
```
### Create the Dataset
Next, create the `Dataset` resource using the `create()` method for the `TabularDataset` class, which takes the following parameters:
- `display_name`: The human readable name for the `Dataset` resource.
- `gcs_source`: A list of one or more dataset index file to import the data items into the `Dataset` resource.
This operation may take several minutes.
```
dataset = aip.TabularDataset.create(
display_name="Bank Marketing" + "_" + TIMESTAMP,
gcs_source=[IMPORT_FILE]
)
print(dataset.resource_name)
```
## Train the model
Now train an AutoML tabular binary classification model using your Vertex `Dataset` resource. To train the model, do the following steps:
1. Create an Vertex training pipeline for the `Dataset` resource.
2. Execute the pipeline to start the training.
### Create and run training pipeline
To train an AutoML tabular binary classification model, you perform two steps: 1) create a training pipeline, and 2) run the pipeline.
#### Create training pipeline
An AutoML training pipeline is created with the `AutoMLTabularTrainingJob` class, with the following parameters:
- `display_name`: The human readable name for the `TrainingJob` resource.
- `optimization_prediction_type`: The type task to train the model for.
- `classification`: A tabuar classification model.
- `regression`: A tabular regression model.
- `forecasting`: A tabular forecasting model.
- `column_transformations`: (Optional): Transformations to apply to the input columns
- `optimization_objective`: The optimization objective to minimize or maximize.
- `minimize-log-loss`
```
dag = aip.AutoMLTabularTrainingJob(
display_name="bank_" + TIMESTAMP,
optimization_prediction_type="classification",
optimization_objective="minimize-log-loss"
)
```
#### Run the training pipeline
Next, you run the DAG to start the training job by invoking the method `run()`, with the following parameters:
- `dataset`: The `Dataset` resource to train the model.
- `model_display_name`: The human readable name for the trained model.
- `target_column`: The name of the column to train as the label.
- `training_fraction_split`: The percentage of the dataset to use for training.
- `validation_fraction_split`: The percentage of the dataset to use for validation.
- `test_fraction_split`: The percentage of the dataset to use for test (holdout data).
- `budget_milli_node_hours`: (optional) Maximum training time specified in unit of millihours (1000 = hour).
- `disable_early_stopping`: If `True`, training maybe completed before using the entire budget if the service believes it cannot further improve on the model objective measurements.
The `run` method when completed returns the `Model` resource.
The execution of the training pipeline will take upto 20 minutes.
```
model = dag.run(
dataset=dataset,
target_column=label_column,
model_display_name="bank_" + TIMESTAMP,
training_fraction_split=0.6,
validation_fraction_split=0.2,
test_fraction_split=0.2,
budget_milli_node_hours=1000,
disable_early_stopping=False
)
```
## Deploy the model
Next, deploy your `Model` resource to an `Endpoint` resource for online prediction. To deploy the `Model` resource, you invoke the `deploy()` method. This call will create an `Endpoint` resource automatically.
The method returns the created `Endpoint` resource.
The `deploy()` method takes the following arguments:
- `machine_type`: The type of compute machine.
```
endpoint = model.deploy(machine_type="n1-standard-4")
```
## Make a online prediction request
Now do a online prediction to your deployed model.
### Make test item
You will use synthetic data as a test data item. Don't be concerned that we are using synthetic data -- we just want to demonstrate how to make a prediction.
```
INSTANCE = {"Age": '58', "Job": "managment", "MaritalStatus": "married", "Education": "teritary", "Default": "no",
"Balance": '2143', "Housing": "yes", "Loan": "no", "Contact": "unknown", "Day": '5', "Month": "may",
"Duration": '261', "Campaign": '1', "PDays": '-1', "Previous": "0", "POutcome": "unknown"}
```
### Make the prediction
Now that your `Model` resource is deployed to an `Endpoint` resource, one can do online predictions by sending prediction requests to the `Endpoint` resource.
#### Request
The format of each instance is:
{[feature_list] }
Since the `predict()` method can take multiple items (instances), send your single test item as a list of one test item.
#### Response
The response from the `predict()` call is a Python dictionary with the following entries:
- `ids`: The internal assigned unique identifiers for each prediction request.
- TODO
- `deployed_model_id`: The Vertex identifier for the deployed `Model` resource which did the predictions.
```
instances_list = [INSTANCE]
prediction = endpoint.predict(instances_list)
print(prediction)
```
## Undeploy the model
When you are done doing predictions, you undeploy the `Model` resource from the `Endpoint` resouce. This deprovisions all compute resources and ends billing for the deployed model.
```
endpoint.undeploy_all()
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex dataset object
try:
if delete_dataset and 'dataset' in globals():
dataset.delete()
except Exception as e:
print(e)
# Delete the model using the Vertex model object
try:
if delete_model and 'model' in globals():
model.delete()
except Exception as e:
print(e)
# Delete the endpoint using the Vertex endpoint object
try:
if delete_endpoint and 'model' in globals():
endpoint.delete()
except Exception as e:
print(e)
# Delete the batch prediction job using the Vertex batch prediction object
try:
if delete_batchjob and 'model' in globals():
batch_predict_job.delete()
except Exception as e:
print(e)
if delete_bucket and 'BUCKET_NAME' in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
# Data Visualization with Python and Jupyter
In this module of the course, we will use some of the libraries available with Python and Jupyter to examine our data set. In order to better understand the data, we can use visualizations such as charts, plots, and graphs. We'll use some commont tools such as [`matplotlib`](https://matplotlib.org/users/index.html) and [`seaborn`](https://seaborn.pydata.org/index.html) and gather some statistical insights into our data.
We'll continue to use the [`insurance.csv`](https://www.kaggle.com/noordeen/insurance-premium-prediction/download) file from you project assets, so if you have not already [`downloaded this file`](https://www.kaggle.com/noordeen/insurance-premium-prediction/download) to your local machine, and uploaded it to your project, do that now.
## Table of Contents
1. [Using the Jupyter notebook](#jupyter)<br>
2. [Load the data](#data)<br>
3. [Visualize Data](#visualize)<br>
4. [Understand Data](#understand)<br>
<a id="jupyter"></a>
## 1. Using the Jupyter notebook
### Jupyter cells
When you are editing a cell in Jupyter notebook, you need to re-run the cell by pressing **`<Shift> + <Enter>`**. This will allow changes you made to be available to other cells.
Use **`<Enter>`** to make new lines inside a cell you are editing.
#### Code cells
Re-running will execute any statements you have written. To edit an existing code cell, click on it.
#### Markdown cells
Re-running will render the markdown text. To edit an existing markdown cell, double-click on it.
<hr>
### Common Jupyter operations
Near the top of the Jupyter notebook page, Jupyter provides a row of menu options (`File`, `Edit`, `View`, `Insert`, ...) and a row of tool bar icons (disk, plus sign, scissors, 2 files, clipboard and file, up arrow, ...).
#### Inserting and removing cells
- Use the "plus sign" icon to insert a cell below the currently selected cell
- Use "Insert" -> "Insert Cell Above" from the menu to insert above
#### Clear the output of all cells
- Use "Kernel" -> "Restart" from the menu to restart the kernel
- click on "clear all outputs & restart" to have all the output cleared
#### Save your notebook file locally
- Clear the output of all cells
- Use "File" -> "Download as" -> "IPython Notebook (.ipynb)" to download a notebook file representing your session
<hr>
<a id="data"></a>
## 2.0 Load the data
A lot of data is **structured data**, which is data that is organized and formatted so it is easily readable, for example a table with variables as columns and records as rows, or key-value pairs in a noSQL database. As long as the data is formatted consistently and has multiple records with numbers, text and dates, you can probably read the data with [Pandas](https://pandas.pydata.org/pandas-docs/stable/index.html), an open-source Python package providing high-performance data manipulation and analysis.
### 2.1 Load our data as a pandas data frame
**<font color='red'><< FOLLOW THE INSTRUCTIONS BELOW TO LOAD THE DATASET >></font>**
* Highlight the cell below by clicking it.
* Click the `10/01` "Find data" icon in the upper right of the notebook.
* Add the locally uploaded file `insurance.csv` by choosing the `Files` tab. Then choose the `insurance.csv`. Click `Insert to code` and choose `Insert Pandas DataFrame`.
* The code to bring the data into the notebook environment and create a Pandas DataFrame will be added to the cell below.
* Run the cell
```
# Place cursor below and insert the Pandas DataFrame for the Insurance Expense data
```
### 2.2 Update the variable for our Pandas dataframe
We'll use the Pandas naming convention df for our DataFrame. Make sure that the cell below uses the name for the dataframe used above. For the locally uploaded file it should look like df_data_1 or df_data_2 or df_data_x.
**<font color='red'><< UPDATE THE VARIABLE ASSIGNMENT TO THE VARIABLE GENERATED ABOVE. >></font>**
```
# Replace data_df_1 with the variable name generated above.
df = df_data_1
```
<a id="visualize"></a>
## 3.0 Visualize Data
Pandas uses [`Matplotlib`](https://matplotlib.org/users/index.html) as the default for visualisations.
In addition, we'll use [`Numpy`](https://numpy.org), which is "The fundamental package for scientific computing with Python".
The conventions when using Jupyter notebooks is to import numpy as `np` and to import matplotlib.pyplot as `plt`. You can call these variables whatever you want, but you will often see them done this way.
Import the packages and also add the magic line starting with `%` to output the charts within the notebook. This is what is known as a [`magic command`](https://ipython.readthedocs.io/en/stable/interactive/magics.html).
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
%matplotlib inline
```
### 3.1 Seaborn
Seaborn is a Python data visualization library based on matplotlib. It is an easy to use visualisation package that works well with Pandas DataFrames.
Below are a few examples using Seaborn.
Refer to this [documentation](https://seaborn.pydata.org/index.html) for information on lots of plots you can create.
```
import seaborn as sns
```
### 3.2 Statistical description
We can use the Pandas method [`describe()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.describe.html) to get some statistics that will later be seen in our visualizations. This will include numeric data, but exclude the categorical fields.
```
df.describe()
```
### Question 1: Is there relationship between BMI and insurance expenses?
We'll explore the data by asking a series of questions (hypothesis). The use of plots can help us to find relationships and correlations.
[`Body Mass Index`](https://www.nhlbi.nih.gov/health/educational/lose_wt/BMI/bmicalc.htm) (BMI) is a measure of body fat based on height and weight that applies to adult men and women. It is often correlated with health outcomes, so let's use a [`Seaborn jointplot`](http://seaborn.pydata.org/generated/seaborn.jointplot.html) with a scatterplot to see if that holds for our data.
```
sns.jointplot(x=df["expenses"], y=df["bmi"], kind="scatter")
plt.show()
```
#### Answer:
It doesn't not appear that there is a good correlation between BMI and the expenses for these patients. We see from the histogram on the right that BMI is normally distributed, and from the histogram on top we see that Expenses are clustered around the lower amounts. It does not look like BMI would be a good predictor of the expenses.
### Question 2: Is there relationship between gender and insurance expenses?
Our next hypothesis might be that there is a correlation between gender and expenses. We can use the [`Seaborn boxplot`](https://seaborn.pydata.org/generated/seaborn.boxplot.html). A boxplot uses quartiles to show how the data is distributed, and will give us a good comparison between the 2 categories represented by `gender`. The horizontal line through our boxes is the median value. The area above the median line is the 3rd quartile, representing the values of the 50th-75th percentiles, and the area below the median line is the 2nd quartile, representing the values of the 25th-50th percentiles. The rest of the data is collapse into lines called "whiskers" and outliers are plotted as single points.
```
plt.figure(figsize = (5, 5))
sns.boxplot(x = 'sex', y = 'expenses', data = df)
```
#### Answer:
On average claims from male and female are the same, and both have approximately the same median (the value in the middle of the distribution. The 3rd quartile is "fatter" for the males, meaning there is a broader distribution of values, and it skews to a higher amount. The 4th quartile also skews higher for the males, so this category contains more of the higher expenses.
### Question 3: Is there relationship between region and claim amount?
Perhaps there is a correlation between the various regions and the insurance expenses. We can once again use a series of boxplots to see the differences betweent the regions.
```
plt.figure(figsize = (10, 5))
sns.boxplot(x = 'region', y = 'expenses', data = df)
```
#### Answer:
In this case we see that the median values across regions are nearly the same. There is some variation for the distribution of expense values, and the southeast reagion has more of the higher values in the 3rd and 4th quartile. The differences aren't particularly large, however, and it is unlikely that region could be a good predictor of expenses.
### Question: Is there relationships between claim amount between smokers and non-smokers?
Given the overwhelming evidence that smoking causes mortality (death) and morbidity (disease), we might guess that there is a relationship betweem insurance claims and smoking.
Let's use a boxplot to examine this.
```
plt.figure(figsize = (5, 5))
sns.boxplot(x = 'smoker', y = 'expenses', data = df)
```
#### Answer:
We can see that the mean, and indeed the entire interquartile range from 25% to 75% is much higher in expense for the smokers than for the non-smokers. It looks like whether or not an individual is a smoker could be a good predictor of insurance expenses.
### Question: is the smoker group well represented?
We'll want to make sure that we have a pretty good sample size for both groups.
```
# make the plot a little bigger
countplt, ax = plt.subplots(figsize = (10,7))
ax = sns.countplot(x='smoker', data=df)
```
#### Answer:
Yes, it looks like smokers are a large enough group to be statistically significant.
### Question: Is there relationship between claim amount and age?
It seems reasonable to assume that there might be different insurance costs for different age groups. For example, older adults tend to require more health care.
Since this is continuous data, let's use a scatter plot to investigate.
```
sns.jointplot(x=df['expenses'], y=df['age'], kind='scatter')
plt.show()
```
#### Answer:
Yes, it does look like Claim amounts increase with age. Furthermore, there are interesting bands around the expenses for `$1,200`, up to `$3,000`, and above `$3,000`.
<a id="understand"></a>
## 4.0 Understand data
Now that we have had a look at the data, let's bring some of this information together.
In order to look at the relationship between multiple variables, we can use the [`Seaborn pairplot()`](https://seaborn.pydata.org/generated/seaborn.pairplot.html) method. This will plot each of the variables of the data set on both the x and y axes, in every possible combination. From this we can quickly see patterns that indicate the relationship between the variables.
We'll use the `hue` to color one of the features in the plot to compare it to the other 2 variables.
### 4.1 Impact of Smoking
See which variable correlate with smoking. `Red` indicates a smoker.
```
claim_pplot=df[['age', 'bmi', 'children', 'smoker', 'expenses']]
claim_pplot.head()
sns.pairplot(claim_pplot, kind="scatter", hue = "smoker" , markers=["o", "s"], palette="Set1")
plt.show()
```
#### Analyis
We can see some interesting things from these plots. Whereas older people tend to have more expenses, we can see from `age` vs. `expenses` that smoking is a more dominant feature. The same holds for `BMI` vs `expenses`.
### 4.2 Impact of Gender
What is the correlation between the features and gender. `Red` is female, `Blue` is male.
```
claim_pplot=df[['age', 'bmi', 'children', 'sex', 'expenses']]
claim_pplot.head()
sns.pairplot(claim_pplot, kind="scatter", hue = "sex" , markers=["o", "s"], palette="Set1")
plt.show()
```
#### Analysis:
Gender has very little impact of the expenses.
#### REGION IMPACT
```
claim_pplot=df[['age', 'bmi', 'children', 'region', 'expenses']]
claim_pplot.head()
sns.pairplot(claim_pplot, kind="scatter", hue = "region" , markers=["o", "s","x","+"], palette="Set1")
plt.show()
```
#### Analysis:
Region does have some imact on the expenses, which can be seen in the `age` vs. `expenses` chart where the `northeast` region appears in the lowest band more commonly, followed by the `northwest` region, and the `southeast` region is clearly higher and more prevelant in the highest band.
### Show correlations
We can quantify the correlations between features of the data set using [`Pandas corr()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.corr.html) method. This will output a table with a numberical value for the correlation coefficient.
```
df[['age', 'sex','bmi', 'children', 'smoker', 'region', 'expenses']].corr(method='pearson')
```
#### Analysis:
We can see from the numerical correlation coefficient that there is little relationship amongst the numerical features.
## Summary:
From our visual analysis of the data, we see that the best predictor of insurance claim expenses is whether or not the individual is a smoker.
| github_jupyter |
# Sampler statistics
When checking for convergence or when debugging a badly behaving
sampler, it is often helpful to take a closer look at what the
sampler is doing. For this purpose some samplers export
statistics for each generated sample.
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sb
import pandas as pd
import pymc3 as pm
%matplotlib inline
```
As a minimal example we sample from a standard normal distribution:
```
model = pm.Model()
with model:
mu1 = pm.Normal("mu1", mu=0, sigma=1, shape=10)
with model:
step = pm.NUTS()
trace = pm.sample(2000, tune=1000, init=None, step=step, cores=2)
```
NUTS provides the following statistics:
```
trace.stat_names
```
- `mean_tree_accept`: The mean acceptance probability for the tree that generated this sample. The mean of these values across all samples but the burn-in should be approximately `target_accept` (the default for this is 0.8).
- `diverging`: Whether the trajectory for this sample diverged. If there are many diverging samples, this usually indicates that a region of the posterior has high curvature. Reparametrization can often help, but you can also try to increase `target_accept` to something like 0.9 or 0.95.
- `energy`: The energy at the point in phase-space where the sample was accepted. This can be used to identify posteriors with problematically long tails. See below for an example.
- `energy_error`: The difference in energy between the start and the end of the trajectory. For a perfect integrator this would always be zero.
- `max_energy_error`: The maximum difference in energy along the whole trajectory.
- `depth`: The depth of the tree that was used to generate this sample
- `tree_size`: The number of leafs of the sampling tree, when the sample was accepted. This is usually a bit less than $2 ^ \text{depth}$. If the tree size is large, the sampler is using a lot of leapfrog steps to find the next sample. This can for example happen if there are strong correlations in the posterior, if the posterior has long tails, if there are regions of high curvature ("funnels"), or if the variance estimates in the mass matrix are inaccurate. Reparametrisation of the model or estimating the posterior variances from past samples might help.
- `tune`: This is `True`, if step size adaptation was turned on when this sample was generated.
- `step_size`: The step size used for this sample.
- `step_size_bar`: The current best known step-size. After the tuning samples, the step size is set to this value. This should converge during tuning.
- `model_logp`: The model log-likelihood for this sample.
If the name of the statistic does not clash with the name of one of the variables, we can use indexing to get the values. The values for the chains will be concatenated.
We can see that the step sizes converged after the 1000 tuning samples for both chains to about the same value. The first 2000 values are from chain 1, the second 2000 from chain 2.
```
plt.plot(trace['step_size_bar'])
```
The `get_sampler_stats` method provides more control over which values should be returned, and it also works if the name of the statistic is the same as the name of one of the variables. We can use the `chains` option, to control values from which chain should be returned, or we can set `combine=False` to get the values for the individual chains:
```
sizes1, sizes2 = trace.get_sampler_stats('depth', combine=False)
fig, (ax1, ax2) = plt.subplots(2, 1, sharex=True, sharey=True)
ax1.plot(sizes1)
ax2.plot(sizes2)
accept = trace.get_sampler_stats('mean_tree_accept', burn=1000)
sb.distplot(accept, kde=False)
accept.mean()
```
Find the index of all diverging transitions:
```
trace['diverging'].nonzero()
```
It is often useful to compare the overall distribution of the
energy levels with the change of energy between successive samples.
Ideally, they should be very similar:
```
energy = trace['energy']
energy_diff = np.diff(energy)
sb.distplot(energy - energy.mean(), label='energy')
sb.distplot(energy_diff, label='energy diff')
plt.legend()
```
If the overall distribution of energy levels has longer tails, the efficiency of the sampler will deteriorate quickly.
## Multiple samplers
If multiple samplers are used for the same model (e.g. for continuous and discrete variables), the exported values are merged or stacked along a new axis.
Note that for the `model_logp` sampler statistic, only the last column (i.e. `trace.get_sampler_stat('model_logp')[-1]`) will be the overall model logp.
```
model = pm.Model()
with model:
mu1 = pm.Bernoulli("mu1", p=0.8)
mu2 = pm.Normal("mu2", mu=0, sigma=1, shape=10)
with model:
step1 = pm.BinaryMetropolis([mu1])
step2 = pm.Metropolis([mu2])
trace = pm.sample(10000, init=None, step=[step1, step2], cores=2, tune=1000)
trace.stat_names
```
Both samplers export `accept`, so we get one acceptance probability for each sampler:
```
trace.get_sampler_stats('accept')
```
| github_jupyter |
# Package Import
```
import numpy as np
import pandas as pd
from sklearn import preprocessing
from sklearn import model_selection
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import precision_recall_fscore_support
from collections import Counter
from imblearn.under_sampling import (ClusterCentroids, RandomUnderSampler,
NearMiss,
InstanceHardnessThreshold,
CondensedNearestNeighbour,
EditedNearestNeighbours,
RepeatedEditedNearestNeighbours,
AllKNN,
NeighbourhoodCleaningRule,
OneSidedSelection)
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.ensemble import GradientBoostingClassifier
```
# Data Cleaning
```
# read raw data
csv = pd.read_csv('NYPD_Arrests_Data__Historic_.csv')
csv.head(3)
# Drop na drop duplicates
data = csv[['ARREST_DATE','PD_DESC','OFNS_DESC','LAW_CAT_CD', 'ARREST_BORO','AGE_GROUP','PERP_SEX','PERP_RACE']].drop_duplicates().dropna()
data = data.reset_index(drop=True)
data['LAW_CAT_CD'].unique()
#Level of offense: felony, misdemeanor, violation
# Because data description did not explain I, we only analyze F, M, V in our project
# drop I
# Get names of indexes for which column CAT_CD is I
indexNames = data[ data['LAW_CAT_CD'] == 'I' ].index
# Delete these row indexes from dataFrame
data.drop(indexNames , inplace=True)
data = data.reset_index(drop=True)
data['ARREST_BORO'].unique()
#Borough of arrest. B(Bronx), S(Staten Island), K(Brooklyn), M(Manhattan), Q(Queens)
data['OFNS_DESC'].unique()
# extract year and month
data['YEAR'] = pd.DatetimeIndex(data['ARREST_DATE']).year
data['MONTH'] = pd.DatetimeIndex(data['ARREST_DATE']).month
# keep only the data from 2008.1 to 2011.7
data3 = data.loc[(data['YEAR'] == 2008)|(data['YEAR'] == 2009)|(data['YEAR'] == 2010)]
data4 = data.loc[(data['YEAR'] == 2011)]
data5 = data4.loc[(data4['MONTH'].isin([1,2,3,4,5,6,7]))]
data2= pd.concat([data3,data5])
data2=data2.reset_index(drop=True)
# save the cleaned data
data2.to_csv('NYCCrime_TimeModified.csv')
```
# Modeling
```
data2 = pd.read_csv('NYCCrime_TimeModified.csv')
data2
Counter(data2['LAW_CAT_CD'])
data2.info()
def labeling(org_label):
if org_label == 'V':
return 0
elif org_label == 'M':
return 1
else:
return 2
##Level of offense: felony (2), misdemeanor (1), violation (0)
data2['LAW_CAT_LABEL'] = data2['LAW_CAT_CD'].apply(labeling)
# Get feature feature space by dropping useless feature
to_drop = ['ARREST_DATE','PD_DESC','OFNS_DESC','LAW_CAT_CD','YEAR','MONTH','LAW_CAT_LABEL','Unnamed: 0']
X = data2.drop(to_drop, axis=1)
y = data2['LAW_CAT_LABEL']
# factors
X['ARREST_BORO'] = preprocessing.LabelEncoder().fit_transform(X['ARREST_BORO'])
X['AGE_GROUP'] = preprocessing.LabelEncoder().fit_transform(X['AGE_GROUP'])
X['PERP_SEX'] = preprocessing.LabelEncoder().fit_transform(X['PERP_SEX'])
X['PERP_RACE'] = preprocessing.LabelEncoder().fit_transform(X['PERP_RACE'])
y
X.duplicated()
# X has duplication
# Splite data into training and testing
# Reserve 25% for testing
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=0.25)
Counter(y_train)
# Random Forest
classifier_RF = RandomForestClassifier()
# fit RF model
classifier_RF.fit(X_train, y_train)
# Prediction of test data
classifier_RF.predict(X_test)
# Accuracy of test data
classifier_RF.score(X_test, y_test)
# get precision, recall, fscore
y_pred = classifier_RF.predict(X_test)
precision_recall_fscore_support(y_test, y_pred,labels=[0,1,2])
```
# Undersampling
```
rus = RandomUnderSampler(random_state=0)
X_resampled, y_resampled = rus.fit_resample(X_train, y_train)
print(sorted(Counter(y_resampled).items()))
# fit model after undersamling
classifier_RF.fit(X_resampled, y_resampled)
# Prediction of test data
classifier_RF.predict(X_test)
# Accuracy of test data
classifier_RF.score(X_test, y_test)
y_pred = classifier_RF.predict(X_test)
precision_recall_fscore_support(y_test, y_pred,labels=[0,1,2])
```
# Gradient Boosting Decision Tree
```
# set learning rate list
lr_list = [0.05, 0.075, 0.1, 0.25, 0.5, 0.75, 1]
for learning_rate in lr_list:
gb_clf = GradientBoostingClassifier(n_estimators=20, learning_rate=learning_rate, max_features=2, max_depth=3, random_state=0)
gb_clf.fit(X_resampled, y_resampled)
print("Learning rate: ", learning_rate)
print("Accuracy score (training): {0:.3f}".format(gb_clf.score(X_resampled, y_resampled)))
print("Accuracy score (validation): {0:.3f}".format(gb_clf.score(X_test, y_test)))
# select the learning rate with the highest accuracy to fit the GBDT model, lr is 0.25
gb_clf2 = GradientBoostingClassifier(n_estimators=20, learning_rate=0.25, max_features=2, max_depth=2, random_state=0)
gb_clf2.fit(X_resampled, y_resampled)
predictions = gb_clf2.predict(X_test)
print("Confusion Matrix:")
print(confusion_matrix(y_test, predictions))
print("Classification Report")
print(classification_report(y_test, predictions))
```
| github_jupyter |
# Operations on word vectors
Welcome to your first assignment of this week!
Because word embeddings are very computionally expensive to train, most ML practitioners will load a pre-trained set of embeddings.
**After this assignment you will be able to:**
- Load pre-trained word vectors, and measure similarity using cosine similarity
- Use word embeddings to solve word analogy problems such as Man is to Woman as King is to ______.
- Modify word embeddings to reduce their gender bias
Let's get started! Run the following cell to load the packages you will need.
```
import numpy as np
from w2v_utils import *
```
Next, lets load the word vectors. For this assignment, we will use 50-dimensional GloVe vectors to represent words. Run the following cell to load the `word_to_vec_map`.
```
words, word_to_vec_map = read_glove_vecs('../../readonly/glove.6B.50d.txt')
```
You've loaded:
- `words`: set of words in the vocabulary.
- `word_to_vec_map`: dictionary mapping words to their GloVe vector representation.
You've seen that one-hot vectors do not do a good job cpaturing what words are similar. GloVe vectors provide much more useful information about the meaning of individual words. Lets now see how you can use GloVe vectors to decide how similar two words are.
# 1 - Cosine similarity
To measure how similar two words are, we need a way to measure the degree of similarity between two embedding vectors for the two words. Given two vectors $u$ and $v$, cosine similarity is defined as follows:
$$\text{CosineSimilarity(u, v)} = \frac {u . v} {||u||_2 ||v||_2} = cos(\theta) \tag{1}$$
where $u.v$ is the dot product (or inner product) of two vectors, $||u||_2$ is the norm (or length) of the vector $u$, and $\theta$ is the angle between $u$ and $v$. This similarity depends on the angle between $u$ and $v$. If $u$ and $v$ are very similar, their cosine similarity will be close to 1; if they are dissimilar, the cosine similarity will take a smaller value.
<img src="images/cosine_sim.png" style="width:800px;height:250px;">
<caption><center> **Figure 1**: The cosine of the angle between two vectors is a measure of how similar they are</center></caption>
**Exercise**: Implement the function `cosine_similarity()` to evaluate similarity between word vectors.
**Reminder**: The norm of $u$ is defined as $ ||u||_2 = \sqrt{\sum_{i=1}^{n} u_i^2}$
```
# GRADED FUNCTION: cosine_similarity
def cosine_similarity(u, v):
"""
Cosine similarity reflects the degree of similariy between u and v
Arguments:
u -- a word vector of shape (n,)
v -- a word vector of shape (n,)
Returns:
cosine_similarity -- the cosine similarity between u and v defined by the formula above.
"""
distance = 0.0
### START CODE HERE ###
# Compute the dot product between u and v (≈1 line)
dot = np.dot(u,v)
# Compute the L2 norm of u (≈1 line)
norm_u = np.sqrt(sum(np.square(u)))
# Compute the L2 norm of v (≈1 line)
norm_v = np.sqrt(sum(np.square(v)))
# Compute the cosine similarity defined by formula (1) (≈1 line)
cosine_similarity = np.divide(dot,(norm_u*norm_v))
### END CODE HERE ###
return cosine_similarity
father = word_to_vec_map["father"]
mother = word_to_vec_map["mother"]
ball = word_to_vec_map["ball"]
crocodile = word_to_vec_map["crocodile"]
france = word_to_vec_map["france"]
italy = word_to_vec_map["italy"]
paris = word_to_vec_map["paris"]
rome = word_to_vec_map["rome"]
print("cosine_similarity(father, mother) = ", cosine_similarity(father, mother))
print("cosine_similarity(ball, crocodile) = ",cosine_similarity(ball, crocodile))
print("cosine_similarity(france - paris, rome - italy) = ",cosine_similarity(france - paris, rome - italy))
```
**Expected Output**:
<table>
<tr>
<td>
**cosine_similarity(father, mother)** =
</td>
<td>
0.890903844289
</td>
</tr>
<tr>
<td>
**cosine_similarity(ball, crocodile)** =
</td>
<td>
0.274392462614
</td>
</tr>
<tr>
<td>
**cosine_similarity(france - paris, rome - italy)** =
</td>
<td>
-0.675147930817
</td>
</tr>
</table>
After you get the correct expected output, please feel free to modify the inputs and measure the cosine similarity between other pairs of words! Playing around the cosine similarity of other inputs will give you a better sense of how word vectors behave.
## 2 - Word analogy task
In the word analogy task, we complete the sentence <font color='brown'>"*a* is to *b* as *c* is to **____**"</font>. An example is <font color='brown'> '*man* is to *woman* as *king* is to *queen*' </font>. In detail, we are trying to find a word *d*, such that the associated word vectors $e_a, e_b, e_c, e_d$ are related in the following manner: $e_b - e_a \approx e_d - e_c$. We will measure the similarity between $e_b - e_a$ and $e_d - e_c$ using cosine similarity.
**Exercise**: Complete the code below to be able to perform word analogies!
```
# GRADED FUNCTION: complete_analogy
def complete_analogy(word_a, word_b, word_c, word_to_vec_map):
"""
Performs the word analogy task as explained above: a is to b as c is to ____.
Arguments:
word_a -- a word, string
word_b -- a word, string
word_c -- a word, string
word_to_vec_map -- dictionary that maps words to their corresponding vectors.
Returns:
best_word -- the word such that v_b - v_a is close to v_best_word - v_c, as measured by cosine similarity
"""
# convert words to lower case
word_a, word_b, word_c = word_a.lower(), word_b.lower(), word_c.lower()
### START CODE HERE ###
# Get the word embeddings e_a, e_b and e_c (≈1-3 lines)
e_a, e_b, e_c = word_to_vec_map[word_a],word_to_vec_map[word_b],word_to_vec_map[word_c]
### END CODE HERE ###
words = word_to_vec_map.keys()
max_cosine_sim = -100 # Initialize max_cosine_sim to a large negative number
best_word = None # Initialize best_word with None, it will help keep track of the word to output
# loop over the whole word vector set
for w in words:
# to avoid best_word being one of the input words, pass on them.
if w in [word_a, word_b, word_c] :
continue
### START CODE HERE ###
# Compute cosine similarity between the vector (e_b - e_a) and the vector ((w's vector representation) - e_c) (≈1 line)
cosine_sim = cosine_similarity((e_b - e_a),(word_to_vec_map[w]-e_c))
# If the cosine_sim is more than the max_cosine_sim seen so far,
# then: set the new max_cosine_sim to the current cosine_sim and the best_word to the current word (≈3 lines)
if cosine_sim > max_cosine_sim:
max_cosine_sim = cosine_sim
best_word = w
### END CODE HERE ###
return best_word
```
Run the cell below to test your code, this may take 1-2 minutes.
```
triads_to_try = [('italy', 'italian', 'spain'), ('india', 'delhi', 'japan'), ('man', 'woman', 'boy'), ('small', 'smaller', 'large')]
for triad in triads_to_try:
print ('{} -> {} :: {} -> {}'.format( *triad, complete_analogy(*triad,word_to_vec_map)))
```
**Expected Output**:
<table>
<tr>
<td>
**italy -> italian** ::
</td>
<td>
spain -> spanish
</td>
</tr>
<tr>
<td>
**india -> delhi** ::
</td>
<td>
japan -> tokyo
</td>
</tr>
<tr>
<td>
**man -> woman ** ::
</td>
<td>
boy -> girl
</td>
</tr>
<tr>
<td>
**small -> smaller ** ::
</td>
<td>
large -> larger
</td>
</tr>
</table>
Once you get the correct expected output, please feel free to modify the input cells above to test your own analogies. Try to find some other analogy pairs that do work, but also find some where the algorithm doesn't give the right answer: For example, you can try small->smaller as big->?.
### Congratulations!
You've come to the end of this assignment. Here are the main points you should remember:
- Cosine similarity a good way to compare similarity between pairs of word vectors. (Though L2 distance works too.)
- For NLP applications, using a pre-trained set of word vectors from the internet is often a good way to get started.
Even though you have finished the graded portions, we recommend you take a look too at the rest of this notebook.
Congratulations on finishing the graded portions of this notebook!
## 3 - Debiasing word vectors (OPTIONAL/UNGRADED)
In the following exercise, you will examine gender biases that can be reflected in a word embedding, and explore algorithms for reducing the bias. In addition to learning about the topic of debiasing, this exercise will also help hone your intuition about what word vectors are doing. This section involves a bit of linear algebra, though you can probably complete it even without being expert in linear algebra, and we encourage you to give it a shot. This portion of the notebook is optional and is not graded.
Lets first see how the GloVe word embeddings relate to gender. You will first compute a vector $g = e_{woman}-e_{man}$, where $e_{woman}$ represents the word vector corresponding to the word *woman*, and $e_{man}$ corresponds to the word vector corresponding to the word *man*. The resulting vector $g$ roughly encodes the concept of "gender". (You might get a more accurate representation if you compute $g_1 = e_{mother}-e_{father}$, $g_2 = e_{girl}-e_{boy}$, etc. and average over them. But just using $e_{woman}-e_{man}$ will give good enough results for now.)
```
g = word_to_vec_map['woman'] - word_to_vec_map['man']
print(g)
```
Now, you will consider the cosine similarity of different words with $g$. Consider what a positive value of similarity means vs a negative cosine similarity.
```
print ('List of names and their similarities with constructed vector:')
# girls and boys name
name_list = ['john', 'marie', 'sophie', 'ronaldo', 'priya', 'rahul', 'danielle', 'reza', 'katy', 'yasmin']
for w in name_list:
print (w, cosine_similarity(word_to_vec_map[w], g))
```
As you can see, female first names tend to have a positive cosine similarity with our constructed vector $g$, while male first names tend to have a negative cosine similarity. This is not suprising, and the result seems acceptable.
But let's try with some other words.
```
print('Other words and their similarities:')
word_list = ['lipstick', 'guns', 'science', 'arts', 'literature', 'warrior','doctor', 'tree', 'receptionist',
'technology', 'fashion', 'teacher', 'engineer', 'pilot', 'computer', 'singer']
for w in word_list:
print (w, cosine_similarity(word_to_vec_map[w], g))
```
Do you notice anything surprising? It is astonishing how these results reflect certain unhealthy gender stereotypes. For example, "computer" is closer to "man" while "literature" is closer to "woman". Ouch!
We'll see below how to reduce the bias of these vectors, using an algorithm due to [Boliukbasi et al., 2016](https://arxiv.org/abs/1607.06520). Note that some word pairs such as "actor"/"actress" or "grandmother"/"grandfather" should remain gender specific, while other words such as "receptionist" or "technology" should be neutralized, i.e. not be gender-related. You will have to treat these two type of words differently when debiasing.
### 3.1 - Neutralize bias for non-gender specific words
The figure below should help you visualize what neutralizing does. If you're using a 50-dimensional word embedding, the 50 dimensional space can be split into two parts: The bias-direction $g$, and the remaining 49 dimensions, which we'll call $g_{\perp}$. In linear algebra, we say that the 49 dimensional $g_{\perp}$ is perpendicular (or "othogonal") to $g$, meaning it is at 90 degrees to $g$. The neutralization step takes a vector such as $e_{receptionist}$ and zeros out the component in the direction of $g$, giving us $e_{receptionist}^{debiased}$.
Even though $g_{\perp}$ is 49 dimensional, given the limitations of what we can draw on a screen, we illustrate it using a 1 dimensional axis below.
<img src="images/neutral.png" style="width:800px;height:300px;">
<caption><center> **Figure 2**: The word vector for "receptionist" represented before and after applying the neutralize operation. </center></caption>
**Exercise**: Implement `neutralize()` to remove the bias of words such as "receptionist" or "scientist". Given an input embedding $e$, you can use the following formulas to compute $e^{debiased}$:
$$e^{bias\_component} = \frac{e \cdot g}{||g||_2^2} * g\tag{2}$$
$$e^{debiased} = e - e^{bias\_component}\tag{3}$$
If you are an expert in linear algebra, you may recognize $e^{bias\_component}$ as the projection of $e$ onto the direction $g$. If you're not an expert in linear algebra, don't worry about this.
<!--
**Reminder**: a vector $u$ can be split into two parts: its projection over a vector-axis $v_B$ and its projection over the axis orthogonal to $v$:
$$u = u_B + u_{\perp}$$
where : $u_B = $ and $ u_{\perp} = u - u_B $
!-->
```
def neutralize(word, g, word_to_vec_map):
"""
Removes the bias of "word" by projecting it on the space orthogonal to the bias axis.
This function ensures that gender neutral words are zero in the gender subspace.
Arguments:
word -- string indicating the word to debias
g -- numpy-array of shape (50,), corresponding to the bias axis (such as gender)
word_to_vec_map -- dictionary mapping words to their corresponding vectors.
Returns:
e_debiased -- neutralized word vector representation of the input "word"
"""
### START CODE HERE ###
# Select word vector representation of "word". Use word_to_vec_map. (≈ 1 line)
e = word_to_vec_map[word]
# Compute e_biascomponent using the formula give above. (≈ 1 line)
e_biascomponent = (np.dot(e,g)*g)/(sum(np.square(g)))
def neutralize(word, g, word_to_vec_map):
"""
Removes the bias of "word" by projecting it on the space orthogonal to the bias axis.
This function ensures that gender neutral words are zero in the gender subspace.
Arguments:
word -- string indicating the word to debias
g -- numpy-array of shape (50,), corresponding to the bias axis (such as gender)
word_to_vec_map -- dictionary mapping words to their corresponding vectors.
Returns:
e_debiased -- neutralized word vector representation of the input "word"
"""
### START CODE HERE ###
# Select word vector representation of "word". Use word_to_vec_map. (≈ 1 line)
e = word_to_vec_map[word]
# Compute e_biascomponent using the formula give above. (≈ 1 line)
# Correct!
e_biascomponent = np.divide(np.dot(e,g),np.linalg.norm(g)**2) * g
# Wrong!
#e_biascomponent = (np.dot(e,g))/(sum(np.square(g)))*g
#e_biascomponent = np.divide(np.dot(e,g),sum(np.square(g)))* g
# Neutralize e by substracting e_biascomponent from it
# e_debiased should be equal to its orthogonal projection. (≈ 1 line)
e_debiased = e - e_biascomponent
### END CODE HERE ###
return e_debiased
# Neutralize e by substracting e_biascomponent from it
# e_debiased should be equal to its orthogonal projection. (≈ 1 line)
e_debiased = e - e_biascomponent
### END CODE HERE ###
return e_debiased
e = "receptionist"
print("cosine similarity between " + e + " and g, before neutralizing: ", cosine_similarity(word_to_vec_map["receptionist"], g))
e_debiased = neutralize("receptionist", g, word_to_vec_map)
print("cosine similarity between " + e + " and g, after neutralizing: ", cosine_similarity(e_debiased, g))
```
**Expected Output**: The second result is essentially 0, up to numerical roundof (on the order of $10^{-17}$).
<table>
<tr>
<td>
**cosine similarity between receptionist and g, before neutralizing:** :
</td>
<td>
0.330779417506
</td>
</tr>
<tr>
<td>
**cosine similarity between receptionist and g, after neutralizing:** :
</td>
<td>
-3.26732746085e-17
</tr>
</table>
### 3.2 - Equalization algorithm for gender-specific words
Next, lets see how debiasing can also be applied to word pairs such as "actress" and "actor." Equalization is applied to pairs of words that you might want to have differ only through the gender property. As a concrete example, suppose that "actress" is closer to "babysit" than "actor." By applying neutralizing to "babysit" we can reduce the gender-stereotype associated with babysitting. But this still does not guarantee that "actor" and "actress" are equidistant from "babysit." The equalization algorithm takes care of this.
The key idea behind equalization is to make sure that a particular pair of words are equi-distant from the 49-dimensional $g_\perp$. The equalization step also ensures that the two equalized steps are now the same distance from $e_{receptionist}^{debiased}$, or from any other work that has been neutralized. In pictures, this is how equalization works:
<img src="images/equalize10.png" style="width:800px;height:400px;">
The derivation of the linear algebra to do this is a bit more complex. (See Bolukbasi et al., 2016 for details.) But the key equations are:
$$ \mu = \frac{e_{w1} + e_{w2}}{2}\tag{4}$$
$$ \mu_{B} = \frac {\mu \cdot \text{bias_axis}}{||\text{bias_axis}||_2^2} *\text{bias_axis}
\tag{5}$$
$$\mu_{\perp} = \mu - \mu_{B} \tag{6}$$
$$ e_{w1B} = \frac {e_{w1} \cdot \text{bias_axis}}{||\text{bias_axis}||_2^2} *\text{bias_axis}
\tag{7}$$
$$ e_{w2B} = \frac {e_{w2} \cdot \text{bias_axis}}{||\text{bias_axis}||_2^2} *\text{bias_axis}
\tag{8}$$
$$e_{w1B}^{corrected} = \sqrt{ |{1 - ||\mu_{\perp} ||^2_2} |} * \frac{e_{\text{w1B}} - \mu_B} {|(e_{w1} - \mu_{\perp}) - \mu_B)|} \tag{9}$$
$$e_{w2B}^{corrected} = \sqrt{ |{1 - ||\mu_{\perp} ||^2_2} |} * \frac{e_{\text{w2B}} - \mu_B} {|(e_{w2} - \mu_{\perp}) - \mu_B)|} \tag{10}$$
$$e_1 = e_{w1B}^{corrected} + \mu_{\perp} \tag{11}$$
$$e_2 = e_{w2B}^{corrected} + \mu_{\perp} \tag{12}$$
**Exercise**: Implement the function below. Use the equations above to get the final equalized version of the pair of words. Good luck!
```
def equalize(pair, bias_axis, word_to_vec_map):
"""
Debias gender specific words by following the equalize method described in the figure above.
Arguments:
pair -- pair of strings of gender specific words to debias, e.g. ("actress", "actor")
bias_axis -- numpy-array of shape (50,), vector corresponding to the bias axis, e.g. gender
word_to_vec_map -- dictionary mapping words to their corresponding vectors
Returns
e_1 -- word vector corresponding to the first word
e_2 -- word vector corresponding to the second word
"""
### START CODE HERE ###
# Step 1: Select word vector representation of "word". Use word_to_vec_map. (≈ 2 lines)
w1, w2 = (pair[0],pair[1])
e_w1, e_w2 = word_to_vec_map[w1], word_to_vec_map[w2] # note [] not (), dictionary key
# Step 2: Compute the mean of e_w1 and e_w2 (≈ 1 line)
mu = (e_w1+e_w2)/2
# Step 3: Compute the projections of mu over the bias axis and the orthogonal axis (≈ 2 lines)
mu_B = np.divide((np.dot(mu, bias_axis)*bias_axis),(np.linalg.norm(bias_axis)**2))
mu_orth = mu -mu_B
# Step 4: Use equations (7) and (8) to compute e_w1B and e_w2B (≈2 lines)
e_w1B = np.divide((np.dot(e_w1,bias_axis)*bias_axis),(np.linalg.norm(bias_axis)**2))
e_w2B = np.divide(np.dot(e_w2,bias_axis)*bias_axis,(np.linalg.norm(bias_axis)**2))
# Step 5: Adjust the Bias part of e_w1B and e_w2B using the formulas (9) and (10) given above (≈2 lines)
corrected_e_w1B = np.sqrt(np.absolute(1-mu_orth))*((e_w1B -mu_B)/(e_w1B -mu_orth)-mu_B)
corrected_e_w2B = np.sqrt(np.absolute(1-mu_orth))*((e_w2B -mu_B)/(e_w2B -mu_orth)-mu_B)
# Step 6: Debias by equalizing e1 and e2 to the sum of their corrected projections (≈2 lines)
e1 = corrected_e_w1B + mu_orth
e2 = corrected_e_w2B + mu_orth
### END CODE HERE ###
return e1, e2
print("cosine similarities before equalizing:")
print("cosine_similarity(word_to_vec_map[\"man\"], gender) = ", cosine_similarity(word_to_vec_map["man"], g))
print("cosine_similarity(word_to_vec_map[\"woman\"], gender) = ", cosine_similarity(word_to_vec_map["woman"], g))
print()
e1, e2 = equalize(("man", "woman"), g, word_to_vec_map)
print("cosine similarities after equalizing:")
print("cosine_similarity(e1, gender) = ", cosine_similarity(e1, g))
print("cosine_similarity(e2, gender) = ", cosine_similarity(e2, g))
e1, e2 = equalize(("man", "woman"), g, word_to_vec_map)
print("cosine similarities after equalizing:")
print("cosine_similarity(e1, gender) = ", cosine_similarity(e1, g))
print("cosine_similarity(e2, gender) = ", cosine_similarity(e2, g))
print("done")
```
**Expected Output**:
cosine similarities before equalizing:
<table>
<tr>
<td>
**cosine_similarity(word_to_vec_map["man"], gender)** =
</td>
<td>
-0.117110957653
</td>
</tr>
<tr>
<td>
**cosine_similarity(word_to_vec_map["woman"], gender)** =
</td>
<td>
0.356666188463
</td>
</tr>
</table>
cosine similarities after equalizing:
<table>
<tr>
<td>
**cosine_similarity(u1, gender)** =
</td>
<td>
-0.700436428931
</td>
</tr>
<tr>
<td>
**cosine_similarity(u2, gender)** =
</td>
<td>
0.700436428931
</td>
</tr>
</table>
Please feel free to play with the input words in the cell above, to apply equalization to other pairs of words.
These debiasing algorithms are very helpful for reducing bias, but are not perfect and do not eliminate all traces of bias. For example, one weakness of this implementation was that the bias direction $g$ was defined using only the pair of words _woman_ and _man_. As discussed earlier, if $g$ were defined by computing $g_1 = e_{woman} - e_{man}$; $g_2 = e_{mother} - e_{father}$; $g_3 = e_{girl} - e_{boy}$; and so on and averaging over them, you would obtain a better estimate of the "gender" dimension in the 50 dimensional word embedding space. Feel free to play with such variants as well.
### Note
* difference between / and np.devide
* difference between * and np.dot
* The L1 norm that is calculated as the sum of the absolute values of the vector.
* The L2 norm that is calculated as the square root of the sum of the squared vector values. np.linalg.norm
* Reference :https://machinelearningmastery.com/vector-norms-machine-learning/
### Congratulations
You have come to the end of this notebook, and have seen a lot of the ways that word vectors can be used as well as modified.
Congratulations on finishing this notebook!
**References**:
- The debiasing algorithm is from Bolukbasi et al., 2016, [Man is to Computer Programmer as Woman is to
Homemaker? Debiasing Word Embeddings](https://papers.nips.cc/paper/6228-man-is-to-computer-programmer-as-woman-is-to-homemaker-debiasing-word-embeddings.pdf)
- The GloVe word embeddings were due to Jeffrey Pennington, Richard Socher, and Christopher D. Manning. (https://nlp.stanford.edu/projects/glove/)
| github_jupyter |
<a href="https://colab.research.google.com/github/Ignvz/copert_5/blob/main/Factores_emision.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#If used from google drive
from google.colab import drive
drive.mount('/content/drive')
%cd "/content/drive/MyDrive/Factores de emision"
!ls
import pandas as pd
pd.options.mode.chained_assignment = None # default='warn'
import numpy as np
def copert5(alpha, beta, gamma, delta, epsilon, zeta, hta, vm, redFactor):
if epsilon*vm**2+zeta*vm+hta == 0:
return 0
else:
return (1-redFactor)*(alpha*vm**2 + beta*vm + gamma + delta/vm)/(epsilon * vm**2 + zeta*vm+ hta)
#cargando. excels
#xls = pd.ExcelFile('Copert5.xlsx')
xls = pd.ExcelFile('1.A.3.b.i-iv Road Transport Appendix 4 Emission Factors 2021.xlsx')
df = pd.read_excel(xls,'HOT_EMISSIONS_PARAMETERS', index_col=None)
#Filtrando usando REGEX a las categorías vehiculares a utilizar
df = df.loc[
((df['Fuel'] == 'Petrol') | (df['Fuel'] == 'Diesel')) &
df.Segment.str.contains('^Medium') | #VLP, TAX
df.Segment.str.contains("^N1-II$") | #VLC
df.Segment.str.contains('^Art*|^Rigid 28 *|^Rigid >32*') | #CAM PES RIGID>28 t and articulated
df.Segment.str.contains('^Rigid 14 *|^Rigid 20 *|^Rigid 26 *') |#CAM MED RIGID 14-28 t
df.Segment.str.contains('^Rigid <=7,5 *|^Rigid 7,5 -*|^Rigid 12 *') |#CAM LIV
df.Segment.str.contains('^Urban Buses Standard 15 *') | #Bus RIG
df.Segment.str.contains('^Urban Buses Articulated >18 t') | #BUS ART
df.Segment.str.contains('^Coaches Standard <=18 t') | #BUS INT PROV
df.Segment.str.contains("^Motorcycles 2-stroke >50 *") #MOT
]
#Cambiando a la clasificacion a utilizar
df.loc[df['Segment'] == 'Medium', 'Segment'] = 'VLP'
df.loc[df['Segment'] == 'N1-II', 'Segment'] = 'VLC'
df.loc[df.Segment.str.contains('^Rigid <=7,5 *|^Rigid 7,5 -*|^Rigid 12 *'), 'Segment'] = 'CAM LIV'
df.loc[df.Segment.str.contains('^Rigid 14 *|^Rigid 20 *|^Rigid 26 *'), 'Segment'] = 'CAM MED'
df.loc[df.Segment.str.contains('^Art*|^Rigid 28 *|^Rigid >32*'), 'Segment'] = 'CAM PES'
df.loc[df.Segment.str.contains('^Urban Buses Standard 15 *'), 'Segment'] = 'BUS RIG'
df.loc[df.Segment.str.contains('^Urban Buses Articulated >18 t'), 'Segment'] = 'BUS ART'
df.loc[df.Segment.str.contains('^Coaches Standard <=18 t'), 'Segment'] = 'BUS INT PROV'
df.loc[df.Segment.str.contains("^Motorcycles 2-stroke >50 *"), 'Segment'] = 'MOT'
#df.to_excel('C5_reducido.xlsx',index = False)
#df.loc[df.Segment.str.contains('^CAM MED*')]
#df = pd.read_excel('C5_reducido_prueba.xlsx')
df = df.loc[ ((df['Load'].isnull())) | #todas las pendientes y las que no tienen pendiente
((df['Load'] == 0.5))] #carga 50% y las que su carga no está definida
#df.to_excel('C5_reducido_prueba.xlsx',index = False)
pd.unique(df["Load"])
df = df.loc[(df["Mode"] == "Highway") | (df["Mode"].isnull()) | (df["Mode"] == "Urban Peak") | (df["Mode"] == "Hot")]
df["Mode"].unique()
#Seteando velocidad, cada tipo de veh. tiene una velocidad máxima y mínima
# BUS INT PROV + CAM LIV/MED velocidad max 90
# CAM PES velocidad max 80
velMin = 20
velMax = 100
df["velMin"] = np.where(df["Min Speed [km/h]"] > velMin, #verificando si es menor que el valor min
df["Min Speed [km/h]"], velMin)
df["velMax"] = np.where((df["Segment"] =="CAM PES"),80, velMax) #seteando velmax como 80 para los CAM PES
df["velMax"] = np.where((df["Segment"] == "BUS INT PROV") | (df["Segment"] == "CAM LIV") | (df["Segment"] == "CAM MED"),
90, df["velMax"])
df["velMax"] = np.where(df["Max Speed [km/h]"] < df["velMax"], #verificando si es mayor que el valor max
df["Max Speed [km/h]"], df["velMax"])
#df.to_excel("revisar.xlsx", index=False)
#df.loc[df["Segment"] == "CAM PES"]["velMax"]
#Cambiando clasificaciones
df.loc[df['Euro Standard'] == 'PRE ECE', 'Euro Standard'] = 'No cat'
df.loc[df['Euro Standard'] == 'Euro 1', 'Euro Standard'] = 'E1'
df.loc[df['Euro Standard'] == 'Euro 2', 'Euro Standard'] = 'E2'
df.loc[df['Euro Standard'] == 'Euro 3', 'Euro Standard'] = 'E3'
df.loc[df['Euro Standard'] == 'Euro 4', 'Euro Standard'] = 'E4'
df.loc[df['Euro Standard'] == 'Euro 5', 'Euro Standard'] = 'E5'
df.loc[df['Euro Standard'] == 'Euro 6 a/b/c', 'Euro Standard'] = 'E6'
df.loc[df['Euro Standard'] == 'Euro 6 d', 'Euro Standard'] = 'E6'
df.loc[df['Euro Standard'] == 'Conventional', 'Euro Standard'] = 'No cat'
df.loc[df['Euro Standard'] == 'Euro I', 'Euro Standard'] = 'E1'
df.loc[df['Euro Standard'] == 'Euro II', 'Euro Standard'] = 'E2'
df.loc[df['Euro Standard'] == 'Euro III', 'Euro Standard'] = 'E3'
df.loc[df['Euro Standard'] == 'Euro IV', 'Euro Standard'] = 'E4'
df.loc[df['Euro Standard'] == 'Euro V', 'Euro Standard'] = 'E5'
df.loc[df['Euro Standard'] == 'Euro VI A/B/C', 'Euro Standard'] = 'E6' #al parecer no hay diferencia en los parámetros estudiados
df.loc[df['Euro Standard'] == 'Euro VI D/E', 'Euro Standard'] = 'E6'
df.loc[df['Fuel'] == 'Petrol', 'Fuel'] = 'Gasolina'
#df.to_excel('C5_reducido_prueba.xlsx',index = False)
df = df.loc[
(df["Euro Standard"] == "No cat") |
(df["Euro Standard"] == "E1") |
(df["Euro Standard"] == "E2") |
(df["Euro Standard"] == "E3") |
(df["Euro Standard"] == "E4") |
(df["Euro Standard"] == "E5") |
(df["Euro Standard"] == "E6") |
(df["Euro Standard"] == "E6")
]
df = df.loc[
#Todo lo en blanco
(df['Technology'].isnull()) |
#Passenger Cars
((df['Category'] == 'Passenger Cars') &
(df['Technology'] == 'PFI') | (df['Technology'] == 'DPF')) |
#Light Commercial Vehicles
((df['Category'] == 'Light Commercial Vehicles') &
(df['Technology'] == 'PFI') | (df['Technology'] == 'DPF')) |
#Buses
((df['Category'] == 'Buses') &
(df['Technology'] == 'EGR') | (df['Technology'] == 'DPF+SCR')) |
#Heavy Duty Trucks
((df['Category'] == 'Heavy Duty Trucks') &
(df['Technology'] == 'EGR') | (df['Technology'] == 'DPF+SCR'))
]
alpha = df[["Alpha"]].astype("float")
beta = df[["Beta"]].astype("float")
gamma = df[["Gamma"]].astype("float")
delta = df[["Delta"]].astype("float")
epsilon = df[["Epsilon"]].astype("float")
zeta = df[["Zita"]].astype("float")
hta = df[["Hta"]].astype("float")
vmUrb = df[["velMin"]].astype("float")
vmInt = df[["velMax"]].astype("float")
redFactor = df[["Reduction Factor [%]"]].astype("float")
#copert5(alpha, beta, gamma, delta, epsilon, zeta, hta, vm, redFactor)
discriminante = (epsilon.values*vmUrb.values**2+zeta.values*vmUrb.values+
hta.values)
df["EF[g/km]_ECF[MJ/km]_Urb"] = np.where(discriminante == 0, 0, (1 - redFactor.values)*((alpha.values * vmUrb.values**2 + beta.values * vmUrb.values
+ gamma.values + (delta.values/vmUrb.values)) / (discriminante)))
discriminante = (epsilon.values*vmInt.values**2+zeta.values*vmInt.values+
hta.values)
df["EF[g/km]_ECF[MJ/km]_Int"] = np.where(discriminante == 0, 0, (1 - redFactor.values)*((alpha.values * vmInt.values**2 + beta.values * vmInt.values
+ gamma.values + (delta.values/vmInt.values)) / (discriminante)))
#Calculando PM para camiones/bus EURO 5 a EURO 6
consumo_Urb = df.loc[
((df["Category"] == "Heavy Duty Trucks") | (df["Category"] == "Buses")) & (df["Pollutant"] == "EC") &
((df["Euro Standard"] == "E5") | (df["Euro Standard"] == "E6"))]
consumo_Int = df.loc[((df["Category"] == "Heavy Duty Trucks") | (df["Category"] == "Buses")) & (df["Pollutant"] == "EC") &
((df["Euro Standard"] == "E5") | (df["Euro Standard"] == "E6"))]
pm_E4E5 = df.loc[((df["Category"] == "Heavy Duty Trucks") | (df["Category"] == "Buses")) & (df["Pollutant"] == "PM Exhaust [g/kWh]")]
#consumo_Urb["EF[g/km]_ECF[MJ/km]_Urb"] * pmUrb["EF[g/km]_ECF[MJ/km]_Urb"]
pm_E4E5 = pm_E4E5.groupby(["Segment","Fuel","Pollutant","Euro Standard", "Technology", "Road Slope"], as_index=False).sum()
consumo_Urb = consumo_Urb.groupby(["Segment","Fuel","Pollutant", "Euro Standard", "Technology", "Road Slope"], as_index=False).sum()
consumo_Int = consumo_Int.groupby(["Segment","Fuel","Pollutant", "Euro Standard", "Technology", "Road Slope"], as_index=False).sum()
pm_E4E5["EF[g/km]_ECF[MJ/km]_Urb"] = consumo_Urb["EF[g/km]_ECF[MJ/km]_Urb"]*0.278*pm_E4E5["EF[g/km]_ECF[MJ/km]_Urb"]
pm_E4E5["EF[g/km]_ECF[MJ/km]_Int"] = consumo_Int["EF[g/km]_ECF[MJ/km]_Int"]*0.278*pm_E4E5["EF[g/km]_ECF[MJ/km]_Int"]
cols = list(df.columns)
#df = df[cols[0:9]+cols[11:19]]
dfCO = df.loc[df['Pollutant'] == 'CO']
dfNOx = df.loc[df['Pollutant'] == 'NOx']
dfVOC = df.loc[df['Pollutant'] == 'VOC']
dfPM = df.loc[df['Pollutant'] == 'PM Exhaust']
dfCH4 = df.loc[df['Pollutant'] == 'CH4']
dfCO2 = df.loc[df['Pollutant'] == 'EC']
dfN2O = df.loc[df['Pollutant'] == 'N2O']
dfNH3 = df.loc[df['Pollutant'] == 'NH3']
#Calculo CO2 y SOx
gasDensidad = 0.730 #Ton/m3
gasPcalor = 11200 #Kcal/kg
dieselPcalor = 10900 #Kcal/kg
dieselDensidad = 0.840 #ton/m3
tJouleAtCal = 2.39*10**(-1) #de joule a cal
kCalLt = (gasDensidad*1000*gasPcalor*0.001)
kCalLt = (dieselDensidad*1000*dieselPcalor*0.001)
######## Fin Datos ##########
####### Urbano #######
FCgas_urb = (1000*dfCO2[["EF[g/km]_ECF[MJ/km]_Urb"]]*tJouleAtCal)/(gasPcalor/1000) #gr/km
FCdies_urb = (1000*dfCO2[["EF[g/km]_ECF[MJ/km]_Urb"]]*tJouleAtCal)/(dieselPcalor/1000) #gr/km
FC_urb = np.where(dfCO2[["Fuel"]]=="Gasolina", FCgas_urb, FCdies_urb) #diferenciando diesel y gasolina gr/km
co2Calc_urb = np.where(dfCO2[["Fuel"]] == "Gasolina",44.0011*(FC_urb/(12.011+1.008*1.8)), 44.0011*(FC_urb/(12.011+1.008*2)))
soxCalc_urb = 2*5*(10**-5)*FC_urb
dfCO2[["km/lt_urb"]] = np.where(dfCO2[["Fuel"]]=="Gasolina", (FC_urb/(gasDensidad*10**3))**-1,(FC_urb/(dieselDensidad*10**3))**-1)
dfCO2[["CO2 g/km_urb"]] = co2Calc_urb
dfCO2[["SOx g/km_urb"]] = soxCalc_urb
#### Interurbano ####
FCgas_int = (1000*dfCO2[["EF[g/km]_ECF[MJ/km]_Int"]]*tJouleAtCal)/(gasPcalor/1000) #gr/km
FCdies_int = (1000*dfCO2[["EF[g/km]_ECF[MJ/km]_Int"]]*tJouleAtCal)/(dieselPcalor/1000) #gr/km
FC_int = np.where(dfCO2[["Fuel"]]=="Gasolina", FCgas_int, FCdies_int) #diferenciando diesel y gasolina gr/km
co2Calc_int = np.where(dfCO2[["Fuel"]] == "Gasolina",44.0011*(FC_int/(12.011+1.008*1.8)), 44.0011*(FC_int/(12.011+1.008*2)))
soxCalc_int = 2*5*(10**-5)*FC_int
dfCO2[["km/lt_int"]] = np.where(dfCO2[["Fuel"]]=="Gasolina", (FC_int/(gasDensidad*10**3))**-1,(FC_int/(dieselDensidad*10**3))**-1)
dfCO2[["CO2 g/km_int"]] = co2Calc_int
dfCO2[["SOx g/km_int"]] = soxCalc_int
#Una vez reducida a lo que se utilizara, se separan dataframes diferentes por contaminante.
#df = pd.read_excel('C5_reducido_prueba.xlsx')
dfCO.to_excel('CO_reducido.xlsx',index = False)
dfNOx.to_excel('NOx_reducido.xlsx',index = False)
dfVOC.to_excel('VOC_reducido.xlsx',index = False)
dfPM.to_excel('PM_reducido.xlsx',index = False)
dfCH4.to_excel('CH4_reducido.xlsx',index = False)
dfCO2.to_excel('CO2_reducido.xlsx',index = False)
def formater(gf,cont):
gf["Gas"] = cont
gf["Unidad"] = "[g/km]"
urb_gf = gf.iloc[:,[0,1,2,22,24,25]]
int_gf = gf.iloc[:,[0,1,2,23,24,25]]
urb_gf["Ambito"] = "Urbano"
int_gf["Ambito"] = "Interurbano"
urb_gf.columns = ["Modo","Motorizacion", "Norma", "2014", "Gas", "Unidad", "Ambito"]
urb_gf = urb_gf[["Ambito", "Modo", "Motorizacion", "Norma", "Gas", "Unidad", "2014"]]
int_gf.columns = ["Modo","Motorizacion", "Norma", "2014", "Gas", "Unidad", "Ambito"]
int_gf = int_gf[["Ambito", "Modo", "Motorizacion", "Norma", "Gas", "Unidad", "2014"]]
gf2 = pd.concat([urb_gf,int_gf], axis=0)
new_col = "R1"
gf2.insert(loc=0, column="Region", value=new_col)
final=gf2.copy()
for i in range(2,17):
new_col = "R"+str(i)
gf2["Region"]=new_col
final = pd.concat([final,gf2],axis=0)
return final
def formater2(gf, cont):
gf["Gas"] = cont
gf["Unidad"] = "[g/km]"
gf_urb = gf.iloc[:,[0,1,2,3,5,6]]
gf_int = gf.iloc[:,[0,1,2,4,5,6]]
gf_urb["Ambito"] = "Urbano"
gf_int["Ambito"] = "Interurbano"
gf_urb.columns = ["Modo","Motorizacion", "Norma", "2014", "Gas", "Unidad", "Ambito"]
gf_urb = gf_urb[["Ambito", "Modo", "Motorizacion", "Norma", "Gas", "Unidad", "2014"]]
gf_int.columns = ["Modo","Motorizacion", "Norma", "2014", "Gas", "Unidad", "Ambito"]
gf_int = gf_int[["Ambito", "Modo", "Motorizacion", "Norma", "Gas", "Unidad", "2014"]]
gf2 = pd.concat([gf_urb,gf_int], axis=0)
new_col = "R1"
gf2.insert(loc=0, column="Region", value=new_col)
final=gf2.copy()
for i in range(2,17):
new_col = "R"+str(i)
gf2["Region"]=new_col
final = pd.concat([final,gf2],axis=0)
return final
### columns_text = ["Region", "Ambito", "Modo", "Motorizacion", "Norma", "Gas", "Unidad"]
#dfNOx[(dfNOx["Segment"]=="BUS ART") & (dfNOx["Euro Standard"]=="E1")]["EF[g/km]_ECF[MJ/km]_Urb"].mean()
gfNOx = dfNOx.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
gfCO = dfCO.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
gfVOC = dfVOC.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
gfPM = dfPM.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
gfCH4 = dfCH4.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
gkpm = pm_E4E5.groupby(["Segment", "Pollutant", "Fuel", 'Euro Standard'],as_index=False).mean()
gfN2O = dfN2O.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
gfNH3 = dfNH3.groupby(["Segment","Fuel","Euro Standard"],as_index=False).mean()
emPME4E5 = gkpm.iloc[:,[0,2,3,23,24]]
#formater(gfNOx, "NOx").to_excel("NOx.xlsx", index=False)
#formater(gfCO, "CO").to_excel("CO.xlsx", index=False)
#formater(gfVOC, "VOC").to_excel("VOC.xlsx", index=False)
#formater(gfPM, "PM25").to_excel("PM25.xlsx", index=False)
#formater(gfCH4, "CH4").to_excel("CH4.xlsx", index=False)
gfNOx = formater(gfNOx, "NOx")
gfCO = formater(gfCO, "CO")
gfVOC = formater(gfVOC, "VOC")
gfCH4 = formater(gfCH4, "CH4")
gfPM = formater(gfPM, "PM25")
gfPM2 = formater2(emPME4E5, "PM25")
gfN2O = formater(gfN2O, "N2O")
gfNH3 = formater(gfNH3, "NH3")
#pd.concat([gfNOx,gfCO,gfCH4,gfVOC, gfPM], axis=0).to_excel("FE.xlsx", index=False)
```
## Calculate N2O and NH3 VLP/VLC
```
#Calculando N2O para VLC/VLP
#ecuacion [a*kilometraje + b]*EFbase
kilometraje = 180000
xls = pd.ExcelFile("input/N2O_VLCVLP.xlsx")
dfN2OVL = pd.read_excel(xls,"0-30", index_col=None)
dfN2OVL["N2O"] = (dfN2OVL["a"]*kilometraje+ dfN2OVL["b"])*dfN2OVL["Base [mg/km]"]/1000
gfN2O_2 = formater2(dfN2OVL.iloc[:,[0,1,2,6,6]],"N2O")
gfN2O_f = pd.concat([gfN2O,gfN2O_2], axis=0)
xls2 = pd.ExcelFile("input/NH3_VLCVLP.xlsx")
dfNH3VL = pd.read_excel(xls2,"0-30", index_col=None)
dfNH3VL["NH3"] = (dfNH3VL["a"]*kilometraje + dfNH3VL["b"])*dfNH3VL["Base [mg/km]"]/1000
gfNH3_2 = formater2(dfNH3VL.iloc[:,[0,1,2,6,6]],"N2O")
gfNH3_f = pd.concat([gfNH3,gfNH3_2], axis=0)
gk = df.groupby(["Segment", "Pollutant", "Fuel", 'Euro Standard'],as_index=False).mean()
gk[["EF[g/km]_ECF[MJ/km]_Urb","EF[g/km]_ECF[MJ/km]_Int", "velMax", "velMin"]].to_excel("Resumen_categoria_combustible.xlsx",index = True)
#df.to_excel('C5_reducido_prueba.xlsx',index = False)
#df2 = pd.read_excel('PM_reducido.xlsx')
gk2 = dfCO2.groupby(["Segment", "Pollutant", "Fuel", 'Euro Standard'],as_index=False).mean()
#gk2.to_excel("ResumenCO2.xlsx", index=True)
#gk[["velMin", "velMax", "EF[g/km]_ECF[MJ/km]_Urb", "EF[g/km]_ECF[MJ/km]_Int"]]
kmLt = gk2.iloc[:,[0,2,3,25,28]]
emCO2 = gk2.iloc[:,[0,2,3,26,29]]
emSOx = gk2.iloc[:,[0,2,3,27,30]]
gfSOx = formater2(emSOx, "SOx")
gfCO2 = formater2(emCO2, "CO2")
gfkmLt = formater2(kmLt, "km/lt")
#(df[df["Segment"]=="CAM PES"]["Max Speed [km/h]"])
gf_concat = pd.concat([gfNOx,gfCO,gfCH4,gfVOC, gfPM, gfPM2, gfSOx, gfCO2, gfkmLt, gfNH3_f, gfN2O_f], axis=0)
#final.to_excel("NOx.xlsx", index=False)
n = 2060-2014
years = (pd.concat([gf_concat[["2014"]]] * n, axis=1, ignore_index=True)).rename(lambda x: str(2015 + x), axis=1)
pd.concat([gf_concat,years], axis=1).to_excel("FE.xlsx", index=False)
```
## Calculate BC
```
df2 = pd.read_excel('input/factores_bc.xlsx')
df2.sort_values("Modo")
pm_concat = pd.concat([gfPM,gfPM2], axis=0)
df3 = pm_concat.merge(df2,left_on=["Modo","Motorizacion","Norma"],right_on=["Modo","Motorizacion","Norma"],how="left")
df3["BC_f"] = df3["2014"] * df3["BC"]
df2
```
# Archivo Velocidades
Ahora realizar el mismo proceso, pero utilizando un excel con las velocidades para cada región y año
```
df2 = pd.read_excel("input/Velocidades.xlsx")
bdf = df2.merge(df, left_on=["Modo", "Motorizacion", "Norma"], right_on=["Segment","Fuel","Euro Standard"], how="left")
alpha = bdf[["Alpha"]].astype("float")
beta = bdf[["Beta"]].astype("float")
gamma = bdf[["Gamma"]].astype("float")
delta = bdf[["Delta"]].astype("float")
epsilon = bdf[["Epsilon"]].astype("float")
zeta = bdf[["Zita"]].astype("float")
hta = bdf[["Hta"]].astype("float")
redFactor = bdf[["Reduction Factor [%]"]].astype("float")
bdf.columns
vel = bdf.iloc[:,7:len(df2.columns)]
bdf.iloc[:,[0,1,2,3,4,-24]]
vel2 = np.where(vel.gt(bdf["Max Speed [km/h]"], axis=0),bdf[["Max Speed [km/h]"]],vel)
vel2 = pd.DataFrame(vel2)
vel2.columns = vel.columns
discriminante = (epsilon.values*vel2**2+zeta.values*vel2+hta.values)
EFs = np.where(discriminante == 0, 0, (1 - redFactor.values)*((alpha.values * vel2**2 + beta.values * vel2
+ gamma.values + (delta.values/vel2)) / (discriminante)))
EFs = pd.DataFrame(EFs)
EFs.columns = vel.columns
EFs = pd.concat([bdf.iloc[:,[0,1,2,3,4,-24]],EFs],axis=1)
EFs = EFs.groupby(["Region","Ambito","Modo","Motorizacion",'Norma',"Pollutant"],as_index=False).mean()
#Calculando CO2, SOx, km/lt
ECs_gas=EFs[(EFs["Pollutant"]=="EC") & (EFs["Motorizacion"] == "Gasolina")].iloc[:,6:-1]
ECs_diesel = EFs[(EFs["Pollutant"]=="EC") & (EFs["Motorizacion"] == "Diesel")].iloc[:,6:-1]
FCgas = (1000*ECs_gas*tJouleAtCal)/(gasPcalor/1000) #gr/km
FCdies = (1000*ECs_diesel*tJouleAtCal)/(dieselPcalor/1000) #gr/km
CO2_gas = 44.0011*(FCgas/(12.011+1.008*1.8))
CO2_dies = 44.0011*(FCdies/(12.011+1.008*2))
SOx_gas = 2*5*(10**-5)*FCgas
SOx_dies = 2*5*(10**-5)*FCdies
kmltGas = (FCgas/(gasDensidad*10**3))**-1
kmltDies = (FCdies/(dieselDensidad*10**3))**-1
namesG = EFs[(EFs["Pollutant"]=="EC") & (EFs["Motorizacion"] == "Gasolina")].iloc[:,0:6]
namesD = EFs[(EFs["Pollutant"]=="EC") & (EFs["Motorizacion"] == "Diesel")].iloc[:,0:6]
CO2 = pd.concat([pd.concat([namesG,CO2_gas],axis=1), pd.concat([namesD,CO2_dies],axis=1)],axis=0)
CO2["Pollutant"] = "CO2"
SOx = pd.concat([pd.concat([namesG,SOx_gas],axis=1), pd.concat([namesD,SOx_dies],axis=1)],axis=0)
SOx["Pollutant"] = "SOx"
kmLt = pd.concat([pd.concat([namesG,kmltGas],axis=1), pd.concat([namesD,kmltDies],axis=1)],axis=0)
kmLt["Pollutant"] = "[km/lt]"
kmLt
#Fixing g/kWh PM to gr/km
consumo = EFs.loc[
((EFs.Modo.str.contains("CAM*")) | (EFs.Modo.str.contains("BUS*"))) & (EFs["Pollutant"] == "EC") &
((EFs["Norma"] == "E5") | (EFs["Norma"] == "E6"))]
pme4e5 = EFs.loc[
((EFs.Modo.str.contains("CAM*")) | (EFs.Modo.str.contains("BUS*"))) & (EFs["Pollutant"] == "PM Exhaust [g/kWh]")]
pme4e5_2 = pme4e5.iloc[:,6:-1]*consumo.iloc[:,6:-1].values*0.278
namesPM = pme4e5.iloc[:,0:6]
pme4e5_3 = pd.concat([namesPM,pme4e5_2],axis= 1, ignore_index=False)
pme4e5_3["Pollutant"] = "PM Exhaust"
kilometraje = pd.read_excel("input/kilometraje.xlsx")
xls = pd.ExcelFile("input/N2O_VLCVLP.xlsx")
dfN2OVL = pd.read_excel(xls,"0-30", index_col=None)
#dfN2OVL["N2O"] = (dfN2OVL["a"]*kilometraje+ dfN2OVL["b"])*dfN2OVL["Base [mg/km]"]/1000
km = kilometraje.loc[((kilometraje["Modo"]=="VLP") | (kilometraje["Modo"]=="VLC"))]
mkm = km.merge(dfN2OVL, left_on=["Modo", "Motorizacion", "Norma"], right_on=["Modo","Motorizacion","Norma"], how="left")
N2O = mkm.iloc[:,7:54].multiply(mkm["a"],axis="index").add(mkm["b"],axis=0).multiply(mkm["Base [mg/km]"],axis="index")/1000
N2O = pd.concat([mkm.iloc[:,0:6],N2O],axis=1)
N2O["Pollutant"]="N2O"
xls2 = pd.ExcelFile("input/NH3_VLCVLP.xlsx")
dfNH3VL = pd.read_excel(xls2,"0-30", index_col=None)
mkm = km.merge(dfNH3VL, left_on=["Modo", "Motorizacion", "Norma"], right_on=["Modo","Motorizacion","Norma"], how="left")
NH3 = mkm.iloc[:,7:54].multiply(mkm["a"],axis="index").add(mkm["b"],axis=0).multiply(mkm["Base [mg/km]"],axis="index")/1000
NH3 = pd.concat([mkm.iloc[:,0:6],NH3],axis=1)
NH3["Pollutant"]="NH3"
pd.concat([EFs,CO2,SOx,kmLt,pme4e5_3,N2O,NH3],axis=0).to_excel("EF.xlsx")
```
| github_jupyter |
# Horse or Human? In-graph training loop Assignment
This assignment lets you practice how to train a Keras model on the [horses_or_humans](https://www.tensorflow.org/datasets/catalog/horses_or_humans) dataset with the entire training process performed in graph mode. These steps include:
- loading batches
- calculating gradients
- updating parameters
- calculating validation accuracy
- repeating the loop until convergence
## Setup
Import TensorFlow 2.0:
```
from __future__ import absolute_import, division, print_function, unicode_literals
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_hub as hub
import matplotlib.pyplot as plt
```
### Prepare the dataset
Load the horses to human dataset, splitting 80% for the training set and 20% for the test set.
```
splits, info = tfds.load('horses_or_humans', as_supervised=True, with_info=True, split=['train[:80%]', 'train[80%:]', 'test'], data_dir='./data')
(train_examples, validation_examples, test_examples) = splits
num_examples = info.splits['train'].num_examples
num_classes = info.features['label'].num_classes
BATCH_SIZE = 32
IMAGE_SIZE = 224
```
## Pre-process an image (please complete this section)
You'll define a mapping function that resizes the image to a height of 224 by 224, and normalizes the pixels to the range of 0 to 1. Note that pixels range from 0 to 255.
- You'll use the following function: [tf.image.resize](https://www.tensorflow.org/api_docs/python/tf/image/resize) and pass in the (height,width) as a tuple (or list).
- To normalize, divide by a floating value so that the pixel range changes from [0,255] to [0,1].
```
# Create a autograph pre-processing function to resize and normalize an image
### START CODE HERE ###
@tf.function
def map_fn(img, label):
image_height = 224
image_width = 224
### START CODE HERE ###
# resize the image
img = tf.image.resize(img, (image_height, image_width))
# normalize the image
img /= 255.0
### END CODE HERE
return img, label
## TEST CODE:
test_image, test_label = list(train_examples)[0]
test_result = map_fn(test_image, test_label)
print(test_result[0].shape)
print(test_result[1].shape)
del test_image, test_label, test_result
```
**Expected Output:**
```
(224, 224, 3)
()
```
## Apply pre-processing to the datasets (please complete this section)
Apply the following steps to the training_examples:
- Apply the `map_fn` to the training_examples
- Shuffle the training data using `.shuffle(buffer_size=)` and set the buffer size to the number of examples.
- Group these into batches using `.batch()` and set the batch size given by the parameter.
Hint: You can look at how validation_examples and test_examples are pre-processed to get a sense of how to chain together multiple function calls.
```
# Prepare train dataset by using preprocessing with map_fn, shuffling and batching
def prepare_dataset(train_examples, validation_examples, test_examples, num_examples, map_fn, batch_size):
### START CODE HERE ###
train_ds = train_examples.map(map_fn).shuffle(buffer_size=num_examples).batch(batch_size)
### END CODE HERE ###
valid_ds = validation_examples.map(map_fn).batch(batch_size)
test_ds = test_examples.map(map_fn).batch(batch_size)
return train_ds, valid_ds, test_ds
train_ds, valid_ds, test_ds = prepare_dataset(train_examples, validation_examples, test_examples, num_examples, map_fn, BATCH_SIZE)
## TEST CODE:
test_train_ds = list(train_ds)
print(len(test_train_ds))
print(test_train_ds[0][0].shape)
del test_train_ds
```
**Expected Output:**
```
26
(32, 224, 224, 3)
```
### Define the model
```
MODULE_HANDLE = 'data/resnet_50_feature_vector'
model = tf.keras.Sequential([
hub.KerasLayer(MODULE_HANDLE, input_shape=(IMAGE_SIZE, IMAGE_SIZE, 3)),
tf.keras.layers.Dense(num_classes, activation='softmax')
])
model.summary()
```
## Define optimizer: (please complete these sections)
Define the [Adam optimizer](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adam) that is in the tf.keras.optimizers module.
```
def set_adam_optimizer():
### START CODE HERE ###
# Define the adam optimizer
optimizer = tf.keras.optimizers.Adam()
### END CODE HERE ###
return optimizer
## TEST CODE:
test_optimizer = set_adam_optimizer()
print(type(test_optimizer))
del test_optimizer
```
**Expected Output:**
```
<class 'tensorflow.python.keras.optimizer_v2.adam.Adam'>
```
## Define the loss function (please complete this section)
Define the loss function as the [sparse categorical cross entropy](https://www.tensorflow.org/api_docs/python/tf/keras/losses/SparseCategoricalCrossentropy) that's in the tf.keras.losses module. Use the same function for both training and validation.
```
def set_sparse_cat_crossentropy_loss():
### START CODE HERE ###
# Define object oriented metric of Sparse categorical crossentropy for train and val loss
train_loss = tf.keras.losses.SparseCategoricalCrossentropy()
val_loss = tf.keras.losses.SparseCategoricalCrossentropy()
### END CODE HERE ###
return train_loss, val_loss
## TEST CODE:
test_train_loss, test_val_loss = set_sparse_cat_crossentropy_loss()
print(type(test_train_loss))
print(type(test_val_loss))
del test_train_loss, test_val_loss
```
**Expected Output:**
```
<class 'tensorflow.python.keras.losses.SparseCategoricalCrossentropy'>
<class 'tensorflow.python.keras.losses.SparseCategoricalCrossentropy'>
```
## Define the acccuracy function (please complete this section)
Define the accuracy function as the [spare categorical accuracy](https://www.tensorflow.org/api_docs/python/tf/keras/metrics/SparseCategoricalAccuracy) that's contained in the tf.keras.metrics module. Use the same function for both training and validation.
```
def set_sparse_cat_crossentropy_accuracy():
### START CODE HERE ###
# Define object oriented metric of Sparse categorical accuracy for train and val accuracy
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy()
val_accuracy = tf.keras.metrics.SparseCategoricalAccuracy()
### END CODE HERE ###
return train_accuracy, val_accuracy
## TEST CODE:
test_train_accuracy, test_val_accuracy = set_sparse_cat_crossentropy_accuracy()
print(type(test_train_accuracy))
print(type(test_val_accuracy))
del test_train_accuracy, test_val_accuracy
```
**Expected Output:**
```
<class 'tensorflow.python.keras.metrics.SparseCategoricalAccuracy'>
<class 'tensorflow.python.keras.metrics.SparseCategoricalAccuracy'>
```
Call the three functions that you defined to set the optimizer, loss and accuracy
```
optimizer = set_adam_optimizer()
train_loss, val_loss = set_sparse_cat_crossentropy_loss()
train_accuracy, val_accuracy = set_sparse_cat_crossentropy_accuracy()
```
### Define the training loop (please complete this section)
In the training loop:
- Get the model predictions: use the model, passing in the input `x`
- Get the training loss: Call `train_loss`, passing in the true `y` and the predicted `y`.
- Calculate the gradient of the loss with respect to the model's variables: use `tape.gradient` and pass in the loss and the model's `trainable_variables`.
- Optimize the model variables using the gradients: call `optimizer.apply_gradients` and pass in a `zip()` of the two lists: the gradients and the model's `trainable_variables`.
- Calculate accuracy: Call `train_accuracy`, passing in the true `y` and the predicted `y`.
```
# this code uses the GPU if available, otherwise uses a CPU
device = '/gpu:0' if tf.config.list_physical_devices('GPU') else '/cpu:0'
EPOCHS = 2
# Custom training step
def train_one_step(model, optimizer, x, y, train_loss, train_accuracy):
'''
Trains on a batch of images for one step.
Args:
model (keras Model) -- image classifier
optimizer (keras Optimizer) -- optimizer to use during training
x (Tensor) -- training images
y (Tensor) -- training labels
train_loss (keras Loss) -- loss object for training
train_accuracy (keras Metric) -- accuracy metric for training
'''
with tf.GradientTape() as tape:
### START CODE HERE ###
# Run the model on input x to get predictions
predictions = model(x)
# Compute the training loss using `train_loss`, passing in the true y and the predicted y
loss = train_loss(y, predictions)
# Using the tape and loss, compute the gradients on model variables using tape.gradient
grads = tape.gradient(loss, model.trainable_weights)
# Zip the gradients and model variables, and then apply the result on the optimizer
optimizer.apply_gradients(zip(grads, model.trainable_weights))
# Call the train accuracy object on ground truth and predictions
train_accuracy(y, predictions)
### END CODE HERE
return loss
## TEST CODE:
def base_model():
inputs = tf.keras.layers.Input(shape=(2))
x = tf.keras.layers.Dense(64, activation='relu')(inputs)
outputs = tf.keras.layers.Dense(1, activation='sigmoid')(x)
model = tf.keras.Model(inputs=inputs, outputs=outputs)
return model
test_model = base_model()
test_optimizer = set_adam_optimizer()
test_image = tf.ones((2,2))
test_label = tf.ones((1,))
test_train_loss, _ = set_sparse_cat_crossentropy_loss()
test_train_accuracy, _ = set_sparse_cat_crossentropy_accuracy()
test_result = train_one_step(test_model, test_optimizer, test_image, test_label, test_train_loss, test_train_accuracy)
print(test_result)
del test_result, test_model, test_optimizer, test_image, test_label, test_train_loss, test_train_accuracy
```
**Expected Output:**
You will see a Tensor with the same shape and dtype. The value might be different.
```
tf.Tensor(0.6931472, shape=(), dtype=float32)
```
## Define the 'train' function (please complete this section)
You'll first loop through the training batches to train the model. (Please complete these sections)
- The `train` function will use a for loop to iteratively call the `train_one_step` function that you just defined.
- You'll use `tf.print` to print the step number, loss, and train_accuracy.result() at each step. Remember to use tf.print when you plan to generate autograph code.
Next, you'll loop through the batches of the validation set to calculation the validation loss and validation accuracy. (This code is provided for you). At each iteration of the loop:
- Use the model to predict on x, where x is the input from the validation set.
- Use val_loss to calculate the validation loss between the true validation 'y' and predicted y.
- Use val_accuracy to calculate the accuracy of the predicted y compared to the true y.
Finally, you'll print the validation loss and accuracy using tf.print. (Please complete this section)
- print the final `loss`, which is the validation loss calculated by the last loop through the validation dataset.
- Also print the val_accuracy.result().
**HINT**
If you submit your assignment and see this error for your stderr output:
```
Cannot convert 1e-07 to EagerTensor of dtype int64
```
Please check your calls to train_accuracy and val_accuracy to make sure that you pass in the true and predicted values in the correct order (check the documentation to verify the order of parameters).
```
# Decorate this function with tf.function to enable autograph on the training loop
@tf.function
def train(model, optimizer, epochs, device, train_ds, train_loss, train_accuracy, valid_ds, val_loss, val_accuracy):
'''
Performs the entire training loop. Prints the loss and accuracy per step and epoch.
Args:
model (keras Model) -- image classifier
optimizer (keras Optimizer) -- optimizer to use during training
epochs (int) -- number of epochs
train_ds (tf Dataset) -- the train set containing image-label pairs
train_loss (keras Loss) -- loss function for training
train_accuracy (keras Metric) -- accuracy metric for training
valid_ds (Tensor) -- the val set containing image-label pairs
val_loss (keras Loss) -- loss object for validation
val_accuracy (keras Metric) -- accuracy metric for validation
'''
step = 0
loss = 0.0
for epoch in range(epochs):
for x, y in train_ds:
# training step number increments at each iteration
step += 1
with tf.device(device_name=device):
### START CODE HERE ###
# Run one training step by passing appropriate model parameters
# required by the function and finally get the loss to report the results
loss = train_one_step(model, optimizer, x, y, train_loss, train_accuracy)
### END CODE HERE ###
# Use tf.print to report your results.
# Print the training step number, loss and accuracy
tf.print('Step', step,
': train loss', loss,
'; train accuracy', train_accuracy.result())
with tf.device(device_name=device):
for x, y in valid_ds:
# Call the model on the batches of inputs x and get the predictions
y_pred = model(x)
loss = val_loss(y, y_pred)
val_accuracy(y, y_pred)
# Print the validation loss and accuracy
### START CODE HERE ###
tf.print('val loss', loss, '; val accuracy', val_accuracy.result())
### END CODE HERE ###
```
Run the `train` function to train your model! You should see the loss generally decreasing and the accuracy increasing.
**Note**: **Please let the training finish before submitting** and **do not** modify the next cell. It is required for grading. This will take around 5 minutes to run.
```
train(model, optimizer, EPOCHS, device, train_ds, train_loss, train_accuracy, valid_ds, val_loss, val_accuracy)
```
# Evaluation
You can now see how your model performs on test images. First, let's load the test dataset and generate predictions:
```
test_imgs = []
test_labels = []
predictions = []
with tf.device(device_name=device):
for images, labels in test_ds:
preds = model(images)
preds = preds.numpy()
predictions.extend(preds)
test_imgs.extend(images.numpy())
test_labels.extend(labels.numpy())
```
Let's define a utility function for plotting an image and its prediction.
```
# Utilities for plotting
class_names = ['horse', 'human']
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
img = np.squeeze(img)
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
# green-colored annotations will mark correct predictions. red otherwise.
if predicted_label == true_label:
color = 'green'
else:
color = 'red'
# print the true label first
print(true_label)
# show the image and overlay the prediction
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
```
### Plot the result of a single image
Choose an index and display the model's prediction for that image.
```
# Visualize the outputs
# you can modify the index value here from 0 to 255 to test different images
index = 87
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(index, predictions, test_labels, test_imgs)
plt.show()
```
| github_jupyter |
# Baselines for Bayes Minimum Risk Pruning
```
# here we comparet the performance of
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score, recall_score,\
accuracy_score, f1_score, classification_report, confusion_matrix, roc_auc_score
import numpy as np
from joblib import Parallel, delayed
from c45 import C45
mainDir = "./ahmedDatasets/"
```
### Zoo Data
```
zooDir = mainDir + "Zoo/zoo.data"
zooData = pd.read_csv(zooDir,delimiter=",",header=None)
zooData.head()
zooTree = C45()
zooTrainX, zooTestX, zooTrainY, zooTestY = train_test_split(zooData[range(1,17)],
zooData[17],
test_size=0.4)
zooTree.fit(zooTrainX, zooTrainY)
accuracy_score(zooTree.predict(zooTestX),zooTestY)
zooRoot = zooTree.getTree()
%%time
import copy
import pickle
def makePhishFile(root,data,outCol):
terminals = []
d = dict()
reverseD = dict()
def isLeaf(node):
children = [child for child in node]
return len(children) == 0
def modifyDict(root,terminals,d,reverseD):
if (isLeaf(root)):
d[root] = []
else:
children = [child for child in root]
for child in children:
if d.get(root) is None:
d[root] = [child]
else:
d[root].append(child)
if reverseD.get(child) is None:
reverseD[child] = [root]
else:
reverseD[child].append(root)
if (not isLeaf(child)):
modifyDict(child,terminals,d,reverseD)
else:
modifyDict(child,
terminals.append(child),
d,
reverseD)
def populateDict(mainRoot,d,reverseD,terminals,df,output):
root = copy.deepcopy(mainRoot)
top = root
modifyDict(root,terminals,d,reverseD)
return np.array([d,
top,
reverseD,
terminals,output,
df], dtype=object)
values = populateDict(root,d,reverseD,terminals,data, outCol)
pickle.dump(values, open('uciPhishGraph.p','wb'))
makePhishFile(zooRoot,zooData,17)
%%time
import sys
import pickle
import time
import numpy as np
from functools import reduce
'''
Takes in a dictionary representation of node: [list of node's neighbors],
Each node has only 1 parent.
'''
def getCategoryProportions(colData):
categoryDict = dict()
for i in range(len(colData)):
if (categoryDict.get(colData[i]) is None):
categoryDict[colData[i]] = [[i],1]
else:
indices = categoryDict[colData[i]][0]
numPerClass = categoryDict[colData[i]][1]
categoryDict[colData[i]] = [indices+[i],numPerClass+1]
# classDict is a dictionary containing categories as keys
# and the values are two-element lists where the first element is
# the list of indices where that category exists and the second
# element is the proportion of the category in the column of data
outputClasses = []
for key in categoryDict:
outputClasses.append(key)
categoryDict[key] = [categoryDict[key][0],
float(categoryDict[key][1])/len(colData)]
return (categoryDict, outputClasses)
class Node:
def __init__(self, children, splitFeature,
splitChildren, output, label = None, depth = 0):
self.children = children
self.splitFeature = splitFeature
self.splitChildren = splitChildren
self.output = output
self.label = label
self.depth = depth
class Graph(object):
def __init__(self, dagList, parentList, root, terminals,outAttribute,trainData):
self.root = root
self.terminals = terminals
self.dagList = dagList
self.parentList = parentList
self.outAttribute = outAttribute
self.trainData = trainData
def preprocessNodesBelow(self):
nodeDict = dict()
i = 0
for node in self.dagList:
lstNodes = []
self.lookBelow(node,lstNodes)
nodeDict[node] = set(lstNodes)
i += 1
return nodeDict
def lookBelow(self, node, lstNodes):
for child in self.dagList[node]:
lstNodes.append(child)
self.lookBelow(child, lstNodes)
# destructively modifies the graph
# we prune the graph over all training examples
# when "turning" a parent into a leaf, the dagList[parent] = []
def runPruning(self,par=True):
def modifyDict(node,catProps):
if len(node.output) != 0:
catProps[node] = getCategoryProportions(node.output)
else:
catProps[node] = None,None
parents = self.parentList
currTerminals = set(self.terminals)
nodeBelowDict = self.preprocessNodesBelow()
self.catProps = dict()
_ = [modifyDict(node,self.catProps) \
for node in self.dagList]
if par:
(currTerminals) = \
self.parPruning(currTerminals,
nodeBelowDict)
else:
(currGraph, parents, currTerminals) = \
self.seqPruning(currGraph,
currFrontier,
parents,
currTerminals,
nodeBelowDict)
# leaves should be kept track of.
def getBayesRisk(self, node):
risk = 0
examples = node.output
for trueClass in examples:
probDict, outputClasses = self.catProps[node]
if probDict is None and outputClasses is None:
return 0
for _,c in enumerate(outputClasses):
if trueClass != c:
risk += abs(float(trueClass)-float(c))*(probDict[c][1])
return risk
# when the root is reached, all leaves are considered.
def seqPruning(self,currGraph,currFrontier,parents, currTerminals, nodeBelowDict):
def getRisks(node,leaves):
return (self.getBayesRisk(node),\
sum(map(lambda leaf: self.getBayesRisk(leaf), leaves)))
def riskFrontier(node, currTerminals, nodeBelowDict):
leaves = \
list(nodeBelowDict[node].intersection(currTerminals))
(parentRisk,leavesRisk) = getRisks(node,leaves)
if (parentRisk < leavesRisk):
currGraph[node] = []
currTerminals = currTerminals.difference(leaves)
currTerminals.add(node)
numRound = 0
while (len(currFrontier) > 1):
start = time.time()
_ = [riskFrontier(node, currTerminals, nodeBelowDict) \
for node in currFrontier]
nextFrontier = set()
# moving onto the next frontier from current frontier
for node in currFrontier:
if node == self.root:
parent = node
else:
parent = self.parentList[node][0]
nextFrontier.add(parent)
currFrontier = nextFrontier
print("{} & {} & {}".format(numRound,\
len(currFrontier),\
time.time()-start))
numRound += 1
return (currGraph, parents, currTerminals)
def parPruning(self,currTerminals, nodeBelowDict):
def getRisks(node,leaves):
return (self.getBayesRisk(node),\
sum(map(lambda leaf: self.getBayesRisk(leaf), leaves)))
def riskFrontier(node, currTerminals, nodeBelowDict):
leaves = \
list(nodeBelowDict[node].intersection(currTerminals))
(parentRisk,leavesRisk) = getRisks(node,leaves)
if (parentRisk < leavesRisk):
self.dagList[node] = []
currTerminals = currTerminals.difference(leaves)
currTerminals.add(node)
numRound = 0
while (len(self.terminals) > 1):
start = time.time()
_ = Parallel(n_jobs=50,prefer="threads")(delayed(riskFrontier)(node, currTerminals, nodeBelowDict) \
for node in self.terminals)
nextFrontier = set()
# moving onto the next frontier from current frontier
for node in self.terminals:
if node == self.root:
parent = node
else:
parent = self.parentList[node][0]
nextFrontier.add(parent)
self.terminals = nextFrontier
print("{} & {} & {}".format(numRound,\
len(self.terminals),\
time.time()-start))
numRound += 1
return currTerminals
def addToGraph(currGraph, node, d):
d[node] = currGraph[node]
for otherNode in currGraph[node]:
addToGraph(currGraph, otherNode, d)
print("Benchmarking...")
with open('uciPhishGraph.p','rb') as f:
graphData = pickle.load(f)
dagList,root,parentList,terminals,outAttribute,trainData = graphData
g = Graph(dagList, parentList, root, terminals, outAttribute, trainData)
print("Starting...")
g.runPruning()
currGraph = g.dagList
newGraph = dict()
addToGraph(currGraph, g.root, newGraph)
import sys
import pickle
import time
import numpy as np
from functools import reduce
import multiprocessing as mp
from multiprocessing import Manager
'''
Takes in a dictionary representation of node: [list of node's neighbors],
Each node has only 1 parent.
'''
def getCategoryProportions(colData):
categoryDict = dict()
for i in range(len(colData)):
if (categoryDict.get(colData[i]) is None):
categoryDict[colData[i]] = [[i],1]
else:
indices = categoryDict[colData[i]][0]
numPerClass = categoryDict[colData[i]][1]
categoryDict[colData[i]] = [indices+[i],numPerClass+1]
# classDict is a dictionary containing categories as keys
# and the values are two-element lists where the first element is
# the list of indices where that category exists and the second
# element is the proportion of the category in the column of data
outputClasses = []
for key in categoryDict:
outputClasses.append(key)
categoryDict[key] = [categoryDict[key][0],
float(categoryDict[key][1])/len(colData)]
return (categoryDict, outputClasses)
class Node:
def __init__(self, children, splitFeature,
splitChildren, output, label = None, depth = 0):
self.children = children
self.splitFeature = splitFeature
self.splitChildren = splitChildren
self.output = output
self.label = label
self.depth = depth
class Graph(object):
def __init__(self, dagList, parentList, root, terminals,outAttribute,trainData):
self.root = root
self.terminals = terminals
self.dagList = dagList
self.parentList = parentList
self.outAttribute = outAttribute
self.trainData = trainData
def preprocessNodesBelow(self):
nodeDict = dict()
i = 0
for node in self.dagList:
lstNodes = []
self.lookBelow(node,lstNodes)
nodeDict[node] = set(lstNodes)
i += 1
return nodeDict
def lookBelow(self, node, lstNodes):
for child in self.dagList[node]:
lstNodes.append(child)
self.lookBelow(child, lstNodes)
# destructively modifies the graph
# we prune the graph over all training examples
# when "turning" a parent into a leaf, the dagList[parent] = []
def runPruning(self,par=True):
def modifyDict(node,catProps):
if len(node.attrib) != 0:
catProps[node] = getCategoryProportions(node.attrib['son_category'])
else:
catProps[node] = None,None
parents = self.parentList
currTerminals = set(self.terminals)
nodeBelowDict = self.preprocessNodesBelow()
self.catProps = dict()
_ = [modifyDict(node,self.catProps) \
for node in self.dagList]
if par:
currTerminals = \
self.parPruning(currTerminals,
nodeBelowDict)
else:
(currGraph, parents, currTerminals) = \
self.seqPruning(currGraph,
currFrontier,
parents,
currTerminals,
nodeBelowDict)
# leaves should be kept track of.
def getBayesRisk(self, node):
risk = 0
if len(node.attrib) == 0:
return sum([self.getBayesRisk(child) for child in node])
examples = node.attrib['son_category']
for trueClass in examples:
probDict, outputClasses = self.catProps[node]
if probDict is None and outputClasses is None:
return 0
for _,c in enumerate(outputClasses):
if trueClass != c:
risk += abs(float(trueClass)-float(c))*(probDict[c][1])
return risk
# when the root is reached, all leaves are considered.
def seqPruning(self,currGraph,currFrontier,parents, currTerminals, nodeBelowDict):
def getRisks(node,leaves):
return (self.getBayesRisk(node),\
sum(map(lambda leaf: self.getBayesRisk(leaf), leaves)))
def riskFrontier(node, currTerminals, nodeBelowDict):
leaves = \
list(nodeBelowDict[node].intersection(currTerminals))
(parentRisk,leavesRisk) = getRisks(node,leaves)
if (parentRisk < leavesRisk):
currGraph[node] = []
currTerminals = currTerminals.difference(leaves)
currTerminals.add(node)
numRound = 0
while (len(currFrontier) > 1):
start = time.time()
_ = [riskFrontier(node, currTerminals, nodeBelowDict) \
for node in currFrontier]
nextFrontier = set()
# moving onto the next frontier from current frontier
for node in currFrontier:
if node == self.root:
parent = node
else:
parent = self.parentList[node][0]
nextFrontier.add(parent)
currFrontier = nextFrontier
print("{} & {} & {}".format(numRound,\
len(currFrontier),\
time.time()-start))
numRound += 1
return (currGraph, parents, currTerminals)
def parPruning(self,currTerminals, nodeBelowDict):
def getRisks(node,leaves):
return (self.getBayesRisk(node),\
sum(map(lambda leaf: self.getBayesRisk(leaf), leaves)))
def riskFrontier(node, currTerminals, nodeBelowDict):
leaves = \
list(nodeBelowDict[node].intersection(currTerminals))
(parentRisk,leavesRisk) = getRisks(node,leaves)
if (parentRisk > leavesRisk):
self.dagList[node] = []
currTerminals = currTerminals.difference(leaves)
currTerminals.add(node)
numRound = 0
while (len(currFrontier) > 1):
start = time.time()
_ = Parallel(n_jobs=50,prefer="threads")(delayed(riskFrontier)(node, currTerminals, nodeBelowDict) \
for node in self.terminals)
nextFrontier = set()
# moving onto the next frontier from current frontier
for node in self.terminals:
if node == self.root:
parent = node
else:
parent = self.parentList[node][0]
nextFrontier.add(parent)
self.terminals = nextFrontier
print("{} & {} & {}".format(numRound,\
len(self.terminals),\
time.time()-start))
numRound += 1
return (currGraph, currTerminals)
def addToGraph(currGraph, node, d):
d[node] = currGraph[node]
for otherNode in currGraph[node]:
addToGraph(currGraph, otherNode, d)
#ray.init()
print("Benchmarking...")
with open('uciPhishGraph.p','rb') as f:
graphData = pickle.load(f)
dagList,root,parentList,terminals,outAttribute,trainData = graphData
print(dagList)
g = Graph(dagList, parentList, root, terminals, outAttribute, trainData)
print("Starting...")
g.runPruning()
currGraph = g.dagList
newGraph = dict()
addToGraph(currGraph, g.root, newGraph)
print(newGraph)
```
### Iris Data
```
irisDir = mainDir + "Iris/iris.data"
irisData = pd.read_csv(irisDir, delimiter = ",", header=None)
irisData.head()
irisesDict = {'Iris-setosa':0, 'Iris-versicolor':1, 'Iris-virginica':2}
irisData[5] = irisData[4].apply(lambda row: irisesDict[row])
irisData.head()
irisTree = C45()
irisTrainX, irisTestX, irisTrainY, irisTestY = train_test_split(irisData[[0,1,2,3]],
irisData[5], test_size=0.4)
irisTree.fit(irisTrainX, irisTrainY)
accuracy_score(irisTree.predict(irisTestX),irisTestY)
irisRoot = irisTree.getTree()
testData = pd.read_csv("test.data",delimiter=",")
testData["class"] = testData["class"].apply(lambda row: 1 if row == "Yes" else 0)
testTree = C45()
testTree.fit(testData[['a','b','c']],testData['class'])
# using the makePhishFile to update the uciPhishGraph.p file:
makePhishFile(irisRoot,irisData)
print("Benchmarking...")
with open('uciPhishGraph.p','rb') as f:
graphData = pickle.load(f)
dagList,root,parentList,terminals,outAttribute,trainData = graphData
print(dagList)
g = Graph(dagList, parentList, root, terminals, outAttribute, trainData)
print("Starting...")
currGraph,_,currTerminals = g.runPruning()
newGraph = dict()
addToGraph(currGraph, g.root, newGraph)
print(newGraph)
```
### Diabetes Data
```
diabetesDir = mainDir + "Diabetes/Diabetes-Data/"
for i in range(1,71):
dataDiabetesDir = diabetesDir
if i < 10:
dataDiabetesDir += "data-0{}".format(i)
else:
dataDiabetesDir += "data-{}".format(i)
dataDiabetesI = pd.read_csv(dataDiabetesDir, delimiter="\t",header=None)
print(dataDiabetesDir)
print(dataDiabetesI[0])
break
```
### Labor Data
```
laborDir = mainDir+"Labor/"
laborData = pd.read_csv(laborDir+"laborTrain.data")
laborData
```
| github_jupyter |
```
# Libraries
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from datetime import datetime
from matplotlib_venn import venn2
from collections import Counter
from pprint import pprint
%matplotlib inline
# Dataset
print(os.listdir('./dataset'))
```
## Registered individuals
----
```
# Registered users dataset
ind = pd.read_excel('./dataset/devfest-registered.xlsx', dtype=object)
print('Individuals')
print('Shape: ', ind.shape)
# Data head
ind.head(3)
# Description
print('Columns')
print(ind.columns)
# Column - Name
print('Name')
print('Unique value count: ', ind['Name'].nunique())
```
#### Column - Email
Email ID is the most important column in the data set. It's the primary identifier for unique individuals.
```
# Column - Email
print('Email')
print('Unique value count: ', ind['Email'].nunique())
# Column - City
print('City')
print('Unique value count: ', ind['City'].nunique())
# Column - Graduation year
print('Graduation Year')
print('Unique value count: ', ind['Graduation Year'].nunique())
pprint(Counter(ind['Graduation Year']))
# Graduation Year plot
# Countplot
gyear = ind['Graduation Year'].copy()
gyear = gyear.fillna('Missing').sort_values(ascending=True)
fig= plt.figure(figsize=(18,6))
sns.countplot(gyear)
# Column - College
print('College')
print('Unique value count: ', ind['College'].nunique())
# Column - Resume
print('Resume')
print('Uploaded resume count: ', ind.shape[0] - ind['Resume'].isna().sum())
print((ind.shape[0] - ind['Resume'].isna().sum())/ind.shape[0] * 100, '% of the individuals uploaded a resume' )
# Column - Gender
print('Gender')
print('Unique value count: ', ind['Gender'].nunique())
pprint(Counter(ind['Gender']))
# Gender type plot
# Countplot
gen = ind['Gender'].copy()
gen = gen.fillna('Missing')
sns.countplot(gen)
# Column - Phone Number
print('Phone Number')
print('Unique value count: ', ind['Phone Number'].nunique())
# Convert phone numbers to 10 digit standard format
# Parameters:
# data - phone number (any format)
# Return:
# Parsed 10 digit phone number
def parse_mobno(data):
data = str(data)
return data[-10:]
# Parsing phone number
# It is really important to parse the phone number in a 10 digit format to enable the script to send out emails
print('Sample: ', ind['Phone Number'][0])
print('\nParsing...\n')
ind['Phone Number'] = ind['Phone Number'].apply(parse_mobno)
print('Sample: ', ind['Phone Number'][0])
# Column - Stream
print('Stream')
print('Unique value count: ', ind['Stream'].nunique())
```
#### Column - Registered At
This is an important column that saves the timestamp of when the user registered on the Hackerearth portal
```
# Column - Registered At
print('Registered At')
print('Unique value count: ', ind['Registered At'].nunique())
# Convert str to datetime
# Parameters:
# data - String data format
# Return:
# datetime object
def convert_to_datetime(data):
return datetime.strptime(data[:-4], '%b %d, %Y %I:%M %p')
# Type conversion to datatime
print('Type of Registered At: ', type(ind['Registered At'][0]))
print('Sample: ', ind['Registered At'][0])
print('\nConverting...\n')
ind['Registered At'] = ind['Registered At'].apply(convert_to_datetime)
print('Type of Registered At: ', type(ind['Registered At'][0]))
print('Sample: ', ind['Registered At'][0])
# Splitting datetime into date and time
ind['date'] = [d.date() for d in ind['Registered At']]
ind['time'] = [d.time() for d in ind['Registered At']]
ind.head(2)
# Date analysis
print('Start date: ', ind['Registered At'].min())
print('End date: ', ind['Registered At'].max())
# Group by date
day_count = ind.groupby(['date'])['Email'].count()
print('Day count')
print(day_count[:5])
# Day wise registrations plot
fig= plt.figure(figsize=(18,6))
days = list(day_count.keys())
colors = [(x, x, 0.75) for x in [i for i in np.linspace(1,0,len(days))]]
plt.bar(days, day_count, color=colors)
plt.title('Daily registraion count')
plt.xlabel('Date')
plt.ylabel('Registration count')
plt.show()
# Day wise total registrations plot
fig= plt.figure(figsize=(18,6))
plt.plot(days, np.array(day_count).cumsum(), 'red')
plt.title('Cumulative registrations count')
plt.xlabel('Date')
plt.ylabel('Total Registrations')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/moh2236945/Natural-language-processing/blob/master/Multichannel_CNN_Model_for_Text_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
```
model can be expanded by using multiple parallel convolutional neural networks that read the source document using different kernel sizes. This, in effect, creates a multichannel convolutional neural network for text that reads text with different n-gram sizes (groups of words).
Movie Review Dataset
Data Preparation
In this section, we will look at 3 things:
Separation of data into training and test sets.
Loading and cleaning the data to remove punctuation and numbers.
Prepare all reviews and save to file.
```
from string import punctuation
from os import listdir
from nltk.corpus import stopwords
from pickle import dump
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# turn a doc into clean tokens
def clean_doc(doc):
# split into tokens by white space
tokens = doc.split()
# remove punctuation from each token
table = str.maketrans('', '', punctuation)
tokens = [w.translate(table) for w in tokens]
# remove remaining tokens that are not alphabetic
tokens = [word for word in tokens if word.isalpha()]
# filter out stop words
stop_words = set(stopwords.words('english'))
tokens = [w for w in tokens if not w in stop_words]
# filter out short tokens
tokens = [word for word in tokens if len(word) > 1]
tokens = ' '.join(tokens)
return tokens
# load all docs in a directory
def process_docs(directory, is_trian):
documents = list()
# walk through all files in the folder
for filename in listdir(directory):
# skip any reviews in the test set
if is_trian and filename.startswith('cv9'):
continue
if not is_trian and not filename.startswith('cv9'):
continue
# create the full path of the file to open
path = directory + '/' + filename
# load the doc
doc = load_doc(path)
# clean doc
tokens = clean_doc(doc)
# add to list
documents.append(tokens)
return documents
# save a dataset to file
def save_dataset(dataset, filename):
dump(dataset, open(filename, 'wb'))
print('Saved: %s' % filename)
# load all training reviews
negative_docs = process_docs('txt_sentoken/neg', True)
positive_docs = process_docs('txt_sentoken/pos', True)
trainX = negative_docs + positive_docs
trainy = [0 for _ in range(900)] + [1 for _ in range(900)]
save_dataset([trainX,trainy], 'train.pkl')
# load all test reviews
negative_docs = process_docs('txt_sentoken/neg', False)
positive_docs = process_docs('txt_sentoken/pos', False)
testX = negative_docs + positive_docs
testY = [0 for _ in range(100)] + [1 for _ in range(100)]
save_dataset([testX,testY], 'test.pkl')
from nltk.corpus import stopwords
import string
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# turn a doc into clean tokens
def clean_doc(doc):
# split into tokens by white space
tokens = doc.split()
# remove punctuation from each token
table = str.maketrans('', '', string.punctuation)
tokens = [w.translate(table) for w in tokens]
# remove remaining tokens that are not alphabetic
tokens = [word for word in tokens if word.isalpha()]
# filter out stop words
stop_words = set(stopwords.words('english'))
tokens = [w for w in tokens if not w in stop_words]
# filter out short tokens
tokens = [word for word in tokens if len(word) > 1]
return tokens
# load the document
filename = 'txt_sentoken/pos/cv000_29590.txt'
text = load_doc(filename)
tokens = clean_doc(text)
print(tokens)
# load all docs in a directory
def process_docs(directory, is_trian):
documents = list()
# walk through all files in the folder
for filename in listdir(directory):
# skip any reviews in the test set
if is_trian and filename.startswith('cv9'):
continue
if not is_trian and not filename.startswith('cv9'):
continue
# create the full path of the file to open
path = directory + '/' + filename
# load the doc
doc = load_doc(path)
# clean doc
tokens = clean_doc(doc)
# add to list
documents.append(tokens)
return documents
negative_docs = process_docs('txt_sentoken/neg', True)
trainy = [0 for _ in range(900)] + [1 for _ in range(900)]
testY = [0 for _ in range(100)] + [1 for _ in range(100)]
def save_dataset(dataset, filename):
dump(dataset, open(filename, 'wb'))
print('Saved: %s' % filename)
# load all test reviews
negative_docs = process_docs('txt_sentoken/neg', False)
positive_docs = process_docs('txt_sentoken/pos', False)
testX = negative_docs + positive_docs
testY = [0 for _ in range(100)] + [1 for _ in range(100)]
save_dataset([testX,testY], 'test.pkl')
```
develop a multichannel convolutional neural network for the sentiment analysis prediction problem.
This section is divided into 3 part
```
# load a clean dataset
def load_dataset(filename):
return load(open(filename, 'rb'))
trainLines, trainLabels = load_dataset('train.pkl')
# fit a tokenizer
def create_tokenizer(lines):
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# calculate the maximum document length
def max_length(lines):
return max([len(s.split()) for s in lines])
# calculate vocabulary size
vocab_size = len(tokenizer.word_index) + 1
# calculate vocabulary size
vocab_size = len(tokenizer.word_index) + 1
# encode a list of lines
def encode_text(tokenizer, lines, length):
# integer encode
encoded = tokenizer.texts_to_sequences(lines)
# pad encoded sequences
padded = pad_sequences(encoded, maxlen=length, padding='post')
return padded
# encode a list of lines
def encode_text(tokenizer, lines, length):
# integer encode
encoded = tokenizer.texts_to_sequences(lines)
# pad encoded sequences
padded = pad_sequences(encoded, maxlen=length, padding='post')
return padded
# define the model
def define_model(length, vocab_size):
# channel 1
inputs1 = Input(shape=(length,))
embedding1 = Embedding(vocab_size, 100)(inputs1)
conv1 = Conv1D(filters=32, kernel_size=4, activation='relu')(embedding1)
drop1 = Dropout(0.5)(conv1)
pool1 = MaxPooling1D(pool_size=2)(drop1)
flat1 = Flatten()(pool1)
# channel 2
inputs2 = Input(shape=(length,))
embedding2 = Embedding(vocab_size, 100)(inputs2)
conv2 = Conv1D(filters=32, kernel_size=6, activation='relu')(embedding2)
drop2 = Dropout(0.5)(conv2)
pool2 = MaxPooling1D(pool_size=2)(drop2)
flat2 = Flatten()(pool2)
# channel 3
inputs3 = Input(shape=(length,))
embedding3 = Embedding(vocab_size, 100)(inputs3)
conv3 = Conv1D(filters=32, kernel_size=8, activation='relu')(embedding3)
drop3 = Dropout(0.5)(conv3)
pool3 = MaxPooling1D(pool_size=2)(drop3)
flat3 = Flatten()(pool3)
# merge
merged = concatenate([flat1, flat2, flat3])
# interpretation
dense1 = Dense(10, activation='relu')(merged)
outputs = Dense(1, activation='sigmoid')(dense1)
model = Model(inputs=[inputs1, inputs2, inputs3], outputs=outputs)
# compile
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# summarize
print(model.summary())
plot_model(model, show_shapes=True, to_file='multichannel.png')
return model
# load training dataset
trainLines, trainLabels = load_dataset('train.pkl')
# create tokenizer
tokenizer = create_tokenizer(trainLines)
# calculate max document length
length = max_length(trainLines)
# calculate vocabulary size
vocab_size = len(tokenizer.word_index) + 1
print('Max document length: %d' % length)
print('Vocabulary size: %d' % vocab_size)
# encode data
trainX = encode_text(tokenizer, trainLines, length)
print(trainX.shape)
# define model
model = define_model(length, vocab_size)
# fit model
model.fit([trainX,trainX,trainX], array(trainLabels), epochs=10, batch_size=16)
# save the model
model.save('model.h5')
#Evaluation
# load datasets
trainLines, trainLabels = load_dataset('train.pkl')
testLines, testLabels = load_dataset('test.pkl')
# create tokenizer
tokenizer = create_tokenizer(trainLines)
# calculate max document length
length = max_length(trainLines)
# calculate vocabulary size
vocab_size = len(tokenizer.word_index) + 1
print('Max document length: %d' % length)
print('Vocabulary size: %d' % vocab_size)
# encode data
trainX = encode_text(tokenizer, trainLines, length)
testX = encode_text(tokenizer, testLines, length)
print(trainX.shape, testX.shape)
```
| github_jupyter |
<img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# NASA - Sea level
<a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/NASA/NASA_Sea_level.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
**Tags:** #nasa #naas #opendata #analytics #plotly
**Author:** [Colyn TIDMAN](https://www.linkedin.com/in/colyntidman/), [Dylan PICHON](https://www.linkedin.com/in/dylan-pichon/)
Sea level rise is caused primarily by two factors related to global warming: the added water from melting ice sheets and glaciers and the expansion of seawater as it warms. The first graph tracks the change in sea level since 1993 as observed by satellites.
The second graph, derived from coastal tide gauge and satellite data, shows how much sea level changed from about 1900 to 2018. Items with pluses (+) are factors that cause global mean sea level to increase, while minuses (-) are variables that cause sea levels to decrease. These items are displayed at the time they were affecting sea level.
The data shown are the latest available, with a four- to five-month lag needed for processing.
* You now need to create an Earthdata account to access NASA's sea level data. Register for free by clicking on 'Get data : http'. Once logged in you will access the data.
Website : https://climate.nasa.gov/vital-signs/sea-level/
Data source: Satellite sea level observations.
Credit: NASA's Goddard Space Flight Center
## Input
### Import libraries
```
import pandas
import plotly.graph_objects as go
```
### Path of the source
Data source : nasa_sea_levels.txt downloaded earlier
```
uri_nasa_sea_level = "nasa-sea-level-data.txt"
```
## Model
### Read the csv and create the table
```
df = pandas.read_csv(uri_nasa_sea_level, engine="python", comment='HDR',delim_whitespace=True, names=["A","B","Year + Fraction","D","E","F","G", "H","I","J","K","Smoothed GMSL (mm)",])
df.head(10)
```
Now lets only get the information we want and convert year + fraction to date
```
new_df = pandas.DataFrame(df, columns=['Year + Fraction', 'Smoothed GMSL (mm)'])
dates = []
values = []
ref = 0
for i, row in new_df.iterrows():
#date formating
date_split = str(row['Year + Fraction']).split('.')
year = date_split[0]
fraction = '0.' + date_split[1]
float_fraction = float(fraction)
date = year + "-1-1"
date_delta = 365 * float_fraction
value = pandas.to_datetime(date) + pandas.to_timedelta(date_delta, unit='D')
dates.append(value)
#value formating
#to stay inline with the graph visible on nasa's website, we need to have 0 as our first value
if i == 0:
ref = row['Smoothed GMSL (mm)']
val = row['Smoothed GMSL (mm)'] - ref
values.append(val)
new_df['Date'] = dates
new_df['Value'] = values
new_df.head()
```
## Output
### Land-Ocean Temperature Index - Visualization
```
fig = go.Figure(layout_title="<b>Sea Level variation since 1993 (mm)</b>")
fig.add_trace(go.Scatter(
x = new_df["Date"],
y = new_df["Value"],
name="Delta",
))
fig.update_layout(
autosize=False,
width=1300,
height=700,
plot_bgcolor='rgb(250,250,250)',
)
fig.add_annotation(y=6, x='2020-1-1',
text="Data source: Satellite sea level observations.<br> Credit: NASA's Goddard Space Flight Center",
showarrow=False,
)
fig.update_yaxes(title_text="Sea Height Variation (mm)")
fig.update_xaxes(title_text="Year", tickangle=60)
fig.add_hline(y=0.0)
fig.update_layout(title_x=0.5)
fig.show()
```
| github_jupyter |
# Introduction
In a prior notebook, documents were partitioned by assigning them to the domain with the highest Dice similarity of their term and structure occurrences. The occurrences of terms and structures in each domain is what we refer to as the domain "archetype." Here, we'll assess whether the observed similarity between documents and the archetype is greater than expected by chance. This would indicate that information in the framework generalizes well to individual documents.
# Load the data
```
import pandas as pd
import numpy as np
import sys
sys.path.append("..")
import utilities
from style import style
framework = "data-driven_k09"
version = 190325 # Document-term matrix version
suffix = "" # Suffix for term lists
clf = "_lr" # Classifier used to generate the framework
n_iter = 1000 # Iterations for null distribution
dx = [0.38, 0.38, 0.37, 0.39, 0.37, 0.38, 0.32, 0.34, 0.37] # Nudges for plotted means
ds = 0.11 # Nudges for plotted stars
alpha = 0.001 # Significance level for statistical comparisons
```
## Brain activation coordinates
```
act_bin = utilities.load_coordinates()
print("Document N={}, Structure N={}".format(
act_bin.shape[0], act_bin.shape[1]))
```
## Document-term matrix
```
dtm_bin = utilities.load_doc_term_matrix(version=version, binarize=True)
print("Document N={}, Term N={}".format(
dtm_bin.shape[0], dtm_bin.shape[1]))
```
## Domain archetypes
```
from collections import OrderedDict
lists, circuits = utilities.load_framework(framework, suffix=suffix, clf=clf)
words = sorted(list(set(lists["TOKEN"])))
structures = sorted(list(set(act_bin.columns)))
domains = list(OrderedDict.fromkeys(lists["DOMAIN"]))
archetypes = pd.DataFrame(0.0, index=words+structures, columns=domains)
for dom in domains:
for word in lists.loc[lists["DOMAIN"] == dom, "TOKEN"]:
archetypes.loc[word, dom] = 1.0
for struct in structures:
archetypes.loc[struct, dom] = circuits.loc[struct, dom]
archetypes[archetypes > 0.0] = 1.0
print("Term & Structure N={}, Domain N={}".format(
archetypes.shape[0], archetypes.shape[1]))
```
## Document structure-term vectors
```
pmids = dtm_bin.index.intersection(act_bin.index)
len(pmids)
dtm_words = dtm_bin.loc[pmids, words]
act_structs = act_bin.loc[pmids, structures]
docs = dtm_words.copy()
docs[structures] = act_structs.copy()
docs.head()
```
## Document splits
```
splits = {}
splits["discovery"] = [int(pmid.strip()) for pmid in open("../data/splits/train.txt")]
splits["replication"] = [int(pmid.strip()) for pmid in open("../data/splits/validation.txt")]
splits["replication"] += [int(pmid.strip()) for pmid in open("../data/splits/test.txt")]
for split, pmids in splits.items():
print("{:12s} N={}".format(split.title(), len(pmids)))
```
## Document assignments
```
doc2dom_df = pd.read_csv("../partition/data/doc2dom_{}{}.csv".format(framework, clf),
header=None, index_col=0)
doc2dom = {int(pmid): str(dom.values[0]) for pmid, dom in doc2dom_df.iterrows()}
dom2docs = {dom: {split: [] for split in ["discovery", "replication"]} for dom in domains}
for doc, dom in doc2dom.items():
for split, split_pmids in splits.items():
if doc in splits[split]:
dom2docs[dom][split].append(doc)
```
# Compute similarity to archetype
```
from scipy.spatial.distance import cdist
```
## Observed values
```
pmid_list, split_list, dom_list, obs_list = [], [], [], []
for split, split_pmids in splits.items():
split_list += [split] * len(split_pmids)
for dom in domains:
dom_pmids = dom2docs[dom][split]
dom_vecs = docs.loc[dom_pmids].values
dom_arche = archetypes[dom].values.reshape(1, archetypes.shape[0])
dom_sims = 1.0 - cdist(dom_vecs, dom_arche, metric="dice")
pmid_list += dom_pmids
dom_list += [dom] * len(dom_sims)
obs_list += list(dom_sims[:,0])
df_obs = pd.DataFrame({"PMID": pmid_list, "SPLIT": split_list,
"DOMAIN": dom_list, "OBSERVED": obs_list})
df_obs.to_csv("data/arche_obs_{}{}.csv".format(framework, clf))
df_obs.head()
```
## Null distributions
```
import os
df_null = {}
for split, split_pmids in splits.items():
print("Processing {} split (N={} documents)".format(split, len(split_pmids)))
file_null = "data/arche_null_{}{}_{}_{}iter.csv".format(framework, clf, split, n_iter)
if not os.path.isfile(file_null):
df_null[split] = np.zeros((len(domains), n_iter))
for n in range(n_iter):
null = np.random.choice(range(len(docs.columns)),
size=len(docs.columns), replace=False)
for i, dom in enumerate(domains):
dom_pmids = dom2docs[dom][split]
dom_vecs = docs.loc[dom_pmids].values
dom_arche = archetypes.values[null,i].reshape(1, archetypes.shape[0])
df_null[split][i,n] = 1.0 - np.mean(cdist(dom_vecs, dom_arche, metric="dice"))
if n % int(n_iter / 10.0) == 0:
print("----- Processed {} iterations".format(n))
df_null[split] = pd.DataFrame(df_null[split], index=domains, columns=range(n_iter))
df_null[split].to_csv(file_null)
print("")
else:
df_null[split] = pd.read_csv(file_null, index_col=0, header=0)
```
## Interleave splits to facilitate plotting
```
df_null_interleaved = pd.DataFrame()
null_idx = []
for dom in domains:
for split in ["discovery", "replication"]:
df_null_interleaved = df_null_interleaved.append(df_null[split].loc[dom])
null_idx.append(dom + "_" + split)
df_null_interleaved.index = null_idx
df_null_interleaved.head()
```
## Bootstrap distributions
```
df_boot = {}
for split, split_pmids in splits.items():
print("Processing {} split (N={} documents)".format(split, len(split_pmids)))
file_boot = "data/arche_boot_{}{}_{}_{}iter.csv".format(framework, clf, split, n_iter)
if not os.path.isfile(file_boot):
df_boot[split] = np.zeros((len(domains), n_iter))
for n in range(n_iter):
boot = np.random.choice(range(len(docs.columns)),
size=len(docs.columns), replace=True)
for i, dom in enumerate(domains):
dom_pmids = dom2docs[dom][split]
dom_vecs = docs.loc[dom_pmids].values[:,boot]
dom_arche = archetypes.values[boot,i].reshape(1, archetypes.shape[0])
df_boot[split][i,n] = 1.0 - np.mean(cdist(dom_vecs, dom_arche, metric="dice"))
if n % int(n_iter / 10.0) == 0:
print("----- Processed {} iterations".format(n))
df_boot[split] = pd.DataFrame(df_boot[split], index=domains, columns=range(n_iter))
df_boot[split].to_csv(file_boot)
print("")
else:
df_boot[split] = pd.read_csv(file_boot, index_col=0, header=0)
```
# Perform significance testing
```
from statsmodels.stats import multitest
df_stat = {}
for split, split_pmids in splits.items():
df_stat[split] = pd.DataFrame(index=domains)
pval = []
for dom in domains:
dom_obs = df_obs.loc[(df_obs["SPLIT"] == split) & (df_obs["DOMAIN"] == dom), "OBSERVED"].mean()
df_stat[split].loc[dom, "OBSERVED"] = dom_obs
dom_null = df_null[split].loc[dom].values
p = np.sum(dom_null >= dom_obs) / float(n_iter)
pval.append(p)
df_stat[split].loc[dom, "P"] = p
df_stat[split]["FDR"] = multitest.multipletests(pval, method="fdr_bh")[1]
for dom in domains:
if df_stat[split].loc[dom, "FDR"] < alpha:
df_stat[split].loc[dom, "STARS"] = "*"
else:
df_stat[split].loc[dom, "STARS"] = ""
df_stat[split] = df_stat[split].loc[domains, ["OBSERVED", "P", "FDR", "STARS"]]
df_stat[split].to_csv("data/arche_mean_{}{}_{}.csv".format(framework, clf, split))
print("-" * 65 + "\n" + split.upper() + "\n" + "-" * 65)
print(df_stat[split])
print("")
```
# Plot results
```
%matplotlib inline
palette = style.palettes["data-driven"] + style.palettes["dsm"]
utilities.plot_split_violins(framework, domains, df_obs, df_null_interleaved, df_stat,
palette, metric="arche", dx=dx, figsize=(3.5, 2.1),
ylim=[-0.25,1], yticks=[-0.25,0,0.25,0.5,0.75,1],
interval=0.999, alphas=[0], suffix=clf)
```
| github_jupyter |
# Deep Convolutional GANs
In this notebook, you'll build a GAN using convolutional layers in the generator and discriminator. This is called a Deep Convolutional GAN, or DCGAN for short. The DCGAN architecture was first explored in 2016 and has seen impressive results in generating new images; you can read the [original paper, here](https://arxiv.org/pdf/1511.06434.pdf).
You'll be training DCGAN on the [Street View House Numbers](http://ufldl.stanford.edu/housenumbers/) (SVHN) dataset. These are color images of house numbers collected from Google street view. SVHN images are in color and much more variable than MNIST.
<img src='assets/svhn_dcgan.png' width=80% />
So, our goal is to create a DCGAN that can generate new, realistic-looking images of house numbers. We'll go through the following steps to do this:
* Load in and pre-process the house numbers dataset
* Define discriminator and generator networks
* Train these adversarial networks
* Visualize the loss over time and some sample, generated images
#### Deeper Convolutional Networks
Since this dataset is more complex than our MNIST data, we'll need a deeper network to accurately identify patterns in these images and be able to generate new ones. Specifically, we'll use a series of convolutional or transpose convolutional layers in the discriminator and generator. It's also necessary to use batch normalization to get these convolutional networks to train.
Besides these changes in network structure, training the discriminator and generator networks should be the same as before. That is, the discriminator will alternate training on real and fake (generated) images, and the generator will aim to trick the discriminator into thinking that its generated images are real!
```
# import libraries
import matplotlib.pyplot as plt
import numpy as np
import pickle as pkl
%matplotlib inline
```
## Getting the data
Here you can download the SVHN dataset. It's a dataset built-in to the PyTorch datasets library. We can load in training data, transform it into Tensor datatypes, then create dataloaders to batch our data into a desired size.
```
import torch
from torchvision import datasets
from torchvision import transforms
# Tensor transform
transform = transforms.ToTensor()
# SVHN training datasets
svhn_train = datasets.SVHN(root='data/', split='train', download=True, transform=transform)
batch_size = 128
num_workers = 0
# build DataLoaders for SVHN dataset
train_loader = torch.utils.data.DataLoader(dataset=svhn_train,
batch_size=batch_size,
shuffle=True,
num_workers=num_workers)
```
### Visualize the Data
Here I'm showing a small sample of the images. Each of these is 32x32 with 3 color channels (RGB). These are the real, training images that we'll pass to the discriminator. Notice that each image has _one_ associated, numerical label.
```
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.transpose(images[idx], (1, 2, 0)))
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### Pre-processing: scaling from -1 to 1
We need to do a bit of pre-processing; we know that the output of our `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
```
# current range
img = images[0]
print('Min: ', img.min())
print('Max: ', img.max())
# helper scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
# assume x is scaled to (0, 1)
# scale to feature_range and return scaled x
min, max = feature_range
x = x * (max - min) + min
return x
# scaled range
scaled_img = scale(img)
print('Scaled min: ', scaled_img.min())
print('Scaled max: ', scaled_img.max())
```
---
# Define the Model
A GAN is comprised of two adversarial networks, a discriminator and a generator.
## Discriminator
Here you'll build the discriminator. This is a convolutional classifier like you've built before, only without any maxpooling layers.
* The inputs to the discriminator are 32x32x3 tensor images
* You'll want a few convolutional, hidden layers
* Then a fully connected layer for the output; as before, we want a sigmoid output, but we'll add that in the loss function, [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss), later
<img src='assets/conv_discriminator.png' width=80%/>
For the depths of the convolutional layers I suggest starting with 32 filters in the first layer, then double that depth as you add layers (to 64, 128, etc.). Note that in the DCGAN paper, they did all the downsampling using only strided convolutional layers with no maxpooling layers.
You'll also want to use batch normalization with [nn.BatchNorm2d](https://pytorch.org/docs/stable/nn.html#batchnorm2d) on each layer **except** the first convolutional layer and final, linear output layer.
#### Helper `conv` function
In general, each layer should look something like convolution > batch norm > leaky ReLU, and so we'll define a function to put these layers together. This function will create a sequential series of a convolutional + an optional batch norm layer. We'll create these using PyTorch's [Sequential container](https://pytorch.org/docs/stable/nn.html#sequential), which takes in a list of layers and creates layers according to the order that they are passed in to the Sequential constructor.
Note: It is also suggested that you use a **kernel_size of 4** and a **stride of 2** for strided convolutions.
```
import torch.nn as nn
import torch.nn.functional as F
# helper conv function
def conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a convolutional layer, with optional batch normalization.
"""
layers = []
conv_layer = nn.Conv2d(in_channels, out_channels,
kernel_size, stride, padding, bias=False)
# append conv layer
layers.append(conv_layer)
if batch_norm:
# append batchnorm layer
layers.append(nn.BatchNorm2d(out_channels))
# using Sequential container
return nn.Sequential(*layers)
class Discriminator(nn.Module):
def __init__(self, conv_dim=32):
super(Discriminator, self).__init__()
# complete init function
self.conv_dim = conv_dim
# 32x32 input
self.conv1 = conv(3, conv_dim, 4, batch_norm=False) # first layer, no batch_norm
# 16x16 out
self.conv2 = conv(conv_dim, conv_dim*2, 4)
# 8x8 out
self.conv3 = conv(conv_dim*2, conv_dim*4, 4)
# 4x4 out
# final, fully-connected layer
self.fc = nn.Linear(conv_dim*4*4*4, 1)
def forward(self, x):
# all hidden layers + leaky relu activation
out = F.leaky_relu(self.conv1(x), 0.2)
out = F.leaky_relu(self.conv2(out), 0.2)
out = F.leaky_relu(self.conv3(out), 0.2)
# flatten
out = out.view(-1, self.conv_dim*4*4*4)
# final output layer
out = self.fc(out)
return out
```
## Generator
Next, you'll build the generator network. The input will be our noise vector `z`, as before. And, the output will be a $tanh$ output, but this time with size 32x32 which is the size of our SVHN images.
<img src='assets/conv_generator.png' width=80% />
What's new here is we'll use transpose convolutional layers to create our new images.
* The first layer is a fully connected layer which is reshaped into a deep and narrow layer, something like 4x4x512.
* Then, we use batch normalization and a leaky ReLU activation.
* Next is a series of [transpose convolutional layers](https://pytorch.org/docs/stable/nn.html#convtranspose2d), where you typically halve the depth and double the width and height of the previous layer.
* And, we'll apply batch normalization and ReLU to all but the last of these hidden layers. Where we will just apply a `tanh` activation.
#### Helper `deconv` function
For each of these layers, the general scheme is transpose convolution > batch norm > ReLU, and so we'll define a function to put these layers together. This function will create a sequential series of a transpose convolutional + an optional batch norm layer. We'll create these using PyTorch's Sequential container, which takes in a list of layers and creates layers according to the order that they are passed in to the Sequential constructor.
Note: It is also suggested that you use a **kernel_size of 4** and a **stride of 2** for transpose convolutions.
```
# helper deconv function
def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a transposed-convolutional layer, with optional batch normalization.
"""
# create a sequence of transpose + optional batch norm layers
layers = []
transpose_conv_layer = nn.ConvTranspose2d(in_channels, out_channels,
kernel_size, stride, padding, bias=False)
# append transpose convolutional layer
layers.append(transpose_conv_layer)
if batch_norm:
# append batchnorm layer
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
class Generator(nn.Module):
def __init__(self, z_size, conv_dim=32):
super(Generator, self).__init__()
# complete init function
self.conv_dim = conv_dim
# first, fully-connected layer
self.fc = nn.Linear(z_size, conv_dim*4*4*4)
# transpose conv layers
self.t_conv1 = deconv(conv_dim*4, conv_dim*2, 4)
self.t_conv2 = deconv(conv_dim*2, conv_dim, 4)
self.t_conv3 = deconv(conv_dim, 3, 4, batch_norm=False)
def forward(self, x):
# fully-connected + reshape
out = self.fc(x)
out = out.view(-1, self.conv_dim*4, 4, 4) # (batch_size, depth, 4, 4)
# hidden transpose conv layers + relu
out = F.relu(self.t_conv1(out))
out = F.relu(self.t_conv2(out))
# last layer + tanh activation
out = self.t_conv3(out)
out = F.tanh(out)
return out
```
## Build complete network
Define your models' hyperparameters and instantiate the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.
```
# define hyperparams
conv_dim = 32
z_size = 100
# define discriminator and generator
D = Discriminator(conv_dim)
G = Generator(z_size=z_size, conv_dim=conv_dim)
print(D)
print()
print(G)
```
### Training on GPU
Check if you can train on GPU. If you can, set this as a variable and move your models to GPU.
> Later, we'll also move any inputs our models and loss functions see (real_images, z, and ground truth labels) to GPU as well.
```
train_on_gpu = torch.cuda.is_available()
if train_on_gpu:
# move models to GPU
G.cuda()
D.cuda()
print('GPU available for training. Models moved to GPU')
else:
print('Training on CPU.')
```
---
## Discriminator and Generator Losses
Now we need to calculate the losses. And this will be exactly the same as before.
### Discriminator Losses
> * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`.
* Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.
The losses will by binary cross entropy loss with logits, which we can get with [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss). This combines a `sigmoid` activation function **and** and binary cross entropy loss in one function.
For the real images, we want `D(real_images) = 1`. That is, we want the discriminator to classify the the real images with a label = 1, indicating that these are real. The discriminator loss for the fake data is similar. We want `D(fake_images) = 0`, where the fake images are the _generator output_, `fake_images = G(z)`.
### Generator Loss
The generator loss will look similar only with flipped labels. The generator's goal is to get `D(fake_images) = 1`. In this case, the labels are **flipped** to represent that the generator is trying to fool the discriminator into thinking that the images it generates (fakes) are real!
```
def real_loss(D_out, smooth=False):
batch_size = D_out.size(0)
# label smoothing
if smooth:
# smooth, real labels = 0.9
labels = torch.ones(batch_size)*0.9
else:
labels = torch.ones(batch_size) # real labels = 1
# move labels to GPU if available
if train_on_gpu:
labels = labels.cuda()
# binary cross entropy with logits loss
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
def fake_loss(D_out):
batch_size = D_out.size(0)
labels = torch.zeros(batch_size) # fake labels = 0
if train_on_gpu:
labels = labels.cuda()
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
```
## Optimizers
Not much new here, but notice how I am using a small learning rate and custom parameters for the Adam optimizers, This is based on some research into DCGAN model convergence.
### Hyperparameters
GANs are very sensitive to hyperparameters. A lot of experimentation goes into finding the best hyperparameters such that the generator and discriminator don't overpower each other. Try out your own hyperparameters or read [the DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf) to see what worked for them.
```
import torch.optim as optim
# params
lr = 0.0002
beta1=0.5
beta2=0.999 # default value
# Create optimizers for the discriminator and generator
d_optimizer = optim.Adam(D.parameters(), lr, [beta1, beta2])
g_optimizer = optim.Adam(G.parameters(), lr, [beta1, beta2])
```
---
## Training
Training will involve alternating between training the discriminator and the generator. We'll use our functions `real_loss` and `fake_loss` to help us calculate the discriminator losses in all of the following cases.
### Discriminator training
1. Compute the discriminator loss on real, training images
2. Generate fake images
3. Compute the discriminator loss on fake, generated images
4. Add up real and fake loss
5. Perform backpropagation + an optimization step to update the discriminator's weights
### Generator training
1. Generate fake images
2. Compute the discriminator loss on fake images, using **flipped** labels!
3. Perform backpropagation + an optimization step to update the generator's weights
#### Saving Samples
As we train, we'll also print out some loss statistics and save some generated "fake" samples.
**Evaluation mode**
Notice that, when we call our generator to create the samples to display, we set our model to evaluation mode: `G.eval()`. That's so the batch normalization layers will use the population statistics rather than the batch statistics (as they do during training), *and* so dropout layers will operate in eval() mode; not turning off any nodes for generating samples.
```
import pickle as pkl
# training hyperparams
num_epochs = 50
# keep track of loss and generated, "fake" samples
samples = []
losses = []
print_every = 300
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# train the network
for epoch in range(num_epochs):
for batch_i, (real_images, _) in enumerate(train_loader):
batch_size = real_images.size(0)
# important rescaling step
real_images = scale(real_images)
# ============================================
# TRAIN THE DISCRIMINATOR
# ============================================
d_optimizer.zero_grad()
# 1. Train with real images
# Compute the discriminator losses on real images
if train_on_gpu:
real_images = real_images.cuda()
D_real = D(real_images)
d_real_loss = real_loss(D_real)
# 2. Train with fake images
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
# move x to GPU, if available
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
D_fake = D(fake_images)
d_fake_loss = fake_loss(D_fake)
# add up loss and perform backprop
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# =========================================
# TRAIN THE GENERATOR
# =========================================
g_optimizer.zero_grad()
# 1. Train with fake images and flipped labels
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
# using flipped labels!
D_fake = D(fake_images)
g_loss = real_loss(D_fake) # use real loss to flip labels
# perform backprop
g_loss.backward()
g_optimizer.step()
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, num_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# generate and save sample, fake images
G.eval() # for generating samples
if train_on_gpu:
fixed_z = fixed_z.cuda()
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
```
## Training loss
Here we'll plot the training losses for the generator and discriminator, recorded after each epoch.
```
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
```
## Generator samples from training
Here we can view samples of images from the generator. We'll look at the images we saved during training.
```
# helper function for viewing a list of passed in sample images
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach().cpu().numpy()
img = np.transpose(img, (1, 2, 0))
img = ((img +1)*255 / (2)).astype(np.uint8) # rescale to pixel range (0-255)
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((32,32,3)))
_ = view_samples(-1, samples)
```
| github_jupyter |
# Model Analysis Project
In this project we analyse the Solow model. We want to examine how a higher labor supply will affect the level of productivity. Thus we extend the Solow model with $\lambda$ which indicades the fraction of the population in the labor force.
Furthermore, we adjust the model such that the level of technology increases within every period, and we analyse how this will affect the economy and the level of productivity.
Import and set magics:
```
import numpy as np
from scipy import optimize
import sympy as sm
from IPython.display import display
%matplotlib inline
import matplotlib.pyplot as plt
from matplotlib import cm
plt.style.use('seaborn-whitegrid')
plt.rcParams.update({'font.size': 12})
%matplotlib inline
import matplotlib.pyplot as plt # baseline modul
from mpl_toolkits.mplot3d import Axes3D # for 3d figures
plt.style.use('seaborn-whitegrid') # whitegrid nice with 3d
```
# The Solow model with labor supply
We consider the Solow model with an adjustable supply of labor. The model consists of the following equations:
$$Y_t = BK_t^{\alpha}L_t^{1-\alpha}, 0<\alpha<1$$
$$S_t = sY_t, 0 < s < 1$$
$$K_{t+1}-K_t = S_t\delta K_t, 0<\delta<1$$
where $K_t$ is capital today, $K_{t+1}$ is capital tomorrow, $S_t$ is savings and $Y_t$ is the output of the economy.
We also have $L_t$ which is the supply of labor, and $N_t$ which is the size of the population today, which are defined by the following:
$$L_t=\lambda N_t, 0<\lambda<1$$
$$N_{t+1}=(1+n)N_t$$
where lambda is the share of the population in work. The population is growing with a constant rate of $n$ in every period.
We consider capital, labor and output pr. capita (not pr. worker):
$$y_t = \frac{Y_t}{N_t} = \lambda^{(1-\alpha)}Bk_t^{\alpha}$$ with $$k_t = \frac{K_t}{N_t}$$
This will give us the following transformation equation of capital:
$$k_{t+1} = \frac{1}{(1+n)}(sB\lambda^{1-\alpha}k_t^{\alpha}+(1-\delta)k_t)$$
## Solving the model
The first step in solving the model is to find steady state of capital. This means solving for $k_t = k_{t+1} = k^{*}$
First we define all of the necessary symbols and equations of our model:
```
k = sm.symbols('k')
n = sm.symbols('n')
s = sm.symbols('s')
B = sm.symbols('B')
lam = sm.symbols('lambda')
delta = sm.symbols('delta')
alpha = sm.symbols('alpha')
steadystateK = sm.Eq(k,(s*B*lam**(1-alpha)*k**(alpha)+(1-delta)*k)/((1+n)))
```
We solve this using sympy:
```
steadystateKstar = sm.solve(steadystateK,k)[0]
steadystateKstar
```
We see that the level of capital in steady state is positively dependent of the savings $s$, $B$ and the share of the population that is working. The level of capital in steady state is negatively dependent of the growth in population as well as the rate of attrition of capital $\delta$, with the condition of stability of $\delta+n>0$.
We also want to find the steady state of the output of the economy $y^{*}$.
We know that $y_t = \lambda^{(1-\alpha)}Bk_t^{\alpha}$.
To solve for $y^{*}$ we need to insert $k^{*}$ into the equation.
```
#saving the steady state for k
ssk = sm.lambdify((B,lam,alpha,delta,n,s),steadystateKstar)
y = sm.symbols('y')
steadystateY = sm.Eq(y,lam**(1-alpha)*B*steadystateKstar**alpha)
steadystateYstar = sm.solve(steadystateY,y)[0]
steadystateYstar
```
We note that output is positively dependent of the same parameters as capital.
### How will the supply of labor affect the output of the economy? (Theoretically)
To show how the supply of labor affects the output in steady state we need to take a look at lambda - the share of population in work. First we look at the elasticity of lambda. To do this we need to rewrite the steady state for $y^{*}$:
$$y^{*} = B\lambda^{(1-\alpha)}(\frac{B\lambda^{(1-\alpha)}s}{(\delta + n)})^{\frac{\alpha}{(1-\alpha)}}$$
$$y^{*} = \frac{B\lambda^{(1-\alpha)}B^{\frac{\alpha}{(1-\alpha)}}\lambda^{\alpha}s^{\frac{\alpha}{(1-\alpha)}}}{(\delta + n)^\frac{\alpha}{{(1-\alpha)}}}$$
$$y^{*} = \frac{B^{\frac{1}{(1-\alpha)}}\lambda s^{\frac{\alpha}{(1-\alpha)}}}{(\delta + n)^\frac{\alpha}{{(1-\alpha)}}}$$
$$y^{*} = B^{\frac{1}{(1-\alpha)}}\lambda(\frac{s}{(\delta + n)})^{\frac{\alpha}{(1-\alpha)}}$$
```
ely = sm.Eq(y,B**(1/(1-alpha))*lam*(s/(delta+n))**(alpha/(1-alpha)))
ely
```
To find the elasticity of $y^{*}$ with respect to lambda we take the log on both sides:
$$ln(y^{*}) = \frac{1}{1-\alpha}ln(B) + ln(\lambda) + \frac{\alpha}{1-\alpha}(ln(s) - ln(n+\delta)) $$
We also find the elasticity of $y^{*}$ with respect to lambda with sympy:
```
y_eq = B**(1/(1-alpha))*lam*(s/(delta+n))**(alpha/(1-alpha))
print(f'The elasticity is {sm.diff(y_eq,lam)*lam/y_eq}')
```
We find that the elasticity of $y^{*}$ with respect to lambda is 1. This means that an increase in $\lambda$ with 1 pct. will result in an increase in $y^{*}$ with 1 pct.
### How will the supply of labor affect the output of the economy? (Example)
To give an example of what the outcome of the steady state could be, we have tried to insert plausible values of the parameters. To see the effect of an increase in lambda, we have calculated the steady state value of capital two times given a share of workers being 0.6 and 0.75 which leads to the following two results:
```
print(f'The first solution with lampda = 0.6 is: {ssk(1,0.6,1/3,0.1,0.25,0.2):.3f}')
print(f'The second solution with lampda = 0.75 is: {ssk(1,0.75,1/3,0.1,0.25,0.2):.3f}')
```
As expected we get a higher level of capital the higher the share of workers relative to the population. We will illustrate this relationship between $y^{*}$ and $\lambda$:
```
alpha=1/3
delta=0.1
g=0.02
n=0.25
s=0.2
B=1
lam_plot = np.linspace(0,1,100)
yss_plot = (B**(1/(1-alpha))*lam_plot*(s/(delta+n))**(alpha/(1-alpha)))
#Plotting different value of lambda from 0 to 1 against the value of tecnolegy adjusted capital steady state
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.set_xlabel("Share of population in the laborforce")
ax.set_ylabel("$y^\star$")
ax.set_title('Figure 1: The effect of $\lambda$ on the steady state output', fontsize=13)
ax.plot(lam_plot,yss_plot)
plt.show;
```
From this we clearly see the 1:1 relationship between an incease in $\lambda$ and the output pr. capita of the economy in steady state. Outside of steady state the relationship between the share of the population in the labor force ($\lambda$) and the output will be different.
The output (outside ss) in the economy in pr. capita is:
$y_t = \lambda^{(1-\alpha)}Bk_t^{\alpha}$
We calculate the elasticity of $y_t$ with respect to $\lambda$ with sympy:
```
y_t = B*lam**(1-alpha)*k**(alpha)
print(f'The elasticity is {sm.diff(y_t,lam)*lam/y_t}')
```
This means that an increase in $\lambda$ by 1 pct. will result in an increase in $y^{*}$ with 0.667 pct. The elasticity is now smaller than in steady state, since some of the production will be used as savings in the capital.
## Visualization of the two solutions
### The transition to steady state in a phase diagram
We want to show the transition to steady state in a phase diagram with the two different share of workers. We do this by simulating two different transations from time 0 to steady state.
```
#Transformation equation of capital
def k_trans(alpha,delta,lam,B,s,n,k_t):
return 1/(1+n)*(s*B*lam**(1-alpha)*k_t**alpha+(1-delta)*k_t)
#Steady state for k
def ss_k(alpha,delta,lam,B,s,n):
return (B*lam**(1-alpha)*s/(delta+n))**(1/(1-alpha))
# Setting the parameters
alpha=1/3
delta=0.1
lam_one=0.6
lam_two=0.75
B=1
n=0.25
s=0.2
k0 = 0.01
#Time periods
T = 20
index = range(T)
# Number of values in the grid
N = 1000
# Capital in period t
k_tone = np.linspace(0,0.4,N)
k_ttwo = np.linspace(0,0.4,N)
# Capital in period t+1
k_trans_one = np.zeros(N)
k_trans_two = np.zeros(N)
for i,k in enumerate(k_tone):
k_trans_one[i] = k_trans(alpha,delta,lam_one,B,s,n,k)
for i,k in enumerate(k_ttwo):
k_trans_two[i] = k_trans(alpha,delta,lam_two,B,s,n,k)
fig, axes = plt.subplots(1, 2, sharex='col', figsize=(13, 4))
ax1 = axes[0]
ax2 = axes[1]
#Phase diagram 1:
ax1.plot(k_tone,k_trans_one, color='0.3',label=r'$k_{t+1} = f(k_t)$')
ax1.plot(k_tone,k_tone,color='0.3',linestyle=':', label=r'$45^{\circ}$ degree line')
ax1.set_ylabel('$k_{t+1}$')
ax1.set_xlabel('$k_t$')
ax1.set_xlim(0,0.35)
ax1.set_ylim(0,0.35)
ax1.legend(loc=4,frameon=True)
ax1.set_title('Figure 2.a: Phase diagram with $\lambda$ = 0.6', fontsize=13)
#Phase diagram 2:
ax2.plot(k_ttwo,k_trans_two, color='0.3',label=r'$k_{t+1} = f(k_t)$')
ax2.plot(k_ttwo,k_ttwo,color='0.3',linestyle=':', label=r'$45^{\circ}$ degree line')
ax2.set_ylabel('$k_{t+1}$')
ax2.set_xlabel('$k_t$')
ax2.set_xlim(0,0.35)
ax2.set_ylim(0,0.35)
ax2.legend(loc=4,frameon=True)
ax2.set_title('Figure 2.b: Phase diagram with $\lambda$ = 0.75', fontsize=13)
plt.show()
```
In the two phase diagrams we see the transations from time 0 to steady state with two different values for $\lambda$. We clearly see that the steady state value of capital pr. capita is higher with a higher share of workers. An increase in $\lambda$ will not just benefit the economy temporarily but also in the long run.
### A simulation of convergence of capital to steady state
```
#Graph 3
k_level_one = np.zeros(len(index))
k_level_one[0] = k0
for t in index[1:]:
k_level_one[t] = k_trans(alpha,delta,lam_one,B,s,n,k_level_one[t-1])
k_level_two = np.zeros(len(index))
k_level_two[0] = k0
for t in index[1:]:
k_level_two[t] = k_trans(alpha,delta,lam_two,B,s,n,k_level_two[t-1])
fig, ax = plt.subplots(sharex='col', figsize=(6.5, 4))
ax.plot(k_level_one, color='0.3',label=r'lambda = 0.6')
ax.plot(k_level_two,color='0.3',linestyle=':', label=r'lambda = 0.75')
ax.set_ylabel('$k_{t}$')
ax.set_xlabel('t')
ax.legend(loc=4,frameon=True)
ax.set_title('Figure 3: Simulated convergence of capital to steady state', fontsize=13)
plt.show()
```
Figure 3 illustrates how capital converges to steady state with $\lambda$ being 0.6 and 0.75. Not only is the steady state level of $\lambda = 0.75$ higher, the level of capital in this economy is also higher during the time it converges to steady state. We see that in $t = 15$ the level of capital with $\lambda = 0.75$ is approximately 0.06 larger than the level of capital with $\lambda = 0.6$.
# The Solow model with labor supply and exogeneous technological progress
Instead of a fixed level of technology we now consider an economy with growing technology. The level of technology increases in every period beacuse of more R&D. The model changes such that
$$Y_t = K_t^{\alpha}(A_tL_t)^{1-\alpha}, 0<\alpha<1$$
$$A_{t+1}=(1+g)A_t$$
We note that the level of technology increases by $g$ in every period. Other than this the model is the same as earlier.
Now we consider technologically adjusted capital, labor and output pr. capita:
$$\tilde{y}_t = \frac{Y_t}{A_tN_t} = \lambda^{(1-\alpha)}\tilde{k}_t^{\alpha}$$ with $$\tilde{k}_t = \frac{K_t}{A_tN_t}$$
This gives us the following transformation equation of capital:
$$\tilde{k}_{t+1} = \frac{1}{(1+n)(1+g)}(s\lambda^{1-\alpha}\tilde{k}_t^{\alpha}+(1-\delta)\tilde{k}_t)$$
## Solving the model analytically
The analytic solution of steady state of capital is solved:
```
k = sm.symbols('k')
n = sm.symbols('n')
s = sm.symbols('s')
lam = sm.symbols('lambda')
delta = sm.symbols('delta')
alpha = sm.symbols('alpha')
g = sm.symbols('g')
steadystateK_tilde = sm.Eq(k,(s*lam**(1-alpha)*k**(alpha)+(1-delta)*k)/((1+n)*(1+g)))
kss = sm.solve(steadystateK_tilde,k)[0]
kss
```
## Solving the model numerically
The numerical solution of steady state of capital is solved:
```
# Setting the parameters again
alpha=1/3
delta=0.1
lam_one=0.6
lam_two=0.75
g=0.02
n=0.25
s=0.2
def solver(lam,s,g,n,alpha,delta):
func = lambda k: k**alpha
k_tilde_func = lambda k_tilde: k_tilde - (s*lam**(1-alpha)*func(k_tilde)+(1-delta)*k_tilde)/((1+n)*(1+g))
solve = optimize.root_scalar(k_tilde_func,bracket=[0.1,100],method='bisect')
return solve
solution1 = solver(lam_one,s,g,n,alpha,delta)
solution2 = solver(lam_two,s,g,n,alpha,delta)
print(f'The numerical solution with lambda = 0.6 is : {solution1.root:.3f}')
print(f'The numerical solution with lambda = 0.75 is : {solution2.root:.3f}')
```
The steady state level of capital is now smaller in both cases since we look at technologically adjusted capital pr. capita and not just capital pr. capita as we did earlier. We notice that the increase in capital given an increase in $\lambda$ is smaller in this economy.
## Visualization of the second model
To get a better idea of the impact on the steady state value from the parameters, we plot the share of the population in the labor force against the technologically adjusted capital pr. capita:
```
lam_plot = np.linspace(0,1,100)
kss_plot = (lam_plot**(1-alpha)*s/(delta+g*n+g+n))**1/(1-alpha)
#Plotting different value of lambda from 0 to 1 against the value of tecnolegy adjusted capital steady state
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.set_xlabel("Share of population in the laborforce")
ax.set_ylabel("$k_{tilde}^\star$")
ax.set_title('Figure 4: The effect of $\lambda$ on the steady state of capital', fontsize=13)
ax.plot(lam_plot,kss_plot)
plt.show;
```
We find that a higher value of $\lambda$ also increases technologically adjusted capital pr. capita.
We combine this with different values for the populations growth:
```
def kss_plot(lam,n,alpha=1/3,delta=0.1,g=0.02,s=0.2):
ksteady = (lam**(1-alpha)*s/(delta+g*n+g+n))**1/(1-alpha)
return ksteady
n_plot = np.linspace(0,1,100)
lam_plot = np.linspace(0,1,100)
x1_values_alt,x2_values_alt = np.meshgrid(lam_plot,n_plot,indexing='ij')
u_values_alt = kss_plot(x1_values_alt,x2_values_alt,alpha=1/3,delta=0.1,g=0.02,s=0.2)
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_surface(x2_values_alt,x1_values_alt,u_values_alt,cmap=cm.jet)
ax.set_xlabel("n")
ax.set_ylabel("$lambda$")
ax.set_zlabel("$k_{tilde}^\star$")
ax.set_title('Figure 5: The effect of $n$ and $\lambda$ on the steady state of capital', fontsize=13);
```
As we expected the highest level of technologically adjusted capital pr. capita is found with a high level og $\lambda$ and no population growth. Since we look at the capital pr. capita an increase in the population will result in more pepole to "share" the capital with.
# Conclusion
The conclusion of this project is that a higher share of workers will result in a higher level of capital in steady state. In steady state we find a 1:1 relationship between an incease in $\lambda$ and the output pr. capita of the economy. This applies to both an economy with a fixed level of technology and an economy with an increasing development of technology over time. The difference is however, that an economy with a fixed level of technology will see a larger positive effect on capital pr. capita than an economy with an increasing development of technology will see on technologically adjusted capital pr. capita. This is primarily because of the fact that the level of technology is adjusted.
Thus if an economy is in a crisis where they have problems with a decreasing level of productivity which leads to no economic growth, a possible solution to this could be to raise the supply of labor. The supply of labor could be increased through lower taxation on income or if the unemployment benefits were lowered.
| github_jupyter |
# Tutorial: Computing with shapes of landmarks in Kendall shape spaces
Lead author: Nina Miolane.
In this tutorial, we show how to use geomstats to perform a shape data analysis. Specifically, we aim to study the difference between two groups of data:
- optical nerve heads that correspond to normal eyes,
- optical nerve heads that correspond to glaucoma eyes.
We wish to investigate if there is a difference in these two groups, and if this difference is a difference in sizes of the optical nerve heads, or a difference in shapes (where the size has been quotiented out).
<img src="figures/optic_nerves.png" />
## Set up
```
import os
import sys
import warnings
sys.path.append(os.path.dirname(os.getcwd()))
warnings.filterwarnings("ignore")
%matplotlib inline
import matplotlib.colors as colors
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits.mplot3d.art3d import Poly3DCollection
import geomstats.backend as gs
import geomstats.datasets.utils as data_utils
from geomstats.geometry.pre_shape import PreShapeSpace, KendallShapeMetric
```
We import the dataset of the optical nerve heads from 22 images of Rhesus monkeys’ eyes (11 monkeys), available in [[PE2015]](#References).
For each monkey, an experimental glaucoma was introduced in one eye, while the second
eye was kept as control. One seeks to observe differences between the glaucoma and the
control eyes. On each image, 5 anatomical landmarks were recorded:
- 1st landmark: superior aspect of the retina,
- 2nd landmark: side of the retina closest to the temporal bone of the skull,
- 3rd landmark: nose side of the retina,
- 4th landmark: inferior point,
- 5th landmark: optical nerve head deepest point.
Label 0 refers to a normal eye, and Label 1 to an eye with glaucoma.
```
nerves, labels, monkeys = data_utils.load_optical_nerves()
print(nerves.shape)
print(labels)
print(monkeys)
```
We extract the landmarks' sets corresponding to the two eyes' nerves of the first monkey, with their corresponding labels.
```
two_nerves = nerves[monkeys == 0]
print(two_nerves.shape)
two_labels = labels[monkeys == 0]
print(two_labels)
label_to_str = {0: "Normal nerve", 1: "Glaucoma nerve"}
label_to_color = {
0: (102 / 255, 178 / 255, 255 / 255, 1.0),
1: (255 / 255, 178 / 255, 102 / 255, 1.0),
}
fig = plt.figure()
ax = Axes3D(fig)
ax.set_xlim((2000, 4000))
ax.set_ylim((1000, 5000))
ax.set_zlim((-600, 200))
for nerve, label in zip(two_nerves, two_labels):
x = nerve[:, 0]
y = nerve[:, 1]
z = nerve[:, 2]
verts = [list(zip(x, y, z))]
poly = Poly3DCollection(verts, alpha=0.5)
color = label_to_color[int(label)]
poly.set_color(colors.rgb2hex(color))
poly.set_edgecolor("k")
ax.add_collection3d(poly)
patch_0 = mpatches.Patch(color=label_to_color[0], label=label_to_str[0], alpha=0.5)
patch_1 = mpatches.Patch(color=label_to_color[1], label=label_to_str[1], alpha=0.5)
plt.legend(handles=[patch_0, patch_1], prop={"size": 14})
plt.show()
```
We first try to detect if there are two groups of optical nerve heads, based on the 3D coordinates of the landmarks sets.
```
from geomstats.geometry.euclidean import EuclideanMetric
nerves_vec = nerves.reshape(22, -1)
eucl_metric = EuclideanMetric(nerves_vec.shape[-1])
eucl_dist = eucl_metric.dist_pairwise(nerves_vec)
plt.figure()
plt.imshow(eucl_dist);
```
We do not see any two clear clusters.
We want to investigate if there is a difference between these two groups of shapes - normal nerve versus glaucoma nerve - or if the main difference is merely relative to the global size of the landmarks' sets.
```
m_ambient = 3
k_landmarks = 5
preshape = PreShapeSpace(m_ambient=m_ambient, k_landmarks=k_landmarks)
matrices_metric = preshape.embedding_metric
sizes = matrices_metric.norm(preshape.center(nerves))
plt.figure(figsize=(6, 4))
for label, col in label_to_color.items():
label_sizes = sizes[labels == label]
plt.hist(label_sizes, color=col, label=label_to_str[label], alpha=0.5, bins=10)
plt.axvline(gs.mean(label_sizes), color=col)
plt.legend(fontsize=14)
plt.title("Sizes of optical nerves", fontsize=14);
```
The vertical lines represent the sample mean of each group (normal/glaucoma).
```
plt.figure(figsize=(6, 4))
plt.hist(sizes[labels == 1] - sizes[labels == 0], alpha=0.5)
plt.axvline(0, color="black")
plt.title(
"Difference in size of optical nerve between glaucoma and normal eyes", fontsize=14
);
```
We perform a hypothesis test, testing if the two samples of sizes have the same average. We use the t-test for related samples, since the sample elements are paired: two eyes for each monkey.
```
from scipy import stats
signif_level = 0.05
tstat, pvalue = stats.ttest_rel(sizes[labels == 0], sizes[labels == 1])
print(pvalue < signif_level)
```
There is a significative difference, in optical nerve eyes' sizes, between the glaucoma and normal eye.
We want to investigate if there is a difference in shapes, where the size component has been quotiented out.
We project the data to the Kendall pre-shape space, which:
- centers the nerve landmark sets so that they share the same barycenter,
- normalizes the sizes of the landmarks' sets to 1.
```
nerves_preshape = preshape.projection(nerves)
print(nerves_preshape.shape)
print(preshape.belongs(nerves_preshape))
print(gs.isclose(matrices_metric.norm(nerves_preshape), 1.0))
```
In order to quotient out the 3D orientation component, we align the landmark sets in the preshape space.
```
base_point = nerves_preshape[0]
nerves_shape = preshape.align(point=nerves_preshape, base_point=base_point)
```
The Kendall metric is a Riemannian metric that takes this alignment into account. It corresponds to the metric of the Kendall shape space, which is the manifold defined as the preshape space quotient by the action of the rotation in m_ambient dimensions, here in 3 dimensions.
```
kendall_metric = KendallShapeMetric(m_ambient=m_ambient, k_landmarks=k_landmarks)
```
We can use it to perform a tangent PCA in the Kendall shape space, and determine if we see a difference in the shapes of the optical nerves.
```
from geomstats.learning.pca import TangentPCA
tpca = TangentPCA(kendall_metric)
tpca.fit(nerves_shape)
plt.plot(tpca.explained_variance_ratio_)
plt.xlabel("Number of principal tangent components", size=14)
plt.ylabel("Fraction of explained variance", size=14);
```
Two principal components already describe around 60% of the variance. We plot the data projected in the tangent space defined by these two principal components.
```
X = tpca.transform(nerves_shape)
plt.figure(figsize=(12, 12))
for label, col in label_to_color.items():
mask = labels == label
plt.scatter(X[mask, 0], X[mask, 1], color=col, s=100, label=label_to_str[label])
plt.legend(fontsize=14)
for label, x, y in zip(monkeys, X[:, 0], X[:, 1]):
plt.annotate(
label,
xy=(x, y),
xytext=(-20, 20),
textcoords="offset points",
ha="right",
va="bottom",
bbox=dict(boxstyle="round,pad=0.5", fc="white", alpha=0.5),
arrowprops=dict(arrowstyle="->", connectionstyle="arc3,rad=0"),
)
plt.show()
```
The indices represent the monkeys' indices.
In contrast to the above study focusing on the optical nerves' sizes, visual inspection does not reveal any clusters between the glaucoma and normal optical nerves' shapes. We also do not see any obvious pattern between the two optical nerves of the same monkey.
This shows that the difference between the optical nerve heads mainly resides in the over sizes of the optical nerves.
```
dist_pairwise = kendall_metric.dist_pairwise(nerves_shape)
print(dist_pairwise.shape)
plt.figure()
plt.imshow(dist_pairwise);
```
We try a agglomerative hierarchical clustering to investigate if we can cluster in the Kendall shape space.
```
from geomstats.learning.agglomerative_hierarchical_clustering import (
AgglomerativeHierarchicalClustering,
)
clustering = AgglomerativeHierarchicalClustering(distance="precomputed", n_clusters=2)
clustering.fit(dist_pairwise)
predicted_labels = clustering.labels_
print("True labels:", labels)
print("Predicted labels:", predicted_labels)
accuracy = gs.sum(labels == predicted_labels) / len(labels)
print(f"Accuracy: {accuracy:.2f}")
```
The accuracy is barely above the accuracy of a random classifier, that would assign 0 or 1 with probably 0.5 to each of the shapes. This confirms that the difference that exists between the two groups is mostly due to the landmarks' set size and not their shapes.
## References
.. [PE2015] Patrangenaru and L. Ellingson. Nonparametric Statistics on Manifolds and Their Applications to Object Data, 2015. https://doi.org/10.1201/b18969
| github_jupyter |
# Reinforcement Learning
## Importing packages
We import several familiar libraries like numpy and random.
We also provide you an implementation of an MDP in MDP_FrozenLake.py and with a way to visualize your learning and your environments in vis_ex1ab.py.
```
import numpy as np
import random
import matplotlib.pyplot as plt
from FrozenLake import FrozenLakeEnv
plt.rcParams.update({'figure.max_open_warning': 0})
```
# Dynamic Programming
## The MDP
The problem is provided by the Berkeley Deep RL Course Homeworks: https://github.com/berkeleydeeprlcourse/homework
The task is to reach the goal state <b>G</b>, starting from initial state <b>S</b>, without falling into Hole <b>H</b> and die miserably. That, means we have to reach the goal traversing the <b>F</b> tiles. Remember that in the Frozenworld Problem the ground is slippery, meaning if you move right, you can also move upward or downward as well (the transition model is non-deterministic).
The map of the MDP is the following:
S F F F
F H F H
F F F H
H F F G
The possible actions are the following:
LEFT = 0
DOWN = 1
RIGHT = 2
UP = 3
## Value Iteration
The basis for every decision making process is to measure, if the state $s$ following an action $a$ is good or not. If we know the transition model and reward function of the MDP, we can achieve this by estimating the <b>optimal</b> State-Value Function $V$, assigning a value to each state $s$.
We can approximate the optimal value function using Dynamic Programming. Dynamic programming is tabular approach, which saves a value for each state. This table is created in __\_\_init\_\___ using numpy. Note that it is a good practice to initialize with $V(s) = 0$ for all states.
We calculate the values using the bellman optimality equation. The adaption for the <b>Value Iteration</b> approach from lecture one is as follows:
$V \left( s \right) = \underset{a \in \mathcal{A}}{max} \underset{ s' \in \mathcal{S}, r}{\sum} \mathcal{P}\left( s' , r | s, a \right) \left[ r + \gamma V \left( s' \right) \right] $
As you probably can imagine, $V$ should be updated accordingly in \_\_value_iteration\_\_ using the formula above. However, one update won't converge to the actual values. This means, we have to iterate all states, multiple times, until the values are sufficiently accurate. Basically, you have to implement the evaluation part of the algorithm from lecture one in this function.
<b>ATTENTION</b>: An importan aspect to consider is that special care needs to be taken when we are at a terminal state (e.g., in the Goal <b>G</b> or Hole <b>H</b> states of the Frozen Lake MDP). Here, no matter what action we execute, we cannot "escape" this state, so the Value Iteration implementation should be adapted accordingly.
## The Value Iteration Agent
```
class DP_Agent:
def __init__(self, discount_factor):
self.g = discount_factor
self.MDP = FrozenLakeEnv()
self.MDP.seed(0)
self.terminals = [5,7,11,12,15]
# MDP.p_a: uniform action probability (25% chance an action is selected)
# MDP.p_s_: uniform next-state probability
# MDP.P_ss_(s,a): set of all possible states when taking action a in state s
# MDP.R(s): reward in state s
# MDP.A: set of all possible actions,e.g. full action space
# MDP.S_(s,a): next state when taking action a in state s
# 4x4 gridworld:
#"S F F F",
#"F H F H",
#"F F F H",
#"H F F G"
#S: Start (constant starting position, reward=0)
#F: Ice (introduces stochastic action, reward=0)
#H: Hole (ends episode, reward=0)
#G: Goal (ends episode, reward=1)
# states = {0,1,...,15}
self.V = np.zeros(shape=(16), dtype=float)
# Dynamic Programming
def one_step_lookahead(self, state, V):
"""
Helper function to calculate the value for all action in a given state.
Args:
state: The state to consider (int)
V: The value to use as an estimator, Vector of length env.nS
Returns:
A vector of length env.nA containing the expected value of each action.
"""
A = np.zeros(self.MDP.nA)
for a in range(self.MDP.nA):
for prob, next_state, reward, done in self.MDP.P[state][a]:
A[a] += prob * (reward + self.g * self.V[next_state])
return A
def update_DP(self):
policy = np.zeros([self.MDP.nS])
# Update each state...
for s in range(self.MDP.nS):
# Do a one-step lookahead to find the best action
A = self.one_step_lookahead(s, self.V)
best_action_value = np.max(A)
best_action = np.argmax(A)
# Update the value function. Ref: Sutton book eq. 4.10.
self.V[s] = best_action_value
policy[s] = best_action
return policy, self.V
```
## Base Loop
```
agentDP = DP_Agent(discount_factor=0.95)
Vs = [np.zeros(agentDP.MDP.nS)]
pis = []
for iterations in range(20):
policy, V = agentDP.update_DP()
Vs.append(V.copy())
pis.append(policy)
```
## Visualization
```
for (V, pi) in zip(Vs, pis):
plt.figure(figsize=(3,3))
plt.imshow(V.reshape(4,4), cmap='gray', interpolation='none', clim=(0,1))
ax = plt.gca()
ax.set_xticks(np.arange(4)-.5)
ax.set_yticks(np.arange(4)-.5)
ax.set_xticklabels([])
ax.set_yticklabels([])
Y, X = np.mgrid[0:4, 0:4]
a2uv = {0: (-1, 0), 1:(0, -1), 2:(1,0), 3:(-1, 0)}
Pi = pi.reshape(4,4)
for y in range(4):
for x in range(4):
a = Pi[y, x]
u, v = a2uv[a]
plt.arrow(x, y,u*.3, -v*.3, color='m', head_width=0.1, head_length=0.1)
plt.text(x, y, str(agentDP.MDP.unwrapped.desc[y,x].item().decode()),
color='g', size=12, verticalalignment='center',
horizontalalignment='center', fontweight='bold')
plt.grid(color='b', lw=2, ls='-')
plt.figure()
plt.plot(Vs)
plt.title("Values of different states");
```
| github_jupyter |
# Udacity. Deep Reingorcement Learning : Collaboration and Competition
### Markus Buchholz
```
from unityagents import UnityEnvironment
import numpy as np
env = UnityEnvironment(file_name='./Tennis_Linux/Tennis.x86_64')
```
## BRAIN
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
## MODEL
```
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
def hidden_init(layer):
fan_in = layer.weight.data.size()[0]
lim = 1. / np.sqrt(fan_in)
return (-lim, lim)
class Actor(nn.Module):
"""Actor (Policy) Model."""
def __init__(self, state_size, action_size, seed, fc1_units=200, fc2_units=150):
"""Initialize parameters and build model.
Params
======
state_size (int): Dimension of each state
action_size (int): Dimension of each action
seed (int): Random seed
fc1_units (int): Number of nodes in first hidden layer
fc2_units (int): Number of nodes in second hidden layer
"""
super(Actor, self).__init__()
self.seed = torch.manual_seed(seed)
self.fc1 = nn.Linear(state_size, fc1_units)
self.fc2 = nn.Linear(fc1_units, fc2_units)
self.fc3 = nn.Linear(fc2_units, action_size)
self.reset_parameters()
def reset_parameters(self):
self.fc1.weight.data.uniform_(*hidden_init(self.fc1))
self.fc2.weight.data.uniform_(*hidden_init(self.fc2))
self.fc3.weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state):
"""Build an actor (policy) network that maps states -> actions."""
x = F.relu(self.fc1(state))
x = F.relu(self.fc2(x))
return F.tanh(self.fc3(x))
class Critic(nn.Module):
"""Critic (Value) Model."""
def __init__(self, state_size, action_size, seed, fcs1_units=400, fc2_units=300):
"""Initialize parameters and build model.
Params
======
state_size (int): Dimension of each state
action_size (int): Dimension of each action
seed (int): Random seed
fcs1_units (int): Number of nodes in the first hidden layer
fc2_units (int): Number of nodes in the second hidden layer
"""
super(Critic, self).__init__()
self.seed = torch.manual_seed(seed)
self.fcs1 = nn.Linear(state_size, fcs1_units)
self.fc2 = nn.Linear(fcs1_units+action_size, fc2_units)
self.fc3 = nn.Linear(fc2_units, 1)
self.reset_parameters()
def reset_parameters(self):
self.fcs1.weight.data.uniform_(*hidden_init(self.fcs1))
self.fc2.weight.data.uniform_(*hidden_init(self.fc2))
self.fc3.weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state, action):
"""Build a critic (value) network that maps (state, action) pairs -> Q-values."""
xs = F.relu(self.fcs1(state))
x = torch.cat((xs, action), dim=1)
x = F.relu(self.fc2(x))
return self.fc3(x)
```
## AGENT AND CRITIC
```
import numpy as np
import random
import copy
from collections import namedtuple, deque
import torch
import torch.nn.functional as F
import torch.optim as optim
BUFFER_SIZE = int(1e6) # replay buffer size
BATCH_SIZE = 512 # minibatch size
GAMMA = 0.99 # discount factor
TAU = 1e-3 # for soft update of target parameters
LR_ACTOR = 1e-3 # learning rate of the actor
LR_CRITIC = 1e-3 # learning rate of the critic
WEIGHT_DECAY = 0 # L2 weight decay
eps_start= 1.0
eps_end = 0.01
eps_decay = 1e-6
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
class Agent():
def __init__(self, state_size, action_size, num_agents, random_seed):
self.state_size = state_size
self.action_size = action_size
self.seed = random_seed
# Actor Network (w/ Target Network)
self.actor_local = Actor(self.state_size, self.action_size, self.seed).to(device)
self.actor_target = Actor(self.state_size, self.action_size, self.seed).to(device)
self.actor_optimizer = optim.Adam(self.actor_local.parameters(), lr=LR_ACTOR)
# Critic Network (w/ Target Network)
self.critic_local = Critic(self.state_size, self.action_size, self.seed).to(device)
self.critic_target = Critic(self.state_size, self.action_size, self.seed).to(device)
self.critic_optimizer = optim.Adam(self.critic_local.parameters(), lr=LR_CRITIC, weight_decay=WEIGHT_DECAY)
# Noise process
self.noise = OUNoise((num_agents,action_size), random_seed)
# Replay memory
self.memory = ReplayBuffer(action_size, BUFFER_SIZE, BATCH_SIZE, random_seed)
def step(self, state, action, reward, next_state, done):
"""Save experience in replay memory, and use random sample from buffer to learn."""
# Save experience / reward
self.memory.add(state, action, reward, next_state, done)
# Learn, if enough samples are available in memory
if len(self.memory) > BATCH_SIZE:
experiences = self.memory.sample()
self.learn(experiences, GAMMA)
def act(self, state, add_noise=True):
"""Returns actions for given state as per current policy."""
state = torch.from_numpy(state).float().to(device)
self.actor_local.eval()
with torch.no_grad():
action = self.actor_local(state).cpu().data.numpy()
self.actor_local.train()
if add_noise:
action += self.noise.sample()
return np.clip(action, -1, 1)
def reset(self):
self.noise.reset()
def learn(self, experiences, gamma):
"""Update policy and value parameters using given batch of experience tuples.
Q_targets = r + γ * critic_target(next_state, actor_target(next_state))
where:
actor_target(state) -> action
critic_target(state, action) -> Q-value
Params
======
experiences (Tuple[torch.Tensor]): tuple of (s, a, r, s', done) tuples
gamma (float): discount factor
"""
states, actions, rewards, next_states, dones = experiences
# ---------------------------- update critic ---------------------------- #
# Get predicted next-state actions and Q values from target models
actions_next = self.actor_target(next_states)
Q_targets_next = self.critic_target(next_states, actions_next)
# Compute Q targets for current states (y_i)
Q_targets = rewards + (gamma * Q_targets_next * (1 - dones))
# Compute critic loss
Q_expected = self.critic_local(states, actions)
critic_loss = F.mse_loss(Q_expected, Q_targets)
# Minimize the loss
self.critic_optimizer.zero_grad()
critic_loss.backward()
self.critic_optimizer.step()
# ---------------------------- update actor ---------------------------- #
# Compute actor loss
actions_pred = self.actor_local(states)
actor_loss = -self.critic_local(states, actions_pred).mean()
# Minimize the loss
self.actor_optimizer.zero_grad()
actor_loss.backward()
self.actor_optimizer.step()
# ----------------------- update target networks ----------------------- #
self.soft_update(self.critic_local, self.critic_target, TAU)
self.soft_update(self.actor_local, self.actor_target, TAU)
def soft_update(self, local_model, target_model, tau):
"""Soft update model parameters.
θ_target = τ*θ_local + (1 - τ)*θ_target
Params
======
local_model: PyTorch model (weights will be copied from)
target_model: PyTorch model (weights will be copied to)
tau (float): interpolation parameter
"""
for target_param, local_param in zip(target_model.parameters(), local_model.parameters()):
target_param.data.copy_(tau*local_param.data + (1.0-tau)*target_param.data)
class OUNoise:
"""Ornstein-Uhlenbeck process."""
def __init__(self, size, seed, mu=0., theta=0.15, sigma=0.2):
"""Initialize parameters and noise process."""
self.mu = mu * np.ones(size)
self.theta = theta
self.sigma = sigma
self.seed = random.seed(seed)
self.size = size
self.reset()
def reset(self):
"""Reset the internal state (= noise) to mean (mu)."""
self.state = copy.copy(self.mu)
def sample(self):
"""Update internal state and return it as a noise sample."""
x = self.state
dx = self.theta * (self.mu - x) + self.sigma * np.random.standard_normal(self.size)
self.state = x + dx
return self.state
class ReplayBuffer:
"""Fixed-size buffer to store experience tuples."""
def __init__(self, action_size, buffer_size, batch_size, seed):
"""Initialize a ReplayBuffer object.
Params
======
buffer_size (int): maximum size of buffer
batch_size (int): size of each training batch
"""
self.action_size = action_size
self.memory = deque(maxlen=buffer_size) # internal memory (deque)
self.batch_size = batch_size
self.experience = namedtuple("Experience", field_names=["state", "action", "reward", "next_state", "done"])
self.seed = random.seed(seed)
def add(self, state, action, reward, next_state, done):
"""Add a new experience to memory."""
e = self.experience(state, action, reward, next_state, done)
self.memory.append(e)
def sample(self):
"""Randomly sample a batch of experiences from memory."""
experiences = random.sample(self.memory, k=self.batch_size)
states = torch.from_numpy(np.vstack([e.state for e in experiences if e is not None])).float().to(device)
actions = torch.from_numpy(np.vstack([e.action for e in experiences if e is not None])).float().to(device)
rewards = torch.from_numpy(np.vstack([e.reward for e in experiences if e is not None])).float().to(device)
next_states = torch.from_numpy(np.vstack([e.next_state for e in experiences if e is not None])).float().to(device)
dones = torch.from_numpy(np.vstack([e.done for e in experiences if e is not None]).astype(np.uint8)).float().to(device)
return (states, actions, rewards, next_states, dones)
def __len__(self):
"""Return the current size of internal memory."""
return len(self.memory)
```
## EXAMINE THE STATE AND ACTION SPACES
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
```
## TRAIN DDPG AGENTS
```
num_agents = len(env_info.agents)
agent = Agent(state_size=state_size, action_size=action_size, num_agents=num_agents, random_seed=0)
def ddpg(n_episodes=3000):
total_scores_deque = deque(maxlen=100)
total_scores = []
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name]
states = env_info.vector_observations
scores = np.zeros(num_agents)
agent.reset()
while True:
actions = agent.act(states)
env_info = env.step(actions)[brain_name]
next_states = env_info.vector_observations # get the next state
rewards = env_info.rewards # get the reward
dones = env_info.local_done # see if episode has finished
for state, action, reward, next_state, done in zip(states, actions, rewards, next_states, dones):
agent.step(state, action, reward, next_state, done) # agent takes an action
scores += rewards # update the score
states = next_states # update the state
if np.any(dones): # exit loop if episode finished
break
max_score = np.max(scores)
total_scores_deque.append(max_score)
total_scores.append(max_score)
total_average_score = np.mean(total_scores_deque)
if i_episode % 10 == 0: # print every 10
print('\rEpisode {}\tTotal Average Score: {:.2f}'.format(i_episode, total_average_score))
if total_average_score >= 0.5 and i_episode >= 100:
print('Problem Solved after {} epsisodes. Total Average score: {:.2f}'.format(i_episode, total_average_score))
torch.save(agent.actor_local.state_dict(), 'checkpoint_MA_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_MA_critic.pth')
break
return total_scores
scores = ddpg()
```
## PRINT TRAIN STATISTCS
```
%matplotlib inline
import matplotlib.pyplot as plt
#def __init__(self, state_size, action_size, random_seed):
#agent = Agent(state_size=state_size, action_size=action_size, random_seed=0)
#scores = ddpg(agent)
# plot the scores
average_score = 0.5
success = [average_score] * len(scores)
fig = plt.figure(figsize=(30,20))
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores,np.arange(len(scores)), success)
plt.ylabel('Score', fontsize = 33)
plt.xlabel('Episode #', fontsize = 33)
plt.title('Train DDPG in Multi Agent Environment', fontsize = 33)
plt.gca().legend(('actual score','average'), fontsize = 23)
plt.show()
%matplotlib inline
import pandas as pd
import numpy as np
df = pd.DataFrame(scores, columns=['score'])
df.plot.box(figsize =(30,20), fontsize =33)
```
## WATCH THE AGENTS IN ACTION
```
# Load the saved weights into Pytorch model
agent.actor_local.load_state_dict(torch.load('checkpoint_actor.pth', map_location='cpu'))
agent.critic_local.load_state_dict(torch.load('checkpoint_critic.pth', map_location='cpu'))
for i in range(100): # play game for 5 episodes
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
states = env_info.vector_observations # get the current state (for each agent)
scores = np.zeros(num_agents) # initialize the score (for each agent)
while True:
actions = agent.act(states) # select actions from loaded model agent
actions = np.clip(actions, -1, 1) # all actions between -1 and 1
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/HowEarthEngineWorks/ClientVsServer.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/HowEarthEngineWorks/ClientVsServer.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=HowEarthEngineWorks/ClientVsServer.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/HowEarthEngineWorks/ClientVsServer.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
### CNN German Newset
This notebook follows the steps of the CNN notebook presented in class.
The result seem to be not very good, so I will not go into further details.
Even after 10 epochs the average f1-score is only about 0.5.
Maybe using a more complicated CNN architecture would help but I suspect the number of examples in the dataset to be to small for such a deep learning technique.
```
!pip install 'fhnw-nlp-utils>=0.1.3'
!pip install pyarrow fastparquet fasttext
from fhnw.nlp.utils.storage import load_dataframe
from fhnw.nlp.utils.storage import download
from fhnw.nlp.utils.colab import runs_on_colab
import numpy as np
import pandas as pd
import tensorflow as tf
print("Tensorflow version:", tf.__version__)
#physical_devices = tf.config.list_physical_devices('GPU')
#tf.config.experimental.set_memory_growth(physical_devices[0], True)
gpu_devices = tf.config.experimental.list_physical_devices('GPU')
for device in gpu_devices:
tf.config.experimental.set_memory_growth(device, True)
print("GPU is", "available" if tf.config.list_physical_devices("GPU") else "NOT AVAILABLE")
# Use prepared data
file = "data/german_news_articles_original_train_and_test_tokenized.parq"
data_all = load_dataframe(file)
data_all.sample(3)
from fhnw.nlp.utils.params import get_train_test_split, extract_vocabulary_and_set,create_label_binarizer_and_set, extract_text_vectorization_and_set
# used standard params from class
# Maybe increasing cnn_num_conv_pooling_layers would help
params = {
"verbose": True,
"shuffle": True,
"batch_size": 16,
"X_column_name": "text_clean",
"y_column_name": "label",
"embedding_type": "fasttext",
#"embedding_type": "bytepair",
"embedding_dim": 300,
"embedding_mask_zero": True,
"embedding_trainable": False,
#"embedding_input_sequence_length": output_sequence_length if 'output_sequence_length' in locals() or 'output_sequence_length' in globals() else None,
"embedding_fasttext_model": "cc.de.300.bin",
"embedding_word2vec_model_url": "https://cloud.devmount.de/d2bc5672c523b086/german.model",
"embedding_spacy_model": "de_core_news_md",
"embedding_tensorflow_hub_url": "https://tfhub.dev/google/nnlm-de-dim128-with-normalization/2",
"cnn_num_conv_pooling_layers": 2,
"model_type": "cnn",
}
create_label_binarizer_and_set(params, data_all)
extract_vocabulary_and_set(params, data_all)
extract_text_vectorization_and_set(params)
%%time
from fhnw.nlp.utils.params import extract_embedding_layer_and_set, dataframe_to_dataset
extract_embedding_layer_and_set(params)
data_train_orig = data_all.loc[(data_all["split"] == "train")]
data_test_orig = data_all.loc[(data_all["split"] == "test")]
data_train, data_test = get_train_test_split(params, data_train_orig)
dataset_train = dataframe_to_dataset(params, data_train)
dataset_test = dataframe_to_dataset(params, data_test)
for text, labels in dataset_train.take(1):
print(text)
print(labels)
from fhnw.nlp.utils.params import build_model_cnn, compile_model, train_model
# Use CNN model
model = build_model_cnn(params)
model.summary()
compile_model(params, model)
# Load the TensorBoard notebook extension
#%load_ext tensorboard
def train_model_local(params, model, dataset_train, dataset_val):
"""Performs the model training
Parameters
----------
params: dict
The dictionary containing the parameters
model: model
The keras model
dataset_train: tf Dataset
The dataset for training
dataset_val; tf Dataset
The dataset for validation
Returns
-------
history
The training history
"""
import os
import datetime
from tensorflow import keras
training_epochs = params.get("training_epochs", 5)
training_logdir = params.get("training_logdir", None)
if training_logdir is None:
training_logdir = os.path.join("logs", datetime.datetime.now().strftime("%Y%m%d-%H%M%S"))
checkpoint_path = "training_2/cp-{epoch:04d}.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
cp_callback = tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_path,
verbose=1,
save_weights_only=True,
save_freq=5*params['batch_size'])
tensorboard_callback = keras.callbacks.TensorBoard(training_logdir, histogram_freq=1)
history = model.fit(
dataset_train,
validation_data=dataset_val,
callbacks=[cp_callback],
epochs=training_epochs)
return history
# %tensorboard --logdir logs/
# Training is fast enough so increase number of epochs
params["training_epochs"] = 10
history = train_model_local(params, model, dataset_train, dataset_test)
from fhnw.nlp.utils.ploting import plot_history
from fhnw.nlp.utils.params import save_model
# accuracy seems to tend to 0.5
save_model(params, model, history)
plot_history(history)
# Unfreeze all layers (i.e. make embeddings trainable)
model.trainable = True
model.summary()
from fhnw.nlp.utils.params import re_compile_model
re_compile_model(params, model)
params["training_epochs"] = 10
history = train_model_local(params, model, dataset_train, dataset_test)
save_model(params, model, history)
plot_history(history)
from fhnw.nlp.utils.ploting import report_classification_results
report_classification_results(params, data_train_orig, model)
```
| github_jupyter |
## Overfitting Exercise
In this exercise, we'll build a model that, as you'll see, dramatically overfits the training data. This will allow you to see what overfitting can "look like" in practice.
```
import os
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
```
For this exercise, we'll use gradient boosted trees. In order to implement this model, we'll use the XGBoost package.
```
! pip install xgboost
import xgboost as xgb
```
Here, we define a few helper functions.
```
# number of rows in a dataframe
def nrow(df):
return(len(df.index))
# number of columns in a dataframe
def ncol(df):
return(len(df.columns))
# flatten nested lists/arrays
flatten = lambda l: [item for sublist in l for item in sublist]
# combine multiple arrays into a single list
def c(*args):
return(flatten([item for item in args]))
```
In this exercise, we're going to try to predict the returns of the S&P 500 ETF. This may be a futile endeavor, since many experts consider the S&P 500 to be essentially unpredictable, but it will serve well for the purpose of this exercise. The following cell loads the data.
```
df = pd.read_csv("SPYZ.csv")
```
As you can see, the data file has four columns, `Date`, `Close`, `Volume` and `Return`.
```
df.head()
n = nrow(df)
```
Next, we'll form our predictors/features. In the cells below, we create four types of features. We also use a parameter, `K`, to set the number of each type of feature to build. With a `K` of 25, 100 features will be created. This should already seem like a lot of features, and alert you to the potential that the model will be overfit.
```
predictors = []
# we'll create a new DataFrame to hold the data that we'll use to train the model
# we'll create it from the `Return` column in the original DataFrame, but rename that column `y`
model_df = pd.DataFrame(data = df['Return']).rename(columns = {"Return" : "y"})
# IMPORTANT: this sets how many of each of the following four predictors to create
K = 25
```
Now, you write the code to create the four types of predictors.
```
for L in range(1,K+1):
# this predictor is just the return L days ago, where L goes from 1 to K
# these predictors will be named `R1`, `R2`, etc.
pR = "".join(["R",str(L)])
predictors.append(pR)
for i in range(K+1,n):
# TODO: fill in the code to assign the return from L days before to the ith row of this predictor in `model_df`
model_df.loc[i, pR] = df.loc[i-L,'Return']
# this predictor is the return L days ago, squared, where L goes from 1 to K
# these predictors will be named `Rsq1`, `Rsq2`, etc.
pR2 = "".join(["Rsq",str(L)])
predictors.append(pR2)
for i in range(K+1,n):
# TODO: fill in the code to assign the squared return from L days before to the ith row of this predictor
# in `model_df`
model_df.loc[i, pR2] = (df.loc[i-L,'Return']) ** 2
# this predictor is the log volume L days ago, where L goes from 1 to K
# these predictors will be named `V1`, `V2`, etc.
pV = "".join(["V",str(L)])
predictors.append(pV)
for i in range(K+1,n):
# TODO: fill in the code to assign the log of the volume from L days before to the ith row of this predictor
# in `model_df`
# Add 1 to the volume before taking the log
model_df.loc[i, pV] = math.log(1.0 + df.loc[i-L,'Volume'])
# this predictor is the product of the return and the log volume from L days ago, where L goes from 1 to K
# these predictors will be named `RV1`, `RV2`, etc.
pRV = "".join(["RV",str(L)])
predictors.append(pRV)
for i in range(K+1,n):
# TODO: fill in the code to assign the product of the return and the log volume from L days before to the
# ith row of this predictor in `model_df`
model_df.loc[i, pRV] = model_df.loc[i, pR] * model_df.loc[i, pV]
```
Let's take a look at the predictors we've created.
```
model_df.iloc[100:105,:]
```
Next, we create a DataFrame that holds the recent volatility of the ETF's returns, as measured by the standard deviation of a sliding window of the past 20 days' returns.
```
vol_df = pd.DataFrame(data = df[['Return']])
for i in range(K+1,n):
# TODO: create the code to assign the standard deviation of the return from the time period starting
# 20 days before day i, up to the day before day i, to the ith row of `vol_df`
vol_df.loc[i, 'vol'] = np.std(vol_df.loc[(i-20):(i-1),'Return'])
```
Let's take a quick look at the result.
```
vol_df.iloc[100:105,:]
```
Now that we have our data, we can start thinking about training a model.
```
# for training, we'll use all the data except for the first K days, for which the predictors' values are NaNs
model = model_df.iloc[K:n,:]
```
In the cell below, first split the data into train and test sets, and then split off the targets from the predictors.
```
# Split data into train and test sets
train_size = 2.0/3.0
breakpoint = round(nrow(model) * train_size)
# TODO: fill in the code to split off the chunk of data up to the breakpoint as the training set, and
# assign the rest as the test set.
training_data = model.iloc[1:breakpoint,:]
test_data = model.loc[breakpoint : nrow(model),]
# TODO: Split training data and test data into targets (Y) and predictors (X), for the training set and the test set
X_train = training_data.iloc[:,1:ncol(training_data)]
Y_train = training_data.iloc[:,0]
X_test = test_data.iloc[:,1:ncol(training_data)]
Y_test = test_data.iloc[:,0]
```
Great, now that we have our data, let's train the model.
```
# DMatrix is a internal data structure that used by XGBoost which is optimized for both memory efficiency
# and training speed.
dtrain = xgb.DMatrix(X_train, Y_train)
# Train the XGBoost model
param = { 'max_depth':20, 'silent':1 }
num_round = 20
xgModel = xgb.train(param, dtrain, num_round)
```
Now let's predict the returns for the S&P 500 ETF in both the train and test periods. If the model is successful, what should the train and test accuracies look like? What would be a key sign that the model has overfit the training data?
Todo: Before you run the next cell, write down what you expect to see if the model is overfit.
```
# Make the predictions on the test data
preds_train = xgModel.predict(xgb.DMatrix(X_train))
preds_test = xgModel.predict(xgb.DMatrix(X_test))
```
Let's quickly look at the mean squared error of the predictions on the training and testing sets.
```
# TODO: Calculate the mean squared error on the training set
msetrain = sum((preds_train-Y_train)**2)/len(preds_train)
msetrain
# TODO: Calculate the mean squared error on the test set
msetest = sum((preds_test-Y_test)**2)/len(preds_test)
msetest
```
Looks like the mean squared error on the test set is an order of magnitude greater than on the training set. Not a good sign. Now let's do some quick calculations to gauge how this would translate into performance.
```
# combine prediction arrays into a single list
predictions = c(preds_train, preds_test)
responses = c(Y_train, Y_test)
# as a holding size, we'll take predicted return divided by return variance
# this is mean-variance optimization with a single asset
vols = vol_df.loc[K:n,'vol']
position_size = predictions / vols ** 2
# TODO: Calculate pnl. Pnl in each time period is holding * realized return.
performance = position_size * responses
# plot simulated performance
plt.plot(np.cumsum(performance))
plt.ylabel('Simulated Performance')
plt.axvline(x=breakpoint, c = 'r')
plt.show()
```
Our simulated returns accumulate throughout the training period, but they are absolutely flat in the testing period. The model has no predictive power whatsoever in the out-of-sample period.
Can you think of a few reasons our simulation of performance is unrealistic?
```
# TODO: Answer the above question.
```
1. We left out any accounting of trading costs. If we had included trading costs, the performance in the out-of-sample period would be downward!
2. We didn't account for any time for trading. It's most conservative to assume that we would make trades on the day following our calculation of position size to take, and realize returns the day after that, such that there's a two-day delay between holding size calculation and realized return.
If you need a little assistance, check out the [solution](overfitting_exercise_solution.ipynb).
| github_jupyter |
```
#https://towardsdatascience.com/categorical-encoding-using-label-encoding-and-one-hot-encoder-911ef77fb5bd
#https://www.kaggle.com/ldfreeman3/a-data-science-framework-to-achieve-99-accuracy
#https://www.kaggle.com/leonardolima/titanic-competition-notebook-0-80-top-10
!git clone https://gitlab.com/mirsakhawathossain/pha-ml.git
!cd pha-ml
!ls
import pandas as pd
asteroid_data = pd.read_csv('pha-ml/Dataset/dataset.csv',index_col=['spkid']).sort_index(axis=0)
asteroid_data.head(5)
asteroid_data.tail(5)
asteroid_data.isnull().sum()
asteroid_data.shape
dataframe=asteroid_data.drop(columns=['name','epoch','prefix','orbit_id','id','pha','full_name','pdes','diameter_sigma','epoch_mjd','epoch_cal','equinox','neo','sigma_e','sigma_a','sigma_q','sigma_i','sigma_om','sigma_om','sigma_w','sigma_ma','sigma_ad','sigma_n','sigma_tp','sigma_per','rms','per_y','moid','tp','tp_cal'])
dataframe.head(5)
dataframe.tail(5)
dataframe.isnull().sum()
dataframe['class'].value_counts()
dataframe.info()
dataframe['diameter']=dataframe['diameter'].astype('float64')
dataframe['class']=dataframe['class'].astype('category')
dataframe.info()
dataframe['diameter'].fillna(dataframe.groupby('class')['diameter'].transform('median'),inplace=True)
dataframe['H'].fillna(dataframe.groupby('class')['H'].transform('median'),inplace=True)
dataframe['albedo'].fillna(dataframe.groupby('class')['albedo'].transform('median'),inplace=True)
dataframe['ma'].fillna(dataframe.groupby('class')['ma'].transform('median'),inplace=True)
dataframe['ad'].fillna(dataframe.groupby('class')['ad'].transform('median'),inplace=True)
dataframe['per'].fillna(dataframe.groupby('class')['per'].transform('median'),inplace=True)
dataframe['moid_ld'].fillna(dataframe.groupby('class')['moid_ld'].transform('median'),inplace=True)
dataframe.groupby('class')['diameter'].transform('median')
dataframe.head(5)
dataframe.tail(5)
dataframe.isnull().sum()
dataframe[['diameter','albedo','ad','per']]=dataframe[['diameter','albedo','ad','per']].fillna(value=dataframe[['diameter','albedo','ad','per']].median())
dataframe.isnull().sum()
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set()
correlation=dataframe.corr()
correlation
sns.heatmap(dataframe.corr(),cmap='coolwarm',annot=True)
dataframe.describe()
df_asteroid=dataframe[['class','diameter']]
df_asteroid.head(5)
df_asteroid.groupby('class')['diameter'].mean().plot(kind='pie',figsize=(20,15),title='Mean Diameter of Asteroids')
plt.show()
df_asteroid.groupby('class')['diameter'].mean().plot(kind='bar',figsize=(20,15),title='Mean Diameter of Asteroids')
plt.show()
from sklearn.preprocessing import LabelEncoder
labelencoder = LabelEncoder()
dataframe['class_cat']=labelencoder.fit_transform(dataframe['class'])
dataframe.head(5)
dataframe['class_cat'].value_counts()
X=dataframe.drop(['class','class_cat'],axis=1)
y=dataframe['class_cat']
X.shape
y.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=4)
print(X_train.shape)
print(X_test.shape)
X_test.isnull().sum()
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=5)
knn.fit(X_train, y_train)
y_pred=knn.predict(X_test)
from sklearn import metrics
print(metrics.accuracy_score(y_test,y_pred))
neighbors_settings = list(range(1,26))
scores = []
for k in neighbors_settings:
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train,y_train)
y_pred = knn.predict(X_test)
scores.append(metrics.accuracy_score(y_test,y_pred))
plt.plot(neighbors_settings,scores)
plt.xlabel('Value of K for KNN')
plt.ylabel('Testing Accuracy')
```
| github_jupyter |
# AOC 2020 Day 11
Seat layout input is a grid:
```
L.LL.LL.LL
LLLLLLL.LL
L.L.L..L..
LLLL.LL.LL
L.LL.LL.LL
L.LLLLL.LL
..L.L.....
LLLLLLLLLL
L.LLLLLL.L
L.LLLLL.LL
```
Grid entries can be one of:
- `L` - empty seat
- `#` - occupied seat
- `.` - floor
Rules are based on adjacent seats, 8 surrounding seats ala chess king moves (U,D,L,R,UL,UR,DL,DR).
Rules to apply are:
1. If a seat is empty (L) and there are no occupied seats adjacent to it, the seat becomes occupied.
2. If a seat is occupied (#) and four or more seats adjacent to it are also occupied, the seat becomes empty.
3. Otherwise, the seat's state does not change.
Floor seats do not change.
When rules are applied they stabilize, at that point count occupied seats, for this grid it's `37`.
Input after one round of rules:
```
#.##.##.##
#######.##
#.#.#..#..
####.##.##
#.##.##.##
#.#####.##
..#.#.....
##########
#.######.#
#.#####.##
```
After second round:
```
#.LL.L#.##
#LLLLLL.L#
L.L.L..L..
#LLL.LL.L#
#.LL.LL.LL
#.LLLL#.##
..L.L.....
#LLLLLLLL#
#.LLLLLL.L
#.#LLLL.##
```
Eventual stable state after 3 more rounds:
```
#.#L.L#.##
#LLL#LL.L#
L.#.L..#..
#L##.##.L#
#.#L.LL.LL
#.#L#L#.##
..L.L.....
#L#L##L#L#
#.LLLLLL.L
#.#L#L#.##
```
```
SampleInput="""L.LL.LL.LL
LLLLLLL.LL
L.L.L..L..
LLLL.LL.LL
L.LL.LL.LL
L.LLLLL.LL
..L.L.....
LLLLLLLLLL
L.LLLLLL.L
L.LLLLL.LL"""
def load_seat_map(input):
result = []
for line in input.split('\n'):
if line != '':
row = [seat for seat in line]
result.append(row)
return result
sample_map0 = load_seat_map(SampleInput)
sample_map0
def print_seat_map(seat_map):
max_row = len(seat_map)
max_col = len(seat_map[0])
for r in seat_map:
print("".join(r))
print("max_row: {}, max_col: {}".format(max_row, max_col))
def adjacent_seats(seat_map, row, col):
"""Return a list of all adjacent seats for position row, column in seat_map"""
adj = []
max_row = len(seat_map)
max_col = len(seat_map[0])
for r in range(row-1,row+2):
for c in range(col-1,col+2):
# skip invalid co-ordinates
if r < 0:
continue
if c < 0:
continue
if r >= max_row:
continue
if c >= max_col:
continue
if r == row and c == col:
continue
#print("({},{}): {}".format(r,c, seat_map[r][c]))
adj.append(seat_map[r][c])
return adj
print("adjacent to row {}, col {} :- {}".format(0, 0, adjacent_seats(sample_map0, 0, 0)))
print("adjacent to row {}, col {} :- {}".format(1, 1, adjacent_seats(sample_map0, 1, 1)))
print("adjacent to row {}, col {} :- {}".format(9, 0, adjacent_seats(sample_map0, 9, 0)))
def apply_rules(seat_map):
"""Apply rules to seat_map"""
result = []
max_row = len(seat_map)
max_col = len(seat_map[0])
# create blank result grid
for r in range(0, max_row):
result.append([])
for c in range(0, max_col):
result[r].append('')
# apply rules
# 1. If a seat is empty (L) and there are no occupied seats adjacent to it, the seat becomes occupied.
# 2. If a seat is occupied (#) and four or more seats adjacent to it are also occupied, the seat becomes empty.
# 3. Otherwise, the seat's state does not change.
for r in range(0, max_row):
for c in range(0, max_col):
seat = seat_map[r][c]
neighbours = adjacent_seats(seat_map, r, c)
if seat == '.':
result[r][c] = '.'
if seat == 'L' and '#' not in neighbours:
result[r][c] = '#'
if seat == '#' and neighbours.count('#') >= 4:
result[r][c] = 'L'
elif seat == '#':
result[r][c] = '#'
return(result)
expected_sample_map1 = load_seat_map("""#.##.##.##
#######.##
#.#.#..#..
####.##.##
#.##.##.##
#.#####.##
..#.#.....
##########
#.######.#
#.#####.##""")
assert apply_rules(sample_map0) == expected_sample_map1, "invalid result!"
expected_sample_map2 = load_seat_map("""#.LL.L#.##
#LLLLLL.L#
L.L.L..L..
#LLL.LL.L#
#.LL.LL.LL
#.LLLL#.##
..L.L.....
#LLLLLLLL#
#.LLLLLL.L
#.#LLLL.##""")
assert apply_rules(apply_rules(sample_map0)) == expected_sample_map2, "invalid result!"
def part1(input):
last_map = load_seat_map(input)
while apply_rules(last_map) != last_map:
last_map = apply_rules(last_map)
result = 0
for row in last_map:
result += row.count('#')
return result
assert part1(SampleInput) == 37, "invalid result - got {}".format(part1(SampleInput))
part1(SampleInput)
day11 = open("./inputs/day11").read()
day11_map = load_seat_map(day11)
#print_seat_map(day11_map)
#apply_rules(day11_map)
part1(day11)
```
## part 2
Updated adjacency rules, not just nearest neighbor, any neighbor you can see in any direction.
For example below the empty seat sees 8 neighbors:
```.......#.
...#.....
.#.......
.........
..#L....#
....#....
.........
#........
...#.....
```
Whereas for below the left most empty seat only sees one empty seat (to its right):
```.............
.L.L.#.#.#.#.
.............
```
And for this empty seat below it sees no neighbors:
```.##.##.
#.#.#.#
##...##
...L...
##...##
#.#.#.#
.##.##.
```
So basically sudoku "seeing" ... does any value in same row, column or diagonal have a '#' or a 'L'?
```
def look(seat_map, row, col, row_offset, col_offset):
"""
'Look' from row,col in direction specified by row/col_offset
Return the first key thing we encounter, will be one of:
- 'L' - hit an empty seat
- '#' - hit an occupied seat
- None - hit edge of the grid
"""
min_row = 0
min_col = 0
max_row = len(seat_map)
max_col = len(seat_map[0])
seen = None
r = row
c = col
while not seen:
r = r + row_offset
c = c + col_offset
if r < min_row or c < min_col or r >= max_row or c >= max_col:
break
s = seat_map[r][c]
#print("Looking at ({}, {}): {}".format(r, c, s))
if s == '.':
continue # skip blank
seen = s
return seen
test_look_grid1 = load_seat_map(""".L.L.#.#.#.#.
.............""")
assert look(test_look_grid1, 0, 1, 0, 1) == 'L', "expected 'L', got '{}'".format(look(test_look_grid1, 0, 1, 0, 1))
test_look_grid2 = load_seat_map(""".......#.
...#.....
.#.......
.........
..#L....#
....#....
.........
#........
...#.....""")
test_grid = test_look_grid2
print_seat_map(test_grid)
print()
s = (4, 3)
print("starting at ({},{}): {}".format(s[0], s[1], test_grid[s[0]][s[1]]))
for r in (-1, 0, 1):
for c in (-1, 0, 1):
if (r, c) != (0, 0):
print("looking from ({},{}):{} in direction ({}, {})".format(s[0], s[1], test_grid[s[0]][s[1]],r, c))
assert look(test_grid, s[0], s[1], r, c) == '#', "expected '#', got '{}'".format(look(test_grid, s[0], s[1], r, c))
print("Ok, saw None")
if (r, c) == (0,0):
print("Not feeling introspective so not looking inwards (0,0)")
test_look_grid3 = load_seat_map(""".##.##.
#.#.#.#
##...##
...L...
##...##
#.#.#.#
.##.##.""")
test_grid = test_look_grid3
print_seat_map(test_grid)
print()
s = (3, 3)
print("starting at ({},{}): {}".format(s[0], s[1], test_grid[s[0]][s[1]]))
for r in (-1, 0, 1):
for c in (-1, 0, 1):
if (r, c) != (0, 0):
print("looking from ({},{}):{} in direction ({}, {})".format(s[0], s[1], test_grid[s[0]][s[1]],r, c))
assert look(test_grid, s[0], s[1], r, c) == None, "expected None, got '{}'".format(look(test_grid, s[0], s[1], r, c))
print("Ok, saw None")
if (r, c) == (0,0):
print("Not feeling introspective so not looking inwards (0,0)")
def new_adj_seats(seat_map, row, col):
adj = []
for r in (-1, 0, 1):
for c in (-1, 0, 1):
if (r, c) != (0, 0):
saw = look(seat_map, row, col, r, c)
if saw:
adj.append(saw)
return adj
def new_apply_rules(seat_map):
"""Apply rules to seat_map"""
result = []
max_row = len(seat_map)
max_col = len(seat_map[0])
# create blank result grid
for r in range(0, max_row):
result.append([])
for c in range(0, max_col):
result[r].append('')
# apply rules
# 1. If a seat is empty (L) and there are no occupied seats adjacent to it, the seat becomes occupied.
# 2. If a seat is occupied (#) and five or more seats adjacent to it are also occupied, the seat becomes empty.
# 3. Otherwise, the seat's state does not change.
for r in range(0, max_row):
for c in range(0, max_col):
seat = seat_map[r][c]
neighbours = new_adj_seats(seat_map, r, c)
if seat == '.':
result[r][c] = '.'
elif seat == 'L' and '#' not in neighbours:
result[r][c] = '#'
elif seat == '#' and neighbours.count('#') >= 5:
result[r][c] = 'L'
else:
#print("final else case, seat: {}, neighbours: {}".format(seat, neighbours))
result[r][c] = seat
return result
def part2(input):
last_map = load_seat_map(input)
while new_apply_rules(last_map) != last_map:
last_map = new_apply_rules(last_map)
#print("Stable map:")
#print_seat_map(last_map)
result = 0
for row in last_map:
result += row.count('#')
return result
print("Starting with sample input:")
sample_map0 = load_seat_map(SampleInput)
print_seat_map(sample_map0)
expected_sample_map1_p2 = load_seat_map("""#.##.##.##
#######.##
#.#.#..#..
####.##.##
#.##.##.##
#.#####.##
..#.#.....
##########
#.######.#
#.#####.##""")
print("observed result from new_apply_rules(sample input)")
print_seat_map(new_apply_rules(sample_map0))
print()
print("expected result from new_apply_rules(sample input)")
print_seat_map(expected_sample_map1_p2)
assert new_apply_rules(sample_map0) == expected_sample_map1_p2, "invalid result!"
expected_sample_map2_p2 = load_seat_map("""#.LL.LL.L#
#LLLLLL.LL
L.L.L..L..
LLLL.LL.LL
L.LL.LL.LL
L.LLLLL.LL
..L.L.....
LLLLLLLLL#
#.LLLLLL.L
#.LLLLL.L#""")
assert new_apply_rules(new_apply_rules(sample_map0)) == expected_sample_map2_p2, "invalid result!"
p2_sample = part2(SampleInput)
assert p2_sample == 26, "invalid result - got {}".format(p2_sample)
p2_sample
part2(day11)
```
| github_jupyter |
# Create a Learner for inference
```
from fastai.gen_doc.nbdoc import *
```
In this tutorial, we'll see how the same API allows you to create an empty [`DataBunch`](/basic_data.html#DataBunch) for a [`Learner`](/basic_train.html#Learner) at inference time (once you have trained your model) and how to call the `predict` method to get the predictions on a single item.
```
jekyll_note("""As usual, this page is generated from a notebook that you can find in the <code>docs_src</code> folder of the
<a href="https://github.com/fastai/fastai">fastai repo</a>. We use the saved models from <a href="/tutorial.data.html">this tutorial</a> to
have this notebook run quickly.""")
```
## Vision
To quickly get acces to all the vision functionality inside fastai, we use the usual import statements.
```
from fastai.vision import *
```
### A classification problem
Let's begin with our sample of the MNIST dataset.
```
mnist = untar_data(URLs.MNIST_TINY)
tfms = get_transforms(do_flip=False)
```
It's set up with an imagenet structure so we use it to split our training and validation set, then labelling.
```
data = (ImageItemList.from_folder(mnist)
.split_by_folder()
.label_from_folder()
.add_test_folder('test')
.transform(tfms, size=32)
.databunch()
.normalize(imagenet_stats))
```
Now that our data has been properly set up, we can train a model. We already did in the [look at your data tutorial](/tutorial.data.html) so we'll just load our saved results here.
```
learn = create_cnn(data, models.resnet18).load('mini_train')
```
Once everything is ready for inference, we just have to call `learn.export` to save all the information of our [`Learner`](/basic_train.html#Learner) object for inference: the stuff we need in the [`DataBunch`](/basic_data.html#DataBunch) (transforms, classes, normalization...), the model with its weights and all the callbacks our [`Learner`](/basic_train.html#Learner) was using. Everything will be in a file named `export.pkl` in the folder `learn.path`. If you deploy your model on a different machine, this is the file you'll need to copy.
```
learn.export()
```
To create the [`Learner`](/basic_train.html#Learner) for inference, you'll need to use the [`load_learner`](/basic_train.html#load_learner) function. Note that you don't have to specify anything: it remembers the classes, the transforms you used or the normalization in the data, the model, its weigths... The only argument needed is the folder where the 'export.pkl' file is.
```
learn = load_learner(mnist)
```
You can now get the predictions on any image via `learn.predict`.
```
img = data.train_ds[0][0]
learn.predict(img)
```
It returns a tuple of three things: the object predicted (with the class in this instance), the underlying data (here the corresponding index) and the raw probabilities. You can also do inference on a larger set of data by adding a *test set*. This is done by passing an [`ItemList`](/data_block.html#ItemList) to [`load_learner`](/basic_train.html#load_learner).
```
learn = load_learner(mnist, test=ImageItemList.from_folder(mnist/'test'))
preds,y = learn.get_preds(ds_type=DatasetType.Test)
preds[:5]
```
### A multi-label problem
Now let's try these on the planet dataset, which is a little bit different in the sense that each image can have multiple tags (and not just one label).
```
planet = untar_data(URLs.PLANET_TINY)
planet_tfms = get_transforms(flip_vert=True, max_lighting=0.1, max_zoom=1.05, max_warp=0.)
```
Here each images is labelled in a file named `labels.csv`. We have to add [`train`](/train.html#train) as a prefix to the filenames, `.jpg` as a suffix and indicate that the labels are separated by spaces.
```
data = (ImageItemList.from_csv(planet, 'labels.csv', folder='train', suffix='.jpg')
.random_split_by_pct()
.label_from_df(label_delim=' ')
.transform(planet_tfms, size=128)
.databunch()
.normalize(imagenet_stats))
```
Again, we load the model we saved in [look at your data tutorial](/tutorial.data.html).
```
learn = create_cnn(data, models.resnet18).load('mini_train')
```
Then we can export it before loading it for inference.
```
learn.export()
learn = load_learner(planet)
```
And we get the predictions on any image via `learn.predict`.
```
img = data.train_ds[0][0]
learn.predict(img)
```
Here we can specify a particular threshold to consider the predictions to be correct or not. The default is `0.5`, but we can change it.
```
learn.predict(img, thresh=0.3)
```
### A regression example
For the next example, we are going to use the [BIWI head pose](https://data.vision.ee.ethz.ch/cvl/gfanelli/head_pose/head_forest.html#db) dataset. On pictures of persons, we have to find the center of their face. For the fastai docs, we have built a small subsample of the dataset (200 images) and prepared a dictionary for the correspondance fielname to center.
```
biwi = untar_data(URLs.BIWI_SAMPLE)
fn2ctr = pickle.load(open(biwi/'centers.pkl', 'rb'))
```
To grab our data, we use this dictionary to label our items. We also use the [`PointsItemList`](/vision.data.html#PointsItemList) class to have the targets be of type [`ImagePoints`](/vision.image.html#ImagePoints) (which will make sure the data augmentation is properly applied to them). When calling [`transform`](/tabular.transform.html#tabular.transform) we make sure to set `tfm_y=True`.
```
data = (PointsItemList.from_folder(biwi)
.random_split_by_pct(seed=42)
.label_from_func(lambda o:fn2ctr[o.name])
.transform(get_transforms(), tfm_y=True, size=(120,160))
.databunch()
.normalize(imagenet_stats))
```
As before, the road to inference is pretty straightforward: load the model we trained before, export the [`Learner`](/basic_train.html#Learner) then load it for production.
```
learn = create_cnn(data, models.resnet18, lin_ftrs=[100], ps=0.05).load('mini_train');
learn.export()
learn = load_learner(biwi)
```
And now we can a prediction on an image.
```
img = data.valid_ds[0][0]
learn.predict(img)
```
To visualize the predictions, we can use the [`Image.show`](/vision.image.html#Image.show) method.
```
img.show(y=learn.predict(img)[0])
```
### A segmentation example
Now we are going to look at the [camvid dataset](http://mi.eng.cam.ac.uk/research/projects/VideoRec/CamVid/) (at least a small sample of it), where we have to predict the class of each pixel in an image. Each image in the 'images' subfolder as an equivalent in 'labels' that is its segmentations mask.
```
camvid = untar_data(URLs.CAMVID_TINY)
path_lbl = camvid/'labels'
path_img = camvid/'images'
```
We read the classes in 'codes.txt' and the function maps each image filename with its corresponding mask filename.
```
codes = np.loadtxt(camvid/'codes.txt', dtype=str)
get_y_fn = lambda x: path_lbl/f'{x.stem}_P{x.suffix}'
```
The data block API allows us to uickly get everything in a [`DataBunch`](/basic_data.html#DataBunch) and then we can have a look with `show_batch`.
```
data = (SegmentationItemList.from_folder(path_img)
.random_split_by_pct()
.label_from_func(get_y_fn, classes=codes)
.transform(get_transforms(), tfm_y=True, size=128)
.databunch(bs=16, path=camvid)
.normalize(imagenet_stats))
```
As before, we load our model, export the [`Learner`](/basic_train.html#Learner) then create a new one with [`load_learner`](/basic_train.html#load_learner).
```
learn = unet_learner(data, models.resnet18).load('mini_train');
learn.export()
learn = load_learner(camvid)
```
And now we can a prediction on an image.
```
img = data.train_ds[0][0]
learn.predict(img);
```
To visualize the predictions, we can use the [`Image.show`](/vision.image.html#Image.show) method.
```
img.show(y=learn.predict(img)[0])
```
## Text
Next application is text, so let's start by importing everything we'll need.
```
from fastai.text import *
```
### Language modelling
First let's look a how to get a language model ready for inference. Since we'll load the model trained in the [visualize data tutorial](/tutorial.data.html), we load the vocabulary used there.
```
imdb = untar_data(URLs.IMDB_SAMPLE)
vocab = Vocab(pickle.load(open(imdb/'tmp'/'itos.pkl', 'rb')))
data_lm = (TextList.from_csv(imdb, 'texts.csv', cols='text', vocab=vocab)
.random_split_by_pct()
.label_for_lm()
.databunch())
```
Like in vision, we just have to type `learn.export()` after loading our pretrained model to save all the information inside the [`Learner`](/basic_train.html#Learner) we'll need. In this case, this includes all the vocabulary we created. The only difference is that we will specify a filename, since we have several model in the same path (language model and classifier).
```
learn = language_model_learner(data_lm).load('mini_train_lm', with_opt=False);
learn.export(fname = 'export_lm.pkl')
```
Now let's define our inference learner.
```
learn = load_learner(imdb, fname = 'export_lm.pkl')
```
Then we can predict with the usual method, here we can specify how many words we want the model to predict.
```
learn.predict('This is a simple test of', n_words=20)
```
### Classification
Now let's see a classification example. We have to use the same vocabulary as for the language model if we want to be able to use the encoder we saved.
```
data_clas = (TextList.from_csv(imdb, 'texts.csv', cols='text', vocab=vocab)
.split_from_df(col='is_valid')
.label_from_df(cols='label')
.databunch(bs=42))
```
Again we export the [`Learner`](/basic_train.html#Learner) where we load our pretrained model.
```
learn = text_classifier_learner(data_clas).load('mini_train_clas', with_opt=False);
learn.export(fname = 'export_clas.pkl')
```
Now let's use [`load_learner`](/basic_train.html#load_learner).
```
learn = load_learner(imdb, fname = 'export_clas.pkl')
```
Then we can predict with the usual method.
```
learn.predict('I really loved that movie!')
```
## Tabular
Last application brings us to tabular data. First let's import everything we'll need.
```
from fastai.tabular import *
```
We'll use a sample of the [adult dataset](https://archive.ics.uci.edu/ml/datasets/adult) here. Once we read the csv file, we'll need to specify the dependant variable, the categorical variables, the continuous variables and the processors we want to use.
```
adult = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(adult/'adult.csv')
dep_var = 'salary'
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race', 'sex', 'native-country']
cont_names = ['education-num', 'hours-per-week', 'age', 'capital-loss', 'fnlwgt', 'capital-gain']
procs = [FillMissing, Categorify, Normalize]
```
Then we can use the data block API to grab everything together.
```
data = (TabularList.from_df(df, path=adult, cat_names=cat_names, cont_names=cont_names, procs=procs)
.split_by_idx(valid_idx=range(800,1000))
.label_from_df(cols=dep_var)
.databunch())
```
We define a [`Learner`](/basic_train.html#Learner) object that we fit and then save the model.
```
learn = tabular_learner(data, layers=[200,100], metrics=accuracy)
learn.fit(1, 1e-2)
learn.save('mini_train')
```
As in the other applications, we just have to type `learn.export()` to save everything we'll need for inference (here it includes the inner state of each processor).
```
learn.export()
```
Then we create a [`Learner`](/basic_train.html#Learner) for inference like before.
```
learn = load_learner(adult)
```
And we can predict on a row of dataframe that has the right `cat_names` and `cont_names`.
```
learn.predict(df.iloc[0])
```
| github_jupyter |
This notebook is developed using the `Python 3 (Data Science)` kernel on an `ml.t3.medium` instance.
```
import sagemaker
import json
import boto3
role = sagemaker.get_execution_role()
sess = sagemaker.Session()
region = sess.boto_region_name
bucket = sess.default_bucket()
prefix = 'sagemaker-studio-book/chapter05'
from time import gmtime, strftime
import time
!wget -q https://github.com/le-scientifique/torchDatasets/raw/master/dbpedia_csv.tar.gz
!tar -xzf dbpedia_csv.tar.gz
!head dbpedia_csv/train.csv -n 3
!grep -i "automatic electric" dbpedia_csv/train.csv
!cat dbpedia_csv/classes.txt
d_label = {}
with open('dbpedia_csv/classes.txt') as f:
for i, label in enumerate(f.readlines()):
d_label[str(i + 1)] = label.strip()
print(d_label)
import nltk
nltk.download('punkt')
def transform_text(row):
cur_row = []
label = f'__label__{d_label[row[0]]}' # Prefix the index-ed label with __label__
cur_row.append(label)
cur_row.extend(nltk.word_tokenize(row[1].lower()))
cur_row.extend(nltk.word_tokenize(row[2].lower()))
return cur_row
from random import shuffle
import multiprocessing
from multiprocessing import Pool
import csv
def preprocess(input_file, output_file, keep=1):
all_rows = []
with open(input_file, 'r') as csvinfile:
csv_reader = csv.reader(csvinfile, delimiter=',')
for row in csv_reader:
all_rows.append(row)
shuffle(all_rows)
all_rows = all_rows[: int(keep * len(all_rows))]
pool = Pool(processes=multiprocessing.cpu_count())
transformed_rows = pool.map(transform_text, all_rows)
pool.close()
pool.join()
with open(output_file, 'w') as csvoutfile:
csv_writer = csv.writer(csvoutfile, delimiter=' ', lineterminator='\n')
csv_writer.writerows(transformed_rows)
%%time
preprocess('dbpedia_csv/train.csv', 'dbpedia_csv/dbpedia.train', keep=0.2)
preprocess('dbpedia_csv/test.csv', 'dbpedia_csv/dbpedia.validation')
!head -n 1 dbpedia_csv/dbpedia.train
image=sagemaker.image_uris.retrieve(framework='blazingtext',
region=region,
version='1')
print(image)
s3_output_location = f's3://{bucket}/{prefix}/output'
estimator = sagemaker.estimator.Estimator(
image,
role,
instance_count=1,
instance_type='ml.c5.2xlarge',
volume_size=30,
max_run=360000,
input_mode='File',
enable_sagemaker_metrics=True,
output_path=s3_output_location,
hyperparameters={
'mode': 'supervised',
'epochs': 20,
'min_count': 2,
'learning_rate': 0.05,
'vector_dim': 10,
'early_stopping': True,
'patience': 4,
'min_epochs': 5,
'word_ngrams': 2,
},
)
train_channel = prefix + '/train'
validation_channel = prefix + '/validation'
sess.upload_data(path='dbpedia_csv/dbpedia.train', bucket=bucket, key_prefix=train_channel)
sess.upload_data(path='dbpedia_csv/dbpedia.validation', bucket=bucket, key_prefix=validation_channel)
s3_train_data = f's3://{bucket}/{train_channel}'
s3_validation_data = f's3://{bucket}/{validation_channel}'
print(s3_train_data)
print(s3_validation_data)
data_channels = {'train': s3_train_data,
'validation': s3_validation_data}
exp_datetime = strftime('%Y-%m-%d-%H-%M-%S', gmtime())
jobname = f'dbpedia-blazingtext-{exp_datetime}'
estimator.fit(inputs=data_channels,
job_name=jobname,
logs=True)
estimator.model_data
!aws s3 cp {estimator.model_data} ./dbpedia_csv/
%%sh
cd dbpedia_csv/
tar -zxf model.tar.gz
```
-------
```
!pip install -q sagemaker-experiments
from smexperiments.experiment import Experiment
from smexperiments.trial import Trial
from botocore.exceptions import ClientError
experiment_name = 'dbpedia-text-classification'
try:
experiment = Experiment.create(
experiment_name=experiment_name,
description='Training a text classification model using dbpedia dataset.')
except ClientError as e:
print(f'{experiment_name} experiment already exists! Reusing the existing experiment.')
for lr in [0.1, 0.01, 0.001]:
exp_datetime = strftime('%Y-%m-%d-%H-%M-%S', gmtime())
jobname = f'dbpedia-blazingtext-{exp_datetime}'
# Creating a new trial for the experiment
exp_trial = Trial.create(
experiment_name=experiment_name,
trial_name=jobname)
experiment_config={
'ExperimentName': experiment_name,
'TrialName': exp_trial.trial_name,
'TrialComponentDisplayName': 'Training'}
estimator = sagemaker.estimator.Estimator(
image,
role,
instance_count=1,
instance_type='ml.c5.2xlarge',
volume_size=30,
max_run=360000,
input_mode='File',
enable_sagemaker_metrics=True,
output_path=s3_output_location,
hyperparameters={
'mode': 'supervised',
'epochs': 40,
'min_count': 2,
'learning_rate': lr,
'vector_dim': 10,
'early_stopping': True,
'patience': 4,
'min_epochs': 5,
'word_ngrams': 2},
)
estimator.fit(
inputs=data_channels,
job_name=jobname,
experiment_config=experiment_config,
wait=False)
print(f'Submitted training job {jobname}')
```
| github_jupyter |
# JAGS example in PyMC3
This notebook attempts to solve the same problem that has been solved manually in [w02-04b-mcmc-demo-continuous.ipynb](http://localhost:8888/notebooks/w02-04b-mcmc-demo-continuous.ipynb), but using PyMC3 instead of JAGS as demonstrated in the course video.
## Problem Definition
Data is for personnel change from last year to this year for 10 companies. Model is defined as follows:
$$y_i | \mu \overset{iid}{\sim} N(\mu, 1)$$
$$\mu \sim t(0, 1, 1)$$
where y<sub>i</sub> represents personnel change for company i, and the distribution of y<sub>i</sub> given $\mu$ is a Normal distribution with mean $\mu$ and variance 1. Prior distribution of $\mu$ is a t distribution with location 0, scale parameter 1, and degrees of freedom 1 (also known as Cauchy's distribution).
Model is not conjugate, thus posterior is not a standard form that we can conveniently sample. To get posterior samples, we will need to setup a Markov chain, whose stationery distribution is the posterior distribution we want.
Main difference with manually solved example is that we don't need to compute the analytical form of the posterior for our simulation.
## PyMC3 Solution
JAGS usage follows the 4 step process:
* Specify model -- this is the first 2 lines in the `with model` block in cell 3.
* Setup model -- this is the observed attribute in y_obs where real values of y are plugged in.
* Run MCMC sampler -- the block under the `run MCMC sampler` command. The call to `update` and `coda.sample` is merged into a single `pm.sample` call with separate `n_iter` and `n_tune` variables. The `step` attribute is set to Metropolis-Hastings as that is the preferred sampler in the course, PyMC3 default is the NUTS sampler.
* Post-processing -- whatever we do with `trace["mu"]` after the sampling is done.
```
import matplotlib.pyplot as plt
import numpy as np
import pymc3 as pm
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
y = np.array([1.2, 1.4, -0.5, 0.3, 0.9, 2.3, 1.0, 0.1, 1.3, 1.9])
n_iter = 1000
n_tune = 500
with pm.Model() as model:
# model specification, and setup (set observed=y)
mu = pm.StudentT("mu", nu=1, mu=0, sigma=1)
y_obs = pm.Normal("y_obs", mu=mu, sigma=1, observed=y)
# run MCMC sampler
step = pm.Metropolis() # PyMC3 default is NUTS, course uses Metropolis-Hastings
trace = pm.sample(n_iter, tune=n_tune, step=step)
# post-processing
mu_sims = trace["mu"]
print("mu_sims :", mu_sims)
print("len(mu_sims): {:d}".format(len(mu_sims)))
_ = pm.traceplot(trace)
_ = pm.traceplot(trace, combined=True)
pm.summary(trace)
```
## Reference
* [Markov Chain Monte Carlo for Bayesian Inference - the Metropolis Algorithm](https://www.quantstart.com/articles/Markov-Chain-Monte-Carlo-for-Bayesian-Inference-The-Metropolis-Algorithm/)
| github_jupyter |
Importando as bibliotecas
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.formula.api as smf
import seaborn as sns
import random
import scipy.stats
import datetime
pd.set_option("display.max_rows", 18, "display.max_columns", 18)
```
Preparando uma base de dados geral
```
#verificando a quantidade de dados por arquivo
rows_aluno = sum(1 for line in open('DM_ALUNO.csv')) - 1
rows_curso = sum(1 for line in open('DM_CURSO.csv')) - 1
rows_docente = sum(1 for line in open('DM_DOCENTE.csv')) - 1
print(rows_aluno, rows_curso, rows_docente)
#para importar um sample de um df muito grande
colunas_aluno = ['CO_IES','CO_ALUNO','TP_CATEGORIA_ADMINISTRATIVA','TP_SEXO','CO_OCDE_AREA_GERAL','CO_OCDE_AREA_ESPECIFICA','CO_OCDE_AREA_DETALHADA','CO_OCDE','CO_CURSO','NU_ANO_INGRESSO','TP_SITUACAO','NU_ANO_NASCIMENTO','NU_MES_NASCIMENTO','NU_DIA_NASCIMENTO','IN_MATRICULA', 'IN_CONCLUINTE', 'IN_APOIO_SOCIAL', 'IN_BOLSA_ESTAGIO', 'NU_ANO_INGRESSO']
n = sum(1 for line in open('DM_ALUNO.csv')) - 1
s = 1200000
skip = sorted(random.sample(range(1, n+1), n-s))
#importação com o skip jogando fora o numero de linhas selecionadas aleatoriamente
df_aluno = pd.read_csv('DM_ALUNO.csv', sep="|", skiprows = skip, encoding = "latin1", usecols=colunas_aluno)
df_aluno
df_aluno.info()
#importando apenas as colunas necessárias do arquivo com os dados dos cursos
colunas_curso = ['CO_IES','DT_INICIO_FUNCIONAMENTO','CO_UF', 'CO_CURSO','CO_OCDE_AREA_GERAL','CO_OCDE_AREA_ESPECIFICA','CO_OCDE_AREA_DETALHADA','CO_OCDE']
df_curso = pd.read_csv('DM_CURSO.CSV', usecols = colunas_curso, sep = "|", encoding='latin1')
df_curso
#importando os dados referentes as UF
df_estado = pd.read_csv('estados.csv', sep=';')
df_estado = df_estado.drop(columns = ['Unnamed: 0'])
df_estado
#importando o df com as informações dos cursos de acordo com a OCDE
df_ocde = pd.read_csv('TB_AUX_AREA_OCDE.CSV', sep="|", engine='python')
df_ocde
# Merge nos df's referentes aos alunos e cursos para uma base geral com dados para todos os alunos
df_curso_completo = pd.merge(df_curso, df_ocde, how='left')
df_curso_completo = pd.merge(df_curso_completo, df_estado, how='left')
df_aluno_completo = pd.merge(df_aluno, df_curso_completo, how='left')
del df_aluno
df_aluno_completo
colunas_docente = ['TP_REGIME_TRABALHO','IN_BOLSA_PESQUISA','IN_ATUACAO_PESQUISA','TP_ESCOLARIDADE','CO_IES','TP_CATEGORIA_ADMINISTRATIVA','CO_DOCENTE']
df_docente = pd.read_csv('DM_DOCENTE.CSV', usecols=colunas_docente, sep="|")
df_docente
```
# 1) Tabelas com número de universidades públicas e privadas por estado e número de alunos por universidade
```
lista_ins = {1: 'Pública', 2: 'Pública', 3: 'Pública', 4: 'Privada', 5: 'Privada', 6: 'Privada', 7: 'Privada'}
df_aluno_completo['Pub/Priv'] = df_aluno_completo['TP_CATEGORIA_ADMINISTRATIVA'].map(lista_ins)
df_aluno_completo
group1 = df_aluno_completo.groupby(['NOME_UF', 'Pub/Priv'])
tabela1 = group1.agg({'CO_IES': 'nunique', 'CO_ALUNO': 'nunique'})
tabela1.unstack()
```
# 2) Pergunta-se: é verdade que existe menos mulheres nos cursos de exatas? Explique com os dados.
```
df_ocde
# Verificando qual categoria da OCDE se refere a cursos de exatas
group2 = df_ocde.groupby('CO_OCDE_AREA_GERAL')
group2.agg({'NO_OCDE_AREA_GERAL': list})
df_aluno_completo
# filtrando o curso de exatas
prim = df_aluno_completo[df_aluno_completo['CO_OCDE_AREA_GERAL'] > 3]
curso_exatas = prim[prim['CO_OCDE_AREA_GERAL'] < 6]
curso_exatas
# Verificando se o data frame curso_exatas tem apenas as duas categorias de cursos desejadas
curso_exatas.nunique()
# se tratando de um evento binomial, para elucidar melhor a resolução, iremos substituir 2 para 0 na ocorrência de ser homem
curso_exatas['TP_SEXO_B'] = curso_exatas['TP_SEXO'].replace(2, 0)
curso_exatas.drop(['TP_SEXO'], axis =1)
# contagem de mulheres em cursos de exatas
curso_exatas.TP_SEXO_B.value_counts()
# No caso de uma distribuição binominal, a probabilidade de um evento é a própria média
prop_mulheres = 77177/(168860+77177)
prop_mulheres
plt.hist(curso_exatas['TP_SEXO_B'], label = '0 - Homem \n1 - Mulher')
plt.title('Mulheres em curso de exatas')
plt.legend();
# fazendo o teste de hipótese para uma distribuição binomial, sendo que a hipótese nula é de que homens e mulheres tem a mesma
# proporção em cursos de exatas e a alternativa é de que existem menos mulheres em cursos de exatas, ou seja, um teste unicaudal
scipy.stats.binom_test(77177, n=(168860+77177), p=0.5, alternative='less')
```
A hipótese nula de que não existe diferença na presença de homens e mulheres em cursos na área de exatas é rejeitada com um nível de confiança de 95%, ja que o p value do teste é menor que 5% ou 0,05 e a hipótese alternativa, de que existem menos mulheres em cursos de exatas é aceita.
# 3) Quantos cursos novos abrem por ano?
```
df_curso_completo
df_curso_completo['DT_INICIO_FUNCIONAMENTO'] = df_curso_completo['DT_INICIO_FUNCIONAMENTO'].astype('str')
df_curso_completo['ANO_FUNC'] = df_curso_completo['DT_INICIO_FUNCIONAMENTO'].apply(lambda x: x[-4:])
df_curso_completo
df_questao3 = df_curso_completo.copy()
questao3 = df_questao3.groupby(['ANO_FUNC'], as_index=False).agg({'CO_CURSO': 'count'})
questao3 = questao3[questao3.ANO_FUNC != 'nan']
questao3['ANO_FUNC'] = questao3['ANO_FUNC'].astype('int')
questao3 = questao3[questao3['ANO_FUNC']>1999]
questao3 = questao3.drop(questao3.index[19])
questao3
```
# 4) Se usarmos a taxa de concluientes de um curso como variável de dificuldade dos cursos, eles tem ficado mais faceis ou mais duros ao longo do tempo? Quais as dificuldades para uma afirmação dessas?
```
df_aluno_completo
df_questao4 = df_aluno_completo[['CO_CURSO', 'CO_ALUNO', 'NU_ANO_INGRESSO', 'IN_CONCLUINTE']]
df_questao4
teste = df_questao4.groupby(['CO_CURSO', 'NU_ANO_INGRESSO'], as_index=False).agg({'IN_CONCLUINTE': 'mean', 'CO_ALUNO': 'count'}) #rodar depois
teste
teste2 = teste[teste['CO_ALUNO']>10]
teste2 = teste2[teste2['NU_ANO_INGRESSO']>2000]
teste2 = teste2.round(2)
teste2
teste3 = teste2.groupby(['CO_CURSO', 'NU_ANO_INGRESSO']).agg({'IN_CONCLUINTE': list})
teste3.unstack()
cond1 = teste2['CO_CURSO'] == 5001384
df_graph1 = teste2.loc[(cond1)]
fig, ax = plt.subplots(1,1)
x = df_graph1['NU_ANO_INGRESSO']
y = df_graph1['IN_CONCLUINTE']
plt.title('Taxa de concluintes de por ano - curso 5001384')
plt.bar(x,y, color = 'pink');
```
# 5) Rode uma regressão multipla que explique o abandono dos cursos, será que professores mais/menos preparados influência nessas taxas?
```
# calculando uma média de instrução para os docentes de cada IES
docente_ies = df_docente.groupby(['CO_IES'])
media_docente = docente_ies.agg({'TP_ESCOLARIDADE': 'mean'})
media_docente.reset_index(inplace = True)
media_docente
df_aluno_5 = pd.read_csv('DM_ALUNO.csv', sep="|", skiprows = skip, encoding = "latin1")
df_aluno_5
df_aluno_5 = pd.merge(df_aluno_5, media_docente, how='left')
df_aluno_5.corr().round(2)['TP_SITUACAO'].sort_values(ascending=False)
teste_questao5 = df_aluno_5[['CO_IES', 'TP_SITUACAO', 'IN_CONCLUINTE', 'QT_CARGA_HORARIA_INTEG', 'TP_MOBILIDADE_ACADEMICA_INTERN', 'IN_APOIO_SOCIAL', 'IN_INGRESSO_TOTAL', 'TP_ESCOLARIDADE']]
teste_questao5
lista_st = {2: 'normal', 3: 'abandono', 4: 'abandono', 5: 'abandono', 6: 'normal', 7: 'normal'}
lista_st2 = {'normal': 0, 'abandono': 1}
lista_st3 = {1: 'sim', 2: 'sim'}
lista_st4 = {'sim': 1}
teste_questao5['situacao'] = teste_questao5['TP_SITUACAO'].map(lista_st)
teste_questao5['TX_ABANDONO'] = teste_questao5['situacao'].map(lista_st2)
teste_questao5['intercambio'] = teste_questao5['TP_MOBILIDADE_ACADEMICA_INTERN'].map(lista_st3)
teste_questao5['TX_INTERCAMBIO'] = teste_questao5['intercambio'].map(lista_st4)
teste_questao5
questao5 = teste_questao5.groupby(['CO_IES'], as_index=False).agg({'IN_CONCLUINTE': 'mean', 'QT_CARGA_HORARIA_INTEG': 'mean', 'TP_ESCOLARIDADE': 'mean', 'TX_INTERCAMBIO': 'mean', 'IN_APOIO_SOCIAL': 'mean', 'IN_INGRESSO_TOTAL': 'mean', 'TX_ABANDONO': 'mean' })
questao5
questao5 = questao5.fillna(0)
questao5['TX_INTERCAMBIO_DUMMY'] = questao5['TX_INTERCAMBIO'].apply(lambda x: 1 if x>0.5 else 0)
questao5 = pd.get_dummies(questao5, columns =['TX_INTERCAMBIO_DUMMY'])
questao5
questao5['TX_ABANDONO_LOG'] = np.log1p(questao5['TX_ABANDONO'])
questao5['IN_APOIO_SOCIAL_LOG'] = np.log1p(questao5['IN_APOIO_SOCIAL'])
questao5['TP_ESCOLARIDADE_LOG'] = np.log1p(questao5['TP_ESCOLARIDADE'])
questao5['IN_INGRESSO_TOTAL_LOG'] = np.log1p(questao5['IN_INGRESSO_TOTAL'])
questao5['QT_CARGA_HORARIA_INTEG_LOG'] = np.log1p(questao5['QT_CARGA_HORARIA_INTEG'])
questao5['IN_CONCLUINTE_LOG'] = np.log1p(questao5['IN_CONCLUINTE'])
questao5
questao5.corr()
regressao5 = questao5[['TX_ABANDONO_LOG', 'IN_APOIO_SOCIAL_LOG', 'TP_ESCOLARIDADE_LOG', 'IN_INGRESSO_TOTAL_LOG', 'QT_CARGA_HORARIA_INTEG_LOG', 'IN_CONCLUINTE_LOG', 'TX_INTERCAMBIO_DUMMY_0', 'TX_INTERCAMBIO_DUMMY_1']]
regressao5
sns.pairplot(data=regressao5)
#rodando a regressão
function = 'TX_ABANDONO_LOG ~ IN_APOIO_SOCIAL_LOG + IN_INGRESSO_TOTAL_LOG + QT_CARGA_HORARIA_INTEG_LOG + IN_CONCLUINTE_LOG + TP_ESCOLARIDADE_LOG + TX_INTERCAMBIO_DUMMY_0 + TX_INTERCAMBIO_DUMMY_1'
model = smf.ols(formula=function, data=regressao5).fit()
model.summary()
```
# 6) Quais os cursos com maior crescimento de matriculas por região? E quais os com maior queda? Como você explicaria isso.
```
df_aluno_completo
df_questao6 =df_aluno_completo.copy()
questao6 = df_questao6[df_questao6['NU_ANO_INGRESSO']>2000]
questao6 = questao6.groupby(['Região', 'NO_OCDE'])['IN_MATRICULA'].agg(['sum', 'count']).reset_index([0,1])
questao6['tx_matricula'] = questao6['sum'] / questao6['count']
questao6.sort_values(by=['Região','sum','tx_matricula'],ascending=False, inplace=True)
questao6 = questao6[questao6['count']>100]
questao6
maiores = []
menores = []
for reg in questao6['Região'].unique():
t = questao6.loc[questao6['Região']==reg]
M = t.nlargest (5, columns=['tx_matricula'])
m = t.nsmallest(5, columns=['tx_matricula'])
maiores.append(M)
menores.append(m)
lista1 = pd.concat(maiores).sort_values(by=['Região','tx_matricula'], ascending = False)
lista2 = pd.concat(menores).sort_values(by=['Região','tx_matricula'], ascending = False)
pd.set_option("display.max_rows", 25)
lista1
pd.set_option("display.max_rows", 25)
lista2
```
# 7) Construa uma variável "Signo" dos estudantes e explique porque ela é correlacionada com a variável "probabilidade de formação" (construir)
```
idx = pd.date_range('2018-01-01', periods=365, freq='D')
ts = pd.Series(range(len(idx)), index=idx)
teste7 = pd.DataFrame(ts)
teste7 = datas.reset_index()
teste7 = datas.rename(columns={'index':'anomesdia'})
teste7 = datas.rename(columns={'0':'fora'})
teste7['anomesdia'] = teste7['anomesdia'].astype('str')
signos = []
cap = 'Capricornio'
aq = 'Aquario'
pe = 'Peixes'
ar = 'Aries'
to = 'Touro'
ge = 'Gemeos'
ca = 'Cancer'
le = 'Leao'
vi = 'Virgem'
li = 'Libra'
es = 'Escorpiao'
sa = 'Sagitario'
for x in teste7['anomesdia']:
if x > '01-01' and x < '01-21':
signos.append(cap)
elif x > '01-21'and x < '02-20':
signos.append(aq)
elif x > '02-19' and x < '03-21':
signos.append(pe)
elif x > '03-20' and x < '04-21':
signos.append(ar)
elif x > '04-20' and x < '05-21':
signos.append(to)
elif x > '05-20' and x < '06-21':
signos.append(ge)
elif x > '06-20' and x < '07-22':
signos.append(ca)
elif x > '07-21' and x < '08-23':
signos.append(le)
elif x > '08-22' and x < '09-23':
signos.append(vi)
elif x > '09-22' and x < '10-23':
signos.append(li)
elif x > '10-22' and x < '11-22':
signos.append(es)
elif x > '11-21' and x < '12-22':
signos.append(sa)
else:
signos.append(cap)
teste7['signo'] = signos
teste7['anomesdia'] = teste7['anomesdia'].apply(lambda x: x[-5:])
teste7 = teste7[['anomesdia', 'signo']]
teste7 = pd.DataFrame(teste7)
teste7.nunique()
df_aluno7 = df_aluno_completo.copy()
df_aluno7['NU_ANO_NASCIMENTO'] = df_aluno7['NU_ANO_NASCIMENTO'].astype('str')
df_aluno7['NU_MES_NASCIMENTO'] = df_aluno7['NU_MES_NASCIMENTO'].astype('str')
df_aluno7['NU_DIA_NASCIMENTO'] = df_aluno7['NU_DIA_NASCIMENTO'].astype('str')
df_aluno7['data_nasc'] = df_aluno7['NU_ANO_NASCIMENTO']+'-'+df_aluno7['NU_MES_NASCIMENTO']+'-'+df_aluno7['NU_DIA_NASCIMENTO']
df_aluno7['data_nasc'] = pd.to_datetime(df_aluno7['data_nasc'])
df_aluno7['data_nasc'] = df_aluno7['data_nasc'].astype('str')
df_aluno7['data_signo'] = df_aluno7['data_nasc'].apply(lambda x: x[-5:])
df_aluno7 = pd.merge(df_aluno7, teste7, how='left', left_on=['data_signo'] , right_on=['anomesdia'])
df_aluno7
df_aluno7.drop(['diames','anomesdia'])
df_aluno7
```
| github_jupyter |
# Importing libraries
```
import sys, os, re, csv, subprocess, operator
import pandas as pd
from urllib.request import urlopen
import urllib.request
from bs4 import BeautifulSoup
```
# Configure repository and directories
```
userhome = os.path.expanduser('~')
txt_file = open(userhome + r"/DifferentDiffAlgorithms/SZZ/code_document/project_identity.txt", "r")
pid = txt_file.read().split('\n')
project = pid[0]
bugidentifier = pid[1]
repository = userhome + r'/DifferentDiffAlgorithms/SZZ/datasource/' + project + '/'
analyze_dir = userhome + r'/DifferentDiffAlgorithms/SZZ/projects_analyses/' + project + '/'
print ("Project name = %s" % project)
print ("Project key = %s" % bugidentifier)
```
# Load textfile contains bug-ids
```
txtfile = open(analyze_dir + "01_bug_ids_extraction/candidate_bug_ids.txt", "r")
bug_links = txtfile.read().split('\n')
print ("Found " + str(len(bug_links)) + " bug_ids")
```
# Finding affected versions by bug ids
```
error_links = []
affected_version = []
for a,b in enumerate(bug_links):
link = "https://issues.apache.org/jira/browse/" + b
sys.stdout.write("\r%i " %(a+1) + "Extracting: " + b)
sys.stdout.flush()
try:
page = urllib.request.urlopen(link)
soup = BeautifulSoup(page, 'html.parser')
aff_version = soup.find('span', attrs={'id':'versions-val'}).text.replace("\n",'').replace(" M",'-M').replace(" ",'').replace(".x",'.').split(",")
aff_version = sorted(aff_version)
aff_version.insert(0,b)
affected_version.append(aff_version)
except:
error_links.append(b)
print("\nExtraction has been completed.")
print (error_links)
#Repeat the process if there are still some affected versions by bug_ids haven't been captured due to network problems
errorlinks = []
if error_links != []:
for c,d in enumerate(error_links):
link = "https://issues.apache.org/jira/browse/" + d
sys.stdout.write("\r%i " %(c+1) + "Extracting: " + d)
sys.stdout.flush()
try:
page = urllib.request.urlopen(link)
soup = BeautifulSoup(page, 'html.parser')
types = soup.find('span', attrs={'id':'versions-val'}).text.replace("\n",'').replace(" M",'-M').replace(" ",'').replace(".x",'.').split(",")
types = sorted(types)
types.insert(0, d)
affected_version.append(types)
except:
errorlinks.append(d)
print ("\nExtraction is complete")
print (errorlinks)
affected_version.sort()
#Finding the earliest version affected by the bug ids
earliest_version = []
for num, affver in enumerate(affected_version):
earliest_version.append(affver[:2])
earliest_version.sort()
for early in earliest_version:
print (early)
```
# Defining the function for git command
```
def execute_command(cmd, work_dir):
#Executes a shell command in a subprocess, waiting until it has completed.
pipe = subprocess.Popen(cmd, shell=True, cwd=work_dir, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
(out, error) = pipe.communicate()
return out, error
pipe.wait()
```
# Finding the versions related with earliest version
```
related_version = []
for n, item in enumerate(earliest_version):
if "." in item[1]:
git_cmd = 'git tag -l "*' + item[1] + '*"'
temp = str(execute_command(git_cmd, repository)).replace("b'",'').replace("(",'').replace(")",'').split("\\n")
del temp[len(temp)-1]
if temp == []:
temp = [item[1].replace("Java-SCA-","")]
else:
temp = ['None']
temp.insert(0, item[0])
related_version.append(temp)
for xx in related_version:
print (xx)
```
# Finding the date release for affected version
```
date_release = []
for n, item in enumerate(related_version):
sys.stdout.write("\rFinding datetime for version {}: {}".format(n+1, item[0]))
sys.stdout.flush()
if item[1] != "None":
for m in range(1, len(item)):
git_cmd = "git log -1 --format=%ai " + item[m]
temp = str(execute_command(git_cmd, repository)).replace("b'",'').replace("(",'').replace(")",'').split("\\n")
del temp[len(temp)-1]
temp = temp[0].split(" ")
if temp[0] != "',":
temp.insert(0,item[0])
temp.insert(1,item[m])
date_release.append(temp)
date_release = sorted(date_release, key=operator.itemgetter(0, 2))
"""else:
date_release.append(item)"""
date_release = sorted(date_release, key=operator.itemgetter(0), reverse=True)
print ("\nThe process is finish")
#save in CSV file
with open(analyze_dir + '04_affected_versions/affected_version.csv','w') as csvfile:
writers = csv.writer(csvfile)
writers.writerow(['bug_id','earliest_affected_version','date_release','time_release','tz'])
for item in date_release:
writers.writerow(item)
df = pd.read_csv(analyze_dir + '04_affected_versions/affected_version.csv')
df
earliest_vers = df.groupby('bug_id', as_index=False).first()
earliest_vers = earliest_vers.sort_values(['date_release', 'time_release', 'earliest_affected_version'], ascending=True)
earliest_vers.to_csv(analyze_dir + '04_affected_versions/earliest_version.csv', index=False)
earliest_vers
```
# Joining 2 csv files: list of annotated files and earliest affected versions
```
colname = ['bug_id','bugfix_commitID','parent_id','filepath','diff_myers_file','diff_histogram_file','blame_myers_file','blame_histogram_file',
'#deletions_myers','#deletions_histogram']
filedata = pd.read_csv(analyze_dir + '03_annotate/01_annotated_files/listof_diff_n_annotated_files/diff_n_blame_combination_files.csv')
filedata = filedata[colname]
details = filedata.join(earliest_vers.set_index('bug_id')[['earliest_affected_version','date_release']], on='bug_id')
details.to_csv(analyze_dir + '04_affected_versions/affected_version_for_identified_files.csv', index=False)
print ("Affected version for identified files has been created")
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.